JP5836271B2 - グリセロールを含まない発酵によるエタノール生成 - Google Patents
グリセロールを含まない発酵によるエタノール生成 Download PDFInfo
- Publication number
- JP5836271B2 JP5836271B2 JP2012521593A JP2012521593A JP5836271B2 JP 5836271 B2 JP5836271 B2 JP 5836271B2 JP 2012521593 A JP2012521593 A JP 2012521593A JP 2012521593 A JP2012521593 A JP 2012521593A JP 5836271 B2 JP5836271 B2 JP 5836271B2
- Authority
- JP
- Japan
- Prior art keywords
- cell
- seq
- dependent
- glycerol
- gene
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
- LFQSCWFLJHTTHZ-UHFFFAOYSA-N Ethanol Chemical compound CCO LFQSCWFLJHTTHZ-UHFFFAOYSA-N 0.000 title claims description 219
- PEDCQBHIVMGVHV-UHFFFAOYSA-N Glycerine Chemical compound OCC(O)CO PEDCQBHIVMGVHV-UHFFFAOYSA-N 0.000 title claims description 189
- 238000000855 fermentation Methods 0.000 title claims description 34
- 230000004151 fermentation Effects 0.000 title claims description 32
- 238000004519 manufacturing process Methods 0.000 title description 21
- 210000004027 cell Anatomy 0.000 claims description 118
- 108090000623 proteins and genes Proteins 0.000 claims description 106
- 150000007523 nucleic acids Chemical group 0.000 claims description 75
- 230000000694 effects Effects 0.000 claims description 67
- 230000001419 dependent effect Effects 0.000 claims description 63
- 210000005253 yeast cell Anatomy 0.000 claims description 60
- 238000000034 method Methods 0.000 claims description 51
- 102000004190 Enzymes Human genes 0.000 claims description 47
- 108090000790 Enzymes Proteins 0.000 claims description 47
- 229930027945 nicotinamide-adenine dinucleotide Natural products 0.000 claims description 39
- BOPGDPNILDQYTO-NNYOXOHSSA-N nicotinamide-adenine dinucleotide Chemical compound C1=CCC(C(=O)N)=CN1[C@H]1[C@H](O)[C@H](O)[C@@H](COP(O)(=O)OP(O)(=O)OC[C@@H]2[C@H]([C@@H](O)[C@@H](O2)N2C3=NC=NC(N)=C3N=C2)O)O1 BOPGDPNILDQYTO-NNYOXOHSSA-N 0.000 claims description 39
- 240000004808 Saccharomyces cerevisiae Species 0.000 claims description 38
- 102000004169 proteins and genes Human genes 0.000 claims description 38
- 235000014680 Saccharomyces cerevisiae Nutrition 0.000 claims description 36
- QTBSBXVTEAMEQO-UHFFFAOYSA-M Acetate Chemical compound CC([O-])=O QTBSBXVTEAMEQO-UHFFFAOYSA-M 0.000 claims description 32
- 230000015572 biosynthetic process Effects 0.000 claims description 31
- 102000039446 nucleic acids Human genes 0.000 claims description 29
- 108020004707 nucleic acids Proteins 0.000 claims description 29
- 108091028043 Nucleic acid sequence Proteins 0.000 claims description 28
- 108010081577 aldehyde dehydrogenase (NAD(P)+) Proteins 0.000 claims description 27
- 150000001720 carbohydrates Chemical class 0.000 claims description 26
- WQZGKKKJIJFFOK-GASJEMHNSA-N Glucose Natural products OC[C@H]1OC(O)[C@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-GASJEMHNSA-N 0.000 claims description 25
- 239000008103 glucose Substances 0.000 claims description 25
- 238000003786 synthesis reaction Methods 0.000 claims description 24
- 235000014633 carbohydrates Nutrition 0.000 claims description 22
- 102100036669 Glycerol-3-phosphate dehydrogenase [NAD(+)], cytoplasmic Human genes 0.000 claims description 20
- 101001009678 Homo sapiens Glycerol-3-phosphate dehydrogenase, mitochondrial Proteins 0.000 claims description 16
- 102100030395 Glycerol-3-phosphate dehydrogenase, mitochondrial Human genes 0.000 claims description 15
- 101001072574 Homo sapiens Glycerol-3-phosphate dehydrogenase [NAD(+)], cytoplasmic Proteins 0.000 claims description 15
- 238000002360 preparation method Methods 0.000 claims description 14
- 230000002829 reductive effect Effects 0.000 claims description 13
- 108010021809 Alcohol dehydrogenase Proteins 0.000 claims description 12
- 102000007698 Alcohol dehydrogenase Human genes 0.000 claims description 12
- 108010049926 Acetate-CoA ligase Proteins 0.000 claims description 11
- 102000008146 Acetate-CoA ligase Human genes 0.000 claims description 11
- 230000035772 mutation Effects 0.000 claims description 10
- 235000000346 sugar Nutrition 0.000 claims description 10
- 239000002029 lignocellulosic biomass Substances 0.000 claims description 9
- 108010041921 Glycerolphosphate Dehydrogenase Proteins 0.000 claims description 8
- 241000235070 Saccharomyces Species 0.000 claims description 8
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 claims description 7
- SRBFZHDQGSBBOR-IOVATXLUSA-N D-xylopyranose Chemical compound O[C@@H]1COC(O)[C@H](O)[C@H]1O SRBFZHDQGSBBOR-IOVATXLUSA-N 0.000 claims description 6
- PYMYPHUHKUWMLA-UHFFFAOYSA-N arabinose Natural products OCC(O)C(O)C(O)C=O PYMYPHUHKUWMLA-UHFFFAOYSA-N 0.000 claims description 6
- SRBFZHDQGSBBOR-UHFFFAOYSA-N beta-D-Pyranose-Lyxose Natural products OC1COC(O)C(O)C1O SRBFZHDQGSBBOR-UHFFFAOYSA-N 0.000 claims description 6
- 230000002950 deficient Effects 0.000 claims description 6
- 229920002488 Hemicellulose Polymers 0.000 claims description 5
- 239000001814 pectin Substances 0.000 claims description 5
- 229920001277 pectin Polymers 0.000 claims description 5
- 235000010987 pectin Nutrition 0.000 claims description 5
- 230000021736 acetylation Effects 0.000 claims description 4
- 238000006640 acetylation reaction Methods 0.000 claims description 4
- WQZGKKKJIJFFOK-VFUOTHLCSA-N beta-D-glucose Chemical compound OC[C@H]1O[C@@H](O)[C@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-VFUOTHLCSA-N 0.000 claims description 4
- 239000001913 cellulose Substances 0.000 claims description 4
- 229920002678 cellulose Polymers 0.000 claims description 4
- 238000012217 deletion Methods 0.000 claims description 4
- 230000037430 deletion Effects 0.000 claims description 4
- 101710088194 Dehydrogenase Proteins 0.000 claims description 3
- 101150004714 GPP1 gene Proteins 0.000 claims description 3
- 101150059691 GPP2 gene Proteins 0.000 claims description 3
- XYZZKVRWGOWVGO-UHFFFAOYSA-N Glycerol-phosphate Chemical compound OP(O)(O)=O.OCC(O)CO XYZZKVRWGOWVGO-UHFFFAOYSA-N 0.000 claims description 3
- 108090000608 Phosphoric Monoester Hydrolases Proteins 0.000 claims description 3
- 102000004160 Phosphoric Monoester Hydrolases Human genes 0.000 claims description 3
- PYMYPHUHKUWMLA-WDCZJNDASA-N arabinose Chemical compound OC[C@@H](O)[C@@H](O)[C@H](O)C=O PYMYPHUHKUWMLA-WDCZJNDASA-N 0.000 claims description 3
- OWEGMIWEEQEYGQ-UHFFFAOYSA-N 100676-05-9 Natural products OC1C(O)C(O)C(CO)OC1OCC1C(O)C(O)C(O)C(OC2C(OC(O)C(O)C2O)CO)O1 OWEGMIWEEQEYGQ-UHFFFAOYSA-N 0.000 claims description 2
- WQZGKKKJIJFFOK-QTVWNMPRSA-N D-mannopyranose Chemical compound OC[C@H]1OC(O)[C@@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-QTVWNMPRSA-N 0.000 claims description 2
- 229930091371 Fructose Natural products 0.000 claims description 2
- 239000005715 Fructose Substances 0.000 claims description 2
- RFSUNEUAIZKAJO-ARQDHWQXSA-N Fructose Chemical compound OC[C@H]1O[C@](O)(CO)[C@@H](O)[C@@H]1O RFSUNEUAIZKAJO-ARQDHWQXSA-N 0.000 claims description 2
- GUBGYTABKSRVRQ-PICCSMPSSA-N Maltose Natural products O[C@@H]1[C@@H](O)[C@H](O)[C@@H](CO)O[C@@H]1O[C@@H]1[C@@H](CO)OC(O)[C@H](O)[C@H]1O GUBGYTABKSRVRQ-PICCSMPSSA-N 0.000 claims description 2
- 229930006000 Sucrose Natural products 0.000 claims description 2
- CZMRCDWAGMRECN-UGDNZRGBSA-N Sucrose Chemical compound O[C@H]1[C@H](O)[C@@H](CO)O[C@@]1(CO)O[C@@H]1[C@H](O)[C@@H](O)[C@H](O)[C@@H](CO)O1 CZMRCDWAGMRECN-UGDNZRGBSA-N 0.000 claims description 2
- WQZGKKKJIJFFOK-PHYPRBDBSA-N alpha-D-galactose Chemical compound OC[C@H]1O[C@H](O)[C@H](O)[C@@H](O)[C@H]1O WQZGKKKJIJFFOK-PHYPRBDBSA-N 0.000 claims description 2
- GUBGYTABKSRVRQ-QUYVBRFLSA-N beta-maltose Chemical compound OC[C@H]1O[C@H](O[C@H]2[C@H](O)[C@@H](O)[C@H](O)O[C@@H]2CO)[C@H](O)[C@@H](O)[C@@H]1O GUBGYTABKSRVRQ-QUYVBRFLSA-N 0.000 claims description 2
- 229930182830 galactose Natural products 0.000 claims description 2
- 150000004676 glycans Chemical class 0.000 claims description 2
- 230000003301 hydrolyzing effect Effects 0.000 claims description 2
- 229920001282 polysaccharide Polymers 0.000 claims description 2
- 239000005017 polysaccharide Substances 0.000 claims description 2
- 239000005720 sucrose Substances 0.000 claims description 2
- QTBSBXVTEAMEQO-UHFFFAOYSA-N Acetic acid Chemical compound CC(O)=O QTBSBXVTEAMEQO-UHFFFAOYSA-N 0.000 description 69
- 229940088598 enzyme Drugs 0.000 description 43
- 235000018102 proteins Nutrition 0.000 description 33
- IKHGUXGNUITLKF-UHFFFAOYSA-N Acetaldehyde Chemical compound CC=O IKHGUXGNUITLKF-UHFFFAOYSA-N 0.000 description 28
- 108090000765 processed proteins & peptides Proteins 0.000 description 27
- ZSLZBFCDCINBPY-ZSJPKINUSA-N acetyl-CoA Chemical compound O[C@@H]1[C@H](OP(O)(O)=O)[C@@H](COP(O)(=O)OP(O)(=O)OCC(C)(C)[C@@H](O)C(=O)NCCC(=O)NCCSC(=O)C)O[C@H]1N1C2=NC=NC(N)=C2N=C1 ZSLZBFCDCINBPY-ZSJPKINUSA-N 0.000 description 26
- 229920001184 polypeptide Polymers 0.000 description 24
- 102000004196 processed proteins & peptides Human genes 0.000 description 24
- 239000013615 primer Substances 0.000 description 23
- 239000013598 vector Substances 0.000 description 23
- 239000000047 product Substances 0.000 description 20
- 238000006243 chemical reaction Methods 0.000 description 18
- 241000588724 Escherichia coli Species 0.000 description 16
- 239000002609 medium Substances 0.000 description 16
- 230000012010 growth Effects 0.000 description 15
- 239000013612 plasmid Substances 0.000 description 14
- 235000001014 amino acid Nutrition 0.000 description 12
- 150000001413 amino acids Chemical class 0.000 description 12
- 108020004705 Codon Proteins 0.000 description 11
- 229940100228 acetyl coenzyme a Drugs 0.000 description 11
- 101150024975 mhpF gene Proteins 0.000 description 11
- 238000003752 polymerase chain reaction Methods 0.000 description 11
- QVGXLLKOCUKJST-UHFFFAOYSA-N atomic oxygen Chemical compound [O] QVGXLLKOCUKJST-UHFFFAOYSA-N 0.000 description 10
- 101150056052 dmpF gene Proteins 0.000 description 10
- 239000001301 oxygen Substances 0.000 description 10
- 229910052760 oxygen Inorganic materials 0.000 description 10
- 108091033319 polynucleotide Proteins 0.000 description 10
- 102000040430 polynucleotide Human genes 0.000 description 10
- 239000002157 polynucleotide Substances 0.000 description 10
- 230000008569 process Effects 0.000 description 10
- 230000009467 reduction Effects 0.000 description 10
- 239000002028 Biomass Substances 0.000 description 9
- 150000001875 compounds Chemical class 0.000 description 9
- 239000002773 nucleotide Substances 0.000 description 9
- 125000003729 nucleotide group Chemical group 0.000 description 9
- 229920000642 polymer Polymers 0.000 description 9
- 241000894007 species Species 0.000 description 9
- 239000002699 waste material Substances 0.000 description 9
- 108020004414 DNA Proteins 0.000 description 8
- 241000589774 Pseudomonas sp. Species 0.000 description 8
- 238000006460 hydrolysis reaction Methods 0.000 description 8
- 238000013518 transcription Methods 0.000 description 8
- 230000035897 transcription Effects 0.000 description 8
- 238000012795 verification Methods 0.000 description 8
- 101100393309 Saccharomyces cerevisiae (strain ATCC 204508 / S288c) GPD2 gene Proteins 0.000 description 7
- 239000000284 extract Substances 0.000 description 7
- 101150087371 gpd1 gene Proteins 0.000 description 7
- 230000007062 hydrolysis Effects 0.000 description 7
- 230000010354 integration Effects 0.000 description 7
- 230000037361 pathway Effects 0.000 description 7
- 238000011160 research Methods 0.000 description 7
- 101100393304 Saccharomyces cerevisiae (strain ATCC 204508 / S288c) GPD1 gene Proteins 0.000 description 6
- 125000003275 alpha amino acid group Chemical group 0.000 description 6
- 230000009604 anaerobic growth Effects 0.000 description 6
- 238000003745 diagnosis Methods 0.000 description 6
- 239000013604 expression vector Substances 0.000 description 6
- 239000012978 lignocellulosic material Substances 0.000 description 6
- 238000002703 mutagenesis Methods 0.000 description 6
- 231100000350 mutagenesis Toxicity 0.000 description 6
- 230000009466 transformation Effects 0.000 description 6
- 101150002721 GPD2 gene Proteins 0.000 description 5
- 102000000587 Glycerolphosphate Dehydrogenase Human genes 0.000 description 5
- 239000006227 byproduct Substances 0.000 description 5
- 230000015556 catabolic process Effects 0.000 description 5
- 238000010276 construction Methods 0.000 description 5
- 238000005516 engineering process Methods 0.000 description 5
- 230000002255 enzymatic effect Effects 0.000 description 5
- 238000001704 evaporation Methods 0.000 description 5
- 230000008020 evaporation Effects 0.000 description 5
- 239000013613 expression plasmid Substances 0.000 description 5
- 238000012239 gene modification Methods 0.000 description 5
- 230000005017 genetic modification Effects 0.000 description 5
- 235000013617 genetically modified food Nutrition 0.000 description 5
- 238000003780 insertion Methods 0.000 description 5
- 230000037431 insertion Effects 0.000 description 5
- 239000003550 marker Substances 0.000 description 5
- 244000005700 microbiome Species 0.000 description 5
- 150000002772 monosaccharides Chemical class 0.000 description 5
- 230000003287 optical effect Effects 0.000 description 5
- 238000010405 reoxidation reaction Methods 0.000 description 5
- 239000000758 substrate Substances 0.000 description 5
- 108091032973 (ribonucleotides)n+m Proteins 0.000 description 4
- IJGRMHOSHXDMSA-UHFFFAOYSA-N Atomic nitrogen Chemical compound N#N IJGRMHOSHXDMSA-UHFFFAOYSA-N 0.000 description 4
- CURLTUGMZLYLDI-UHFFFAOYSA-N Carbon dioxide Chemical compound O=C=O CURLTUGMZLYLDI-UHFFFAOYSA-N 0.000 description 4
- 108091026890 Coding region Proteins 0.000 description 4
- 108020004511 Recombinant DNA Proteins 0.000 description 4
- 230000007613 environmental effect Effects 0.000 description 4
- 230000002068 genetic effect Effects 0.000 description 4
- 230000002779 inactivation Effects 0.000 description 4
- 239000000463 material Substances 0.000 description 4
- 230000004048 modification Effects 0.000 description 4
- 238000012986 modification Methods 0.000 description 4
- 150000002972 pentoses Chemical class 0.000 description 4
- 230000010076 replication Effects 0.000 description 4
- 238000013519 translation Methods 0.000 description 4
- 239000003643 water by type Substances 0.000 description 4
- OKTJSMMVPCPJKN-UHFFFAOYSA-N Carbon Chemical compound [C] OKTJSMMVPCPJKN-UHFFFAOYSA-N 0.000 description 3
- 108700010070 Codon Usage Proteins 0.000 description 3
- 108091034117 Oligonucleotide Proteins 0.000 description 3
- 108700026244 Open Reading Frames Proteins 0.000 description 3
- 240000008042 Zea mays Species 0.000 description 3
- 235000005824 Zea mays ssp. parviglumis Nutrition 0.000 description 3
- 235000002017 Zea mays subsp mays Nutrition 0.000 description 3
- -1 acetate Chemical class 0.000 description 3
- 101150014383 adhE gene Proteins 0.000 description 3
- 238000004458 analytical method Methods 0.000 description 3
- 238000003556 assay Methods 0.000 description 3
- 230000001588 bifunctional effect Effects 0.000 description 3
- 230000008238 biochemical pathway Effects 0.000 description 3
- 229910052799 carbon Inorganic materials 0.000 description 3
- 239000003795 chemical substances by application Substances 0.000 description 3
- 235000005822 corn Nutrition 0.000 description 3
- 238000002474 experimental method Methods 0.000 description 3
- 239000007789 gas Substances 0.000 description 3
- 102000006602 glyceraldehyde-3-phosphate dehydrogenase Human genes 0.000 description 3
- 108020004445 glyceraldehyde-3-phosphate dehydrogenase Proteins 0.000 description 3
- 150000002402 hexoses Chemical class 0.000 description 3
- 238000004128 high performance liquid chromatography Methods 0.000 description 3
- 101150003679 hsaG gene Proteins 0.000 description 3
- 230000001939 inductive effect Effects 0.000 description 3
- 238000005259 measurement Methods 0.000 description 3
- 108020004999 messenger RNA Proteins 0.000 description 3
- 230000004060 metabolic process Effects 0.000 description 3
- 229920001542 oligosaccharide Polymers 0.000 description 3
- 150000002482 oligosaccharides Chemical class 0.000 description 3
- 230000003647 oxidation Effects 0.000 description 3
- 238000007254 oxidation reaction Methods 0.000 description 3
- 239000013600 plasmid vector Substances 0.000 description 3
- 230000002441 reversible effect Effects 0.000 description 3
- 239000000126 substance Substances 0.000 description 3
- 150000008163 sugars Chemical class 0.000 description 3
- FQVLRGLGWNWPSS-BXBUPLCLSA-N (4r,7s,10s,13s,16r)-16-acetamido-13-(1h-imidazol-5-ylmethyl)-10-methyl-6,9,12,15-tetraoxo-7-propan-2-yl-1,2-dithia-5,8,11,14-tetrazacycloheptadecane-4-carboxamide Chemical compound N1C(=O)[C@@H](NC(C)=O)CSSC[C@@H](C(N)=O)NC(=O)[C@H](C(C)C)NC(=O)[C@H](C)NC(=O)[C@@H]1CC1=CN=CN1 FQVLRGLGWNWPSS-BXBUPLCLSA-N 0.000 description 2
- HFKQINMYQUXOCH-UHFFFAOYSA-N 4-hydroxy-2-oxopentanoic acid Chemical compound CC(O)CC(=O)C(O)=O HFKQINMYQUXOCH-UHFFFAOYSA-N 0.000 description 2
- 102100034035 Alcohol dehydrogenase 1A Human genes 0.000 description 2
- 241000219310 Beta vulgaris subsp. vulgaris Species 0.000 description 2
- 241001453380 Burkholderia Species 0.000 description 2
- 108020004638 Circular DNA Proteins 0.000 description 2
- 241000193403 Clostridium Species 0.000 description 2
- 102000053602 DNA Human genes 0.000 description 2
- 241000196324 Embryophyta Species 0.000 description 2
- 241000588722 Escherichia Species 0.000 description 2
- 241001646716 Escherichia coli K-12 Species 0.000 description 2
- 241000233866 Fungi Species 0.000 description 2
- 108700007698 Genetic Terminator Regions Proteins 0.000 description 2
- 101000892220 Geobacillus thermodenitrificans (strain NG80-2) Long-chain-alcohol dehydrogenase 1 Proteins 0.000 description 2
- 101000780443 Homo sapiens Alcohol dehydrogenase 1A Proteins 0.000 description 2
- 108010044467 Isoenzymes Proteins 0.000 description 2
- QNAYBMKLOCPYGJ-REOHCLBHSA-N L-alanine Chemical compound C[C@H](N)C(O)=O QNAYBMKLOCPYGJ-REOHCLBHSA-N 0.000 description 2
- JVTAAEKCZFNVCJ-UHFFFAOYSA-M Lactate Chemical compound CC(O)C([O-])=O JVTAAEKCZFNVCJ-UHFFFAOYSA-M 0.000 description 2
- 240000006024 Lactobacillus plantarum Species 0.000 description 2
- 235000013965 Lactobacillus plantarum Nutrition 0.000 description 2
- 241000186359 Mycobacterium Species 0.000 description 2
- UFWIBTONFRDIAS-UHFFFAOYSA-N Naphthalene Chemical compound C1=CC=CC2=CC=CC=C21 UFWIBTONFRDIAS-UHFFFAOYSA-N 0.000 description 2
- 108091005461 Nucleic proteins Proteins 0.000 description 2
- 241000320412 Ogataea angusta Species 0.000 description 2
- 102000004316 Oxidoreductases Human genes 0.000 description 2
- 108090000854 Oxidoreductases Proteins 0.000 description 2
- 238000012408 PCR amplification Methods 0.000 description 2
- 240000000220 Panda oleosa Species 0.000 description 2
- 235000016496 Panda oleosa Nutrition 0.000 description 2
- ISWSIDIOOBJBQZ-UHFFFAOYSA-N Phenol Chemical compound OC1=CC=CC=C1 ISWSIDIOOBJBQZ-UHFFFAOYSA-N 0.000 description 2
- 241000235648 Pichia Species 0.000 description 2
- 241000589516 Pseudomonas Species 0.000 description 2
- 241001327108 Pseudomonas sp. CF600 Species 0.000 description 2
- LCTONWCANYUPML-UHFFFAOYSA-M Pyruvate Chemical compound CC(=O)C([O-])=O LCTONWCANYUPML-UHFFFAOYSA-M 0.000 description 2
- 229920002472 Starch Polymers 0.000 description 2
- 235000021536 Sugar beet Nutrition 0.000 description 2
- 239000002253 acid Substances 0.000 description 2
- 235000004279 alanine Nutrition 0.000 description 2
- 125000000539 amino acid group Chemical group 0.000 description 2
- 230000001580 bacterial effect Effects 0.000 description 2
- 239000001569 carbon dioxide Substances 0.000 description 2
- 229910002092 carbon dioxide Inorganic materials 0.000 description 2
- YCIMNLLNPGFGHC-UHFFFAOYSA-N catechol Chemical compound OC1=CC=CC=C1O YCIMNLLNPGFGHC-UHFFFAOYSA-N 0.000 description 2
- 238000010367 cloning Methods 0.000 description 2
- 238000004590 computer program Methods 0.000 description 2
- 238000012790 confirmation Methods 0.000 description 2
- 238000009792 diffusion process Methods 0.000 description 2
- GNGACRATGGDKBX-UHFFFAOYSA-N dihydroxyacetone phosphate Chemical compound OCC(=O)COP(O)(O)=O GNGACRATGGDKBX-UHFFFAOYSA-N 0.000 description 2
- 229910001873 dinitrogen Inorganic materials 0.000 description 2
- ZUOUZKKEUPVFJK-UHFFFAOYSA-N diphenyl Chemical compound C1=CC=CC=C1C1=CC=CC=C1 ZUOUZKKEUPVFJK-UHFFFAOYSA-N 0.000 description 2
- 239000012634 fragment Substances 0.000 description 2
- 238000001727 in vivo Methods 0.000 description 2
- 230000001965 increasing effect Effects 0.000 description 2
- 230000002401 inhibitory effect Effects 0.000 description 2
- JVTAAEKCZFNVCJ-UHFFFAOYSA-N lactic acid Chemical compound CC(O)C(O)=O JVTAAEKCZFNVCJ-UHFFFAOYSA-N 0.000 description 2
- 229940072205 lactobacillus plantarum Drugs 0.000 description 2
- 239000002207 metabolite Substances 0.000 description 2
- 238000002156 mixing Methods 0.000 description 2
- 238000010369 molecular cloning Methods 0.000 description 2
- 238000005457 optimization Methods 0.000 description 2
- 230000036284 oxygen consumption Effects 0.000 description 2
- 101150093025 pepA gene Proteins 0.000 description 2
- 229920001296 polysiloxane Polymers 0.000 description 2
- 239000002994 raw material Substances 0.000 description 2
- 238000012552 review Methods 0.000 description 2
- AWUCVROLDVIAJX-GSVOUGTGSA-N sn-glycerol 3-phosphate Chemical compound OC[C@@H](O)COP(O)(O)=O AWUCVROLDVIAJX-GSVOUGTGSA-N 0.000 description 2
- 235000019698 starch Nutrition 0.000 description 2
- KDYFGRWQOYBRFD-UHFFFAOYSA-L succinate(2-) Chemical compound [O-]C(=O)CCC([O-])=O KDYFGRWQOYBRFD-UHFFFAOYSA-L 0.000 description 2
- 230000001131 transforming effect Effects 0.000 description 2
- 230000009261 transgenic effect Effects 0.000 description 2
- OILXMJHPFNGGTO-UHFFFAOYSA-N (22E)-(24xi)-24-methylcholesta-5,22-dien-3beta-ol Natural products C1C=C2CC(O)CCC2(C)C2C1C1CCC(C(C)C=CC(C)C(C)C)C1(C)CC2 OILXMJHPFNGGTO-UHFFFAOYSA-N 0.000 description 1
- RQOCXCFLRBRBCS-UHFFFAOYSA-N (22E)-cholesta-5,7,22-trien-3beta-ol Natural products C1C(O)CCC2(C)C(CCC3(C(C(C)C=CCC(C)C)CCC33)C)C3=CC=C21 RQOCXCFLRBRBCS-UHFFFAOYSA-N 0.000 description 1
- VEMLQICWTSVKQH-BTVCFUMJSA-N (2r,3s,4r,5r)-2,3,4,5,6-pentahydroxyhexanal;propane-1,2,3-triol Chemical compound OCC(O)CO.OC[C@@H](O)[C@@H](O)[C@H](O)[C@@H](O)C=O VEMLQICWTSVKQH-BTVCFUMJSA-N 0.000 description 1
- 102000040650 (ribonucleotides)n+m Human genes 0.000 description 1
- OSJPPGNTCRNQQC-UWTATZPHSA-N 3-phospho-D-glyceric acid Chemical compound OC(=O)[C@H](O)COP(O)(O)=O OSJPPGNTCRNQQC-UWTATZPHSA-N 0.000 description 1
- 108010076069 4-hydroxy-2-ketovalerate aldolase Proteins 0.000 description 1
- 102100031126 6-phosphogluconolactonase Human genes 0.000 description 1
- 108010029731 6-phosphogluconolactonase Proteins 0.000 description 1
- OQMZNAMGEHIHNN-UHFFFAOYSA-N 7-Dehydrostigmasterol Natural products C1C(O)CCC2(C)C(CCC3(C(C(C)C=CC(CC)C(C)C)CCC33)C)C3=CC=C21 OQMZNAMGEHIHNN-UHFFFAOYSA-N 0.000 description 1
- 101150105694 ACS1 gene Proteins 0.000 description 1
- 101150050888 ACS2 gene Proteins 0.000 description 1
- 101150078509 ADH2 gene Proteins 0.000 description 1
- 101150026777 ADH5 gene Proteins 0.000 description 1
- 101710194330 Acetyl-coenzyme A synthetase 2 Proteins 0.000 description 1
- 101100112372 Acinetobacter baylyi (strain ATCC 33305 / BD413 / ADP1) catM gene Proteins 0.000 description 1
- 241000186361 Actinobacteria <class> Species 0.000 description 1
- 102100034042 Alcohol dehydrogenase 1C Human genes 0.000 description 1
- 102100039702 Alcohol dehydrogenase class-3 Human genes 0.000 description 1
- 108010025188 Alcohol oxidase Proteins 0.000 description 1
- 102100034044 All-trans-retinol dehydrogenase [NAD(+)] ADH1B Human genes 0.000 description 1
- 102100031795 All-trans-retinol dehydrogenase [NAD(+)] ADH4 Human genes 0.000 description 1
- 101710193111 All-trans-retinol dehydrogenase [NAD(+)] ADH4 Proteins 0.000 description 1
- 241000609240 Ambelania acida Species 0.000 description 1
- 244000048280 Amelanchier denticulata Species 0.000 description 1
- 235000010992 Amelanchier denticulata Nutrition 0.000 description 1
- 239000004382 Amylase Substances 0.000 description 1
- 102000013142 Amylases Human genes 0.000 description 1
- 108010065511 Amylases Proteins 0.000 description 1
- 101100504210 Arabidopsis thaliana GLDP1 gene Proteins 0.000 description 1
- 101100504216 Arabidopsis thaliana GLDP2 gene Proteins 0.000 description 1
- 101100330294 Arabidopsis thaliana OASC gene Proteins 0.000 description 1
- 241000235349 Ascomycota Species 0.000 description 1
- 101100460671 Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) norA gene Proteins 0.000 description 1
- 101100317631 Aspergillus tubingensis xynA gene Proteins 0.000 description 1
- 241000726110 Azoarcus Species 0.000 description 1
- 101100162670 Bacillus subtilis (strain 168) amyE gene Proteins 0.000 description 1
- 101100277447 Bacillus subtilis (strain 168) degQ gene Proteins 0.000 description 1
- 241000894006 Bacteria Species 0.000 description 1
- 241000221198 Basidiomycota Species 0.000 description 1
- 101710085076 Beta-galactosidase LacA Proteins 0.000 description 1
- 108091003079 Bovine Serum Albumin Proteins 0.000 description 1
- 241000722885 Brettanomyces Species 0.000 description 1
- 101150092493 CREA gene Proteins 0.000 description 1
- 244000025254 Cannabis sativa Species 0.000 description 1
- 102100035882 Catalase Human genes 0.000 description 1
- 108010053835 Catalase Proteins 0.000 description 1
- 108010059892 Cellulase Proteins 0.000 description 1
- 241000207199 Citrus Species 0.000 description 1
- 241000193454 Clostridium beijerinckii Species 0.000 description 1
- 241000186570 Clostridium kluyveri Species 0.000 description 1
- RGJOEKWQDUBAIZ-IBOSZNHHSA-N CoASH Chemical compound O[C@@H]1[C@H](OP(O)(O)=O)[C@@H](COP(O)(=O)OP(O)(=O)OCC(C)(C)[C@@H](O)C(=O)NCCC(=O)NCCS)O[C@H]1N1C2=NC=NC(N)=C2N=C1 RGJOEKWQDUBAIZ-IBOSZNHHSA-N 0.000 description 1
- 229920000742 Cotton Polymers 0.000 description 1
- 101000796894 Coturnix japonica Alcohol dehydrogenase 1 Proteins 0.000 description 1
- 108010051219 Cre recombinase Proteins 0.000 description 1
- 239000003155 DNA primer Substances 0.000 description 1
- 102000016928 DNA-directed DNA polymerase Human genes 0.000 description 1
- 108010014303 DNA-directed DNA polymerase Proteins 0.000 description 1
- 102000004163 DNA-directed RNA polymerases Human genes 0.000 description 1
- 108090000626 DNA-directed RNA polymerases Proteins 0.000 description 1
- 101100317179 Dictyostelium discoideum vps26 gene Proteins 0.000 description 1
- 101100273894 Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) celB gene Proteins 0.000 description 1
- 101100407639 Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) prtB gene Proteins 0.000 description 1
- 101710121765 Endo-1,4-beta-xylanase Proteins 0.000 description 1
- 241000224431 Entamoeba Species 0.000 description 1
- 241000224432 Entamoeba histolytica Species 0.000 description 1
- YQYJSBFKSSDGFO-UHFFFAOYSA-N Epihygromycin Natural products OC1C(O)C(C(=O)C)OC1OC(C(=C1)O)=CC=C1C=C(C)C(=O)NC1C(O)C(O)C2OCOC2C1O YQYJSBFKSSDGFO-UHFFFAOYSA-N 0.000 description 1
- DNVPQKQSNYMLRS-NXVQYWJNSA-N Ergosterol Natural products CC(C)[C@@H](C)C=C[C@H](C)[C@H]1CC[C@H]2C3=CC=C4C[C@@H](O)CC[C@]4(C)[C@@H]3CC[C@]12C DNVPQKQSNYMLRS-NXVQYWJNSA-N 0.000 description 1
- 108091092566 Extrachromosomal DNA Proteins 0.000 description 1
- LLQPHQFNMLZJMP-UHFFFAOYSA-N Fentrazamide Chemical compound N1=NN(C=2C(=CC=CC=2)Cl)C(=O)N1C(=O)N(CC)C1CCCCC1 LLQPHQFNMLZJMP-UHFFFAOYSA-N 0.000 description 1
- 230000005526 G1 to G0 transition Effects 0.000 description 1
- 101150108358 GLAA gene Proteins 0.000 description 1
- 108700039691 Genetic Promoter Regions Proteins 0.000 description 1
- 108010073178 Glucan 1,4-alpha-Glucosidase Proteins 0.000 description 1
- 102100022624 Glucoamylase Human genes 0.000 description 1
- 108010015776 Glucose oxidase Proteins 0.000 description 1
- 239000004366 Glucose oxidase Substances 0.000 description 1
- 108010018962 Glucosephosphate Dehydrogenase Proteins 0.000 description 1
- 235000014477 Gustavia superba Nutrition 0.000 description 1
- 241000238631 Hexapoda Species 0.000 description 1
- 101000780463 Homo sapiens Alcohol dehydrogenase 1C Proteins 0.000 description 1
- 101000959452 Homo sapiens Alcohol dehydrogenase class-3 Proteins 0.000 description 1
- 101000775437 Homo sapiens All-trans-retinol dehydrogenase [NAD(+)] ADH4 Proteins 0.000 description 1
- 101000799318 Homo sapiens Long-chain-fatty-acid-CoA ligase 1 Proteins 0.000 description 1
- 101000780205 Homo sapiens Long-chain-fatty-acid-CoA ligase 5 Proteins 0.000 description 1
- 101000780202 Homo sapiens Long-chain-fatty-acid-CoA ligase 6 Proteins 0.000 description 1
- 101000801742 Homo sapiens Triosephosphate isomerase Proteins 0.000 description 1
- GRRNUXAQVGOGFE-UHFFFAOYSA-N Hygromycin-B Natural products OC1C(NC)CC(N)C(O)C1OC1C2OC3(C(C(O)C(O)C(C(N)CO)O3)O)OC2C(O)C(CO)O1 GRRNUXAQVGOGFE-UHFFFAOYSA-N 0.000 description 1
- 229930010555 Inosine Natural products 0.000 description 1
- UGQMRVRMYYASKQ-KQYNXXCUSA-N Inosine Chemical compound O[C@@H]1[C@H](O)[C@@H](CO)O[C@H]1N1C2=NC=NC(O)=C2N=C1 UGQMRVRMYYASKQ-KQYNXXCUSA-N 0.000 description 1
- 241000588748 Klebsiella Species 0.000 description 1
- 241000588747 Klebsiella pneumoniae Species 0.000 description 1
- 241000235649 Kluyveromyces Species 0.000 description 1
- 241000235058 Komagataella pastoris Species 0.000 description 1
- ROHFNLRQFUQHCH-YFKPBYRVSA-N L-leucine Chemical compound CC(C)C[C@H](N)C(O)=O ROHFNLRQFUQHCH-YFKPBYRVSA-N 0.000 description 1
- ROHFNLRQFUQHCH-UHFFFAOYSA-N Leucine Natural products CC(C)CC(N)C(O)=O ROHFNLRQFUQHCH-UHFFFAOYSA-N 0.000 description 1
- 102100033995 Long-chain-fatty-acid-CoA ligase 1 Human genes 0.000 description 1
- 102100034337 Long-chain-fatty-acid-CoA ligase 6 Human genes 0.000 description 1
- 241000406668 Loxodonta cyclotis Species 0.000 description 1
- 102100024295 Maltase-glucoamylase Human genes 0.000 description 1
- 241000187492 Mycobacterium marinum Species 0.000 description 1
- 241000187917 Mycobacterium ulcerans Species 0.000 description 1
- 239000000020 Nitrocellulose Substances 0.000 description 1
- 241000142651 Pelotomaculum thermopropionicum Species 0.000 description 1
- 108091005804 Peptidases Proteins 0.000 description 1
- 102000010292 Peptide Elongation Factor 1 Human genes 0.000 description 1
- 108010077524 Peptide Elongation Factor 1 Proteins 0.000 description 1
- 102000002508 Peptide Elongation Factors Human genes 0.000 description 1
- 108010068204 Peptide Elongation Factors Proteins 0.000 description 1
- 108091093037 Peptide nucleic acid Proteins 0.000 description 1
- 108091000080 Phosphotransferase Proteins 0.000 description 1
- 241000193632 Piromyces sp. Species 0.000 description 1
- 241000219000 Populus Species 0.000 description 1
- 239000004365 Protease Substances 0.000 description 1
- 108010009736 Protein Hydrolysates Proteins 0.000 description 1
- 241000589776 Pseudomonas putida Species 0.000 description 1
- 229920001131 Pulp (paper) Polymers 0.000 description 1
- 108700005075 Regulator Genes Proteins 0.000 description 1
- 102100037486 Reverse transcriptase/ribonuclease H Human genes 0.000 description 1
- 108091028664 Ribonucleotide Proteins 0.000 description 1
- 101100010928 Saccharolobus solfataricus (strain ATCC 35092 / DSM 1617 / JCM 11322 / P2) tuf gene Proteins 0.000 description 1
- 241000235343 Saccharomycetales Species 0.000 description 1
- 241001326564 Saccharomycotina Species 0.000 description 1
- 241000235060 Scheffersomyces stipitis Species 0.000 description 1
- 238000002105 Southern blotting Methods 0.000 description 1
- 241000191967 Staphylococcus aureus Species 0.000 description 1
- KDYFGRWQOYBRFD-UHFFFAOYSA-N Succinic acid Natural products OC(=O)CCC(O)=O KDYFGRWQOYBRFD-UHFFFAOYSA-N 0.000 description 1
- 108700005078 Synthetic Genes Proteins 0.000 description 1
- 101150001810 TEAD1 gene Proteins 0.000 description 1
- 101150074253 TEF1 gene Proteins 0.000 description 1
- 108700009124 Transcription Initiation Site Proteins 0.000 description 1
- 102100029898 Transcriptional enhancer factor TEF-1 Human genes 0.000 description 1
- GSEJCLTVZPLZKY-UHFFFAOYSA-N Triethanolamine Chemical group OCCN(CCO)CCO GSEJCLTVZPLZKY-UHFFFAOYSA-N 0.000 description 1
- 102100033598 Triosephosphate isomerase Human genes 0.000 description 1
- 108020005202 Viral DNA Proteins 0.000 description 1
- 241000700605 Viruses Species 0.000 description 1
- 241000235017 Zygosaccharomyces Species 0.000 description 1
- 241000235029 Zygosaccharomyces bailii Species 0.000 description 1
- JLCPHMBAVCMARE-UHFFFAOYSA-N [3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-hydroxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methyl [5-(6-aminopurin-9-yl)-2-(hydroxymethyl)oxolan-3-yl] hydrogen phosphate Polymers Cc1cn(C2CC(OP(O)(=O)OCC3OC(CC3OP(O)(=O)OCC3OC(CC3O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c3nc(N)[nH]c4=O)C(COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3CO)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cc(C)c(=O)[nH]c3=O)n3cc(C)c(=O)[nH]c3=O)n3ccc(N)nc3=O)n3cc(C)c(=O)[nH]c3=O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)O2)c(=O)[nH]c1=O JLCPHMBAVCMARE-UHFFFAOYSA-N 0.000 description 1
- 230000002378 acidificating effect Effects 0.000 description 1
- 150000007513 acids Chemical class 0.000 description 1
- 101150063416 add gene Proteins 0.000 description 1
- 239000000654 additive Substances 0.000 description 1
- 230000000996 additive effect Effects 0.000 description 1
- 230000002411 adverse Effects 0.000 description 1
- 239000002154 agricultural waste Substances 0.000 description 1
- 150000001299 aldehydes Chemical class 0.000 description 1
- 108010028144 alpha-Glucosidases Proteins 0.000 description 1
- AVKUERGKIZMTKX-NJBDSQKTSA-N ampicillin Chemical compound C1([C@@H](N)C(=O)N[C@H]2[C@H]3SC([C@@H](N3C2=O)C(O)=O)(C)C)=CC=CC=C1 AVKUERGKIZMTKX-NJBDSQKTSA-N 0.000 description 1
- 229960000723 ampicillin Drugs 0.000 description 1
- 101150069712 amyA gene Proteins 0.000 description 1
- 101150009288 amyB gene Proteins 0.000 description 1
- 235000019418 amylase Nutrition 0.000 description 1
- 238000000137 annealing Methods 0.000 description 1
- 239000002518 antifoaming agent Substances 0.000 description 1
- 101150075954 apeB gene Proteins 0.000 description 1
- 238000013459 approach Methods 0.000 description 1
- 150000001491 aromatic compounds Chemical class 0.000 description 1
- 239000010905 bagasse Substances 0.000 description 1
- 238000006065 biodegradation reaction Methods 0.000 description 1
- 239000004305 biphenyl Substances 0.000 description 1
- 235000010290 biphenyl Nutrition 0.000 description 1
- 229940098773 bovine serum albumin Drugs 0.000 description 1
- 239000000872 buffer Substances 0.000 description 1
- KDYFGRWQOYBRFD-NUQCWPJISA-N butanedioic acid Chemical compound O[14C](=O)CC[14C](O)=O KDYFGRWQOYBRFD-NUQCWPJISA-N 0.000 description 1
- 150000007942 carboxylates Chemical class 0.000 description 1
- 150000001732 carboxylic acid derivatives Chemical class 0.000 description 1
- 101150053553 catR gene Proteins 0.000 description 1
- 101150072516 cbhA gene Proteins 0.000 description 1
- 108091092356 cellular DNA Proteins 0.000 description 1
- 108091092328 cellular RNA Proteins 0.000 description 1
- 229940106157 cellulase Drugs 0.000 description 1
- 238000005119 centrifugation Methods 0.000 description 1
- 230000008859 change Effects 0.000 description 1
- 230000007073 chemical hydrolysis Effects 0.000 description 1
- 239000007795 chemical reaction product Substances 0.000 description 1
- 239000003153 chemical reaction reagent Substances 0.000 description 1
- 230000002759 chromosomal effect Effects 0.000 description 1
- 235000020971 citrus fruits Nutrition 0.000 description 1
- 238000003776 cleavage reaction Methods 0.000 description 1
- RGJOEKWQDUBAIZ-UHFFFAOYSA-N coenzime A Natural products OC1C(OP(O)(O)=O)C(COP(O)(=O)OP(O)(=O)OCC(C)(C)C(O)C(=O)NCCC(=O)NCCS)OC1N1C2=NC=NC(N)=C2N=C1 RGJOEKWQDUBAIZ-UHFFFAOYSA-N 0.000 description 1
- 239000005515 coenzyme Substances 0.000 description 1
- 239000005516 coenzyme A Substances 0.000 description 1
- 229940093530 coenzyme a Drugs 0.000 description 1
- 230000000295 complement effect Effects 0.000 description 1
- 238000007796 conventional method Methods 0.000 description 1
- 239000012228 culture supernatant Substances 0.000 description 1
- 210000004748 cultured cell Anatomy 0.000 description 1
- 210000000172 cytosol Anatomy 0.000 description 1
- 230000006378 damage Effects 0.000 description 1
- 238000006114 decarboxylation reaction Methods 0.000 description 1
- 230000003247 decreasing effect Effects 0.000 description 1
- 239000007857 degradation product Substances 0.000 description 1
- 238000006731 degradation reaction Methods 0.000 description 1
- 239000008367 deionised water Substances 0.000 description 1
- 229910021641 deionized water Inorganic materials 0.000 description 1
- 238000004925 denaturation Methods 0.000 description 1
- 230000036425 denaturation Effects 0.000 description 1
- 239000005547 deoxyribonucleotide Substances 0.000 description 1
- 125000002637 deoxyribonucleotide group Chemical group 0.000 description 1
- KDTSHFARGAKYJN-UHFFFAOYSA-N dephosphocoenzyme A Natural products OC1C(O)C(COP(O)(=O)OP(O)(=O)OCC(C)(C)C(O)C(=O)NCCC(=O)NCCS)OC1N1C2=NC=NC(N)=C2N=C1 KDTSHFARGAKYJN-UHFFFAOYSA-N 0.000 description 1
- 238000001514 detection method Methods 0.000 description 1
- 238000011161 development Methods 0.000 description 1
- 230000018109 developmental process Effects 0.000 description 1
- 150000002016 disaccharides Chemical group 0.000 description 1
- 238000004821 distillation Methods 0.000 description 1
- 238000009837 dry grinding Methods 0.000 description 1
- 101150095359 eglA gene Proteins 0.000 description 1
- 101150038498 eglB gene Proteins 0.000 description 1
- 238000004520 electroporation Methods 0.000 description 1
- 239000003623 enhancer Substances 0.000 description 1
- 229940007078 entamoeba histolytica Drugs 0.000 description 1
- 230000007071 enzymatic hydrolysis Effects 0.000 description 1
- 238000006047 enzymatic hydrolysis reaction Methods 0.000 description 1
- 238000001952 enzyme assay Methods 0.000 description 1
- DNVPQKQSNYMLRS-SOWFXMKYSA-N ergosterol Chemical compound C1[C@@H](O)CC[C@]2(C)[C@H](CC[C@]3([C@H]([C@H](C)/C=C/[C@@H](C)C(C)C)CC[C@H]33)C)C3=CC=C21 DNVPQKQSNYMLRS-SOWFXMKYSA-N 0.000 description 1
- 210000003527 eukaryotic cell Anatomy 0.000 description 1
- 238000011156 evaluation Methods 0.000 description 1
- 238000012262 fermentative production Methods 0.000 description 1
- 235000013305 food Nutrition 0.000 description 1
- 230000004927 fusion Effects 0.000 description 1
- 230000006251 gamma-carboxylation Effects 0.000 description 1
- 238000004868 gas analysis Methods 0.000 description 1
- 101150056490 gdp1 gene Proteins 0.000 description 1
- 238000012826 global research Methods 0.000 description 1
- 229940116332 glucose oxidase Drugs 0.000 description 1
- 235000019420 glucose oxidase Nutrition 0.000 description 1
- 125000000291 glutamic acid group Chemical group N[C@@H](CCC(O)=O)C(=O)* 0.000 description 1
- 108010032776 glycerol-1-phosphatase Proteins 0.000 description 1
- 230000034659 glycolysis Effects 0.000 description 1
- 230000002414 glycolytic effect Effects 0.000 description 1
- 230000013595 glycosylation Effects 0.000 description 1
- 238000006206 glycosylation reaction Methods 0.000 description 1
- 101150073906 gpdA gene Proteins 0.000 description 1
- 101150095733 gpsA gene Proteins 0.000 description 1
- 239000003102 growth factor Substances 0.000 description 1
- 239000001963 growth medium Substances 0.000 description 1
- 238000002744 homologous recombination Methods 0.000 description 1
- 230000006801 homologous recombination Effects 0.000 description 1
- 230000033444 hydroxylation Effects 0.000 description 1
- 238000005805 hydroxylation reaction Methods 0.000 description 1
- GRRNUXAQVGOGFE-NZSRVPFOSA-N hygromycin B Chemical compound O[C@@H]1[C@@H](NC)C[C@@H](N)[C@H](O)[C@H]1O[C@H]1[C@H]2O[C@@]3([C@@H]([C@@H](O)[C@@H](O)[C@@H](C(N)CO)O3)O)O[C@H]2[C@@H](O)[C@@H](CO)O1 GRRNUXAQVGOGFE-NZSRVPFOSA-N 0.000 description 1
- 229940097277 hygromycin b Drugs 0.000 description 1
- 238000000126 in silico method Methods 0.000 description 1
- 238000011534 incubation Methods 0.000 description 1
- 238000011090 industrial biotechnology method and process Methods 0.000 description 1
- 239000003112 inhibitor Substances 0.000 description 1
- 238000011081 inoculation Methods 0.000 description 1
- 229910052816 inorganic phosphate Inorganic materials 0.000 description 1
- 229960003786 inosine Drugs 0.000 description 1
- 230000003834 intracellular effect Effects 0.000 description 1
- 229930027917 kanamycin Natural products 0.000 description 1
- 229960000318 kanamycin Drugs 0.000 description 1
- SBUJHOSQTJFQJX-NOAMYHISSA-N kanamycin Chemical compound O[C@@H]1[C@@H](O)[C@H](O)[C@@H](CN)O[C@@H]1O[C@H]1[C@H](O)[C@@H](O[C@@H]2[C@@H]([C@@H](N)[C@H](O)[C@@H](CO)O2)O)[C@H](N)C[C@@H]1N SBUJHOSQTJFQJX-NOAMYHISSA-N 0.000 description 1
- 229930182823 kanamycin A Natural products 0.000 description 1
- 239000004310 lactic acid Substances 0.000 description 1
- 235000014655 lactic acid Nutrition 0.000 description 1
- 230000004576 lipid-binding Effects 0.000 description 1
- 239000007788 liquid Substances 0.000 description 1
- 238000012423 maintenance Methods 0.000 description 1
- 210000004962 mammalian cell Anatomy 0.000 description 1
- 239000011159 matrix material Substances 0.000 description 1
- 230000007246 mechanism Effects 0.000 description 1
- 230000002503 metabolic effect Effects 0.000 description 1
- 230000037353 metabolic pathway Effects 0.000 description 1
- 229910052751 metal Inorganic materials 0.000 description 1
- 239000002184 metal Substances 0.000 description 1
- 239000000203 mixture Substances 0.000 description 1
- 235000013379 molasses Nutrition 0.000 description 1
- 238000001823 molecular biology technique Methods 0.000 description 1
- 238000012544 monitoring process Methods 0.000 description 1
- OYKBQNOPCSXWBL-SNAWJCMRSA-N n-hydroxy-3-[(e)-3-(hydroxyamino)-3-oxoprop-1-enyl]benzamide Chemical compound ONC(=O)\C=C\C1=CC=CC(C(=O)NO)=C1 OYKBQNOPCSXWBL-SNAWJCMRSA-N 0.000 description 1
- 230000007935 neutral effect Effects 0.000 description 1
- 229920001220 nitrocellulos Polymers 0.000 description 1
- 229910052757 nitrogen Inorganic materials 0.000 description 1
- 238000000424 optical density measurement Methods 0.000 description 1
- LPNBBFKOUUSUDB-UHFFFAOYSA-N p-toluic acid Chemical compound CC1=CC=C(C(O)=O)C=C1 LPNBBFKOUUSUDB-UHFFFAOYSA-N 0.000 description 1
- 230000006506 pH homeostasis Effects 0.000 description 1
- 101150100654 pacC gene Proteins 0.000 description 1
- 239000010893 paper waste Substances 0.000 description 1
- 230000036961 partial effect Effects 0.000 description 1
- 101150035909 pepB gene Proteins 0.000 description 1
- 101150094986 pepC gene Proteins 0.000 description 1
- 239000008363 phosphate buffer Substances 0.000 description 1
- 102000020233 phosphotransferase Human genes 0.000 description 1
- 230000035479 physiological effects, processes and functions Effects 0.000 description 1
- 230000008488 polyadenylation Effects 0.000 description 1
- 235000010482 polyoxyethylene sorbitan monooleate Nutrition 0.000 description 1
- 229920000053 polysorbate 80 Polymers 0.000 description 1
- 239000011148 porous material Substances 0.000 description 1
- 239000008057 potassium phosphate buffer Substances 0.000 description 1
- 101150023641 ppc gene Proteins 0.000 description 1
- 210000001236 prokaryotic cell Anatomy 0.000 description 1
- 101150115781 prtT gene Proteins 0.000 description 1
- 238000002708 random mutagenesis Methods 0.000 description 1
- 239000011541 reaction mixture Substances 0.000 description 1
- 230000006798 recombination Effects 0.000 description 1
- 238000005215 recombination Methods 0.000 description 1
- 238000004064 recycling Methods 0.000 description 1
- 230000001105 regulatory effect Effects 0.000 description 1
- 230000003362 replicative effect Effects 0.000 description 1
- 230000001718 repressive effect Effects 0.000 description 1
- 230000004044 response Effects 0.000 description 1
- 239000002336 ribonucleotide Substances 0.000 description 1
- 125000002652 ribonucleotide group Chemical group 0.000 description 1
- 150000003839 salts Chemical class 0.000 description 1
- 238000005070 sampling Methods 0.000 description 1
- 238000012807 shake-flask culturing Methods 0.000 description 1
- 230000037432 silent mutation Effects 0.000 description 1
- 238000002741 site-directed mutagenesis Methods 0.000 description 1
- 239000008107 starch Substances 0.000 description 1
- 230000001954 sterilising effect Effects 0.000 description 1
- 238000004659 sterilization and disinfection Methods 0.000 description 1
- 239000010907 stover Substances 0.000 description 1
- 239000010902 straw Substances 0.000 description 1
- 230000019635 sulfation Effects 0.000 description 1
- 238000005670 sulfation reaction Methods 0.000 description 1
- 239000006228 supernatant Substances 0.000 description 1
- 230000009469 supplementation Effects 0.000 description 1
- 238000012360 testing method Methods 0.000 description 1
- 238000009283 thermal hydrolysis Methods 0.000 description 1
- 231100000419 toxicity Toxicity 0.000 description 1
- 230000001988 toxicity Effects 0.000 description 1
- 230000005026 transcription initiation Effects 0.000 description 1
- 238000010361 transduction Methods 0.000 description 1
- 230000026683 transduction Effects 0.000 description 1
- 230000014621 translational initiation Effects 0.000 description 1
- 230000001960 triggered effect Effects 0.000 description 1
- 238000009281 ultraviolet germicidal irradiation Methods 0.000 description 1
- 239000013603 viral vector Substances 0.000 description 1
- 238000004065 wastewater treatment Methods 0.000 description 1
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Chemical compound O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 1
- 238000001238 wet grinding Methods 0.000 description 1
- 101150077833 xlnA gene Proteins 0.000 description 1
- 101150021205 xlnB gene Proteins 0.000 description 1
- 101150068226 xlnC gene Proteins 0.000 description 1
- 101150011516 xlnD gene Proteins 0.000 description 1
- 101150034760 xlnR gene Proteins 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/80—Vectors or expression systems specially adapted for eukaryotic hosts for fungi
- C12N15/81—Vectors or expression systems specially adapted for eukaryotic hosts for fungi for yeasts
- C12N15/815—Vectors or expression systems specially adapted for eukaryotic hosts for fungi for yeasts for yeasts other than Saccharomyces
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N1/00—Microorganisms, e.g. protozoa; Compositions thereof; Processes of propagating, maintaining or preserving microorganisms or compositions thereof; Processes of preparing or isolating a composition containing a microorganism; Culture media therefor
- C12N1/14—Fungi; Culture media therefor
- C12N1/16—Yeasts; Culture media therefor
- C12N1/18—Baker's yeast; Brewer's yeast
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/0004—Oxidoreductases (1.)
- C12N9/0006—Oxidoreductases (1.) acting on CH-OH groups as donors (1.1)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/0004—Oxidoreductases (1.)
- C12N9/0008—Oxidoreductases (1.) acting on the aldehyde or oxo group of donors (1.2)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/93—Ligases (6)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/02—Preparation of oxygen-containing organic compounds containing a hydroxy group
- C12P7/04—Preparation of oxygen-containing organic compounds containing a hydroxy group acyclic
- C12P7/06—Ethanol, i.e. non-beverage
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/02—Preparation of oxygen-containing organic compounds containing a hydroxy group
- C12P7/04—Preparation of oxygen-containing organic compounds containing a hydroxy group acyclic
- C12P7/06—Ethanol, i.e. non-beverage
- C12P7/08—Ethanol, i.e. non-beverage produced as by-product or from waste or cellulosic material substrate
- C12P7/10—Ethanol, i.e. non-beverage produced as by-product or from waste or cellulosic material substrate substrate containing cellulosic material
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y101/00—Oxidoreductases acting on the CH-OH group of donors (1.1)
- C12Y101/01—Oxidoreductases acting on the CH-OH group of donors (1.1) with NAD+ or NADP+ as acceptor (1.1.1)
- C12Y101/01001—Alcohol dehydrogenase (1.1.1.1)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y101/00—Oxidoreductases acting on the CH-OH group of donors (1.1)
- C12Y101/01—Oxidoreductases acting on the CH-OH group of donors (1.1) with NAD+ or NADP+ as acceptor (1.1.1)
- C12Y101/01008—Glycerol-3-phosphate dehydrogenase (NAD+) (1.1.1.8)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y102/00—Oxidoreductases acting on the aldehyde or oxo group of donors (1.2)
- C12Y102/01—Oxidoreductases acting on the aldehyde or oxo group of donors (1.2) with NAD+ or NADP+ as acceptor (1.2.1)
- C12Y102/0101—Acetaldehyde dehydrogenase (acetylating) (1.2.1.10)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y602/00—Ligases forming carbon-sulfur bonds (6.2)
- C12Y602/01—Acid-Thiol Ligases (6.2.1)
- C12Y602/01001—Acetate-CoA ligase (6.2.1.1)
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y02—TECHNOLOGIES OR APPLICATIONS FOR MITIGATION OR ADAPTATION AGAINST CLIMATE CHANGE
- Y02E—REDUCTION OF GREENHOUSE GAS [GHG] EMISSIONS, RELATED TO ENERGY GENERATION, TRANSMISSION OR DISTRIBUTION
- Y02E50/00—Technologies for the production of fuel of non-fossil origin
- Y02E50/10—Biofuels, e.g. bio-diesel
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y02—TECHNOLOGIES OR APPLICATIONS FOR MITIGATION OR ADAPTATION AGAINST CLIMATE CHANGE
- Y02P—CLIMATE CHANGE MITIGATION TECHNOLOGIES IN THE PRODUCTION OR PROCESSING OF GOODS
- Y02P20/00—Technologies relating to chemical industry
- Y02P20/50—Improvements relating to the production of bulk chemicals
- Y02P20/52—Improvements relating to the production of bulk chemicals using catalysts, e.g. selective catalysts
Landscapes
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- Health & Medical Sciences (AREA)
- Engineering & Computer Science (AREA)
- Wood Science & Technology (AREA)
- Zoology (AREA)
- Genetics & Genomics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Biotechnology (AREA)
- Microbiology (AREA)
- Biomedical Technology (AREA)
- Mycology (AREA)
- Medicinal Chemistry (AREA)
- Molecular Biology (AREA)
- General Chemical & Material Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Botany (AREA)
- Tropical Medicine & Parasitology (AREA)
- Virology (AREA)
- Physics & Mathematics (AREA)
- Biophysics (AREA)
- Plant Pathology (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
Description
アセチルコエンザイムA+NADH+H+<−>アセトアルデヒド+NAD++コエンザイムA。
菌株の構築および維持
使用したサッカロマイセス・セレビシエ株(表1)は、CEN.PKファミリー由来であり、遺伝学および生理学を組み合わせた研究に好適なバックグランドとして以前確認された(ファン・ダイケン(van Dijken)ら(2000年)エンザイム・アンド・マイクロバイアル・テクノロジー第26巻:p.706−714)。
attBl配列およびattB2配列をそれぞれ含むプライマー対mhpF−FW(5’−GGGGACAAGTTTGTACAAAAAAGCAGGCTATGAGTAAGCGTAAAGTCGCCATTATCGG−3’[配列番号3])およびmhpF−RV(5’−GGGGACCACTTTGTACAAGAAAGCTGGGTGTTCATGCCGCTTCTCCTGCCTTGC−3’、[配列番号4])を用いて、E.コリmhpF遺伝子(EMBL受託番号Y09555.7)を、E.コリK12菌株JM109ゲノムDNAからPCR増幅した。製造者の仕様に従いPhusion(登録商標)ホット・スタート・ハイフィデリティーDNAポリメラーゼ(Hot Start High−Fidelity DNA Polymerase)(フィンザイム・オイ(Finnzymes Oy)、エスポー(Espoo)、フィンランド)を用いて、以下の条件のBiometra TGradientサーモサイクラー(バイオメトラ(Biometra)、ゲッティンゲン(Goettingen)、ドイツ)でポリメラーゼ連鎖反応(PCR)を行った:変性98℃で10秒、アニーリング30秒、伸長72℃で25サイクル。1011bpのPCR産物を、Gateway(登録商標)クローニングテクノロジー(cloning technology)(インビトロジェン、カールズバッド、カリフォルニア、米国)を用いてクローニングした。BP反応によりプラスミドpDONR221を用いてエントリークローンを作製し、プラスミドpUD64と命名した。このエントリークローン、および多コピープラスミドpAG426GPD−ccdB(アドジーン(Addgene)、ケンブリッジ、マサチューセッツ州、米国)から、LR反応を利用して酵母発現プラスミドpUDE43を構築した。Z−Competent(商標)E.トランスフォーメーションキット(Transformation Kit)(ザイモリサーチ・コーポレーション(Zymoresearch Corporation)、オレンジ、米国)に従い、コンピテントなE.コリK12菌株JM109への組換え反応産物の形質転換を行い、アンピシリン(100mg l−1)あるいはカナマイシン(50mg.l−1)のどちらかを含むLB培地に蒔いた。酵母形質転換は、バークら(メソッズ・イン・イースト・ジェネティックス(2000年)、コールド・スプリング・ハーバー・ラボラトリー・プレス、プレーンビュー、ニューヨーク)に従い実施した。酵母発現プラスミドによる形質転換後、細胞を合成培地に蒔いた。多コピープラスミドpUDE43の挿入が成功したかを、クローニング用のプライマー対を用いてPCR診断により確認した。
合成培地(46)を用いて30℃で振盪フラスコ培養を行った。培地のpHは、滅菌の前に2MのKOHで6.0に調整した。500mL振盪フラスコ中に20g l−1のグルコースを含む100mlの培地に、(1mlの)凍結した保存培養物を接種して、前培養液を調製した。Innova(登録商標)インキュベーターシェーカー(200rpm、ニューブランズウィックサイエンティフィック(New Brunswick Scientific)、ニュージャージー州、米国)を用いた30℃で24時間のインキュベーション後、培養物をバイオリアクターに移した。
予め秤量したニトロセルロースフィルター(細孔サイズ0.45μm;ゲルマンラボラトリー(Gelman Laboratory)、アナーバー(Ann Arbor)、米国)により所定の時間間隔で培養サンプル(10ml)を濾過した。培地の除去後、フィルターを脱イオン水で洗浄し、電子レンジ(ボッシュ(Bosch)、シュトゥットガルト(Stuttgart)、ドイツ)で350W、20min乾燥させ、秤量した。2反復測定のばらつきは1%未満であった。さらに、Novaspec(登録商標)II分光光度計を用いて波長660nmの光学密度の測定値により培養増殖もモニターした。
排気ガスを冷却器(condensor)(2℃)で冷却し、パーマピュアドライヤー(Permapure dryer)タイプMD−110−48P−4(パーマピュア、トムズリバー(Toms River)、米国)で乾燥させた。酸素および二酸化炭素の濃度をNGA 2000アナライザー(ローズマウント・アナリティカル(Rosemount Analytical)、オービル(Orrville)、米国)で判定した。排気ガス流速および二酸化炭素生成速度を、以前記載されたように判定した(3)。これらのバイオマス特有の速度を算出する際、培養サンプルの除去により起こる体積変化の補正を行った。
バイオラッド(Biorad)HPX 87Hカラム(バイオラッド、ハーキュリーズ(Hercules)、米国)を含むウォーターズアライアンス(Waters Alliance)2690HPLC(ウォーターズ、ミルフォード(Milford)、米国)を用いたHPLC解析で、培養サンプルの遠心分離により得られた上清を、グルコース、酢酸、コハク酸、乳酸、グリセロールおよびエタノールについて解析した。カラムは、0.5g l−1のH2SO4を用いて60℃、0.6ml min−1の流速で溶出した。検出は、ウォーターズ2410屈折率検出器、およびウォーターズ2487 UV検出器により行った。さらに、酵素的測定キット(アールバイオファーム(R−Biopharm)AG、ダルムシュタット(Darmstadt)、ドイツ)を用いて最初と最後のグリセロール濃度を判定した。窒素ガスをスパージするバイオレアクターでの培養の過程で、オフガスによりエタノールのかなりの部分が失われる。これを補正するため、無菌合成培地を用いて同一条件下、異なる有効容積で運転したバイオレアクターでエタノール蒸発のカイネティクスを解析した。得られた容積依存性のエタノール蒸発定数(この構成の場合、リットル単位の容積で割った商0.0080に相当し、h−1で表す)を用いて、培養上清におけるエタノール濃度のHPLC測定値を補正し、サンプリングにより起こる容積の変化を考慮した。
以前記載されたように指数関数的に増殖する嫌気回分培養から、NAD+依存性アセトアルデヒドデヒドロゲナーゼ(アセチル化)活性アッセイのための細胞抽出物を調製した(アボット(Abbott)ら、アプライド・アンド・エンバイロンメンタル・マイクロバイオロジー、第75巻:p.2320−2325)。340nmでNADHの酸化をモニタリングすることにより、NAD+依存性アセトアルデヒドデヒドロゲナーゼ(アセチル化)活性を30℃で測定した。反応混合物(全容積1ml)は、50mMのリン酸カリウム緩衝液(pH7.5)、15mMのNADHおよび細胞抽出物を含んでいた。0.5mMのアセチルコエンザイムAを添加して反応を開始した。グリセロール3−リン酸デヒドロゲナーゼ(EC1.1.1.8)活性の判定では、リン酸塩緩衝液をトリエタノールアミン緩衝液(10mM、pH5)(5、19)と交換したこと以外は上記のように細胞抽出物を調製した。グリセロール−3−リン酸デヒドロゲナーゼ活性は、以前記載されたように細胞抽出物を用いて30℃でアッセイした(ブロムバーグ(Blomberg)およびアドラー(Adler)(1989年)、ジャーナル・オブ・バクテリオロジー第171巻:p.1087−1092。反応速度は、添加した細胞抽出物の量に比例した。タンパク質濃度は、ウシ血清アルブミンを標準物質としてローリー(Lowry)法により判定した(ローリーet al(1951年)ジャーナル・オブ・バイオロジカル・ケミストリー第193巻:p.265−275)。
嫌気回分培養における増殖および産物形成
原栄養性の基準菌株S.セレビシエIME076(GPD1 GPD2)の培養物に2.0g l−1の酢酸を補充した場合、比増殖速度(0.32h−1)は、酢酸の非存在下で増殖させた培養物で報告された比増殖速度(0.34h−1)、カイパー(Kuyper)ら(2005年)FEMSイーストリサーチ第5巻:p.399−409と同一であった。酢酸の添加によりバイオマスの収率は若干低下し、その結果、酢酸の非存在下で増殖した培養物と比較してグルコースのグリセロールの収率も低下した(図2、表3)。
本研究は、化学量論的には、S.セレビシエの嫌気的増殖のレドックスシンクとしてのグリセロールの役割を、アセテートのエタノールへのNADH依存性還元における線形経路で十分に置き換えられるという原理の証拠となる。これは、リグノセルロース加水分解物など酢酸を含む供給原料からの大規模エタノール生成の興味深い視点を与える。
嫌気回分培養における増殖および生成物の形成
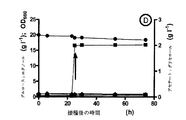
gpd1Δ gpd2ΔS.セレビシエ株におけるシュードモナス・エスピーCF600dmpF遺伝子の発現の結果は、同じ菌株におけるE.コリmhpF遺伝子の発現で得られた結果と類似していた。嫌気性回分発酵では、菌株IMZ132(gpd1Δ gpd2ΔmhpF)と類似して、2.0g l−1の酢酸を含む培地の補充に伴いdmpF遺伝子の機能発現は指数増殖し、比増殖速度は0.11h−1であった(図1)。菌株IMZ130(gpd1Δ gpd2ΔdmpF)の回分時間(55h)は、菌株IMZ132(40h)よりやや長かった。培養中、グルコースの初濃度20g l−1は完全に消化される一方、グリセロールの形成が観察された。同時に、アセテートは初濃度2.1g l−1から最終濃度1.6g l−1に消化された。エタノールが主な有機生成物であり、グルコースのエタノール収率は0.48g g−1(エタノール蒸発で補正した収率)を示した。同じ条件下で増殖させた基準菌株の培養で観察されたのと同様に、少量のスクシネートおよびラクテートが生成された。
配列表
SEQ ID NO: 1
E. coli mhpF gene Acetaldehyde dehydrogenase, acylating
ATGAGTAAGCGTAAAGTCGCCATTATCGGTTCTGGCAACATTGGTACCGATCTGATGATTAAAATTTTGCGTCACGGTCAGCATCTGGAGATGGCGGTGATGGTTGGCATTGATCCTCAGTCCGACGGTCTGGCGCGCGCCAGACGTATGGGCGTCGCCACCACCCATGAAGGGGTGATCGGACTGATGAACATGCCTGAATTTGCTGATATCGACATTGTATTTGATGCGACCAGCGCCGGTGCTCATGTGAAAAACGATGCCGCTTTACGCGAAGCGAAACCGGATATTCGCTTAATTGACCTGACGCCTGCTGCCATCGGCCCTTACTGCGTGCCGGTGGTTAACCTCGAGGCGAACGTCGATCAACTGAACGTCAACATGGTCACCTGCGGCGGCCAGGCCACCATTCCAATGGTGGCGGCAGTTTCACGCGTGGCGCGTGTTCATTACGCCGAAATTATCGCTTCTATCGCCAGTAAATCTGCCGGACCTGGCACGCGTGCCAATATCGATGAATTTACGGAAACCACTTCCCGAGCCATTGAAGTGGTGGGCGGCGCGGCAAAAGGGAAGGCGATTATTGTGCTTAACCCAGCA
GAGCCACCGTTGATGATGCGTGACACGGTGTATGTATTGAGCGACGAAGCTTCACAAGATGATATCGAAGCCTCAATCAATGAAATGGCTGAGGCGGTGCAGGCTTACGTACCGGGTTATCGCCTGAAACAGCGCGTGCAGTTTGAAGTTATCCCGCAGGATAAACCGGTCAATTTACCGGGCGTGGGGCAATTCTCCGGACTGAAAACAGCGGTCTGGCTGGAAGTCGAAGGCGCAGCGCATTATCTGCCTGCCTATGCGGGCAACCTCGACATTATGACTTCCAGTGCGCTGGCGACAGCGGAAAAAATGGCCCAGTCACTGGCGCGCAAGGCAGGAGAAGCGGCATGA
SEQ ID NO: 2
E. coli Acetaldehyde dehydrogenase OS=Escherichia coli (strain K12) GN=mhpF PE=1 SV=1
MSKRKVAIIGSGNIGTDLMIKILRHGQHLEMAVMVGIDPQSDGLARARRMGVATTHEGVIGLMNMPEFADIDIVFDATSAGAHVKNDAALREAKPDIRLIDLTPAAIGPYCVPVVNLEANVDQLNVNMVTCGGQATIPMVAAVSRVARVHYAEIIASIASKSAGPGTRANIDEFTETTSRAIEVVGGAAKGKAIIVLNPAEPPLMMRDTVYVLSDEASQDDIEASINEMAEAVQAYVPGYRLKQRVQFEVIPQDKPVNLPGVGQFSGLKTAVWLEVEGAAHYLPAYAGNLDIMTSSALATAEKMAQSLARKAGEAA
SEQ ID NO: 3
primer mhpF-FW
GGGGACAAGTTTGTACAAAAAAGCAGGCTATGAGTAAGCGTAAAGTCGCCATTATCGG
SEQ ID NO: 4
primer mhpF-RV GGGGACCACTTTGTACAAGAAAGCTGGGTGTTCATGCCGCTTCTCCTGCCTTGC
SEQ ID NO: 5 GPD1/YDL022W fw gene disruption primer
TTGTACACCCCCCCCCTCCACAAACACAAATATTGATAATATAAAcagctgaagcttcgtacgc
SEQ ID NO: 6 GPD1/YDL022W rv gene disruption primer
AATCTAATCTTCATGTAGATCTAATTCTTCAATCATGTCCGGCGgcataggccactagtggatctg
SEQ ID NO: 7 GPD2/YOL059W fw gene disruption primer
TCAATTCTCTTTCCCTTTCCTTTTCCTTCGCTCCCCTTCCTTATC ccaggctgaagcttcgtacg
SEQ ID NO: 8 GPD2/YOL059W rv gene disruption primer
GTGTCTATTCGTCATCGATGTCTAGCTCTTCAATCATCTCCGGTAGgcataggccactagtggatc
SEQ ID NO: 9 GPD1/YDL022W fw verification primer
CCCACCCACACCACCAATAC
SEQ ID NO: 10 GPD1/YDL022Wrv verification primer
CGGACGCCAGATGCTAGAAG
SEQ ID NO: 11 GPD2/YOL059W fw verification primer
GTTCAGCAGCTCTTCTCTAC
SEQ ID NO: 12 GPD2/YOL059W rv verification primer
CCAAATGCGACATGAGTCAC
SEQ ID NO: 13 GPD1/YDL022W fw disruption cassette specific verification primer
CGCACGTCAAGACTGTCAAG
SEQ ID NO: 14 GPD1/YDL022W rv disruption cassette specific verification primer
TCGTATGTGAATGCTGGTCG
SEQ ID NO: 15 GPD2/YOL059W fw disruption cassette specific verification primer
CGCACGTCAAGACTGTCAAG
SEQ ID NO: 16 GPD2/YOL059W rv disruption cassette specific verification primer
TCGTATGTGAATGCTGGTCG
SEQ ID NO: 17 S. cerevisiae ACS 1 gene
ATGTCGCCCTCTGCCGTACAATCATCAAAACTAGAAGAACAGTCAAGTGAAATTGACAAG
TTGAAAGCAAAAATGTCCCAGTCTGCCGCCACTGCGCAGCAGAAGAAGGAACATGAGTAT
GAACATTTGACTTCGGTCAAGATCGTGCCACAACGGCCCATCTCAGATAGACTGCAGCCC
GCAATTGCTACCCACTATTCTCCACACTTGGACGGGTTGCAGGACTATCAGCGCTTGCAC
AAGGAGTCTATTGAAGACCCTGCTAAGTTCTTCGGTTCTAAAGCTACCCAATTTTTAAAC
TGGTCTAAGCCATTCGATAAGGTGTTCATCCCAGACCCTAAAACGGGCAGGCCCTCCTTC
CAGAACAATGCATGGTTCCTCAACGGCCAATTAAACGCCTGTTACAACTGTGTTGACAGA
CATGCCTTGAAGACTCCTAACAAGAAAGCCATTATTTTCGAAGGTGACGAGCCTGGCCAA
GGCTATTCCATTACCTACAAGGAACTACTTGAAGAAGTTTGTCAAGTGGCACAAGTGCTG
ACTTACTCTATGGGCGTTCGCAAGGGCGATACTGTTGCCGTGTACATGCCTATGGTCCCA
GAAGCAATCATAACCTTGTTGGCCATTTCCCGTATCGGTGCCATTCACTCCGTAGTCTTT
GCCGGGTTTTCTTCCAACTCCTTGAGAGATCGTATCAACGATGGGGACTCTAAAGTTGTC
ATCACTACAGATGAATCCAACAGAGGTGGTAAAGTCATTGAGACTAAAAGAATTGTTGAT
GACGCGCTAAGAGAGACCCCAGGCGTGAGACACGTCTTGGTTTATAGAAAGACCAACAAT
CCATCTGTTGCTTTCCATGCCCCCAGAGATTTGGATTGGGCAACAGAAAAGAAGAAATAC
AAGACCTACTATCCATGCACACCCGTTGATTCTGAGGATCCATTATTCTTGTTGTATACG
TCTGGTTCTACTGGTGCCCCCAAGGGTGTTCAACATTCTACCGCAGGTTACTTGCTGGGA
GCTTTGTTGACCATGCGCTACACTTTTGACACTCACCAAGAAGACGTTTTCTTCACAGCT
GGAGACATTGGCTGGATTACAGGCCACACTTATGTGGTTTATGGTCCCTTACTATATGGT
TGTGCCACTTTGGTCTTTGAAGGGACTCCTGCGTACCCAAATTACTCCCGTTATTGGGAT
ATTATTGATGAACACAAAGTCACCCAATTTTATGTTGCGCCAACTGCTTTGCGTTTGTTG
AAAAGAGCTGGTGATTCCTACATCGAAAATCATTCCTTAAAATCTTTGCGTTGCTTGGGT
TCGGTCGGTGAGCCAATTGCTGCTGAAGTTTGGGAGTGGTACTCTGAAAAAATAGGTAAA
AATGAAATCCCCATTGTAGACACCTACTGGCAAACAGAATCTGGTTCGCATCTGGTCACC
CCGCTGGCTGGTGGTGTTACACCAATGAAACCGGGTTCTGCCTCATTCCCCTTCTTCGGT
ATTGATGCAGTTGTTCTTGACCCTAACACTGGTGAAGAACTTAACACCAGCCACGCAGAG
GGTGTCCTTGCCGTCAAAGCTGCATGGCCATCATTTGCAAGAACTATTTGGAAAAATCAT
GATAGGTATCTAGACACTTATTTGAACCCTTACCCTGGCTACTATTTCACTGGTGATGGT
GCTGCAAAGGATAAGGATGGTTATATCTGGATTTTGGGTCGTGTAGACGATGTGGTGAAC
GTCTCTGGTCACCGTCTGTCTACCGCTGAAATTGAGGCTGCTATTATCGAAGATCCAATT
GTGGCCGAGTGTGCTGTTGTCGGATTCAACGATGACTTGACTGGTCAAGCAGTTGCTGCA
TTTGTGGTGTTGAAAAACAAATCTAGTTGGTCCACCGCAACAGATGATGAATTACAAGAT
ATCAAGAAGCATTTGGTCTTTACTGTTAGAAAAGACATCGGGCCATTTGCCGCACCAAAA
TTGATCATTTTAGTGGATGACTTGCCCAAGACAAGATCCGGCAAAATTATGAGACGTATT
TTAAGAAAAATCCTAGCAGGAGAAAGTGACCAACTAGGCGACGTTTCTACATTGTCAAAC
CCTGGCATTGTTAGACATCTAATTGATTCGGTCAAGTTGTAA
SEQ ID NO: 18 S. cerevisiae ACS 2 gene
ATGACAATCAAGGAACATAAAGTAGTTTATGAAGCTCACAACGTAAAGGCTCTTAAGGCT
CCTCAACATTTTTACAACAGCCAACCCGGCAAGGGTTACGTTACTGATATGCAACATTAT
CAAGAAATGTATCAACAATCTATCAATGAGCCAGAAAAATTCTTTGATAAGATGGCTAAG
GAATACTTGCATTGGGATGCTCCATACACCAAAGTTCAATCTGGTTCATTGAACAATGGT
GATGTTGCATGGTTTTTGAACGGTAAATTGAATGCATCATACAATTGTGTTGACAGACAT
GCCTTTGCTAATCCCGACAAGCCAGCTTTGATCTATGAAGCTGATGACGAATCCGACAAC
AAAATCATCACATTTGGTGAATTACTCAGAAAAGTTTCCCAAATCGCTGGTGTCTTAAAA
AGCTGGGGCGTTAAGAAAGGTGACACAGTGGCTATCTATTTGCCAATGATTCCAGAAGCG
GTCATTGCTATGTTGGCTGTGGCTCGTATTGGTGCTATTCACTCTGTTGTCTTTGCTGGG
TTCTCCGCTGGTTCGTTGAAAGATCGTGTCGTTGACGCTAATTCTAAAGTGGTCATCACT
TGTGATGAAGGTAAAAGAGGTGGTAAGACCATCAACACTAAAAAAATTGTTGACGAAGGT
TTGAACGGAGTCGATTTGGTTTCCCGTATCTTGGTTTTCCAAAGAACTGGTACTGAAGGT
ATTCCAATGAAGGCCGGTAGAGATTACTGGTGGCATGAGGAGGCCGCTAAGCAGAGAACT
TACCTACCTCCTGTTTCATGTGACGCTGAAGATCCTCTATTTTTATTATACACTTCCGGT
TCCACTGGTTCTCCAAAGGGTGTCGTTCACACTACAGGTGGTTATTTATTAGGTGCCGCT
TTAACAACTAGATACGTTTTTGATATTCACCCAGAAGATGTTCTCTTCACTGCCGGTGAC
GTCGGCTGGATCACGGGTCACACCTATGCTCTATATGGTCCATTAACCTTGGGTACCGCC
TCAATAATTTTCGAATCCACTCCTGCCTACCCAGATTATGGTAGATATTGGAGAATTATC
CAACGTCACAAGGCTACCCATTTCTATGTGGCTCCAACTGCTTTAAGATTAATCAAACGT
GTAGGTGAAGCCGAAATTGCCAAATATGACACTTCCTCATTACGTGTCTTGGGTTCCGTC
GGTGAACCAATCTCTCCAGACTTATGGGAATGGTATCATGAAAAAGTGGGTAACAAAAAC
TGTGTCATTTGTGACACTATGTGGCAAACAGAGTCTGGTTCTCATTTAATTGCTCCTTTG
GCAGGTGCTGTCCCAACAAAACCTGGTTCTGCTACCGTGCCATTCTTTGGTATTAACGCT
TGTATCATTGACCCTGTTACAGGTGTGGAATTAGAAGGTAATGATGTCGAAGGTGTCCTT
GCCGTTAAATCACCATGGCCATCAATGGCTAGATCTGTTTGGAACCACCACGACCGTTAC
ATGGATACTTACTTGAAACCTTATCCTGGTCACTATTTCACAGGTGATGGTGCTGGTAGA
GATCATGATGGTTACTACTGGATCAGGGGTAGAGTTGACGACGTTGTAAATGTTTCCGGT
CATAGATTATCCACATCAGAAATTGAAGCATCTATCTCAAATCACGAAAACGTCTCGGAA
GCTGCTGTTGTCGGTATTCCAGATGAATTGACCGGTCAAACCGTCGTTGCATATGTTTCC
CTAAAAGATGGTTATCTACAAAACAACGCTACTGAAGGTGATGCAGAACACATCACACCA
GATAATTTACGTAGAGAATTGATCTTACAAGTTAGGGGTGAGATTGGTCCTTTCGCCTCA
CCAAAAACCATTATTCTAGTTAGAGATCTACCAAGAACAAGGTCAGGAAAGATTATGAGA
AGAGTTCTAAGAAAGGTTGCTTCTAACGAAGCCGAACAGCTAGGTGACCTAACTACTTTG
GCCAACCCAGAAGTTGTACCTGCCATCATTTCTGCTGTAGAGAACCAATTTTTCTCTCAA
AAAAAGAAATAA
SEQ ID NO: 19 S. cerevisiae ADH1
ATGTCTATCCCAGAAACTCAAAAAGGTGTTATCTTCTACGAATCCCACGGTAAGTTGGAA
TACAAAGATATTCCAGTTCCAAAGCCAAAGGCCAACGAATTGTTGATCAACGTTAAATAC
TCTGGTGTCTGTCACACTGACTTGCACGCTTGGCACGGTGACTGGCCATTGCCAGTTAAG
CTACCATTAGTCGGTGGTCACGAAGGTGCCGGTGTCGTTGTCGGCATGGGTGAAAACGTT
AAGGGCTGGAAGATCGGTGACTACGCCGGTATCAAATGGTTGAACGGTTCTTGTATGGCC
TGTGAATACTGTGAATTGGGTAACGAATCCAACTGTCCTCACGCTGACTTGTCTGGTTAC
ACCCACGACGGTTCTTTCCAACAATACGCTACCGCTGACGCTGTTCAAGCCGCTCACATT
CCTCAAGGTACCGACTTGGCCCAAGTCGCCCCCATCTTGTGTGCTGGTATCACCGTCTAC
AAGGCTTTGAAGTCTGCTAACTTGATGGCCGGTCACTGGGTTGCTATCTCCGGTGCTGCT
GGTGGTCTAGGTTCTTTGGCTGTTCAATACGCCAAGGCTATGGGTTACAGAGTCTTGGGT
ATTGACGGTGGTGAAGGTAAGGAAGAATTATTCAGATCCATCGGTGGTGAAGTCTTCATT
GACTTCACTAAGGAAAAGGACATTGTCGGTGCTGTTCTAAAGGCCACTGACGGTGGTGCT
CACGGTGTCATCAACGTTTCCGTTTCCGAAGCCGCTATTGAAGCTTCTACCAGATACGTT
AGAGCTAACGGTACCACCGTTTTGGTCGGTATGCCAGCTGGTGCCAAGTGTTGTTCTGAT
GTCTTCAACCAAGTCGTCAAGTCCATCTCTATTGTTGGTTCTTACGTCGGTAACAGAGCT
GACACCAGAGAAGCTTTGGACTTCTTCGCCAGAGGTTTGGTCAAGTCTCCAATCAAGGTT
GTCGGCTTGTCTACCTTGCCAGAAATTTACGAAAAGATGGAAAAGGGTCAAATCGTTGGT
AGATACGTTGTTGACACTTCTAAATAA
SEQ ID NO: 20 S. cerevisiae ADH2
ATGTCTATTCCAGAAACTCAAAAAGCCATTATCTTCTACGAATCCAACGGCAAGTTGGAG
CATAAGGATATCCCAGTTCCAAAGCCAAAGCCCAACGAATTGTTAATCAACGTCAAGTAC
TCTGGTGTCTGCCACACCGATTTGCACGCTTGGCATGGTGACTGGCCATTGCCAACTAAG
TTACCATTAGTTGGTGGTCACGAAGGTGCCGGTGTCGTTGTCGGCATGGGTGAAAACGTT
AAGGGCTGGAAGATCGGTGACTACGCCGGTATCAAATGGTTGAACGGTTCTTGTATGGCC
TGTGAATACTGTGAATTGGGTAACGAATCCAACTGTCCTCACGCTGACTTGTCTGGTTAC
ACCCACGACGGTTCTTTCCAAGAATACGCTACCGCTGACGCTGTTCAAGCCGCTCACATT
CCTCAAGGTACTGACTTGGCTGAAGTCGCGCCAATCTTGTGTGCTGGTATCACCGTATAC
AAGGCTTTGAAGTCTGCCAACTTGAGAGCAGGCCACTGGGCGGCCATTTCTGGTGCTGCT
GGTGGTCTAGGTTCTTTGGCTGTTCAATATGCTAAGGCGATGGGTTACAGAGTCTTAGGT
ATTGATGGTGGTCCAGGAAAGGAAGAATTGTTTACCTCGCTCGGTGGTGAAGTATTCATC
GACTTCACCAAAGAGAAGGACATTGTTAGCGCAGTCGTTAAGGCTACCAACGGCGGTGCC
CACGGTATCATCAATGTTTCCGTTTCCGAAGCCGCTATCGAAGCTTCTACCAGATACTGT
AGGGCGAACGGTACTGTTGTCTTGGTTGGTTTGCCAGCCGGTGCAAAGTGCTCCTCTGAT
GTCTTCAACCACGTTGTCAAGTCTATCTCCATTGTCGGCTCTTACGTGGGGAACAGAGCT
GATACCAGAGAAGCCTTAGATTTCTTTGCCAGAGGTCTAGTCAAGTCTCCAATAAAGGTA
GTTGGCTTATCCAGTTTACCAGAAATTTACGAAAAGATGGAGAAGGGCCAAATTGCTGGT
AGATACGTTGTTGACACTTCTAAATAA
SEQ ID NO: 21 S. cerevisiae ADH3
ATGTTGAGAACGTCAACATTGTTCACCAGGCGTGTCCAACCAAGCCTATTTTCTAGAAAC
ATTCTTAGATTGCAATCCACAGCTGCAATCCCTAAGACTCAAAAAGGTGTCATCTTTTAT
GAGAATAAGGGGAAGCTGCATTACAAAGATATCCCTGTCCCCGAGCCTAAGCCAAATGAA
ATTTTAATCAACGTTAAATATTCTGGTGTATGTCACACCGATTTACATGCTTGGCACGGC
GATTGGCCATTACCTGTTAAACTACCATTAGTAGGTGGTCATGAAGGTGCTGGTGTAGTT
GTCAAACTAGGTTCCAATGTCAAGGGCTGGAAAGTCGGTGATTTAGCAGGTATCAAATGG
CTGAACGGTTCTTGTATGACATGCGAATTCTGTGAATCAGGTCATGAATCAAATTGTCCA
GATGCTGATTTATCTGGTTACACTCATGATGGTTCTTTCCAACAATTTGCGACCGCTGAT
GCTATTCAAGCCGCCAAAATTCAACAGGGTACCGACTTGGCCGAAGTAGCCCCAATATTA
TGTGCTGGTGTTACTGTATATAAAGCACTAAAAGAGGCAGACTTGAAAGCTGGTGACTGG
GTTGCCATCTCTGGTGCTGCAGGTGGCTTGGGTTCCTTGGCCGTTCAATATGCAACTGCG
ATGGGTTACAGAGTTCTAGGTATTGATGCAGGTGAGGAAAAGGAAAAACTTTTCAAGAAA
TTGGGGGGTGAAGTATTCATCGACTTTACTAAAACAAAGAATATGGTTTCTGACATTCAA
GAAGCTACCAAAGGTGGCCCTCATGGTGTCATTAACGTTTCCGTTTCTGAAGCCGCTATT
TCTCTATCTACGGAATATGTTAGACCATGTGGTACCGTCGTTTTGGTTGGTTTGCCCGCT
AACGCCTACGTTAAATCAGAGGTATTCTCTCATGTGGTGAAGTCCATCAATATCAAGGGT
TCTTATGTTGGTAACAGAGCTGATACGAGAGAAGCCTTAGACTTCTTTAGCAGAGGTTTG
ATCAAATCACCAATCAAAATTGTTGGATTATCTGAATTACCAAAGGTTTATGACTTGATG
GAAAAGGGCAAGATTTTGGGTAGATACGTCGTCGATACTAGTAAATAA
SEQ ID NO: 22 S. cerevisiae ADH4
ATGTCTTCCGTTACTGGGTTTTACATTCCACCAATCTCTTTCTTTGGTGAAGGTGCTTTA
GAAGAAACCGCTGATTACATCAAAAACAAGGATTACAAAAAGGCTTTGATCGTTACTGAT
CCTGGTATTGCAGCTATTGGTCTCTCCGGTAGAGTCCAAAAGATGTTGGAAGAACGTGAC
TTAAACGTTGCTATCTATGACAAAACTCAACCAAACCCAAATATTGCCAATGTCACAGCT
GGTTTGAAGGTTTTGAAGGAACAAAACTCTGAAATTGTTGTTTCCATTGGTGGTGGTTCT
GCTCACGACAATGCTAAGGCCATTGCTTTATTGGCTACTAACGGTGGGGAAATCGGAGAC
TATGAAGGTGTCAATCAATCTAAGAAGGCTGCTTTACCACTATTTGCCATCAACACTACT
GCTGGTACTGCTTCCGAAATGACCAGATTCACTATTATCTCTAATGAAGAAAAGAAAATC
AAGATGGCTATCATTGACAACAACGTCACTCCAGCTGTTGCTGTCAACGATCCATCTACC
ATGTTTGGTTTGCCACCTGCTTTGACTGCTGCTACTGGTCTAGATGCTTTGACTCACTGT
ATCGAAGCTTATGTTTCCACCGCCTCTAACCCAATCACCGATGCCTGTGCTTTGAAGGGT
ATTGATTTGATCAATGAAAGCTTAGTCGCTGCATACAAAGACGGTAAAGACAAGAAGGCC
AGAACTGACATGTGTTACGCTGAATACTTGGCAGGTATGGCTTTCAACAATGCTTCTCTA
GGTTATGTTCATGCCCTTGCTCATCAACTTGGTGGTTTCTACCACTTGCCTCATGGTGTT
TGTAACGCTGTCTTGTTGCCTCATGTTCAAGAGGCCAACATGCAATGTCCAAAGGCCAAG
AAGAGATTAGGTGAAATTGCTTTGCATTTCGGTGCTTCTCAAGAAGATCCAGAAGAAACC
ATCAAGGCTTTGCACGTTTTAAACAGAACCATGAACATTCCAAGAAACTTGAAAGAATTA
GGTGTTAAAACCGAAGATTTTGAAATTTTGGCTGAACACGCCATGCATGATGCCTGCCAT
TTGACTAACCCAGTTCAATTCACCAAAGAACAAGTGGTTGCCATTATCAAGAAAGCCTAT
GAATATTAA
SEQ ID NO: 23 S. cerevisiae ADH5
ATGCCTTCGCAAGTCATTCCTGAAAAACAAAAGGCTATTGTCTTTTATGAGACAGATGGA
AAATTGGAATATAAAGACGTCACAGTTCCGGAACCTAAGCCTAACGAAATTTTAGTCCAC
GTTAAATATTCTGGTGTTTGTCATAGTGACTTGCACGCGTGGCACGGTGATTGGCCATTT
CAATTGAAATTTCCATTAATCGGTGGTCACGAAGGTGCTGGTGTTGTTGTTAAGTTGGGA
TCTAACGTTAAGGGCTGGAAAGTCGGTGATTTTGCAGGTATAAAATGGTTGAATGGGACT
TGCATGTCCTGTGAATATTGTGAAGTAGGTAATGAATCTCAATGTCCTTATTTGGATGGT
ACTGGCTTCACACATGATGGTACTTTTCAAGAATACGCAACTGCCGATGCCGTTCAAGCT
GCCCATATTCCACCAAACGTCAATCTTGCTGAAGTTGCCCCAATCTTGTGTGCAGGTATC
ACTGTTTATAAGGCGTTGAAAAGAGCCAATGTGATACCAGGCCAATGGGTCACTATATCC
GGTGCATGCGGTGGCTTGGGTTCTCTGGCAATCCAATACGCCCTTGCTATGGGTTACAGG
GTCATTGGTATCGATGGTGGTAATGCCAAGCGAAAGTTATTTGAACAATTAGGCGGAGAA
ATATTCATCGATTTCACGGAAGAAAAAGACATTGTTGGTGCTATAATAAAGGCCACTAAT
GGCGGTTCTCATGGAGTTATTAATGTGTCTGTTTCTGAAGCAGCTATCGAGGCTTCTACG
AGGTATTGTAGGCCCAATGGTACTGTCGTCCTGGTTGGTATGCCAGCTCATGCTTACTGC
AATTCCGATGTTTTCAATCAAGTTGTAAAATCAATCTCCATCGTTGGATCTTGTGTTGGA
AATAGAGCTGATACAAGGGAGGCTTTAGATTTCTTCGCCAGAGGTTTGATCAAATCTCCG
ATCCACTTAGCTGGCCTATCGGATGTTCCTGAAATTTTTGCAAAGATGGAGAAGGGTGAA
ATTGTTGGTAGATATGTTGTTGAGACTTCTAAATGA
SEQ ID NO 24: S. cerevisiae GPD1
ATGTCTGCTGCTGCTGATAGATTAAACTTAACTTCCGGCCACTTGAATGCTGGTAGAAAG
AGAAGTTCCTCTTCTGTTTCTTTGAAGGCTGCCGAAAAGCCTTTCAAGGTTACTGTGATT
GGATCTGGTAACTGGGGTACTACTATTGCCAAGGTGGTTGCCGAAAATTGTAAGGGATAC
CCAGAAGTTTTCGCTCCAATAGTACAAATGTGGGTGTTCGAAGAAGAGATCAATGGTGAA
AAATTGACTGAAATCATAAATACTAGACATCAAAACGTGAAATACTTGCCTGGCATCACT
CTACCCGACAATTTGGTTGCTAATCCAGACTTGATTGATTCAGTCAAGGATGTCGACATC
ATCGTTTTCAACATTCCACATCAATTTTTGCCCCGTATCTGTAGCCAATTGAAAGGTCAT
GTTGATTCACACGTCAGAGCTATCTCCTGTCTAAAGGGTTTTGAAGTTGGTGCTAAAGGT
GTCCAATTGCTATCCTCTTACATCACTGAGGAACTAGGTATTCAATGTGGTGCTCTATCT
GGTGCTAACATTGCCACCGAAGTCGCTCAAGAACACTGGTCTGAAACAACAGTTGCTTAC
CACATTCCAAAGGATTTCAGAGGCGAGGGCAAGGACGTCGACCATAAGGTTCTAAAGGCC
TTGTTCCACAGACCTTACTTCCACGTTAGTGTCATCGAAGATGTTGCTGGTATCTCCATC
TGTGGTGCTTTGAAGAACGTTGTTGCCTTAGGTTGTGGTTTCGTCGAAGGTCTAGGCTGG
GGTAACAACGCTTCTGCTGCCATCCAAAGAGTCGGTTTGGGTGAGATCATCAGATTCGGT
CAAATGTTTTTCCCAGAATCTAGAGAAGAAACATACTACCAAGAGTCTGCTGGTGTTGCT
GATTTGATCACCACCTGCGCTGGTGGTAGAAACGTCAAGGTTGCTAGGCTAATGGCTACT
TCTGGTAAGGACGCCTGGGAATGTGAAAAGGAGTTGTTGAATGGCCAATCCGCTCAAGGT
TTAATTACCTGCAAAGAAGTTCACGAATGGTTGGAAACATGTGGCTCTGTCGAAGACTTC
CCATTATTTGAAGCCGTATACCAAATCGTTTACAACAACTACCCAATGAAGAACCTGCCG
GACATGATTGAAGAATTAGATCTACATGAAGATTAG
SEQ ID NO: 25: S. cerevisiae GPD2
ATGCTTGCTGTCAGAAGATTAACAAGATACACATTCCTTAAGCGAACGCATCCGGTGTTA
TATACTCGTCGTGCATATAAAATTTTGCCTTCAAGATCTACTTTCCTAAGAAGATCATTA
TTACAAACACAACTGCACTCAAAGATGACTGCTCATACTAATATCAAACAGCACAAACAC
TGTCATGAGGACCATCCTATCAGAAGATCGGACTCTGCCGTGTCAATTGTACATTTGAAA
CGTGCGCCCTTCAAGGTTACAGTGATTGGTTCTGGTAACTGGGGGACCACCATCGCCAAA
GTCATTGCGGAAAACACAGAATTGCATTCCCATATCTTCGAGCCAGAGGTGAGAATGTGG
GTTTTTGATGAAAAGATCGGCGACGAAAATCTGACGGATATCATAAATACAAGACACCAG
AACGTTAAATATCTACCCAATATTGACCTGCCCCATAATCTAGTGGCCGATCCTGATCTT
TTACACTCCATCAAGGGTGCTGACATCCTTGTTTTCAACATCCCTCATCAATTTTTACCA
AACATAGTCAAACAATTGCAAGGCCACGTGGCCCCTCATGTAAGGGCCATCTCGTGTCTA
AAAGGGTTCGAGTTGGGCTCCAAGGGTGTGCAATTGCTATCCTCCTATGTTACTGATGAG
TTAGGAATCCAATGTGGCGCACTATCTGGTGCAAACTTGGCACCGGAAGTGGCCAAGGAG
CATTGGTCCGAAACCACCGTGGCTTACCAACTACCAAAGGATTATCAAGGTGATGGCAAG
GATGTAGATCATAAGATTTTGAAATTGCTGTTCCACAGACCTTACTTCCACGTCAATGTC
ATCGATGATGTTGCTGGTATATCCATTGCCGGTGCCTTGAAGAACGTCGTGGCACTTGCA
TGTGGTTTCGTAGAAGGTATGGGATGGGGTAACAATGCCTCCGCAGCCATTCAAAGGCTG
GGTTTAGGTGAAATTATCAAGTTCGGTAGAATGTTTTTCCCAGAATCCAAAGTCGAGACC
TACTATCAAGAATCCGCTGGTGTTGCAGATCTGATCACCACCTGCTCAGGCGGTAGAAAC
GTCAAGGTTGCCACATACATGGCCAAGACCGGTAAGTCAGCCTTGGAAGCAGAAAAGGAA
TTGCTTAACGGTCAATCCGCCCAAGGGATAATCACATGCAGAGAAGTTCACGAGTGGCTA
CAAACATGTGAGTTGACCCAAGAATTCCCATTATTCGAGGCAGTCTACCAGATAGTCTAC
AACAACGTCCGCATGGAAGACCTACCGGAGATGATTGAAGAGCTAGACATCGATGACGAA
TAG
SEQ ID NO 26: S. cerevisiae GPP1
ATGCCTTTGACCACAAAACCTTTATCTTTGAAAATCAACGCCGCTCTATTCGATGTTGAC
GGTACCATCATCATCTCTCAACCAGCCATTGCTGCTTTCTGGAGAGATTTCGGTAAAGAC
AAGCCTTACTTCGATGCCGAACACGTTATTCACATCTCTCACGGTTGGAGAACTTACGAT
GCCATTGCCAAGTTCGCTCCAGACTTTGCTGATGAAGAATACGTTAACAAGCTAGAAGGT
GAAATCCCAGAAAAGTACGGTGAACACTCCATCGAAGTTCCAGGTGCTGTCAAGTTGTGT
AATGCTTTGAACGCCTTGCCAAAGGAAAAATGGGCTGTCGCCACCTCTGGTACCCGTGAC
ATGGCCAAGAAATGGTTCGACATTTTGAAGATCAAGAGACCAGAATACTTCATCACCGCC
AATGATGTCAAGCAAGGTAAGCCTCACCCAGAACCATACTTAAAGGGTAGAAACGGTTTG
GGTTTCCCAATTAATGAACAAGACCCATCCAAATCTAAGGTTGTTGTCTTTGAAGACGCA
CCAGCTGGTATTGCTGCTGGTAAGGCTGCTGGCTGTAAAATCGTTGGTATTGCTACCACT
TTCGATTTGGACTTCTTGAAGGAAAAGGGTTGTGACATCATTGTCAAGAACCACGAATCT
ATCAGAGTCGGTGAATACAACGCTGAAACCGATGAAGTCGAATTGATCTTTGATGACTAC
TTATACGCTAAGGATGACTTGTTGAAATGGTAA
SEQ ID NO: 27 S. cerevisiae GPP2
ATGGGATTGACTACTAAACCTCTATCTTTGAAAGTTAACGCCGCTTTGTTCGACGTCGAC
GGTACCATTATCATCTCTCAACCAGCCATTGCTGCATTCTGGAGGGATTTCGGTAAGGAC
AAACCTTATTTCGATGCTGAACACGTTATCCAAGTCTCGCATGGTTGGAGAACGTTTGAT
GCCATTGCTAAGTTCGCTCCAGACTTTGCCAATGAAGAGTATGTTAACAAATTAGAAGCT
GAAATTCCGGTCAAGTACGGTGAAAAATCCATTGAAGTCCCAGGTGCAGTTAAGCTGTGC
AACGCTTTGAACGCTCTACCAAAAGAGAAATGGGCTGTGGCAACTTCCGGTACCCGTGAT
ATGGCACAAAAATGGTTCGAGCATCTGGGAATCAGGAGACCAAAGTACTTCATTACCGCT
AATGATGTCAAACAGGGTAAGCCTCATCCAGAACCATATCTGAAGGGCAGGAATGGCTTA
GGATATCCGATCAATGAGCAAGACCCTTCCAAATCTAAGGTAGTAGTATTTGAAGACGCT
CCAGCAGGTATTGCCGCCGGAAAAGCCGCCGGTTGTAAGATCATTGGTATTGCCACTACT
TTCGACTTGGACTTCCTAAAGGAAAAAGGCTGTGACATCATTGTCAAAAACCACGAATCC
ATCAGAGTTGGCGGCTACAATGCCGAAACAGACGAAGTTGAATTCATTTTTGACGACTAC
TTATATGCTAAGGACGATCTGTTGAAATGGTAA
SEQ ID NO: 28 dmpF ORF synthetic codon optimized for Saccharomyces cerevisiae. Reference sequence: Pseudomonas sp. CF600
EMBL Accession number: X60835.1
ATGAATCAGAAACTGAAAGTAGCTATCATAGGTTCCGGTAATATCGGAAC
AGACCTGATGATTAAGGTACTGCGTAATGCAAAGTACTTAGAAATGGGCG
CGATGGTCGGTATCGATGCAGCCTCTGATGGACTGGCCAGAGCTCAAAGA
ATGGGCGTTACGACTACTTATGCAGGTGTAGAAGGGCTAATCAAGCTTCC
TGAATTTGCAGACATAGATTTCGTCTTCGATGCTACATCTGCATCAGCCC
ACGTTCAGAACGAGGCTTTATTAAGACAAGCTAAACCTGGTATTAGATTG
ATCGACCTTACCCCGGCGGCAATCGGTCCTTACTGTGTCCCCGTAGTTAA
TCTCGAGGAACATTTGGGCAAGTTGAACGTTAACATGGTTACTTGCGGTG
GCCAAGCTACTATTCCGATGGTCGCAGCTGTCTCACGTGTAGCCAAAGTC
CATTATGCTGAGATTGTTGCTTCTATTTCAAGCAAGAGTGCCGGACCTGG
AACCAGAGCCAATATAGATGAATTCACTGAGACAACCAGTAAAGCCATAG
AAGTTATTGGTGGTGCTGCAAAGGGTAAAGCTATAATTATTATGAACCCA
GCTGAACCACCATTGATTATGAGGGATACGGTGTATGTGCTTTCCGCCGC
CGCTGATCAAGCCGCTGTCGCAGCTTCTGTGGCTGAAATGGTTCAAGCGG
TTCAAGCATACGTGCCAGGCTATAGGTTAAAACAACAGGTTCAGTTTGAC
GTGATTCCCGAGTCCGCGCCACTAAACATCCCCGGTTTGGGGAGATTCAG
CGGGTTGAAAACAAGTGTGTTCCTAGAAGTAGAAGGTGCTGCTCATTATT
TGCCAGCATACGCAGGAAACTTAGATATTATGACTTCCGCAGCGTTAGCT
ACAGCCGAACGTATGGCGCAATCAATGTTGAATGCATAG
SEQ ID NO: 29 dmpF ORF
MNQKLKVAIIGSGNIGTDLMIKVLRNAKYLEMGAMVGIDAASDGLARAQR
MGVTTTYAGVEGLIKLPEFADIDFVFDATSASAHVQNEALLRQAKPGIRL
IDLTPAAIGPYCVPVVNLEEHLGKLNVNMVTCGGQATIPMVAAVSRVAKV
HYAEIVASISSKSAGPGTRANIDEFTETTSKAIEVIGGAAKGKAIIIMNP
AEPPLIMRDTVYVLSAAADQAAVAASVAEMVQAVQAYVPGYRLKQQVQFD
VIPESAPLNIPGLGRFSGLKTSVFLEVEGAAHYLPAYAGNLDIMTSAALA
TAERMAQSMLNA
SEQ ID NO: 30 primer dmpF-FW
CATTGATTGCGCCATACG
SEQ ID NO: 31 primer dmpF-RV
CCGGTAATATCGGAACAGAC
SEQ ID NO: 32 Wild type sequence of dmpF gene from Pseudomonas sp. CF600. EMBL Accession number: X60835.1
ATGAACCAGAAACTCAAAGTCGCGATCATCGGTTCGGGCAATATCGGCACCGACCTGATGATCAAGGTGCTGCGCAACGCCAAGTACCTGGAAATGGGCGCCATGGTCGGCATCGACGCCGCCTCCGACGGCCTGGCCCGCGCCCAGCGCATGGGCGTGACGACCACCTATGCCGGCGTCGAAGGGCTGATCAAGCTGCCCGAATTCGCCGACATCGATTTCGTCTTCGACGCCACCTCGGCCAGTGCCCACGTGCAGAACGAGGCGCTGCTGCGCCAGGCCAAACCTGGCATCCGCCTGATCGACCTGACCCCGGCGGCCATCGGCCCGTACTGCGTGCCGGTGGTCAATCTGGAGGAGCACCTCGGCAAGCTCAACGTCAACATGGTTACCTGCGGCGGCCAGGCGACCATCCCGATGGTCGCCGCGGTCTCCCGTGTGGCCAAGGTCCATTACGCCGAGATCGTCGCCTCGATCAGCAGCAAGTCGGCCGGACCCGGCACCCGCGCCAACATCGACGAGTTCACCGAGACCACCAGCAAAGCCATCGAAGTGATCGGTGGTGCGGCCAAGGGCAAGGCGATCATCATCATGAACCCGGCTGAGCCGCCGCTGATCATGCGCGACACCGTGTATGTGCTGTCCGCCGCCGCCGATCAGGCCGCCGTCGCGGCCTCGGTGGCGGAAATGGTTCAGGCGGTGCAGGCCTACGTGCCCGGCTATCGCCTGAAGCAGCAGGTGCAGTTCGACGTGATCCCCGAGTCCGCGCCGCTGAACATCCCCGGTCTCGGCCGGTTCAGCGGGTTGAAGACCTCGGTGTTCCTCGAAGTCGAAGGCGCCGCCCATTACCTGCCGGCCTACGCCGGCAACCTCGACATCATGACCTCCGCCGCGCTGGCTACCGCCGAGCGTATGGCGCAGTCGATGTTGAACGCCTGA
Claims (14)
- 組換え酵母細胞であって、NAD + 依存性アセチル化アセトアルデヒドデヒドロゲナーゼ(EC1.2.1.10)活性を有するタンパク質をコードする1つまたは複数の組換え核酸配列を含み、NADH依存性グリセロール合成に必要な酵素活性を欠損しているか、あるいは、その対応する野生型酵母細胞と比較してNADH依存性グリセロール合成に対する酵素活性が低下している、酵母細胞。
- 前記細胞は、配列番号2もしくは配列番号29で表されるNAD + 依存性アセチル化アセトアルデヒドデヒドロゲナーゼ、または配列番号2もしくは配列番号29の機能的ホモログをコードする1つまたは複数の異種核酸配列を含み、前記ホモログは配列番号2もしくは配列番号29との配列同一性が少なくとも90%である、請求項1に記載の細胞。
- 前記細胞は、前記デヒドロゲナーゼをコードする核酸配列であって、配列番号1、配列番号28もしくは配列番号32に記載の配列または配列番号1、配列番号28もしくは配列番号32の機能的ホモログを含む、核酸配列を含み、前記ホモログは配列番号1、配列番号28もしくは配列番号32との配列同一性が少なくとも90%である、請求項2に記載の細胞。
- 前記細胞はNAD + 依存性グリセロール3−リン酸デヒドロゲナーゼ活性を含まないか、もしくは、NAD + 依存性グリセロール3−リン酸デヒドロゲナーゼ活性がその対応する野生型細胞と比較して低下しており、および/または、前記細胞はグリセロールリン酸ホスファターゼ活性を含まないか、もしくは、グリセロールリン酸ホスファターゼ活性がその対応する野生型細胞と比較して低下している、請求項1〜3のいずれか一項に記載の細胞。
- 前記細胞のゲノムは、GPD1、GPD2、GPP1およびGPP2の群から選択される少なくとも1つの遺伝子に突然変異を含み、前記突然変異はノックアウト突然変異でもよく、前記ノックアウト突然変異は前記細胞に対応する野生型酵母遺伝子に対する、前記遺伝子の少なくとも1つの完全な欠失でもよい、請求項4に記載の細胞。
- 前記細胞はNAD + 依存性グリセロール3−リン酸デヒドロゲナーゼ(EC1.1.1.8)をコードする遺伝子を含まない、請求項4または5に記載の細胞。
- アセチルコエンザイムAシンテターゼ活性(EC6.2.1.1)を有するタンパク質をコードする1つまたは複数の核酸配列、およびNAD + 依存性アルコールデヒドロゲナーゼ活性(EC1.1.1.1)を有するタンパク質をコードする1つまたは複数の核酸配列を含む、請求項1〜6にいずれか一項に記載の細胞。
- 前記細胞はサッカロマイセス科から選択される酵母細胞である、請求項1〜7のいずれか一項に記載の細胞。
- 前記細胞は2009年7月16日にセントラルビューロー・フォー・シュメルカルチャーズに寄託された寄託番号CBS125049のサッカロマイセス・セレビシエ株の細胞である、請求項1〜8のいずれか一項に記載の細胞。
- エタノールの調製のための、請求項1〜9のいずれか一項に記載の細胞の使用。
- アセテートと、グルコース、フルクトース、スクロース、マルトース、キシロース、アラビノース、ガラクトースおよびマンノースの群から選択される発酵性糖質とからエタノールを調製することを含み、前記調製は酵母細胞を用いて嫌気条件下で行われ、前記細胞はアセチルコエンザイムAシンテターゼ活性、およびNAD + 依存性アセチル化アセトアルデヒドデヒドロゲナーゼ活性を含み、前記細胞は、NADH依存性グリセロール合成に必要な酵素活性を欠損している細胞、およびその対応する野生型酵母細胞と比較してNADH依存性グリセロール合成に対する酵素活性が低下している細胞の群から選択される細胞である、エタノールを調製する方法。
- 前記調製は発酵培地内で行われ、前記発酵培地は前記アセテートおよび前記発酵性糖質を含み、前記発酵性糖質に対する前記アセテートのモル比が0.7以下である、請求項11に記載の方法。
- 前記発酵性糖質の少なくとも一部および前記アセテートの少なくとも一部はリグノセルロース、セルロース、ヘミセルロースおよびペクチンの群から選択される多糖を加水分解することにより得られる、請求項11または12に記載の方法。
- リグノセルロース系バイオマスは加水分解され、それにより前記発酵性糖質およびアセテートを得る、請求項13に記載の方法。
Applications Claiming Priority (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP09166360A EP2277989A1 (en) | 2009-07-24 | 2009-07-24 | Fermentative glycerol-free ethanol production |
| EP09166360.9 | 2009-07-24 | ||
| PCT/NL2010/050475 WO2011010923A1 (en) | 2009-07-24 | 2010-07-23 | Fermentative glycerol-free ethanol production |
Publications (3)
| Publication Number | Publication Date |
|---|---|
| JP2013500006A JP2013500006A (ja) | 2013-01-07 |
| JP2013500006A5 JP2013500006A5 (ja) | 2013-08-22 |
| JP5836271B2 true JP5836271B2 (ja) | 2015-12-24 |
Family
ID=41460190
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| JP2012521593A Active JP5836271B2 (ja) | 2009-07-24 | 2010-07-23 | グリセロールを含まない発酵によるエタノール生成 |
Country Status (15)
| Country | Link |
|---|---|
| US (10) | US8795998B2 (ja) |
| EP (4) | EP2277989A1 (ja) |
| JP (1) | JP5836271B2 (ja) |
| CN (1) | CN102712895B (ja) |
| AU (1) | AU2010275085A1 (ja) |
| BR (1) | BR112012001642B1 (ja) |
| CA (1) | CA2768897C (ja) |
| DK (1) | DK2456851T3 (ja) |
| EA (1) | EA201270187A1 (ja) |
| ES (1) | ES2706653T3 (ja) |
| MX (1) | MX2012001062A (ja) |
| PL (1) | PL2456851T3 (ja) |
| UA (1) | UA107467C2 (ja) |
| WO (1) | WO2011010923A1 (ja) |
| ZA (1) | ZA201200703B (ja) |
Families Citing this family (87)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| EP2277989A1 (en) | 2009-07-24 | 2011-01-26 | Technische Universiteit Delft | Fermentative glycerol-free ethanol production |
| CA2798452C (en) * | 2010-05-05 | 2019-09-03 | Mascoma Corporation | Detoxification of biomass derived acetate via metabolic conversion to ethanol, acetone, isopropanol, or ethyl acetate |
| WO2011149353A1 (en) * | 2010-05-27 | 2011-12-01 | C5 Yeast Company B.V. | Yeast strains engineered to produce ethanol from acetic acid and glycerol |
| WO2011153516A2 (en) | 2010-06-03 | 2011-12-08 | Mascoma Corporation | Yeast expressing saccharolytic enzymes for consolidated bioprocessing using starch and cellulose |
| BR122019017739B1 (pt) | 2011-04-05 | 2021-06-22 | Lallemand Hungary Liquidity Management Llc | Micro-organismo recombinante compreendendo uma deleção de enzimas nativas que atuam para produzir glicerol e/ou regular a síntese de glicerol e vias metabólicas sintéticas para converter uma fonte de carboidrato a etanol |
| US9890398B2 (en) | 2011-10-24 | 2018-02-13 | Toyota Jidosha Kabushiki Kaisha | Method for producing ethanol using recombinant yeast |
| CN104039974B (zh) | 2011-11-09 | 2016-10-12 | 阿迈瑞斯公司 | 乙酰辅酶a衍生的类异戊二烯的生产 |
| CA2857498A1 (en) | 2011-11-30 | 2013-09-26 | Mascoma Corporation | Engineering an increase in ethanol production by altering cofactor specificity |
| KR20140099251A (ko) * | 2011-11-30 | 2014-08-11 | 디에스엠 아이피 어셋츠 비.브이. | 아세트산 및 글리세롤로부터 에탄올을 생성하도록 합성된 이스트 스트레인 |
| WO2014033019A1 (en) * | 2012-08-28 | 2014-03-06 | Dsm Ip Assets B.V. | Yeast strains engineered to produce ethanol from acetate |
| WO2014033018A1 (en) * | 2012-08-28 | 2014-03-06 | Dsm Ip Assets B.V. | Yeast strains engineered to produce ethanol from acetate |
| CA2881772C (en) | 2012-08-29 | 2023-03-21 | Lallemand Hungary Liquidity Management Llc | Expression of enzymes in yeast for lignocellulose derived oligomer cbp |
| EP2917343A2 (en) * | 2012-11-09 | 2015-09-16 | Lallemand Hungary Liquidity Management LLC | Method for acetate consumption during ethanolic fermentation of cellulosic feedstocks |
| AU2014219510A1 (en) | 2013-02-22 | 2015-08-27 | Technische Universiteit Delft | Recombinant micro-organism for use in method with increased product yield |
| BR112015019777B1 (pt) * | 2013-02-27 | 2022-12-06 | Toyota Jidosha Kabushiki Kaisha | Método para produzir etanol com o uso de saccharomyces cerevisiae recombinante |
| HUE045200T2 (hu) | 2013-03-14 | 2019-12-30 | Du Pont | Glicerin-3-foszfát dehidrogenáz butanol elõállítására |
| US9850512B2 (en) | 2013-03-15 | 2017-12-26 | The Research Foundation For The State University Of New York | Hydrolysis of cellulosic fines in primary clarified sludge of paper mills and the addition of a surfactant to increase the yield |
| EP2970864B1 (en) * | 2013-03-15 | 2020-10-28 | Lallemand Hungary Liquidity Management LLC | Methods for regulating nitrogen metabolism during the production of ethanol from corn by metabolically engineered yeast strains |
| WO2014180820A2 (en) * | 2013-05-08 | 2014-11-13 | Dsm Ip Assets B.V. | Gpd- yeast strains with improved osmotolerance |
| FI3033413T4 (fi) | 2013-08-15 | 2023-08-31 | Menetelmiä tuotesaannon ja tuotannon parantamiseksi mikro-organismissa glyserolikierrätyksellä | |
| AR097480A1 (es) | 2013-08-29 | 2016-03-16 | Dsm Ip Assets Bv | Células de levadura convertidoras de glicerol y ácido acético con una conversión de ácido acético mejorada |
| BR112016009416A2 (pt) | 2013-10-28 | 2017-10-03 | Danisco Us Inc | Levedura seca ativa geneticamente modificada de grande escala |
| KR102255306B1 (ko) * | 2013-11-15 | 2021-05-25 | 삼성전자주식회사 | 아세트알데히드 데히드로게나제를 포함하는 락테이트 생산능을 갖는 유전적으로 조작된 효모 세포, 그를 제조하는 방법 및 그를 사용하여 락테이트를 생산하는 방법 |
| US9951363B2 (en) | 2014-03-14 | 2018-04-24 | The Research Foundation for the State University of New York College of Environmental Science and Forestry | Enzymatic hydrolysis of old corrugated cardboard (OCC) fines from recycled linerboard mill waste rejects |
| WO2015143324A1 (en) * | 2014-03-21 | 2015-09-24 | Novozymes A/S | Processes for producing ethanol and yeast |
| DK3122876T3 (da) | 2014-03-28 | 2021-02-08 | Danisco Us Inc | Ændret værtscellebane for forbedret ethanolproduktion |
| KR20160012561A (ko) * | 2014-07-24 | 2016-02-03 | 삼성전자주식회사 | Erg5의 활성이 증가되도록 유전적으로 조작된, 내산성을 갖는 효모 세포 및 그를 이용하여 락테이트를 생산하는 방법 |
| AU2016235802B2 (en) | 2015-03-20 | 2022-03-31 | Microbiogen Pty. Ltd. | Processes for producing ethanol and ethanol producing yeast |
| ES2755680T3 (es) | 2015-05-22 | 2020-04-23 | Dsm Ip Assets Bv | Célula de levadura que consume acetato |
| US11136600B2 (en) | 2015-10-06 | 2021-10-05 | Dsm Ip Assets B.V. | Eukaryotic cell with increased production of fermentation product |
| US20190338256A1 (en) * | 2016-04-28 | 2019-11-07 | Danisco Us Inc. | Redox balancing in yeast |
| US10689670B2 (en) | 2016-06-14 | 2020-06-23 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| JP6447583B2 (ja) * | 2016-06-24 | 2019-01-09 | トヨタ自動車株式会社 | 組換え酵母、及びそれを用いたエタノールの製造方法 |
| WO2018089333A1 (en) | 2016-11-09 | 2018-05-17 | Danisco Us Inc. | Yeast with improved alcohol production |
| CN110366594B (zh) | 2016-12-16 | 2023-12-15 | 丹尼斯科美国公司 | 双功能磷酸转酮酶-磷酸转乙酰酶融合多肽 |
| BR112019011093A2 (pt) * | 2016-12-20 | 2019-10-22 | Dsm Ip Assets Bv | cepas de levedura aperfeiçoadas para a produção de etanol |
| EP3571218A1 (en) | 2017-01-18 | 2019-11-27 | Danisco US Inc. | Modified yeast cells that overexpress a dna polymerase subunit |
| WO2018176021A1 (en) | 2017-03-24 | 2018-09-27 | Danisco Us Inc | Reduction of acetate and glycerol in modified yeast having an exogenous ethanol-producing pathway |
| WO2018194892A1 (en) | 2017-04-17 | 2018-10-25 | Danisco Us Inc | Selected acetyl-coa synthase enzymes for reduction of acetate production in yeast |
| US20200131591A1 (en) | 2017-06-06 | 2020-04-30 | Danisco Us Inc. | Yeast with improved alcohol production |
| CA3064143A1 (en) | 2017-06-13 | 2018-12-20 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| JP6879111B2 (ja) | 2017-08-02 | 2021-06-02 | トヨタ自動車株式会社 | 組換え酵母及びこれを用いたエタノールの製造方法 |
| UY37835A (es) | 2017-08-08 | 2019-03-29 | Danisco Us Inc | Producción incrementada de etanol por levaduras que albergan alelos del activador de transcripción constitutivo mal |
| CA3073387A1 (en) | 2017-08-29 | 2019-03-07 | Danisco Us Inc | Modified yeast comprising glucose-specific, atp-mediated transporters |
| CA3073329A1 (en) | 2017-09-26 | 2019-04-04 | Dsm Ip Assets B.V. | Improved process for ethanol production |
| US11414683B2 (en) | 2017-09-26 | 2022-08-16 | Dsm Ip Assets B.V. | Acetic acid consuming strain |
| US11274310B2 (en) | 2017-09-29 | 2022-03-15 | Dsm Ip Assets B.V. | Yeast cells for glycerol free ethanol production |
| EP3700920A1 (en) | 2017-10-24 | 2020-09-02 | Danisco US Inc. | Yeast with improved alcohol production |
| WO2019110492A1 (en) | 2017-12-08 | 2019-06-13 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| WO2019110491A1 (en) | 2017-12-08 | 2019-06-13 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| AU2019222480A1 (en) * | 2018-02-15 | 2020-10-08 | Microbiogen Pty. Ltd. | Improved yeast for ethanol production |
| US20210040474A1 (en) * | 2018-03-06 | 2021-02-11 | Danisco Us Inc. | Yeast with improved alcohol production under high dissolved solids conditions |
| EP3762499B1 (en) | 2018-03-06 | 2023-12-20 | Danisco Us Inc | Reduction in acetate production by yeast over-expressing pab1 |
| WO2019173217A1 (en) | 2018-03-06 | 2019-09-12 | Danisco Us Inc | Compositions and methods for increasing ethanol production by yeast using gcy1 and dak1 |
| WO2019195382A1 (en) | 2018-04-06 | 2019-10-10 | Danisco Us Inc | Increased alcohol production from yeast producing an increased amount of active hac1 protein |
| UY38179A (es) | 2018-04-12 | 2019-11-29 | Danisco Us Inc | Levadura que sobreexpresa proteína fosfatasas asociadas con la vía hog |
| CN112513261A (zh) | 2018-05-25 | 2021-03-16 | 丹尼斯科美国公司 | 富马酸还原酶过表达导致酵母中增加的发酵速率 |
| EP3802784A1 (en) | 2018-05-30 | 2021-04-14 | Danisco US Inc. | Over-expression of transcriptional activator/repressor gis1 in yeast for increased ethanol production |
| EP3830282A1 (en) | 2018-07-27 | 2021-06-09 | Danisco US Inc. | Increased alcohol production from yeast producing an increased amount of active crz1 protein |
| US20210395756A1 (en) | 2018-09-28 | 2021-12-23 | Danisco Us Inc | Over expression of ribonucleotide reductase inhibitor in yeast for increased ethanol production |
| US20210332091A1 (en) | 2018-09-28 | 2021-10-28 | Danisco Us Inc | Over-expression of gds1 in yeast for increased ethanol and decreased acetate production |
| WO2020068907A1 (en) | 2018-09-28 | 2020-04-02 | Danisco Us Inc | Selected phosphotransacetylase genes for increased ethanol production in engineered yeast |
| WO2020146357A1 (en) | 2019-01-08 | 2020-07-16 | Danisco Us Inc | Hybrid yeast with increased ethanol production |
| CN113795503A (zh) | 2019-03-14 | 2021-12-14 | 丹尼斯科美国公司 | 细胞色素b2在酵母中过表达用于增加乙醇生产 |
| WO2020186254A1 (en) | 2019-03-14 | 2020-09-17 | Danisco Us Inc | Over-expression of fumarate-succinate transporter in yeast for increased ethanol and reduced acetate production |
| CA3144631A1 (en) | 2019-06-24 | 2020-12-30 | Danisco Us Inc | Disruption of cdc42 effectors in yeast for increased alcohol and lysine production |
| WO2020263720A1 (en) | 2019-06-24 | 2020-12-30 | Danisco Us Inc | Modified yeast and method for increasing lysine content in fermentation co-products |
| BR112021026542A2 (pt) | 2019-06-28 | 2022-05-17 | Danisco Us Inc | Células de levedura modificadas que superexpressam as proteínas endógenas selecionadas |
| BR112022001828A2 (pt) | 2019-08-01 | 2022-06-28 | Danisco Us Inc | Superexpressão de pho13 para produção de etanol aumentada por levedura |
| BR112022001859A2 (pt) | 2019-08-01 | 2022-06-28 | Danisco Us Inc | Superexpressão de adh5p para produção de etanol aumentada por levedura |
| WO2021041141A1 (en) | 2019-08-29 | 2021-03-04 | Danisco Us Inc | Expression of beta-glucosidase in yeast for improved ethanol production |
| US20220380813A1 (en) | 2019-11-08 | 2022-12-01 | Dsm Ip Assets B.V. | Process for producing ethanol |
| MX2022006340A (es) | 2019-11-26 | 2022-10-07 | Danisco Us Inc | Reducción de la producción de acetato mediante una levadura que sobreexpresa polipéptidos mig. |
| WO2021150911A1 (en) | 2020-01-23 | 2021-07-29 | Danisco Us Inc | Increased ethanol production by overexpression of jid1 in yeast |
| EP4225929A1 (en) | 2020-10-05 | 2023-08-16 | DSM IP Assets B.V. | Saccharomyces yeast cell and fermentation process using such |
| AR125830A1 (es) | 2021-05-10 | 2023-08-16 | Danisco Us Inc | Producción de etanol aumentada mediante sobreexpresión de kgd2 en levaduras |
| WO2023285281A1 (en) | 2021-07-12 | 2023-01-19 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| EP4370651A1 (en) | 2021-07-12 | 2024-05-22 | Danisco Us Inc | Recombinant yeast cell |
| EP4370691A1 (en) | 2021-07-12 | 2024-05-22 | Danisco US Inc. | Recombinant yeast cell |
| WO2023285294A1 (en) | 2021-07-12 | 2023-01-19 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| WO2023285279A1 (en) | 2021-07-12 | 2023-01-19 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| WO2023285280A1 (en) | 2021-07-12 | 2023-01-19 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| WO2023076323A1 (en) | 2021-10-26 | 2023-05-04 | Danisco Us Inc. | Reduction in acetate produced by yeast with reduced expression of rsf2 or tda9 |
| WO2023079048A1 (en) | 2021-11-04 | 2023-05-11 | Dsm Ip Assets B.V. | Process for the production of ethanol and recombinant yeast cell |
| WO2023079049A1 (en) | 2021-11-04 | 2023-05-11 | Dsm Ip Assets B.V. | Variant polypeptide and recombinant yeast cell |
| WO2023079050A1 (en) | 2021-11-04 | 2023-05-11 | Dsm Ip Assets B.V. | Recombinant yeast cell |
| WO2023208762A2 (en) | 2022-04-25 | 2023-11-02 | Dsm Ip Assets B.V. | Mutant yeast cell and process for the production of ethanol |
Family Cites Families (26)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US7041814B1 (en) | 1998-02-18 | 2006-05-09 | Genome Therapeutics Corporation | Nucleic acid and amino acid sequences relating to Enterobacter cloacae for diagnostics and therapeutics |
| US6562958B1 (en) | 1998-06-09 | 2003-05-13 | Genome Therapeutics Corporation | Nucleic acid and amino acid sequences relating to Acinetobacter baumannii for diagnostics and therapeutics |
| WO2000003021A2 (en) * | 1998-07-10 | 2000-01-20 | Jens Nielsen | Metabolically engineered microbial cell comprising a modified redox activity |
| US6476209B1 (en) | 2000-11-28 | 2002-11-05 | Genesis Research & Development Corporation Ltd. | Polynucleotides, materials incorporating them, and methods for using them |
| US7026463B2 (en) | 1999-08-13 | 2006-04-11 | Matthew Glenn | Polynucleotides and polypeptides, materials incorporating them and methods for using them |
| FR2807446B1 (fr) | 2000-04-11 | 2005-05-06 | Agronomique Inst Nat Rech | Genomes de lactococcus lactis, polypeptides et utilisations |
| EP2311855A3 (en) | 2000-10-06 | 2011-05-11 | Novozymes Inc. | Bacillus licheniformis YvnA negative strain |
| US20070020625A1 (en) | 2001-02-07 | 2007-01-25 | Eric Duchaud | Sequence of the photorhabdus luminescens strain tt01 genome and uses |
| US7253001B2 (en) | 2002-03-19 | 2007-08-07 | Forskarpatent I Syd Ab | Metabolic engineering for improved xylose utilisation of Saccharomyces cerevisiae |
| SE0200857D0 (sv) * | 2002-03-19 | 2002-03-19 | Forskarpatent I Syd Ab | Metabolic engineering for improved xylose utilisation of saccharomyces cerevisiae |
| JP4372408B2 (ja) | 2002-11-14 | 2009-11-25 | 三菱レイヨン株式会社 | ロドコッカス(Rhodococcus)属細菌組換え体、及びそれを用いた光学活性体の製造方法 |
| GB0227435D0 (en) | 2002-11-25 | 2002-12-31 | Univ Denmark Tech Dtu | Metabolically engineered micro-organisms having reduced production of undesired metobolic products |
| WO2004085627A1 (en) | 2003-03-26 | 2004-10-07 | Forskarpatent I Syd Ab | New saccharomyces cerevisiae strains utilizing xylose |
| JP5796925B2 (ja) | 2004-07-16 | 2015-10-21 | ディーエスエム アイピー アセッツ ビー.ブイ. | キシロース発酵性真核細胞の代謝工学 |
| PL2351772T3 (pl) | 2005-02-18 | 2017-01-31 | Glaxosmithkline Biologicals Sa | Białka i kwasy nukleinowe z bakterii Escherichia coli związanej z zapaleniem opon mózgowo-rdzeniowych/posocznicą |
| US20090053797A1 (en) | 2005-08-19 | 2009-02-26 | Yoichiro Shiba | Genetically modified host cells and use of same for producing isoprenoid compounds |
| ZA200803755B (en) | 2005-10-26 | 2009-12-30 | Du Pont | Fermentive production of four carbon alcohols |
| US20080293101A1 (en) | 2006-07-27 | 2008-11-27 | Peters Matthew W | Engineered microorganisms for increasing product yield in biotransformations, related methods and systems |
| EP2097528A2 (en) | 2006-10-31 | 2009-09-09 | DSMIP Assets B.V. | Butanol production in a eukaryotic cell |
| CN101939433B (zh) | 2007-07-23 | 2015-05-20 | 帝斯曼知识产权资产管理有限公司 | 酵母中生产乙酰-CoA的酶 |
| EP2060632A1 (en) | 2007-10-29 | 2009-05-20 | Technische Universität Berlin | Method of modifying a yeast cell for the production of ethanol |
| WO2009078973A2 (en) | 2007-12-13 | 2009-06-25 | Glycos Biotechnologies, Incorporated | Microbial conversion of oils and fatty acids to high-value chemicals |
| DE102008004253B4 (de) * | 2008-01-14 | 2011-07-28 | Butalco Gmbh | Gesteigerte Produktion von Acetyl-Coenzym A |
| BRPI0913667A2 (pt) | 2008-02-20 | 2016-08-02 | Nagarjuna Fertilizers And Chemicals Ltd | microorganismos geneticamente transformados com aumento simultâneo de potencial de redução e atividades de enzima redutiva para fermentação de biomassa |
| SI2262901T1 (sl) * | 2008-03-05 | 2019-02-28 | Genomatica, Inc. | Primarni alkohol producirajoči organizmi |
| EP2277989A1 (en) * | 2009-07-24 | 2011-01-26 | Technische Universiteit Delft | Fermentative glycerol-free ethanol production |
-
2009
- 2009-07-24 EP EP09166360A patent/EP2277989A1/en not_active Withdrawn
-
2010
- 2010-07-23 MX MX2012001062A patent/MX2012001062A/es active IP Right Grant
- 2010-07-23 ES ES10740412T patent/ES2706653T3/es active Active
- 2010-07-23 CN CN201080042861.6A patent/CN102712895B/zh active Active
- 2010-07-23 EP EP10740412.1A patent/EP2456851B1/en active Active
- 2010-07-23 EP EP20217514.7A patent/EP3828261A1/en active Pending
- 2010-07-23 EP EP18179439.7A patent/EP3476931A1/en active Pending
- 2010-07-23 PL PL10740412T patent/PL2456851T3/pl unknown
- 2010-07-23 AU AU2010275085A patent/AU2010275085A1/en not_active Abandoned
- 2010-07-23 BR BR112012001642A patent/BR112012001642B1/pt active IP Right Grant
- 2010-07-23 JP JP2012521593A patent/JP5836271B2/ja active Active
- 2010-07-23 WO PCT/NL2010/050475 patent/WO2011010923A1/en active Application Filing
- 2010-07-23 DK DK10740412.1T patent/DK2456851T3/en active
- 2010-07-23 US US13/061,695 patent/US8795998B2/en active Active
- 2010-07-23 EA EA201270187A patent/EA201270187A1/ru unknown
- 2010-07-23 UA UAA201201698A patent/UA107467C2/uk unknown
- 2010-07-23 CA CA2768897A patent/CA2768897C/en active Active
-
2012
- 2012-01-27 ZA ZA2012/00703A patent/ZA201200703B/en unknown
-
2014
- 2014-02-25 US US14/189,989 patent/US9528117B2/en active Active
-
2016
- 2016-05-13 US US15/154,199 patent/US20160312246A1/en not_active Abandoned
-
2018
- 2018-05-03 US US15/970,641 patent/US10738317B2/en active Active
-
2019
- 2019-05-10 US US16/409,197 patent/US10533181B2/en active Active
-
2020
- 2020-03-27 US US16/833,342 patent/US11214810B2/en active Active
- 2020-06-04 US US16/893,065 patent/US10883110B2/en active Active
-
2021
- 2021-02-15 US US17/176,040 patent/US11174489B2/en active Active
- 2021-02-15 US US17/176,073 patent/US11326175B2/en active Active
- 2021-11-19 US US17/531,218 patent/US20220135989A1/en active Pending
Also Published As
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11174489B2 (en) | Fermentative glycerol-free ethanol production | |
| EP2663645B1 (en) | Yeast strains engineered to produce ethanol from glycerol | |
| JP5608999B2 (ja) | キシロースを利用して有用物質を生産する方法 | |
| US20210310013A1 (en) | Acetate consuming yeast cell | |
| WO2018172328A1 (en) | Improved glycerol free ethanol production | |
| KR20170041768A (ko) | 아세토인을 제조하는 방법 | |
| EP3359655B1 (en) | Eukaryotic cell with increased production of fermentation product | |
| US8748140B2 (en) | Xylose-fermenting microorganism | |
| CA3074001A1 (en) | Acetic acid consuming strain | |
| SI24955A (sl) | Modificirana 6-fosfofrukto-1-kinaza, ki rekombinantnim celicam kvasovke Saccharomyces cerevisiae omogoča fermentativno uporabo pentoznih sladkorjev |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A523 Effective date: 20130703 |
|
| A621 | Written request for application examination |
Free format text: JAPANESE INTERMEDIATE CODE: A621 Effective date: 20130703 |
|
| A131 | Notification of reasons for refusal |
Free format text: JAPANESE INTERMEDIATE CODE: A131 Effective date: 20141111 |
|
| A601 | Written request for extension of time |
Free format text: JAPANESE INTERMEDIATE CODE: A601 Effective date: 20150210 |
|
| RD03 | Notification of appointment of power of attorney |
Free format text: JAPANESE INTERMEDIATE CODE: A7423 Effective date: 20150210 |
|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A523 Effective date: 20150216 |
|
| A711 | Notification of change in applicant |
Free format text: JAPANESE INTERMEDIATE CODE: A711 Effective date: 20150306 |
|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A821 Effective date: 20150306 |
|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A523 Effective date: 20150511 |
|
| TRDD | Decision of grant or rejection written | ||
| A01 | Written decision to grant a patent or to grant a registration (utility model) |
Free format text: JAPANESE INTERMEDIATE CODE: A01 Effective date: 20151006 |
|
| A61 | First payment of annual fees (during grant procedure) |
Free format text: JAPANESE INTERMEDIATE CODE: A61 Effective date: 20151102 |
|
| R150 | Certificate of patent or registration of utility model |
Ref document number: 5836271 Country of ref document: JP Free format text: JAPANESE INTERMEDIATE CODE: R150 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |