WO2018199275A1 - 卵巣腫瘍の検出のためのキット、デバイス及び方法 - Google Patents
卵巣腫瘍の検出のためのキット、デバイス及び方法 Download PDFInfo
- Publication number
- WO2018199275A1 WO2018199275A1 PCT/JP2018/017125 JP2018017125W WO2018199275A1 WO 2018199275 A1 WO2018199275 A1 WO 2018199275A1 JP 2018017125 W JP2018017125 W JP 2018017125W WO 2018199275 A1 WO2018199275 A1 WO 2018199275A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- mir
- hsa
- gene
- seq
- polynucleotide
- Prior art date
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- B—PERFORMING OPERATIONS; TRANSPORTING
- B01—PHYSICAL OR CHEMICAL PROCESSES OR APPARATUS IN GENERAL
- B01L—CHEMICAL OR PHYSICAL LABORATORY APPARATUS FOR GENERAL USE
- B01L7/00—Heating or cooling apparatus; Heat insulating devices
- B01L7/52—Heating or cooling apparatus; Heat insulating devices with provision for submitting samples to a predetermined sequence of different temperatures, e.g. for treating nucleic acid samples
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12M—APPARATUS FOR ENZYMOLOGY OR MICROBIOLOGY; APPARATUS FOR CULTURING MICROORGANISMS FOR PRODUCING BIOMASS, FOR GROWING CELLS OR FOR OBTAINING FERMENTATION OR METABOLIC PRODUCTS, i.e. BIOREACTORS OR FERMENTERS
- C12M1/00—Apparatus for enzymology or microbiology
- C12M1/34—Measuring or testing with condition measuring or sensing means, e.g. colony counters
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6813—Hybridisation assays
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6869—Methods for sequencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/112—Disease subtyping, staging or classification
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/158—Expression markers
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/178—Oligonucleotides characterized by their use miRNA, siRNA or ncRNA
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2800/00—Detection or diagnosis of diseases
- G01N2800/70—Mechanisms involved in disease identification
- G01N2800/7023—(Hyper)proliferation
- G01N2800/7028—Cancer
Definitions
- the present invention relates to a kit or device for detecting an ovarian tumor comprising a specific miRNA, or a nucleic acid capable of specifically binding to its complementary strand, used for examining whether or not the subject has an ovarian tumor. And a method for detecting an ovarian tumor, comprising measuring the expression level of the miRNA using the nucleic acid.
- the ovary is a female reproductive organ that produces eggs, one on each side of the uterus.
- the oviduct is a tube until the egg released from the ovary moves to the uterus, where the fertilized egg is implanted and grows into a fetus.
- the ovary also has the function of secreting female hormones, estrogen and progesterone.
- the ovary is considered to be an organ in which tumors are likely to occur, and ovarian tumors are roughly classified into superficial epithelial / stromal tumors, sex cord / stromal tumors, and germ cell tumors based on their origin.
- Ovarian tumors are classified into benign, borderline malignant, and malignant according to the grade of malignancy, and borderline malignant and malignant ovarian tumors are called ovarian cancer.
- the incidence of ovarian cancer in Japan as disclosed by the National Cancer Center Information Center for Cancer Control in Japan was 9,384 in 2012, or 1 in 87 Japanese women, and ovarian cancer.
- the number of deaths in 2014 reached 4,840 in 2014.
- the degree of progression of ovarian cancer and fallopian tube cancer is determined by whether the tumor is unilateral or bilateral, intraperitoneal dissemination, lymph node metastasis, distant metastasis, and size, etc. Stages IA, IB, IC (IC1, IC2, IC3), IIA, IIB, IIIA1, IIIA2, IIIB, IIIC, IVA, IVB.
- the five-year relative survival rate for ovarian cancer depends largely on the degree of cancer progression, 92% if the cancer is localized, 73% if the cancer is in the surrounding area, and distant metastatic cancer. If yes, it is reported as 29%. Therefore, since early detection of ovarian cancer leads to an improvement in survival rate, provision of a means enabling early detection is strongly desired.
- the treatment of ovarian cancer is basically a surgical treatment, but drug therapy is used in combination with the degree of progression, metastasis, general condition, and classification of ovarian cancer.
- drug therapy is used in combination with the degree of progression, metastasis, general condition, and classification of ovarian cancer.
- stage I early stage ovarian cancer it may be possible to remove only the ovaries and fallopian tubes on the side where the cancer is found, rather than removing the uterus, ovaries, and fallopian tubes. The possibility of subsequent childbirth can be maintained.
- Patent Documents 1 to 4 and Non-Patent Documents 1 to 3 ovarian tumors are detected based on the expression level of microRNA (miRNA) in biological samples including blood. There is a report.
- miRNA microRNA
- Patent Document 1 discloses miR-191, miR-24, miR-320, miR-328, miR-625-3p, miR-483-5p, miR-92a, miR-1290 and the like in plasma. Methods for detecting ovarian cancer and predicting prognosis are shown.
- Patent Document 2 discloses miR-135a-3 in blood as a biomarker for gynecological cancer.
- Patent Document 3 discloses miR-125a-3p, miR-211-3 and the like as biomarkers for ovarian cancer using human sample tissues.
- Patent Document 4 describes endometriosis, endometriosis-derived ovarian cancer and serous ovary using miR-191-5p, miR-652-5p, miR-744-5p, miR-1246, etc. in plasma. Shows how to determine cancer.
- Non-Patent Document 1 includes miR-92a, miR-128, miR-191-5p, miR-296-5p, miR-320a, miR-625-3p, etc. in the blood as circulating extracellular miRNAs related to ovarian cancer. It is shown.
- Non-patent document 2 shows that the expression levels of miR-22, miR-45, etc. are different as a result of analyzing miRNA derived from serum samples of ovarian cancer and benign disease patients and healthy persons using a next-generation sequencer. Yes.
- Non-Patent Document 3 includes miR-23a / b, miR-92, miR-125a, miR-296-5p, miR-320, miR-422a, miR-663a, etc. in the ER of ovarian cancer patients Has been shown to do.
- Non-patent document 4 reports that blood marker CA-125 has a sensitivity of 77.4% for detecting ovarian cancer and a specificity of 70.8%.
- An object of the present invention is to find a novel ovarian tumor marker and to provide a method capable of effectively detecting an ovarian tumor using a nucleic acid for detecting the marker.
- CA-125 is a protein in the blood and is said to increase in ovarian tumor patients, but it is more likely to increase for reasons different from ovarian tumors, while CA- Because some patients do not increase 125 levels, the CA-125 test is not an effective screening test. As described above, according to Non-Patent Document 4, it is reported that the sensitivity of CA-125 to detect ovarian cancer was 77.4% and the specificity was 70.8%.
- Patent Document 1 discloses that ovary using miRNA such as miR-191, miR-24, miR-320, miR-328, miR-625-3p, miR-483-5p, miR-92a, miR-1290 in plasma. Although a method for detecting cancer and a method for predicting prognosis have been shown, these miRNAs are also expressed in the control group and are not suitable for methods for detecting the presence or absence of ovarian cancer. In addition, the number of specimens implemented is only a few tens, and the marker reliability is low.
- miRNA such as miR-191, miR-24, miR-320, miR-328, miR-625-3p, miR-483-5p, miR-92a, miR-1290 in plasma.
- miRNA such as miR-135a-3 in blood is shown as a biomarker for gynecological cancer, but only miRNA expression difference related to endometrial cancer is shown, and ovarian cancer is discriminated. There is no description of detection performance such as specific accuracy, sensitivity, specificity, etc., and industrial practicality is poor.
- miRNA such as miR-125a-3p and miR-211-3 is shown as a biomarker for ovarian cancer using a human sample tissue. This process is not preferable as an inspection method because the physical burden on the patient is heavy. In addition, there is no description about detection performance such as specific accuracy, sensitivity, specificity, etc. for discriminating ovarian cancer, and industrial practicality is poor.
- Patent Document 4 describes endometriosis, endometriosis-derived ovarian cancer and serous fluid using miRNA such as miR-191-5p, miR-652-5p, miR-744-5p, miR-1246 in plasma. Shows how to detect ovarian cancer, but it does not show how ovarian cancer is differentiated from other cancers, so the risk of misdetecting other cancers as ovarian cancer There is. In addition, the number of specimens implemented is only a few tens, and the marker reliability is low.
- Non-Patent Document 1 includes miR-92a, miR-128, miR-191-5p, miR-296-5p, miR-320a, miR-625-3p, etc. in the blood as circulating extracellular miRNAs related to ovarian cancer. It is shown. However, the detection sensitivity of ovarian epithelial cancer by the combination of miR-205 and let-7f is as low as 62.4%, and there is a concern that ovarian cancer may be overlooked. In addition, the performance of distinguishing ovarian cancer from other miRNAs and healthy people is partially shown, but other cancer patients are not included in the study, so other cancers are misidentified as ovarian cancer. There is a risk of separation, leading to unnecessary additional examinations and delaying the detection of the original cancer, which is undesirable.
- Non-patent document 2 shows that miRNAs derived from serum samples of ovarian cancer, benign disease patients, and healthy persons are analyzed by next-generation sequencers, and that there is a difference in the expression levels of miRNAs such as miR-22 and miR-45.
- detection performance such as specific accuracy, sensitivity, specificity, etc. for discriminating ovarian cancer, and it is poor in industrial practicality.
- Non-Patent Document 3 discloses that miRNAs such as miR-23a / b, miR-92, miR-125a, miR-296-5p, miR-320, miR-422a, miR-663a are present in the body fluid endoplasmic reticulum of ovarian cancer patients.
- miRNAs such as miR-23a / b, miR-92, miR-125a, miR-296-5p, miR-320, miR-422a, miR-663a are present in the body fluid endoplasmic reticulum of ovarian cancer patients.
- detection performance such as specific accuracy, sensitivity, specificity, etc. for discriminating ovarian cancer, and it is poor in industrial practicality.
- the performance of existing tumor markers is low, and the performance and detection methods are not specifically shown for markers at the research stage.
- it is not preferable to collect ovarian tissue for measuring a tumor marker because of its high invasiveness to patients. Therefore, there is a need for a highly accurate ovarian cancer marker that can be detected from blood that can be collected in a minimally invasive manner and that can correctly distinguish ovarian tumor patients from ovarian tumor patients and healthy individuals from healthy individuals.
- microRNA microRNA
- a microRNA (miRNA) gene marker that can be used as a marker for detecting ovarian tumors from blood that can be collected in a minimally invasive manner, and a nucleic acid for detecting this marker.
- a probe that can specifically bind to the marker and a primer that amplifies the marker
- ovarian tumors can be detected significantly, preferably specifically As a result, the present invention has been completed.
- the present invention has the following features.
- miR-4675 is hsa-miR-4675
- miR-4783-3p is hsa-miR-4783-3p
- miR-1228-5p is hsa-miR-1228-5p
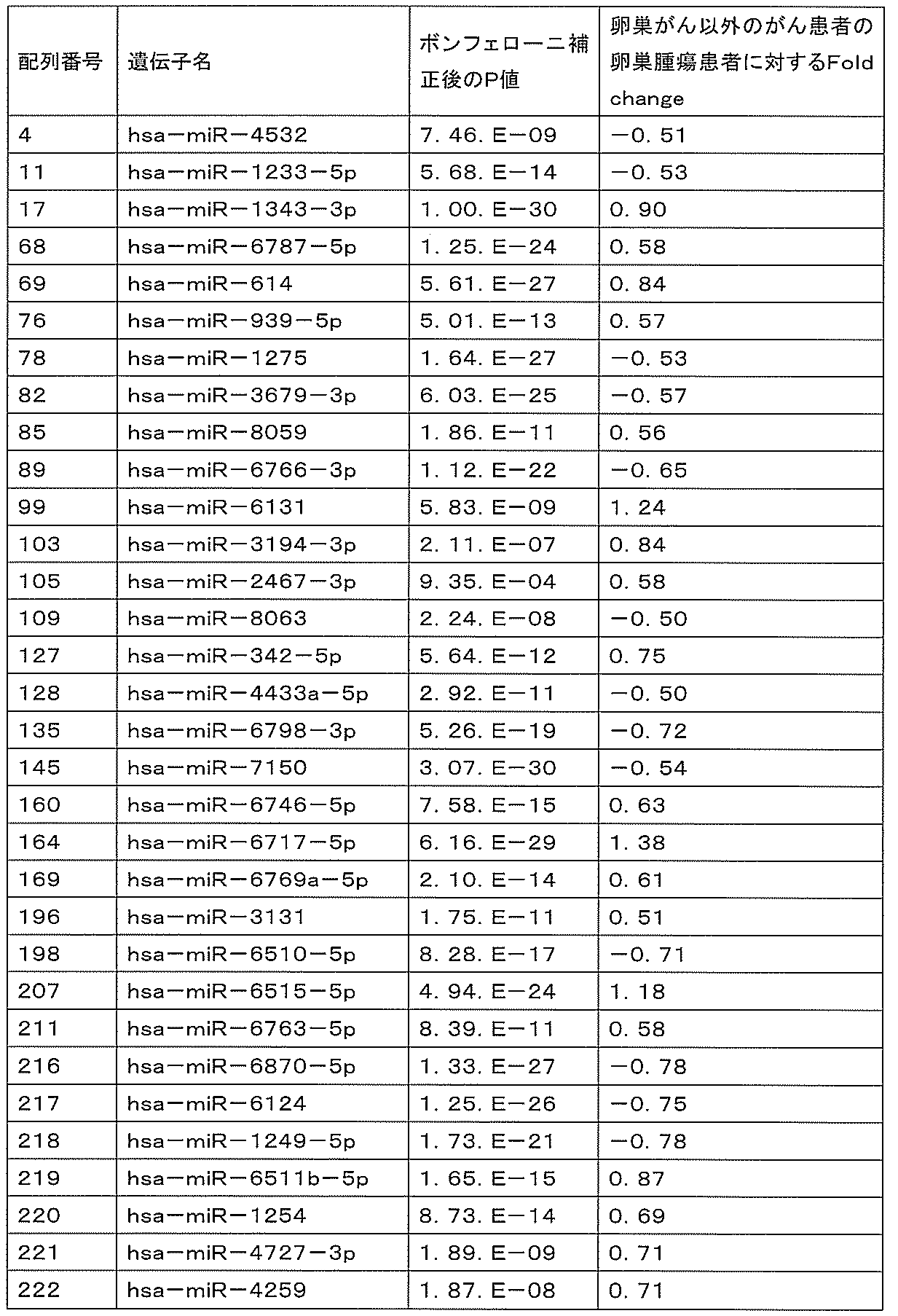
- miR-4532 Is hsa-miR-4532
- miR-6802-5p is hsa-miR-6802-5p
- miR-6784-5p is hsa-miR-6784-5p
- miR-3940-5p is hsa-miR -3940-5p
- miR-1307-3p is hsa-miR-1307-3p
- miR-8073 is hsa-miR-8073
- miR-3184-5p is hsa-miR-3184-5p
- MiR-1233-5p is hsa-miR-1233-5p
- miR-608 Is hsa-miR-6088
- MiR-675-5p is hsa-miR-675-5p
- miR-711 is hsa-miR-711
- miR-6875-5p is hsa-miR-6875-5p
- miR-3160 -5p is hsa-miR-3160-5p
- miR-1908-5p is hs MiR-1908-5p
- miR-6726-5p is hsa-miR-6726-5p
- miR-1913 is hsa-miR-1913
- miR-8071 is hsa-miR-8071
- miR -3648 is hsa-miR-3648
- miR-4732-5p is hsa-miR-4732-5p
- miR-4787-5p is hsa-miR-4787-5p
- miR-3717 is hsa-miR -3917
- miR-619-5p is
- miR-6076 is hsa-miR-6076
- miR-6515-5p is hsa-miR-6515-5p
- miR-6820-5p is hsa-miR-6820-5p
- MiR-4634 is hsa-miR-4634
- miR-187-5p is hsa-miR-187-5p
- miR-6673-5p is hsa-miR-6673-5p
- miR-1908-3p Is hsa-miR-1908-3p
- miR-1181 is hsa-miR- 1181
- miR-6878-5p is hsa-miR-6878-5p
- miR-5010-5p is hsa-miR-5010-5p
- miR-6870-5p is hsa-miR-6870-5p.
- MiR-6124 is hsa-miR-6124, miR-1249-5p is hsa-miR-1249-5p, miR-6511b-5p is hsa-miR-6511b-5p, and miR-1254 is hsa-miR-1254, miR-4727-3p is hsa-miR-4727-3p, miR-4259 is hsa-miR-4259, miR-4771 is hsa-miR-4771, miR- 3622a-5p is hsa-miR-3622a-5p, and miR- 480 is hsa-miR-4480, miR-4740-5p is hsa-miR-4740-5p, miR-6777-5p is hsa-miR-6777-5p, and miR-6794-5p is hsa- miR-6794-5p, miR-4687-3p is hsa-mi
- polynucleotide according to any one of the following (a) to (e): (A) a polynucleotide comprising the nucleotide sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268 or a nucleotide sequence wherein u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more consecutive sequences Its fragments, including (B) a polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268, (C) a polynucleotide comprising a nucleotide sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268, or a nucleotide sequence complementary to the nucleotide sequence in which u is t in the nucleotide sequence, variants thereof and derivatives thereof Or fragments thereof containing 15 or more consecutive bases, (D) a polynucle
- the kit is another ovarian tumor marker, miR-320a, miR-663a, miR-328-5p, miR-128-2-5p, miR-125a-3p, miR-191-5p, miR -92b-5p, miR-296-5p, miR-1246, miR-92a-2-5p, miR-128-1-5p, miR-1290, miR-211-3p, miR-744-5p, miR-135a -3p, miR-451a, miR-625-3p, miR-92a-3p, miR-422a, miR-483-5p, miR-652-5p, miR-24-3p, miR-23b-3p, miR-23a At least one polynucleotide selected from -3p, miR-92b-3p, miR-22-3p, or the poly Further comprising a complementary strand capable of specifically binding a nucleic acid of Kureochido kit according to any one of (1) to (3).
- miR-320a is hsa-miR-320a
- miR-663a is hsa-miR-663a
- miR-328-5p is hsa-miR-328-5p
- miR-128-2-5p Is hsa-miR-128-2-5p
- miR-125a-3p is hsa-miR-125a-3p
- miR-191-5p is hsa-miR-191-5p
- miR-92b-5p Is hsa-miR-92b-5p
- miR-296-5p is hsa-miR-296-5p
- miR-1246 is hsa-miR-1246
- miR-92a-2-5p is hsa-miR -92a-2-5p
- miR-128-1-5p is hsa-miR-128-1-5p
- miR-1290 is sa-miR-1290
- polynucleotide according to any one of the following (f) to (j): (F) a polynucleotide comprising the base sequence represented by any one of SEQ ID NOS: 248 to 250, 252 to 267, and 269 to 275, or a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or Its fragments containing 15 or more consecutive bases, (G) a polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, (H) a polynucleotide comprising a nucleotide sequence represented by any one of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, or a nucleotide sequence complementary to the nucleotide sequence in which u is t in the nucleotide sequence, and variations thereof Body, derivatives thereof, or fragments thereof comprising the
- the kit is capable of specifically binding at least two polynucleotides selected from all the ovarian tumor markers according to (1) or (2), or each of the complementary strands of the polynucleotides.
- the kit according to any one of (1) to (6), comprising two nucleic acids.
- miR-4675 is hsa-miR-4675
- miR-4783-3p is hsa-miR-4783-3p
- miR-1228-5p is hsa-miR-1228-5p
- miR-4532 Is hsa-miR-4532
- miR-6802-5p is hsa-miR-6802-5p
- miR-6784-5p is hsa-miR-6784-5p
- miR-3940-5p is hsa-miR -3940-5p
- miR-1307-3p is hsa-miR-1307-3p
- miR-8073 is hsa-miR-8073
- miR-3184-5p is hsa-miR-3184-5p
- MiR-1233-5p is hsa-miR-1233-5p
- miR-608 Is hsa-miR-6088
- MiR-675-5p is hsa-miR-675-5p
- miR-711 is hsa-miR-711
- miR-6875-5p is hsa-miR-6875-5p
- miR-3160 -5p is hsa-miR-3160-5p
- miR-1908-5p is hs MiR-1908-5p
- miR-6726-5p is hsa-miR-6726-5p
- miR-1913 is hsa-miR-1913
- miR-8071 is hsa-miR-8071
- miR -3648 is hsa-miR-3648
- miR-4732-5p is hsa-miR-4732-5p
- miR-4787-5p is hsa-miR-4787-5p
- miR-3717 is hsa-miR -3917
- miR-619-5p is
- miR-6076 is hsa-miR-6076
- miR-6515-5p is hsa-miR-6515-5p
- miR-6820-5p is hsa-miR-6820-5p
- MiR-4634 is hsa-miR-4634
- miR-187-5p is hsa-miR-187-5p
- miR-6673-5p is hsa-miR-6673-5p
- miR-1908-3p Is hsa-miR-1908-3p
- miR-1181 is hsa-miR- 1181
- miR-6878-5p is hsa-miR-6878-5p
- miR-5010-5p is hsa-miR-5010-5p
- miR-6870-5p is hsa-miR-6870-5p.
- MiR-6124 is hsa-miR-6124, miR-1249-5p is hsa-miR-1249-5p, miR-6511b-5p is hsa-miR-6511b-5p, and miR-1254 is hsa-miR-1254, miR-4727-3p is hsa-miR-4727-3p, miR-4259 is hsa-miR-4259, miR-4771 is hsa-miR-4771, miR- 3622a-5p is hsa-miR-3622a-5p, and miR- 480 is hsa-miR-4480, miR-4740-5p is hsa-miR-4740-5p, miR-6777-5p is hsa-miR-6777-5p, and miR-6794-5p is hsa- miR-6794-5p, miR-4687-3p is hsa-mi
- polynucleotide according to any one of the following (a) to (e): (A) a polynucleotide comprising the nucleotide sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268 or a nucleotide sequence wherein u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more consecutive sequences Its fragments, including (B) a polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268, (C) a polynucleotide comprising a nucleotide sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268, or a nucleotide sequence complementary to the nucleotide sequence in which u is t in the nucleotide sequence, variants thereof and derivatives thereof Or fragments thereof containing 15 or more consecutive bases, (D) a polyn
- the device is another ovarian tumor marker, miR-320a, miR-663a, miR-328-5p, miR-128-2-5p, miR-125a-3p, miR-191-5p, miR -92b-5p, miR-296-5p, miR-1246, miR-92a-2-5p, miR-128-1-5p, miR-1290, miR-211-3p, miR-744-5p, miR-135a -3p, miR-451a, miR-625-3p, miR-92a-3p, miR-422a, miR-483-5p, miR-652-5p, miR-24-3p, miR-23b-3p, miR-23a At least one polynucleotide selected from -3p, miR-92b-3p, miR-22-3p, or the polynucleotide Further comprising a complementary strand capable of specifically binding a nucleic acid of the nucleotide A device according to any one of (8) to (10).
- miR-320a is hsa-miR-320a
- miR-663a is hsa-miR-663a
- miR-328-5p is hsa-miR-328-5p
- miR-128-2-5p Is hsa-miR-128-2-5p
- miR-125a-3p is hsa-miR-125a-3p
- miR-191-5p is hsa-miR-191-5p
- miR-92b-5p Is hsa-miR-92b-5p
- miR-296-5p is hsa-miR-296-5p
- miR-1246 is hsa-miR-1246
- miR-92a-2-5p is hsa-miR -92a-2-5p
- miR-128-1-5p is hsa-miR-128-1-5p
- the device is capable of specifically binding to at least two polynucleotides selected from all ovarian tumor markers according to (8) or (9), or each of the complementary strands of the polynucleotides.
- the device according to any one of (8) to (15), comprising one nucleic acid.
- Measure the expression level of the target gene in a body sample and teach the gene expression level of a sample derived from a subject who is known to have ovarian tumor and the gene expression level of a sample derived from a subject not suffering from ovarian tumor
- the target residue in the specimen from the subject is described.
- a method for detecting an ovarian tumor comprising substituting the expression level of a gene and thereby evaluating the presence or absence of the ovarian tumor in vitro.
- miR-4675 is hsa-miR-4675
- miR-4783-3p is hsa-miR-4783-3p
- miR-1228-5p is hsa-miR-1228-5p
- miR-4532 Is hsa-miR-4532
- miR-6802-5p is hsa-miR-6802-5p
- miR-6784-5p is hsa-miR-6784-5p
- miR-3940-5p is hsa-miR -3940-5p
- miR-1307-3p is hsa-miR-1307-3p
- miR-8073 is hsa-miR-8073
- miR-3184-5p is hsa-miR-3184-5p
- MiR-1233-5p is hsa-miR-1233-5p
- miR-60 8 is hsa-miR-6088
- polynucleotide according to any one of the following (a) to (e): (A) a polynucleotide comprising the nucleotide sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268, or a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more A fragment thereof containing a continuous base of (B) a polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268, (C) a polynucleotide comprising a base sequence complementary to the base sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268, or a base sequence in which u is t in the base sequence, variants thereof, A derivative thereof, or a fragment thereof comprising 15 or more consecutive bases, (D) a polynucleotide comprising a base sequence complementary to the
- the kit or device is another ovarian tumor marker, miR-320a, miR-663a, miR-328-5p, miR-128-2-5p, miR-125a-3p, miR-191-5p MiR-92b-5p, miR-296-5p, miR-1246, miR-92a-2-5p, miR-128-1-5p, miR-1290, miR-211-3p, miR-744-5p, miR -135a-3p, miR-451a, miR-625-3p, miR-92a-3p, miR-422a, miR-483-5p, miR-652-5p, miR-24-3p, miR-23b-3p, miR At least one polynucleotide selected from -23a-3p, miR-92b-3p, miR-22-3p Or further comprising a complementary strand capable of specifically binding a nucleic acid of the polynucleotide, the method according to any one of (17) to (21).
- miR-320a is hsa-miR-320a
- miR-663a is hsa-miR-663a
- miR-328-5p is hsa-miR-328-5p
- miR-128-2-5p Is hsa-miR-128-2-5p
- miR-125a-3p is hsa-miR-125a-3p
- miR-191-5p is hsa-miR-191-5p
- miR-92b-5p Is hsa-miR-92b-5p
- miR-296-5p is hsa-miR-296-5p
- miR-1246 is hsa-miR-1246
- miR-92a-2-5p is hsa-miR -92a-2-5p
- miR-128-1-5p is hsa-miR-128-1-5p
- polynucleotide according to any one of the following (f) to (j): (F) a polynucleotide comprising the nucleotide sequence represented by any of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, or a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, or a derivative thereof Or fragments thereof containing 15 or more consecutive bases, (G) a polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, (H) a polynucleotide comprising a base sequence complementary to the base sequence represented by any of SEQ ID NOS: 248 to 250, 252 to 267, and 269 to 275, or the base sequence in which u is t in the base sequence, A variant thereof, a derivative thereof, or a fragment thereof comprising 15 or more consecutive bases,
- polynucleotide is used for nucleic acids including RNA, DNA, and RNA / DNA (chimera).
- the DNA includes any of cDNA, genomic DNA, and synthetic DNA.
- the RNA includes total RNA, mRNA, rRNA, miRNA, siRNA, snoRNA, snRNA, non-coding RNA, and synthetic RNA.
- synthetic DNA and “synthetic RNA” are artificially generated using, for example, an automatic nucleic acid synthesizer based on a predetermined base sequence (which may be either a natural sequence or a non-natural sequence). It refers to the prepared DNA and RNA.
- non-natural sequence is intended to be used in a broad sense, and is a sequence (for example, including one or more nucleotide substitutions, deletions, insertions and / or additions) that differs from the natural sequence. That is, it includes a mutant sequence), a sequence containing one or more modified nucleotides (ie, a modified sequence), and the like.
- the polynucleotide is used interchangeably with the nucleic acid.
- a “fragment” is a polynucleotide having a base sequence of a continuous part of a polynucleotide, and desirably has a length of 15 bases or more, preferably 17 bases or more, more preferably 19 bases or more. .
- “gene” includes not only RNA and double-stranded DNA, but also each single-stranded DNA such as positive strand (or sense strand) or complementary strand (or antisense strand) constituting the same. It is intended to be used. The length is not particularly limited. Therefore, in this specification, unless otherwise specified, “gene” refers to double-stranded DNA including human genomic DNA, single-stranded DNA (positive strand), and single-stranded DNA having a sequence complementary to the positive strand (complementary). Strand), cDNA, microRNA (miRNA), and fragments thereof, the human genome, and transcripts thereof.
- the “gene” is not limited to a “gene” represented by a specific nucleotide sequence (or sequence number), but also RNAs having biological functions equivalent to RNA encoded by these, for example, homologs (ie, homologs). Or an ortholog), variants such as genetic polymorphisms, and “nucleic acids” encoding derivatives.
- the “nucleic acid” encoding such a homologue, variant or derivative is, for example, the base sequence represented by any one of SEQ ID NOs: 1 to 842 under the stringent conditions described later, or the base A “nucleic acid” having a base sequence that hybridizes with a complementary sequence of the base sequence in which u is t in the sequence can be mentioned.
- the “gene” does not ask whether the functional region is different, and may include, for example, an expression control region, a coding region, an exon, or an intron. Further, the “gene” may be contained in the cell, may be released outside the cell and may be present alone, or may be encapsulated in a vesicle called an exosome.
- exosome or “exosome” is a vesicle encapsulated in a lipid bilayer secreted from a cell. Exosomes are derived from multivesicular endosomes, and when released to the extracellular environment, they may contain biological substances such as “genes” such as RNA and DNA and proteins. It is known that exosomes are contained in body fluids such as blood, serum, plasma, serum and lymph.
- RNA refers to RNA synthesized using a DNA sequence of a gene as a template.
- RNA is synthesized in such a manner that RNA polymerase binds to a site called a promoter located upstream of the gene and ribonucleotides are bound to the 3 'end so as to be complementary to the DNA base sequence.
- This RNA includes not only the gene itself but also the entire sequence from the transcription start point to the end of the poly A sequence, including the expression control region, coding region, exon or intron.
- microRNA is a protein complex that is transcribed as a hairpin-like RNA precursor, cleaved by a dsRNA cleaving enzyme having RNase III cleaving activity, and called RISC. 15-25 base non-coding RNA that is incorporated into and is involved in the translational repression of mRNA.
- miRNA used herein includes not only “miRNA” represented by a specific base sequence (or sequence number) but also a precursor of the “miRNA” (pre-miRNA, pri-miRNA).
- miRNAs that are biologically equivalent to the miRNAs encoded by them, such as homologues (ie, homologs or orthologs), variants such as genetic polymorphisms, and derivatives that encode derivatives.
- the “miRNA” encoding such precursor, homologue, variant or derivative can be specifically identified by miRBase release 21 (http://www.mirbase.org/) and described later. Examples thereof include “miRNA” having a base sequence that hybridizes with a complementary sequence of any one of the specific base sequences represented by any of SEQ ID NOS: 1 to 842 under stringent conditions.
- miRNA used herein may be a gene product of a miR gene, and such a gene product is a mature miRNA (for example, 15 to 15 involved in the suppression of translation of mRNA as described above). 25-base, or 19-25 base non-coding RNA) or miRNA precursors (eg, pre-miRNA or pri-miRNA as described above).
- the “probe” includes a polynucleotide used for specifically detecting RNA produced by gene expression or a polynucleotide derived therefrom and / or a polynucleotide complementary thereto.
- the “primer” includes a continuous polynucleotide and / or a complementary polynucleotide that specifically recognizes and amplifies RNA generated by gene expression or a polynucleotide derived therefrom.
- a complementary polynucleotide is a polynucleotide comprising a base sequence defined by any one of SEQ ID NOs: 1 to 842 or a base sequence in which u is t in the base sequence.
- the base sequence is complementary to the full-length sequence or a partial sequence thereof (for convenience, this is referred to as the positive strand) based on the base pair relationship such as A: T (U), G: C.
- A T (U)
- G C.
- such a complementary strand is not limited to the case where it forms a completely complementary sequence with the target positive strand base sequence, but has a complementary relationship that allows hybridization with the target normal strand under stringent conditions. There may be.
- stringent conditions means a nucleic acid probe that is detectable to a greater extent than other sequences (eg, average of background measurements + standard error of background measurements ⁇ measured value of 2 or more). ) And the conditions for hybridizing to the target sequence. Stringent conditions are sequence-dependent and depend on the environment in which hybridization is performed. By controlling the stringency of the hybridization and / or wash conditions, target sequences that are 100% complementary to the nucleic acid probe can be identified. Specific examples of “stringent conditions” will be described later.
- the “Tm value” means a temperature at which a double-stranded portion of a polynucleotide is denatured into a single strand and the double strand and the single strand are present in a ratio of 1: 1.
- the term “variant” refers to a natural variant caused by polymorphism, mutation, or the like, a base sequence represented by a sequence number, or u in the base sequence in the case of a nucleic acid. Mutant including deletion, substitution, addition or insertion of 1, 2 or 3 or more (for example, 1 to several) bases in the base sequence or a partial sequence thereof, or any sequence of SEQ ID NOs: 1 to 275 A nucleotide sequence of the precursor RNA of (i), a nucleotide sequence in which u is t in the nucleotide sequence, or a mutant comprising a deletion, substitution, addition or insertion of one or more bases in a partial sequence thereof, Alternatively, there is a variant that exhibits a percent identity of about 90% or more, about 95% or more, about 97% or more, about 98% or more, or about 99% or more with each of the base sequences or a partial sequence thereof It refers to a polynucleotide or a nucleic acid.
- “several” means an integer of about 10, 9, 8, 7, 6, 5, 4, 3 or 2.
- a mutant can be prepared using a well-known technique such as site-directed mutagenesis or PCR-based mutagenesis.
- % identity is based on BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and FASTA (http://www.genome.jp/tools/fasta/).
- a protein or gene search system can be used to determine with or without introducing a gap (Zheng Zhang et al., 2000, J. Comput. Biol., 7, p203-214; Altschul, SF, et al., 1990, Journal of Molecular Biology, Vol. 215, p403-410; Pearson, WR, et al., 1988, Proc. Natl. Acad. Sci. USA, Volume 85, p2444- 448).
- the term “derivative” refers to a modified nucleic acid, a non-limiting group such as a labeled derivative such as a fluorophore, a modified nucleotide (for example, a halogen, an alkyl such as methyl, an alkoxy such as methoxy, a group such as thio, carboxymethyl, etc.
- a derivative containing PNA peptide nucleic acid; Nielsen, PE, etc.
- a polynucleotide selected from the above-mentioned ovarian tumor marker miRNA, or a “nucleic acid” capable of specifically binding to a complementary strand of the polynucleotide, or a “nucleic acid” capable of detecting the polynucleotide A nucleic acid synthesized or prepared, for example, containing a “nucleic acid probe” or “primer” to detect the presence or absence of an ovarian tumor in a subject, or whether or not an ovarian tumor is affected, Used directly or indirectly to diagnose the presence or extent of ovarian tumor improvement, sensitivity to treatment of ovarian tumors, or to screen for candidate substances useful for prevention, improvement or treatment of ovarian tumors.
- the term “detection” can be replaced by the term inspection, measurement, detection or decision support. Further, in this specification, the term “evaluation” is used in a meaning including supporting diagnosis or evaluation based on a test result or a measurement result.
- subject includes humans, primates including chimpanzees, pet animals such as dogs and cats, livestock animals such as cows, horses, sheep and goats, rodents such as mice and rats, Mammals such as animals raised in zoos.
- a preferred subject is a human.
- the “subject” is sometimes referred to as a “subject”.
- “Healthy person” also means an animal or subject that is such a mammal and does not suffer from the cancer to be detected.
- a preferred healthy person is a human.
- ovarian cancer is a malignant tumor that develops in the ovary, and includes, for example, epithelial ovarian cancer that arises from the mucosal epithelium and germ cell interstitial ovarian cancer.
- an “ovarian benign tumor” is a benign tumor that occurs in the ovary, and includes, for example, mucinous cystadenoma, serous cystadenoma, mature teratoma, and fibroma.
- ovarian tumor is a term encompassing the above “ovarian cancer” and “ovarian benign tumor”.
- P or “P value” refers to the probability that, in a statistical test, a statistic more extreme than the statistic actually calculated from the data under the null hypothesis is observed. Indicates. Therefore, it can be considered that the smaller the “P” or “P value”, the more significant the difference between the comparison objects.
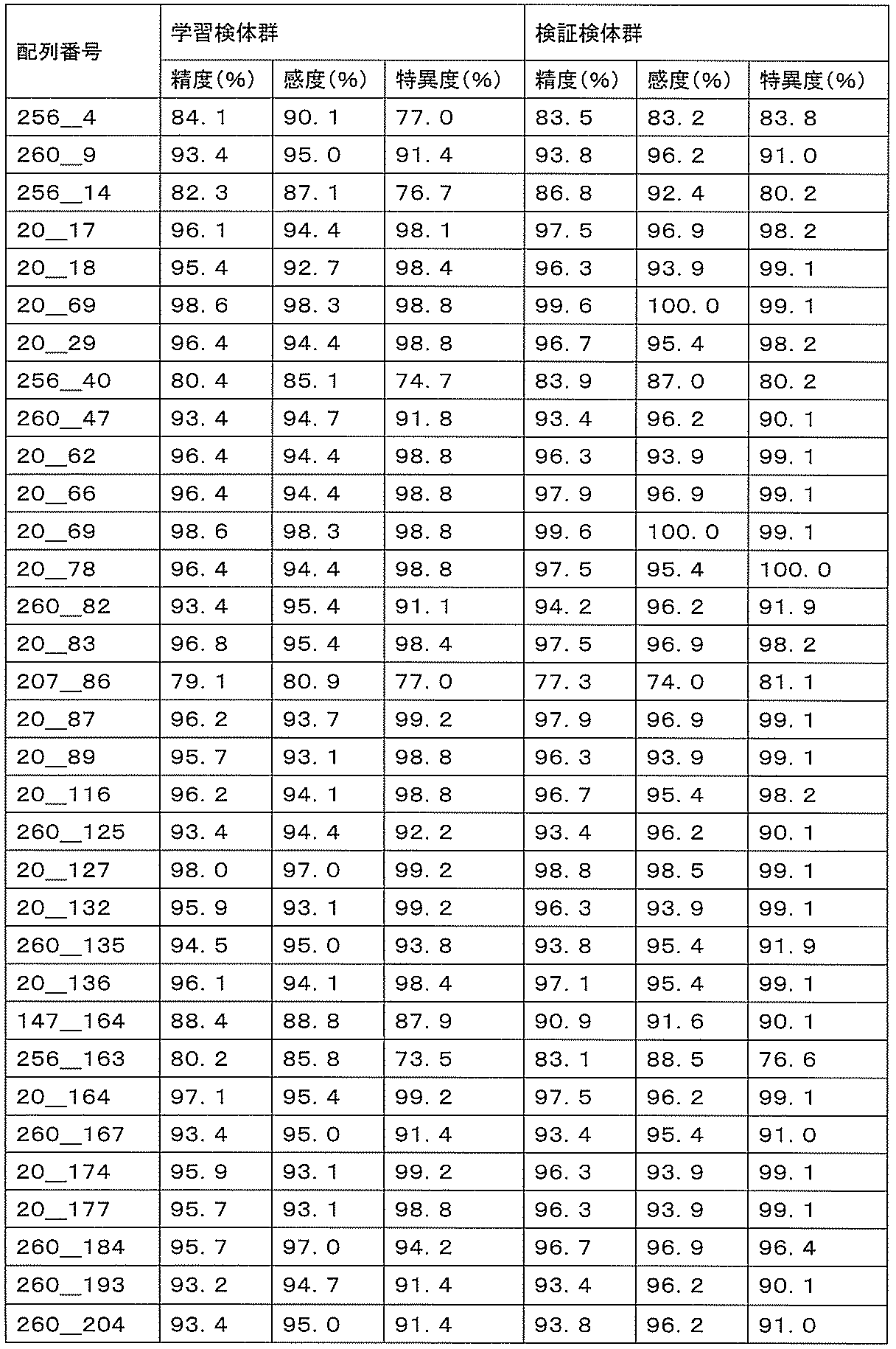
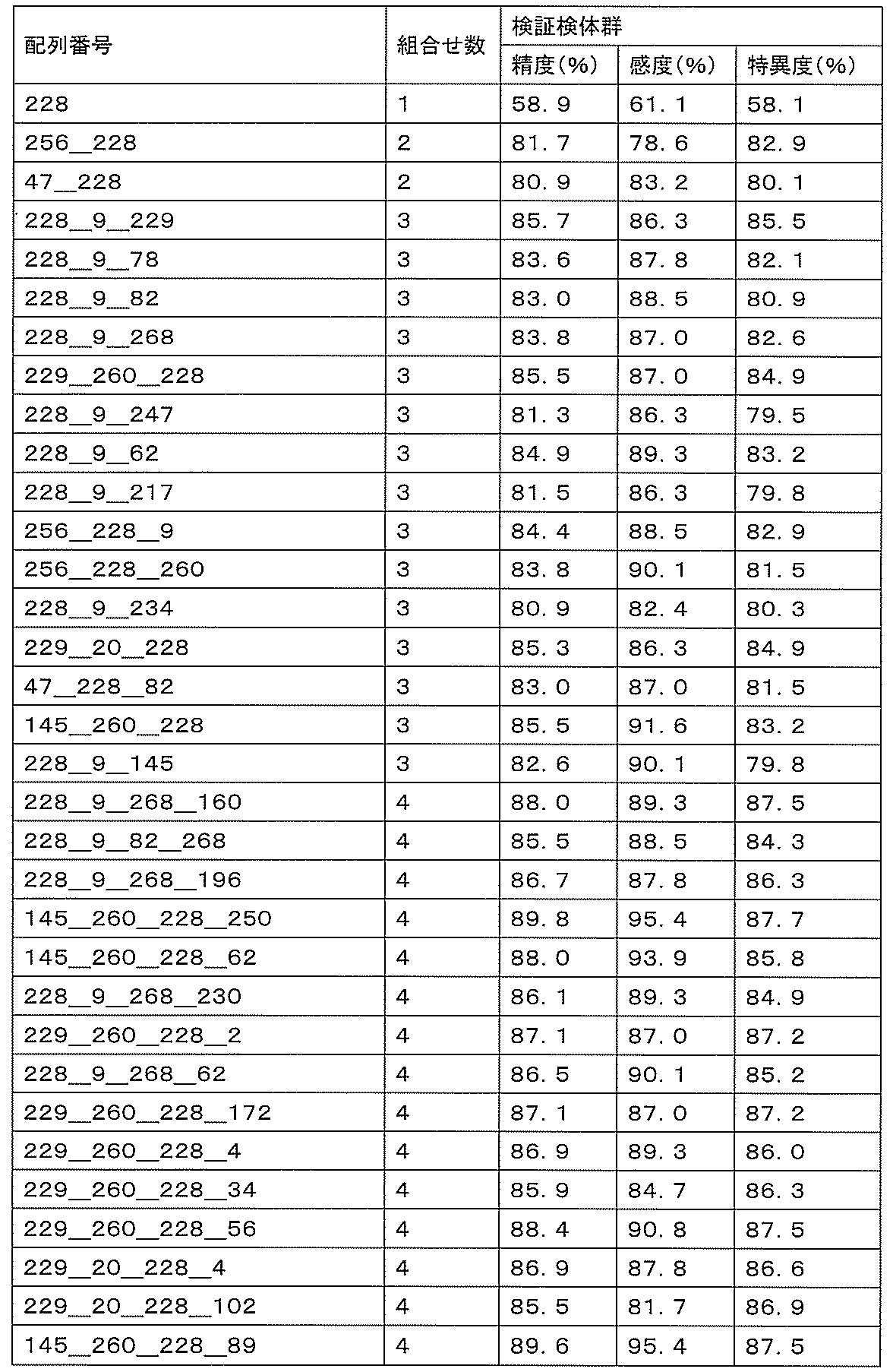
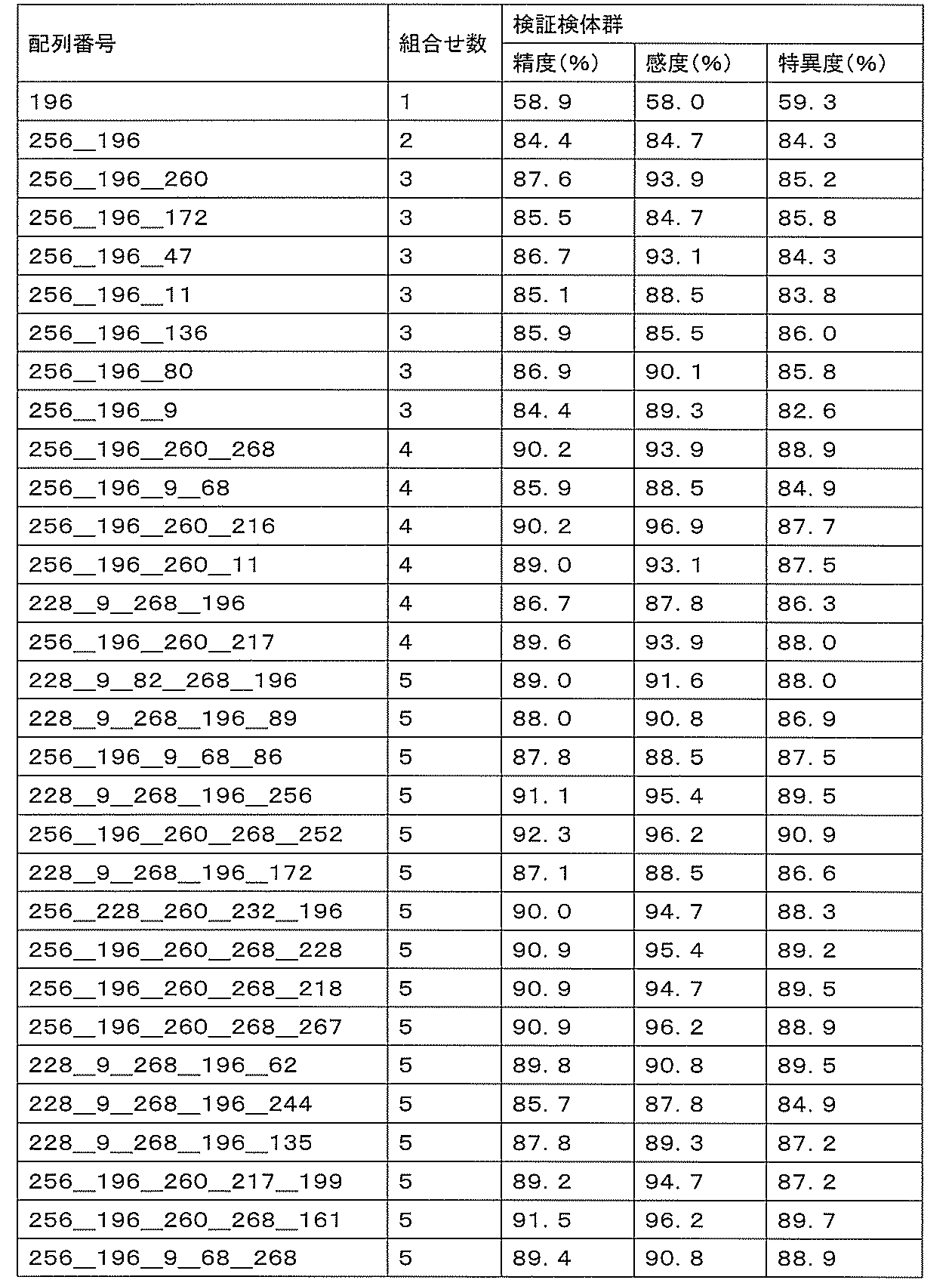
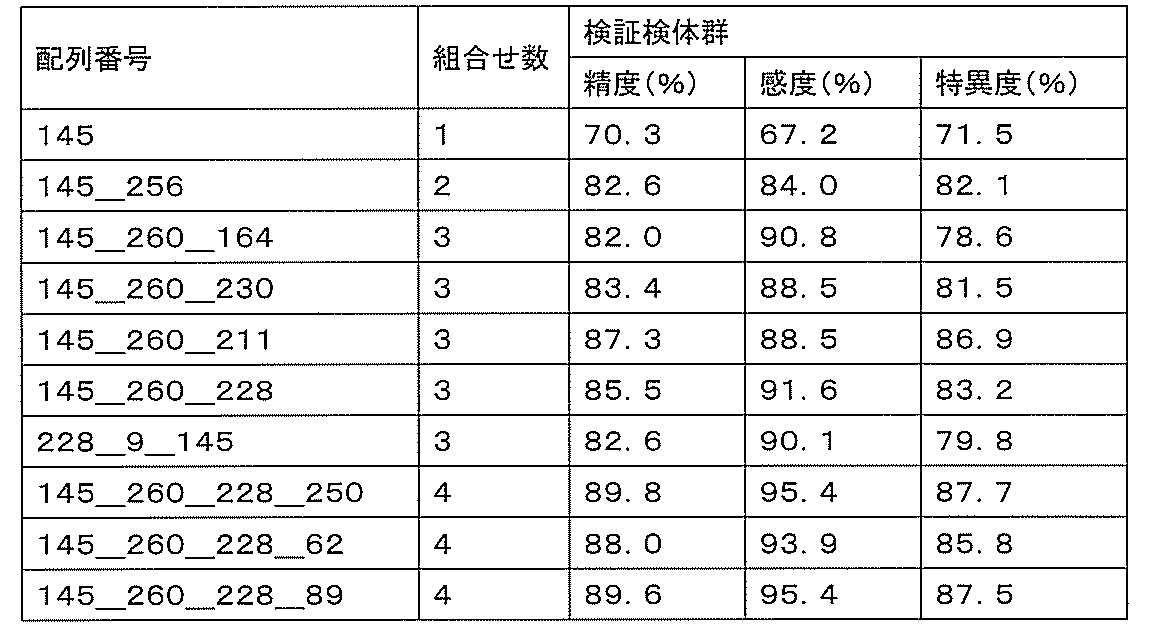
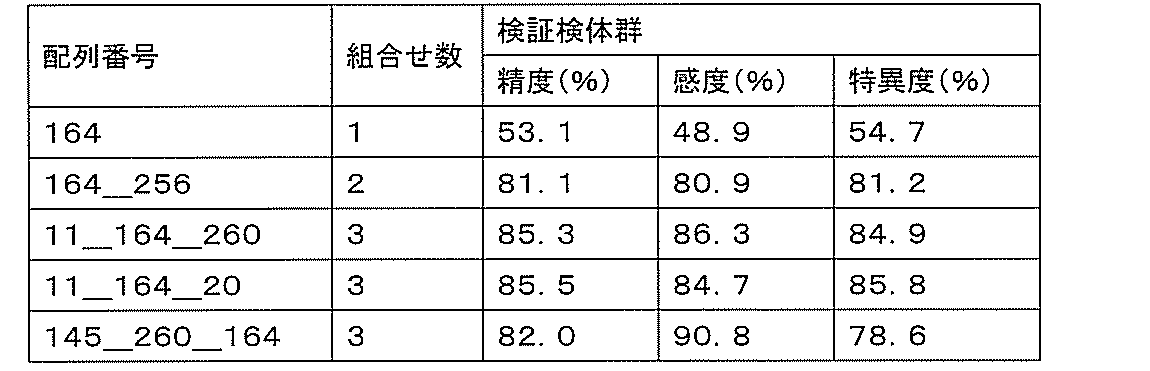
- sensitivity means a value of (number of true positives) / (number of true positives + number of false negatives). High sensitivity makes it possible to detect ovarian cancer early, leading to complete removal of the cancerous part and a reduction in the recurrence rate.
- specificity means (number of true negatives) / (number of true negatives + number of false positives). If the specificity is high, it is possible to prevent unnecessary additional tests by misidentifying a healthy person as an ovarian cancer patient, thereby reducing the burden on the patient and medical costs.
- accuracy means a value of (number of true positives + number of true negatives) / (number of all cases). The accuracy indicates the rate at which the discrimination results for all the samples are correct, and is a first index for evaluating the detection performance.
- the “specimen” to be determined, detected or diagnosed changes the expression of the gene of the present invention as ovarian cancer develops, ovarian cancer progresses, and the therapeutic effect on ovarian cancer is exerted.
- tissue and biomaterial Specifically, ovarian tissue and fallopian tubes, lymph nodes and surrounding organs, organs suspected of metastasis, skin, and body fluids such as blood, urine, saliva, sweat, tissue exudate, serum prepared from blood, plasma, In addition, it refers to feces and hair. Furthermore, it refers to a biological sample extracted from these, specifically genes such as RNA and miRNA.
- hsa-miR-4675 gene or “hsa-miR-4675” refers to the hsa-miR-4675 gene (miRBase Accession No. MIMAT0019757) described in SEQ ID NO: 1 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4675 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-4675 “hsa-mir-4675” (miRBase Accession No. MI0017306, SEQ ID NO: 276) having a hairpin-like structure as a precursor is known.
- hsa-miR-4783-3p gene or “hsa-miR-4783-3p” refers to the hsa-miR-4783-3p gene (miRBase Accession No. 2) described in SEQ ID NO: 2. MIMAT0019947) and other species homologues or orthologues are included.
- the hsa-miR-4783-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4783-3p” is known as “hsa-mir-4783” (miRBase Accession No. MI0017428, SEQ ID NO: 277) having a hairpin-like structure as a precursor.
- hsa-miR-1228-5p gene or “hsa-miR-1228-5p” refers to the hsa-miR-1228-5p gene described in SEQ ID NO: 3 (miRBase Accession No. MIMAT0005582) and other species homologues or orthologues are included.
- the hsa-miR-1228-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, 28, p328-336.
- “Hsa-miR-1228-5p” is known as “hsa-mir-1228” (miRBase Accession No. MI0006318, SEQ ID NO: 278) having a hairpin-like structure as a precursor.
- hsa-miR-4532 gene or “hsa-miR-4532” refers to the hsa-miR-4532 gene (miRBase Accession No. MIMAT0019071) described in SEQ ID NO: 4 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4532 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-mir-4532 miRBase Accession No. MI0016899, SEQ ID NO: 279 having a hairpin-like structure as a precursor is known.
- hsa-miR-6802-5p gene or “hsa-miR-6802-5p” refers to the hsa-miR-6802-5p gene (miRBase Accession No. 5) described in SEQ ID NO: 5. MIMAT0027504) and other species homologues or orthologues are included.
- the hsa-miR-6802-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6802-5p” is known as “hsa-mir-6802” (miRBase Accession No. MI0022647, SEQ ID NO: 280) having a hairpin-like structure as a precursor.
- hsa-miR-6784-5p gene or “hsa-miR-6784-5p” refers to the hsa-miR-6784-5p gene described in SEQ ID NO: 6 (miRBase Accession No. MIMAT0027468) and other species homologues or orthologues are included.
- the hsa-miR-6784-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6784-5p” “hsa-mir-6784” (miRBase Accession No. MI0022629, SEQ ID NO: 281) having a hairpin-like structure as a precursor is known.
- hsa-miR-3940-5p gene or “hsa-miR-3940-5p” refers to the hsa-miR-3940-5p gene (miRBase Accession No. 5) described in SEQ ID NO: 7. MIMAT0019229) and other species homologues or orthologues are included.
- the hsa-miR-3940-5p gene can be obtained by the method described in Liao JY et al., 2010, PLoS One, 5, e10563.
- “hsa-miR-3940-5p” is known as “hsa-mir-3940” (miRBase Accession No.
- hsa-miR-1307-3p gene or “hsa-miR-1307-3p” refers to the hsa-miR-1307-3p gene (miRBase Accession No. 5) described in SEQ ID NO: 8. MIMAT0005951) and other species homologues or orthologues are included.
- the hsa-miR-1307-3p gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- “hsa-miR-1307-3p” “hsa-mir-1307” (miRBase Accession No. MI0006444, SEQ ID NO: 283) having a hairpin-like structure as a precursor is known.
- hsa-miR-8073 gene or “hsa-miR-8073” refers to the hsa-miR-8073 gene (miRBase Accession No. MIMAT0031000) described in SEQ ID NO: 9 or other biological species. Homologs or orthologs are included.
- the hsa-miR-8073 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487.
- hsa-miR-8073 “hsa-mir-8073” (miRBase Accession No. MI0025909, SEQ ID NO: 284) having a hairpin-like structure as a precursor is known.
- hsa-miR-3184-5p gene or “hsa-miR-3184-5p” refers to the hsa-miR-3184-5p gene described in SEQ ID NO: 10 (miRBase Accession No. MIMAT0015064) and other species homologs or orthologs.
- the hsa-miR-3184-5p gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- “Hsa-miR-3184-5p” is known as “hsa-mir-3184” (miRBase Accession No. MI0014226, SEQ ID NO: 285), which has a hairpin-like structure as a precursor.
- hsa-miR-1233-5p gene or “hsa-miR-1233-5p” refers to the hsa-miR-1233-5p gene (miRBase Accession No. 5) described in SEQ ID NO: 11. MIMAT0022943) and other species homologues or orthologues are included.
- the hsa-miR-1233-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, 28, p328-336.
- “Hsa-miR-1233-5p” has a hairpin-like structure as a precursor thereof, “hsa-mir-1233-1, hsa-mir-12233-2” (miRBase Accession No. MI0006323, MI0015973, SEQ ID NO: 286, 287) are known.
- hsa-miR-6088 gene or “hsa-miR-6088” refers to the hsa-miR-6088 gene (miRBase Accession No. MIMAT0023713) described in SEQ ID NO: 12 and other biological species. Homologs or orthologs are included.
- the hsa-miR-6088 gene can be obtained by the method described in Yoo JK et al., 2012, Stem Cells Dev, 21, p2049-2057.
- hsa-miR-6088 is known as “hsa-mir-6088” (miRBase Accession No. MI0020365, SEQ ID NO: 288) having a hairpin-like structure as a precursor.
- hsa-miR-5195-3p gene or “hsa-miR-5195-3p” refers to the hsa-miR-5195-3p gene described in SEQ ID NO: 13 (miRBase Accession No. MIMAT0021127) and other species homologs or orthologs are included.
- the hsa-miR-5195-3p gene can be obtained by the method described in Schotte D et al., 2011, Leukemia, 25, p1389-1399.
- “hsa-miR-5195-3p” is known as “hsa-mir-5195” (miRBase Accession No. MI0018174, SEQ ID NO: 289) having a hairpin-like structure as a precursor.
- hsa-miR-320b gene or “hsa-miR-320b” refers to the hsa-miR-320b gene (miRBase Accession No. MIMAT0005792) described in SEQ ID NO: 14 and other biological species. A homolog or ortholog is included.
- the hsa-miR-320b gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, 16, p1299-1298.
- “Hsa-miR-320b” has a hairpin-like structure as a precursor thereof, “hsa-mir-320b-1, hsa-mir-320b-2” (miRBase Accession No. MI0003776, MI0003839, SEQ ID NO: 290, 291) is known.
- hsa-miR-4649-5p gene or “hsa-miR-4649-5p” refers to the hsa-miR-4649-5p gene described in SEQ ID NO: 15 (miRBase Accession No. MIMAT0019711) and other species homologues or orthologues are included.
- the hsa-miR-4649-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4649-5p” “hsa-mir-4649” (miRBase Accession No. MI0017276, SEQ ID NO: 292) having a hairpin-like structure as a precursor is known.
- hsa-miR-6800-5p gene or “hsa-miR-6800-5p” refers to the hsa-miR-6800-5p gene described in SEQ ID NO: 16 (miRBase Accession No. MIMAT0027500) and other species homologs or orthologs.
- the hsa-miR-6800-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6800-5p “hsa-mir-6800” (miRBase Accession No. MI0022645, SEQ ID NO: 293) having a hairpin-like structure as a precursor is known.
- hsa-miR-1343-3p gene or “hsa-miR-1343-3p” refers to the hsa-miR-1343-3p gene described in SEQ ID NO: 17 (miRBase Accession No. MIMAT0019776) and other species homologs or orthologs.
- the hsa-miR-1343-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-1343-3p “hsa-mir-1343” (miRBase Accession No. MI0017320, SEQ ID NO: 294) having a hairpin-like structure as a precursor is known.
- hsa-miR-4730 gene or “hsa-miR-4730” refers to the hsa-miR-4730 gene (miRBase Accession No. MIMAT0019852) described in SEQ ID NO: 18 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4730 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-4730 “hsa-mir-4730” (miRBase Accession No. MI0017367, SEQ ID NO: 295) having a hairpin-like structure as a precursor is known.
- hsa-miR-685-5p gene or “hsa-miR-685-5p” refers to the hsa-miR-6885-5p gene (miRBase Accession No. MIMAT0027670) and other species homologs or orthologs.
- the hsa-miR-685-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6885-5p” is known as “hsa-mir-6885” (miRBase Accession No. MI0022732, SEQ ID NO: 296) having a hairpin-like structure as a precursor.
- hsa-miR-5100 gene or “hsa-miR-5100” refers to the hsa-miR-5100 gene (miRBase Accession No. MIMAT0022259) described in SEQ ID NO: 20 or other biological species. Homologs or orthologs are included.
- the hsa-miR-5100 gene can be obtained by the method described in Tandon M, et al., 2012, Oral Dis, 18, p127-131.
- hsa-miR-5100 is known as “hsa-mir-5100” (miRBase Accession No. MI0019116, SEQ ID NO: 297), which has a hairpin-like structure as a precursor.
- hsa-miR-1203 gene or “hsa-miR-1203” refers to the hsa-miR-1203 gene (miRBase Accession No. MIMAT0005866) described in SEQ ID NO: 21 and other species. Homologs or orthologs are included.
- the hsa-miR-1203 gene can be obtained by the method described in Marton S, et al., 2008, Leukemia, Vol. 22, p330-338.
- “Hsa-miR-1203” is known as “hsa-mir-1203” (miRBase Accession No. MI0006335, SEQ ID NO: 298), which has a hairpin-like structure as a precursor.
- hsa-miR-6756-5p gene or “hsa-miR-6756-5p” refers to the hsa-miR-6756-5p gene described in SEQ ID NO: 22 (miRBase Accession No. MIMAT0027412) and other species homologues or orthologues are included.
- the hsa-miR-6756-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6756-5p “hsa-mir-6756” (miRBase Accession No. MI0022601, SEQ ID NO: 299) having a hairpin-like structure as a precursor is known.
- hsa-miR-373-5p gene or “hsa-miR-373-5p” refers to the hsa-miR-373-5p gene described in SEQ ID NO: 23 (miRBase Accession No. MIMAT000025) and other species homologues or orthologues are included.
- the hsa-miR-373-5p gene can be obtained by the method described in Suh MR et al., 2004, Dev Biol, 270, p488-498.
- hsa-miR-373-5p “hsa-mir-373” (miRBase Accession No. MI0000811, SEQ ID NO: 300) having a hairpin-like structure as a precursor is known.
- hsa-miR-1268a gene or “hsa-miR-1268a” refers to the hsa-miR-1268a gene (miRBase Accession No. MIMAT0005922) described in SEQ ID NO: 24 and other biological species. Homologs or orthologs are included.
- the hsa-miR-1268a gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- “hsa-miR-1268a” is known as “hsa-mir-1268a” (miRBase Accession No. MI0006405, SEQ ID NO: 301) having a hairpin-like structure as a precursor.
- hsa-miR-1260b gene or “hsa-miR-1260b” refers to the hsa-miR-1260b gene (miRBase Accession No. MIMAT0015041) described in SEQ ID NO: 25 and other biological species. Homologs or orthologs are included.
- the hsa-miR-1260b gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- “hsa-miR-1260b” is known as “hsa-mir-1260b” (miRBase Accession No. MI0014197, SEQ ID NO: 302), which has a hairpin-like structure as a precursor.
- hsa-miR-4258 gene or “hsa-miR-4258” refers to the hsa-miR-4258 gene (miRBase Accession No. MIMAT0016879) described in SEQ ID NO: 26 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4258 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- hsa-miR-4258 “hsa-mir-4258” (miRBase Accession No. MI0015857, SEQ ID NO: 303) having a hairpin-like structure as a precursor is known.
- hsa-miR-4697-5p gene or “hsa-miR-4697-5p” refers to the hsa-miR-4697-5p gene described in SEQ ID NO: 27 (miRBase Accession No. MIMAT0019791) and other species homologues or orthologues are included.
- the hsa-miR-4697-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4697-5p” “hsa-mir-4697” (miRBase Accession No. MI0017330, SEQ ID NO: 304) having a hairpin-like structure as a precursor is known.
- hsa-miR-1469 gene or “hsa-miR-1469” refers to the hsa-miR-1469 gene (miRBase Accession No. MIMAT0007347) described in SEQ ID NO: 28 or other biological species. Homologs or orthologs are included.
- the hsa-miR-1469 gene can be obtained by the method described in Kawaji H et al., 2008, BMC Genomics, Vol. 9, p157.
- “Hsa-miR-1469” is known as “hsa-mir-1469” (miRBase Accession No. MI00000074, SEQ ID NO: 305) having a hairpin-like structure as a precursor.
- hsa-miR-4515 gene or “hsa-miR-4515” refers to the hsa-miR-4515 gene (miRBase Accession No. MIMAT0019052) described in SEQ ID NO: 29 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4515 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- “hsa-miR-4515” is known as “hsa-mir-4515” (miRBase Accession No. MI0016881, SEQ ID NO: 306) having a hairpin-like structure as a precursor.
- hsa-miR-6861-5p gene or “hsa-miR-6861-5p” refers to the hsa-miR-6861-5p gene (miRBase Accession No. MIMAT0027623) and other species homologues or orthologues are included.
- the hsa-miR-6861-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6861-5p “hsa-mir-6861” (miRBase Accession No. MI0022708, SEQ ID NO: 307) having a hairpin-like structure as a precursor is known.
- hsa-miR-6821-5p gene or “hsa-miR-6821-5p” refers to the hsa-miR-6821-5p gene (miRBase Accession No. MIMAT0027542) and other species homologues or orthologues are included.
- the hsa-miR-6682-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “Hsa-miR-6821-5p” is known as “hsa-mir-6821” (miRBase Accession No. MI0022666, SEQ ID NO: 308), which has a hairpin-like structure as a precursor.
- hsa-miR-575 gene or “hsa-miR-575” refers to the hsa-miR-575 gene (miRBase Accession No. MIMAT0003240) described in SEQ ID NO: 32 or other biological species. Homologs or orthologs are included.
- the hsa-miR-575 gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692.
- “hsa-miR-575” is known as “hsa-mir-575” (miRBase Accession No. MI0003582, SEQ ID NO: 309) having a hairpin-like structure as a precursor.
- hsa-miR-6805-5p gene or “hsa-miR-6805-5p” refers to the hsa-miR-6805-5p gene described in SEQ ID NO: 33 (miRBase Accession No. MIMAT0027510) and other species homologs or orthologs.
- the hsa-miR-6805-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6805-5p” is known as “hsa-mir-6805” (miRBase Accession No. MI0022650, SEQ ID NO: 310) having a hairpin-like structure as a precursor.
- hsa-miR-4758-5p gene or “hsa-miR-4758-5p” refers to the hsa-miR-4758-5p gene described in SEQ ID NO: 34 (miRBase Accession No. MIMAT0019903) and other species homologues or orthologues are included.
- the hsa-miR-4758-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4758-5p” is known as “hsa-mir-4758” (miRBase Accession No. MI0017399, SEQ ID NO: 311) having a hairpin-like structure as a precursor.
- hsa-miR-3663-3p gene or “hsa-miR-3663-3p” refers to the hsa-miR-3663-3p gene described in SEQ ID NO: 35 (miRBase Accession No. MIMAT0018085) and other species homologs or orthologs.
- the hsa-miR-3663-3p gene can be obtained by the method described in Liao JY et al., 2010, PLoS One, 5, e10563.
- “hsa-miR-3663-3p” is known as “hsa-mir-3663” (miRBase Accession No. MI0016064, SEQ ID NO: 312) having a hairpin-like structure as a precursor.
- hsa-miR-4530 gene or “hsa-miR-4530” refers to the hsa-miR-4530 gene (miRBase Accession No. MIMAT0019069) described in SEQ ID NO: 36 or other species. Homologs or orthologs are included.
- the hsa-miR-4530 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-mir-4530 (miRBase Accession No. MI0016897, SEQ ID NO: 313) having a hairpin-like structure as a precursor is known.
- hsa-miR-6798-5p gene or “hsa-miR-6798-5p” refers to the hsa-miR-6798-5p gene described in SEQ ID NO: 37 (miRBase Accession No. MIMAT0027496) and other species homologs or orthologs.
- the hsa-miR-6798-5p gene can be obtained by the method described in Ladwig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6798-5p “hsa-mir-6798” (miRBase Accession No. MI0022643, SEQ ID NO: 314) having a hairpin-like structure as a precursor is known.
- hsa-miR-6781-5p gene or “hsa-miR-6781-5p” refers to the hsa-miR-6781-5p gene described in SEQ ID NO: 38 (miRBase Accession No. MIMAT0027462) and other species homologues or orthologues are included.
- the hsa-miR-6781-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6781-5p “hsa-mir-6781” (miRBase Accession No. MI0022626, SEQ ID NO: 315) having a hairpin-like structure as a precursor is known.
- hsa-miR-885-3p gene or “hsa-miR-885-3p” refers to the hsa-miR-885-3p gene described in SEQ ID NO: 39 (miRBase Accession No. MIMAT0004948) and other species homologs or orthologs.
- the hsa-miR-885-3p gene can be obtained by the method described in Berezikov E, et al., 2006, Genome Res, 16, p1299-1298.
- hsa-miR-885-3p “hsa-mir-885” (miRBase Accession No. MI0005560, SEQ ID NO: 316) having a hairpin-like structure as a precursor is known.
- hsa-miR-1273g-3p gene or “hsa-miR-1273g-3p” refers to the hsa-miR-1273g-3p gene described in SEQ ID NO: 40 (miRBase Accession No. MIMAT0022742) and other species homologues or orthologues are included.
- the hsa-miR-1273g-3p gene can be obtained by the method described in Reshmi G et al., 2011, Genomics, 97, p333-340.
- hsa-miR-1273g-3p “hsa-mir-1273g” (miRBase Accession No. MI0018003, SEQ ID NO: 317) having a hairpin-like structure as a precursor is known.
- hsa-miR-4787-3p gene or “hsa-miR-4787-3p” refers to the hsa-miR-4787-3p gene described in SEQ ID NO: 41 (miRBase Accession No. MIMAT0019957) and other species homologs or orthologs.
- the hsa-miR-4787-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4787-3p” “hsa-mir-4787” (miRBase Accession No. MI0017434, SEQ ID NO: 318) having a hairpin-like structure as a precursor is known.
- hsa-miR-4454 gene or “hsa-miR-4454” refers to the hsa-miR-4454 gene (miRBase Accession No. MIMAT0018976) described in SEQ ID NO: 42 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4454 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-mir-4454 (miRBase Accession No. MI0016800, SEQ ID NO: 319) having a hairpin-like structure as a precursor is known.
- hsa-miR-4706 gene or “hsa-miR-4706” refers to the hsa-miR-4706 gene (miRBase Accession No. MIMAT0019806) described in SEQ ID NO: 43 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4706 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-4706 “hsa-mir-4706” (miRBase Accession No. MI0017339, SEQ ID NO: 320) having a hairpin-like structure as a precursor is known.
- hsa-miR-1249-3p gene or “hsa-miR-1249-3p” refers to the hsa-miR-1249-3p gene described in SEQ ID NO: 44 (miRBase Accession No. MIMAT0005901) and other species homologues or orthologues are included.
- the hsa-miR-1249-3p gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- “hsa-miR-1249-3p” “hsa-mir-1249” (miRBase Accession No. MI0006384, SEQ ID NO: 321) having a hairpin-like structure as a precursor is known.
- hsa-miR-887-3p gene or “hsa-miR-887-3p” refers to the hsa-miR-887-3p gene described in SEQ ID NO: 45 (miRBase Accession No. MIMAT0004951) and other species homologs or orthologs.
- the hsa-miR-887-3p gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, 16: p1299-1298.
- hsa-miR-887-3p “hsa-mir-887” (miRBase Accession No. MI0005562, SEQ ID NO: 322) having a hairpin-like structure as a precursor is known.
- hsa-miR-6786-5p gene or “hsa-miR-6786-5p” refers to the hsa-miR-6786-5p gene described in SEQ ID NO: 46 (miRBase Accession No. MIMAT0027472) and other species homologues or orthologues are included.
- the hsa-miR-6786-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6786-5p “hsa-mir-6786” (miRBase Accession No. MI0022631, SEQ ID NO: 323) having a hairpin-like structure as a precursor is known.
- hsa-miR-1238-5p gene or “hsa-miR-1238-5p” refers to the hsa-miR-1238-5p gene described in SEQ ID NO: 47 (miRBase Accession No. MIMAT0022947) and other species homologues or orthologues are included.
- the hsa-miR-1238-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, 28, p328-336.
- “hsa-miR-1238-5p” is known as “hsa-mir-1238” (miRBase Accession No. MI0006328, SEQ ID NO: 324) having a hairpin-like structure as a precursor.
- hsa-miR-6749-5p gene or “hsa-miR-6749-5p” refers to the hsa-miR-6749-5p gene described in SEQ ID NO: 48 (miRBase Accession No. MIMAT0027398) and other species homologs or orthologs.
- the hsa-miR-6749-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6749-5p “hsa-mir-6749” (miRBase Accession No. MI0022594, SEQ ID NO: 325) having a hairpin-like structure as a precursor is known.
- hsa-miR-6729-5p gene or “hsa-miR-6729-5p” refers to the hsa-miR-6729-5p gene described in SEQ ID NO: 49 (miRBase Accession No. MIMAT0027359) and other species homologues or orthologues are included.
- the hsa-miR-6729-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6729-5p “hsa-mir-6729” (miRBase Accession No. MI0022574, SEQ ID NO: 326) having a hairpin-like structure as a precursor is known.
- hsa-miR-6825-5p gene or “hsa-miR-6825-5p” refers to the hsa-miR-6825-5p gene described in SEQ ID NO: 50 (miRBase Accession No. MIMAT0027550) and other species homologues or orthologues are included.
- the hsa-miR-6825-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6825-5p” is known as “hsa-mir-6825” (miRBase Accession No. MI0022670, SEQ ID NO: 327) having a hairpin-like structure as a precursor.
- hsa-miR-663b gene or “hsa-miR-663b” refers to the hsa-miR-663b gene (miRBase Accession No. MIMAT0005867) described in SEQ ID NO: 51 and other biological species. Homologs or orthologs are included.
- the hsa-miR-663b gene can be obtained by the method described in Takada S et al., 2008, Leukemia, Vol. 22, p1274-1278.
- hsa-miR-663b is known as “hsa-mir-663b” (miRBase Accession No. MI0006336, SEQ ID NO: 328) having a hairpin-like structure as a precursor.
- hsa-miR-6858-5p gene or “hsa-miR-6858-5p” refers to the hsa-miR-6858-5p gene described in SEQ ID NO: 52 (miRBase Accession No. MIMAT0027616) and other species homologs or orthologs.
- the hsa-miR-6858-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6858-5p “hsa-mir-6858” (miRBase Accession No. MI0022704, SEQ ID NO: 329) having a hairpin-like structure as a precursor is known.
- hsa-miR-4690-5p gene or “hsa-miR-4690-5p” refers to the hsa-miR-4690-5p gene described in SEQ ID NO: 53 (miRBase Accession No. MIMAT0019779) and other species homologs or orthologs.
- the hsa-miR-4690-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4690-5p” “hsa-mir-4690” (miRBase Accession No. MI0017323, SEQ ID NO: 330) having a hairpin-like structure as a precursor is known.
- hsa-miR-6765-5p gene or “hsa-miR-6765-5p” refers to the hsa-miR-6765-5p gene described in SEQ ID NO: 54 (miRBase Accession No. MIMAT0027430) and other species homologs or orthologs.
- the hsa-miR-6765-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6765-5p “hsa-mir-6765” (miRBase Accession No. MI0022610, SEQ ID NO: 331) having a hairpin-like structure as a precursor is known.
- hsa-miR-4710 gene or “hsa-miR-4710” refers to the hsa-miR-4710 gene (miRBase Accession No. MIMAT0019815) described in SEQ ID NO: 55 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4710 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4710” is known as “hsa-mir-4710” (miRBase Accession No. MI0017344, SEQ ID NO: 332) having a hairpin-like structure as a precursor.
- hsa-miR-6775-5p gene or “hsa-miR-6775-5p” refers to the hsa-miR-6775-5p gene described in SEQ ID NO: 56 (miRBase Accession No. MIMAT0027450) and other species homologs or orthologs.
- the hsa-miR-6775-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6775-5p” is known as “hsa-mir-6775” (miRBase Accession No. MI0022620, SEQ ID NO: 333) having a hairpin-like structure as a precursor.
- hsa-miR-371a-5p gene or “hsa-miR-371a-5p” refers to the hsa-miR-371a-5p gene described in SEQ ID NO: 57 (miRBase Accession No. MIMAT0004687) and other species homologues or orthologues are included.
- the hsa-miR-371a-5p gene can be obtained by the method described in Suh MR et al., 2004, Dev Biol, 270, p488-498.
- “hsa-miR-371a-5p” is known as “hsa-mir-371a” (miRBase Accession No. MI000079, SEQ ID NO: 334) having a hairpin-like structure as a precursor.
- hsa-miR-6816-5p gene or “hsa-miR-6816-5p” refers to the hsa-miR-6816-5p gene described in SEQ ID NO: 58 (miRBase Accession No. MIMAT0027532) and other species homologues or orthologues are included.
- the hsa-miR-6816-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6816-5p “hsa-mir-6816” (miRBase Accession No. MI0022661, SEQ ID NO: 335) having a hairpin-like structure as a precursor is known.
- hsa-miR-296-3p gene or “hsa-miR-296-3p” refers to the hsa-miR-296-3p gene described in SEQ ID NO: 59 (miRBase Accession No. MIMAT0004679) and other species homologs or orthologs.
- the hsa-miR-296-3p gene can be obtained by the method described in Houbavy HB et al., 2003, Dev Cell, Vol. 5, p351-358.
- “Hsa-miR-296-3p” is known as “hsa-mir-296” (miRBase Accession No. MI000047, SEQ ID NO: 336) having a hairpin-like structure as a precursor.
- hsa-miR-7777 gene or “hsa-miR-7777” refers to the hsa-miR-7777 gene (miRBase Accession No. MIMAT0031180) described in SEQ ID NO: 60 and other biological species. Homologs or orthologs are included.
- the hsa-miR-7777 gene can be obtained by the method described in Velthut-Meikas A et al., 2013, Endocrinol, Vol. 27, p1128-1141.
- hsa-miR-7777 “hsa-mir-7777” (miRBase Accession No. MI0025753, SEQ ID NO: 337) having a hairpin-like structure as a precursor is known.
- hsa-miR-8069 gene or “hsa-miR-8069” refers to the hsa-miR-8069 gene (miRBase Accession No. MIMAT0030996) described in SEQ ID NO: 61 or other biological species. Homologs or orthologs are included.
- the hsa-miR-8069 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487.
- “Hsa-miR-8069” has a hairpin-like structure as a precursor thereof, “hsa-mir-8069-1, hsa-mir-8069-2” (miRBase Accession No. MI0025905, MI0031519, SEQ ID NO: 338, 339) is known.
- hsa-miR-6515-3p gene or “hsa-miR-6515-3p” refers to the hsa-miR-6515-3p gene described in SEQ ID NO: 62 (miRBase Accession No. MIMAT0025487) and other species homologues or orthologues are included.
- the hsa-miR-6515-3p gene can be obtained by the method described in Joyce CE et al., 2011, Hum Mol Genet, 20, p4025-4040.
- hsa-mir-6515 (miRBase Accession No. MI0022227, SEQ ID NO: 340) having a hairpin-like structure as a precursor is known.
- hsa-miR-4687-5p gene or “hsa-miR-4687-5p” refers to the hsa-miR-4687-5p gene described in SEQ ID NO: 63 (miRBase Accession No. MIMAT0019774) and other species homologues or orthologues are included.
- the hsa-miR-4687-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4687-5p” “hsa-mir-4687” (miRBase Accession No. MI0017319, SEQ ID NO: 341) having a hairpin-like structure as a precursor is known.
- hsa-miR-1343-5p gene or “hsa-miR-1343-5p” refers to the hsa-miR-1343-5p gene described in SEQ ID NO: 64 (miRBase Accession No. MIMAT0027038) and other species homologs or orthologs.
- the hsa-miR-1343-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-1343-5p” is known as “hsa-mir-1343” (miRBase Accession No. MI0017320, SEQ ID NO: 294), which has a hairpin-like structure as a precursor.
- hsa-miR-7110-5p gene or “hsa-miR-7110-5p” refers to the hsa-miR-7110-5p gene described in SEQ ID NO: 65 (miRBase Accession No. MIMAT0028117) and other species homologues or orthologues are included.
- the hsa-miR-7110-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-7110-5p” is known as “hsa-mir-7110” (miRBase Accession No. MI0022961, SEQ ID NO: 342) having a hairpin-like structure as a precursor.
- hsa-miR-4525 gene or “hsa-miR-4525” refers to the hsa-miR-4525 gene (miRBase Accession No. MIMAT0019064) described in SEQ ID NO: 66 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4525 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- “Hsa-miR-4525” is known as “hsa-mir-4525” (miRBase Accession No. MI0016892, SEQ ID NO: 343) having a hairpin-like structure as a precursor.
- hsa-miR-3158-5p gene or “hsa-miR-3158-5p” refers to the hsa-miR-3158-5p gene described in SEQ ID NO: 67 (miRBase Accession No. MIMAT0019211) and other species homologues or orthologues are included.
- the hsa-miR-3158-5p gene can be obtained by the method described in Creighton CJ et al., 2010, PLoS One, 5, e9637.
- “Hsa-miR-3158-5p” has a hairpin-like structure as a precursor thereof, “hsa-mir-3158-1, hsa-mir-3158-2” (miRBase Accession No. MI0014186, MI0014187, SEQ ID NO: 344, 345) are known.
- hsa-miR-6787-5p gene or “hsa-miR-6787-5p” refers to the hsa-miR-6787-5p gene described in SEQ ID NO: 68 (miRBase Accession No. MIMAT0027474) and other species homologues or orthologues are included.
- the hsa-miR-6787-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6787-5p “hsa-mir-6787” (miRBase Accession No. MI0022632, SEQ ID NO: 346) having a hairpin-like structure as a precursor is known.
- hsa-miR-614 gene or “hsa-miR-614” refers to the hsa-miR-614 gene (miRBase Accession No. MIMAT0003282) described in SEQ ID NO: 69 or other biological species. Homologs or orthologs are included.
- the hsa-miR-614 gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci US A, 103, p3687-3692.
- “hsa-miR-614” is known as “hsa-mir-614” (miRBase Accession No. MI0003627, SEQ ID NO: 347) having a hairpin-like structure as a precursor.
- hsa-miR-4689 gene or “hsa-miR-4687” refers to the hsa-miR-4689 gene (miRBase Accession No. MIMAT0019778) described in SEQ ID NO: 70 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4689 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4689” is known as “hsa-mir-4689” (miRBase Accession No. MI0017322, SEQ ID NO: 348) having a hairpin-like structure as a precursor.
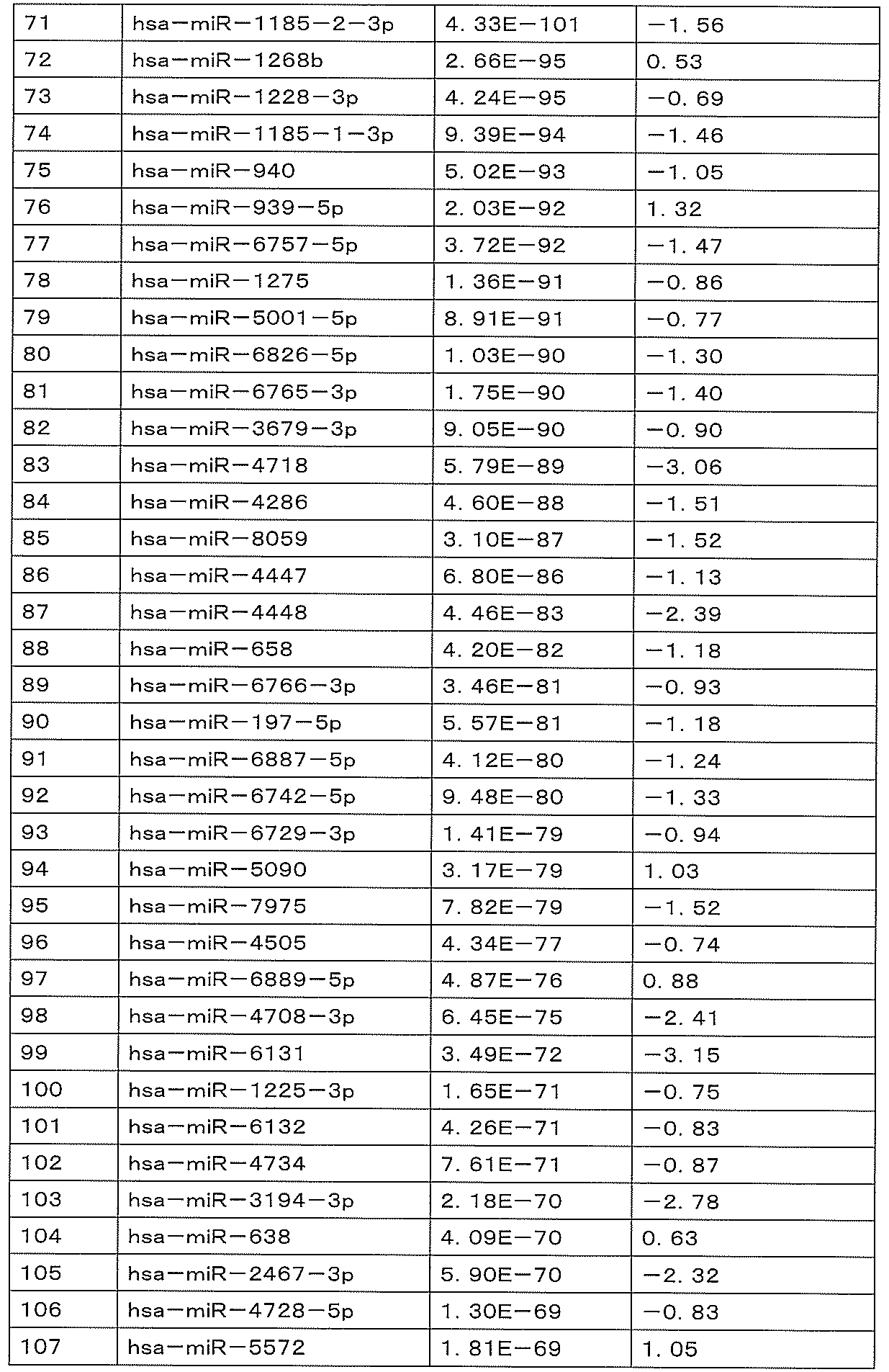
- hsa-miR-1185-2-3p gene or “hsa-miR-1185-2-3p” as used herein refers to the hsa-miR-1185-2-3p described in SEQ ID NO: 71. Genes (miRBBase Accession No. MIMAT0022713) and other species homologues or orthologues are included.
- the hsa-miR-1185-2-3p gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, 16, p1299-1298.
- “hsa-miR-1185-2-3p” is known as “hsa-mir-1185-2” (miRBase Accession No. MI0003821, SEQ ID NO: 349) having a hairpin-like structure as a precursor.
- hsa-miR-1268b gene or “hsa-miR-1268b” refers to the hsa-miR-1268b gene (miRBase Accession No. MIMAT0018925) described in SEQ ID NO: 72 or other species. A homolog or ortholog is included.
- the hsa-miR-1268b gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- “hsa-miR-1268b” is known as “hsa-mir-1268b” (miRBase Accession No. MI0016748, SEQ ID NO: 350) having a hairpin-like structure as a precursor.
- hsa-miR-1228-3p gene or “hsa-miR-1228-3p” refers to the hsa-miR-1228-3p gene described in SEQ ID NO: 73 (miRBase Accession No. MIMAT0005583) and other species homologs or orthologs are included.
- the hsa-miR-1228-3p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, 28, p328-336.
- hsa-mir-1228 (miRBase Accession No. MI0006318, SEQ ID NO: 278) having a hairpin-like structure as a precursor is known.
- hsa-miR-1185-1-3p gene or “hsa-miR-1185-1-3p” as used herein refers to the hsa-miR-1185-1-3p described in SEQ ID NO: 74. Genes (miRBBase Accession No. MIMAT0022838) and other species homologues or orthologues are included.
- the hsa-miR-1185-1-3p gene can be obtained by the method described in Berezikov E, et al., 2006, Genome Res, 16, p1299-1298.
- hsa-mir-1185-1 miRBase Accession No. MI0003844, SEQ ID NO: 351 having a hairpin-like structure as a precursor is known.
- hsa-miR-940 gene or “hsa-miR-940” refers to the hsa-miR-940 gene (miRBase Accession No. MIMAT0004983) described in SEQ ID NO: 75 and other biological species. Homologs or orthologs are included.
- the hsa-miR-940 gene can be obtained by the method described in Lui WO et al., 2007, Cancer Res, Vol. 67, p6031-43.
- “hsa-miR-940” is known as “hsa-mir-940” (miRBase Accession No. MI0005762, SEQ ID NO: 352) having a hairpin-like structure as a precursor.
- hsa-miR-939-5p gene or “hsa-miR-939-5p” refers to the hsa-miR-939-5p gene described in SEQ ID NO: 76 (miRBase Accession No. MIMAT0004982) and other species homologues or orthologues are included.
- the hsa-miR-939-5p gene can be obtained by the method described in Lui WO et al., 2007, Cancer Res, 67, p6031-6043.
- hsa-miR-939-5p “hsa-mir-939” (miRBase Accession No. MI0005761, SEQ ID NO: 353) having a hairpin-like structure as a precursor is known.
- hsa-miR-6757-5p gene or “hsa-miR-6757-5p” refers to the hsa-miR-6757-5p gene described in SEQ ID NO: 77 (miRBase Accession No. MIMAT0027414) and other species homologues or orthologues are included.
- the hsa-miR-6757-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6757-5p” is known as “hsa-mir-6757” (miRBase Accession No. MI0022602, SEQ ID NO: 354) having a hairpin-like structure as a precursor.
- hsa-miR-1275 gene or “hsa-miR-1275” refers to the hsa-miR-1275 gene (miRBase Accession No. MIMAT0005929) described in SEQ ID NO: 78 and other biological species. Homologs or orthologs are included.
- the hsa-miR-1275 gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- “hsa-miR-1275” “hsa-mir-1275” (miRBase Accession No. MI0006415, SEQ ID NO: 355) having a hairpin-like structure as a precursor is known.
- hsa-miR-5001-5p gene or “hsa-miR-5001-5p” refers to the hsa-miR-5001-5p gene described in SEQ ID NO: 79 (miRBase Accession No. MIMAT0021021) and other species homologues or orthologues are included.
- the hsa-miR-5001-5p gene can be obtained by the method described in Hansen TB et al., 2011, RNA Biol, 8, p378-383.
- “hsa-miR-5001-5p” is known as “hsa-mir-5001” (miRBase Accession No. MI0017867, SEQ ID NO: 356) having a hairpin-like structure as a precursor.
- hsa-miR-6826-5p gene or “hsa-miR-6826-5p” refers to the hsa-miR-6826-5p gene described in SEQ ID NO: 80 (miRBase Accession No. MIMAT0027552) and other species homologues or orthologues are included.
- the hsa-miR-6826-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6826-5p “hsa-mir-6826” (miRBase Accession No. MI0022671, SEQ ID NO: 357) having a hairpin-like structure as a precursor is known.
- hsa-miR-6765-3p gene or “hsa-miR-6765-3p” refers to the hsa-miR-6765-3p gene described in SEQ ID NO: 81 (miRBase Accession No. MIMAT0027431) and other species homologues or orthologues are included.
- the hsa-miR-6765-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6765-3p “hsa-mir-6765” (miRBase Accession No. MI0022610, SEQ ID NO: 331) having a hairpin-like structure as a precursor is known.
- hsa-miR-3679-3p gene or “hsa-miR-3679-3p” refers to the hsa-miR-3679-3p gene described in SEQ ID NO: 82 (miRBase Accession No. MIMAT0018105) and other species homologues or orthologues are included.
- the hsa-miR-3679-3p gene can be obtained by the method described in Creighton CJ et al., 2010, PLoS One, 5, e9637.
- hsa-miR-3679-3p “hsa-mir-3679” (miRBase Accession No. MI0016080, SEQ ID NO: 358) having a hairpin-like structure as a precursor is known.
- hsa-miR-4718 gene or “hsa-miR-4718” refers to the hsa-miR-4718 gene (miRBase Accession No. MIMAT0019831) described in SEQ ID NO: 83 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4718 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-4718 “hsa-mir-4718” (miRBase Accession No. MI0017353, SEQ ID NO: 359) having a hairpin-like structure as a precursor is known.
- hsa-miR-4286 gene or “hsa-miR-4286” refers to the hsa-miR-4286 gene (miRBase Accession No. MIMAT0016916) described in SEQ ID NO: 84 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4286 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- hsa-miR-4286 “hsa-mir-4286” (miRBase Accession No. MI0015894, SEQ ID NO: 360) having a hairpin-like structure as a precursor is known.
- hsa-miR-8059 gene or “hsa-miR-8059” refers to the hsa-miR-8059 gene (miRBase Accession No. MIMAT0030986) described in SEQ ID NO: 85 or other biological species. Homologs or orthologs are included.
- the hsa-miR-8059 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, 39, p480-487.
- “hsa-miR-8059” is known as “hsa-mir-8059” (miRBase Accession No. MI0025895, SEQ ID NO: 361) having a hairpin-like structure as a precursor.
- hsa-miR-4447 gene or “hsa-miR-4447” refers to the hsa-miR-4447 gene (miRBase Accession No. MIMAT0018966) described in SEQ ID NO: 86 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4447 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-4447 “hsa-mir-4447” (miRBase Accession No. MI0016790, SEQ ID NO: 362) having a hairpin-like structure as a precursor is known.
- hsa-miR-4448 gene or “hsa-miR-4448” refers to the hsa-miR-4448 gene (miRBase Accession No. MIMAT0018967) described in SEQ ID NO: 87 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4448 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- “hsa-miR-4448” is known as “hsa-mir-4448” (miRBase Accession No. MI0016791, SEQ ID NO: 363) having a hairpin-like structure as a precursor.
- hsa-miR-658 gene or “hsa-miR-658” refers to the hsa-miR-658 gene (miRBase Accession No. MIMAT0003336) described in SEQ ID NO: 88 or other biological species. Homologs or orthologs are included.
- the hsa-miR-658 gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692.
- “hsa-miR-658” is known as “hsa-mir-658” (miRBase Accession No. MI0003682, SEQ ID NO: 364) having a hairpin-like structure as a precursor.
- hsa-miR-6766-3p gene or “hsa-miR-6766-3p” refers to the hsa-miR-6766-3p gene described in SEQ ID NO: 89 (miRBase Accession No. MIMAT0027433) and other species homologues or orthologues are included.
- the hsa-miR-6766-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6766-3p “hsa-mir-6766” (miRBase Accession No. MI0022611, SEQ ID NO: 365) having a hairpin-like structure as a precursor is known.
- hsa-miR-197-5p gene or “hsa-miR-197-5p” refers to the hsa-miR-197-5p gene described in SEQ ID NO: 90 (miRBase Accession No. MIMAT0022691) and other species homologs or orthologs are included.
- the hsa-miR-197-5p gene can be obtained by the method described in Lagos-Quintana M et al., 2003, RNA, Vol. 9, p175-179.
- hsa-mir-197 miRBase Accession No. MI0000239, SEQ ID NO: 366 having a hairpin-like structure as a precursor is known.
- hsa-miR-687-5p gene or “hsa-miR-6687-5p” refers to the hsa-miR-6887-5p gene (miRBase Accession No. MIMAT0027674) and other species homologues or orthologues are included.
- the hsa-miR-6687-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-687-5p” is known as “hsa-mir-6687” (miRBase Accession No. MI0022734, SEQ ID NO: 367) having a hairpin-like structure as a precursor.
- hsa-miR-6742-5p gene or “hsa-miR-6742-5p” refers to the hsa-miR-6742-5p gene described in SEQ ID NO: 92 (miRBase Accession No. MIMAT0027385) and other species homologues or orthologues are included.
- the hsa-miR-6742-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6742-5p” is known as “hsa-mir-6742” (miRBase Accession No. MI0022587, SEQ ID NO: 368) having a hairpin-like structure as a precursor.
- hsa-miR-6729-3p gene or “hsa-miR-6729-3p” refers to the hsa-miR-6729-3p gene described in SEQ ID NO: 93 (miRBase Accession No. MIMAT0027360) and other species homologs or orthologs.
- the hsa-miR-6729-3p gene can be obtained by the method described in Ladewig E, et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6729-3p “hsa-mir-6729” (miRBase Accession No. MI0022574, SEQ ID NO: 326) having a hairpin-like structure as a precursor is known.
- hsa-miR-5090 gene or “hsa-miR-5090” refers to the hsa-miR-5090 gene (miRBase Accession No. MIMAT0021082) described in SEQ ID NO: 94 or other biological species. Homologs or orthologs are included.
- the hsa-miR-5090 gene can be obtained by the method described in Ding N et al., 2011, J Radiat Res, 52, p425-432.
- “hsa-miR-5090” is known as “hsa-mir-5090” (miRBase Accession No. MI0017979, SEQ ID NO: 369) having a hairpin-like structure as a precursor.
- hsa-miR-7975 gene or “hsa-miR-7975” refers to the hsa-miR-7975 gene (miRBase Accession No. MIMAT0031178) described in SEQ ID NO: 95 or other biological species. Homologs or orthologs are included.
- the hsa-miR-7975 gene can be obtained by the method described in Velthut-Meikas A et al., 2013, Mol Endocrinol, 27, p1128-1141.
- hsa-miR-7975 “hsa-mir-7975” (miRBase Accession No. MI0025751, SEQ ID NO: 370) having a hairpin-like structure as a precursor is known.
- hsa-miR-4505 gene or “hsa-miR-4505” refers to the hsa-miR-4505 gene (miRBase Accession No. MIMAT0019041) described in SEQ ID NO: 96 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4505 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-4505 “hsa-mir-4505” (miRBase Accession No. MI0016868, SEQ ID NO: 371) having a hairpin-like structure as a precursor is known.
- hsa-miR-6889-5p gene or “hsa-miR-6889-5p” refers to the hsa-miR-6889-5p gene described in SEQ ID NO: 97 (miRBase Accession No. MIMAT0027678) and other species homologs or orthologs.
- the hsa-miR-6889-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6889-5p” is known as “hsa-mir-6889” (miRBase Accession No. MI0022736, SEQ ID NO: 372) having a hairpin-like structure as a precursor.
- hsa-miR-4708-3p gene or “hsa-miR-4708-3p” refers to the hsa-miR-4708-3p gene described in SEQ ID NO: 98 (miRBase Accession No. MIMAT0019810) and other species homologues or orthologues are included.
- the hsa-miR-4708-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4708-3p” is known as “hsa-mir-4708” (miRBase Accession No. MI0017341, SEQ ID NO: 373) having a hairpin-like structure as a precursor.
- hsa-miR-6131 gene or “hsa-miR-6131” refers to the hsa-miR-6131 gene (miRBase Accession No. MIMAT0024615) described in SEQ ID NO: 99 or other biological species. Homologs or orthologs are included.
- the hsa-miR-6131 gene can be obtained by the method described in Dannemann M et al., 2012, Genome Biol Evol, Vol. 4, p552-564.
- “hsa-miR-6131” is known as “hsa-mir-6131” (miRBase Accession No. MI0021276, SEQ ID NO: 374) having a hairpin-like structure as a precursor.
- hsa-miR-1225-3p gene or “hsa-miR-1225-3p” refers to the hsa-miR-1225-3p gene described in SEQ ID NO: 100 (miRBase Accession No. MIMAT0005573) and other species homologues or orthologues are included.
- the hsa-miR-1225-3p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, 28, p328-336.
- hsa-mir-1225 (miRBase Accession No. MI0006311, SEQ ID NO: 375) having a hairpin-like structure as a precursor is known.
- hsa-miR-6132 gene or “hsa-miR-6132” refers to the hsa-miR-6132 gene (miRBase Accession No. MIMAT0024616) described in SEQ ID NO: 101 or other biological species. Homologs or orthologs are included.
- the hsa-miR-6132 gene can be obtained by the method described in Dannemann M et al., 2012, Genome Biol Evol, Vol. 4, p552-564.
- “hsa-miR-6132” is known as “hsa-mir-6132” (miRBase Accession No. MI0021277, SEQ ID NO: 376) having a hairpin-like structure as a precursor.
- hsa-miR-4734 gene or “hsa-miR-4734” refers to the hsa-miR-4734 gene (miRBase Accession No. MIMAT0019859) described in SEQ ID NO: 102 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4734 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4734” is known as “hsa-mir-4734” (miRBase Accession No. MI0017371, SEQ ID NO: 377) having a hairpin-like structure as a precursor.
- hsa-miR-3194-3p gene or “hsa-miR-3194-3p” refers to the hsa-miR-3194-3p gene described in SEQ ID NO: 103 (miRBase Accession No. MIMAT0019218) and other species homologues or orthologues are included.
- the hsa-miR-3194-3p gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- “hsa-miR-3194-3p” is known as “hsa-mir-3194” (miRBase Accession No. MI0014239, SEQ ID NO: 378) having a hairpin-like structure as a precursor.
- hsa-miR-638 gene or “hsa-miR-638” refers to the hsa-miR-638 gene (miRBase Accession No. MIMAT0003308) described in SEQ ID NO: 104 or other biological species. A homolog or ortholog is included.
- the hsa-miR-638 gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692.
- “hsa-miR-638” is known as “hsa-mir-638” (miRBase Accession No. MI0003653, SEQ ID NO: 379) having a hairpin-like structure as a precursor.
- hsa-miR-2467-3p gene or “hsa-miR-2467-3p” refers to the hsa-miR-2467-3p gene described in SEQ ID NO: 105 (miRBase Accession No. MIMAT0019953) and other species homologs or orthologs.
- the hsa-miR-2467-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-2467-3p” is known as “hsa-mir-2467” (miRBase Accession No. MI0017432, SEQ ID NO: 380) having a hairpin-like structure as a precursor.
- hsa-miR-4728-5p gene or “hsa-miR-4728-5p” refers to the hsa-miR-4728-5p gene described in SEQ ID NO: 106 (miRBase Accession No. MIMAT0019849) and other species homologues or orthologues are included.
- the hsa-miR-4728-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4728-5p” “hsa-mir-4728” (miRBase Accession No. MI0017365, SEQ ID NO: 381) having a hairpin-like structure as a precursor is known.
- hsa-miR-5572 gene or “hsa-miR-5572” refers to the hsa-miR-5572 gene (miRBase Accession No. MIMAT0022260) described in SEQ ID NO: 107 or other biological species. A homolog or ortholog is included.
- the hsa-miR-5572 gene can be obtained by the method described in Tandon M, et al., 2012, Oral Dis, 18, p127-131.
- hsa-miR-5572 “hsa-mir-5572” (miRBase Accession No. MI0019117, SEQ ID NO: 382) having a hairpin-like structure as a precursor is known.
- hsa-miR-6789-5p gene or “hsa-miR-6789-5p” refers to the hsa-miR-6789-5p gene described in SEQ ID NO: 108 (miRBase Accession No. MIMAT0027478) and other species homologs or orthologs.
- the hsa-miR-6789-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6789-5p “hsa-mir-6789” (miRBase Accession No. MI0022634, SEQ ID NO: 383) having a hairpin-like structure as a precursor is known.
- hsa-miR-8063 gene or “hsa-miR-8063” refers to the hsa-miR-8063 gene (miRBase Accession No. MIMAT0030990) described in SEQ ID NO: 109 or other biological species. Homologs or orthologs are included.
- the hsa-miR-8063 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487.
- hsa-miR-8063 “hsa-mir-8063” (miRBase Accession No. MI0025899, SEQ ID NO: 384) having a hairpin-like structure as a precursor is known.
- hsa-miR-4429 gene or “hsa-miR-4429” refers to the hsa-miR-4429 gene (miRBase Accession No. MIMAT0018944) described in SEQ ID NO: 110 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4429 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-mir-4429 miRBase Accession No. MI0016768, SEQ ID NO: 385 having a hairpin-like structure as a precursor is known.
- hsa-miR-6840-3p gene or “hsa-miR-6840-3p” refers to the hsa-miR-6840-3p gene described in SEQ ID NO: 111 (miRBase Accession No. MIMAT0027583) and other species homologs or orthologs.
- the hsa-miR-6840-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-mir-6840 miRBase Accession No. MI0022686, SEQ ID NO: 386 having a hairpin-like structure as a precursor is known.
- hsa-miR-4476 gene or “hsa-miR-4476” refers to the hsa-miR-4476 gene (miRBase Accession No. MIMAT0019003) described in SEQ ID NO: 112 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4476 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-4476 “hsa-mir-4476” (miRBase Accession No. MI0016828, SEQ ID NO: 387) having a hairpin-like structure as a precursor is known.
- hsa-miR-675-5p gene or “hsa-miR-675-5p” refers to the hsa-miR-675-5p gene described in SEQ ID NO: 113 (miRBase Accession No. MIMAT0004284) and other species homologues or orthologues are included.
- the hsa-miR-675-5p gene can be obtained by the method described in Cai X et al., 2007, RNA, Vol. 13, p313-316.
- “hsa-miR-675-5p” is known as “hsa-mir-675” (miRBase Accession No. MI0005416, SEQ ID NO: 388) having a hairpin-like structure as a precursor.
- hsa-miR-711 gene or “hsa-miR-711” refers to the hsa-miR-711 gene (miRBase Accession No. MIMAT0012734) described in SEQ ID NO: 114 or other biological species. Homologs or orthologs are included.
- the hsa-miR-711 gene can be obtained by the method described in Artzi S et al., 2008, BMC Bioinformatics, Vol. 9, p39.
- “hsa-miR-711” is known as “hsa-mir-711” (miRBase Accession No. MI0012488, SEQ ID NO: 389) having a hairpin-like structure as a precursor.
- hsa-miR-6875-5p gene or “hsa-miR-6875-5p” refers to the hsa-miR-6875-5p gene described in SEQ ID NO: 115 (miRBase Accession No. MIMAT0027650) and other species homologues or orthologues are included.
- the hsa-miR-6875-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6875-5p “hsa-mir-6875” (miRBase Accession No. MI0022722, SEQ ID NO: 390) having a hairpin-like structure as a precursor is known.
- hsa-miR-3160-5p gene or “hsa-miR-3160-5p” refers to the hsa-miR-3160-5p gene described in SEQ ID NO: 116 (miRBase Accession No. MIMAT0019212) and other species homologues or orthologues are included.
- the hsa-miR-3160-5p gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- “Hsa-miR-3160-5p” has a hairpin-like structure as its precursor, “hsa-mir-3160-1, hsa-mir-3160-2” (miRBase Accession No. MI0014189, MI0014190, SEQ ID NO: 391, 392) are known.
- hsa-miR-1908-5p gene or “hsa-miR-1908-5p” refers to the hsa-miR-1908-5p gene described in SEQ ID NO: 117 (miRBase Accession No. MIMAT0007881) and other species homologues or orthologues are included.
- the hsa-miR-1908-5p gene can be obtained by the method described in Bar M et al., 2008, Stem Cells, Vol. 26, p2496-2505.
- hsa-miR-1908-5p “hsa-mir-1908” (miRBase Accession No. MI0008329, SEQ ID NO: 393) having a hairpin-like structure as a precursor is known.
- hsa-miR-6726-5p gene or “hsa-miR-6726-5p” refers to the hsa-miR-6726-5p gene described in SEQ ID NO: 118 (miRBase Accession No. MIMAT0027353) and other species homologues or orthologues are included.
- the hsa-miR-6726-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6726-5p” is known as “hsa-mir-6726” (miRBase Accession No. MI0022571, SEQ ID NO: 394) having a hairpin-like structure as a precursor.
- hsa-miR-1913 gene or “hsa-miR-1913” refers to the hsa-miR-1913 gene (miRBase Accession No. MIMAT0007888) described in SEQ ID NO: 119 or other biological species. Homologs or orthologs are included.
- the hsa-miR-1913 gene can be obtained by the method described in Bar M et al., 2008, Stem Cells, 26, p2496-2505.
- “hsa-miR-1913” is known as “hsa-mir-1913” (miRBase Accession No. MI0008334, SEQ ID NO: 395) having a hairpin-like structure as a precursor.
- hsa-miR-8071 gene or “hsa-miR-8071” refers to the hsa-miR-8071 gene (miRBase Accession No. MIMAT0030998) described in SEQ ID NO: 120 or other biological species. Homologs or orthologs are included.
- the hsa-miR-8071 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487.
- “Hsa-miR-8071” has a hairpin-like structure as its precursor, “hsa-mir-8071-1, hsa-mir-8071-2” (miRBase Accession No. MI0025907, MI0026417, SEQ ID NO: 396, 397) is known.
- hsa-miR-3648 gene or “hsa-miR-3648” refers to the hsa-miR-3648 gene (miRBase Accession No. MIMAT0018068) described in SEQ ID NO: 121 and other biological species. Homologs or orthologs are included.
- the hsa-miR-3648 gene can be obtained by the method described in Meiri E et al., 2010, Nucleic Acids Res, 38, p6234-6246.
- “Hsa-miR-3648” has a hairpin-like structure as its precursor, “hsa-mir-3648-1, hsa-mir-3648-2” (miRBase Accession No. MI0016048, MI0031512, SEQ ID NO: 398, 399) is known.
- hsa-miR-4732-5p gene or “hsa-miR-4732-5p” refers to the hsa-miR-4732-5p gene (miRBase Accession No. MIMAT0019855) and other species homologs or orthologs.
- the hsa-miR-4732-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “Hsa-miR-4732-5p” is known as “hsa-mir-4732” (miRBase Accession No. MI0017369, SEQ ID NO: 400) having a hairpin-like structure as a precursor.
- hsa-miR-4787-5p gene or “hsa-miR-4787-5p” refers to the hsa-miR-4787-5p gene described in SEQ ID NO: 123 (miRBase Accession No. MIMAT0019956) and other species homologues or orthologues are included.
- the hsa-miR-4787-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4787-5p” “hsa-mir-4787” (miRBase Accession No. MI0017434, SEQ ID NO: 318) having a hairpin-like structure as a precursor is known.
- hsa-miR-3717 gene or “hsa-miR-3717” refers to the hsa-miR-3919 gene (miRBase Accession No. MIMAT0018191) described in SEQ ID NO: 124 or other biological species. Homologs or orthologs are included.
- the hsa-miR-3913 gene can be obtained by the method described in Creighton CJ et al., 2010, PLoS One, 5, e9637.
- hsa-miR-3917 is known as “hsa-mir-3917” (miRBase Accession No. MI0016423, SEQ ID NO: 401) having a hairpin-like structure as a precursor.
- hsa-miR-619-5p gene or “hsa-miR-619-5p” refers to the hsa-miR-619-5p gene (miRBase Accession No. MIMAT0026622) and other species homologues or orthologues are included.
- the hsa-miR-619-5p gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, 103, p3687-3692.
- “hsa-miR-619-5p” is known as “hsa-mir-619” (miRBase Accession No. MI0003633, SEQ ID NO: 402) having a hairpin-like structure as a precursor.
- hsa-miR-1231 gene or “hsa-miR-1231” refers to the hsa-miR-1231 gene (miRBase Accession No. MIMAT0005586) described in SEQ ID NO: 126 or other biological species. Homologs or orthologs are included.
- the hsa-miR-1231 gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, 28, p328-336.
- hsa-miR-1231 is known as “hsa-mir-1231” (miRBase Accession No. MI0006321, SEQ ID NO: 403) having a hairpin-like structure as a precursor.
- hsa-miR-342-5p gene or “hsa-miR-342-5p” refers to the hsa-miR-342-5p gene described in SEQ ID NO: 127 (miRBase Accession No. MIMAT0004694) and other species homologues or orthologues are included.
- the hsa-miR-342-5p gene can be obtained by the method described in Kim J et al., 2004, Proc Natl Acad Sci USA, Vol. 101, p360-365.
- “hsa-miR-342-5p” “hsa-mir-342” (miRBase Accession No. MI000005, SEQ ID NO: 404) having a hairpin-like structure as a precursor is known.
- hsa-miR-4433a-5p gene or “hsa-miR-4433a-5p” refers to the hsa-miR-4433a-5p gene described in SEQ ID NO: 128 (miRBase Accession No. MIMAT0020956) and other species homologues or orthologues are included.
- the hsa-miR-4433a-5p gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- “hsa-miR-4433a-5p” is known as “hsa-mir-4433a” (miRBase Accession No. MI0016773, SEQ ID NO: 405) having a hairpin-like structure as a precursor.
- hsa-miR-6766-5p gene or “hsa-miR-6766-5p” refers to the hsa-miR-6766-5p gene described in SEQ ID NO: 129 (miRBase Accession No. MIMAT0027432) and other species homologues or orthologues are included.
- the hsa-miR-6766-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6766-5p “hsa-mir-6766” (miRBase Accession No. MI0022611, SEQ ID NO: 365) having a hairpin-like structure as a precursor is known.
- hsa-miR-4707-5p gene or “hsa-miR-4707-5p” refers to the hsa-miR-4707-5p gene described in SEQ ID NO: 130 (miRBase Accession No. MIMAT0019807) and other species homologues or orthologues are included.
- the hsa-miR-4707-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4707-5p” “hsa-mir-4707” (miRBase Accession No. MI0017340, SEQ ID NO: 406) having a hairpin-like structure as a precursor is known.
- hsa-miR-7114-5p gene or “hsa-miR-7114-5p” refers to the hsa-miR-7114-5p gene described in SEQ ID NO: 131 (miRBase Accession No. MIMAT0028125) and other species homologues or orthologues are included.
- the hsa-miR-7114-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-7114-5p “hsa-mir-7114” (miRBase Accession No. MI0022965, SEQ ID NO: 407) having a hairpin-like structure as a precursor is known.
- hsa-miR-6872-3p gene or “hsa-miR-6872-3p” refers to the hsa-miR-6872-3p gene described in SEQ ID NO: 132 (miRBase Accession No. MIMAT0027645) and other species homologs or orthologs.
- the hsa-miR-6872-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “Hsa-miR-6872-3p” is known as “hsa-mir-6872” (miRBase Accession No. MI0022719, SEQ ID NO: 408) having a hairpin-like structure as a precursor.
- hsa-miR-6780b-5p gene or “hsa-miR-6780b-5p” refers to the hsa-miR-6780b-5p gene described in SEQ ID NO: 133 (miRBase Accession No. MIMAT0027572) and other species homologues or orthologues are included.
- the hsa-miR-6780b-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6780b-5p” is known as “hsa-mir-6780b” (miRBase Accession No. MI0022681, SEQ ID NO: 409) having a hairpin-like structure as a precursor.
- hsa-miR-7845-5p gene or “hsa-miR-7845-5p” refers to the hsa-miR-7845-5p gene described in SEQ ID NO: 134 (miRBase Accession No. MIMAT0030420) and other species homologs or orthologs.
- the hsa-miR-7845-5p gene can be obtained by the method described in Ple H et al., 2012, PLoS One, Vol. 7, e50746.
- hsa-miR-7845-5p “hsa-mir-7845” (miRBase Accession No. MI0025515, SEQ ID NO: 410) having a hairpin-like structure as a precursor is known.
- hsa-miR-6798-3p gene or “hsa-miR-6798-3p” refers to the hsa-miR-6798-3p gene described in SEQ ID NO: 135 (miRBase Accession No. MIMAT0027497) and other species homologues or orthologues are included.
- the hsa-miR-6798-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6798-3p “hsa-mir-6798” (miRBase Accession No. MI0022643, SEQ ID NO: 314) having a hairpin-like structure as a precursor is known.
- hsa-miR-665 gene or “hsa-miR-665” refers to the hsa-miR-665 gene (miRBase Accession No. MIMAT0004952) described in SEQ ID NO: 136 and other biological species. Homologs or orthologs are included.
- the hsa-miR-665 gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, 16, p1299-1298.
- “hsa-miR-665” is known as “hsa-mir-665” (miRBase Accession No. MI0005563, SEQ ID NO: 411) having a hairpin-like structure as a precursor.
- hsa-miR-6848-5p gene or “hsa-miR-6848-5p” refers to the hsa-miR-6848-5p gene described in SEQ ID NO: 137 (miRBase Accession No. MIMAT0027596) and other species homologs or orthologs.
- the hsa-miR-6848-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6848-5p “hsa-mir-6848” (miRBase Accession No. MI0022694, SEQ ID NO: 412) having a hairpin-like structure as a precursor is known.
- hsa-miR-5008-5p gene or “hsa-miR-5008-5p” refers to the hsa-miR-5008-5p gene described in SEQ ID NO: 138 (miRBase Accession No. MIMAT0021039) and other species homologs or orthologs.
- the hsa-miR-5008-5p gene can be obtained by the method described in Hansen TB et al., 2011, RNA Biol, Vol. 8, p378-383.
- hsa-miR-5008-5p “hsa-mir-5008” (miRBase Accession No. MI0017876, SEQ ID NO: 413) having a hairpin-like structure as a precursor is known.
- hsa-miR-4294 gene or “hsa-miR-4294” refers to the hsa-miR-4294 gene (miRBase Accession No. MIMAT0016849) described in SEQ ID NO: 139 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4294 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- hsa-miR-4294 “hsa-mir-4294” (miRBase Accession No. MI0015827, SEQ ID NO: 414) having a hairpin-like structure as a precursor is known.
- hsa-miR-6511a-5p gene or “hsa-miR-6511a-5p” refers to the hsa-miR-6511a-5p gene described in SEQ ID NO: 140 (miRBase Accession No. MIMAT0025478) and other species homologues or orthologues are included.
- the hsa-miR-6511a-5p gene can be obtained by the method described in Joyce CE et al., 2011, Hum Mol Genet, 20, p4025-4040.
- hsa-miR-6511a-5p has a hairpin-like structure as its precursor, “hsa-mir-6651a-1, hsa-mir-6511a-2, hsa-mir-6651a-3, hsa-mir”.
- -6511a-4 "(miRBase Accession No. MI0022223, MI0023564, MI0023565, MI0023566, SEQ ID NO: 415, 416, 417, 418) is known.
- hsa-miR-4435 gene or “hsa-miR-4435” refers to the hsa-miR-4435 gene (miRBase Accession No. MIMAT0018951) described in SEQ ID NO: 141 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4435 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-4435 has a hairpin-like structure as its precursor, “hsa-mir-4435-1, hsa-mir-4435-2” (miRBase Accession No. MI0016775, MI0016777, SEQ ID NO: 419, 420) is known.
- hsa-miR-4747-3p gene or “hsa-miR-4747-3p” refers to the hsa-miR-4747-3p gene described in SEQ ID NO: 142 (miRBase Accession No. MIMAT0019883) and other species homologs or orthologs.
- the hsa-miR-4747-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4747-3p” “hsa-mir-4747” (miRBase Accession No. MI0017386, SEQ ID NO: 421) having a hairpin-like structure as a precursor is known.
- hsa-miR-6880-3p gene or “hsa-miR-6880-3p” refers to the hsa-miR-6880-3p gene described in SEQ ID NO: 143 (miRBase Accession No. MIMAT0027661) and other species homologues or orthologues are included.
- the hsa-miR-6880-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6880-3p “hsa-mir-6880” (miRBase Accession No. MI0022727, SEQ ID NO: 422) having a hairpin-like structure as a precursor is known.
- hsa-miR-6869-5p gene or “hsa-miR-6869-5p” refers to the hsa-miR-6869-5p gene described in SEQ ID NO: 144 (miRBase Accession No. MIMAT0027638) and other species homologs or orthologs.
- the hsa-miR-6869-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6869-5p “hsa-mir-6869” (miRBase Accession No. MI0022716, SEQ ID NO: 423) having a hairpin-like structure as a precursor is known.
- hsa-miR-7150 gene or “hsa-miR-7150” refers to the hsa-miR-7150 gene (miRBase Accession No. MIMAT0028211) described in SEQ ID NO: 145 or other biological species. Homologs or orthologs are included.
- the hsa-miR-7150 gene can be obtained by the method described in Oulas A et al., 2009, Nucleic Acids Res, 37, p3276-3287.
- hsa-miR-7150 “hsa-mir-7150” (miRBase Accession No. MI0023610, SEQ ID NO: 424) having a hairpin-like structure as a precursor is known.
- hsa-miR-1260a gene or “hsa-miR-1260a” refers to the hsa-miR-1260a gene (miRBase Accession No. MIMAT0005911) described in SEQ ID NO: 146 or other biological species. A homolog or ortholog is included.
- the hsa-miR-1260a gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- “hsa-miR-1260a” is known as “hsa-mir-1260a” (miRBase Accession No. MI0006394, SEQ ID NO: 425), which has a hairpin-like structure as a precursor.
- hsa-miR-6877-5p gene or “hsa-miR-6877-5p” refers to the hsa-miR-6877-5p gene described in SEQ ID NO: 147 (miRBase Accession No. MIMAT0027654) and other species homologs or orthologs.
- the hsa-miR-6877-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6877-5p” is known as “hsa-mir-6877” (miRBase Accession No. MI0022724, SEQ ID NO: 426) having a hairpin-like structure as a precursor.
- hsa-miR-6721-5p gene or “hsa-miR-6721-5p” refers to the hsa-miR-6721-5p gene described in SEQ ID NO: 148 (miRBase Accession No. MIMAT0025852) and other species homologs or orthologs.
- the hsa-miR-6721-5p gene can be obtained by the method described in Li Y et al., 2012, Gene, 497, p330-335.
- hsa-miR-6721-5p “hsa-mir-6721” (miRBase Accession No. MI0022556, SEQ ID NO: 427) having a hairpin-like structure as a precursor is known.
- hsa-miR-4656 gene or “hsa-miR-4656” refers to the hsa-miR-4656 gene (miRBase Accession No. MIMAT0019723) described in SEQ ID NO: 149 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4656 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4656” is known as “hsa-mir-4656” (miRBase Accession No. MI0017284, SEQ ID NO: 428) having a hairpin-like structure as a precursor.
- hsa-miR-1229-5p gene or “hsa-miR-1229-5p” refers to the hsa-miR-1229-5p gene (miRBase Accession No. MIMAT0022942) and other species homologs or orthologs are included.
- the hsa-miR-1229-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, 28, p328-336.
- hsa-mir-1229 (miRBase Accession No. MI0006319, SEQ ID NO: 429) having a hairpin-like structure as a precursor is known.
- hsa-miR-4433a-3p gene or “hsa-miR-4433a-3p” refers to the hsa-miR-4433a-3p gene described in SEQ ID NO: 151 (miRBase Accession No. MIMAT0018949) and other species homologues or orthologues are included.
- the hsa-miR-4433a-3p gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- “hsa-miR-4433a-3p” “hsa-mir-4433a” (miRBase Accession No. MI0016773, SEQ ID NO: 405) having a hairpin-like structure as a precursor is known.
- hsa-miR-4274 gene or “hsa-miR-4274” refers to the hsa-miR-4274 gene (miRBase Accession No. MIMAT0016906) described in SEQ ID NO: 152 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4274 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- hsa-miR-4274 “hsa-mir-4274” (miRBase Accession No. MI0015884, SEQ ID NO: 430) having a hairpin-like structure as a precursor is known.
- hsa-miR-4419b gene or “hsa-miR-4419b” refers to the hsa-miR-4419b gene (miRBase Accession No. MIMAT0019034) described in SEQ ID NO: 153 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4419b gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-4419b “hsa-mir-4419b” (miRBase Accession No. MI0016861, SEQ ID NO: 431) having a hairpin-like structure as a precursor is known.
- hsa-miR-4675 gene or “hsa-miR-4675” refers to the hsa-miR-4675 gene (miRBase Accession No. MIMAT0019756) described in SEQ ID NO: 154 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4675 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-4675 “hsa-mir-4675” (miRBase Accession No. MI0017305, SEQ ID NO: 432) having a hairpin-like structure as a precursor is known.
- hsa-miR-6893-5p gene or “hsa-miR-6893-5p” refers to the hsa-miR-6893-5p gene described in SEQ ID NO: 155 (miRBase Accession No. MIMAT0027686) and other species homologues or orthologues are included.
- the hsa-miR-6893-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6893-5p “hsa-mir-6893” (miRBase Accession No. MI0022740, SEQ ID NO: 433) having a hairpin-like structure as a precursor is known.
- hsa-miR-6663-3p gene or “hsa-miR-6663-3p” refers to the hsa-miR-67663-3p gene (miRBase Accession No. MIMAT0027427) and other species homologues or orthologues are included.
- the hsa-miR-6673-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “Hsa-miR-6663-3p” is known as “hsa-mir-6673” (miRBase Accession No. MI0022608, SEQ ID NO: 434) having a hairpin-like structure as a precursor.
- hsa-miR-6762-5p gene or “hsa-miR-6762-5p” refers to the hsa-miR-6762-5p gene described in SEQ ID NO: 157 (miRBase Accession No. MIMAT0027424) and other species homologs or orthologs are included.
- the hsa-miR-6762-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6762-5p “hsa-mir-6762” (miRBase Accession No. MI0022607, SEQ ID NO: 435) having a hairpin-like structure as a precursor is known.
- hsa-miR-6738-5p gene or “hsa-miR-6738-5p” refers to the hsa-miR-6738-5p gene described in SEQ ID NO: 158 (miRBase Accession No. MIMAT0027377) and other species homologues or orthologues are included.
- the hsa-miR-6738-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6738-5p” is known as “hsa-mir-6738” (miRBase Accession No. MI0022583, SEQ ID NO: 436) having a hairpin-like structure as a precursor.
- hsa-miR-4513 gene or “hsa-miR-4513” refers to the hsa-miR-4513 gene (miRBase Accession No. MIMAT0019050) described in SEQ ID NO: 159 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4513 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-4513 “hsa-mir-4513” (miRBase Accession No. MI0016879, SEQ ID NO: 437) having a hairpin-like structure as a precursor is known.
- hsa-miR-6746-5p gene or “hsa-miR-6746-5p” refers to the hsa-miR-6746-5p gene described in SEQ ID NO: 160 (miRBase Accession No. MIMAT0027392) and other species homologues or orthologues are included.
- the hsa-miR-6746-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6746-5p” is known as “hsa-mir-6746” (miRBase Accession No. MI0022591, SEQ ID NO: 438) having a hairpin-like structure as a precursor.
- hsa-miR-6880-5p gene or “hsa-miR-6880-5p” refers to the hsa-miR-6880-5p gene described in SEQ ID NO: 161 (miRBase Accession No. MIMAT0027660) and other species homologs or orthologs.
- the hsa-miR-6880-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6880-5p “hsa-mir-6880” (miRBase Accession No. MI0022727, SEQ ID NO: 422) having a hairpin-like structure as a precursor is known.
- hsa-miR-4736 gene or “hsa-miR-4736” refers to the hsa-miR-4736 gene (miRBase Accession No. MIMAT0019862) described in SEQ ID NO: 162 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4736 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-4736 “hsa-mir-4736” (miRBase Accession No. MI0017373, SEQ ID NO: 439) having a hairpin-like structure as a precursor is known.
- hsa-miR-718 gene or “hsa-miR-718” refers to the hsa-miR-718 gene (miRBase Accession No. MIMAT0012735) described in SEQ ID NO: 163 or other biological species. Homologs or orthologs are included.
- the hsa-miR-718 gene can be obtained by the method described in Artzi S et al., 2008, BMC Bioinformatics, Vol. 9, p39.
- hsa-miR-718 “hsa-mir-718” (miRBase Accession No. MI0012489, SEQ ID NO: 440) having a hairpin-like structure as a precursor is known.
- hsa-miR-6717-5p gene or “hsa-miR-6717-5p” refers to the hsa-miR-6717-5p gene (miRBase Accession No. MIMAT00258446) and other species homologs or orthologs.
- the hsa-miR-6717-5p gene can be obtained by the method described in Li Y et al., 2012, Gene, 497, p330-335.
- hsa-miR-6717-5p “hsa-mir-6717” (miRBase Accession No. MI0022551, SEQ ID NO: 441) having a hairpin-like structure as a precursor is known.
- hsa-miR-7847-3p gene or “hsa-miR-7847-3p” refers to the hsa-miR-7847-3p gene described in SEQ ID NO: 165 (miRBase Accession No. MIMAT0030422) and other species homologues or orthologues are included.
- the hsa-miR-7847-3p gene can be obtained by the method described in Ple H et al., 2012, PLoS One, Vol. 7, e50746.
- hsa-miR-7847-3p “hsa-mir-7847” (miRBase Accession No. MI0025517, SEQ ID NO: 442) having a hairpin-like structure as a precursor is known.
- hsa-miR-760 gene or “hsa-miR-760” refers to the hsa-miR-760 gene (miRBase Accession No. MIMAT0004957) described in SEQ ID NO: 166 or other biological species. A homolog or ortholog is included.
- the hsa-miR-760 gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, 16, p1299-1298.
- “hsa-miR-760” is known as “hsa-mir-760” (miRBase Accession No. MI0005567, SEQ ID NO: 443) having a hairpin-like structure as a precursor.
- hsa-miR-1199-5p gene or “hsa-miR-1199-5p” refers to the hsa-miR-1199-5p gene described in SEQ ID NO: 167 (miRBase Accession No. MIMAT0031119) and other species homologues or orthologues are included.
- the hsa-miR-1199-5p gene can be obtained by the method described in Salvi A et al., 2013, Int J Oncol, 42, p391-402.
- hsa-miR-1199-5p “hsa-mir-1199” (miRBase Accession No. MI0020340, SEQ ID NO: 444) having a hairpin-like structure as a precursor is known.
- hsa-miR-6813-5p gene or “hsa-miR-6813-5p” refers to the hsa-miR-6813-5p gene described in SEQ ID NO: 168 (miRBase Accession No. MIMAT0027526) and other species homologues or orthologues are included.
- the hsa-miR-6813-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6813-5p” is known as “hsa-mir-6683” (miRBase Accession No. MI0022658, SEQ ID NO: 445) having a hairpin-like structure as a precursor.
- hsa-miR-6769a-5p gene or “hsa-miR-6769a-5p” refers to the hsa-miR-6769a-5p gene described in SEQ ID NO: 169 (miRBase Accession No. MIMAT0027438) and other species homologs or orthologs.
- the hsa-miR-6769a-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6769a-5p” is known as “hsa-mir-6769a” (miRBase Accession No. MI0022614, SEQ ID NO: 446) having a hairpin-like structure as a precursor.
- hsa-miR-1193 gene or “hsa-miR-1193” refers to the hsa-miR-1193 gene (miRBase Accession No. MIMAT0015049) described in SEQ ID NO: 170 or other biological species. Homologs or orthologs are included.
- the hsa-miR-1193 gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- “hsa-miR-1193” is known as “hsa-mir-1193” (miRBase Accession No. MI0014205, SEQ ID NO: 447) having a hairpin-like structure as a precursor.
- hsa-miR-7108-3p gene or “hsa-miR-7108-3p” refers to the hsa-miR-7108-3p gene described in SEQ ID NO: 171 (miRBase Accession No. MIMAT0028114) and other species homologs or orthologs.
- the hsa-miR-7108-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-7108-3p” is known as “hsa-mir-7108” (miRBase Accession No. MI0022959, SEQ ID NO: 448) having a hairpin-like structure as a precursor.
- hsa-miR-6741-5p gene or “hsa-miR-6741-5p” refers to the hsa-miR-6741-5p gene described in SEQ ID NO: 172 (miRBase Accession No. MIMAT0027383) and other species homologs or orthologs are included.
- the hsa-miR-6741-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6741-5p “hsa-mir-6741” (miRBase Accession No. MI0022586, SEQ ID NO: 449) having a hairpin-like structure as a precursor is known.
- hsa-miR-4298 gene or “hsa-miR-4298” refers to the hsa-miR-4298 gene (miRBase Accession No. MIMAT0016852) described in SEQ ID NO: 173 and other biological species. Homologs or orthologs are included.
- the hsa-miR-4298 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- hsa-miR-4298 “hsa-mir-4298” (miRBase Accession No. MI0015830, SEQ ID NO: 450) having a hairpin-like structure as a precursor is known.
- hsa-miR-6696-3p gene or “hsa-miR-6696-3p” refers to the hsa-miR-6696-3p gene described in SEQ ID NO: 174 (miRBase Accession No. MIMAT0027493) and other species homologues or orthologues are included.
- the hsa-miR-6969-3p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6696-3p” is known as “hsa-mir-6696” (miRBase Accession No. MI0022641, SEQ ID NO: 451) having a hairpin-like structure as a precursor.
- hsa-miR-4750-5p gene or “hsa-miR-4750-5p” refers to the hsa-miR-4750-5p gene described in SEQ ID NO: 175 (miRBase Accession No. MIMAT0019887) and other species homologues or orthologues are included.
- the hsa-miR-4750-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-4750-5p “hsa-mir-4750” (miRBase Accession No. MI0017389, SEQ ID NO: 452) having a hairpin-like structure as a precursor is known.
- hsa-miR-6785-5p gene or “hsa-miR-6785-5p” refers to the hsa-miR-6785-5p gene described in SEQ ID NO: 176 (miRBase Accession No. MIMAT0027470) and other species homologues or orthologues are included.
- the hsa-miR-6785-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6785-5p “hsa-mir-6785” (miRBase Accession No. MI0022630, SEQ ID NO: 453) having a hairpin-like structure as a precursor is known.
- hsa-miR-1292-3p gene or “hsa-miR-1292-3p” refers to the hsa-miR-1292-3p gene described in SEQ ID NO: 177 (miRBase Accession No. MIMAT0022948) and other species homologs or orthologs.
- the hsa-miR-1292-3p gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- “hsa-miR-1292-3p” is known as “hsa-mir-1292” (miRBase Accession No. MI0006433, SEQ ID NO: 454) having a hairpin-like structure as a precursor.
- hsa-miR-4749-3p gene or “hsa-miR-4749-3p” refers to the hsa-miR-4749-3p gene described in SEQ ID NO: 178 (miRBase Accession No. MIMAT0019886) and other species homologues or orthologues are included.
- the hsa-miR-4749-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4749-3p” is known as “hsa-mir-4749” (miRBase Accession No. MI0017388, SEQ ID NO: 455) having a hairpin-like structure as a precursor.
- hsa-miR-6800-3p gene or “hsa-miR-6800-3p” refers to the hsa-miR-6800-3p gene described in SEQ ID NO: 179 (miRBase Accession No. MIMAT0027501) and other species homologues or orthologues are included.
- the hsa-miR-6800-3p gene can be obtained by the method described in Ladewig E, et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6800-3p” is known as “hsa-mir-6800” (miRBase Accession No. MI0022645, SEQ ID NO: 293) having a hairpin-like structure as a precursor.
- hsa-miR-4722-5p gene or “hsa-miR-4722-5p” refers to the hsa-miR-4722-5p gene described in SEQ ID NO: 180 (miRBase Accession No. MIMAT0019836) and other species homologs or orthologs are included.
- the hsa-miR-4722-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4722-5p” “hsa-mir-4722” (miRBase Accession No. MI0017357, SEQ ID NO: 456) having a hairpin-like structure as a precursor is known.
- hsa-miR-4746-3p gene or “hsa-miR-4746-3p” refers to the hsa-miR-4746-3p gene described in SEQ ID NO: 181 (miRBase Accession No. MIMAT0019881) and other species homologues or orthologues are included.
- the hsa-miR-4746-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4746-3p” “hsa-mir-4746” (miRBase Accession No. MI0017385, SEQ ID NO: 457) having a hairpin-like structure as a precursor is known.
- hsa-miR-4450 gene or “hsa-miR-4450” refers to the hsa-miR-4450 gene (miRBase Accession No. MIMAT0018971) described in SEQ ID NO: 182 or other species. Homologs or orthologs are included.
- the hsa-miR-4450 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-4450 is known as “hsa-mir-4450” (miRBase Accession No. MI0016795, SEQ ID NO: 458) having a hairpin-like structure as a precursor.
- hsa-miR-6695-5p gene or “hsa-miR-6695-5p” refers to the hsa-miR-6695-5p gene described in SEQ ID NO: 183 (miRBase Accession No. MIMAT0027490) and other species homologs or orthologs.
- the hsa-miR-6695-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6695-5p “hsa-mir-6695” (miRBase Accession No. MI0022640, SEQ ID NO: 459) having a hairpin-like structure as a precursor is known.
- hsa-miR-365a-5p gene or “hsa-miR-365a-5p” refers to the hsa-miR-365a-5p gene described in SEQ ID NO: 184 (miRBase Accession No. MIMAT0009199) and other species homologues or orthologues are included.
- the hsa-miR-365a-5p gene can be obtained by the method described in Xie X et al., 2005, Nature, 434, p338-345.
- hsa-mir-365a miRBase Accession No. MI000067, SEQ ID NO: 460 having a hairpin-like structure as a precursor is known.
- hsa-miR-498 gene or “hsa-miR-498” refers to the hsa-miR-498 gene (miRBase Accession No. MIMAT0002824) described in SEQ ID NO: 185 or other biological species. Homologs or orthologs are included.
- the hsa-miR-498 gene can be obtained by the method described in Bentwich I et al., 2005, Nat Genet, 37, p766-770.
- hsa-miR-498 “hsa-mir-498” (miRBase Accession No. MI0003142, SEQ ID NO: 461) having a hairpin-like structure as a precursor is known.
- hsa-miR-6797-5p gene or “hsa-miR-6797-5p” refers to the hsa-miR-6797-5p gene described in SEQ ID NO: 186 (miRBase Accession No. MIMAT0027494) and other species homologues or orthologues are included.
- the hsa-miR-6797-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6797-5p” is known as “hsa-mir-6797” (miRBase Accession No. MI0022642, SEQ ID NO: 462) having a hairpin-like structure as a precursor.
- hsa-miR-1470 gene or “hsa-miR-1470” refers to the hsa-miR-1470 gene (miRBase Accession No. MIMAT0007348) described in SEQ ID NO: 187 or other biological species. A homolog or ortholog is included.
- the hsa-miR-1470 gene can be obtained by the method described in Kawaji H et al., 2008, BMC Genomics, Vol. 9, p157.
- hsa-miR-1470 “hsa-mir-1470” (miRBase Accession No. MI00000075, SEQ ID NO: 463) having a hairpin-like structure as a precursor is known.
- hsa-miR-6851-5p gene or “hsa-miR-6851-5p” refers to the hsa-miR-6851-5p gene described in SEQ ID NO: 188 (miRBase Accession No. MIMAT0027602) and other species homologues or orthologues are included.
- the hsa-miR-6651-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6851-5p “hsa-mir-6851” (miRBase Accession No. MI0022697, SEQ ID NO: 464) having a hairpin-like structure as a precursor is known.
- hsa-miR-1247-3p gene or “hsa-miR-1247-3p” refers to the hsa-miR-1247-3p gene (miRBase Accession No. MIMAT0022721) and other species homologues or orthologues are included.
- the hsa-miR-1247-3p gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- “hsa-miR-1247-3p” is known as “hsa-mir-1247” (miRBase Accession No. MI0006382, SEQ ID NO: 465) having a hairpin-like structure as a precursor.
- hsa-miR-5196-5p gene or “hsa-miR-5196-5p” refers to the hsa-miR-5196-5p gene described in SEQ ID NO: 190 (miRBase Accession No. MIMAT0021128) and other species homologs or orthologs.
- the hsa-miR-5196-5p gene can be obtained by the method described in Schotte D, et al., 2011, Leukemia, 25, p1389-1399.
- hsa-miR-5196-5p “hsa-mir-5196” (miRBase Accession No. MI0018175, SEQ ID NO: 466) having a hairpin-like structure as a precursor is known.
- hsa-miR-208a-5p gene or “hsa-miR-208a-5p” refers to the hsa-miR-208a-5p gene described in SEQ ID NO: 191 (miRBase Accession No. MIMAT0026474) and other species homologues or orthologues are included.
- the hsa-miR-208a-5p gene can be obtained by the method described in Lagos-Quintana M et al., 2003, RNA, Vol. 9, p175-179.
- “hsa-miR-208a-5p” “hsa-mir-208a” (miRBase Accession No. MI000000251, SEQ ID NO: 467) having a hairpin-like structure as a precursor is known.
- hsa-miR-6842-5p gene or “hsa-miR-6842-5p” refers to the hsa-miR-6842-5p gene described in SEQ ID NO: 192 (miRBase Accession No. MIMAT0027586) and other species homologues or orthologues are included.
- the hsa-miR-6842-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6842-5p “hsa-mir-6842” (miRBase Accession No. MI0022688, SEQ ID NO: 468) having a hairpin-like structure as a precursor is known.
- hsa-miR-150-3p gene or “hsa-miR-150-3p” refers to the hsa-miR-150-3p gene described in SEQ ID NO: 193 (miRBase Accession No. MIMAT0004610) and other species homologues or orthologues are included.
- the hsa-miR-150-3p gene can be obtained by the method described in Lagos-Quintana M et al., 2002, Curr Biol, Vol. 12, p735-739.
- “hsa-miR-150-3p” is known as “hsa-mir-150” (miRBase Accession No. MI0000479, SEQ ID NO: 469) having a hairpin-like structure as a precursor.
- hsa-miR-4534 gene or “hsa-miR-4534” refers to the hsa-miR-4534 gene (miRBase Accession No. MIMAT0019073) described in SEQ ID NO: 194 or other biological species. A homolog or ortholog is included.
- the hsa-miR-4534 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- “hsa-miR-4534” is known as “hsa-mir-4534” (miRBase Accession No. MI0016901, SEQ ID NO: 470) having a hairpin-like structure as a precursor.
- hsa-miR-3135b gene or “hsa-miR-3135b” refers to the hsa-miR-3135b gene (miRBase Accession No. MIMAT0018985) described in SEQ ID NO: 195 or other biological species. Homologs or orthologs are included.
- the hsa-miR-3135b gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-3135b is known as “hsa-mir-3135b” (miRBase Accession No. MI0016809, SEQ ID NO: 471) having a hairpin-like structure as a precursor.
- hsa-miR-3131 gene or “hsa-miR-3131” refers to the hsa-miR-3131 gene (miRBase Accession No. MIMAT0014996) described in SEQ ID NO: 196 or other species. Homologs or orthologs are included.
- the hsa-miR-3131 gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- “hsa-miR-3131” is known as “hsa-mir-3131” (miRBase Accession No. MI0014151, SEQ ID NO: 472) having a hairpin-like structure as a precursor.
- hsa-miR-4792 gene or “hsa-miR-4792” refers to the hsa-miR-4792 gene (miRBase Accession No. MIMAT0019964) described in SEQ ID NO: 197 or other biological species. A homolog or ortholog is included.
- the hsa-miR-4792 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “Hsa-miR-4792” is known as “hsa-mir-4792” (miRBase Accession No. MI0017439, SEQ ID NO: 473), which has a hairpin-like structure as a precursor.
- hsa-miR-6510-5p gene or “hsa-miR-6510-5p” refers to the hsa-miR-6510-5p gene described in SEQ ID NO: 198 (miRBase Accession No. MIMAT0025476) and other species homologs or orthologs.
- the hsa-miR-6510-5p gene can be obtained by the method described in Joyce CE et al., 2011, Hum Mol Genet, 20, p4025-4040.
- “Hsa-miR-6510-5p” is known as “hsa-mir-6510” (miRBase Accession No. MI0022222, SEQ ID NO: 474) having a hairpin-like structure as a precursor.
- hsa-miR-504-3p gene or “hsa-miR-504-3p” refers to the hsa-miR-504-3p gene (miRBase Accession No. 199 described in SEQ ID NO: 199). MIMAT0026612) and other species homologues or orthologues are included.
- the hsa-miR-504-3p gene can be obtained by the method described in Bentwich I et al., 2005, Nat Genet, 37, p766-770.
- “hsa-miR-504-3p” is known as “hsa-mir-504” (miRBase Accession No. MI0003189, SEQ ID NO: 475), which has a hairpin-like structure as a precursor.
- hsa-miR-3619-3p gene or “hsa-miR-3619-3p” refers to the hsa-miR-3619-3p gene (miRBase Accession No. MIMAT0019219) and other species homologues or orthologues are included.
- the hsa-miR-3619-3p gene can be obtained by the method described in Witten D et al., 2010, BMC Biol, Vol. 8, p58.
- hsa-mir-3619 (miRBase Accession No. MI0016009, SEQ ID NO: 476) having a hairpin-like structure as a precursor is known.
- hsa-miR-671-5p gene or “hsa-miR-671-5p” refers to the hsa-miR-671-5p gene described in SEQ ID NO: 201 (miRBase Accession No. MIMAT0003880) and other species homologs or orthologs.
- the hsa-miR-671-5p gene can be obtained by the method described in Berezikov E, et al., 2006, Genome Res, 16: p1299-1298.
- “hsa-miR-671-5p” is known as “hsa-mir-671” (miRBase Accession No. MI0003760, SEQ ID NO: 477) having a hairpin-like structure as a precursor.
- hsa-miR-4667-5p gene or “hsa-miR-4667-5p” refers to the hsa-miR-4667-5p gene described in SEQ ID NO: 202 (miRBase Accession No. MIMAT0019743) and other species homologues or orthologues are included.
- the hsa-miR-4667-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4667-5p” “hsa-mir-4667” (miRBase Accession No. MI0017297, SEQ ID NO: 478) having a hairpin-like structure as a precursor is known.
- hsa-miR-4430 gene or “hsa-miR-4430” refers to the hsa-miR-4430 gene (miRBase Accession No. MIMAT0018945) described in SEQ ID NO: 203 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4430 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-mir-4430 (miRBase Accession No. MI0016769, SEQ ID NO: 479) having a hairpin-like structure as a precursor is known.
- hsa-miR-3195 gene or “hsa-miR-3195” refers to the hsa-miR-3195 gene (miRBase Accession No. MIMAT0015079) described in SEQ ID NO: 204 and other biological species. Homologs or orthologs are included.
- the hsa-miR-3195 gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- hsa-miR-3195 “hsa-mir-3195” (miRBase Accession No. MI0014240, SEQ ID NO: 480) having a hairpin-like structure as a precursor is known.
- hsa-miR-3679-5p gene or “hsa-miR-3679-5p” refers to the hsa-miR-3679-5p gene described in SEQ ID NO: 205 (miRBase Accession No. MIMAT0018104) and other species homologues or orthologues are included.
- the hsa-miR-3679-5p gene can be obtained by the method described in Creighton CJ et al., 2010, PLoS One, 5, e9637.
- hsa-miR-3679-5p “hsa-mir-3679” (miRBase Accession No. MI0016080, SEQ ID NO: 358) having a hairpin-like structure as a precursor is known.
- hsa-miR-6076 gene or “hsa-miR-6076” refers to the hsa-miR-6076 gene (miRBase Accession No. MIMAT0023701) described in SEQ ID NO: 206 or other biological species. Homologs or orthologs are included.
- the hsa-miR-6076 gene can be obtained by the method described in Voellenkle C et al., 2012, RNA, 18, p472-484.
- hsa-miR-6076 “hsa-mir-6076” (miRBase Accession No. MI0020353, SEQ ID NO: 481) having a hairpin-like structure as a precursor is known.
- hsa-miR-6515-5p gene or “hsa-miR-6515-5p” refers to the hsa-miR-6515-5p gene described in SEQ ID NO: 207 (miRBase Accession No. MIMAT0025486) and other species homologues or orthologues are included.
- the hsa-miR-6515-5p gene can be obtained by the method described in Joyce CE et al., 2011, Hum Mol Genet, 20, p4025-4040.
- “hsa-miR-6515-5p” is known as “hsa-mir-6515” (miRBase Accession No. MI0022227, SEQ ID NO: 340) having a hairpin-like structure as a precursor.
- hsa-miR-6820-5p gene or “hsa-miR-6820-5p” refers to the hsa-miR-6820-5p gene described in SEQ ID NO: 208 (miRBase Accession No. MIMAT0027540) and other species homologs or orthologs.
- the hsa-miR-6820-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6820-5p “hsa-mir-6820” (miRBase Accession No. MI0022665, SEQ ID NO: 482) having a hairpin-like structure as a precursor is known.
- hsa-miR-4634 gene or “hsa-miR-4634” refers to the hsa-miR-4634 gene (miRBase Accession No. MIMAT0019691) described in SEQ ID NO: 209 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4634 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- hsa-miR-4634 “hsa-mir-4634” (miRBase Accession No. MI0017261, SEQ ID NO: 483) having a hairpin-like structure as a precursor is known.
- hsa-miR-187-5p gene or “hsa-miR-187-5p” refers to the hsa-miR-187-5p gene described in SEQ ID NO: 210 (miRBase Accession No. MIMAT0004561) and other species homologs or orthologs are included.
- the hsa-miR-187-5p gene can be obtained by the method described in Lim LP et al., 2003, Science, 299, p1540.
- “hsa-miR-187-5p” is known as “hsa-mir-187” (miRBase Accession No. MI000000274, SEQ ID NO: 484) having a hairpin-like structure as a precursor.
- hsa-miR-6663-5p gene or “hsa-miR-6663-5p” refers to the hsa-miR-6663-5p gene described in SEQ ID NO: 211 (miRBase Accession No. MIMAT0027426) and other species homologues or orthologues are included.
- the hsa-miR-6673-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6673-5p “hsa-mir-6663” (miRBase Accession No. MI0022608, SEQ ID NO: 434) having a hairpin-like structure as a precursor is known.
- hsa-miR-1908-3p gene or “hsa-miR-1908-3p” refers to the hsa-miR-1908-3p gene (miRBase Accession No. MIMAT0026916) and other species homologues or orthologues are included.
- the hsa-miR-1908-3p gene can be obtained by the method described in Bar M, et al., 2008, Stem Cells, 26, p2496-2505.
- “hsa-miR-1908-3p” is known as “hsa-mir-1908” (miRBase Accession No. MI0008329, SEQ ID NO: 393) having a hairpin-like structure as a precursor.
- hsa-miR-1181 gene or “hsa-miR-1181” refers to the hsa-miR-1181 gene (miRBase Accession No. MIMAT0005826) described in SEQ ID NO: 213 or other biological species. Homologs or orthologs are included.
- the hsa-miR-1181 gene can be obtained by the method described in Subramanian S et al., 2008, Oncogene, 27, p2015-2026.
- hsa-miR-1181 “hsa-mir-1181” (miRBase Accession No. MI0006274, SEQ ID NO: 485), which has a hairpin-like structure as a precursor, is known.
- hsa-miR-6782-5p gene or “hsa-miR-6782-5p” refer to the hsa-miR-6782-5p gene described in SEQ ID NO: 214 (miRBase Accession No. MIMAT0027464) and other species homologues or orthologues are included.
- the hsa-miR-6782-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6782-5p” is known as “hsa-mir-6782” (miRBase Accession No. MI0022627, SEQ ID NO: 486), which has a hairpin-like structure as a precursor.
- hsa-miR-5010-5p gene or “hsa-miR-5010-5p” refers to the hsa-miR-5010-5p gene described in SEQ ID NO: 215 (miRBase Accession No. MIMAT0021043) and other species homologs or orthologs.
- the hsa-miR-5010-5p gene can be obtained by the method described in Hansen TB et al., 2011, RNA Biol, 8, p378-383.
- “hsa-miR-5010-5p” is known as “hsa-mir-5010” (miRBase Accession No. MI0017878, SEQ ID NO: 487), which has a hairpin-like structure as a precursor.
- hsa-miR-6870-5p gene or “hsa-miR-6870-5p” refers to the hsa-miR-6870-5p gene described in SEQ ID NO: 216 (miRBase Accession No. MIMAT0027640) and other species homologues or orthologues are included.
- the hsa-miR-6870-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6870-5p “hsa-mir-6870” (miRBase Accession No. MI0022717, SEQ ID NO: 488) having a hairpin-like structure as a precursor is known.
- hsa-miR-6124 gene or “hsa-miR-6124” refers to the hsa-miR-6124 gene (miRBase Accession No. MIMAT0024597) described in SEQ ID NO: 217 or other biological species. Homologs or orthologs are included.
- the hsa-miR-6124 gene can be obtained by the method described in Smith JL et al., 2012, J Virol, 86, p5278-5287.
- hsa-miR-6124 “hsa-mir-6124” (miRBase Accession No. MI0021258, SEQ ID NO: 489), which has a hairpin-like structure as a precursor, is known.
- hsa-miR-1249-5p gene or “hsa-miR-1249-5p” refers to the hsa-miR-1249-5p gene described in SEQ ID NO: 218 (miRBase Accession No. MIMAT0032029) and other species homologues or orthologues are included.
- the hsa-miR-1249-5p gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- “hsa-miR-1249-5p” “hsa-mir-1249” (miRBase Accession No. MI0006384, SEQ ID NO: 321) having a hairpin-like structure as a precursor is known.
- hsa-miR-6511b-5p gene or “hsa-miR-6511b-5p” refers to the hsa-miR-6511b-5p gene described in SEQ ID NO: 219 (miRBase Accession No. MIMAT0025847) and other species homologs or orthologs.
- the hsa-miR-6511b-5p gene can be obtained by the method described in Li Y et al., 2012, Gene, 497, p330-335.
- “Hsa-miR-6511b-5p” has a hairpin-like structure as a precursor thereof, “hsa-mir-6651b-1, hsa-mir-6511b-2” (miRBase Accession No. MI0022552, MI0023431, SEQ ID NO: 490, 491) are known.
- hsa-miR-1254 gene or “hsa-miR-1254” refers to the hsa-miR-1254 gene (miRBase Accession No. MIMAT0005905) described in SEQ ID NO: 220 or other biological species. A homolog or ortholog is included.
- the hsa-miR-1254 gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621. “Hsa-miR-1254” has a hairpin-like structure as a precursor thereof, “hsa-mir-1254-1, hsa-mir-1254-2” (miRBase Accession No. MI0006388, MI0016747, SEQ ID NO: 492, 493) is known.
- hsa-miR-4727-3p gene or “hsa-miR-4727-3p” refers to the hsa-miR-4727-3p gene described in SEQ ID NO: 221 (miRBase Accession No. MIMAT0019848) and other species homologs or orthologs.
- the hsa-miR-4727-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4727-3p” “hsa-mir-4727” (miRBase Accession No. MI0017364, SEQ ID NO: 494) having a hairpin-like structure as a precursor is known.
- hsa-miR-4259 gene or “hsa-miR-4259” refers to the hsa-miR-4259 gene (miRBase Accession No. MIMAT0016880) described in SEQ ID NO: 222 and other biological species. A homolog or ortholog is included.
- the hsa-miR-4259 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- “hsa-miR-4259” is known as “hsa-mir-4259” (miRBase Accession No. MI0015858, SEQ ID NO: 495) having a hairpin-like structure as a precursor.
- hsa-miR-4771 gene or “hsa-miR-4771” refers to the hsa-miR-4771 gene (miRBase Accession No. MIMAT0019925) described in SEQ ID NO: 223 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4771 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “Hsa-miR-4771” has “hsa-mir-4771-1, hsa-mir-4771-2” (miRBase Accession No. MI0017412, MI0017413, SEQ ID NO: 496, which takes a hairpin-like structure as a precursor thereof. 497) is known.
- hsa-miR-3622a-5p gene or “hsa-miR-3622a-5p” refers to the hsa-miR-3622a-5p gene described in SEQ ID NO: 224 (miRBase Accession No. MIMAT0018003) and other species homologs or orthologs.
- the hsa-miR-3622a-5p gene can be obtained by the method described in Witten D et al., 2010, BMC Biol, Vol. 8, p58.
- hsa-mir-3622a “miRBase Accession No. MI0016013, SEQ ID NO: 498) having a hairpin-like structure as a precursor is known.
- hsa-miR-4480 gene or “hsa-miR-4480” refers to the hsa-miR-4480 gene (miRBase Accession No. MIMAT0019014) described in SEQ ID NO: 225 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4480 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- “Hsa-miR-4480” is known as “hsa-mir-4480” (miRBase Accession No. MI0016841, SEQ ID NO: 499) having a hairpin-like structure as a precursor.
- hsa-miR-4740-5p gene or “hsa-miR-4740-5p” refers to the hsa-miR-4740-5p gene described in SEQ ID NO: 226 (miRBase Accession No. MIMAT0019869) and other species homologues or orthologues are included.
- the hsa-miR-4740-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4740-5p” is known as “hsa-mir-4740” (miRBase Accession No. MI0017378, SEQ ID NO: 500) having a hairpin-like structure as a precursor.
- hsa-miR-6777-5p gene or “hsa-miR-6777-5p” refers to the hsa-miR-6777-5p gene described in SEQ ID NO: 227 (miRBase Accession No. MIMAT0027454) and other species homologues or orthologues are included.
- the hsa-miR-6777-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6777-5p “hsa-mir-6777” (miRBase Accession No. MI0022622, SEQ ID NO: 501) having a hairpin-like structure as a precursor is known.
- hsa-miR-6794-5p gene or “hsa-miR-6794-5p” refers to the hsa-miR-6794-5p gene described in SEQ ID NO: 228 (miRBase Accession No. MIMAT0027488) and other species homologues or orthologues are included.
- the hsa-miR-6794-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6794-5p” is known as “hsa-mir-6794” (miRBase Accession No. MI0022639, SEQ ID NO: 502) having a hairpin-like structure as a precursor.
- hsa-miR-4687-3p gene or “hsa-miR-4687-3p” refers to the hsa-miR-4687-3p gene described in SEQ ID NO: 229 (miRBase Accession No. MIMAT0019775) and other species homologs or orthologs.
- the hsa-miR-4687-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4687-3p” “hsa-mir-4687” (miRBase Accession No. MI0017319, SEQ ID NO: 341) having a hairpin-like structure as a precursor is known.
- hsa-miR-6743-5p gene or “hsa-miR-6743-5p” refers to the hsa-miR-6743-5p gene described in SEQ ID NO: 230 (miRBase Accession No. MIMAT0027387) and other species homologues or orthologues are included.
- the hsa-miR-6743-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6743-5p” is known as “hsa-mir-6743” (miRBase Accession No. MI0022588, SEQ ID NO: 503) having a hairpin-like structure as a precursor.
- hsa-miR-6671-5p gene or “hsa-miR-6771-5p” refers to the hsa-miR-6671-5p gene described in SEQ ID NO: 231 (miRBase Accession No. MIMAT0027442) and other species homologues or orthologues are included.
- the hsa-miR-6671-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6711-5p” is known as “hsa-mir-6711” (miRBase Accession No. MI0022616, SEQ ID NO: 504) having a hairpin-like structure as a precursor.
- hsa-miR-3141 gene or “hsa-miR-3141” refers to the hsa-miR-3141 gene (miRBase Accession No. MIMAT0015010) described in SEQ ID NO: 232 or other biological species. Homologs or orthologs are included.
- the hsa-miR-3141 gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- “hsa-miR-3141” is known as “hsa-mir-3141” (miRBase Accession No. MI0014165, SEQ ID NO: 505), which has a hairpin-like structure as a precursor.
- hsa-miR-3162-5p gene or “hsa-miR-3162-5p” refers to the hsa-miR-3162-5p gene described in SEQ ID NO: 233 (miRBase Accession No. MIMAT0015036) and other species homologs or orthologs.
- the hsa-miR-3162-5p gene can be obtained by the method described in Stark MS et al., 2010, PLoS One, 5, e9685.
- “hsa-miR-3162-5p” “hsa-mir-3162” (miRBase Accession No. MI0014192, SEQ ID NO: 506) having a hairpin-like structure as a precursor is known.
- hsa-miR-4271 gene or “hsa-miR-4271” refers to the hsa-miR-4271 gene (miRBase Accession No. MIMAT0016901) described in SEQ ID NO: 234 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4271 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- hsa-miR-4271 “hsa-mir-4271” (miRBase Accession No. MI0015879, SEQ ID NO: 507) having a hairpin-like structure as a precursor is known.
- hsa-miR-1227-5p gene or “hsa-miR-1227-5p” refers to the hsa-miR-1227-5p gene (miRBase Accession No. MIMAT0022941) and other species homologues or orthologues are included.
- the hsa-miR-1227-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, 28, p328-336.
- hsa-mir-1227 (miRBase Accession No. MI0006316, SEQ ID NO: 508) having a hairpin-like structure as a precursor is known.
- hsa-miR-4257 gene or “hsa-miR-4257” refers to the hsa-miR-4257 gene (miRBase Accession No. MIMAT0016878) described in SEQ ID NO: 236 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4257 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- hsa-miR-4257 is known as “hsa-mir-4257” (miRBase Accession No. MI0015856, SEQ ID NO: 509) having a hairpin-like structure as a precursor.
- hsa-miR-4270 gene or “hsa-miR-4270” refers to the hsa-miR-4270 gene (miRBase Accession No. MIMAT0016900) described in SEQ ID NO: 237 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4270 gene can be obtained by the method described in Goff LA et al., 2009, PLoS One, Vol. 4, e7192.
- hsa-miR-4270 “hsa-mir-4270” (miRBase Accession No. MI0015878, SEQ ID NO: 510) having a hairpin-like structure as a precursor is known.
- hsa-miR-4516 gene or “hsa-miR-4516” refers to the hsa-miR-4516 gene (miRBase Accession No. MIMAT0019053) described in SEQ ID NO: 238 or other biological species. Homologs or orthologs are included.
- the hsa-miR-4516 gene can be obtained by the method described in Jim DD et al., 2010, Blood, 116, e118-e127.
- hsa-miR-4516 “hsa-mir-4516” (miRBase Accession No. MI0016882, SEQ ID NO: 511) having a hairpin-like structure as a precursor is known.
- hsa-miR-4651 gene or “hsa-miR-4651” refers to the hsa-miR-4651 gene (miRBase Accession No. MIMAT0019715) described in SEQ ID NO: 239 or other biological species. A homolog or ortholog is included.
- the hsa-miR-4651 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4651” is known as “hsa-mir-4651” (miRBase Accession No. MI0017279, SEQ ID NO: 512) having a hairpin-like structure as a precursor.
- hsa-miR-4725-3p gene or “hsa-miR-4725-3p” refers to the hsa-miR-4725-3p gene described in SEQ ID NO: 240 (miRBase Accession No. MIMAT0019844) and other species homologs or orthologs.
- the hsa-miR-4725-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, 71, p78-86.
- “hsa-miR-4725-3p” is known as “hsa-mir-4725” (miRBase Accession No. MI0017362, SEQ ID NO: 513) having a hairpin-like structure as a precursor.
- hsa-miR-6125 gene or “hsa-miR-6125” refers to the hsa-miR-6125 gene (miRBase Accession No. MIMAT0024598) described in SEQ ID NO: 241 or other biological species. A homolog or ortholog is included.
- the hsa-miR-6125 gene can be obtained by the method described in Smith JL et al., 2012, J Virol, 86, p5278-5287.
- “hsa-miR-6125” is known as “hsa-mir-6125” (miRBase Accession No. MI0021259, SEQ ID NO: 514) having a hairpin-like structure as a precursor.
- hsa-miR-6732-5p gene or “hsa-miR-6732-5p” refers to the hsa-miR-6732-5p gene described in SEQ ID NO: 242 (miRBase Accession No. MIMAT0027365) and other species homologs or orthologs.
- the hsa-miR-6732-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6732-5p” is known as “hsa-mir-6732” (miRBase Accession No. MI0022577, SEQ ID NO: 515), which has a hairpin-like structure as a precursor.
- hsa-miR-6791-5p gene or “hsa-miR-6791-5p” refers to the hsa-miR-6791-5p gene described in SEQ ID NO: 243 (miRBase Accession No. MIMAT0027482) and other species homologues or orthologues are included.
- the hsa-miR-6791-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6791-5p” is known as “hsa-mir-6791” (miRBase Accession No. MI0022636, SEQ ID NO: 516) having a hairpin-like structure as a precursor.
- hsa-miR-6819-5p gene or “hsa-miR-6819-5p” refers to the hsa-miR-6819-5p gene described in SEQ ID NO: 244 (miRBase Accession No. MIMAT0027538) and other species homologs or orthologs.
- the hsa-miR-6819-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-6819-5p” is known as “hsa-mir-6819” (miRBase Accession No. MI0022664, SEQ ID NO: 517) having a hairpin-like structure as a precursor.
- hsa-miR-6891-5p gene or “hsa-miR-6891-5p” refer to the hsa-miR-6891-5p gene described in SEQ ID NO: 245 (miRBase Accession No. MIMAT0027682) and other species homologues or orthologues are included.
- the hsa-miR-6891-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- hsa-miR-6891-5p “hsa-mir-6891” (miRBase Accession No. MI0022738, SEQ ID NO: 518) having a hairpin-like structure as a precursor is known.
- hsa-miR-7108-5p gene or “hsa-miR-7108-5p” refers to the hsa-miR-7108-5p gene described in SEQ ID NO: 246 (miRBase Accession No. MIMAT0028113) and other species homologues or orthologues are included.
- the hsa-miR-7108-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-7108-5p” is known as “hsa-mir-7108” (miRBase Accession No. MI0022959, SEQ ID NO: 448) having a hairpin-like structure as a precursor.
- hsa-miR-7109-5p gene or “hsa-miR-7109-5p” refers to the hsa-miR-7109-5p gene described in SEQ ID NO: 247 (miRBase Accession No. MIMAT0028115) and other species homologs or orthologs.
- the hsa-miR-7109-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645.
- “hsa-miR-7109-5p” is known as “hsa-mir-7109” (miRBase Accession No. MI0022960, SEQ ID NO: 519) having a hairpin-like structure as a precursor.
- hsa-miR-320a gene or “hsa-miR-320a” refers to the hsa-miR-320a gene (miRBase Accession No. MIMAT000010) described in SEQ ID NO: 248 or other biological species. Homologs or orthologs are included.
- the hsa-miR-320a gene can be obtained by the method described in Michael MZ et al., 2003, Mol Cancer Res, Vol. 1, p882-891.
- hsa-miR-320a “hsa-mir-320a” (miRBase Accession No. MI000042, SEQ ID NO: 520) having a hairpin-like structure as a precursor is known.
- hsa-miR-663a gene or “hsa-miR-663a” refers to the hsa-miR-663a gene (miRBase Accession No. MIMAT0003326) described in SEQ ID NO: 249 or other biological species. Homologs or orthologs are included.
- the hsa-miR-663a gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692.
- “hsa-miR-663a” is known as “hsa-mir-663a” (miRBase Accession No. MI0003672, SEQ ID NO: 521) having a hairpin-like structure as a precursor.
- hsa-miR-328-5p gene or “hsa-miR-328-5p” refers to the hsa-miR-328-5p gene (miRBase Accession No. MIMAT0026486) and other species homologues or orthologues are included.
- the hsa-miR-328-5p gene can be obtained by the method described in Kim J et al., 2004, Proc Natl Acad Sci USA, Vol. 101, p360-365.
- hsa-miR-328-5p “hsa-mir-328” (miRBase Accession No. MI000004, SEQ ID NO: 522) having a hairpin-like structure as a precursor is known.
- hsa-miR-642b-3p gene or “hsa-miR-642b-3p” refers to the hsa-miR-642b-3p gene described in SEQ ID NO: 251 (miRBase Accession No. MIMAT0018444) and other species homologues or orthologues are included.
- the hsa-miR-642b-3p gene can be obtained by the method described in Witten D et al., 2010, BMC Biol, Vol. 8, p58.
- hsa-mir-642b miRBase Accession No. MI0016685, SEQ ID NO: 523 having a hairpin-like structure as a precursor is known.
- hsa-miR-128-2-5p gene or “hsa-miR-128-2-5p” refers to the hsa-miR-128-2-5p set forth in SEQ ID NO: 252. Genes (miRBBase Accession No. MIMAT0031095) and other species homologues or orthologues are included.
- the hsa-miR-128-2-5p gene can be obtained by the method described in Lagos-Quintana M et al., 2002, Curr Biol, Vol. 12, p735-739.
- “hsa-miR-128-2-5p” is known as “hsa-mir-128-2” (miRBase Accession No. MI000027, SEQ ID NO: 524) having a hairpin-like structure as a precursor.
- hsa-miR-125a-3p gene or “hsa-miR-125a-3p” refers to the hsa-miR-125a-3p gene described in SEQ ID NO: 253 (miRBase Accession No. MIMAT0004602) and other species homologues or orthologues are included.
- the hsa-miR-125a-3p gene can be obtained by the method described in Lagos-Quintana M et al., 2002, Curr Biol, 12, p735-739.
- “hsa-miR-125a-3p” “hsa-mir-125a” (miRBase Accession No. MI000069, SEQ ID NO: 525) having a hairpin-like structure as a precursor is known.
- hsa-miR-191-5p gene or “hsa-miR-191-5p” refers to the hsa-miR-191-5p gene described in SEQ ID NO: 254 (miRBase Accession No. MIMAT000040) and other species homologues or orthologues are included.
- the hsa-miR-191-5p gene can be obtained by the method described in Lagos-Quintana M et al., 2003, RNA, Vol. 9, p175-179.
- “hsa-miR-191-5p” is known as “hsa-mir-191” (miRBase Accession No. MI000065, SEQ ID NO: 526) having a hairpin-like structure as a precursor.
- hsa-miR-92b-5p gene or “hsa-miR-92b-5p” refers to the hsa-miR-92b-5p gene described in SEQ ID NO: 255 (miRBase Accession No. MIMAT0004792) and other species homologs or orthologs are included.
- the hsa-miR-92b-5p gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, 103, p3687-3692.
- “hsa-miR-92b-5p” is known as “hsa-mir-92b” (miRBase Accession No. MI0003560, SEQ ID NO: 527) having a hairpin-like structure as a precursor.
- hsa-miR-296-5p gene or “hsa-miR-296-5p” refers to the hsa-miR-296-5p gene described in SEQ ID NO: 256 (miRBase Accession No. MIMAT060690) and other species homologues or orthologues are included.
- the hsa-miR-296-5p gene can be obtained by the method described in Houbavy HB et al., 2003, Dev Cell, Vol. 5, p351-358.
- hsa-mir-296 miRBase Accession No. MI000047, SEQ ID NO: 336) having a hairpin-like structure as a precursor is known.
- hsa-miR-1246 gene or “hsa-miR-1246” refers to the hsa-miR-1246 gene (miRBase Accession No. MIMAT0005898) described in SEQ ID NO: 257 or other biological species. Homologs or orthologs are included.
- the hsa-miR-1246 gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- hsa-miR-1246 “hsa-mir-1246” (miRBase Accession No. MI0006381, SEQ ID NO: 528) having a hairpin-like structure as a precursor is known.
- hsa-miR-92a-2-5p gene or “hsa-miR-92a-2-5p” refers to the hsa-miR-92a-2-5p set forth in SEQ ID NO: 258. Genes (miRBBase Accession No. MIMAT0004508) and other species homologues or orthologues are included.
- the hsa-miR-92a-2-5p gene can be obtained by the method described in Murelatos Z et al., 2002, Genes Dev, 16, p720-728.
- “hsa-miR-92a-2-5p” is known as “hsa-mir-92a-2” (miRBase Accession No. MI00000094, SEQ ID NO: 529) having a hairpin-like structure as a precursor.
- hsa-miR-128-1-5p gene or “hsa-miR-128-1-5p” refers to the hsa-miR-128-1-5p set forth in SEQ ID NO: 259. Genes (miRBBase Accession No. MIMAT0026477) and other species homologues or orthologues are included.
- the hsa-miR-128-1-5p gene can be obtained by the method described in Lagos-Quintana M et al., 2002, Curr Biol, Vol. 12, p735-739.
- “hsa-miR-128-1-5p” is known as “hsa-mir-128-1” (miRBase Accession No. MI0000447, SEQ ID NO: 530) having a hairpin-like structure as a precursor.
- hsa-miR-1290 gene or “hsa-miR-1290” refers to the hsa-miR-1290 gene (miRBase Accession No. MIMAT0005880) described in SEQ ID NO: 260 and other biological species. Homologs or orthologs are included.
- the hsa-miR-1290 gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, 18, p610-621.
- hsa-miR-1290 “hsa-mir-1290” (miRBase Accession No. MI0006352, SEQ ID NO: 531) having a hairpin-like structure as a precursor is known.
- hsa-miR-211-3p gene or “hsa-miR-211-3p” as used herein refers to the hsa-miR-211-3p gene described in SEQ ID NO: 261 (miRBase Accession No. MIMAT0022694) and other species homologues or orthologues are included.
- the hsa-miR-211-3p gene can be obtained by the method described in Lim LP et al., 2003, Science, 299, p1540.
- “hsa-miR-211-3p” is known as “hsa-mir-211” (miRBase Accession No. MI000027, SEQ ID NO: 532) having a hairpin-like structure as a precursor.
- hsa-miR-744-5p gene or “hsa-miR-744-5p” refers to the hsa-miR-744-5p gene described in SEQ ID NO: 262 (miRBase Accession No. MIMAT0004945) and other species homologs or orthologs.
- the hsa-miR-744-5p gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, 16, p1299-1298.
- “hsa-miR-744-5p” is known as “hsa-mir-744” (miRBase Accession No. MI0005559, SEQ ID NO: 533) having a hairpin-like structure as a precursor.
- hsa-miR-135a-3p gene or “hsa-miR-135a-3p” refers to the hsa-miR-135a-3p gene described in SEQ ID NO: 263 (miRBase Accession No. MIMAT0004595) and other species homologs or orthologs.
- the hsa-miR-135a-3p gene can be obtained by the method described in Lagos-Quintana M, et al., 2002, Curr Biol, 12, p735-739.
- “hsa-miR-135a-3p” is known as “hsa-mir-135a-1” (miRBase Accession No. MI000052, SEQ ID NO: 534) having a hairpin-like structure as a precursor.
- hsa-miR-451a gene or “hsa-miR-451a” refers to the hsa-miR-451a gene (miRBase Accession No. MIMAT0001631) described in SEQ ID NO: 264 or other biological species. Homologs or orthologs are included.
- the hsa-miR-451a gene can be obtained by the method described in Altvia Y et al., 2005, Nucleic Acids Res, 33, p2697-2706.
- hsa-miR-451a “hsa-mir-451a” (miRBase Accession No. MI0001729, SEQ ID NO: 535) having a hairpin-like structure as a precursor is known.
- hsa-miR-625-3p gene or “hsa-miR-625-3p” refers to the hsa-miR-625-3p gene described in SEQ ID NO: 265 (miRBase Accession No. MIMAT0004808) and other species homologues or orthologues are included.
- the hsa-miR-625-3p gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692.
- “hsa-miR-625-3p” is known as “hsa-mir-625” (miRBase Accession No. MI0003639, SEQ ID NO: 536) having a hairpin-like structure as a precursor.
- hsa-miR-92a-3p gene or “hsa-miR-92a-3p” refers to the hsa-miR-92a-3p gene described in SEQ ID NO: 266 (miRBase Accession No. MIMAT00000092) and other species homologs or orthologs.
- the hsa-miR-92a-3p gene can be obtained by the method described in Murelatos Z et al., 2002, Genes Dev, 16, p720-728.
- “Hsa-miR-92a-3p” has a hairpin-like structure as its precursor, “hsa-mir-92a-1, hsa-mir-92a-2” (miRBase Accession No. MI00000093, MI00000094, SEQ ID NO: 537, 529).
- hsa-miR-422a gene or “hsa-miR-422a” refers to the hsa-miR-422a gene (miRBase Accession No. MIMAT0001339) described in SEQ ID NO: 267 or other biological species. Homologs or orthologs are included.
- the hsa-miR-422a gene can be obtained by the method described in Kasashima K et al., 2004, Biochem Biophys Res Commun, 322, p403-410.
- hsa-miR-422a “hsa-mir-422a” (miRBase Accession No. MI0001444, SEQ ID NO: 538) having a hairpin-like structure as a precursor is known.
- hsa-miR-642a-3p gene or “hsa-miR-642a-3p” refer to the hsa-miR-642a-3p gene described in SEQ ID NO: 268 (miRBase Accession No. MIMAT0020924) and other species homologs or orthologs.
- the hsa-miR-642a-3p gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, 103, p3687-3692.
- “hsa-miR-642a-3p” is known as “hsa-mir-642a” (miRBase Accession No. MI0003657, SEQ ID NO: 539) having a hairpin-like structure as a precursor.
- hsa-miR-483-5p gene or “hsa-miR-483-5p” refers to the hsa-miR-483-5p gene described in SEQ ID NO: 269 (miRBase Accession No. MIMAT0004761) and other species homologues or orthologues are included.
- the hsa-miR-483-5p gene can be obtained by the method described in Fu H et al., 2005, FEBS Lett, 579, p3849-3854.
- “hsa-miR-483-5p” “hsa-mir-483” (miRBase Accession No. MI0002467, SEQ ID NO: 540) having a hairpin-like structure as a precursor is known.
- hsa-miR-652-5p gene or “hsa-miR-652-5p” refers to the hsa-miR-652-5p gene described in SEQ ID NO: 270 (miRBase Accession No. MIMAT0022709) and other species homologues or orthologues are included.
- the hsa-miR-652-5p gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692.
- “Hsa-miR-652-5p” is known as “hsa-mir-652” (miRBase Accession No. MI0003667, SEQ ID NO: 541) having a hairpin-like structure as a precursor.
- hsa-miR-24-3p gene or “hsa-miR-24-3p” refers to the hsa-miR-24-3p gene described in SEQ ID NO: 271 (miRBase Accession No. MIMAT00000080) and other species homologs or orthologs.
- the hsa-miR-24-3p gene can be obtained by the method described in Lagos-Quintana M et al., 2001, Science, 294, p853-858.
- “Hsa-miR-24-3p” has a hairpin-like structure as its precursor, “hsa-mir-24-1, hsa-mir-24-2” (miRBase Accession No. MI00000080, MI00000081, SEQ ID NO: 542, 543) are known.
- hsa-miR-23b-3p gene or “hsa-miR-23b-3p” refers to the hsa-miR-23b-3p gene described in SEQ ID NO: 272 (miRBase Accession No. MIMAT000018) and other species homologues or orthologues are included.
- the hsa-miR-23b-3p gene can be obtained by the method described in Lagos-Quintana M et al., 2002, Curr Biol, 12, p735-739.
- “hsa-miR-23b-3p” is known as “hsa-mir-23b” (miRBase Accession No. MI0000439, SEQ ID NO: 544) having a hairpin-like structure as a precursor.
- hsa-miR-23a-3p gene or “hsa-miR-23a-3p” refers to the hsa-miR-23a-3p gene described in SEQ ID NO: 273 (miRBase Accession No. MIMAT0000078) and other species homologues or orthologues are included.
- the hsa-miR-23a-3p gene can be obtained by the method described in Lagos-Quintana M et al., 2001, Science, 294, p853-858.
- “hsa-miR-23a-3p” “hsa-mir-23a” (miRBase Accession No. MI0000079, SEQ ID NO: 545) having a hairpin-like structure as a precursor is known.
- hsa-miR-92b-3p gene or “hsa-miR-92b-3p” refers to the hsa-miR-92b-3p gene described in SEQ ID NO: 274 (miRBase Accession No. MIMAT0003218) and other species homologues or orthologues are included.
- the hsa-miR-92b-3p gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692.
- hsa-miR-92b-3p is known as “hsa-mir-92b” (miRBase Accession No. MI0003560, SEQ ID NO: 527) having a hairpin-like structure as a precursor.
- hsa-miR-22-3p gene or “hsa-miR-22-3p” refers to the hsa-miR-22-3p gene described in SEQ ID NO: 275 (miRBase Accession No. MIMAT0000077) and other species homologs or orthologs.
- the hsa-miR-22-3p gene can be obtained by the method described in Lagos-Quintana M et al., 2001, Science, 294, p853-858.
- “hsa-miR-22-3p” is known as “hsa-mir-22” (miRBase Accession No. MI0000078, SEQ ID NO: 546) having a hairpin-like structure as a precursor.
- isomiR miRNA RD. Et al., 2008, Genome Res., Vol. 18, p.610-621.
- miRBBase Release 21 in addition to the nucleotide sequence represented by any of SEQ ID NOs: 1 to 275, a number of variants and fragments of the nucleotide sequence represented by any of SEQ ID NOs: 547 to 842 called isomiR are also shown. Yes.
- mutants can also be obtained as miRNA having the nucleotide sequence represented by any of SEQ ID NOs: 1 to 275. That is, SEQ ID NOs: 2, 3, 4, 8, 11, 14, 17, 18, 20, 24, 25, 29, 34, 36, 39, 40, 41, 42, 43, 44, 45, 51 of the present invention.
- polynucleotides that are isomiRs of SEQ ID NOs: 1 to 275 registered in miRBBase.
- examples of the polynucleotide containing the base sequence represented by any one of SEQ ID NOs: 1 to 275 include the polynucleotides represented by any one of SEQ ID NOs: 276 to 546, which are precursors.
- nucleic acid such as a nucleic acid probe or primer used in the present invention binds to a specific target nucleic acid and cannot substantially bind to another nucleic acid.
- the present invention makes it possible to detect ovarian tumors easily and with high accuracy. For example, using a measurement value of the expression level of one to several miRNAs in blood, serum and / or plasma of a patient that can be collected in a minimally invasive manner, it is easy to detect whether the patient is an ovarian tumor or not. Can do.
- This figure shows hsa-miR-4433a-5p represented by SEQ ID NO: 128 generated from hsa-mir-4433a represented by SEQ ID NO: 405, and hsa-miR represented by SEQ ID NO: 151.
- the horizontal line in the figure shows the threshold (7.75) for discriminating both groups, optimized by Fisher's discriminant analysis.
- the dotted line in the figure indicates a discrimination boundary for discriminating both groups having a discrimination score of 0.
- Subjects who were not affected by ovarian tumors selected as the test specimen group 120 healthy subjects, 111 benign bone and soft tissue tumors and 111 benign disease patients, and 120 cancer patients other than ovarian cancer
- Hsa-miR-296-5p SEQ ID NO: 256
- hsa-miR-3131 SEQ ID NO: 196
- hsa-miR-1290 SEQ ID NO: 260
- hsa-miR- in serum of ovarian tumor patients 131
- a discriminant 1.390xhsa-miR-296-5p- in which the measured value of the expression level of 642a-3p (SEQ ID NO: 268) and hsa-miR-128-2-5p (SEQ ID NO: 252) was created in the learning sample group 1.095xhsa-miR-3131 + 0.201xhsa-miR-12
- Ovarian tumor target nucleic acid An ovarian tumor marker for detecting the presence and / or absence of an ovarian tumor or ovarian tumor cell using a nucleic acid such as a nucleic acid probe or primer for ovarian tumor detection as defined above of the present invention
- the main target nucleic acids as hsa-miR-4675, hsa-miR-4783-3p, hsa-miR-1228-5p, hsa-miR-4532, hsa-miR-6802-5p, hsa-miR-6784 -5p, hsa-miR-3940-5p, hsa-miR-1307-3p, hsa-miR-8073, hsa-miR-3184-5p, hsa-miR-1233-5p, hsa-miR-6088, hsa-miR -5195-3
- ovarian tumor markers that can be combined with these miRNAs: hsa-miR-320a, hsa-miR-663a, hsa-miR-328-5p, hsa-miR-128-2-5p, hsa-miR -125a-3p, hsa-miR-191-5p, hsa-miR-92b-5p, hsa-miR-296-5p, hsa-miR-1246, hsa-miR-92a-2-5p, hsa-miR-128 -1-5p, hsa-miR-1290, hsa-miR-211-3p, hsa-miR-744-5p, hsa-miR-135a-3p, hsa-miR-451a, hsa-miR-625-3p, hsa -MiR-92a-3p
- the miRNA includes, for example, a human gene containing a base sequence represented by any of SEQ ID NOs: 1 to 275 (ie, hsa-miR-4675, hsa-miR-4783-3p, hsa-miR-1228, respectively).
- a preferred target nucleic acid is a human gene comprising the base sequence represented by any of SEQ ID NOs: 1 to 842, a transcription product thereof, more preferably the transcription product, ie, miRNA, its precursor RNA, pri-miRNA or pre- miRNA.
- the first target gene is the hsa-miR-4675 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the second target gene is the hsa-miR-4783-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the third target gene is the hsa-miR-1228-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the fourth target gene is the hsa-miR-4532 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the fifth target gene is the hsa-miR-6802-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the sixth target gene is the hsa-miR-6784-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the seventh target gene is the hsa-miR-3940-5p gene, their homologues, their transcription products, or their mutants or derivatives. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the eighth target gene is the hsa-miR-1307-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the ninth target gene is the hsa-miR-8073 gene, their homologs, their transcripts, or their mutants or derivatives. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the tenth target gene is the hsa-miR-3184-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the eleventh target gene is the hsa-miR-1233-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the twelfth target gene is the hsa-miR-6088 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the thirteenth target gene is the hsa-miR-5195-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the fourteenth target gene is the hsa-miR-320b gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the fifteenth target gene is the hsa-miR-4649-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the sixteenth target gene is the hsa-miR-6800-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 17th target gene is the hsa-miR-1343-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 18th target gene is the hsa-miR-4730 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the nineteenth target gene is the hsa-miR-6885-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the twentieth target gene is the hsa-miR-5100 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 21st target gene is the hsa-miR-1203 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 22nd target gene is the hsa-miR-6756-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 23rd target gene is the hsa-miR-373-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 24th target gene is the hsa-miR-1268a gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 25th target gene is the hsa-miR-1260b gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the twenty-sixth target gene is the hsa-miR-4258 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 27th target gene is the hsa-miR-4697-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the twenty-eighth target gene is the hsa-miR-1469 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 29th target gene is the hsa-miR-4515 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 30th target gene is the hsa-miR-6861-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the thirty-first target gene is the hsa-miR-6821-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the thirty-second target gene is the hsa-miR-575 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 33rd target gene is the hsa-miR-6805-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 34th target gene is the hsa-miR-4758-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 35th target gene is the hsa-miR-3663-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the thirty-sixth target gene is the hsa-miR-4530 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 37th target gene is the hsa-miR-6798-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 38th target gene is the hsa-miR-6781-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 39th target gene is the hsa-miR-885-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 40th target gene is the hsa-miR-1273g-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 41st target gene is the hsa-miR-4787-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the forty-second target gene is the hsa-miR-4454 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 43rd target gene is the hsa-miR-4706 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 44th target gene is the hsa-miR-1249-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 45th target gene is the hsa-miR-887-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 46th target gene is the hsa-miR-6786-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 47th target gene is the hsa-miR-1238-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 48th target gene is the hsa-miR-6749-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 49th target gene is the hsa-miR-6729-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 50th target gene is the hsa-miR-6825-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 51st target gene is the hsa-miR-663b gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 52nd target gene is the hsa-miR-6858-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 53rd target gene is the hsa-miR-4690-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 54th target gene is the hsa-miR-6765-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 55th target gene is the hsa-miR-4710 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 56th target gene is the hsa-miR-6775-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 57th target gene is the hsa-miR-371a-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- 58 target genes are the hsa-miR-6816-5p gene, their homologs, their transcripts, or their mutants or derivatives. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 59th target gene is the hsa-miR-296-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 60th target gene is the hsa-miR-7777 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 61st target gene is the hsa-miR-8069 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 62nd target gene is the hsa-miR-6515-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 63rd target gene is the hsa-miR-4687-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 64th target gene is the hsa-miR-1343-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 65th target gene is the hsa-miR-7110-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 66th target gene is the hsa-miR-4525 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 67th target gene is the hsa-miR-3158-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 68th target gene is the hsa-miR-6787-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 69th target gene is an hsa-miR-614 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 70th target gene is the hsa-miR-4687 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 71st target gene is the hsa-miR-1185-2-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 72nd target gene is the hsa-miR-1268b gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 73rd target gene is the hsa-miR-1228-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 74th target gene is the hsa-miR-1185-1-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 75th target gene is an hsa-miR-940 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 76th target gene is an hsa-miR-939-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 77th target gene is the hsa-miR-6757-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 78th target gene is the hsa-miR-1275 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 79th target gene is the hsa-miR-5001-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 80th target gene is the hsa-miR-6826-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 81st target gene is the hsa-miR-6765-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 82nd target gene is the hsa-miR-3679-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 83rd target gene is the hsa-miR-4718 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 84th target gene is the hsa-miR-4286 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 85th target gene is the hsa-miR-8059 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 86th target gene is the hsa-miR-4447 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 87th target gene is the hsa-miR-4448 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 88th target gene is the hsa-miR-658 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 89th target gene is the hsa-miR-6766-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 90th target gene is an hsa-miR-197-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 91st target gene is the hsa-miR-6687-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 92nd target gene is the hsa-miR-6742-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 93rd target gene is the hsa-miR-6729-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 94th target gene is an hsa-miR-5090 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 95th target gene is the hsa-miR-7975 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 96th target gene is an hsa-miR-4505 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 97th target gene is the hsa-miR-6889-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 98th target gene is the hsa-miR-4708-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 99th target gene is the hsa-miR-6131 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 100th target gene is the hsa-miR-1225-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 101st target gene is the hsa-miR-6132 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 102nd target gene is the hsa-miR-4734 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 103rd target gene is the hsa-miR-3194-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 104th target gene is the hsa-miR-638 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 105th target gene is the hsa-miR-2467-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 106th target gene is the hsa-miR-4728-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 107th target gene is the hsa-miR-5572 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 108th target gene is the hsa-miR-6789-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 109th target gene is the hsa-miR-8063 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 110th target gene is the hsa-miR-4429 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 111th target gene is the hsa-miR-6840-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 112th target gene is the hsa-miR-4476 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 113th target gene is the hsa-miR-675-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 114th target gene is the hsa-miR-711 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 115th target gene is the hsa-miR-6875-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 116th target gene is the hsa-miR-3160-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 117th target gene is the hsa-miR-1908-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 118th target gene is the hsa-miR-6726-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 119th target gene is the hsa-miR-1913 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 120th target gene is the hsa-miR-8071 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 121st target gene is the hsa-miR-3648 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 122nd target gene is the hsa-miR-4732-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 123rd target gene is the hsa-miR-4787-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 124th target gene is the hsa-miR-3913 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 125th target gene is the hsa-miR-619-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 126th target gene is the hsa-miR-1231 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 127th target gene is the hsa-miR-342-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 128th target gene is the hsa-miR-4433a-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 129th target gene is the hsa-miR-6766-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 130th target gene is the hsa-miR-4707-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 131st target gene is the hsa-miR-7114-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 132nd target gene is the hsa-miR-6872-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 133rd target gene is the hsa-miR-6780b-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 134th target gene is the hsa-miR-7845-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 135th target gene is the hsa-miR-6798-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 136th target gene is the hsa-miR-665 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 137th target gene is the hsa-miR-6848-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 138th target gene is the hsa-miR-5008-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 139th target gene is the hsa-miR-4294 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 140th target gene is the hsa-miR-6511a-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 141st target gene is the hsa-miR-4435 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 142nd target gene is the hsa-miR-4747-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the target gene 143 is the hsa-miR-6880-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 144th target gene is the hsa-miR-6869-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 145th target gene is the hsa-miR-7150 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 146th target gene is the hsa-miR-1260a gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 147th target gene is the hsa-miR-6877-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 148th target gene is the hsa-miR-6721-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 149th target gene is the hsa-miR-4656 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 150th target gene is the hsa-miR-1229-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 151st target gene is the hsa-miR-4433a-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 152th target gene is the hsa-miR-4274 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 153rd target gene is the hsa-miR-4419b gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 154th target gene is the hsa-miR-4675 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 155th target gene is the hsa-miR-6893-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 156th target gene is the hsa-miR-6673-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 157th target gene is the hsa-miR-6762-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 158th target gene is the hsa-miR-6738-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 159th target gene is the hsa-miR-4513 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 160th target gene is the hsa-miR-6746-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 161st target gene is the hsa-miR-6880-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 162nd target gene is the hsa-miR-4736 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 163rd target gene is an hsa-miR-718 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 164th target gene is the hsa-miR-6717-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 165th target gene is the hsa-miR-7847-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 166th target gene is an hsa-miR-760 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 167th target gene is the hsa-miR-1199-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 168th target gene is the hsa-miR-6813-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 169th target gene is the hsa-miR-6769a-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 170th target gene is the hsa-miR-1193 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 171st target gene is the hsa-miR-7108-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 172nd target gene is the hsa-miR-6741-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 173rd target gene is the hsa-miR-4298 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 174th target gene is the hsa-miR-6969-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 175th target gene is the hsa-miR-4750-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 176th target gene is the hsa-miR-6785-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 177th target gene is the hsa-miR-1292-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 178th target gene is the hsa-miR-4749-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 179th target gene is the hsa-miR-6800-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 180th target gene is the hsa-miR-4722-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the target gene 181 is the hsa-miR-4746-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 182nd target gene is the hsa-miR-4450 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 183rd target gene is the hsa-miR-6695-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 184th target gene is the hsa-miR-365a-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 185th target gene is the hsa-miR-498 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 186th target gene is the hsa-miR-6797-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 187th target gene is the hsa-miR-1470 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 188th target gene is the hsa-miR-6851-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 189th target gene is the hsa-miR-1247-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 190th target gene is the hsa-miR-5196-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 191st target gene is the hsa-miR-208a-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 192nd target gene is the hsa-miR-6842-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 193rd target gene is the hsa-miR-150-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 194th target gene is the hsa-miR-4534 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 195th target gene is the hsa-miR-3135b gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 196th target gene is the hsa-miR-3131 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 197th target gene is the hsa-miR-4792 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 198th target gene is the hsa-miR-6510-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 199th target gene is the hsa-miR-504-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 200th target gene is the hsa-miR-3619-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 201st target gene is the hsa-miR-671-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 202nd target gene is the hsa-miR-4667-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 203rd target gene is the hsa-miR-4430 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 204th target gene is the hsa-miR-3195 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 205th target gene is the hsa-miR-3679-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 206th target gene is an hsa-miR-6076 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 207th target gene is the hsa-miR-6515-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 208th target gene is the hsa-miR-6820-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 209th target gene is the hsa-miR-4634 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 210th target gene is the hsa-miR-187-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 211st target gene is the hsa-miR-6673-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 212th target gene is the hsa-miR-1908-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the target gene 213 is the hsa-miR-1181 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 214th target gene is the hsa-miR-6782-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 215th target gene is an hsa-miR-5010-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 216th target gene is the hsa-miR-6870-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 217th target gene is the hsa-miR-6124 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 218th target gene is the hsa-miR-1249-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 219th target gene is the hsa-miR-6511b-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 220th target gene is the hsa-miR-1254 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the target gene 221 is the hsa-miR-4727-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 222nd target gene is the hsa-miR-4259 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 223rd target gene is the hsa-miR-4771 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 224th target gene is the hsa-miR-3622a-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 225th target gene is the hsa-miR-4480 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 226th target gene is the hsa-miR-4740-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 227th target gene is the hsa-miR-6777-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 228th target gene is the hsa-miR-6794-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 229th target gene is the hsa-miR-4687-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 230th target gene is the hsa-miR-6743-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the target gene 231 is the hsa-miR-6671-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 232th target gene is the hsa-miR-3141 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 233rd target gene is the hsa-miR-3162-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 234th target gene is the hsa-miR-4271 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 235th target gene is the hsa-miR-1227-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 236th target gene is the hsa-miR-4257 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 237th target gene is the hsa-miR-4270 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 238th target gene is the hsa-miR-4516 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 239th target gene is the hsa-miR-4651 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 240th target gene is the hsa-miR-4725-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the target gene 241 is the hsa-miR-6125 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 242nd target gene is the hsa-miR-6732-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 243rd target gene is the hsa-miR-6791-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 244th target gene is the hsa-miR-6819-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 245th target gene is the hsa-miR-6891-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 246th target gene is the hsa-miR-7108-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the target gene 247 is the hsa-miR-7109-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 248th target gene is the hsa-miR-320a gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 1.
- the 249th target gene is the hsa-miR-663a gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Non-patent Document 3.
- the 250th target gene is the hsa-miR-328-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 1.
- the target gene 251 is the hsa-miR-642b-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 252nd target gene is the hsa-miR-128-2-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- the 253rd target gene is the hsa-miR-125a-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 3.
- the 254th target gene is the hsa-miR-191-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 1.
- the 255th target gene is the hsa-miR-92b-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Non-patent Document 3).
- the 256th target gene is the hsa-miR-296-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Non-patent Document 3).
- the 257th target gene is the hsa-miR-1246 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a gene or a transcription product thereof can serve as a marker for ovarian tumors (Patent Document 4).
- the 258th target gene is the hsa-miR-92a-2-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- the 259th target gene is the hsa-miR-128-1-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- the 260th target gene is an hsa-miR-1290 gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 1.
- the target gene 261 is the hsa-miR-211-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 3.
- the 262th target gene is the hsa-miR-744-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a gene or a transcription product thereof can serve as a marker for ovarian tumors.
- the 263rd target gene is an hsa-miR-135a-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- Patent Document 2 There has been known a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors.
- the 264th target gene is the hsa-miR-451a gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- the 265th target gene is the hsa-miR-625-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 1.
- the 266th target gene is the hsa-miR-92a-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 1.
- the 267th target gene is the hsa-miR-422a gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Non-patent Document 3.
- the 268th target gene is the hsa-miR-642a-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof. To date, there has been no report that changes in the expression of genes or their transcripts can serve as markers for ovarian tumors.
- the 269th target gene is the hsa-miR-483-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 1.
- the 270th target gene is the hsa-miR-652-5p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a gene or a transcription product thereof can serve as a marker for ovarian tumors (Patent Document 4).
- the 271st target gene is an hsa-miR-24-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Patent Document 1.
- the 272nd target gene is an hsa-miR-23b-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Non-patent Document 3.
- the 273rd target gene is an hsa-miR-23a-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Non-patent Document 3.
- the 274th target gene is the hsa-miR-92b-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- a report that changes in the expression of a gene or a transcription product thereof can serve as a marker for ovarian tumors Non-patent Document 3).
- the 275th target gene is the hsa-miR-22-3p gene, a homologue thereof, a transcription product thereof, or a mutant or derivative thereof.
- the present invention relates to a marker for detecting an ovarian tumor or diagnosing an ovarian tumor, comprising at least one of the target nucleic acids.
- the invention relates to the use of at least one of the above target nucleic acids for detecting ovarian tumors or for diagnosing ovarian tumors.
- a nucleic acid for detection of ovarian tumor for example, a nucleic acid probe or primer that can be used for diagnosing an ovarian tumor, is a human-derived hsa- miR-4675, hsa-miR-4783-3p, hsa-miR-1228-5p, hsa-miR-4532, hsa-miR-6802-5p, hsa-miR-6784-5p, hsa-miR-3940-5p, hsa-miR-1307-3p, hsa-miR-8073, hsa-miR-3184-5p, hsa-miR-1233-5p, hsa-miR-6088, hsa-miR-5195-3p, hsa-miR-320b, hsa-miR-4649-5p,
- the target nucleic acid may be used in healthy subjects, benign bone and soft tissue tumors and patients with benign disease (or disease animals), and subjects with ovarian tumors compared to subjects with cancers other than ovarian cancer.
- the kit or device of the present invention can be used for body fluids derived from subjects suspected of having ovarian tumors (eg, humans) and healthy subjects, benign bone and soft tissue tumors and patients with benign diseases (or disease animals), and ovarian cancer.
- the nucleic acid probe or primer usable in the present invention specifically binds to, for example, a polynucleotide comprising a base sequence represented by at least one of SEQ ID NOs: 1 to 247, 251 and 268, or a complementary strand of the polynucleotide. It is a possible nucleic acid probe or a primer for amplifying a polynucleotide comprising a base sequence represented by at least one of SEQ ID NOs: 1 to 247, 251 and 268.
- the nucleic acid probe or primer that can be used in the present invention further includes, for example, a polynucleotide comprising a base sequence represented by at least one of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, or a complementary strand of the polynucleotide Or a primer for amplifying a polynucleotide consisting of a base sequence represented by at least one of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275 .
- the nucleic acid probe or primer is a polynucleotide group comprising a base sequence represented by any of SEQ ID NOs: 1 to 842, or a base sequence in which u is t in the base sequence, and
- the complementary polynucleotide group, the polynucleotide group that hybridizes with DNA comprising a base sequence complementary to the base sequence under stringent conditions (described later), the complementary polynucleotide group, and the polynucleotide group It includes a combination of one or a plurality of polynucleotides selected from the group of polynucleotides containing 15 or more, preferably 17 or more consecutive bases in the base sequence. These polynucleotides can be used as nucleic acid probes and primers for detecting the ovarian tumor marker, which is a target nucleic acid.
- examples of the nucleic acid probe or primer that can be used in the present invention are one or a plurality of polynucleotides selected from the group consisting of any of the following polynucleotides (a) to (e).
- A a polynucleotide comprising any one of SEQ ID NOS: 1 to 247, 251 and 268, or a nucleotide sequence wherein u is t in the nucleotide sequence, a variant thereof, a derivative thereof, 15 or more A fragment thereof containing a continuous base of
- B a polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268,
- C a polynucleotide comprising a base sequence complementary to the base sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268, or a base sequence in which u is t in the base sequence, variants thereof, A derivative thereof, or a fragment thereof comprising
- the nucleic acid probe or primer that can be used in the present invention further includes the following (f) to (j) in addition to at least one polynucleotide selected from any of the above-mentioned polynucleotides (a) to (e): Any of the polynucleotides shown can be included.
- Any of the above polynucleotides or fragments thereof used in the present invention may be DNA or RNA.
- the above-mentioned polynucleotide that can be used in the present invention can be prepared using a general technique such as a DNA recombination technique, a PCR method, or a method using a DNA / RNA automatic synthesizer.
- DNA recombination techniques and PCR methods are described, for example, in Ausubel et al., Current Protocols in Molecular Biology, John Willy & Sons, US (1993); Sambrook et al., Molecular Cloning A Laboratory, Ltd., 1998. Technology can be used.
- Such a nucleic acid probe or primer can be chemically synthesized using an automatic DNA synthesizer.
- the phosphoramidite method is used for this synthesis, and single-stranded DNA of up to about 100 bases can be automatically synthesized by this method.
- Automatic DNA synthesizers are commercially available from, for example, Polygen, ABI, Applied BioSystems, and the like.
- the polynucleotide of the present invention can also be prepared by a cDNA cloning method.
- a cDNA cloning method for example, microRNA Cloning Kit Wako can be used as the cDNA cloning technique.
- the sequence of the nucleic acid probe and primer for detecting the polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 1 to 275 does not exist in the living body as miRNA or a precursor thereof.
- the base sequences represented by SEQ ID NO: 128 and SEQ ID NO: 151 are generated from the precursor represented by SEQ ID NO: 405, and this precursor has a hairpin-like structure as shown in FIG.
- the base sequences represented by SEQ ID NO: 128 and SEQ ID NO: 151 have mismatch sequences with each other. For this reason, a completely complementary base sequence to the base sequence represented by SEQ ID NO: 121 or 151 is not naturally generated in vivo. Therefore, the nucleic acid probe and primer for detecting the base sequence represented by any one of SEQ ID NOs: 1 to 275 can have an artificial base sequence that does not exist in the living body.
- Ovarian tumor detection kit or device The present invention also provides a polynucleotide (including a variant, fragment, or derivative) that can be used as a nucleic acid probe or primer in the present invention for measuring a target nucleic acid that is an ovarian tumor marker.
- a kit or device for detecting ovarian tumors comprising one or more of:
- the target nucleic acid that is an ovarian tumor marker in the present invention is preferably selected from the following group A.
- Group A hsa-miR-4675, hsa-miR-4783-3p, hsa-miR-1228-5p, hsa-miR-4532, hsa-miR-6802-5p, hsa-miR-6784-5p, hsa-miR-3940- 5p, hsa-miR-1307-3p, hsa-miR-8073, hsa-miR-3184-5p, hsa-miR-1233-5p, hsa-miR-6088, hsa-miR-5195-3p, hsa-miR- 320b, hsa-miR-4649-5p, hsa-miR-6800-5p, hsa-miR-1343-3p,
- the additional target nucleic acid that can optionally be used for the measurement is preferably selected from group B below.
- Group B hsa-miR-320a, hsa-miR-663a, hsa-miR-328-5p, hsa-miR-128-2-5p, hsa-miR-125a-3p, hsa-miR-191-5p, hsa-miR- 92b-5p, hsa-miR-296-5p, hsa-miR-1246, hsa-miR-92a-2-5p, hsa-miR-128-1-5p, hsa-miR-1290, hsa-miR-211- 3p, hsa-miR-744-5p, hsa-miR-135a-3p, hsa-miR-451a, hsa-miR-625-3p, h
- the kit or device of the present invention is selected from a nucleic acid capable of specifically binding to the target nucleic acid that is the ovarian tumor marker or a nucleic acid for detecting the target nucleic acid, preferably the polynucleotides described in 2 above.
- a nucleic acid capable of specifically binding to the target nucleic acid that is the ovarian tumor marker or a nucleic acid for detecting the target nucleic acid preferably the polynucleotides described in 2 above.
- the kit or device of the present invention includes, for example, a base sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268, or a base sequence in which u is t in the base sequence ( Or a polynucleotide comprising (or consisting of) a complementary sequence thereof, a polynucleotide that hybridizes with these polynucleotides under stringent conditions, or a sequence of 15 or more of these polynucleotide sequences. At least one variant or fragment containing the selected base can be included.
- the kit or device of the present invention further includes, for example, a base sequence represented by any one of SEQ ID NOS: 248 to 250, 252 to 267, and 269 to 275, or a base sequence in which u is t in the base sequence ( Or a polynucleotide comprising (or consisting of) a complementary sequence thereof, a polynucleotide that hybridizes with these polynucleotides under stringent conditions, or a sequence of 15 or more of these polynucleotide sequences.
- a base sequence represented by any one of SEQ ID NOS: 248 to 250, 252 to 267, and 269 to 275 or a base sequence in which u is t in the base sequence ( Or a polynucleotide comprising (or consisting of) a complementary sequence thereof, a polynucleotide that hybridizes with these polynucleotides under stringent conditions, or a sequence of 15 or more of these polynucleotide
- the fragment that can be included in the kit or device of the present invention is, for example, one or more, preferably two or more polynucleotides selected from the group consisting of (1) and (2) below: (1) Sequence A polynucleotide comprising 15 or more consecutive bases in a base sequence represented by any one of Nos. 1 to 247, 251 and 268, wherein u is t or a complementary sequence thereof. (2) a polynucleotide comprising 15 or more consecutive bases in a base sequence represented by any one of SEQ ID NOS: 248 to 250, 252 to 267, and 269 to 275, wherein u is t or a complementary sequence thereof .
- the polynucleotide is a polynucleotide comprising a nucleotide sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268, or a nucleotide sequence in which u is t in the nucleotide sequence, its complementary A polynucleotide comprising a sequence, a polynucleotide that hybridizes with these polynucleotides under stringent conditions, or a variant containing 15 or more, preferably 17 or more, more preferably 19 or more consecutive bases thereof.
- the polynucleotide comprises a base sequence represented by any one of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, or a base sequence in which u is t in the base sequence.
- the fragment may be a polynucleotide comprising 15 or more, preferably 17 or more, more preferably 19 or more consecutive bases.
- the size of a polynucleotide fragment is, for example, 15 to less than the total number of bases in the sequence, 17 to less than the total number of bases in the sequence, and 19 to less than the total number of bases in the sequence.
- the number of bases in the range is, for example, 15 to less than the total number of bases in the sequence, 17 to less than the total number of bases in the sequence, and 19 to less than the total number of bases in the sequence. The number of bases in the range.
- kits or device of the present invention examples include, for example, one or two of the polynucleotides having the base sequences represented by SEQ ID NOs: 1 to 275 shown in Table 1 above. 3, 4, 4, 6, 7, 8, 9, 10, or more can be listed, but these are only examples and other All of the various possible combinations are intended to be encompassed by the present invention.
- the target nucleic acid combination in the kit or device for discriminating between ovarian tumor patients and healthy individuals includes, for example, the above two polynucleotides consisting of the base sequences represented by the sequence numbers shown in Table 1
- the combination of the above can be mentioned. More preferably, a combination of two or more of the above-mentioned polynucleotides consisting of the base sequence represented by the SEQ ID NO shown in Table 2 can be mentioned.
- any two or more of the above polynucleotides consisting of the base sequences represented by SEQ ID NOs: 1 to 203 and 248 to 268 can be combined. Among these, it is preferable to select at least one polynucleotide having a base sequence represented by SEQ ID NOs: 1 to 203, 251 and 268 newly found.
- a base sequence represented by a sequence number shown in Table 1 A combination of two or more of the above polynucleotides. More preferably, a combination of two or more of the above-mentioned polynucleotides consisting of the base sequences represented by the SEQ ID NOs shown in Table 6 can be mentioned.
- a base sequence represented by a sequence number shown in Table 1 A combination of two or more of the above polynucleotides. More preferably, two or more of the above-mentioned polynucleotides comprising the base sequence represented by the SEQ ID NO shown in Table 10 can be combined.
- any two of the polynucleotides selected from the group consisting of the nucleotide sequences can be combined. .
- cancer patients other than ovarian cancer For example, SEQ ID NOs: 2, 3, 4, 9, 11, 13, 15, 20, 33, 34, 38, 40, 44, 47, 56, 62, 68, 77, 78, 80, 82, 86, 89, 90, 91, 102, 104, 109, 117, 118, 135, 136, 145, 150, 157, 160, 161, 164, 169, 172, 196, 199, 211, 216, 217, 218, 220, 228, 229, 230, 231, 232, 233, 234, 235, 236, 237, 238, 239, 240, 241, 242, 243, 244, 245, 246, 24 , 250, 252, 256, 260, 267, 268, at least one polynucleotide
- ovarian tumor patients can be distinguished not only from healthy subjects but also from cancer patients other than ovarian cancer. It is more preferable to use a plurality (ie, 2 or more) of polynucleotides selected from nucleotide group 1.
- polynucleotides that are target nucleic acids having cancer types that can distinguish ovarian tumor patients from not only healthy subjects but also cancer patients other than ovarian cancer Among the combinations of a plurality of selected polynucleotides, for example, a group consisting of polynucleotides of SEQ ID NOs: 228, 9, 196, 229, 145 and 164 (hereinafter, this group is referred to as “specific polynucleotide group 2”).
- a combination comprising at least one polynucleotide selected from is more preferred.
- the number of combinations of the polynucleotides that are the target nucleic acids having the above cancer types is 1, or 2, 3, 4, 5, 6, 7, 8, 9, It may be any of 10 or more. A combination of two or more is preferable.
- the target nucleic acid combinations include, but are not limited to, a polynucleotide consisting of the base sequence represented by SEQ ID NO: 228 or a complementary sequence thereof, and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof.
- the combination with the polynucleotide which consists of a sequence is illustrated.
- target nucleic acids for example, but not limited to, a polynucleotide consisting of the base sequence represented by SEQ ID NO: 9 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- target nucleic acids for example, without limitation, a polynucleotide consisting of the base sequence represented by SEQ ID NO: 196 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- a polynucleotide consisting of the base sequence represented by SEQ ID NO: 196 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- Examples of combinations with polynucleotides comprising sequences are given below.
- (1) Combination of SEQ ID NOS: 256, 196 (2) Combination of SEQ ID NOS: 256, 196, 260 (3) Combination of SEQ ID NOS: 256, 196, 172 (4) Combination of SEQ ID NOS: 256, 196, 47 (5) Sequence Combination of Nos.
- target nucleic acids for example, without limitation, a polynucleotide consisting of the base sequence represented by SEQ ID NO: 229 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- a polynucleotide consisting of the base sequence represented by SEQ ID NO: 229 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- Examples of combinations with polynucleotides comprising sequences are given below.
- target nucleic acids for example, without limitation, a polynucleotide consisting of the base sequence represented by SEQ ID NO: 145 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- a polynucleotide consisting of the base sequence represented by SEQ ID NO: 145 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- Examples of combinations with polynucleotides comprising sequences are given below.
- target nucleic acids for example, without limitation, a polynucleotide consisting of the base sequence represented by SEQ ID NO: 164 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- a polynucleotide consisting of the base sequence represented by SEQ ID NO: 164 or a complementary sequence thereof and a polynucleotide selected from the specific polynucleotide group 1 or a complementary sequence thereof
- Examples of combinations with polynucleotides comprising sequences are given below. (1) Combination of SEQ ID NO: 11, 164, 260 (2) Combination of SEQ ID NO: 11, 164, 20 (3) Combination of SEQ ID NO: 145, 260, 164
- kits or devices of the present invention include known polynucleotides that allow ovarian tumor detection.
- polynucleotides that may be found in the future can be included.
- the kit or device of the present invention includes an antibody for measuring a known marker for ovarian tumor testing such as CA-125, in addition to the polynucleotide of the present invention described above and a variant or fragment thereof. Can be made.
- polynucleotides included in the kit of the present invention, and variants or fragments thereof can be packaged individually or in any combination in different containers.
- the kit of the present invention can include a kit for extracting nucleic acid (for example, total RNA) from body fluids, cells or tissues, a fluorescent substance for labeling, an enzyme and medium for nucleic acid amplification, instructions for use, and the like.
- nucleic acid for example, total RNA
- the device of the present invention is a device for measuring a cancer marker in which a nucleic acid such as a polynucleotide, a variant thereof, a derivative thereof, or a fragment thereof in the present invention described above is bound or attached to a solid phase, for example. is there.
- a nucleic acid such as a polynucleotide, a variant thereof, a derivative thereof, or a fragment thereof in the present invention described above is bound or attached to a solid phase, for example. is there.
- the material of the solid phase are plastic, paper, glass silicon, and the like. From the viewpoint of ease of processing, the preferable material of the solid phase is plastic.
- the shape of the solid phase is arbitrary, for example, a square shape, a round shape, a strip shape, a film shape and the like.
- Examples of the device of the present invention include a device for measurement by a hybridization technique, and specific examples include a blotting device, a nucleic acid array (for example, a
- the nucleic acid array technology uses a high-density dispenser called a spotter or arrayer on the surface of a solid phase that has been subjected to surface treatment such as introduction of functional groups such as L-lysine coat, amino group, and carboxyl group as necessary.
- the method of spotting nucleic acids the method of spraying nucleic acids onto a solid phase using an inkjet that ejects fine droplets from a nozzle with a piezoelectric element, the method of sequentially synthesizing nucleotides on a solid phase, etc.
- an array such as a chip is prepared by binding or attaching each of the nucleic acids one by one, and the target nucleic acid is measured using hybridization using this array.
- the kit or device of the present invention comprises at least one, preferably at least 2, more preferably at least 3, most preferably at least 5 to all polynucleotides of the above group A ovarian tumor marker miRNA, or A nucleic acid capable of specifically binding to each of the complementary strands of the polynucleotide;
- the kit or device of the present invention may further optionally include at least one, preferably at least 2, more preferably at least 3, most preferably at least 5 to all of the miRNAs that are Group B ovarian tumor markers as described above.
- a nucleic acid that can specifically bind to each of a polynucleotide or a complementary strand of the polynucleotide can be included.
- the kit or device of the present invention can be used for the detection of the following 4 ovarian tumors.
- the present invention further includes hsa-miR-4675, hsa in a sample using the above-described nucleic acid (or the kit or device of the present invention described in 3. above) that can be used in the present invention.
- the expression level of the gene in the specimen for example, for a specimen such as blood, serum, plasma, etc. collected from a subject suspected of suffering from ovarian tumor and a subject not suffering from ovarian tumor, the expression level of the gene in the specimen, If there is a difference in the expression level of the target nucleic acid in the sample using the control expression level of the subject not suffering from ovarian tumor (eg, comparing both expression levels), the subject Can be assessed as having
- the above method of the present invention enables early diagnosis of cancer with minimal invasiveness, high sensitivity and specificity, thereby leading to early treatment and improvement of prognosis, as well as monitoring disease aversion and surgical Enables monitoring of the effectiveness of radiotherapeutic and chemotherapeutic treatments.
- RNA extraction reagent liquid sample kit As a method for extracting genes derived from ovarian tumors from specimens such as blood, serum, plasma, etc. of the present invention, for extracting RNA in 3D-Gene (registered trademark) RNA extraction reagent liquid sample kit (Toray Industries, Japan) Although it is particularly preferable to adjust by adding a reagent, a general acidic phenol method (Acid Guanidinium-Phenol-Chloroform (AGPC) method) or Trizol (registered trademark) (Life Technologies) may be used. Alternatively, it may be prepared by adding an RNA extraction reagent containing acidic phenol such as Trizol (life technologies) or Isogen (Nippon Gene, Japan). Furthermore, kits such as miRNeasy (registered trademark) Mini Kit (Qiagen) can be used, but are not limited to these methods.
- AGPC Acid Guanidinium-Phenol-Chloroform
- Trizol registered trademark
- kits such as miRNeasy (registered trademark
- the present invention also provides the use of the kit or device of the present invention for in vitro detection of an expression product of an ovarian tumor-derived miRNA gene in a specimen derived from a subject.
- the kit or device is used as described above, and includes a single or any possible combination of polynucleotides that can be used in the present invention.
- the polynucleotide contained in the kit or device of the present invention can be used as a probe or primer.
- a primer Life Technologies' TaqMan (registered trademark) MicroRNA Assays, Qiagen's miScript PCR System, and the like can be used, but are not limited thereto.
- the gene expression level is determined by a quantitative technique such as a Northern blot method, Southern blot method, in situ hybridization method, Northern hybridization method, Southern hybridization method, or quantitative RT-PCR method.
- a quantitative technique such as a Northern blot method, Southern blot method, in situ hybridization method, Northern hybridization method, Southern hybridization method, or quantitative RT-PCR method.
- a known method for specifically detecting a specific gene such as an amplification technique and a method using a next-generation sequencer, it can be performed according to a conventional method, for example, using the primer or the probe.
- body fluid such as blood, serum, plasma, urine, etc. of the subject is collected according to the type of detection method used.
- total RNA prepared by the above-described method may be used, or various polynucleotides containing cDNA prepared based on the RNA may be used.
- the method, kit or device of the present invention is useful for diagnosis of ovarian tumors or detection of the presence or absence of morbidity. Specifically, in the detection of ovarian tumors using the method, kit or device, it is detected by the method using a specimen such as blood, serum, plasma, urine or the like from a subject suspected of suffering from ovarian tumors. Or by detecting the expression level of a gene detected using a nucleic acid probe or primer included in the kit or device in vitro.
- the expression level of a polynucleotide comprising, for example, the nucleotide sequence represented by one or more of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, is not a subject having an ovarian tumor (ie, a control animal) May be evaluated as having an ovarian tumor if it is statistically significantly higher than the expression level in a sample of blood, serum, plasma, urine, etc. it can.
- a test sample derived from a subject does not contain an ovarian tumor, or a method for detecting the presence of an ovarian tumor is performed by subject blood or serum.
- a body fluid such as plasma or urine is collected, and the expression level of the target gene (or target nucleic acid) contained therein is determined according to one or more polynucleotides (variants, fragments) selected from the polynucleotide group of the present invention. Or the like.) Or the presence of an ovarian tumor by measurement using a method.
- the ovarian tumor detection method of the present invention is a known or developing ovarian cancer-related therapeutic agent (for example, cisplatin, Including phosphamide, doxorubicin, etoposide, carboplatin, paclitaxel, combinations thereof, etc.), it can also be used for evaluating or diagnosing whether or not the disease has been improved or the degree of improvement.
- ovarian cancer-related therapeutic agent for example, cisplatin, Including phosphamide, doxorubicin, etoposide, carboplatin, paclitaxel, combinations thereof, etc.
- the method of the present invention includes, for example, the following steps (a), (b) and (c): (A) contacting a specimen from a subject with a polynucleotide of a kit or device of the present invention in vitro; (B) measuring the expression level of the target nucleic acid in the specimen using the polynucleotide as a nucleic acid probe or primer; (C) evaluating the presence or absence of ovarian tumors (cells) in the subject based on the results of (b); Can be included.
- the present invention provides miR-4675, miR-4783-3p, miR-1228-5p, miR-4532, miR-6802-5p, miR-6784-5p, miR-3940-5p, miR-1307. -3p, miR-8073, miR-3184-5p, miR-1233-5p, miR-6088, miR-5195-3p, miR-320b, miR-4649-5p, miR-6800-5p, miR-1343-3p , MiR-4730, miR-685-5p, miR-5100, miR-1203, miR-6756-5p, miR-373-5p, miR-1268a, miR-1260b, miR-4258, miR-4597-5p, miR -1469, miR-4515, miR-686 -5p, miR-6821-5p, miR-575, miR-6805-5p, miR-4758-5p, miR-3663-3p, miR-4530, miR
- evaluation is evaluation support based on the result of an in vitro examination that is not judged by a doctor.
- miR-4675 is hsa-miR-4675
- miR-4783-3p is hsa-miR-4783-3p
- miR-1228-5p is hsa.
- miR-4532 is hsa-miR-4532
- miR-6802-5p is hsa-miR-6802-5p
- miR-6784-5p is hsa-miR-6784-5p
- MiR-3940-5p is hsa-miR-3940-5p
- miR-1307-3p is hsa-miR-1307-3p
- miR-8073 is hsa-miR-8073
- miR-3184 ⁇ 5p is hsa-miR-3184-5p
- miR-1233-5p is hsa-m R-1233-5p
- miR-6088 is hsa-miR-6088
- miR-5195-3p is hsa-miR-5195-3p
- miR-320b is hsa-miR-320b
- miR- 4649-5p is hsa-miR-4649-5p
- miR-638 is hsa-miR-638
- MiR-2467-3p is hsa-miR-2467-3p
- miR-4728-5p is hsa-miR-4728-5p
- miR-5572 is hsa-miR-5572
- miR-6789- 5p is hsa-miR-6789-5p
- miR-8063 is hsa-miR-8063
- miR-4429 is hsa-miR-4429
- miR-6840-3p is hsa-miR-6840-3p
- MiR-4476 is hsa-miR-4476
- miR-675-5p is hsa-miR-675-5p
- miR-711 is hsa-miR-711
- miR-6875-5p is hsa- miR-6875-5p
- miR-3160-5p is hsa-
- the nucleic acid (eg, probe or primer) in the method of the present invention is, for example, a polynucleotide shown in any of the following (a) to (e): (A) a polynucleotide comprising the nucleotide sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268, or a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more A fragment thereof containing a continuous base of (B) a polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268, (C) a polynucleotide comprising a base sequence complementary to the base sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268, or a base sequence in which u is t in the base sequence, variants thereof, A derivative thereof, or a fragment thereof comprising
- the nucleic acid in the method of the present invention further includes miR-320a, miR-663a, miR-328-5p, miR-128-2-5p, miR-125a-3p, miR-191-5p, miR-92b-5p, miR-296-5p, miR-1246, miR-92a-2-5p, miR-128-1-5p, miR-1290, miR-211-3p, miR-744-5p, miR-135a-3p, miR- 451a, miR-625-3p, miR-92a-3p, miR-422a, miR-483-5p, miR-652-5p, miR-24-3p, miR-23b-3p, miR-23a-3p, miR- 92b-3p, at least one polynucleotide selected from miR-22-3p, or the polynucleotide It can include complementary strand capable of specifically binding nucleic acids.
- miR-320a is hsa-miR-320a
- miR-663a is hsa-miR-663a
- miR-328-5p is hsa-miR-328-5p
- miR-128-2 -5p is hsa-miR-128-2-5p
- miR-125a-3p is hsa-miR-125a-3p
- miR-191-5p is hsa-miR-191-5p
- miR-92b -5p is hsa-miR-92b-5p
- miR-296-5p is hsa-miR-296-5p
- miR-1246 is hsa-miR-1246
- miR-92a-2-5p is hsa MiR-92a-2-5p
- miR-128-1-5p is hsa-miR-128-1-5p
- miR-1 90 is hsa-miR-1290
- the nucleic acid is, for example, a polynucleotide shown in any of the following (f) to (j): (F) a polynucleotide comprising the nucleotide sequence represented by any of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, or a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, or a derivative thereof Or fragments thereof containing 15 or more consecutive bases, (G) a polynucleotide comprising the base sequence represented by any of SEQ ID NOs: 248 to 250, 252 to 267, and 269 to 275, (H) a polynucleotide comprising a base sequence complementary to the base sequence represented by any of SEQ ID NOS: 248 to 250, 252 to 267, and 269 to 275, or the base sequence in which u is t in the base sequence, A variant thereof, a derivative thereof
- specimens prepared from a biological tissue of a subject preferably ovarian tissue or fallopian tube tissue
- body fluid such as blood, serum, plasma, urine and the like.
- an RNA-containing sample prepared from the tissue a sample containing a polynucleotide further prepared therefrom, a body fluid such as blood, serum, plasma, urine, a part or all of a biological tissue of a subject, a biopsy, etc.
- a biological tissue or the like extracted by surgery from which a specimen for measurement can be prepared.
- a subject refers to a mammal such as, but not limited to, a human, monkey, mouse, rat, and the like, preferably a human.
- the steps can be changed according to the type of specimen used as a measurement target.
- the method for detecting an ovarian tumor is, for example, the following steps (a), (b) and (c): (A) RNA prepared from a subject sample (wherein for the quantitative RT-PCR of step (b), for example, the 3 ′ end of the RNA may be polyadenylated) or complementary transcribed therefrom Binding a polynucleotide (cDNA) to the polynucleotide of the kit of the invention, (B) Measure RNA derived from a specimen bound to the polynucleotide or cDNA synthesized from the RNA by hybridization using the polynucleotide as a nucleic acid probe or by quantitative RT-PCR using the polynucleotide as a primer. Step to do, (C) evaluating the presence or absence of ovarian tumor (or ovarian tumor-derived gene) based on the measurement result of (b) above, Can be included.
- A RNA prepared from a subject sample (wherein for the quantitative RT-PCR
- hybridization methods In order to measure the expression level of a target gene according to the present invention, or to detect, test, evaluate or diagnose ovarian tumors (or genes derived from ovarian tumors) in vitro, for example, various hybridization methods are used. Can do. As such a hybridization method, for example, Northern blot method, Southern blot method, DNA chip analysis method, in situ hybridization method, Northern hybridization method, Southern hybridization method and the like can be used. Further, a PCR method such as a quantitative RT-PCR method can be used in combination with or as an alternative to the hybridization method.
- nucleic acid probes complementary strand radioisotopes (such as 32 P, 33 P, 35 S ) or a fluorescent substance labeled with a, subject organism it was transferred like a nylon membrane by a conventional method
- a signal derived from the formed DNA / RNA duplex label is detected with a radiation detector (BAS-1800II (Fuji Film Co., Ltd.)
- a fluorescence detector STORM 865 (GE Healthcare), etc.
- RNA derived from a living tissue of a subject is collected, cDNA is prepared according to a 3′-end polyadenylation treatment or a reverse transcription method using a specific primer, and using this as a template, the region of each target gene marker is determined.
- a pair of primers from the detection kit of the present invention consisting of a positive strand and a reverse strand that bind to the cDNA are hybridized with cDNA and subjected to PCR by a conventional method so that amplification is possible.
- a method for detecting a double-stranded DNA or a double-stranded DNA can be exemplified.
- a method for detecting single-stranded or double-stranded DNA a method in which the PCR is performed using a primer previously labeled with a radioisotope or a fluorescent substance, a PCR product is electrophoresed on an agarose gel, and ethidium is used.
- a method for detecting double-stranded DNA by staining with bromide or the like, a method for detecting the produced single-stranded or double-stranded DNA by transferring it to a nylon membrane or the like according to a conventional method, and hybridizing with a labeled nucleic acid probe. Can be included.
- RNA chip or DNA chip in which the detection kit or device of the present invention is attached to a substrate (solid phase) as a nucleic acid probe (single strand or double strand) is used.
- the region where the nucleic acid probe is attached is called a probe spot, and the region where the nucleic acid probe is not attached is called a blank spot.
- a gene group immobilized on a substrate generally has a name such as a nucleic acid chip, a nucleic acid array, or a microarray.
- a DNA or RNA array includes a DNA or RNA macroarray and a DNA or RNA microarray.
- the term “chip” includes the array.
- 3D-Gene (registered trademark) Human miRNA Oligo chip Toray Industries, Inc., Japan
- 3D-Gene registered trademark Human miRNA Oligo chip
- the measurement of the DNA chip is not limited.
- a signal derived from a label of a detection kit or device is detected by an image detector (Typhoon 9410 (GE Healthcare), 3D-Gene (registered trademark) scanner (Toray Industries, Inc., The method of detecting and measuring can be illustrated.
- stringent conditions refers to the extent to which a nucleic acid probe is detectable relative to other sequences as described above (eg, average of background measurements + standard of background measurements). (Measurement value of error x 2 or more)).
- Stringent conditions are defined by hybridization and subsequent washing.
- the hybridization conditions are not limited, but for example, 30 to 60 ° C. and 1 to 24 hours in a solution containing SSC, surfactant, formamide, dextran sulfate, blocking agent and the like.
- 1 ⁇ SSC is an aqueous solution (pH 7.0) containing 150 mM sodium chloride and 15 mM sodium citrate, and the surfactant includes SDS (sodium dodecyl sulfate), Triton, or Tween.
- More preferable hybridization conditions include 3 to 10 ⁇ SSC and 0.1 to 1% SDS.
- Washing conditions after hybridization include, for example, a solution containing 0.5 ⁇ SSC at 30 ° C. and 0.1% SDS, and 0.2 at 30 ° C. There may be mentioned conditions such as continuous washing with a solution containing x SSC and 0.1% SDS and a 0.05 x SSC solution at 30 ° C. It is desirable that the complementary strand maintain a hybridized state with the target positive strand even when washed under such conditions.
- a complementary strand a strand consisting of a base sequence that is completely complementary to the target positive strand base sequence, and at least 80%, preferably at least 85%, more preferably, the strand. Examples thereof include a chain consisting of a base sequence having at least 90% or at least 95% identity.
- a PCR buffer having a composition such as 10 mM Tris-HCL (pH 8.3), 50 mM KCL, 1 to 2 mM MgCl 2 is used.
- the treatment may be performed for 15 seconds to 1 minute at a Tm value calculated from the primer sequence +5 to 10 ° C.
- Tm value 2 ⁇ (number of adenine residues + number of thymine residues) + 4 ⁇ (number of guanine residues + number of cytosine residues).
- TaqMan registered trademark
- MicroRNA Assays Life Technologies
- LNA registered trademark
- MicroRNA PCR Exiqon
- Ncode registered trademark
- miRNA qRT-PCT kit A commercially available measurement kit specially devised to quantitatively measure miRNA, such as (Invitrogen) may be used.
- the gene expression level may be measured using a sequencer in addition to the hybridization method described above.
- a sequencer any of the DNA sequencer as the first generation based on the Sanger method, the second generation with a short read size, and the third generation with a long read size can be used (the second generation and the third generation). Also referred to herein as “next generation sequencer”, including generational sequencers).
- Ion Proton / Ion PGM / Ion S5 / S5 XL (Thermo Fisher Scientific)
- PacBio RS II / Sequel Pacific Bioscience
- nanopore sequencer a commercially available measurement kit specially devised to measure miRNA using MinION (Oxford Nanopore Technologies) or the like may be used.
- the next-generation sequence is a method for acquiring sequence information using a next-generation sequencer, and is characterized in that a huge number of sequence reactions can be performed in parallel as compared to the Sanger method (for example, Rick Kamps et al. , Int. J. Mol. Sci., 2017, 18 (2), p.308 and Int. Neurolol. J., 2016, 20 (Suppl. 2), S76-83).
- the next generation sequence for miRNA first, an adapter sequence having a predetermined base sequence is added, and then, before or after the addition of the sequence, the total RNA is reverse-transcribed to cDNA.
- a cDNA derived from a specific target miRNA may be enriched or concentrated by PCR or using a probe or the like.
- the details of the subsequent sequencing step vary depending on the type of the next-generation sequencer, but typically, the sequencing reaction is performed by connecting to a substrate via an adapter sequence and using the adapter sequence as a priming site. For details of the sequence reaction, see, for example, Rick Kamps et al. See (above). Finally, data output is performed. In this step, a collection of sequence information (reads) obtained by the sequence reaction is obtained. For example, in a next-generation sequence, a target miRNA can be identified based on sequence information, and the expression level can be measured based on the number of reads having the sequence of the target miRNA.
- Calculation of gene expression level is not limited, but for example, statistical analysis of gene expression microarray data (Speed T., Chapman and HalliCensalCensitiveChemistralus. Et al., Blackwell publishing) can be used in the present invention.
- a spot can be regarded as a detection spot.
- the average value of the measured value of the blank spot can be regarded as the background, and can be subtracted from the measured value of the probe spot to obtain the gene expression level.
- the missing value of the gene expression level is excluded from the analysis target, preferably replaced with the minimum value of the gene expression level in each DNA chip, or more preferably 0.1 from the logarithmic value of the minimum value of the gene expression level. Can be replaced with the subtracted value.
- the number of samples to be measured is 20% or more, preferably 50% or more, more preferably 80% or more, 2 6, preferably 2 8, more preferably 2 Only genes having a gene expression level of the 10th power or higher can be selected as an analysis target. Examples of normalization of gene expression levels include, but are not limited to, global normalization and quantile normalization (Bolstad, B. M. et al., 2003, Bioinformatics, Vol. 19, p185-193).
- the present invention also includes, for example, the gene marker (or target nucleic acid) of the present invention, a nucleic acid (or a detection or diagnostic polynucleotide) for detecting the gene marker, a kit, a device (for example, a chip), or a combination thereof.
- a device for example, a chip
- a combination thereof Use to measure the expression level of a target gene or gene in a specimen derived from a subject, and a specimen derived from a subject known to have an ovarian tumor (eg, a patient) and a subject not suffering from an ovarian tumor
- the discriminant (discriminant function) described above was created as a teacher sample and can discriminate the presence or absence of ovarian tumors. Substituting the expression level of the target gene in a subject-derived specimen, thereby evaluating the presence or absence of the ovarian tumor, in the subject Detect (or, to aid detection) to provide a method.
- the present invention further includes, for example, the gene marker (or target nucleic acid) of the present invention, a nucleic acid (or a polynucleotide for detection or diagnosis) for detecting the gene marker, a kit, a device (for example, a chip), or their
- the combination is used to measure in vitro the expression level of a target gene in a plurality of specimens that are known to determine or evaluate that a subject contains and / or does not contain an ovarian tumor.
- the third step is to measure in vitro
- the discriminant obtained in the second step is the third step.
- the method is provided comprising four steps.
- the target gene can be detected by, for example, the polynucleotide for detection or diagnosis, the polynucleotide contained in the kit or device, and a variant or fragment thereof.
- the discriminant is any discriminant analysis method that can create a discriminant that discriminates the presence or absence of an ovarian tumor, for example, Fisher discriminant analysis, nonlinear discriminant analysis by Mahalanobis distance, neural Although a discriminant can be created using a network, Support Vector Machine (SVM), etc., it is not limited to these specific examples.
- SVM Support Vector Machine
- Linear discriminant analysis is a method of discriminating group membership using Formula 1 as a discriminant when the boundary of grouping is a straight line or a hyperplane.
- x is an explanatory variable
- w is a coefficient of the explanatory variable
- w 0 is a constant term.
- the value obtained by the discriminant is called a discriminant score, and the measured value of a newly given data set is substituted into the discriminant as an explanatory variable, and the grouping can be discriminated by the code of the discriminant score.
- Fisher's discriminant analysis which is a type of linear discriminant analysis, is a dimension reduction method for selecting a dimension suitable for class discrimination. Focusing on the variance of composite variables, the data with the same label is used. Combining highly discriminating synthetic variables by minimizing the variance (Venables, W. N. et al., Modern Applied Statistics with S. Fourth edition. Springer., 2002).
- a projection direction w that maximizes Equation 2 is obtained.
- ⁇ is the average of inputs
- ng is the number of data belonging to class g
- ⁇ g is the average of inputs of data belonging to class g.
- the numerator and denominator are the inter-class variance and intra-class variance when the data is projected in the direction of the vector w, respectively, and the discriminant coefficient wi is obtained by maximizing this ratio.
- the Mahalanobis distance is calculated by Equation 3 in consideration of data correlation, and can be used as a nonlinear discriminant analysis for discriminating a group having a close Mahalanobis distance from each group as a belonging group.
- ⁇ is the center vector of each group
- S ⁇ 1 is the inverse matrix of the variance-covariance matrix of that group.
- the center vector is calculated from the explanatory variable x, and an average vector or a median vector can be used.
- a boundary surface called a hyperplane is used to correctly classify the data set into a known grouping, with specific data items in the data set with a known grouping as explanatory variables and the grouping to be classified as an objective variable. And determine a discriminant for classifying data using the boundary surface.
- the discriminant can determine the grouping by substituting the measured value of the newly given data set into the discriminant as an explanatory variable. Further, the discrimination result at this time may be a group to be classified, may be a probability of being classified into a group to be classified, or may be a distance from a hyperplane.
- a method for dealing with a non-linear problem a method is known in which a feature vector is non-linearly transformed into a higher dimension and linear identification is performed in the space.
- An expression in which the inner product of two elements in a non-linearly mapped space is expressed only by the input in the original space is called a kernel.
- a kernel a linear kernel, RBF (Radial Basis Function) Kernel and Gaussian kernel.
- the optimal discriminant that is, the discriminant, can be constructed only by calculating the kernel while avoiding the calculation of the features in the mapped space while actually mapping in high dimensions by the kernel (for example, Hideki Aso et al.
- C-support vector classification (C-SVC), a kind of SVM method, creates a hyperplane by learning with two explanatory variables to determine which group an unknown data set falls into (C. Cortes et al., 1995, Machine Learning, 20, pp. 273-297).
- ovarian tumor patients are divided into two groups: ovarian tumor patients and subjects not suffering from ovarian tumors.
- ovarian histology can be used as a standard for determining that the subject is suffering from ovarian tumor or not suffering from ovarian tumor.
- a data set (hereinafter referred to as “learning sample group”) composed of comprehensive gene expression levels of the two groups of serum-derived specimens is prepared, and there is a clear difference in gene expression levels between the two groups.
- the discriminant by C-SVC is determined with the gene as the explanatory variable and the grouping as the target variable (eg, -1 and +1).
- Equation 4 is an objective function to be optimized, where e is all input vectors, y is an objective variable, a is a Lagrange undetermined multiplier vector, Q is a positive definite matrix, and C is a parameter for adjusting the constraint condition.
- Equation 5 is the discriminant finally obtained, and the group to which it belongs can be determined by the sign of the value obtained by the discriminant.
- x is a support vector
- y is a label indicating group membership
- a is a corresponding coefficient
- b is a constant term
- K is a kernel function.
- Equation 6 the RBF kernel defined by Equation 6 can be used.
- x represents a support vector
- ⁇ represents a kernel parameter that adjusts the complexity of the hyperplane.
- methods such as neural networks, k-neighbors, decision trees, and logistic regression analysis are selected as methods for determining or evaluating whether a subject-derived specimen contains or does not contain ovarian tumors can do.
- the method of the invention comprises, for example, the following steps (a), (b) and (c): (A) the expression level of the target gene in a specimen that is already known to be derived from an ovarian tumor patient and is a subject that does not contain an ovarian tumor; Measuring using a device (eg a DNA chip), (B) creating a discriminant of the above formulas 1 to 3, 5 and 6 from the measured value of the expression level measured in (a), (C) The expression level of the target gene in the specimen derived from the subject is measured using the detection or diagnostic polynucleotide, kit or device (for example, DNA chip) according to the present invention, and the discrimination created in (b) Substituting them into the formula and determining or assessing that the subject contains or does not contain an ovarian tumor based on the results obtained, or the ovarian tumor-derived expression level is affected by the ovarian tumor A step of evaluating relative to a control from a non-subject (eg, including a
- x in the formulas 1 to 3, 5 and 6 is an explanatory variable, and is obtained by measuring a polynucleotide selected from the polynucleotides as the target nucleic acid described in Section 2 above or a fragment thereof.
- the explanatory variable for determining the presence or absence of the ovarian tumor of the present invention is a gene expression level selected from the following (1) or (2).
- (1) Measured by any of nucleic acids (eg, DNA, RNA, etc.) containing 15 or more consecutive bases in the base sequence represented by any one of SEQ ID NOs: 1 to 247, 251 and 268 or its complementary sequence Gene expression level in sera of ovarian tumor patients and subjects not suffering from ovarian tumors.
- nucleic acids eg, DNA, RNA, etc.
- SEQ ID NOS: 248 to 250, 252 to 267, and 269 to 275, or a complementary sequence thereof Gene expression level in serum of ovarian tumor patients and subjects not suffering from ovarian tumors as measured by
- a discriminant using one or more gene expression levels as explanatory variables is necessary. is there.
- the expression level between two groups of ovarian tumor patient groups and subject groups not suffering from ovarian tumors is clearly defined. It is necessary to use genes with differences in discriminants.
- the gene used for the explanatory variable of the discriminant is determined as follows. First, using the data set of the exhaustive gene expression level of the ovarian tumor patient group as the learning group and the exhaustive gene expression level of the subject group not affected by the ovarian tumor, the P value of the t-test, which is a parametric analysis, non-parametric analysis Using the P value of Mann-Whitney U test or the P value of Wilcoxon test, the difference in expression level of each gene between the two groups is determined.
- Bonferroni correction for example, by multiplying the P value obtained by the test by the number of test repetitions, that is, the number of genes used in the analysis, and comparing it with the desired significance level, the first type of error in the entire test is obtained. Probability of occurrence can be suppressed.
- the absolute value (Fold change) of the median expression ratio of each gene expression level is not between the gene expression level of the ovarian tumor patient group and the gene expression level of the subject group not suffering from ovarian tumor instead of the test. It is also possible to select a gene to be calculated and used as an explanatory variable of the discriminant equation. Alternatively, the ROC curve may be created using the gene expression levels of the ovarian tumor patient group and the subject group not affected by the ovarian tumor, and the gene used as the explanatory variable of the discriminant may be selected based on the AUROC value.
- a discriminant that can be calculated by the above-described various methods is created using an arbitrary number of genes having a large difference in gene expression level obtained here.
- a method of constructing a discriminant that obtains the maximum discriminating accuracy for example, a method of constructing a discriminant with any combination of genes satisfying the significance level of the P value, or a gene used to create a discriminant, gene expression There is a method in which evaluation is repeated while increasing one by one in descending order of the amount of difference (Furey TS. Et al., 2000, Bioinformatics., Vol. 16, p906-14).
- the discrimination result of the group to which the user belongs is calculated. That is, the diagnostic gene set and ovary that can detect a more unbiased ovarian tumor by evaluating the discriminant constructed using the found diagnostic gene set and the diagnostic gene set in an independent sample group A method for discriminating tumors can be found.
- the Split-sample method for evaluating the discriminating performance (generalization) of the discriminant is preferable to use the Split-sample method for evaluating the discriminating performance (generalization) of the discriminant. That is, the data set is divided into a learning sample group and a test sample group, the gene selection and the discriminant formula are made by statistical test in the learning sample group, and the test sample group is discriminated by the discriminant formula and the test sample group The accuracy, sensitivity, and specificity are calculated using the true group to which the group belongs, and the discrimination performance is evaluated. On the other hand, without dividing the data set, the selection of genes and the creation of discriminants by statistical tests are performed using all the samples, and the newly prepared samples are discriminated by the discriminants to increase the accuracy, sensitivity and specificity. It is also possible to calculate and evaluate the discrimination performance.
- the present invention relates to polynucleotides for diagnosing or detecting diseases useful for diagnosis and treatment of ovarian tumors, methods for detecting ovarian tumors using the polynucleotides, kits for detecting or diagnosing ovarian tumors containing the polynucleotides, and Provide a device.
- a diagnostic gene that shows an accuracy exceeding the diagnostic method for ovarian tumors using the existing tumor marker CA-125 and to create a discriminant
- a known tumor such as CA-125 is used.
- the gene expressed in the serum from patients whose final ovarian tumor was revealed by close examination such as computed tomography using contrast medium, and ovary By comparing expressed genes in serum from subjects not suffering from tumors, diagnostic gene sets and discriminants that show an accuracy exceeding CA-125 can be constructed.
- one or more of the above polynucleotides based on the base sequence represented by any of SEQ ID NOs: 1 to 247, 251 and 268 as described above, and optionally SEQ ID NOs: 248 to 250, 252 to 267 And any combination from one or more of the above-mentioned polynucleotides based on the nucleotide sequence represented by any of 269 to 275 is used as a diagnostic gene set.
- a discriminant is constructed using the expression level of the diagnostic gene set in the specimen derived from the ovarian tumor patient and the specimen derived from the subject not suffering from the ovarian tumor. As a result, by measuring the expression level of the diagnostic gene set of an unknown sample, it is possible with a maximum accuracy of 100% that the subject from which the unknown sample is derived contains or does not contain an ovarian tumor. Can be distinguished.
- breast cancer, biliary tract cancer, pancreatic cancer, colon cancer, esophageal cancer, stomach cancer, liver cancer, lung cancer patients are collectively referred to as “cancer patients other than ovarian cancer”.
- RNA extraction reagent liquid sample kit Toray Industries, Inc. (Japan)
- total RNA was obtained according to the protocol specified by the company.
- RNA obtained from the serum of a total of 1,602 people including 400 healthy subjects, 368 benign bone and soft tissue tumors and mammary benign disease patients, 434 ovarian tumor patients, and 400 cancer patients other than ovarian cancer were used as specimens.
- 3D-Gene (registered trademark) miRNA Labeling kit Toray Industries, Inc. was used to fluorescently label miRNA based on the protocol established by the company.
- 3D-Gene Human miRNA Oligo chip (Toray Industries, Inc.) equipped with a probe having a sequence complementary to 2,565 kinds of miRNAs among miRNAs registered in miRBBase release 21 ) And hybridization and washing after hybridization under stringent conditions based on the protocol defined by the company.
- the DNA chip was scanned using a 3D-Gene (registered trademark) scanner (Toray Industries, Inc.), an image was acquired, and the fluorescence intensity was digitized with 3D-Gene (registered trademark) Extraction (Toray Industries, Inc.).
- the digitized fluorescence intensity was converted to a logarithmic value with a base of 2 to obtain the gene expression level, the blank value was subtracted, and the missing value was replaced with a signal value of 0.1.
- comprehensive miRNA gene expression levels were obtained for the sera of 400 healthy subjects, 368 patients with benign bone and soft tissue tumors and mammary diseases, 434 ovarian tumor patients, and 400 cancer patients other than ovarian cancer. It was. Next, 70% of the samples in each sample group were assigned to the learning sample group, and 30% of the samples were assigned to the test sample group.
- 280 healthy individuals, 257 patients with benign bone and soft tissue tumors and benign diseases, 303 patients with ovarian tumors, and 280 cancer patients other than ovarian cancer were used as learning sample groups.
- the test specimen group was 111 patients with benign disease, 131 patients with ovarian tumors, and 120 cancer patients other than ovarian cancer.
- the calculation and statistical analysis using the expressed gene expression level of miRNA were performed in R language 3.3.1 (R Core Team (2016). R: A language and environment for statistical computing, R Foundation for Statistical Computation, URL https://www.R-project.org/.) And MASS package 7.3.45 (Venables, WN & Ripley, BD (2002) Modern Applied Statistics with S. D Springer, New York, ISBN 0-387-95457-0).
- Example 1 ⁇ Selection of ovarian tumor gene marker by comparison with healthy subjects and evaluation method of ovarian tumor discrimination performance by single gene marker>
- miRNAs showing significant differences in expression levels between ovarian tumor patients and healthy subjects were selected as ovarian tumor gene markers, and a discriminant based on one or two gene markers was created using a learning sample group.
- determination performance of the ovarian tumor patient and healthy subject by said gene marker was evaluated by calculating the discrimination
- the miRNA expression levels of the learning sample group and the test sample group obtained in the above reference example were combined and normalized by global normalization.
- the gene for diagnosis was selected.
- genes having a gene expression level of 2 6 or more in 50% or more of specimens were selected in either the ovarian tumor patient group or the healthy subject group .
- a two-tailed t-test assuming equal variance is performed, and Bonferroni corrected P value is 0.01.
- the genes that were less than were selected.
- genes whose ovarian tumor patient group and healthy group log expression converted difference (fold change) absolute value is greater than 0.5 are detected as ovarian tumors. Were selected as diagnostic markers for healthy individuals, and the results are shown in Table 2.
- a gene newly found as a marker for examining the presence or absence of ovarian tumors is a polynucleotide comprising a nucleotide sequence represented by SEQ ID NOs: 1 to 203.
- the discriminant is created by inputting the gene expression level of the polynucleotide comprising the nucleotide sequence represented by any of SEQ ID NOs: 1 to 203 and 248 to 268 found above into Equation 2, and the calculated learning
- Table 3 shows the discriminant coefficients and constant terms at that time.
- the accuracy, sensitivity and specificity of the test sample group were calculated, and the discriminating performance of the selected polynucleotide was verified with an independent sample (Table 3).
- the target value of ovarian tumor detection was calculated using the threshold value (7.75) for discriminating both groups set in the learning sample group. In this case, 127 cases are true positive, 117 cases are true negative, 3 cases are false positives, and 4 cases are false negatives. From these values, the detection performance is 97.2% accuracy, 96.9% sensitivity, and 97.5% specificity. Obtained.
- the detection performance of all the polynucleotides shown in SEQ ID NOs: 1 to 203 and 248 to 268 was calculated and listed in Table 3.
- Non-patent Document 4 a polynucleotide comprising these sequences alone has a sensitivity higher than that of CA-125. It was proved that ovarian tumors could be identified.
- SEQ ID NO indicates a combination of a plurality of used polynucleotides by SEQ ID NO.
- all polynucleotides consisting of the nucleotide sequences represented by SEQ ID NOs: 1 to 203 and 248 to 268 can be used for ovarian tumor patients and healthy individuals by using a combination of one, two or more. It was proved that the gene can be distinguished with high accuracy.
- Example 2 ⁇ Selection of ovarian tumor gene marker by comparison with benign bone and soft tissue tumor and breast benign disease patient and evaluation method of ovarian tumor discrimination performance by single gene marker>
- miRNAs showing significant differences in expression levels in ovarian tumors, benign bone and soft tissue tumors and milk benign diseases are selected as ovarian tumor gene markers, and one or two gene markers are used using a learning sample group. After creating the discriminant, the discrimination performance of the test sample group was calculated to evaluate the discrimination performance of ovarian tumors, benign bone and soft tissue tumors and milk benign diseases by the above gene markers.
- the miRNA expression levels of the learning sample group and the test sample group obtained in the above reference example were combined and normalized by global normalization.
- the gene for diagnosis was selected.
- gene expression level of 2 6 or more in 50% or more specimens in either ovarian tumor patient group or benign bone and soft tissue tumor group and breast benign disease patient group Only genes with a were selected.
- a two-tailed t-test assuming equal variance is performed and Bonferroni correction is performed. Genes with a P value of less than 0.01 were selected.
- the absolute value of the difference between the logarithmically transformed gene expression (Fold change) between the ovarian tumor patient group and the benign bone and soft tissue tumor group and the benign disease patient group is 0.
- Genes greater than 5 were selected as diagnostic markers for ovarian tumors, benign bone and soft tissue tumors and milk benign diseases and the results are listed in Table 6.
- genes newly found as markers for examining the presence or absence of ovarian tumors are SEQ ID NOs: 4, 9, 14, 17, 18, 20, 29, 40, 47, 62, 66, 69, 78, 82. 83, 86, 87, 89, 116, 125, 127, 132, 135, 136, 147, 163, 164, 167, 174, 177, 184, 193, 204-215, a polynucleotide comprising the nucleotide sequence It is.
- a discriminant for discriminating the presence or absence of ovarian tumors by Fisher's discriminant analysis was created. That is, SEQ ID NOs: 4, 9, 14, 17, 18, 20, 29, 40, 47, 62, 66, 69, 78, 82, 83, 86, 87, 89, 116, 125, found above 127, 132, 135, 136, 147, 163, 164, 167, 174, 177, 184, 193, 204-215, 248, 256, 260, 264, 265, 268, 269 and 270
- the discriminant is created by inputting the gene expression level of the polynucleotide comprising the base sequence into Formula 2, and the calculated accuracy, sensitivity and specificity in the learning group are shown in Table 7. Table 8 shows the discriminant coefficients and constant terms at that time.
- the accuracy, sensitivity and specificity of the test sample group were calculated, and the discriminating performance of the selected polynucleotide was verified with an independent sample (Table 7).
- the target value for detecting ovarian tumors was calculated using a threshold (11.59) for discriminating both groups set in the learning sample group.
- Example 3 ⁇ Selection of ovarian tumor gene marker by comparison with cancer patients other than ovarian cancer and evaluation method of ovarian tumor discrimination performance by single gene marker>
- the miRNA is selected as an ovarian tumor gene marker
- a discriminant is created using one or two gene markers using the learning sample group, and then the discrimination accuracy of the test sample group is calculated.
- the discrimination accuracy of the test sample group is calculated.
- the miRNA expression levels of the learning sample group and the test sample group obtained in the above reference example were combined and normalized by global normalization.
- the gene for diagnosis was selected.
- the gene expression level of 2 to the 6th power or higher in 50% or more specimens in either the ovarian tumor patient group or the cancer patient group other than ovarian cancer Only those genes that had it were selected.
- a two-tailed t-test assuming equal variance was performed and Bonferroni correction was performed.
- a gene having a P value of less than 0.01 was selected.
- the absolute value of the difference in the expression level of the logarithmically transformed gene expression (Fold change) between the ovarian tumor patient group and the cancer patient group other than ovarian cancer is 0.5.
- Larger genes were selected as diagnostic markers for cancers other than ovarian tumors and ovarian cancer, and the results are listed in Table 10.
- genes newly found as markers for examining the presence of ovarian tumors are SEQ ID NOs: 4, 11, 17, 68, 69, 76, 78, 82, 85, 89, 99, 103, 105, 109. 127, 128, 135, 145, 160, 164, 169, 196, 198, 207, 211, 216 to 227.
- the accuracy, sensitivity, and specificity of the test sample group were calculated, and the discriminating performance of the selected polynucleotide was verified with an independent sample (Table 11).
- the target value of ovarian tumor detection was calculated using the threshold value (8.07) for discriminating both groups set in the learning sample group. 97 cases of true positive, 74 cases of true negative, 46 cases of false positive, and 34 cases of false negative. From these values, the detection performance is as follows: accuracy 68.1%, sensitivity 74.0%, specificity 61.7%. Obtained.
- SEQ ID NOs: 4, 11, 17, 68, 69, 76, 78, 82, 85, 89, 99, 103, 105, 109, 127, 128, 135, 145, 160, 164, 169, 196, 198 207, 211, 216-227 have the sensitivity of 65.6%, 62.6%, 74.0%, 74.8%, 67.9%, 63.4%, 74 respectively.
- the number of all polynucleotides is 1, 2 or more. This is a gene or nucleic acid that can distinguish ovarian tumor patients and cancer patients other than ovarian cancer with high accuracy.
- Example 4 ⁇ Evaluation method of ovarian tumor discrimination performance by combining multiple genetic markers>
- healthy subjects benign bone and soft tissue tumors and patients with benign diseases, and cancer patients other than ovarian cancer (ie, suffering from ovarian tumors) used in Examples 1, 2 and 3 above.
- cancer patients other than ovarian cancer ie, suffering from ovarian tumors
- Non-subject subjects were defined as negative sample groups, and ovarian tumor patients as positive sample groups, and combinations of genetic markers with high ovarian tumor discrimination performance were searched.
- test specimen groups the miRNA expression levels in the sera of 120 healthy individuals, 111 benign bone and soft tissue tumors and 111 benign disease patients, 131 ovarian tumor patients, and 120 cancer patients other than ovarian cancer were used.
- gene expression should be greater than or equal to 2 6 in more than 50% of specimens in either ovarian tumor patients or non-ovarian tumor subjects. Only genes with quantity were selected.
- Specificity polynucleotide group 1 includes SEQ ID NOs: 2, 3, 4, 9, 11, 13, 15, 20, 33, 34, 38, 40, 44, 47, 56, 62, obtained in Example 1.
- nucleotides hsa-miR-6794-5p, hsa-miR-4687-3p, hsa-miR-6743-5p, hsa-mi -6771-5p, hsa-miR-3141,
- SEQ ID NOS: 228, 229, 230, 231, 232, 233, 234, 235, 236, 237, 238, 239, 240, 241, 242, 243, 244, 245 which are genes newly found in this example
- Each of the polynucleotides consisting of the nucleotide sequences represented by 246, 247 is a gene newly found as a marker for examining the presence or absence of an ovarian tumor.
- polynucleotides for example, four polynucleotides having the base sequences represented by SEQ ID NOs: 260, 256, 20, 89 have sensitivity of 87.0%, 80.9%, 77. 9% and 77.9% (Table 15).
- a polynucleotide comprising these sequences alone has a sensitivity higher than that of CA-125. It was proved that ovarian tumors could be identified.
- the discriminant created by combining the measurement values of the base sequences represented by SEQ ID NOs: 256, 196, 260, 268 and 252 was used for 303 ovarian tumor patients in the learning sample group and 817 subjects not suffering from ovarian tumors.
- the upper part of FIG. 3 shows that the discrimination score is significantly separated. This result was also reproducible in the test sample group (bottom of FIG. 3).
- 94 formulas with a discrimination accuracy of 80% or more in both the learning group and the verification group are 94 formulas, of which 93 formulas are target nucleic acids as explanatory variables. , SEQ ID NO: 228, 9, 196, 229, 145, or 164, or a polynucleotide consisting of a complementary sequence thereof.
- the base represented by SEQ ID NO: 228, 9, 196, 229, 145 or 164 which is included in the specific polynucleotide group 1 among the combination of a plurality of polynucleotides selected from the specific polynucleotide group 1
- the presence or absence of ovarian tumors can be specifically detected with high accuracy It was identifiable.
- the discrimination accuracy when measured using a polynucleotide comprising the base sequence represented by SEQ ID NO: 228 or its complementary sequence as the target nucleic acid is shown in Table 16-1.
- Table 16-1 When measured using a combination of one, two, three, four or five genes including a polynucleotide consisting of the base sequence represented by SEQ ID NO: 228 or a complementary sequence thereof, a test sample group, respectively The highest accuracy was 58.9%, 81.7%, 85.5%, 89.8%, 91.3%.
- Table 16-2 shows the discrimination accuracy when measured using a polynucleotide consisting of the base sequence represented by SEQ ID NO: 9 or its complementary sequence as the target nucleic acid.
- a test sample group When measured using a combination of one, two, three, four, or five genes including a polynucleotide consisting of the base sequence represented by SEQ ID NO: 9 or a complementary sequence thereof, a test sample group, respectively The highest accuracy was 66.8%, 79.0%, 85.7%, 89.8%, and 91.1%.
- Table 16-3 shows the discrimination accuracy when measured using a polynucleotide consisting of the base sequence represented by SEQ ID NO: 196 or its complementary sequence as the target nucleic acid.
- a test sample group When measured using a combination of one, two, three, four, or five genes including a polynucleotide consisting of the base sequence represented by SEQ ID NO: 196 or its complementary sequence, a test sample group, respectively The highest accuracy was 58.9%, 84.4%, 87.6%, 90.2%, and 92.3%.
- Table 16-4 shows the discrimination accuracy when measured using a polynucleotide consisting of the base sequence represented by SEQ ID NO: 229 or its complementary sequence as the target nucleic acid.
- a test sample group When measured using a combination of one, two, three, four or five genes including a polynucleotide consisting of the base sequence represented by SEQ ID NO: 229 or its complementary sequence, a test sample group, respectively The highest accuracy was 74.9%, 81.5%, 85.9%, 89.4%, 91.3%.
- Table 16-5 shows the discrimination accuracy when measured using a polynucleotide consisting of the base sequence represented by SEQ ID NO: 145 or its complementary sequence as the target nucleic acid.
- the precision was 70.3%, 82.6%, 87.3%, and 89.8%.
- Table 16-6 shows the discrimination accuracy when measured using a polynucleotide consisting of the base sequence represented by SEQ ID NO: 164 or its complementary sequence as the target nucleic acid.
- the maximum accuracy in the test sample group is 53.1. %, 81.1%, and 85.5%.
- Specific polynucleotide groups 1 and 2 obtained in this example can specifically distinguish ovarian tumor patients from healthy individuals, benign bone and soft tissue tumors and patients with benign disease, and cancer patients other than ovarian cancer. It can be said to be a gene group.
- a higher ovarian tumor discrimination performance can be obtained when a plurality of polynucleotides are combined as a target nucleic acid than a single polynucleotide or a smaller number of polynucleotides.
- the combination of a plurality of polynucleotides is not limited to the above combination, and any combination of a plurality of polynucleotides can be used for detection of ovarian tumors.
- kits, device and method of the present invention it is possible to detect ovarian tumors more sensitively than existing tumor markers, so that it is possible to detect ovarian tumors early, and as a result Early treatment, such as chemotherapy, radiation therapy, immunotherapy, molecular targeted therapy, resection surgery with high chance of complete cure, etc. will be possible, leading to a significant improvement in survival rate.
- an ovarian tumor can be detected effectively by a simple and inexpensive method, early detection, diagnosis and treatment of the ovarian tumor become possible. Further, according to the method of the present invention, ovarian tumors can be detected in a minimally invasive manner using patient blood, so that ovarian tumors can be detected easily and rapidly. All publications, patents and patent applications cited herein are hereby incorporated by reference in their entirety.
Abstract
Description
すなわち、本発明は、以下の特徴を有する。
(1)卵巣腫瘍マーカーである、miR-4675、miR-4783-3p、miR-1228-5p、miR-4532、miR-6802-5p、miR-6784-5p、miR-3940-5p、miR-1307-3p、miR-8073、miR-3184-5p、miR-1233-5p、miR-6088、miR-5195-3p、miR-320b、miR-4649-5p、miR-6800-5p、miR-1343-3p、miR-4730、miR-6885-5p、miR-5100、miR-1203、miR-6756-5p、miR-373-5p、miR-1268a、miR-1260b、miR-4258、miR-4697-5p、miR-1469、miR-4515、miR-6861-5p、miR-6821-5p、miR-575、miR-6805-5p、miR-4758-5p、miR-3663-3p、miR-4530、miR-6798-5p、miR-6781-5p、miR-885-3p、miR-1273g-3p、miR-4787-3p、miR-4454、miR-4706、miR-1249-3p、miR-887-3p、miR-6786-5p、miR-1238-5p、miR-6749-5p、miR-6729-5p、miR-6825-5p、miR-663b、miR-6858-5p、miR-4690-5p、miR-6765-5p、miR-4710、miR-6775-5p、miR-371a-5p、miR-6816-5p、miR-296-3p、miR-7977、miR-8069、miR-6515-3p、miR-4687-5p、miR-1343-5p、miR-7110-5p、miR-4525、miR-3158-5p、miR-6787-5p、miR-614、miR-4689、miR-1185-2-3p、miR-1268b、miR-1228-3p、miR-1185-1-3p、miR-940、miR-939-5p、miR-6757-5p、miR-1275、miR-5001-5p、miR-6826-5p、miR-6765-3p、miR-3679-3p、miR-4718、miR-4286、miR-8059、miR-4447、miR-4448、miR-658、miR-6766-3p、miR-197-5p、miR-6887-5p、miR-6742-5p、miR-6729-3p、miR-5090、miR-7975、miR-4505、miR-6889-5p、miR-4708-3p、miR-6131、miR-1225-3p、miR-6132、miR-4734、miR-3194-3p、miR-638、miR-2467-3p、miR-4728-5p、miR-5572、miR-6789-5p、miR-8063、miR-4429、miR-6840-3p、miR-4476、miR-675-5p、miR-711、miR-6875-5p、miR-3160-5p、miR-1908-5p、miR-6726-5p、miR-1913、miR-8071、miR-3648、miR-4732-5p、miR-4787-5p、miR-3917、miR-619-5p、miR-1231、miR-342-5p、miR-4433a-5p、miR-6766-5p、miR-4707-5p、miR-7114-5p、miR-6872-3p、miR-6780b-5p、miR-7845-5p、miR-6798-3p、miR-665、miR-6848-5p、miR-5008-5p、miR-4294、miR-6511a-5p、miR-4435、miR-4747-3p、miR-6880-3p、miR-6869-5p、miR-7150、miR-1260a、miR-6877-5p、miR-6721-5p、miR-4656、miR-1229-5p、miR-4433a-3p、miR-4274、miR-4419b、miR-4674、miR-6893-5p、miR-6763-3p、miR-6762-5p、miR-6738-5p、miR-4513、miR-6746-5p、miR-6880-5p、miR-4736、miR-718、miR-6717-5p、miR-7847-3p、miR-760、miR-1199-5p、miR-6813-5p、miR-6769a-5p、miR-1193、miR-7108-3p、miR-6741-5p、miR-4298、miR-6796-3p、miR-4750-5p、miR-6785-5p、miR-1292-3p、miR-4749-3p、miR-6800-3p、miR-4722-5p、miR-4746-3p、miR-4450、miR-6795-5p、miR-365a-5p、miR-498、miR-6797-5p、miR-1470、miR-6851-5p、miR-1247-3p、miR-5196-5p、miR-208a-5p、miR-6842-5p、miR-150-3p、miR-4534、miR-3135b、miR-3131、miR-4792、miR-6510-5p、miR-504-3p、miR-3619-3p、miR-671-5p、miR-4667-5p、miR-4430、miR-3195、miR-3679-5p、miR-6076、miR-6515-5p、miR-6820-5p、miR-4634、miR-187-5p、miR-6763-5p、miR-1908-3p、miR-1181、miR-6782-5p、miR-5010-5p、miR-6870-5p、miR-6124、miR-1249-5p、miR-6511b-5p、miR-1254、miR-4727-3p、miR-4259、miR-4771、miR-3622a-5p、miR-4480、miR-4740-5p、miR-6777-5p、miR-6794-5p、miR-4687-3p、miR-6743-5p、miR-6771-5p、miR-3141、miR-3162-5p、miR-4271、miR-1227-5p、miR-4257、miR-4270、miR-4516、miR-4651、miR-4725-3p、miR-6125、miR-6732-5p、miR-6791-5p、miR-6819-5p、miR-6891-5p、miR-7108-5p、miR-7109-5p、miR-642b-3p及びmiR-642a-3pから選択される少なくとも1つのポリヌクレオチド、又は当該ポリヌクレオチドの相補鎖と特異的に結合可能な核酸を含む、卵巣腫瘍の検出用キット。
であり、miR-6800-3pがhsa-miR-6800-3pであり、miR-722-5pがhsa-miR-4722-5pであり、miR-4746-3pがhsa-miR-4746-3pであり、miR-4450がhsa-miR-4450であり、miR-6795-5pがhsa-miR-6795-5pであり、miR-365a-5pがhsa-miR-365a-5pであり、miR-498がhsa-miR-498であり、miR-6797-5pがhsa-miR-6797-5pであり、miR-1470がhsa-miR-1470であり、miR-6851-5pがhsa-miR-6851-5pであり、miR-1247-3pがhsa-miR-1247-3pであり、miR-5196-5pがhsa-miR-5196-5pであり、miR-208a-5pがhsa-miR-208a-5pであり、miR-6842-5pがhsa-miR-6842-5pであり、miR-150-3pがhsa-miR-150-3pであり、miR-4534がhsa-miR-4534であり、miR-3135bがhsa-miR-3135bであり、miR-3131がhsa-miR-3131であり、miR-4792がhsa-miR-4792であり、miR-6510-5pがhsa-miR-6510-5pであり、miR-504-3pがhsa-miR-504-3pであり、miR-3619-3pがhsa-miR-3619-3pであり、miR-671-5pがhsa-miR-671-5pであり、miR-4667-5pがhsa-miR-4667-5pであり、miR-4430がhsa-miR-4430であり、miR-3195がhsa-miR-3195であり、miR-3679-5pがhsa-miR-3679-5pであり、miR-6076がhsa-miR-6076であり、miR-6515-5pがhsa-miR-6515-5pであり、miR-6820-5pがhsa-miR-6820-5pであり、miR-4634がhsa-miR-4634であり、miR-187-5pがhsa-miR-187-5pであり、miR-6763-5pがhsa-miR-6763-5pであり、miR-1908-3pがhsa-miR-1908-3pであり、miR-1181がhsa-miR-1181であり、miR-6782-5pがhsa-miR-6782-5pであり、miR-5010-5pがhsa-miR-5010-5pであり、miR-6870-5pがhsa-miR-6870-5pであり、miR-6124がhsa-miR-6124であり、miR-1249-5pがhsa-miR-1249-5pであり、miR-6511b-5pがhsa-miR-6511b-5pであり、miR-1254がhsa-miR-1254であり、miR-4727-3pがhsa-miR-4727-3pであり、miR-4259がhsa-miR-4259であり、miR-4771がhsa-miR-4771であり、miR-3622a-5pがhsa-miR-3622a-5pであり、miR-4480がhsa-miR-4480であり、miR-4740-5pがhsa-miR-4740-5pであり、miR-6777-5pがhsa-miR-6777-5pであり、miR-6794-5pがhsa-miR-6794-5pであり、miR-4687-3pがhsa-miR-4687-3pであり、miR-6743-5pがhsa-miR-6743-5pであり、miR-6771-5pがhsa-miR-6771-5pであり、miR-3141がhsa-miR-3141であり、miR-3162-5pがhsa-miR-3162-5pであり、miR-4271がhsa-miR-4271であり、miR-1227-5pがhsa-miR-1227-5pであり、miR-4257がhsa-miR-4257であり、miR-4270がhsa-miR-4270であり、miR-4516がhsa-miR-4516であり、miR-4651がhsa-miR-4651であり、miR-4725-3pがhsa-miR-4725-3pであり、miR-6125がhsa-miR-6125であり、miR-6732-5pがhsa-miR-6732-5pであり、miR-6791-5pがhsa-miR-6791-5pであり、miR-6819-5pがhsa-miR-6819-5pであり、miR-6891-5pがhsa-miR-6891-5pであり、miR-7108-5pがhsa-miR-7108-5pであり、miR-7109-5pがhsa-miR-7109-5pであり、miR-642b-3pがhsa-miR-642b-3pであり、miR-642a-3pがhsa-miR-642a-3pである、(1)に記載のキット。
(a)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~247、251及び268のいずれかで表される塩基配列を含むポリヌクレオチド、
(c)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、及び
(e)上記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(1)又は(2)に記載のキット。
(f)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列を含むポリヌクレオチド、
(h)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、及び
(j)上記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(4)又は(5)に記載のキット。
であり、miR-6800-3pがhsa-miR-6800-3pであり、miR-722-5pがhsa-miR-4722-5pであり、miR-4746-3pがhsa-miR-4746-3pであり、miR-4450がhsa-miR-4450であり、miR-6795-5pがhsa-miR-6795-5pであり、miR-365a-5pがhsa-miR-365a-5pであり、miR-498がhsa-miR-498であり、miR-6797-5pがhsa-miR-6797-5pであり、miR-1470がhsa-miR-1470であり、miR-6851-5pがhsa-miR-6851-5pであり、miR-1247-3pがhsa-miR-1247-3pであり、miR-5196-5pがhsa-miR-5196-5pであり、miR-208a-5pがhsa-miR-208a-5pであり、miR-6842-5pがhsa-miR-6842-5pであり、miR-150-3pがhsa-miR-150-3pであり、miR-4534がhsa-miR-4534であり、miR-3135bがhsa-miR-3135bであり、miR-3131がhsa-miR-3131であり、miR-4792がhsa-miR-4792であり、miR-6510-5pがhsa-miR-6510-5pであり、miR-504-3pがhsa-miR-504-3pであり、miR-3619-3pがhsa-miR-3619-3pであり、miR-671-5pがhsa-miR-671-5pであり、miR-4667-5pがhsa-miR-4667-5pであり、miR-4430がhsa-miR-4430であり、miR-3195がhsa-miR-3195であり、miR-3679-5pがhsa-miR-3679-5pであり、miR-6076がhsa-miR-6076であり、miR-6515-5pがhsa-miR-6515-5pであり、miR-6820-5pがhsa-miR-6820-5pであり、miR-4634がhsa-miR-4634であり、miR-187-5pがhsa-miR-187-5pであり、miR-6763-5pがhsa-miR-6763-5pであり、miR-1908-3pがhsa-miR-1908-3pであり、miR-1181がhsa-miR-1181であり、miR-6782-5pがhsa-miR-6782-5pであり、miR-5010-5pがhsa-miR-5010-5pであり、miR-6870-5pがhsa-miR-6870-5pであり、miR-6124がhsa-miR-6124であり、miR-1249-5pがhsa-miR-1249-5pであり、miR-6511b-5pがhsa-miR-6511b-5pであり、miR-1254がhsa-miR-1254であり、miR-4727-3pがhsa-miR-4727-3pであり、miR-4259がhsa-miR-4259であり、miR-4771がhsa-miR-4771であり、miR-3622a-5pがhsa-miR-3622a-5pであり、miR-4480がhsa-miR-4480であり、miR-4740-5pがhsa-miR-4740-5pであり、miR-6777-5pがhsa-miR-6777-5pであり、miR-6794-5pがhsa-miR-6794-5pであり、miR-4687-3pがhsa-miR-4687-3pであり、miR-6743-5pがhsa-miR-6743-5pであり、miR-6771-5pがhsa-miR-6771-5pであり、miR-3141がhsa-miR-3141であり、miR-3162-5pがhsa-miR-3162-5pであり、miR-4271がhsa-miR-4271であり、miR-1227-5pがhsa-miR-1227-5pであり、miR-4257がhsa-miR-4257であり、miR-4270がhsa-miR-4270であり、miR-4516がhsa-miR-4516であり、miR-4651がhsa-miR-4651であり、miR-4725-3pがhsa-miR-4725-3pであり、miR-6125がhsa-miR-6125であり、miR-6732-5pがhsa-miR-6732-5pであり、miR-6791-5pがhsa-miR-6791-5pであり、miR-6819-5pがhsa-miR-6819-5pであり、miR-6891-5pがhsa-miR-6891-5pであり、miR-7108-5pがhsa-miR-7108-5pであり、miR-7109-5pがhsa-miR-7109-5pであり、miR-642b-3pがhsa-miR-642b-3pであり、miR-642a-3pがhsa-miR-642a-3pである、(8)に記載のデバイス。
(a)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~247、251及び268のいずれかで表される塩基配列を含むポリヌクレオチド、
(c)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、及び
(e)上記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(8)又は(9)に記載のデバイス。
(f)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列を含むポリヌクレオチド、
(h)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、及び
(j)上記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(11)又は(12)に記載のデバイス。
pであり、miR-6800-3pがhsa-miR-6800-3pであり、miR-722-5pがhsa-miR-4722-5pであり、miR-4746-3pがhsa-miR-4746-3pであり、miR-4450がhsa-miR-4450であり、miR-6795-5pがhsa-miR-6795-5pであり、miR-365a-5pがhsa-miR-365a-5pであり、miR-498がhsa-miR-498であり、miR-6797-5pがhsa-miR-6797-5pであり、miR-1470がhsa-miR-1470であり、miR-6851-5pがhsa-miR-6851-5pであり、miR-1247-3pがhsa-miR-1247-3pであり、miR-5196-5pがhsa-miR-5196-5pであり、miR-208a-5pがhsa-miR-208a-5pであり、miR-6842-5pがhsa-miR-6842-5pであり、miR-150-3pがhsa-miR-150-3pであり、miR-4534がhsa-miR-4534であり、miR-3135bがhsa-miR-3135bであり、miR-3131がhsa-miR-3131であり、miR-4792がhsa-miR-4792であり、miR-6510-5pがhsa-miR-6510-5pであり、miR-504-3pがhsa-miR-504-3pであり、miR-3619-3pがhsa-miR-3619-3pであり、miR-671-5pがhsa-miR-671-5pであり、miR-4667-5pがhsa-miR-4667-5pであり、miR-4430がhsa-miR-4430であり、miR-3195がhsa-miR-3195であり、miR-3679-5pがhsa-miR-3679-5pであり、miR-6076がhsa-miR-6076であり、miR-6515-5pがhsa-miR-6515-5pであり、miR-6820-5pがhsa-miR-6820-5pであり、miR-4634がhsa-miR-4634であり、miR-187-5pがhsa-miR-187-5pであり、miR-6763-5pがhsa-miR-6763-5pであり、miR-1908-3pがhsa-miR-1908-3pであり、miR-1181がhsa-miR-1181であり、miR-6782-5pがhsa-miR-6782-5pであり、miR-5010-5pがhsa-miR-5010-5pであり、miR-6870-5pがhsa-miR-6870-5pであり、miR-6124がhsa-miR-6124であり、miR-1249-5pがhsa-miR-1249-5pであり、miR-6511b-5pがhsa-miR-6511b-5pであり、miR-1254がhsa-miR-1254であり、miR-4727-3pがhsa-miR-4727-3pであり、miR-4259がhsa-miR-4259であり、miR-4771がhsa-miR-4771であり、miR-3622a-5pがhsa-miR-3622a-5pであり、miR-4480がhsa-miR-4480であり、miR-4740-5pがhsa-miR-4740-5pであり、miR-6777-5pがhsa-miR-6777-5pであり、miR-6794-5pがhsa-miR-6794-5pであり、miR-4687-3pがhsa-miR-4687-3pであり、miR-6743-5pがhsa-miR-6743-5pであり、miR-6771-5pがhsa-miR-6771-5pであり、miR-3141がhsa-miR-3141であり、miR-3162-5pがhsa-miR-3162-5pであり、miR-4271がhsa-miR-4271であり、miR-1227-5pがhsa-miR-1227-5pであり、miR-4257がhsa-miR-4257であり、miR-4270がhsa-miR-4270であり、miR-4516がhsa-miR-4516であり、miR-4651がhsa-miR-4651であり、miR-4725-3pがhsa-miR-4725-3pであり、miR-6125がhsa-miR-6125であり、miR-6732-5pがhsa-miR-6732-5pであり、miR-6791-5pがhsa-miR-6791-5pであり、miR-6819-5pがhsa-miR-6819-5pであり、miR-6891-5pがhsa-miR-6891-5pであり、miR-7108-5pがhsa-miR-7108-5pであり、miR-7109-5pがhsa-miR-7109-5pであり、miR-642b-3pがhsa-miR-642b-3pであり、miR-642a-3pがhsa-miR-642a-3pである、(17)~(19)のいずれかに記載の方法。
(a)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~247、251及び268のいずれかで表される塩基配列を含むポリヌクレオチド、
(c)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、及び
(e)上記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(17)~(19)のいずれかに記載の方法。
(f)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列を含むポリヌクレオチド、
(h)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、及び
(j)上記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(22)又は(23)に記載の方法。
本明細書中で使用する用語は、以下の定義を有する。
ヌクレオチド、ポリヌクレオチド、DNA、RNAなどの略号による表示は、「塩基配列又はアミノ酸配列を含む明細書等の作成のためのガイドライン」(日本国特許庁編)及び当技術分野における慣用に従うものとする。
本明細書において「Tm値」とは、ポリヌクレオチドの二本鎖部分が一本鎖へと変性し、二本鎖と一本鎖が1:1の比で存在する温度を意味する。
本明細書で使用される「hsa-miR-1307-3p遺伝子」又は「hsa-miR-1307-3p」という用語は、配列番号8に記載のhsa-miR-1307-3p遺伝子(miRBase Accession No.MIMAT0005951)やその他生物種ホモログもしくはオーソログなどが包含される。hsa-miR-1307-3p遺伝子は、Morin RDら、2008年、Genome Res、18巻、p610-621に記載される方法によって得ることができる。また、「hsa-miR-1307-3p」は、その前駆体としてヘアピン様構造をとる「hsa-mir-1307」(miRBase Accession No.MI0006444、配列番号283)が知られている。
1.卵巣腫瘍の標的核酸
本発明の上記定義の卵巣腫瘍検出用の核酸プローブ又はプライマーなどの核酸を使用して、卵巣腫瘍又は卵巣腫瘍細胞の存在及び/又は不存在を検出するための、卵巣腫瘍マーカーとしての主要な標的核酸には、hsa-miR-4675、hsa-miR-4783-3p、hsa-miR-1228-5p、hsa-miR-4532、hsa-miR-6802-5p、hsa-miR-6784-5p、hsa-miR-3940-5p、hsa-miR-1307-3p、hsa-miR-8073、hsa-miR-3184-5p、hsa-miR-1233-5p、hsa-miR-6088、hsa-miR-5195-3p、hsa-miR-320b、hsa-miR-4649-5p、hsa-miR-6800-5p、hsa-miR-1343-3p、hsa-miR-4730、hsa-miR-6885-5p、hsa-miR-5100、hsa-miR-1203、hsa-miR-6756-5p、hsa-miR-373-5p、hsa-miR-1268a、hsa-miR-1260b、hsa-miR-4258、hsa-miR-4697-5p、hsa-miR-1469、hsa-miR-4515、hsa-miR-6861-5p、hsa-miR-6821-5p、hsa-miR-575、hsa-miR-6805-5p、hsa-miR-4758-5p、hsa-miR-3663-3p、hsa-miR-4530、hsa-miR-6798-5p、hsa-miR-6781-5p、hsa-miR-885-3p、hsa-miR-1273g-3p、hsa-miR-4787-3p、hsa-miR-4454、hsa-miR-4706、hsa-miR-1249-3p、hsa-miR-887-3p、hsa-miR-6786-5p、hsa-miR-1238-5p、hsa-miR-6749-5p、hsa-miR-6729-5p、hsa-miR-6825-5p、hsa-miR-663b、hsa-miR-6858-5p、hsa-miR-4690-5p、hsa-miR-6765-5p、hsa-miR-4710、hsa-miR-6775-5p、hsa-miR-371a-5p、hsa-miR-6816-5p、hsa-miR-296-3p、hsa-miR-7977、hsa-miR-8069、hsa-miR-6515-3p、hsa-miR-4687-5p、hsa-miR-1343-5p、hsa-miR-7110-5p、hsa-miR-4525、hsa-miR-3158-5p、hsa-miR-6787-5p、hsa-miR-614、hsa-miR-4689、hsa-miR-1185-2-3p、hsa-miR-1268b、hsa-miR-1228-3p、hsa-miR-1185-1-3p、hsa-miR-940、hsa-miR-939-5p、hsa-miR-6757-5p、hsa-miR-1275、hsa-miR-5001-5p、hsa-miR-6826-5p、hsa-miR-6765-3p、hsa-miR-3679-3p、hsa-miR-4718、hsa-miR-4286、hsa-miR-8059、hsa-miR-4447、hsa-miR-4448、hsa-miR-658、hsa-miR-6766-3p、hsa-miR-197-5p、hsa-miR-6887-5p、hsa-miR-6742-5p、hsa-miR-6729-3p、hsa-miR-5090、hsa-miR-7975、hsa-miR-4505、hsa-miR-6889-5p、hsa-miR-4708-3p、hsa-miR-6131、hsa-miR-1225-3p、hsa-miR-6132、hsa-miR-4734、hsa-miR-3194-3p、hsa-miR-638、hsa-miR-2467-3p、hsa-miR-4728-5p、hsa-miR-5572、hsa-miR-6789-5p、hsa-miR-8063、hsa-miR-4429、hsa-miR-6840-3p、hsa-miR-4476、hsa-miR-675-5p、hsa-miR-711、hsa-miR-6875-5p、hsa-miR-3160-5p、hsa-miR-1908-5p、hsa-miR-6726-5p、hsa-miR-1913、hsa-miR-8071、hsa-miR-3648、hsa-miR-4732-5p、hsa-miR-4787-5p、hsa-miR-3917、hsa-miR-619-5p、hsa-miR-1231、hsa-miR-342-5p、hsa-miR-4433a-5p、hsa-miR-6766-5p、hsa-miR-4707-5p、hsa-miR-7114-5p、hsa-miR-6872-3p、hsa-miR-6780b-5p、hsa-miR-7845-5p、hsa-miR-6798-3p、hsa-miR-665、hsa-miR-6848-5p、hsa-miR-5008-5p、hsa-miR-4294、hsa-miR-6511a-5p、hsa-miR-4435、hsa-miR-4747-3p、hsa-miR-6880-3p、hsa-miR-6869-5p、hsa-miR-7150、hsa-miR-1260a、hsa-miR-6877-5p、hsa-miR-6721-5p、hsa-miR-4656、hsa-miR-1229-5p、hsa-miR-4433a-3p、hsa-miR-4274、hsa-miR-4419b、hsa-miR-4674、hsa-miR-6893-5p、hsa-miR-6763-3p、hsa-miR-6762-5p、hsa-miR-6738-5p、hsa-miR-4513、hsa-miR-6746-5p、hsa-miR-6880-5p、hsa-miR-4736、hsa-miR-718、hsa-miR-6717-5p、hsa-miR-7847-3p、hsa-miR-760、hsa-miR-1199-5p、hsa-miR-6813-5p、hsa-miR-6769a-5p、hsa-miR-1193、hsa-miR-7108-3p、hsa-miR-6741-5p、hsa-miR-4298、hsa-miR-6796-3p、hsa-miR-4750-5p、hsa-miR-6785-5p、hsa-miR-1292-3p、hsa-miR-4749-3p、hsa-miR-6800-3p、hsa-miR-4722-5p、hsa-miR-4746-3p、hsa-miR-4450、hsa-miR-6795-5p、hsa-miR-365a-5p、hsa-miR-498、hsa-miR-6797-5p、hsa-miR-1470、hsa-miR-6851-5p、hsa-miR-1247-3p、hsa-miR-5196-5p、hsa-miR-208a-5p、hsa-miR-6842-5p、hsa-miR-150-3p、hsa-miR-4534、hsa-miR-3135b、hsa-miR-3131、hsa-miR-4792、hsa-miR-6510-5p、hsa-miR-504-3p、hsa-miR-3619-3p、hsa-miR-671-5p、hsa-miR-4667-5p、hsa-miR-4430、hsa-miR-3195、hsa-miR-3679-5p、hsa-miR-6076、hsa-miR-6515-5p、hsa-miR-6820-5p、hsa-miR-4634、hsa-miR-187-5p、hsa-miR-6763-5p、hsa-miR-1908-3p、hsa-miR-1181、hsa-miR-6782-5p、hsa-miR-5010-5p、hsa-miR-6870-5p、hsa-miR-6124、hsa-miR-1249-5p、hsa-miR-6511b-5p、hsa-miR-1254、hsa-miR-4727-3p、hsa-miR-4259、hsa-miR-4771、hsa-miR-3622a-5p、hsa-miR-4480、hsa-miR-4740-5p、hsa-miR-6777-5p、hsa-miR-6794-5p、hsa-miR-4687-3p、hsa-miR-6743-5p、hsa-miR-6771-5p、hsa-miR-3141、hsa-miR-3162-5p、hsa-miR-4271、hsa-miR-1227-5p、hsa-miR-4257、hsa-miR-4270、hsa-miR-4516、hsa-miR-4651、hsa-miR-4725-3p、hsa-miR-6125、hsa-miR-6732-5p、hsa-miR-6791-5p、hsa-miR-6819-5p、hsa-miR-6891-5p、hsa-miR-7108-5p、hsa-miR-7109-5p、hsa-miR-642b-3p及びhsa-miR-642a-3pからなる群から選択される少なくとも1つのmiRNAが含まれる。さらにこれらのmiRNAと組み合わせることができる他の卵巣腫瘍マーカー、すなわち、hsa-miR-320a、hsa-miR-663a、hsa-miR-328-5p、hsa-miR-128-2-5p、hsa-miR-125a-3p、hsa-miR-191-5p、hsa-miR-92b-5p、hsa-miR-296-5p、hsa-miR-1246、hsa-miR-92a-2-5p、hsa-miR-128-1-5p、hsa-miR-1290、hsa-miR-211-3p、hsa-miR-744-5p、hsa-miR-135a-3p、hsa-miR-451a、hsa-miR-625-3p、hsa-miR-92a-3p、hsa-miR-422a、hsa-miR-483-5p、hsa-miR-652-5p、hsa-miR-24-3p、hsa-miR-23b-3p、hsa-miR-23a-3p、hsa-miR-92b-3p、hsa-miR-22-3pからなる群から選択される少なくとも1つのmiRNAも標的核酸として好ましく用いることができる。
本発明において、卵巣腫瘍を検出するための核酸、例えば卵巣腫瘍を診断するために使用可能な核酸プローブ又はプライマーは、卵巣腫瘍の標的核酸としての、ヒト由来のhsa-miR-4675、hsa-miR-4783-3p、hsa-miR-1228-5p、hsa-miR-4532、hsa-miR-6802-5p、hsa-miR-6784-5p、hsa-miR-3940-5p、hsa-miR-1307-3p、hsa-miR-8073、hsa-miR-3184-5p、hsa-miR-1233-5p、hsa-miR-6088、hsa-miR-5195-3p、hsa-miR-320b、hsa-miR-4649-5p、hsa-miR-6800-5p、hsa-miR-1343-3p、hsa-miR-4730、hsa-miR-6885-5p、hsa-miR-5100、hsa-miR-1203、hsa-miR-6756-5p、hsa-miR-373-5p、hsa-miR-1268a、hsa-miR-1260b、hsa-miR-4258、hsa-miR-4697-5p、hsa-miR-1469、hsa-miR-4515、hsa-miR-6861-5p、hsa-miR-6821-5p、hsa-miR-575、hsa-miR-6805-5p、hsa-miR-4758-5p、hsa-miR-3663-3p、hsa-miR-4530、hsa-miR-6798-5p、hsa-miR-6781-5p、hsa-miR-885-3p、hsa-miR-1273g-3p、hsa-miR-4787-3p、hsa-miR-4454、hsa-miR-4706、hsa-miR-1249-3p、hsa-miR-887-3p、hsa-miR-6786-5p、hsa-miR-1238-5p、hsa-miR-6749-5p、hsa-miR-6729-5p、hsa-miR-6825-5p、hsa-miR-663b、hsa-miR-6858-5p、hsa-miR-4690-5p、hsa-miR-6765-5p、hsa-miR-4710、hsa-miR-6775-5p、hsa-miR-371a-5p、hsa-miR-6816-5p、hsa-miR-296-3p、hsa-miR-7977、hsa-miR-8069、hsa-miR-6515-3p、hsa-miR-4687-5p、hsa-miR-1343-5p、hsa-miR-7110-5p、hsa-miR-4525、hsa-miR-3158-5p、hsa-miR-6787-5p、hsa-miR-614、hsa-miR-4689、hsa-miR-1185-2-3p、hsa-miR-1268b、hsa-miR-1228-3p、hsa-miR-1185-1-3p、hsa-miR-940、hsa-miR-939-5p、hsa-miR-6757-5p、hsa-miR-1275、hsa-miR-5001-5p、hsa-miR-6826-5p、hsa-miR-6765-3p、hsa-miR-3679-3p、hsa-miR-4718、hsa-miR-4286、hsa-miR-8059、hsa-miR-4447、hsa-miR-4448、hsa-miR-658、hsa-miR-6766-3p、hsa-miR-197-5p、hsa-miR-6887-5p、hsa-miR-6742-5p、hsa-miR-6729-3p、hsa-miR-5090、hsa-miR-7975、hsa-miR-4505、hsa-miR-6889-5p、hsa-miR-4708-3p、hsa-miR-6131、hsa-miR-1225-3p、hsa-miR-6132、hsa-miR-4734、hsa-miR-3194-3p、hsa-miR-638、hsa-miR-2467-3p、hsa-miR-4728-5p、hsa-miR-5572、hsa-miR-6789-5p、hsa-miR-8063、hsa-miR-4429、hsa-miR-6840-3p、hsa-miR-4476、hsa-miR-675-5p、hsa-miR-711、hsa-miR-6875-5p、hsa-miR-3160-5p、hsa-miR-1908-5p、hsa-miR-6726-5p、hsa-miR-1913、hsa-miR-8071、hsa-miR-3648、hsa-miR-4732-5p、hsa-miR-4787-5p、hsa-miR-3917、hsa-miR-619-5p、hsa-miR-1231、hsa-miR-342-5p、hsa-miR-4433a-5p、hsa-miR-6766-5p、hsa-miR-4707-5p、hsa-miR-7114-5p、hsa-miR-6872-3p、hsa-miR-6780b-5p、hsa-miR-7845-5p、hsa-miR-6798-3p、hsa-miR-665、hsa-miR-6848-5p、hsa-miR-5008-5p、hsa-miR-4294、hsa-miR-6511a-5p、hsa-miR-4435、hsa-miR-4747-3p、hsa-miR-6880-3p、hsa-miR-6869-5p、hsa-miR-7150、hsa-miR-1260a、hsa-miR-6877-5p、hsa-miR-6721-5p、hsa-miR-4656、hsa-miR-1229-5p、hsa-miR-4433a-3p、hsa-miR-4274、hsa-miR-4419b、hsa-miR-4674、hsa-miR-6893-5p、hsa-miR-6763-3p、hsa-miR-6762-5p、hsa-miR-6738-5p、hsa-miR-4513、hsa-miR-6746-5p、hsa-miR-6880-5p、hsa-miR-4736、hsa-miR-718、hsa-miR-6717-5p、hsa-miR-7847-3p、hsa-miR-760、hsa-miR-1199-5p、hsa-miR-6813-5p、hsa-miR-6769a-5p、hsa-miR-1193、hsa-miR-7108-3p、hsa-miR-6741-5p、hsa-miR-4298、hsa-miR-6796-3p、hsa-miR-4750-5p、hsa-miR-6785-5p、hsa-miR-1292-3p、hsa-miR-4749-3p、hsa-miR-6800-3p、hsa-miR-4722-5p、hsa-miR-4746-3p、hsa-miR-4450、hsa-miR-6795-5p、hsa-miR-365a-5p、hsa-miR-498、hsa-miR-6797-5p、hsa-miR-1470、hsa-miR-6851-5p、hsa-miR-1247-3p、hsa-miR-5196-5p、hsa-miR-208a-5p、hsa-miR-6842-5p、hsa-miR-150-3p、hsa-miR-4534、hsa-miR-3135b、hsa-miR-3131、hsa-miR-4792、hsa-miR-6510-5p、hsa-miR-504-3p、hsa-miR-3619-3p、hsa-miR-671-5p、hsa-miR-4667-5p、hsa-miR-4430、hsa-miR-3195、hsa-miR-3679-5p、hsa-miR-6076、hsa-miR-6515-5p、hsa-miR-6820-5p、hsa-miR-4634、hsa-miR-187-5p、hsa-miR-6763-5p、hsa-miR-1908-3p、hsa-miR-1181、hsa-miR-6782-5p、hsa-miR-5010-5p、hsa-miR-6870-5p、hsa-miR-6124、hsa-miR-1249-5p、hsa-miR-6511b-5p、hsa-miR-1254、hsa-miR-4727-3p、hsa-miR-4259、hsa-miR-4771、hsa-miR-3622a-5p、hsa-miR-4480、hsa-miR-4740-5p、hsa-miR-6777-5p、hsa-miR-6794-5p、hsa-miR-4687-3p、hsa-miR-6743-5p、hsa-miR-6771-5p、hsa-miR-3141、hsa-miR-3162-5p、hsa-miR-4271、hsa-miR-1227-5p、hsa-miR-4257、hsa-miR-4270、hsa-miR-4516、hsa-miR-4651、hsa-miR-4725-3p、hsa-miR-6125、hsa-miR-6732-5p、hsa-miR-6791-5p、hsa-miR-6819-5p、hsa-miR-6891-5p、hsa-miR-7108-5p、hsa-miR-7109-5p、hsa-miR-642b-3p及びhsa-miR-642a-3p、あるいはそれらの組み合わせ、並びに、それらと場合により組み合わせることができる、hsa-miR-320a、hsa-miR-663a、hsa-miR-328-5p、hsa-miR-128-2-5p、hsa-miR-125a-3p、hsa-miR-191-5p、hsa-miR-92b-5p、hsa-miR-296-5p、hsa-miR-1246、hsa-miR-92a-2-5p、hsa-miR-128-1-5p、hsa-miR-1290、hsa-miR-211-3p、hsa-miR-744-5p、hsa-miR-135a-3p、hsa-miR-451a、hsa-miR-625-3p、hsa-miR-92a-3p、hsa-miR-422a、hsa-miR-483-5p、hsa-miR-652-5p、hsa-miR-24-3p、hsa-miR-23b-3p、hsa-miR-23a-3p、hsa-miR-92b-3p、hsa-miR-22-3p、あるいはそれらの組み合わせ、それらの同族体、それらの転写産物、あるいはそれらの変異体又は誘導体の存在、発現量又は存在量を定性的及び/又は定量的に測定することを可能にする。
(a)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、からなるポリヌクレオチド、その変異体、その誘導体、は15以上の連続した塩基を含むその断片、
(b)配列番号1~247、251及び268のいずれかで表される塩基配列を含むポリヌクレオチド、
(c)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、並びに、
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド。
(f)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列を含むポリヌクレオチド、
(h)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、並びに、
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド。
本発明はまた、卵巣腫瘍マーカーである標的核酸を測定するための、本発明において核酸プローブ又はプライマーとして使用可能なポリヌクレオチド(これには、変異体、断片、又は誘導体を含みうる。)の1つ又は複数を含む卵巣腫瘍検出用キット又はデバイスを提供する。
群A:
hsa-miR-4675、hsa-miR-4783-3p、hsa-miR-1228-5p、hsa-miR-4532、hsa-miR-6802-5p、hsa-miR-6784-5p、hsa-miR-3940-5p、hsa-miR-1307-3p、hsa-miR-8073、hsa-miR-3184-5p、hsa-miR-1233-5p、hsa-miR-6088、hsa-miR-5195-3p、hsa-miR-320b、hsa-miR-4649-5p、hsa-miR-6800-5p、hsa-miR-1343-3p、hsa-miR-4730、hsa-miR-6885-5p、hsa-miR-5100、hsa-miR-1203、hsa-miR-6756-5p、hsa-miR-373-5p、hsa-miR-1268a、hsa-miR-1260b、hsa-miR-4258、hsa-miR-4697-5p、hsa-miR-1469、hsa-miR-4515、hsa-miR-6861-5p、hsa-miR-6821-5p、hsa-miR-575、hsa-miR-6805-5p、hsa-miR-4758-5p、hsa-miR-3663-3p、hsa-miR-4530、hsa-miR-6798-5p、hsa-miR-6781-5p、hsa-miR-885-3p、hsa-miR-1273g-3p、hsa-miR-4787-3p、hsa-miR-4454、hsa-miR-4706、hsa-miR-1249-3p、hsa-miR-887-3p、hsa-miR-6786-5p、hsa-miR-1238-5p、hsa-miR-6749-5p、hsa-miR-6729-5p、hsa-miR-6825-5p、hsa-miR-663b、hsa-miR-6858-5p、hsa-miR-4690-5p、hsa-miR-6765-5p、hsa-miR-4710、hsa-miR-6775-5p、hsa-miR-371a-5p、hsa-miR-6816-5p、hsa-miR-296-3p、hsa-miR-7977、hsa-miR-8069、hsa-miR-6515-3p、hsa-miR-4687-5p、hsa-miR-1343-5p、hsa-miR-7110-5p、hsa-miR-4525、hsa-miR-3158-5p、hsa-miR-6787-5p、hsa-miR-614、hsa-miR-4689、hsa-miR-1185-2-3p、hsa-miR-1268b、hsa-miR-1228-3p、hsa-miR-1185-1-3p、hsa-miR-940、hsa-miR-939-5p、hsa-miR-6757-5p、hsa-miR-1275、hsa-miR-5001-5p、hsa-miR-6826-5p、hsa-miR-6765-3p、hsa-miR-3679-3p、hsa-miR-4718、hsa-miR-4286、hsa-miR-8059、hsa-miR-4447、hsa-miR-4448、hsa-miR-658、hsa-miR-6766-3p、hsa-miR-197-5p、hsa-miR-6887-5p、hsa-miR-6742-5p、hsa-miR-6729-3p、hsa-miR-5090、hsa-miR-7975、hsa-miR-4505、hsa-miR-6889-5p、hsa-miR-4708-3p、hsa-miR-6131、hsa-miR-1225-3p、hsa-miR-6132、hsa-miR-4734、hsa-miR-3194-3p、hsa-miR-638、hsa-miR-2467-3p、hsa-miR-4728-5p、hsa-miR-5572、hsa-miR-6789-5p、hsa-miR-8063、hsa-miR-4429、hsa-miR-6840-3p、hsa-miR-4476、hsa-miR-675-5p、hsa-miR-711、hsa-miR-6875-5p、hsa-miR-3160-5p、hsa-miR-1908-5p、hsa-miR-6726-5p、hsa-miR-1913、hsa-miR-8071、hsa-miR-3648、hsa-miR-4732-5p、hsa-miR-4787-5p、hsa-miR-3917、hsa-miR-619-5p、hsa-miR-1231、hsa-miR-342-5p、hsa-miR-4433a-5p、hsa-miR-6766-5p、hsa-miR-4707-5p、hsa-miR-7114-5p、hsa-miR-6872-3p、hsa-miR-6780b-5p、hsa-miR-7845-5p、hsa-miR-6798-3p、hsa-miR-665、hsa-miR-6848-5p、hsa-miR-5008-5p、hsa-miR-4294、hsa-miR-6511a-5p、hsa-miR-4435、hsa-miR-4747-3p、hsa-miR-6880-3p、hsa-miR-6869-5p、hsa-miR-7150、hsa-miR-1260a、hsa-miR-6877-5p、hsa-miR-6721-5p、hsa-miR-4656、hsa-miR-1229-5p、hsa-miR-4433a-3p、hsa-miR-4274、hsa-miR-4419b、hsa-miR-4674、hsa-miR-6893-5p、hsa-miR-6763-3p、hsa-miR-6762-5p、hsa-miR-6738-5p、hsa-miR-4513、hsa-miR-6746-5p、hsa-miR-6880-5p、hsa-miR-4736、hsa-miR-718、hsa-miR-6717-5p、hsa-miR-7847-3p、hsa-miR-760、hsa-miR-1199-5p、hsa-miR-6813-5p、hsa-miR-6769a-5p、hsa-miR-1193、hsa-miR-7108-3p、hsa-miR-6741-5p、hsa-miR-4298、hsa-miR-6796-3p、hsa-miR-4750-5p、hsa-miR-6785-5p、hsa-miR-1292-3p、hsa-miR-4749-3p、hsa-miR-6800-3p、hsa-miR-4722-5p、hsa-miR-4746-3p、hsa-miR-4450、hsa-miR-6795-5p、hsa-miR-365a-5p、hsa-miR-498、hsa-miR-6797-5p、hsa-miR-1470、hsa-miR-6851-5p、hsa-miR-1247-3p、hsa-miR-5196-5p、hsa-miR-208a-5p、hsa-miR-6842-5p、hsa-miR-150-3p、hsa-miR-4534、hsa-miR-3135b、hsa-miR-3131、hsa-miR-4792、hsa-miR-6510-5p、hsa-miR-504-3p、hsa-miR-3619-3p、hsa-miR-671-5p、hsa-miR-4667-5p、hsa-miR-4430、hsa-miR-3195、hsa-miR-3679-5p、hsa-miR-6076、hsa-miR-6515-5p、hsa-miR-6820-5p、hsa-miR-4634、hsa-miR-187-5p、hsa-miR-6763-5p、hsa-miR-1908-3p、hsa-miR-1181、hsa-miR-6782-5p、hsa-miR-5010-5p、hsa-miR-6870-5p、hsa-miR-6124、hsa-miR-1249-5p、hsa-miR-6511b-5p、hsa-miR-1254、hsa-miR-4727-3p、hsa-miR-4259、hsa-miR-4771、hsa-miR-3622a-5p、hsa-miR-4480、hsa-miR-4740-5p、hsa-miR-6777-5p、hsa-miR-6794-5p、hsa-miR-4687-3p、hsa-miR-6743-5p、hsa-miR-6771-5p、hsa-miR-3141、hsa-miR-3162-5p、hsa-miR-4271、hsa-miR-1227-5p、hsa-miR-4257、hsa-miR-4270、hsa-miR-4516、hsa-miR-4651、hsa-miR-4725-3p、hsa-miR-6125、hsa-miR-6732-5p、hsa-miR-6791-5p、hsa-miR-6819-5p、hsa-miR-6891-5p、hsa-miR-7108-5p、hsa-miR-7109-5p、hsa-miR-642b-3p及びhsa-miR-642a-3p。
群B:
hsa-miR-320a、hsa-miR-663a、hsa-miR-328-5p、hsa-miR-128-2-5p、hsa-miR-125a-3p、hsa-miR-191-5p、hsa-miR-92b-5p、hsa-miR-296-5p、hsa-miR-1246、hsa-miR-92a-2-5p、hsa-miR-128-1-5p、hsa-miR-1290、hsa-miR-211-3p、hsa-miR-744-5p、hsa-miR-135a-3p、hsa-miR-451a、hsa-miR-625-3p、hsa-miR-92a-3p、hsa-miR-422a、hsa-miR-483-5p、hsa-miR-652-5p、hsa-miR-24-3p、hsa-miR-23b-3p、hsa-miR-23a-3p、hsa-miR-92b-3p、hsa-miR-22-3p。
(2)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列においてuがtである塩基配列又はその相補的配列において、15以上の連続した塩基を含むポリヌクレオチド。
(1)配列番号256、228の組合せ
(2)配列番号47、228の組合せ
(3)配列番号228、9、229の組合せ
(4)配列番号228、9、78の組合せ
(5)配列番号228、9、82の組合せ
(6)配列番号228、9、268の組合せ
(7)配列番号229、260、228の組合せ
(8)配列番号228、9、247の組合せ
(9)配列番号228、9、62の組合せ
(10)配列番号228、9、217の組合せ
(11)配列番号256、228、9の組合せ
(12)配列番号256、228、260の組合せ
(13)配列番号228、9、234の組合せ
(14)配列番号229、20、228の組合せ
(15)配列番号47、228、82の組合せ
(16)配列番号145、260、228の組合せ
(17)配列番号228、9、145の組合せ
(18)配列番号228、9、268、160の組合せ
(19)配列番号228、9、82、268の組合せ
(20)配列番号228、9、268、196の組合せ
(21)配列番号145、260、228、250の組合せ
(22)配列番号145、260、228、62の組合せ
(23)配列番号228、9、268、230の組合せ
(24)配列番号229、260、228、2の組合せ
(25)配列番号228、9、268、62の組合せ
(26)配列番号229、260、228、172の組合せ
(27)配列番号229、260、228、4の組合せ
(28)配列番号229、260、228、34の組合せ
(29)配列番号229、260、228、56の組合せ
(30)配列番号229、20、228、4の組合せ
(31)配列番号229、20、228、102の組合せ
(32)配列番号145、260、228、89の組合せ
(33)配列番号228、9、229、82の組合せ
(34)配列番号228、9、78、62の組合せ
(35)配列番号229、260、228、157の組合せ
(36)配列番号229、260、228、231の組合せ
(37)配列番号256、228、260、232の組合せ
(38)配列番号228、9、229、117の組合せ
(39)配列番号228、9、229、62の組合せ
(40)配列番号228、9、78、172の組合せ
(41)配列番号228、9、78、3の組合せ
(42)配列番号228、9、82、268、196の組合せ
(43)配列番号229、20、228、4、172の組合せ
(44)配列番号228、9、82、268、233の組合せ
(45)配列番号228、9、268、196、89の組合せ
(46)配列番号228、9、268、160、82の組合せ
(47)配列番号228、9、268、196、256の組合せ
(48)配列番号228、9、78、62、238の組合せ
(49)配列番号228、9、268、196、172の組合せ
(50)配列番号256、228、260、232、196の組合せ
(51)配列番号228、9、268、160、230の組合せ
(52)配列番号256、196、260、268、228の組合せ
(53)配列番号228、9、268、196、62の組合せ
(54)配列番号228、9、268、196、244の組合せ
(55)配列番号228、9、268、196、135の組合せ
(56)配列番号229、260、228、2、217の組合せ
(57)配列番号228、9、268、62、233の組合せ
(58)配列番号229、260、228、172、230の組合せ
(59)配列番号229、260、228、56、33の組合せ
(60)配列番号229、260、228、56、231の組合せ
(61)配列番号229、260、228、231、91の組合せ
(62)配列番号228、9、268、160、245の組合せ
(63)配列番号228、9、82、268、237の組合せ
(64)配列番号228、9、82、268、236の組合せ
(1)配列番号228、9、229の組合せ
(2)配列番号228、9、78の組合せ
(3)配列番号228、9、82の組合せ
(4)配列番号228、9、268の組合せ
(5)配列番号228、9、247の組合せ
(6)配列番号228、9、62の組合せ
(7)配列番号228、9、217の組合せ
(8)配列番号256、228、9の組合せ
(9)配列番号228、9、234の組合せ
(10)配列番号228、9、145の組合せ
(11)配列番号256、196、9の組合せ
(12)配列番号228、9、268、160の組合せ
(13)配列番号256、196、9、68の組合せ
(14)配列番号228、9、82、268の組合せ
(15)配列番号228、9、268、196の組合せ
(16)配列番号228、9、268、230の組合せ
(17)配列番号228、9、268、62の組合せ
(18)配列番号228、9、229、82の組合せ
(19)配列番号228、9、78、62の組合せ
(20)配列番号228、9、229、117の組合せ
(21)配列番号228、9、229、62の組合せ
(22)配列番号228、9、78、172の組合せ
(23)配列番号228、9、78、3の組合せ
(24)配列番号228、9、82、268、196の組合せ
(25)配列番号228、9、82、268、233の組合せ
(26)配列番号228、9、268、196、89の組合せ
(27)配列番号228、9、268、160、82の組合せ
(28)配列番号256、196、9、68、86の組合せ
(29)配列番号228、9、268、196、256の組合せ
(30)配列番号228、9、78、62、238の組合せ
(31)配列番号228、9、268、196、172の組合せ
(32)配列番号228、9、268、160、230の組合せ
(33)配列番号228、9、268、196、62の組合せ
(34)配列番号228、9、268、196、244の組合せ
(35)配列番号228、9、268、196、135の組合せ
(36)配列番号228、9、268、62、233の組合せ
(37)配列番号228、9、268、160、245の組合せ
(38)配列番号256、196、9、68、268の組合せ
(39)配列番号228、9、82、268、237の組合せ
(40)配列番号228、9、82、268、236の組合せ
(1)配列番号256、196の組合せ
(2)配列番号256、196、260の組合せ
(3)配列番号256、196、172の組合せ
(4)配列番号256、196、47の組合せ
(5)配列番号256、196、11の組合せ
(6)配列番号256、196、136の組合せ
(7)配列番号256、196、80の組合せ
(8)配列番号256、196、9の組合せ
(9)配列番号256、196、260、268の組合せ
(10)配列番号256、196、9、68の組合せ
(11)配列番号256、196、260、216の組合せ
(12)配列番号256、196、260、11の組合せ
(13)配列番号228、9、268、196の組合せ
(14)配列番号256、196、260、217の組合せ
(15)配列番号228、9、82、268、196の組合せ
(16)配列番号228、9、268、196、89の組合せ
(17)配列番号256、196、9、68、86の組合せ
(18)配列番号228、9、268、196、256の組合せ
(19)配列番号256、196、260、268、252の組合せ
(20)配列番号228、9、268、196、172の組合せ
(21)配列番号256、228、260、232、196の組合せ
(22)配列番号256、196、260、268、228の組合せ
(23)配列番号256、196、260、268、218の組合せ
(24)配列番号256、196、260、268、267の組合せ
(25)配列番号228、9、268、196、62の組合せ
(26)配列番号228、9、268、196、244の組合せ
(27)配列番号228、9、268、196、135の組合せ
(28)配列番号256、196、260、217、199の組合せ
(29)配列番号256、196、260、268、161の組合せ
(30)配列番号256、196、9、68、268の組合せ
(1)配列番号228、9、229の組合せ
(2)配列番号229、260、228の組合せ
(3)配列番号229、260、230の組合せ
(4)配列番号229、260、169の組合せ
(5)配列番号229、260、160の組合せ
(6)配列番号229、20、228の組合せ
(7)配列番号229、260、228、2の組合せ
(8)配列番号229、260、169、2の組合せ
(9)配列番号229、260、228、172の組合せ
(10)配列番号229、260、228、4の組合せ
(11)配列番号229、260、228、34の組合せ
(12)配列番号229、260、228、56の組合せ
(13)配列番号229、20、228、4の組合せ
(14)配列番号229、20、228、102の組合せ
(15)配列番号228、9、229、82の組合せ
(16)配列番号229、260、228、157の組合せ
(17)配列番号229、260、228、231の組合せ
(18)配列番号228、9、229、117の組合せ
(19)配列番号228、9、229、62の組合せ
(20)配列番号229、20、228、4、172の組合せ
(21)配列番号229、260、228、2、217の組合せ
(22)配列番号229、260、228、172、230の組合せ
(23)配列番号229、260、228、56、33の組合せ
(24)配列番号229、260、228、56、231の組合せ
(25)配列番号229、260、228、231、91の組合せ
(1)配列番号145、260、164の組合せ
(2)配列番号145、260、230の組合せ
(3)配列番号145、260、211の組合せ
(4)配列番号145、260、228の組合せ
(5)配列番号228、9、145の組合せ
(6)配列番号145、260、228、250の組合せ
(7)配列番号145、260、228、62の組合せ
(8)配列番号145、260、228、89の組合せ
(1)配列番号11、164、260の組合せ
(2)配列番号11、164、20の組合せ
(3)配列番号145、260、164の組合せ
本発明はさらに、本発明で使用可能な上記の核酸(或いは、例えば上記3.で説明した本発明のキット又はデバイス)を用いて、検体中のhsa-miR-4675、hsa-miR-4783-3p、hsa-miR-1228-5p、hsa-miR-4532、hsa-miR-6802-5p、hsa-miR-6784-5p、hsa-miR-3940-5p、hsa-miR-1307-3p、hsa-miR-8073、hsa-miR-3184-5p、hsa-miR-1233-5p、hsa-miR-6088、hsa-miR-5195-3p、hsa-miR-320b、hsa-miR-4649-5p、hsa-miR-6800-5p、hsa-miR-1343-3p、hsa-miR-4730、hsa-miR-6885-5p、hsa-miR-5100、hsa-miR-1203、hsa-miR-6756-5p、hsa-miR-373-5p、hsa-miR-1268a、hsa-miR-1260b、hsa-miR-4258、hsa-miR-4697-5p、hsa-miR-1469、hsa-miR-4515、hsa-miR-6861-5p、hsa-miR-6821-5p、hsa-miR-575、hsa-miR-6805-5p、hsa-miR-4758-5p、hsa-miR-3663-3p、hsa-miR-4530、hsa-miR-6798-5p、hsa-miR-6781-5p、hsa-miR-885-3p、hsa-miR-1273g-3p、hsa-miR-4787-3p、hsa-miR-4454、hsa-miR-4706、hsa-miR-1249-3p、hsa-miR-887-3p、hsa-miR-6786-5p、hsa-miR-1238-5p、hsa-miR-6749-5p、hsa-miR-6729-5p、hsa-miR-6825-5p、hsa-miR-663b、hsa-miR-6858-5p、hsa-miR-4690-5p、hsa-miR-6765-5p、hsa-miR-4710、hsa-miR-6775-5p、hsa-miR-371a-5p、hsa-miR-6816-5p、hsa-miR-296-3p、hsa-miR-7977、hsa-miR-8069、hsa-miR-6515-3p、hsa-miR-4687-5p、hsa-miR-1343-5p、hsa-miR-7110-5p、hsa-miR-4525、hsa-miR-3158-5p、hsa-miR-6787-5p、hsa-miR-614、hsa-miR-4689、hsa-miR-1185-2-3p、hsa-miR-1268b、hsa-miR-1228-3p、hsa-miR-1185-1-3p、hsa-miR-940、hsa-miR-939-5p、hsa-miR-6757-5p、hsa-miR-1275、hsa-miR-5001-5p、hsa-miR-6826-5p、hsa-miR-6765-3p、hsa-miR-3679-3p、hsa-miR-4718、hsa-miR-4286、hsa-miR-8059、hsa-miR-4447、hsa-miR-4448、hsa-miR-658、hsa-miR-6766-3p、hsa-miR-197-5p、hsa-miR-6887-5p、hsa-miR-6742-5p、hsa-miR-6729-3p、hsa-miR-5090、hsa-miR-7975、hsa-miR-4505、hsa-miR-6889-5p、hsa-miR-4708-3p、hsa-miR-6131、hsa-miR-1225-3p、hsa-miR-6132、hsa-miR-4734、hsa-miR-3194-3p、hsa-miR-638、hsa-miR-2467-3p、hsa-miR-4728-5p、hsa-miR-5572、hsa-miR-6789-5p、hsa-miR-8063、hsa-miR-4429、hsa-miR-6840-3p、hsa-miR-4476、hsa-miR-675-5p、hsa-miR-711、hsa-miR-6875-5p、hsa-miR-3160-5p、hsa-miR-1908-5p、hsa-miR-6726-5p、hsa-miR-1913、hsa-miR-8071、hsa-miR-3648、hsa-miR-4732-5p、hsa-miR-4787-5p、hsa-miR-3917、hsa-miR-619-5p、hsa-miR-1231、hsa-miR-342-5p、hsa-miR-4433a-5p、hsa-miR-6766-5p、hsa-miR-4707-5p、hsa-miR-7114-5p、hsa-miR-6872-3p、hsa-miR-6780b-5p、hsa-miR-7845-5p、hsa-miR-6798-3p、hsa-miR-665、hsa-miR-6848-5p、hsa-miR-5008-5p、hsa-miR-4294、hsa-miR-6511a-5p、hsa-miR-4435、hsa-miR-4747-3p、hsa-miR-6880-3p、hsa-miR-6869-5p、hsa-miR-7150、hsa-miR-1260a、hsa-miR-6877-5p、hsa-miR-6721-5p、hsa-miR-4656、hsa-miR-1229-5p、hsa-miR-4433a-3p、hsa-miR-4274、hsa-miR-4419b、hsa-miR-4674、hsa-miR-6893-5p、hsa-miR-6763-3p、hsa-miR-6762-5p、hsa-miR-6738-5p、hsa-miR-4513、hsa-miR-6746-5p、hsa-miR-6880-5p、hsa-miR-4736、hsa-miR-718、hsa-miR-6717-5p、hsa-miR-7847-3p、hsa-miR-760、hsa-miR-1199-5p、hsa-miR-6813-5p、hsa-miR-6769a-5p、hsa-miR-1193、hsa-miR-7108-3p、hsa-miR-6741-5p、hsa-miR-4298、hsa-miR-6796-3p、hsa-miR-4750-5p、hsa-miR-6785-5p、hsa-miR-1292-3p、hsa-miR-4749-3p、hsa-miR-6800-3p、hsa-miR-4722-5p、hsa-miR-4746-3p、hsa-miR-4450、hsa-miR-6795-5p、hsa-miR-365a-5p、hsa-miR-498、hsa-miR-6797-5p、hsa-miR-1470、hsa-miR-6851-5p、hsa-miR-1247-3p、hsa-miR-5196-5p、hsa-miR-208a-5p、hsa-miR-6842-5p、hsa-miR-150-3p、hsa-miR-4534、hsa-miR-3135b、hsa-miR-3131、hsa-miR-4792、hsa-miR-6510-5p、hsa-miR-504-3p、hsa-miR-3619-3p、hsa-miR-671-5p、hsa-miR-4667-5p、hsa-miR-4430、hsa-miR-3195、hsa-miR-3679-5p、hsa-miR-6076、hsa-miR-6515-5p、hsa-miR-6820-5p、hsa-miR-4634、hsa-miR-187-5p、hsa-miR-6763-5p、hsa-miR-1908-3p、hsa-miR-1181、hsa-miR-6782-5p、hsa-miR-5010-5p、hsa-miR-6870-5p、hsa-miR-6124、hsa-miR-1249-5p、hsa-miR-6511b-5p、hsa-miR-1254、hsa-miR-4727-3p、hsa-miR-4259、hsa-miR-4771、hsa-miR-3622a-5p、hsa-miR-4480、hsa-miR-4740-5p、hsa-miR-6777-5p、hsa-miR-6794-5p、hsa-miR-4687-3p、hsa-miR-6743-5p、hsa-miR-6771-5p、hsa-miR-3141、hsa-miR-3162-5p、hsa-miR-4271、hsa-miR-1227-5p、hsa-miR-4257、hsa-miR-4270、hsa-miR-4516、hsa-miR-4651、hsa-miR-4725-3p、hsa-miR-6125、hsa-miR-6732-5p、hsa-miR-6791-5p、hsa-miR-6819-5p、hsa-miR-6891-5p、hsa-miR-7108-5p、hsa-miR-7109-5p、hsa-miR-642b-3p及びhsa-miR-642a-3pで表される卵巣腫瘍由来の遺伝子の発現量、並びに場合により、hsa-miR-320a、hsa-miR-663a、hsa-miR-328-5p、hsa-miR-128-2-5p、hsa-miR-125a-3p、hsa-miR-191-5p、hsa-miR-92b-5p、hsa-miR-296-5p、hsa-miR-1246、hsa-miR-92a-2-5p、hsa-miR-128-1-5p、hsa-miR-1290、hsa-miR-211-3p、hsa-miR-744-5p、hsa-miR-135a-3p、hsa-miR-451a、hsa-miR-625-3p、hsa-miR-92a-3p、hsa-miR-422a、hsa-miR-483-5p、hsa-miR-652-5p、hsa-miR-24-3p、hsa-miR-23b-3p、hsa-miR-23a-3p、hsa-miR-92b-3p、hsa-miR-22-3pで表される卵巣腫瘍由来の遺伝子の発現量の1つ以上(例えば発現プロフィール)をin vitroで測定し、該測定された発現量(及び任意に同様に測定された健常者の対照発現量)を用いて被験体が卵巣腫瘍に罹患しているか否かをin vitroで評価することを含む、卵巣腫瘍の検出方法を提供する。本方法において、例えば、卵巣腫瘍の罹患が疑われる被験体と、卵巣腫瘍に罹患していない被験体とから採取した血液、血清、血漿等の検体について、検体中の上記遺伝子の発現量と、卵巣腫瘍に罹患していない被験体の対照発現量とを用いて、(例えば両発現量を比較して、)当該検体中の標的核酸の発現量に差がある場合、被験体が、卵巣腫瘍に罹患していると評価することができる。
(a)被験体由来の検体を、in vitroで、本発明のキット又はデバイスのポリヌクレオチドと接触させるステップ、
(b)検体中の標的核酸の発現量を、上記ポリヌクレオチドを核酸プローブ又はプライマーとして用いて測定するステップ、
(c)(b)の結果をもとに、当該被験体中の卵巣腫瘍(細胞)の存在又は不存在を評価するステップ、
を含むことができる。
749-3pがhsa-miR-4749-3pであり、miR-6800-3pがhsa-miR-6800-3pであり、miR-4722-5pがhsa-miR-4722-5pであり、miR-4746-3pがhsa-miR-4746-3pであり、miR-4450がhsa-miR-4450であり、miR-6795-5pがhsa-miR-6795-5pであり、miR-365a-5pがhsa-miR-365a-5pであり、miR-498がhsa-miR-498であり、miR-6797-5pがhsa-miR-6797-5pであり、miR-1470がhsa-miR-1470であり、miR-6851-5pがhsa-miR-6851-5pであり、miR-1247-3pがhsa-miR-1247-3pであり、miR-5196-5pがhsa-miR-5196-5pであり、miR-208a-5pがhsa-miR-208a-5pであり、miR-6842-5pがhsa-miR-6842-5pであり、miR-150-3pがhsa-miR-150-3pであり、miR-4534がhsa-miR-4534であり、miR-3135bがhsa-miR-3135bであり、miR-3131がhsa-miR-3131であり、miR-4792がhsa-miR-4792であり、miR-6510-5pがhsa-miR-6510-5pであり、miR-504-3pがhsa-miR-504-3pであり、miR-3619-3pがhsa-miR-3619-3pであり、miR-671-5pがhsa-miR-671-5pであり、miR-4667-5pがhsa-miR-4667-5pであり、miR-4430がhsa-miR-4430であり、miR-3195がhsa-miR-3195であり、miR-3679-5pがhsa-miR-3679-5pであり、miR-6076がhsa-miR-6076であり、miR-6515-5pがhsa-miR-6515-5pであり、miR-6820-5pがhsa-miR-6820-5pであり、miR-4634がhsa-miR-4634であり、miR-187-5pがhsa-miR-187-5pであり、miR-6763-5pがhsa-miR-6763-5pであり、miR-1908-3pがhsa-miR-1908-3pであり、miR-1181がhsa-miR-1181であり、miR-6782-5pがhsa-miR-6782-5pであり、miR-5010-5pがhsa-miR-5010-5pであり、miR-6870-5pがhsa-miR-6870-5pであり、miR-6124がhsa-miR-6124であり、miR-1249-5pがhsa-miR-1249-5pであり、miR-6511b-5pがhsa-miR-6511b-5pであり、miR-1254がhsa-miR-1254であり、miR-4727-3pがhsa-miR-4727-3pであり、miR-4259がhsa-miR-4259であり、miR-4771がhsa-miR-4771であり、miR-3622a-5pがhsa-miR-3622a-5pであり、miR-4480がhsa-miR-4480であり、miR-4740-5pがhsa-miR-4740-5pであり、miR-6777-5pがhsa-miR-6777-5pであり、miR-6794-5pがhsa-miR-6794-5pであり、miR-4687-3pがhsa-miR-4687-3pであり、miR-6743-5pがhsa-miR-6743-5pであり、miR-6771-5pがhsa-miR-6771-5pであり、miR-3141がhsa-miR-3141であり、miR-3162-5pがhsa-miR-3162-5pであり、miR-4271がhsa-miR-4271であり、miR-1227-5pがhsa-miR-1227-5pであり、miR-4257がhsa-miR-4257であり、miR-4270がhsa-miR-4270であり、miR-4516がhsa-miR-4516であり、miR-4651がhsa-miR-4651であり、miR-4725-3pがhsa-miR-4725-3pであり、miR-6125がhsa-miR-6125であり、miR-6732-5pがhsa-miR-6732-5pであり、miR-6791-5pがhsa-miR-6791-5pであり、miR-6819-5pがhsa-miR-6819-5pであり、miR-6891-5pがhsa-miR-6891-5pであり、miR-7108-5pがhsa-miR-7108-5pであり、miR-7109-5pがhsa-miR-7109-5pであり、miR-642b-3pがhsa-miR-642b-3pであり、miR-642a-3pがhsa-miR-642a-3pである。
(a)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~247、251及び268のいずれかで表される塩基配列を含むポリヌクレオチド、
(c)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択される。
(f)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列を含むポリヌクレオチド、
(h)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されてもよい。
(a)被験体の検体から調製されたRNA(ここでステップ(b)の定量RT-PCRのために、例えばRNAの3'末端はポリアデニル化されてもよい。)又はそれから転写された相補的ポリヌクレオチド(cDNA)を、本発明のキットのポリヌクレオチドと結合させるステップ、
(b)当該ポリヌクレオチドに結合した検体由来のRNA又は当該RNAから合成されたcDNAを、上記ポリヌクレオチドを核酸プローブとして用いるハイブリダイゼーションによって、あるいは上記ポリヌクレオチドをプライマーとして用いる定量RT-PCRによって、測定するステップ、
(c)上記(b)の測定結果に基づいて、卵巣腫瘍(又は卵巣腫瘍由来の遺伝子)の存在又は不存在を評価するステップ、
を含むことができる。
(a)卵巣腫瘍患者由来であること及び卵巣腫瘍を含まない被験体であることが既に知られている検体中の標的遺伝子の発現量を、本発明による検出用もしくは診断用ポリヌクレオチド、キット又はデバイス(例えばDNAチップ)を用いて測定するステップ、
(b)(a)で測定された発現量の測定値から、上記の式1~3、5及び6の判別式を作成するステップ、
(c)被験体由来の検体中の当該標的遺伝子の発現量を、本発明による検出用もしくは診断用ポリヌクレオチド、キット又はデバイス(例えばDNAチップ)を用いて測定し、(b)で作成した判別式にそれらを代入して、得られた結果に基づいて被験体が卵巣腫瘍を含むこと又は卵巣腫瘍を含まないことを決定又は評価する、或いは卵巣腫瘍由来発現量を、卵巣腫瘍に罹患していない被験体(例えば健常者を含む。)由来の対照と比較し評価する、ステップを含むことができる。
(1)配列番号1~247、251及び268のいずれかで表される塩基配列又はその相補的配列において、15以上の連続した塩基を含む核酸(例えばDNA、RNAなど)のいずれかによって測定される卵巣腫瘍患者及び卵巣腫瘍に罹患していない被験者の血清における遺伝子発現量。
(2)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列又はその相補的配列において、15以上の連続した塩基を含む核酸(例えばDNA、RNAなど)のいずれかによって測定される卵巣腫瘍患者及び卵巣腫瘍に罹患していない被験者の血清における遺伝子発現量。
<検体の採取>
インフォームドコンセントを得た健常者400人、良性骨軟部腫瘍患者327人、乳良性疾患患者41人、卵巣がん患者406人、卵巣良性腫瘍患者28人、乳がん・胆道がん・膵臓がん・大腸がん・食道がん・胃がん・肝臓がん・肺がん患者各50人からベノジェクトII真空採血管VP-AS109K60(テルモ株式会社(日本))を用いてそれぞれ血清を採取した。なお、以下では、乳がん・胆道がん・膵臓がん・大腸がん・食道がん・胃がん・肝臓がん・肺がん患者を包括して「卵巣がん以外のがん患者」とする。
検体として上記健常者400人、良性骨軟部腫瘍及び乳良性疾患患者368人、卵巣腫瘍患者434人、卵巣がん以外のがん患者400人の合計1,602人からそれぞれ得られた血清300μLから、3D-Gene(登録商標)RNA extraction reagent from liquid sample kit(東レ株式会社(日本))中のRNA抽出用試薬を用いて、同社の定めるプロトコールに従ってtotal RNAを得た。
検体として上記健常者400人、良性骨軟部腫瘍及び乳良性疾患患者368人、卵巣腫瘍患者434人、卵巣がん以外のがん患者400人の合計1,602人の血清から得たtotal RNAに対して、3D-Gene(登録商標)miRNA Labeling kit(東レ株式会社)を用いて同社が定めるプロトコールに基づいてmiRNAを蛍光標識した。オリゴDNAチップとして、miRBase release 21に登録されているmiRNAの中で、2,565種のmiRNAと相補的な配列を有するプローブを搭載した3D-Gene(登録商標)Human miRNA Oligo chip(東レ株式会社)を用い、同社が定めるプロトコールに基づいてストリンジェントな条件でハイブリダイゼーション及びハイブリダイゼーション後の洗浄を行った。DNAチップを3D-Gene(登録商標)スキャナー(東レ株式会社)を用いてスキャンし、画像を取得して3D-Gene(登録商標)Extraction(東レ株式会社)にて蛍光強度を数値化した。数値化された蛍光強度を、底が2の対数値に変換して遺伝子発現量とし、ブランク値の減算を行い、欠損値はシグナル値0.1で置換した。その結果、健常者400人、良性骨軟部腫瘍及び乳良性疾患患者368人、卵巣腫瘍患者434人、卵巣がん以外のがん患者400人の血清に対する、網羅的なmiRNAの遺伝子発現量を得た。次に、各検体群の70%の検体を学習検体群に、30%の検体をテスト検体群に振り分けた。すなわち、健常者280人、良性骨軟部腫瘍及び乳良性疾患患者257人、卵巣腫瘍患者303人、卵巣がん以外のがん患者280人を学習検体群として、健常者120人、良性骨軟部腫瘍及び乳良性疾患患者111人、卵巣腫瘍患者131人、卵巣がん以外のがん患者120人をテスト検体群とした。数値化されたmiRNAの遺伝子発現量を用いた計算及び統計解析は、R言語3.3.1(R Core Team(2016).R:A language and environment for statistical computing.R Foundation for Statistical Computing、Vienna、Austria.URL https://www.R-project.org/.)及びMASSパッケージ7.3.45(Venables、W.N.&Ripley、B.D.(2002)Modern Applied Statistics with S.Fourth Edition.Springer、New York.ISBN 0-387-95457-0)を用いて実施した。
<健常者との比較による卵巣腫瘍遺伝子マーカーの選定と単独の遺伝子マーカーによる卵巣腫瘍判別性能の評価方法>
本実施例では、卵巣腫瘍患者と健常者において発現量に有意差が見られるmiRNAを卵巣腫瘍遺伝子マーカーとして選定し、学習検体群を用いて1個または2個の遺伝子マーカーによる判別式を作成した上で、テスト検体群の判別精度を計算することで、上記の遺伝子マーカーによる卵巣腫瘍患者と健常者の判別性能を評価した。
<良性骨軟部腫瘍及び乳良性疾患患者との比較による卵巣腫瘍遺伝子マーカーの選定と単独の遺伝子マーカーによる卵巣腫瘍判別性能の評価方法>
本実施例では、卵巣腫瘍と良性骨軟部腫瘍及び乳良性疾患において発現量に有意差が見られるmiRNAを卵巣腫瘍遺伝子マーカーとして選定し、学習検体群を用いて1個または2個の遺伝子マーカーによる判別式を作成した上で、テスト検体群の判別精度を計算することで、上記の遺伝子マーカーによる卵巣腫瘍と良性骨軟部腫瘍及び乳良性疾患の判別性能を評価した。
<卵巣がん以外のがん患者との比較による卵巣腫瘍遺伝子マーカーの選定と単独の遺伝子マーカーによる卵巣腫瘍の判別性能の評価方法>
本実施例では、卵巣腫瘍と卵巣がん以外のがんである、乳がん・胆道がん・膵臓がん・大腸がん・食道がん・胃がん・肝臓がん・肺がんにおいて発現量に有意差が見られるmiRNAを卵巣腫瘍遺伝子マーカーとして選定し、学習検体群を用いて1個または2個の遺伝子マーカーによる判別式を作成した上で、テスト検体群の判別精度を計算することで、上記の遺伝子マーカーによる卵巣腫瘍と卵巣がん以外のがんの判別性能を評価した。
<複数の遺伝子マーカーの組合せによる卵巣腫瘍判別性能の評価方法>
本実施例では、上記実施例1、実施例2及び実施例3で用いた健常者、良性骨軟部腫瘍及び乳良性疾患患者及び卵巣がん以外のがん患者(すなわち、卵巣腫瘍に罹患していない被験者)を陰性検体群、卵巣腫瘍患者を陽性検体群として、卵巣腫瘍の判別性能が高い遺伝子マーカーの組み合せを探索した。
本明細書で引用した全ての刊行物、特許及び特許出願はそのまま引用により本明細書に組み入れられるものとする。
Claims (18)
- 卵巣腫瘍マーカーである、miR-4675、miR-4783-3p、miR-1228-5p、miR-4532、miR-6802-5p、miR-6784-5p、miR-3940-5p、miR-1307-3p、miR-8073、miR-3184-5p、miR-1233-5p、miR-6088、miR-5195-3p、miR-320b、miR-4649-5p、miR-6800-5p、miR-1343-3p、miR-4730、miR-6885-5p、miR-5100、miR-1203、miR-6756-5p、miR-373-5p、miR-1268a、miR-1260b、miR-4258、miR-4697-5p、miR-1469、miR-4515、miR-6861-5p、miR-6821-5p、miR-575、miR-6805-5p、miR-4758-5p、miR-3663-3p、miR-4530、miR-6798-5p、miR-6781-5p、miR-885-3p、miR-1273g-3p、miR-4787-3p、miR-4454、miR-4706、miR-1249-3p、miR-887-3p、miR-6786-5p、miR-1238-5p、miR-6749-5p、miR-6729-5p、miR-6825-5p、miR-663b、miR-6858-5p、miR-4690-5p、miR-6765-5p、miR-4710、miR-6775-5p、miR-371a-5p、miR-6816-5p、miR-296-3p、miR-7977、miR-8069、miR-6515-3p、miR-4687-5p、miR-1343-5p、miR-7110-5p、miR-4525、miR-3158-5p、miR-6787-5p、miR-614、miR-4689、miR-1185-2-3p、miR-1268b、miR-1228-3p、miR-1185-1-3p、miR-940、miR-939-5p、miR-6757-5p、miR-1275、miR-5001-5p、miR-6826-5p、miR-6765-3p、miR-3679-3p、miR-4718、miR-4286、miR-8059、miR-4447、miR-4448、miR-658、miR-6766-3p、miR-197-5p、miR-6887-5p、miR-6742-5p、miR-6729-3p、miR-5090、miR-7975、miR-4505、miR-6889-5p、miR-4708-3p、miR-6131、miR-1225-3p、miR-6132、miR-4734、miR-3194-3p、miR-638、miR-2467-3p、miR-4728-5p、miR-5572、miR-6789-5p、miR-8063、miR-4429、miR-6840-3p、miR-4476、miR-675-5p、miR-711、miR-6875-5p、miR-3160-5p、miR-1908-5p、miR-6726-5p、miR-1913、miR-8071、miR-3648、miR-4732-5p、miR-4787-5p、miR-3917、miR-619-5p、miR-1231、miR-342-5p、miR-4433a-5p、miR-6766-5p、miR-4707-5p、miR-7114-5p、miR-6872-3p、miR-6780b-5p、miR-7845-5p、miR-6798-3p、miR-665、miR-6848-5p、miR-5008-5p、miR-4294、miR-6511a-5p、miR-4435、miR-4747-3p、miR-6880-3p、miR-6869-5p、miR-7150、miR-1260a、miR-6877-5p、miR-6721-5p、miR-4656、miR-1229-5p、miR-4433a-3p、miR-4274、miR-4419b、miR-4674、miR-6893-5p、miR-6763-3p、miR-6762-5p、miR-6738-5p、miR-4513、miR-6746-5p、miR-6880-5p、miR-4736、miR-718、miR-6717-5p、miR-7847-3p、miR-760、miR-1199-5p、miR-6813-5p、miR-6769a-5p、miR-1193、miR-7108-3p、miR-6741-5p、miR-4298、miR-6796-3p、miR-4750-5p、miR-6785-5p、miR-1292-3p、miR-4749-3p、miR-6800-3p、miR-4722-5p、miR-4746-3p、miR-4450、miR-6795-5p、miR-365a-5p、miR-498、miR-6797-5p、miR-1470、miR-6851-5p、miR-1247-3p、miR-5196-5p、miR-208a-5p、miR-6842-5p、miR-150-3p、miR-4534、miR-3135b、miR-3131、miR-4792、miR-6510-5p、miR-504-3p、miR-3619-3p、miR-671-5p、miR-4667-5p、miR-4430、miR-3195、miR-3679-5p、miR-6076、miR-6515-5p、miR-6820-5p、miR-4634、miR-187-5p、miR-6763-5p、miR-1908-3p、miR-1181、miR-6782-5p、miR-5010-5p、miR-6870-5p、miR-6124、miR-1249-5p、miR-6511b-5p、miR-1254、miR-4727-3p、miR-4259、miR-4771、miR-3622a-5p、miR-4480、miR-4740-5p、miR-6777-5p、miR-6794-5p、miR-4687-3p、miR-6743-5p、miR-6771-5p、miR-3141、miR-3162-5p、miR-4271、miR-1227-5p、miR-4257、miR-4270、miR-4516、miR-4651、miR-4725-3p、miR-6125、miR-6732-5p、miR-6791-5p、miR-6819-5p、miR-6891-5p、miR-7108-5p、miR-7109-5p、miR-642b-3p及びmiR-642a-3pから選択される少なくとも1つのポリヌクレオチド、又は当該ポリヌクレオチドの相補鎖と特異的に結合可能な核酸を含む、卵巣腫瘍の検出用キット。
- 前記核酸が、下記の(a)~(e)のいずれかに示すポリヌクレオチド:
(a)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~247、251及び268のいずれかで表される塩基配列を含むポリヌクレオチド、
(c)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項1に記載のキット。 - 前記キットが、別の卵巣腫瘍マーカーである、miR-320a、miR-663a、miR-328-5p、miR-128-2-5p、miR-125a-3p、miR-191-5p、miR-92b-5p、miR-296-5p、miR-1246、miR-92a-2-5p、miR-128-1-5p、miR-1290、miR-211-3p、miR-744-5p、miR-135a-3p、miR-451a、miR-625-3p、miR-92a-3p、miR-422a、miR-483-5p、miR-652-5p、miR-24-3p、miR-23b-3p、miR-23a-3p、miR-92b-3p、miR-22-3pから選択される少なくとも1つのポリヌクレオチド、又は当該ポリヌクレオチドの相補鎖と特異的に結合可能な核酸をさらに含む、請求項1又は2に記載のキット。
- 前記核酸が、下記の(f)~(j)のいずれかに示すポリヌクレオチド:
(f)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列を含むポリヌクレオチド、
(h)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項3に記載のキット。 - 卵巣腫瘍マーカーである、miR-4675、miR-4783-3p、miR-1228-5p、miR-4532、miR-6802-5p、miR-6784-5p、miR-3940-5p、miR-1307-3p、miR-8073、miR-3184-5p、miR-1233-5p、miR-6088、miR-5195-3p、miR-320b、miR-4649-5p、miR-6800-5p、miR-1343-3p、miR-4730、miR-6885-5p、miR-5100、miR-1203、miR-6756-5p、miR-373-5p、miR-1268a、miR-1260b、miR-4258、miR-4697-5p、miR-1469、miR-4515、miR-6861-5p、miR-6821-5p、miR-575、miR-6805-5p、miR-4758-5p、miR-3663-3p、miR-4530、miR-6798-5p、miR-6781-5p、miR-885-3p、miR-1273g-3p、miR-4787-3p、miR-4454、miR-4706、miR-1249-3p、miR-887-3p、miR-6786-5p、miR-1238-5p、miR-6749-5p、miR-6729-5p、miR-6825-5p、miR-663b、miR-6858-5p、miR-4690-5p、miR-6765-5p、miR-4710、miR-6775-5p、miR-371a-5p、miR-6816-5p、miR-296-3p、miR-7977、miR-8069、miR-6515-3p、miR-4687-5p、miR-1343-5p、miR-7110-5p、miR-4525、miR-3158-5p、miR-6787-5p、miR-614、miR-4689、miR-1185-2-3p、miR-1268b、miR-1228-3p、miR-1185-1-3p、miR-940、miR-939-5p、miR-6757-5p、miR-1275、miR-5001-5p、miR-6826-5p、miR-6765-3p、miR-3679-3p、miR-4718、miR-4286、miR-8059、miR-4447、miR-4448、miR-658、miR-6766-3p、miR-197-5p、miR-6887-5p、miR-6742-5p、miR-6729-3p、miR-5090、miR-7975、miR-4505、miR-6889-5p、miR-4708-3p、miR-6131、miR-1225-3p、miR-6132、miR-4734、miR-3194-3p、miR-638、miR-2467-3p、miR-4728-5p、miR-5572、miR-6789-5p、miR-8063、miR-4429、miR-6840-3p、miR-4476、miR-675-5p、miR-711、miR-6875-5p、miR-3160-5p、miR-1908-5p、miR-6726-5p、miR-1913、miR-8071、miR-3648、miR-4732-5p、miR-4787-5p、miR-3917、miR-619-5p、miR-1231、miR-342-5p、miR-4433a-5p、miR-6766-5p、miR-4707-5p、miR-7114-5p、miR-6872-3p、miR-6780b-5p、miR-7845-5p、miR-6798-3p、miR-665、miR-6848-5p、miR-5008-5p、miR-4294、miR-6511a-5p、miR-4435、miR-4747-3p、miR-6880-3p、miR-6869-5p、miR-7150、miR-1260a、miR-6877-5p、miR-6721-5p、miR-4656、miR-1229-5p、miR-4433a-3p、miR-4274、miR-4419b、miR-4674、miR-6893-5p、miR-6763-3p、miR-6762-5p、miR-6738-5p、miR-4513、miR-6746-5p、miR-6880-5p、miR-4736、miR-718、miR-6717-5p、miR-7847-3p、miR-760、miR-1199-5p、miR-6813-5p、miR-6769a-5p、miR-1193、miR-7108-3p、miR-6741-5p、miR-4298、miR-6796-3p、miR-4750-5p、miR-6785-5p、miR-1292-3p、miR-4749-3p、miR-6800-3p、miR-4722-5p、miR-4746-3p、miR-4450、miR-6795-5p、miR-365a-5p、miR-498、miR-6797-5p、miR-1470、miR-6851-5p、miR-1247-3p、miR-5196-5p、miR-208a-5p、miR-6842-5p、miR-150-3p、miR-4534、miR-3135b、miR-3131、miR-4792、miR-6510-5p、miR-504-3p、miR-3619-3p、miR-671-5p、miR-4667-5p、miR-4430、miR-3195、miR-3679-5p、miR-6076、miR-6515-5p、miR-6820-5p、miR-4634、miR-187-5p、miR-6763-5p、miR-1908-3p、miR-1181、miR-6782-5p、miR-5010-5p、miR-6870-5p、miR-6124、miR-1249-5p、miR-6511b-5p、miR-1254、miR-4727-3p、miR-4259、miR-4771、miR-3622a-5p、miR-4480、miR-4740-5p、miR-6777-5p、miR-6794-5p、miR-4687-3p、miR-6743-5p、miR-6771-5p、miR-3141、miR-3162-5p、miR-4271、miR-1227-5p、miR-4257、miR-4270、miR-4516、miR-4651、miR-4725-3p、miR-6125、miR-6732-5p、miR-6791-5p、miR-6819-5p、miR-6891-5p、miR-7108-5p、miR-7109-5p、miR-642b-3p及びmiR-642a-3pから選択される少なくとも1つのポリヌクレオチド、又は当該ポリヌクレオチドの相補鎖と特異的に結合可能な核酸を含む、卵巣腫瘍の検出用デバイス。
- 前記核酸が、下記の(a)~(e)のいずれかに示すポリヌクレオチド:
(a)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~247、251及び268のいずれかで表される塩基配列を含むポリヌクレオチド、
(c)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~247、251及び268のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項5に記載のデバイス。 - 前記デバイスが、別の卵巣腫瘍マーカーである、miR-320a、miR-663a、miR-328-5p、miR-128-2-5p、miR-125a-3p、miR-191-5p、miR-92b-5p、miR-296-5p、miR-1246、miR-92a-2-5p、miR-128-1-5p、miR-1290、miR-211-3p、miR-744-5p、miR-135a-3p、miR-451a、miR-625-3p、miR-92a-3p、miR-422a、miR-483-5p、miR-652-5p、miR-24-3p、miR-23b-3p、miR-23a-3p、miR-92b-3p、miR-22-3pから選択される少なくとも1つのポリヌクレオチド、又は当該ポリヌクレオチドの相補鎖と特異的に結合可能な核酸をさらに含む、請求項5又は6に記載のデバイス。
- 前記核酸が、下記の(f)~(j)のいずれかに示すポリヌクレオチド:
(f)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列を含むポリヌクレオチド、
(h)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列もしくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項7に記載のデバイス。 - 前記デバイスが、ハイブリダイゼーション技術による測定のためのデバイスである、請求項5~8のいずれか1項に記載のデバイス。
- 前記ハイブリダイゼーション技術が、核酸アレイ技術である、請求項9に記載のデバイス。
- 被験体の検体において、卵巣腫瘍マーカーである、miR-4675、miR-4783-3p、miR-1228-5p、miR-4532、miR-6802-5p、miR-6784-5p、miR-3940-5p、miR-1307-3p、miR-8073、miR-3184-5p、miR-1233-5p、miR-6088、miR-5195-3p、miR-320b、miR-4649-5p、miR-6800-5p、miR-1343-3p、miR-4730、miR-6885-5p、miR-5100、miR-1203、miR-6756-5p、miR-373-5p、miR-1268a、miR-1260b、miR-4258、miR-4697-5p、miR-1469、miR-4515、miR-6861-5p、miR-6821-5p、miR-575、miR-6805-5p、miR-4758-5p、miR-3663-3p、miR-4530、miR-6798-5p、miR-6781-5p、miR-885-3p、miR-1273g-3p、miR-4787-3p、miR-4454、miR-4706、miR-1249-3p、miR-887-3p、miR-6786-5p、miR-1238-5p、miR-6749-5p、miR-6729-5p、miR-6825-5p、miR-663b、miR-6858-5p、miR-4690-5p、miR-6765-5p、miR-4710、miR-6775-5p、miR-371a-5p、miR-6816-5p、miR-296-3p、miR-7977、miR-8069、miR-6515-3p、miR-4687-5p、miR-1343-5p、miR-7110-5p、miR-4525、miR-3158-5p、miR-6787-5p、miR-614、miR-4689、miR-1185-2-3p、miR-1268b、miR-1228-3p、miR-1185-1-3p、miR-940、miR-939-5p、miR-6757-5p、miR-1275、miR-5001-5p、miR-6826-5p、miR-6765-3p、miR-3679-3p、miR-4718、miR-4286、miR-8059、miR-4447、miR-4448、miR-658、miR-6766-3p、miR-197-5p、miR-6887-5p、miR-6742-5p、miR-6729-3p、miR-5090、miR-7975、miR-4505、miR-6889-5p、miR-4708-3p、miR-6131、miR-1225-3p、miR-6132、miR-4734、miR-3194-3p、miR-638、miR-2467-3p、miR-4728-5p、miR-5572、miR-6789-5p、miR-8063、miR-4429、miR-6840-3p、miR-4476、miR-675-5p、miR-711、miR-6875-5p、miR-3160-5p、miR-1908-5p、miR-6726-5p、miR-1913、miR-8071、miR-3648、miR-4732-5p、miR-4787-5p、miR-3917、miR-619-5p、miR-1231、miR-342-5p、miR-4433a-5p、miR-6766-5p、miR-4707-5p、miR-7114-5p、miR-6872-3p、miR-6780b-5p、miR-7845-5p、miR-6798-3p、miR-665、miR-6848-5p、miR-5008-5p、miR-4294、miR-6511a-5p、miR-4435、miR-4747-3p、miR-6880-3p、miR-6869-5p、miR-7150、miR-1260a、miR-6877-5p、miR-6721-5p、miR-4656、miR-1229-5p、miR-4433a-3p、miR-4274、miR-4419b、miR-4674、miR-6893-5p、miR-6763-3p、miR-6762-5p、miR-6738-5p、miR-4513、miR-6746-5p、miR-6880-5p、miR-4736、miR-718、miR-6717-5p、miR-7847-3p、miR-760、miR-1199-5p、miR-6813-5p、miR-6769a-5p、miR-1193、miR-7108-3p、miR-6741-5p、miR-4298、miR-6796-3p、miR-4750-5p、miR-6785-5p、miR-1292-3p、miR-4749-3p、miR-6800-3p、miR-4722-5p、miR-4746-3p、miR-4450、miR-6795-5p、miR-365a-5p、miR-498、miR-6797-5p、miR-1470、miR-6851-5p、miR-1247-3p、miR-5196-5p、miR-208a-5p、miR-6842-5p、miR-150-3p、miR-4534、miR-3135b、miR-3131、miR-4792、miR-6510-5p、miR-504-3p、miR-3619-3p、miR-671-5p、miR-4667-5p、miR-4430、miR-3195、miR-3679-5p、miR-6076、miR-6515-5p、miR-6820-5p、miR-4634、miR-187-5p、miR-6763-5p、miR-1908-3p、miR-1181、miR-6782-5p、miR-5010-5p、miR-6870-5p、miR-6124、miR-1249-5p、miR-6511b-5p、miR-1254、miR-4727-3p、miR-4259、miR-4771、miR-3622a-5p、miR-4480、miR-4740-5p、miR-6777-5p、miR-6794-5p、miR-4687-3p、miR-6743-5p、miR-6771-5p、miR-3141、miR-3162-5p、miR-4271、miR-1227-5p、miR-4257、miR-4270、miR-4516、miR-4651、miR-4725-3p、miR-6125、miR-6732-5p、miR-6791-5p、miR-6819-5p、miR-6891-5p、miR-7108-5p、miR-7109-5p、miR-642b-3p、miR-642a-3pから選択される少なくとも1つのポリヌクレオチドの発現量を測定し、該測定された発現量を用いて被験体が卵巣腫瘍に罹患していること、又は卵巣腫瘍に罹患していないことをin vitroで評価することを含む、卵巣腫瘍の検出方法。
- 卵巣腫瘍マーカーである、miR-4675、miR-4783-3p、miR-1228-5p、miR-4532、miR-6802-5p、miR-6784-5p、miR-3940-5p、miR-1307-3p、miR-8073、miR-3184-5p、miR-1233-5p、miR-6088、miR-5195-3p、miR-320b、miR-4649-5p、miR-6800-5p、miR-1343-3p、miR-4730、miR-6885-5p、miR-5100、miR-1203、miR-6756-5p、miR-373-5p、miR-1268a、miR-1260b、miR-4258、miR-4697-5p、miR-1469、miR-4515、miR-6861-5p、miR-6821-5p、miR-575、miR-6805-5p、miR-4758-5p、miR-3663-3p、miR-4530、miR-6798-5p、miR-6781-5p、miR-885-3p、miR-1273g-3p、miR-4787-3p、miR-4454、miR-4706、miR-1249-3p、miR-887-3p、miR-6786-5p、miR-1238-5p、miR-6749-5p、miR-6729-5p、miR-6825-5p、miR-663b、miR-6858-5p、miR-4690-5p、miR-6765-5p、miR-4710、miR-6775-5p、miR-371a-5p、miR-6816-5p、miR-296-3p、miR-7977、miR-8069、miR-6515-3p、miR-4687-5p、miR-1343-5p、miR-7110-5p、miR-4525、miR-3158-5p、miR-6787-5p、miR-614、miR-4689、miR-1185-2-3p、miR-1268b、miR-1228-3p、miR-1185-1-3p、miR-940、miR-939-5p、miR-6757-5p、miR-1275、miR-5001-5p、miR-6826-5p、miR-6765-3p、miR-3679-3p、miR-4718、miR-4286、miR-8059、miR-4447、miR-4448、miR-658、miR-6766-3p、miR-197-5p、miR-6887-5p、miR-6742-5p、miR-6729-3p、miR-5090、miR-7975、miR-4505、miR-6889-5p、miR-4708-3p、miR-6131、miR-1225-3p、miR-6132、miR-4734、miR-3194-3p、miR-638、miR-2467-3p、miR-4728-5p、miR-5572、miR-6789-5p、miR-8063、miR-4429、miR-6840-3p、miR-4476、miR-675-5p、miR-711、miR-6875-5p、miR-3160-5p、miR-1908-5p、miR-6726-5p、miR-1913、miR-8071、miR-3648、miR-4732-5p、miR-4787-5p、miR-3917、miR-619-5p、miR-1231、miR-342-5p、miR-4433a-5p、miR-6766-5p、miR-4707-5p、miR-7114-5p、miR-6872-3p、miR-6780b-5p、miR-7845-5p、miR-6798-3p、miR-665、miR-6848-5p、miR-5008-5p、miR-4294、miR-6511a-5p、miR-4435、miR-4747-3p、miR-6880-3p、miR-6869-5p、miR-7150、miR-1260a、miR-6877-5p、miR-6721-5p、miR-4656、miR-1229-5p、miR-4433a-3p、miR-4274、miR-4419b、miR-4674、miR-6893-5p、miR-6763-3p、miR-6762-5p、miR-6738-5p、miR-4513、miR-6746-5p、miR-6880-5p、miR-4736、miR-718、miR-6717-5p、miR-7847-3p、miR-760、miR-1199-5p、miR-6813-5p、miR-6769a-5p、miR-1193、miR-7108-3p、miR-6741-5p、miR-4298、miR-6796-3p、miR-4750-5p、miR-6785-5p、miR-1292-3p、miR-4749-3p、miR-6800-3p、miR-4722-5p、miR-4746-3p、miR-4450、miR-6795-5p、miR-365a-5p、miR-498、miR-6797-5p、miR-1470、miR-6851-5p、miR-1247-3p、miR-5196-5p、miR-208a-5p、miR-6842-5p、miR-150-3p、miR-4534、miR-3135b、miR-3131、miR-4792、miR-6510-5p、miR-504-3p、miR-3619-3p、miR-671-5p、miR-4667-5p、miR-4430、miR-3195、miR-3679-5p、miR-6076、miR-6515-5p、miR-6820-5p、miR-4634、miR-187-5p、miR-6763-5p、miR-1908-3p、miR-1181、miR-6782-5p、miR-5010-5p、miR-6870-5p、miR-6124、miR-1249-5p、miR-6511b-5p、miR-1254、miR-4727-3p、miR-4259、miR-4771、miR-3622a-5p、miR-4480、miR-4740-5p、miR-6777-5p、miR-6794-5p、miR-4687-3p、miR-6743-5p、miR-6771-5p、miR-3141、miR-3162-5p、miR-4271、miR-1227-5p、miR-4257、miR-4270、miR-4516、miR-4651、miR-4725-3p、miR-6125、miR-6732-5p、miR-6791-5p、miR-6819-5p、miR-6891-5p、miR-7108-5p、miR-7109-5p、miR-642b-3p、miR-642a-3pから選択される少なくとも1つ又は少なくとも2つのポリヌクレオチド、又は該ポリヌクレオチドの相補鎖のそれぞれと特異的に結合可能な核酸を含む、請求項1~4のいずれか1項に記載のキット又は請求項5~10のいずれか1項に記載のデバイスを用いて、被験体の検体における標的核酸の発現量を測定し、該測定された発現量と、同様に測定された健常者の対照発現量とを用いて被験体が卵巣腫瘍に罹患していること、又は卵巣腫瘍に罹患していないことをin vitroで評価することを含む、卵巣腫瘍の検出方法。
- 卵巣腫瘍マーカーである、miR-4675、miR-4783-3p、miR-1228-5p、miR-4532、miR-6802-5p、miR-6784-5p、miR-3940-5p、miR-1307-3p、miR-8073、miR-3184-5p、miR-1233-5p、miR-6088、miR-5195-3p、miR-320b、miR-4649-5p、miR-6800-5p、miR-1343-3p、miR-4730、miR-6885-5p、miR-5100、miR-1203、miR-6756-5p、miR-373-5p、miR-1268a、miR-1260b、miR-4258、miR-4697-5p、miR-1469、miR-4515、miR-6861-5p、miR-6821-5p、miR-575、miR-6805-5p、miR-4758-5p、miR-3663-3p、miR-4530、miR-6798-5p、miR-6781-5p、miR-885-3p、miR-1273g-3p、miR-4787-3p、miR-4454、miR-4706、miR-1249-3p、miR-887-3p、miR-6786-5p、miR-1238-5p、miR-6749-5p、miR-6729-5p、miR-6825-5p、miR-663b、miR-6858-5p、miR-4690-5p、miR-6765-5p、miR-4710、miR-6775-5p、miR-371a-5p、miR-6816-5p、miR-296-3p、miR-7977、miR-8069、miR-6515-3p、miR-4687-5p、miR-1343-5p、miR-7110-5p、miR-4525、miR-3158-5p、miR-6787-5p、miR-614、miR-4689、miR-1185-2-3p、miR-1268b、miR-1228-3p、miR-1185-1-3p、miR-940、miR-939-5p、miR-6757-5p、miR-1275、miR-5001-5p、miR-6826-5p、miR-6765-3p、miR-3679-3p、miR-4718、miR-4286、miR-8059、miR-4447、miR-4448、miR-658、miR-6766-3p、miR-197-5p、miR-6887-5p、miR-6742-5p、miR-6729-3p、miR-5090、miR-7975、miR-4505、miR-6889-5p、miR-4708-3p、miR-6131、miR-1225-3p、miR-6132、miR-4734、miR-3194-3p、miR-638、miR-2467-3p、miR-4728-5p、miR-5572、miR-6789-5p、miR-8063、miR-4429、miR-6840-3p、miR-4476、miR-675-5p、miR-711、miR-6875-5p、miR-3160-5p、miR-1908-5p、miR-6726-5p、miR-1913、miR-8071、miR-3648、miR-4732-5p、miR-4787-5p、miR-3917、miR-619-5p、miR-1231、miR-342-5p、miR-4433a-5p、miR-6766-5p、miR-4707-5p、miR-7114-5p、miR-6872-3p、miR-6780b-5p、miR-7845-5p、miR-6798-3p、miR-665、miR-6848-5p、miR-5008-5p、miR-4294、miR-6511a-5p、miR-4435、miR-4747-3p、miR-6880-3p、miR-6869-5p、miR-7150、miR-1260a、miR-6877-5p、miR-6721-5p、miR-4656、miR-1229-5p、miR-4433a-3p、miR-4274、miR-4419b、miR-4674、miR-6893-5p、miR-6763-3p、miR-6762-5p、miR-6738-5p、miR-4513、miR-6746-5p、miR-6880-5p、miR-4736、miR-718、miR-6717-5p、miR-7847-3p、miR-760、miR-1199-5p、miR-6813-5p、miR-6769a-5p、miR-1193、miR-7108-3p、miR-6741-5p、miR-4298、miR-6796-3p、miR-4750-5p、miR-6785-5p、miR-1292-3p、miR-4749-3p、miR-6800-3p、miR-4722-5p、miR-4746-3p、miR-4450、miR-6795-5p、miR-365a-5p、miR-498、miR-6797-5p、miR-1470、miR-6851-5p、miR-1247-3p、miR-5196-5p、miR-208a-5p、miR-6842-5p、miR-150-3p、miR-4534、miR-3135b、miR-3131、miR-4792、miR-6510-5p、miR-504-3p、miR-3619-3p、miR-671-5p、miR-4667-5p、miR-4430、miR-3195、miR-3679-5p、miR-6076、miR-6515-5p、miR-6820-5p、miR-4634、miR-187-5p、miR-6763-5p、miR-1908-3p、miR-1181、miR-6782-5p、miR-5010-5p、miR-6870-5p、miR-6124、miR-1249-5p、miR-6511b-5p、miR-1254、miR-4727-3p、miR-4259、miR-4771、miR-3622a-5p、miR-4480、miR-4740-5p、miR-6777-5p、miR-6794-5p、miR-4687-3p、miR-6743-5p、miR-6771-5p、miR-3141、miR-3162-5p、miR-4271、miR-1227-5p、miR-4257、miR-4270、miR-4516、miR-4651、miR-4725-3p、miR-6125、miR-6732-5p、miR-6791-5p、miR-6819-5p、miR-6891-5p、miR-7108-5p、miR-7109-5p、miR-642b-3p、miR-642a-3pから選択される少なくとも1つ又は少なくとも2つのポリヌクレオチド、又は当該ポリヌクレオチドの相補鎖のそれぞれと特異的に結合可能な核酸を含む、請求項1~4のいずれか1項に記載のキット又は請求項5~10いずれか1項に記載のデバイスを用いて被験体の検体における標的遺伝子の発現量を測定し、卵巣腫瘍を有することが既知である被験体由来の検体の遺伝子発現量と卵巣腫瘍に罹患していない被験体由来の検体の遺伝子発現量を教師サンプルとして作成された、かつ卵巣腫瘍の存在または不存在を区別的に判別することが可能である判別式に、前記被験体由来の検体における標的遺伝子の発現量を代入し、それによって、卵巣腫瘍の存在又は不存在をin vitroで評価することを含む、卵巣腫瘍の検出方法。
- 前記核酸が、下記の(a)~(e)のいずれかに示すポリヌクレオチド:
(a)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~247、251及び268のいずれかで表される塩基配列を含むポリヌクレオチド、
(c)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~247、251及び268のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項11~13のいずれか1項に記載の方法。 - 前記キットまたはデバイスが、別の卵巣腫瘍マーカーである、miR-320a、miR-663a、miR-328-5p、miR-128-2-5p、miR-125a-3p、miR-191-5p、miR-92b-5p、miR-296-5p、miR-1246、miR-92a-2-5p、miR-128-1-5p、miR-1290、miR-211-3p、miR-744-5p、miR-135a-3p、miR-451a、miR-625-3p、miR-92a-3p、miR-422a、miR-483-5p、miR-652-5p、miR-24-3p、miR-23b-3p、miR-23a-3p、miR-92b-3p、miR-22-3pから選択される少なくとも1つのポリヌクレオチド、又は当該ポリヌクレオチドの相補鎖と特異的に結合可能な核酸をさらに含む、請求項11~14のいずれか1項に記載の方法。
- 前記核酸が、下記の(f)~(j)のいずれかに示すポリヌクレオチド:
(f)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列を含むポリヌクレオチド、
(h)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号248~250、252~267及び269~275のいずれかで表される塩基配列、もしくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項15に記載の方法。 - 前記被験体が、ヒトである、請求項11~16のいずれか1項に記載の方法。
- 前記検体が、血液、血清又は血漿である、請求項11~17のいずれか1項に記載の方法。
Priority Applications (10)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP18790377.8A EP3628736A4 (en) | 2017-04-28 | 2018-04-27 | KIT, DEVICE AND METHOD FOR DETECTION OF OVARIAL TUMOR |
| KR1020197029058A KR20200002809A (ko) | 2017-04-28 | 2018-04-27 | 난소 종양의 검출을 위한 키트, 디바이스, 및 방법 |
| CN201880027774.XA CN110546263B (zh) | 2017-04-28 | 2018-04-27 | 用于检测卵巢肿瘤的试剂盒、装置和方法 |
| US16/608,673 US10975444B2 (en) | 2017-04-28 | 2018-04-27 | Kit, device, and method for detecting ovarian tumor |
| BR112019021215-9A BR112019021215A2 (pt) | 2017-04-28 | 2018-04-27 | Kit para detecção de tumor ovariano, dispositivo para detecção de tumor ovariano e métodos para detectar tumor ovariano |
| CA3059480A CA3059480A1 (en) | 2017-04-28 | 2018-04-27 | Kit, device, and method for detecting ovarian tumor |
| JP2019514645A JP7165329B2 (ja) | 2017-04-28 | 2018-04-27 | 卵巣腫瘍の検出のためのキット、デバイス及び方法 |
| US17/219,000 US20210230706A1 (en) | 2017-04-28 | 2021-03-31 | Kit, device, and method for detecting ovarian tumor |
| JP2022164429A JP7440875B2 (ja) | 2017-04-28 | 2022-10-13 | 卵巣腫瘍の検出のためのキット、デバイス及び方法 |
| JP2024017115A JP2024042098A (ja) | 2017-04-28 | 2024-02-07 | 卵巣腫瘍の検出のためのキット、デバイス及び方法 |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2017-090799 | 2017-04-28 | ||
| JP2017090799 | 2017-04-28 |
Related Child Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US16/608,673 A-371-Of-International US10975444B2 (en) | 2017-04-28 | 2018-04-27 | Kit, device, and method for detecting ovarian tumor |
| US17/219,000 Division US20210230706A1 (en) | 2017-04-28 | 2021-03-31 | Kit, device, and method for detecting ovarian tumor |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2018199275A1 true WO2018199275A1 (ja) | 2018-11-01 |
Family
ID=63919756
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2018/017125 WO2018199275A1 (ja) | 2017-04-28 | 2018-04-27 | 卵巣腫瘍の検出のためのキット、デバイス及び方法 |
Country Status (8)
| Country | Link |
|---|---|
| US (2) | US10975444B2 (ja) |
| EP (1) | EP3628736A4 (ja) |
| JP (3) | JP7165329B2 (ja) |
| KR (1) | KR20200002809A (ja) |
| CN (1) | CN110546263B (ja) |
| BR (1) | BR112019021215A2 (ja) |
| CA (1) | CA3059480A1 (ja) |
| WO (1) | WO2018199275A1 (ja) |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2020032228A1 (ja) * | 2018-08-10 | 2020-02-13 | 東レ株式会社 | 前立腺がんの検出のためのキット、デバイス及び方法 |
| WO2021132547A1 (ja) * | 2019-12-25 | 2021-07-01 | 東レ株式会社 | 検査方法、検査装置、学習方法、学習装置、検査プログラムおよび学習プログラム |
| WO2021201092A1 (ja) * | 2020-03-31 | 2021-10-07 | 東レ株式会社 | 海馬萎縮を検出するためのキット又はデバイス及び方法 |
| WO2023068318A1 (ja) * | 2021-10-20 | 2023-04-27 | 東レ株式会社 | 卵巣がんと卵巣良性腫瘍とを判別するためのキット、デバイス及び方法 |
Families Citing this family (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP7306633B2 (ja) * | 2017-06-29 | 2023-07-11 | 東レ株式会社 | 肺がんの検出のためのキット、デバイス及び方法 |
| CN110499367B (zh) * | 2019-08-09 | 2022-11-22 | 深圳市第二人民医院 | 生物标志物及其应用 |
| CN111778339B (zh) * | 2020-08-16 | 2021-01-15 | 徐州医科大学 | 与食管癌相关的miRNA标志物及其应用 |
| CN112255334B (zh) * | 2020-09-28 | 2022-06-17 | 复旦大学 | 用于区分交界性和恶性卵巢肿瘤的小分子标志物及其应用 |
| US20240002949A1 (en) * | 2020-11-12 | 2024-01-04 | Uniwersytet Medyczny W Bialymstoku | Panel of mirna biomarkers for diagnosis of ovarian cancer, method for in vitro diagnosis of ovarian cancer, uses of panel of mirna biomarkers for in vitro diagnosis of ovarian cancer and test for in vitro diagnosis of ovarian cancer |
| CN112641797B (zh) * | 2020-12-30 | 2021-12-17 | 温州医科大学 | 抑制结直肠癌生长、转移的靶标与诊断标志物及其应用 |
| CA3221494A1 (en) * | 2021-06-09 | 2022-12-15 | Andrew Zhang | Cancer detection method, kit, and system |
| CN115804843A (zh) * | 2021-09-15 | 2023-03-17 | 中国药科大学 | Skor1基因作为靶点在2型糖尿病和/或炎症治疗中的应用 |
| WO2023171012A1 (en) * | 2022-03-09 | 2023-09-14 | Craif Inc. | Methods for categorizing disease outcomes |
| WO2023219447A1 (ko) * | 2022-05-12 | 2023-11-16 | 서울대학교병원 | 세포외 소포체 유래 mirna를 검출하는 제제를 포함하는 난소암 진단용 조성물 |
Citations (9)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2010154843A (ja) | 2008-06-27 | 2010-07-15 | Keio Gijuku | バイオマーカーとしてのマイクロrnaを用いた婦人科がんの診断・治療選択 |
| JP2010538610A (ja) | 2007-09-06 | 2010-12-16 | ジ・オハイオ・ステイト・ユニバーシティ・リサーチ・ファウンデイション | ヒト卵巣癌中のマイクロrnaシグネチャー |
| US20120309645A1 (en) * | 2010-02-05 | 2012-12-06 | Febit Holding Gmbh | miRNA IN THE DIAGNOSIS OF OVARIAN CANCER |
| WO2013148151A1 (en) | 2012-03-29 | 2013-10-03 | University Of Pittsburgh - Of The Commonwealth System Of Higher Education | Plasma microribonucleic acids as biomarkers for endometriosis and endometriosis-associated ovarian cancer |
| WO2015190586A1 (ja) * | 2014-06-13 | 2015-12-17 | 東レ株式会社 | 大腸がんの検出キット又はデバイス及び検出方法 |
| WO2015194535A1 (ja) * | 2014-06-16 | 2015-12-23 | 東レ株式会社 | 胃がんの検出キット又はデバイス及び検出方法 |
| WO2015194615A1 (ja) * | 2014-06-18 | 2015-12-23 | 東レ株式会社 | 肝臓がんの検出キット又はデバイス及び検出方法 |
| US20160312301A1 (en) | 2013-12-20 | 2016-10-27 | The Feinstein Institute For Medical Research | Microrna biomarkers for ovarian cancer |
| JP2017090799A (ja) | 2015-11-16 | 2017-05-25 | 株式会社ジャパンディスプレイ | 表示装置 |
Family Cites Families (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| IL179285A (en) * | 2004-05-14 | 2011-04-28 | Rosetta Genomics Ltd | Micrornas and uses thereof |
| JP2013148151A (ja) | 2012-01-19 | 2013-08-01 | Seiko Epson Corp | 減速機、減速機一体モーター、ロボットアーム、及びロボット |
| PL2922554T3 (pl) * | 2012-11-26 | 2022-06-20 | Modernatx, Inc. | Na zmodyfikowany na końcach |
| WO2014210341A2 (en) * | 2013-06-27 | 2014-12-31 | Institute For Systems Biology | Products and methods relating to micro rnas and cancer |
| JP2019508063A (ja) * | 2016-01-27 | 2019-03-28 | オンコラス, インコーポレイテッド | 腫瘍溶解性ウイルスベクター及びその使用 |
-
2018
- 2018-04-27 KR KR1020197029058A patent/KR20200002809A/ko not_active Application Discontinuation
- 2018-04-27 WO PCT/JP2018/017125 patent/WO2018199275A1/ja unknown
- 2018-04-27 CA CA3059480A patent/CA3059480A1/en active Pending
- 2018-04-27 JP JP2019514645A patent/JP7165329B2/ja active Active
- 2018-04-27 BR BR112019021215-9A patent/BR112019021215A2/pt not_active Application Discontinuation
- 2018-04-27 EP EP18790377.8A patent/EP3628736A4/en active Pending
- 2018-04-27 CN CN201880027774.XA patent/CN110546263B/zh active Active
- 2018-04-27 US US16/608,673 patent/US10975444B2/en active Active
-
2021
- 2021-03-31 US US17/219,000 patent/US20210230706A1/en active Pending
-
2022
- 2022-10-13 JP JP2022164429A patent/JP7440875B2/ja active Active
-
2024
- 2024-02-07 JP JP2024017115A patent/JP2024042098A/ja active Pending
Patent Citations (9)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2010538610A (ja) | 2007-09-06 | 2010-12-16 | ジ・オハイオ・ステイト・ユニバーシティ・リサーチ・ファウンデイション | ヒト卵巣癌中のマイクロrnaシグネチャー |
| JP2010154843A (ja) | 2008-06-27 | 2010-07-15 | Keio Gijuku | バイオマーカーとしてのマイクロrnaを用いた婦人科がんの診断・治療選択 |
| US20120309645A1 (en) * | 2010-02-05 | 2012-12-06 | Febit Holding Gmbh | miRNA IN THE DIAGNOSIS OF OVARIAN CANCER |
| WO2013148151A1 (en) | 2012-03-29 | 2013-10-03 | University Of Pittsburgh - Of The Commonwealth System Of Higher Education | Plasma microribonucleic acids as biomarkers for endometriosis and endometriosis-associated ovarian cancer |
| US20160312301A1 (en) | 2013-12-20 | 2016-10-27 | The Feinstein Institute For Medical Research | Microrna biomarkers for ovarian cancer |
| WO2015190586A1 (ja) * | 2014-06-13 | 2015-12-17 | 東レ株式会社 | 大腸がんの検出キット又はデバイス及び検出方法 |
| WO2015194535A1 (ja) * | 2014-06-16 | 2015-12-23 | 東レ株式会社 | 胃がんの検出キット又はデバイス及び検出方法 |
| WO2015194615A1 (ja) * | 2014-06-18 | 2015-12-23 | 東レ株式会社 | 肝臓がんの検出キット又はデバイス及び検出方法 |
| JP2017090799A (ja) | 2015-11-16 | 2017-05-25 | 株式会社ジャパンディスプレイ | 表示装置 |
Non-Patent Citations (70)
| Title |
|---|
| ALTSCHUL, S.F. ET AL., JOURNAL OF MOLECULAR BIOLOGY, vol. 215, 1990, pages 403 - 410 |
| ALTUVIA Y ET AL., NUCLEIC ACIDS RES, vol. 33, 2005, pages 2697 - 2706 |
| ARTZI S ET AL., BMC BIOINFORMATICS, vol. 9, 2008, pages 39 |
| AUSUBEL ET AL.: "Current Protocols in Molecular Biology", 1993, JOHN WILLEY & SONS |
| BAR M ET AL., STEM CELLS, vol. 26, 2008, pages 2496 - 2505 |
| BENTWICH I ET AL., NAT GENET, vol. 37, 2005, pages 766 - 770 |
| BEREZIKOV E ET AL., GENOME RES, vol. 16, 2006, pages 1289 - 1298 |
| BEREZIKOV E ET AL., MOL CELL, vol. 28, 2007, pages 328 - 336 |
| BOLSTAD, B. M. ET AL., BIOINFORMATICS, vol. 19, 2003, pages 185 - 193 |
| C. CORTES ET AL., MACHINE LEARNING, vol. 20, 1995, pages 273 - 297 |
| CAI X ET AL., RNA, vol. 13, 2007, pages 313 - 316 |
| CREIGHTON CJ ET AL., PLOS ONE, vol. 5, 2010, pages e10563 |
| CUMMINS JM ET AL., PROC NATL ACAD SCI USA, vol. 103, 2006, pages 3687 - 3692 |
| DANNEMANN M ET AL., GENOME BIOL EVOL, vol. 4, 2012, pages 552 - 564 |
| DING N ET AL., J RADIAT RES, vol. 52, 2011, pages 425 - 432 |
| FU H ET AL., FEBS LETT, vol. 579, 2005, pages 3849 - 3854 |
| FUREY TS. ET AL., BIOINFORMATICS, vol. 16, 2000, pages 906 - 14 |
| GIUSTI I. ET AL., BIOMED RESEARCH INTERNATIONAL, vol. 2013, 2013, pages 703048 |
| GOFF LA ET AL., PLOS ONE, vol. 4, 2009, pages e7192 |
| HANSEN TB ET AL., RNA BIOL, vol. 8, 2011, pages 378 - 383 |
| HIDEKI ASO ET AL.: "Frontier of Statistical Science 6", 2004, IWANAMI SHOTEN, PUBLISHERS, article "Statistics of pattern recognition and learning - New concepts and approaches" |
| HOUBAVIY HB ET AL., DEV CELL, vol. 5, 2003, pages 351 - 358 |
| INT. NEUROUROL. J., vol. 20, no. 2, 2016, pages 76 - 83 |
| JI T. ET AL., ASIAN PACIFIC JOURNAL OF CANCER PREVENTION, vol. 15, no. 4, 2014, pages 1739 |
| JIMA DD ET AL., BLOOD, vol. 116, 2010, pages e118 - e127 |
| JOYCE CE ET AL., HUM MOL GENET, vol. 20, 2011, pages 4025 - 4040 |
| KASASHIMA K ET AL., BIOCHEM BIOPHYS RES COMMUN, vol. 322, 2004, pages 403 - 410 |
| KAWAJI H ET AL., BMC GENOMICS, vol. 9, 2008, pages 157 |
| KIM J ET AL., PROC NATL ACAD SCI USA, vol. 101, 2004, pages 360 - 365 |
| KIM YW. ET AL., PLOS ONE, vol. 7, no. 9, 2012, pages e50746 |
| LADEWIG E ET AL., GENOME RES, vol. 22, 2012, pages 1634 - 1645 |
| LAGOS-QUINTANA M ET AL., CURR BIOL, vol. 12, 2002, pages 735 - 739 |
| LAGOS-QUINTANA M ET AL., RNA, vol. 9, 2003, pages 175 - 179 |
| LAGOS-QUINTANA M ET AL., SCIENCE, vol. 294, 2001, pages 853 - 858 |
| LI Y ET AL., GENE, vol. 497, 2012, pages 330 - 335 |
| LIM LP ET AL., SCIENCE, vol. 299, 2003, pages 1540 |
| MEIRI E ET AL., NUCLEIC ACIDS RES, vol. 38, 2010, pages 6234 - 6246 |
| MICHAEL MZ ET AL., MOL CANCER RES, vol. 1, 2003, pages 882 - 891 |
| MORIN RD ET AL., GENOME RES, vol. 18, 2008, pages 610 - 621 |
| MORIN RD. ET AL., GENOME RES., vol. 18, 2008, pages 610 - 621 |
| MOURELATOS Z ET AL., GENES DEV, vol. 16, 2002, pages 720 - 728 |
| NAKAMURA K. ET AL., MOLECULAR CANCER, vol. 15, no. 1, 2016, pages 48 |
| NELLO CRISTIANINI ET AL.: "Introduction to SVM", 2008, KYORITSU SHUPPAN CO., LTD. |
| NIELSEN, P.E. ET AL., SCIENCE, vol. 254, 1991, pages 1497 - 500 |
| OBIKA, S. ET AL., TETRAHEDRON LETT., vol. 39, 1998, pages 5401 - 5404 |
| OULAS A ET AL., NUCLEIC ACIDS RES, vol. 37, 2009, pages 3276 - 3287 |
| PEARSON, W.R. ET AL., PROC. NATL. ACAD. SCI. U. S. A., vol. 85, 1988, pages 2444 - 2448 |
| PERSSON H ET AL., CANCER RES, vol. 67, 2007, pages 6031 - 6043 |
| PERSSON H ET AL., CANCER RES, vol. 71, 2011, pages 78 - 86 |
| RESHMI G ET AL., GENOMICS, vol. 97, 2011, pages 333 - 340 |
| RICK KAMPS ET AL., INT. J. MOL. SCI., vol. 18, no. 2, 2017, pages 308 |
| SALVI A ET AL., INT J ONCOL, vol. 42, 2013, pages 391 - 402 |
| SAMBROOK ET AL.: "Molecular Cloning - A Laboratory Manual", 1989, COLD SPRING HARBOR LABORATORY PRESS |
| SAMBROOK, J.RUSSEL, D.: "Molecular Cloning, A LABORATORY MANUAL", vol. 1, 15 January 2001, COLD SPRING HARBOR LABORATORY PRESS |
| SCHOTTE D ET AL., LEUKEMIA, vol. 25, 2011, pages 1389 - 1399 |
| SMITH JL ET AL., J VIROL, vol. 86, 2012, pages 5278 - 5287 |
| SUBRAMANIAN S ET AL., ONCOGENE, vol. 27, 2008, pages 2015 - 2026 |
| SUH MR ET AL., DEV BIOL, vol. 270, 2004, pages 488 - 498 |
| TAKADA S ET AL., LEUKEMIA, vol. 22, 2008, pages 1274 - 1278 |
| TAKAFUMI KANAMORI ET AL.: "Pattern Recognition", 2009, KYORITSU SHUPPAN CO., LTD. |
| TANDON M ET AL., ORAL DIS, vol. 18, 2012, pages 127 - 131 |
| VELTHUT-MEIKAS A ET AL., MOL ENDOCRINOL, vol. 27, 2013, pages 1128 - 1141 |
| VENABLES, W. N.RIPLEY, B. D.: "Modern Applied Statistics with S", 2002, SPRINGER |
| VOELLENKLE C ET AL., RNA, vol. 18, 2012, pages 472 - 484 |
| WANG HJ ET AL., SHOCK, vol. 39, 2013, pages 480 - 487 |
| WITTEN D ET AL., BMC BIOL, vol. 8, 2010, pages 58 |
| XIE X ET AL., NATURE, vol. 434, 2005, pages 338 - 345 |
| YASUSHI NAGATA ET AL.: "Basics of statistical multiple comparison methods", 2007, SCIENTIST PRESS CO., LTD. |
| YOO JK ET AL., STEM CELLS DEV, vol. 21, 2012, pages 2049 - 2057 |
| ZHENG ZHANG ET AL., J. COMPUT. BIOL., vol. 7, 2000, pages 203 - 214 |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2020032228A1 (ja) * | 2018-08-10 | 2020-02-13 | 東レ株式会社 | 前立腺がんの検出のためのキット、デバイス及び方法 |
| WO2021132547A1 (ja) * | 2019-12-25 | 2021-07-01 | 東レ株式会社 | 検査方法、検査装置、学習方法、学習装置、検査プログラムおよび学習プログラム |
| WO2021201092A1 (ja) * | 2020-03-31 | 2021-10-07 | 東レ株式会社 | 海馬萎縮を検出するためのキット又はデバイス及び方法 |
| WO2023068318A1 (ja) * | 2021-10-20 | 2023-04-27 | 東レ株式会社 | 卵巣がんと卵巣良性腫瘍とを判別するためのキット、デバイス及び方法 |
Also Published As
| Publication number | Publication date |
|---|---|
| US20200140956A1 (en) | 2020-05-07 |
| BR112019021215A2 (pt) | 2020-06-02 |
| EP3628736A4 (en) | 2021-06-09 |
| JP7165329B2 (ja) | 2022-11-04 |
| US20210230706A1 (en) | 2021-07-29 |
| EP3628736A1 (en) | 2020-04-01 |
| US10975444B2 (en) | 2021-04-13 |
| JPWO2018199275A1 (ja) | 2020-03-26 |
| CN110546263B (zh) | 2024-03-05 |
| JP7440875B2 (ja) | 2024-02-29 |
| KR20200002809A (ko) | 2020-01-08 |
| CA3059480A1 (en) | 2018-11-01 |
| CN110546263A (zh) | 2019-12-06 |
| JP2022188256A (ja) | 2022-12-20 |
| JP2024042098A (ja) | 2024-03-27 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP7440875B2 (ja) | 卵巣腫瘍の検出のためのキット、デバイス及び方法 | |
| JP7397449B2 (ja) | 大腸がんの検出キット又はデバイス及び検出方法 | |
| JP7383269B2 (ja) | 早期膵がん又は膵がん前駆病変の検出キット又はデバイス及び検出方法 | |
| US20230193402A1 (en) | Kit, device, and method for detecting lung cancer | |
| JP6925125B2 (ja) | 胃がんの検出キット又はデバイス及び検出方法 | |
| WO2015182781A1 (ja) | 膵臓がんの検出キット又はデバイス及び検出方法 | |
| WO2015194627A1 (ja) | 食道がんの検出キット又はデバイス及び検出方法 | |
| WO2015194610A1 (ja) | 肺がんの検出キット又はデバイス及び検出方法 | |
| WO2015190584A1 (ja) | 前立腺がんの検出キット又はデバイス及び検出方法 | |
| US20240060142A1 (en) | Kit, device, and method for detecting bladder cancer |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 18790377 Country of ref document: EP Kind code of ref document: A1 |
|
| ENP | Entry into the national phase |
Ref document number: 2019514645 Country of ref document: JP Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 20197029058 Country of ref document: KR Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 3059480 Country of ref document: CA |
|
| REG | Reference to national code |
Ref country code: BR Ref legal event code: B01A Ref document number: 112019021215 Country of ref document: BR |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2018790377 Country of ref document: EP Effective date: 20191128 |
|
| ENP | Entry into the national phase |
Ref document number: 2018790377 Country of ref document: EP Effective date: 20191128 |
|
| ENP | Entry into the national phase |
Ref document number: 112019021215 Country of ref document: BR Kind code of ref document: A2 Effective date: 20191009 |