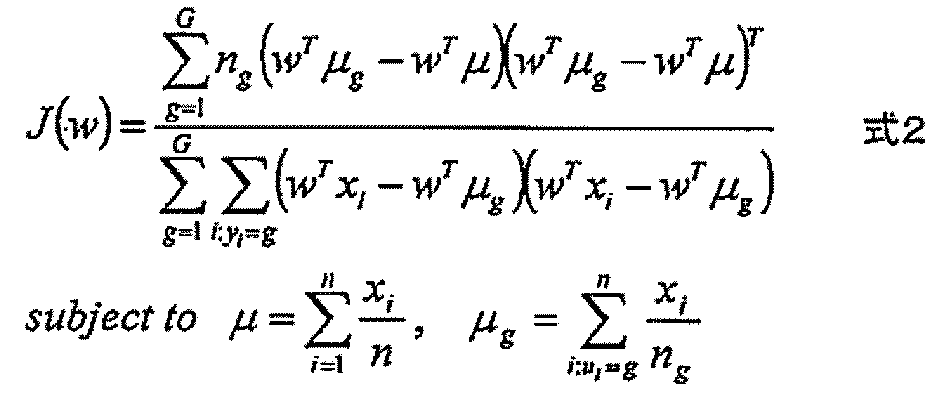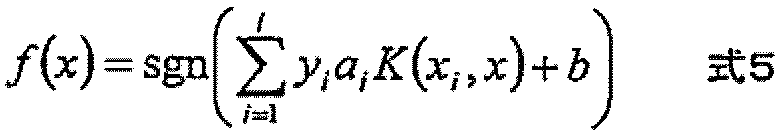WO2021201092A1 - 海馬萎縮を検出するためのキット又はデバイス及び方法 - Google Patents
海馬萎縮を検出するためのキット又はデバイス及び方法 Download PDFInfo
- Publication number
- WO2021201092A1 WO2021201092A1 PCT/JP2021/013807 JP2021013807W WO2021201092A1 WO 2021201092 A1 WO2021201092 A1 WO 2021201092A1 JP 2021013807 W JP2021013807 W JP 2021013807W WO 2021201092 A1 WO2021201092 A1 WO 2021201092A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- mir
- hsa
- gene
- seq
- polynucleotide
- Prior art date
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6813—Hybridisation assays
- C12Q1/6834—Enzymatic or biochemical coupling of nucleic acids to a solid phase
- C12Q1/6837—Enzymatic or biochemical coupling of nucleic acids to a solid phase using probe arrays or probe chips
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/158—Expression markers
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/178—Oligonucleotides characterized by their use miRNA, siRNA or ncRNA
Definitions
- the present invention relates to a kit or device for detecting hippocampal atrophy containing a specific miRNA or a nucleic acid that can specifically bind to a complementary strand thereof, which can be used for examining hippocampal atrophy of a subject's brain, and the miRNA.
- the present invention relates to a method for detecting hippocampal atrophy, which comprises measuring the expression level.
- the hippocampus is a part of the region called the hippocampal formation that exists in the limbic system, and is an organ of the brain located deep in the temporal lobe of the cerebral cortex. Hippocampal physiology is associated with intellectual function, memory, and spatial grasping ability. The hippocampus has the role of storing memory until new memory (approximate memory) is established as long-term memory. Therefore, when the hippocampus is atrophied or damaged, memory formation is impaired and cognitive memory impairment occurs.
- Hippocampal atrophy is caused by the loss of nerve cells that make up the hippocampus. It is known that the brain including the hippocampus is atrophied as a whole due to aging, but Alzheimer-type dementia, frontotemporal lobar degeneration, and depression are diseases in which the hippocampus is particularly significantly atrophied due to causes other than aging. Diseases, post-traumatic stress disorders, schizophrenia, etc.
- the test result of the atrophy degree in the hippocampal region is treated as one piece of information (N: Neurodegeneration) for the diagnosis of Alzheimer's disease.
- N Neurodegeneration
- a general method for examining the hippocampus is diagnostic imaging by brain MRI or the like.
- FDG-PET which is an diagnostic imaging method for measuring glucose metabolism in the brain, may be used as a method for measuring nerve cell death, which is the cause of hippocampal atrophy (Non-Patent Document 1).
- Non-Patent Document 2 a method of minimally invasively measuring amyloid ⁇ , which is a cause of the onset of Alzheimer's disease, from a small amount of plasma by immunoprecipitation and mass spectrometry has been reported (Non-Patent Document 2). Also, as a blood test, by measuring the miRNA expression level of 451 species (hsa-miR-1228-5p, hsa-miR-6088, hsa-miR-4525, hsa-miR-4471, etc.) in the serum sample. It has been reported that patients with Alzheimer's disease can be detected with a sensitivity of 81% and a specificity of 79% (Patent Document 1).
- Alzheimer's disease is performed by measuring the expression level of miRNA such as hsa-miR-486-3p contained in blood components. There is a report that it is possible to detect a patient (Patent Document 2).
- Patent Document 3 there is a report that it is possible to detect a patient with Alzheimer's disease by measuring hsa-miR-6805-5p or the like in a blood sample.
- Patent Document 4 There is a report that it is possible to detect patients with Alzheimer's disease and mild dementia by measuring hsa-miR-320b or the like in a blood sample.
- Non-Patent Document 3 it has been reported that the expression level of miRNA such as hsa-miR-663a contained in serum is significantly reduced in patients with Alzheimer's disease as compared with healthy subjects.
- Patent Document 5 There is a report that it can be detected by measuring hsa-miR-760 and hsa-miR-887-3p contained in cerebrospinal fluid.
- Patent Document 6 There is a report that it is possible to detect a dementia patient by measuring hsa-miR-4463 or the like in a sample.
- neurodegenerative diseases such as Alzheimer-type dementia, hsa-miR-128-1-5p, hsa-miR-128-2-5p, hsa-contained in microvesicles in body fluids. It has been reported that patients with Alzheimer's disease can be detected by measuring the expression levels of miRNAs such as miR-187-5p, hsa-miR-711, and hsa-miR-328-5p (Patent Document 7). ..
- Non-Patent Document 4 it has been reported that the expression levels of miRNAs such as hsa-miR-4674 and hsa-miR-6722-3p contained in brain tissue significantly fluctuate in patients with Alzheimer's disease.
- Non-Patent Document 5 Korean Patent Document 5
- the generalized hippocampal atrophy index value for the quantitative value of the hippocampal volume is No.
- some facilities do not have an environment for brain MRI imaging, and in addition, it is difficult for people with magnetic metal in their bodies or people with claustrophobia to undergo MRI examinations. Therefore, it is not universally available to everyone and has the disadvantage of lacking in convenience.
- Non-Patent Document 1 introduces an FDG-PET examination as a brain imaging examination, but the facilities where the FDG-PET examination can be performed are limited in terms of equipment, and although safety is guaranteed, a radiological substance is used. There is a danger.
- the brain imaging method for detecting hippocampal atrophy and synaptic degeneration mainly has problems in convenience, safety, and quantitativeness for patients.
- Non-Patent Document 2 Since the inspection method described in Non-Patent Document 2 is limited to measurement by a mass spectrometer, advanced measurement technique is required. In addition, it is unclear how the mechanism of action of APP669-771 (amyloid ⁇ protein) to be measured fluctuates, and no direct relationship with hippocampal atrophy can be found.
- APP669-771 amyloid ⁇ protein
- Patent Document 1 since patients with Alzheimer's disease classified by a doctor's diagnosis based on the clinical findings of the patients are positive, stratified tests cannot be performed, and a direct relationship with hippocampal atrophy is found from them. It is not possible.
- Non-Patent Document 4 In the method described in Non-Patent Document 4, it is necessary to use the brain tissue as a sample, and it cannot be used as a simple test.
- Non-Patent Document 5 does not directly show the relationship between the nucleic acid and hippocampal atrophy, and it cannot be said that the disclosed miRNA can be used as a marker for hippocampal atrophy.
- the method of directly showing hippocampal atrophy is brain imaging, but at this stage, its use is limited due to the burden on patients and equipment, and its convenience and versatility are low.
- a blood test or the like is desirable as a simple biochemical diagnosis with low invasiveness, but there is no method capable of detecting hippocampal atrophy in a minimally invasive manner in the current marker test. It is desired to develop a diagnostic marker that can correctly determine whether or not the hippocampus is atrophied using a blood sample.
- An object of the present invention is to provide a hippocampal atrophy marker that enables detection of hippocampal atrophy without imaging MRI, and a means and method that can objectively and effectively detect hippocampal atrophy using the marker. be.
- the present inventors have found a hippocampal atrophy marker capable of detecting the degree of atrophy in the hippocampal region, which is closely related to dementia, and have completed the present invention based on the marker.
- the present invention includes the following aspects. (1) Kaiba atrophy markers, miR-3131, miR-6757-5p, miR-4706, miR-5001-5p, miR-3180-3p, miR-642b-3p, miR-4655-5p, miR-6819 -5p, miR-937-5p, miR-4688, miR-6471-5p, miR-7107-5p, miR-4271, miR-1229-5p, miR-4707-5p, miR-6808-5p, miR-4656 , MiR-6076, miR-6762-5p, miR-7109-5p, miR-6732-5p, miR-3195, miR-7150, miR-642a-3p, miR-1249-5p, miR-3185, miR-4689 , MiR-3141, miR-6840-3p, miR-3135b, miR-1914-3p, miR-4446-3p, miR-4433b-3p, miR-6877-5p, miR
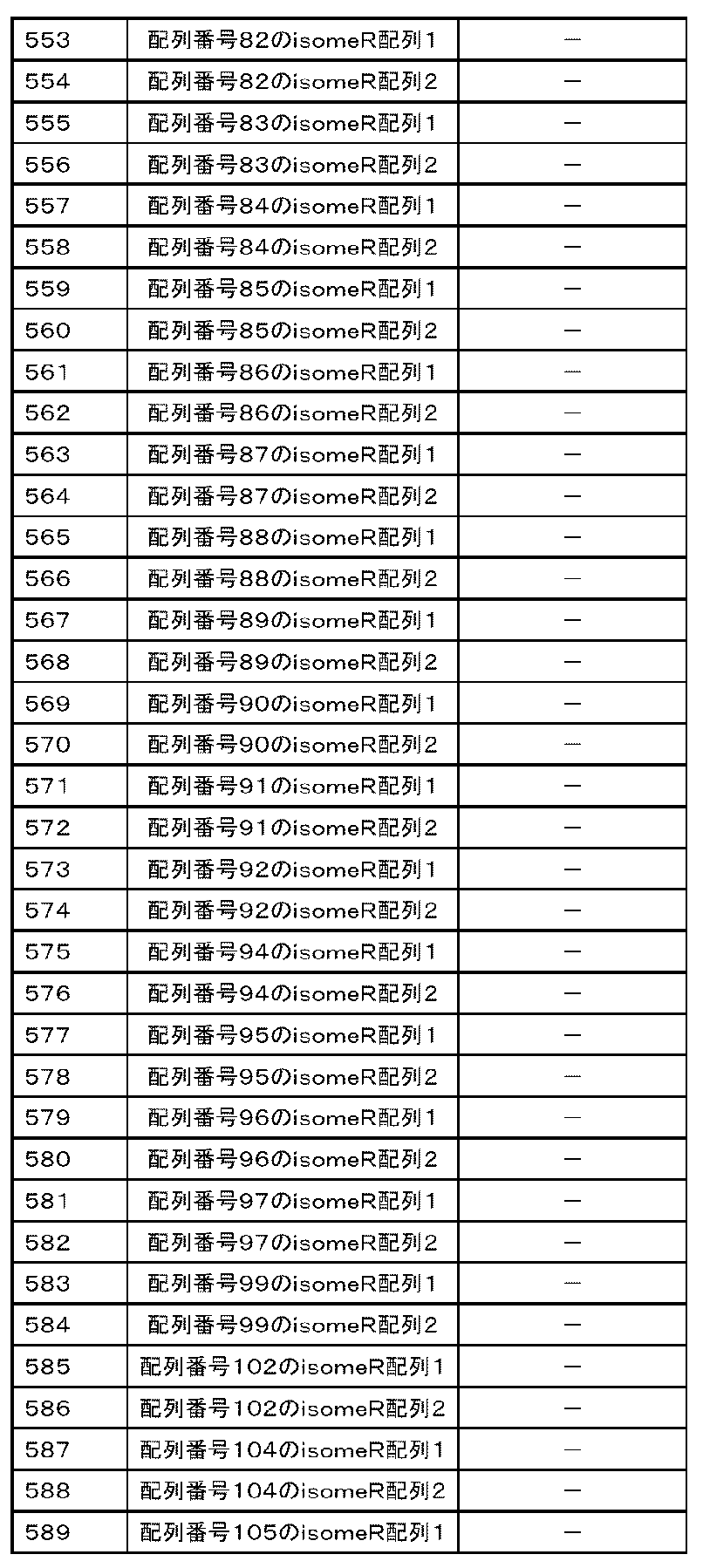
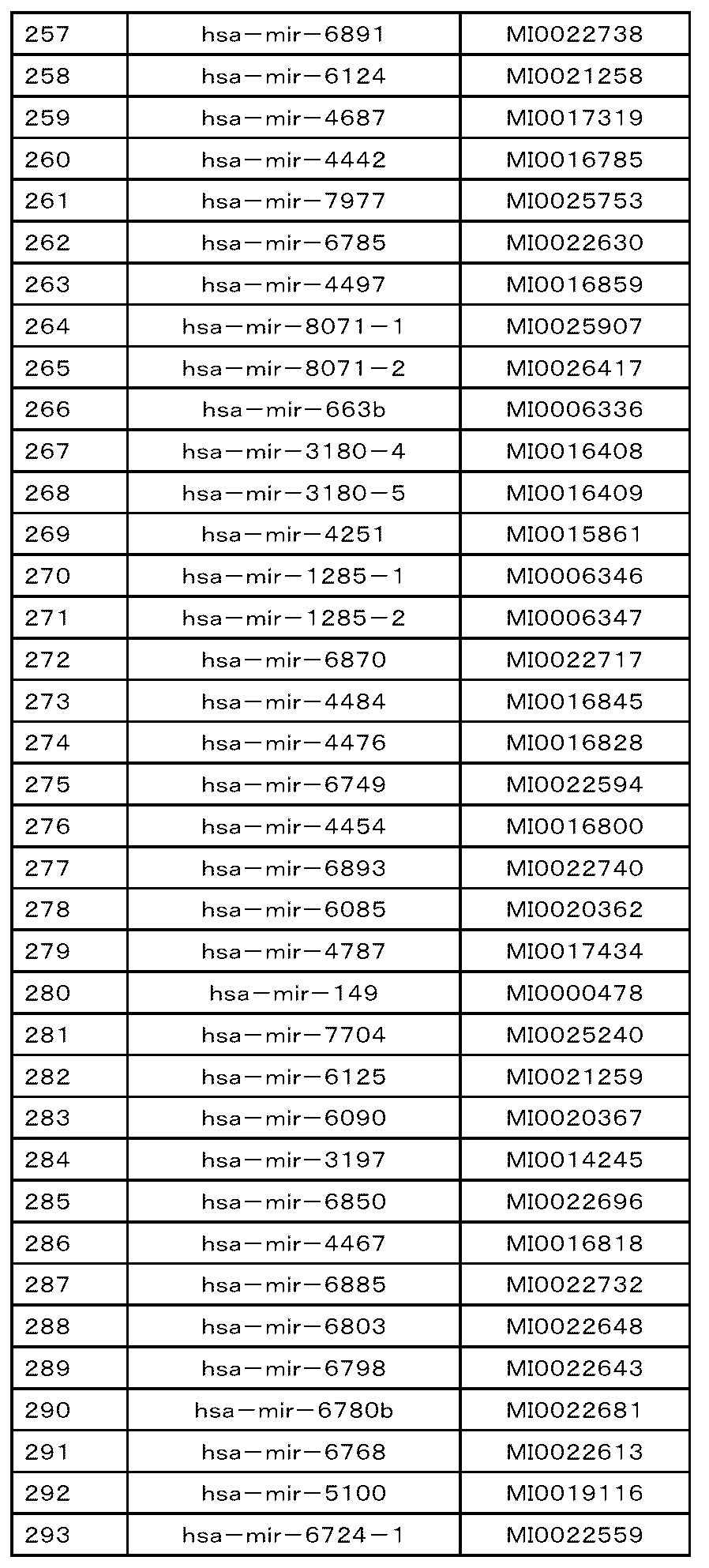
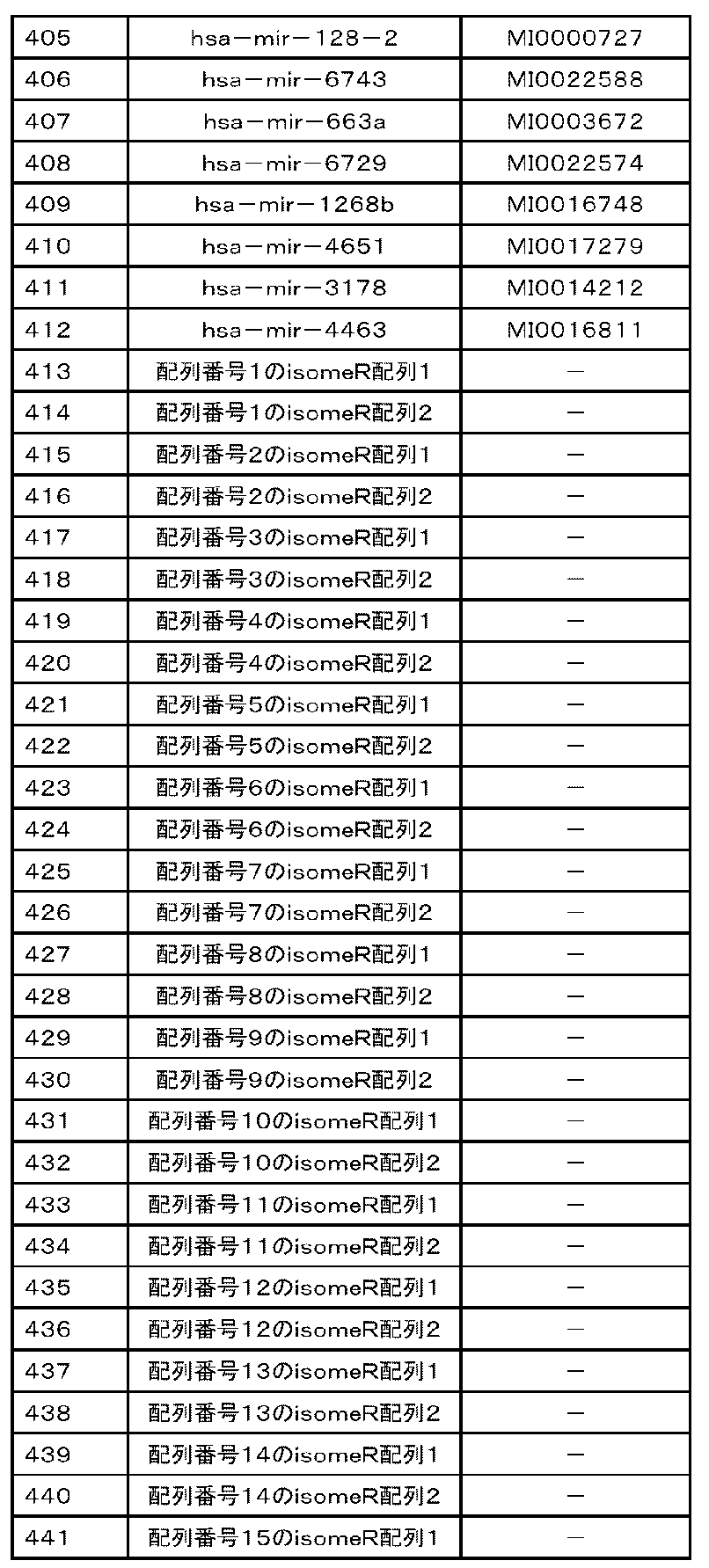
- the nucleic acid is the following polynucleotides (a) to (e): (A) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases. That fragment, (B) A polynucleotide containing the base sequence represented by any of SEQ ID NOs: 1 to 109, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (C) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more. Its fragment, which contains consecutive bases of (D) A polynucleotide containing a nucleotide sequence represented by any of SEQ ID NOs: 1 to 109 or a nucleotide sequence complementary to a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more.
- kits according to (1) which is a polynucleotide selected from the group consisting of.
- the kit is another hippocampal atrophy marker, miR-1228-5p, miR-760, miR-187-5p, miR-7111-5p, miR-6088, miR-6805-3p, miR-4640. -5p, miR-6721-5p, miR-6880-5p, miR-711, miR-128-1-5p, miR-4525, miR-486-3p, miR-6756-5p, miR-1260b, miR-3184 -5p, miR-6075, miR-204-3p, miR-4728-5p, miR-4534, miR-4758-5p, miR-8063, miR-6863-3p, miR-6789-5p, miR-744-5p , MiR-1909-3p, miR-887-3p, miR-4745-5p, miR-4433a-3p, miR-5090, miR-296-5p, miR-939-5p, miR-3648, miR-3196, miR -67
- the nucleic acid is the following polynucleotides (f) to (j): (F) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases. That fragment, (G) A polynucleotide containing a base sequence represented by any of SEQ ID NOs: 110 to 200, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (H) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more. Its fragment, which contains consecutive bases of (I) A polynucleotide containing a nucleotide sequence represented by any of SEQ ID NOs: 110 to 200 or a nucleotide sequence complementary to a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more.
- kits according to (3) which is a polynucleotide selected from the group consisting of.
- Kaiba atrophy markers miR-3131, miR-6757-5p, miR-4706, miR-5001-5p, miR-3180-3p, miR-642b-3p, miR-4655-5p, miR-6819 -5p, miR-937-5p, miR-4688, miR-6471-5p, miR-7107-5p, miR-4271, miR-1229-5p, miR-4707-5p, miR-6808-5p, miR-4656 , MiR-6076, miR-6762-5p, miR-7109-5p, miR-6732-5p, miR-3195, miR-7150, miR-642a-3p, miR-1249-5p, miR-3185, miR-4689 , MiR-3141, miR-6840-3p, miR-3135b, miR-1914-3p, miR-4446-3p, miR-4433b-3p, miR-6877-5p, miR-6884-5p, miR-3620-5
- the nucleic acid is the following polynucleotides (a) to (e): (A) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases. That fragment, (B) A polynucleotide containing the base sequence represented by any of SEQ ID NOs: 1 to 109, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (C) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more. Its fragment, which contains consecutive bases of (D) A polynucleotide containing a nucleotide sequence represented by any of SEQ ID NOs: 1 to 109 or a nucleotide sequence complementary to a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more.
- the device according to (5) which is a polynucleotide selected from the group consisting of.
- the device is another hippocampal atrophy marker, miR-1228-5p, miR-760, miR-187-5p, miR-7111-5p, miR-6088, miR-6805-3p, miR-4640. -5p, miR-6721-5p, miR-6880-5p, miR-711, miR-128-1-5p, miR-4525, miR-486-3p, miR-6756-5p, miR-1260b, miR-3184 -5p, miR-6075, miR-204-3p, miR-4728-5p, miR-4534, miR-4758-5p, miR-8063, miR-6863-3p, miR-6789-5p, miR-744-5p , MiR-1909-3p, miR-887-3p, miR-4745-5p, miR-4433a-3p, miR-5090, miR-296-5p, miR-939-5p, miR-3648, miR-3196, miR -
- the nucleic acid is the following polynucleotides (f) to (j): (F) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases. That fragment, (G) A polynucleotide containing a base sequence represented by any of SEQ ID NOs: 110 to 200, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (H) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more. Its fragment, which contains consecutive bases of (I) A polynucleotide containing a nucleotide sequence represented by any of SEQ ID NOs: 110 to 200 or a nucleotide sequence complementary to a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more.
- the device according to (7) which is a polynucleotide selected from the group consisting of.
- the hippocampal atrophy markers miR-3131, miR-6757-5p, miR-4706, miR-5001-5p, miR-3180-3p, miR-642b-3p, miR-4655 -5p, miR-6819-5p, miR-937-5p, miR-4688, miR-6471-5p, miR-7107-5p, miR-4271, miR-1229-5p, miR-4707-5p, miR-6808 -5p, miR-4656, miR-6076, miR-6762-5p, miR-7109-5p, miR-6732-5p, miR-3195, miR-7150, miR-642a-3p, miR-1249-5p, miR -3185, miR-4689, miR-3141, miR-6840-3p, miR-3135b, miR-1914-3p, miR-4446-3p, miR-4433b-3p, miR-6877-5p, miR-6
- the expression level of the polynucleotide is measured using the kit according to any one of (1) to (4) or the device according to any one of (5) to (10), according to (11). The method described.
- the measured expression level of the polynucleotide is the expression level of the polynucleotide in a sample derived from a subject whose hippocampus is known to be atrophied and that the hippocampal is not atrophied.
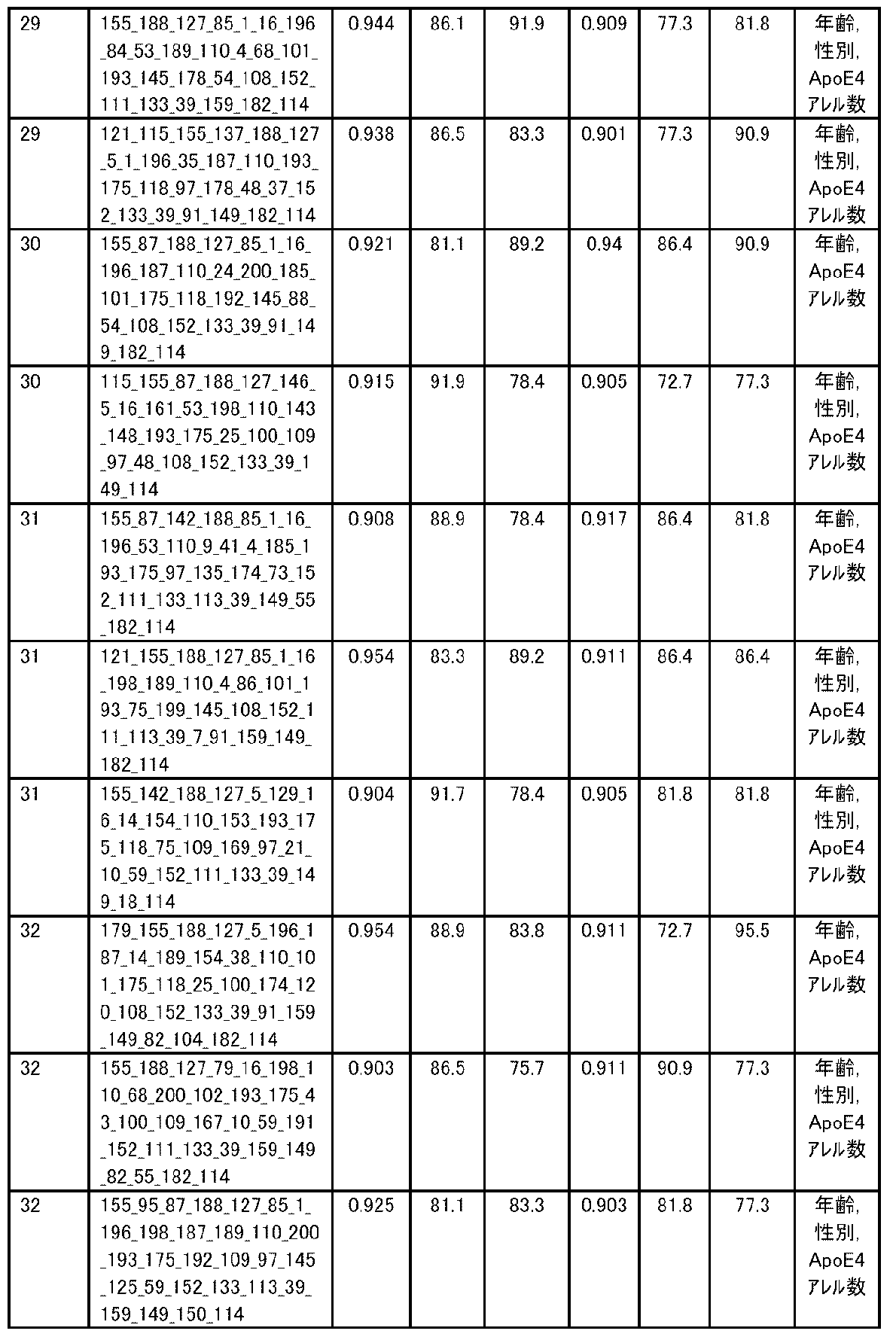
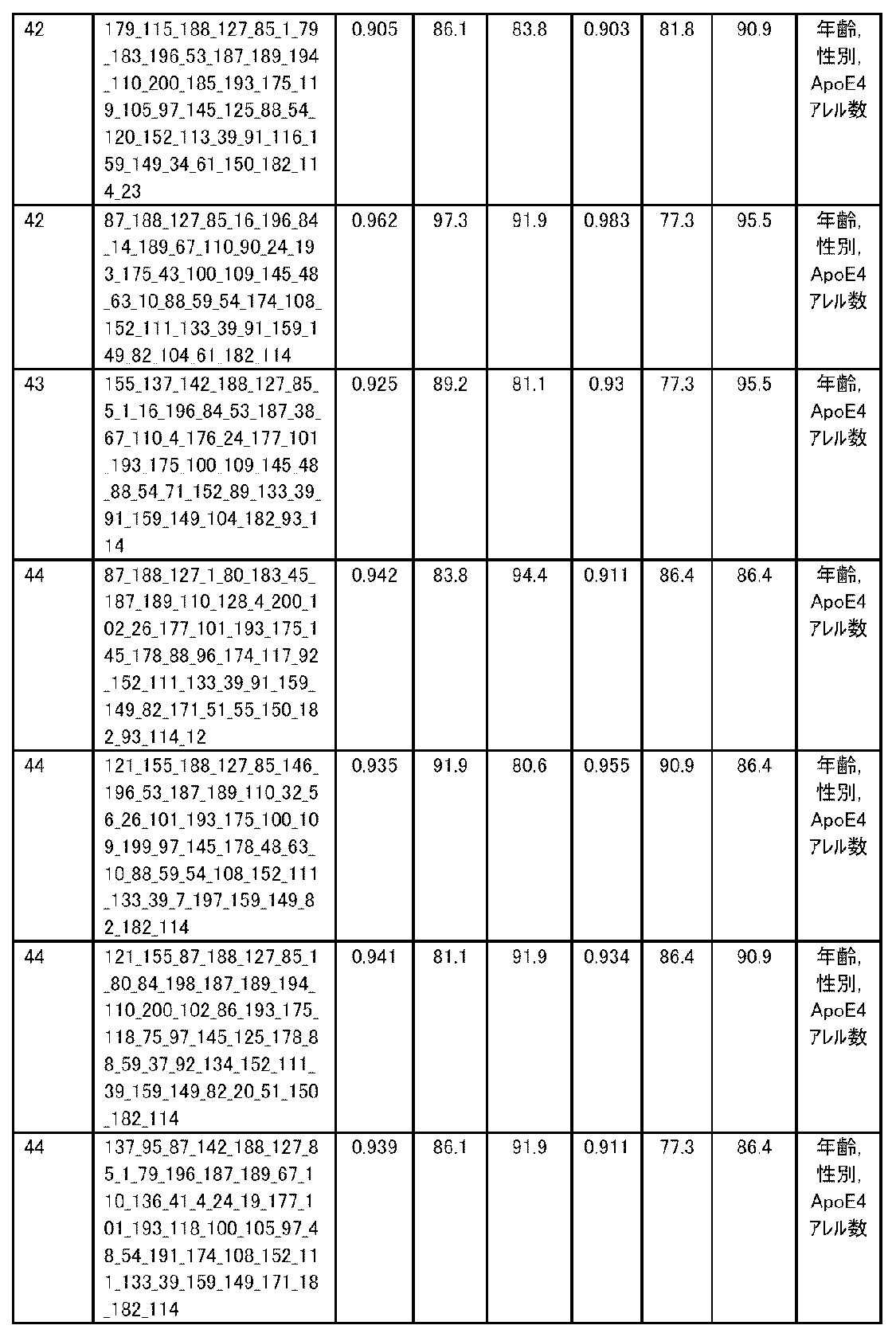
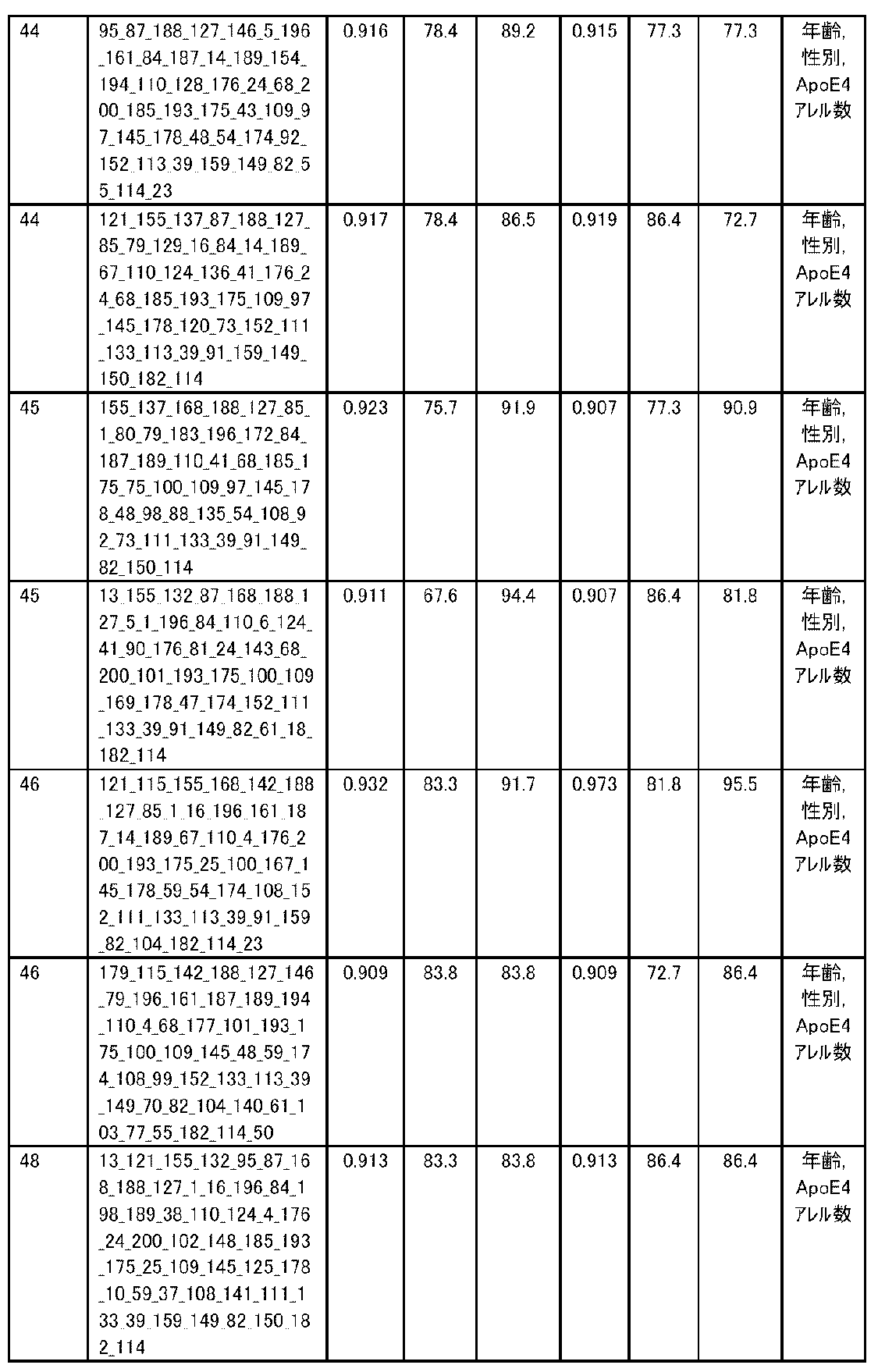
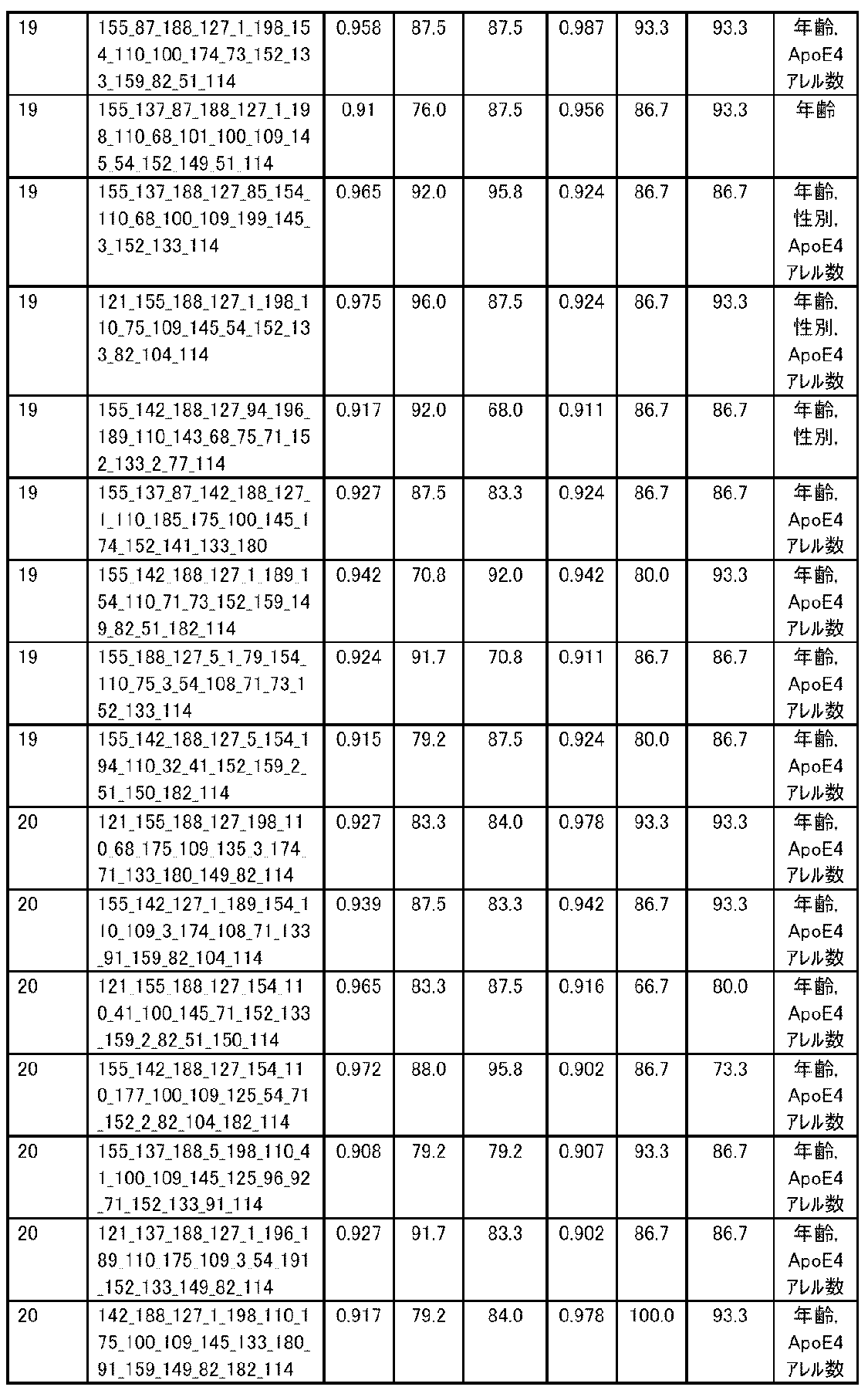
- the expression level of the polynucleotide in the sample derived from the subject is substituted into a discriminant formula capable of distinguishing atrophy or non-atrophy of the hippocampus prepared as a teacher sample, thereby causing atrophy or non-atrophy of the hippocampus.
- the nucleic acid is the following polynucleotides (a) to (e): (A) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases. That fragment, (B) A polynucleotide containing the base sequence represented by any of SEQ ID NOs: 1 to 109, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (C) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more. Its fragment, which contains consecutive bases of (D) A polynucleotide containing a nucleotide sequence represented by any of SEQ ID NOs: 1 to 109 or a nucleotide sequence complementary to a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more.
- the kit or device is another hippocampal atrophy marker, miR-1228-5p, miR-760, miR-187-5p, miR-7111-5p, miR-6088, miR-6805-3p, miR. -4640-5p, miR-6721-5p, miR-6880-5p, miR-711, miR-128-1-5p, miR-4525, miR-486-3p, miR-6756-5p, miR-1260b, miR -3184-5p, miR-6075, miR-204-3p, miR-4728-5p, miR-4534, miR-4758-5p, miR-8063, miR-6863-3p, miR-6789-5p, miR-744 -5p, miR-1909-3p, miR-887-3p, miR-4745-5p, miR-4433a-3p, miR-5090, miR-296-5p, miR-939-5p, miR-3648, miR-3196 , Mi
- the nucleic acid is the following polynucleotides (f) to (j): (F) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases. That fragment, (G) A polynucleotide containing a base sequence represented by any of SEQ ID NOs: 110 to 200, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (H) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more. Its fragment, which contains consecutive bases of (I) A polynucleotide containing a nucleotide sequence represented by any of SEQ ID NOs: 110 to 200 or a nucleotide sequence complementary to a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more.
- the term "polynucleotide” is used for nucleic acids including RNA, DNA, and RNA / DNA (chimera).
- the DNA includes any of cDNA, genomic DNA, and synthetic DNA.
- the RNA includes all of total RNA, mRNA, rRNA, miRNA, siRNA, snoRNA, snRNA, non-coding RNA and synthetic RNA.
- synthetic DNA and “synthetic RNA” are artificially based on a predetermined base sequence (either a natural sequence or a non-natural sequence), for example, using an automatic nucleic acid synthesizer. Refers to the produced DNA and RNA.
- non-natural sequence is intended to be used in a broad sense and is different from the natural sequence, for example, a sequence containing substitutions, deletions, insertions and / or additions of one or more nucleotides. That is, it includes a mutant sequence), a sequence containing one or more modified nucleotides (ie, a modified sequence), and the like.
- polynucleotides are used interchangeably with nucleic acids. Polynucleotides herein include two or more base nucleotides.
- fragment is a polynucleotide having a base sequence of a continuous portion of a polynucleotide, and preferably has a length of 15 bases or more, preferably 17 bases or more, and more preferably 19 bases or more. ..
- the term "gene” includes not only RNA and double-stranded DNA, but also single-stranded DNA such as the positive strand (or sense strand) or complementary strand (or antisense strand) that compose it. It is used with the intention of doing so. Moreover, the length is not particularly limited.
- the term "gene” as used herein refers to double-stranded DNA containing human genomic DNA, single-stranded DNA (positive strand), and single-stranded DNA having a sequence complementary to the positive strand (complementary). Includes any of strands), cDNA, microRNAs (miRNAs), and fragments thereof, the human genome, and their transcripts. Further, the “gene” is not only a “gene” represented by a specific base sequence (or SEQ ID NO:), but also an RNA having a biological function equivalent to that of the RNA encoded by these, for example, a homologue (that is, a homologue).
- Orthologs variants such as gene polymorphisms
- “nucleic acids” encoding derivatives are included.
- Specific examples of the "nucleic acid” encoding such a homologue, variant or derivative include the base sequence represented by any of SEQ ID NOs: 1 to 740 or the base sequence thereof under stringent conditions described later.
- a "nucleic acid” having a base sequence that hybridizes with a complementary sequence of a base sequence in which u is t can be mentioned.
- the “gene” does not ask whether the functional region is different, and may include, for example, an expression control region, a coding region, an exon, or an intron.
- the "gene” may be contained in a cell, may be released extracellularly and exist alone, or may be contained in a vesicle called an exosome.
- exosome or “exosome” is a vesicle wrapped in a lipid bilayer membrane secreted from a cell.
- Exosomes are derived from polyvesicular endosomes, and when released into the extracellular environment, they may contain "genes” such as RNA and DNA and biological substances such as proteins. Exosomes are known to be contained in body fluids such as blood, serum, plasma, cerebrospinal fluid, and lymph.
- RNA refers to RNA synthesized using the DNA sequence of a gene as a template.
- RNA is synthesized in such a way that RNA polymerase binds to a site called a promoter upstream of a gene and ribonucleotides are bound to the 3'end so as to be complementary to the base sequence of DNA.
- This RNA includes not only the gene itself, but also the entire sequence from the transcription start site to the end of the poly A sequence, including the expression control region, coding region, exon or intron.
- microRNA in the present specification is transcribed as an RNA precursor having a hairpin-like structure, cleaved by dsRNA cleaving enzyme having RNase III cleaving activity, and is a protein complex called RISC.
- RISC protein complex
- the "miRNA” used in the present specification is not only a so-called “miRNA” represented by a specific base sequence (or SEQ ID NO:), but also a precursor (pre-miRNA, tri-miRNA) of the "miRNA”.
- MiRNAs that have the same biological function as miRNAs represented by a particular base sequence such as their homologues (ie, homologs or orthologs), variants such as gene polymorphisms, and derivatives.
- a particular base sequence or SEQ ID NO:
- homologues ie, homologs or orthologs
- variants such as gene polymorphisms, and derivatives.
- miRNA also includes “miRNA”.
- the "miRNA” which is such a precursor, homologue, variant or derivative can be specifically identified by, for example, “miRBase” (version 21), and is sequenced under stringent conditions described later. Examples thereof include “miRNA” having a base sequence that hybridizes with a complementary sequence of any specific base sequence represented by any of Nos. 1 to 740.
- miRBase version 21
- miRNA version 21
- miRNA precursors eg, pre-miRNAs or pre-miRNAs as described above.
- probe refers to a polynucleotide, for example, a polynucleotide used for specifically detecting RNA generated by expression of a gene or a polynucleotide derived from the same, and / or a polynucleotide complementary thereto. Including.
- a "primer” is a continuous poly that can specifically recognize (that is, specifically bind to) a polynucleotide, for example, RNA generated by expression of a gene or a polynucleotide derived from the polynucleotide, and can amplify the polynucleotide. Includes nucleotides and / or polynucleotides complementary thereto.
- the "complementary polynucleotide” (reverse chain) is a full-length sequence of a polynucleotide consisting of a base sequence defined by any one of SEQ ID NOs: 1 to 740 or a base sequence in which u is t in the base sequence.
- a polynucleotide having a basically complementary relationship with respect to its partial sequence here, for convenience, this is referred to as a normal chain
- a base pair relationship such as A: T (U) or G: C.
- such a “complementary polynucleotide” is not limited to the case where it forms a completely complementary sequence with the base sequence of the target positive chain, and can hybridize with the target positive chain under stringent conditions. It may have a complementary relationship.
- complementary strand means a single-stranded polynucleotide consisting of a base sequence that is completely (at a 100% level) complementary to a specific single-stranded polynucleotide.
- stringent condition means that a polynucleotide such as a nucleic acid probe or nucleic acid primer is more detectable than other polynucleotides (for example, the average of background measurements + background measurement).
- a condition for hybridizing to the target polynucleotide with a standard error (measured value of 2 or more).
- Stringent conditions are sequence-dependent and vary depending on the environment in which hybridization takes place. By controlling the stringency of hybridization and / or washing conditions, target polynucleotides that are complementary (eg, 100% complementary) to the polynucleotide can be defined. Specific examples of “stringent conditions” will be described later.
- the "Tm value” means the temperature at which the double-stranded portion of the polynucleotide is denatured into a single strand and the double-stranded and single-stranded are present in a ratio of 1: 1.
- the term "variant" refers to a natural variant due to polymorphism, mutation, etc., or a SEQ ID NO: (in the present invention, for example, any of SEQ ID NOs: 1 to 740) in the case of a nucleic acid. Deletion, substitution, addition or insertion of 1, 2 or 3 or more (for example, 1 to several) bases in the represented base sequence or the base sequence in which u is t in the base sequence, or a subsequence thereof.
- RNA premature miRNA
- the base sequence of the precursor RNA (premature miRNA) of the base sequence of the relevant SEQ ID NO the base sequence in which u is t in the base sequence, or a partial sequence thereof.
- It can be a mutant, or a variant that is a nucleic acid that hybridizes with a polynucleotide or oligonucleotide containing the base sequence or a partial sequence thereof under stringent conditions as defined above.
- an oligonucleotide means a polynucleotide having a length of 2 to 100 bases.
- “several” or “several kinds” means an integer of about 10, 9, 8, 7, 6, 5, 4, 3 or 2 or kinds.
- the mutant can be prepared by using a well-known technique such as a site-specific mutagenesis method or a mutagenesis method using a PCR method.
- % identity is based on BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and FASTA (http://www.genome.jp/tools/fasta/). It can be determined with or without a gap introduced using a protein or gene retrieval system (Zheng Zhang et al., 2000, J. Comput. Biol., Vol. 7, p203-214; Altschul, S.F. et al., 1990, Journal of Molecular Biology, Vol. 215, p403-410; Pearson, WR et al., 1988, Proc. Natl. Acad. Sci. U.S.A.,. Vol. 85, p2444-2448).
- the term "derivative" refers to a modified nucleic acid, but not limited to, for example, a labeled derivative such as a fluorescent group, a modified nucleotide (for example, an alkyl such as halogen or methyl, an alkoxy such as methoxy, a group such as thio or carboxymethyl. Nucleotides containing nucleotides and bases, nucleotides containing double bond saturation, deaminoization, nucleotides that have undergone substitution of oxygen molecules with sulfur molecules, etc.), PNA (peptide nucleic acid; Nielsen, PE. Et al., 1991, Science, Vol. 254, p1497-500), LNA (locked nucleic acid; Obika, S. et al., 1998, Tetrahedron Lett., Vol. 39, p5401-5404).
- a labeled derivative such as a fluorescent group
- a modified nucleotide for example, an alkyl such as halogen or methyl, an
- a polynucleotide selected from the group of miRNAs that are markers of hippocampus atrophy, or a "nucleic acid” that can specifically bind to a complementary strand of the polynucleotide is a synthesized or prepared nucleic acid, and specifically. Includes “nucleic acid probe” or "primer”.
- the nucleic acid is used to detect the presence or absence of hippocampal atrophy in a subject, to diagnose the presence or absence of improvement in hippocampal atrophy, the degree of improvement, susceptibility to treatment of hippocampal atrophy, or to prevent, improve or treat hippocampal atrophy. It can be used directly or indirectly to screen for useful candidate substances.
- the nucleic acid is a transcript consisting of a nucleotide sequence represented by any of SEQ ID NOs: 1 to 740 associated with hippocampal atrophy in a living body, particularly a sample such as a body fluid such as blood or urine, or a cDNA synthetic nucleic acid thereof, or a complement thereof.
- oligonucleotides and polynucleotides capable of specifically recognizing and binding chains serve as probes for detecting the genes / miRNAs expressed in vivo, tissues, cells, etc. based on the above properties, and primers for amplifying the genes / miRNAs expressed in vivo. Can be effectively used as.
- detection as used herein may be replaced by the terms inspection, measurement, detection or determination support.
- evaluation is used herein to include supporting diagnosis or evaluation based on a test result or a measurement result.
- subject includes humans, primates including monkeys such as chimpanzees and gorillas, pet animals such as dogs and cats, domestic animals such as cows, horses, sheep and goats, mice and rats. Means mammals such as rodents and animals kept in zoos. A preferred subject is a human.
- normal hippocampus "subject with hippocampal atrophy” and “subject with non-hippocampal atrophy” are also such mammals, and the hippocampal atrophy to be detected is not specified. Means no animal.
- Preferred hippocampal normal subjects, subjects with hippocampal atrophy, and subjects with non-hippocampal atrophy are humans.
- the terms “subject” and “patient” can be used interchangeably, but the more typical "patient” is human.
- the "hippocampus” used in the present specification is located in the medial part of the temporal lobe of the cerebrum and at the base of the inferior horn of the lateral ventricle, and is a cortical part indispensable for the formation of overt memory such as episodic memory.
- hippocampal atrophy refers to a condition in which the volume of an organ or tissue that has grown to a normal volume has decreased due to various causes.
- hippocampal atrophy means that the size of the hippocampus is reduced to the extent that it adversely affects hippocampal function. Hippocampal atrophy typically means that the volume of the hippocampus decreases until the volume ratio (%) of the hippocampus to the whole brain is 0.6% or less. Hippocampal atrophy generally progresses progressively with aging, as described above. Diseases associated with hippocampal atrophy other than aging include, but are not limited to, Alzheimer-type dementia, frontotemporal lobar degeneration, depression, post-traumatic stress disorder, and schizophrenia.
- the "hippocampal atrophy degree” used in the present specification means the degree of hippocampal atrophy in a patient, and can be determined using the hippocampal volume ratio (%) to the whole brain as an index.
- hippocampal non-atrophic patients are those whose hippocampal volume ratio to total brain (ratio of hippocampal volume to total brain volume) is 0.63% or more, 0.64% or more, 0. Includes .68% or more, or 0.86% or more.
- hippocampal atrophy patients are those whose hippocampal volume ratio to total brain (hippocampal volume ratio to total brain volume) is 0.60% or less, 0.59% or less, and 0. Includes .57% or less, or 0.38% or less.
- Alzheimer's disease and “Alzheimer's disease” are dementias in which accumulation of amyloid ⁇ and phosphorylated tau are observed in the brain, and neurodegeneration characterized by hippocampal atrophy is caused. Is.
- patients with mild dementia are patients who do not develop dementia but have signs, specifically, a dementia test (MMSE) score of 25 or less. Refers to the patient.
- MMSE dementia test
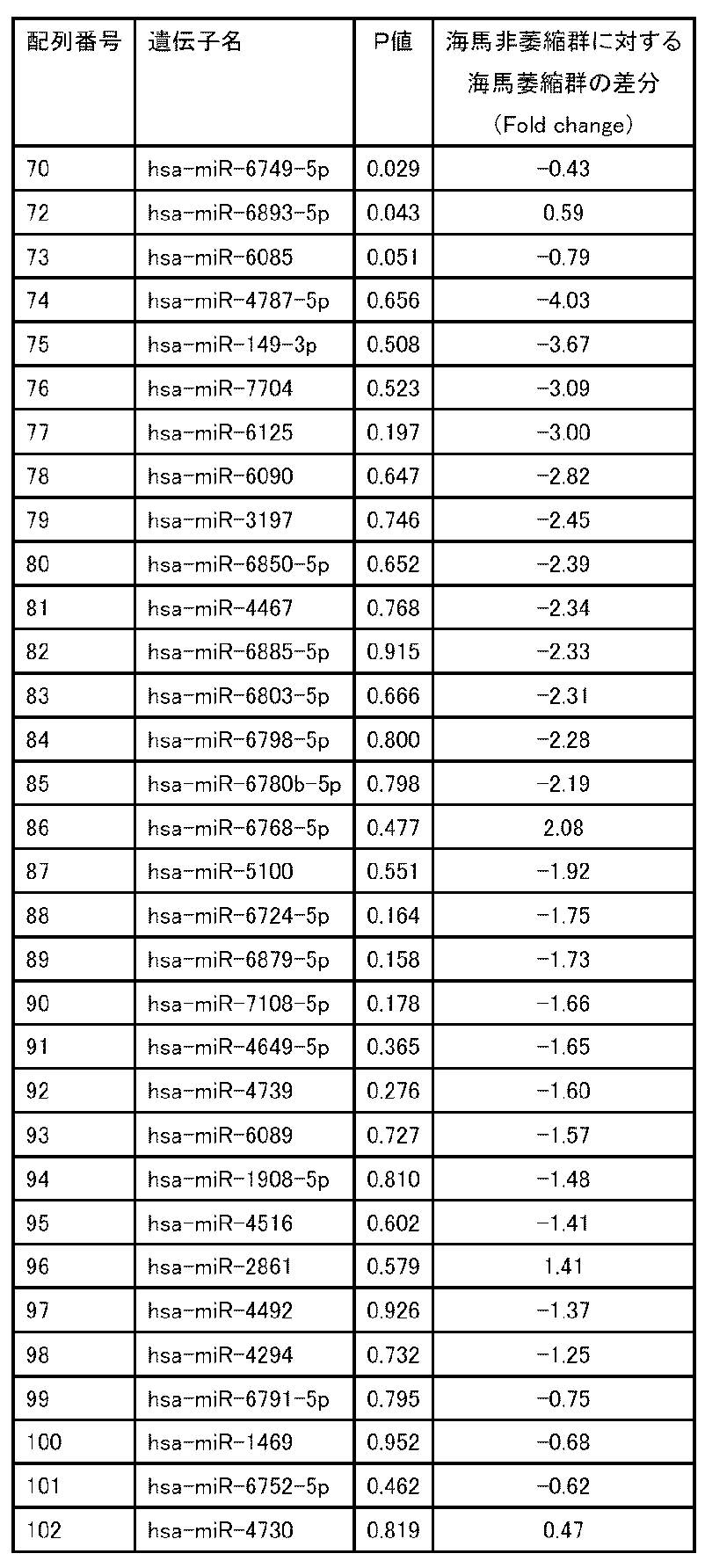
- P or "P value” is the probability that a statistic that is more extreme than the statistic actually calculated from the data under the null hypothesis will be observed in a statistical test. Is shown. Therefore, the smaller the "P” or "P value", the more significant the difference can be considered between the comparison targets.
- sensitivity means a value of (number of true positives) / (number of true positives + number of false negatives). Higher sensitivity enables early detection of hippocampal atrophy and early medical intervention.
- specificity means (number of true negatives) / (number of true negatives + number of false positives). If the specificity is high, non-hippocampal atrophy patients (healthy subjects) such as those with normal cognitive function can be prevented from performing unnecessary additional tests by misidentifying them as hippocampal atrophy patients, leading to reduction of the burden on patients and reduction of medical expenses. ..
- accuracy means a value of (number of true positives + number of true negatives) / (total number of cases). The accuracy indicates the ratio of the discrimination results to all the samples being correct, and is the first index for evaluating the discrimination performance.
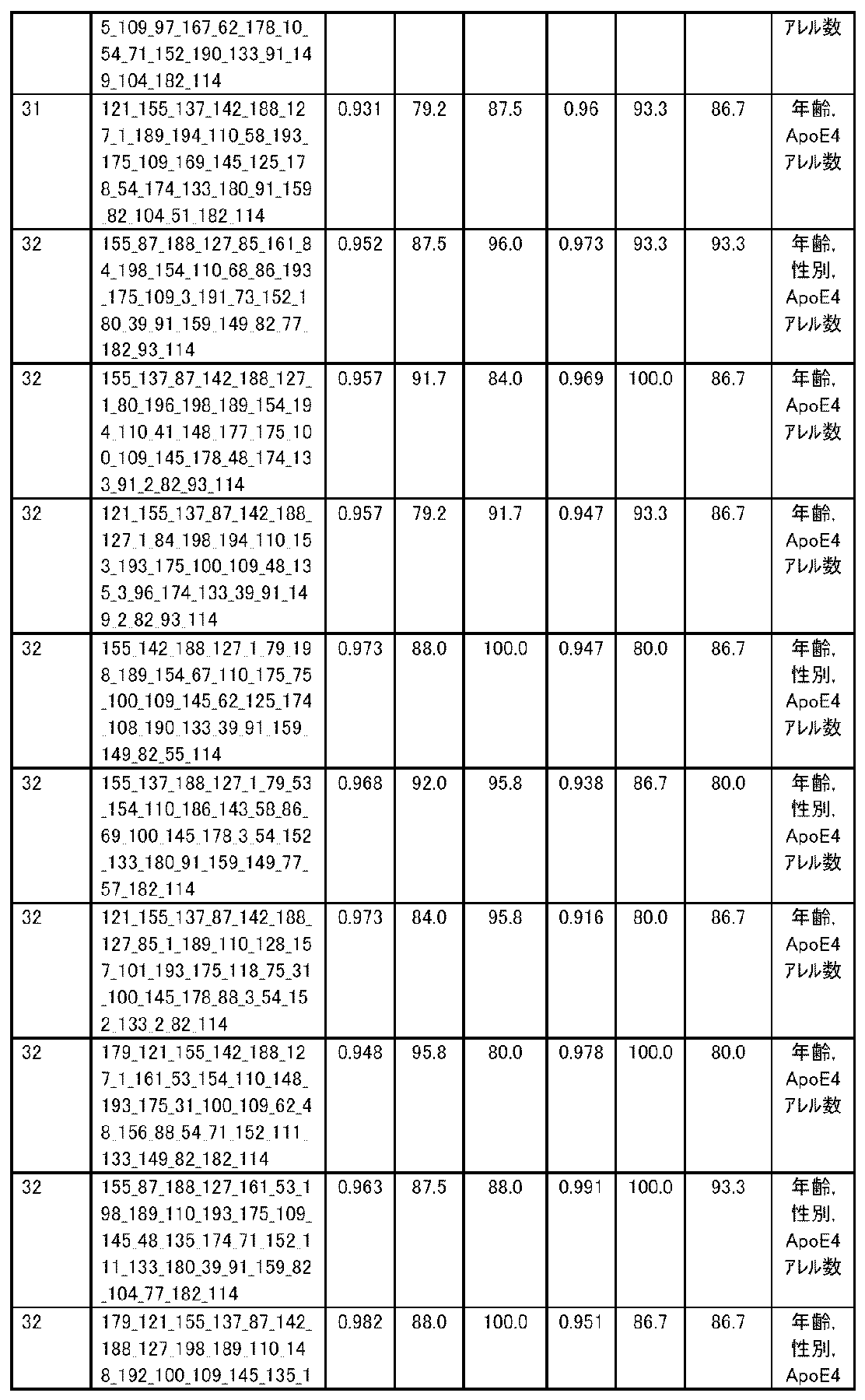
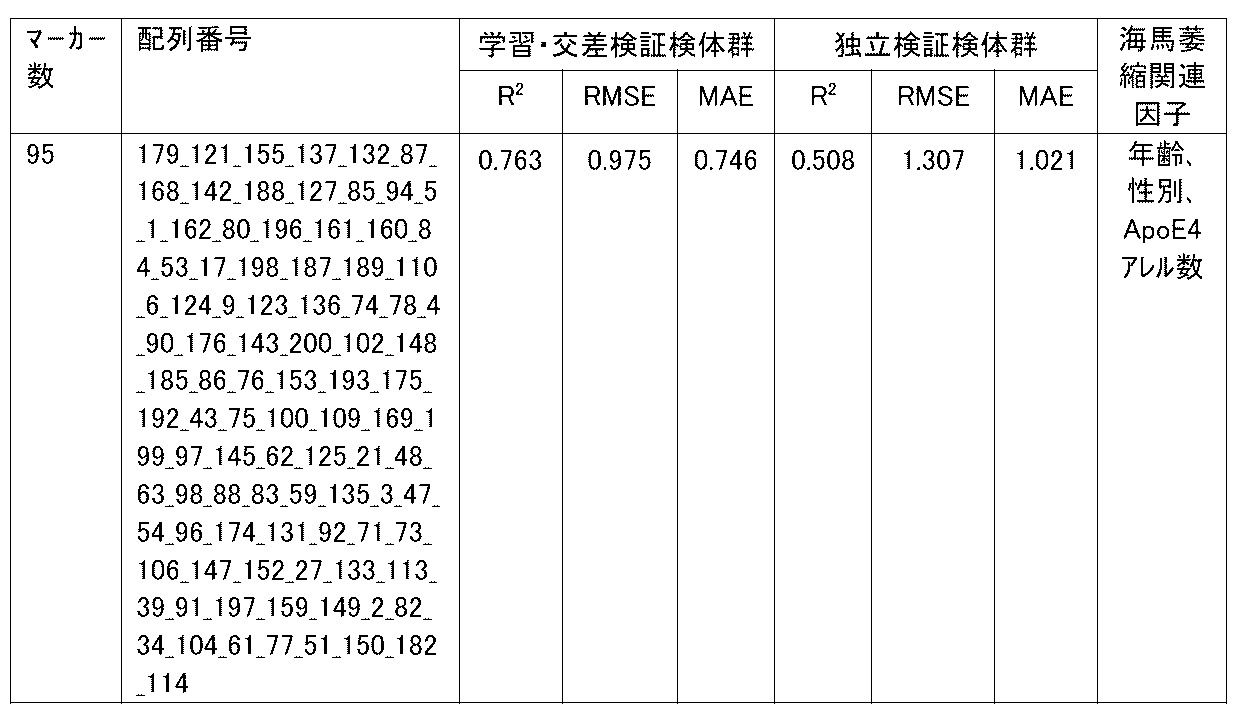
- R 2 is the value of the square of the correlation coefficient R.
- the correlation coefficient R is an index showing the correlation between two variables.
- RMSE is an abbreviation for root mean squared error (root mean square error) and is one of the indexes for evaluating the error of the regression equation model. The closer the observed value and the predicted value are, the smaller the RMSE becomes.
- MAE is an abbreviation for mean absolute error (mean absolute error), and is one of the indexes for evaluating the error of the regression equation model like RMSE. It becomes smaller as the observed value and the predicted value get closer.
- the "specimen" to be determined, detected or diagnosed is the gene / miRNA of the hippocampal atrophy marker of the present invention according to the presence or absence of hippocampal atrophy, the progression of hippocampal atrophy, and the exertion of a therapeutic effect on hippocampal atrophy.
- the specimens are brain tissue and nerve cells, nerve tissue, cerebrospinal fluid, bone marrow fluid, organs, skin, body fluid such as blood, urine, saliva, sweat, and tissue exudate, serum prepared from blood, and plasma. , Stool, hair, etc.
- the sample extracted from these may be specifically RNA, miRNA, or the like.
- hsa-miR-3131 gene or “hsa-miR-3131” refers to the hsa-miR-3131 gene (miRBase Accession No. MIMAT0014996) shown in SEQ ID NO: 1 and other species. Includes homologs or orthologs.
- the hsa-miR-3131 gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-3131", “hsa-mir-3131” (miRBase Accession No. MI0014151, SEQ ID NO: 201) having a hairpin-like structure is known.
- hsa-miR-6757-5p gene or "hsa-miR-6757-5p” used herein refers to the hsa-miR-6757-5p gene (miRBase Accession No. MIMAT0027414) and other species such as homologs or orthologs.
- the hsa-miR-6757-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6757-5p", “hsa-mir-6757” (miRBase Accession No. MI0022602, SEQ ID NO: 202) having a hairpin-like structure is known.
- hsa-miR-4706 gene or "hsa-miR-4706” refers to the hsa-miR-4706 gene (miRBase Accession No. MIMAT0019806) set forth in SEQ ID NO: 3 and other species. Includes homologs or orthologs.
- the hsa-miR-4706 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4706", “hsa-mir-4706” (miRBase Accession No. MI0017339, SEQ ID NO: 203) having a hairpin-like structure is known.
- hsa-miR-5001-5p gene or "hsa-miR-5001-5p” refers to the hsa-miR-5001-5p gene (miRBase Accession No. Includes MIMAT0021021) and other species homologs or orthologs.
- the hsa-miR-5001-5p gene can be obtained by the method described in Hansen TB et al., 2011, RNA Biol, Volume 8, p378-383. Further, as the precursor of "hsa-miR-5001-5p", “hsa-mir-5001" (miRBase Accession No. MI0017867, SEQ ID NO: 204) having a hairpin-like structure is known.
- hsa-miR-3180-3p gene or "hsa-miR-3180-3p” refers to the hsa-miR-3180-3p gene (miRBase Accession No. 5) set forth in SEQ ID NO: 5. MIMAT0015058) and other species such as homologs or orthologs are included.
- the hsa-miR-3180-3p gene can be obtained by the method described in Creighton CJ et al., 2010, PLos One, Volume 5, e9637.
- hsa-miR-3180-3p has a hairpin-like structure as a precursor thereof, "hsa-mir-3180-1, hsa-mir-3180-2, hsa-mir-3180-3” (miRBase Accession). No. MI0014214, MI0014215, MI0014217, SEQ ID NOs: 205, 206, 207) are known.
- hsa-miR-642b-3p gene or "hsa-miR-642b-3p” used herein refer to the hsa-miR-642b-3p gene (miRBase Accession No. 6) set forth in SEQ ID NO: 6. Includes MIMAT0018444) and other species homologs or orthologs.
- the hsa-miR-642b-3p gene can be obtained by the method described in Witten D et al., 2010, BMC Biol, Volume 8, p58. Further, as the precursor of "hsa-miR-642b-3p", “hsa-mir-642b” (miRBase Accession No. MI0016685, SEQ ID NO: 208) having a hairpin-like structure is known.
- hsa-miR-4655-5p gene or "hsa-miR-4655-5p” refers to the hsa-miR-4655-5p gene (miRBase Accession No. Includes MIMAT0019721) and other species homologs or orthologs.
- the hsa-miR-4655-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4655-5p", "hsa-mir-4655" (miRBase Accession No. MI0017283, SEQ ID NO: 209) having a hairpin-like structure is known.
- hsa-miR-6819-5p gene or "hsa-miR-6819-5p” refers to the hsa-miR-6819-5p gene (miRBase Accession No. MIMAT0027538) and other species such as homologs or orthologs are included.
- the hsa-miR-6819-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6819-5p", “hsa-mir-6819” (miRBase Accession No. MI0022664, SEQ ID NO: 210) having a hairpin-like structure is known.
- hsa-miR-937-5p gene or "hsa-miR-937-5p” refers to the hsa-miR-937-5p gene (miRBase Accession No. MIMAT0022938) and other species such as homologs or orthologs.
- the hsa-miR-937-5p gene can be obtained by the method described in Lui WO et al., 2007, Cancer Res, Vol. 67, p6031-6043. Further, as the precursor of "hsa-miR-937-5p", "hsa-mir-937” (miRBase Accession No. MI0005759, SEQ ID NO: 211) having a hairpin-like structure is known.
- hsa-miR-4688 gene or “hsa-miR-4688” refers to the hsa-miR-4688 gene (miRBase Accession No. MIMAT0019777) set forth in SEQ ID NO: 10 and other species. Includes homologs or orthologs.
- the hsa-miR-4688 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4688", “hsa-mir-4688” (miRBase Accession No. MI0017321, SEQ ID NO: 212) having a hairpin-like structure is known.
- hsa-miR-6741-5p gene or "hsa-miR-6741-5p” used herein refer to the hsa-miR-6741-5p gene (miRBase Accession No. 1) set forth in SEQ ID NO: 11. Includes MIMAT0027383) and other species homologs or orthologs.
- the hsa-miR-6741-5p gene can be obtained by the method described in Ladewig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6741-5p", “hsa-mir-6741” (miRBase Accession No. MI0022586, SEQ ID NO: 213) having a hairpin-like structure is known.
- hsa-miR-7107-5p gene or "hsa-miR-7107-5p” refers to the hsa-miR-7107-5p gene (miRBase Accession No. MIMAT0028111) and other species such as homologs or orthologs are included.
- the hsa-miR-7107-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-7107-5p", “hsa-mir-7107” (miRBase Accession No. MI0022958, SEQ ID NO: 214) having a hairpin-like structure is known.
- hsa-miR-4271 gene or “hsa-miR-4271” refers to the hsa-miR-4271 gene (miRBase Accession No. MIMAT0016901) set forth in SEQ ID NO: 13 and other species. Includes homologs or orthologs.
- the hsa-miR-4271 gene can be obtained by the method described in Goff LA et al., 2009, PLos One, Volume 4, e7192. Further, as the precursor of "hsa-miR-4271", “hsa-mir-4271” (miRBase Accession No. MI0015879, SEQ ID NO: 215) having a hairpin-like structure is known.
- hsa-miR-1229-5p gene or "hsa-miR-1229-5p” refers to the hsa-miR-1229-5p gene (miRBase Accession No. MIMAT0022942) and other species such as homologs or orthologs.
- the hsa-miR-1229-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, Vol. 28, p328-336. Further, as the precursor of "hsa-miR-1229-5p", “hsa-mir-1229” (miRBase Accession No. MI0006319, SEQ ID NO: 216) having a hairpin-like structure is known.
- hsa-miR-4707-5p gene or "hsa-miR-4707-5p” used herein refers to the hsa-miR-4707-5p gene (miRBase Accession No. MIMAT0019807) and other species such as homologs or orthologs are included.
- the hsa-miR-4707-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4707-5p", “hsa-mir-4707” (miRBase Accession No. MI0017340, SEQ ID NO: 217) having a hairpin-like structure is known.
- hsa-miR-6808-5p gene or "hsa-miR-6808-5p” refers to the hsa-miR-6808-5p gene (miRBase Accession No. MIMAT0027516) and other species such as homologs or orthologs are included.
- the hsa-miR-6808-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, “hsa-miR-6808-5p” has a hairpin-like structure as a precursor thereof, and "hsa-mir-6808” (miRBase Accession No. MI0022653, SEQ ID NO: 218) is known.
- hsa-miR-4656 gene or “hsa-miR-4656” refers to the hsa-miR-4656 gene (miRBase Accession No. MIMAT0019723) set forth in SEQ ID NO: 17 and other species. Includes homologs or orthologs.
- the hsa-miR-4656 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4656", “hsa-mir-4656” (miRBase Accession No. MI0017284, SEQ ID NO: 219) having a hairpin-like structure is known.
- hsa-miR-6076 gene or “hsa-miR-6076” refers to the hsa-miR-6076 gene (miRBase Accession No. MIMAT0023701) set forth in SEQ ID NO: 18 and other species. Includes homologs or orthologs.
- the hsa-miR-6076 gene can be obtained by the method described in Voellenkle C et al., 2012, RNA, Vol. 18, p472-484. Further, as the precursor of "hsa-miR-6076", “hsa-mir-6076” (miRBase Accession No. MI0020353, SEQ ID NO: 220) having a hairpin-like structure is known.
- hsa-miR-6762-5p gene or "hsa-miR-6762-5p” refers to the hsa-miR-6762-5p gene (miRBase Accession No. MIMAT0027424) and other species such as homologs or orthologs are included.
- the hsa-miR-6762-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6762-5p", “hsa-mir-6762” (miRBase Accession No. MI0022607, SEQ ID NO: 221) having a hairpin-like structure is known.
- hsa-miR-7109-5p gene or "hsa-miR-7109-5p” refers to the hsa-miR-7109-5p gene (miRBase Accession No. MIMAT0028115) and other species such as homologs or orthologs.
- the hsa-miR-7109-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-7109-5p", “hsa-mir-7109” (miRBase Accession No. MI0022960, SEQ ID NO: 222) having a hairpin-like structure is known.
- hsa-miR-6732-5p gene or "hsa-miR-6732-5p” used herein refers to the hsa-miR-6732-5p gene (miRBase Accession No. Includes MIMAT0027365) and other species homologs or orthologs.
- the hsa-miR-6732-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6732-5p", "hsa-mir-6732" (miRBase Accession No. MI0022575, SEQ ID NO: 223) having a hairpin-like structure is known.
- hsa-miR-3195 gene or “hsa-miR-3195” refers to the hsa-miR-3195 gene (miRBase Accession No. MIMAT0015079) set forth in SEQ ID NO: 22 and other species. Includes homologs or orthologs.
- the hsa-miR-3195 gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-3195", “hsa-mir-3195" (miRBase Accession No. MI0014240, SEQ ID NO: 224) having a hairpin-like structure is known.
- hsa-miR-7150 gene or “hsa-miR-7150” refers to the hsa-miR-7150 gene (miRBase Accession No. MIMAT0028211) and other species set forth in SEQ ID NO: 23. Includes homologs or orthologs.
- the hsa-miR-7150 gene can be obtained by the method described in Ouras A et al., 2009, Nucleic Acids Res, Vol. 37, p3276-3287. Further, as the precursor of "hsa-miR-7150", “hsa-mir-7150” (miRBase Accession No. MI0023610, SEQ ID NO: 225) having a hairpin-like structure is known.
- hsa-miR-642a-3p gene or "hsa-miR-642a-3p” used herein refer to the hsa-miR-642a-3p gene (miRBase Accession No. MIMAT0020924) and other species such as homologs or orthologs.
- the hsa-miR-642a-3p gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-1692. Further, as the precursor of "hsa-miR-642a-3p", “hsa-mir-642a” (miRBase Accession No. MI0003657, SEQ ID NO: 226) having a hairpin-like structure is known.
- hsa-miR-1249-5p gene or "hsa-miR-1249-5p” refers to the hsa-miR-1249-5p gene (miRBase Accession No. Includes MIMAT0032029) and other species homologs or orthologs.
- the hsa-miR-1249-5p gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, Vol. 18, p610-621. Further, as the precursor of "hsa-miR-1249-5p", “hsa-mir-1249” (miRBase Accession No. MI0006384, SEQ ID NO: 227) having a hairpin-like structure is known.
- hsa-miR-3185 gene or “hsa-miR-3185” refers to the hsa-miR-3185 gene (miRBase Accession No. MIMAT0015065) set forth in SEQ ID NO: 26 and other species. Includes homologs or orthologs.
- the hsa-miR-3185 gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-3185”, “hsa-mir-3185” (miRBase Accession No. MI0014227, SEQ ID NO: 228) having a hairpin-like structure is known.
- hsa-miR-4689 gene or "hsa-miR-4689” refers to the hsa-miR-4689 gene (miRBase Accession No. MIMAT0019778) set forth in SEQ ID NO: 27 and other species. Includes homologs or orthologs.
- the hsa-miR-4689 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4689", “hsa-mir-4689” (miRBase Accession No. MI0017322, SEQ ID NO: 229) having a hairpin-like structure is known.
- hsa-miR-3141 gene or “hsa-miR-3141” refers to the hsa-miR-3141 gene (miRBase Accession No. MIMAT0015010) set forth in SEQ ID NO: 28 and other species. Includes homologs or orthologs.
- the hsa-miR-3141 gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-3141", “hsa-mir-3141” (miRBase Accession No. MI0014165, SEQ ID NO: 230) having a hairpin-like structure is known.
- hsa-miR-6840-3p gene or "hsa-miR-6840-3p” refers to the hsa-miR-6840-3p gene (miRBase Accession No. Includes MIMAT0027583) and other species homologs or orthologs.
- the hsa-miR-6840-3p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6840-3p", "hsa-mir-6840" (miRBase Accession No. MI0022686, SEQ ID NO: 231) having a hairpin-like structure is known.
- hsa-miR-3135b gene or “hsa-miR-3135b” refers to the hsa-miR-3135b gene (miRBase Accession No. MIMAT0018985) set forth in SEQ ID NO: 30 and other species. Includes homologs or orthologs.
- the hsa-miR-3135b gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-3135b", “hsa-mir-3135b” (miRBase Accession No. MI0016809, SEQ ID NO: 232) having a hairpin-like structure is known.
- hsa-miR-1914-3p gene or "hsa-miR-1914-3p” used herein refer to the hsa-miR-1914-3p gene (miRBase Accession No. MIMAT0007890) and other species such as homologs or orthologs are included.
- the hsa-miR-1914-3p gene can be obtained by the method described in Bar M et al., 2008, Stem Cells, Vol. 26, p2496-2505. Further, as the precursor of "hsa-miR-1914-3p", “hsa-mir-1914" (miRBase Accession No. MI0008335, SEQ ID NO: 233) having a hairpin-like structure is known.
- hsa-miR-4446-3p gene or "hsa-miR-4446-3p” refers to the hsa-miR-4446-3p gene (miRBase Accession No. MIMAT0018965) and other species such as homologs or orthologs are included.
- the hsa-miR-4446-3p gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4446-3p", “hsa-mir-4446” (miRBase Accession No. MI0016789, SEQ ID NO: 234) having a hairpin-like structure is known.
- hsa-miR-4433b-3p gene or "hsa-miR-4433b-3p” used herein refer to the hsa-miR-4433b-3p gene (miRBase Accession No. 3p) set forth in SEQ ID NO: 33. Includes MIMAT0030414) and other species homologs or orthologs.
- the hsa-miR-4433b-3p gene can be obtained by the method described in Ple H et al., 2012, PLos One, Volume 7, e50746.
- hsa-miR-4433b-3p has a hairpin-like structure as a precursor thereof, and "hsa-mir-4433b” (miRBase Accession No. MI0025511, SEQ ID NO: 235) is known.
- hsa-miR-6877-5p gene or "hsa-miR-6877-5p” used herein refers to the hsa-miR-6877-5p gene (miRBase Accession No. 3) set forth in SEQ ID NO: 34. MIMAT0027654) and other species such as homologs or orthologs.
- the hsa-miR-6877-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6877-5p", “hsa-mir-6877” (miRBase Accession No. MI0022724, SEQ ID NO: 236) having a hairpin-like structure is known.
- hsa-miR-6884-5p gene or "hsa-miR-6884-5p” used herein refers to the hsa-miR-6884-5p gene (miRBase Accession No. 3) set forth in SEQ ID NO: 35. MIMAT0027596) and other species such as homologs or orthologs are included.
- the hsa-miR-6884-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6884-5p", “hsa-mir-6884" (miRBase Accession No. MI0022694, SEQ ID NO: 237) having a hairpin-like structure is known.
- hsa-miR-3620-5p gene or "hsa-miR-3620-5p” refers to the hsa-miR-3620-5p gene (miRBase Accession No. MIMAT0022967) and other species such as homologs or orthologs are included.
- the hsa-miR-3620-5p gene can be obtained by the method described in Witten D et al., 2010, BMC Biol, Volume 8, p58. Further, as the precursor of "hsa-miR-3620-5p", “hsa-mir-3620” (miRBase Accession No. MI0016011, SEQ ID NO: 238) having a hairpin-like structure is known.
- hsa-miR-6825-5p gene or "hsa-miR-6825-5p” used herein refers to the hsa-miR-6825-5p gene (miRBase Accession No. MIMAT0027550) and other species such as homologs or orthologs are included.
- the hsa-miR-6825-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6825-5p", "hsa-mir-6825” (miRBase Accession No. MI0022670, SEQ ID NO: 239) having a hairpin-like structure is known.
- hsa-miR-5739 gene or “hsa-miR-5739” refers to the hsa-miR-5739 gene (miRBase Accession No. MIMAT0023116) and other species set forth in SEQ ID NO: 38. Includes homologs or orthologs.
- the hsa-miR-5739 gene was described by Yoo JK et al., 2011, Biochem Biophyss Res Commun. It can be obtained by the method described in Volume 415, p258-262. Further, as the precursor of "hsa-miR-5739", “hsa-mir-5739” (miRBase Accession No. MI0019412, SEQ ID NO: 240) having a hairpin-like structure is known.
- hsa-miR-3663-3p gene or "hsa-miR-3663-3p” used herein refer to the hsa-miR-3663-3p gene (miRBase Accession No. 3p) set forth in SEQ ID NO: 39. MIMAT0018085) and other species such as homologs or orthologs.
- the hsa-miR-3663-3p gene can be obtained by the method described in Liao JY et al., 2010, PLos One, Volume 5, e10563. Further, as the precursor of "hsa-miR-3663-3p", “hsa-mir-3663” (miRBase Accession No. MI0016064, SEQ ID NO: 241) having a hairpin-like structure is known.
- hsa-miR-4695-5p gene or "hsa-miR-4695-5p” refers to the hsa-miR-4695-5p gene (miRBase Accession No. MIMAT0019788) and other species such as homologs or orthologs are included.
- the hsa-miR-4695-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as a precursor of "hsa-miR-4695-5p", "hsa-mir-4695" (miRBase Accession No. MI0017328, SEQ ID NO: 242) having a hairpin-like structure is known.
- hsa-miR-3162-5p gene or "hsa-miR-3162-5p” refers to the hsa-miR-3162-5p gene (miRBase Accession No. MIMAT0015036) and other species such as homologs or orthologs are included.
- the hsa-miR-3162-5p gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-3162-5p", “hsa-mir-3162” (miRBase Accession No. MI0014192, SEQ ID NO: 243) having a hairpin-like structure is known.
- hsa-miR-3679-5p gene or "hsa-miR-3679-5p” refers to the hsa-miR-3679-5p gene (miRBase Accession No. MIMAT0018104) and other species such as homologs or orthologs are included.
- the hsa-miR-3679-5p gene can be obtained by the method described in Creighton CJ et al., 2010, PLOS One, Volume 5, e9637. Further, as the precursor of "hsa-miR-3679-5p", “hsa-mir-3679” (miRBase Accession No. MI0016080, SEQ ID NO: 244) having a hairpin-like structure is known.
- hsa-miR-8059 gene or “hsa-miR-8059” refers to the hsa-miR-8059 gene (miRBase Accession No. MIMAT0030986) set forth in SEQ ID NO: 43 and other species. Includes homologs or orthologs.
- the hsa-miR-8059 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487. Further, as the precursor of "hsa-miR-8059", “hsa-mir-8059” (miRBase Accession No. MI0025895, SEQ ID NO: 245) having a hairpin-like structure is known.
- hsa-miR-7110-5p gene or "hsa-miR-7110-5p” refers to the hsa-miR-7110-5p gene (miRBase Accession No. Includes MIMAT0028117) and other species homologs or orthologs.
- the hsa-miR-7110-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-7110-5p", “hsa-mir-7110” (miRBase Accession No. MI0022961, SEQ ID NO: 246) having a hairpin-like structure is known.
- hsa-miR-1275 gene or “hsa-miR-1275” refers to the hsa-miR-1275 gene (miRBase Accession No. MIMAT0005929) set forth in SEQ ID NO: 45 and other species. Includes homologs or orthologs.
- the hsa-miR-1275 gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, Vol. 18, p610-621. Further, as the precursor of "hsa-miR-1275", “hsa-mir-1275” (miRBase Accession No. MI0006415, SEQ ID NO: 247) having a hairpin-like structure is known.
- hsa-miR-6779-5p gene or "hsa-miR-6779-5p” refers to the hsa-miR-6779-5p gene (miRBase Accession No. MIMAT0027458) and other species such as homologs or orthologs.
- the hsa-miR-6779-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6779-5p", “hsa-mir-6779” (miRBase Accession No. MI0022624, SEQ ID NO: 248) having a hairpin-like structure is known.
- hsa-miR-197-5p gene or "hsa-miR-197-5p” refers to the hsa-miR-197-5p gene (miRBase Accession No. Includes MIMAT0022691) and other species homologs or orthologs.
- the hsa-miR-197-5p gene can be obtained by the method described in Lagos-Quintana M et al., 2003, RNA, Volume 9, p175-179. Further, as a precursor of "hsa-miR-197-5p", "hsa-mir-197" (miRBase Accession No. MI0000239, SEQ ID NO: 249) having a hairpin-like structure is known.
- hsa-miR-6845-5p gene or "hsa-miR-6845-5p” used herein refers to the hsa-miR-6845-5p gene (miRBase Accession No. MIMAT0027590) and other species such as homologs or orthologs.
- the hsa-miR-6845-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6845-5p", “hsa-mir-6845” (miRBase Accession No. MI0022691, SEQ ID NO: 250) having a hairpin-like structure is known.
- hsa-miR-4327 gene or “hsa-miR-4327” refers to the hsa-miR-4327 gene (miRBase Accession No. MIMAT00168889) set forth in SEQ ID NO: 49 and other species. Includes homologs or orthologs.
- the hsa-miR-4327 gene can be obtained by the method described in Goff LA et al., 2009, PLos One, Volume 4, e7192. Further, as the precursor of "hsa-miR-4327", “hsa-mir-4327” (miRBase Accession No. MI0015867, SEQ ID NO: 251) having a hairpin-like structure is known.
- hsa-miR-4723-5p gene or "hsa-miR-4723-5p” refers to the hsa-miR-4723-5p gene (miRBase Accession No. MIMAT0019838) and other species such as homologs or orthologs are included.
- the hsa-miR-4723-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4723-5p", “hsa-mir-4723” (miRBase Accession No. MI0017359, SEQ ID NO: 252) having a hairpin-like structure is known.
- hsa-miR-4530 gene or “hsa-miR-4530” refers to the hsa-miR-4530 gene (miRBase Accession No. MIMAT0019069) set forth in SEQ ID NO: 51 and other species. Includes homologs or orthologs.
- the hsa-miR-4530 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4530", “hsa-mir-4530” (miRBase Accession No. MI0016897, SEQ ID NO: 253) having a hairpin-like structure is known.
- hsa-miR-6771-5p gene or "hsa-miR-6771-5p” refers to the hsa-miR-6771-5p gene (miRBase Accession No.) set forth in SEQ ID NO: 52. Includes MIMAT0027442) and other species homologs or orthologs.
- the hsa-miR-6771-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6771-5p", “hsa-mir-6771” (miRBase Accession No. MI0022616, SEQ ID NO: 254) having a hairpin-like structure is known.
- hsa-miR-614 gene or “hsa-miR-614" refers to the hsa-miR-614 gene (miRBase Accession No. MIMAT0003282) and other species set forth in SEQ ID NO: 53. Includes homologs or orthologs.
- the hsa-miR-614 gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692. Further, as the precursor of "hsa-miR-614", “hsa-mir-614" (miRBase Accession No. MI0003627, SEQ ID NO: 255) having a hairpin-like structure is known.
- hsa-miR-92a-2-5p gene or "hsa-miR-92a-2-5p” is described in SEQ ID NO: 54, hsa-miR-92a-2-5p. Includes genes (miRBase Accession No. MIMAT0004508) and other species homologs or orthologs.
- the hsa-miR-92a-2-5p gene can be obtained by the method described in Maurelatos Z et al., 2002, Genes Dev, Vol. 16, p720-728. Further, as the precursor of "hsa-miR-92a-2-5p", “hsa-mir-92a-2” (miRBase Accession No. MI00000094, SEQ ID NO: 256) having a hairpin-like structure is known.
- hsa-miR-6891-5p gene or "hsa-miR-6891-5p” used herein refers to the hsa-miR-6891-5p gene (miRBase Accession No. Includes MIMAT0027682) and other species homologs or orthologs.
- the hsa-miR-6891-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6891-5p", "hsa-mir-6891” (miRBase Accession No. MI0022738, SEQ ID NO: 257) having a hairpin-like structure is known.
- hsa-miR-6124 gene or “hsa-miR-6124” refers to the hsa-miR-6124 gene (miRBase Accession No. MIMAT00245957) and other species set forth in SEQ ID NO: 56. Includes homologs or orthologs.
- the hsa-miR-6124 gene can be obtained by the method described in Smith JL et al., 2012, J. Virol, Vol. 86, p5278-5287. Further, as the precursor of "hsa-miR-6124", “hsa-mir-6124" (miRBase Accession No. MI0021258, SEQ ID NO: 258) having a hairpin-like structure is known.
- hsa-miR-4678-3p gene or "hsa-miR-4678-3p” used herein refer to the hsa-miR-4678-3p gene (miRBase Accession No.) set forth in SEQ ID NO: 57. MIMAT0019775) and other species such as homologs or orthologs are included.
- the hsa-miR-4678-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4678-3p", “hsa-mir-4678” (miRBase Accession No. MI0017319, SEQ ID NO: 259) having a hairpin-like structure is known.
- hsa-miR-4442 gene or “hsa-miR-4442” refers to the hsa-miR-4442 gene (miRBase Accession No. MIMAT0018960) and other species set forth in SEQ ID NO: 58. Includes homologs or orthologs.
- the hsa-miR-4442 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4442", “hsa-mir-4442” (miRBase Accession No. MI0016785, SEQ ID NO: 260) having a hairpin-like structure is known.
- hsa-miR-7977 gene or “hsa-miR-7977” refers to the hsa-miR-7977 gene (miRBase Accession No. MIMAT0031180) and other species set forth in SEQ ID NO: 59. Includes homologs or orthologs.
- the hsa-miR-7977 gene can be obtained by the method described in Verthut-Meikas A et al., 2013, Mol Endocrinol, online version. Further, as the precursor of "hsa-miR-7977", “hsa-mir-7977” (miRBase Accession No. MI0025753, SEQ ID NO: 261) having a hairpin-like structure is known.
- hsa-miR-6785-5p gene or "hsa-miR-6785-5p” refers to the hsa-miR-6785-5p gene (miRBase Accession No. MIMAT0027470) and other species such as homologs or orthologs are included.
- the hsa-miR-6785-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6785-5p", “hsa-mir-6785” (miRBase Accession No. MI0022630, SEQ ID NO: 262) having a hairpin-like structure is known.
- hsa-miR-4497 gene or “hsa-miR-4497” refers to the hsa-miR-4497 gene (miRBase Accession No. MIMAT0019032) set forth in SEQ ID NO: 61 and other species. Includes homologs or orthologs.
- the hsa-miR-4497 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4497", “hsa-mir-4497” (miRBase Accession No. MI0016859, SEQ ID NO: 263) having a hairpin-like structure is known.
- hsa-miR-8071 gene or “hsa-miR-8071” refers to the hsa-miR-8071 gene (miRBase Accession No. MIMAT0030998) set forth in SEQ ID NO: 62 and other species. Includes homologs or orthologs.
- the hsa-miR-8071 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487.
- hsa-miR-8071 has a hairpin-like structure as a precursor thereof, "hsa-mir-8071-1, hsa-mir-8071-2” (miRBase Accession No. MI0025907, MI0026417, SEQ ID NO: 264, 265) is known.
- hsa-miR-663b gene or “hsa-miR-663b” refers to the hsa-miR-663b gene (miRBase Accession No. MIMAT0005867) set forth in SEQ ID NO: 63 and other species. Includes homologs or orthologs.
- the hsa-miR-663b gene can be obtained by the method described in Takada S et al., 2008, Leukemia, Vol. 22, p1274-1278. Further, as the precursor of "hsa-miR-663b", “hsa-mir-663b” (miRBase Accession No. MI0006336, SEQ ID NO: 266) having a hairpin-like structure is known.
- hsa-miR-3180 gene or “hsa-miR-3180” refers to the hsa-miR-3180 gene (miRBase Accession No. MIMAT0018178) set forth in SEQ ID NO: 64 and other species. Includes homologs or orthologs.
- the hsa-miR-3180 gene can be obtained by the method described in Creighton CJ et al., 2010, PLos One, Volume 5, e9637.
- "hsa-miR-3180” has a hairpin-like structure as a precursor thereof, "hsa-mir-3180-4, hsa-mir-3180-5" (miRBase Accession No. MI0016408, MI0016409, SEQ ID NO: 266, 268) is known.
- hsa-miR-4251 gene or “hsa-miR-4251” refers to the hsa-miR-4251 gene (miRBase Accession No. MIMAT0016883) set forth in SEQ ID NO: 65 and other species. Includes homologs or orthologs.
- the hsa-miR-4251 gene can be obtained by the method described in Goff LA et al., 2009, PLos One, Volume 4, e7192. Further, as the precursor of "hsa-miR-4251", “hsa-mir-4251” (miRBase Accession No. MI0015861, SEQ ID NO: 269) having a hairpin-like structure is known.
- hsa-miR-1285-3p gene or "hsa-miR-1285-3p” used herein refer to the hsa-miR-1285-3p gene (miRBase Accession No. Includes MIMAT0005876) and other species homologs or orthologs.
- the hsa-miR-1285-3p gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, Vol. 18, p610-621.
- “hsa-miR-1285-3p” has a hairpin-like structure as a precursor thereof, "hsa-mir-1285-1, hsa-mir-1285-2” (miRBase Accession No. MI0006346, MI0006347, SEQ ID NO: 270, 271) are known.
- hsa-miR-6870-5p gene or "hsa-miR-6870-5p” used herein refers to the hsa-miR-6870-5p gene (miRBase Accession No. MIMAT0027640) and other species such as homologs or orthologs.
- the hsa-miR-6870-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6870-5p", “hsa-mir-6870” (miRBase Accession No. MI0022717, SEQ ID NO: 272) having a hairpin-like structure is known.
- hsa-miR-4484 gene or "hsa-miR-4484" refers to the hsa-miR-4484 gene (miRBase Accession No. MIMAT0019018) and other species set forth in SEQ ID NO: 68. Includes homologs or orthologs.
- the hsa-miR-4484 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4484", "hsa-mir-4484" (miRBase Accession No. MI0016845, SEQ ID NO: 273) having a hairpin-like structure is known.
- hsa-miR-4476 gene or “hsa-miR-4476” refers to the hsa-miR-4476 gene (miRBase Accession No. MIMAT0019003) set forth in SEQ ID NO: 69 and other species. Includes homologs or orthologs.
- the hsa-miR-4476 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4476", “hsa-mir-4476” (miRBase Accession No. MI0016828, SEQ ID NO: 274) having a hairpin-like structure is known.
- hsa-miR-6794-5p gene or "hsa-miR-6794-5p” refers to the hsa-miR-6794-5p gene (miRBase Accession No.) set forth in SEQ ID NO: 70. MIMAT0027398) and other species such as homologs or orthologs.
- the hsa-miR-6794-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, “hsa-miR-6794-5p” has a hairpin-like structure as a precursor thereof, and "hsa-mir-6949" (miRBase Accession No. MI0022594, SEQ ID NO: 275) is known.
- hsa-miR-4454 gene or "hsa-miR-4454" refers to the hsa-miR-4454 gene (miRBase Accession No. MIMAT0018976) set forth in SEQ ID NO: 71 and other species. Includes homologs or orthologs.
- the hsa-miR-4454 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4454", “hsa-mir-4454" (miRBase Accession No. MI0016800, SEQ ID NO: 276) having a hairpin-like structure is known.
- hsa-miR-6893-5p gene or "hsa-miR-6893-5p” refers to the hsa-miR-6893-5p gene (miRBase Accession No.) set forth in SEQ ID NO: 72. Includes MIMAT0027686) and other species homologs or orthologs.
- the hsa-miR-6893-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6893-5p", “hsa-mir-6893” (miRBase Accession No. MI0022740, SEQ ID NO: 277) having a hairpin-like structure is known.
- hsa-miR-6085 gene or “hsa-miR-6085” refers to the hsa-miR-6085 gene (miRBase Accession No. MIMAT0023710) and other species set forth in SEQ ID NO: 73. Includes homologs or orthologs.
- the hsa-miR-6085 gene can be obtained by the method described in Voellenkle C et al., 2012, RNA, Vol. 18, p472-484. Further, as the precursor of "hsa-miR-6085", “hsa-mir-6085” (miRBase Accession No. MI0020362, SEQ ID NO: 278) having a hairpin-like structure is known.
- hsa-miR-4787-5p gene or "hsa-miR-4787-5p” refers to the hsa-miR-4787-5p gene (miRBase Accession No. MIMAT0019956) and other species such as homologs or orthologs are included.
- the hsa-miR-4787-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4787-5p", “hsa-mir-4787” (miRBase Accession No. MI0017434, SEQ ID NO: 279) having a hairpin-like structure is known.
- hsa-miR-149-3p gene or "hsa-miR-149-3p” refers to the hsa-miR-149-3p gene (miRBase Accession No. Includes MIMAT0004609) and other species homologs or orthologs.
- the hsa-miR-149-3p gene can be obtained by the method described in Lagos-Quintana M et al., 2002, Curr Biol, Vol. 12, p735-739. Further, as a precursor of "hsa-miR-149-3p", "hsa-mir-149” (miRBase Accession No. MI0000478, SEQ ID NO: 280) having a hairpin-like structure is known.
- hsa-miR-7704 gene or "hsa-miR-7704" refers to the hsa-miR-7704 gene (miRBase Accession No. MIMAT0030019) set forth in SEQ ID NO: 76 and other species. Includes homologs or orthologs.
- the hsa-miR-7704 gene can be obtained by the method described in Swamisan S et al., 2013, Biochem Biophyss Res Commun, Vol. 434, p228-234. Further, as the precursor of "hsa-miR-7704", "hsa-mir-7704" (miRBase Accession No. MI0025240, SEQ ID NO: 281) having a hairpin-like structure is known.
- hsa-miR-6125 gene or “hsa-miR-6125” refers to the hsa-miR-6125 gene (miRBase Accession No. MIMAT0024598) set forth in SEQ ID NO: 77 and other species. Includes homologs or orthologs.
- the hsa-miR-6125 gene can be obtained by the method described in Smith JL et al., 2012, J. Virol, Vol. 86, p5278-5287. Further, as the precursor of "hsa-miR-6125", “hsa-mir-6125” (miRBase Accession No. MI0021259, SEQ ID NO: 282) having a hairpin-like structure is known.
- hsa-miR-6090 gene or “hsa-miR-6090” refers to the hsa-miR-6090 gene (miRBase Accession No. MIMAT0023715) and other species set forth in SEQ ID NO: 78. Includes homologs or orthologs.
- the hsa-miR-6090 gene can be obtained by the method described in Yoo JK et al., 2012, Stem Cells Dev, Vol. 21, p2049-2057. Further, as the precursor of "hsa-miR-6090", “hsa-mir-6090” (miRBase Accession No. MI0020367, SEQ ID NO: 283) having a hairpin-like structure is known.
- hsa-miR-3197 gene or “hsa-miR-3197” refers to the hsa-miR-3197 gene (miRBase Accession No. MIMAT0015082) set forth in SEQ ID NO: 79 and other species. Includes homologs or orthologs.
- the hsa-miR-3197 gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-3197", “hsa-mir-3197" (miRBase Accession No. MI0014245, SEQ ID NO: 284) having a hairpin-like structure is known.
- hsa-miR-6850-5p gene or "hsa-miR-6850-5p” refers to the hsa-miR-6850-5p gene (miRBase Accession No. MIMAT0027600) and other species such as homologs or orthologs.
- the hsa-miR-6850-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6850-5p", “hsa-mir-6850” (miRBase Accession No. MI0022696, SEQ ID NO: 285) having a hairpin-like structure is known.
- hsa-miR-4467 gene or “hsa-miR-4467” refers to the hsa-miR-4467 gene (miRBase Accession No. MIMAT0018994) set forth in SEQ ID NO: 81 and other species. Includes homologs or orthologs.
- the hsa-miR-4467 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4467", “hsa-mir-4467” (miRBase Accession No. MI0016818, SEQ ID NO: 286) having a hairpin-like structure is known.
- hsa-miR-6885-5p gene or "hsa-miR-6885-5p” refers to the hsa-miR-6885-5p gene (miRBase Accession No.) set forth in SEQ ID NO: 82. MIMAT0027670) and other species such as homologs or orthologs.
- the hsa-miR-6885-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6885-5p", “hsa-mir-6885” (miRBase Accession No. MI0022732, SEQ ID NO: 287) having a hairpin-like structure is known.
- hsa-miR-6803-5p gene or "hsa-miR-6803-5p” refers to the hsa-miR-6803-5p gene (miRBase Accession No. MIMAT0027506) and other species such as homologs or orthologs are included.
- the hsa-miR-6803-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6803-5p", “hsa-mir-6803" (miRBase Accession No. MI0022648, SEQ ID NO: 288) having a hairpin-like structure is known.
- hsa-miR-6798-5p gene or "hsa-miR-6798-5p” refers to the hsa-miR-6798-5p gene (miRBase Accession No. MIMAT0027496) and other species such as homologs or orthologs are included.
- the hsa-miR-6798-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6798-5p", “hsa-mir-6798” (miRBase Accession No. MI0022643, SEQ ID NO: 289) having a hairpin-like structure is known.
- hsa-miR-6780b-5p gene or "hsa-miR-6780b-5p” used herein refers to the hsa-miR-6780b-5p gene (miRBase Accession No. Includes MIMAT0027572) and other species homologs or orthologs.
- the hsa-miR-6780b-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, “hsa-miR-6780b-5p” has a hairpin-like structure as a precursor thereof, and "hsa-mir-6780b” (miRBase Accession No. MI0022681, SEQ ID NO: 290) is known.
- hsa-miR-6768-5p gene or "hsa-miR-6768-5p” refers to the hsa-miR-6768-5p gene (miRBase Accession No. MIMAT0027436) and other species such as homologs or orthologs.
- the hsa-miR-6768-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6768-5p", “hsa-mir-6768” (miRBase Accession No. MI0022613, SEQ ID NO: 291) having a hairpin-like structure is known.
- hsa-miR-5100 gene or “hsa-miR-5100” refers to the hsa-miR-5100 gene (miRBase Accession No. MIMAT0022259) set forth in SEQ ID NO: 87 and other species. Includes homologs or orthologs.
- the hsa-miR-5100 gene can be obtained by the method described in Tandon M et al., 2012, Oral Dis, Vol. 18, p127-131. Further, as the precursor of "hsa-miR-5100", “hsa-mir-5100” (miRBase Accession No. MI0019116, SEQ ID NO: 292) having a hairpin-like structure is known.
- hsa-miR-6724-5p gene or "hsa-miR-6724-5p” refers to the hsa-miR-6724-5p gene (miRBase Accession No. MIMAT0025856) and other species such as homologs or orthologs.
- the hsa-miR-6724-5p gene can be obtained by the method described in Li Y et al., 2012, Gene, Vol. 497, p330-335.
- hsa-miR-6724-5p has a hairpin-like structure as a precursor thereof, "hsa-mir-6724-1, hsa-mir-6724-2, hsa-mir-6724-3, hsa-mir”.
- -6724-4 "(miRBase Accession No. MI0022559, MI0031516, MI0031517, MI0031518, SEQ ID NO: 293, 294, 295, 296) is known.
- hsa-miR-6879-5p gene or "hsa-miR-6879-5p” refers to the hsa-miR-6879-5p gene (miRBase Accession No. MIMAT0027658) and other species such as homologs or orthologs are included.
- the hsa-miR-6879-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6879-5p", “hsa-mir-6879” (miRBase Accession No. MI0022726, SEQ ID NO: 297) having a hairpin-like structure is known.
- hsa-miR-7108-5p gene or "hsa-miR-7108-5p” refers to the hsa-miR-7108-5p gene (miRBase Accession No. MIMAT0028113) and other species such as homologs or orthologs are included.
- the hsa-miR-7108-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, “hsa-miR-7108-5p” has a hairpin-like structure as a precursor thereof, and "hsa-mir-7108" (miRBase Accession No. MI0022959, SEQ ID NO: 298) is known.
- hsa-miR-4649-5p gene or "hsa-miR-4649-5p” refers to the hsa-miR-4649-5p gene (miRBase Accession No. Includes MIMAT0019711) and other species homologs or orthologs.
- the hsa-miR-4649-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4649-5p", “hsa-mir-4649” (miRBase Accession No. MI0017276, SEQ ID NO: 299) having a hairpin-like structure is known.
- hsa-miR-4739 gene or "hsa-miR-4739” refers to the hsa-miR-4739 gene (miRBase Accession No. MIMAT0019868) set forth in SEQ ID NO: 92 and other species. Includes homologs or orthologs.
- the hsa-miR-4739 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4739", “hsa-mir-4739” (miRBase Accession No. MI0017377, SEQ ID NO: 300) having a hairpin-like structure is known.
- hsa-miR-6089 gene or “hsa-miR-6089” refers to the hsa-miR-6089 gene (miRBase Accession No. MIMAT0023714) set forth in SEQ ID NO: 93 and other species. Includes homologs or orthologs.
- the hsa-miR-6089 gene can be obtained by the method described in Yoo JK et al., 2012, Stem Cells Dev, Vol. 21, p2049-2057.
- hsa-miR-6089 has a hairpin-like structure as a precursor thereof, "hsa-mir-6089-1, hsa-mir-6089-2” (miRBase Accession No. MI0020366, MI0023563, SEQ ID NO: 301, 302) is known.
- hsa-miR-1908-5p gene or "hsa-miR-1908-5p” refers to the hsa-miR-1908-5p gene (miRBase Accession No. Includes MIMAT0007881) and other species homologs or orthologs.
- the hsa-miR-1908-5p gene can be obtained by the method described in Bar M et al., 2008, Stem Cells, Vol. 26, p2496-2505. Further, as the precursor of "hsa-miR-1908-5p", "hsa-mir-1908" (miRBase Accession No. MI0008329, SEQ ID NO: 303) having a hairpin-like structure is known.
- hsa-miR-4516 gene or “hsa-miR-4516” refers to the hsa-miR-4516 gene (miRBase Accession No. MIMAT0019053) set forth in SEQ ID NO: 95 and other species. Includes homologs or orthologs.
- the hsa-miR-4516 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4516", “hsa-mir-4516” (miRBase Accession No. MI0016882, SEQ ID NO: 304) having a hairpin-like structure is known.
- hsa-miR-2861 gene or “hsa-miR-2861” refers to the hsa-miR-2861 gene (miRBase Accession No. MIMAT0013802) set forth in SEQ ID NO: 96 and other species. Includes homologs or orthologs.
- the hsa-miR-2861 gene can be obtained by the method described in Li H et al., 2009, J Clin Invest, 119, p3666-3677. Further, as the precursor of "hsa-miR-2861", “hsa-mir-2861” (miRBase Accession No. MI0013006, SEQ ID NO: 305) having a hairpin-like structure is known.
- hsa-miR-4492 gene or “hsa-miR-4492” refers to the hsa-miR-4492 gene (miRBase Accession No. MIMAT0019027) set forth in SEQ ID NO: 97 and other species. Includes homologs or orthologs.
- the hsa-miR-4492 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4492", “hsa-mir-4492” (miRBase Accession No. MI0016854, SEQ ID NO: 306) having a hairpin-like structure is known.
- hsa-miR-4294 gene or "hsa-miR-4294" refers to the hsa-miR-4294 gene (miRBase Accession No. MIMAT0016849) set forth in SEQ ID NO: 98 and other species. Includes homologs or orthologs.
- the hsa-miR-4294 gene can be obtained by the method described in Goff LA et al., 2009, PLos One, Volume 4, e7192. Further, as the precursor of "hsa-miR-4294", “hsa-mir-4294" (miRBase Accession No. MI0015827, SEQ ID NO: 307) having a hairpin-like structure is known.
- hsa-miR-6791-5p gene or "hsa-miR-6791-5p” used herein refers to the hsa-miR-6791-5p gene (miRBase Accession No. Includes MIMAT0027482) and other species homologs or orthologs.
- the hsa-miR-6791-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6791-5p", "hsa-mir-6791” (miRBase Accession No. MI0022636, SEQ ID NO: 308) having a hairpin-like structure is known.
- hsa-miR-1469 gene or “hsa-miR-1469” refers to the hsa-miR-1469 gene (miRBase Accession No. MIMAT0007347) and other species set forth in SEQ ID NO: 100. Includes homologs or orthologs.
- the hsa-miR-1469 gene can be obtained by the method described in Kawaji H et al., 2008, BMC Genomics, Vol. 9, p157. Further, as the precursor of "hsa-miR-1469", “hsa-mir-1469” (miRBase Accession No. MI0007074, SEQ ID NO: 309) having a hairpin-like structure is known.
- hsa-miR-6752-5p gene or "hsa-miR-6752-5p” used herein refers to the hsa-miR-6752-5p gene (miRBase Accession No. 1) set forth in SEQ ID NO: 101. MIMAT0027404) and other species such as homologs or orthologs.
- the hsa-miR-6752-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6752-5p", “hsa-mir-6752” (miRBase Accession No. MI0022595, SEQ ID NO: 310) having a hairpin-like structure is known.
- hsa-miR-4730 gene or “hsa-miR-4730” refers to the hsa-miR-4730 gene (miRBase Accession No. MIMAT0019852) set forth in SEQ ID NO: 102 and other species. Includes homologs or orthologs.
- the hsa-miR-4730 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4730", “hsa-mir-4730” (miRBase Accession No. MI0017367, SEQ ID NO: 311) having a hairpin-like structure is known.
- hsa-miR-6126 gene or “hsa-miR-6126” refers to the hsa-miR-6126 gene (miRBase Accession No. MIMAT0024599) set forth in SEQ ID NO: 103 and other species. Includes homologs or orthologs.
- the hsa-miR-6126 gene can be obtained by the method described in Smith JL et al., 2012, J. Virol, Vol. 86, p5278-5287. Further, as the precursor of "hsa-miR-6126", “hsa-mir-6126” (miRBase Accession No. MI0021260, SEQ ID NO: 312) having a hairpin-like structure is known.
- hsa-miR-6869-5p gene or "hsa-miR-6869-5p” refers to the hsa-miR-6869-5p gene (miRBase Accession No. MIMAT0027638) and other species such as homologs or orthologs are included.
- the hsa-miR-6869-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6869-5p", “hsa-mir-6869” (miRBase Accession No. MI0022716, SEQ ID NO: 313) having a hairpin-like structure is known.
- hsa-miR-1268a gene or “hsa-miR-1268a” refers to the hsa-miR-1268a gene (miRBase Accession No. MIMAT0005922) set forth in SEQ ID NO: 105 and other species. Includes homologs or orthologs.
- the hsa-miR-1268a gene can be obtained by the method described in Morin RD et al., 2008, Genome Res, Vol. 18, p610-621. Further, as the precursor of "hsa-miR-1268a", “hsa-mir-1268a” (miRBase Accession No. MI0006405, SEQ ID NO: 314) having a hairpin-like structure is known.
- hsa-miR-6799-5p gene or "hsa-miR-6799-5p” refers to the hsa-miR-6799-5p gene (miRBase Accession No. MIMAT0027498) and other species such as homologs or orthologs are included.
- the hsa-miR-6799-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6799-5p", “hsa-mir-6799” (miRBase Accession No. MI0022644, SEQ ID NO: 315) having a hairpin-like structure is known.
- hsa-miR-8069 gene or “hsa-miR-8069” refers to the hsa-miR-8069 gene (miRBase Accession No. MIMAT0030996) set forth in SEQ ID NO: 107 and other species. Includes homologs or orthologs.
- the hsa-miR-8069 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487.
- hsa-miR-8069 has a hairpin-like structure as a precursor thereof, "hsa-mir-8069-1, hsa-mir-8069-2” (miRBase Accession No. MI0025905, MI0031518, SEQ ID NO: 316, 317) is known.
- hsa-miR-3621 gene or “hsa-miR-3621” refers to the hsa-miR-3621 gene (miRBase Accession No. MIMAT0018002) set forth in SEQ ID NO: 108 and other species. Includes homologs or orthologs.
- the hsa-miR-3621 gene can be obtained by the method described in Witten D et al., 2010, BMC Biol, Volume 8, p58. Further, as the precursor of "hsa-miR-3621", “hsa-mir-3621” (miRBase Accession No. MI0016012, SEQ ID NO: 318) having a hairpin-like structure is known.
- hsa-miR-4763-3p gene or "hsa-miR-4763-3p” used herein refer to the hsa-miR-4763-3p gene (miRBase Accession No. MIMAT0019913) and other species such as homologs or orthologs are included.
- the hsa-miR-4763-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4763-3p", “hsa-mir-4763” (miRBase Accession No. MI0017404, SEQ ID NO: 319) having a hairpin-like structure is known.
- hsa-miR-1228-5p gene or "hsa-miR-1228-5p” refers to the hsa-miR-1228-5p gene (miRBase Accession No. Includes MIMAT0005582) and other species homologs or orthologs.
- the hsa-miR-1228-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, Vol. 28, p328-336. Further, as the precursor of "hsa-miR-1228-5p", "hsa-mir-1228” (miRBase Accession No. MI0006318, SEQ ID NO: 320) having a hairpin-like structure is known.
- hsa-miR-760 gene or “hsa-miR-760” refers to the hsa-miR-760 gene (miRBase Accession No. MIMAT0004957) set forth in SEQ ID NO: 111 and other species. Includes homologs or orthologs.
- the hsa-miR-760 gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, Vol. 16, pp. 1289-1298. Further, as the precursor of "hsa-miR-760", “hsa-mir-760” (miRBase Accession No. MI0005567, SEQ ID NO: 321) having a hairpin-like structure is known.
- hsa-miR-187-5p gene or "hsa-miR-187-5p” refers to the hsa-miR-187-5p gene (miRBase Accession No. Includes MIMAT0004561) and other species homologs or orthologs.
- the hsa-miR-187-5p gene can be obtained by the method described in Lim LP et al., 2003, Science, Vol. 299, p1540. Further, as the precursor of "hsa-miR-187-5p", “hsa-mir-187” (miRBase Accession No. MI0000274, SEQ ID NO: 322) having a hairpin-like structure is known.
- hsa-miR-7111-5p gene or "hsa-miR-7111-5p” refers to the hsa-miR-7111-5p gene (miRBase Accession No. Includes MIMAT0028119) and other species homologs or orthologs.
- the hsa-miR-7111-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-7111-5p", "hsa-mir-7111” (miRBase Accession No. MI0022962, SEQ ID NO: 323) having a hairpin-like structure is known.
- hsa-miR-6088 gene or “hsa-miR-6088” refers to the hsa-miR-6088 gene (miRBase Accession No. MIMAT0023713) set forth in SEQ ID NO: 114 and other species. Includes homologs or orthologs.
- the hsa-miR-6088 gene can be obtained by the method described in Yoo JK et al., 2012, Stem Cells Dev, Vol. 21, p2049-2057. Further, as the precursor of "hsa-miR-6088", “hsa-mir-6088” (miRBase Accession No. MI0020365, SEQ ID NO: 324) having a hairpin-like structure is known.
- hsa-miR-6805-3p gene or "hsa-miR-6805-3p” refers to the hsa-miR-6805-3p gene (miRBase Accession No. Includes MIMAT0027511) and other species homologs or orthologs.
- the hsa-miR-6805-3p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6805-3p", "hsa-mir-6805" (miRBase Accession No. MI0022650, SEQ ID NO: 325) having a hairpin-like structure is known.
- hsa-miR-4640-5p gene or "hsa-miR-4640-5p” refers to the hsa-miR-4640-5p gene (miRBase Accession No. MIMAT0019699) and other species such as homologs or orthologs are included.
- the hsa-miR-4640-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4640-5p", “hsa-mir-4640” (miRBase Accession No. MI0017267, SEQ ID NO: 326) having a hairpin-like structure is known.
- hsa-miR-6721-5p gene or "hsa-miR-6721-5p” refers to the hsa-miR-6721-5p gene (miRBase Accession No. MIMAT0025852) and other species such as homologs or orthologs are included.
- the hsa-miR-6721-5p gene can be obtained by the method described in Li Y et al., 2012, Gene, Vol. 497, p330-335. Further, as the precursor of "hsa-miR-6721-5p", “hsa-mir-6721” (miRBase Accession No. MI0022556, SEQ ID NO: 327) having a hairpin-like structure is known.
- hsa-miR-6880-5p gene or "hsa-miR-6880-5p” refers to the hsa-miR-6880-5p gene (miRBase Accession No. MIMAT0027660) and other species such as homologs or orthologs.
- the hsa-miR-6880-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6880-5p", “hsa-mir-6880” (miRBase Accession No. MI0022727, SEQ ID NO: 328) having a hairpin-like structure is known.
- hsa-miR-711 gene or “hsa-miR-711” refers to the hsa-miR-711 gene (miRBase Accession No. MIMAT0012734) set forth in SEQ ID NO: 119 and other species. Includes homologs or orthologs.
- the hsa-miR-711 gene can be obtained by the method described in Artzi S et al., 2008, BMC Bioinformatics, Volume 9, p39. Further, as the precursor of "hsa-miR-711", “hsa-mir-711” (miRBase Accession No. MI0012488, SEQ ID NO: 329) having a hairpin-like structure is known.
- hsa-miR-128-1-5p gene or "hsa-miR-128-1-5p” is described in SEQ ID NO: 120, hsa-miR-128-1-5p. Includes genes (miRBase Accession No. MIMAT0026477) and other species homologs or orthologs.
- the hsa-miR-128-1-5p gene can be obtained by the method described in Lagos-Quintana M et al., 2002, Curr Biol, Vol. 12, p735-739. Further, as the precursor of "hsa-miR-128-1-5p", “hsa-mir-128-1” (miRBase Accession No. MI0000447, SEQ ID NO: 330) having a hairpin-like structure is known.
- hsa-miR-4525 gene or "hsa-miR-4525” refers to the hsa-miR-4525 gene (miRBase Accession No. MIMAT00190604) set forth in SEQ ID NO: 121 and other species. Includes homologs or orthologs.
- the hsa-miR-4525 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4525", “hsa-mir-4525” (miRBase Accession No. MI0016892, SEQ ID NO: 331) having a hairpin-like structure is known.
- hsa-miR-486-3p gene or "hsa-miR-486-3p” refers to the hsa-miR-486-3p gene (miRBase Accession No. Includes MIMAT0004762) and other species homologs or orthologs.
- the hsa-miR-486-3p gene can be obtained by the method described in Fu H et al., 2005, FEBS Lett, Vol. 579, p3849-3854.
- “hsa-miR-486-3p” has a hairpin-like structure as a precursor thereof, "hsa-mir-486-1, hsa-mir-486-2” (miRBase Accession No. MI0002470, MI0023622, SEQ ID NO: 332,333) are known.
- hsa-miR-6756-5p gene or "hsa-miR-6756-5p” as used herein refers to the hsa-miR-6756-5p gene (miRBase Accession No. Includes MIMAT0027412) and other species homologs or orthologs.
- the hsa-miR-6756-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6756-5p", “hsa-mir-6756” (miRBase Accession No. MI0022601, SEQ ID NO: 334) having a hairpin-like structure is known.
- hsa-miR-1260b gene or “hsa-miR-1260b” refers to the hsa-miR-1260b gene (miRBase Accession No. MIMAT0015041) set forth in SEQ ID NO: 124 and other species. Includes homologs or orthologs.
- the hsa-miR-1260b gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-1260b", “hsa-mir-1260b” (miRBase Accession No. MI0014197, SEQ ID NO: 335) having a hairpin-like structure is known.
- hsa-miR-3184-5p gene or "hsa-miR-3184-5p” refers to the hsa-miR-3184-5p gene (miRBase Accession No. MIMAT0015064) and other species such as homologs or orthologs.
- the hsa-miR-3184-5p gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685.
- “hsa-mir-3184" as the precursor of "hsa-miR-3184-5p
- hsa-mir-3184 having a hairpin-like structure is known.
- hsa-miR-6075 gene or “hsa-miR-6075” refers to the hsa-miR-6075 gene (miRBase Accession No. MIMAT0023700) and other species set forth in SEQ ID NO: 126. Includes homologs or orthologs.
- the hsa-miR-6075 gene can be obtained by the method described in Voellenkle C et al., 2012, RNA, Vol. 18, p472-484. Further, as the precursor of "hsa-miR-6075", “hsa-mir-6075” (miRBase Accession No. MI0020352, SEQ ID NO: 337) having a hairpin-like structure is known.
- hsa-miR-204-3p gene or "hsa-miR-204-3p” refers to the hsa-miR-204-3p gene (miRBase Accession No. Includes MIMAT0022693) and other species homologs or orthologs.
- the hsa-miR-204-3p gene can be obtained by the method described in Lim LP et al., 2003, Science, Volume 299, p1540. Further, as a precursor of "hsa-miR-204-3p", "hsa-mir-204" (miRBase Accession No. MI00000284, SEQ ID NO: 338) having a hairpin-like structure is known.
- hsa-miR-4728-5p gene or "hsa-miR-4728-5p” refers to the hsa-miR-4728-5p gene (miRBase Accession No. MIMAT0019849) and other species such as homologs or orthologs are included.
- the hsa-miR-4728-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4728-5p", “hsa-mir-4728” (miRBase Accession No. MI0017365, SEQ ID NO: 339) having a hairpin-like structure is known.
- hsa-miR-4534 gene or “hsa-miR-4534” refers to the hsa-miR-4534 gene (miRBase Accession No. MIMAT0019073) set forth in SEQ ID NO: 129 and other species. Includes homologs or orthologs.
- the hsa-miR-4534 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4534", “hsa-mir-4534” (miRBase Accession No. MI0016901, SEQ ID NO: 340) having a hairpin-like structure is known.
- hsa-miR-4758-5p gene or "hsa-miR-4758-5p” refers to the hsa-miR-4758-5p gene (miRBase Accession No. Includes MIMAT0019903) and other species homologs or orthologs.
- the hsa-miR-4758-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4758-5p", “hsa-mir-4758” (miRBase Accession No. MI0017399, SEQ ID NO: 341) having a hairpin-like structure is known.
- hsa-miR-8063 gene or “hsa-miR-8063” refers to the hsa-miR-8063 gene (miRBase Accession No. MIMAT0030990) set forth in SEQ ID NO: 131 and other species. Includes homologs or orthologs.
- the hsa-miR-8063 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487. Further, as the precursor of "hsa-miR-8063", “hsa-mir-8063” (miRBase Accession No. MI0025899, SEQ ID NO: 342) having a hairpin-like structure is known.
- hsa-miR-6863-3p gene or "hsa-miR-6863-3p” refers to the hsa-miR-6863-3p gene (miRBase Accession No. MIMAT0027575) and other species such as homologs or orthologs.
- the hsa-miR-6863-3p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6863-3p", “hsa-mir-6863” (miRBase Accession No. MI0022682, SEQ ID NO: 343) having a hairpin-like structure is known.
- hsa-miR-6789-5p gene or "hsa-miR-6789-5p” refers to the hsa-miR-6789-5p gene (miRBase Accession No. MIMAT0027478) and other species such as homologs or orthologs are included.
- the hsa-miR-6789-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6789-5p", “hsa-mir-6789” (miRBase Accession No. MI0022634, SEQ ID NO: 344) having a hairpin-like structure is known.
- hsa-miR-744-5p gene or "hsa-miR-744-5p” refers to the hsa-miR-744-5p gene (miRBase Accession No. MIMAT0004945) and other species such as homologs or orthologs.
- the hsa-miR-744-5p gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, Vol. 16, p1289-1298. Further, as the precursor of "hsa-miR-744-5p", "hsa-mir-744" (miRBase Accession No. MI0005559, SEQ ID NO: 345) having a hairpin-like structure is known.
- hsa-miR-1909-3p gene or "hsa-miR-1909-3p” used herein refer to the hsa-miR-1909-3p gene (miRBase Accession No. Includes MIMAT0007883) and other species homologs or orthologs.
- the hsa-miR-1909-3p gene can be obtained by the method described in Bar M et al., 2008, Stem Cells, Vol. 26, p2496-2505. Further, as the precursor of "hsa-miR-1909-3p", “hsa-mir-1909” (miRBase Accession No. MI0008330, SEQ ID NO: 346) having a hairpin-like structure is known.
- hsa-miR-887-3p gene or "hsa-miR-887-3p” refers to the hsa-miR-887-3p gene (miRBase Accession No. Includes MIMAT0004951) and other species homologs or orthologs.
- the hsa-miR-887-3p gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, Vol. 16, p1289-1298. Further, as the precursor of "hsa-miR-887-3p", “hsa-mir-887” (miRBase Accession No. MI0005562, SEQ ID NO: 347) having a hairpin-like structure is known.
- hsa-miR-44745-5p gene or "hsa-miR-44745-5p” refers to the hsa-miR-44745-5p gene (miRBase Accession No. MIMAT0019878) and other species such as homologs or orthologs are included.
- the hsa-miR-4745-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4745-5p", “hsa-mir-4745” (miRBase Accession No. MI0017384, SEQ ID NO: 348) having a hairpin-like structure is known.
- hsa-miR-4433a-3p gene or "hsa-miR-4433a-3p” used herein refer to the hsa-miR-4433a-3p gene (miRBase Accession No. 3p) set forth in SEQ ID NO: 138. MIMAT0018949) and other species such as homologs or orthologs.
- the hsa-miR-4433a-3p gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4433a-3p", “hsa-mir-4433a” (miRBase Accession No. MI0016773, SEQ ID NO: 349) having a hairpin-like structure is known.
- hsa-miR-5090 gene or “hsa-miR-5090” refers to the hsa-miR-5090 gene (miRBase Accession No. MIMAT0021082) set forth in SEQ ID NO: 139 and other species. Includes homologs or orthologs.
- the hsa-miR-5090 gene can be obtained by the method described in Ding N et al., 2011, J Radiat Res, Vol. 52, p425-432. Further, as the precursor of "hsa-miR-5090", “hsa-mir-5090” (miRBase Accession No. MI0017979, SEQ ID NO: 350) having a hairpin-like structure is known.
- hsa-miR-296-5p gene or "hsa-miR-296-5p” refers to the hsa-miR-296-5p gene (miRBase Accession No. MIMAT0000690) and other species such as homologs or orthologs are included.
- the hsa-miR-296-5p gene can be obtained by the method described in Hubaviy HB et al., 2003, Dev Cell, Volume 5, p351-358. Further, as the precursor of "hsa-miR-296-5p", "hsa-mir-296” (miRBase Accession No. MI0000747, SEQ ID NO: 351) having a hairpin-like structure is known.
- hsa-miR-939-5p gene or "hsa-miR-939-5p” refers to the hsa-miR-939-5p gene (miRBase Accession No. Includes MIMAT0004982) and other species homologs or orthologs.
- the hsa-miR-939-5p gene can be obtained by the method described in Lui WO et al., 2007, Cancer Res, Vol. 67, p6031-6043. Further, as the precursor of "hsa-miR-939-5p", “hsa-mir-939” (miRBase Accession No. MI0005761, SEQ ID NO: 352) having a hairpin-like structure is known.
- hsa-miR-3648 gene or "hsa-miR-3648” refers to the hsa-miR-3648 gene (miRBase Accession No. MIMAT0018068) set forth in SEQ ID NO: 142 and other species. Includes homologs or orthologs.
- the hsa-miR-3648 gene can be obtained by the method described in Meiri E et al., 2010, Nucleic Acids Res, Vol. 38, pp. 6234-6246.
- hsa-miR-3648 has a hairpin-like structure as a precursor thereof, "hsa-mir-3648-1, hsa-mir-3648-2” (miRBase Accession No. MI0016048, MI0031512, SEQ ID NO: 353, 354) is known.
- hsa-miR-3196 gene or “hsa-miR-3196” refers to the hsa-miR-3196 gene (miRBase Accession No. MIMAT0015080) set forth in SEQ ID NO: 143 and other species. Includes homologs or orthologs.
- the hsa-miR-3196 gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-3196", “hsa-mir-3196” (miRBase Accession No. MI0014241, SEQ ID NO: 355) having a hairpin-like structure is known.
- hsa-miR-6722-3p gene or "hsa-miR-6722-3p” used herein refers to the hsa-miR-6722-3p gene (miRBase Accession No. MIMAT0025854) and other species such as homologs or orthologs.
- the hsa-miR-6722-3p gene can be obtained by the method described in Li Y et al., 2012, Gene, Vol. 497, p330-335. Further, as the precursor of "hsa-miR-6722-3p", “hsa-mir-6722” (miRBase Accession No. MI0022557, SEQ ID NO: 356) having a hairpin-like structure is known.
- hsa-miR-6805-5p gene or "hsa-miR-6805-5p” refers to the hsa-miR-6805-5p gene (miRBase Accession No. Includes MIMAT0027510) and other species homologs or orthologs.
- the hsa-miR-6805-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6805-5p", "hsa-mir-6805" (miRBase Accession No. MI0022650, SEQ ID NO: 325) having a hairpin-like structure is known.
- hsa-miR-1202 gene or “hsa-miR-1202” refers to the hsa-miR-1202 gene (miRBase Accession No. MIMAT00055865) set forth in SEQ ID NO: 146 and other species. Includes homologs or orthologs.
- the hsa-miR-1202 gene can be obtained by the method described in Marton S et al., 2008, Leukemia, Vol. 22, p330-338. Further, as the precursor of "hsa-miR-1202", “hsa-mir-1202" (miRBase Accession No. MI0006334, SEQ ID NO: 357) having a hairpin-like structure is known.
- hsa-miR-6775-5p gene or "hsa-miR-6775-5p” refers to the hsa-miR-6775-5p gene (miRBase Accession No. MIMAT0027450) and other species such as homologs or orthologs.
- the hsa-miR-6775-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6775-5p", “hsa-mir-6775” (miRBase Accession No. MI0022620, SEQ ID NO: 358) having a hairpin-like structure is known.
- hsa-miR-6087 gene or “hsa-miR-6087” refers to the hsa-miR-6087 gene (miRBase Accession No. MIMAT0023712) set forth in SEQ ID NO: 148 and other species. Includes homologs or orthologs.
- the hsa-miR-6087 gene can be obtained by the method described in Yoo JK et al., 2012, Stem Cells Dev, Vol. 21, p2049-2057. Further, as the precursor of "hsa-miR-6087", “hsa-mir-6087” (miRBase Accession No. MI0020364, SEQ ID NO: 359) having a hairpin-like structure is known.
- hsa-miR-6765-5p gene or "hsa-miR-6765-5p” used herein refers to the hsa-miR-6765-5p gene (miRBase Accession No. MIMAT0027430) and other species such as homologs or orthologs are included.
- the hsa-miR-6765-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6765-5p", “hsa-mir-6765” (miRBase Accession No. MI0022610, SEQ ID NO: 360) having a hairpin-like structure is known.
- hsa-miR-6875-5p gene or "hsa-miR-6875-5p” refers to the hsa-miR-6875-5p gene (miRBase Accession No. MIMAT0027650) and other species such as homologs or orthologs are included.
- the hsa-miR-6875-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6875-5p", "hsa-mir-6875” (miRBase Accession No. MI0022722, SEQ ID NO: 361) having a hairpin-like structure is known.
- hsa-miR-4674 gene or "hsa-miR-4674" refers to the hsa-miR-4674 gene (miRBase Accession No. MIMAT0019756) set forth in SEQ ID NO: 151 and other species. Includes homologs or orthologs.
- the hsa-miR-4674 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4674", “hsa-mir-4674" (miRBase Accession No. MI0017305, SEQ ID NO: 362) having a hairpin-like structure is known.
- hsa-miR-1233-5p gene or "hsa-miR-1233-5p” refers to the hsa-miR-1233-5p gene (miRBase Accession No. MIMAT0022943) and other species such as homologs or orthologs.
- the hsa-miR-1233-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, Vol. 28, p328-336.
- "hsa-miR-1233-5p” has a hairpin-like structure as a precursor thereof, "hsa-mir-1233-1, hsa-mir-1233-2” (miRBase Accession No. MI0006323, MI0015973, SEQ ID NO: 363,364) are known.
- hsa-miR-7114-5p gene or "hsa-miR-7114-5p” refers to the hsa-miR-7114-5p gene (miRBase Accession No. MIMAT0028125) and other species such as homologs or orthologs are included.
- the hsa-miR-7114-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-7114-5p", “hsa-mir-7114" (miRBase Accession No. MI0022965, SEQ ID NO: 365) having a hairpin-like structure is known.
- hsa-miR-5787 gene or “hsa-miR-5787” refers to the hsa-miR-5787 gene (miRBase Accession No. MIMAT0023252) and other species set forth in SEQ ID NO: 154. Includes homologs or orthologs.
- the hsa-miR-5787 gene can be obtained by the method described in Yoo H et al., 2011, Biochem Biophyss Res Commun, Vol. 415, p567-572. Further, as the precursor of "hsa-miR-5787", “hsa-mir-5787” (miRBase Accession No. MI0019977, SEQ ID NO: 366) having a hairpin-like structure is known.
- hsa-miR-8072 gene or “hsa-miR-8072” refers to the hsa-miR-8072 gene (miRBase Accession No. MIMAT0030999) set forth in SEQ ID NO: 155 and other species. Includes homologs or orthologs.
- the hsa-miR-8072 gene can be obtained by the method described in Wang HJ et al., 2013, Shock, Vol. 39, p480-487. Further, as the precursor of "hsa-miR-8072", “hsa-mir-8072” (miRBase Accession No. MI0025908, SEQ ID NO: 367) having a hairpin-like structure is known.
- hsa-miR-3619-3p gene or "hsa-miR-3619-3p” refers to the hsa-miR-3619-3p gene (miRBase Accession No. Includes MIMAT0019219) and other species homologs or orthologs.
- the hsa-miR-3619-3p gene can be obtained by the method described in Witten D et al., 2010, BMC Biol, Volume 8, p58.
- hsa-mir-3619 as the precursor of "hsa-miR-3619-3p
- hsa-mir-3619 having a hairpin-like structure is known.
- hsa-miR-4632-5p gene or "hsa-miR-4632-5p” used herein refers to the hsa-miR-4632-5p gene (miRBase Accession No. MIMAT0022977) and other species such as homologs or orthologs are included.
- the hsa-miR-4632-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4632-5p", "hsa-mir-4632” (miRBase Accession No. MI0017259, SEQ ID NO: 369) having a hairpin-like structure is known.
- hsa-miR-6800-5p gene or "hsa-miR-6800-5p” refers to the hsa-miR-6800-5p gene (miRBase Accession No. MIMAT0027500) and other species such as homologs or orthologs.
- the hsa-miR-6800-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6800-5p", “hsa-mir-6800” (miRBase Accession No. MI0022645, SEQ ID NO: 370) having a hairpin-like structure is known.
- hsa-miR-4634 gene or "hsa-miR-4634” refers to the hsa-miR-4634 gene (miRBase Accession No. MIMAT0019691) set forth in SEQ ID NO: 159 and other species. Includes homologs or orthologs.
- the hsa-miR-4634 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4634", “hsa-mir-4634” (miRBase Accession No. MI0017261, SEQ ID NO: 371) having a hairpin-like structure is known.
- hsa-miR-4486 gene or “hsa-miR-4486” refers to the hsa-miR-4486 gene (miRBase Accession No. MIMAT0019020) and other species set forth in SEQ ID NO: 160. Includes homologs or orthologs.
- the hsa-miR-4486 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4486", “hsa-mir-4486” (miRBase Accession No. MI0016847, SEQ ID NO: 372) having a hairpin-like structure is known.
- hsa-miR-6727-5p gene or "hsa-miR-6727-5p” refers to the hsa-miR-6727-5p gene (miRBase Accession No. Includes MIMAT0027355) and other species homologs or orthologs.
- the hsa-miR-6727-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, “hsa-miR-6727-5p” has a hairpin-like structure as a precursor thereof, and "hsa-mir-6727” (miRBase Accession No. MI0022572, SEQ ID NO: 373) is known.
- hsa-miR-4505 gene or "hsa-miR-4505" refers to the hsa-miR-4505 gene (miRBase Accession No. MIMAT0019041) set forth in SEQ ID NO: 162 and other species. Includes homologs or orthologs.
- the hsa-miR-4505 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4505", “hsa-mir-4505" (miRBase Accession No. MI0016868, SEQ ID NO: 374) having a hairpin-like structure is known.
- hsa-miR-4725-3p gene or "hsa-miR-4725-3p” used herein refer to the hsa-miR-4725-3p gene (miRBase Accession No. MIMAT0019844) and other species such as homologs or orthologs are included.
- the hsa-miR-4725-3p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4725-3p", “hsa-mir-4725” (miRBase Accession No. MI0017362, SEQ ID NO: 375) having a hairpin-like structure is known.
- hsa-miR-1538 gene or “hsa-miR-1538” refers to the hsa-miR-1538 gene (miRBase Accession No. MIMAT0007400) and other species set forth in SEQ ID NO: 164. Includes homologs or orthologs.
- the hsa-miR-1538 gene can be obtained by the method described in Azuma-Mukai A et al., 2008, Proc Natl Acad Sci USA, Vol. 105, pp. 7964-7769. Further, as the precursor of "hsa-miR-1538", “hsa-mir-1538” (miRBase Accession No. MI0007259, SEQ ID NO: 376) having a hairpin-like structure is known.
- hsa-miR-320b gene or "hsa-miR-320b” refers to the hsa-miR-320b gene (miRBase Accession No. MIMAT0005792) set forth in SEQ ID NO: 165 and other species. Includes homologs or orthologs.
- the hsa-miR-320b gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, Vol. 16, p1289-1298.
- hsa-miR-320b has a hairpin-like structure as a precursor thereof, "hsa-mir-320b-1, hsa-mir-320b-2" (miRBase Accession No. MI0003776, MI0003839, SEQ ID NO: 377, 378) is known.
- hsa-miR-1955-5p gene or "hsa-miR-1955-5p” refers to the hsa-miR-915-5p gene (miRBase Accession No.) set forth in SEQ ID NO: 166. Includes MIMAT0007891) and other species homologs or orthologs.
- the hsa-miR-1915-5p gene can be obtained by the method described in Bar M et al., 2008, Stem Cells, Vol. 26, p2496-2505. Further, as the precursor of "hsa-miR-915-5p", "hsa-mir-1915" (miRBase Accession No. MI0008336, SEQ ID NO: 379) having a hairpin-like structure is known.
- hsa-miR-328-5p gene or "hsa-miR-328-5p” refers to the hsa-miR-328-5p gene (miRBase Accession No. MIMAT0026486) and other species such as homologs or orthologs are included.
- the hsa-miR-328-5p gene can be obtained by the method described in Kim J et al., 2004, Proc Natl Acad Sci USA, Vol. 101, p360-365. Further, as the precursor of "hsa-miR-328-5p", “hsa-mir-328” (miRBase Accession No. MI00000804, SEQ ID NO: 380) having a hairpin-like structure is known.
- hsa-miR-6820-5p gene or "hsa-miR-6820-5p” refers to the hsa-miR-6820-5p gene (miRBase Accession No. MIMAT0027540) and other species such as homologs or orthologs.
- the hsa-miR-6820-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6820-5p", "hsa-mir-6820” (miRBase Accession No. MI0022665, SEQ ID NO: 381) having a hairpin-like structure is known.
- hsa-miR-6726-5p gene or "hsa-miR-6726-5p” refers to the hsa-miR-6726-5p gene (miRBase Accession No. Includes MIMAT0027353) and other species homologs or orthologs.
- the hsa-miR-6726-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6726-5p", “hsa-mir-6726” (miRBase Accession No. MI0022571, SEQ ID NO: 382) having a hairpin-like structure is known.
- hsa-miR-3665 gene or “hsa-miR-3665" refers to the hsa-miR-3665 gene (miRBase Accession No. MIMAT0018087) set forth in SEQ ID NO: 170 and other species. Includes homologs or orthologs.
- the hsa-miR-3665 gene can be obtained by the method described in Xie X et al., 2005, Nature, Vol. 434, p338-345. Further, as the precursor of "hsa-miR-3665", “hsa-mir-3665" (miRBase Accession No. MI0016066, SEQ ID NO: 383) having a hairpin-like structure is known.
- hsa-miR-638 gene or “hsa-miR-638” refers to the hsa-miR-638 gene (miRBase Accession No. MIMAT0003308) set forth in SEQ ID NO: 171 and other species. Includes homologs or orthologs.
- the hsa-miR-638 gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-1692. Further, as the precursor of "hsa-miR-638", “hsa-mir-638” (miRBase Accession No. MI0003653, SEQ ID NO: 384) having a hairpin-like structure is known.
- hsa-miR-762 gene or “hsa-miR-762” refers to the hsa-miR-762 gene (miRBase Accession No. MIMAT0010313) set forth in SEQ ID NO: 172 and other species. Includes homologs or orthologs.
- the hsa-miR-762 gene can be obtained by the method described in Berezikov E et al., 2006, Genome Res, Vol. 16, pp. 1289-1298. Further, as the precursor of "hsa-miR-762", “hsa-mir-762” (miRBase Accession No. MI0003892, SEQ ID NO: 385) having a hairpin-like structure is known.
- hsa-miR-4466 gene or “hsa-miR-4466” refers to the hsa-miR-4466 gene (miRBase Accession No. MIMAT0018993) and other species set forth in SEQ ID NO: 173. Includes homologs or orthologs.
- the hsa-miR-4466 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4466", “hsa-mir-4466” (miRBase Accession No. MI0016817, SEQ ID NO: 386) having a hairpin-like structure is known.
- hsa-miR-3940-5p gene or "hsa-miR-3940-5p” refers to the hsa-miR-3940-5p gene (miRBase Accession No. MIMAT0019229) and other species such as homologs or orthologs are included.
- the hsa-miR-3940-5p gene can be obtained by the method described in Liao JY et al., 2010, PLos One, Volume 5, e10563. Further, "hsa-miR-3940-5p” has a hairpin-like structure as a precursor thereof, and "hsa-mir-3940" (miRBase Accession No. MI00165591, SEQ ID NO: 387) is known.
- hsa-miR-1237-5p gene or "hsa-miR-1237-5p” refers to the hsa-miR-1237-5p gene (miRBase Accession No. Includes MIMAT0022946) and other species homologs or orthologs.
- the hsa-miR-1237-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, Vol. 28, p328-336.
- “hsa-miR-1237-5p” has a hairpin-like structure as a precursor thereof, and "hsa-mir-1237” (miRBase Accession No. MI0006327, SEQ ID NO: 388) is known.
- hsa-miR-575 gene or "hsa-miR-575" refers to the hsa-miR-575 gene (miRBase Accession No. MIMAT0003240) set forth in SEQ ID NO: 176 and other species. Includes homologs or orthologs.
- the hsa-miR-575 gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-1692. Further, as the precursor of "hsa-miR-575", “hsa-mir-575" (miRBase Accession No. MI0003582, SEQ ID NO: 389) having a hairpin-like structure is known.
- hsa-miR-3656 gene or “hsa-miR-3656” refers to the hsa-miR-3656 gene (miRBase Accession No. MIMAT0018076) set forth in SEQ ID NO: 177 and other species. Includes homologs or orthologs.
- the hsa-miR-3656 gene can be obtained by the method described in Meiri E et al., 2010, Nucleic Acids Res, Vol. 38, pp. 6234-6246. Further, as the precursor of "hsa-miR-3656", “hsa-mir-3656” (miRBase Accession No. MI0016056, SEQ ID NO: 390) having a hairpin-like structure is known.
- hsa-miR-4488 gene or “hsa-miR-4488” refers to the hsa-miR-4488 gene (miRBase Accession No. MIMAT0019022) set forth in SEQ ID NO: 178 and other species. Includes homologs or orthologs.
- the hsa-miR-4488 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4488", “hsa-mir-4488” (miRBase Accession No. MI0016849, SEQ ID NO: 391) having a hairpin-like structure is known.
- hsa-miR-4281 gene or “hsa-miR-4281” refers to the hsa-miR-4281 gene (miRBase Accession No. MIMAT0016907) set forth in SEQ ID NO: 179 and other species. Includes homologs or orthologs.
- the hsa-miR-4281 gene can be obtained by the method described in Goff LA et al., 2009, PLos One, Volume 4, e7192. Further, as the precursor of "hsa-miR-4281", “hsa-mir-4281” (miRBase Accession No. MI0015885, SEQ ID NO: 392) having a hairpin-like structure is known.
- hsa-miR-6781-5p gene or "hsa-miR-6781-5p” used herein refers to the hsa-miR-6781-5p gene (miRBase Accession No. Includes MIMAT0027462) and other species homologs or orthologs.
- the hsa-miR-6781-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6781-5p", "hsa-mir-6781” (miRBase Accession No. MI0022626, SEQ ID NO: 393) having a hairpin-like structure is known.
- hsa-miR-4532 gene or "hsa-miR-4532” refers to the hsa-miR-4532 gene (miRBase Accession No. MIMAT0019071) set forth in SEQ ID NO: 181 and other species. Includes homologs or orthologs.
- the hsa-miR-4532 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4532", “hsa-mir-4532” (miRBase Accession No. MI0016899, SEQ ID NO: 394) having a hairpin-like structure is known.
- hsa-miR-4665-5p gene or "hsa-miR-4665-5p” refers to the hsa-miR-4665-5p gene (miRBase Accession No. MIMAT0019739) and other species such as homologs or orthologs are included.
- the hsa-miR-4665-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, "hsa-miR-4665-5p” has a hairpin-like structure as a precursor thereof, and "hsa-mir-4665" (miRBase Accession No. MI0017295, SEQ ID NO: 395) is known.
- hsa-miR-6816-5p gene or "hsa-miR-6816-5p” refers to the hsa-miR-6816-5p gene (miRBase Accession No. MIMAT0027532) and other species such as homologs or orthologs.
- the hsa-miR-6816-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6816-5p", “hsa-mir-6816” (miRBase Accession No. MI0022661, SEQ ID NO: 396) having a hairpin-like structure is known.
- hsa-miR-4508 gene or “hsa-miR-4508” refers to the hsa-miR-4508 gene (miRBase Accession No. MIMAT00190405) set forth in SEQ ID NO: 184 and other species. Includes homologs or orthologs.
- the hsa-miR-4508 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4508", “hsa-mir-4508” (miRBase Accession No. MI0016872, SEQ ID NO: 397) having a hairpin-like structure is known.
- hsa-miR-6784-5p gene or "hsa-miR-6784-5p” as used herein refers to the hsa-miR-6784-5p gene (miRBase Accession No. MIMAT0027468) and other species such as homologs or orthologs are included.
- the hsa-miR-6784-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6784-5p", “hsa-mir-6784" (miRBase Accession No. MI0022629, SEQ ID NO: 398) having a hairpin-like structure is known.
- hsa-miR-6786-5p gene or "hsa-miR-6786-5p” as used herein refers to the hsa-miR-6786-5p gene (miRBase Accession No. Includes MIMAT0027472) and other species homologs or orthologs.
- the hsa-miR-6786-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6786-5p", "hsa-mir-6786” (miRBase Accession No. MI0022631, SEQ ID NO: 399) having a hairpin-like structure is known.
- hsa-miR-4471 gene or “hsa-miR-4471” refers to the hsa-miR-4471 gene (miRBase Accession No. MIMAT0019871) set forth in SEQ ID NO: 187 and other species. Includes homologs or orthologs.
- the hsa-miR-4741 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4471", “hsa-mir-4471” (miRBase Accession No. MI0017379, SEQ ID NO: 400) having a hairpin-like structure is known.
- hsa-miR-1343-5p gene or "hsa-miR-1343-5p” as used herein refers to the hsa-miR-1343-5p gene (miRBase Accession No. Includes MIMAT0027038) and other species homologs or orthologs.
- the hsa-miR-1343-5p gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-1343-5p", “hsa-mir-1343” (miRBase Accession No. MI0017320, SEQ ID NO: 401) having a hairpin-like structure is known.
- hsa-miR-1227-5p gene or "hsa-miR-1227-5p” refers to the hsa-miR-1227-5p gene (miRBase Accession No. Includes MIMAT0022941) and other species homologs or orthologs.
- the hsa-miR-1227-5p gene can be obtained by the method described in Berezikov E et al., 2007, Mol Cell, Vol. 28, p328-336. Further, as the precursor of "hsa-miR-1227-5p", “hsa-mir-1227” (miRBase Accession No. MI0006316, SEQ ID NO: 402) having a hairpin-like structure is known.
- hsa-miR-4734 gene or “hsa-miR-4734” refers to the hsa-miR-4734 gene (miRBase Accession No. MIMAT0019859) set forth in SEQ ID NO: 190 and other species. Includes homologs or orthologs.
- the hsa-miR-4734 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4734", “hsa-mir-4734” (miRBase Accession No. MI0017371, SEQ ID NO: 403) having a hairpin-like structure is known.
- hsa-miR-3960 gene or “hsa-miR-3960” refers to the hsa-miR-3960 gene (miRBase Accession No. MIMAT0019337) set forth in SEQ ID NO: 191 and other species. Includes homologs or orthologs.
- the hsa-miR-3960 gene can be obtained by the method described in Hu R et al., 2011, J Biol Chem, Vol. 286, p12328-12339. Further, as the precursor of "hsa-miR-3960", “hsa-mir-3960” (miRBase Accession No. MI0016964, SEQ ID NO: 404) having a hairpin-like structure is known.
- hsa-miR-128-2-5p gene or "hsa-miR-128-2-5p” is described in SEQ ID NO: 192, hsa-miR-128-2-5p. Includes genes (miRBase Accession No. MIMAT0031095) and other species homologs or orthologs.
- the hsa-miR-128-2-5p gene can be obtained by the method described in Lagos-Quintana M et al., 2002, Curr Biol, Vol. 12, p735-739. Further, as the precursor of "hsa-miR-128-2-5p", “hsa-mir-128-2” (miRBase Accession No. MI0000727, SEQ ID NO: 405) having a hairpin-like structure is known.
- hsa-miR-6743-5p gene or "hsa-miR-6743-5p” used herein refer to the hsa-miR-6743-5p gene (miRBase Accession No.) set forth in SEQ ID NO: 193. MIMAT0027387) and other species such as homologs or orthologs are included.
- the hsa-miR-6743-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6743-5p", “hsa-mir-6734" (miRBase Accession No. MI0022588, SEQ ID NO: 406) having a hairpin-like structure is known.
- hsa-miR-663a gene or “hsa-miR-663a” refers to the hsa-miR-663a gene (miRBase Accession No. MIMAT0003326) set forth in SEQ ID NO: 194 and other species. Includes homologs or orthologs.
- the hsa-miR-663a gene can be obtained by the method described in Cummins JM et al., 2006, Proc Natl Acad Sci USA, Vol. 103, p3687-3692. Further, as the precursor of "hsa-miR-663a”, “hsa-mir-663a” (miRBase Accession No. MI0003672, SEQ ID NO: 407) having a hairpin-like structure is known.
- hsa-miR-6729-5p gene or "hsa-miR-6729-5p” used herein refers to the hsa-miR-6729-5p gene (miRBase Accession No. Includes MIMAT0027359) and other species homologs or orthologs.
- the hsa-miR-6729-5p gene can be obtained by the method described in Ladywig E et al., 2012, Genome Res, Vol. 22, p1634-1645. Further, as the precursor of "hsa-miR-6729-5p", “hsa-mir-6729” (miRBase Accession No. MI0022574, SEQ ID NO: 408) having a hairpin-like structure is known.
- hsa-miR-1915-3p gene or "hsa-miR-1915-3p” used herein refer to the hsa-miR-1915-3p gene (miRBase Accession No. Includes MIMAT0007892) and other species homologs or orthologs.
- the hsa-miR-1915-3p gene can be obtained by the method described in Bar M et al., 2008, Stem Cells, Vol. 26, p2496-2505. Further, as the precursor of "hsa-miR-1915-3p", "hsa-mir-1915" (miRBase Accession No. MI0008336, SEQ ID NO: 379) having a hairpin-like structure is known.
- hsa-miR-1268b gene or "hsa-miR-1268b” refers to the hsa-miR-1268b gene (miRBase Accession No. MIMAT0018925) set forth in SEQ ID NO: 197 and other species. Includes homologs or orthologs.
- the hsa-miR-1268b gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-1268b", “hsa-mir-1268b” (miRBase Accession No. MI0016748, SEQ ID NO: 409) having a hairpin-like structure is known.
- hsa-miR-4651 gene or “hsa-miR-4651” refers to the hsa-miR-4651 gene (miRBase Accession No. MIMAT0019715) set forth in SEQ ID NO: 198 and other species. Includes homologs or orthologs.
- the hsa-miR-4651 gene can be obtained by the method described in Persson H et al., 2011, Cancer Res, Vol. 71, p78-86. Further, as the precursor of "hsa-miR-4651", “hsa-mir-4651” (miRBase Accession No. MI0017279, SEQ ID NO: 410) having a hairpin-like structure is known.
- hsa-miR-3178 gene or “hsa-miR-3178” refers to the hsa-miR-3178 gene (miRBase Accession No. MIMAT0015055) set forth in SEQ ID NO: 199 and other species. Includes homologs or orthologs.
- the hsa-miR-3178 gene can be obtained by the method described in Stark MS et al., 2010, PLos One, Volume 5, e9685. Further, as the precursor of "hsa-miR-3178", “hsa-mir-3178” (miRBase Accession No. MI0014212, SEQ ID NO: 411) having a hairpin-like structure is known.
- hsa-miR-4463 or “hsa-miR-4463” refers to the hsa-miR-4463 gene (miRBase Accession No. MIMAT0018987) set forth in SEQ ID NO: 200 and other species. Includes homologs or orthologs.
- the hsa-miR-4463 gene can be obtained by the method described in Jima DD et al., 2010, Blood, Volume 116, e118-e127. Further, as the precursor of "hsa-miR-4463", “hsa-mir-4463” (miRBase Accession No. MI0016811, SEQ ID NO: 412) having a hairpin-like structure is known.
- a mature miRNA When a mature miRNA is excised as a mature miRNA from an RNA precursor having a hairpin-like structure, one to several bases before and after the sequence are excised short or long, or base substitution occurs and mutation occurs. It may become a body, which is referred to as isomiR (Morin RD. et al., 2008, Genome Res., Vol. 18, p. 610-621).
- miRBase version 21
- a number of mutations in the base sequence represented by any of SEQ ID NOs: 413 to 740 called isomiR corresponding thereto.
- Body and fragments are also shown. These mutants can also be obtained as variants of miRNA having the nucleotide sequence represented by any of SEQ ID NOs: 1 to 200.
- a polynucleotide variant consisting of a nucleotide sequence in which u is t in the nucleotide sequence, for example, registered in miRBase (Version 21).
- the longest variants present are SEQ ID NOs: 413, 415, 417, 419, 421, 423, 425, 427, 249, 431, 433, 435, 437, 439, 441, 443, 445, 447, 449, 451 and 453.
- the shortest variants are SEQ ID NOs: 414, 416, 418, 420, 422, 424, 426, 428, 430, 432, 434, 436, 438, 440, 442, 444, 446, 448, respectively.
- isomiR polynucleotides corresponding to miRNAs having the nucleotide sequences represented by any of SEQ ID NOs: 1 to 200 are registered in miRBase.
- the polynucleotide containing the base sequence represented by any of SEQ ID NOs: 1 to 200 the polynucleotide represented by any of SEQ ID NOs: 201 to 412, which are precursors, can be mentioned.
- the name of the gene / miRNA consisting of the nucleotide sequences represented by SEQ ID NOs: 1 to 740 and the miRBase Accession No. (Registration number) is shown in Table 1.
- nucleic acid probe or primer used in the present invention binds to a specific target nucleic acid and cannot substantially bind to other nucleic acids.
- hippocampal atrophy can be detected with high accuracy (or AUC), sensitivity, and specificity, and a hippocampal atrophy person and a normal hippocampal person (non-hippocampal atrophy person) can be distinguished.
- FIG. 1 shows hsa-mir-1915 represented by SEQ ID NO: 379, which is a precursor, hsa-miR-915-5p represented by SEQ ID NO: 166 generated from the precursor, and hsa represented by SEQ ID NO: 196.
- the relationship of the base sequence of -miR-1915-3p is shown.
- 2A and 2B are discriminant score diagrams of the learning sample group (A) and the cross-validation sample group (B) obtained in Example 6.
- 2C and 2D are ROC curves of the learning sample group (C) and the cross-validation sample group (D).
- FIG. 3 is a regression line obtained by Kaiba quantitative values and explanatory variables obtained in Example 7- (2).
- FIG. 3A is a learning / cross-validation sample group
- FIG. 3B is an independent validation sample group.
- Nucleic Acids for Kaiba Atrophy Targets as Kaiba Atrophy Markers for Detecting Kaiba Atrophy, or Kaiba Atrophy or Non-Atrophy Using Nucleic Acids for Detecting Kaiba Atrophy as defined above in the present invention, such as nucleic acid probes or primers.
- Nucleic acids are miR-3131, miR-6757-5p, miR-4706, miR-5001-5p, miR-3180-3p, miR-642b-3p, miR-4655-5p, miR-6819-5p, miR-937.
- miR-4688 miR-6471-5p, miR-7107-5p, miR-4271, miR-1229-5p, miR-4707-5p, miR-6808-5p, miR-4656, miR-6076, miR -6762-5p, miR-7109-5p, miR-6732-5p, miR-3195, miR-7150, miR-642a-3p, miR-1249-5p, miR-3185, miR-4689, miR-3141, miR -6840-3p, miR-3135b, miR-1914-3p, miR-4446-3p, miR-4433b-3p, miR-6877-5p, miR-6884-5p, miR-3620-5p, miR-6825-5p , MiR-5739, miR-3663-3p, miR-4695-5p, miR-3162-5p, miR-3679-5p, miR-8059, miR-7110-5p, miR-1275, mi
- hippocampal atrophy markers examples include hsa-miR-3131, hsa-miR-6757-5p, hsa-miR-4706, hsa-miR-5001-5p, hsa-miR-3180.
- hippocampal atrophy markers specifically miR-1228-5p, miR-760, miR-187-5p, miR-7111-5p, miR-6088, miR-6805-3p, miR-4640-5p, miR-6721-5p, miR-6880-5p, miR-711, miR-128-1-5p, miR-4525, miR-486-3p, miR-6756-5p, miR-1260b, miR-3184-5p, miR-6075, miR-204-3p, miR-4728-5p, miR-4534, miR-4758-5p, miR-8063, miR-6863-3p, miR-6789-5p, miR- 744-5p, miR-1909-3p, miR-887-3p, miR-4745-5p, miR-4433a-3p, miR-5090, miR-296-5p, miR-939-5p, miR-3648, miR- 3196
- hippocampal atrophy markers examples include hsa-miR-1228-5p, hsa-miR-760, hsa-miR-187-5p, hsa-miR-7111-5p, hsa-miR-6088, hsa.
- the above miRNAs include human miRNA polynucleotides / genes, hsa-miR-3131, hsa-miR-6757-5p, hsa-miR-4706, hsa-miR-5001-5p, hsa-miR-3180-3p.
- a preferred target nucleic acid is a human miRNA polynucleotide / gene comprising the nucleotide sequence represented by any of SEQ ID NOs: 1 to 200, a transcript thereof, more preferably the transcript, i.e. miRNA, a pre-RNA thereof. It is miRNA or pre-miRNA.
- the first target gene is the hsa-miR-3131 gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the second target gene is the hsa-miR-6757-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the third target gene is the hsa-miR-4706 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the fourth target gene is the hsa-miR-5001-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the fifth target gene is the hsa-miR-3180-3p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the sixth target gene is the hsa-miR-642b-3p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the seventh target gene is the hsa-miR-4655-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the eighth target gene is the hsa-miR-6819-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the ninth target gene is the hsa-miR-937-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the tenth target gene is the hsa-miR-4688 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the eleventh target gene is the hsa-miR-6741-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the twelfth target gene is the hsa-miR-7107-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the thirteenth target gene is the hsa-miR-4271 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 14th target gene is the hsa-miR-1229-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the fifteenth target gene is the hsa-miR-4707-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 16th target gene is the hsa-miR-6808-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 17th target gene is the hsa-miR-4656 gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the eighteenth target gene is the hsa-miR-6076 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 19th target gene is the hsa-miR-6762-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the twentieth target gene is the hsa-miR-7109-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 21st target gene is the hsa-miR-6732-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 22nd target gene is the hsa-miR-3195 gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 23rd target gene is the hsa-miR-7150 gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 24th target gene is the hsa-miR-642a-3p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 25th target gene is the hsa-miR-1249-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 26th target gene is the hsa-miR-3185 gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 27th target gene is the hsa-miR-4689 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 28th target gene is the hsa-miR-3141 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 29th target gene is the hsa-miR-6840-3p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the thirtieth target gene is the hsa-miR-3135b gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 31st target gene is the hsa-miR-1914-3p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 32nd target gene is the hsa-miR-4446-3p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 33rd target gene is the hsa-miR-4433b-3p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 34th target gene is the hsa-miR-6877-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 35th target gene is the hsa-miR-6884-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 36th target gene is the hsa-miR-3620-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 37th target gene is the hsa-miR-6825-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 38th target gene is the hsa-miR-5739 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 39th target gene is the hsa-miR-3663-3p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 40th target gene is the hsa-miR-4695-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 41st target gene is the hsa-miR-3162-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 42nd target gene is the hsa-miR-3679-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 43rd target gene is the hsa-miR-8059 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 44th target gene is the hsa-miR-7110-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 45th target gene is the hsa-miR-1275 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 46th target gene is the hsa-miR-6779-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 47th target gene is the hsa-miR-197-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 48th target gene is the hsa-miR-6845-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 49th target gene is the hsa-miR-4327 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 50th target gene is the hsa-miR-4723-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 51st target gene is the hsa-miR-4530 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 52nd target gene is the hsa-miR-6771-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 53rd target gene is the hsa-miR-614 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 54th target gene is the hsa-miR-92a-2-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 55th target gene is the hsa-miR-6891-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 56th target gene is the hsa-miR-6124 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 57th target gene is the hsa-miR-4678-3p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 58th target gene is the hsa-miR-4442 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 59th target gene is the hsa-miR-7977 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 60th target gene is the hsa-miR-6785-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 61st target gene is the hsa-miR-4497 gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 62nd target gene is the hsa-miR-8071 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 63rd target gene is the hsa-miR-663b gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 64th target gene is the hsa-miR-3180 gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 65th target gene is the hsa-miR-4251 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 66th target gene is the hsa-miR-1285-3p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 67th target gene is the hsa-miR-6870-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 68th target gene is the hsa-miR-4484 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 69th target gene is the hsa-miR-4476 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 70th target gene is the hsa-miR-6794-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 71st target gene is the hsa-miR-4454 gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 72nd target gene is the hsa-miR-6893-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 73rd target gene is the hsa-miR-6085 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 74th target gene is the hsa-miR-4787-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 75th target gene is the hsa-miR-149-3p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 76th target gene is the hsa-miR-7704 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 77th target gene is the hsa-miR-6125 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 78th target gene is the hsa-miR-6090 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 79th target gene is the hsa-miR-3197 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 80th target gene is the hsa-miR-6850-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 81st target gene is the hsa-miR-4467 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 82nd target gene is the hsa-miR-6885-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 83rd target gene is the hsa-miR-6803-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 84th target gene is the hsa-miR-6798-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 85th target gene is the hsa-miR-6780b-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 86th target gene is the hsa-miR-6768-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 87th target gene is the hsa-miR-5100 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 88th target gene is the hsa-miR-6724-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 89th target gene is the hsa-miR-6879-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 90th target gene is the hsa-miR-7108-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 91st target gene is the hsa-miR-4649-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 92nd target gene is the hsa-miR-4739 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 93rd target gene is the hsa-miR-6089 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 94th target gene is the hsa-miR-1908-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 95th target gene is the hsa-miR-4516 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 96th target gene is the hsa-miR-2861 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 97th target gene is the hsa-miR-4492 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 98th target gene is the hsa-miR-4294 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 99th target gene is the hsa-miR-6791-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 100th target gene is the hsa-miR-1469 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 101st target gene is the hsa-miR-6752-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 102nd target gene is the hsa-miR-4730 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 103rd target gene is the hsa-miR-6126 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 104th target gene is the hsa-miR-6869-5p gene, their homologues, their transcripts, or their variants or derivatives. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 105th target gene is the hsa-miR-1268a gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 106th target gene is the hsa-miR-6799-5p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 107th target gene is the hsa-miR-8069 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 108th target gene is the hsa-miR-3621 gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 109th target gene is the hsa-miR-4763-3p gene, their homologues, their transcripts, or variants or derivatives thereof. There are no known reports that changes in the expression of genes or transcripts thereof can be markers of hippocampal atrophy.
- the 110th target gene is the hsa-miR-1228-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 111th target gene is the hsa-miR-760 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported so far that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 5).
- the 112th target gene is the hsa-miR-187-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported so far that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 7).
- the 113th target gene is the hsa-miR-7111-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 114th target gene is the hsa-miR-6088 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 115th target gene is the hsa-miR-6805-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 116th target gene is the hsa-miR-4640-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 117th target gene is the hsa-miR-6721-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 118th target gene is the hsa-miR-6880-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 119th target gene is the hsa-miR-711 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported so far that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 7).
- the 120th target gene is the hsa-miR-128-1-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported so far that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 7).
- the 121st target gene is the hsa-miR-4525 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 122nd target gene is the hsa-miR-486-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 2).
- the 123rd target gene is the hsa-miR-6756-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 124th target gene is the hsa-miR-1260b gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 125th target gene is the hsa-miR-3184-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 126th target gene is the hsa-miR-6075 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 127th target gene is the hsa-miR-204-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Non-Patent Document 5).
- the 128th target gene is the hsa-miR-4728-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 129th target gene is the hsa-miR-4534 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 130th target gene is the hsa-miR-4758-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 131st target gene is the hsa-miR-8063 gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 132nd target gene is the hsa-miR-6863-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 133rd target gene is the hsa-miR-6789-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 134th target gene is the hsa-miR-744-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 135th target gene is the hsa-miR-1909-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 136th target gene is the hsa-miR-887-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported so far that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 5).
- the 137th target gene is the hsa-miR-4745-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 138th target gene is the hsa-miR-4433a-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 139th target gene is the hsa-miR-5090 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 140th target gene is the hsa-miR-296-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 141st target gene is the hsa-miR-939-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 142nd target gene is the hsa-miR-3648 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 143rd target gene is the hsa-miR-3196 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 144th target gene is the hsa-miR-6722-3p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Non-Patent Document 4).
- the 145th target gene is the hsa-miR-6805-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 3).
- the 146th target gene is the hsa-miR-1202 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 147th target gene is the hsa-miR-6775-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 148th target gene is the hsa-miR-6087 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 149th target gene is the hsa-miR-6765-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 150th target gene is the hsa-miR-6875-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 151st target gene is the hsa-miR-4674 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Non-Patent Document 4).
- the 152nd target gene is the hsa-miR-1233-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 153rd target gene is the hsa-miR-7114-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 154th target gene is the hsa-miR-5787 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 155th target gene is the hsa-miR-8072 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 156th target gene is the hsa-miR-3619-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 157th target gene is the hsa-miR-4632-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 158th target gene is the hsa-miR-6800-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 159th target gene is the hsa-miR-4634 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 160th target gene is the hsa-miR-4486 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 161st target gene is the hsa-miR-6727-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 162nd target gene is the hsa-miR-4505 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 163rd target gene is the hsa-miR-4725-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 164th target gene is the hsa-miR-1538 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 165th target gene is the hsa-miR-320b gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 4).
- the 166th target gene is the hsa-miR-1955-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 167th target gene is the hsa-miR-328-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported so far that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 7).
- the 168th target gene is the hsa-miR-6820-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 169th target gene is the hsa-miR-6726-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 170th target gene is the hsa-miR-3665 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 171st target gene is the hsa-miR-638 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 172nd target gene is the hsa-miR-762 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 173rd target gene is the hsa-miR-4466 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 174th target gene is the hsa-miR-3940-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 175th target gene is the hsa-miR-1237-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 176th target gene is the hsa-miR-575 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 177th target gene is the hsa-miR-3656 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 178th target gene is the hsa-miR-4488 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 179th target gene is the hsa-miR-4281 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 180th target gene is the hsa-miR-6781-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 181st target gene is the hsa-miR-4532 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 182nd target gene is the hsa-miR-4665-5p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 183rd target gene is the hsa-miR-6816-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 184th target gene is the hsa-miR-4508 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 185th target gene is the hsa-miR-6784-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 186th target gene is the hsa-miR-6786-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 187th target gene is the hsa-miR-4471 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 188th target gene is the hsa-miR-1343-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 189th target gene is the hsa-miR-1227-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 190th target gene is the hsa-miR-4734 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 191st target gene is the hsa-miR-3960 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 192nd target gene is the hsa-miR-128-2-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported so far that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 7).
- the 193rd target gene is the hsa-miR-6743-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 194th target gene is the hsa-miR-663a gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Non-Patent Document 3).
- the 195th target gene is the hsa-miR-6729-5p gene, their homologues, their transcripts, or their variants or derivatives. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 196th target gene is the hsa-miR-1915-3p gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 197th target gene is the hsa-miR-1268b gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 198th target gene is the hsa-miR-4651 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 199th target gene is the hsa-miR-3178 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 1).
- the 200th target gene is the hsa-miR-4463 gene, their homologues, their transcripts, or variants or derivatives thereof. It has been reported that changes in the expression of genes or transcripts thereof can be markers for hippocampal atrophy and diseases characterized by hippocampal atrophy (Patent Document 6).
- nucleic acid for detecting hippocampal atrophy for example, a nucleic acid that can be used for diagnosing hippocampal atrophy, for example, a nucleic acid probe or a primer is used as a target nucleic acid for hippocampal atrophy (kaiba atrophy marker).
- nucleic acids that can be used to detect hippocampal atrophy or to diagnose hippocampal atrophy are human-derived hsa-miR-3131, hsa-miR-6757-5p.
- the nucleic acid can be a nucleic acid.
- the nucleic acid can be a nucleic acid that can specifically bind to the polynucleotide that is the miRNA or the complementary strand of the polynucleotide and amplify it.
- the primer may be a primer set that can specifically bind to the polynucleotide that is the miRNA or the complementary strand of the polynucleotide and amplify it, for example, a primer pair.
- hippocampal atrophy marker miRNA is another hippocampal atrophy marker miR-1228-5p, miR-760, miR-187-5p, miR-7111-5p, miR-6088, miR-6805-3p, miR-4640.
- the above-mentioned nucleic acid for detecting hippocampal atrophy can detect a hippocampal atrophy marker, and the above-mentioned miRNA which is a hippocampal atrophy marker, for example, SEQ ID NOs: 1 to 200, alone or in combination of two or more thereof.
- a hippocampal atrophy marker for example, SEQ ID NOs: 1 to 200, alone or in combination of two or more thereof.
- the nucleic acids that can be used in the present invention are specific to a polynucleotide consisting of a nucleotide sequence represented by at least one of SEQ ID NOs: 1 to 109, or a complementary strand of the polynucleotide. It is a nucleic acid probe capable of binding to a nucleic acid, or a primer capable of amplifying a polynucleotide consisting of a nucleotide sequence represented by at least one of SEQ ID NOs: 1 to 109.
- the nucleic acids that can be used in the present invention are further associated with a polynucleotide consisting of the nucleotide sequence represented by at least one of SEQ ID NOs: 110-200, or a complementary strand of the polynucleotide.
- a nucleic acid probe capable of specifically binding, or a primer capable of amplifying a polynucleotide consisting of a nucleotide sequence represented by at least one of SEQ ID NOs: 110 to 200 can be further included.
- nucleic acids that can be used in the present invention include the base sequence represented by any of SEQ ID NOs: 1 to 740, or the base sequence in which u is t in the base sequence.
- a group of polynucleotides and their complementary polynucleotides a group of polynucleotides that hybridize with DNA consisting of a base sequence complementary to the base sequence under stringent conditions (described later), and a group of polynucleotides complementary thereto, and It contains a combination of one or more polynucleotides selected from a group of polynucleotides containing 15 or more, preferably 17 or more consecutive bases in the base sequence of those polynucleotide groups.
- These polynucleotides can be used as nucleic acids for detecting miRNA, which is a target nucleic acid, which is a marker for hippocampal atrophy, for example, a nucleic acid probe or a primer.
- examples of nucleic acids that can be used in the present invention are one or more polynucleotides selected from the group consisting of the following polynucleotides (a)-(e):
- A A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence in which u is t in the base sequence, a mutant thereof, and a derivative thereof contain 15 or more consecutive bases. That fragment
- (C) A polynucleotide consisting of a base sequence complementary to the base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 The fragment containing the above contiguous bases
- (D) A polynucleotide containing a base sequence complementary to the base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 Fragments containing the above contiguous bases
- (E) A polynucleotide that hybridizes with any of the polynucleotides (a) to (d) above under stringent conditions.
- polynucleotides (a) to (e) are polynucleotides consisting of or derived from the base sequence represented by any of SEQ ID NOs: 1 to 109.
- Nucleic acids that can be used in the present invention further include at least one polynucleotide selected from the group consisting of the above polynucleotides (a) to (e), as well as the following polynucleotide (f). It can contain at least one polynucleotide selected from the group consisting of (j): (F) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases.
- (G) A polynucleotide containing a base sequence represented by any of SEQ ID NOs: 110 to 200, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (H) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more.
- polynucleotides (f) to (j) are polynucleotides consisting of or derived from the base sequence represented by any of SEQ ID NOs: 110 to 200.
- the nucleic acid used in the present invention may be DNA or RNA.
- the above-mentioned nucleic acid that can be used in the present invention can be prepared by using general techniques such as DNA recombination technique, PCR method, and method using an automatic DNA / RNA synthesizer.
- DNA recombination technology and PCR method for example, Ausube et al., Molecular Protocols in Molecular Biology, John Willey & Sons, US (1993); Sambrook et al., Molecular Cloning A Labor The techniques described in can be used.
- hsa-miR-3131 Human-derived hsa-miR-3131, hsa-miR-6757-5p, hsa-miR-4706, hsa-miR-5001-5p, hsa-miR-3180- consisting of the nucleotide sequences represented by SEQ ID NOs: 1 to 200.
- nucleic acids such as nucleic acid probes or primers
- nucleic acid synthesizer can be chemically synthesized using an automatic nucleic acid synthesizer.
- the phosphoramidite method is generally used for this synthesis, and single-stranded DNA up to about 100 bases can be automatically synthesized by this method.
- the nucleic acid automatic synthesizer is commercially available from, for example, Polygen, ABI, Applied Biosystems, and the like.
- the polynucleotide of the present invention can also be prepared by the cDNA cloning method.
- the cDNA cloning technique for example, microRNA Cloning Kit Wako or the like can be used.
- Nucleic acids for detecting a polynucleotide consisting of a nucleotide sequence represented by any of SEQ ID NOs: 1 to 200 are those that do not exist in vivo as miRNA or a precursor thereof. It may be there.
- the nucleotide sequences represented by SEQ ID NO: 166 and SEQ ID NO: 196 are generated from the precursor represented by SEQ ID NO: 379, which has a hairpin-like structure as shown in FIG.
- the base sequences represented by SEQ ID NO: 166 and SEQ ID NO: 196 have mismatch sequences with each other.
- nucleic acid for detecting the base sequence represented by any of SEQ ID NOs: 1 to 200 for example, the nucleic acid probe and the primer may have an artificial base sequence that does not exist in the living body.
- the present invention provides one or more of the nucleic acid for detecting hippocampal atrophy in the present invention, for example, a polynucleotide that can be used as a nucleic acid probe or primer for measuring a target nucleic acid that is a marker for hippocampal atrophy.
- a kit or device for detecting hippocampal atrophy including.
- the target nucleic acid which is a hippocampal atrophy marker in the present invention, is preferably selected from at least one of the following group A.
- Group A miR-3131, miR-6757-5p, miR-4706, miR-5001-5p, miR-3180-3p, miR-642b-3p, miR-4655-5p, miR-6819-5p, miR-937-5p, miR-4688, miR-6471-5p, miR-7107-5p, miR-4271, miR-1229-5p, miR-4707-5p, miR-6808-5p, miR-4656, miR-6076, miR-6762- 5p, miR-7109-5p, miR-6732-5p, miR-3195, miR-7150, miR-642a-3p, miR-1249-5p, miR-3185, miR-4689, miR-3141, miR-6840- 3p, miR-3135b, miR-1914-3p, miR-4
- Group B miR-1228-5p, miR-760, miR-187-5p, miR-7111-5p, miR-6088, miR-6805-3p, miR-4640-5p, miR-6721-5p, miR-6880-5p, miR-711, miR-128-1-5p, miR-4525, miR-486-3p, miR-6756-5p, miR-1260b, miR-3184-5p, miR-6075, miR-204-3p, miR- 4728-5p, miR-4534, miR-4758-5p, miR-8063, miR-6863-3p, miR-6789-5p, miR-744-5p, miR-1909-3p, miR-887-3p, miR- 4745-5p, miR-4433a-3p, miR-5090, miR-296-5p, miR
- the kit or device of the present invention is derived from a nucleic acid that can specifically bind to the target nucleic acid that is the above-mentioned hippocampal atrophy marker or the complementary strand of the polynucleotide, preferably a polynucleotide that is a miRNA described in the above group. Includes one or more polynucleotides of choice or variants thereof.
- the above-mentioned target nucleic acid causes hippocampal atrophy when compared with a sample of a normal subject in which the hippocampus is not atrophied (also referred to as “non-hippocampal non-atrophic subject (subject)” or “non-hippocampal atrophic subject” in the present specification).
- a sample of a patient also referred to as “hippocampal atrophy patient”, “hippocampal atrophy subject (subject)” or “hippocampal atrophy person” in the present specification
- the expression level thereof depending on the type of the target nucleic acid. Increases or decreases (hereinafter referred to as "increase / decrease").
- the kit or device of the present invention measures the expression level of the target nucleic acid in a body fluid derived from a subject to be tested for hippocampal atrophy, for example, a subject suspected of having hippocampal atrophy (for example, a human) and a body fluid derived from a non-hippocampal atrophy subject. It can be effectively used to compare them and detect hippocampal atrophy.
- the kit or device of the present invention can contain at least one nucleic acid for detecting hippocampal atrophy, for example, a nucleic acid probe or primer.
- the kit or device of the present invention comprises (or consists of) a nucleotide sequence represented by any of SEQ ID NOs: 1 to 109, or a nucleotide sequence in which u is t in the nucleotide sequence.
- at least one fragment can be contained.
- the kit or device of the present invention further comprises (or consists of) a nucleotide sequence represented by any of SEQ ID NOs: 110 to 200, or a nucleotide sequence in which u is t in the nucleotide sequence.
- a nucleotide sequence represented by any of SEQ ID NOs: 110 to 200, or a nucleotide sequence in which u is t in the nucleotide sequence.
- the fragment that can be included in the kit or device of the present invention is, for example, one or more, preferably two or more polynucleotides selected from the group consisting of (1) and (2) below. .. (1) A polynucleotide containing 15 or more consecutive base sequences in a base sequence in which u is t in the base sequence represented by any of SEQ ID NOs: 1 to 109 or a complementary sequence thereof. (2) A polynucleotide containing 15 or more consecutive base sequences in a base sequence in which u is t in the base sequence represented by any of SEQ ID NOs: 110 to 200 or a complementary sequence thereof.
- the polynucleotide is a polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence in which u is t in the base sequence, or a polynucleotide consisting of a complementary sequence thereof.
- the polynucleotide is a polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence in which u is t in the base sequence, or a polynucleotide consisting of a complementary sequence thereof.
- the fragment can be a polynucleotide containing 15 or more, preferably 17 or more, more preferably 19 or more consecutive bases.
- the size of a polynucleotide fragment is, for example, in the base sequence of each polynucleotide, from 15 to less than the total number of bases in the sequence, from 17 to less than the total number of bases in the sequence, and from 19 to less than the total number of bases in the sequence.
- the number of bases in the range such as.
- the kit or device of the present invention may contain one nucleic acid (polynucleotide) that specifically binds to the polynucleotide that is the above-mentioned hippocampal atrophy marker (target nucleic acid) or the complementary strand of the polynucleotide, or 2 It may contain more than one combination.
- nucleic acid polynucleotide
- target nucleic acid target nucleic acid
- 2 complementary strand of the polynucleotide
- the combination of the polynucleotides contained in the kit or device of the present invention that can specifically bind to the target nucleic acid specifically comprises the base sequences represented by SEQ ID NOs: 1 to 740 shown in Table 1. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more (eg, 11, 12) of the above polynucleotides derived from or derived from them.
- the target nucleic acid of the nucleic acid eg, nucleic acid probe or primer contained in the kit or device of the invention is at least one polynucleotide (eg, miR-) selected from the hippocampal atrophy markers of Group A above.
- the target nucleic acid of the nucleic acid eg, nucleic acid probe or primer contained in the kit or device of the present invention is at least one polynucleotide selected from the above-mentioned group A hippocampal atrophy markers and its selection. It may be a combination of at least one polynucleotide selected from the above-mentioned group A hippocampal atrophy markers excluding the above-mentioned polynucleotides and at least one polynucleotide selected from the above-mentioned group B hippocampal atrophy markers.
- the target nucleic acid of the nucleic acid (eg, a nucleic acid probe or primer) contained in the kit or device of the invention is at least one polynucleotide selected from the above group A hippocampal atrophy markers and the above. It may be a combination with at least one polynucleotide selected from the Kaiba atrophy markers of Group B.
- the target nucleic acid in the kit or device of the invention is at least one polynucleotide selected from the above-mentioned group A hippocampal atrophy markers and the above-mentioned group A hippocampal atrophy excluding the selected polynucleotide. It may be a combination of a marker and at least one polynucleotide selected from the group consisting of the hippocampal atrophy markers of Group B above.
- the combination of hippocampal atrophy markers is shown in SEQ ID NOs: 1 to 740 shown in Table 1. It is desirable to combine two or more of the polynucleotides consisting of the represented base sequence.
- the combination of hippocampal atrophy markers (target nucleic acids) is 2 or more, 3 or more, 4 or more, 5 or more of the above polynucleotides consisting of the nucleotide sequences represented by the SEQ ID NOs: Tables 3 to 6.
- any two or more of the above polynucleotides consisting of the nucleotide sequences represented by SEQ ID NOs: 1 to 200 may be combined.
- the combination preferably comprises at least one polynucleotide selected from the group of polynucleotides consisting of the nucleotide sequences represented by SEQ ID NOs: 1-109, which are newly found as markers of hippocampal atrophy.
- kits or devices for discriminating a hippocampal atrophied person as a non-coarma atrophied person in the present invention as a combination of hippocampal atrophy markers (target nucleic acids), SEQ ID NOs: 1-3, 68, 110, 111, 114 It is desirable to combine any two of the above polynucleotides consisting of the nucleotide sequences represented by 127, 135, 137, 142, 149, 152 and 155, or the corresponding miRNA or human miRNA.
- the combination is at least one polynucleotide selected from the group of polynucleotides consisting of the nucleotide sequences represented by SEQ ID NOs: 1-3 and 68 newly discovered as markers of hippocampal atrophy, or the corresponding miRNAs or humans. It preferably contains miRNA.
- kits or device for distinguishing a hippocampal atrophied person from a hippocampal non-atrophied person in the present invention as a combination of hippocampal atrophy markers (target nucleic acids), SEQ ID NOs: 1, 26, 65, 66, 71, 110 , 112, 114, 125-127, 162 and 164-166, or any two of the above polynucleotides consisting of the nucleotide sequences represented by, or the corresponding miRNA or human miRNA.
- target nucleic acids target nucleic acids
- SEQ ID NOs: 1, 26, 65, 66, 71, 110 , 112, 114, 125-127, 162 and 164-166 or any two of the above polynucleotides consisting of the nucleotide sequences represented by, or the corresponding miRNA or human miRNA.
- the combination is at least one polynucleotide selected from the group of polynucleotides consisting of the nucleotide sequences represented by SEQ ID NOs: 1, 26, 65, 66 and 71 newly discovered as markers of hippocampal atrophy, or the corresponding polynucleotides. It is preferable to include miRNA or human miRNA.
- hippocampal atrophy markers target nucleic acids
- any two of the above polynucleotides consisting of the nucleotide sequence represented by the above, or the corresponding miRNA or human miRNA is desirable to combine any two of the above polynucleotides consisting of the nucleotide sequence represented by the above, or the corresponding miRNA or human miRNA.
- the combination is a newly discovered marker for hippocampal atrophy, SEQ ID NOs: 1-7, 9-14, 16-21, 23-26, 30-32, 34, 35, 37-48, 50-56, 58- At least one polynucleotide selected from the group of polynucleotides consisting of the nucleotide sequences represented by 63, 67-73, 75, 77-93, 95-106 and 108-109, or the corresponding miRNA or human miRNA. It is preferable to include it.
- kits or devices for distinguishing a hippocampal atrophied person from a hippocampal non-atrophied person in the present invention as a combination of hippocampal atrophy markers (target nucleic acids), SEQ ID NOs: 1 to 5, 9, 10, 12, 13 , 16-18, 20, 21, 24-26, 29-32, 37-39, 41, 42, 44, 45, 47-59, 61-63, 67-71, 73-75, 77-88, 90 To 94, 96 to 102, 104 to 111, 113, 114, 117, 118, 121, 123, 125, 127, 128, 133 to 138, 141 to 143, 145 to 150, 152 to 159, 161-163, 167.
- hippocampal atrophy markers target nucleic acids
- any two of the above polynucleotides consisting of the nucleotide sequences represented by ⁇ 169, 171-180, 182, 183, 185 ⁇ 194 and 196 ⁇ 200, or the corresponding miRNA or human miRNA.
- the combination is a newly discovered marker for hippocampal atrophy in SEQ ID NOs: 1-5, 9, 10, 12, 13, 16-18, 20, 21, 24-26, 29-32, 37-39, 41, From a group of polynucleotides consisting of nucleotide sequences represented by 42, 44, 45, 47-59, 61-63, 67-71, 73-75, 77-88, 90-94, 96-102, 104-109. It preferably comprises at least one polynucleotide selected, or a corresponding miRNA or human miRNA.
- kits or devices for distinguishing a hippocampal atrophied person from a hippocampal non-atrophied person in the present invention as a combination of hippocampal atrophy markers (target nucleic acids), SEQ ID NO: 155, 188, 127, 5, 198, 189. , 110, 176, 143, 102, 148, 101, 75, 109, 199, 145, 54, 174, 106, 152, 133, 2, 20, 51 and 114. It is desirable to combine any two of these, or the corresponding miRNA or human miRNA.
- the combination is selected from the group of polynucleotides consisting of the nucleotide sequences represented by SEQ ID NOs: 2, 5, 20, 51, 54, 75, 101, 102, 106, and 109 newly discovered as Kaiba atrophy markers. It is preferred to include at least one polynucleotide to be made, or the corresponding miRNA or human miRNA.
- kits or devices for distinguishing a hippocampal atrophied person from a hippocampal non-atrophied person in the present invention as a combination of hippocampal atrophy markers (target nucleic acids), SEQ ID NOs: 1 to 6, 9, 17, 21, 27.
- the combination is a newly discovered marker of hippocampal atrophy in SEQ ID NOs: 1-6, 9, 17, 21, 27, 34, 39, 43, 47-48, 51, 53-54, 59, 61-63, At least one selected from the group of polynucleotides consisting of the nucleotide sequences represented by 71, 73-78, 80, 82-88, 90-92, 94, 96-98, 100, 102, 104, 106 and 109. It preferably contains a polynucleotide, or a corresponding miRNA or human miRNA.
- the number of combinations of polynucleotides, which are hippocampal atrophy markers (target nucleic acids) for discriminating the above hippocampal atrophy is 1, 2, 3, 4, 5, 6, 6, 7, 8, and so on. 9, 10 or more (eg, 11, 12, 13, 14, 15, 16, 16, 17, 18, 19, 20, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40 , 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, or 70 or more, 80 or more, 90 or more, or 100 or more) It may be either. It is preferably a combination of two or more.
- hippocampal atrophy marker used in the present invention
- the polynucleotide consisting of the nucleotide sequence represented by SEQ ID NO: 110 and the hippocampal atrophy marker selected from Tables 3 to 6
- the combination with the polynucleotide is illustrated.
- the combination of hippocampal atrophy markers (target nucleic acids) may be a combination of miRNA or human miRNA corresponding to the following combinations.
- hippocampal atrophy marker used in the present invention
- a polynucleotide consisting of the nucleotide sequence represented by SEQ ID NO: 39 and a polynucleotide of the hippocampal atrophy marker selected from Tables 3 to 6 The combination of is illustrated below.
- the combination of hippocampal atrophy markers (target nucleic acids) may be a combination of miRNA or human miRNA corresponding to the following combinations.
- hippocampal atrophy marker used in the present invention
- the combination with nucleotides is illustrated below.
- the combination of hippocampal atrophy markers (target nucleic acids) may be a combination of miRNA or human miRNA corresponding to the following combinations.
- Combination (84) Combination of SEQ ID NOs: 155, 87, 188, 127, 1, 161, 198, 110, 175, 109, 169, 145, 178, 10, 92, 133, 159, 82, 104, 114 (85).
- the combination of the hippocampal atrophy marker (target nucleic acid) used in the present invention the polynucleotide consisting of the nucleotide sequence represented by SEQ ID NO: 1 and the polypharmaceutical atrophy marker poly selected from Tables 3 to 6 Examples of combinations with nucleotides or polynucleotides consisting of complementary sequences thereof are shown below.
- the combination of hippocampal atrophy markers (target nucleic acids) may be a combination of miRNA or human miRNA corresponding to the following combinations.
- Combination (16) Combination number 121, 188, 127, 189, 154, 110, 102, 177, 100, 109, 3, 54, 174, 143, 159, 2, 82, 104, 114 Combination (17) SEQ ID NO: Combination of 137, 142, 188, 127, 161, 110, 75, 109, 145, 178, 10, 96, 174, 133, 39, 159, 82, 104, 114 (18) SEQ ID NO: 87, 188, 127, Combination of 196, 161, 154, 110, 100, 109, 135, 71, 73, 152, 159, 2, 82, 182, 114 (19) SEQ ID NO: 155, 87, 188, 127, 196, 194, 110, Combination of 4, 193, 175, 109, 145, 178, 48, 10, 59, 71, 159, 82, 114 (20) SEQ ID NO: 155, 142, 188, 127,
- Combination (32) Combination of SEQ ID NO: 155, 137, 87, 188, 127, 196, 189, 110, 193, 100, 109, 48, 71, 73, 152, 133, 91, 159, 2, 82, 114 ( 33) Combination of SEQ ID NOs: 155, 142, 188, 127, 53, 189, 154, 110, 78, 185, 75, 109, 145, 174, 152, 133, 2, 82, 93, 114 (34) SEQ ID NO: Combination of 155, 188, 127, 196, 154, 110, 176, 102, 69, 175, 75, 100, 109, 145, 54, 108, 73, 133, 159, 82, 114 (35) SEQ ID NO: 121, Combination of 155, 137, 87, 142, 188, 127, 30, 196, 154, 110, 100, 109, 145, 135, 71,
- hippocampal atrophy marker used in the present invention
- a polynucleotide consisting of the nucleotide sequence represented by SEQ ID NO: 100 and a hippocampal atrophy marker selected from Tables 3 to 6 are used.
- the combination with the polynucleotide is illustrated below.
- the combination of hippocampal atrophy markers (target nucleic acids) may be a combination of miRNA or human miRNA corresponding to the following combinations.
- hippocampal atrophy marker used in the present invention
- the combination with nucleotides is illustrated below.
- the combination of hippocampal atrophy markers (target nucleic acids) may be a combination of miRNA or human miRNA corresponding to the following combinations.
- a combination of a polynucleotide consisting of the base represented by SEQ ID NO: 5 or a complementary sequence thereof and a polynucleotide selected from Tables 3 to 6 or a polynucleotide consisting of a complementary sequence thereof is described below. Illustrate to.
- a combination of a polynucleotide consisting of the base represented by SEQ ID NO: 77 or a complementary sequence thereof and a polynucleotide selected from Tables 3 to 6 or a polynucleotide consisting of a complementary sequence thereof is described below. Illustrate to.
- hippocampal atrophy marker used in the present invention
- the combination with nucleotides is illustrated below.
- the combination of hippocampal atrophy markers (target nucleic acids) may be a combination of miRNA or human miRNA corresponding to the following combinations.
- hippocampal atrophy marker used in the present invention
- the polynucleotide consisting of the base represented by SEQ ID NO: 4 and the polynucleotide of the hippocampal atrophy marker selected from Tables 3 to 6 The combination with is illustrated below.
- the combination of hippocampal atrophy markers (target nucleic acids) may be a combination of miRNA or human miRNA corresponding to the following combinations.
- the kit or device of the present invention is a nucleic acid capable of specifically binding to the target nucleic acid which is the above-mentioned hippocampal atrophy marker, preferably the polynucleotides of the above-mentioned hippocampal atrophy marker described in groups A and B.
- the kit or device of the present invention enables detection of hippocampal atrophy in addition to the nucleic acid (nucleic acid for detecting hippocampal atrophy) that can specifically bind to the polynucleotide that is the marker of hippocampal atrophy in the present invention described above.
- Nucleic acids such as other known polynucleotides or polynucleotides that may be found in the future can also be included.
- the kit or device of the present invention contains known dementia test markers such as amyloid ⁇ protein and tau protein, in addition to the nucleic acid that can specifically bind to the polynucleotide that is the hippocampal atrophy marker in the present invention described above.
- Imaging tests such as MRI, CT, SPECT, PET, MIBG myocardial scintigraphy that visualize those markers may also be used in combination with the kits or devices of the invention.
- kit or device of the present invention can also be used in combination with a neuropsychological test or test such as the MMSE.
- the nucleic acids for detecting hippocampal atrophy contained in the kit of the present invention can be packaged in different containers individually or in any combination.
- the kit of the present invention is at least one selected from reagents for extracting nucleic acids (for example, total RNA) from samples such as body fluids, cells or tissues, labeling fluorescent substances, nucleic acid amplification enzymes and media, instructions for use, and the like. May include one.
- nucleic acids for example, total RNA
- samples such as body fluids, cells or tissues, labeling fluorescent substances, nucleic acid amplification enzymes and media, instructions for use, and the like. May include one.
- the device of the present invention may be a device for measuring or detecting a hippocampal atrophy marker in which the nucleic acid for detecting hippocampal atrophy in the present invention is bound or adhered to a solid phase, for example.
- a solid phase for example.
- the material of the solid phase may be plastic, paper, glass silicon, or the like. From the viewpoint of ease of processing, the preferred solid phase material may be plastic.
- the shape of the solid phase is arbitrary, for example, a square shape, a round shape, a strip shape, a film shape, or the like.
- the device of the present invention may be, for example, a device for measurement by a hybridization technique, and specific examples thereof include a blotting device, a nucleic acid array (for example, a microarray, a DNA chip, an RNA chip, etc.).
- a blotting device for example, a blotting device, a nucleic acid array (for example, a microarray, a DNA chip, an RNA chip, etc.).
- Nucleic acid arrays are high-density dispensers called spotters or layerers on the surface of a solid phase (substrate) that has been subjected to surface treatment such as introduction of functional groups such as L-lysine coat, amino group, and carboxyl group as needed.
- a method of spotting nucleic acid (single-stranded or double-stranded nucleic acid probe) using ) Can be produced by binding or adhering nucleic acids one by one by using a method such as spraying on a solid phase or sequentially performing nucleotide synthesis on the solid phase.
- the target nucleic acid can be measured by utilizing hybridization using this nucleic acid array.
- the kit or device of the present invention comprises at least one, preferably at least two, more preferably at least three, most preferably at least five to all polynucleotides of the miRNAs that are markers of hippocampal atrophy in Group A above. It contains a nucleic acid that can specifically bind to each of the complementary strands of the polynucleotide.
- the kit or device of the present invention comprises at least one, preferably at least two, more preferably at least three, most preferably at least five to all polyRNAs, which are markers of hippocampal atrophy in Group B above. Nucleic acids that can specifically bind to each of the nucleotides or complementary strands of the polynucleotide can be further included.
- the kit or device of the present invention is, for example, the following 4. It can be suitably used for a method for detecting hippocampal atrophy. That is, the present invention also provides kits and devices for detecting or diagnosing hippocampal atrophy.
- hippocampal atrophy marker target nucleic acid
- its combination described above for the kit or device of the present invention are also applied to the method, use, etc. according to the present invention.
- the expression level of the polynucleotide which is a marker for hippocampal atrophy in the present invention, is measured in a sample of a subject, and the hippocampus of the subject is atrophied using the measured expression level.
- methods for detecting hippocampal atrophy including assessing the presence or absence.
- the present invention also measures the expression level of the polynucleotide, which is a hippocampal atrophy marker in the present invention, in the sample of the subject, and uses the measured expression level to determine whether or not the hippocampus of the subject is atrophied.
- the present invention measures the expression level of a polynucleotide, which is a marker for hippocampal atrophy in the present invention, in a sample of a subject, and evaluates whether or not the hippocampus of the subject is atrophied using the measured expression level.
- a polynucleotide which is a marker for hippocampal atrophy in the present invention
- the present invention presents the expression level of at least one hippocampal atrophy marker gene or miRNA selected from group A and, optionally, the expression of at least one hippocampal atrophy marker gene or miRNA selected from group B in the sample.
- the present invention relates to a method for detecting hippocampal atrophy, which comprises measuring the amount and evaluating the expression level thereof as to whether or not the hippocampus of a subject is atrophic (for example, evaluated in vitro).
- the method for detecting hippocampal atrophy of the present invention may also be a method for detecting nerve cell death in the hippocampus.
- the methods of the invention allow for a minimally invasive, sensitive and specific quantitative and objective diagnosis of hippocampal atrophy, which results in early medical intervention and suppression of progression, as well as monitoring disease exacerbations. It is preferable to be able to monitor the effectiveness of surgical and medication treatments.
- the expression level of the hippocampal atrophy marker gene or miRNA is, for example, a nucleic acid (for example, a probe or a primer) capable of specifically binding to the polynucleotide which is the hippocampal atrophy marker in the present invention, or the kit of the present invention. Alternatively, it can be measured using a device, but is not limited thereto.
- the present invention is derived from a subject-derived nucleic acid (nucleic acid for detecting hippocampal atrophy; for example, a probe or primer) that can specifically bind to a polynucleotide that is a marker of hippocampal atrophy in the present invention, or a kit or device of the present invention. Also provided is the use for in vitro detection of a target nucleic acid of a hippocampal atrophy marker in a sample (eg, a miRNA precursor that is an expression product of a miRNA gene or a miRNA derived from it).
- a target nucleic acid of a hippocampal atrophy marker in a sample (eg, a miRNA precursor that is an expression product of a miRNA gene or a miRNA derived from it).
- the method of the present invention is useful for diagnosing the degree of hippocampal atrophy or detecting the presence or absence of hippocampal atrophy.
- the detection of hippocampal atrophy of the present invention refers to a nucleic acid for detecting hippocampal atrophy (eg, a nucleic acid probe or primer), eg, the present invention, for a sample obtained from a subject to be tested for hippocampal atrophy, eg, a subject suspected of having hippocampal atrophy.
- hippocampal atrophy marker gene / miRNA in vitro using a nucleic acid for detecting hippocampal atrophy (for example, a nucleic acid probe or a primer) contained in the kit or device of.
- a nucleic acid for detecting hippocampal atrophy for example, a nucleic acid probe or a primer
- the method for detecting hippocampal atrophy according to the present invention is a known or developmental therapeutic agent (as a non-limiting example, Aricept, for the purpose of treating or ameliorating hippocampal atrophy or a disease associated therewith, for example, in a hippocampal atrophy subject. It can also be used to evaluate or diagnose the presence or absence or degree of improvement in hippocampal atrophy or disease when GE Aricept, memantine, galantamine, or rivastigmine, or a combination thereof) is administered.
- Aricept a known or developmental therapeutic agent
- the method of the invention is described, for example, in steps (a), (b) and (c) below:
- B A step of measuring the expression level of a hippocampal atrophy marker (target nucleic acid) in a sample using the nucleic acid as a nucleic acid probe or primer.
- C From the results of (b), the step of evaluating the presence or absence (presence or absence) of hippocampal atrophy (nerve cell death) in the subject, or obtaining an index for the evaluation. Can be included.
- the present invention presents the hippocampal atrophy markers miR-3131, miR-6757-5p, miR-4706, miR-5001-5p, miR-3180-3p, miR-642b-3p, miR- 4655-5p, miR-6819-5p, miR-937-5p, miR-4688, miR-6471-5p, miR-7107-5p, miR-4271, miR-1229-5p, miR-4707-5p, miR- 6808-5p, miR-4656, miR-6076, miR-6762-5p, miR-7109-5p, miR-6732-5p, miR-3195, miR-7150, miR-642a-3p, miR-1249-5p, miR-3185, miR-4689, miR-3141, miR-6840-3p, miR-3135b, miR-1914-3p, miR-4446-3p, miR-4433b-3p, miR-6877-5p, miR-6884- 5p,
- the expression level of a polynucleotide (for example, 1 or 2 or more, preferably 3 or more, 10 or more, more preferably 3 to 30 polynucleotides; target nucleic acid) is measured, and the measured expression level is used.
- a method for detecting hippocampal atrophy which comprises assessing whether or not a subject's hippocampus is atrophic.
- the methods of the invention are miR-3131, miR-6757-5p, miR-4706, miR-5001-5p, miR-3180-3p, miR-642b-3p, miR-4655-5p, miR. -6819-5p, miR-937-5p, miR-4688, miR-6471-5p, miR-7107-5p, miR-4271, miR-1229-5p, miR-4707-5p, miR-6808-5p, miR -4656, miR-6076, miR-6762-5p, miR-7109-5p, miR-6732-5p, miR-3195, miR-7150, miR-642a-3p, miR-1249-5p, miR-3185, miR -4689, miR-3141, miR-6840-3p, miR-3135b, miR-1914-3p, miR-4446-3p, miR-4433b-3p, miR-6877-5p, miR-6884-5p,
- miR-3131 is hsa-miR-3131
- miR-6757-5p is hsa-miR-6757-5p
- miR-4706 is hsa-miR-4706.
- miR-5001-5p is hsa-miR-5001-5p
- miR-3180-3p is hsa-miR-3180-3p
- miR-642b-3p is hsa-miR-642b-3p, and so on.
- miR-4655-5p is hsa-miR-4655-5p
- miR-6819-5p is hsa-miR-6819-5p
- miR-937-5p is hsa-miR-937-5p
- miR- 4688 is hsa-miR-4688
- miR-6471-5p is hsa-miR-6741-5p
- miR-7107-5p is hsa-miR-7107-5p
- miR-4271 is hsa-miR-.
- miR-1229-5p is hsa-miR-1229-5p
- miR-4707-5p is hsa-miR-4707-5p
- miR-6808-5p is hsa-miR-6808-5p.
- miR-4656 is hsa-miR-4656
- miR-6076 is hsa-miR-6076
- miR-6762-5p is hsa-miR-6762-5p
- miR-7109-5p is hsa- miR-7109-5p
- miR-6732-5p is hsa-miR-6732-5p
- miR-3195 is hsa-miR-3195
- miR-7150 is hsa-miR-7150
- miR- 642a-3p is hsa-miR-642a-3p
- miR-1249-5p is hsa-miR-1249-5p
- miR-3185 is hsa-miR-3185
- miR-4689 is hsa-miR- 4689
- miR-3141 is hsa-miR-3141
- miR-6840-3p is hsa-miR-6840-3p
- miR-7110-5p miR-1275 is hsa-miR-1275
- miR-6779-5p is hsa-miR-6779-5p
- miR-197-5p is hsa-miR-197-5p Yes
- miR-6845-5p is hsa-miR-6845-5p
- miR-4327 is hsa-miR-4327
- miR-4723-5p is hsa-miR-4723-5p
- miR-4530 is.
- miR-6771-5p is hsa-miR-6771-5p
- miR-614 is hsa-miR-614
- miR-92a-2-5p is hsa-miR-92a- 2-5p
- miR-6891-5p is hsa-miR-6891-5p
- miR-6124 is hsa-miR-6124
- miR-4687-3p is hsa-miR-4487-3p, and so on.
- miR-4442 is hsa-miR-4442
- miR-7977 is hsa-miR-7977
- miR-6785-5p is hsa-miR-6785-5p
- miR-4497 is hsa-miR-4497.
- miR-8071 is hsa-miR-8071
- miR-663b is hsa-miR-663b
- miR-3180 is hsa-miR-3180
- miR-4251 is hsa-miR-4251, and so on.
- miR-1285-3p is hsa-miR-1285-3p
- miR-6870-5p is hsa-miR-6870-5p
- miR-4484 is hsa-miR-4484
- miR-4476 is hsa-. It is miR-4476 and miR-6.
- miR-4454 is hsa-miR-4454
- miR-6893-5p is hsa-miR-6893-5p
- miR-6085 is hsa-miR- 6085
- miR-4787-5p is hsa-miR-4787-5p
- miR-149-3p is hsa-miR-149-3p
- miR-7704 is hsa-miR-7704
- miR- 6125 is hsa-miR-6125
- miR-6090 is hsa-miR-6090
- miR-3197 is hsa-miR-3197
- miR-6850-5p is hsa-miR-6850-5p, and so on.
- miR-4467 is hsa-miR-4467
- miR-6885-5p is hsa-miR-6885-5p
- miR-6803-5p is hsa-miR-6803-5p
- miR-6798-5p is.
- miR-6780b-5p is hsa-miR-6780b-5p
- miR-6768-5p is hsa-miR-6768-5p
- miR-5100 is hsa-miR- It is 5100
- miR-6724-5p is hsa-miR-6724-5p
- miR-6879-5p is hsa-miR-6879-5p
- miR-7108-5p is hsa-miR-7108-5p.
- miR-4649-5p is hsa-miR-4649-5p
- miR-4739 is hsa-miR-4739
- miR-6089 is hsa-miR-6089
- miR-1908-5p is hsa-. It is miR-1908-5p
- miR-4516 is hsa-miR-4516
- miR-2861 is hsa-miR-2861
- miR-4492 is hsa-miR-4492
- miR-4294 is hsa-.
- miR-6791-5p is hsa-miR-6791-5p
- miR-1469 is hsa-miR-1469
- miR-6752-5p is hsa-miR-6752-5p
- miR-4730 is hsa-miR-4730
- miR-6126 is hsa-miR-6126
- miR-6869-5p is hsa-miR-6869-5p
- miR-1268a is hsa-miR-1268.
- miR-6799-5p is hsa-miR-6799-5p
- miR-8069 is hsa-miR-8069
- miR-3621 is hsa-miR-3621
- miR-4763-3p is It may be hsa-miR-4763-3p.
- Nucleic acids for detecting hippocampal atrophy used for measuring the expression level in a preferred embodiment of the method of the present invention include the following polynucleotides (a) to (e): (A) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases. That fragment, (B) A polynucleotide containing the base sequence represented by any of SEQ ID NOs: 1 to 109, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (C) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 1 to 109 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more. Its fragment, which contains consecutive bases of (D) A polynucleotide containing a nucleotide sequence represented by any of SEQ ID NOs: 1 to 109 or a nucleotide sequence complementary to a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more.
- the nucleic acids for detecting hippocampal atrophy used to measure expression levels in the methods of the invention are further miR-1228-5p, miR-760, miR-187-. 5p, miR-7111-5p, miR-6088, miR-6805-3p, miR-4640-5p, miR-6721-5p, miR-6880-5p, miR-711, miR-1281-5p, miR- 4525, miR-486-3p, miR-6756-5p, miR-1260b, miR-3184-5p, miR-6075, miR-204-3p, miR-4728-5p, miR-4534, miR-4758-5p, miR-8063, miR-6863-3p, miR-6789-5p, miR-744-5p, miR-1909-3p, miR-887-3p, miR-4745-5p, miR-4433a-3p, miR-5090, miR-296-5
- such additional nucleic acids for detecting hippocampal atrophy include the polynucleotides (f)-(j) below: (F) Containing a nucleotide consisting of the base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence in which u is t in the base sequence, a mutant thereof, a derivative thereof, or 15 or more consecutive bases. That fragment, (G) A polynucleotide containing a base sequence represented by any of SEQ ID NOs: 110 to 200, a variant thereof, a derivative thereof, or a fragment thereof containing 15 or more consecutive bases.
- (H) A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 110 to 200 or a base sequence complementary to a base sequence in which u is t in the base sequence, a variant thereof, a derivative thereof, or 15 or more. Its fragment, which contains consecutive bases of (I) A polynucleotide containing a nucleotide sequence represented by any of SEQ ID NOs: 110 to 200 or a nucleotide sequence complementary to a nucleotide sequence in which u is t in the nucleotide sequence, a variant thereof, a derivative thereof, or 15 or more.
- the specimen used in the method of the present invention may be any specimen of the subject and includes biological tissue (preferably brain tissue or nerve tissue) derived from the subject, body fluid or cell, or a specimen prepared from the tissue. do.
- the sample is, for example, an RNA-containing sample prepared from the biological tissue, a sample containing a polynucleotide further prepared from the sample, a body fluid such as blood, serum, plasma, urine, or a part or all of the biological tissue of the subject. It is a living tissue collected by such means or surgically removed. From these samples, a sample to be measured can be prepared by a conventional method.
- Total RNA may be prepared from a sample and used for measuring the expression level
- various polynucleotides such as cDNA may be prepared from RNA such as total RNA and used for measuring the expression level.
- the hippocampal atrophy marker gene / miRNA may be extracted from the sample.
- a sample such as blood, serum, plasma, etc.
- 3D-Gene (registered trademark) RNA extension reagent from simple sample kit (Toray Co., Ltd.)
- It is particularly preferable to use the reagent for RNA extraction in the medium but it is particularly preferable to use a general acidic phenol method (Acid Guanidinium-Phenol-Chloroform (AGPC) method), Trizol (registered trademark) (life technologies), Isogen (Nippon Gene), etc.
- An RNA extraction reagent containing the acidic phenol of the above may be used, or a kit such as miRNeasy (registered trademark) Mini Kit (Qiagen) may be used, but the present invention is not limited thereto.
- the subject to which the method of the present invention is applied is a mammal as defined above, and may be, for example, non-limitingly human, monkey, mouse, rat, etc., preferably human.
- the steps may be changed according to the type of polynucleotide to be measured.
- the method for detecting hippocampal atrophy is, for example, the following steps (a), (b) and (c): (A) RNA prepared from subject specimens (where, for quantitative RT-PCR in step (b), eg, the 3'end of RNA may be polyadenylated, or either or both. An arbitrary sequence may be added to the end by a ligation method or the like), or a polynucleotide (cDNA) consisting of a complementary sequence thereof synthesized by reverse transcription is added to a nucleic acid for detecting hippocampal atrophy, for example, the kit of the present invention.
- RNA prepared from subject specimens where, for quantitative RT-PCR in step (b), eg, the 3'end of RNA may be polyadenylated, or either or both.
- An arbitrary sequence may be added to the end by a ligation method or the like), or a polynucleotide (cDNA) consisting of a complementary sequence thereof synthesized by reverse
- RNA derived from a sample bound to the nucleic acid or cDNA synthesized from the RNA is hybridized using the nucleic acid as a nucleic acid probe, or nucleic acid amplification using the nucleic acid as a primer, for example, quantitative RT-PCR.
- Steps to measure eg, quantify
- C A step of evaluating the presence or absence of hippocampal atrophy (or hippocampal atrophy / neuronal cell death, marker gene / miRNA) based on the measurement result of (b) above. Can be included.
- hybridization techniques can be used to measure the expression level according to the present invention.
- Such hybridization techniques are not limited to the following, but include, for example, nucleic acid array techniques such as Northern blotting (Northern hybridization method), Southern blotting (Southern hybridization method), and DNA chip analysis method, in situ hybridization. The law etc. can be mentioned.
- a PCR method such as quantitative RT-PCR, a nucleic acid amplification technique such as the LAMP method, or a next-generation sequencing method can be used.
- the presence or absence of expression of the hippocampal atrophy marker miRNA corresponding to the nucleic acid probe and its gene, and the expression level thereof are detected and measured.
- the nucleic acid probe of the present invention (for example, a complementary strand to the hippocampal atrophy marker miRNA) is labeled with a radioisotope (32 P, 33 P, 35 S, etc.) or a label such as a fluorescent substance, and is always labeled.
- RNA derived from the subject's sample transferred to a membrane such as a nylon membrane By hybridizing with RNA derived from the subject's sample transferred to a membrane such as a nylon membrane according to the method, a signal derived from the formed DNA / RNA double-stranded label (radioactive isotope, fluorescent substance, etc.) is detected by radiation.
- a method of detecting and measuring with a detector BAS-1800II (Fuji Film Co., Ltd.), etc.) or a fluorescence detector (STORM 865 (GE Healthcare Co., Ltd.), etc.
- BAS-1800II Fluji Film Co., Ltd.
- STORM 865 GE Healthcare Co., Ltd.
- the quantitative RT-PCR method for example, using the above-mentioned primers that can be used in the present invention, the presence or absence of expression of the hippocampal atrophy marker miRNA corresponding to the primers and the expression level thereof and the expression level thereof are detected and measured. can do.
- RNA derived from a sample of a subject can be recovered, the 3'end is polyadenylated, cDNA is prepared from the polyadenylated RNA according to a conventional method, and this can be included in the detection kit or device of the present invention as a template.
- a primer pair containing the primers of the present invention (preferably consisting of a normal chain sequence and a reverse chain sequence in which each primer specifically binds to the above-mentioned cDNA) is hybridized with the cDNA, and the PCR method is carried out by a conventional method.
- a method for detecting the single-stranded or double-stranded DNA obtained can be exemplified.
- a method for detecting single-stranded or double-stranded DNA a method for detecting a labeled signal by performing the above PCR using a primer labeled with a radioactive isotope or a fluorescent substance in advance, or an agarose gel for the PCR product.
- a method of detecting double-stranded DNA by staining with ethidium bromide or the like, and transferring the obtained single-stranded or double-stranded DNA to a membrane such as a nylon membrane according to a conventional method and hybridizing with a nucleic acid probe examples thereof include a method of detecting and detecting (Northern blot method).
- Nucleic acid array technology uses, for example, a nucleic acid for detecting hippocampal atrophy of the present invention, for example, a nucleic acid array in which a nucleic acid probe (single-stranded or double-stranded) contained in the kit or device of the present invention is immobilized on a solid phase (substrate). Use. The region where the nucleic acid probe is fixed is called a probe spot, and the region where the nucleic acid probe is not attached is called a blank spot.
- a nucleic acid probe single-stranded or double-stranded
- Nucleic acid groups fixed on a solid phase (substrate) generally have names such as nucleic acid chips (DNA chips or RNA chips), nucleic acid arrays (DNA arrays or RNA arrays), and microarrays (DNA microarrays or RNA microarrays).
- the term "nucleic acid array” is used as a general term for them.
- the DNA or RNA array includes a DNA or RNA macroarray and a DNA or RNA microarray.
- 3D-Gene (registered trademark) Human miRNA Oligo chip can be used as the DNA chip, but the DNA chip is not limited thereto.
- the measurement using the DNA chip is not limited to the following, but for example, a signal derived from a nucleic acid label for detecting hippocampal atrophy is detected by an image detector (Typhoon 9410 (GE Healthcare), 3D-Gene (registered trademark)). A method of detecting and measuring with a scanner (Toray Industries, Inc.), etc. can be exemplified.
- nucleic acid eg, a nucleic acid for detecting hippocampal atrophy of the invention
- background measurements e.g., a condition that enables hybridization to the target nucleic acid by the average of the values + the standard error of the background measurement value ⁇ the measurement value of 2 or more.
- Stringent conditions are defined by the reaction conditions of hybridization and subsequent washing.
- the hybridization condition is not limited to the following, but is a condition for reacting at 30 ° C. to 60 ° C. in a solution containing SSC, a surfactant, formamide, dextran sulfate, a blocking agent, etc. for 1 to 24 hours.
- 1 ⁇ SSC is an aqueous solution (pH 7.0) containing 150 mM sodium chloride and 15 mM sodium citrate
- the surfactant includes SDS (sodium dodecyl sulfate), Triton, Tween, and the like.
- Hybridization conditions more preferably include 3 to 10 ⁇ SSC and 0.1 to 1% SDS.
- washing conditions after hybridization are, for example, a solution containing 0.5 ⁇ SSC at 30 ° C. and 0.1% SDS, and 0.2 at 30 ° C. Conditions such as a solution containing ⁇ SSC and 0.1% SDS and continuous washing with a 0.05 ⁇ SSC solution at 30 ° C. can be mentioned. It is preferable that the complementary polynucleotide maintains a hybridized state with the positive chain of interest even when washed under such conditions.
- any nucleic acid amplification method such as PCR method or LAMP method can be used.
- PCR method any nucleic acid amplification method
- LAMP method for example, TaqMan (registered trademark) MicroRNA Assays of Life Technologies, miScript PCR System of Qiagen, LAMP method primer and the like can be used.
- the nucleic acid amplification method used may be used, but the method is not limited thereto.
- reaction conditions when carrying out PCR using the primer which is the nucleic acid for detecting hippocampal atrophy of the present invention of the present invention include, for example, 10 mM Tris-HCL (pH 8.3), 50 mM KCL, and 1 to 2 mM MgCl 2.
- treatment may be performed at a Tm value of +5 to 10 ° C. calculated from the base sequence of the primer for about 15 seconds to 1 minute.
- Tm value 2 ⁇ (number of adenine residues + number of thymine residues) + 4 ⁇ (number of guanine residues + number of cytosine residues).
- TaqMan registered trademark
- MicroRNA Assays Life Technologies
- LNA registered trademark
- MicroRNA PCR Exiqon
- Ncode registered trademark
- miRNA qRT-PCT kit A commercially available measurement kit specially devised for quantitatively measuring miRNA, such as (Invitrogen), may be used.
- the expression level may be measured by using a sequencer in addition to or as an alternative method to the above hybridization method.
- a sequencer any of a first-generation DNA sequencer based on the Sanger method, a second-generation sequencer having a short read size, and a third-generation sequencer having a long read size can be used.
- the second-generation and third-generation sequencers are also included, and are also referred to as "next-generation sequencers".
- Misseq / Hiseq / NexSeq (Illumina), Ion Proton / Ion PGM / Ion S5 / S5 XL (Thermo Fisher Scientific), PacBio RS II / Sequence ( Pacific Bioscience), Nanopores, etc.
- a commercially available measurement kit specially devised for measuring miRNA may be used by using MinION (Oxford Nanopore Technologies) or the like.
- the next-generation sequence is a base sequence determination technique using a next-generation sequencer, and is characterized in that a huge number of sequence reactions can be executed in parallel as compared with the Sanger method (for example, Rick Kamps et al., See Int. J. Mol. Sci., 2017, 18 (2), p. 308 and Int. Neurour. J., 2016, 20 (Suppl. 2), S76-83).
- the step of adding an adapter sequence having a predetermined base sequence to both ends of the sample-derived miRNA or its cDNA and before or before the addition of the adapter sequence.
- RNA total RNA
- cDNA derived from a specific target miRNA may be amplified by a nucleic acid amplification method such as PCR or concentrated using a probe or the like.
- a nucleic acid amplification method such as PCR or concentrated using a probe or the like.
- the details of the subsequent sequencing steps vary depending on the type of next-generation sequencer, but typically the cDNA is linked to the substrate via an adapter sequence and the adapter sequence is used as the priming site for the sequencing primer for the sequencing reaction.
- the base sequence For details of the sequence reaction and the like, for example, Rick Kamps et al. See (above). Finally, the data is output.
- a set of sequence information (reads) obtained by the sequence reaction and its analysis data can be obtained.
- the target miRNA can be identified based on the obtained sequence information, and the expression level thereof can be determined based on the number of reads having the base sequence of the target miRNA.
- the method for determining the expression level (for example, miRNA amount or gene expression level) in the present invention is not limited to the following, but is, for example, Static analog of gene expression microrary data (Speed T., Chapman and Hall / CRC), and A. Examples thereof include a method based on statistical processing described in's guide Microarray gene expression dataanalysis (written by Caston HC et al., Blackwell publishing). For example, it is obtained by adding a value of 2 times, preferably 3 times, more preferably 6 times the standard deviation of the measured value of the blank spot to the average value of the measured value of the blank spot on the nucleic acid array (DNA chip, etc.). A probe spot having a signal value equal to or higher than that value can be used as a detection spot.
- the average value of the measured values of the blank spot can be used as the background, subtracted from the measured value of the probe spot, and the obtained value can be used as the expression level.
- the missing value of the expression level is excluded from the analysis target, preferably replaced with the minimum value of the expression level obtained on each nucleic acid array (DNA chip, etc.), or more preferably the minimum value of the expression level. It can be replaced by a value obtained by subtracting 0.1 from the logarithmic value.
- 20% or more preferably 50% or more, more preferably 80% or more of the number of measurement samples is 2 to the 6th power, preferably 2 to the 8th power, more preferably.
- genes / miRNAs having an expression level of 2 to the 10th power or higher can be selected as analysis targets.
- the normalization of the expression level is not limited, and examples thereof include the methods described in global normalization and quantile normalization (Bolstad, BM et al., 2003, Bioinformatics, Vol. 19, p185-193). To be done.
- the method of the invention uses the hippocampus of a subject using the expression levels measured as described above and the control expression levels of the subject in which the hippocampus is not atrophied in the same manner.
- the method for detecting hippocampal atrophy is, for example, a sample (eg, blood, serum, etc.) collected from a subject to be tested for hippocampal atrophy, for example, a subject suspected to have hippocampal atrophy and a subject having no hippocampal atrophy (control).
- a subject to be tested for hippocampal atrophy including measuring the expression level of the gene / miRNA of the specific hippocampal atrophy marker (target nucleic acid) of the present invention and comparing both expression levels in a sample such as plasma), for example. If there is a difference (eg, statistically significant difference) between the expression level in a subject suspected of having hippocampal atrophy and the control expression level in a subject (control) in which the hippocampal is not atrophied, the hippocampal atrophy in the subject. Therefore, it can be evaluated that the subject's hippocampus is atrophied.
- a difference eg, statistically significant difference
- the subjects to be tested for hippocampal atrophy for example, the subject suspected to have hippocampal atrophy and the subject without hippocampal atrophy (control), which are compared with each other, are preferably the same species.
- evaluating whether or not the hippocampus is atrophied may be evaluation support based on the result of an in vitro test that is not judged by a doctor.
- the expression level of the polynucleotide consisting of the base sequence represented by at least one of, and optionally the base sequence represented by at least one of SEQ ID NOs: 110 to 200, is the blood of a subject in which the hippocampus is not atrophied.
- the expression level is high or low (preferably statistically significantly high or low) in the sample such as serum, plasma, urine, etc.
- the difference from the hippocampal group allows it to be evaluated that the hippocampus is atrophied in a subject to be tested for the hippocampal atrophy, for example, a subject suspected of having hippocampal atrophy.
- the expression level of the hippocampal atrophy marker is measured in the sample of the subject, the expression level in the sample of the subject (or patient) known to have atrophy of the hippocampus, and the atrophy of the hippocampus.
- the above was measured by a discriminant formula (discrimination function) capable of discriminating between atrophy and non-atrophy of the hippocampus prepared as a teacher sample based on the expression level in the sample of a subject known not to be atrophied.
- Substituting expression levels in a subject including assessing hippocampal atrophy or non-atrophy (whether or not the subject's hippocampus is atrophied) or obtaining an indicator of hippocampal atrophy or non-atrophy.
- a method for detecting hippocampal atrophy is provided.
- a discrimination score that correlates with the degree of hippocampal atrophy (such as one based on the volume ratio (%) of the hippocampus to the whole brain) can be mentioned.
- the creation of a discriminant is not limited to the following, but for example, marker candidates are selected based on statistically significant differences such as the Lasso method, difference (Fold Change), and p-value, and they are subjected to logistic regression analysis. And a method of constructing as a discriminant using Fisher's discriminant analysis method or the like.
- the present invention measures the expression level of the hippocampal atrophy marker in a sample of a subject, and the subject (or patient) whose hippocampus is known to be atrophied (for example, by icobrin measurement).
- a discriminant formula (discrimination) capable of distinguishing atrophy or non-atrophy of the hippocampus, which was created using the expression levels of each of the specimens of the sample and the sample of the subject known not to have atrophy of the hippocampus as a teacher sample.
- a method of determining the degree of hippocampal atrophy in a subject comprising substituting the expression level measured above into a function) and thereby assessing the degree of hippocampal atrophy.
- the present invention also measures the expression level of the hippocampal atrophy marker in the sample of the subject, and the sample of the subject (or the patient) whose hippocampus is known to be atrophied and the hippocampus are not atrophied.
- the expression measured above is used in a discriminant formula (discrimination function) capable of discriminating between atrophy and non-atrophy of the hippocampus prepared by using each expression level of a subject's sample known to be a teacher sample.
- a discriminant formula discrimination function
- Provided are methods of assisting in determining hippocampal atrophy in a subject, or obtaining an index of hippocampal atrophy in a subject, comprising substituting an amount and thereby obtaining an index of hippocampal atrophy.
- an index of the hippocampal atrophy degree for example, the volume ratio (%) of the hippocampus to the whole brain can be mentioned.
- the present invention measures the expression level of the hippocampal atrophy marker in a sample of a subject, and a sample of a subject (or a patient) whose hippocampus is known to be atrophic and a hippocampus are used.
- Substituting the expression level measured above into a regression equation that can predict the hippocampal volume created using the expression level of each sample of a subject known to be non-atrophic as a teacher sample, Provides a method of predicting hippocampal volume in a subject, including assessing hippocampal volume.
- the creation of the regression equation is not limited to the following, but for example, a lasso regression analysis or a regression analysis using the partial least squares method may be used.
- the present invention measures the expression level of the above-mentioned hippocampal atrophy marker in a plurality of subjects known to have atrophy of the hippocampus and a plurality of subjects known to have no atrophy of the hippocampus in vitro.
- First step the second step of creating a discrimination formula using the expression level of the hippocampal atrophy marker obtained in the first step as a teacher sample, and the expression level of the hippocampal atrophy marker in the sample of the subject.
- the third step measured in vitro by the same method as in step 1 the expression level of the hippocampal atrophy marker obtained in the third step is substituted into the discrimination formula obtained in the second step for discrimination.
- a fourth step may be included to evaluate whether the hippocampus of the subject whose expression level was measured in the third step is atrophied.
- the discriminant formula capable of discriminating between atrophy and non-shrinkage of the hippocampus is an arbitrary discriminant analysis method, for example, linear discriminant analysis such as Fisher's discriminant analysis, non-linear discriminant analysis by Maharanobis distance, and neural network. , C-SVC, etc.
- Support Vector Machine (SVM) Logistic Regression Analysis (In particular, LASSO (Last Absolute Shrinkage and Selection Operator) Method, Ridge Regression, Logistic Regression Using Ridge Regression, Logistic Regression Using Multiple logistic regression using), k-neighborhood method, decision tree, partial least squared (PLS) regression, etc.
- SVM Support Vector Machine
- Logistic Regression Analysis In particular, LASSO (Last Absolute Shrinkage and Selection Operator) Method, Ridge Regression, Logistic Regression Using Ridge Regression, Logistic Regression Using Multiple logistic regression using
- PLS partial least squared
- the linear discriminant analysis is a method of discriminating the affiliation of a group by using Equation 1 as a discriminant when the boundary of grouping is a straight line or a hyperplane.
- x is an explanatory variable
- w is a coefficient of the explanatory variable
- w 0 is a constant term.
- the value obtained by the discriminant is called a discriminant score, and the measured value of the newly given data set can be substituted into the discriminant as an explanatory variable, and the grouping can be discriminated by the code of the discriminant score.
- Fisher's discriminant analysis which is a type of linear discriminant analysis, is a dimension reduction method for selecting a dimension suitable for class discrimination, focusing on the variance of synthetic variables and data having the same label.
- a highly discriminant synthetic variable is constructed by minimizing the variance of (Venables, W.N. et al., Modeln Applied Statistics with S. Fourth edition., Springer., 2002).
- the projection direction w that maximizes Equation 2 is obtained.
- ⁇ is the average of the inputs
- ng is the number of data belonging to the class g
- ⁇ g is the average of the inputs of the data belonging to the class g.
- the numerator and denominator are the interclass variance and intraclass variance when the data is projected in the direction of the vector w, and the discriminant coefficient wi is obtained by maximizing this ratio (Kanamori Takafumi et al., "Pattern”. Recognition, Kyoritsu Publishing (Tokyo, Japan) (2009), Richard O. et al., Pattern Classification Section Edition., Willey-Interscience, 2000).
- the Mahalanobis distance is calculated by Equation 3 in consideration of the correlation of data, and can be used as a nonlinear discriminant analysis for discriminating a group having a close Mahalanobis distance from each group as a belonging group.
- ⁇ is the center vector of each group
- S -1 is the inverse matrix of the variance-covariance matrix of that group.
- the center vector is calculated from the explanatory variable x, and an average vector, a median vector, or the like can be used.
- SVM is V.I. It is a discriminant analysis method devised by Vapnik (The Nature of Statistical Learning Theory, Springer, 1995).
- a boundary surface called a hyperplane for correctly classifying the data set into a known grouping with a specific data item of a data set whose grouping to be classified is known as an explanatory variable and a grouping to be classified as an objective variable.
- the discrimination result at this time may be a group to be classified, a probability of being classified into a group to be classified, or a distance from a hyperplane.
- SVM as a method for dealing with a non-linear problem, a method of non-linearly transforming a feature vector to a higher dimension and performing linear discrimination in the space is known.
- An expression in which the inner product of two elements in a non-linearly mapped space is expressed only by the input in the original space is called a kernel.
- a linear kernel RBF (Radial Basis Function) Kernel and Gaussian kernel can be mentioned.
- C-support vector classification which is a type of SVM method, learns with two groups of explanatory variables to create a hyperplane, and determines which group the unknown data set is classified into.
- C-SVC discriminant An example of calculating the C-SVC discriminant that can be used by the method of the present invention is shown below.
- all subjects are grouped into two groups: patients with hippocampal atrophy and subjects without hippocampal atrophy.
- a numerical value obtained by quantifying the result of brain imaging (MRI) by icobrain can be used.
- a data set (hereinafter referred to as a learning sample group) consisting of comprehensive expression levels of the two divided groups of samples (for example, samples derived from serum) was prepared, and there was a clear difference in the expression levels between the two groups.
- the discriminant by C-SVC is determined by using the hippocampal atrophy marker in which the above is observed as the explanatory variable and the grouping as the objective variable (for example, -1 and +1).
- Equation 4 is an objective function to be optimized, where e is all input vectors, y is an objective variable, a is a Lagrange undetermined multiplier vector, Q is a positive-definite matrix, and C is a parameter for adjusting constraints.
- Equation 5 is the finally obtained discriminant, and the group to which it belongs can be determined by the sign of the value obtained by the discriminant.
- x is a support vector
- y is a label indicating the affiliation of a group
- a is a corresponding coefficient
- b is a constant term
- K is a kernel function.
- Equation 6 the RBF kernel defined by Equation 6 can be used.
- x is the support vector and ⁇ is the kernel parameter that adjusts the complexity of the hyperplane.
- Logistic regression is a multivariate analysis method in which one categorical variable (binary variable) is used as an objective variable and its occurrence probability is predicted using a plurality of explanatory variables, and is expressed by the following equation 7.
- the Lasso (Last Absolute Shockage and Selection Operator) method is one of the variable selection and adjustment methods when a large number of observed variables exist, and was proposed by Tibshirani (Tibshirani R., 1996, J.R., J.R., 1996). Ser B, Vol. 58, p267-88). Ridge regression is the oldest proposed regularization method and was proposed by Hoerl (Hoell, E.E., 1970, Technologies., Vol. 12, p. 55-67). The Elastic net is a model that linearly combines the Lasso method and Ridge regression, and was proposed by Zou (Zou, H., 2005, JR Stat Soc Ser B, Vol. 67, p. 301-320).
- the Lasso method has a feature of suppressing overfitting to the model and estimating some regression coefficients to 0 by introducing a penalty term when estimating the regression coefficient.
- Ridge regression can also estimate the coefficients of the model, and even when the explanatory variables are larger than the number of samples, it is possible to select more variables than the number of samples. If there is a strong correlation between the explanatory variables, only one of the highly correlated variables remains in the Lasso method and the others are estimated to be zero, but in Ridge regression both can be selected.
- the Elastic net can appropriately select variables even for highly correlated explanatory variables that are difficult to select by the Lasso method, and can generate a dimension-reduced model.
- the regression coefficient is estimated so as to maximize the log-likelihood function represented by Equation 8.
- the Principal Component Analysis method eliminates the correlation between variables of quantitative data described by many variables and reduces it to a small number of uncorrelated synthetic variables with as little information loss as possible. Pearson (PEARSON, K., 1901, Freedom Variable Magazine, Series 6, 2 (11), p559-572) and Hotelling (HOTELLING, H., 1933, Journal of Digital) 24, p417-441, p498-520).
- the principal component score obtained as a result of the principal component analysis can indicate the magnitude of the contribution to a certain data point.
- the method of the present invention is described in, for example, the following steps (a), (b) and (c):
- (A) The expression level of the hippocampal atrophy marker in the sample of the subject known to have atrophy of the hippocampus and the sample of the subject known not to have atrophy of the hippocampus is the nucleic acid for detecting hippocampus atrophy of the present invention. Steps to measure using (eg, probes or primers), kits or devices,
- B) A step of creating a discriminant of the above formulas 1 to 3, 5 and 6 from the measured value of the expression level measured in (a).
- (C) The expression level of the hippocampal atrophy marker in the sample of the (unknown) subject to be tested for hippocampal atrophy is measured using the hippocampal atrophy detection nucleic acid (for example, probe or primer), kit or device of the present invention. Then, by substituting it into the discriminant formula prepared in (b), it is determined or evaluated that the hippocampus of the subject is atrophied or not atrophied based on the obtained result, or the hippocampus is not atrophied. A step in which marker expression from an unknown subject with suspected atrophy is evaluated by comparing it with a control from a subject in which the hippocampus is not atrophied. Can be included.
- x in the formulas 1 to 3, 5 and 6 is an explanatory variable and includes a measured value of the expression level of the hippocampal atrophy marker.
- the explanatory variable for discriminating between the subject in which the hippocampus is atrophied and the subject in which the hippocampus is not atrophied in the present invention may be a measured value of the expression level of the hippocampal atrophy marker, for example, the following. May be the expression level of: A polynucleotide consisting of a base sequence represented by any of SEQ ID NOs: 1 to 109 and 110 to 200, a base sequence in which u is t in the base sequence, or a base sequence complementary thereto, or variants or derivatives thereof.
- a sample of a subject with atrophy of the hippocampus and a subject with no atrophy of the hippocampus measured using a nucleic acid for detecting hippocampal atrophy, which is a fragment thereof (for example, DNA) containing 15 or more consecutive nucleotides (for example, DNA). , Serum) expression level.
- one or more of the above can be used to determine or evaluate whether or not the subject has hippocampal atrophy (whether or not the subject's hippocampus is atrophied).
- a discriminant formula using the expression level of the hippocampal atrophy marker as an explanatory variable is required.
- the hippocampal atrophy patient group hippocampal atrophy subject group
- the subject group in which the hippocampus is not atrophied normal group / It is preferable to use a hippocampal atrophy marker having a clear difference in the expression level between the two groups consisting of the hippocampal non-atrophy subject group) in the discriminant formula.
- the hippocampal atrophy marker used as the explanatory variable of the discriminant is determined as follows. First, using the comprehensive expression level of the hippocampal atrophy patient group as the learning group and the comprehensive expression level of the subject group without hippocampal atrophy as a data set, the P value of the t-test, which is a parametric analysis, and Mann, which is a nonparametric analysis. -Using the P value of the Whitney U test or the P value of the Wilcoxon test, the magnitude of the difference in the expression level of each hippocampal atrophy marker between the two groups is determined.
- the risk rate (significance level) of the P value obtained by the test is smaller than, for example, 5%, 1% or 0.01%, it can be regarded as statistically significant.
- Bonferroni correction for example, the P value obtained by the test is multiplied by the number of repetitions of the test, that is, the number of hippocampal atrophy markers used in the analysis, and compared with the desired significance level. The probability of error can be suppressed.
- the absolute value (Fold change) of the median expression ratio of each expression level was calculated and discriminated between the expression level of the hippocampal atrophy patient group and the expression level of the subject group in which the hippocampal atrophy was not performed.
- the hippocampal atrophy marker used as the explanatory variable of the equation may be selected.
- a ROC curve may be created using the expression levels of the hippocampal atrophy patient group and the subject group in which the hippocampus is not atrophied, and the hippocampal atrophy marker used as an explanatory variable of the discriminant may be selected based on the AUROC value.
- a discriminant that can be calculated by the above-mentioned various methods is created by using an arbitrary number of hippocampal atrophy markers having a large difference in the expression level obtained here.
- a method of constructing a discriminant that obtains the maximum discrimination accuracy for example, a method of constructing a discriminant with any combination of hippocampal atrophy markers that satisfy the significance level of the P value, or a hippocampal atrophy marker used for creating a discriminant. Is repeatedly evaluated while increasing one by one in descending order of the difference in expression level (Furey TS. et al., 2000, Bioinformatics., Vol. 16, p906-14).
- the expression level of another independent hippocampal atrophy patient or subject whose hippocampus is not atrophied is substituted into the explanatory variable, and the group to which this independent hippocampal atrophy patient or subject whose hippocampus is not atrophied belongs.
- the discrimination result of is calculated. That is, for diagnostic purposes, more unbiased hippocampal atrophy can be detected by evaluating the discriminant formula constructed using the found diagnostic hippocampal atrophy marker set and the diagnostic hippocampal atrophy marker set in an independent sample group. Hippocampal atrophy marker sets and methods for discriminating hippocampal atrophy can be found.
- At least one of at least one hippocampal atrophy-related factor other than the above-mentioned hippocampal atrophy marker for example, age, gender, and the number of ApoE4 alleles (eg, age; age and gender; Age and number of ApoE4 alleles; or age, gender, and number of ApoE4 alleles) may be included.
- the hippocampus is known to decrease and atrophy by about 1% every other year with age. There is also a report that the size of each part of the brain differs between men and women, and the hippocampus is larger in women.
- the Split-sample method for evaluating the discrimination performance (generalization) of the discriminant. That is, the data set is divided into a learning sample group and a verification sample group, and the learning sample group selects the hippocampal atrophy marker by statistical test and creates a discriminant formula, and the result and verification of the discriminant sample group discriminated by the discriminant formula. The accuracy, sensitivity, and specificity are calculated using the true group to which the sample group belongs, and the discrimination performance is evaluated.
- the hippocampal atrophy markers are selected and the discriminant is created by statistical test using all the samples, and the newly prepared sample is discriminated by the discriminant, and the accuracy, sensitivity, and accuracy, and It is also possible to calculate the specificity and evaluate the discriminant performance.
- the present invention is also for the detection of hippocampal atrophy of a polynucleotide that is at least one miRNA selected from Group A above and, optionally, a polynucleotide that is at least one miRNA selected from Group B above. Also provided for use as a Kaiba atrophy marker.
- Example 1 Serum was collected from a total of 1,126 human subjects (Table 2) who gave informed consent at the National Center for Geriatrics and Gerontology using BD Vacutainer blood collection tube 219 AFBZX00109000 (BD Co., Ltd. (Japan)).
- RNA extraction reagent in 3D-Gene (registered trademark) RNA extraction reagent from liquid sample kit (Toray Industries, Inc. (Japan) was used. Total RNA was obtained according to the protocol set by the company.
- the oligo DNA chip was scanned using a 3D-Gene (registered trademark) scanner (Toray Industries, Inc.), images were acquired, and the fluorescence intensity was quantified by 3D-Gene (registered trademark) Extension (Toray Industries, Inc.). The quantified fluorescence intensity was converted to a logarithmic value having a base of 2 to obtain the expression level, the blank value was subtracted, and the missing value was replaced with a signal value of 0.1. As a result, a comprehensive miRNA expression level was obtained for the sera of the above 1,126 persons.
- each sample group was divided into a learning sample group, a cross-validation sample group, and an independent verification sample group as shown in each of the following examples.
- Calculations and statistical analysis using the quantified miRNA expression level are performed in R language 3.5.2 (R Core Team (2016). R: A language and environmental for static commuting. R Foundation for Austria. URL https: // www. R-project. Org /.) And MASS Package 7.3.45 (Venables, W.N. & Ripley, BD (2002) Modern Applied Statistics. It was carried out using Springer, New York. ISBN 0-387-95457-0).
- DNA was extracted from peripheral blood cells of the above 264 subjects, and total sequence analysis was performed using a next-generation sequencer. Of these, the gene polymorphism of the APOE gene was analyzed, and in particular, the number of ApoE4 alleles was measured.
- Example 2 ⁇ Comparison between hippocampal atrophy group and non-atrophy group with a single miRNA>
- two groups with different hippocampal atrophy degrees using the hippocampal volume ratio to the whole brain as an index
- a single miRNA showing a significant difference in the expression level was selected as a hippocampal atrophy marker.
- the hippocampal atrophy degree thresholds are divided into four trials, and a hippocampal atrophy group and a hippocampal non-atrophy group are set at each threshold value to select a marker. was done.
- each marker the expression level of each miRNA obtained in Example 1 above was corrected using an endogenous control (miR-149-3p, miR-2861, miR-4463). Furthermore, in order to obtain more reliable diagnostic markers, 210 miRNAs having an expression level of 2 to the 4th power or more in 90% or more of the samples were selected in either the hippocampal atrophy group or the non-hippocampal atrophy group. Was analyzed.
- hippocampal atrophy group 1 with a hippocampal volume ratio of 0.60% or less to the whole brain and hippocampal non-atrophic group 1 with a hippocampal volume ratio of 0.63% or more to the whole brain
- the expression level between the hippocampal atrophy group 1 (132 samples) with a value of 0.60% or less and the hippocampal non-atrophy group 1 (132 samples) with a hippocampal volume ratio of 0.63% or more to the whole brain is statistically significant.
- a two-sided t-test assuming equal dispersion was performed to calculate the P value.
- the P value is less than 0.1, or the absolute value of the log-converted gene expression level difference (Fold Change) between the hippocampal atrophy group and the hippocampal non-atrophy group is estimated as a measurement error.
- MiRNAs / genes greater than 2 were selected and the results are shown in Table 3.
- the miRNA newly found as a marker for examining the presence or absence of hippocampal atrophy is a polynucleotide consisting of the nucleotide sequences represented by SEQ ID NOs: 1 to 64.
- Example 2- (1) Comparison between hippocampal atrophy group 2 in which the hippocampal volume ratio to the whole brain is 0.59% or less and hippocampal non-atrophic group 2 in which the hippocampal volume ratio to the whole brain is 0.64% or more.
- An analysis was performed in which Example 2- (1) was deepened by comparing two extremely different groups. Hippocampal atrophy group 2 (105 samples) with a lower threshold and a hippocampal volume ratio of 0.59% or less to the whole brain, and a hippocampal volume ratio to the whole brain with a higher threshold of 0.64.
- a comparison was made between two groups of hippocampal non-atrophic group 2 (105 samples), which was% or more.
- the miRNA newly found as a marker for examining the presence or absence of hippocampal atrophy is a polynucleotide consisting of the nucleotide sequences represented by SEQ ID NOs: 65 to 69 and 71.
- Example 2- (2) Comparison between the hippocampal atrophy group 3 in which the hippocampal volume ratio to the whole brain is 0.57% or less and the hippocampal non-atrophic group 3 in which the hippocampal volume ratio to the whole brain is 0.68% or more.
- An analysis was performed in which Example 2- (2) was deepened by comparing two extremely different groups. Hippocampal atrophy group 3 (88 samples) with a lower threshold and a hippocampal volume ratio of 0.57% or less to the whole brain, and a hippocampal volume ratio to the whole brain with a higher threshold of 0.68.
- a comparison was made between two groups of hippocampal non-atrophic group 3 (88 samples), which was% or more.
- miRNA / which has a P value of less than 0.1 or an absolute value of Fold Change of 0.2 or more in comparison between the two groups. Genes were selected and the results are shown in Table 5.
- the miRNA newly found as a marker for examining the presence or absence of hippocampal atrophy is a polynucleotide consisting of the nucleotide sequences represented by SEQ ID NOs: 70 and 72 to 107.
- Example 2- (3) was deepened by comparing two extremely different groups. Hippocampal atrophy group 4 (5 samples) with an extremely low threshold and a hippocampal volume ratio of 0.38% or less to the whole brain, and a hippocampal volume ratio of 0 to the whole brain with an extremely high threshold. A comparison was made between two groups of hippocampal non-atrophic group 4 (5 samples) of .86% or more.
- hsa-miR-3621 and hsa-miR-4763-3p were newly selected.
- the miRNAs corresponding to these are polynucleotides consisting of the nucleotide sequences of SEQ ID NOs: 108 and 109, respectively, and have been newly found as markers for examining the presence or absence of hippocampal atrophy (Kaiba atrophy markers).
- All the polynucleotides consisting of the nucleotide sequences represented by SEQ ID NOs: 1 to 200 above are markers that can independently distinguish between a subject with hippocampal atrophy and a subject with non-hippocampal atrophy.
- Example 3 ⁇ Discrimination of hippocampal atrophy patients by combination of a small number (2 to 5 types) of miRNA markers (discrimination by logistic regression analysis)>
- the discriminant performance is evaluated in the independent verification sample group (Table 7), and the miRNA / gene used in the discriminant with high performance.
- Table 7 independent verification sample group
- the hippocampal atrophy group 3 (88 samples in which the volume ratio of the hippocampus to the whole brain is 0.57% or less) and the hippocampus non-atrophy group 3 (relative to the whole brain) used in Example 2- (3).
- a miRNA marker capable of discriminating the degree of hippocampal atrophy was searched for using 88 specimens having a hippocampal volume ratio of 0.68% or more).
- 73 samples were assigned to the learning / cross-validation sample group in each of the hippocampal atrophy group and the non-hippocampal non-atrophy group, and the remaining 15 samples were assigned to the independent validation sample group (Table 7).
- the sample was further divided into three groups, both the hippocampal atrophy group and the non-validated hippocampal group, two-thirds of which were divided into the learning sample group (learning group), and one-third of which was cross-validated.
- the sample group (cross-validation group) was used.
- the cross-validation sample group was replaced three times, and each time the cross-validation sample group was replaced, a discriminant formula was constructed using the learning sample group. Furthermore, the division method was changed 10 times.
- both groups (kaiba atrophy group and non-kaiba atrophy group) set in the learning / cross-validation sample group are discriminated with respect to the expression level of miRNA consisting of the nucleotide sequences represented by SEQ ID NOs: 110 and 114.
- the hit rate of hippocampal atrophy was calculated using the threshold (0.5), AUC 0.862, sensitivity 80.0%, and specificity 86.7% were obtained as the discrimination performance in the independent verification sample group.
- the discrimination performance in the independent verification sample group was AUC 0.898, sensitivity 73.3%, and so on. A specificity of 93.3% was obtained.
- the discrimination performance in the independent verification sample group is AUC 0.92 and sensitivity 80.0. %, Specificity 86.7% was obtained.
- the discrimination performance in the independent verification sample group is AUC 0.902, sensitivity 86. 2.7% and 73.3% specificity were obtained.
- Example 4 ⁇ Discrimination of hippocampal atrophy patients by a combination of a small number (5 or 6 types) of miRNA markers (discriminant analysis by Fisher)>
- a discriminant formula was created for a combination of 2 to 5 types of miRNA markers by the Lasso method, but in order to further improve the discriminant performance of the two groups, discriminant analysis was performed using Fisher's discriminant analysis in this example. rice field.
- Example 4- (1) the hippocampal atrophy group 1 used in Example 2- (1) (132 samples in which the volume ratio of the hippocampus to the whole brain is 0.60% or less). And the non-hippocampal atrophy group (132 samples in which the volume ratio of the hippocampus to the whole brain was 0.64% or more) was searched for a miRNA marker capable of discriminating the degree of hippocampal atrophy.
- 110 samples which is five-sixths of the total in each of the hippocampal atrophy group and the hippocampal non-atrophy group, are distributed to the learning / cross-validation sample group, and the remaining 22 samples, which is one-sixth, are distributed to the independent validation sample group. (Table 9).
- the discriminant formula constructed using them is The threshold of the discrimination score for discriminating between the hippocampal atrophy group and the hippocampal non-atrophy group is set to 0 (zero), and when the hit rate of hippocampal atrophy is calculated using this formula, the discrimination performance in the independent verification sample group is AUC 0.855, sensitivity 80.0%, specificity 80.0% were obtained.
- the combination of miRNA markers represented by the SEQ ID NO: included in the discriminant having an AUC of 0.8 or more in the cross-validation sample group and the independent validation sample group, and the calculated AUC and sensitivity in each group.
- Table 10 Specificity are shown in Table 10.
- 6 types of markers for example, regarding the expression level of miRNA consisting of the nucleotide sequences represented by SEQ ID NOs: 26, 66, 110, 112, 114 and 165 in the learning / cross-validation sample group, a discrimination formula constructed using them.
- Example 5 ⁇ Discrimination of hippocampal atrophy patients using a combination of 5 or more miRNA markers and hippocampal atrophy-related factors (discrimination by logistic regression analysis)>
- a logistic regression analysis using the Lasso method was performed for the purpose of examining variations when the number of markers used for discrimination was not limited.
- improvement of discrimination performance was examined by adding age, gender, and the number of ApoE4 alleles as explanatory variables to the discriminant as hippocampal atrophy-related factors.
- Example 4 The same sample as in Example 4 was divided into the same sample group and number of samples as in Table 9 and used. However, the division and distribution into the learning / cross-validation sample group and the independent verification sample group were performed in 30 ways by randomly exchanging the samples. Logistic regression analysis using the Lasso method was performed using the measured values of the expression levels of 210 miRNAs analyzed in Example 2 and the age, sex, and number of ApoE4 alleles of the subjects as explanatory variables, and whether or not the hippocampus was atrophic. We constructed a discrimination formula to determine whether or not. In the constructed discriminant, the accuracy, sensitivity, and specificity were calculated using the learning / cross-validation sample group to evaluate the discriminant performance, and the discriminant performance was further verified using the independent validation sample group.
- the combination of miRNA markers represented by the SEQ ID NO: included in the discriminant in which the AUC in the cross-validation sample group and the independent validation sample group is 0.9 or more, and the calculated AUC and sensitivity in each group.
- the specificity is shown in Table 11.
- the score threshold was set to 0 (zero), and the accuracy rate of hippocampal atrophy was calculated using this formula.
- the discrimination performance in the independent verification sample group was AUC 0.986, sensitivity 86.4%, and specificity. Degree 95.5% was obtained.
- miRNA markers used in the discrimination formula having an AUC of 0.8 or more SEQ ID NOs: 1-28, 30-64, 67-73, 75- A polynucleotide consisting of the nucleotide sequences represented by 122, 124, 125, 127 to 156, 158 to 163, and 167 to 200 is selected, and when these are arbitrarily combined, the hippocampal atrophy group and the hippocampal non-atrophy group can be distinguished. It has been shown.
- Example 6 ⁇ Discrimination of two groups with more extremely different hippocampal volumes (logistic regression discrimination) using a combination of multiple miRNA markers and hippocampal atrophy-related factors>
- Objective 1 To perform a deepening analysis of Example 5 by comparing two groups whose hippocampal atrophy degree is extremely different from that of Example 5.
- Objective 2 To examine whether a marker set with an AUC exceeding 0.9 can be obtained even when the number of samples used for learning and verification in the construction of the discriminant is reduced.
- Example 3 The same specimens as in Example 3, that is, 88 specimens in which the hippocampal volume ratio to the whole brain was 0.57% or less in the hippocampal atrophy group, and 0.68% or more in the hippocampal volume ratio to the whole brain in the hippocampal non-atrophic group.
- a certain 88 samples were divided into a learning / cross-validation sample group and an independent validation sample group in the same manner as in Table 7. This learning / cross-validation sample group and the independent validation sample group were sorted 30 times by randomly exchanging the samples. Further, as in Example 3, the learning / cross-validation sample group was divided into a learning sample group (learning group) and a cross-validation sample group.
- Logistic regression analysis using the Lasso method was performed using the measured values of the expression levels of 210 miRNAs analyzed in Example 2 and the age, sex, and number of ApoE4 alleles of the subjects as explanatory variables, and whether the hippocampal atrophy was performed.
- a discrimination formula was constructed to determine whether or not it was present. In the constructed discriminant, the accuracy, sensitivity, and specificity were calculated using the learning / cross-validation sample group to evaluate the discrimination performance, and the discrimination performance was further verified using the independent sample verification group.
- the combination of miRNA markers represented by the SEQ ID NO: included in the discriminant in which the AUC in the cross-validation sample group and the independent validation sample group is 0.9 or more, and the calculated AUC and sensitivity in each group.
- the specificity is shown in Table 12.
- the discrimination formula constructed using them sets the threshold of the discrimination score for discriminating between the hippocampal atrophy group and the hippocampal non-atrophy group as 0.5.
- AUC 1.0, sensitivity 100.0%, and specificity 100.0% were obtained as the discrimination performance in the independent verification sample group.
- the discriminant score charts (A, B) and the ROC (Receiver Operating Characteristic) curve (C, D) as the analysis results are shown in FIG.
- the discrimination score (score; score) takes a numerical value of 0 to 1, and the dotted line (discrimination score 0.5) in the figure indicates the discrimination boundary for discriminating whether or not the hippocampus is atrophied.
- Discriminant score The discriminant score shown in the figure is a discriminant score prepared using the above 13 types of miRNA (SEQ ID NO: 155, 137, 188, 127, 196, 154, 110, 128, 100, 152, 133, 159 and 114). It was calculated by substituting the measured value of the miRNA expression level of each sample with the age, sex, and number of ApoE4 alleles into the formula.
- FIGS. 2A and 2B show the obtained discrimination score (score) on the vertical axis and the sample group on the horizontal axis.
- the ROC curve is represented by the sensitivity calculated using the miRNA marker on the vertical axis and the specificity on the horizontal axis (FIGS. 2C and D).
- markers used in the discriminant formula having an AUC of 0.8 or higher with respect to the discrimination performance in the cross-validation sample group and the independent validation sample group SEQ ID NOs: 1 to 5, 9 to 13, 16 to 18, 20, 21, 23 to 26, 29-32, 37-39, 41, 42, 44, 45, 47-63, 67-75, 77-88, 90-94, 96-102, 104-111, 113, 114, 117, 118, From the base sequences represented by 120, 121, 123-125, 127-129, 133-138, 141-143, 145-150, 152-159, 161-163, 167-169, 171-183, 185-200. Polynucleotides were selected and showed that when they were arbitrarily combined, patients with hippocampal atrophy and those without hippocampal atrophy could be distinguished.
- the discriminant was constructed using two groups of samples with extremely different hippocampal atrophy degrees, but the discriminant score obtained from the constructed and verified discriminant was actually obtained from the brain MRI image.
- the ratio of hippocampal volume to the whole brain was highly correlated.
- the expression level and age of miRNA consisting of the nucleotide sequences represented by SEQ ID NOs: 155, 137, 188, 127, 196, 154, 110, 128, 100, 152, 133, 159 and 114 in the learning / cross-validation sample group.
- the discriminant score obtained from the discriminant constructed by combining the gender and the number of ApoE4 alleles was correlated with the degree of hippocampal atrophy.
- the volume ratio of the hippocampus to the whole brain is 0.69% or more, and when the discrimination score is -2 or more and less than 0, all.
- the volume ratio of the hippocampus to the brain is 0.61% or more and less than 0.69%, and the volume ratio of the hippocampus to the whole brain is 0.53% or more and less than 0.61% when the discrimination score is 0 or more and less than 2.
- the volume ratio of the hippocampus to the whole brain when the discrimination score is 2 or more and less than 4 is 0.45% or more and less than 0.53, and the volume ratio of the hippocampus to the whole brain when the discrimination score is 4 or more is 0.45.
- Multiple types of polynucleotides consisting of the base sequences represented by ⁇ 143, 145 to 150, 152 to 159, 161 to 163, 167 to 169, 171 to 180, 182, 183, 185 to 194, and 196 to 200 are used. When combined, it was shown that subjects with hippocampal atrophy and subjects without hippocampal atrophy can be accurately discriminated (AUC 0.9 or higher).
- Example 7 ⁇ Determination of hippocampal atrophy and prediction of hippocampal volume by regression equation construction>
- Example 13 Of the total of 264 samples selected in Example 1 and used for the discriminant analysis of hippocampal atrophy, 132 samples, which is half of the total, were included in the learning / cross-validation sample group, and 132 samples, which was the remaining half, were independent. It was assigned to the verification sample group (Table 13).
- the miRNA expression level was corrected in the same manner as in Example 2.
- Lasso regression analysis was performed on the expression level, age, sex, and number of ApoE4 alleles of 210 types of miRNA analyzed in Example 2, a regression equation that can predict the volume of the hippocampus was constructed, and an independent verification sample was obtained. We verified the discrimination performance of the regression equation constructed using the group.
- regression equation was constructed using the expression level of miRNA comprising the nucleotide sequence represented by 133,2,20,51 and 114, R 2 as regression performance in learning and cross validation sample group is 0.621, the RMSE 1.187, MAE is 0.940, regression performance in independent validation sample group R 2 is 0.488, RMSE is 1.393, MAE was 1.117.
- the discrimination score obtained from this regression equation was highly correlated with the quantitative value of the hippocampal volume actually obtained from the brain MRI image.
- the discriminant score obtained from the formula was correlated with the quantitative value of hippocampal volume and the degree of hippocampal atrophy.
- the volume ratio of the hippocampus to the whole brain when the discrimination score obtained from the discrimination formula is less than 6 is less than 0.44%, and the volume ratio of the hippocampus to the whole brain when the discrimination score is 6 or more and less than 8.
- the hippocampal volume ratio to the whole brain is 0.59% or more and less than 0.75%, and the discrimination score is 10 or more.
- the hippocampal volume ratio to the whole brain in the case can be predicted to be 0.75% or more.
- the miRNA expression level was corrected in the same manner as in Example 2.
- FIG. 3 shows a regression line based on the hippocampal volume (quantitative value) and explanatory variables obtained in Example 7- (2).
- the discriminant score obtained from the discriminant constructed by combining the expression level and age of miRNA consisting of the base sequence, sex, and the number of ApoE4 alleles was correlated with the quantitative value of hippocampal volume and the degree of hippocampal atrophy.
- the volume ratio of the hippocampus to the whole brain is less than 0.48%, and when the discrimination score is 6 or more and less than 8, the volume ratio of the hippocampus is less than 0.48%.
- the hippocampal volume ratio is 0.48% or more and less than 0.60%, and the hippocampal volume ratio to the whole brain is 0.60% or more and less than 0.71% when the discrimination score is 8 or more and less than 10. It can be predicted that the hippocampal volume ratio to the whole brain is 0.71% or more when the score is 10 or more.
- hippocampal atrophy degree of the subject can be predicted based on miRNA measurement, and the hippocampal atrophy reserve army with a relatively small hippocampal atrophy degree can be detected.
- hippocampal atrophy can be easily and objectively and quantitatively detected without any difference between facilities of doctors and medical institutions.
- the degree of hippocampal atrophy of a subject can be easily determined by using a measured value of the expression level of one or more to several miRNAs in the blood, serum, and / or plasma of a subject that can be collected in a minimally invasive manner as an index. ..
- hippocampal atrophy can be detected at an early stage, leading to early diagnosis of dementia and the like.
- the diagnostic marker of the present invention it is possible to correctly determine whether or not the hippocampus is atrophied using a blood sample, and thus it can be expected that the progression of hippocampal atrophy is suppressed by medical intervention.
- the present invention is 1. It is an objective method in which there is no difference in test results between medical facilities, equipment, and doctors. It can be applied to patients who cannot undergo MRI. The degree of atrophy can be determined gradually, and 4. It is possible to provide an advantageous inspection method with low invasiveness.
Landscapes
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Health & Medical Sciences (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Genetics & Genomics (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Analytical Chemistry (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Molecular Biology (AREA)
- Biochemistry (AREA)
- Biophysics (AREA)
- Physics & Mathematics (AREA)
- Microbiology (AREA)
- General Health & Medical Sciences (AREA)
- Immunology (AREA)
- Biomedical Technology (AREA)
- Pathology (AREA)
- Plant Pathology (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
Description
(1)海馬萎縮マーカーである、miR-3131、miR-6757-5p、miR-4706、miR-5001-5p、miR-3180-3p、miR-642b-3p、miR-4655-5p、miR-6819-5p、miR-937-5p、miR-4688、miR-6741-5p、miR-7107-5p、miR-4271、miR-1229-5p、miR-4707-5p、miR-6808-5p、miR-4656、miR-6076、miR-6762-5p、miR-7109-5p、miR-6732-5p、miR-3195、miR-7150、miR-642a-3p、miR-1249-5p、miR-3185、miR-4689、miR-3141、miR-6840-3p、miR-3135b、miR-1914-3p、miR-4446-3p、miR-4433b-3p、miR-6877-5p、miR-6848-5p、miR-3620-5p、miR-6825-5p、miR-5739、miR-3663-3p、miR-4695-5p、miR-3162-5p、miR-3679-5p、miR-8059、miR-7110-5p、miR-1275、miR-6779-5p、miR-197-5p、miR-6845-5p、miR-4327、miR-4723-5p、miR-4530、miR-6771-5p、miR-614、miR-92a-2-5p、miR-6891-5p、miR-6124、miR-4687-3p、miR-4442、miR-7977、miR-6785-5p、miR-4497、miR-8071、miR-663b、miR-3180、miR-4251、miR-1285-3p、miR-6870-5p、miR-4484、miR-4476、miR-6749-5p、miR-4454、miR-6893-5p、miR-6085、miR-4787-5p、miR-149-3p、miR-7704、miR-6125、miR-6090、miR-3197、miR-6850-5p、miR-4467、miR-6885-5p、miR-6803-5p、miR-6798-5p、miR-6780b-5p、miR-6768-5p、miR-5100、miR-6724-5p、miR-6879-5p、miR-7108-5p、miR-4649-5p、miR-4739、miR-6089、miR-1908-5p、miR-4516、miR-2861、miR-4492、miR-4294、miR-6791-5p、miR-1469、miR-6752-5p、miR-4730、miR-6126、miR-6869-5p、miR-1268a、miR-6799-5p、miR-8069、miR-3621及びmiR-4763-3pからなる群から選択される少なくとも1つのポリヌクレオチド、又は該ポリヌクレオチドの相補鎖と、特異的に結合可能な核酸を含む、海馬萎縮の検出用キット。
(a)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~109のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(c)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(1)に記載のキット。
(f)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号110~200のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(h)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(3)に記載のキット。
(a)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~109のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(c)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(5)に記載のデバイス。
(f)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号110~200のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(h)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(7)に記載のデバイス。
(a)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~109のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(c)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(12)又は(13)に記載の方法。
(f)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号110~200のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(h)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、(15)に記載の方法。
本明細書中で使用する用語は、以下の定義を有する。
1.海馬萎縮の標的核酸
本発明の上記定義の海馬萎縮検出用の核酸、例えば核酸プローブ又はプライマーを使用して、海馬萎縮、又は海馬の萎縮若しくは非萎縮を検出するための、海馬萎縮マーカーとしての標的核酸は、miR-3131、miR-6757-5p、miR-4706、miR-5001-5p、miR-3180-3p、miR-642b-3p、miR-4655-5p、miR-6819-5p、miR-937-5p、miR-4688、miR-6741-5p、miR-7107-5p、miR-4271、miR-1229-5p、miR-4707-5p、miR-6808-5p、miR-4656、miR-6076、miR-6762-5p、miR-7109-5p、miR-6732-5p、miR-3195、miR-7150、miR-642a-3p、miR-1249-5p、miR-3185、miR-4689、miR-3141、miR-6840-3p、miR-3135b、miR-1914-3p、miR-4446-3p、miR-4433b-3p、miR-6877-5p、miR-6848-5p、miR-3620-5p、miR-6825-5p、miR-5739、miR-3663-3p、miR-4695-5p、miR-3162-5p、miR-3679-5p、miR-8059、miR-7110-5p、miR-1275、miR-6779-5p、miR-197-5p、miR-6845-5p、miR-4327、miR-4723-5p、miR-4530、miR-6771-5p、miR-614、miR-92a-2-5p、miR-6891-5p、miR-6124、miR-4687-3p、miR-4442、miR-7977、miR-6785-5p、miR-4497、miR-8071、miR-663b、miR-3180、miR-4251、miR-1285-3p、miR-6870-5p、miR-4484、miR-4476、miR-6749-5p、miR-4454、miR-6893-5p、miR-6085、miR-4787-5p、miR-149-3p、miR-7704、miR-6125、miR-6090、miR-3197、miR-6850-5p、miR-4467、miR-6885-5p、miR-6803-5p、miR-6798-5p、miR-6780b-5p、miR-6768-5p、miR-5100、miR-6724-5p、miR-6879-5p、miR-7108-5p、miR-4649-5p、miR-4739、miR-6089、miR-1908-5p、miR-4516、miR-2861、miR-4492、miR-4294、miR-6791-5p、miR-1469、miR-6752-5p、miR-4730、miR-6126、miR-6869-5p、miR-1268a、miR-6799-5p、miR-8069、miR-3621及びmiR-4763-3pからなる群から選択される少なくとも1つのmiRNAであるポリヌクレオチドであり得る。
本発明において、海馬萎縮の検出用の核酸、例えば海馬萎縮を診断するために使用可能な核酸、例えば核酸プローブ又はプライマーは、海馬萎縮の標的核酸(海馬萎縮マーカー)としての、miR-3131、miR-6757-5p、miR-4706、miR-5001-5p、miR-3180-3p、miR-642b-3p、miR-4655-5p、miR-6819-5p、miR-937-5p、miR-4688、miR-6741-5p、miR-7107-5p、miR-4271、miR-1229-5p、miR-4707-5p、miR-6808-5p、miR-4656、miR-6076、miR-6762-5p、miR-7109-5p、miR-6732-5p、miR-3195、miR-7150、miR-642a-3p、miR-1249-5p、miR-3185、miR-4689、miR-3141、miR-6840-3p、miR-3135b、miR-1914-3p、miR-4446-3p、miR-4433b-3p、miR-6877-5p、miR-6848-5p、miR-3620-5p、miR-6825-5p、miR-5739、miR-3663-3p、miR-4695-5p、miR-3162-5p、miR-3679-5p、miR-8059、miR-7110-5p、miR-1275、miR-6779-5p、miR-197-5p、miR-6845-5p、miR-4327、miR-4723-5p、miR-4530、miR-6771-5p、miR-614、miR-92a-2-5p、miR-6891-5p、miR-6124、miR-4687-3p、miR-4442、miR-7977、miR-6785-5p、miR-4497、miR-8071、miR-663b、miR-3180、miR-4251、miR-1285-3p、miR-6870-5p、miR-4484、miR-4476、miR-6749-5p、miR-4454、miR-6893-5p、miR-6085、miR-4787-5p、miR-149-3p、miR-7704、miR-6125、miR-6090、miR-3197、miR-6850-5p、miR-4467、miR-6885-5p、miR-6803-5p、miR-6798-5p、miR-6780b-5p、miR-6768-5p、miR-5100、miR-6724-5p、miR-6879-5p、miR-7108-5p、miR-4649-5p、miR-4739、miR-6089、miR-1908-5p、miR-4516、miR-2861、miR-4492、miR-4294、miR-6791-5p、miR-1469、miR-6752-5p、miR-4730、miR-6126、miR-6869-5p、miR-1268a、miR-6799-5p、miR-8069、miR-3621及びmiR-4763-3pからなる群から選択される少なくとも1つのmiRNAであるポリヌクレオチド又は該ポリヌクレオチドの相補鎖と、特異的に結合する核酸であり得る。
(a)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、は15以上の連続した塩基を含むその断片、
(b)配列番号1~109のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(c)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列、に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、並びに、
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド。
(f)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号110~200のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(h)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、並びに、
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド。
本発明は、海馬萎縮マーカーである標的核酸を測定するための、本発明における海馬萎縮検出用核酸、例えば核酸プローブ又はプライマーとして使用可能なポリヌクレオチドの1つ又は複数を含む海馬萎縮検出用キット又はデバイスを提供する。
群A:
miR-3131、miR-6757-5p、miR-4706、miR-5001-5p、miR-3180-3p、miR-642b-3p、miR-4655-5p、miR-6819-5p、miR-937-5p、miR-4688、miR-6741-5p、miR-7107-5p、miR-4271、miR-1229-5p、miR-4707-5p、miR-6808-5p、miR-4656、miR-6076、miR-6762-5p、miR-7109-5p、miR-6732-5p、miR-3195、miR-7150、miR-642a-3p、miR-1249-5p、miR-3185、miR-4689、miR-3141、miR-6840-3p、miR-3135b、miR-1914-3p、miR-4446-3p、miR-4433b-3p、miR-6877-5p、miR-6848-5p、miR-3620-5p、miR-6825-5p、miR-5739、miR-3663-3p、miR-4695-5p、miR-3162-5p、miR-3679-5p、miR-8059、miR-7110-5p、miR-1275、miR-6779-5p、miR-197-5p、miR-6845-5p、miR-4327、miR-4723-5p、miR-4530、miR-6771-5p、miR-614、miR-92a-2-5p、miR-6891-5p、miR-6124、miR-4687-3p、miR-4442、miR-7977、miR-6785-5p、miR-4497、miR-8071、miR-663b、miR-3180、miR-4251、miR-1285-3p、miR-6870-5p、miR-4484、miR-4476、miR-6749-5p、miR-4454、miR-6893-5p、miR-6085、miR-4787-5p、miR-149-3p、miR-7704、miR-6125、miR-6090、miR-3197、miR-6850-5p、miR-4467、miR-6885-5p、miR-6803-5p、miR-6798-5p、miR-6780b-5p、miR-6768-5p、miR-5100、miR-6724-5p、miR-6879-5p、miR-7108-5p、miR-4649-5p、miR-4739、miR-6089、miR-1908-5p、miR-4516、miR-2861、miR-4492、miR-4294、miR-6791-5p、miR-1469、miR-6752-5p、miR-4730、miR-6126、miR-6869-5p、miR-1268a、miR-6799-5p、miR-8069、miR-3621及びmiR-4763-3p
群B:
miR-1228-5p、miR-760、miR-187-5p、miR-7111-5p、miR-6088、miR-6805-3p、miR-4640-5p、miR-6721-5p、miR-6880-5p、miR-711、miR-128-1-5p、miR-4525、miR-486-3p、miR-6756-5p、miR-1260b、miR-3184-5p、miR-6075、miR-204-3p、miR-4728-5p、miR-4534、miR-4758-5p、miR-8063、miR-6836-3p、miR-6789-5p、miR-744-5p、miR-1909-3p、miR-887-3p、miR-4745-5p、miR-4433a-3p、miR-5090、miR-296-5p、miR-939-5p、miR-3648、miR-3196、miR-6722-3p、miR-6805-5p、miR-1202、miR-6775-5p、miR-6087、miR-6765-5p、miR-6875-5p、miR-4674、miR-1233-5p、miR-7114-5p、miR-5787、miR-8072、miR-3619-3p、miR-4632-5p、miR-6800-5p、miR-4634、miR-4486、miR-6727-5p、miR-4505、miR-4725-3p、miR-1538、miR-320b、miR-1915-5p、miR-328-5p、miR-6820-5p、miR-6726-5p、miR-3665、miR-638、miR-762、miR-4466、miR-3940-5p、miR-1237-5p、miR-575、miR-3656、miR-4488、miR-4281、miR-6781-5p、miR-4532、miR-4665-5p、miR-6816-5p、miR-4508、miR-6784-5p、miR-6786-5p、miR-4741、miR-1343-5p、miR-1227-5p、miR-4734、miR-3960、miR-128-2-5p、miR-6743-5p、miR-663a、miR-6729-5p、miR-1915-3p、miR-1268b、miR-4651、miR-3178及びmiR-4463
(1)配列番号1~109のいずれかで表される塩基配列においてuがtである塩基配列又はその相補的配列において、15以上の連続した塩基配列を含むポリヌクレオチド。
(2)配列番号110~200のいずれかで表される塩基配列においてuがtである塩基配列又はその相補的配列において、15以上の連続した塩基配列を含むポリヌクレオチド。
(1)配列番号110、114の組み合わせ
(2)配列番号110、114、1の組み合わせ
(3)配列番号110、114、135の組み合わせ
(4)配列番号110、114、155の組み合わせ
(5)配列番号110、114、137の組み合わせ
(6)配列番号110、114、155、135の組み合わせ
(7)配列番号110、114、137、111の組み合わせ
(8)配列番号110、114、155、2の組み合わせ
(9)配列番号110、114、142、2の組み合わせ
(10)配列番号110、114、1、149の組み合わせ
(11)配列番号110、114、1、135の組み合わせ
(12)配列番号110、114、155、68の組み合わせ
(13)配列番号110、114、2、135の組み合わせ
(14)配列番号110、114、155、135、2の組み合わせ
(15)配列番号110、114、155、137、2の組み合わせ
(16)配列番号110、114、3、2、142の組み合わせ
(17)配列番号110、114、142、135、3の組み合わせ
(18)配列番号110、114、127、1、149の組み合わせ
(19)配列番号110、114、127、1、135の組み合わせ
(20)配列番号110、114、155、68、152の組み合わせ
(21)配列番号110、114、68、135、2の組み合わせ
(22)配列番号110、114、142、1、2の組み合わせ
(23)配列番号127、1、162、114、110の組み合わせ
(24)配列番号1、126、125、114、110の組み合わせ
(25)配列番号66、110、165、65、166、164の組み合わせ
(26)配列番号66、110、165、26、112、114の組み合わせ
(27)配列番号1、110、165、65、164、71の組み合わせ
(1)配列番号155、87、188、127、85、5、1、110、193、175、118、152、133、39、159、149、150、114の組み合わせ
(2)配列番号115、188、127、85、1、16、110、41、24、200、26、193、175、108、152、113、39、159、114の組み合わせ
(3)配列番号155、188、127、146、5、1、16、110、200、101、193、175、145、125、152、111、39、114の組み合わせ
(4)配列番号155、87、188、127、85、1、183、196、14、189、110、24、101、193、175、118、178、152、39、150、114の組み合わせ
(5)配列番号179、155、142、188、127、146、1、189、110、4、200、193、175、118、63、152、111、180、39、114の組み合わせ
(6)配列番号87、188、127、85、5、1、16、196、14、38、110、4、200、193、175、109、145、152、39、159、149、114の組み合わせ
(7)配列番号155、87、142、188、127、85、5、1、189、154、110、41、193、175、167、10、152、111、133、39、171、150、114の組み合わせ
(8)配列番号179、188、127、5、16、154、110、193、175、118、100、109、10、88、152、39、91、159、149、82、104、182、114の組み合わせ
(9)配列番号155、188、127、1、183、187、110、185、86、193、175、109、145、88、3、120、152、133、39、149、182、114の組み合わせ
(10)配列番号155、87、142、188、127、85、1、16、110、26、193、175、118、97、174、152、111、133、113、39、159、114の組み合わせ
(11)配列番号155、87、188、127、1、110、9、24、200、193、43、109、145、125、178、135、174、152、133、39、91、149、182、114の組み合わせ
(12)配列番号155、87、127、5、1、198、189、110、41、200、193、175、75、31、109、59、174、152、111、133、113、39、91、114の組み合わせ
(13)配列番号155、87、188、127、5、1、16、187、110、176、193、175、145、125、88、152、89、141、133、39、149、150、114の組み合わせ
(14)配列番号179、95、188、127、85、5、129、16、196、154、110、200、193、175、118、63、174、108、113、39、91、159、34、114の組み合わせ
(15)配列番号155、87、188、127、85、5、1、16、196、198、38、110、24、200、193、88、96、152、111、39、159、149、182、114の組み合わせ
(16)配列番号179、121、155、87、188、127、5、187、14、189、194、110、58、200、193、118、145、125、59、152、111、113、39、159、149、114の組み合わせ
(17)配列番号121、155、87、188、127、1、189、67、110、4、176、193、75、145、125、88、59、108、152、111、39、159、149、182、114、23の組み合わせ
(18)配列番号155、137、87、168、188、196、189、110、41、4、24、58、101、193、175、145、152、111、113、39、91、61、51、55、182、114の組み合わせ
(19)配列番号155、87、188、127、85、1、16、84、187、110、4、26、193、175、118、109、88、108、152、111、39、159、149、182、114の組み合わせ
(20)配列番号179、155、132、168、188、127、5、1、14、189、110、124、4、24、200、193、175、97、145、174、108、152、133、39、114の組み合わせ
(21)配列番号142、188、85、146、1、196、187、189、110、58、101、193、175、25、105、54、120、152、111、113、39、91、159、149、61、182、114の組み合わせ
(22)配列番号121、155、95、188、127、80、198、189、110、200、101、193、175、75、145、88、120、152、133、39、91、149、34、171、140、150、114の組み合わせ
(23)配列番号155、188、127、85、1、16、196、84、53、189、110、4、68、101、193、145、178、54、108、152、111、133、39、159、182、114の組み合わせ
(24)配列番号121、115、155、137、188、127、5、1、196、35、187、110、193、175、118、97、178、48、37、152、133、39、91、149、182、114の組み合わせ
(25)配列番号155、87、188、127、85、1、16、196、187、110、24、200、185、101、175、118、192、145、88、54、108、152、133、39、91、149、182、114の組み合わせ
(26)配列番号115、155、87、188、127、146、5、16、161、53、198、110、143、148、193、175、25、100、109、97、48、108、152、133、39、149、114の組み合わせ
(27)配列番号155、87、142、188、85、1、16、196、53、110、9、41、4、185、193、175、97、135、174、73、152、111、133、113、39、149、55、182、114の組み合わせ
(28)配列番号121、155、188、127、85、1、16、198、189、110、4、86、101、193、75、199、145、108、152、111、113、39、7、91、159、149、182、114の組み合わせ
(29)配列番号155、142、188、127、5、129、16、14、154、110、153、193、175、118、75、109、169、97、21、10、59、152、111、133、39、149、18、114の組み合わせ
(30)配列番号179、155、188、127、5、196、187、14、189、154、38、110、101、175、118、25、100、174、120、108、152、133、39、91、159、149、82、104、182、114の組み合わせ
(31)配列番号155、188、127、79、16、198、110、68、200、102、193、175、43、100、109、167、10、59、191、152、111、133、39、159、149、82、55、182、114の組み合わせ
(32)配列番号155、95、87、188、127、85、1、196、198、187、189、110、200、193、175、192、109、97、145、125、59、152、133、113、39、159、149、150、114の組み合わせ
(33)配列番号121、155、188、127、1、198、187、189、154、110、193、175、118、31、109、145、59、54、174、108、152、133、113、39、91、159、149、2、55、182、114の組み合わせ
(34)配列番号121、155、87、188、127、85、1、79、16、196、187、110、176、24、101、193、175、109、199、145、178、88、152、111、133、39、149、150、182、114、23の組み合わせ
(35)配列番号155、188、127、1、196、189、110、32、41、143、185、193、175、97、145、178、54、174、108、152、111、133、39、91、149、61、18、55、182、114の組み合わせ
(36)配列番号121、155、95、188、127、196、53、187、189、194、110、128、101、193、175、25、109、97、145、174、152、111、113、39、91、159、149、104、182、114の組み合わせ
(37)配列番号155、137、168、188、127、16、196、187、154、110、6、44、9、200、148、101、193、175、109、145、178、48、152、133、113、39、77、55、182、114の組み合わせ
(38)配列番号155、87、188、127、85、1、16、196、84、17、14、110、176、24、200、193、175、75、109、97、145、174、108、152、141、39、159、149、182、114の組み合わせ
(39)配列番号155、87、188、127、85、1、16、196、14、110、176、185、193、175、118、109、199、145、48、108、152、141、133、39、159、149、55、182、114、23の組み合わせ
(40)配列番号179、121、155、95、188、127、85、1、129、196、187、110、176、193、175、31、145、178、48、88、47、54、174、120、108、133、39、91、149、114の組み合わせ
(41)配列番号179、115、155、87、188、127、85、146、5、1、45、84、187、189、110、128、56、200、185、86、193、175、118、145、48、59、152、113、39、159、149、114の組み合わせ
(42)配列番号179、95、142、188、127、5、1、129、16、187、110、193、175、118、100、109、97、125、63、92、152、133、180、39、91、159、149、104、171、150、182、114の組み合わせ
(43)配列番号155、137、142、188、127、85、196、189、110、136、41、143、148、193、175、118、199、97、145、178、88、117、108、133、39、91、159、149、18、182、114の組み合わせ
(44)配列番号13、155、137、87、188、127、85、196、187、110、136、41、24、200、101、193、175、118、97、145、48、152、111、133、113、39、91、149、55、182、114の組み合わせ
(45)配列番号155、188、127、85、5、1、79、84、187、67、110、41、176、175、118、109、97、145、54、108、152、111、133、113、39、91、159、149、171、182、114の組み合わせ
(46)配列番号155、87、188、127、85、146、1、16、196、187、14、110、200、26、101、193、175、97、145、48、120、108、152、141、111、39、159、149、77、182、114の組み合わせ
(47)配列番号155、168、188、127、85、5、1、16、196、84、187、189、110、128、136、176、200、193、175、118、109、145、125、152、89、133、39、149、46、182、114の組み合わせ
(48)配列番号179、121、115、155、188、127、1、16、196、161、84、67、110、124、4、176、24、148、193、175、109、145、178、48、117、152、39、159、149、182、114の組み合わせ
(49)配列番号115、95、87、188、127、85、5、1、16、196、198、187、110、176、101、193、175、118、109、97、125、178、120、108、152、141、39、159、149、182、114の組み合わせ
(50)配列番号179、87、188、127、85、30、1、84、198、187、110、193、175、109、145、125、54、174、120、163、152、133、113、39、91、159、149、82、55、182、114の組み合わせ
(51)配列番号115、155、137、168、188、127、5、1、79、196、187、189、194、110、136、193、175、118、109、199、145、88、152、133、39、91、149、18、46、182、114の組み合わせ
(52)配列番号121、155、188、127、85、1、196、187、110、176、24、200、26、101、193、175、118、25、100、169、145、125、178、54、92、152、111、113、39、91、159、55、114の組み合わせ
(53)配列番号155、95、188、127、85、1、16、196、187、189、110、136、176、69、101、193、175、118、25、75、109、97、167、145、108、152、111、39、159、149、171、182、114の組み合わせ
(54)配列番号121、115、155、168、188、127、85、5、1、84、198、187、14、189、110、193、109、97、145、54、120、152、111、113、39、91、159、149、104、150、182、114、12の組み合わせ
(55)配列番号155、188、127、85、146、5、1、80、183、196、189、110、177、193、175、109、169、97、167、145、59、174、152、111、133、39、91、149、82、103、52、114の組み合わせ
(56)配列番号121、168、188、127、85、16、45、198、187、189、110、4、176、200、148、86、193、175、97、145、125、174、152、113、39、91、159、149、70、82、182、114の組み合わせ
(57)配列番号121、155、188、127、85、5、1、129、187、194、110、176、24、86、26、193、118、100、125、54、174、120、152、133、39、91、149、104、171、150、182、114の組み合わせ
(58)配列番号155、142、188、127、85、1、80、183、196、187、110、124、200、101、193、25、97、145、125、59、54、174、120、108、152、133、39、91、159、82、104、61、150、114の組み合わせ
(59)配列番号121、155、95、87、188、127、85、16、196、187、194、110、136、176、200、185、101、193、175、109、145、59、108、152、111、113、39、91、159、149、46、55、182、114の組み合わせ
(60)配列番号155、87、142、188、127、85、146、5、1、16、45、196、67、110、101、193、175、118、25、109、145、178、63、54、174、108、152、39、91、159、149、55、182、114の組み合わせ
(61)配列番号155、188、127、85、5、1、138、196、84、187、110、24、148、177、101、193、175、100、145、178、88、54、71、152、133、39、91、149、82、61、51、55、182、114の組み合わせ
(62)配列番号115、155、87、168、188、127、85、1、16、196、187、110、124、41、176、24、148、153、193、175、109、145、10、88、59、120、152、133、39、91、149、2、104、182、114の組み合わせ
(63)配列番号87、188、127、1、79、183、196、187、189、110、6、32、24、148、86、193、175、118、100、109、88、135、108、92、152、111、113、39、149、55、150、182、114の組み合わせ
(64)配列番号121、155、142、188、127、85、1、16、172、53、187、110、101、193、118、75、109、199、97、145、125、63、88、54、152、42、39、159、149、82、55、114、23の組み合わせ
(65)配列番号155、188、127、85、146、5、1、196、84、187、110、200、193、175、118、75、97、145、125、178、59、54、106、163、152、133、39、91、159、82、34、182、114の組み合わせ
(66)配列番号121、95、188、127、85、5、79、84、187、189、110、4、193、175、119、25、97、145、10、88、108、152、111、133、113、39、91、159、149、171、150、182、114の組み合わせ
(67)配列番号115、155、188、127、85、1、16、196、161、187、189、110、176、24、101、193、175、118、31、199、145、48、54、152、111、133、39、149、82、104、51、182、114の組み合わせ
(68)配列番号179、13、155、87、188、127、85、1、79、183、189、110、81、24、193、175、199、167、125、88、59、174、108、152、111、113、39、159、149、70、150、114、23の組み合わせ
(69)配列番号155、87、188、127、85、1、16、196、161、84、187、14、38、124、109、145、178、48、54、96、174、120、108、152、141、39、91、159、149、82、77、55、182、93、114の組み合わせ
(70)配列番号155、87、168、188、127、85、1、196、187、110、136、24、200、148、193、175、118、109、178、120、108、152、111、133、39、91、159、149、82、171、18、55、182、114の組み合わせ
(71)配列番号155、87、142、188、127、85、80、161、84、187、189、194、110、101、193、175、25、199、97、145、120、152、111、133、39、91、159、149、103、77、184、182、114、23の組み合わせ
(72)配列番号155、95、87、188、85、5、1、196、84、53、187、110、90、176、148、193、175、118、25、145、48、88、96、174、108、152、133、39、91、149、171、55、182、114の組み合わせ
(73)配列番号155、95、188、127、85、146、1、45、196、187、189、194、110、128、68、200、86、177、101、193、175、118、109、199、37、92、152、133、39、159、149、104、182、114の組み合わせ
(74)配列番号121、155、95、168、188、127、5、16、196、172、53、187、14、189、38、67、110、68、193、175、145、178、174、152、133、39、91、159、149、82、20、182、114、12の組み合わせ
(75)配列番号95、188、127、85、1、16、196、161、14、110、32、24、175、118、100、109、97、145、178、10、59、54、108、92、111、133、180、113、39、91、159、149、184、182、114、40の組み合わせ
(76)配列番号155、95、87、142、188、127、85、5、1、196、84、198、187、110、41、68、200、177、101、193、175、118、25、109、97、145、152、111、133、113、39、159、149、55、182、114の組み合わせ
(77)配列番号121、155、188、127、85、1、79、16、196、187、189、110、124、193、175、75、169、97、145、178、21、63、54、108、71、152、111、133、39、91、159、149、55、114、23の組み合わせ
(78)配列番号121、155、188、127、85、5、1、79、160、84、187、14、110、32、176、200、193、175、118、109、97、145、178、63、10、88、152、111、133、39、91、149、77、182、114の組み合わせ
(79)配列番号13、155、87、168、188、127、85、196、187、189、194、110、78、24、200、177、101、193、175、118、109、125、174、108、73、152、111、133、39、159、149、171、52、182、114の組み合わせ
(80)配列番号121、137、142、188、127、85、5、1、196、84、187、189、38、110、4、185、177、101、193、199、145、88、3、174、152、111、133、180、113、39、91、159、149、171、140、182、114の組み合わせ
(81)配列番号121、95、188、127、85、5、1、196、187、189、194、110、128、4、26、177、101、193、175、31、109、97、145、62、88、108、152、111、133、39、91、159、149、82、171、182、114の組み合わせ
(82)配列番号121、188、127、85、30、1、16、196、187、189、11、154、194、110、58、185、101、193、175、118、25、100、88、54、152、111、180、39、91、159、61、77、20、55、182、114、23の組み合わせ
(83)配列番号155、87、188、127、1、138、16、196、187、154、38、110、200、101、193、175、25、100、199、145、178、96、174、92、152、111、133、113、39、91、159、149、82、77、182、114、23の組み合わせ
(84)配列番号87、142、188、127、85、146、5、1、196、35、189、38、67、110、9、4、177、101、193、175、118、109、145、63、88、54、108、133、39、91、159、149、72、82、51、182、114の組み合わせ
(85)配列番号155、137、188、127、1、16、196、161、35、187、189、38、67、110、181、200、148、101、193、175、118、109、145、125、174、120、152、141、111、133、39、91、159、82、55、114の組み合わせ
(86)配列番号155、132、87、188、127、1、196、161、53、198、187、189、110、181、136、4、176、143、200、26、193、175、75、97、145、54、174、37、163、152、111、39、70、55、182、114の組み合わせ
(87)配列番号179、121、155、188、127、5、1、79、16、196、187、14、67、110、176、24、143、86、175、109、145、48、88、47、54、174、120、152、133、39、91、159、149、82、182、114の組み合わせ
(88)配列番号121、87、142、188、127、85、45、196、187、189、194、110、176、24、102、86、101、193、118、100、109、145、88、174、108、71、99、152、141、111、133、39、91、159、149、182、114、23の組み合わせ
(89)配列番号121、155、87、188、127、1、16、198、187、189、38、67、194、110、4、176、24、200、185、193、175、118、192、109、167、145、125、63、120、108、152、133、180、39、91、159、149、182の組み合わせ
(90)配列番号121、115、155、137、168、188、127、85、1、79、138、16、196、187、67、110、136、4、101、193、175、118、109、97、145、48、54、120、108、152、133、39、91、159、149、18、182、114の組み合わせ
(91)配列番号137、188、127、85、146、1、16、196、161、189、110、56、58、148、193、175、118、25、100、109、145、178、48、174、120、108、152、141、133、39、91、149、82、77、55、150、114の組み合わせ
(92)配列番号155、87、168、188、127、85、1、196、84、189、110、124、176、24、200、185、69、175、25、169、145、125、178、96、120、117、108、152、111、133、39、91、159、149、82、182、114の組み合わせ
(93)配列番号155、95、188、127、85、5、1、16、187、194、110、9、41、4、153、193、175、100、109、145、62、21、48、10、54、152、111、133、39、91、197、159、149、104、171、182、114の組み合わせ
(94)配列番号87、188、127、85、5、1、80、196、189、154、110、124、78、186、90、177、193、175、118、31、109、97、145、88、59、54、120、108、134、152、111、133、39、91、149、104、140、182、114の組み合わせ
(95)配列番号179、155、95、188、127、146、5、129、16、196、84、187、14、189、110、32、68、26、193、175、109、145、178、59、54、174、120、152、133、39、91、149、82、104、61、182、93、114、23の組み合わせ
(96)配列番号121、115、155、87、188、127、85、1、80、161、187、14、110、4、24、200、193、109、97、145、125、178、63、54、174、152、111、133、39、91、159、149、70、82、61、55、182、114の組み合わせ
(97)配列番号155、137、95、87、188、127、85、146、1、196、189、38、194、110、32、176、193、175、118、109、125、178、63、59、54、96、191、174、120、152、141、133、39、91、149、82、55、182、114の組み合わせ
(98)配列番号179、155、87、142、188、127、85、1、80、189、110、6、56、185、153、26、193、175、109、62、178、10、88、59、135、54、108、92、152、111、133、180、113、39、91、159、149、150、114、60の組み合わせ
(99)配列番号179、115、188、127、85、1、79、183、196、53、187、189、194、110、200、185、193、175、119、105、97、145、125、88、54、120、152、113、39、91、116、159、149、34、61、150、182、114、23の組み合わせ
(100)配列番号87、188、127、85、16、196、84、14、189、67、110、90、24、193、175、43、100、109、145、48、63、10、88、59、54、174、108、152、111、133、39、91、159、149、82、104、61、182、114の組み合わせ
(101)配列番号155、137、142、188、127、85、5、1、16、196、84、53、187、38、67、110、4、176、24、177、101、193、175、100、109、145、48、88、54、71、152、89、133、39、91、159、149、104、182、93、114の組み合わせ
(102)配列番号87、188、127、1、80、183、45、187、189、110、128、4、200、102、26、177、101、193、175、145、178、88、96、174、117、92、152、111、133、39、91、159、149、82、171、51、55、150、182、93、114、12の組み合わせ
(103)配列番号121、155、188、127、85、146、196、53、187、189、110、32、56、26、101、193、175、100、109、199、97、145、178、48、63、10、88、59、54、108、152、111、133、39、7、197、159、149、82、182、114の組み合わせ
(104)配列番号121、155、87、188、127、85、1、80、84、198、187、189、194、110、200、102、86、193、175、118、75、97、145、125、178、88、59、37、92、134、152、111、39、159、149、82、20、51、150、182、114の組み合わせ
(105)配列番号137、95、87、142、188、127、85、1、79、196、187、189、67、110、136、41、4、24、19、177、101、193、118、100、105、97、48、54、191、174、108、152、111、133、39、159、149、171、18、182、114の組み合わせ
(106)配列番号95、87、188、127、146、5、196、161、84、187、14、189、154、194、110、128、176、24、68、200、185、193、175、43、109、97、145、178、48、54、174、92、152、113、39、159、149、82、55、114、23の組み合わせ
(107)配列番号121、155、137、87、188、127、85、79、129、16、84、14、189、67、110、124、136、41、176、24、68、185、193、175、109、97、145、178、120、73、152、111、133、113、39、91、159、149、150、182、114の組み合わせ
(108)配列番号155、137、168、188、127、85、1、80、79、183、196、172、84、187、189、110、41、68、185、175、75、100、109、97、145、178、48、98、88、135、54、108、92、73、111、133、39、91、149、82、150、114の組み合わせ
(109)配列番号13、155、132、87、168、188、127、5、1、196、84、110、6、124、41、90、176、81、24、143、68、200、101、193、175、100、109、169、178、47、174、152、111、133、39、91、149、82、61、18、182、114の組み合わせ
(110)配列番号121、115、155、168、142、188、127、85、1、16、196、161、187、14、189、67、110、4、176、200、193、175、25、100、167、145、178、59、54、174、108、152、111、133、113、39、91、159、82、104、182、114、23の組み合わせ
(111)配列番号179、115、142、188、127、146、79、196、161、187、189、194、110、4、68、177、101、193、175、100、109、145、48、59、174、108、99、152、133、113、39、149、70、82、104、140、61、103、77、55、182、114、50の組み合わせ
(112)配列番号13、121、155、132、95、87、168、188、127、1、16、196、84、198、189、38、110、124、4、176、24、200、102、148、185、193、175、25、109、145、125、178、10、59、37、108、141、111、133、39、159、149、82、150、182、114の組み合わせ
(113)配列番号179、121、95、188、127、85、1、79、16、84、53、198、187、38、194、110、136、78、4、176、193、175、75、100、109、97、145、48、88、59、54、37、152、111、113、39、91、159、149、82、171、46、150、182、114の組み合わせ
(114)配列番号179、115、155、87、168、188、127、85、1、16、196、198、14、110、176、24、143、68、200、193、175、118、109、97、125、178、88、83、59、47、54、120、108、163、111、133、39、91、159、149、82、46、55、182、114の組み合わせ
(1)配列番号127、1、110の組み合わせ
(2)配列番号1、110、3、114の組み合わせ
(3)配列番号137、1、110、111、114の組み合わせ
(4)配列番号155、127、5、1、154、110の組み合わせ
(5)配列番号155、87、1、154、110、73、133、149、114の組み合わせ
(6)配列番号30、1、110、128、145、3、159、114の組み合わせ
(7)配列番号127、5、1、154、110、3、73、114の組み合わせ
(8)配列番号155、5、1、110、100、145、54、133、114の組み合わせ
(9)配列番号188、5、1、38、110、152、133、159、114の組み合わせ
(10)配列番号155、1、110、100、145、54、152、133、159、114の組み合わせ
(11)配列番号137、188、127、5、1、110、145、3、152、180、114の組み合わせ
(12)配列番号155、188、1、110、75、3、71、152、133、159、149、114の組み合わせ
(13)配列番号179、121、137、5、1、183、110、185、3、54、152、114の組み合わせ
(14)配列番号155、188、127、30、5、1、53、110、109、145、152、159の組み合わせ
(15)配列番号155、188、127、1、196、189、110、109、71、152、133、149、114の組み合わせ
(16)配列番号155、137、127、1、110、193、175、109、3、152、133、180、114の組み合わせ
(17)配列番号155、142、1、154、110、100、109、145、152、133、39、104、114の組み合わせ
(18)配列番号155、137、188、127、5、1、187、189、110、26、109、71、152、114の組み合わせ
(19)配列番号155、188、127、1、189、110、78、185、100、71、152、133、149、114の組み合わせ
(20)配列番号121、155、188、1、79、198、154、110、100、109、54、133、104、114の組み合わせ
(21)配列番号155、142、188、127、1、196、189、110、175、100、109、145、3、174、114の組み合わせ
(22)配列番号121、155、188、127、1、110、4、100、109、145、71、152、133、82、114の組み合わせ
(23)配列番号155、142、188、127、1、189、110、100、109、174、71、133、57、51、114の組み合わせ
(24)配列番号155、188、127、1、154、110、58、100、109、54、71、73、133、149、114の組み合わせ
(25)配列番号155、142、188、127、1、189、110、100、145、174、152、133、149、51、114の組み合わせ
(26)配列番号179、155、137、188、127、1、138、110、193、100、145、10、152、51、114の組み合わせ
(27)配列番号155、142、188、30、1、53、110、109、145、54、96、174、152、2、51、114の組み合わせ
(28)配列番号155、188、127、30、1、189、194、110、100、109、145、3、71、152、133、82、114の組み合わせ
(29)配列番号155、137、188、127、85、30、1、110、175、100、145、54、71、152、159、114の組み合わせ
(30)配列番号155、188、127、1、110、78、185、175、100、145、152、133、149、82、51の組み合わせ
(31)配列番号155、188、127、1、198、110、86、175、100、109、174、152、133、159、149、93、114の組み合わせ
(32)配列番号155、188、127、1、110、175、145、3、152、133、180、39、159、149、77、114の組み合わせ
(33)配列番号155、188、127、5、1、196、53、154、110、58、54、152、133、82、114の組み合わせ
(34)配列番号179、188、127、1、110、102、175、75、100、109、3、54、152、190、133、159、2の組み合わせ
(35)配列番号155、137、188、127、1、196、189、110、109、145、3、73、152、133、2、82、114の組み合わせ
(36)配列番号155、188、127、1、198、110、68、175、75、100、54、71、73、133、149、77、114の組み合わせ
(37)配列番号121、155、87、188、127、30、1、161、110、193、109、145、135、71、152、82、114の組み合わせ
(38)配列番号155、87、188、127、1、198、154、110、100、174、73、152、133、159、82、51、114の組み合わせ
(39)配列番号155、137、87、188、127、1、198、110、68、101、100、109、145、54、152、149、51、114の組み合わせ
(40)配列番号121、155、188、127、1、198、110、75、109、145、54、152、133、82、104、114の組み合わせ
(41)配列番号155、137、87、142、188、127、1、110、185、175、100、145、174、152、141、133、180の組み合わせ
(42)配列番号155、142、188、127、1、189、154、110、71、73、152、159、149、82、51、182、114の組み合わせ
(43)配列番号155、188、127、5、1、79、154、110、75、3、54、108、71、73、152、133、114の組み合わせ
(44)配列番号155、142、127、1、189、154、110、109、3、174、108、71、133、91、159、82、104、114の組み合わせ
(45)配列番号121、137、188、127、1、196、189、110、175、109、3、54、191、152、133、149、82、114の組み合わせ
(46)配列番号142、188、127、1、198、110、175、100、109、145、133、180、91、159、149、82、182、114の組み合わせ
(47)配列番号121、155、137、188、127、85、5、1、110、100、109、3、54、73、152、133、159、114の組み合わせ
(48)配列番号155、188、127、1、138、198、110、68、100、109、48、3、71、152、141、133、91、182の組み合わせ
(49)配列番号155、137、142、188、127、1、110、109、174、71、152、111、133、180、159、149、51、114の組み合わせ
(50)配列番号155、137、188、127、30、1、161、154、194、110、178、10、152、111、133、180、114の組み合わせ
(51)配列番号179、155、188、127、1、161、110、175、109、62、125、10、54、71、152、133、180、114の組み合わせ
(52)配列番号155、188、127、1、110、193、175、145、48、10、71、152、141、133、180、182、114の組み合わせ
(53)配列番号155、137、87、188、127、1、189、194、110、193、100、10、111、133、180、159、149、114の組み合わせ
(54)配列番号155、188、5、1、154、110、109、3、54、71、73、163、152、133、180、82、104、114の組み合わせ
(55)配列番号155、137、188、127、30、1、17、110、193、145、71、152、111、133、180、77、150、114の組み合わせ
(56)配列番号155、137、87、142、188、127、1、198、110、58、175、109、10、152、133、149、82、93、114の組み合わせ
(57)配列番号155、188、127、5、1、196、110、68、193、109、10、54、152、133、180、82、114の組み合わせ
(58)配列番号155、142、188、127、1、110、75、109、145、10、3、71、152、133、82、51、114の組み合わせ
(59)配列番号121、155、142、188、127、1、161、154、110、175、109、71、73、152、133、149、51、114の組み合わせ
(60)配列番号155、137、188、127、5、1、198、154、110、193、109、3、71、152、180、159、82、104、114の組み合わせ
(61)配列番号121、155、137、142、188、127、1、154、110、175、109、108、133、159、149、82、77、114の組み合わせ
(62)配列番号155、188、127、1、189、110、26、193、175、109、145、73、152、141、133、149、82、77、114の組み合わせ
(63)配列番号155、188、127、1、162、198、189、154、110、75、100、109、54、71、152、133、82、104、114の組み合わせ
(64)配列番号121、155、137、188、127、1、110、185、69、100、109、3、174、71、133、180、159、82、114の組み合わせ
(65)配列番号179、155、188、127、1、154、110、86、193、175、75、109、145、71、133、159、82、114の組み合わせ
(66)配列番号155、87、188、127、1、84、110、193、175、100、109、71、152、133、149、82、104、77、114の組み合わせ
(67)配列番号155、137、188、127、1、110、175、100、145、10、135、3、71、152、133、180、149、77、114の組み合わせ
(68)配列番号155、142、188、127、1、189、154、110、175、109、145、125、174、71、152、133、91、82、114の組み合わせ
(69)配列番号121、142、188、127、5、1、189、154、110、100、109、145、125、174、108、71、152、82、114の組み合わせ
(70)配列番号155、137、87、188、127、1、198、110、185、175、100、109、145、92、152、133、180、149、114の組み合わせ
(71)配列番号155、137、87、188、127、1、161、198、110、100、109、73、152、141、133、149、182、114の組み合わせ
(72)配列番号155、142、188、1、79、198、154、110、78、185、175、100、10、152、133、180、159、51、114の組み合わせ
(73)配列番号155、137、142、188、127、1、196、189、110、78、143、200、185、145、152、133、180、159の組み合わせ
(74)配列番号155、137、188、127、1、198、110、41、101、109、145、174、73、152、159、82、51、182、114の組み合わせ
(75)配列番号155、142、188、127、1、80、189、110、185、153、109、145、178、174、111、133、82、104、114の組み合わせ
(76)配列番号179、155、142、188、127、1、198、194、110、68、185、101、100、109、96、152、133、180、149の組み合わせ
(77)配列番号121、155、87、188、127、5、1、198、194、110、68、193、145、152、180、159、149、182、114の組み合わせ
(78)配列番号155、137、127、1、198、110、175、100、109、54、152、133、180、39、91、149、2、82、114の組み合わせ
(79)配列番号121、155、137、87、142、188、127、1、198、110、56、193、109、10、3、152、180、104、182、114の組み合わせ
(80)配列番号121、137、87、188、127、1、187、189、110、193、109、145、135、3、152、133、159、2、82、114の組み合わせ
(81)配列番号155、137、188、127、1、79、198、110、86、100、109、3、174、71、152、133、91、149、82、114の組み合わせ
(82)配列番号155、137、188、127、1、198、110、193、100、109、145、98、3、71、152、133、159、82、150、114の組み合わせ
(83)配列番号155、137、188、127、1、110、153、193、75、109、88、174、71、152、133、180、149、82、51、114の組み合わせ
(84)配列番号155、87、188、127、1、161、198、110、175、109、169、145、178、10、92、133、159、82、104、114の組み合わせ
(85)配列番号155、87、142、188、127、1、154、194、110、101、75、100、109、145、152、133、180、82、114の組み合わせ
(86)配列番号155、137、87、127、1、110、185、175、173、109、145、133、180、91、159、149、104、77、114の組み合わせ
(87)配列番号155、142、188、127、1、196、110、58、185、175、75、100、109、145、125、3、91、2、82、114の組み合わせ
(88)配列番号121、137、142、188、127、1、154、110、175、100、109、145、88、3、54、174、152、133、82、182、114の組み合わせ
(89)配列番号121、137、87、188、127、1、53、110、185、175、109、152、133、91、159、149、82、104、77、150、114の組み合わせ
(90)配列番号155、142、188、127、30、1、196、198、189、154、110、109、145、10、174、152、133、159、82、114の組み合わせ
(91)配列番号179、155、188、127、1、198、110、101、175、118、109、145、3、37、73、133、91、149、82、77、114の組み合わせ
(92)配列番号155、137、87、188、127、1、196、154、110、68、26、178、71、152、133、180、91、149、82、114の組み合わせ
(93)配列番号121、155、137、188、127、1、79、198、110、86、75、100、109、3、152、133、149、2、82、182、114の組み合わせ
(94)配列番号155、137、188、127、5、1、84、198、194、110、75、100、109、145、48、71、152、133、82、114の組み合わせ
(95)配列番号155、142、188、127、1、79、196、154、110、41、31、109、145、48、54、174、108、71、133、82、114の組み合わせ
(96)配列番号121、155、188、127、1、161、198、189、110、78、100、178、71、152、133、159、149、104、182、114の組み合わせ
(97)配列番号155、188、127、1、79、189、110、176、175、109、145、48、71、73、152、133、91、159、82、77、114の組み合わせ
(98)配列番号155、87、142、188、127、1、196、17、189、110、153、101、175、100、109、133、180、149、51、114の組み合わせ
(99)配列番号121、155、188、127、85、1、198、189、154、110、109、145、125、178、71、152、141、133、180、159、114の組み合わせ
(100)配列番号155、188、127、1、198、187、110、68、175、100、109、145、125、10、3、71、152、133、82、182、114の組み合わせ
(101)配列番号121、155、87、142、188、127、1、84、110、26、75、100、71、106、152、180、159、149、2、114の組み合わせ
(102)配列番号155、142、188、127、1、189、154、110、78、185、153、177、109、178、54、174、92、133、82、57、114の組み合わせ
(103)配列番号155、142、188、127、146、1、189、154、110、185、153、175、100、109、145、92、152、133、82、104、114の組み合わせ
(104)配列番号155、137、188、5、1、196、198、154、110、193、109、145、135、3、152、159、104、171、182、114の組み合わせ
(105)配列番号121、155、188、127、1、198、110、175、75、109、145、3、96、71、152、133、180、104、51、182、114の組み合わせ
(106)配列番号121、155、137、188、127、30、1、79、110、58、193、109、145、156、71、152、133、149、182、114の組み合わせ
(107)配列番号121、155、87、127、1、158、161、84、110、175、100、109、48、3、54、190、133、91、159、82、114の組み合わせ
(108)配列番号155、188、127、1、196、198、110、185、175、75、100、109、145、152、133、39、91、149、82、182、114の組み合わせ
(109)配列番号155、137、142、188、127、1、196、161、154、110、185、109、199、145、178、3、71、133、82、114、12の組み合わせ
(110)配列番号121、155、137、87、188、127、5、1、198、189、194、110、75、100、109、145、3、152、159、82、114の組み合わせ
(111)配列番号155、137、87、188、127、85、1、198、154、110、128、185、101、145、10、3、152、180、159、149、51、114の組み合わせ
(112)配列番号155、137、142、188、127、1、161、194、110、101、100、145、178、3、54、152、133、180、149、51、182、114の組み合わせ
(113)配列番号155、87、188、127、1、161、198、189、110、185、193、175、109、145、73、133、159、149、82、61、18、114の組み合わせ
(114)配列番号121、155、87、188、127、1、16、161、198、154、110、109、145、178、10、174、71、133、159、2、82、114の組み合わせ
(115)配列番号121、155、188、127、1、161、189、110、185、177、100、109、125、71、133、159、149、82、104、182、114の組み合わせ
(116)配列番号121、137、188、127、30、1、110、128、193、175、100、199、145、10、152、133、180、159、149、104、51、114の組み合わせ
(117)配列番号155、188、127、85、1、161、198、154、110、175、100、109、178、54、92、152、133、82、104、77、51、114の組み合わせ
(118)配列番号155、142、188、127、146、1、198、189、38、110、68、193、192、100、109、145、10、152、133、159、149、82、114の組み合わせ
(119)配列番号155、137、142、188、127、1、198、189、154、110、68、192、100、109、145、152、133、159、149、82、114の組み合わせ
(120)配列番号155、142、188、127、1、194、110、9、200、148、86、109、145、125、174、92、163、152、159、149、82、114の組み合わせ
(121)配列番号121、155、137、188、127、1、161、198、110、86、109、135、3、71、152、133、180、149、82、104、182、114の組み合わせ
(122)配列番号155、142、188、127、146、1、189、110、143、101、175、75、109、145、135、174、152、133、159、82、182、114の組み合わせ
(123)配列番号155、137、87、142、188、127、30、1、196、198、110、58、193、175、109、145、73、133、180、149、182、114の組み合わせ
(124)配列番号155、137、188、127、1、189、110、78、185、175、100、109、145、178、152、133、180、159、149、82、182、114の組み合わせ
(125)配列番号155、87、142、188、127、1、53、198、110、193、175、109、97、3、152、133、39、91、159、149、82、93、114の組み合わせ
(126)配列番号121、155、137、87、188、127、1、84、198、110、102、175、75、100、109、145、54、152、133、149、77、114の組み合わせ
(127)配列番号121、155、137、188、127、5、1、84、189、154、194、110、75、3、152、133、180、159、149、104、77、114の組み合わせ
(128)配列番号155、137、87、188、127、1、196、161、110、143、86、175、100、109、63、71、133、39、91、159、149、82、114の組み合わせ
(129)配列番号121、155、137、87、188、127、1、196、110、9、193、100、109、135、54、71、73、111、133、149、104、182、114の組み合わせ
(130)配列番号155、137、87、188、127、1、53、198、189、110、153、175、100、109、145、10、88、190、133、180、91、149、114の組み合わせ
(131)配列番号121、155、87、142、188、127、1、196、198、110、128、41、101、175、100、109、145、54、152、150、182、93、114の組み合わせ
(132)配列番号179、155、87、142、188、127、5、1、198、154、194、110、193、175、199、145、3、71、152、133、159、82、114の組み合わせ
(133)配列番号121、155、87、127、1、189、154、110、193、100、109、10、3、152、133、180、159、2、82、104、182、114の組み合わせ
(134)配列番号155、142、188、127、1、162、196、154、110、200、101、193、175、109、125、71、133、180、149、82、77、114の組み合わせ
(135)配列番号155、137、188、127、1、161、198、154、194、110、100、145、178、3、71、73、152、133、159、149、82、114の組み合わせ
(136)配列番号179、155、87、188、127、5、1、154、110、200、193、175、100、109、145、3、133、91、159、149、82、182、114の組み合わせ
(137)配列番号179、121、155、87、188、127、1、161、198、154、143、193、175、192、100、109、71、133、149、104、182、114の組み合わせ
(138)配列番号155、137、87、188、127、1、198、189、38、194、110、175、100、109、145、10、152、133、149、82、57、114の組み合わせ
(139)配列番号155、137、87、142、188、127、1、154、194、110、193、175、145、156、54、152、133、180、159、149、77、57、114の組み合わせ
(140)配列番号155、137、188、127、5、1、84、154、110、175、192、100、109、145、10、108、152、133、180、159、149、82、114の組み合わせ
(141)配列番号155、188、127、146、1、196、189、194、110、200、86、193、100、109、152、39、159、149、82、77、182、114の組み合わせ
(142)配列番号155、188、127、146、1、196、161、154、38、194、110、56、78、68、175、100、73、152、133、159、149、182、114の組み合わせ
(143)配列番号155、137、87、142、188、127、5、1、84、189、154、194、110、175、75、145、152、133、180、159、77、114の組み合わせ
(144)配列番号121、137、87、142、188、127、1、154、194、110、128、185、175、100、83、71、73、152、133、149、82、182、114の組み合わせ
(145)配列番号155、188、127、5、1、84、154、110、176、58、102、148、109、145、135、3、54、191、152、133、180、82、114の組み合わせ
(146)配列番号155、137、142、188、127、5、1、196、84、189、154、110、109、145、135、96、174、152、133、180、91、159、82、114の組み合わせ
(147)配列番号121、155、137、87、188、127、1、198、189、110、26、177、175、100、109、145、48、98、54、108、133、159、82、93、114の組み合わせ
(148)配列番号179、155、142、188、127、1、154、194、110、56、175、109、145、88、3、174、73、152、133、82、171、150、114の組み合わせ
(149)配列番号179、155、137、142、188、127、1、84、189、110、148、193、109、145、88、3、71、152、180、159、82、150、114の組み合わせ
(150)配列番号179、155、137、87、142、188、127、5、1、161、198、110、41、100、109、145、174、71、152、159、149、82、182、114の組み合わせ
(151)配列番号155、107、188、127、1、79、198、110、58、100、109、145、125、10、54、174、106、152、133、82、57、51、114の組み合わせ
(152)配列番号155、137、87、142、188、127、1、53、198、110、41、185、175、100、109、169、49、3、54、71、133、149、182、114の組み合わせ
(153)配列番号121、155、137、87、188、127、1、198、110、58、100、109、3、54、191、71、133、180、91、149、2、77、182、114の組み合わせ
(154)配列番号155、137、87、142、188、127、1、198、110、26、101、175、109、145、178、152、133、180、149、61、150、182、114の組み合わせ
(155)配列番号121、155、87、188、127、1、162、110、86、193、175、109、145、3、71、152、133、159、149、82、77、150、114の組み合わせ
(156)配列番号121、142、188、127、1、196、154、110、56、153、177、109、21、10、3、54、174、71、163、159、82、104、182、114の組み合わせ
(157)配列番号155、137、188、127、85、1、138、110、86、26、193、175、100、145、156、10、88、3、152、111、2、51、182、114の組み合わせ
(158)配列番号155、137、188、127、1、189、110、136、68、26、193、175、100、105、145、3、71、152、111、133、82、104、77、114の組み合わせ
(159)配列番号155、137、87、188、127、1、161、198、154、110、68、185、175、100、109、178、10、54、108、71、133、82、104、114の組み合わせ
(160)配列番号121、155、137、87、188、127、1、161、189、194、110、186、200、100、145、3、191、108、133、91、159、149、82、114の組み合わせ
(161)配列番号121、155、188、127、1、198、154、110、185、86、193、175、109、145、178、135、3、73、152、133、159、82、182、114の組み合わせ
(162)配列番号121、155、137、188、127、1、161、53、198、110、102、175、100、109、178、54、174、71、133、82、150、182、114の組み合わせ
(163)配列番号155、142、188、127、85、1、161、189、154、110、185、175、109、145、178、54、174、71、133、91、82、57、93、114の組み合わせ
(164)配列番号121、155、137、142、188、127、1、198、110、193、175、100、145、174、71、106、152、159、149、82、55、182、93、114の組み合わせ
(165)配列番号121、155、188、127、146、30、1、189、38、110、68、193、175、192、100、145、152、133、180、159、149、150、114の組み合わせ
(166)配列番号155、137、87、188、127、1、84、198、154、38、110、176、109、48、71、152、111、190、133、91、159、149、182、93、114の組み合わせ
(167)配列番号121、155、137、87、142、188、127、1、198、110、9、175、100、109、145、54、174、152、133、149、82、104、182、114の組み合わせ
(168)配列番号121、155、188、127、1、196、172、198、154、110、145、48、135、3、152、133、39、159、82、104、51、182、114の組み合わせ
(169)配列番号155、188、127、146、5、1、189、110、200、185、26、193、100、109、145、96、71、152、133、159、82、77、114の組み合わせ
(170)配列番号121、155、188、127、1、198、110、123、41、58、175、100、109、145、125、21、54、147、152、133、149、82、182、114の組み合わせ
(171)配列番号155、137、87、142、188、127、1、198、110、193、175、100、109、145、178、10、3、54、174、152、133、91、149、114の組み合わせ
(172)配列番号155、87、188、127、1、196、161、198、154、110、186、68、200、175、100、109、48、73、152、133、159、77、182、114の組み合わせ
(173)配列番号121、155、87、188、127、1、196、189、110、200、175、109、145、48、3、54、191、71、133、159、149、82、61、77、114の組み合わせ
(174)配列番号155、137、142、188、127、1、79、198、154、110、75、100、109、145、178、96、174、92、152、133、180、91、159、182、114の組み合わせ
(175)配列番号121、155、137、87、142、188、127、1、161、154、110、175、100、109、145、62、3、174、133、180、91、159、82、114の組み合わせ
(176)配列番号155、137、188、127、1、183、196、198、189、154、38、194、110、68、175、100、109、145、152、133、159、149、82、77、114の組み合わせ
(177)配列番号155、142、188、127、1、138、196、189、154、110、185、101、175、75、100、109、145、62、178、3、133、149、82、114の組み合わせ
(178)配列番号121、155、137、87、188、127、1、196、198、110、175、100、109、145、3、71、152、133、159、82、77、150、182、93、114の組み合わせ
(179)配列番号155、87、142、188、127、1、189、110、193、175、109、145、62、178、3、71、133、180、91、159、149、82、104、182、93、114の組み合わせ
(180)配列番号179、155、137、87、188、127、30、1、161、198、189、154、110、175、100、109、145、178、71、133、39、91、159、82、114の組み合わせ
(181)配列番号179、121、155、142、188、127、1、161、172、53、198、110、175、100、109、105、3、54、133、149、82、77、182、93、114の組み合わせ
(182)配列番号179、121、155、188、127、5、1、154、110、68、193、175、100、109、145、10、3、108、152、133、91、159、149、82、114の組み合わせ
(183)配列番号179、121、155、142、188、127、1、110、128、175、100、109、125、178、10、135、3、174、92、133、42、149、82、182、114の組み合わせ
(184)配列番号121、155、188、127、85、5、1、79、198、154、110、145、135、54、71、73、152、133、39、82、104、51、182、114の組み合わせ
(185)配列番号121、155、87、142、188、127、1、161、198、154、110、185、193、109、178、48、174、152、133、159、2、82、104、114の組み合わせ
(186)配列番号155、142、188、127、1、198、189、110、193、175、100、109、62、48、191、174、71、152、133、91、159、149、82、104、182、114の組み合わせ
(187)配列番号155、87、142、188、127、85、30、1、198、194、110、78、58、148、105、145、3、174、152、133、180、159、149、57、114の組み合わせ
(188)配列番号121、155、137、87、188、127、1、161、53、198、194、110、102、175、173、100、109、3、73、133、149、82、104、93、114の組み合わせ
(189)配列番号121、155、137、87、188、127、1、196、198、189、154、110、185、109、178、3、54、174、71、152、133、149、82、114の組み合わせ
(190)配列番号155、87、188、127、5、1、196、161、198、110、68、185、100、109、145、63、54、37、108、133、159、2、82、104、114の組み合わせ
(191)配列番号155、142、188、85、1、138、196、53、198、189、110、44、101、175、100、145、152、180、159、149、2、82、51、182、93、114の組み合わせ
(192)配列番号155、137、87、188、127、1、161、198、189、154、194、110、143、193、175、109、145、3、111、133、39、91、149、104、93、114の組み合わせ
(193)配列番号179、121、155、188、127、1、198、110、200、148、175、75、100、109、145、125、63、54、174、92、190、133、91、82、104の組み合わせ
(194)配列番号121、155、142、188、94、1、161、154、110、185、175、75、100、109、3、96、174、108、92、152、133、180、91、2、82、114の組み合わせ
(195)配列番号121、155、137、87、188、127、1、84、198、110、148、175、75、100、109、145、178、3、54、174、152、133、180、91、149、114の組み合わせ
(196)配列番号121、155、87、188、127、1、196、198、110、148、185、75、109、145、3、54、96、174、71、152、133、91、159、2、82、114の組み合わせ
(197)配列番号155、142、188、127、94、1、84、189、194、110、9、185、175、75、109、145、88、135、71、152、190、133、180、82、182、114の組み合わせ
(198)配列番号155、87、142、188、127、1、79、161、198、154、110、100、109、178、174、92、152、133、91、159、149、82、55、93、114の組み合わせ
(199)配列番号179、155、137、87、188、127、1、196、161、198、189、110、86、175、100、109、108、152、133、39、159、149、82、77、93、114の組み合わせ
(200)配列番号155、188、127、1、45、194、110、157、68、200、86、193、75、109、145、125、3、152、133、159、149、2、82、51、150、114の組み合わせ
(201)配列番号121、155、137、87、127、1、84、198、189、110、185、177、193、100、109、62、10、88、191、133、91、159、82、104、182、114の組み合わせ
(202)配列番号121、155、87、142、188、127、1、196、172、198、110、102、26、193、175、109、145、62、71、73、133、149、82、51、114の組み合わせ
(203)配列番号13、121、87、142、188、127、30、1、198、154、110、175、192、109、145、178、10、174、152、180、91、159、149、82、182、114の組み合わせ
(204)配列番号155、137、87、142、188、127、1、79、196、84、198、110、175、100、109、145、63、152、190、133、180、91、159、149、93、114の組み合わせ
(205)配列番号155、87、188、127、1、196、84、198、110、128、56、185、175、109、178、3、174、133、180、39、159、149、82、77、182、114の組み合わせ
(206)配列番号155、87、142、188、127、5、1、79、196、161、154、110、41、58、102、175、100、109、145、48、71、133、91、82、93、114の組み合わせ
(207)配列番号155、188、127、1、194、110、200、86、193、175、100、109、145、125、88、3、71、152、133、180、39、159、149、77、114、50の組み合わせ
(208)配列番号179、155、137、87、188、127、85、1、161、84、110、148、153、101、175、109、178、63、71、152、133、91、159、149、77、114の組み合わせ
(209)配列番号155、137、87、188、127、1、161、198、110、193、175、100、109、97、178、48、54、174、108、152、133、159、82、61、114の組み合わせ
(210)配列番号179、155、137、142、188、127、1、161、198、189、110、58、175、100、109、10、3、108、73、152、133、159、82、77、182、114の組み合わせ
(211)配列番号121、155、87、142、188、127、1、161、198、154、110、193、175、192、100、109、169、73、163、133、159、149、82、77、182、93、114の組み合わせ
(212)配列番号179、155、87、188、127、1、161、198、189、154、194、110、68、86、175、25、199、152、133、180、149、2、104、77、114の組み合わせ
(213)配列番号155、137、87、142、188、127、1、198、154、110、68、86、75、100、54、174、92、152、111、190、133、180、159、57、114の組み合わせ
(214)配列番号155、87、188、127、1、196、161、198、110、26、193、175、100、109、145、178、48、71、73、152、133、149、82、104、182、114の組み合わせ
(215)配列番号155、87、188、127、1、161、198、110、185、175、25、100、109、178、54、108、152、141、133、91、149、82、104、182、114の組み合わせ
(216)配列番号155、87、142、188、127、5、1、198、154、110、148、101、193、175、109、178、48、135、73、152、133、91、159、82、51、182、114の組み合わせ
(217)配列番号155、87、188、127、146、30、1、196、198、154、38、110、193、175、100、167、178、54、133、180、159、2、77、18、20、182、114の組み合わせ
(218)配列番号121、155、87、142、188、127、146、1、196、198、189、154、38、110、177、100、109、178、174、152、133、91、159、82、182、114の組み合わせ
(219)配列番号87、188、127、1、196、161、84、53、198、110、200、193、175、109、48、135、54、174、71、133、180、91、159、82、104、182、114の組み合わせ
(220)配列番号155、142、188、127、1、53、110、81、185、153、101、175、173、109、117、152、133、180、39、91、159、149、2、82、182、114の組み合わせ
(221)配列番号121、155、87、188、127、1、196、161、198、154、110、58、175、109、145、125、178、3、191、92、152、133、149、82、93、114の組み合わせ
(222)配列番号155、137、87、142、188、127、1、198、154、110、175、100、109、145、178、3、47、54、133、39、91、149、82、104、182、114の組み合わせ
(223)配列番号179、155、142、188、127、5、1、161、198、154、110、143、69、193、175、173、109、145、178、10、174、71、159、82、93、114の組み合わせ
(224)配列番号179、155、137、87、188、127、1、161、84、194、110、185、175、109、178、63、71、152、133、180、91、159、149、82、77、182、114の組み合わせ
(225)配列番号155、87、142、188、127、30、1、198、154、110、176、100、109、145、48、135、3、71、152、180、39、149、82、51、93、114の組み合わせ
(226)配列番号121、155、137、87、188、127、1、198、194、110、193、175、100、109、199、145、10、3、71、133、149、2、82、77、93、114の組み合わせ
(227)配列番号121、155、137、87、127、1、79、198、154、110、185、175、109、145、178、10、3、152、91、159、149、82、104、52、182、114の組み合わせ
(228)配列番号121、155、137、87、142、127、1、194、110、86、26、193、175、25、75、109、145、3、174、152、133、180、39、159、82、182、114の組み合わせ
(229)配列番号155、137、142、188、127、1、196、161、198、110、143、185、153、175、109、145、62、178、48、174、73、152、133、159、82、104、114の組み合わせ
(230)配列番号155、137、142、188、127、30、1、198、194、110、9、58、86、175、100、167、145、98、152、133、180、159、82、104、51、150、114の組み合わせ
(231)配列番号121、155、87、188、127、1、79、161、198、154、110、78、175、100、109、54、174、71、152、133、180、149、82、182、93、114の組み合わせ
(232)配列番号155、87、142、188、127、1、80、161、53、187、189、154、110、175、75、31、109、62、48、54、191、92、71、133、149、82、114の組み合わせ
(233)配列番号155、137、87、188、127、5、1、161、84、198、194、110、175、100、109、145、3、133、180、159、149、82、104、150、182、114の組み合わせ
(234)配列番号155、137、87、142、188、127、1、53、198、189、154、110、175、192、100、109、145、125、178、54、174、133、159、82、182、114の組み合わせ
(235)配列番号155、142、188、127、146、1、53、189、154、38、194、110、185、175、100、109、145、135、3、96、174、71、133、159、82、182、114の組み合わせ
(236)配列番号155、188、127、5、1、198、110、176、68、175、145、125、98、10、3、71、152、111、133、39、159、149、82、104、51、150、114の組み合わせ
(237)配列番号155、137、142、188、127、146、1、189、194、110、143、200、193、175、100、109、145、156、10、152、180、91、159、82、77、182、114の組み合わせ
(238)配列番号179、155、142、188、127、1、196、198、189、110、44、78、148、185、175、100、109、105、167、145、62、152、133、39、82、114の組み合わせ
(239)配列番号179、87、142、188、127、1、196、161、53、198、154、110、58、153、175、109、10、3、141、133、180、91、149、82、182、114の組み合わせ
(240)配列番号121、155、87、188、127、1、79、158、196、161、198、38、110、102、175、100、109、135、54、108、71、133、149、82、104、114の組み合わせ
(241)配列番号121、155、137、87、188、127、94、1、84、194、110、148、193、175、75、100、109、145、178、3、54、152、133、149、2、82、77、114の組み合わせ
(242)配列番号155、137、87、188、127、1、161、198、154、110、185、175、109、145、178、63、108、71、152、133、91、159、149、82、104、77、150、114の組み合わせ
(243)配列番号155、142、188、127、30、1、161、189、110、153、193、175、109、145、62、178、98、3、191、71、152、133、39、149、82、51、182、114の組み合わせ
(244)配列番号121、155、137、142、188、127、30、1、196、161、198、110、193、175、100、109、10、174、152、133、180、91、149、82、51、182、114の組み合わせ
(245)配列番号155、137、87、142、188、127、1、196、161、198、110、78、90、193、175、100、109、145、71、73、133、159、149、82、51、182、114の組み合わせ
(246)配列番号155、137、87、188、127、1、79、196、161、189、110、153、175、100、109、145、62、54、191、108、71、73、133、91、159、149、82、114の組み合わせ
(247)配列番号179、155、142、188、127、1、79、158、189、154、110、4、185、175、100、109、145、54、174、71、152、133、159、82、51、93、114の組み合わせ
(248)配列番号179、155、142、188、127、1、80、198、187、189、110、148、175、100、109、97、145、135、54、174、92、152、133、149、82、51、114の組み合わせ
(249)配列番号179、121、155、142、188、127、5、1、194、110、9、4、75、100、109、145、125、178、3、54、92、152、133、91、82、182、114の組み合わせ
(250)配列番号179、155、188、127、1、161、198、154、110、58、153、26、175、192、109、97、145、178、10、174、152、133、91、159、104、93、114の組み合わせ
(251)配列番号155、188、127、1、79、84、189、110、78、176、58、185、175、75、100、109、145、48、3、152、141、133、39、159、82、77、51、114の組み合わせ
(252)配列番号179、155、87、188、127、1、79、196、198、189、110、153、193、175、100、109、3、152、190、133、39、91、159、149、82、61、182、114の組み合わせ
(253)配列番号155、87、142、188、127、30、1、198、189、154、194、110、68、86、193、175、100、109、97、145、10、152、133、159、77、57、182、114の組み合わせ
(254)配列番号121、155、137、87、188、127、1、198、110、56、193、175、192、100、109、145、178、48、10、83、71、152、133、180、159、149、182、114の組み合わせ
(255)配列番号121、155、87、188、127、1、161、198、189、154、38、110、175、109、48、108、71、73、152、133、91、159、82、77、182、114、12の組み合わせ
(256)配列番号121、155、137、87、188、127、1、198、110、176、185、175、109、48、10、3、54、73、133、180、91、159、149、82、51、182、93、114の組み合わせ
(257)配列番号179、155、87、142、188、127、1、183、53、198、189、110、68、148、175、100、109、3、99、152、133、180、39、159、149、2、82、182、114の組み合わせ
(258)配列番号155、137、87、142、188、127、1、79、196、198、189、38、110、75、109、145、10、174、92、71、152、190、133、180、91、104、182、114の組み合わせ
(259)配列番号121、155、137、87、188、127、1、196、172、84、194、110、200、102、86、193、75、109、3、71、133、159、149、2、82、77、182、114の組み合わせ
(260)配列番号121、155、87、142、188、127、94、1、196、161、198、189、110、102、175、100、109、145、48、54、174、39、91、159、82、77、182、114の組み合わせ
(261)配列番号155、87、188、127、30、1、79、196、161、154、38、194、110、86、26、100、145、3、54、174、71、152、133、39、159、149、104、57、114の組み合わせ
(262)配列番号155、137、142、188、127、1、196、161、53、198、189、154、38、110、68、102、175、100、109、97、98、54、133、91、149、82、182、114の組み合わせ
(263)配列番号155、137、87、142、188、127、85、1、79、198、189、154、110、148、185、100、109、145、135、174、71、152、133、91、159、149、82、182、114の組み合わせ
(264)配列番号155、137、87、142、188、127、146、30、1、79、53、189、154、110、192、100、109、145、48、174、71、152、133、159、82、150、182、93、114の組み合わせ
(265)配列番号121、155、137、87、142、188、127、1、110、177、193、175、192、100、109、178、54、73、152、111、133、180、91、159、149、104、51、150、114の組み合わせ
(266)配列番号155、137、87、142、188、127、1、198、110、41、185、193、175、109、97、167、62、178、10、54、71、152、190、133、91、149、104、182、114の組み合わせ
(267)配列番号121、155、137、142、188、127、1、189、194、110、58、193、175、109、169、145、125、178、54、174、133、180、91、159、82、104、51、182、114の組み合わせ
(268)配列番号155、137、87、142、188、127、1、80、196、198、189、154、194、110、41、148、177、175、100、109、145、178、48、174、133、91、2、82、93、114の組み合わせ
(269)配列番号121、155、137、87、142、188、127、1、84、198、194、110、153、193、175、100、109、48、135、3、96、174、133、39、91、149、2、82、93、114の組み合わせ
(270)配列番号155、142、188、127、1、79、198、189、154、67、110、175、75、100、109、145、62、125、174、108、190、133、39、91、159、149、82、55、114の組み合わせ
(271)配列番号155、137、188、127、1、79、53、154、110、186、143、58、86、69、100、145、178、3、54、152、133、180、91、159、149、77、57、182、114の組み合わせ
(272)配列番号121、155、137、87、142、188、127、85、1、189、110、128、157、101、193、175、118、75、31、100、145、178、88、3、54、152、133、2、82、114の組み合わせ
(273)配列番号179、121、155、142、188、127、1、161、53、154、110、148、193、175、31、100、109、62、48、156、88、54、71、152、111、133、149、82、182、114の組み合わせ
(274)配列番号121、155、87、188、127、1、161、198、110、128、200、148、185、175、192、75、100、109、145、48、174、152、133、180、149、82、104、182、114の組み合わせ
(275)配列番号155、137、188、127、5、1、196、53、198、154、110、74、68、101、175、192、109、145、10、3、108、71、133、159、149、82、77、150、182、93、114の組み合わせ
(276)配列番号155、137、87、127、5、1、84、198、110、185、153、175、75、100、109、145、178、10、174、71、141、190、133、180、39、91、149、182、93、114の組み合わせ
(277)配列番号121、155、87、142、188、127、94、1、161、198、187、110、185、175、109、167、145、178、54、152、133、39、91、159、149、2、82、61、51、114の組み合わせ
(278)配列番号121、155、137、142、188、127、5、1、161、198、189、154、110、185、153、177、175、100、109、178、10、54、174、152、133、91、159、149、82、104、114の組み合わせ
(279)配列番号179、155、137、142、188、127、1、198、187、154、110、176、143、200、175、100、109、97、145、62、178、48、108、71、152、111、133、91、159、104、114の組み合わせ
(280)配列番号155、142、188、127、1、161、198、189、154、110、78、143、175、100、109、178、10、3、191、174、108、71、73、152、133、149、2、82、104、182、114の組み合わせ
(281)配列番号121、155、87、188、127、1、29、161、53、198、187、110、185、153、175、100、109、145、178、48、59、152、141、133、180、91、149、82、104、114の組み合わせ
(282)配列番号155、142、188、127、1、189、154、110、78、193、175、75、100、109、145、62、178、96、174、108、92、71、134、152、133、159、82、104、93、114の組み合わせ
(283)配列番号121、155、87、142、188、127、1、161、198、189、194、110、185、193、175、75、100、109、125、54、191、174、152、190、133、91、159、82、77、182、93、114の組み合わせ
(284)配列番号155、137、87、188、127、1、79、198、189、110、193、175、75、109、145、178、48、98、135、3、71、133、39、91、159、149、2、82、77、93、114の組み合わせ
(285)配列番号179、121、155、137、87、188、127、1、79、196、161、198、154、110、148、86、175、100、109、145、174、92、152、133、159、149、2、82、182、93、114の組み合わせ
(286)配列番号179、121、87、142、188、127、5、1、183、161、154、110、153、175、109、145、125、178、156、3、54、174、71、133、159、149、82、104、150、93、114の組み合わせ
(287)配列番号121、155、137、87、188、127、1、196、198、189、194、110、68、193、175、100、109、169、145、10、3、174、152、133、39、91、159、149、82、77、57、182、114の組み合わせ
(288)配列番号155、137、87、188、127、1、80、161、198、189、110、176、102、185、175、100、109、97、145、54、92、152、133、180、91、149、104、61、18、57、93、114の組み合わせ
(289)配列番号121、155、137、87、188、127、1、161、198、189、38、110、185、175、100、109、145、62、125、178、48、10、3、108、71、152、133、91、159、104、77、182、93、114の組み合わせ
(290)配列番号179、121、155、87、188、127、1、79、161、172、198、110、148、185、153、175、109、145、178、48、54、191、174、71、73、152、133、39、91、149、82、51、93、114の組み合わせ
(291)配列番号121、155、137、87、188、127、1、79、198、189、38、194、110、143、68、26、177、101、175、25、192、100、109、63、135、54、108、71、152、133、91、82、104、171、77、182、114の組み合わせ
(1)配列番号155、142、188、127、198、154、110、75、100、3、133、149、82、114の組み合わせ
(2)配列番号155、5、198、110、75、109、54、174、71、152、133、91、82、114の組み合わせ
(3)配列番号155、188、127、110、9、101、175、54、71、111、133、2、82、114の組み合わせ
(4)配列番号155、142、188、127、154、110、100、109、96、174、71、133、2、82、114の組み合わせ
(5)配列番号155、142、188、127、198、154、110、68、109、3、54、71、111、82、114の組み合わせ
(6)配列番号155、188、127、110、68、200、109、145、10、83、71、152、180、159、82、114の組み合わせ
(7)配列番号155、137、188、127、5、194、110、100、109、156、3、71、152、133、82、150、114の組み合わせ
(8)配列番号155、142、188、127、154、110、75、31、100、109、3、71、152、133、2、82、114の組み合わせ
(9)配列番号121、155、188、127、198、110、68、175、109、135、3、174、71、133、180、149、82、114の組み合わせ
(10)配列番号121、155、188、127、154、110、41、100、145、71、152、133、159、2、82、51、150、114の組み合わせ
(11)配列番号155、142、188、127、154、110、177、100、109、125、54、71、152、2、82、104、182、114の組み合わせ
(12)配列番号155、188、127、30、198、189、110、193、175、100、3、71、133、159、2、82、182、114の組み合わせ
(13)配列番号155、137、142、188、127、154、110、193、175、109、88、135、3、54、152、133、82、114の組み合わせ
(14)配列番号155、137、188、127、110、101、193、25、109、145、10、3、152、133、180、159、2、82、114の組み合わせ
(15)配列番号155、87、188、127、161、198、38、110、101、175、100、109、10、54、71、133、91、82、114の組み合わせ
(16)配列番号121、188、127、189、154、110、102、177、100、109、3、54、174、133、159、2、82、104、114の組み合わせ
(17)配列番号137、142、188、127、161、110、75、109、145、178、10、96、174、133、39、159、82、104、114の組み合わせ
(18)配列番号87、188、127、196、161、154、110、100、109、135、71、73、152、159、2、82、182、114の組み合わせ
(19)配列番号155、87、188、127、196、194、110、4、193、175、109、145、178、48、10、59、71、159、82、114の組み合わせ
(20)配列番号155、142、188、127、196、198、154、110、101、75、109、3、174、71、152、133、159、2、82、114の組み合わせ
(21)配列番号155、137、188、127、5、198、189、154、110、128、193、100、109、3、152、133、82、182、114の組み合わせ
(22)配列番号121、155、188、127、5、79、198、110、100、145、59、54、108、73、152、133、159、2、82、114の組み合わせ
(23)配列番号121、155、87、188、127、196、110、175、75、145、48、3、152、159、82、104、77、57、114の組み合わせ
(24)配列番号155、87、142、188、127、198、154、110、185、109、178、54、174、152、133、159、82、77、51、114の組み合わせ
(25)配列番号87、142、188、127、161、198、187、110、175、100、109、145、178、174、133、159、2、82、93、114の組み合わせ
(26)配列番号155、137、87、188、127、189、154、194、110、200、148、86、193、3、152、159、82、104、77、93、114の組み合わせ
(27)配列番号121、155、142、188、127、196、110、185、26、175、100、109、145、88、3、71、152、2、82、114の組み合わせ
(28)配列番号179、155、137、142、188、127、110、109、145、125、10、88、3、174、152、180、113、82、51、150、114の組み合わせ
(29)配列番号155、87、142、188、127、198、194、110、136、68、175、109、145、3、71、73、152、159、82、77、114の組み合わせ
(30)配列番号121、155、142、188、127、53、189、110、200、193、75、109、145、3、152、133、180、82、104、114の組み合わせ
(31)配列番号155、137、87、188、127、5、198、154、110、193、175、100、145、178、152、91、159、2、82、77、114の組み合わせ
(32)配列番号155、137、87、188、127、196、189、110、193、100、109、48、71、73、152、133、91、159、2、82、114の組み合わせ
(33)配列番号155、142、188、127、53、189、154、110、78、185、75、109、145、174、152、133、2、82、93、114の組み合わせ
(34)配列番号155、188、127、196、154、110、176、102、69、175、75、100、109、145、54、108、73、133、159、82、114の組み合わせ
(35)配列番号121、155、137、87、142、188、127、30、196、154、110、100、109、145、135、71、152、133、149、2、82、114の組み合わせ
(36)配列番号155、137、87、188、127、196、187、110、102、193、175、100、109、145、125、152、149、2、82、104、114の組み合わせ
(37)配列番号155、137、87、188、127、196、189、194、110、68、200、175、100、145、10、152、133、180、159、82、104、114の組み合わせ
(38)配列番号121、155、188、127、198、110、193、75、100、109、145、48、152、133、180、159、2、82、104、182、114の組み合わせ
(39)配列番号155、87、188、127、110、185、193、109、145、10、3、71、73、180、91、159、149、82、51、182、93、114の組み合わせ
(40)配列番号155、87、188、127、196、161、198、110、193、175、109、178、174、152、133、91、159、82、51、93、114の組み合わせ
(41)配列番号155、87、188、127、198、110、175、109、145、178、48、10、3、152、133、91、159、2、82、51、114の組み合わせ
(42)配列番号121、155、188、127、196、189、110、68、193、100、109、145、135、3、71、152、133、180、159、82、182、114の組み合わせ
(43)配列番号155、137、87、188、127、53、189、194、110、101、193、175、100、109、145、3、152、133、149、82、114の組み合わせ
(44)配列番号155、142、188、127、154、110、68、148、193、175、145、156、10、152、133、180、159、149、82、77、182、114の組み合わせ
(45)配列番号121、155、87、188、127、198、194、110、109、135、3、191、152、133、39、159、149、82、104、51、150、114の組み合わせ
(46)配列番号121、155、87、142、188、127、196、198、154、110、4、175、109、167、191、73、113、159、149、2、82、182、114の組み合わせ
(47)配列番号179、155、137、87、188、127、196、198、194、110、86、75、100、109、152、133、180、91、149、2、82、182、114の組み合わせ
(48)配列番号155、87、188、127、161、198、110、153、175、192、109、145、178、10、152、133、91、149、82、104、182、114の組み合わせ
(49)配列番号155、137、87、142、188、127、189、154、110、175、100、109、145、178、10、3、174、152、133、91、82、93、114の組み合わせ
(50)配列番号155、87、142、188、127、94、138、189、110、193、109、145、10、135、3、71、152、133、159、2、82、182、114の組み合わせ
(51)配列番号142、188、127、196、189、154、110、185、193、109、145、178、88、3、152、133、180、159、82、150、182、114の組み合わせ
(52)配列番号121、155、87、188、127、85、198、154、38、110、185、193、109、145、125、178、3、152、133、159、82、182、114の組み合わせ
(53)配列番号179、121、155、87、142、188、127、161、172、198、110、193、175、145、54、73、152、159、149、82、93、114の組み合わせ
(54)配列番号179、155、142、188、127、138、198、110、9、56、175、100、109、145、10、54、174、141、133、82、182、114の組み合わせ
(55)配列番号121、155、87、142、188、127、198、110、109、167、145、21、48、3、174、71、152、133、149、2、82、182、114の組み合わせ
(56)配列番号179、155、87、142、188、127、79、198、110、102、175、100、109、97、145、3、174、141、91、159、149、2、82、114の組み合わせ
(57)配列番号121、155、87、142、188、127、5、196、198、110、102、69、175、192、109、145、88、54、174、2、82、52、114の組み合わせ
(58)配列番号155、142、188、127、183、198、110、128、200、185、175、75、100、178、152、133、180、159、149、82、77、51、114の組み合わせ
(59)配列番号155、137、87、142、188、127、79、53、198、110、193、100、109、62、135、174、152、133、39、91、82、51、182、114の組み合わせ
(60)配列番号155、137、188、127、196、198、110、68、86、193、100、109、145、88、73、152、39、159、149、2、82、182、114の組み合わせ
(61)配列番号155、87、188、127、30、198、110、9、175、109、48、10、3、96、73、152、133、91、159、149、2、82、51、93、114の組み合わせ
(62)配列番号121、155、87、142、188、127、79、196、198、110、101、175、109、178、3、54、152、133、39、159、149、82、150、182、114の組み合わせ
(63)配列番号155、87、188、127、196、161、198、110、68、185、86、193、175、109、145、178、10、152、141、133、149、82、51、182、114の組み合わせ
(64)配列番号179、155、137、188、127、5、79、198、189、154、110、177、193、100、109、145、178、3、152、133、149、82、104、150、114の組み合わせ
(65)配列番号155、87、188、127、5、196、161、84、198、38、110、68、175、109、145、3、73、133、180、91、159、149、82、104、114の組み合わせ
(66)配列番号121、155、137、87、142、188、127、183、196、84、189、154、110、143、145、135、71、152、91、159、2、82、182、114の組み合わせ
(67)配列番号121、155、137、87、188、127、161、198、154、110、69、175、100、3、71、73、152、133、91、159、82、182、93、114の組み合わせ
(68)配列番号179、121、155、188、127、84、198、110、78、68、148、185、175、100、109、174、108、152、141、133、91、149、82、182、93、114の組み合わせ
(69)配列番号155、137、87、142、127、196、198、189、38、110、68、175、100、109、145、174、71、152、133、91、159、2、82、77、182、114の組み合わせ
(70)配列番号155、137、87、188、127、198、110、68、185、177、175、100、145、63、3、174、71、152、91、159、2、82、77、51、114の組み合わせ
(71)配列番号155、137、142、188、127、198、189、110、56、176、193、175、75、100、97、145、174、73、152、133、159、82、77、182、114の組み合わせ
(72)配列番号179、121、155、137、87、188、127、196、198、154、110、193、175、25、100、109、178、48、3、152、133、91、82、182、114の組み合わせ
(73)配列番号155、137、87、188、127、196、84、198、154、194、110、68、175、73、152、133、180、159、149、2、82、104、77、182、114の組み合わせ
(74)配列番号155、137、142、188、127、138、53、198、110、41、175、75、109、145、3、174、71、152、133、180、159、149、2、82、51、182、114の組み合わせ
(75)配列番号179、155、137、87、188、127、161、198、189、110、4、100、109、145、178、10、3、54、174、133、39、91、159、149、82、51、114の組み合わせ
(76)配列番号121、155、137、87、142、188、127、80、198、110、101、193、175、100、109、145、54、152、111、133、159、149、82、182、93、114の組み合わせ
(77)配列番号121、155、137、87、188、127、161、189、38、110、56、193、175、109、62、178、3、191、133、180、91、159、149、82、77、114の組み合わせ
(78)配列番号121、155、142、188、127、5、196、161、198、154、110、26、175、100、109、145、63、10、54、152、133、159、2、82、104、114の組み合わせ
(79)配列番号155、87、142、188、127、146、158、196、53、198、110、68、101、25、100、109、145、3、54、152、180、91、159、82、182、114、12の組み合わせ
(80)配列番号155、137、87、188、127、30、79、161、53、198、194、110、185、175、100、109、97、174、71、133、159、149、2、82、77、182、114の組み合わせ
(81)配列番号121、155、137、142、188、127、161、198、187、194、110、148、26、177、175、109、145、178、10、71、152、133、91、159、149、82、182、114の組み合わせ
(82)配列番号121、155、87、142、188、127、196、161、198、189、38、110、200、175、75、100、109、169、125、71、152、133、91、149、82、182、114の組み合わせ
(83)配列番号121、155、87、142、188、127、30、161、198、187、110、101、193、175、100、109、145、178、10、152、111、133、39、149、82、51、182、114の組み合わせ
(84)配列番号155、142、188、127、53、198、154、110、176、69、193、192、100、109、145、48、10、152、133、39、91、159、149、82、77、182、114の組み合わせ
(85)配列番号121、155、87、142、188、127、198、110、102、101、175、100、109、169、145、3、174、108、71、152、133、91、159、2、82、57、182、114の組み合わせ
(86)配列番号121、155、137、87、142、188、127、196、161、198、110、41、193、175、173、75、100、109、97、71、73、133、91、159、149、82、77、114の組み合わせ
(87)配列番号155、137、142、188、127、196、198、189、110、78、185、175、100、109、145、3、174、73、152、141、133、91、159、149、82、51、182、114の組み合わせ
(88)配列番号179、155、87、188、127、85、79、196、198、187、154、110、86、175、109、145、88、3、174、71、134、152、39、159、82、57、114の組み合わせ
(89)配列番号121、155、87、188、127、198、110、102、185、86、175、109、145、178、96、71、190、133、180、91、159、149、82、20、93、114、12の組み合わせ
(90)配列番号179、155、137、188、127、5、198、154、110、68、200、148、86、175、109、145、71、152、190、180、39、159、149、2、82、150、182、114の組み合わせ
(91)配列番号155、87、142、188、127、161、84、189、110、186、24、58、175、100、109、3、96、174、133、39、91、159、2、82、104、182、93、114の組み合わせ
(92)配列番号155、137、87、168、188、127、161、53、198、187、110、101、193、175、100、109、10、152、141、111、133、42、91、159、82、51、93、114の組み合わせ
(93)配列番号155、87、188、127、85、161、84、198、154、110、68、86、193、175、109、3、191、73、152、180、39、91、159、149、82、77、182、93、114の組み合わせ
(94)配列番号155、87、188、127、161、53、198、189、110、193、175、109、145、48、135、174、71、152、111、133、180、39、91、159、82、104、77、182、114の組み合わせ
(95)配列番号121、155、87、142、188、127、94、79、196、198、189、154、110、102、175、75、109、145、62、48、3、174、108、133、197、159、82、77、150、114の組み合わせ
(96)配列番号179、155、142、188、127、79、196、161、53、198、189、154、110、148、69、100、109、145、54、174、92、152、180、113、91、159、149、2、82、77、57、114の組み合わせ
(97)配列番号155、137、87、142、188、127、146、196、161、17、198、187、194、110、56、200、148、86、193、109、54、73、152、159、149、2、82、104、77、150、182、114の組み合わせ
(98)配列番号179、155、137、87、142、188、127、172、84、53、154、38、110、86、153、193、175、75、100、109、145、125、156、3、54、133、180、39、91、159、149、2、82、77、93、114の組み合わせ
(1)配列番号155、87、188、127、110、68、185、100、145、152、180、114の組み合わせ
(2)配列番号155、137、188、127、196、154、110、128、100、152、133、159、114の組み合わせ
(3)配列番号121、155、188、196、198、110、193、100、145、3、152、133、39、182、114の組み合わせ
(4)配列番号121、155、137、188、194、110、100、109、145、21、71、152、180、2、182、114の組み合わせ
(5)配列番号155、188、127、196、154、110、193、100、109、145、62、3、54、152、133、180、114の組み合わせ
(6)配列番号155、137、188、127、85、154、110、68、100、109、199、145、3、152、133、114の組み合わせ
(7)配列番号155、137、188、5、198、110、41、100、109、145、125、96、92、71、152、133、91、114の組み合わせ
(8)配列番号155、142、188、127、5、154、110、175、25、100、109、54、133、180、91、159、2、114の組み合わせ
(9)配列番号155、137、188、127、110、68、75、100、109、145、135、152、111、133、113、182、114の組み合わせ
(10)配列番号121、155、142、188、127、5、198、154、110、100、109、145、54、71、163、152、2、114の組み合わせ
(11)配列番号155、137、142、188、127、85、5、154、110、128、193、175、100、145、152、133、51、182、114の組み合わせ
(12)配列番号155、87、142、188、127、196、154、110、193、100、109、145、3、174、71、152、159、114の組み合わせ
(13)配列番号155、87、188、127、198、110、68、101、25、192、100、145、3、152、180、159、149、114の組み合わせ
(14)配列番号155、188、127、189、110、68、86、193、192、100、135、3、71、152、133、180、159、77、114の組み合わせ
(15)配列番号121、155、87、188、127、79、53、154、110、185、175、100、3、54、73、152、133、93、114の組み合わせ
(16)配列番号155、137、188、127、198、189、194、110、200、175、75、100、10、152、133、180、159、149、2、182、114の組み合わせ
(17)配列番号121、155、137、188、127、194、110、148、100、109、145、125、88、135、3、152、91、159、149、77、114の組み合わせ
(18)配列番号121、155、188、127、154、194、110、176、200、69、100、145、174、71、73、152、133、159、149、77、51、114の組み合わせ
(19)配列番号142、188、127、138、189、154、110、26、193、100、109、178、10、88、152、133、159、104、171、77、182、114の組み合わせ
(20)配列番号155、137、188、127、146、183、196、198、110、200、193、25、100、109、152、133、180、91、159、149、77、114の組み合わせ
(21)配列番号121、155、188、127、198、154、110、26、193、175、100、109、145、88、3、54、174、71、152、159、182、114の組み合わせ
(22)配列番号179、155、137、142、188、127、79、196、198、154、110、102、148、101、75、100、109、145、3、54、152、2、114の組み合わせ
(23)配列番号155、137、87、188、127、85、162、196、198、110、68、175、75、100、145、3、54、71、152、133、159、55、114、12の組み合わせ
(24)配列番号155、137、188、127、183、196、110、68、86、193、175、100、145、135、71、73、152、133、180、159、149、2、77、51、182の組み合わせ
(25)配列番号155、142、188、127、5、194、110、68、86、26、193、175、75、100、145、10、3、71、152、111、133、149、77、150、114の組み合わせ
(26)配列番号121、155、87、142、188、127、138、196、161、198、189、110、56、185、101、100、10、73、152、133、180、149、70、51、182、114の組み合わせ
(27)配列番号155、137、142、188、127、146、196、53、154、110、41、68、200、175、100、109、97、145、63、152、190、180、91、159、182、114の組み合わせ
(28)配列番号179、155、188、127、146、198、187、38、110、200、193、175、100、109、199、178、71、152、133、180、91、159、149、77、51、93、114の組み合わせ
(29)配列番号155、137、142、188、127、53、189、110、41、200、175、25、75、31、100、125、156、54、71、106、152、133、180、159、2、77、182、114の組み合わせ
(30)配列番号121、155、188、127、5、196、84、198、187、189、110、78、102、148、86、177、175、100、109、145、3、174、133、180、149、2、182、114の組み合わせ
(31)配列番号179、121、155、137、87、142、188、127、198、189、110、148、192、100、109、145、135、174、152、133、180、113、91、159、149、77、18、182、114の組み合わせ
(1)配列番号110、3、114の組み合わせ
(2)配列番号188、5、110、3、114の組み合わせ
(3)配列番号155、142、110、68、3、71、114の組み合わせ
(4)配列番号155、188、127、110、145、3、152、180、159、114の組み合わせ
(5)配列番号155、188、127、110、148、193、145、3、71、152、180、159、2、114の組み合わせ
(6)配列番号155、137、87、188、127、196、194、110、101、145、3、152、133、149、182、114の組み合わせ
(7)配列番号155、142、188、127、5、196、154、110、68、193、109、3、71、133、180、77、114の組み合わせ
(8)配列番号155、137、142、188、127、85、30、5、196、110、193、145、135、3、152、2、182、114の組み合わせ
(9)配列番号155、137、87、107、188、127、198、154、110、41、185、193、109、178、3、152、133、91、149、104、51、93、114の組み合わせ
(10)配列番号155、87、188、127、183、198、189、154、194、110、56、58、175、75、109、199、145、3、174、73、152、133、159、149、57、93、114の組み合わせ
(11)配列番号121、155、87、142、188、79、53、198、154、110、185、101、175、75、109、169、125、3、174、92、71、152、141、133、91、149、104、114の組み合わせ
(12)配列番号155、137、87、142、188、127、146、196、198、189、154、38、110、148、177、109、135、3、108、152、180、39、91、159、149、2、182、114の組み合わせ
(1)配列番号137、127、5、110、145、152、114の組み合わせ
(2)配列番号155、137、87、188、5、194、110、68、109、145、71、152、2、114の組み合わせ
(3)配列番号155、137、188、127、5、110、68、145、71、73、152、159、149、114の組み合わせ
(4)配列番号155、137、188、127、30、5、110、193、71、152、180、159、149、2、51、182、114の組み合わせ
(5)配列番号155、142、188、127、5、154、194、110、32、41、152、159、2、51、150、182、114の組み合わせ
(6)配列番号155、188、127、5、84、198、110、185、75、109、145、71、152、180、159、2、51、114の組み合わせ
(7)配列番号179、121、155、142、188、127、5、84、198、189、154、194、110、78、143、175、75、145、135、47、174、71、73、152、133、104、182、114の組み合わせ
(1)配列番号155、137、188、127、196、110、75、145、71、73、152、133、2、77、114の組み合わせ
(2)配列番号155、142、188、127、94、196、189、110、143、68、75、71、152、133、2、77、114の組み合わせ
(3)配列番号155、87、188、127、161、198、38、110、185、109、10、152、180、91、159、104、77、182、114の組み合わせ
(1)配列番号155、188、127、5、198、189、110、176、143、102、148、101、75、109、199、145、54、174、106、152、133、2、20、51、114の組み合わせ
(1)配列番号179、121、155、137、132、87、168、142、188、127、85、94、5、1、162、80、196、161、160、84、53、17、198、187、189、110、6、124、9、123、136、74、78、4、90、176、143、200、102、148、185、86、76、153、193、175、192、43、75、100、109、169、199、97、145、62、125、21、48、63、98、88、83、59、135、3、47、54、96、174、131、92、71、73、106、147、152、27、133、113、39、91、197、159、149、2、82、34、104、61、77、51、150、182、114の組み合わせ
本発明は、被験体の検体において、本発明における海馬萎縮マーカーであるポリヌクレオチドの発現量を測定し、該測定された発現量を用いて、被験体の海馬が萎縮しているか否かを評価することを含む、海馬萎縮の検出方法を提供する。
(a)被験体由来の検体を、in vitroで、例えば、本発明のキット又はデバイス中の海馬萎縮検出用の核酸と接触させるステップ、
(b)検体中の海馬萎縮マーカー(標的核酸)の発現量を、上記核酸を核酸プローブ又はプライマーとして用いて測定するステップ、
(c)(b)の結果から、当該被験体における海馬萎縮(神経細胞死)の有無(存在又は不存在)を評価するか、又はその評価の指標を得るステップ、
を含むことができる。
(a)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~109のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(c)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドであってよい。
(f)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号110~200のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(h)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドであってよい。
(a)被験体の検体から調製されたRNA(ここで、ステップ(b)の定量RT-PCRのために、例えばRNAの3’末端はポリアデニル化されていてもよく、又はいずれか若しくは両方の末端に任意の配列がライゲーション法などで付加されていてもよい)又はそれから逆転写により合成されたその相補配列からなるポリヌクレオチド(cDNA)を、海馬萎縮検出用の核酸、例えば、本発明のキット又はデバイスに含まれる海馬萎縮検出用の核酸と接触させ結合させるステップ、
(b)当該核酸に結合した検体由来のRNA又は当該RNAから合成されたcDNAを、当該核酸を核酸プローブとして用いるハイブリダイゼーションによって、あるいは、当該核酸をプライマーとして用いる核酸増幅、例えば定量RT-PCRによって測定(例えば、定量)するステップ、
(c)上記(b)の測定結果に基づいて、海馬萎縮(又は海馬萎縮/神経細胞死の、マーカー遺伝子/miRNA)の有無を評価するステップ、
を含むことができる。
(a)海馬が萎縮していることが既知の被験体の検体及び海馬が萎縮していないことが既知の被験体の検体における上記海馬萎縮マーカーの発現量を、本発明の海馬萎縮検出用核酸(例えば、プローブ又はプライマー)、キット又はデバイスを用いて測定するステップ、
(b)(a)で測定された発現量の測定値から、上記の式1~3、5及び6の判別式を作成するステップ、
(c)海馬萎縮について試験すべき(未知の)被験体の検体における上記海馬萎縮マーカーの発現量を、本発明の海馬萎縮検出用核酸(例えば、プローブ又はプライマー)、キット又はデバイスを用いて測定し、(b)で作成した判別式にそれを代入して、得られた結果に基づいて被験体の海馬が萎縮していること又は海馬が萎縮していないことを決定又は評価する、或いは海馬萎縮が疑われる未知の被験者由来のマーカー発現量を海馬が萎縮していない被験者由来の対照と比較し評価する、ステップ、
を含むことができる。ここで、式1~3、5及び6の式中のxは説明変数であり、上記海馬萎縮マーカーの発現量の測定値を含む。具体的には本発明の海馬が萎縮している被験体と海馬が萎縮していない被験体を判別するための説明変数は、上記海馬萎縮マーカーの発現量の測定値であってよく、例えば下記の発現量であってよい:
配列番号1~109、及び110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列又はそれに相補的な塩基配列からなるポリヌクレオチド、又はその変異体、誘導体、もしくは15以上の連続した塩基を含むその断片(例えば、DNA)である海馬萎縮検出用核酸を用いて測定される海馬が萎縮している被験体及び海馬が萎縮していない被験体の検体(例えば、血清)における発現量。
<検体の採取>
国立長寿医療研究センターにてインフォームドコンセントを得たヒト被験者合計1,126人(表2)からBD バキュティナ採血管219AFBZX00109000(BD株式会社(日本))を用いてそれぞれ血清を採取した。
検体として上記の合計1,126人それぞれから得られた血清300μLから、3D-Gene(登録商標)RNA extraction reagent from liquid sample kit(東レ株式会社(日本))中のRNA抽出用試薬を用いて、同社の定めるプロトコールに従ってtotal RNAを得た。
検体として上記の合計1,126人の血清のそれぞれから得たtotal RNA中のmiRNAを、3D-Gene(登録商標) miRNA Labeling kit(東レ株式会社)を用いて同社が定めるプロトコールに基づいて蛍光標識した。オリゴDNAチップとして、miRBase release 21に登録されているmiRNAのうち、2,565種のmiRNAのそれぞれと相補的な配列を有するプローブを搭載した3D-Gene(登録商標) Human miRNA Oligo chip(東レ株式会社)を用い、同社が定めるプロトコールに基づいてストリンジェントな条件下でハイブリダイゼーション及びハイブリダイゼーション後の洗浄を行った。オリゴDNAチップを3D-Gene(登録商標)スキャナー(東レ株式会社)を用いてスキャンし、画像を取得して3D-Gene(登録商標)Extraction(東レ株式会社)にて蛍光強度を数値化した。数値化された蛍光強度を、底が2の対数値に変換して発現量とし、ブランク値の減算を行い、欠損値はシグナル値0.1で置換した。その結果、上記の1,126人の血清に対する、網羅的なmiRNAの発現量を得た。
上記の264人の被験者について、脳のMRI画像を取得し、この画像を脳画像解析プログラムIcobrain(Icometrix社)を用いて数値化し、脳の各部位の正規化された容積(体積)の定量値を得た。この定量値に基づいて全脳に対する海馬の容積割合を算出した。
上記の264人の被験者について、末梢血細胞からDNAを抽出し、次世代シークエンサーを用いて全配列解析を実施した。このうち、APOE遺伝子の遺伝子多型を解析し、特にApoE4アレル数を測定した。
<単一のmiRNAによる海馬萎縮群と非萎縮群の比較>
本実施例では、海馬萎縮度(全脳に対する海馬の容積割合を指標とする)の異なる2群を比較し発現量に有意差が見られる単一のmiRNAを海馬萎縮マーカーとして選定した。下記の実施例2-(1)~2-(4)では、海馬萎縮度の閾値を4通りに分けて試行し、各々の閾値において海馬萎縮群と海馬非萎縮群を設定してマーカーの選定を行った。
全脳に対する海馬の容積割合が0.60%以下の海馬萎縮群1(132検体)と全脳に対する海馬の容積割合が0.63%以上の海馬非萎縮群1(132検体)の2群間の発現量に統計的有意差があるmiRNA/遺伝子を評価するため、等分散を仮定した両側t検定を行いP値を算出した。マーカーとして、P値が0.1未満となるか、又は、海馬萎縮群と海馬非萎縮群の対数変換した遺伝子発現量の差分(Fold Change)の絶対値が、測定誤差として推定される0.2よりも大きい、miRNA/遺伝子を選択し、結果を表3に記載した。
海馬萎縮度がより極端に異なる2群を比較することで実施例2-(1)を深化させた解析を行った。閾値をより低く設定した、全脳に対する海馬の容積割合が0.59%以下である海馬萎縮群2(105検体)と、閾値をより高く設定した、全脳に対する海馬の容積割合が0.64%以上である海馬非萎縮群2(105検体)の2群間で比較を行った。
海馬萎縮度がより極端に異なる2群を比較することで実施例2-(2)を深化させた解析を行った。閾値をより低く設定した、全脳に対する海馬の容積割合が0.57%以下である海馬萎縮群3(88検体)と、閾値をより高く設定した、全脳に対する海馬の容積割合が0.68%以上である海馬非萎縮群3(88検体)の2群間で比較を行った。
海馬萎縮度がより極端に異なる2群を比較することで実施例2-(3)を深化させた解析を行った。閾値を極端に低く設定した、全脳に対する海馬の容積割合が0.38%以下である海馬萎縮群4(5検体)と、閾値を極端に高く設定した、全脳に対する海馬の容積割合が0.86%以上である海馬非萎縮群4(5検体)の2群間で比較を行った。
<少数(2~5種)のmiRNAマーカーの組み合わせによる海馬萎縮患者の判別(ロジスティック回帰分析による判別)>
本実施例では、2~5種のmiRNAマーカーからなる組み合わせについて判別式を作成した上で、独立検証検体群(表7)において判別性能を評価し、性能が高い判別式に用いられるmiRNA/遺伝子を抽出することで、海馬萎縮を検出可能なmiRNAマーカーの組み合わせを取得した。
<少数(5、6種)のmiRNAマーカーの組み合わせによる海馬萎縮患者の判別(フィッシャーの判別分析による判別)>
実施例3ではLASSO法により2~5種のmiRNAマーカーからなる組み合わせについて判別式を作成したが、2群の判別性能をより向上させるため、本実施例ではフィッシャーの判別分析を用いて判別を行った。
上記学習・交差検証検体群を用いて、LASSO法によりmiRNA発現量の相関性が高い類似マーカーを除外した上でP値上位20種のmiRNAマーカーを選択した。この20種のうち5種のmiRNAの組み合わせを総当たりで用いてフィッシャー判別手法により、海馬が萎縮しているか否かを判別する判別式を構築した。学習・交差検証検体群で構築した判別式の判別性能を評価するため精度、感度、及び特異度を算出し、さらに独立検証検体群を用いてその判別式の判別性能を検証した。
さらにマーカー数を6種に増やした場合に判別式の判別性能が向上するか検証した。上記学習・交差検証検体群を用いて、LASSO法によりmiRNA発現量の相関性が高い類似マーカーを除外した上でP値上位40種のmiRNAマーカーを選択した。その他は実施例4-(1)と同様に行った。
<5種以上のmiRNAマーカーと海馬萎縮関連因子の組み合わせを用いた海馬萎縮患者の判別(ロジスティック回帰分析による判別)>
本実施例では、判別に用いるマーカー数を限定しない場合のバリエーションを検討することを目的としてLASSO法を用いたロジスティック回帰分析を行った。また海馬萎縮関連因子として年齢、性別、及びApoE4アレル数を説明変数として判別式に加えることで、判別性能の向上を検討した。
<複数のmiRNAマーカーと海馬萎縮関連因子の組み合わせを用いた、海馬容積がより極端に異なる2群の判別(ロジスティック回帰の判別)>
実施例6では、以下の二つを目的として解析を行った:
目的1:海馬萎縮度が実施例5よりも極端に異なる2群を比較することで実施例5を深化させた解析を行うこと、
目的2:判別式構築において学習、検証に用いる検体数を減らした場合にも、AUCが0.9を超えるマーカーセットが得られるか検討を行うこと。
<回帰式構築による海馬萎縮度の判別及び海馬容積の予測>
本実施例では、海馬の容積を予測可能なより定量性の高いマーカーを選択した。具体的には、海馬萎縮群と海馬非萎縮群の2群比較ではなく、miRNA発現量、年齢、性別、及びApoE4アレル数を説明変数として、目的変数である海馬の容積値を予測するための線形回帰分析を行った。
上記学習・交差検証検体群を用いてLASSO回帰により平均二乗誤差(MSE)が最小になるようなλの最適値を求め、学習・交差検証検体群を10分割したうちの1分割で交差検証を行い、最適λをとった式を独立検証群で評価し、回帰式の性能(相関係数R2、RMSE、MAE)を算出した。
本実施例では、上記学習・交差検証検体群を用いて部分的最小二乗法(PLS)を用いた回帰分析による座標変換を行った。説明変数の値を平均と標準偏差を用いてZスコア(Z-score)化し、miRNAや年齢因子が互いに無相関になるよう線形変換した変数を使って回帰した。説明変数を変数重要度射影(VIP)の閾値(=0.6)により選択し、学習・交差検証検体群のうちの1検体で交差検証を行い、最適な変数を用いたモデルを独立検証検体群で評価し、回帰性能(相関係数R2、RMSE、MAE)を算出した。
Claims (18)
- 海馬萎縮マーカーである、miR-3131、miR-6757-5p、miR-4706、miR-5001-5p、miR-3180-3p、miR-642b-3p、miR-4655-5p、miR-6819-5p、miR-937-5p、miR-4688、miR-6741-5p、miR-7107-5p、miR-4271、miR-1229-5p、miR-4707-5p、miR-6808-5p、miR-4656、miR-6076、miR-6762-5p、miR-7109-5p、miR-6732-5p、miR-3195、miR-7150、miR-642a-3p、miR-1249-5p、miR-3185、miR-4689、miR-3141、miR-6840-3p、miR-3135b、miR-1914-3p、miR-4446-3p、miR-4433b-3p、miR-6877-5p、miR-6848-5p、miR-3620-5p、miR-6825-5p、miR-5739、miR-3663-3p、miR-4695-5p、miR-3162-5p、miR-3679-5p、miR-8059、miR-7110-5p、miR-1275、miR-6779-5p、miR-197-5p、miR-6845-5p、miR-4327、miR-4723-5p、miR-4530、miR-6771-5p、miR-614、miR-92a-2-5p、miR-6891-5p、miR-6124、miR-4687-3p、miR-4442、miR-7977、miR-6785-5p、miR-4497、miR-8071、miR-663b、miR-3180、miR-4251、miR-1285-3p、miR-6870-5p、miR-4484、miR-4476、miR-6749-5p、miR-4454、miR-6893-5p、miR-6085、miR-4787-5p、miR-149-3p、miR-7704、miR-6125、miR-6090、miR-3197、miR-6850-5p、miR-4467、miR-6885-5p、miR-6803-5p、miR-6798-5p、miR-6780b-5p、miR-6768-5p、miR-5100、miR-6724-5p、miR-6879-5p、miR-7108-5p、miR-4649-5p、miR-4739、miR-6089、miR-1908-5p、miR-4516、miR-2861、miR-4492、miR-4294、miR-6791-5p、miR-1469、miR-6752-5p、miR-4730、miR-6126、miR-6869-5p、miR-1268a、miR-6799-5p、miR-8069、miR-3621及びmiR-4763-3pからなる群から選択される少なくとも1つのポリヌクレオチド、又は該ポリヌクレオチドの相補鎖と、特異的に結合可能な核酸を含む、海馬萎縮の検出用キット。
- 前記核酸が、下記の(a)~(e)のポリヌクレオチド:
(a)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~109のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(c)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項1に記載のキット。 - 前記キットが、別の海馬萎縮マーカーである、miR-1228-5p、miR-760、miR-187-5p、miR-7111-5p、miR-6088、miR-6805-3p、miR-4640-5p、miR-6721-5p、miR-6880-5p、miR-711、miR-128-1-5p、miR-4525、miR-486-3p、miR-6756-5p、miR-1260b、miR-3184-5p、miR-6075、miR-204-3p、miR-4728-5p、miR-4534、miR-4758-5p、miR-8063、miR-6836-3p、miR-6789-5p、miR-744-5p、miR-1909-3p、miR-887-3p、miR-4745-5p、miR-4433a-3p、miR-5090、miR-296-5p、miR-939-5p、miR-3648、miR-3196、miR-6722-3p、miR-6805-5p、miR-1202、miR-6775-5p、miR-6087、miR-6765-5p、miR-6875-5p、miR-4674、miR-1233-5p、miR-7114-5p、miR-5787、miR-8072、miR-3619-3p、miR-4632-5p、miR-6800-5p、miR-4634、miR-4486、miR-6727-5p、miR-4505、miR-4725-3p、miR-1538、miR-320b、miR-1915-5p、miR-328-5p、miR-6820-5p、miR-6726-5p、miR-3665、miR-638、miR-762、miR-4466、miR-3940-5p、miR-1237-5p、miR-575、miR-3656、miR-4488、miR-4281、miR-6781-5p、miR-4532、miR-4665-5p、miR-6816-5p、miR-4508、miR-6784-5p、miR-6786-5p、miR-4741、miR-1343-5p、miR-1227-5p、miR-4734、miR-3960、miR-128-2-5p、miR-6743-5p、miR-663a、miR-6729-5p、miR-1915-3p、miR-1268b、miR-4651、miR-3178及びmiR-4463からなる群から選択される少なくとも1つのポリヌクレオチド、又は該ポリヌクレオチドの相補鎖と、特異的に結合可能な核酸をさらに含む、請求項1又は2に記載のキット。
- 前記核酸が、下記の(f)~(j)のポリヌクレオチド:
(f)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号110~200のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(h)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項3に記載のキット。 - 海馬萎縮マーカーである、miR-3131、miR-6757-5p、miR-4706、miR-5001-5p、miR-3180-3p、miR-642b-3p、miR-4655-5p、miR-6819-5p、miR-937-5p、miR-4688、miR-6741-5p、miR-7107-5p、miR-4271、miR-1229-5p、miR-4707-5p、miR-6808-5p、miR-4656、miR-6076、miR-6762-5p、miR-7109-5p、miR-6732-5p、miR-3195、miR-7150、miR-642a-3p、miR-1249-5p、miR-3185、miR-4689、miR-3141、miR-6840-3p、miR-3135b、miR-1914-3p、miR-4446-3p、miR-4433b-3p、miR-6877-5p、miR-6848-5p、miR-3620-5p、miR-6825-5p、miR-5739、miR-3663-3p、miR-4695-5p、miR-3162-5p、miR-3679-5p、miR-8059、miR-7110-5p、miR-1275、miR-6779-5p、miR-197-5p、miR-6845-5p、miR-4327、miR-4723-5p、miR-4530、miR-6771-5p、miR-614、miR-92a-2-5p、miR-6891-5p、miR-6124、miR-4687-3p、miR-4442、miR-7977、miR-6785-5p、miR-4497、miR-8071、miR-663b、miR-3180、miR-4251、miR-1285-3p、miR-6870-5p、miR-4484、miR-4476、miR-6749-5p、miR-4454、miR-6893-5p、miR-6085、miR-4787-5p、miR-149-3p、miR-7704、miR-6125、miR-6090、miR-3197、miR-6850-5p、miR-4467、miR-6885-5p、miR-6803-5p、miR-6798-5p、miR-6780b-5p、miR-6768-5p、miR-5100、miR-6724-5p、miR-6879-5p、miR-7108-5p、miR-4649-5p、miR-4739、miR-6089、miR-1908-5p、miR-4516、miR-2861、miR-4492、miR-4294、miR-6791-5p、miR-1469、miR-6752-5p、miR-4730、miR-6126、miR-6869-5p、miR-1268a、miR-6799-5p、miR-8069、miR-3621及びmiR-4763-3pからなる群から選択される少なくとも1つのポリヌクレオチド、又は該ポリヌクレオチドの相補鎖と、特異的に結合可能な核酸を含む、海馬萎縮の検出用デバイス。
- 前記核酸が、下記の(a)~(e)のポリヌクレオチド:
(a)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~109のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(c)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項5に記載のデバイス。 - 前記デバイスが、別の海馬萎縮マーカーである、miR-1228-5p、miR-760、miR-187-5p、miR-7111-5p、miR-6088、miR-6805-3p、miR-4640-5p、miR-6721-5p、miR-6880-5p、miR-711、miR-128-1-5p、miR-4525、miR-486-3p、miR-6756-5p、miR-1260b、miR-3184-5p、miR-6075、miR-204-3p、miR-4728-5p、miR-4534、miR-4758-5p、miR-8063、miR-6836-3p、miR-6789-5p、miR-744-5p、miR-1909-3p、miR-887-3p、miR-4745-5p、miR-4433a-3p、miR-5090、miR-296-5p、miR-939-5p、miR-3648、miR-3196、miR-6722-3p、miR-6805-5p、miR-1202、miR-6775-5p、miR-6087、miR-6765-5p、miR-6875-5p、miR-4674、miR-1233-5p、miR-7114-5p、miR-5787、miR-8072、miR-3619-3p、miR-4632-5p、miR-6800-5p、miR-4634、miR-4486、miR-6727-5p、miR-4505、miR-4725-3p、miR-1538、miR-320b、miR-1915-5p、miR-328-5p、miR-6820-5p、miR-6726-5p、miR-3665、miR-638、miR-762、miR-4466、miR-3940-5p、miR-1237-5p、miR-575、miR-3656、miR-4488、miR-4281、miR-6781-5p、miR-4532、miR-4665-5p、miR-6816-5p、miR-4508、miR-6784-5p、miR-6786-5p、miR-4741、miR-1343-5p、miR-1227-5p、miR-4734、miR-3960、miR-128-2-5p、miR-6743-5p、miR-663a、miR-6729-5p、miR-1915-3p、miR-1268b、miR-4651、miR-3178及びmiR-4463からなる群から選択される少なくとも1つのポリヌクレオチド、又は該ポリヌクレオチドの相補鎖と、特異的に結合可能な核酸をさらに含む、請求項5又は6に記載のデバイス。
- 前記核酸が、下記の(f)~(j)のポリヌクレオチド:
(f)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号110~200のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(h)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項7に記載のデバイス。 - 前記デバイスが、ハイブリダイゼーション技術による測定のためのデバイスである、請求項5~8のいずれか1項に記載のデバイス。
- 前記ハイブリダイゼーション技術が、核酸アレイ技術である、請求項9に記載のデバイス。
- 被験体の検体において、海馬萎縮マーカーである、miR-3131、miR-6757-5p、miR-4706、miR-5001-5p、miR-3180-3p、miR-642b-3p、miR-4655-5p、miR-6819-5p、miR-937-5p、miR-4688、miR-6741-5p、miR-7107-5p、miR-4271、miR-1229-5p、miR-4707-5p、miR-6808-5p、miR-4656、miR-6076、miR-6762-5p、miR-7109-5p、miR-6732-5p、miR-3195、miR-7150、miR-642a-3p、miR-1249-5p、miR-3185、miR-4689、miR-3141、miR-6840-3p、miR-3135b、miR-1914-3p、miR-4446-3p、miR-4433b-3p、miR-6877-5p、miR-6848-5p、miR-3620-5p、miR-6825-5p、miR-5739、miR-3663-3p、miR-4695-5p、miR-3162-5p、miR-3679-5p、miR-8059、miR-7110-5p、miR-1275、miR-6779-5p、miR-197-5p、miR-6845-5p、miR-4327、miR-4723-5p、miR-4530、miR-6771-5p、miR-614、miR-92a-2-5p、miR-6891-5p、miR-6124、miR-4687-3p、miR-4442、miR-7977、miR-6785-5p、miR-4497、miR-8071、miR-663b、miR-3180、miR-4251、miR-1285-3p、miR-6870-5p、miR-4484、miR-4476、miR-6749-5p、miR-4454、miR-6893-5p、miR-6085、miR-4787-5p、miR-149-3p、miR-7704、miR-6125、miR-6090、miR-3197、miR-6850-5p、miR-4467、miR-6885-5p、miR-6803-5p、miR-6798-5p、miR-6780b-5p、miR-6768-5p、miR-5100、miR-6724-5p、miR-6879-5p、miR-7108-5p、miR-4649-5p、miR-4739、miR-6089、miR-1908-5p、miR-4516、miR-2861、miR-4492、miR-4294、miR-6791-5p、miR-1469、miR-6752-5p、miR-4730、miR-6126、miR-6869-5p、miR-1268a、miR-6799-5p、miR-8069、miR-3621及びmiR-4763-3pからなる群から選択される少なくとも1つのポリヌクレオチドの発現量を測定し、該測定された発現量を用いて、被験体の海馬が萎縮しているか否かを評価することを含む、海馬萎縮の検出方法。
- 請求項1~4のいずれか1項に記載のキット又は請求項5~10のいずれか1項に記載のデバイスを用いて、前記ポリヌクレオチドの発現量を測定する、請求項11に記載の方法。
- 測定された前記ポリヌクレオチドの前記発現量を、海馬が萎縮していることが既知である被験体由来の検体における前記ポリヌクレオチドの発現量と海馬が萎縮していないことが既知である被験体由来の検体における前記ポリヌクレオチドの発現量を教師サンプルとして作成された海馬の萎縮又は非萎縮を区別的に判別することが可能な判別式に代入し、それによって、海馬の萎縮又は非萎縮を評価することを含む、請求項12に記載の方法。
- 前記核酸が、下記の(a)~(e)のポリヌクレオチド:
(a)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(b)配列番号1~109のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(c)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(d)配列番号1~109のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(e)前記(a)~(d)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項12又は13に記載の方法。 - 前記キット又はデバイスが、別の海馬萎縮マーカーである、miR-1228-5p、miR-760、miR-187-5p、miR-7111-5p、miR-6088、miR-6805-3p、miR-4640-5p、miR-6721-5p、miR-6880-5p、miR-711、miR-128-1-5p、miR-4525、miR-486-3p、miR-6756-5p、miR-1260b、miR-3184-5p、miR-6075、miR-204-3p、miR-4728-5p、miR-4534、miR-4758-5p、miR-8063、miR-6836-3p、miR-6789-5p、miR-744-5p、miR-1909-3p、miR-887-3p、miR-4745-5p、miR-4433a-3p、miR-5090、miR-296-5p、miR-939-5p、miR-3648、miR-3196、miR-6722-3p、miR-6805-5p、miR-1202、miR-6775-5p、miR-6087、miR-6765-5p、miR-6875-5p、miR-4674、miR-1233-5p、miR-7114-5p、miR-5787、miR-8072、miR-3619-3p、miR-4632-5p、miR-6800-5p、miR-4634、miR-4486、miR-6727-5p、miR-4505、miR-4725-3p、miR-1538、miR-320b、miR-1915-5p、miR-328-5p、miR-6820-5p、miR-6726-5p、miR-3665、miR-638、miR-762、miR-4466、miR-3940-5p、miR-1237-5p、miR-575、miR-3656、miR-4488、miR-4281、miR-6781-5p、miR-4532、miR-4665-5p、miR-6816-5p、miR-4508、miR-6784-5p、miR-6786-5p、miR-4741、miR-1343-5p、miR-1227-5p、miR-4734、miR-3960、miR-128-2-5p、miR-6743-5p、miR-663a、miR-6729-5p、miR-1915-3p、miR-1268b、miR-4651、miR-3178及びmiR-4463からなる群から選択される少なくとも1つのポリヌクレオチド、又は該ポリヌクレオチドの相補鎖と、特異的に結合可能な核酸をさらに含む、請求項12~14のいずれか1項に記載の方法。
- 前記核酸が、下記の(f)~(j)のポリヌクレオチド:
(f)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(g)配列番号110~200のいずれかで表される塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(h)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列からなるポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、
(i)配列番号110~200のいずれかで表される塩基配列若しくは当該塩基配列においてuがtである塩基配列に相補的な塩基配列を含むポリヌクレオチド、その変異体、その誘導体、又は15以上の連続した塩基を含むその断片、及び
(j)前記(f)~(i)のいずれかのポリヌクレオチドとストリンジェントな条件下でハイブリダイズするポリヌクレオチド、
からなる群から選択されるポリヌクレオチドである、請求項15に記載の方法。 - 前記被験体が、ヒトである、請求項11~16のいずれか1項に記載の方法。
- 前記検体が、血液、血清又は血漿である、請求項11~17のいずれか1項に記載の方法。
Priority Applications (6)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US17/915,682 US20230212675A1 (en) | 2020-03-31 | 2021-03-31 | Kit or device and method for detecting hippocampal atrophy |
| CA3178889A CA3178889A1 (en) | 2020-03-31 | 2021-03-31 | Kit or device and method for detecting hippocampal atrophy |
| EP21780961.5A EP4130259A1 (en) | 2020-03-31 | 2021-03-31 | Kit or device and method for detecting hippocampal atrophy |
| CN202180026096.7A CN115485378A (zh) | 2020-03-31 | 2021-03-31 | 用于检测海马萎缩的试剂盒或器件以及方法 |
| KR1020227034809A KR20220160602A (ko) | 2020-03-31 | 2021-03-31 | 해마 위축을 검출하기 위한 키트 또는 디바이스 및 방법 |
| JP2022512615A JPWO2021201092A1 (ja) | 2020-03-31 | 2021-03-31 |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2020-064383 | 2020-03-31 | ||
| JP2020064383 | 2020-03-31 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2021201092A1 true WO2021201092A1 (ja) | 2021-10-07 |
Family
ID=77927525
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2021/013807 WO2021201092A1 (ja) | 2020-03-31 | 2021-03-31 | 海馬萎縮を検出するためのキット又はデバイス及び方法 |
Country Status (7)
| Country | Link |
|---|---|
| US (1) | US20230212675A1 (ja) |
| EP (1) | EP4130259A1 (ja) |
| JP (1) | JPWO2021201092A1 (ja) |
| KR (1) | KR20220160602A (ja) |
| CN (1) | CN115485378A (ja) |
| CA (1) | CA3178889A1 (ja) |
| WO (1) | WO2021201092A1 (ja) |
Citations (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20140272993A1 (en) | 2013-03-15 | 2014-09-18 | The Translational Genomics Research Institute | Method of sequencing a full microrna profile from cerebrospinal fluid |
| JP2017184642A (ja) | 2016-04-01 | 2017-10-12 | 株式会社ヘルシーパス | 認知症マーカー、それを用いた認知症の評価方法、評価試薬および評価キット |
| WO2018199275A1 (ja) * | 2017-04-28 | 2018-11-01 | 東レ株式会社 | 卵巣腫瘍の検出のためのキット、デバイス及び方法 |
| WO2019004436A1 (ja) * | 2017-06-29 | 2019-01-03 | 東レ株式会社 | 肺がんの検出のためのキット、デバイス及び方法 |
| WO2019014457A1 (en) | 2017-07-12 | 2019-01-17 | Texas Tech University System | MICROARN-455-3P AS A PERIPHERAL BIOMARKER OF ALZHEIMER'S DISEASE |
| WO2019048500A1 (en) | 2017-09-05 | 2019-03-14 | Amoneta Diagnostics | NON-CODING RNA (RNANC) FOR THE DIAGNOSIS OF COGNITIVE DISORDERS |
| US20190136320A1 (en) | 2016-04-25 | 2019-05-09 | Instytut Biologii Doswiadczalnej Im. Marcelego Nenckiego Polska Akademia Nauk | Microrna biomarkers in blood for diagnosis of alzheimer's disease |
| US20190249250A1 (en) | 2015-11-20 | 2019-08-15 | Braindtech S.R.L. | Microglia Microvesicles Contained MicroRNA-Based Methods For The Diagnosis, Prognosis And Treatment Monitoring Of Neurological, Neurodegenerative And Inflammation-Based Diseases |
| WO2019159884A1 (ja) | 2018-02-13 | 2019-08-22 | 東レ株式会社 | 認知症の検出のためのキット又はデバイス及び方法 |
| JP2020064383A (ja) | 2018-10-16 | 2020-04-23 | 三菱重工業株式会社 | リスク特定装置、リスク特定方法、およびプログラム |
-
2021
- 2021-03-31 EP EP21780961.5A patent/EP4130259A1/en active Pending
- 2021-03-31 WO PCT/JP2021/013807 patent/WO2021201092A1/ja unknown
- 2021-03-31 KR KR1020227034809A patent/KR20220160602A/ko active Search and Examination
- 2021-03-31 JP JP2022512615A patent/JPWO2021201092A1/ja active Pending
- 2021-03-31 CA CA3178889A patent/CA3178889A1/en active Pending
- 2021-03-31 CN CN202180026096.7A patent/CN115485378A/zh active Pending
- 2021-03-31 US US17/915,682 patent/US20230212675A1/en active Pending
Patent Citations (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20140272993A1 (en) | 2013-03-15 | 2014-09-18 | The Translational Genomics Research Institute | Method of sequencing a full microrna profile from cerebrospinal fluid |
| US20190249250A1 (en) | 2015-11-20 | 2019-08-15 | Braindtech S.R.L. | Microglia Microvesicles Contained MicroRNA-Based Methods For The Diagnosis, Prognosis And Treatment Monitoring Of Neurological, Neurodegenerative And Inflammation-Based Diseases |
| JP2017184642A (ja) | 2016-04-01 | 2017-10-12 | 株式会社ヘルシーパス | 認知症マーカー、それを用いた認知症の評価方法、評価試薬および評価キット |
| US20190136320A1 (en) | 2016-04-25 | 2019-05-09 | Instytut Biologii Doswiadczalnej Im. Marcelego Nenckiego Polska Akademia Nauk | Microrna biomarkers in blood for diagnosis of alzheimer's disease |
| WO2018199275A1 (ja) * | 2017-04-28 | 2018-11-01 | 東レ株式会社 | 卵巣腫瘍の検出のためのキット、デバイス及び方法 |
| WO2019004436A1 (ja) * | 2017-06-29 | 2019-01-03 | 東レ株式会社 | 肺がんの検出のためのキット、デバイス及び方法 |
| WO2019014457A1 (en) | 2017-07-12 | 2019-01-17 | Texas Tech University System | MICROARN-455-3P AS A PERIPHERAL BIOMARKER OF ALZHEIMER'S DISEASE |
| WO2019048500A1 (en) | 2017-09-05 | 2019-03-14 | Amoneta Diagnostics | NON-CODING RNA (RNANC) FOR THE DIAGNOSIS OF COGNITIVE DISORDERS |
| WO2019159884A1 (ja) | 2018-02-13 | 2019-08-22 | 東レ株式会社 | 認知症の検出のためのキット又はデバイス及び方法 |
| JP2020064383A (ja) | 2018-10-16 | 2020-04-23 | 三菱重工業株式会社 | リスク特定装置、リスク特定方法、およびプログラム |
Non-Patent Citations (71)
| Title |
|---|
| ALTSCHUL, S.F. ET AL., JOURNAL OF MOLECULAR BIOLOGY, vol. 215, 1990, pages 403 - 410 |
| ARTZI S ET AL., BMC BIOINFORMATICS, vol. 9, 2008, pages 39 |
| AUSUBEL ET AL.: "Current Protocols in Molecular Biology", 1993, JOHN WILLEY & SONS, US |
| AZUMA-MUKAI, A ET AL., PROC. NATL. ACAD. SCI. U.S.A., vol. 105, 2008, pages 7964 - 7969 |
| BAR M ET AL., STEM CELLS, vol. 26, 2008, pages 2496 - 2505 |
| BEREZIKOV E ET AL., GENOME RES., vol. 16, 2006, pages 1289 - 1298 |
| BEREZIKOV E ET AL., MOL. CELL, vol. 28, 2007, pages 328 - 336 |
| BHAGAT R. ET AL., CELL DEATH DIFFER., no. 10, November 2018 (2018-11-01), pages 1837 - 1854 |
| BOLSTAD, B.M. ET AL., BIOINFORMATICS, vol. 19, 2003, pages 185 - 193 |
| C. CORTES ET AL., MACHINE LEARNING, vol. 20, 1995, pages 273 - 297 |
| CREIGHTON CJ ET AL., PLOS ONE, vol. 5, 2010, pages e10563 |
| CUMMINS JM ET AL., PROC. NATL. ACAD. SCI., U.S.A., vol. 103, 2006, pages 3687 - 3692 |
| DANIEL H.S. ET AL., THE JOURNAL OF NUCLEAR MEDICINE, vol. 45, no. 4, April 2004 (2004-04-01) |
| DING N ET AL., J. RADIAT. RES., vol. 52, 2011, pages 425 - 432 |
| EMRAH DUZEL ET AL., ALZHEMER'S & DEMENTIA MONITORING, vol. 10, 2018, pages 782 - 790 |
| FU H ET AL., FEBS LETT., vol. 579, 2005, pages 3849 - 3854 |
| FUREY TS. ET AL., BIOINFORMATICS, vol. 16, 2000, pages 906 - 14 |
| GOFF LA ET AL., PLOS ONE, vol. 4, 2009, pages e7192 |
| GRASSO M. ET AL., NEUROBIOL. AGING., vol. 84, December 2019 (2019-12-01), pages 240 |
| HANSEN TB ET AL., RNA BIOL., vol. 8, 2011, pages 378 - 383 |
| HARUO HANYU, TETSUICHI ASANO, DAIJI KOGURE, HIROFUMI SAKURAI, TOSHIHIKO IWAMOTO, MASARU TAKASAKI: "Relationship between hippocampal lesions and cerebral cortex function in Alzheimer's disease", JOURNAL OF THE JAPAN GERIATRICS SOCIETY, vol. 37, no. 11, 25 November 2000 (2000-11-25), pages 921 - 927, XP055977417, ISSN: 0300-9173, DOI: 10.3143/geriatrics.37.921 * |
| HIDEKI ASO ET AL.: "Frontier of Statistical Science 6", 2004, IWANAMI SHOTEN, article "Statistics of pattern recognition and learning - New concepts and approaches" |
| HOERL, A.E., TECHNOMETRICS, vol. 12, 1970, pages 55 - 67 |
| HOTELLING, H., JOURNAL OF EDUCATIONAL PSYCHOLOGY, vol. 24, 1933, pages 417 - 441 |
| HOUBAVIY HB ET AL., DEV. CELL, vol. 5, 2003, pages 351 - 358 |
| HU R ET AL., J. BIOL. CHEM., vol. 286, 2011, pages 12328 - 12339 |
| INT. NEUROUROL. J., vol. 20, 2016, pages S76 - 83 |
| JIMA DD ET AL., BLOOD, vol. 116, 2010, pages el 18 - el27 |
| KAWAJI H ET AL., BMC GENOMICS, vol. 9, 2008, pages 157 |
| KIM J ET AL., PROC. NATL. ACAD. SCI. U.S.A., vol. 101, 2004, pages 360 - 365 |
| KUMAR S. ET AL., HUM. MOL. GENET., vol. 26, no. 19, 1 October 2017 (2017-10-01), pages 3808 - 3822 |
| LADEWIG E ET AL., GENOME RES., vol. 22, 2012, pages 1634 - 1645 |
| LAGOS-QUINTANA M ET AL., CURR. BIOL., vol. 12, 2002, pages 735 - 739 |
| LAGOS-QUINTANA M ET AL., RNA, vol. 9, 2003, pages 175 - 179 |
| LI H ET AL., J. CLIN. INVEST., vol. 119, 2009, pages 3666 - 3677 |
| LI Y ET AL., GENE, vol. 497, 2012, pages 330 - 335 |
| LIM LP ET AL., SCIENCE, vol. 299, 2003, pages 1540 |
| LUI WO ET AL., CANCER RES., vol. 67, 2007, pages 6031 - 6043 |
| MARTON S ET AL., LEUKEMIA, vol. 22, 2008, pages 1274 - 1278 |
| MEIRI E ET AL., NUCLEIC ACIDS RES., vol. 38, 2010, pages 6234 - 6246 |
| MORIN RD. ET AL., GENOME RES., vol. 18, 2008, pages 610 - 621 |
| MOURELATOS Z ET AL., GENES DEV., vol. 16, 2002, pages 720 - 728 |
| NAKAMURA A. ET AL., NATURE, vol. 554, 31 January 2018 (2018-01-31), pages 249 - 254 |
| NELLO CRISTIANINI ET AL.: "Introduction to SVM", 2008, KYORITSU SHUPPAN CO., LTD. |
| NIELSEN, P.E. ET AL., SCIENCE, vol. 254, 1991, pages 1497 - 500 |
| OBIKA, S. ET AL., TETRAHEDRON LETT., vol. 39, pages 5401 - 5404 |
| OULAS A. ET AL., NUCLEIC ACIDS RES., vol. 37, 2009, pages 3276 - 3287 |
| PEARSON, K., PHILOSOPHICAL MAGAZINE, vol. 2, no. 11, 1901, pages 559 - 572 |
| PEARSON, W.R. ET AL., PROC. NATL. ACAD. SCI. U.S.A., vol. 85, 1988, pages 2444 - 2448 |
| PERSSON H ET AL., CANCER RES., vol. 71, 2011, pages 78 - 86 |
| PERSSON H ET AL., CANCER RES., vol. 71, pages 78 - 86 |
| PLE H ET AL., PLOS ONE, vol. 7, 2012, pages e50746 |
| RICK KAMPS ET AL., INT. J. MOL. SCI., vol. 18, no. 2, 2017, pages 308 |
| SAMBROOK ET AL.: "Molecular Cloning: A Laboratory Manual", 1989, COLD SPRING HARBOR LABORATORY PRESS |
| SAMBROOK, J.RUSSEL, D.: "Molecular Cloning, A LABORATORY MANUAL", vol. 1, 15 January 2001, COLD SPRING HARBOR LABORATORY PRESS |
| SMITH JL ET AL., J. VIROL., vol. 86, 2012, pages 5278 - 5287 |
| SWAMINATHAN S ET AL., BIOCHEM. BIOPHYS. RES. COMMUN., vol. 434, 2013, pages 228 - 234 |
| TAKAFUMI KANAMORI ET AL.: "Pattern Recognition", 2009, KYORITSU SHUPPAN CO., LTD. |
| TANDON M ET AL., ORAL DIS., vol. 18, 2012, pages 127 - 131 |
| TIBSHIRANI R., J R STAT SOC SER B, vol. 58, 1996, pages 267 - 88 |
| VELTHUT-MEIKAS A ET AL., MOL. ENDOCRINOL., 2013 |
| VENABLES, W. N. ET AL.: "Modern Applied Statistics with S", 2002, SPRINGER |
| VOELLENKLE C ET AL., RNA, vol. 18, 2012, pages 472 - 484 |
| WANG HJ ET AL., SHOCK, vol. 39, 2013, pages 480 - 487 |
| WITTEN D ET AL., BMC BIOL., vol. 8, 2010, pages 58 |
| XIE X ET AL., NATURE, vol. 434, 2005, pages 338 - 345 |
| YASUSHI NAGATA ET AL.: "Basics of statistical multiple comparison methods", 2007, SCIENTIST PRESS CO. |
| YOO JK ET AL., BIOCHEM. BIOPHYS. RES. COMMUN., vol. 415, 2011, pages 567 - 572 |
| YOO JK ET AL., STEM CELLS DEV., vol. 21, 2012, pages 2049 - 2057 |
| ZHENG ZHANG ET AL., J. COMPUT. BIOL., vol. 7, 2000, pages 203 - 214 |
| ZOU, H., J R STAT SOC SER B, vol. 67, 2005, pages 301 - 320 |
Also Published As
| Publication number | Publication date |
|---|---|
| EP4130259A1 (en) | 2023-02-08 |
| CN115485378A (zh) | 2022-12-16 |
| CA3178889A1 (en) | 2021-10-07 |
| US20230212675A1 (en) | 2023-07-06 |
| KR20220160602A (ko) | 2022-12-06 |
| JPWO2021201092A1 (ja) | 2021-10-07 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP7397449B2 (ja) | 大腸がんの検出キット又はデバイス及び検出方法 | |
| JP7448144B2 (ja) | 前立腺がんの検出キット又はデバイス及び検出方法 | |
| JP6970416B2 (ja) | 肺がんの検出キット又はデバイス及び検出方法 | |
| JP7426046B2 (ja) | 胃がんの検出キット又はデバイス及び検出方法 | |
| JP6927701B2 (ja) | 胆道がんの検出キット又はデバイス及び検出方法 | |
| JP2021100411A (ja) | 食道がんの検出キット又はデバイス及び検出方法 | |
| EP3754019B1 (en) | Use of a kit or device and method for detecting dementia | |
| EP3786307A1 (en) | Kit, device, and method for detecting bladder cancer | |
| WO2021201092A1 (ja) | 海馬萎縮を検出するためのキット又はデバイス及び方法 | |
| JP2023076054A (ja) | がん患者の緩和ケア病棟入院の要否を予測するためのキット、デバイス及び方法 | |
| JP2022028438A (ja) | アルツハイマー型認知症発症リスク評価のための方法、キット及びデバイス |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 21780961 Country of ref document: EP Kind code of ref document: A1 |
|
| ENP | Entry into the national phase |
Ref document number: 2022512615 Country of ref document: JP Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 3178889 Country of ref document: CA |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2021780961 Country of ref document: EP Effective date: 20221031 |