WO2018123925A1 - アルポート症候群治療薬 - Google Patents
アルポート症候群治療薬 Download PDFInfo
- Publication number
- WO2018123925A1 WO2018123925A1 PCT/JP2017/046304 JP2017046304W WO2018123925A1 WO 2018123925 A1 WO2018123925 A1 WO 2018123925A1 JP 2017046304 W JP2017046304 W JP 2017046304W WO 2018123925 A1 WO2018123925 A1 WO 2018123925A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- oligonucleotide
- exon
- col4a5
- acceptable salt
- nucleotide sequence
- Prior art date
Links
- 0 CC[C@](*1)(CCOC2[C@]1[n]1c(N=C(N)NC3=O)c3nc1)C2OP(O)(OC)=S Chemical compound CC[C@](*1)(CCOC2[C@]1[n]1c(N=C(N)NC3=O)c3nc1)C2OP(O)(OC)=S 0.000 description 11
- UQWHOARYJZPTLO-VHBSBENZSA-N CC(C)C[C@H](C(C1OC)OP(O)(OC)=O)O[C@H]1[n]1c(ncnc2NC)c2nc1 Chemical compound CC(C)C[C@H](C(C1OC)OP(O)(OC)=O)O[C@H]1[n]1c(ncnc2NC)c2nc1 UQWHOARYJZPTLO-VHBSBENZSA-N 0.000 description 1
- KGWDJBZCVBFVBQ-VCQNEHTGSA-N CC[C@](CCOC12)(C1OP(O)(O)=O)O[C@H]2[n]1c(ncnc2N)c2nc1 Chemical compound CC[C@](CCOC12)(C1OP(O)(O)=O)O[C@H]2[n]1c(ncnc2N)c2nc1 KGWDJBZCVBFVBQ-VCQNEHTGSA-N 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/70—Carbohydrates; Sugars; Derivatives thereof
- A61K31/7088—Compounds having three or more nucleosides or nucleotides
- A61K31/713—Double-stranded nucleic acids or oligonucleotides
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K48/00—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P13/00—Drugs for disorders of the urinary system
- A61P13/12—Drugs for disorders of the urinary system of the kidneys
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/78—Connective tissue peptides, e.g. collagen, elastin, laminin, fibronectin, vitronectin, cold insoluble globulin [CIG]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
- C12N15/1138—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing against receptors or cell surface proteins
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/11—Antisense
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/14—Type of nucleic acid interfering N.A.
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/31—Chemical structure of the backbone
- C12N2310/315—Phosphorothioates
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/32—Chemical structure of the sugar
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/32—Chemical structure of the sugar
- C12N2310/321—2'-O-R Modification
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/32—Chemical structure of the sugar
- C12N2310/323—Chemical structure of the sugar modified ring structure
- C12N2310/3231—Chemical structure of the sugar modified ring structure having an additional ring, e.g. LNA, ENA
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/34—Spatial arrangement of the modifications
- C12N2310/346—Spatial arrangement of the modifications having a combination of backbone and sugar modifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2320/00—Applications; Uses
- C12N2320/30—Special therapeutic applications
- C12N2320/33—Alteration of splicing
Definitions
- the present invention relates to a therapeutic agent for Alport syndrome, and more specifically, an oligonucleotide capable of inducing exon skipping having a truncating mutation and having a base number of multiples of 3 in the COL4A5 gene of Alport syndrome patients, and
- the present invention relates to a medicament (preferably a therapeutic agent for Alport syndrome).
- Alport syndrome is a severe hereditary kidney disease, most of which progresses to end-stage renal disease by the age of 30.
- the most common X chromosome-linked Alport syndrome (XLAS) is caused by an abnormality in the gene COL4A5, which encodes the type 4 collagen ⁇ 5 chain.
- the present inventors have conducted genetic diagnosis of XLAS patients of 300 families or more. As a result, about 15% of families have a truncating mutation that does not express a protein with a complete chain length, such as a nonsense mutation. In that case, it is clearer than a non-truncating mutation such as a missense mutation. It has been reported that it presents a severe clinical picture (Non-patent document 1: Kidney Int. 2013, 85, p.
- Non-Patent Document 2 J Am Soc Nephrol. 2000, 11, p.649-657
- Non-patent Literature 3 Nephrol Dial Transplant, 2002, 17, p.1218-1227.
- Non-Patent Document 4 J Am Soc Nephrol, 2010, Vol. 21, p.876-883)
- the age of progression of end-stage renal failure is 10 years or more faster when it has a truncating mutation than when it has a non-truncating mutation It has been found.
- the object of the present invention is to establish a molecular therapy for Alport syndrome.
- the present inventors established a therapeutic method for inducing skipping of exons having a truncating mutation by administering an antisense oligonucleotide (ASO) to an Alport syndrome patient having a truncating mutation (FIG. 1).
- ASO antisense oligonucleotide
- the COL4A5 gene consists of 35 exons of 44 exons forming a collagenous domain with a base number that is a multiple of 3. Therefore, in patients with truncaing mutations in these exons, this treatment leads to non-truncating mutations. Can be replaced. This makes it possible to delay the advanced age of end-stage renal failure in Alport syndrome patients who present a severe clinical picture. This is the world's first molecular therapy development study for XLAS.
- the gist of the present invention is as follows. (1) An oligonucleotide having a nucleotide sequence of 15 to 30 consisting of a nucleotide sequence complementary to the cDNA of COL4A5 gene, having a truncating mutation in COL4A5 gene of Alport syndrome patient, and having a base number that is a multiple of 3 Said oligonucleotide, pharmaceutically acceptable salt or solvate thereof capable of inducing skipping of an exon.
- the oligonucleotide according to (1) comprising a nucleotide sequence complementary to a part of the nucleotide sequence of an exon having a truncating mutation in the COL4A5 gene of a patient with Alport syndrome and having a base number that is a multiple of 3; A pharmaceutically acceptable salt or solvate thereof.
- the exon having a truncating mutation in the COL4A5 gene of a patient with Alport syndrome and having a base number that is a multiple of 3 is exon 24, 20 or 21 of the COL4A5 gene (1) or (2) Oligonucleotides, pharmaceutically acceptable salts or solvates thereof.
- oligonucleotide according to any one of (5) to (7), a pharmaceutically acceptable salt or solvate thereof, wherein the modification of the phosphodiester bond is phosphorothioate.
- a medicament comprising the oligonucleotide according to any one of (1) to (8), a pharmaceutically acceptable salt or solvate thereof.
- a therapeutic agent for Alport syndrome comprising the oligonucleotide according to any one of (1) to (8), a pharmaceutically acceptable salt or solvate thereof.
- An oligonucleotide having 15-30 bases consisting of a nucleotide sequence complementary to the cDNA of COL4A5 gene, having a truncating mutation in COL4A5 gene of Alport syndrome patient, and having a base number that is a multiple of 3
- a method for treating Alport syndrome comprising administering to a subject a pharmaceutically effective amount of the oligonucleotide, pharmaceutically acceptable salt or solvate thereof capable of inducing skipping of an exon.
- a 15-30 nucleotide oligonucleotide consisting of a nucleotide sequence complementary to the cDNA of COL4A5 gene for use in a method for treating Alport syndrome, wherein the mutation in the COL4A5 gene of a patient with Alport syndrome
- the oligonucleotide which is capable of inducing skipping of an exon having a base number that is a multiple of 3 and a pharmaceutically acceptable salt or solvate thereof.
- the truncaing mutation in a patient having a truncaing mutation in an exon of the COL4A5 gene, the truncaing mutation can be replaced with a non-truncating mutation. It becomes possible to delay.
- the present invention also makes it possible to prevent hearing loss and progression of eye lesions.
- FIG. 1 The schematic diagram explaining the treatment principle by the Alport syndrome therapeutic agent of this invention is shown.
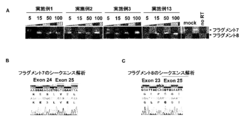
- A Effect of the compounds of Examples 1 to 27 on exon 24 of COL4A5.
- B, C The fragment 1 is a sequence before exon 24 causes skipping, whereas fragment 2 has a sequence that exon 24 skips.
- oligonucleotide for Exon 24 (Example 1, Example 2, Example 3) was introduced into the glomerular epithelial cell podocyte, COL4A5 protein expression (green) increased and Synaptopodin (red) expression increased
- FIG. 7 shows that fragment 7 has a sequence before exon 24 causes skipping, whereas fragment 8 has a sequence that exon 24 skips.
- the present invention is an oligonucleotide having 15 to 30 nucleotides consisting of a nucleotide sequence complementary to cDNA of COL4A5 gene, having a truncating mutation in COL4A5 gene of Alport syndrome patient, and having a base number which is a multiple of 3
- the oligonucleotide, pharmaceutically acceptable salt or solvate thereof, capable of inducing exon skipping is provided.
- the Alport syndrome is preferably X-linked Alport syndrome (XLAS).
- XLAS develops due to an abnormality in the COL4A5 gene, which encodes the type 4 collagen ⁇ 5 chain.
- the COL4A5 gene consists of 35 exons of 44 exons forming a collagenous domain with a base number that is a multiple of 3. Therefore, in patients with truncaing mutations in these exons, treatment with the oligonucleotide of the present invention is applied. Can be substituted for non-truncating mutations.
- a truncating mutation is a mutation in which protein synthesis is interrupted due to a nonsense mutation, a frameshift mutation, a splice site mutation, or the like.
- non-truncating mutations have mutations such as missense mutations and exon deletion mutations, but protein synthesis is performed without shifting the reading frame.
- missense mutation is on an exon that is a multiple of 3
- exon skipping using the oligonucleotide of the present invention results in a shortened protein.
- it is a functioning protein, it can be expected to be mild, and it can be a target for a therapeutic method in which the oligonucleotide of the present invention is administered.
- the exon targeted for skipping by the oligonucleotide of the present invention may have a base number that is a multiple of 3 of the exons (exons 3-46) located in the collagenous domain of the COL4A5 gene. Those having a base number that is a multiple of 3 are exons 3-18, 20-22, 24, 26, 27, 30-35, 38-41, 44-46.
- the oligonucleotide of the present invention may target an exon having a truncating mutation in the COL4A5 gene of a patient with Alport syndrome (particularly XLAS) and having a base number that is a multiple of 3; (Especially XLAS) from consecutive 15-30 nucleotides in the nucleotide sequence of exons 3-18, 20-22, 24, 26, 27, 30-35, 38-41, 44-46 of the patient's COL4A5 gene A nucleotide sequence complementary to the sequence.
- the oligonucleotide (antisense oligonucleotide) of the present invention has an exon having a truncating mutation in the COL4A5 gene of an Alport syndrome (particularly XLAS) patient and having a base number that is a multiple of 3 (for example, exon 3- It may be composed of a nucleotide sequence complementary to a part of the nucleotide sequence of 18, 20-22, 24, 26, 27, 30-35, 38-41, 44-46.
- An oligonucleotide consisting of a nucleotide sequence complementary to a part of the nucleotide sequence of an exon having a truncating mutation and having a base number that is a multiple of 3 in the COL4A5 gene of a patient with Alport syndrome (particularly XLAS) is SEQ ID NO: Examples including all or part of the sequence of any one of 1 to 28, 37 to 41, 51 or 52 (wherein t may be u and u may be t) can do.
- the “part of the sequence” usually means 80% or more of the entire sequence, preferably 85%, more preferably 90%, and most preferably 94%. It is.
- the number of bases in the oligonucleotide of the present invention is suitably 15-30, preferably 15-21, more preferably 16-20.
- the nucleotide constituting the oligonucleotide (antisense oligonucleotide) of the present invention may be any of natural DNA, natural RNA, DNA / RNA chimera, and modifications thereof, but at least one of them is a modified nucleotide. Preferably there is.
- Modified nucleotides include sugar-modified ones (eg, D-ribofuranose is 2'-O-alkylated, D-ribofuranose is 2'-O, -4'-C-alkylenated) ), A phosphodiester bond modified (for example, thioated), a base modified, a combination thereof, and the like.
- oligonucleotide in which at least one D-ribofuranose constituting an antisense oligonucleotide is 2'-O-alkylated or 2'-O, 4'-C-alkylenated have a high binding ability to RNA
- a higher therapeutic effect can be expected than natural nucleotides (ie, oligo DNA and oligo RNA).
- those in which at least one phosphodiester bond constituting the oligonucleotide is thioated are also highly resistant to nucleases, and thus are expected to have a higher therapeutic effect than natural nucleotides (ie, oligo DNA and oligo RNA). it can.
- Oligonucleotides containing both modified sugars and modified phosphates as described above are more resistant to nucleases and thus can be expected to have even higher therapeutic effects.
- examples of sugar modifications include 2′-O-alkylation of D-ribofuranose (eg, 2′-O-methylation, 2′-O-aminoethylation, 2'-O-propylation, 2'-O-allylation, 2'-O-methoxyethylation, 2'-O-butylation, 2'-O-pentylation, 2'-O-propargylation, etc.) 2′-O, 4′-C-alkylenation of D-ribofuranose (eg 2′-O, 4′-C-ethylenation, 2′-O, 4′-C-methyleneation, 2′- O, 4'-C-propylenation, 2'-O, 4'-C-tetramethyleneation, 2'-O, 4'-C-pentamethyleneation, etc.), 2'-deoxy- of D-ribofuranose 2'-C,
- Examples of the modification of the phosphodiester bond for the oligonucleotide include a phosphorothioate bond, a methylphosphonate bond, a methylthiophosphonate bond, a phosphorodithioate bond, and a phosphoramidate bond.
- Examples of base modifications include cytosine 5-methylation, 5-fluorination, 5-bromination, 5-iodination, N4-methylation, thymine 5-demethylation (uracil), 5-fluorination, Examples thereof include 5-bromination, 5-iodination, N6-methylation of adenine, 8-bromination, N2-methylation of guanine, 8-bromination and the like.
- Oligonucleotides are described in the literature (Nucleic Acids Research, 12, 4539 (1984)) using a commercially available synthesizer (for example, model 392 based on the phosphoramidide method of PerkinElmer). It can be synthesized according to the method.
- the phosphoramidite reagents used here are natural nucleosides and 2′-O-methyl nucleosides (ie, 2′-O-methyl guanosine, 2′-O-methyl adenosine, 2′-O-methyl cytidine, Regarding 2′-O-methyluridine, commercially available reagents can be used.
- 2'-O-alkylguanosine, adenosine, cytidine and uridine having 2 to 6 carbon atoms in the alkyl group are as follows.
- 2'-O-aminoethylguanosine, adenosine, cytidine, and uridine can be synthesized according to literature (Blommers et al. Biochemistry (1998), 37, 17714-17725.).
- 2'-O-propylguanosine, adenosine, cytidine, uridine can be synthesized according to the literature (Lesnik, E.A. et al. Biochemistry (1993), 32, 7832-7838.).
- reagents can be used for 2′-O-allylguanosine, adenosine, cytidine, and uridine.
- 2′-O-methoxyethyl guanosine, adenosine, cytidine and uridine can be synthesized according to patent (US6261840) or literature (Martin, P. Helv. Chim. Acta. (1995) 78, 486-504.
- 2′-O-butylguanosine, adenosine, cytidine and uridine can be synthesized according to the literature (Lesnik, E.A. et al. Biochemistry (1993), 32, 7832-7838.).
- 2'-O-pentylguanosine, adenosine, cytidine and uridine can be synthesized according to the literature (Lesnik, E.A. et al. Biochemistry 1993 (1993), 32, 7832-7838.).
- reagents can be used for 2′-O-propargylguanosine, adenosine, cytidine, and uridine.
- 2′-O, 4′-C-methyleneguanosine, adenosine, cytidine, 5-methylcytidine and thymidine according to the method described in WO99 / 14226, 2′-O having 2 to 5 carbon atoms in the alkylene group , 4′-C-alkyleneguanosine, adenosine, cytidine, 5-methylcytidine and thymidine can be produced according to the method described in WO00 / 47599.
- the 2'-deoxy-2'-C, 4'-C-methyleneoxymethyleneated nucleoside of D-ribofuranose was synthesized according to the literature (Wang, G. et al. Tetrahedron (1999), 55, 7707-7724). it can.
- S-cEt (constrained ethyl) can be synthesized according to the literature (Seth, P.P. et al. J.Org.Chem (2010), 75, 1569-1581.).
- AmNA can be synthesized according to literature (Yahara, A. et al., ChemBioChem (2012), 13, 2513-2516.) Or WO2014 / 109384.
- uracil (U) and thymine (T) are interchangeable. Both uracil (U) and thymine (T) can be used for base pairing with the complementary strand of adenine (A).
- a phosphoramidite reagent react with a reagent such as sulfur, tetraethylthiuram disulfide (TETD, Applied Biosystems), Beaucage reagent (Glen Research), or xanthan hydride to make an antisense with a phosphorothioate bond
- TETD tetraethylthiuram disulfide
- Beaucage reagent Glen Research
- xanthan hydride xanthan hydride
- CPG controlled pore glass
- commercially available products having 2′-O-methyl nucleosides bound thereto can be used.
- 2′-O, 4′-C-methyleneguanosine, adenosine, 5-methylcytidine and thymidine according to the method described in WO99 / 14226, 2′-O having 2 to 5 carbon atoms in the alkylene group , 4'-C-alkylene guanosine, adenosine, 5-methylcytidine and thymidine
- the nucleosides prepared according to the method described in WO00 / 47599 according to the literature (Oligonucleotide Synthesis, Edited by MJGait, Oxford University Press, 1984) Can bind to CPG.
- an oligonucleotide having a 2-hydroxyethyl phosphate group bonded to the 3 ′ end can be synthesized.
- an oligonucleotide having a hydroxyalkyl phosphate group or an aminoalkyl phosphate group bonded to the 3 ′ end can be synthesized.
- the oligonucleotide (antisense oligonucleotide) of the present invention can be used for the treatment of Alport syndrome.
- Oligonucleotides may be used in the form of pharmaceutically acceptable salts.
- “Pharmaceutically acceptable salt” refers to a salt of an oligonucleotide (antisense oligonucleotide), such as an alkali metal salt such as sodium salt, potassium salt, lithium salt, calcium salt, Alkaline earth metal salts such as magnesium salts, aluminum salts, iron salts, zinc salts, copper salts, nickel salts, metal salts such as cobalt salts; inorganic salts such as ammonium salts, t-octylamine salts, dibenzylamine Salt, morpholine salt, glucosamine salt, phenylglycine alkyl ester salt, ethylenediamine salt, N-methylglucamine salt, guanidine salt, diethylamine salt, triethylamine salt, dicyclohexylamine salt, N, N'-dibenzylethylenedi
- an alkali metal salt such as
- Oligonucleotides (antisense oligonucleotides) and pharmaceutically acceptable salts thereof may also exist as solvates (eg, hydrates), and may be such solvates. .
- oligonucleotide antisense oligonucleotide
- a pharmaceutically acceptable salt or solvate thereof is used for the treatment of Alport syndrome, itself or an appropriate pharmaceutically acceptable excipient or diluent
- excipients eg, sugar derivatives such as lactose, sucrose, sucrose, mannitol, sorbitol; starch derivatives such as corn starch, potato starch, alpha starch, dextrin; cellulose derivatives such as crystalline cellulose Gum arabic; dextran; organic excipients such as pullulan; silicate derivatives such as light anhydrous silicic acid, synthetic aluminum silicate, calcium silicate, magnesium magnesium magnesium silicate; phosphates such as calcium hydrogen phosphate; calcium carbonate Carbonates such as: inorganic excipients such as sulfates such as calcium sulfate), lubricants (eg, stearic acid; metal stearates such as calcium stearate and magnesium stearate; talc; colloidal silica ; Wax like beeswax, gay wax Boric acid; adipic acid; sulfate such as sodium sulfate; glycol; fumaric acid
- the therapeutic agent of the present invention may contain 0.1 to 250 ⁇ moles / ml oligonucleotide (antisense oligonucleotide), preferably 1 to 50 ⁇ moles / ml oligonucleotide (antisense oligonucleotide), and pharmaceutically Acceptable salts or solvates, 0.02 to 10% w / v carbohydrate or polyhydric alcohol and 0.01 to 0.4% w / v pharmaceutically acceptable surfactant may be included.
- carbohydrate monosaccharides and / or disaccharides are particularly preferable.
- these carbohydrates and polyhydric alcohols include glucose, galactose, mannose, lactose, maltose, mannitol and sorbitol. These may be used alone or in combination.
- the surfactant include polyoxyethylene sorbitan mono-tri-ester, alkylphenyl polyoxyethylene, sodium taurocholate, sodium collate, and polyhydric alcohol ester.
- polyoxyethylene sorbitan mono-tri-esters are particularly preferred, and oleate, laurate, stearate and palmitate are particularly preferred as esters. These may be used alone or in combination.
- the therapeutic agent of the present invention may more preferably contain 0.03 to 0.09 M of a pharmaceutically acceptable neutral salt such as sodium chloride, potassium chloride and / or calcium chloride.
- the therapeutic agent of the present invention can further preferably contain 0.002 to 0.05 M of a pharmaceutically acceptable buffer.
- buffering agents include sodium citrate, sodium glycinate, sodium phosphate, tris (hydroxymethyl) aminomethane. These buffering agents may be used alone or in combination.
- the above therapeutic agent may be supplied in a solution state.
- a solution such as distilled water for injection
- the therapeutic agent of the present invention includes those in a lyophilized state for reconstitution with a solution so that each component is in a predetermined concentration range.
- amino acids such as albumin and glycine may be further contained.
- oligonucleotide antisense oligonucleotide
- a pharmaceutically acceptable salt or solvate thereof is administered to a human, for example, about 0.01 to 100 mg / kg (body weight) per day for an adult, preferably The dosage may be 0.1 to 20 mg / kg (body weight), and it may be subcutaneously, intravenously or intravenously divided into 1 or several times. It can be appropriately changed depending on symptoms, age, administration method, and the like.
- an oligonucleotide (antisense oligonucleotide), a pharmaceutically acceptable salt or solvate thereof to a patient with Alport syndrome can be performed, for example, as follows. That is, an oligonucleotide (antisense oligonucleotide), a pharmaceutically acceptable salt or solvate thereof is produced by a method well known to those skilled in the art, and sterilized by a conventional method, for example, 125 mg / ml for injection Prepare the solution.
- oligonucleotide antisense oligonucleotide
- Administration is performed, for example, at an interval of one week, and thereafter, this treatment is repeated as appropriate while confirming the therapeutic effect by reducing urinary protein, suppressing the progression of renal dysfunction, and the like.
- the therapeutic agent for Alport syndrome of the present invention may be used in combination with other therapeutic agents such as an ACE inhibitor and an angiotensin receptor antagonist.
- Example 1 HO-C m1s -C e2s -C m1s -U m1s -G e2s -G m1s -C m1s -A e2s -A m1s -U m1s -C e2s -C m1s -A m1s -T e2s -C m1s -C m1s- T e2s -G m1t -H (ex24_011) (SEQ ID NO: 1) Synthesis was performed using the phosphoramidite method (Nucleic Acids Research, 12, 4539 (1984) using an automated nucleic acid synthesizer (MerMade 192X manufactured by BioAutomation), and activator solution-3 (0.25 mol / L) as a reagent.
- phenylacetyl disulfide (CARBOSYNTH, product No. FP07495) was used to obtain 0.2 M.
- Nitrile dehydrated, manufactured by Kanto Chemical Co., product No. 01837-05
- pyridine dehydrated, manufactured by Kanto Chemical Co., product No. 11339-05
- Reagents include 2'-O-Me nucleoside phosphoramidites (adenosine product No. ANP-5751, cytidine product No.
- ANP-5752, guanosine product product No. ANP-5753, uridine product No. ANP- No. 5754) manufactured by ChemGenes was used as the non-natural phosphoramidite in Example 14 (5′-O-dimethoxytrityl-2′-O, 4′-C-ethylene-6) of JP-A-2000-297097.
- Example 27 (5'-O-dimethoxytrityl-2'-O, 4'-C-ethylene- 2-N-isobutyrylguanosine-3′-O- (2-cyanoethyl N, N-diisopropyl) phosphoramidite),
- Example 22 (5′-O-dimethoxytrityl-2′-O, 4′-C -Ethylene-4-N- Benzoyl-5-methylcytidine-3'-O- (2-cyanoethyl N, N-diisopropyl) phosphoramidite),
- Example 9 (5'-O-dimethoxytrityl-2'-O, 4'-C-ethylene -5-Methyluridine-3'-O- (2-cyanoethyl N, N-diisopropyl) phosphoramidite)
- the oligomer was excised from the support by treating the protected oligonucleotide analog having the target sequence with 600 ⁇ L of concentrated aqueous ammonia, and the protecting group cyanoethyl group on the phosphorus atom and the protecting group on the nucleobase were removed.
- the mixed solution of oligomers was mixed with 300 ⁇ L of Clarity QSP DNA Loading Buffer (Phenomenex) and charged on Clarity SPE 96-well plate (Phenomenex).
- DCA dichloroacetic acid
- This compound is a reverse phase HPLC (column (Phenomenex, Clarity 2.6 ⁇ Oligo-MS 100A (2.1 ⁇ 50 mm)), A solution: 100 mM hexafluoroisopropanol (HFIP), 8 mM triethylamine aqueous solution, B solution: methanol, B %: 10% ⁇ 25% (4 min, linear gradient); 60 ° C .; 0.5 minmL / min; 260 nm). The compound was identified by negative ion ESI mass spectrometry (calculated value: 6267.73, actual measurement value: 6266.64).
- the base sequence of this compound is a sequence complementary to nucleotide numbers 162604-162621 of Homo sapiens collagen type IV IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG_011977).
- nucleotide number of Homo sapiens collagen type IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG — 011977) is shown.
- the molecular weight in the table indicates an actual measurement value by negative ion ESI mass spectrometry.
- Example 4 HO-C m1s -C e2s -C m1s -T e2s -G m1s -G m1s -C e2s -A m1s -A m1s -U m1s -C m1s -C e2s -A m1s -U m1s -C e2s -C m1s- T e2s -G m1t -H (ex24_c01) (SEQ ID NO: 1)
- the oligomer was excised from the support by treating the protected oligonucleotide analog having the target sequence synthesized under the same conditions as in Example 1 with 600 ⁇ L of concentrated aqueous ammonia, and the protecting group cyanoethyl group on the phosphorus atom and The protecting group on the nucleobase was removed.
- Clarity QSP DNA Loading Buffer 300 ⁇ L was mixed with the mixed solution of oligomers and charged on a Clarity SPE 96 well plate (manufactured by Phenomenex).
- TEAB triethylammonium hydrogen carbonate aqueous solution
- DCA dichloroacetic acid
- This compound was obtained by reverse-phase HPLC (column (Phenomenex, Clarity 2.6 ⁇ m Oligo-MS 100A (2.1 ⁇ 50 mm)), solution A: 100 mM hexafluoroisopropanol (HFIP), 8 mM triethylamine aqueous solution, solution B: methanol, B %: 10% ⁇ 25% (4 min, linear gradient); 60 ° C .; 0.5 mL / min; 260 nm). The compound was identified by negative ion ESI mass spectrometry (calculated value: 6304.76, measured value: 6304.75).
- the base sequence of this compound is a sequence complementary to nucleotide numbers 162604-162621 of Homo sapiens collagen type IV IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG_011977).
- Example 5 to 27 were synthesized in the same manner as Example 4.
- the data of Example 4 and Examples 5 to 27 are shown in Table 2.
- Table 2 ----------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- 4 ex24_c01 cCcTggCaaucCauCcTg 162604 162621 6304.75 1 5 ex24_c02 cCcTggCaaucCaucCTg 162604 162621 6304.74 1 6 ex24_c03 cCcTggCaaTcCauCcTg 162604 162621 6330.76 1 7 ex24_c04 CcCuggCaaTcCauCcTg 162604 162621 6330.76 1 8 ex24_c05 cCcT
- nucleotide number of Homo sapiens collagen type IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG — 011977) is shown.
- the molecular weight in the table indicates an actual measurement value by negative ion ESI mass spectrometry.
- Example 28 HO-U m1s -A e2s -U m1s -A m1s -G e2s -C m1s -U m1s -T e2s -A m1s -C m1s -T e2s -A m1s -G m1s -G e2s -A m1s -G m1s- G e2s -A m1t -H (ex20_001) (SEQ ID NO: 4) Synthesis and purification were performed under the same conditions as in Example 1 to obtain the target compound.
- the base sequence of this compound is a sequence complementary to nucleotide numbers 156301-156318 of Homo sapiens collagen type IV IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG_011977).
- nucleotide number of Homo sapiens collagen type IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG — 011977) is shown.
- the molecular weight in the table indicates an actual measurement value by negative ion ESI mass spectrometry.
- This compound was obtained by reverse-phase HPLC (column (Phenomenex, Clarity 2.6 ⁇ m Oligo-MS 100A (2.1 ⁇ 50 mm)), solution A: 100 mM hexafluoroisopropanol (HFIP), 8 mM triethylamine aqueous solution, solution B: methanol, B %: 10% ⁇ 25% (4 min, linear gradient); 60 ° C .; 0.5 mL / min; 260 nm). The compound was identified by negative ion ESI mass spectrometry (calculated value: 6357.74, measured value: 6357.78).
- the base sequence of this compound is a sequence complementary to nucleotide number 156303-156320 of Homo sapiens collagen type IV IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG_011977).
- nucleotide number of Homo sapiens collagen type IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG — 011977) is shown.
- the molecular weight in the table indicates an actual measurement value by negative ion ESI mass spectrometry.
- HMVEC Skin Microvascular Endothelial Cell
- a primary culture system for falling cells in urine was prepared as follows. Urine ( ⁇ 100 mL) was collected from a patient genetically diagnosed with X-linked Alport syndrome. The supernatant was discarded by centrifugation (1,400 rpm ⁇ 10 min), and the cell mass was washed twice with cold PBS. The cell mass was suspended in a maintenance medium (EGM-MV (Lonza), 20% FBS, penicillin (100 U / ml), streptomycin (1 mg / ml), rifampicin (8 ⁇ g / ml)) at 37 ° C, 5 The cells were cultured in a% CO 2 incubator. After the second passage, the cells were cultured in a maintenance medium without rifampicin.
- the oligonucleotide produced in the example was transfected as follows. The following solutions A and B were prepared and mixed. The above mixture was incubated for 5 minutes at room temperature. The prepared solution was added at 10 ⁇ L per well and incubated for 48 hours.
- RNA extraction RNA was extracted as follows. Cells incubated 48 hours after oligonucleotide transfection were washed once with cold PBS. A cell lysate of SuperPrep Cell Lysis Kit & RT Kit for qPCR (TOYOBO) was added at 50 ⁇ L per well, shaken for 30 seconds, and incubated at room temperature for 5 minutes. 10 ⁇ L of SuperPrep Cell Lysis Kit & RT Kit for qPCR reaction stop solution was added to each well, shaken for 30 seconds, and incubated at room temperature for 2 minutes. A total of 60 ⁇ L of cell lysate was collected and subjected to the reverse transcription reaction described below.
- TOYOBO SuperPrep Cell Lysis Kit & RT Kit for qPCR
- the reverse transcription reaction was performed as follows.
- the reverse transcription reaction master mix 32 ⁇ L of SuperPrep Cell Lysis & RT Kit for qPCR and the cell lysate 8 ⁇ L were mixed.
- the reverse transcription reaction was performed as follows (37 ° C 15 min, 50 ° C 5 min, 98 ° C 5 min, 4 ° C Hold).
- the final product was stored at -30 ° C.
- PCR reaction and fragment sequence analysis PCR reaction and fragment sequence analysis were performed as follows. A PCR reaction solution was prepared as described below.
- PCR reaction was performed as follows (94 ° C. for 5 min, (94 ° C. for 30 sec, 60 ° C for 30 sec, 72 ° C for 30 sec) x 30-38 cycles, 72 ° C for 5 min for 4 ° C. Hold)
- PCR products were analyzed by electrophoresis on 1.8% agarose gel containing ethidium bromide. Each fragment was extracted from the gel with PCR purification kit (QIAGEN) and sequenced using BigDye v1.1. The nucleotide sequence was confirmed with ABI PRISM 3130 Genetic Analyzer (Applied Biosystems) and CLC Main Workbench 6 (CLC bio).
- Immunofluorescence staining of differentiated podocytes was performed as follows. Wash the differentiated podocytes with PBS. The cells were fixed with 4% paraformaldehyde + 4% sucrose / PBS for 10 minutes at room temperature. After washing 3 times with PBS, it was permeabilized with 0.1% Triton X-100 / PBS for 5 minutes at room temperature. After washing 3 times with PBS, it was blocked with 5% Fetal bovine serum / PBS. Primary antibody (rat anti-COL4A5 H53 antibody, Shigei Medical Research Institute; donated by Dr.
- Yoshikazu Sado mouse anti-Synaptopodin antibody, Niigata University graduate School of Medical and Dental Sciences; mouse hybridoma donated by Dr. Naganobu Yao The supernatant was used) at 4 ° C. overnight.
- secondary antibodies Alexa Fluor 488 goat anti-rat IgG, Alexa Fluor 555 goat anti-mouse IgG
- Hoechst33342 was reacted at room temperature for 10 minutes for nuclear staining. After the encapsulation, it was observed with a fluorescence microscope.
- Exon skipping by example compound in urine falling cells from HMVEC or Alport syndrome patients Exon 24 skipping of oligonucleotides to Exon 24 (Example 1 to Example 27) was investigated using HMVEC as shown in FIG. As a result, Exon 24 skipping was observed in the oligonucleotides of Examples 1 to 27 (FIG. 2A). In particular, Example 1, Example 2, Example 3, and Example 13 showed high skipping efficiency. Moreover, when the sequence analysis of the fragment seen by the gel was performed, it was confirmed that the fragment 2 is the sequence which Exon 24 skipped (FIG. 2B and 2C).
- urinary fall cells derived from Alport syndrome patients with mutations in Exon 24 were induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and three types of oligonucleotides against Exon 24 (Examples) In each of Examples 1, 2 and 3), 50 nM was introduced to confirm the effect. Expression of COL4A5 and differentiation podocyte marker Synaptopodin (goat anti-Synaptopodin antibody, SantaCruz) was evaluated.
- oligonucleotide In contrast to non-introduced cells (Mock), the expression of COL4A5 protein (green) increased in oligonucleotide-introduced cells, and the expression of the differentiation podocyte marker Synaptopodin (red) was also confirmed. This suggests that the introduction of oligonucleotide not only contributes to the increased expression of COL4A5 protein, but may also affect the differentiation potential of podocytes.
- Example 28 to 33 Exon 20 skipping of oligonucleotides against Exon 20 (Examples 28 to 33) was examined using COL4A5 highly expressing cells HMVEC.
- the oligonucleotides of Example 28 to Example 33 were subjected to Exon ⁇ ⁇ 20 skipping. 20 skipping was observed.
- Example 31 showed high skipping efficiency.
- Example 20 skipping of oligonucleotides to Exon 20 was examined using HMVEC that highly expresses COL4A5.
- the oligonucleotides of Example 34 to Example 51 were subjected to Exon ⁇ ⁇ 20 skipping. 20 skipping was observed (FIG. 5A).
- Example 43 showed high skipping efficiency.
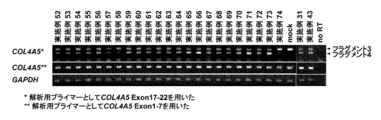
- urinary fall cells derived from Alport syndrome patients having a mutation in Exon 20 were induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and oligonucleotides to Exon 20 (Examples 52 to 52)
- Exon-20 skipping in Example 74 was examined, Exon-20 skipping was observed in all oligonucleotides (FIG. 7A).
- sequence analysis of the fragment seen by the gel was conducted, it was confirmed that the fragment 6 is a sequence skipped by Exon 20 (FIGS. 7B and 7C).
- urinary falling cells derived from Alport syndrome patients having mutations in Exon 24 were induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and oligonucleotides against Exon 24 (Example 1, 2, 3, 13) were introduced into each of 5, 15, ⁇ ⁇ ⁇ 50 or 100 nM and examined for Exon 24 skipping. Exon 24 skipping was observed in all oligonucleotides in a concentration-dependent manner (Fig. 8A). Moreover, when the sequence analysis of the fragment seen by the gel was conducted, it was confirmed that the fragment 8 is a sequence skipped by Exon 24 (FIGS. 8B and 8C).
- Example 75 to 98 The compounds of Examples 75 to 98 were synthesized in the same manner as Example 4. The sequences and data for the compounds of Examples 75 to 128 are listed in Table 5. Table 5 -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- 75 ex24_c25 cCcTggcaaTccauCcTg 162604 162621 6278.72 37 76 ex24_c26 CcCuggcaaTccauCcTg 162604 162621 6278.72 37 77 ex24_c27 cCcuggCaaTccaTccTg 162604 162621 6278.73 37 78 ex24_c28 cCcTggcaauccauCcTg 162604 162621 6252.72 37
- nucleotide number of Homo sapiens collagen type IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG — 011977) is shown.
- the molecular weight in the table indicates an actual measurement value by negative ion ESI mass spectrometry.
- Examples 99 to 139 The compounds of Examples 99 to 139 were synthesized in the same manner as Example 4. Table 6 shows the sequences and data of the compounds of Examples 99 to 139. Table 6 ----------------------------------------------------------------------------------------------------------------------------------------------------------------- 99 ex21_010 cTugGagTccTuuAucAc 156703 156686 6265.65 40 100 ex21_011 gGagTccTuuAucAccTg 156700 156683 6304.68 41 101 ex21_b08 cCuuGgaGucCuuTauCa 156704 156687 6279.59 42 102 ex21_b09 uTggAguCcuTuaTcaCc 156702 156685 62
- nucleotide number of Homo sapiens collagen type IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG — 011977) is shown.
- the molecular weight in the table indicates an actual measurement value by negative ion ESI mass spectrometry.
- Test Example 2 Analysis of exon skipping inducing ability by ENA oligonucleotide
- HMVEC skin microvascular endothelial cell
- urine falling cell primary culture system urine falling cell primary culture system
- kidney glomerular epithelial cells kidney glomerular epithelial cells
- Differentiation induction, transfection of the oligonucleotides produced in the Examples RNA extraction, reverse transcription reaction, PCR reaction and fragment sequence analysis, and immunofluorescence staining of differentiation podocytes were performed.
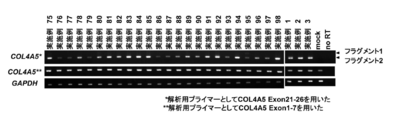
- Exon skipping by example compound in urinary falling cells from HMVEC or Alport syndrome patients Exon 24 skipping of oligonucleotides to Exon 24 (Example 75 to Example 98) was investigated using HMVEC as shown in FIG. As a result, in all the oligonucleotides of Example 75 to Example 98, Fragment 2 indicating that Exon 24 was skipping was detected as in FIG. 2, and Exon 24 skipping was observed.
- urinary fall cells derived from Alport syndrome patients having a mutation in Exon 24 were induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and oligonucleotides against Exon 24 (Example 75 to When Exon 24 skipping in Example 98) was examined, Fragment 2 indicating that Exon 24 was skipped was detected as in FIG. 2, and Exon 24 skipping was observed in all oligonucleotides.
- urinary fall cells derived from Alport syndrome patients having a mutation in Exon 24 were induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and oligonucleotides to Exon 24 (Example 83, In Example 85), 0.05, 0.5, 5, 15, 50, or 100 nM was introduced and examined for Exon 24 skipping. Exon 24 skipping was observed in all oligonucleotides in a concentration-dependent manner.
- urinary fall cells derived from Alport syndrome patients having mutations in Exon 24 were induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and five types of oligonucleotides against Exon 24 (Examples) 1, Example 2, Example 3, Example 83, Example 85) were introduced by 50 nM each, and the effect was confirmed.
- Expression of COL4A5 and differentiation podocyte marker Synaptopodin (goat anti-Synaptopodin antibody, SantaCruz) was evaluated.
- oligonucleotide In contrast to non-introduced cells (Mock), the expression of COL4A5 protein (green) increased in oligonucleotide-introduced cells, and the expression of the differentiation podocyte marker Synaptopodin (red) was also confirmed. This suggests that the introduction of oligonucleotide not only contributes to the increased expression of COL4A5 protein, but may also affect the differentiation potential of podocytes.
- HEK293A human embryonic kidney cell-derived cell line
- HEK293A human embryonic kidney cell-derived cell line
- DMEM maintenance medium
- Thermo Scientific 10% Fetal Bovine Serum
- the cells were peeled off with TrypLE Express (Thermo Scientific) to obtain a cell suspension, and the oligonucleotides produced in the examples as described below were introduced by the reverse transfection method.
- oligonucleotides produced in the examples were transfected as follows.
- the following solutions A and B were prepared and mixed.
- Solution A Opti-MEM Medium (Thermofisher) 4.8 ⁇ L Compound prepared in Example (10 ⁇ M) 0.5 ⁇ L (final concentration 50 nM)
- Solution B Opti-MEM Medium 5 ⁇ L Lipofectamine RNAiMAX (Thermofisher) 0.3 ⁇ L
- the above mixture was incubated for 15 minutes at room temperature.
- the prepared solution was dispensed 10.6 ⁇ L per well of 96 well plate. Cells were seeded on this plate at a density of 2 ⁇ 10 4 cells / well and incubated for 24 hours.
- RNA extraction RNA was extracted as follows. Cells incubated 24 hours after oligonucleotide transfection were washed once with cold PBS. The cell lysate of SuperPrep Cell Lysis Kit & RT Kit for qPCR (TOYOBO) was added at 50 ⁇ L per well, shaken for 30 seconds, and incubated at room temperature for 5 minutes. 10 ⁇ L of SuperPrep Cell Lysis Kit & RT Kit for qPCR reaction stop solution was added to each well, shaken for 30 seconds, and incubated at room temperature for 2 minutes. A total of 60 ⁇ L of cell lysate was collected and subjected to the reverse transcription reaction described below.
- the reverse transcription reaction was performed as follows.
- the reverse transcription reaction master mix 32 ⁇ L of SuperPrep Cell Lysis & RT Kit for qPCR and the cell lysate 8 ⁇ L were mixed.
- the reverse transcription reaction was performed as follows (37 ° C 15 min, 50 ° C 5 min, 98 ° C 5 min, 4 ° C Hold).
- the final product was stored at -20 ° C.
- PCR reaction and fragment sequence analysis PCR reaction and fragment sequence analysis were performed as follows. A PCR reaction solution was prepared as described below.
- PCR reaction was performed as follows (94 ° C. 2 min, (94 ° C. 30 sec, 55 ° C. 30 sec, 68 ° C. 1 min) ⁇ 30-40 cycles, 68 ° C. 10 min, 4 ° C. Hold).
- the primer for analysis was designed as follows.
- PCR products were analyzed with LabChip (registered trademark) GX (CaliperaliLifeSciences). The PCR product was electrophoresed on a 2% agarose gel containing ethidium bromide, and extracted from the gel with NucleoSpin (registered trademark) Gel and PCR Clean-up (MACHEREY-NAGEL). The sequence of the extracted DNA was performed using BigDye (registered trademark) Terminator v3.1. The nucleotide sequence was confirmed with Applied Biosystems 3730xl DNA Analyzer (Life Technologies).
- Example Compound in HEK293A As shown in FIG. 13, when Exon 21 skipping of oligonucleotides to Exon 21 (Example 99 to Example 139) was examined using HEK293A, 21 skipping was observed (FIG. 13A). In addition, sequence analysis was performed on the fragments extracted from the gel, and it was confirmed that fragment 9 was the sequence before skipping and fragment 10 was the sequence skipped by Exon 21 (FIGS. 13B and C).
- Test Example 4 Analysis of ability to induce exon skipping by ENA oligonucleotide
- HMVEC skin microvascular endothelial cell
- urine falling cell primary culture system urine falling cell primary culture system
- kidney glomerular epithelial cells kidney glomerular epithelial cells
- Differentiation induction, transfection of the oligonucleotides produced in the Examples RNA extraction, reverse transcription reaction, PCR reaction and fragment sequence analysis, and immunofluorescence staining of differentiation podocytes were performed.
- the primers for analysis were designed and used as follows.
- Example Compound in HMVEC or Alport Syndrome Patient-Derived Urinary Cells
- Exon 21 skipping of oligonucleotides to Exon 21 was investigated using HMVEC.
- Exon 21 skipping was observed in all oligonucleotides.
- sequence analysis of the fragment seen by the gel was performed, it was confirmed that the fragment 11 is the arrangement
- the oligonucleotide for Exon 21 was used at a final concentration of 5 nM.
- urinary falling cells derived from Alport syndrome patients having a mutation in Exon 21 are induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and oligonucleotides against Exon 21 (Examples 99 to 99)
- oligonucleotides against Exon 21 Examples 99 to 99
- Exon 21 skipping was observed in all oligonucleotides.
- the oligonucleotide for Exon 21 was used at a final concentration of 50 nM.
- urinary falling cells derived from Alport syndrome patients having mutations in Exon 21 were induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and oligonucleotides to Exon 21 (Example 105, In Example 114 and Example 116), 0.05, 0.5, 5, 15, 50 or 100 nM were introduced and examined for Exon 21 skipping. Exon 21 skipping was observed in all oligonucleotides in a concentration-dependent manner.
- urinary fall cells derived from Alport syndrome patients having a mutation in Exon 21 were induced to differentiate into renal glomerular epithelial cell podocytes, which are COL4A5-expressing cells, and three types of oligonucleotides against Exon 21 (Examples) 102, Example 105, Example 114, and Example 116) were introduced at 5 nM each, and the effect was confirmed.
- Expression of COL4A5 and differentiation podocyte marker Synaptopodin (goat anti-Synaptopodin antibody, SantaCruz) was evaluated.
- oligonucleotide In contrast to non-introduced cells (Mock), the expression of COL4A5 protein (green) increased in oligonucleotide-introduced cells, and the expression of the differentiation podocyte marker Synaptopodin (red) was also confirmed. This suggests that the introduction of oligonucleotide not only contributes to the increased expression of COL4A5 protein, but may also affect the differentiation potential of podocytes.
- Example 140 and 141 The compounds of Examples 140 and 141 were synthesized in the same manner as Example 4. The sequences and data for the compounds of Examples 140 and 141 are listed in Table 7. Table 7 --------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------
- nucleotide number of Homo sapiens collagen type IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG — 011977) is shown.
- the molecular weight in the table indicates an actual measurement value by negative ion ESI mass spectrometry.
- Example 142 HO-T e1s -G m1s -G m1s -A e1s -G m1s -U m1s -C m1s -U m1s -U m1s -U m1s -A e1s -U m1s -C m1s -A e1s -C m1s- C m1s -T 1t -H (ex21_Lc29) (SEQ ID NO: 44)
- the compound of Example 142 was synthesized using the phosphoramidite method (Nucleic Acids Research, 12, 4539 (1984).
- the LNA moiety was synthesized using the phosphoramidite body described in WO99 / 14226. .
- This compound is reverse phase HPLC (column (X-Bridge C18 2.5 ⁇ m (4.6 ⁇ 75 mm)), solution A: 100 mM hexafluoroisopropanol (HFIP), 8 mM triethylamine aqueous solution, solution B: methanol, B%: 5 When analyzed at% ⁇ 30% (20 min, linear gradient); 60 ° C .; 1 mL / min; 260 nm), it was eluted at 12.87 minutes. The compound was identified by negative ion ESI mass spectrometry (actual value: 6713.88).
- the base sequence of this compound is a sequence complementary to nucleotide numbers 156701-156684 of Homo sapiens collagen type IV alpha 5 chain (COL4A5) (NCBI-GenBank accession No. NG_011977).
- Example 143 to 280 The compounds of Examples 143 to 280 can be synthesized in the same manner as in Example 140.
- the sequences of the compounds of Examples 143 to 280 are shown in Tables 8 to 11.
- 42 ex21_Lb09 uTggAguCcuTuaTcaCc 156702 156685 43 147 ex21_
- the present invention can be used for the treatment of Alport syndrome.
- ⁇ SEQ ID NO: 1> This shows the nucleotide sequence of the oligonucleotide (ex24_011, ex24_c01, ex24_c02, ex24_c03, ex24_c04, ex24_c05, ex24_c06, ex24_c07, ex24_c08) produced in Examples 1, 4 to 11.
- ⁇ SEQ ID NO: 2> This shows the nucleotide sequence of the oligonucleotide (ex24_b04, ex24_c09, ex24_c10, ex24_c11, ex24_c12, ex24_c13, ex24_c14, ex24_c15, ex24_c16) produced in Examples 2 and 12-19.
- ⁇ SEQ ID NO: 3> This shows the nucleotide sequence of the oligonucleotide (ex24_b05, ex24_c17, ex24_c18, ex24_c19, ex24_c20, ex24_c21, ex24_c22, ex24_c23, ex24_c24) produced in Examples 3 and 20 to 27.
- ⁇ SEQ ID NO: 4> This shows the nucleotide sequence of the oligonucleotide (ex20_001) produced in Example 28.
- ⁇ SEQ ID NO: 5> This shows the nucleotide sequence of the oligonucleotide (ex20_002) produced in Example 29.
- ⁇ SEQ ID NO: 6> This shows the nucleotide sequence of the oligonucleotide (ex20_022) produced in Example 30.
- ⁇ SEQ ID NO: 7> This shows the nucleotide sequence of the oligonucleotide (ex20_023) produced in Example 31.
- ⁇ SEQ ID NO: 8> This shows the nucleotide sequence of the oligonucleotide (ex20_024) produced in Example 32.
- ⁇ SEQ ID NO: 9> This shows the nucleotide sequence of the oligonucleotide (ex20_044) produced in Example 33.
- ⁇ SEQ ID NO: 10> This shows the nucleotide sequence of the oligonucleotide (ex20_b02) produced in Example 34.
- ⁇ SEQ ID NO: 11> This shows the nucleotide sequence of the oligonucleotide (ex20_b03) produced in Example 35.
- ⁇ SEQ ID NO: 12> This shows the nucleotide sequence of the oligonucleotide (ex20_b04) produced in Example 36.
- ⁇ SEQ ID NO: 13> This shows the nucleotide sequence of the oligonucleotide (ex20_b05) produced in Example 37.
- ⁇ SEQ ID NO: 14> This shows the nucleotide sequence of the oligonucleotide (ex20_b07) produced in Example 38.
- ⁇ SEQ ID NO: 15> This shows the nucleotide sequence of the oligonucleotide (ex20_b09) produced in Example 39.
- ⁇ SEQ ID NO: 16> This shows the nucleotide sequence of the oligonucleotide (ex20_b10) produced in Example 40.
- ⁇ SEQ ID NO: 17> This shows the nucleotide sequence of the oligonucleotide (ex20_b11) produced in Example 41.
- ⁇ SEQ ID NO: 18> This shows the nucleotide sequence of the oligonucleotide (ex20_b12) produced in Example 42.
- ⁇ SEQ ID NO: 19> The oligonucleotides prepared in Examples 43 and 63 to 74 (ex20_b13, ex20_c12, ex20_c13, ex20_c14, ex20_c15, ex20_c16, ex20_c17, ex20_c18, ex20_c19, ex20_c20, ex20_c21, ex20_c22, ex20_c22, and the sequence of ex20_c23 are shown).
- ⁇ SEQ ID NO: 20> This shows the nucleotide sequence of the oligonucleotide (ex20_b14) produced in Example 44.
- ⁇ SEQ ID NO: 21> This shows the nucleotide sequence of the oligonucleotide (ex20_b15) produced in Example 45.
- ⁇ SEQ ID NO: 22> This shows the nucleotide sequence of the oligonucleotide (ex20_b16) produced in Example 46.
- ⁇ SEQ ID NO: 23> This shows the nucleotide sequence of the oligonucleotide (ex20_b17) produced in Example 47.
- ⁇ SEQ ID NO: 24> This shows the nucleotide sequence of the oligonucleotide (ex20_b18) produced in Example 48.
- ⁇ SEQ ID NO: 25> This shows the nucleotide sequence of the oligonucleotide (ex20_b19) produced in Example 49.
- ⁇ SEQ ID NO: 26> This shows the nucleotide sequence of the oligonucleotide (ex20_b20) produced in Example 50.
- ⁇ SEQ ID NO: 27> This shows the nucleotide sequence of the oligonucleotide (ex20_b21) produced in Example 51.
- ⁇ SEQ ID NO: 28> This shows the nucleotide sequence of the oligonucleotide (ex20_c01, ex20_c02, ex20_c03, ex20_c04, ex20_c05, ex20_c06, ex20_c07, ex20_c08, ex20_c09, ex20_c10, ex20_c11) produced in Examples 52-62.
- ⁇ SEQ ID NO: 29> This shows the nucleotide sequence of a forward (5'-3 ') primer for analysis of all COL4A5 expression (Exon 1-7).
- ⁇ SEQ ID NO: 30> This shows the nucleotide sequence of an entire COL4A5 expression analysis (Exon 1-7) Reverse (5'-3 ') primer.
- ⁇ SEQ ID NO: 31> This shows the nucleotide sequence of COL4A5 Exon 20 skipping analysis (Exon 17-22) Forward (5'-3 ') primer.
- ⁇ SEQ ID NO: 32> This shows the nucleotide sequence of COL4A5 Exon 20 skipping analysis (Exon 17-22) Reverse (5'-3 ') primer.
- ⁇ SEQ ID NO: 33> This shows the nucleotide sequence of COL4A5 Exon 24 skipping analysis (Exon 21-26) Forward (5'-3 ') primer.
- ⁇ SEQ ID NO: 34> This shows the nucleotide sequence of COL4A5 Exon 24 skipping analysis (Exon 21-26) Reverse (5'-3 ') primer.
- ⁇ SEQ ID NO: 36> This shows the nucleotide sequence of a forward (5'-3 ') primer for analysis of endogenous control gene (GAPDH).
- ⁇ SEQ ID NO: 36> This shows the nucleotide sequence of Reverse (5′-3 ′) primer for analysis of endogenous control gene (GAPDH).
- ⁇ SEQ ID NO: 37> This shows the nucleotide sequence of the oligonucleotide (ex24_c25, ex24_c26, ex24_c27, ex24_c28, ex24_c29, ex24_c30, ex24_c31, ex24_c32) produced in Examples 75-82.
- ⁇ SEQ ID NO: 38> This shows the nucleotide sequence of the oligonucleotide (ex24_c33, ex24_c34, ex24_c35, ex24_c36, ex24_c37, ex24_c38, ex24_c39) produced in Examples 83-90.
- ⁇ SEQ ID NO: 39> This shows the nucleotide sequence of the oligonucleotide (ex24_c41, ex24_c42, ex24_c43, ex24_c44, ex24_c45, ex24_c46, ex24_c47, ex24_c48) produced in Examples 91-98.
- ⁇ SEQ ID NO: 40> This shows the nucleotide sequence of the oligonucleotide (ex21_010, ex21_c10, ex21_c12, ex21_c13, ex21_c14) produced in Examples 99 and 113 to 116.
- ⁇ SEQ ID NO: 41> This shows the nucleotide sequence of the oligonucleotide (ex21_011, ex21_c34, ex21_c35, ex21_c36, ex21_c37) produced in Example 100, 136-139.
- ⁇ SEQ ID NO: 42> This shows the nucleotide sequence of the oligonucleotide (ex21_b08, ex21_c01, ex21_c02, ex21_c03, ex21_c04, ex21_c05, ex21_c06, ex21_c07, ex21_c08, ex21_c09) produced in Examples 101 and 104 to 112.
- ⁇ SEQ ID NO: 43> This shows the nucleotide sequence of the oligonucleotide (ex21_b09, ex21_c15, ex21_c16, ex21_c17, ex21_c18, ex21_c19, ex21_c20, ex21_c21, ex21_c22, ex21_c23) produced in Examples 102 and 117 to 125.
- ⁇ SEQ ID NO: 44> This shows the nucleotide sequence of the oligonucleotide (ex21_b10, ex21_c24, ex21_c25, ex21_c26, ex21_c27, ex21_c28, ex21_c29, ex21_c30, ex21_c31, ex21_c32, ex21_c33) produced in Examples 103 and 126 to 135.
- ⁇ SEQ ID NO: 45> This shows the nucleotide sequence of COL4A5 Exon 21 skipping analysis (Exon 20-23) Forward (5′-3 ′) primer.
- ⁇ SEQ ID NO: 46> This shows the nucleotide sequence of COL4A5 Exon 21 skipping analysis (Exon 20-23) Reverse (5′-3 ′) primer.
- ⁇ SEQ ID NO: 47> This shows the nucleotide sequence of a forward (5′-3 ′) primer for analyzing an endogenous control gene (ACTB).
- ⁇ SEQ ID NO: 48> This shows the nucleotide sequence of Reverse (5′-3 ′) primer for analysis of endogenous control gene (ACTB).
- ⁇ SEQ ID NO: 49> This shows the nucleotide sequence of COL4A5 Exon 21 skipping analysis (Exon 18-24) Forward (5′-3 ′) primer.
- ⁇ SEQ ID NO: 50> This shows the nucleotide sequence of COL4A5 Exon 21 skipping analysis (Exon 18-24) Reverse (5'-3 ') primer.
- ⁇ SEQ ID NO: 51> This shows the nucleotide sequence of the oligonucleotide (ex21_012-2) produced in Example 140.
- ⁇ SEQ ID NO: 52> This shows the nucleotide sequence of the oligonucleotide (ex21_012-4) produced in Example 141.
Abstract
Description
(1)COL4A5遺伝子のcDNAに相補的なヌクレオチド配列からなる塩基数15~30のオリゴヌクレオチドであって、アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのスキッピングを誘導できる前記オリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
(2)アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのヌクレオチド配列の一部に相補的なヌクレオチド配列からなる(1)記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
(3)アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンが、COL4A5遺伝子のエクソン24、20又は21である(1)又は(2)記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
(4)配列番号1~28、37~41、51又は52のいずれかの配列(但し、配列中のtはuであってもよく、uはtであってもよい)の全部又は一部を含む(1)~(3)のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
(5)オリゴヌクレオチドを構成する糖及び/又はリン酸ジエステル結合の少なくとも1個が修飾されている(1)~(4)のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
(6)オリゴヌクレオチドを構成する糖がD-リボフラノースであり、糖の修飾がD-リボフラノースの2’位の水酸基の修飾である(5)記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
(7)糖の修飾がD-リボフラノースの2’-O-アルキル化及び/又は2’-O, 4’-C-アルキレン化である(6)記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
(8)リン酸ジエステル結合の修飾がホスホロチオエートである(5)~(7)のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
(9)(1)~(8)のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物を含む、医薬。
(10)(1)~(8)のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物を含む、アルポート症候群治療薬。
(11)COL4A5遺伝子のcDNAに相補的なヌクレオチド配列からなる塩基数15~30のオリゴヌクレオチドであって、アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのスキッピングを誘導できる前記オリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物を医薬的に有効な量で被験者に投与することを含む、アルポート症候群の治療方法。
(12)アルポート症候群の治療のための、COL4A5遺伝子のcDNAに相補的なヌクレオチド配列からなる塩基数15~30のオリゴヌクレオチドであって、アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのスキッピングを誘導できる前記オリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物の使用。
(13)アルポート症候群を治療する方法に使用するための、COL4A5遺伝子のcDNAに相補的なヌクレオチド配列からなる塩基数15~30のオリゴヌクレオチドであって、アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのスキッピングを誘導できる前記オリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
本明細書は、本願の優先権の基礎である日本国特許出願、特願2016‐254906及び特願2017‐077374の明細書および/または図面に記載される内容を包含する。
また、ミスセンス変異を持ち重症化が懸念されるXLAS患者であって、ミスセンス変異が3の倍数のエクソン上にある場合、本発明のオリゴヌクレオチドを用いてエクソンスキッピングさせると、短縮化したタンパク質になるが、機能するタンパク質であれば、軽症化することも期待でき、本発明のオリゴヌクレオチドを投与する治療法の対象になりうる。
本発明のオリゴヌクレオチドは、アルポート症候群(特に、XLAS)患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンを標的とするものであるとよく、例えば、アルポート症候群(特に、XLAS)患者のCOL4A5遺伝子のエクソン3-18、20-22、24、26、27、30-35、38-41、44-46のヌクレオチド配列中の連続する15~30個のヌクレオチドからなる配列に相補的なヌクレオチド配列を含むものである。すなわち、本発明のオリゴヌクレオチド(アンチセンスオリゴヌクレオチド)は、アルポート症候群(特に、XLAS)患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソン(例えば、エクソン3-18、20-22、24、26、27、30-35、38-41、44-46のヌクレオチド配列の一部に相補的なヌクレオチド配列からなるとよい。
D-リボフラノースの2’-deoxy-2’-C,4’-C-メチレンオキシメチレン化されたヌクレオシドは、文献(Wang,G. et al. Tetrahedron (1999), 55, 7707-7724に従って合成できる。
(実施例1)
HO-Cm1s-Ce2s-Cm1s-Um1s-Ge2s-Gm1s-Cm1s-Ae2s-Am1s-Um1s-Ce2s-Cm1s-Am1s-Te2s-Cm1s-Cm1s-Te2s-Gm1t-H (ex24_011)(配列番号1)
核酸自動合成機(BioAutomation製MerMade 192X)を用い、ホスホロアミダイト法(Nucleic Acids Research, 12, 4539 (1984)を用いて合成を行った。試薬としては、アクチベーター溶液-3 (0.25 mol/L 5-ベンジルチオ-1H-テトラゾール・アセトニトリル溶液、和光純薬工業製、product No. 013-20011)、CAP A for AKTA (1-メチルイミダゾール・アセトニトリル溶液、Sigma-Aldrich製、product No. L040050)、Cap B1 for AKTA (無水酢酸・アセトニトリル溶液、Sigma-Aldrich製、product No. L050050)、Cap B2 for AKTA (ピリジン・アセトニトリル溶液、Sigma-Aldrich製、product No. L050150)、DCA Deblock (ジクロロ酢酸・トルエン溶液、Sigma-Aldrich製、product No. L023050)を用いた。ホスホロチオエート結合を形成するためのチオ化試薬として、0.2 Mになるようにフェニルアセチルジスルフィド(CARBOSYNTH製、product No. FP07495)を、アセトニトリル (脱水、関東化学製、product No. 01837-05)、ピリジン (脱水、関東化学製、product No. 11339-05)1:1(v/v)溶液を用いて溶解して用いた。アミダイト試薬としては、2'-O-Meヌクレオシドのホスホロアミダイト(アデノシン体product No. ANP-5751, シチジン体product No. ANP-5752,グアノシン体product No. ANP-5753, ウリジン体product No. ANP-5754)はChemGenes製のものを用いた。非天然型のホスホロアミダイトは特開2000-297097の実施例14(5'-O-ジメトキシトリチル-2'-O,4'-C-エチレン-6-N-ベンゾイルアデノシン-3'-O-(2-シアノエチル N,N-ジイソプロピル)ホスホロアミダイト)、実施例27 (5'-O-ジメトキシトリチル-2'-O,4'-C-エチレン-2-N-イソブチリルグアノシン-3'-O-(2-シアノエチル N,N-ジイソプロピル)ホスホロアミダイト)、実施例22(5'-O-ジメトキシトリチル-2'-O,4'-C-エチレン-4-N-ベンゾイル-5-メチルシチジン-3'-O-(2-シアノエチル N,N-ジイソプロピル)ホスホロアミダイト)、実施例9(5'-O-ジメトキシトリチル-2'-O,4'-C-エチレン-5-メチルウリジン-3'-O-(2-シアノエチル N,N-ジイソプロピル)ホスホロアミダイト)、の化合物を用いた。固相担体として、Glen Unysupport FC 96ウェルフォーマット0.2 μmol(GlenResearch製)を用い、表記の化合物を合成した。但し、アミダイト体の縮合に要する時間は、約9分とした。
表1
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 分子量 配列番号
---------------------------------------------------------------------
1 ex24_011 cCcuGgcAauCcaTccTg 162604 162621 6276.64 1
2 ex24_b04 cCugGcaAucCauCcuGu 162603 162620 6263.60 2
3 ex24_b05 cTggCaaTccAucCugTc 162602 162619 6291.66 3
---------------------------------------------------------------------
表中の配列において大文字はENA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。表中の分子量は、負イオンESI質量分析による実測値を示す。
(実施例4)
HO-Cm1s-Ce2s-Cm1s-Te2s-Gm1s-Gm1s-Ce2s-Am1s-Am1s-Um1s-Cm1s-Ce2s-Am1s-Um1s-Ce2s-Cm1s-Te2s-Gm1t-H (ex24_c01)(配列番号1)
実施例1と同様の条件で合成した目的配列を有する保護されたオリゴヌクレオチド類縁体を600 μLの濃アンモニア水で処理することによってオリゴマーを支持体から切り出すとともに、リン原子上の保護基シアノエチル基と核酸塩基上の保護基をはずした。オリゴマーの混合溶液を、Clarity QSP DNA Loading Buffer(Phenomenex製)300 μLを混合し、Clarity SPE 96 well plate(Phenomenex製)上にチャージした。Clarity QSP DNA Loading Buffer:水 = 1:1溶液1 mL、0.1 M炭酸水素トリエチルアンモニウム水溶液(TEAB):水 = 8:2溶液 1mL、3%ジクロロ酢酸(DCA)水溶液3 mL、水2 mL、20 mM Tris水溶液1 mLの順に添加した後、20 mM Tris水溶液:アセトニトリル = 9:1溶液にて抽出される成分を集めた。溶媒留去後、目的化合物を得た。本化合物は、逆相HPLC(カラム(Phenomenex, Clarity 2.6 μm Oligo-MS 100A (2.1×50 mm))、A溶液:100 mMヘキサフルオロイソプロパノール(HFIP)、8 mMトリエチルアミン水溶液、B溶液:メタノール、B%:10% → 25%(4min, linear gradient);60℃;0.5 mL/min;260 nm)で分析すると、2.733分に溶出された。化合物は負イオンESI質量分析により同定した(計算値:6304.76、実測値:6304.75)。
表2
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 分子量 配列番号
---------------------------------------------------------------------
4 ex24_c01 cCcTggCaaucCauCcTg 162604 162621 6304.75 1
5 ex24_c02 cCcTggCaaucCaucCTg 162604 162621 6304.74 1
6 ex24_c03 cCcTggCaaTcCauCcTg 162604 162621 6330.76 1
7 ex24_c04 CcCuggCaaTcCauCcTg 162604 162621 6330.76 1
8 ex24_c05 cCcTggCaAuCcAuCcTg 162604 162621 6328.75 1
9 ex24_c06 cCcTggCaaTcCaTcCTg 162604 162621 6356.79 1
10 ex24_c07 CCcTggCaaTcCaTcCTg 162604 162621 6382.80 1
11 ex24_c08 cCcTgGcAaTcCaTcCuG 162604 162621 6340.74 1
12 ex24_c09 cCuggCaaTcCauCcTgu 162603 162620 6305.74 2
13 ex24_c10 CcTggCaauccauCcTgT 162603 162620 6305.74 2
14 ex24_c11 cCuggCaaTcCauCcTgT 162603 162620 6661.74 2
15 ex24_c12 ccTggCaAuCcAuCcTgu 162603 162620 6303.72 2
16 ex24_c13 cCTggCaaTcCauCcTgT 162603 162620 6357.76 2
17 ex24_c14 CcTggCaAuCcAuCcTgu 162603 162620 6329.73 2
18 ex24_c15 cCTggCaaTcCaTCcTgT 162603 162620 6383.78 2
19 ex24_c16 CcTggCaAuCcAuCcTgT 162603 162620 6355.75 2
20 ex24_c17 cTggCaaTccaTcCugTc 162602 162619 6305.73 3
21 ex24_c18 cTggCaauCcaTccTguC 162602 162619 6305.73 3
22 ex24_c19 cTggCaaTcCaTcCugTc 162602 162619 6331.77 3
23 ex24_c20 CTggCaauCcaTcCugTc 162602 162619 6331.75 3
24 ex24_c21 cTggCaAuCcAuCcTgTc 162602 162619 6329.74 3
25 ex24_c22 CTggCaaTccAucCugTC 162602 162619 6343.75 3
26 ex24_c23 CTggCaaTcCaTcCugTC 162602 162619 6383.78 3
27 ex24_c24 cTgGcAaTcCaTcCuGuC 162602 162619 6341.74 3
---------------------------------------------------------------------
表中の配列において大文字はENA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。表中の分子量は、負イオンESI質量分析による実測値を示す。
(実施例28)
HO-Um1s-Ae2s-Um1s-Am1s-Ge2s-Cm1s-Um1s-Te2s-Am1s-Cm1s-Te2s-Am1s-Gm1s-Ge2s-Am1s-Gm1s-Ge2s-Am1t-H (ex20_001)(配列番号4)
実施例1と同様の条件で合成および精製を行い、目的化合物を得た。本化合物は、逆相HPLC(カラム(Phenomenex, Clarity 2.6 μm Oligo-MS 100A (2.1×50 mm))、A溶液:100 mMヘキサフルオロイソプロパノール(HFIP)、8 mMトリエチルアミン水溶液、B溶液:メタノール、B%:10% → 25%(4 min, linear gradient);60℃;0.5 mL/min;260 nm)で分析すると、3.113分に溶出された。化合物は負イオンESI質量分析により同定した(計算値:6401.73、実測値:6401.69)。
表3
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 分子量 配列番号
---------------------------------------------------------------------
28 ex20_001 uAuaGcuTacTagGagGa 156301 156318 6401.69 4
29 ex20_002 gCuuAcuAggAggAauGu 156297 156314 6403.65 5
30 ex20_022 gGagGucCagGaaTggAa 156217 156234 6518.74 6
31 ex20_023 gTccAggAauGgaAauTc 156213 156230 6400.68 7
32 ex20_024 aGgaAugGaaAuuCcaGg 156209 156226 6449.71 8
33 ex20_044 uAacTgcAgcCccTaaGa 156129 156146 6333.72 9
---------------------------------------------------------------------
表中の配列において大文字はENA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。表中の分子量は、負イオンESI質量分析による実測値を示す。
(実施例34~74)
HO-Am1s-Ae2s-Um1s-Am1s-Te2s-Am1s-Gm1s-Ce2s-Um1s-Um1s-Ae2s-Cm1s-Um1s-Ae2s-Gm1s-Gm1s-Ae2s-Gm1t-H (ex20_b02)(配列番号10)
実施例4と同様の条件で合成および精製を行い、目的化合物を得た。本化合物は、逆相HPLC(カラム(Phenomenex, Clarity 2.6 μm Oligo-MS 100A (2.1×50 mm))、A溶液:100 mMヘキサフルオロイソプロパノール(HFIP)、8 mMトリエチルアミン水溶液、B溶液:メタノール、B%:10% → 25%(4 min, linear gradient);60℃;0.5 mL/min;260 nm)で分析すると、3.213分に溶出された。化合物は負イオンESI質量分析により同定した(計算値:6385.74、実測値:6385.78)。
表4
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 分子量 配列番号
---------------------------------------------------------------------
34 ex20_b02 aAuaTagCuuAcuAggAg 156303 156320 6385.78 10
35 ex20_b03 aTauAgcTuaCuaGgaGg 156302 156319 6415.82 11
36 ex20_b04 aTagCuuAcuAggAggAa 156300 156317 6424.80 12
37 ex20_b05 uAgcTuaCuaGgaGgaAu 156299 156316 6401.77 13
38 ex20_b07 cTuaCuaGgaGgaAugTg 156296 156313 6431.74 14
39 ex20_b09 uAcuAggAggAauGugAg 156294 156311 6452.71 15
40 ex20_b10 gAggTccAggAauGgaAa 156216 156233 6488.77 16
41 ex20_b11 aGguCcaGgaAugGaaAu 156215 156232 6449.75 17
42 ex20_b12 gGucCagGaaTggAaaTu 156214 156231 6454.75 18
43 ex20_b13 uCcaGgaAugGaaAuuCc 156212 156229 6360.71 19
44 ex20_b14 cCagGaaTggAaaTucCa 156211 156228 6411.78 20
45 ex20_b15 cAggAauGgaAauTccAg 156210 156227 6409.73 21
46 ex20_b16 cCauAacTgcAgcCccTa 156132 156149 6283.74 22
47 ex20_b17 cAuaAcuGcaGccCcuAa 156131 156148 6265.72 23
48 ex20_b18 aTaaCugCagCccCuaAg 156130 156147 6361.79 24
49 ex20_b19 aAcuGcaGccCcuAagAu 156128 156145 6305.72 25
50 ex20_b20 aCugCagCccCuaAgaTu 156127 156144 6338.76 26
51 ex20_b21 cTgcAgcCccTaaGauTc 156126 156143 6300.73 27
52 ex20_c01 gTcCaggAaTggaaAuTc 156213 156230 6428.77 28
53 ex20_c02 gTcCaggAaTggaaaTuC 156213 156230 6442.78 28
54 ex20_c03 gTCcaggaATggaaaTTc 156213 156230 6442.78 28
55 ex20_c04 gTccAggAaTggaAauTc 156213 156230 6414.75 28
56 ex20_c05 gTccAggAauggaAauTc 156213 156230 6388.73 28
57 ex20_c06 gTcCaggaaTggaaaTuC 156213 156230 6430.78 28
58 ex20_c07 gTcCaggaaTggaaAuTc 156213 156230 6416.77 28
59 ex20_c08 gTccAggaaTggAaaTuc 156213 156230 6402.76 28
60 ex20_c09 gTccAggaauggaAauTc 156213 156230 6376.73 28
61 ex20_c10 gTcCaggaauggaaaTuC 156213 156230 6404.76 28
62 ex20_c11 gTcCaggaaTggaaauTc 156213 156230 6404.77 28
63 ex20_c12 uCcAggAaTggAaAuuCc 156212 156229 6386.75 19
64 ex20_c13 TccAggAauggAaaTucC 156212 156229 6374.74 19
65 ex20_c14 uCcAggaAuggAaAuuCc 156212 156229 6360.73 19
66 ex20_c15 TcCaggAaTggaaaTuCc 156212 156229 6402.77 19
67 ex20_c16 uCcAggAaTggaaaTuCc 156212 156229 6388.77 19
68 ex20_c17 uCcAggAauggAaAuuCc 156212 156229 6360.73 19
69 ex20_c18 uCcAggaauggaAaTuCc 156212 156229 6362.74 19
70 ex20_c19 TcCaggaaTggaaaTuCc 156212 156229 6390.77 19
71 ex20_c20 TcCaggaaTggaaauTcC 156212 156229 6390.77 19
72 ex20_c21 uCcAggaauggaaaTuCc 156212 156229 6350.74 19
73 ex20_c22 uCcaggAauggAaauuCc 156212 156229 6336.73 19
74 ex20_c23 ucCaggAauggAaauTcc 156212 156229 6336.72 19
---------------------------------------------------------------------
表中の配列において大文字はENA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。表中の分子量は、負イオンESI質量分析による実測値を示す。
HMVEC(皮膚微小血管内皮細胞)培養
以下のようにして、HMVECの培養を行った。
正常新生児(単一)由来皮膚微小血管内皮細胞HMVEC(CC-2813、Lonza社)を、維持培地(EGM-2MV; CC-3202、Lonza社)にて培養した。また、3.7x105/10 cm dishの播種密度で継代し、実験には6継代目までの細胞を使用した。96 well plateに2-5x103 cells/wellの密度で細胞を播種し、翌日新鮮維持培地に交換後、後述のように実施例で製造したオリゴヌクレオチドを導入した。
以下のようにして、尿中落下細胞の初代培養系を作製した。
X連鎖型アルポート症候群と遺伝子診断された患者より、尿(~100 mL程度)を採取した。遠心(1,400 rpm x 10 min)により上清を廃棄し、細胞塊をcold PBSによって2回洗浄した。細胞塊を維持培地(EGM-MV (Lonza)、20%FBS、penicillin (100 U/ml)、streptomycin (1 mg/ml)、rifampicin (8 μg/ml))にて懸濁し、37℃、5%CO2インキュベーターにて、培養した。2継代目以降は、rifampicin抜きの維持培地にて、培養した。
以下のようにして、患者由来尿中落下細胞から腎糸球体上皮細胞(ポドサイト)への分化を誘導した。
96 well plateに1x104 cells/wellの密度で細胞を播種し、翌日(分化誘導0日目)、分化誘導培地(DMEM/F12、5%FBS、1xITS supplement、12.5 μM レチノイン酸)に交換した。以後3日ごとに培地を交換した。分化誘導8日目に、後述のように実施例で製造したオリゴヌクレオチドを導入した。
以下のようにして、実施例で製造したオリゴヌクレオチドをトランスフェクションした。
下記A液とB液を作製後、混合した。
上記の混合液を5分間室温でインキュベーションした。作製した溶液を、1 wellにつき10 μLずつ加え、48時間インキュベーションした。
以下のようにしてRNAの抽出を行った。
オリゴヌクレオチドをトランスフェクション後48時間インキュベーションした細胞を、cold PBSにて1回洗浄した。SuperPrep Cell Lysis Kit & RT Kit for qPCR(TOYOBO)の細胞溶解液を、1 wellにつき50 μLずつ加え、30秒間振盪後、室温で5分間インキュベーションした。SuperPrep Cell Lysis Kit & RT Kit for qPCRの反応停止液を、1 wellにつき10 μLずつ加え、30秒間振盪後、室温で2分間インキュベーションした。合計60 μLの細胞溶解液を回収し、後述の逆転写反応を行った。
以下のようにして逆転写反応を行った。
SuperPrep Cell Lysis & RT Kit for qPCRの逆転写反応マスターミックス 32 μLと細胞溶解液 8 μLを混合した。逆転写反応を以下のように行った(37℃ 15 min、50℃ 5 min、98℃ 5 min、4℃ Hold)。最終産物は-30℃にて保存した。
1)全COL4A5発現解析用(Exon 1-7)
Forward (5’-3’): cagaggctgcggcttgctat(配列番号29)
Reverse (5’-3’): ccacgttctcccttggttcca(配列番号30)
2)Exon 20スキッピング解析用(Exon 17-22)
Forward (5’-3’): gggatggtgaaaagggccaaaaag(配列番号31)
Reverse (5’-3’): cctttgtcacctttcactccttgt(配列番号32)
3)Exon 24スキッピング解析用(Exon 21-26)
Forward (5’-3’): caaggagtgaaaggtgacaaaggt(配列番号33)
Reverse (5’-3’): ccctttaggacctggtattcctg(配列番号34)
4)内在性コントロール遺伝子(GAPDH)解析用
Forward (5’-3’): cccttcattgacctcaac(配列番号35)
Reverse (5’-3’): ttcacacccatgacgaac(配列番号36)
以下のように、免疫蛍光染色を行った。
分化ポドサイトをPBSで洗浄し、。4%パラホルムアルデヒド+4%スクロース/PBSで室温10分間、固定した。PBSで3回洗浄後、0.1%Triton X-100/PBSで室温5分間、透過処理した。PBSで3回洗浄後、5% Fetal bovine serum/PBSでブロッキングした。1次抗体(ラット 抗-COL4A5 H53抗体、重井医学研究所; 佐渡義一先生からの供与、マウス 抗-Synaptopodin抗体、新潟大学医歯学総合研究科腎研究センター;矢尾板永信先生から供与されたマウスハイブリドーマ上清を使用)を4℃で一晩反応させた。PBSで3回洗浄後、2次抗体(Alexa Fluor 488 ヤギ 抗-ラットIgG、Alexa Fluor 555 ヤギ 抗-マウスIgG)を室温1時間反応させた。PBSで3回洗浄後、核染色のため、Hoechst33342を室温10分間反応させた。封入後、蛍光顕微鏡で観察した。
図2に示したように、HMVECを用いてExon 24に対するオリゴヌクレオチド(実施例1~実施例27)のExon 24スキッピングについて調べたところ、実施例1~実施例27のオリゴヌクレオチドでExon 24スキッピングが認められた(図2A)。特に、実施例1、実施例2、実施例3、実施例13が高いスキッピング効率を示した。また、ゲルで見られたフラグメントのシークエンス解析を行ったところ、フラグメント2が、Exon 24がスキッピングしている配列であることを確認した(図2B、及び、2C)。
実施例75乃至98の化合物も実施例4と同様に合成した。実施例75乃至128の化合物の配列及びデータについて表5に記載する。
表5
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 分子量 配列番号
---------------------------------------------------------------------
75 ex24_c25 cCcTggcaaTccauCcTg 162604 162621 6278.72 37
76 ex24_c26 CcCuggcaaTccauCcTg 162604 162621 6278.72 37
77 ex24_c27 cCcuggCaaTccaTccTg 162604 162621 6278.73 37
78 ex24_c28 cCcTggcaauccauCcTg 162604 162621 6252.72 37
79 ex24_c29 cCcuggCaaucCauccTg 162604 162621 6252.72 37
80 ex24_c30 ccCuggCaaucCaucCug 162604 162621 6252.70 37
81 ex24_c31 cCcTggcaauccauCcTg 162604 162621 6252.72 37
82 ex24_c32 CccuggCaaucCauccTg 162604 162621 6252.70 37
83 ex24_c33 CcTggcaaTccauccTgT 162603 162620 6279.67 38
84 ex24_c34 CcTggcaauCcauccTgT 162603 162620 6279.71 38
85 ex24_c35 CcTggcaaucCauccTgT 162603 162620 6279.71 38
86 ex24_c36 CcTggcaauccauccTgT 162603 162620 6253.71 38
87 ex24_c37 CcuggCaauccaTccugT 162603 162620 6253.68 38
88 ex24_c38 ccTggcAauccAuccTgu 162603 162620 6225.70 38
89 ex24_c39 ccTggcaaTccaTccugT 162603 162620 6253.69 38
90 ex24_c40 cCuggCaauccaTccugT 162603 162620 6253.69 38
91 ex24_c41 CuggCaaucCaucCuguC 162602 162619 6279.70 39
92 ex24_c42 CuggCaauCcaucCuguC 162602 162619 6279.71 39
93 ex24_c43 cTggCaauCcaucCugTc 162602 162619 6279.71 39
94 ex24_c44 CuggcAauccauCcuguC 162602 162619 6239.63 39
95 ex24_c45 CuggcaAuccaTccuguC 162602 162619 6239.68 39
96 ex24_c46 cTggcaAuccaTccugTc 162602 162619 6239.68 39
97 ex24_c47 cTggcaaTccaTccugTc 162602 162619 6253.69 39
98 ex24_c48 cTggcaaTccauccTgTc 162602 162619 6253.71 39
---------------------------------------------------------------------
表中の配列において大文字はENA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。表中の分子量は、負イオンESI質量分析による実測値を示す。
実施例99乃至139の化合物も実施例4と同様に合成した。実施例99乃至139の化合物の配列及びデータについて表6に記載する。
表6
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 分子量 配列番号
---------------------------------------------------------------------
99 ex21_010 cTugGagTccTuuAucAc 156703 156686 6265.65 40
100 ex21_011 gGagTccTuuAucAccTg 156700 156683 6304.68 41
101 ex21_b08 cCuuGgaGucCuuTauCa 156704 156687 6279.59 42
102 ex21_b09 uTggAguCcuTuaTcaCc 156702 156685 6293.67 43
103 ex21_b10 uGgaGucCuuTauCacCu 156701 156684 6279.67 44
104 ex21_c01 cCuuggAguccTuuauCa 156704 156687 6241.65 42
105 ex21_c02 ccTuggAguccTuuaTca 156704 156687 6241.64 42
106 ex21_c03 ccuTggAguccTuuAuca 156704 156687 6227.63 42
107 ex21_c04 ccTuggagTccuTuaTca 156704 156687 6255.57 42
108 ex21_c05 CcTuggagTccuuuaTcA 156704 156687 6267.66 42
109 ex21_c06 CcTuggaguCcuuuaTcA 156704 156687 6267.64 42
110 ex21_c07 CcTuggagucCuuuaTcA 156704 156687 6267.65 42
111 ex21_c08 cCuTggagTccuuuAuCa 156704 156687 6267.66 42
112 ex21_c09 cCuTggaguCcuuuAuCa 156704 156687 6267.65 42
113 ex21_c10 cTuggAguccuuTaucAc 156703 156686 6227.62 40
114 ex21_c12 CuTggaguCcuuuauCaC 156703 156686 6281.64 40
115 ex21_c13 CuTggagucCuuuauCaC 156703 156686 6281.67 40
116 ex21_c14 CuTggaguccTuuauCaC 156703 156686 6281.66 40
117 ex21_c15 uTggagTccuuTaucaCc 156702 156685 6255.66 43
118 ex21_c16 uTggAguccuuuaTcaCc 156702 156685 6241.63 43
119 ex21_c17 TuggAgucCuuuaTcacC 156702 156685 6267.62 43
120 ex21_c18 TuggAguccTuuaTcacC 156702 156685 6267.58 43
121 ex21_c19 uTggAgucCuuuaTcaCc 156702 156685 6267.65 43
122 ex21_c20 uTggAguccTuuaTcaCc 156702 156685 6267.58 43
123 ex21_c21 uTggagTccTuuaTcaCc 156702 156685 6281.67 43
124 ex21_c22 uTggagTccuTuaTcaCc 156702 156685 6281.66 43
125 ex21_c23 TuggagTccTuuaTcacC 156702 156685 6282.65 43
126 ex21_c24 uggAguCcuuuAucAccu 156701 156684 6213.62 44
127 ex21_c25 uGgaguCcuuuAucacCu 156701 156684 6227.63 44
128 ex21_c26 uggagucCuuTauCacCu 156701 156684 6255.61 44
129 ex21_c27 TggAguccuuuAucAccT 156701 156684 6239.64 44
130 ex21_c28 TggAguCcuuuaTcaCcu 156701 156684 6267.65 44
131 ex21_c29 TggAguccuuuAucAccT 156701 156684 6239.63 44
132 ex21_c30 TggagucCuuTauCacCu 156701 156684 6281.67 44
133 ex21_c31 TggagTccuTuauCaccT 156701 156684 6281.67 44
134 ex21_c32 TggagTccuTuauCacCu 156701 156684 6281.68 44
135 ex21_c33 uggagTccTuTauCacCu 156701 156684 6281.67 44
136 ex21_c34 ggAgucCuuuaTcacCug 156700 156683 6280.67 41
137 ex21_c35 ggAguCcuuTaucAccTg 156700 156683 6292.66 41
138 ex21_c36 gGagucCuuuaTcaccTg 156700 156683 6280.54 41
139 ex21_c37 ggAguCcuTuaTcaCcug 156700 156683 6306.61 41
---------------------------------------------------------------------
表中の配列において大文字はENA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。表中の分子量は、負イオンESI質量分析による実測値を示す。
試験例1と同様に、HMVEC(皮膚微小血管内皮細胞)培養、尿中落下細胞初代培養系作製、腎糸球体上皮細胞(ポドサイト)への分化誘導、実施例で製造したオリゴヌクレオチドをトランスフェクション、RNA抽出、逆転写反応、PCR反応とフラグメント配列解析、及び、分化ポドサイトの免疫蛍光染色の操作を行った。
図9に示したように、HMVECを用いてExon 24に対するオリゴヌクレオチド(実施例75~実施例98)のExon 24スキッピングについて調べたところ、実施例75~実施例98のすべてのオリゴヌクレオチドにおいて、図2と同様にExon 24がスキッピングしていることを示すフラグメント2が検出され、Exon 24スキッピングが認められた。
HEK293A(ヒト胎児腎細胞由来株化細胞)培養
以下のようにして、HEK293Aの培養を行った。
HEK293A (invitrogen社)を、維持培地(DMEM、Thermo Scientific社、10% Fetal Bovine Serum(Thermo Scientific社)を含む)にて10 cm dishまたはT75フラスコで培養した。トランスフェクション直前にTrypLE Express (Thermo Scientific社)により細胞をはがして細胞懸濁液とし、後述のように実施例で製造したオリゴヌクレオチドを、リバーストランスフェクション法にて導入した。
以下のようにして、実施例で製造したオリゴヌクレオチドをトランスフェクションした。
下記A液とB液を作製後、混合した。
A液: Opti-MEM Medium (Thermofisher) 4.8 μL
実施例で製造した化合物(10 μM) 0.5 μL (最終濃度50 nM)
B液: Opti-MEM Medium 5 μL
Lipofectamine RNAiMAX (Thermofisher) 0.3 μL
上記の混合液を15分間室温でインキュベーションした。作製した溶液を、96 well plate 1 wellにつき10.6 μLずつ分注した。このplateに2x104 cells/wellの密度で細胞を播種し24時間インキュベーションした。
以下のようにしてRNAの抽出を行った。
オリゴヌクレオチドをトランスフェクション後24時間インキュベーションした細胞を、cold PBSにて1回洗浄した。SuperPrep Cell Lysis Kit & RT Kit for qPCR(TOYOBO社)の細胞溶解液を、1 wellにつき50 μLずつ加え、30秒間振盪後、室温で5分間インキュベーションした。SuperPrep Cell Lysis Kit & RT Kit for qPCRの反応停止液を、1 wellにつき10 μLずつ加え、30秒間振盪後、室温で2分間インキュベーションした。合計60 μLの細胞溶解液を回収し、後述の逆転写反応を行った。
以下のようにして逆転写反応を行った。
SuperPrep Cell Lysis & RT Kit for qPCRの逆転写反応マスターミックス 32 μLと細胞溶解液 8 μLを混合した。逆転写反応を以下のように行った(37℃ 15 min、50℃ 5 min、98℃ 5 min、4℃ Hold)。最終産物は-20℃にて保存した。
解析用プライマーは下記のように設計した。
1)COL4A5 exon 21スキッピング解析用(Exon 20-23)
Forward (5’-3’): cagttatgggtcctcctggc(配列番号45)
Reverse (5’-3’): agttgcaccagcttgtcctt(配列番号46)
2)内在性コントロール遺伝子(ACTB)解析用
Forward (5’-3’): tggcacccagcacaatgaa(配列番号47)
Reverse (5’-3’): ctaagtcatagtccgcctagaagca(配列番号48)
図13に示したように、HEK293Aを用いてExon 21に対するオリゴヌクレオチド(実施例99~実施例139)のExon 21スキッピングについて調べたところ、すべてのオリゴヌクレオチドでExon 21スキッピングが認められた(図13A)。また、ゲルから抽出したフラグメントのシークエンス解析を行い、フラグメント9がスキッピング前の配列、フラグメント10がExon 21がスキッピングしている配列であることを確認した(図13B, C)。
試験例1と同様に、HMVEC(皮膚微小血管内皮細胞)培養、尿中落下細胞初代培養系作製、腎糸球体上皮細胞(ポドサイト)への分化誘導、実施例で製造したオリゴヌクレオチドをトランスフェクション、RNA抽出、逆転写反応、PCR反応とフラグメント配列解析、及び、分化ポドサイトの免疫蛍光染色の操作を行った。
但し、解析用プライマーは下記のように設計して用いた。
Exon 21スキッピング解析用(Exon 18-24)
Forward (5’-3’): GACCTCCTGGACTTGTAATTCCTA (配列番号49)
Reverse (5’-3’): CTCCTGGAATGCCTGGTAATCCT(配列番号50)
図14Aに示したように、HMVECを用いてExon 21に対するオリゴヌクレオチド(実施例99~実施例139)のExon 21スキッピングについて調べたところ、すべてのオリゴヌクレオチドでExon 21スキッピングが認められた。また、ゲルで見られたフラグメントのシークエンス解析を行ったところ、フラグメント11が、Exon 20がスキッピングしている配列であることを確認した(図14B、及び、14C)。Exon 21に対するオリゴヌクレオチドは、最終濃度5 nMになるように用いた。
実施例140及び141の化合物も実施例4と同様に合成した。実施例140及び141の化合物の配列及びデータについて表7に記載する。
表7
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 分子量 配列番号
---------------------------------------------------------------------
140 ex21_012-2 TccTuuAucAccTgg 156696 156682 5183.57 51
141 ex21_012-4 cTuuAucAccTggAg 156694 156680 5232.56 52
---------------------------------------------------------------------
表中の配列において大文字はENA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。表中の分子量は、負イオンESI質量分析による実測値を示す。
HO-Te1s- Gm1s- Gm1s-Ae1s- Gm1s- Um1s- Cm1s- Cm1s- Um1s- Um1s- Um1s- Ae1s- Um1s- Cm1s- Ae1s- Cm1s- Cm1s- T1t-H (ex21_Lc29)(配列番号44)
実施例142の化合物は、ホスホロアミダイト法(Nucleic Acids Research, 12, 4539 (1984)を用いて合成を行った。LNA部分は、 WO99/14226に記載されたホスホロアミダイト体を用いて合成した。
本化合物は、逆相HPLC(カラム(X-Bridge C18 2.5 μm (4.6×75 mm))、A溶液:100 mMヘキサフルオロイソプロパノール(HFIP)、8 mMトリエチルアミン水溶液、B溶液:メタノール、B%:5% → 30%(20min, linear gradient);60℃;1 mL/min;260nm)で分析すると、12.87分に溶出された。化合物は負イオンESI質量分析により同定した(実測値:6173.88)。
本化合物の塩基配列は、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号156701-156684に相補的な配列である。
実施例143乃至280の化合物も実施例140と同様に合成することができる。実施例143乃至280の化合物の配列について表8乃至11に記載する。
表8
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 配列番号
---------------------------------------------------------------------
143 ex21_L010 cTugGagTccTuuAucAc 156703 156686 40
144 ex21_L011 gGagTccTuuAucAccTg 156700 156683 41
145 ex21_Lb08 cCuuGgaGucCuuTauCa 156704 156687 42
146 ex21_Lb09 uTggAguCcuTuaTcaCc 156702 156685 43
147 ex21_Lb10 uGgaGucCuuTauCacCu 156701 156684 44
148 ex21_Lc01 cCuuggAguccTuuauCa 156704 156687 42
149 ex21_Lc02 ccTuggAguccTuuaTca 156704 156687 42
150 ex21_Lc03 ccuTggAguccTuuAuca 156704 156687 42
151 ex21_Lc04 ccTuggagTccuTuaTca 156704 156687 42
152 ex21_Lc05 CcTuggagTccuuuaTcA 156704 156687 42
153 ex21_Lc06 CcTuggaguCcuuuaTcA 156704 156687 42
154 ex21_Lc07 CcTuggagucCuuuaTcA 156704 156687 42
155 ex21_Lc08 cCuTggagTccuuuAuCa 156704 156687 42
156 ex21_Lc09 cCuTggaguCcuuuAuCa 156704 156687 42
157 ex21_Lc10 cTuggAguccuuTaucAc 156703 156686 40
158 ex21_Lc12 CuTggaguCcuuuauCaC 156703 156686 40
159 ex21_Lc13 CuTggagucCuuuauCaC 156703 156686 40
160 ex21_Lc14 CuTggaguccTuuauCaC 156703 156686 40
161 ex21_Lc15 uTggagTccuuTaucaCc 156702 156685 43
162 ex21_Lc16 uTggAguccuuuaTcaCc 156702 156685 43
163 ex21_Lc17 TuggAgucCuuuaTcacC 156702 156685 43
164 ex21_Lc18 TuggAguccTuuaTcacC 156702 156685 43
165 ex21_Lc19 uTggAgucCuuuaTcaCc 156702 156685 43
166 ex21_Lc20 uTggAguccTuuaTcaCc 156702 156685 43
167 ex21_Lc21 uTggagTccTuuaTcaCc 156702 156685 43
168 ex21_Lc22 uTggagTccuTuaTcaCc 156702 156685 43
169 ex21_Lc23 TuggagTccTuuaTcacC 156702 156685 43
170 ex21_Lc24 uggAguCcuuuAucAccu 156701 156684 44
171 ex21_Lc25 uGgaguCcuuuAucacCu 156701 156684 44
172 ex21_Lc26 uggagucCuuTauCacCu 156701 156684 44
173 ex21_Lc27 TggAguccuuuAucAccT 156701 156684 44
174 ex21_Lc28 TggAguCcuuuaTcaCcu 156701 156684 44
175 ex21_Lc30 TggagucCuuTauCacCu 156701 156684 44
176 ex21_Lc31 TggagTccuTuauCaccT 156701 156684 44
177 ex21_Lc32 TggagTccuTuauCacCu 156701 156684 44
178 ex21_Lc33 uggagTccTuTauCacCu 156701 156684 44
179 ex21_Lc34 ggAgucCuuuaTcacCug 156700 156683 41
180 ex21_Lc35 ggAguCcuuTaucAccTg 156700 156683 41
181 ex21_Lc36 gGagucCuuuaTcaccTg 156700 156683 41
182 ex21_Lc37 ggAguCcuTuaTcaCcug 156700 156683 41
---------------------------------------------------------------------
表中の配列において大文字はLNA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。
表9
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 配列番号
---------------------------------------------------------------------
183 ex24_L011 cCcuGgcAauCcaTccTg 162604 162621 1
184 ex24_Lb04 cCugGcaAucCauCcuGu 162603 162620 2
185 ex24_Lb05 cTggCaaTccAucCugTc 162602 162619 3
186 ex24_Lc01 cCcTggCaaucCauCcTg 162604 162621 1
187 ex24_Lc02 cCcTggCaaucCaucCTg 162604 162621 1
188 ex24_Lc03 cCcTggCaaTcCauCcTg 162604 162621 1
189 ex24_Lc04 CcCuggCaaTcCauCcTg 162604 162621 1
190 ex24_Lc05 cCcTggCaAuCcAuCcTg 162604 162621 1
191 ex24_Lc06 cCcTggCaaTcCaTcCTg 162604 162621 1
192 ex24_Lc07 CCcTggCaaTcCaTcCTg 162604 162621 1
193 ex24_Lc08 cCcTgGcAaTcCaTcCuG 162604 162621 1
194 ex24_Lc09 cCuggCaaTcCauCcTgu 162603 162620 2
195 ex24_Lc10 CcTggCaauccauCcTgT 162603 162620 2
196 ex24_Lc11 cCuggCaaTcCauCcTgT 162603 162620 2
197 ex24_Lc12 ccTggCaAuCcAuCcTgu 162603 162620 2
198 ex24_Lc13 cCTggCaaTcCauCcTgT 162603 162620 2
199 ex24_Lc14 CcTggCaAuCcAuCcTgu 162603 162620 2
200 ex24_Lc15 cCTggCaaTcCaTCcTgT 162603 162620 2
201 ex24_Lc16 CcTggCaAuCcAuCcTgT 162603 162620 2
202 ex24_Lc17 cTggCaaTccaTcCugTc 162602 162619 3
203 ex24_Lc18 cTggCaauCcaTccTguC 162602 162619 3
204 ex24_Lc19 cTggCaaTcCaTcCugTc 162602 162619 3
205 ex24_Lc20 CTggCaauCcaTcCugTc 162602 162619 3
206 ex24_Lc21 cTggCaAuCcAuCcTgTc 162602 162619 3
207 ex24_Lc22 CTggCaaTccAucCugTC 162602 162619 3
208 ex24_Lc23 CTggCaaTcCaTcCugTC 162602 162619 3
209 ex24_Lc24 cTgGcAaTcCaTcCuGuC 162602 162619 3
---------------------------------------------------------------------
表中の配列において大文字はLNA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。
表10
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 配列番号
---------------------------------------------------------------------
210 ex24_Lc25 cCcTggcaaTccauCcTg 162604 162621 37
211 ex24_Lc26 CcCuggcaaTccauCcTg 162604 162621 37
212 ex24_Lc27 cCcuggCaaTccaTccTg 162604 162621 37
213 ex24_Lc28 cCcTggcaauccauCcTg 162604 162621 37
214 ex24_Lc29 cCcuggCaaucCauccTg 162604 162621 37
215 ex24_Lc30 ccCuggCaaucCaucCug 162604 162621 37
216 ex24_Lc31 cCcTggcaauccauCcTg 162604 162621 37
217 ex24_Lc32 CccuggCaaucCauccTg 162604 162621 37
218 ex24_Lc33 CcTggcaaTccauccTgT 162603 162620 38
219 ex24_Lc34 CcTggcaauCcauccTgT 162603 162620 38
220 ex24_Lc35 CcTggcaaucCauccTgT 162603 162620 38
221 ex24_Lc36 CcTggcaauccauccTgT 162603 162620 38
222 ex24_Lc37 CcuggCaauccaTccugT 162603 162620 38
223 ex24_Lc38 ccTggcAauccAuccTgu 162603 162620 38
224 ex24_Lc39 ccTggcaaTccaTccugT 162603 162620 38
225 ex24_Lc40 cCuggCaauccaTccugT 162603 162620 38
226 ex24_Lc41 CuggCaaucCaucCuguC 162602 162619 39
227 ex24_Lc42 CuggCaauCcaucCuguC 162602 162619 39
228 ex24_Lc43 cTggCaauCcaucCugTc 162602 162619 39
229 ex24_Lc44 CuggcAauccauCcuguC 162602 162619 39
230 ex24_Lc45 CuggcaAuccaTccuguC 162602 162619 39
231 ex24_Lc46 cTggcaAuccaTccugTc 162602 162619 39
232 ex24_Lc47 cTggcaaTccaTccugTc 162602 162619 39
233 ex24_Lc48 cTggcaaTccauccTgTc 162602 162619 39
---------------------------------------------------------------------
表中の配列において大文字はLNA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。
表11
---------------------------------------------------------------------
実施例 配列名称 配列 (5’-3’) 開始 終了 配列番号
---------------------------------------------------------------------
234 ex20_L001 uAuaGcuTacTagGagGa 156301 156318 4
235 ex20_L002 gCuuAcuAggAggAauGu 156297 156314 5
236 ex20_L022 gGagGucCagGaaTggAa 156217 156234 6
237 ex20_L023 gTccAggAauGgaAauTc 156213 156230 7
238 ex20_L024 aGgaAugGaaAuuCcaGg 156209 156226 8
239 ex20_L044 uAacTgcAgcCccTaaGa 156129 156146 9
240 ex20_Lb02 aAuaTagCuuAcuAggAg 156303 156320 10
241 ex20_Lb03 aTauAgcTuaCuaGgaGg 156302 156319 11
242 ex20_Lb04 aTagCuuAcuAggAggAa 156300 156317 12
243 ex20_Lb05 uAgcTuaCuaGgaGgaAu 156299 156316 13
244 ex20_Lb07 cTuaCuaGgaGgaAugTg 156296 156313 14
245 ex20_Lb09 uAcuAggAggAauGugAg 156294 156311 15
246 ex20_Lb10 gAggTccAggAauGgaAa 156216 156233 16
247 ex20_Lb11 aGguCcaGgaAugGaaAu 156215 156232 17
248 ex20_Lb12 gGucCagGaaTggAaaTu 156214 156231 18
249 ex20_Lb13 uCcaGgaAugGaaAuuCc 156212 156229 19
250 ex20_Lb14 cCagGaaTggAaaTucCa 156211 156228 20
251 ex20_Lb15 cAggAauGgaAauTccAg 156210 156227 21
252 ex20_Lb16 cCauAacTgcAgcCccTa 156132 156149 22
253 ex20_Lb17 cAuaAcuGcaGccCcuAa 156131 156148 23
254 ex20_Lb18 aTaaCugCagCccCuaAg 156130 156147 24
255 ex20_Lb19 aAcuGcaGccCcuAagAu 156128 156145 25
256 ex20_Lb20 aCugCagCccCuaAgaTu 156127 156144 26
257 ex20_Lb21 cTgcAgcCccTaaGauTc 156126 156143 27
258 ex20_Lc01 gTcCaggAaTggaaAuTc 156213 156230 28
259 ex20_Lc02 gTcCaggAaTggaaaTuC 156213 156230 28
260 ex20_Lc03 gTCcaggaATggaaaTTc 156213 156230 28
261 ex20_Lc04 gTccAggAaTggaAauTc 156213 156230 28
262 ex20_Lc05 gTccAggAauggaAauTc 156213 156230 28
263 ex20_Lc06 gTcCaggaaTggaaaTuC 156213 156230 28
264 ex20_Lc07 gTcCaggaaTggaaAuTc 156213 156230 28
265 ex20_Lc08 gTccAggaaTggAaaTuc 156213 156230 28
266 ex20_Lc09 gTccAggaauggaAauTc 156213 156230 28
267 ex20_Lc10 gTcCaggaauggaaaTuC 156213 156230 28
268 ex20_Lc11 gTcCaggaaTggaaauTc 156213 156230 28
269 ex20_Lc12 uCcAggAaTggAaAuuCc 156212 156229 19
270 ex20_Lc13 TccAggAauggAaaTucC 156212 156229 19
271 ex20_Lc14 uCcAggaAuggAaAuuCc 156212 156229 19
272 ex20_Lc15 TcCaggAaTggaaaTuCc 156212 156229 19
273 ex20_Lc16 uCcAggAaTggaaaTuCc 156212 156229 19
274 ex20_Lc17 uCcAggAauggAaAuuCc 156212 156229 19
275 ex20_Lc18 uCcAggaauggaAaTuCc 156212 156229 19
276 ex20_Lc19 TcCaggaaTggaaaTuCc 156212 156229 19
277 ex20_Lc20 TcCaggaaTggaaauTcC 156212 156229 19
278 ex20_Lc21 uCcAggaauggaaaTuCc 156212 156229 19
270 ex20_Lc22 uCcaggAauggAaauuCc 156212 156229 19
280 ex20_Lc23 ucCaggAauggAaauTcc 156212 156229 19
---------------------------------------------------------------------
表中の配列において大文字はLNA、小文字は2’-OMe-RNAを示す。開始および終了については、Homo sapiens collagen type IV alpha 5 chain (COL4A5)(NCBI-GenBank accession No. NG_011977)のヌクレオチド番号を示す。
本明細書で引用した全ての刊行物、特許および特許出願をそのまま参考として本明細書にとり入れるものとする。
<配列番号2>実施例2、12~19で製造したオリゴヌクレオチド(ex24_b04、ex24_c09、ex24_c10、ex24_c11、ex24_c12、ex24_c13、ex24_c14、ex24_c15、ex24_c16)のヌクレオチド配列を示す。
<配列番号3>実施例3、20~27で製造したオリゴヌクレオチド(ex24_b05、ex24_c17、ex24_c18、ex24_c19、ex24_c20、ex24_c21、ex24_c22、ex24_c23、ex24_c24)のヌクレオチド配列を示す。
<配列番号4>実施例28で製造したオリゴヌクレオチド(ex20_001)のヌクレオチド配列を示す。
<配列番号5>実施例29で製造したオリゴヌクレオチド(ex20_002)のヌクレオチド配列を示す。
<配列番号6>実施例30で製造したオリゴヌクレオチド(ex20_022)のヌクレオチド配列を示す。
<配列番号7>実施例31で製造したオリゴヌクレオチド(ex20_023)のヌクレオチド配列を示す。
<配列番号8>実施例32で製造したオリゴヌクレオチド(ex20_024)のヌクレオチド配列を示す。
<配列番号9>実施例33で製造したオリゴヌクレオチド(ex20_044)のヌクレオチド配列を示す。
<配列番号10>実施例34で製造したオリゴヌクレオチド(ex20_b02)のヌクレオチド配列を示す。
<配列番号11>実施例35で製造したオリゴヌクレオチド(ex20_b03)のヌクレオチド配列を示す。
<配列番号12>実施例36で製造したオリゴヌクレオチド(ex20_b04)のヌクレオチド配列を示す。
<配列番号13>実施例37で製造したオリゴヌクレオチド(ex20_b05)のヌクレオチド配列を示す。
<配列番号14>実施例38で製造したオリゴヌクレオチド(ex20_b07)のヌクレオチド配列を示す。
<配列番号15>実施例39で製造したオリゴヌクレオチド(ex20_b09)のヌクレオチド配列を示す。
<配列番号16>実施例40で製造したオリゴヌクレオチド(ex20_b10)のヌクレオチド配列を示す。
<配列番号17>実施例41で製造したオリゴヌクレオチド(ex20_b11)のヌクレオチド配列を示す。
<配列番号18>実施例42で製造したオリゴヌクレオチド(ex20_b12)のヌクレオチド配列を示す。
<配列番号19>実施例43、63~74で製造したオリゴヌクレオチド(ex20_b13、ex20_c12、ex20_c13、ex20_c14、ex20_c15、ex20_c16、ex20_c17、ex20_c18、ex20_c19、ex20_c20、ex20_c21、ex20_c22、ex20_c23)のヌクレオチド配列を示す。
<配列番号20>実施例44で製造したオリゴヌクレオチド(ex20_b14)のヌクレオチド配列を示す。
<配列番号21>実施例45で製造したオリゴヌクレオチド(ex20_b15)のヌクレオチド配列を示す。
<配列番号22>実施例46で製造したオリゴヌクレオチド(ex20_b16)のヌクレオチド配列を示す。
<配列番号23>実施例47で製造したオリゴヌクレオチド(ex20_b17)のヌクレオチド配列を示す。
<配列番号24>実施例48で製造したオリゴヌクレオチド(ex20_b18)のヌクレオチド配列を示す。
<配列番号25>実施例49で製造したオリゴヌクレオチド(ex20_b19)のヌクレオチド配列を示す。
<配列番号26>実施例50で製造したオリゴヌクレオチド(ex20_b20)のヌクレオチド配列を示す。
<配列番号27>実施例51で製造したオリゴヌクレオチド(ex20_b21)のヌクレオチド配列を示す。
<配列番号28>実施例52~62で製造したオリゴヌクレオチド(ex20_c01、ex20_c02、ex20_c03、ex20_c04、ex20_c05、ex20_c06、ex20_c07、ex20_c08、ex20_c09、ex20_c10、ex20_c11)のヌクレオチド配列を示す。
<配列番号29>全COL4A5発現解析用(Exon 1-7)Forward (5’-3’)プライマーのヌクレオチド配列を示す。
<配列番号30>全COL4A5発現解析用(Exon 1-7)Reverse (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号31>COL4A5 Exon 20スキッピング解析用(Exon 17-22)Forward (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号32>COL4A5 Exon 20スキッピング解析用(Exon 17-22)Reverse (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号33>COL4A5 Exon 24スキッピング解析用(Exon 21-26)Forward (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号34>COL4A5 Exon 24スキッピング解析用(Exon 21-26)Reverse (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号36>内在性コントロール遺伝子(GAPDH)解析用Forward (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号36>内在性コントロール遺伝子(GAPDH)解析用Reverse (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号37>実施例75~82で製造したオリゴヌクレオチド(ex24_c25、ex24_c26、ex24_c27、ex24_c28、ex24_c29、ex24_c30、ex24_c31、ex24_c32)のヌクレオチド配列を示す。
<配列番号38>実施例83~90で製造したオリゴヌクレオチド(ex24_c33、ex24_c34、ex24_c35、ex24_c36、ex24_c37、ex24_c38、ex24_c39)のヌクレオチド配列を示す。
<配列番号39>実施例91~98で製造したオリゴヌクレオチド(ex24_c41、ex24_c42、ex24_c43、ex24_c44、ex24_c45、ex24_c46、ex24_c47、ex24_c48)のヌクレオチド配列を示す。
<配列番号40>実施例99、113~116で製造したオリゴヌクレオチド(ex21_010、ex21_c10、ex21_c12、ex21_c13、ex21_c14)のヌクレオチド配列を示す。
<配列番号41>実施例100、136~139で製造したオリゴヌクレオチド(ex21_011、ex21_c34、ex21_c35、ex21_c36、ex21_c37)のヌクレオチド配列を示す。
<配列番号42>実施例101、104~112で製造したオリゴヌクレオチド(ex21_b08、ex21_c01、ex21_c02、ex21_c03、ex21_c04、ex21_c05、ex21_c06、ex21_c07、ex21_c08、ex21_c09)のヌクレオチド配列を示す。
<配列番号43>実施例102、117~125で製造したオリゴヌクレオチド(ex21_b09、ex21_c15、ex21_c16、ex21_c17、ex21_c18、ex21_c19、ex21_c20、ex21_c21、ex21_c22、ex21_c23)のヌクレオチド配列を示す。
<配列番号44>実施例103、126~135で製造したオリゴヌクレオチド(ex21_b10、ex21_c24、ex21_c25、ex21_c26、ex21_c27、ex21_c28、ex21_c29、ex21_c30、ex21_c31、ex21_c32、ex21_c33)のヌクレオチド配列を示す。
<配列番号45>COL4A5 Exon 21スキッピング解析用(Exon 20-23)Forward (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号46>COL4A5 Exon 21スキッピング解析用(Exon 20-23)Reverse (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号47>内在性コントロール遺伝子(ACTB)解析用Forward (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号48>内在性コントロール遺伝子(ACTB)解析用Reverse (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号49>COL4A5 Exon 21スキッピング解析用(Exon 18-24)Forward (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号50>COL4A5 Exon 21スキッピング解析用(Exon 18-24)Reverse (5’-3’) プライマーのヌクレオチド配列を示す。
<配列番号51>実施例140で製造したオリゴヌクレオチド(ex21_012-2)のヌクレオチド配列を示す。
<配列番号52>実施例141で製造したオリゴヌクレオチド(ex21_012-4)のヌクレオチド配列を示す。
Claims (13)
- COL4A5遺伝子のcDNAに相補的なヌクレオチド配列からなる塩基数15~30のオリゴヌクレオチドであって、アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのスキッピングを誘導できる前記オリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
- アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのヌクレオチド配列の一部に相補的なヌクレオチド配列からなる請求項1記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
- アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンが、COL4A5遺伝子のエクソン24、20又は21である請求項1又は2記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
- 配列番号1~28、37~41、51又は52のいずれかの配列(但し、配列中のtはuであってもよく、uはtであってもよい)の全部又は一部を含む請求項1~3のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
- オリゴヌクレオチドを構成する糖及び/又はリン酸ジエステル結合の少なくとも1個が修飾されている請求項1~4のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
- オリゴヌクレオチドを構成する糖がD-リボフラノースであり、糖の修飾がD-リボフラノースの2’位の水酸基の修飾である請求項5記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
- 糖の修飾がD-リボフラノースの2’-O-アルキル化及び/又は2’-O, 4’-C-アルキレン化である請求項6記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
- リン酸ジエステル結合の修飾がホスホロチオエートである請求項5~7のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
- 請求項1~8のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物を含む、医薬。
- 請求項1~8のいずれかに記載のオリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物を含む、アルポート症候群治療薬。
- COL4A5遺伝子のcDNAに相補的なヌクレオチド配列からなる塩基数15~30のオリゴヌクレオチドであって、アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのスキッピングを誘導できる前記オリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物を医薬的に有効な量で被験者に投与することを含む、アルポート症候群の治療方法。
- アルポート症候群の治療のための、COL4A5遺伝子のcDNAに相補的なヌクレオチド配列からなる塩基数15~30のオリゴヌクレオチドであって、アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのスキッピングを誘導できる前記オリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物の使用。
- アルポート症候群を治療する方法に使用するための、COL4A5遺伝子のcDNAに相補的なヌクレオチド配列からなる塩基数15~30のオリゴヌクレオチドであって、アルポート症候群の患者のCOL4A5遺伝子における、truncating変異を有し、かつ3の倍数の塩基数であるエクソンのスキッピングを誘導できる前記オリゴヌクレオチド、その医薬的に許容できる塩又は溶媒和物。
Priority Applications (7)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP17886231.4A EP3564373A4 (en) | 2016-12-28 | 2017-12-25 | THERAPEUTIC DRUG FOR ALPORT SYNDROME |
| CA3047894A CA3047894A1 (en) | 2016-12-28 | 2017-12-25 | Therapeutic drug for alport's syndrome |
| CN201780081708.6A CN110199023A (zh) | 2016-12-28 | 2017-12-25 | 阿尔波特氏综合症治疗药 |
| US16/467,224 US11230709B2 (en) | 2016-12-28 | 2017-12-25 | Therapeutic drug for Alport's syndrome |
| JP2018559433A JP7072748B2 (ja) | 2016-12-28 | 2017-12-25 | アルポート症候群治療薬 |
| KR1020197019324A KR20190100225A (ko) | 2016-12-28 | 2017-12-25 | 알포트 증후군 치료약 |
| AU2017386771A AU2017386771A1 (en) | 2016-12-28 | 2017-12-25 | Therapeutic drug for Alport's syndrome |
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2016254906 | 2016-12-28 | ||
| JP2016-254906 | 2016-12-28 | ||
| JP2017-077374 | 2017-04-10 | ||
| JP2017077374 | 2017-04-10 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2018123925A1 true WO2018123925A1 (ja) | 2018-07-05 |
Family
ID=62708210
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2017/046304 WO2018123925A1 (ja) | 2016-12-28 | 2017-12-25 | アルポート症候群治療薬 |
Country Status (9)
| Country | Link |
|---|---|
| US (1) | US11230709B2 (ja) |
| EP (1) | EP3564373A4 (ja) |
| JP (1) | JP7072748B2 (ja) |
| KR (1) | KR20190100225A (ja) |
| CN (1) | CN110199023A (ja) |
| AU (1) | AU2017386771A1 (ja) |
| CA (1) | CA3047894A1 (ja) |
| TW (1) | TW201828953A (ja) |
| WO (1) | WO2018123925A1 (ja) |
Citations (8)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JPH0787982A (ja) | 1993-01-29 | 1995-04-04 | Sankyo Co Ltd | 修飾オリゴデオキシリボヌクレオチド |
| WO1998054198A1 (en) | 1997-05-30 | 1998-12-03 | Hybridon, Inc. | Novel sulfur transfer reagents for oligonucleotide synthesis |
| WO1999014226A2 (en) | 1997-09-12 | 1999-03-25 | Exiqon A/S | Bi- and tri-cyclic nucleoside, nucleotide and oligonucleotide analogues |
| WO2000047599A1 (fr) | 1999-02-12 | 2000-08-17 | Sankyo Company, Limited | Nouveaux analogues de nucleosides et d'oligonucleotides |
| US6261840B1 (en) | 2000-01-18 | 2001-07-17 | Isis Pharmaceuticals, Inc. | Antisense modulation of PTP1B expression |
| WO2014109384A1 (ja) | 2013-01-10 | 2014-07-17 | 塩野義製薬株式会社 | 架橋型核酸誘導体の製造方法 |
| JP2016182122A (ja) * | 2002-11-25 | 2016-10-20 | 雅文 松尾 | mRNA前駆体のスプライシングを修飾するENA核酸医薬 |
| JP2017077374A (ja) | 2015-10-20 | 2017-04-27 | 株式会社ニューギン | 遊技機 |
Family Cites Families (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN1871033A (zh) * | 2003-08-07 | 2006-11-29 | 希龙公司 | 作为抗癌治疗靶标的三叶因子3(tff3) |
| WO2009044392A2 (en) * | 2007-10-03 | 2009-04-09 | Quark Pharmaceuticals, Inc. | Novel sirna structures |
| UA116639C2 (uk) | 2012-10-09 | 2018-04-25 | Рег'Юлес Терап'Ютікс Інк. | Способи лікування синдрому альпорта |
| MX2016015267A (es) * | 2014-05-20 | 2017-02-22 | Millennium Pharm Inc | Inhibidores de proteasoma que contienen boro para usarse despues de una terapia de cancer primaria. |
-
2017
- 2017-12-25 WO PCT/JP2017/046304 patent/WO2018123925A1/ja unknown
- 2017-12-25 KR KR1020197019324A patent/KR20190100225A/ko not_active Application Discontinuation
- 2017-12-25 EP EP17886231.4A patent/EP3564373A4/en not_active Withdrawn
- 2017-12-25 AU AU2017386771A patent/AU2017386771A1/en not_active Abandoned
- 2017-12-25 CA CA3047894A patent/CA3047894A1/en not_active Abandoned
- 2017-12-25 CN CN201780081708.6A patent/CN110199023A/zh active Pending
- 2017-12-25 JP JP2018559433A patent/JP7072748B2/ja active Active
- 2017-12-25 US US16/467,224 patent/US11230709B2/en active Active
- 2017-12-28 TW TW106146139A patent/TW201828953A/zh unknown
Patent Citations (9)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JPH0787982A (ja) | 1993-01-29 | 1995-04-04 | Sankyo Co Ltd | 修飾オリゴデオキシリボヌクレオチド |
| WO1998054198A1 (en) | 1997-05-30 | 1998-12-03 | Hybridon, Inc. | Novel sulfur transfer reagents for oligonucleotide synthesis |
| WO1999014226A2 (en) | 1997-09-12 | 1999-03-25 | Exiqon A/S | Bi- and tri-cyclic nucleoside, nucleotide and oligonucleotide analogues |
| WO2000047599A1 (fr) | 1999-02-12 | 2000-08-17 | Sankyo Company, Limited | Nouveaux analogues de nucleosides et d'oligonucleotides |
| JP2000297097A (ja) | 1999-02-12 | 2000-10-24 | Sankyo Co Ltd | 新規ヌクレオシド及びオリゴヌクレオチド類縁体 |
| US6261840B1 (en) | 2000-01-18 | 2001-07-17 | Isis Pharmaceuticals, Inc. | Antisense modulation of PTP1B expression |
| JP2016182122A (ja) * | 2002-11-25 | 2016-10-20 | 雅文 松尾 | mRNA前駆体のスプライシングを修飾するENA核酸医薬 |
| WO2014109384A1 (ja) | 2013-01-10 | 2014-07-17 | 塩野義製薬株式会社 | 架橋型核酸誘導体の製造方法 |
| JP2017077374A (ja) | 2015-10-20 | 2017-04-27 | 株式会社ニューギン | 遊技機 |
Non-Patent Citations (19)
| Title |
|---|
| "NCBI", Database accession no. NG 011977 |
| BLOMMERS ET AL., BIOCHEMISTRY, vol. 37, 1998, pages 17714 - 17725 |
| J AM SOC NEPHROL, vol. 21, 2010, pages 876 - 883 |
| J AM SOC NEPHROL., vol. 11, 2000, pages 649 - 657 |
| J. AM. CHEM. SOC., vol. 112, 1990, pages 1253 |
| JAM SOC NEPHROL, vol. 21, 2010, pages 876 - 883 |
| JAM SOC NEPHROL., vol. 11, 2000, pages 649 - 657 |
| KIDNEY INT., vol. 85, 2013, pages 1208 - 1213 |
| LESNIK, E.A. ET AL., BIOCHEMISTRY, vol. 32, 1993, pages 7832 - 7838 |
| MARTIN, P., HELV. CHIM. ACTA., vol. 78, 1995, pages 486 - 504 |
| NEPHROL DIAL TRANSPLANT, vol. 17, 2002, pages 1218 - 1227 |
| NOZU, KANDAI ET AL.: "Molecular therapeutic strategy using exon skipping therapy for alport syndrome", THE MOTHER AND CHILD HEALTH FOUNDATION: RESEARCH REPORT FOR PROMOTING PEDIATRICS, vol. 26, 2015, pages 16 - 17, XP009515418 * |
| NUCLEIC ACIDS RESEARCH, vol. 12, 1984, pages 4539 |
| See also references of EP3564373A4 |
| SETH, P.P. ET AL., J. ORG. CHEM, vol. 75, 2010, pages 1569 - 1581 |
| SHONO, AKEMI ET AL.: "Alport syndrome-specific treatment method by exon skipping therapy", JAPANESE JOURNAL OF PEDIATRIC NEPHROLOGY, vol. 30, no. 1, May 2017 (2017-05-01), pages 102, XP009515331 * |
| TETRAHEDRON LETTERS, vol. 32, 1991, pages 3005 |
| WANG, G. ET AL., TETRAHEDRON, vol. 55, 1999, pages 7707 - 774 |
| YAHARA, A. ET AL., CHEMBIOCHEM, vol. 13, 2012, pages 2513 - 2516 |
Also Published As
| Publication number | Publication date |
|---|---|
| US11230709B2 (en) | 2022-01-25 |
| US20200080080A1 (en) | 2020-03-12 |
| EP3564373A1 (en) | 2019-11-06 |
| TW201828953A (zh) | 2018-08-16 |
| KR20190100225A (ko) | 2019-08-28 |
| CN110199023A (zh) | 2019-09-03 |
| EP3564373A4 (en) | 2020-12-16 |
| JPWO2018123925A1 (ja) | 2019-10-31 |
| JP7072748B2 (ja) | 2022-05-23 |
| CA3047894A1 (en) | 2018-07-05 |
| AU2017386771A1 (en) | 2019-06-20 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP7007304B2 (ja) | B型肝炎感染症治療用のPAPD5又はPAPD7のmRNA減少用核酸分子 | |
| EP2386636B1 (en) | Ena nucleic acid drugs modifying splicing in mrna precursor | |
| CN106470688B (zh) | 寡聚物和寡聚物缀合物 | |
| KR101985382B1 (ko) | 톨-유사 수용체 기반 면역 반응을 조절하기 위한 면역 조절 올리고뉴클레오타이드(iro) 화합물 | |
| CA3099522A1 (en) | Gapmers and methods of using the same for treatment of muscular dystrophy | |
| WO2018123925A1 (ja) | アルポート症候群治療薬 | |
| US20230119360A1 (en) | Pharmaceutical combination of a therapeutic oligonucleotide targeting hbv and a tlr7 agonist for treatment of hbv | |
| KR20210033004A (ko) | Rtel1의 발현을 조절하기 위한 올리고뉴클레오티드 | |
| WO2021010301A1 (ja) | DUX4 pre-mRNAのスプライシングを変化させるアンチセンスオリゴヌクレオチド | |
| WO2019240164A1 (ja) | 心筋障害治療薬 | |
| JP6029147B2 (ja) | miRNA機能抑制用オリゴヌクレオチド誘導体 | |
| EP4081217A1 (en) | Pharmaceutical combination of antiviral agents targeting hbv and/or an immune modulator for treatment of hbv | |
| CN114574495A (zh) | 核苷类衍生物改性的核酸适体r50 | |
| EP4067489A1 (en) | Method for reducing toxicity of antisense nucleic acids | |
| JPWO2019009299A1 (ja) | α−シヌクレイン発現抑制するENAアンチセンスオリゴヌクレオチド | |
| WO2019111791A1 (ja) | ジストロフィン遺伝子のイントロンリテンションを解消するアンチセンスオリゴヌクレオチド | |
| JP6536911B2 (ja) | CD44遺伝子のバリアントエクソンのスキッピングを誘導し、正常型CD44mRNAの発現を増加させる核酸医薬 | |
| TW202246500A (zh) | 用於抑制 rtel1 表現之增強型寡核苷酸 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 17886231 Country of ref document: EP Kind code of ref document: A1 |
|
| ENP | Entry into the national phase |
Ref document number: 2018559433 Country of ref document: JP Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 2017386771 Country of ref document: AU Date of ref document: 20171225 Kind code of ref document: A Ref document number: 3047894 Country of ref document: CA |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 20197019324 Country of ref document: KR Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 2017886231 Country of ref document: EP Effective date: 20190729 |