JP4177118B2 - 増幅方法 - Google Patents
増幅方法 Download PDFInfo
- Publication number
- JP4177118B2 JP4177118B2 JP2002585667A JP2002585667A JP4177118B2 JP 4177118 B2 JP4177118 B2 JP 4177118B2 JP 2002585667 A JP2002585667 A JP 2002585667A JP 2002585667 A JP2002585667 A JP 2002585667A JP 4177118 B2 JP4177118 B2 JP 4177118B2
- Authority
- JP
- Japan
- Prior art keywords
- ppase
- pyrophosphate
- reaction mixture
- pcr
- polymerase
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Expired - Fee Related
Links
- 238000000034 method Methods 0.000 title claims description 78
- 230000003321 amplification Effects 0.000 title claims description 50
- 238000003199 nucleic acid amplification method Methods 0.000 title claims description 50
- 238000003752 polymerase chain reaction Methods 0.000 claims description 75
- 238000006243 chemical reaction Methods 0.000 claims description 67
- 235000011180 diphosphates Nutrition 0.000 claims description 43
- 239000011541 reaction mixture Substances 0.000 claims description 42
- XPPKVPWEQAFLFU-UHFFFAOYSA-J diphosphate(4-) Chemical compound [O-]P([O-])(=O)OP([O-])([O-])=O XPPKVPWEQAFLFU-UHFFFAOYSA-J 0.000 claims description 40
- 108010006785 Taq Polymerase Proteins 0.000 claims description 34
- 108010014303 DNA-directed DNA polymerase Proteins 0.000 claims description 27
- 102000016928 DNA-directed DNA polymerase Human genes 0.000 claims description 27
- 102000004190 Enzymes Human genes 0.000 claims description 27
- 108090000790 Enzymes Proteins 0.000 claims description 27
- 108010009413 Pyrophosphatases Proteins 0.000 claims description 24
- 102000009609 Pyrophosphatases Human genes 0.000 claims description 22
- 239000000523 sample Substances 0.000 claims description 22
- 241000567139 Aeropyrum pernix Species 0.000 claims description 18
- 239000003153 chemical reaction reagent Substances 0.000 claims description 17
- 108010009595 Inorganic Pyrophosphatase Proteins 0.000 claims description 16
- 102000009617 Inorganic Pyrophosphatase Human genes 0.000 claims description 16
- 125000003275 alpha amino acid group Chemical group 0.000 claims description 16
- 150000007523 nucleic acids Chemical class 0.000 claims description 14
- 108091033319 polynucleotide Proteins 0.000 claims description 13
- 102000040430 polynucleotide Human genes 0.000 claims description 13
- 239000002157 polynucleotide Substances 0.000 claims description 13
- 239000000203 mixture Substances 0.000 claims description 12
- 239000012634 fragment Substances 0.000 claims description 10
- 102000039446 nucleic acids Human genes 0.000 claims description 10
- 108020004707 nucleic acids Proteins 0.000 claims description 10
- 239000000138 intercalating agent Substances 0.000 claims description 8
- 239000002773 nucleotide Substances 0.000 claims description 8
- 125000003729 nucleotide group Chemical group 0.000 claims description 8
- 239000000975 dye Substances 0.000 claims description 7
- 108010001244 Tli polymerase Proteins 0.000 claims description 5
- 230000008569 process Effects 0.000 claims description 4
- FQENQNTWSFEDLI-UHFFFAOYSA-J sodium diphosphate Chemical group [Na+].[Na+].[Na+].[Na+].[O-]P([O-])(=O)OP([O-])([O-])=O FQENQNTWSFEDLI-UHFFFAOYSA-J 0.000 claims description 4
- 235000019818 tetrasodium diphosphate Nutrition 0.000 claims description 4
- 241000617156 archaeon Species 0.000 claims description 3
- 239000007850 fluorescent dye Substances 0.000 claims description 3
- 241000205156 Pyrococcus furiosus Species 0.000 claims description 2
- 241000589499 Thermus thermophilus Species 0.000 claims description 2
- 229910052783 alkali metal Inorganic materials 0.000 claims description 2
- -1 alkali metal pyrophosphate Chemical class 0.000 claims description 2
- 230000015572 biosynthetic process Effects 0.000 claims description 2
- 238000001215 fluorescent labelling Methods 0.000 claims 2
- 238000012408 PCR amplification Methods 0.000 claims 1
- 101900314195 Thermococcus litoralis Inorganic pyrophosphatase Proteins 0.000 claims 1
- 241000589497 Thermus sp. Species 0.000 claims 1
- KWIUHFFTVRNATP-UHFFFAOYSA-N Betaine Natural products C[N+](C)(C)CC([O-])=O KWIUHFFTVRNATP-UHFFFAOYSA-N 0.000 description 55
- KWIUHFFTVRNATP-UHFFFAOYSA-O N,N,N-trimethylglycinium Chemical compound C[N+](C)(C)CC(O)=O KWIUHFFTVRNATP-UHFFFAOYSA-O 0.000 description 55
- 229960003237 betaine Drugs 0.000 description 55
- 239000000047 product Substances 0.000 description 30
- 238000003556 assay Methods 0.000 description 28
- 108020004414 DNA Proteins 0.000 description 20
- 230000000694 effects Effects 0.000 description 14
- 108010064733 Angiotensins Proteins 0.000 description 10
- 239000002585 base Substances 0.000 description 10
- 102000015427 Angiotensins Human genes 0.000 description 9
- 238000002866 fluorescence resonance energy transfer Methods 0.000 description 9
- 238000001514 detection method Methods 0.000 description 8
- 108090000623 proteins and genes Proteins 0.000 description 8
- 239000013598 vector Substances 0.000 description 8
- 108010020713 Tth polymerase Proteins 0.000 description 7
- KCXVZYZYPLLWCC-UHFFFAOYSA-N EDTA Chemical compound OC(=O)CN(CC(O)=O)CCN(CC(O)=O)CC(O)=O KCXVZYZYPLLWCC-UHFFFAOYSA-N 0.000 description 6
- LFQSCWFLJHTTHZ-UHFFFAOYSA-N Ethanol Chemical compound CCO LFQSCWFLJHTTHZ-UHFFFAOYSA-N 0.000 description 6
- 239000007983 Tris buffer Substances 0.000 description 6
- 238000002474 experimental method Methods 0.000 description 6
- LENZDBCJOHFCAS-UHFFFAOYSA-N tris Chemical compound OCC(N)(CO)CO LENZDBCJOHFCAS-UHFFFAOYSA-N 0.000 description 6
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 description 5
- 229920001817 Agar Polymers 0.000 description 5
- 102000053602 DNA Human genes 0.000 description 5
- 108010002747 Pfu DNA polymerase Proteins 0.000 description 5
- 150000001413 amino acids Chemical class 0.000 description 5
- 230000000295 complement effect Effects 0.000 description 5
- 238000004925 denaturation Methods 0.000 description 5
- 230000036425 denaturation Effects 0.000 description 5
- 230000001915 proofreading effect Effects 0.000 description 5
- 102000004169 proteins and genes Human genes 0.000 description 5
- 239000011734 sodium Substances 0.000 description 5
- 238000006467 substitution reaction Methods 0.000 description 5
- QKNYBSVHEMOAJP-UHFFFAOYSA-N 2-amino-2-(hydroxymethyl)propane-1,3-diol;hydron;chloride Chemical compound Cl.OCC(N)(CO)CO QKNYBSVHEMOAJP-UHFFFAOYSA-N 0.000 description 4
- 108091034117 Oligonucleotide Proteins 0.000 description 4
- FAPWRFPIFSIZLT-UHFFFAOYSA-M Sodium chloride Chemical compound [Na+].[Cl-] FAPWRFPIFSIZLT-UHFFFAOYSA-M 0.000 description 4
- 239000008272 agar Substances 0.000 description 4
- 238000011534 incubation Methods 0.000 description 4
- 229930027917 kanamycin Natural products 0.000 description 4
- SBUJHOSQTJFQJX-NOAMYHISSA-N kanamycin Chemical compound O[C@@H]1[C@@H](O)[C@H](O)[C@@H](CN)O[C@@H]1O[C@H]1[C@H](O)[C@@H](O[C@@H]2[C@@H]([C@@H](N)[C@H](O)[C@@H](CO)O2)O)[C@H](N)C[C@@H]1N SBUJHOSQTJFQJX-NOAMYHISSA-N 0.000 description 4
- 229960000318 kanamycin Drugs 0.000 description 4
- 229930182823 kanamycin A Natural products 0.000 description 4
- 239000011777 magnesium Substances 0.000 description 4
- 238000012546 transfer Methods 0.000 description 4
- 238000001712 DNA sequencing Methods 0.000 description 3
- PEDCQBHIVMGVHV-UHFFFAOYSA-N Glycerine Chemical compound OCC(O)CO PEDCQBHIVMGVHV-UHFFFAOYSA-N 0.000 description 3
- 239000006137 Luria-Bertani broth Substances 0.000 description 3
- JLVVSXFLKOJNIY-UHFFFAOYSA-N Magnesium ion Chemical compound [Mg+2] JLVVSXFLKOJNIY-UHFFFAOYSA-N 0.000 description 3
- 108091028043 Nucleic acid sequence Proteins 0.000 description 3
- 238000002944 PCR assay Methods 0.000 description 3
- 229920001213 Polysorbate 20 Polymers 0.000 description 3
- 239000000872 buffer Substances 0.000 description 3
- 230000006862 enzymatic digestion Effects 0.000 description 3
- 239000000499 gel Substances 0.000 description 3
- 238000009396 hybridization Methods 0.000 description 3
- 230000007062 hydrolysis Effects 0.000 description 3
- 238000006460 hydrolysis reaction Methods 0.000 description 3
- BPHPUYQFMNQIOC-NXRLNHOXSA-N isopropyl beta-D-thiogalactopyranoside Chemical compound CC(C)S[C@@H]1O[C@H](CO)[C@H](O)[C@H](O)[C@H]1O BPHPUYQFMNQIOC-NXRLNHOXSA-N 0.000 description 3
- 238000002372 labelling Methods 0.000 description 3
- 229910001425 magnesium ion Inorganic materials 0.000 description 3
- 238000004519 manufacturing process Methods 0.000 description 3
- 239000000256 polyoxyethylene sorbitan monolaurate Substances 0.000 description 3
- 235000010486 polyoxyethylene sorbitan monolaurate Nutrition 0.000 description 3
- 239000011535 reaction buffer Substances 0.000 description 3
- 238000003860 storage Methods 0.000 description 3
- BHNQPLPANNDEGL-UHFFFAOYSA-N 2-(4-octylphenoxy)ethanol Chemical compound CCCCCCCCC1=CC=C(OCCO)C=C1 BHNQPLPANNDEGL-UHFFFAOYSA-N 0.000 description 2
- 241000567147 Aeropyrum Species 0.000 description 2
- 230000004544 DNA amplification Effects 0.000 description 2
- 241000588724 Escherichia coli Species 0.000 description 2
- FFEARJCKVFRZRR-BYPYZUCNSA-N L-methionine Chemical compound CSCC[C@H](N)C(O)=O FFEARJCKVFRZRR-BYPYZUCNSA-N 0.000 description 2
- SEQKRHFRPICQDD-UHFFFAOYSA-N N-tris(hydroxymethyl)methylglycine Chemical compound OCC(CO)(CO)[NH2+]CC([O-])=O SEQKRHFRPICQDD-UHFFFAOYSA-N 0.000 description 2
- 101100244014 Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) ppi-5 gene Proteins 0.000 description 2
- 108020004711 Nucleic Acid Probes Proteins 0.000 description 2
- 239000012807 PCR reagent Substances 0.000 description 2
- ISWSIDIOOBJBQZ-UHFFFAOYSA-N Phenol Chemical compound OC1=CC=CC=C1 ISWSIDIOOBJBQZ-UHFFFAOYSA-N 0.000 description 2
- 229920002684 Sepharose Polymers 0.000 description 2
- 241000205188 Thermococcus Species 0.000 description 2
- 241000205180 Thermococcus litoralis Species 0.000 description 2
- 241000589596 Thermus Species 0.000 description 2
- 229910052770 Uranium Inorganic materials 0.000 description 2
- JLCPHMBAVCMARE-UHFFFAOYSA-N [3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-hydroxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methyl [5-(6-aminopurin-9-yl)-2-(hydroxymethyl)oxolan-3-yl] hydrogen phosphate Polymers Cc1cn(C2CC(OP(O)(=O)OCC3OC(CC3OP(O)(=O)OCC3OC(CC3O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c3nc(N)[nH]c4=O)C(COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3CO)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cc(C)c(=O)[nH]c3=O)n3cc(C)c(=O)[nH]c3=O)n3ccc(N)nc3=O)n3cc(C)c(=O)[nH]c3=O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)O2)c(=O)[nH]c1=O JLCPHMBAVCMARE-UHFFFAOYSA-N 0.000 description 2
- 150000003863 ammonium salts Chemical class 0.000 description 2
- 238000000137 annealing Methods 0.000 description 2
- 230000015556 catabolic process Effects 0.000 description 2
- 238000006731 degradation reaction Methods 0.000 description 2
- 238000005516 engineering process Methods 0.000 description 2
- 238000002955 isolation Methods 0.000 description 2
- 230000007246 mechanism Effects 0.000 description 2
- 229930182817 methionine Natural products 0.000 description 2
- 230000000813 microbial effect Effects 0.000 description 2
- 238000012986 modification Methods 0.000 description 2
- 230000004048 modification Effects 0.000 description 2
- 239000002853 nucleic acid probe Substances 0.000 description 2
- 239000002777 nucleoside Substances 0.000 description 2
- NBIIXXVUZAFLBC-UHFFFAOYSA-K phosphate Chemical compound [O-]P([O-])([O-])=O NBIIXXVUZAFLBC-UHFFFAOYSA-K 0.000 description 2
- 238000011533 pre-incubation Methods 0.000 description 2
- 108091008146 restriction endonucleases Proteins 0.000 description 2
- 238000003757 reverse transcription PCR Methods 0.000 description 2
- 238000007086 side reaction Methods 0.000 description 2
- 239000011780 sodium chloride Substances 0.000 description 2
- VZWGHDYJGOMEKT-UHFFFAOYSA-J sodium pyrophosphate decahydrate Chemical compound O.O.O.O.O.O.O.O.O.O.[Na+].[Na+].[Na+].[Na+].[O-]P([O-])(=O)OP([O-])([O-])=O VZWGHDYJGOMEKT-UHFFFAOYSA-J 0.000 description 2
- 239000000126 substance Substances 0.000 description 2
- RYCLIXPGLDDLTM-UHFFFAOYSA-J tetrapotassium;phosphonato phosphate Chemical compound [K+].[K+].[K+].[K+].[O-]P([O-])(=O)OP([O-])([O-])=O RYCLIXPGLDDLTM-UHFFFAOYSA-J 0.000 description 2
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 2
- SNBCLPGEMZEWLU-QXFUBDJGSA-N 2-chloro-n-[[(2r,3s,5r)-3-hydroxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methyl]acetamide Chemical compound O=C1NC(=O)C(C)=CN1[C@@H]1O[C@H](CNC(=O)CCl)[C@@H](O)C1 SNBCLPGEMZEWLU-QXFUBDJGSA-N 0.000 description 1
- 108010085238 Actins Proteins 0.000 description 1
- 101900352645 Aeropyrum pernix Inorganic pyrophosphatase Proteins 0.000 description 1
- 241000423335 Aeropyrum pernix K1 Species 0.000 description 1
- 101000994875 Arabidopsis thaliana Soluble inorganic pyrophosphatase 3 Proteins 0.000 description 1
- 101000994873 Arabidopsis thaliana Soluble inorganic pyrophosphatase 5 Proteins 0.000 description 1
- 241000894006 Bacteria Species 0.000 description 1
- 108020004635 Complementary DNA Proteins 0.000 description 1
- 102000012410 DNA Ligases Human genes 0.000 description 1
- 108010061982 DNA Ligases Proteins 0.000 description 1
- 230000004568 DNA-binding Effects 0.000 description 1
- AHCYMLUZIRLXAA-SHYZEUOFSA-N Deoxyuridine 5'-triphosphate Chemical compound O1[C@H](COP(O)(=O)OP(O)(=O)OP(O)(O)=O)[C@@H](O)C[C@@H]1N1C(=O)NC(=O)C=C1 AHCYMLUZIRLXAA-SHYZEUOFSA-N 0.000 description 1
- WQZGKKKJIJFFOK-GASJEMHNSA-N Glucose Natural products OC[C@H]1OC(O)[C@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-GASJEMHNSA-N 0.000 description 1
- 241000282412 Homo Species 0.000 description 1
- DGAQECJNVWCQMB-PUAWFVPOSA-M Ilexoside XXIX Chemical compound C[C@@H]1CC[C@@]2(CC[C@@]3(C(=CC[C@H]4[C@]3(CC[C@@H]5[C@@]4(CC[C@@H](C5(C)C)OS(=O)(=O)[O-])C)C)[C@@H]2[C@]1(C)O)C)C(=O)O[C@H]6[C@@H]([C@H]([C@@H]([C@H](O6)CO)O)O)O.[Na+] DGAQECJNVWCQMB-PUAWFVPOSA-M 0.000 description 1
- 241001465754 Metazoa Species 0.000 description 1
- VCUFZILGIRCDQQ-KRWDZBQOSA-N N-[[(5S)-2-oxo-3-(2-oxo-3H-1,3-benzoxazol-6-yl)-1,3-oxazolidin-5-yl]methyl]-2-[[3-(trifluoromethoxy)phenyl]methylamino]pyrimidine-5-carboxamide Chemical compound O=C1O[C@H](CN1C1=CC2=C(NC(O2)=O)C=C1)CNC(=O)C=1C=NC(=NC=1)NCC1=CC(=CC=C1)OC(F)(F)F VCUFZILGIRCDQQ-KRWDZBQOSA-N 0.000 description 1
- 229910019142 PO4 Inorganic materials 0.000 description 1
- VMHLLURERBWHNL-UHFFFAOYSA-M Sodium acetate Chemical compound [Na+].CC([O-])=O VMHLLURERBWHNL-UHFFFAOYSA-M 0.000 description 1
- 241000205101 Sulfolobus Species 0.000 description 1
- 241000205098 Sulfolobus acidocaldarius Species 0.000 description 1
- UZMAPBJVXOGOFT-UHFFFAOYSA-N Syringetin Natural products COC1=C(O)C(OC)=CC(C2=C(C(=O)C3=C(O)C=C(O)C=C3O2)O)=C1 UZMAPBJVXOGOFT-UHFFFAOYSA-N 0.000 description 1
- 241000589500 Thermus aquaticus Species 0.000 description 1
- 241000557720 Thermus brockianus Species 0.000 description 1
- 239000007997 Tricine buffer Substances 0.000 description 1
- NFOVOLLZYFKJCK-UHFFFAOYSA-J [Fe+4].[O-]P([O-])(=O)OP([O-])([O-])=O Chemical class [Fe+4].[O-]P([O-])(=O)OP([O-])([O-])=O NFOVOLLZYFKJCK-UHFFFAOYSA-J 0.000 description 1
- 239000002253 acid Substances 0.000 description 1
- 230000009471 action Effects 0.000 description 1
- BFNBIHQBYMNNAN-UHFFFAOYSA-N ammonium sulfate Chemical compound N.N.OS(O)(=O)=O BFNBIHQBYMNNAN-UHFFFAOYSA-N 0.000 description 1
- 229910052921 ammonium sulfate Inorganic materials 0.000 description 1
- 235000011130 ammonium sulphate Nutrition 0.000 description 1
- 239000012491 analyte Substances 0.000 description 1
- 230000004888 barrier function Effects 0.000 description 1
- 230000008901 benefit Effects 0.000 description 1
- 239000011230 binding agent Substances 0.000 description 1
- 230000003115 biocidal effect Effects 0.000 description 1
- 230000004071 biological effect Effects 0.000 description 1
- 239000002981 blocking agent Substances 0.000 description 1
- 239000007853 buffer solution Substances 0.000 description 1
- 239000006227 byproduct Substances 0.000 description 1
- 238000010804 cDNA synthesis Methods 0.000 description 1
- 239000003054 catalyst Substances 0.000 description 1
- 238000005119 centrifugation Methods 0.000 description 1
- 238000003776 cleavage reaction Methods 0.000 description 1
- 238000010367 cloning Methods 0.000 description 1
- 238000007398 colorimetric assay Methods 0.000 description 1
- 238000004440 column chromatography Methods 0.000 description 1
- 239000013065 commercial product Substances 0.000 description 1
- 230000000052 comparative effect Effects 0.000 description 1
- 239000002299 complementary DNA Substances 0.000 description 1
- 239000002131 composite material Substances 0.000 description 1
- 150000001875 compounds Chemical class 0.000 description 1
- 238000011109 contamination Methods 0.000 description 1
- 238000007796 conventional method Methods 0.000 description 1
- NKLPQNGYXWVELD-UHFFFAOYSA-M coomassie brilliant blue Chemical compound [Na+].C1=CC(OCC)=CC=C1NC1=CC=C(C(=C2C=CC(C=C2)=[N+](CC)CC=2C=C(C=CC=2)S([O-])(=O)=O)C=2C=CC(=CC=2)N(CC)CC=2C=C(C=CC=2)S([O-])(=O)=O)C=C1 NKLPQNGYXWVELD-UHFFFAOYSA-M 0.000 description 1
- 230000001186 cumulative effect Effects 0.000 description 1
- SUYVUBYJARFZHO-RRKCRQDMSA-N dATP Chemical compound C1=NC=2C(N)=NC=NC=2N1[C@H]1C[C@H](O)[C@@H](COP(O)(=O)OP(O)(=O)OP(O)(O)=O)O1 SUYVUBYJARFZHO-RRKCRQDMSA-N 0.000 description 1
- SUYVUBYJARFZHO-UHFFFAOYSA-N dATP Natural products C1=NC=2C(N)=NC=NC=2N1C1CC(O)C(COP(O)(=O)OP(O)(=O)OP(O)(O)=O)O1 SUYVUBYJARFZHO-UHFFFAOYSA-N 0.000 description 1
- RGWHQCVHVJXOKC-SHYZEUOFSA-J dCTP(4-) Chemical compound O=C1N=C(N)C=CN1[C@@H]1O[C@H](COP([O-])(=O)OP([O-])(=O)OP([O-])([O-])=O)[C@@H](O)C1 RGWHQCVHVJXOKC-SHYZEUOFSA-J 0.000 description 1
- HAAZLUGHYHWQIW-KVQBGUIXSA-N dGTP Chemical compound C1=NC=2C(=O)NC(N)=NC=2N1[C@H]1C[C@H](O)[C@@H](COP(O)(=O)OP(O)(=O)OP(O)(O)=O)O1 HAAZLUGHYHWQIW-KVQBGUIXSA-N 0.000 description 1
- 230000001934 delay Effects 0.000 description 1
- 238000012217 deletion Methods 0.000 description 1
- 230000037430 deletion Effects 0.000 description 1
- 230000001419 dependent effect Effects 0.000 description 1
- 239000008121 dextrose Substances 0.000 description 1
- 238000003745 diagnosis Methods 0.000 description 1
- KCFYHBSOLOXZIF-UHFFFAOYSA-N dihydrochrysin Natural products COC1=C(O)C(OC)=CC(C2OC3=CC(O)=CC(O)=C3C(=O)C2)=C1 KCFYHBSOLOXZIF-UHFFFAOYSA-N 0.000 description 1
- 238000010790 dilution Methods 0.000 description 1
- 239000012895 dilution Substances 0.000 description 1
- 201000010099 disease Diseases 0.000 description 1
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 1
- 235000019820 disodium diphosphate Nutrition 0.000 description 1
- GYQBBRRVRKFJRG-UHFFFAOYSA-L disodium pyrophosphate Chemical compound [Na+].[Na+].OP([O-])(=O)OP(O)([O-])=O GYQBBRRVRKFJRG-UHFFFAOYSA-L 0.000 description 1
- 229940038485 disodium pyrophosphate Drugs 0.000 description 1
- VHJLVAABSRFDPM-QWWZWVQMSA-N dithiothreitol Chemical compound SC[C@@H](O)[C@H](O)CS VHJLVAABSRFDPM-QWWZWVQMSA-N 0.000 description 1
- 238000001962 electrophoresis Methods 0.000 description 1
- ZMMJGEGLRURXTF-UHFFFAOYSA-N ethidium bromide Chemical compound [Br-].C12=CC(N)=CC=C2C2=CC=C(N)C=C2[N+](CC)=C1C1=CC=CC=C1 ZMMJGEGLRURXTF-UHFFFAOYSA-N 0.000 description 1
- 229960005542 ethidium bromide Drugs 0.000 description 1
- 238000011156 evaluation Methods 0.000 description 1
- 238000000695 excitation spectrum Methods 0.000 description 1
- 239000011706 ferric diphosphate Substances 0.000 description 1
- 235000007144 ferric diphosphate Nutrition 0.000 description 1
- CADNYOZXMIKYPR-UHFFFAOYSA-B ferric pyrophosphate Chemical compound [Fe+3].[Fe+3].[Fe+3].[Fe+3].[O-]P([O-])(=O)OP([O-])([O-])=O.[O-]P([O-])(=O)OP([O-])([O-])=O.[O-]P([O-])(=O)OP([O-])([O-])=O CADNYOZXMIKYPR-UHFFFAOYSA-B 0.000 description 1
- 229940036404 ferric pyrophosphate Drugs 0.000 description 1
- 238000009472 formulation Methods 0.000 description 1
- 238000012252 genetic analysis Methods 0.000 description 1
- 238000003205 genotyping method Methods 0.000 description 1
- PCHJSUWPFVWCPO-UHFFFAOYSA-N gold Chemical compound [Au] PCHJSUWPFVWCPO-UHFFFAOYSA-N 0.000 description 1
- 239000010931 gold Substances 0.000 description 1
- 229910052737 gold Inorganic materials 0.000 description 1
- 230000036541 health Effects 0.000 description 1
- 238000010438 heat treatment Methods 0.000 description 1
- 238000007849 hot-start PCR Methods 0.000 description 1
- 229910052588 hydroxylapatite Inorganic materials 0.000 description 1
- 238000000338 in vitro Methods 0.000 description 1
- 238000011065 in-situ storage Methods 0.000 description 1
- 239000003112 inhibitor Substances 0.000 description 1
- 230000005764 inhibitory process Effects 0.000 description 1
- 229910052816 inorganic phosphate Inorganic materials 0.000 description 1
- XEEYBQQBJWHFJM-UHFFFAOYSA-N iron Substances [Fe] XEEYBQQBJWHFJM-UHFFFAOYSA-N 0.000 description 1
- 239000011159 matrix material Substances 0.000 description 1
- 229910052751 metal Inorganic materials 0.000 description 1
- 239000002184 metal Substances 0.000 description 1
- 150000002739 metals Chemical class 0.000 description 1
- 238000002156 mixing Methods 0.000 description 1
- 238000012544 monitoring process Methods 0.000 description 1
- 150000004712 monophosphates Chemical class 0.000 description 1
- 229910052755 nonmetal Inorganic materials 0.000 description 1
- 150000002843 nonmetals Chemical class 0.000 description 1
- 150000003833 nucleoside derivatives Chemical class 0.000 description 1
- 125000003835 nucleoside group Chemical group 0.000 description 1
- 244000052769 pathogen Species 0.000 description 1
- 230000001717 pathogenic effect Effects 0.000 description 1
- 239000008188 pellet Substances 0.000 description 1
- XYJRXVWERLGGKC-UHFFFAOYSA-D pentacalcium;hydroxide;triphosphate Chemical compound [OH-].[Ca+2].[Ca+2].[Ca+2].[Ca+2].[Ca+2].[O-]P([O-])([O-])=O.[O-]P([O-])([O-])=O.[O-]P([O-])([O-])=O XYJRXVWERLGGKC-UHFFFAOYSA-D 0.000 description 1
- 125000001997 phenyl group Chemical group [H]C1=C([H])C([H])=C(*)C([H])=C1[H] 0.000 description 1
- 239000010452 phosphate Substances 0.000 description 1
- WRMXOVHLRUVREB-UHFFFAOYSA-N phosphono phosphate;tributylazanium Chemical compound OP(O)(=O)OP([O-])([O-])=O.CCCC[NH+](CCCC)CCCC.CCCC[NH+](CCCC)CCCC WRMXOVHLRUVREB-UHFFFAOYSA-N 0.000 description 1
- 230000026731 phosphorylation Effects 0.000 description 1
- 238000006366 phosphorylation reaction Methods 0.000 description 1
- 239000013600 plasmid vector Substances 0.000 description 1
- 229920002401 polyacrylamide Polymers 0.000 description 1
- 102000054765 polymorphisms of proteins Human genes 0.000 description 1
- 229920001184 polypeptide Polymers 0.000 description 1
- 238000004321 preservation Methods 0.000 description 1
- 102000004196 processed proteins & peptides Human genes 0.000 description 1
- 108090000765 processed proteins & peptides Proteins 0.000 description 1
- 238000000746 purification Methods 0.000 description 1
- 230000001698 pyrogenic effect Effects 0.000 description 1
- 238000010791 quenching Methods 0.000 description 1
- 230000000171 quenching effect Effects 0.000 description 1
- 230000009467 reduction Effects 0.000 description 1
- 230000001172 regenerating effect Effects 0.000 description 1
- 238000011160 research Methods 0.000 description 1
- 230000007017 scission Effects 0.000 description 1
- 230000035945 sensitivity Effects 0.000 description 1
- 238000004904 shortening Methods 0.000 description 1
- 229910052708 sodium Inorganic materials 0.000 description 1
- 239000001632 sodium acetate Substances 0.000 description 1
- 235000017281 sodium acetate Nutrition 0.000 description 1
- 239000001509 sodium citrate Substances 0.000 description 1
- 238000002415 sodium dodecyl sulfate polyacrylamide gel electrophoresis Methods 0.000 description 1
- 239000000243 solution Substances 0.000 description 1
- 238000011895 specific detection Methods 0.000 description 1
- 108010068698 spleen exonuclease Proteins 0.000 description 1
- 239000006228 supernatant Substances 0.000 description 1
- 230000001629 suppression Effects 0.000 description 1
- 238000012360 testing method Methods 0.000 description 1
- 238000005382 thermal cycling Methods 0.000 description 1
- 230000014616 translation Effects 0.000 description 1
- 239000001226 triphosphate Substances 0.000 description 1
- 235000011178 triphosphate Nutrition 0.000 description 1
- UNXRWKVEANCORM-UHFFFAOYSA-N triphosphoric acid Chemical compound OP(O)(=O)OP(O)(=O)OP(O)(O)=O UNXRWKVEANCORM-UHFFFAOYSA-N 0.000 description 1
- HRXKRNGNAMMEHJ-UHFFFAOYSA-K trisodium citrate Chemical compound [Na+].[Na+].[Na+].[O-]C(=O)CC(O)(CC([O-])=O)C([O-])=O HRXKRNGNAMMEHJ-UHFFFAOYSA-K 0.000 description 1
- 229940038773 trisodium citrate Drugs 0.000 description 1
- 238000011144 upstream manufacturing Methods 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6844—Nucleic acid amplification reactions
- C12Q1/6848—Nucleic acid amplification reactions characterised by the means for preventing contamination or increasing the specificity or sensitivity of an amplification reaction
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2521/00—Reaction characterised by the enzymatic activity
- C12Q2521/50—Other enzymatic activities
- C12Q2521/525—Phosphatase
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2525/00—Reactions involving modified oligonucleotides, nucleic acids, or nucleotides
- C12Q2525/10—Modifications characterised by
- C12Q2525/186—Modifications characterised by incorporating a non-extendable or blocking moiety
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2549/00—Reactions characterised by the features used to influence the efficiency or specificity
- C12Q2549/10—Reactions characterised by the features used to influence the efficiency or specificity the purpose being that of reducing false positive or false negative signals
- C12Q2549/101—Hot start
Landscapes
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Health & Medical Sciences (AREA)
- Engineering & Computer Science (AREA)
- Molecular Biology (AREA)
- Biochemistry (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Biophysics (AREA)
- Analytical Chemistry (AREA)
- Physics & Mathematics (AREA)
- Immunology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- General Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
- Enzymes And Modification Thereof (AREA)
Description
(DNA)n残基+dNTP⇔(DNA)n+1残基+PPi
例えばDNAシーケンシング反応で高濃度のPPiが存在すると上記反応は逆方向になることが知られている。これはピロリン酸分解として知られており、熱安定性DNAポリメラーゼを使用して70℃でDNAシーケンシングする場合に問題となることが認められている。DNAシーケンシングで使用するDNAポリメラーゼ製剤に熱安定性PPaseを加えることによりこの問題は解決された。
Taq DNAポリメラーゼを使用し、無機ピロリン酸塩として各種量のピロリン酸四ナトリウム・10水和物(PPi)の存在下に下記試薬を使用して標準の500bpのラムダ鋳型を用いるPCRを実施した。
i)94℃で3.00分間、
ii)94℃で10秒間、50℃で10秒間、72℃で30秒間を20サイクル、
iii)72℃で7分間、
iv)25℃に保温。
1+2 0 PPi
3+4 1mM PPi
5+6 2mM PPi
7+8 3mM PPi
9+10 4mM PPi
11 5mM PPi。
MgはPPiに結合するので、実施例1の結果はMgが過剰のPPiによりキレート化されたためであるとも考えられる。その結果、プライマー伸長を進行させるにはMgが不足したとも考えられる。この可能性を排除するために、各種濃度のマグネシウムイオンの存在下に3mM PPiを使用して実施例1の手順を繰返した。
1+2 1.5mM MgCl2
3+4 5mM MgCl2
5+6 7.5mM MgCl2
7+8 10mM MgCl2
9+10 1.5mM MgCl2+3mM PPi
11+12 5mM MgCl2+3mM PPi
13+14 7.5mM MgCl2+3mM PPi
15 10mM MgCl2+3mM PPi。
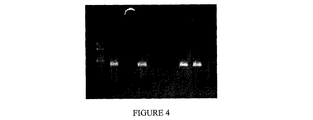
ピロリン酸塩(PPi)を加えた反応混合物にSulfolobus acidocaldarius PPase(Sac PPase)0.2uを加えた以外は実施例1の500bpラムダPCRを繰返した。反応混合物をSac PPase 0.2uの存在下で95℃に5分間保温すると、ピロリン酸塩を分解してPCR反応を進行させるに十分であった。
上段
1+2 1mM PPi+0.2u PPase
3+4 2mM PPi+0.2u PPase
5+6 3mM PPi+0.2u PPase
7+8 4mM PPi+0.2u PPase
9+10 5mM PPi+0.2u PPase
下段
1+2 1mM PPi
3+4 2mM PPi
5+6 3mM PPi
7+8 4mM PPi
9+10 5mM PPi
11+12 0mM PPi。
次に、正しい寸法のPCR産物を生産するために「ホットスタート」反応を必要とするアッセイシステムに本発明の方法を適用した。
i)95℃で2.00分以下、
ii)95℃で15秒間、50℃で30秒間、72℃で30秒間を35サイクル、
iii)72℃で7分間、
iv)25℃に保温。
1 標準TaqポリメラーゼPCR,ベタイン無,ミスプライミング多数;
2 標準TaqポリメラーゼPCR,ベタイン有,明バンドが正しい産物,多少ミスプライミング有;
3 標準TaqポリメラーゼPCR,ベタイン無,3mM PPiと0.2u Sac PPase添加,ミスプライミング皆無,95℃で5分間変性;
4 標準TaqポリメラーゼPCR,ベタイン有,3mM PPiと0.2u Sac PPase添加,正しい産物のみ,95℃で5分間変性;
5+6 95℃で2分間変性以外は3と同じ;
7+8 95℃で2分間変性以外は4と同じ。
0.2u Sac PPaseと3mM PPiを加えたPCR混合物をアンギオテンシンアッセイ実施前に各種時間20℃の室温に放置した場合の影響を調べた。Sac PPaseは熱安定性酵素であるが、室温で低レベルの酵素活性がある可能性があった。その結果、DNAポリメラーゼを抑制/停止させるには反応にPPiが不足し、プライマー伸長が起こり、「ホットスタート」機能が失われる恐れがあった。
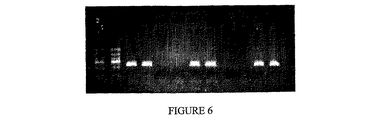
Sac PPaseの代わりに別の市販熱安定性PPase(活性単位の定義は異なる)を使用して同様の反応を行い、実施例4に記載したアッセイを繰返した。結果を図6に示す。同図中、各レーンは以下の反応を表す。
1+2 標準TaqポリメラーゼPCR,ベタイン無;
3+4 標準TaqポリメラーゼPCR,ベタイン有;
5+6 標準TaqポリメラーゼPCR,ベタイン無,3mM PPiと0.2u Sac PPase添加;
7+8 標準TaqポリメラーゼPCR,ベタイン有,3mM PPiと0.2u Sac PPase添加;
9+10 標準TaqポリメラーゼPCR,ベタイン無,3mM PPiと10u*Thermococcus litoralis PPase添加;
11+12 標準TaqポリメラーゼPCR,ベタイン有,3mM PPiと10u*Thermococcus litoralis PPase添加。
本発明の方法と数種の比較アッセイで各種熱安定性DNAポリメラーゼを使用した。使用したポリメラーゼは数種の非プルーフリーディングThermus種DNAポリメラーゼ、プルーフリーディング超好熱始原菌DNAポリメラーゼ、及び非プルーフリーディングDNAポリメラーゼとプルーフリーディングDNAポリメラーゼの混合物である。
レーン
上段
1+2 Taqポリメラーゼ,0mM PPi,PPase無;
3+4 Taqポリメラーゼ,3mM PPi,PPase無;
5+6 Taqポリメラーゼ,3mM PPi,0.2u Sac PPase;
7+8 Tbrポリメラーゼ,0mM PPi,PPase無;
9+10 Tbrポリメラーゼ,3mM PPi,PPase無;
11+12 Tbrポリメラーゼ,3mM PPi,0.2u Sac PPase;
下段
1+2 Tthポリメラーゼ,0mM PPi,PPase無;
3+4 Tthポリメラーゼ,3mM PPi,PPase無;
5+6 Tthポリメラーゼ,3mM PPi,0.2u Sac PPase;
7+8 TspNHポリメラーゼ,0mM PPi,PPase無;
9+10 TspNHポリメラーゼ,3mM PPi,PPase無;
11+12 TspNHポリメラーゼ,3mM PPi,0.2u Sac PPase。
レーン
上段
1+2 Pfuポリメラーゼ,0mM PPi,PPase無;
3+4 Pfuポリメラーゼ,3mM PPi,PPase無;
5+6 Pfuポリメラーゼ,3mM PPi,0.2u Sac PPase;
7+8 9°Nエキソポリメラーゼ,0mM PPi,PPase無;
9+10 9°Nエキソポリメラーゼ,3mM PPi,PPase無;
11+12 9°Nエキソポリメラーゼ,3mM PPi,0.2u Sac PPase;
下段
1+2 VENTポリメラーゼ,0mM PPi,PPase無;
3+4 VENTポリメラーゼ,3mM PPi,PPase無;
5+6 VENTポリメラーゼ,3mM PPi,0.2u Sac PPase。
レーン
1+2 Taqポリメラーゼ,ベタイン無;
3+4 Taqポリメラーゼ,ベタイン有;
5+6 Accuraseポリメラーゼ,ベタイン無;
7+8 Accuraseポリメラーゼ,ベタイン有;
9+10 Tbrポリメラーゼ,ベタイン無;
11+12 Tbrポリメラーゼ,ベタイン有;
13+14 Tthポリメラーゼ,ベタイン無;
15+16 Tthポリメラーゼ,ベタイン有。
対照レーン1〜4(上段)及び12〜16(下段)
レーン
上段
1+2 Taqポリメラーゼ,ベタイン無,3mM PPi,Sac PPase無;
3+4 Taqポリメラーゼ,ベタイン有,3mM PPi,Sac PPase無;
以下、全て3mM PPiと0.2u Sac PPase添加
5+6 Taqポリメラーゼ,ベタイン無;
7+8 Taqポリメラーゼ,ベタイン有;
9+10 Accuraseポリメラーゼ,ベタイン無;
11+12 Accuraseポリメラーゼ,ベタイン有;
13+14 Tbrポリメラーゼ,ベタイン無;
15+16 Tbrポリメラーゼ,ベタイン有。
下段
以下、全て3mM PPiと0.2u Sac PPase添加
1+2 TthポリメラーゼPCR,ベタイン無;
3+4 TthポリメラーゼPCR,ベタイン有;
5+6 TspNHポリメラーゼ,ベタイン無;
7+8 TspNHポリメラーゼ,ベタイン有;
9+10 Pfuポリメラーゼ,ベタイン無;
11+12 Pfuポリメラーゼ,ベタイン有;
13+14 Taqポリメラーゼ対照,ベタイン無,PPi又はPPase無;
15+16 Taqポリメラーゼ対照,ベタイン有,PPi又はPPase無。
当初の結果(図9及び10)によると、化学修飾Taqポリメラーゼ(米国特許第5,677,152号に記載されているように修飾)はベタインの不在下では多少の偽PCR産物を生産するが、ベタインの存在下では正しい産物が得られる。
レーン
1+2 Taqポリメラーゼ,ベタイン無;
3+4 Taqポリメラーゼ,ベタイン有;
5+6 化学修飾Taq,ベタイン無;
7+8 化学修飾Taq,ベタイン有;
9+10 本発明の方法(3mM PPiと2u Sac PPase),ベタイン無;
11+12 本発明の方法(3mM PPiと2u Sac PPase),ベタイン有。
レーン
1+2 抗Taq抗体+Taqポリメラーゼ,ベタイン無;
3+4 抗Taq抗体+Taqポリメラーゼ,ベタイン有。
微生物系統保存施設のカルチャーコレクションからAeropyrum pernixを入手した。無機ピロホスファターゼ酵素をクローニングし、発現させ、精製した。
Aeropyrum pernixのピロホスファターゼ遺伝子を含むゲノム配列を図11に示す。使用したプライマーはAeropyrum pernixのゲノム配列から設計した。これらを5’→3’ として以下に示し、制限部位を太字で示す。
94℃で3分間初期保温、
94℃で10秒間、55℃で10秒間、68℃で45秒間を20サイクル、
72℃で7分間最終保温。
50pM上流プライマー(5’..TGCATGCATATGACAGGCTGTCTGAAAATTG..3’−配列番号18)
50pM下流プライマー(5’..TAAGTGTAAGCTTGACTGTGGGGGCGGTGAAAG..3’−配列番号19)
1.5mM MgCl2
1.25u Accurase DNAポリメラーゼ(カタログ番号AC001,GeneSys Ltd.)
75mM Tris,pH8.8
20mM硫酸アンモニウム
0.1%(w/v)Tween 20
100ng Aeropyrum pernixゲノムDNA。
このクローンをLB24リットルに移した。OD600が約1.5に達したら、0.5mM IPTGを加えて培養を誘導し、更に4時間増殖させた。次に細胞を回収し、細胞ペレットを溶解させた。発現酵素をフェニルセファロースCL4B(Amersham Pharmacia Biotech)、ヒドロキシアパタイト(Bio−rad Laboratories)及び高性能Qセファロース(Amersham Pharmacia Biotech)で標準カラムクロマトグラフィ¬により精製し、最後に20mM Tris−HCl,pH8.0,100mM NaCl,0.5%(v/v)Tween 20,0.5%(v/v)Nonidet P40,0.1mM EDTA,1mMジチオスレイトール及び50%グリセロール中で−20℃で保存した。
A.pernix無機ピロホスファターゼ酵素を使用して本発明の方法を実施した。このアッセイの基本はヒトBアクチン遺伝子の増幅である。
1倍反応緩衝液
dATP、dCTP、dGTP各200μM及びdUTP400μM
0.025u/μl未修飾Taqポリメラーゼ
0.002u/μl Aeropyrum pernix無機ピロホスファターゼ
0.3μM 5’プライマー(5’GAC TCG TCA TAC TCC TGC TTG CT 3’−配列番号22)
0.3μM 3’プライマー(5’CAT TGC CGA CAG GAT GCA GAA 3’−配列番号23)
0.15μM Taqmanプローブ(FAM−ATCCACATCTGCTGGAAGGTGGACAGT−TAMRA−配列番号24)
5mM MgCl2
2mM NaPPi
受動的参照
7.5ngから出発したヒトゲノムDNAの4倍希釈液(2500コピー)。
94℃で3分間初期変性、
94℃で15秒間と60℃で60秒間を40サイクル。
Claims (23)
- 核酸増幅反応を実施するための方法であって、該方法は、
ポリメラーゼ連鎖反応によって増幅反応を実施するために必要な試薬と、プライマー伸長を妨げる量のピロリン酸塩が混合された、PCR反応混合物を形成する段階、
該混合物にPCR反応混合物50μLあたり少なくとも0.04単位の熱安定性ピロホスファターゼ酵素(PPase)を加える段階、及び
該ピロリン酸塩がピロホスファターゼ酵素(PPase)により消化される条件下に該反応混合物をおくことによって増幅反応が実施される段階を含む前記方法。 - 核酸増幅反応を実施するための方法であって、該方法は、
PCR増幅反応を実施するために必要な試薬と、プライマー伸長を妨げる量のピロリン酸塩が混合された、PCR反応混合物を形成する段階、
該混合物に超好熱始原菌から得られる熱安定性ピロホスファターゼ酵素(PPase)を加える段階、及び
該ピロリン酸塩がピロホスファターゼ酵素(PPase)により消化される条件下に該反応混合物をおくことにより増幅反応が実施される段階を含む前記方法。 - 反応混合物がサーマス アクアティカス(Thermus aquaticus)ポリメラーゼ(Taq)、サーマス サーモフィラス(Thermus thermophilus)ポリメラーゼ(Tth)、サーマス(Thermus)種NHポリメラーゼ(TspNH)、サーマス ブロキアヌス(Thermus brockianus)ポリメラーゼ(Tbr)、ピロコッカス フリオサス(Pyrococcus furiosus)ポリメラーゼ(Pfu)、9°N7 エキソDNAポリメラーゼ、及びサーモコッカス リトラリス(Thermococcus litoralis) DNAポリメラーゼから選択されるDNAポリメラーゼを含む請求項1又は2に記載の方法。
- ピロリン酸塩がアルカリ金属ピロリン酸塩である請求項1から3のいずれか一項に記載の方法。
- ピロリン酸塩が式Na4P2O7のピロリン酸四ナトリウムである請求項4に記載の方法。
- ピロリン酸塩が少なくとも0.5mMの濃度で反応混合物中に存在する請求項1から5のいずれか一項に記載の方法。
- ピロリン酸塩が1〜10mMの濃度で存在する請求項6に記載の方法。
- 熱安定性PPaseがスルホロバス アシドカルダリウス(Sulfolobus acidocaldarius)無機ピロホスファターゼ(Sac PPase)、サーモコッカス リトラリス(Thermococcus litoralis)無機ピロホスファターゼ又はアエロピラム ペルニクス(Aeropyrum pernix)無機ピロホスファターゼである請求項1から7のいずれか一項に記載の方法。
- 熱安定性PPaseが配列番号25に示すアミノ酸配列又はその変異体もしくはフラグメントを含む請求項1から8のいずれか一項に記載の方法。
- 熱安定性PPaseを反応混合物の形成直後に添加する請求項8又は9に記載の方法。
- 存在するピロリン酸塩をPPaseで消化するために増幅反応前に50℃以上の温度でインキュベーションする段階を含む請求項10に記載の方法。
- PPaseをPCR反応混合物50μL当たり少なくとも0.04uの濃度で反応混合物に加える請求項2に記載の方法。
- PPaseをPCR反応混合物50μL当たり0.08uの濃度で反応混合物に加える請求項1から12のいずれか一項に記載の方法。
- PPaseをPCR反応混合物50μL当たり0.2〜10uの濃度で反応混合物に加える請求項12又は13に記載の方法。
- 増幅反応を実施するためのキットであって、ピロリン酸塩と、PCR反応混合物50μL当たり少なくとも0.04uとなる量の熱安定性PPaseを含む前記キット。
- 増幅反応で使用するのに必要な1種以上の試薬を含む請求項15に記載のキット。
- 特定ターゲット核酸の増幅を実施するために必要な1種以上のプライマーを更に含む請求項16に記載のキット。
- 1種以上の蛍光標識試薬を更に含む請求項15から17のいずれか一項に記載のキット。
- 蛍光標識試薬がインターカレーター色素、蛍光標識プローブ、蛍光標識プライマー又は蛍光標識ヌクレオチドの1種以上から選択される請求項18に記載のキット。
- 請求項1から14のいずれか一項に記載の増幅反応を実施するための方法における熱安定性ピロホスファターゼ酵素の使用。
- ピロホスファターゼ酵素がアエロピラム ペルニクス(Aeropyrum pernix)から単離した酵素である請求項20に記載の使用。
- ピロホスファターゼ酵素が配列番号26に示すポリヌクレオチド配列又はその変異体もしくはフラグメントによりコードされる請求項20に記載の使用。
- ピロホスファターゼ酵素が配列番号25に示すアミノ酸配列又はその変異体もしくはフラグメントを含む請求項20に記載の使用。
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| GBGB0110501.4A GB0110501D0 (en) | 2001-04-30 | 2001-04-30 | Amplification process |
| PCT/GB2002/001861 WO2002088387A2 (en) | 2001-04-30 | 2002-04-22 | Amplification process |
Publications (3)
| Publication Number | Publication Date |
|---|---|
| JP2004524853A JP2004524853A (ja) | 2004-08-19 |
| JP2004524853A5 JP2004524853A5 (ja) | 2005-12-22 |
| JP4177118B2 true JP4177118B2 (ja) | 2008-11-05 |
Family
ID=9913697
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| JP2002585667A Expired - Fee Related JP4177118B2 (ja) | 2001-04-30 | 2002-04-22 | 増幅方法 |
Country Status (14)
| Country | Link |
|---|---|
| US (2) | US6951744B2 (ja) |
| EP (1) | EP1390532B1 (ja) |
| JP (1) | JP4177118B2 (ja) |
| KR (2) | KR20100005245A (ja) |
| CN (1) | CN1318604C (ja) |
| AR (1) | AR035238A1 (ja) |
| AT (1) | ATE502118T1 (ja) |
| AU (1) | AU2002249444B2 (ja) |
| CA (1) | CA2445864C (ja) |
| DE (1) | DE60239456D1 (ja) |
| GB (2) | GB0110501D0 (ja) |
| NZ (1) | NZ528919A (ja) |
| TW (1) | TWI323284B (ja) |
| WO (1) | WO2002088387A2 (ja) |
Families Citing this family (19)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| GB0112868D0 (en) | 2001-05-25 | 2001-07-18 | Secr Defence | Detection system |
| CN1500887A (zh) * | 2002-10-01 | 2004-06-02 | 松下电器产业株式会社 | 引物伸长反应检测方法、碱基种类判别方法及其装置 |
| US20060051796A1 (en) * | 2004-09-09 | 2006-03-09 | Inga Boell | Real time PCR with the addition of pyrophosphatase |
| EP1634965B1 (en) | 2004-09-09 | 2010-01-20 | Roche Diagnostics GmbH | Real time PCR with the addition of pyrophosphatase |
| GB0502010D0 (en) * | 2005-02-01 | 2005-03-09 | Enigma Diagnostics Ltd | Biochemical reagents and their uses |
| JP5558811B2 (ja) | 2006-06-01 | 2014-07-23 | トリリンク バイオテクノロジーズ | 核酸増幅のための化学的に修飾されたオリゴヌクレオチドプライマー |
| KR101098764B1 (ko) | 2007-10-29 | 2011-12-26 | (주)바이오니아 | 안정화된 핫스타트 pcr용 건조 조성물 |
| GB2473778B (en) | 2008-05-27 | 2013-04-10 | Trilink Biotechnologies | Chemically modified nucleoside 5'-triphosphates for thermally initiated amplification of nucleic acid |
| EP2566959A1 (en) * | 2010-04-30 | 2013-03-13 | Roche Diagniostics GmbH | System and method for purification and use of inorganic pyrophosphatase from aquifex aeolicus |
| US20110312841A1 (en) * | 2010-06-17 | 2011-12-22 | Geneasys Pty Ltd | Fabrication system for lab-on-a-chip (loc) devices with differing application specific functionality |
| KR101870311B1 (ko) * | 2012-03-09 | 2018-06-25 | (주)바이오니아 | 핫스타트 역전사반응 또는 핫스타트 역전사 중합효소 연쇄반응용 조성물 |
| KR102264617B1 (ko) | 2012-06-14 | 2021-06-14 | 라이프 테크놀로지스 코포레이션 | 폴리머라제 연쇄 반응 (pcr)을 위한 신규 조성물, 방법 및 키트 |
| GB201301457D0 (en) | 2013-01-28 | 2013-03-13 | Fluorogenics Ltd | Freeze-dried composition |
| JP6467829B2 (ja) * | 2014-09-03 | 2019-02-13 | 東洋紡株式会社 | 改良されたrt−pcr反応 |
| CN104862288B (zh) * | 2015-05-08 | 2019-10-29 | 苏州大学 | 一种新型温度响应性无机焦磷酸酶偶联物的合成及其在增强聚合酶链式反应中的应用 |
| EP3402880B1 (en) | 2016-01-15 | 2024-01-03 | Thermo Fisher Scientific Baltics UAB | Thermophilic dna polymerase mutants |
| US11618891B2 (en) | 2017-06-26 | 2023-04-04 | Thermo Fisher Scientific Baltics Uab | Thermophilic DNA polymerase mutants |
| JP7234598B2 (ja) * | 2018-01-26 | 2023-03-08 | 東ソー株式会社 | 核酸増幅試薬および当該試薬を用いた核酸増幅制御方法 |
| CN113481180A (zh) * | 2021-07-05 | 2021-10-08 | 吉林大学 | 碱性嗜热无机焦磷酸酶及其在增强聚合酶链式反应和合成udp-半乳糖反应中的应用 |
Family Cites Families (20)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4868103A (en) | 1986-02-19 | 1989-09-19 | Enzo Biochem, Inc. | Analyte detection by means of energy transfer |
| CH670709A5 (ja) * | 1986-04-24 | 1989-06-30 | Univ Moskovsk | |
| US5498523A (en) * | 1988-07-12 | 1996-03-12 | President And Fellows Of Harvard College | DNA sequencing with pyrophosphatase |
| WO1990012111A1 (en) * | 1989-04-12 | 1990-10-18 | President And Fellows Of Harvard College | Improved primer extension reactions |
| KR100236506B1 (ko) | 1990-11-29 | 2000-01-15 | 퍼킨-엘머시터스인스트루먼츠 | 폴리머라제 연쇄 반응 수행 장치 |
| FI923911A (fi) * | 1992-09-01 | 1994-03-02 | Vsevolod Kiselev | DNA-molekylers in vitro-syntes |
| US5565339A (en) | 1992-10-08 | 1996-10-15 | Hoffmann-La Roche Inc. | Compositions and methods for inhibiting dimerization of primers during storage of polymerase chain reaction reagents |
| US5491063A (en) | 1994-09-01 | 1996-02-13 | Hoffmann-La Roche Inc. | Methods for in-solution quenching of fluorescently labeled oligonucleotide probes |
| US5773258A (en) | 1995-08-25 | 1998-06-30 | Roche Molecular Systems, Inc. | Nucleic acid amplification using a reversibly inactivated thermostable enzyme |
| US5665551A (en) * | 1995-09-13 | 1997-09-09 | Roche Molecular Systems, Inc. | Purified nucleic acid encoding a thermostable pyrophosphatase |
| US6291164B1 (en) * | 1996-11-22 | 2001-09-18 | Invitrogen Corporation | Methods for preventing inhibition of nucleic acid synthesis by pyrophosphate |
| GB9716052D0 (en) | 1996-12-06 | 1997-10-01 | Secr Defence | Reaction vessels |
| GB9725197D0 (en) | 1997-11-29 | 1998-01-28 | Secr Defence | Detection system |
| GB9725237D0 (en) | 1997-11-29 | 1998-01-28 | Secr Defence | Amplification system |
| GB9803382D0 (en) | 1998-02-19 | 1998-04-15 | Secr Defence | Detection system |
| US6183998B1 (en) | 1998-05-29 | 2001-02-06 | Qiagen Gmbh Max-Volmer-Strasse 4 | Method for reversible modification of thermostable enzymes |
| GB9812768D0 (en) | 1998-06-13 | 1998-08-12 | Zeneca Ltd | Methods |
| KR100292883B1 (ko) * | 1999-02-09 | 2001-06-15 | 박한오 | 피로포스페이트와 피로포스파타제를 포함하는 핫 스타트 pcr 반응혼합물 |
| AU2001241621B2 (en) | 2000-02-23 | 2005-08-25 | City Of Hope | Pyrophosphorolysis activated polymerization (PAP): application to allele-specific amplification and nucleic acid sequence determination |
| KR20050005626A (ko) * | 2003-07-07 | 2005-01-14 | 주식회사 포스코 | 코크스 오븐에서의 장입탄 밀도 향상방법 |
-
2001
- 2001-04-30 GB GBGB0110501.4A patent/GB0110501D0/en not_active Ceased
-
2002
- 2002-03-28 TW TW091106126A patent/TWI323284B/zh not_active IP Right Cessation
- 2002-04-22 AT AT02718374T patent/ATE502118T1/de not_active IP Right Cessation
- 2002-04-22 WO PCT/GB2002/001861 patent/WO2002088387A2/en active Application Filing
- 2002-04-22 AU AU2002249444A patent/AU2002249444B2/en not_active Ceased
- 2002-04-22 JP JP2002585667A patent/JP4177118B2/ja not_active Expired - Fee Related
- 2002-04-22 KR KR1020097025792A patent/KR20100005245A/ko not_active Application Discontinuation
- 2002-04-22 CA CA2445864A patent/CA2445864C/en not_active Expired - Fee Related
- 2002-04-22 KR KR1020037014191A patent/KR101039563B1/ko not_active IP Right Cessation
- 2002-04-22 GB GB0209052A patent/GB2377991B/en not_active Expired - Fee Related
- 2002-04-22 NZ NZ528919A patent/NZ528919A/en unknown
- 2002-04-22 CN CNB028132726A patent/CN1318604C/zh not_active Expired - Fee Related
- 2002-04-22 EP EP02718374A patent/EP1390532B1/en not_active Expired - Lifetime
- 2002-04-22 DE DE60239456T patent/DE60239456D1/de not_active Expired - Lifetime
- 2002-04-29 AR ARP020101570A patent/AR035238A1/es unknown
- 2002-04-30 US US10/135,807 patent/US6951744B2/en not_active Expired - Fee Related
-
2005
- 2005-08-17 US US11/205,667 patent/US7449312B2/en not_active Expired - Fee Related
Also Published As
| Publication number | Publication date |
|---|---|
| US20060057617A1 (en) | 2006-03-16 |
| CA2445864A1 (en) | 2002-11-07 |
| TWI323284B (en) | 2010-04-11 |
| US20030049655A1 (en) | 2003-03-13 |
| GB2377991B (en) | 2004-01-28 |
| NZ528919A (en) | 2005-07-29 |
| AR035238A1 (es) | 2004-05-05 |
| GB0110501D0 (en) | 2001-06-20 |
| CN1318604C (zh) | 2007-05-30 |
| JP2004524853A (ja) | 2004-08-19 |
| CN1522305A (zh) | 2004-08-18 |
| US6951744B2 (en) | 2005-10-04 |
| DE60239456D1 (de) | 2011-04-28 |
| EP1390532B1 (en) | 2011-03-16 |
| EP1390532A2 (en) | 2004-02-25 |
| ATE502118T1 (de) | 2011-04-15 |
| GB0209052D0 (en) | 2002-05-29 |
| CA2445864C (en) | 2011-08-02 |
| AU2002249444B2 (en) | 2007-07-12 |
| US7449312B2 (en) | 2008-11-11 |
| GB2377991A (en) | 2003-01-29 |
| KR20100005245A (ko) | 2010-01-14 |
| KR101039563B1 (ko) | 2011-06-09 |
| WO2002088387A2 (en) | 2002-11-07 |
| WO2002088387A3 (en) | 2003-12-11 |
| KR20040015216A (ko) | 2004-02-18 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US7449312B2 (en) | Kit for conducting a polymerase chain reaction | |
| US7122355B2 (en) | Composition and method for hot start nucleic acid amplification | |
| JP3421664B2 (ja) | ヌクレオチド塩基の同定方法 | |
| US5487993A (en) | Direct cloning of PCR amplified nucleic acids | |
| JP3140937B2 (ja) | 好熱酵素を用いる鎖置換増幅法 | |
| EP2824189B1 (en) | Composition for hot-start reverse transcription reaction or hot-start reverse transcription polymerase chain reaction | |
| US10787702B2 (en) | Thermolabile exonucleases | |
| US20080064071A1 (en) | Zwitterionic detergents for the storage and use of DNA polymerases | |
| EP0669401A2 (en) | Amplification of long nucleic acid sequences by PCR | |
| AU2002249444A1 (en) | Amplification process | |
| JP2016510601A (ja) | 修飾RNAモノマーを用いたRNaseH系アッセイ | |
| US20160130673A1 (en) | Nucleic acid detection by oligonucleotide probes cleaved by both exonuclease and endonuclease | |
| MX2015000766A (es) | Sondas, cebadores cooperativos y aplicaciones de ellos. | |
| US20120219945A1 (en) | Use of single-stranded binding protein in amplifying target nucleic acid | |
| CN110446791B (zh) | 多核苷酸衔接子及其使用方法 | |
| US20020064837A1 (en) | Method for synthesizing a nucleic acid molecule using a ribonuclease | |
| EP1548112B1 (en) | Method of amplifying nucleic acid, reagent kit for amplifying nucleic acid, method of detecting single nucleotide polymorphism and reagent kit for detecting single nucleotide polymorphism | |
| EP1277841B1 (en) | New composition and method for hot start nucleic acid amplification | |
| JPH0851979A (ja) | Rnaの逆転写方法およびその用途 | |
| WO2002101041A1 (fr) | Procede d'amplification de l'acide nucleique et procede de detection du polymorphisme des nucleotides a l'aide d'un analogue de nucleotide |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A523 Effective date: 20040630 |
|
| A621 | Written request for application examination |
Free format text: JAPANESE INTERMEDIATE CODE: A621 Effective date: 20041118 |
|
| A131 | Notification of reasons for refusal |
Free format text: JAPANESE INTERMEDIATE CODE: A131 Effective date: 20070814 |
|
| A601 | Written request for extension of time |
Free format text: JAPANESE INTERMEDIATE CODE: A601 Effective date: 20071107 |
|
| A602 | Written permission of extension of time |
Free format text: JAPANESE INTERMEDIATE CODE: A602 Effective date: 20071114 |
|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A523 Effective date: 20080214 |
|
| A131 | Notification of reasons for refusal |
Free format text: JAPANESE INTERMEDIATE CODE: A131 Effective date: 20080325 |
|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A523 Effective date: 20080620 |
|
| TRDD | Decision of grant or rejection written | ||
| A01 | Written decision to grant a patent or to grant a registration (utility model) |
Free format text: JAPANESE INTERMEDIATE CODE: A01 Effective date: 20080729 |
|
| A01 | Written decision to grant a patent or to grant a registration (utility model) |
Free format text: JAPANESE INTERMEDIATE CODE: A01 |
|
| A61 | First payment of annual fees (during grant procedure) |
Free format text: JAPANESE INTERMEDIATE CODE: A61 Effective date: 20080821 |
|
| FPAY | Renewal fee payment (event date is renewal date of database) |
Free format text: PAYMENT UNTIL: 20110829 Year of fee payment: 3 |
|
| R150 | Certificate of patent or registration of utility model |
Free format text: JAPANESE INTERMEDIATE CODE: R150 |
|
| FPAY | Renewal fee payment (event date is renewal date of database) |
Free format text: PAYMENT UNTIL: 20110829 Year of fee payment: 3 |
|
| FPAY | Renewal fee payment (event date is renewal date of database) |
Free format text: PAYMENT UNTIL: 20120829 Year of fee payment: 4 |
|
| FPAY | Renewal fee payment (event date is renewal date of database) |
Free format text: PAYMENT UNTIL: 20120829 Year of fee payment: 4 |
|
| FPAY | Renewal fee payment (event date is renewal date of database) |
Free format text: PAYMENT UNTIL: 20130829 Year of fee payment: 5 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| R250 | Receipt of annual fees |
Free format text: JAPANESE INTERMEDIATE CODE: R250 |
|
| LAPS | Cancellation because of no payment of annual fees |