KR20200038528A - Dctn1 단백질과 ret 단백질과의 융합 단백질 - Google Patents
Dctn1 단백질과 ret 단백질과의 융합 단백질 Download PDFInfo
- Publication number
- KR20200038528A KR20200038528A KR1020207007893A KR20207007893A KR20200038528A KR 20200038528 A KR20200038528 A KR 20200038528A KR 1020207007893 A KR1020207007893 A KR 1020207007893A KR 20207007893 A KR20207007893 A KR 20207007893A KR 20200038528 A KR20200038528 A KR 20200038528A
- Authority
- KR
- South Korea
- Prior art keywords
- leu
- glu
- seq
- ala
- ser
- Prior art date
Links
- 101000579425 Homo sapiens Proto-oncogene tyrosine-protein kinase receptor Ret Proteins 0.000 title claims abstract description 211
- 102100028286 Proto-oncogene tyrosine-protein kinase receptor Ret Human genes 0.000 title claims abstract description 207
- 102100036654 Dynactin subunit 1 Human genes 0.000 title claims abstract description 121
- 101000929626 Homo sapiens Dynactin subunit 1 Proteins 0.000 title claims abstract description 120
- 108020001507 fusion proteins Proteins 0.000 title claims description 29
- 102000037865 fusion proteins Human genes 0.000 title claims description 29
- 108091033319 polynucleotide Proteins 0.000 claims abstract description 275
- 102000040430 polynucleotide Human genes 0.000 claims abstract description 275
- 239000002157 polynucleotide Substances 0.000 claims abstract description 275
- 108090000765 processed proteins & peptides Proteins 0.000 claims abstract description 275
- 229920001184 polypeptide Polymers 0.000 claims abstract description 234
- 102000004196 processed proteins & peptides Human genes 0.000 claims abstract description 234
- 238000000034 method Methods 0.000 claims abstract description 122
- 230000014509 gene expression Effects 0.000 claims abstract description 104
- 150000001875 compounds Chemical class 0.000 claims abstract description 95
- 230000000694 effects Effects 0.000 claims abstract description 57
- 238000012216 screening Methods 0.000 claims abstract description 16
- 239000008194 pharmaceutical composition Substances 0.000 claims abstract description 15
- 239000004480 active ingredient Substances 0.000 claims abstract description 12
- 230000004927 fusion Effects 0.000 claims description 189
- 210000004027 cell Anatomy 0.000 claims description 166
- 108090000623 proteins and genes Proteins 0.000 claims description 126
- 150000001413 amino acids Chemical class 0.000 claims description 97
- 239000000523 sample Substances 0.000 claims description 80
- 206010028980 Neoplasm Diseases 0.000 claims description 71
- 201000011510 cancer Diseases 0.000 claims description 65
- 239000002773 nucleotide Substances 0.000 claims description 36
- 125000003729 nucleotide group Chemical group 0.000 claims description 36
- 210000004899 c-terminal region Anatomy 0.000 claims description 35
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 claims description 33
- 238000012360 testing method Methods 0.000 claims description 30
- 238000001514 detection method Methods 0.000 claims description 27
- 230000035755 proliferation Effects 0.000 claims description 26
- 101150077555 Ret gene Proteins 0.000 claims description 21
- 239000013604 expression vector Substances 0.000 claims description 21
- 101150080336 Dctn1 gene Proteins 0.000 claims description 19
- 238000002512 chemotherapy Methods 0.000 claims description 16
- 230000002401 inhibitory effect Effects 0.000 claims description 13
- 230000000692 anti-sense effect Effects 0.000 claims description 12
- 239000003814 drug Substances 0.000 claims description 12
- 229940079593 drug Drugs 0.000 claims description 10
- 230000008569 process Effects 0.000 claims description 8
- 239000003795 chemical substances by application Substances 0.000 claims description 5
- 230000000295 complement effect Effects 0.000 claims description 5
- 238000004519 manufacturing process Methods 0.000 claims description 4
- 239000000090 biomarker Substances 0.000 claims description 3
- 125000003275 alpha amino acid group Chemical group 0.000 claims 10
- YBAFDPFAUTYYRW-UHFFFAOYSA-N N-L-alpha-glutamyl-L-leucine Natural products CC(C)CC(C(O)=O)NC(=O)C(N)CCC(O)=O YBAFDPFAUTYYRW-UHFFFAOYSA-N 0.000 description 50
- WHUUTDBJXJRKMK-UHFFFAOYSA-N Glutamic acid Chemical group OC(=O)C(N)CCC(O)=O WHUUTDBJXJRKMK-UHFFFAOYSA-N 0.000 description 49
- 102000004169 proteins and genes Human genes 0.000 description 36
- 108010055341 glutamyl-glutamic acid Proteins 0.000 description 32
- KOSRFJWDECSPRO-UHFFFAOYSA-N alpha-L-glutamyl-L-glutamic acid Natural products OC(=O)CCC(N)C(=O)NC(CCC(O)=O)C(O)=O KOSRFJWDECSPRO-UHFFFAOYSA-N 0.000 description 31
- 235000018102 proteins Nutrition 0.000 description 30
- 108020004414 DNA Proteins 0.000 description 27
- 108020004459 Small interfering RNA Proteins 0.000 description 27
- VPZXBVLAVMBEQI-UHFFFAOYSA-N glycyl-DL-alpha-alanine Natural products OC(=O)C(C)NC(=O)CN VPZXBVLAVMBEQI-UHFFFAOYSA-N 0.000 description 27
- 108010062796 arginyllysine Proteins 0.000 description 23
- 108010034529 leucyl-lysine Proteins 0.000 description 23
- 108010005233 alanylglutamic acid Proteins 0.000 description 22
- UZGFHWIJWPUPOH-IHRRRGAJSA-N Arg-Leu-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N UZGFHWIJWPUPOH-IHRRRGAJSA-N 0.000 description 20
- 108010079547 glutamylmethionine Proteins 0.000 description 20
- 108010009298 lysylglutamic acid Proteins 0.000 description 20
- 108010054155 lysyllysine Proteins 0.000 description 20
- 239000002609 medium Substances 0.000 description 20
- 108010047495 alanylglycine Proteins 0.000 description 19
- 230000004663 cell proliferation Effects 0.000 description 19
- 108010050848 glycylleucine Proteins 0.000 description 18
- 108010008355 arginyl-glutamine Proteins 0.000 description 17
- 239000000047 product Substances 0.000 description 17
- 108010026333 seryl-proline Proteins 0.000 description 17
- 239000013598 vector Substances 0.000 description 17
- HVHRPWQEQHIQJF-AVGNSLFASA-N Leu-Lys-Glu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O HVHRPWQEQHIQJF-AVGNSLFASA-N 0.000 description 16
- PMGDADKJMCOXHX-UHFFFAOYSA-N L-Arginyl-L-glutamin-acetat Natural products NC(=N)NCCCC(N)C(=O)NC(CCC(N)=O)C(O)=O PMGDADKJMCOXHX-UHFFFAOYSA-N 0.000 description 15
- 108010044940 alanylglutamine Proteins 0.000 description 15
- 239000002299 complementary DNA Substances 0.000 description 15
- 239000000243 solution Substances 0.000 description 15
- VKKYFICVTYKFIO-CIUDSAMLSA-N Arg-Ala-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCCN=C(N)N VKKYFICVTYKFIO-CIUDSAMLSA-N 0.000 description 14
- IDMNOFVUXYYZPF-DKIMLUQUSA-N Ile-Lys-Phe Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)O)N IDMNOFVUXYYZPF-DKIMLUQUSA-N 0.000 description 14
- PNPYKQFJGRFYJE-GUBZILKMSA-N Lys-Ala-Glu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(O)=O PNPYKQFJGRFYJE-GUBZILKMSA-N 0.000 description 14
- 208000024770 Thyroid neoplasm Diseases 0.000 description 14
- 235000001014 amino acid Nutrition 0.000 description 14
- 229940024606 amino acid Drugs 0.000 description 14
- 108010013835 arginine glutamate Proteins 0.000 description 14
- 108010017391 lysylvaline Proteins 0.000 description 14
- 108010031719 prolyl-serine Proteins 0.000 description 14
- 201000002510 thyroid cancer Diseases 0.000 description 14
- 108091032973 (ribonucleotides)n+m Proteins 0.000 description 13
- 241000282414 Homo sapiens Species 0.000 description 13
- KAFOIVJDVSZUMD-UHFFFAOYSA-N Leu-Gln-Gln Natural products CC(C)CC(N)C(=O)NC(CCC(N)=O)C(=O)NC(CCC(N)=O)C(O)=O KAFOIVJDVSZUMD-UHFFFAOYSA-N 0.000 description 13
- 108010060035 arginylproline Proteins 0.000 description 13
- WKOBSJOZRJJVRZ-FXQIFTODSA-N Ala-Glu-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O WKOBSJOZRJJVRZ-FXQIFTODSA-N 0.000 description 12
- OTCJMMRQBVDQRK-DCAQKATOSA-N Arg-Asp-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O OTCJMMRQBVDQRK-DCAQKATOSA-N 0.000 description 12
- LRPXYSGPOBVBEH-IUCAKERBSA-N Glu-Gly-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)NCC(=O)N[C@@H](CC(C)C)C(O)=O LRPXYSGPOBVBEH-IUCAKERBSA-N 0.000 description 12
- FBEJIDRSQCGFJI-GUBZILKMSA-N Glu-Leu-Ser Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O FBEJIDRSQCGFJI-GUBZILKMSA-N 0.000 description 12
- KIEICAOUSNYOLM-NRPADANISA-N Glu-Val-Ala Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C)C(O)=O KIEICAOUSNYOLM-NRPADANISA-N 0.000 description 12
- 241000880493 Leptailurus serval Species 0.000 description 12
- KZNQNBZMBZJQJO-UHFFFAOYSA-N N-glycyl-L-proline Natural products NCC(=O)N1CCCC1C(O)=O KZNQNBZMBZJQJO-UHFFFAOYSA-N 0.000 description 12
- 108091000080 Phosphotransferase Proteins 0.000 description 12
- XQJCEKXQUJQNNK-ZLUOBGJFSA-N Ser-Ser-Ser Chemical compound OC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O XQJCEKXQUJQNNK-ZLUOBGJFSA-N 0.000 description 12
- 108010038633 aspartylglutamate Proteins 0.000 description 12
- 239000000872 buffer Substances 0.000 description 12
- 102000020233 phosphotransferase Human genes 0.000 description 12
- 210000001519 tissue Anatomy 0.000 description 12
- KASDBWKLWJKTLJ-GUBZILKMSA-N Glu-Glu-Met Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCSC)C(O)=O KASDBWKLWJKTLJ-GUBZILKMSA-N 0.000 description 11
- SWQALSGKVLYKDT-UHFFFAOYSA-N Gly-Ile-Ala Natural products NCC(=O)NC(C(C)CC)C(=O)NC(C)C(O)=O SWQALSGKVLYKDT-UHFFFAOYSA-N 0.000 description 11
- NSTUFLGQJCOCDL-UWVGGRQHSA-N Gly-Leu-Arg Chemical compound NCC(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N NSTUFLGQJCOCDL-UWVGGRQHSA-N 0.000 description 11
- XOWMDXHFSBCAKQ-SRVKXCTJSA-N Leu-Ser-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CC(C)C XOWMDXHFSBCAKQ-SRVKXCTJSA-N 0.000 description 11
- LIXBDERDAGNVAV-XKBZYTNZSA-N Thr-Gln-Ser Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(O)=O LIXBDERDAGNVAV-XKBZYTNZSA-N 0.000 description 11
- 108010069020 alanyl-prolyl-glycine Proteins 0.000 description 11
- 230000026731 phosphorylation Effects 0.000 description 11
- 238000006366 phosphorylation reaction Methods 0.000 description 11
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 11
- CCDFBRZVTDDJNM-GUBZILKMSA-N Ala-Leu-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O CCDFBRZVTDDJNM-GUBZILKMSA-N 0.000 description 10
- BEXGZLUHRXTZCC-CIUDSAMLSA-N Arg-Gln-Ser Chemical compound C(C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CO)C(=O)O)N)CN=C(N)N BEXGZLUHRXTZCC-CIUDSAMLSA-N 0.000 description 10
- PDECQIHABNQRHN-GUBZILKMSA-N Asp-Glu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CC(O)=O PDECQIHABNQRHN-GUBZILKMSA-N 0.000 description 10
- DWOGMPWRQQWPPF-GUBZILKMSA-N Asp-Leu-Glu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O DWOGMPWRQQWPPF-GUBZILKMSA-N 0.000 description 10
- 241000588724 Escherichia coli Species 0.000 description 10
- MCAVASRGVBVPMX-FXQIFTODSA-N Gln-Glu-Ala Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O MCAVASRGVBVPMX-FXQIFTODSA-N 0.000 description 10
- QKCZZAZNMMVICF-DCAQKATOSA-N Gln-Leu-Glu Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O QKCZZAZNMMVICF-DCAQKATOSA-N 0.000 description 10
- JJKKWYQVHRUSDG-GUBZILKMSA-N Glu-Ala-Lys Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCCN)C(O)=O JJKKWYQVHRUSDG-GUBZILKMSA-N 0.000 description 10
- VSRCAOIHMGCIJK-SRVKXCTJSA-N Glu-Leu-Arg Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O VSRCAOIHMGCIJK-SRVKXCTJSA-N 0.000 description 10
- IRXNJYPKBVERCW-DCAQKATOSA-N Glu-Leu-Glu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O IRXNJYPKBVERCW-DCAQKATOSA-N 0.000 description 10
- DLISPGXMKZTWQG-IFFSRLJSSA-N Glu-Thr-Val Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C(C)C)C(O)=O DLISPGXMKZTWQG-IFFSRLJSSA-N 0.000 description 10
- OUYCCCASQSFEME-QMMMGPOBSA-N L-tyrosine Chemical compound OC(=O)[C@@H](N)CC1=CC=C(O)C=C1 OUYCCCASQSFEME-QMMMGPOBSA-N 0.000 description 10
- XMBSYZWANAQXEV-UHFFFAOYSA-N N-alpha-L-glutamyl-L-phenylalanine Natural products OC(=O)CCC(N)C(=O)NC(C(O)=O)CC1=CC=CC=C1 XMBSYZWANAQXEV-UHFFFAOYSA-N 0.000 description 10
- 108010079364 N-glycylalanine Proteins 0.000 description 10
- LAYSXAOGWHKNED-XPUUQOCRSA-N Val-Gly-Ser Chemical compound CC(C)[C@H](N)C(=O)NCC(=O)N[C@@H](CO)C(O)=O LAYSXAOGWHKNED-XPUUQOCRSA-N 0.000 description 10
- AEMPCGRFEZTWIF-IHRRRGAJSA-N Val-Leu-Lys Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(O)=O AEMPCGRFEZTWIF-IHRRRGAJSA-N 0.000 description 10
- 108010069205 aspartyl-phenylalanine Proteins 0.000 description 10
- 108010089804 glycyl-threonine Proteins 0.000 description 10
- 108010077515 glycylproline Proteins 0.000 description 10
- 108010089198 phenylalanyl-prolyl-arginine Proteins 0.000 description 10
- 108010090894 prolylleucine Proteins 0.000 description 10
- OUYCCCASQSFEME-UHFFFAOYSA-N tyrosine Natural products OC(=O)C(N)CC1=CC=C(O)C=C1 OUYCCCASQSFEME-UHFFFAOYSA-N 0.000 description 10
- IOFVWPYSRSCWHI-JXUBOQSCSA-N Ala-Thr-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)O)NC(=O)[C@H](C)N IOFVWPYSRSCWHI-JXUBOQSCSA-N 0.000 description 9
- PHONAZGUEGIOEM-GLLZPBPUSA-N Glu-Glu-Thr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O PHONAZGUEGIOEM-GLLZPBPUSA-N 0.000 description 9
- BGWKULMLUIUPKY-BQBZGAKWSA-N Pro-Ser-Gly Chemical compound OC(=O)CNC(=O)[C@H](CO)NC(=O)[C@@H]1CCCN1 BGWKULMLUIUPKY-BQBZGAKWSA-N 0.000 description 9
- NLOAIFSWUUFQFR-CIUDSAMLSA-N Ser-Leu-Asp Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O NLOAIFSWUUFQFR-CIUDSAMLSA-N 0.000 description 9
- KEGBFULVYKYJRD-LFSVMHDDSA-N Thr-Ala-Phe Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 KEGBFULVYKYJRD-LFSVMHDDSA-N 0.000 description 9
- DEGCBBCMYWNJNA-RHYQMDGZSA-N Thr-Pro-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)[C@@H](C)O DEGCBBCMYWNJNA-RHYQMDGZSA-N 0.000 description 9
- 239000000284 extract Substances 0.000 description 9
- 108010056582 methionylglutamic acid Proteins 0.000 description 9
- 239000000203 mixture Substances 0.000 description 9
- 108010073969 valyllysine Proteins 0.000 description 9
- BUANFPRKJKJSRR-ACZMJKKPSA-N Ala-Ala-Gln Chemical compound C[C@H]([NH3+])C(=O)N[C@@H](C)C(=O)N[C@H](C([O-])=O)CCC(N)=O BUANFPRKJKJSRR-ACZMJKKPSA-N 0.000 description 8
- FJVAQLJNTSUQPY-CIUDSAMLSA-N Ala-Ala-Lys Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCCCN FJVAQLJNTSUQPY-CIUDSAMLSA-N 0.000 description 8
- LZRNYBIJOSKKRJ-XVYDVKMFSA-N Ala-Asp-His Chemical compound C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N LZRNYBIJOSKKRJ-XVYDVKMFSA-N 0.000 description 8
- GWFSQQNGMPGBEF-GHCJXIJMSA-N Ala-Asp-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](C)N GWFSQQNGMPGBEF-GHCJXIJMSA-N 0.000 description 8
- MNZHHDPWDWQJCQ-YUMQZZPRSA-N Ala-Leu-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)NCC(O)=O MNZHHDPWDWQJCQ-YUMQZZPRSA-N 0.000 description 8
- DHBKYZYFEXXUAK-ONGXEEELSA-N Ala-Phe-Gly Chemical compound OC(=O)CNC(=O)[C@@H](NC(=O)[C@@H](N)C)CC1=CC=CC=C1 DHBKYZYFEXXUAK-ONGXEEELSA-N 0.000 description 8
- NCQMBSJGJMYKCK-ZLUOBGJFSA-N Ala-Ser-Ser Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O NCQMBSJGJMYKCK-ZLUOBGJFSA-N 0.000 description 8
- NKBQZKVMKJJDLX-SRVKXCTJSA-N Arg-Glu-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O NKBQZKVMKJJDLX-SRVKXCTJSA-N 0.000 description 8
- NXDXECQFKHXHAM-HJGDQZAQSA-N Arg-Glu-Thr Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O NXDXECQFKHXHAM-HJGDQZAQSA-N 0.000 description 8
- LIVXPXUVXFRWNY-CIUDSAMLSA-N Asp-Lys-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(O)=O LIVXPXUVXFRWNY-CIUDSAMLSA-N 0.000 description 8
- 108091026890 Coding region Proteins 0.000 description 8
- IULKWYSYZSURJK-AVGNSLFASA-N Gln-Leu-Lys Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(O)=O IULKWYSYZSURJK-AVGNSLFASA-N 0.000 description 8
- PVBBEKPHARMPHX-DCAQKATOSA-N Glu-Gln-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(N)=O)NC(=O)[C@@H](N)CCC(O)=O PVBBEKPHARMPHX-DCAQKATOSA-N 0.000 description 8
- LGYZYFFDELZWRS-DCAQKATOSA-N Glu-Glu-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CCC(O)=O LGYZYFFDELZWRS-DCAQKATOSA-N 0.000 description 8
- OGNJZUXUTPQVBR-BQBZGAKWSA-N Glu-Gly-Glu Chemical compound OC(=O)CC[C@H](N)C(=O)NCC(=O)N[C@@H](CCC(O)=O)C(O)=O OGNJZUXUTPQVBR-BQBZGAKWSA-N 0.000 description 8
- ZQYZDDXTNQXUJH-CIUDSAMLSA-N Glu-Met-Ala Chemical compound C[C@@H](C(=O)O)NC(=O)[C@H](CCSC)NC(=O)[C@H](CCC(=O)O)N ZQYZDDXTNQXUJH-CIUDSAMLSA-N 0.000 description 8
- CUVBTVWFVIIDOC-YEPSODPASA-N Gly-Thr-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)O)NC(=O)CN CUVBTVWFVIIDOC-YEPSODPASA-N 0.000 description 8
- CQQGCWPXDHTTNF-GUBZILKMSA-N Leu-Ala-Glu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCC(O)=O CQQGCWPXDHTTNF-GUBZILKMSA-N 0.000 description 8
- HASRFYOMVPJRPU-SRVKXCTJSA-N Leu-Arg-Glu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCC(O)=O)C(O)=O HASRFYOMVPJRPU-SRVKXCTJSA-N 0.000 description 8
- FGNQZXKVAZIMCI-CIUDSAMLSA-N Leu-Asp-Cys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CS)C(=O)O)N FGNQZXKVAZIMCI-CIUDSAMLSA-N 0.000 description 8
- VQPPIMUZCZCOIL-GUBZILKMSA-N Leu-Gln-Ala Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(O)=O VQPPIMUZCZCOIL-GUBZILKMSA-N 0.000 description 8
- HQUXQAMSWFIRET-AVGNSLFASA-N Leu-Glu-Lys Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CCCCN HQUXQAMSWFIRET-AVGNSLFASA-N 0.000 description 8
- DSFYPIUSAMSERP-IHRRRGAJSA-N Leu-Leu-Arg Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N DSFYPIUSAMSERP-IHRRRGAJSA-N 0.000 description 8
- BJWKOATWNQJPSK-SRVKXCTJSA-N Leu-Met-Glu Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N BJWKOATWNQJPSK-SRVKXCTJSA-N 0.000 description 8
- HWMZUBUEOYAQSC-DCAQKATOSA-N Lys-Gln-Glu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O HWMZUBUEOYAQSC-DCAQKATOSA-N 0.000 description 8
- RBEATVHTWHTHTJ-KKUMJFAQSA-N Lys-Leu-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(O)=O RBEATVHTWHTHTJ-KKUMJFAQSA-N 0.000 description 8
- IOQWIOPSKJOEKI-SRVKXCTJSA-N Lys-Ser-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O IOQWIOPSKJOEKI-SRVKXCTJSA-N 0.000 description 8
- ZCXQTRXYZOSGJR-FXQIFTODSA-N Pro-Asp-Ser Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O ZCXQTRXYZOSGJR-FXQIFTODSA-N 0.000 description 8
- CGSOWZUPLOKYOR-AVGNSLFASA-N Pro-Pro-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@H]1NCCC1 CGSOWZUPLOKYOR-AVGNSLFASA-N 0.000 description 8
- KWMZPPWYBVZIER-XGEHTFHBSA-N Pro-Ser-Thr Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(O)=O KWMZPPWYBVZIER-XGEHTFHBSA-N 0.000 description 8
- VMLONWHIORGALA-SRVKXCTJSA-N Ser-Leu-Leu Chemical compound CC(C)C[C@@H](C([O-])=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H]([NH3+])CO VMLONWHIORGALA-SRVKXCTJSA-N 0.000 description 8
- AZWNCEBQZXELEZ-FXQIFTODSA-N Ser-Pro-Ser Chemical compound OC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O AZWNCEBQZXELEZ-FXQIFTODSA-N 0.000 description 8
- KERCOYANYUPLHJ-XGEHTFHBSA-N Thr-Pro-Ser Chemical compound C[C@@H](O)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O KERCOYANYUPLHJ-XGEHTFHBSA-N 0.000 description 8
- VUXIQSUQQYNLJP-XAVMHZPKSA-N Thr-Ser-Pro Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CO)C(=O)N1CCC[C@@H]1C(=O)O)N)O VUXIQSUQQYNLJP-XAVMHZPKSA-N 0.000 description 8
- UEHRGZCNLSWGHK-DLOVCJGASA-N Val-Glu-Val Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C(C)C)C(O)=O UEHRGZCNLSWGHK-DLOVCJGASA-N 0.000 description 8
- HGJRMXOWUWVUOA-GVXVVHGQSA-N Val-Leu-Gln Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](C(C)C)N HGJRMXOWUWVUOA-GVXVVHGQSA-N 0.000 description 8
- IECQJCJNPJVUSB-IHRRRGAJSA-N Val-Tyr-Ser Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](Cc1ccc(O)cc1)C(=O)N[C@@H](CO)C(O)=O IECQJCJNPJVUSB-IHRRRGAJSA-N 0.000 description 8
- 108010087924 alanylproline Proteins 0.000 description 8
- 108010001271 arginyl-glutamyl-arginine Proteins 0.000 description 8
- ONIQOQHATWINJY-UHFFFAOYSA-N cabozantinib Chemical compound C=12C=C(OC)C(OC)=CC2=NC=CC=1OC(C=C1)=CC=C1NC(=O)C1(C(=O)NC=2C=CC(F)=CC=2)CC1 ONIQOQHATWINJY-UHFFFAOYSA-N 0.000 description 8
- 108010085059 glutamyl-arginyl-proline Proteins 0.000 description 8
- 108010008237 glutamyl-valyl-glycine Proteins 0.000 description 8
- 108010049041 glutamylalanine Proteins 0.000 description 8
- 108010027668 glycyl-alanyl-valine Proteins 0.000 description 8
- 108010082286 glycyl-seryl-alanine Proteins 0.000 description 8
- 108010025306 histidylleucine Proteins 0.000 description 8
- 230000035772 mutation Effects 0.000 description 8
- 108010005652 splenotritin Proteins 0.000 description 8
- 108010061238 threonyl-glycine Proteins 0.000 description 8
- RLMISHABBKUNFO-WHFBIAKZSA-N Ala-Ala-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)NCC(O)=O RLMISHABBKUNFO-WHFBIAKZSA-N 0.000 description 7
- BLGHHPHXVJWCNK-GUBZILKMSA-N Ala-Gln-Leu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O BLGHHPHXVJWCNK-GUBZILKMSA-N 0.000 description 7
- YHKANGMVQWRMAP-DCAQKATOSA-N Ala-Leu-Arg Chemical compound C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N YHKANGMVQWRMAP-DCAQKATOSA-N 0.000 description 7
- HRGGPWBIMIQANI-GUBZILKMSA-N Asp-Gln-Leu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O HRGGPWBIMIQANI-GUBZILKMSA-N 0.000 description 7
- 102000004190 Enzymes Human genes 0.000 description 7
- 108090000790 Enzymes Proteins 0.000 description 7
- VPKBCVUDBNINAH-GARJFASQSA-N Glu-Arg-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCC(=O)O)N)C(=O)O VPKBCVUDBNINAH-GARJFASQSA-N 0.000 description 7
- HTTSBEBKVNEDFE-AUTRQRHGSA-N Glu-Gln-Val Chemical compound CC(C)[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CCC(=O)O)N HTTSBEBKVNEDFE-AUTRQRHGSA-N 0.000 description 7
- IVGJYOOGJLFKQE-AVGNSLFASA-N Glu-Leu-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCC(=O)O)N IVGJYOOGJLFKQE-AVGNSLFASA-N 0.000 description 7
- UGTHTQWIQKEDEH-BQBZGAKWSA-N L-alanyl-L-prolylglycine zwitterion Chemical compound C[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O UGTHTQWIQKEDEH-BQBZGAKWSA-N 0.000 description 7
- 239000002118 L01XE12 - Vandetanib Substances 0.000 description 7
- ILJREDZFPHTUIE-GUBZILKMSA-N Leu-Asp-Glu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O ILJREDZFPHTUIE-GUBZILKMSA-N 0.000 description 7
- LXKNSJLSGPNHSK-KKUMJFAQSA-N Leu-Leu-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)O)N LXKNSJLSGPNHSK-KKUMJFAQSA-N 0.000 description 7
- BIZNDKMFQHDOIE-KKUMJFAQSA-N Leu-Phe-Asn Chemical compound CC(C)C[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CC(N)=O)C(O)=O)CC1=CC=CC=C1 BIZNDKMFQHDOIE-KKUMJFAQSA-N 0.000 description 7
- NFLFJGGKOHYZJF-BJDJZHNGSA-N Lys-Ala-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCCCN NFLFJGGKOHYZJF-BJDJZHNGSA-N 0.000 description 7
- FKLSMYYLJHYPHH-UWVGGRQHSA-N Pro-Gly-Leu Chemical compound [H]N1CCC[C@H]1C(=O)NCC(=O)N[C@@H](CC(C)C)C(O)=O FKLSMYYLJHYPHH-UWVGGRQHSA-N 0.000 description 7
- 108010079005 RDV peptide Proteins 0.000 description 7
- BTPAWKABYQMKKN-LKXGYXEUSA-N Ser-Asp-Thr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O BTPAWKABYQMKKN-LKXGYXEUSA-N 0.000 description 7
- SDUBQHUJJWQTEU-XUXIUFHCSA-N Val-Ile-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](C(C)C)N SDUBQHUJJWQTEU-XUXIUFHCSA-N 0.000 description 7
- 108010091092 arginyl-glycyl-proline Proteins 0.000 description 7
- 108010047857 aspartylglycine Proteins 0.000 description 7
- 108010078144 glutaminyl-glycine Proteins 0.000 description 7
- 108010057821 leucylproline Proteins 0.000 description 7
- 108010025488 pinealon Proteins 0.000 description 7
- 239000013612 plasmid Substances 0.000 description 7
- 238000002360 preparation method Methods 0.000 description 7
- 108010077112 prolyl-proline Proteins 0.000 description 7
- 108010004914 prolylarginine Proteins 0.000 description 7
- 108010029020 prolylglycine Proteins 0.000 description 7
- -1 pyrimidine compound Chemical class 0.000 description 7
- 108010069117 seryl-lysyl-aspartic acid Proteins 0.000 description 7
- 238000009331 sowing Methods 0.000 description 7
- 238000001890 transfection Methods 0.000 description 7
- 108010035534 tyrosyl-leucyl-alanine Proteins 0.000 description 7
- 229960000241 vandetanib Drugs 0.000 description 7
- UHTHHESEBZOYNR-UHFFFAOYSA-N vandetanib Chemical compound COC1=CC(C(/N=CN2)=N/C=3C(=CC(Br)=CC=3)F)=C2C=C1OCC1CCN(C)CC1 UHTHHESEBZOYNR-UHFFFAOYSA-N 0.000 description 7
- LWUWMHIOBPTZBA-DCAQKATOSA-N Ala-Arg-Lys Chemical compound NC(=N)NCCC[C@H](NC(=O)[C@@H](N)C)C(=O)N[C@@H](CCCCN)C(O)=O LWUWMHIOBPTZBA-DCAQKATOSA-N 0.000 description 6
- NHCPCLJZRSIDHS-ZLUOBGJFSA-N Ala-Asp-Ala Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](C)C(O)=O NHCPCLJZRSIDHS-ZLUOBGJFSA-N 0.000 description 6
- BTYTYHBSJKQBQA-GCJQMDKQSA-N Ala-Asp-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](C)N)O BTYTYHBSJKQBQA-GCJQMDKQSA-N 0.000 description 6
- GGNHBHYDMUDXQB-KBIXCLLPSA-N Ala-Glu-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@H](C)N GGNHBHYDMUDXQB-KBIXCLLPSA-N 0.000 description 6
- PUBLUECXJRHTBK-ACZMJKKPSA-N Ala-Glu-Ser Chemical compound C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(O)=O PUBLUECXJRHTBK-ACZMJKKPSA-N 0.000 description 6
- ZVFVBBGVOILKPO-WHFBIAKZSA-N Ala-Gly-Ala Chemical compound C[C@H](N)C(=O)NCC(=O)N[C@@H](C)C(O)=O ZVFVBBGVOILKPO-WHFBIAKZSA-N 0.000 description 6
- KMGOBAQSCKTBGD-DLOVCJGASA-N Ala-His-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@H](C)N)CC1=CN=CN1 KMGOBAQSCKTBGD-DLOVCJGASA-N 0.000 description 6
- AWZKCUCQJNTBAD-SRVKXCTJSA-N Ala-Leu-Lys Chemical compound C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCCCN AWZKCUCQJNTBAD-SRVKXCTJSA-N 0.000 description 6
- MFMDKJIPHSWSBM-GUBZILKMSA-N Ala-Lys-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O MFMDKJIPHSWSBM-GUBZILKMSA-N 0.000 description 6
- ZBLQIYPCUWZSRZ-QEJZJMRPSA-N Ala-Phe-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@H](C)N)CC1=CC=CC=C1 ZBLQIYPCUWZSRZ-QEJZJMRPSA-N 0.000 description 6
- DCVYRWFAMZFSDA-ZLUOBGJFSA-N Ala-Ser-Ala Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O DCVYRWFAMZFSDA-ZLUOBGJFSA-N 0.000 description 6
- GXCSUJQOECMKPV-CIUDSAMLSA-N Arg-Ala-Gln Chemical compound C[C@H](NC(=O)[C@@H](N)CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(O)=O GXCSUJQOECMKPV-CIUDSAMLSA-N 0.000 description 6
- GFMWTFHOZGLTLC-AVGNSLFASA-N Arg-His-Met Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCSC)C(O)=O GFMWTFHOZGLTLC-AVGNSLFASA-N 0.000 description 6
- LCBSSOCDWUTQQV-SDDRHHMPSA-N Arg-Met-Pro Chemical compound CSCC[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N LCBSSOCDWUTQQV-SDDRHHMPSA-N 0.000 description 6
- BSYKSCBTTQKOJG-GUBZILKMSA-N Arg-Pro-Ala Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C)C(O)=O BSYKSCBTTQKOJG-GUBZILKMSA-N 0.000 description 6
- PJOPLXOCKACMLK-KKUMJFAQSA-N Arg-Tyr-Glu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCC(O)=O)C(O)=O PJOPLXOCKACMLK-KKUMJFAQSA-N 0.000 description 6
- PFOYSEIHFVKHNF-FXQIFTODSA-N Asn-Ala-Arg Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O PFOYSEIHFVKHNF-FXQIFTODSA-N 0.000 description 6
- BHQQRVARKXWXPP-ACZMJKKPSA-N Asn-Asp-Glu Chemical compound C(CC(=O)O)[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CC(=O)N)N BHQQRVARKXWXPP-ACZMJKKPSA-N 0.000 description 6
- GNKVBRYFXYWXAB-WDSKDSINSA-N Asn-Glu-Gly Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O GNKVBRYFXYWXAB-WDSKDSINSA-N 0.000 description 6
- PBVLJOIPOGUQQP-CIUDSAMLSA-N Asp-Ala-Leu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(O)=O PBVLJOIPOGUQQP-CIUDSAMLSA-N 0.000 description 6
- QHAJMRDEWNAIBQ-FXQIFTODSA-N Asp-Arg-Asn Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(O)=O QHAJMRDEWNAIBQ-FXQIFTODSA-N 0.000 description 6
- SARSTIZOZFBDOM-FXQIFTODSA-N Asp-Met-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](C)C(O)=O SARSTIZOZFBDOM-FXQIFTODSA-N 0.000 description 6
- JUWISGAGWSDGDH-KKUMJFAQSA-N Asp-Phe-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)CC(O)=O)CC1=CC=CC=C1 JUWISGAGWSDGDH-KKUMJFAQSA-N 0.000 description 6
- DCXGXDGGXVZVMY-GHCJXIJMSA-N Cys-Asn-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CS DCXGXDGGXVZVMY-GHCJXIJMSA-N 0.000 description 6
- REJJNXODKSHOKA-ACZMJKKPSA-N Gln-Ala-Asp Chemical compound C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)O)NC(=O)[C@H](CCC(=O)N)N REJJNXODKSHOKA-ACZMJKKPSA-N 0.000 description 6
- OYTPNWYZORARHL-XHNCKOQMSA-N Gln-Ala-Pro Chemical compound C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCC(=O)N)N OYTPNWYZORARHL-XHNCKOQMSA-N 0.000 description 6
- NKCZYEDZTKOFBG-GUBZILKMSA-N Gln-Gln-Arg Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O NKCZYEDZTKOFBG-GUBZILKMSA-N 0.000 description 6
- ZQPOVSJFBBETHQ-CIUDSAMLSA-N Gln-Glu-Gln Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O ZQPOVSJFBBETHQ-CIUDSAMLSA-N 0.000 description 6
- SNLOOPZHAQDMJG-CIUDSAMLSA-N Gln-Glu-Glu Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O SNLOOPZHAQDMJG-CIUDSAMLSA-N 0.000 description 6
- IKFZXRLDMYWNBU-YUMQZZPRSA-N Gln-Gly-Arg Chemical compound NC(=O)CC[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CCCN=C(N)N IKFZXRLDMYWNBU-YUMQZZPRSA-N 0.000 description 6
- MLSKFHLRFVGNLL-WDCWCFNPSA-N Gln-Leu-Thr Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O MLSKFHLRFVGNLL-WDCWCFNPSA-N 0.000 description 6
- VDMABHYXBULDGN-LAEOZQHASA-N Gln-Val-Asp Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O VDMABHYXBULDGN-LAEOZQHASA-N 0.000 description 6
- ITYRYNUZHPNCIK-GUBZILKMSA-N Glu-Ala-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(O)=O ITYRYNUZHPNCIK-GUBZILKMSA-N 0.000 description 6
- KKCUFHUTMKQQCF-SRVKXCTJSA-N Glu-Arg-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(O)=O KKCUFHUTMKQQCF-SRVKXCTJSA-N 0.000 description 6
- DYFJZDDQPNIPAB-NHCYSSNCSA-N Glu-Arg-Val Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C(C)C)C(O)=O DYFJZDDQPNIPAB-NHCYSSNCSA-N 0.000 description 6
- NKSGKPWXSWBRRX-ACZMJKKPSA-N Glu-Asn-Cys Chemical compound C(CC(=O)O)[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CS)C(=O)O)N NKSGKPWXSWBRRX-ACZMJKKPSA-N 0.000 description 6
- RQNYYRHRKSVKAB-GUBZILKMSA-N Glu-Cys-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CS)C(=O)N[C@@H](CC(C)C)C(O)=O RQNYYRHRKSVKAB-GUBZILKMSA-N 0.000 description 6
- ALCAUWPAMLVUDB-FXQIFTODSA-N Glu-Gln-Asn Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O ALCAUWPAMLVUDB-FXQIFTODSA-N 0.000 description 6
- HVYWQYLBVXMXSV-GUBZILKMSA-N Glu-Leu-Ala Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(O)=O HVYWQYLBVXMXSV-GUBZILKMSA-N 0.000 description 6
- JHSRJMUJOGLIHK-GUBZILKMSA-N Glu-Met-Glu Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)O)NC(=O)[C@H](CCC(=O)O)N JHSRJMUJOGLIHK-GUBZILKMSA-N 0.000 description 6
- CQGBSALYGOXQPE-HTUGSXCWSA-N Glu-Thr-Phe Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)O)NC(=O)[C@H](CCC(=O)O)N)O CQGBSALYGOXQPE-HTUGSXCWSA-N 0.000 description 6
- PUUYVMYCMIWHFE-BQBZGAKWSA-N Gly-Ala-Arg Chemical compound NCC(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N PUUYVMYCMIWHFE-BQBZGAKWSA-N 0.000 description 6
- JRDYDYXZKFNNRQ-XPUUQOCRSA-N Gly-Ala-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)CN JRDYDYXZKFNNRQ-XPUUQOCRSA-N 0.000 description 6
- TVDHVLGFJSHPAX-UWVGGRQHSA-N Gly-His-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)CN)CC1=CN=CN1 TVDHVLGFJSHPAX-UWVGGRQHSA-N 0.000 description 6
- NNCSJUBVFBDDLC-YUMQZZPRSA-N Gly-Leu-Ser Chemical compound NCC(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O NNCSJUBVFBDDLC-YUMQZZPRSA-N 0.000 description 6
- NVTPVQLIZCOJFK-FOHZUACHSA-N Gly-Thr-Asp Chemical compound [H]NCC(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(O)=O)C(O)=O NVTPVQLIZCOJFK-FOHZUACHSA-N 0.000 description 6
- 102100031181 Glyceraldehyde-3-phosphate dehydrogenase Human genes 0.000 description 6
- IMCHNUANCIGUKS-SRVKXCTJSA-N His-Glu-Arg Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O IMCHNUANCIGUKS-SRVKXCTJSA-N 0.000 description 6
- PUFNQIPSRXVLQJ-IHRRRGAJSA-N His-Val-Lys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CC1=CN=CN1)N PUFNQIPSRXVLQJ-IHRRRGAJSA-N 0.000 description 6
- IDAHFEPYTJJZFD-PEFMBERDSA-N Ile-Asp-Glu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N IDAHFEPYTJJZFD-PEFMBERDSA-N 0.000 description 6
- KUHFPGIVBOCRMV-MNXVOIDGSA-N Ile-Gln-Leu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CC(C)C)C(=O)O)N KUHFPGIVBOCRMV-MNXVOIDGSA-N 0.000 description 6
- PHIXPNQDGGILMP-YVNDNENWSA-N Ile-Glu-Glu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N PHIXPNQDGGILMP-YVNDNENWSA-N 0.000 description 6
- CMNMPCTVCWWYHY-MXAVVETBSA-N Ile-His-Leu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](CC(C)C)C(=O)O)N CMNMPCTVCWWYHY-MXAVVETBSA-N 0.000 description 6
- PRTZQMBYUZFSFA-XEGUGMAKSA-N Ile-Tyr-Gly Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)NCC(=O)O)N PRTZQMBYUZFSFA-XEGUGMAKSA-N 0.000 description 6
- KFKWRHQBZQICHA-STQMWFEESA-N L-leucyl-L-phenylalanine Natural products CC(C)C[C@H](N)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 KFKWRHQBZQICHA-STQMWFEESA-N 0.000 description 6
- GRZSCTXVCDUIPO-SRVKXCTJSA-N Leu-Arg-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(O)=O GRZSCTXVCDUIPO-SRVKXCTJSA-N 0.000 description 6
- KAFOIVJDVSZUMD-DCAQKATOSA-N Leu-Gln-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O KAFOIVJDVSZUMD-DCAQKATOSA-N 0.000 description 6
- ZTLGVASZOIKNIX-DCAQKATOSA-N Leu-Gln-Glu Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N ZTLGVASZOIKNIX-DCAQKATOSA-N 0.000 description 6
- FQZPTCNSNPWHLJ-AVGNSLFASA-N Leu-Gln-Lys Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCCN)C(O)=O FQZPTCNSNPWHLJ-AVGNSLFASA-N 0.000 description 6
- WIDZHJTYKYBLSR-DCAQKATOSA-N Leu-Glu-Glu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O WIDZHJTYKYBLSR-DCAQKATOSA-N 0.000 description 6
- ZFNLIDNJUWNIJL-WDCWCFNPSA-N Leu-Glu-Thr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O ZFNLIDNJUWNIJL-WDCWCFNPSA-N 0.000 description 6
- YFBBUHJJUXXZOF-UWVGGRQHSA-N Leu-Gly-Pro Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)N1CCC[C@H]1C(O)=O YFBBUHJJUXXZOF-UWVGGRQHSA-N 0.000 description 6
- AVEGDIAXTDVBJS-XUXIUFHCSA-N Leu-Ile-Arg Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O AVEGDIAXTDVBJS-XUXIUFHCSA-N 0.000 description 6
- ORWTWZXGDBYVCP-BJDJZHNGSA-N Leu-Ile-Cys Chemical compound SC[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)CC)NC(=O)[C@@H](N)CC(C)C ORWTWZXGDBYVCP-BJDJZHNGSA-N 0.000 description 6
- ZGUMORRUBUCXEH-AVGNSLFASA-N Leu-Lys-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(N)=O)C(O)=O ZGUMORRUBUCXEH-AVGNSLFASA-N 0.000 description 6
- LVTJJOJKDCVZGP-QWRGUYRKSA-N Leu-Lys-Gly Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)NCC(O)=O LVTJJOJKDCVZGP-QWRGUYRKSA-N 0.000 description 6
- GNRPTBRHRRZCMA-RWMBFGLXSA-N Leu-Met-Pro Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCSC)C(=O)N1CCC[C@@H]1C(=O)O)N GNRPTBRHRRZCMA-RWMBFGLXSA-N 0.000 description 6
- JIHDFWWRYHSAQB-GUBZILKMSA-N Leu-Ser-Glu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(O)=O JIHDFWWRYHSAQB-GUBZILKMSA-N 0.000 description 6
- VUBIPAHVHMZHCM-KKUMJFAQSA-N Leu-Tyr-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CO)C(O)=O)CC1=CC=C(O)C=C1 VUBIPAHVHMZHCM-KKUMJFAQSA-N 0.000 description 6
- PBIPLDMFHAICIP-DCAQKATOSA-N Lys-Glu-Glu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O PBIPLDMFHAICIP-DCAQKATOSA-N 0.000 description 6
- VQXAVLQBQJMENB-SRVKXCTJSA-N Lys-Glu-Met Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCSC)C(O)=O VQXAVLQBQJMENB-SRVKXCTJSA-N 0.000 description 6
- FHIAJWBDZVHLAH-YUMQZZPRSA-N Lys-Gly-Ser Chemical compound NCCCC[C@H](N)C(=O)NCC(=O)N[C@@H](CO)C(O)=O FHIAJWBDZVHLAH-YUMQZZPRSA-N 0.000 description 6
- FGMHXLULNHTPID-KKUMJFAQSA-N Lys-His-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CCCCN)C(O)=O)CC1=CN=CN1 FGMHXLULNHTPID-KKUMJFAQSA-N 0.000 description 6
- RIJCHEVHFWMDKD-SRVKXCTJSA-N Lys-Lys-Asn Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(N)=O)C(O)=O RIJCHEVHFWMDKD-SRVKXCTJSA-N 0.000 description 6
- LOGFVTREOLYCPF-RHYQMDGZSA-N Lys-Pro-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CCCCN LOGFVTREOLYCPF-RHYQMDGZSA-N 0.000 description 6
- ZVZRQKJOQQAFCF-ULQDDVLXSA-N Lys-Tyr-Arg Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O ZVZRQKJOQQAFCF-ULQDDVLXSA-N 0.000 description 6
- XDGFFEZAZHRZFR-RHYQMDGZSA-N Met-Leu-Thr Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O XDGFFEZAZHRZFR-RHYQMDGZSA-N 0.000 description 6
- FSTWDRPCQQUJIT-NHCYSSNCSA-N Met-Val-Glu Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)O)NC(=O)[C@H](CCSC)N FSTWDRPCQQUJIT-NHCYSSNCSA-N 0.000 description 6
- SITLTJHOQZFJGG-UHFFFAOYSA-N N-L-alpha-glutamyl-L-valine Natural products CC(C)C(C(O)=O)NC(=O)C(N)CCC(O)=O SITLTJHOQZFJGG-UHFFFAOYSA-N 0.000 description 6
- 108700019961 Neoplasm Genes Proteins 0.000 description 6
- 102000048850 Neoplasm Genes Human genes 0.000 description 6
- BKWJQWJPZMUWEG-LFSVMHDDSA-N Phe-Ala-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CC1=CC=CC=C1 BKWJQWJPZMUWEG-LFSVMHDDSA-N 0.000 description 6
- KZRQONDKKJCAOL-DKIMLUQUSA-N Phe-Leu-Ile Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O KZRQONDKKJCAOL-DKIMLUQUSA-N 0.000 description 6
- DNAXXTQSTKOHFO-QEJZJMRPSA-N Phe-Lys-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CC1=CC=CC=C1 DNAXXTQSTKOHFO-QEJZJMRPSA-N 0.000 description 6
- AJLVKXCNXIJHDV-CIUDSAMLSA-N Pro-Ala-Gln Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(O)=O AJLVKXCNXIJHDV-CIUDSAMLSA-N 0.000 description 6
- DXTOOBDIIAJZBJ-BQBZGAKWSA-N Pro-Gly-Ser Chemical compound [H]N1CCC[C@H]1C(=O)NCC(=O)N[C@@H](CO)C(O)=O DXTOOBDIIAJZBJ-BQBZGAKWSA-N 0.000 description 6
- RFWXYTJSVDUBBZ-DCAQKATOSA-N Pro-Pro-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@H]1NCCC1 RFWXYTJSVDUBBZ-DCAQKATOSA-N 0.000 description 6
- KHRLUIPIMIQFGT-AVGNSLFASA-N Pro-Val-Leu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O KHRLUIPIMIQFGT-AVGNSLFASA-N 0.000 description 6
- PVDTYLHUWAEYGY-CIUDSAMLSA-N Ser-Glu-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O PVDTYLHUWAEYGY-CIUDSAMLSA-N 0.000 description 6
- YQQKYAZABFEYAF-FXQIFTODSA-N Ser-Glu-Gln Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O YQQKYAZABFEYAF-FXQIFTODSA-N 0.000 description 6
- UIGMAMGZOJVTDN-WHFBIAKZSA-N Ser-Gly-Ser Chemical compound OC[C@H](N)C(=O)NCC(=O)N[C@@H](CO)C(O)=O UIGMAMGZOJVTDN-WHFBIAKZSA-N 0.000 description 6
- NNFMANHDYSVNIO-DCAQKATOSA-N Ser-Lys-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O NNFMANHDYSVNIO-DCAQKATOSA-N 0.000 description 6
- OCWWJBZQXGYQCA-DCAQKATOSA-N Ser-Lys-Met Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCSC)C(O)=O OCWWJBZQXGYQCA-DCAQKATOSA-N 0.000 description 6
- GYDFRTRSSXOZCR-ACZMJKKPSA-N Ser-Ser-Glu Chemical compound OC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(O)=O GYDFRTRSSXOZCR-ACZMJKKPSA-N 0.000 description 6
- PURRNJBBXDDWLX-ZDLURKLDSA-N Ser-Thr-Gly Chemical compound C[C@H]([C@@H](C(=O)NCC(=O)O)NC(=O)[C@H](CO)N)O PURRNJBBXDDWLX-ZDLURKLDSA-N 0.000 description 6
- BEBVVQPDSHHWQL-NRPADANISA-N Ser-Val-Glu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O BEBVVQPDSHHWQL-NRPADANISA-N 0.000 description 6
- MQCPGOZXFSYJPS-KZVJFYERSA-N Thr-Ala-Arg Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O MQCPGOZXFSYJPS-KZVJFYERSA-N 0.000 description 6
- JHBHMCMKSPXRHV-NUMRIWBASA-N Thr-Asn-Gln Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N)O JHBHMCMKSPXRHV-NUMRIWBASA-N 0.000 description 6
- LMMDEZPNUTZJAY-GCJQMDKQSA-N Thr-Asp-Ala Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](C)C(O)=O LMMDEZPNUTZJAY-GCJQMDKQSA-N 0.000 description 6
- XIULAFZYEKSGAJ-IXOXFDKPSA-N Thr-Leu-His Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CC1=CNC=N1 XIULAFZYEKSGAJ-IXOXFDKPSA-N 0.000 description 6
- PUEWAXRPXOEQOW-HJGDQZAQSA-N Thr-Met-Gln Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCC(N)=O)C(O)=O PUEWAXRPXOEQOW-HJGDQZAQSA-N 0.000 description 6
- BKIOKSLLAAZYTC-KKHAAJSZSA-N Thr-Val-Asn Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O BKIOKSLLAAZYTC-KKHAAJSZSA-N 0.000 description 6
- RIJPHPUJRLEOAK-JYJNAYRXSA-N Tyr-Gln-His Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)N)O RIJPHPUJRLEOAK-JYJNAYRXSA-N 0.000 description 6
- ZOBLBMGJKVJVEV-BZSNNMDCSA-N Tyr-Lys-Lys Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)O)N)O ZOBLBMGJKVJVEV-BZSNNMDCSA-N 0.000 description 6
- SCBITHMBEJNRHC-LSJOCFKGSA-N Val-Asp-Val Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](C(C)C)C(=O)O)N SCBITHMBEJNRHC-LSJOCFKGSA-N 0.000 description 6
- WDIGUPHXPBMODF-UMNHJUIQSA-N Val-Glu-Pro Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N1CCC[C@@H]1C(=O)O)N WDIGUPHXPBMODF-UMNHJUIQSA-N 0.000 description 6
- DJEVQCWNMQOABE-RCOVLWMOSA-N Val-Gly-Asp Chemical compound CC(C)[C@@H](C(=O)NCC(=O)N[C@@H](CC(=O)O)C(=O)O)N DJEVQCWNMQOABE-RCOVLWMOSA-N 0.000 description 6
- KZSNJWFQEVHDMF-UHFFFAOYSA-N Valine Natural products CC(C)C(N)C(O)=O KZSNJWFQEVHDMF-UHFFFAOYSA-N 0.000 description 6
- 108010008685 alanyl-glutamyl-aspartic acid Proteins 0.000 description 6
- 108010041407 alanylaspartic acid Proteins 0.000 description 6
- 238000011304 droplet digital PCR Methods 0.000 description 6
- 108020004445 glyceraldehyde-3-phosphate dehydrogenase Proteins 0.000 description 6
- 108010023364 glycyl-histidyl-arginine Proteins 0.000 description 6
- 238000011534 incubation Methods 0.000 description 6
- 108010031424 isoleucyl-prolyl-proline Proteins 0.000 description 6
- 108010044056 leucyl-phenylalanine Proteins 0.000 description 6
- 239000013642 negative control Substances 0.000 description 6
- 238000011580 nude mouse model Methods 0.000 description 6
- 108010070643 prolylglutamic acid Proteins 0.000 description 6
- 108010053725 prolylvaline Proteins 0.000 description 6
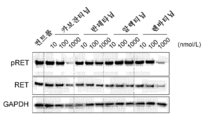
- 238000001262 western blot Methods 0.000 description 6
- PAIHPOGPJVUFJY-WDSKDSINSA-N Ala-Glu-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O PAIHPOGPJVUFJY-WDSKDSINSA-N 0.000 description 5
- HXNNRBHASOSVPG-GUBZILKMSA-N Ala-Glu-Leu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O HXNNRBHASOSVPG-GUBZILKMSA-N 0.000 description 5
- CKLDHDOIYBVUNP-KBIXCLLPSA-N Ala-Ile-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(O)=O)C(O)=O CKLDHDOIYBVUNP-KBIXCLLPSA-N 0.000 description 5
- SUMYEVXWCAYLLJ-GUBZILKMSA-N Ala-Leu-Gln Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O SUMYEVXWCAYLLJ-GUBZILKMSA-N 0.000 description 5
- PVQLRJRPUTXFFX-CIUDSAMLSA-N Ala-Met-Gln Chemical compound CSCC[C@H](NC(=O)[C@H](C)N)C(=O)N[C@@H](CCC(N)=O)C(O)=O PVQLRJRPUTXFFX-CIUDSAMLSA-N 0.000 description 5
- NOZYDJOPOGKUSR-AVGNSLFASA-N Arg-Leu-Met Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCSC)C(O)=O NOZYDJOPOGKUSR-AVGNSLFASA-N 0.000 description 5
- SPIPSJXLZVTXJL-ZLUOBGJFSA-N Asn-Cys-Ser Chemical compound NC(=O)C[C@H](N)C(=O)N[C@@H](CS)C(=O)N[C@@H](CO)C(O)=O SPIPSJXLZVTXJL-ZLUOBGJFSA-N 0.000 description 5
- GQRDIVQPSMPQME-ZPFDUUQYSA-N Asn-Ile-Leu Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(C)C)C(O)=O GQRDIVQPSMPQME-ZPFDUUQYSA-N 0.000 description 5
- XLDMSQYOYXINSZ-QXEWZRGKSA-N Asn-Val-Arg Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)NC(=O)[C@H](CC(=O)N)N XLDMSQYOYXINSZ-QXEWZRGKSA-N 0.000 description 5
- WSWYMRLTJVKRCE-ZLUOBGJFSA-N Asp-Ala-Asp Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(O)=O)C(O)=O WSWYMRLTJVKRCE-ZLUOBGJFSA-N 0.000 description 5
- JNNVNVRBYUJYGS-CIUDSAMLSA-N Asp-Leu-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(O)=O JNNVNVRBYUJYGS-CIUDSAMLSA-N 0.000 description 5
- RJPKQCFHEPPTGL-ZLUOBGJFSA-N Cys-Ser-Asp Chemical compound [H]N[C@@H](CS)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O RJPKQCFHEPPTGL-ZLUOBGJFSA-N 0.000 description 5
- 108010090461 DFG peptide Proteins 0.000 description 5
- KVYVOGYEMPEXBT-GUBZILKMSA-N Gln-Ala-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCC(N)=O KVYVOGYEMPEXBT-GUBZILKMSA-N 0.000 description 5
- ITZWDGBYBPUZRG-KBIXCLLPSA-N Gln-Ile-Ser Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CO)C(O)=O ITZWDGBYBPUZRG-KBIXCLLPSA-N 0.000 description 5
- XFAUJGNLHIGXET-AVGNSLFASA-N Gln-Leu-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O XFAUJGNLHIGXET-AVGNSLFASA-N 0.000 description 5
- LKDIBBOKUAASNP-FXQIFTODSA-N Glu-Ala-Glu Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(O)=O LKDIBBOKUAASNP-FXQIFTODSA-N 0.000 description 5
- VTTSANCGJWLPNC-ZPFDUUQYSA-N Glu-Arg-Ile Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O VTTSANCGJWLPNC-ZPFDUUQYSA-N 0.000 description 5
- BUZMZDDKFCSKOT-CIUDSAMLSA-N Glu-Glu-Glu Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O BUZMZDDKFCSKOT-CIUDSAMLSA-N 0.000 description 5
- SUIAHERNFYRBDZ-GVXVVHGQSA-N Glu-Lys-Val Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C(C)C)C(O)=O SUIAHERNFYRBDZ-GVXVVHGQSA-N 0.000 description 5
- YQAQQKPWFOBSMU-WDCWCFNPSA-N Glu-Thr-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(O)=O YQAQQKPWFOBSMU-WDCWCFNPSA-N 0.000 description 5
- PYTZFYUXZZHOAD-WHFBIAKZSA-N Gly-Ala-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](C)NC(=O)CN PYTZFYUXZZHOAD-WHFBIAKZSA-N 0.000 description 5
- KFMBRBPXHVMDFN-UWVGGRQHSA-N Gly-Arg-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)CN)CCCNC(N)=N KFMBRBPXHVMDFN-UWVGGRQHSA-N 0.000 description 5
- HFXJIZNEXNIZIJ-BQBZGAKWSA-N Gly-Glu-Gln Chemical compound NCC(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O HFXJIZNEXNIZIJ-BQBZGAKWSA-N 0.000 description 5
- NSVOVKWEKGEOQB-LURJTMIESA-N Gly-Pro-Gly Chemical compound NCC(=O)N1CCC[C@H]1C(=O)NCC(O)=O NSVOVKWEKGEOQB-LURJTMIESA-N 0.000 description 5
- OOCFXNOVSLSHAB-IUCAKERBSA-N Gly-Pro-Pro Chemical compound NCC(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 OOCFXNOVSLSHAB-IUCAKERBSA-N 0.000 description 5
- WNGHUXFWEWTKAO-YUMQZZPRSA-N Gly-Ser-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)CN WNGHUXFWEWTKAO-YUMQZZPRSA-N 0.000 description 5
- AQCUAZTZSPQJFF-ZKWXMUAHSA-N Ile-Ala-Gly Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](C)C(=O)NCC(O)=O AQCUAZTZSPQJFF-ZKWXMUAHSA-N 0.000 description 5
- CYHYBSGMHMHKOA-CIQUZCHMSA-N Ile-Ala-Thr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(=O)O)N CYHYBSGMHMHKOA-CIQUZCHMSA-N 0.000 description 5
- ASCFJMSGKUIRDU-ZPFDUUQYSA-N Ile-Arg-Gln Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(O)=O ASCFJMSGKUIRDU-ZPFDUUQYSA-N 0.000 description 5
- IITVUURPOYGCTD-NAKRPEOUSA-N Ile-Pro-Ala Chemical compound CC[C@H](C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C)C(O)=O IITVUURPOYGCTD-NAKRPEOUSA-N 0.000 description 5
- XMYURPUVJSKTMC-KBIXCLLPSA-N Ile-Ser-Gln Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N XMYURPUVJSKTMC-KBIXCLLPSA-N 0.000 description 5
- XBBKIIGCUMBKCO-JXUBOQSCSA-N Leu-Ala-Thr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O XBBKIIGCUMBKCO-JXUBOQSCSA-N 0.000 description 5
- YOZCKMXHBYKOMQ-IHRRRGAJSA-N Leu-Arg-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCCN)C(=O)O)N YOZCKMXHBYKOMQ-IHRRRGAJSA-N 0.000 description 5
- HRTRLSRYZZKPCO-BJDJZHNGSA-N Leu-Ile-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CO)C(O)=O HRTRLSRYZZKPCO-BJDJZHNGSA-N 0.000 description 5
- IWMJFLJQHIDZQW-KKUMJFAQSA-N Leu-Ser-Phe Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 IWMJFLJQHIDZQW-KKUMJFAQSA-N 0.000 description 5
- UCRJTSIIAYHOHE-ULQDDVLXSA-N Leu-Tyr-Arg Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N UCRJTSIIAYHOHE-ULQDDVLXSA-N 0.000 description 5
- VKVDRTGWLVZJOM-DCAQKATOSA-N Leu-Val-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CO)C(O)=O VKVDRTGWLVZJOM-DCAQKATOSA-N 0.000 description 5
- VUTWYNQUSJWBHO-BZSNNMDCSA-N Lys-Leu-Tyr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O VUTWYNQUSJWBHO-BZSNNMDCSA-N 0.000 description 5
- RMOKGALPSPOYKE-KATARQTJSA-N Lys-Thr-Ser Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(O)=O RMOKGALPSPOYKE-KATARQTJSA-N 0.000 description 5
- HOZNVKDCKZPRER-XUXIUFHCSA-N Met-Lys-Ile Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O HOZNVKDCKZPRER-XUXIUFHCSA-N 0.000 description 5
- 239000002033 PVDF binder Substances 0.000 description 5
- FGXIJNMDRCZVDE-KKUMJFAQSA-N Phe-Cys-Lys Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CS)C(=O)N[C@@H](CCCCN)C(=O)O)N FGXIJNMDRCZVDE-KKUMJFAQSA-N 0.000 description 5
- DZZCICYRSZASNF-FXQIFTODSA-N Pro-Ala-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1 DZZCICYRSZASNF-FXQIFTODSA-N 0.000 description 5
- MCWHYUWXVNRXFV-RWMBFGLXSA-N Pro-Leu-Pro Chemical compound CC(C)C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@@H]2CCCN2 MCWHYUWXVNRXFV-RWMBFGLXSA-N 0.000 description 5
- XWCYBVBLJRWOFR-WDSKDSINSA-N Ser-Gln-Gly Chemical compound OC[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)NCC(O)=O XWCYBVBLJRWOFR-WDSKDSINSA-N 0.000 description 5
- SMIDBHKWSYUBRZ-ACZMJKKPSA-N Ser-Glu-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O SMIDBHKWSYUBRZ-ACZMJKKPSA-N 0.000 description 5
- LALNXSXEYFUUDD-GUBZILKMSA-N Ser-Glu-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O LALNXSXEYFUUDD-GUBZILKMSA-N 0.000 description 5
- PTWIYDNFWPXQSD-GARJFASQSA-N Ser-Lys-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCCN)NC(=O)[C@H](CO)N)C(=O)O PTWIYDNFWPXQSD-GARJFASQSA-N 0.000 description 5
- NQZFFLBPNDLTPO-DLOVCJGASA-N Ser-Phe-Ala Chemical compound C[C@@H](C(=O)O)NC(=O)[C@H](CC1=CC=CC=C1)NC(=O)[C@H](CO)N NQZFFLBPNDLTPO-DLOVCJGASA-N 0.000 description 5
- RFKVQLIXNVEOMB-WEDXCCLWSA-N Thr-Leu-Gly Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC(C)C)C(=O)NCC(=O)O)N)O RFKVQLIXNVEOMB-WEDXCCLWSA-N 0.000 description 5
- YJVJPJPHHFOVMG-VEVYYDQMSA-N Thr-Met-Asp Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(=O)O)C(=O)O)N)O YJVJPJPHHFOVMG-VEVYYDQMSA-N 0.000 description 5
- YFOCMOVJBQDBCE-NRPADANISA-N Val-Ala-Glu Chemical compound C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)O)NC(=O)[C@H](C(C)C)N YFOCMOVJBQDBCE-NRPADANISA-N 0.000 description 5
- GVJUTBOZZBTBIG-AVGNSLFASA-N Val-Lys-Arg Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N GVJUTBOZZBTBIG-AVGNSLFASA-N 0.000 description 5
- 108010081404 acein-2 Proteins 0.000 description 5
- 108010029539 arginyl-prolyl-proline Proteins 0.000 description 5
- 239000012472 biological sample Substances 0.000 description 5
- 239000003153 chemical reaction reagent Substances 0.000 description 5
- 238000007796 conventional method Methods 0.000 description 5
- 238000010195 expression analysis Methods 0.000 description 5
- 238000010362 genome editing Methods 0.000 description 5
- XBGGUPMXALFZOT-UHFFFAOYSA-N glycyl-L-tyrosine hemihydrate Natural products NCC(=O)NC(C(O)=O)CC1=CC=C(O)C=C1 XBGGUPMXALFZOT-UHFFFAOYSA-N 0.000 description 5
- 108010044311 leucyl-glycyl-glycine Proteins 0.000 description 5
- 108010073472 leucyl-prolyl-proline Proteins 0.000 description 5
- 108010003700 lysyl aspartic acid Proteins 0.000 description 5
- 108010064235 lysylglycine Proteins 0.000 description 5
- 238000002156 mixing Methods 0.000 description 5
- 238000007857 nested PCR Methods 0.000 description 5
- 108010024607 phenylalanylalanine Proteins 0.000 description 5
- 229920002981 polyvinylidene fluoride Polymers 0.000 description 5
- 108010048818 seryl-histidine Proteins 0.000 description 5
- 230000008685 targeting Effects 0.000 description 5
- GJLXVWOMRRWCIB-MERZOTPQSA-N (2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-acetamido-5-(diaminomethylideneamino)pentanoyl]amino]-3-(4-hydroxyphenyl)propanoyl]amino]-3-(4-hydroxyphenyl)propanoyl]amino]-5-(diaminomethylideneamino)pentanoyl]amino]-3-(1H-indol-3-yl)propanoyl]amino]-6-aminohexanoyl]amino]-6-aminohexanoyl]amino]-6-aminohexanoyl]amino]-6-aminohexanoyl]amino]-6-aminohexanoyl]amino]-6-aminohexanoyl]amino]-6-aminohexanamide Chemical compound C([C@H](NC(=O)[C@H](CCCN=C(N)N)NC(=O)C)C(=O)N[C@@H](CC=1C=CC(O)=CC=1)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CC=1C2=CC=CC=C2NC=1)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(N)=O)C1=CC=C(O)C=C1 GJLXVWOMRRWCIB-MERZOTPQSA-N 0.000 description 4
- BRPMXFSTKXXNHF-IUCAKERBSA-N (2s)-1-[2-[[(2s)-pyrrolidine-2-carbonyl]amino]acetyl]pyrrolidine-2-carboxylic acid Chemical compound OC(=O)[C@@H]1CCCN1C(=O)CNC(=O)[C@H]1NCCC1 BRPMXFSTKXXNHF-IUCAKERBSA-N 0.000 description 4
- QAPSNMNOIOSXSQ-YNEHKIRRSA-N 1-[(2r,4s,5r)-4-[tert-butyl(dimethyl)silyl]oxy-5-(hydroxymethyl)oxolan-2-yl]-5-methylpyrimidine-2,4-dione Chemical compound O=C1NC(=O)C(C)=CN1[C@@H]1O[C@H](CO)[C@@H](O[Si](C)(C)C(C)(C)C)C1 QAPSNMNOIOSXSQ-YNEHKIRRSA-N 0.000 description 4
- CXRCVCURMBFFOL-FXQIFTODSA-N Ala-Ala-Pro Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)N1CCC[C@H]1C(O)=O CXRCVCURMBFFOL-FXQIFTODSA-N 0.000 description 4
- YYSWCHMLFJLLBJ-ZLUOBGJFSA-N Ala-Ala-Ser Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CO)C(O)=O YYSWCHMLFJLLBJ-ZLUOBGJFSA-N 0.000 description 4
- KUDREHRZRIVKHS-UWJYBYFXSA-N Ala-Asp-Tyr Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O KUDREHRZRIVKHS-UWJYBYFXSA-N 0.000 description 4
- LGFCAXJBAZESCF-ACZMJKKPSA-N Ala-Gln-Ala Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(O)=O LGFCAXJBAZESCF-ACZMJKKPSA-N 0.000 description 4
- AWAXZRDKUHOPBO-GUBZILKMSA-N Ala-Gln-Lys Chemical compound C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCCN)C(O)=O AWAXZRDKUHOPBO-GUBZILKMSA-N 0.000 description 4
- CRWFEKLFPVRPBV-CIUDSAMLSA-N Ala-Gln-Met Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCSC)C(O)=O CRWFEKLFPVRPBV-CIUDSAMLSA-N 0.000 description 4
- FUSPCLTUKXQREV-ACZMJKKPSA-N Ala-Glu-Ala Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O FUSPCLTUKXQREV-ACZMJKKPSA-N 0.000 description 4
- BGNLUHXLSAQYRQ-FXQIFTODSA-N Ala-Glu-Gln Chemical compound C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O BGNLUHXLSAQYRQ-FXQIFTODSA-N 0.000 description 4
- XYTNPQNAZREREP-XQXXSGGOSA-N Ala-Glu-Thr Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O XYTNPQNAZREREP-XQXXSGGOSA-N 0.000 description 4
- RZZMZYZXNJRPOJ-BJDJZHNGSA-N Ala-Ile-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](C)N RZZMZYZXNJRPOJ-BJDJZHNGSA-N 0.000 description 4
- OKIKVSXTXVVFDV-MMWGEVLESA-N Ala-Ile-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](C)N OKIKVSXTXVVFDV-MMWGEVLESA-N 0.000 description 4
- SOBIAADAMRHGKH-CIUDSAMLSA-N Ala-Leu-Ser Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O SOBIAADAMRHGKH-CIUDSAMLSA-N 0.000 description 4
- SUHLZMHFRALVSY-YUMQZZPRSA-N Ala-Lys-Gly Chemical compound NCCCC[C@H](NC(=O)[C@@H](N)C)C(=O)NCC(O)=O SUHLZMHFRALVSY-YUMQZZPRSA-N 0.000 description 4
- PMQXMXAASGFUDX-SRVKXCTJSA-N Ala-Lys-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@H](C)N)CCCCN PMQXMXAASGFUDX-SRVKXCTJSA-N 0.000 description 4
- WQLDNOCHHRISMS-NAKRPEOUSA-N Ala-Pro-Ile Chemical compound [H]N[C@@H](C)C(=O)N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)CC)C(O)=O WQLDNOCHHRISMS-NAKRPEOUSA-N 0.000 description 4
- ADSGHMXEAZJJNF-DCAQKATOSA-N Ala-Pro-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@H](C)N ADSGHMXEAZJJNF-DCAQKATOSA-N 0.000 description 4
- HOVPGJUNRLMIOZ-CIUDSAMLSA-N Ala-Ser-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@H](C)N HOVPGJUNRLMIOZ-CIUDSAMLSA-N 0.000 description 4
- MMLHRUJLOUSRJX-CIUDSAMLSA-N Ala-Ser-Lys Chemical compound C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCCCN MMLHRUJLOUSRJX-CIUDSAMLSA-N 0.000 description 4
- OEVCHROQUIVQFZ-YTLHQDLWSA-N Ala-Thr-Ala Chemical compound C[C@H](N)C(=O)N[C@@H]([C@H](O)C)C(=O)N[C@@H](C)C(O)=O OEVCHROQUIVQFZ-YTLHQDLWSA-N 0.000 description 4
- YEBZNKPPOHFZJM-BPNCWPANSA-N Ala-Tyr-Val Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](C(C)C)C(O)=O YEBZNKPPOHFZJM-BPNCWPANSA-N 0.000 description 4
- SBVJJNJLFWSJOV-UBHSHLNASA-N Arg-Ala-Phe Chemical compound NC(=N)NCCC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 SBVJJNJLFWSJOV-UBHSHLNASA-N 0.000 description 4
- KBBKCNHWCDJPGN-GUBZILKMSA-N Arg-Gln-Gln Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O KBBKCNHWCDJPGN-GUBZILKMSA-N 0.000 description 4
- PBSOQGZLPFVXPU-YUMQZZPRSA-N Arg-Glu-Gly Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O PBSOQGZLPFVXPU-YUMQZZPRSA-N 0.000 description 4
- AQPVUEJJARLJHB-BQBZGAKWSA-N Arg-Gly-Ala Chemical compound OC(=O)[C@H](C)NC(=O)CNC(=O)[C@@H](N)CCCN=C(N)N AQPVUEJJARLJHB-BQBZGAKWSA-N 0.000 description 4
- LVMUGODRNHFGRA-AVGNSLFASA-N Arg-Leu-Arg Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O LVMUGODRNHFGRA-AVGNSLFASA-N 0.000 description 4
- OMKZPCPZEFMBIT-SRVKXCTJSA-N Arg-Met-Arg Chemical compound NC(=N)NCCC[C@H](N)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O OMKZPCPZEFMBIT-SRVKXCTJSA-N 0.000 description 4
- JCROZIFVIYMXHM-GUBZILKMSA-N Arg-Met-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](CCSC)NC(=O)[C@@H](N)CCCN=C(N)N JCROZIFVIYMXHM-GUBZILKMSA-N 0.000 description 4
- BDMIFVIWCNLDCT-CIUDSAMLSA-N Asn-Arg-Glu Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(O)=O BDMIFVIWCNLDCT-CIUDSAMLSA-N 0.000 description 4
- HDHZCEDPLTVHFZ-GUBZILKMSA-N Asn-Leu-Glu Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O HDHZCEDPLTVHFZ-GUBZILKMSA-N 0.000 description 4
- ORJQQZIXTOYGGH-SRVKXCTJSA-N Asn-Lys-Leu Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O ORJQQZIXTOYGGH-SRVKXCTJSA-N 0.000 description 4
- NPZJLGMWMDNQDD-GHCJXIJMSA-N Asn-Ser-Ile Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O NPZJLGMWMDNQDD-GHCJXIJMSA-N 0.000 description 4
- XEDQMTWEYFBOIK-ACZMJKKPSA-N Asp-Ala-Glu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(O)=O XEDQMTWEYFBOIK-ACZMJKKPSA-N 0.000 description 4
- FMWHSNJMHUNLAG-FXQIFTODSA-N Asp-Cys-Arg Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CS)C(=O)N[C@H](C(O)=O)CCCN=C(N)N FMWHSNJMHUNLAG-FXQIFTODSA-N 0.000 description 4
- DTNUIAJCPRMNBT-WHFBIAKZSA-N Asp-Gly-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)NCC(=O)N[C@@H](C)C(O)=O DTNUIAJCPRMNBT-WHFBIAKZSA-N 0.000 description 4
- OGTCOKZFOJIZFG-CIUDSAMLSA-N Asp-His-Asp Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(O)=O)C(O)=O OGTCOKZFOJIZFG-CIUDSAMLSA-N 0.000 description 4
- KTTCQQNRRLCIBC-GHCJXIJMSA-N Asp-Ile-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](C)C(O)=O KTTCQQNRRLCIBC-GHCJXIJMSA-N 0.000 description 4
- UMHUHHJMEXNSIV-CIUDSAMLSA-N Asp-Leu-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC(O)=O UMHUHHJMEXNSIV-CIUDSAMLSA-N 0.000 description 4
- PLOKOIJSGCISHE-BYULHYEWSA-N Asp-Val-Asn Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O PLOKOIJSGCISHE-BYULHYEWSA-N 0.000 description 4
- JGLWFWXGOINXEA-YDHLFZDLSA-N Asp-Val-Tyr Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 JGLWFWXGOINXEA-YDHLFZDLSA-N 0.000 description 4
- GSNRZJNHMVMOFV-ACZMJKKPSA-N Cys-Asp-Glu Chemical compound C(CC(=O)O)[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CS)N GSNRZJNHMVMOFV-ACZMJKKPSA-N 0.000 description 4
- HJGUQJJJXQGXGJ-FXQIFTODSA-N Cys-Met-Ser Chemical compound CSCC[C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)[C@H](CS)N HJGUQJJJXQGXGJ-FXQIFTODSA-N 0.000 description 4
- KZZYVYWSXMFYEC-DCAQKATOSA-N Cys-Val-Leu Chemical compound [H]N[C@@H](CS)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O KZZYVYWSXMFYEC-DCAQKATOSA-N 0.000 description 4
- IAZDPXIOMUYVGZ-UHFFFAOYSA-N Dimethylsulphoxide Chemical compound CS(C)=O IAZDPXIOMUYVGZ-UHFFFAOYSA-N 0.000 description 4
- LFQSCWFLJHTTHZ-UHFFFAOYSA-N Ethanol Chemical compound CCO LFQSCWFLJHTTHZ-UHFFFAOYSA-N 0.000 description 4
- NNQHEEQNPQYPGL-FXQIFTODSA-N Gln-Ala-Gln Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(O)=O NNQHEEQNPQYPGL-FXQIFTODSA-N 0.000 description 4
- KWUSGAIFNHQCBY-DCAQKATOSA-N Gln-Arg-Arg Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O KWUSGAIFNHQCBY-DCAQKATOSA-N 0.000 description 4
- PGPJSRSLQNXBDT-YUMQZZPRSA-N Gln-Arg-Gly Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)NCC(O)=O PGPJSRSLQNXBDT-YUMQZZPRSA-N 0.000 description 4
- IXFVOPOHSRKJNG-LAEOZQHASA-N Gln-Asp-Val Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](C(C)C)C(O)=O IXFVOPOHSRKJNG-LAEOZQHASA-N 0.000 description 4
- PKVWNYGXMNWJSI-CIUDSAMLSA-N Gln-Gln-Gln Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O PKVWNYGXMNWJSI-CIUDSAMLSA-N 0.000 description 4
- NPTGGVQJYRSMCM-GLLZPBPUSA-N Gln-Gln-Thr Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O NPTGGVQJYRSMCM-GLLZPBPUSA-N 0.000 description 4
- PNENQZWRFMUZOM-DCAQKATOSA-N Gln-Glu-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O PNENQZWRFMUZOM-DCAQKATOSA-N 0.000 description 4
- MAGNEQBFSBREJL-DCAQKATOSA-N Gln-Glu-Lys Chemical compound C(CCN)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CCC(=O)N)N MAGNEQBFSBREJL-DCAQKATOSA-N 0.000 description 4
- WVUZERSNWGUKJY-BPUTZDHNSA-N Gln-Glu-Trp Chemical compound C1=CC=C2C(=C1)C(=CN2)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CCC(=O)N)N WVUZERSNWGUKJY-BPUTZDHNSA-N 0.000 description 4
- XJKAKYXMFHUIHT-AUTRQRHGSA-N Gln-Glu-Val Chemical compound CC(C)[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CCC(=O)N)N XJKAKYXMFHUIHT-AUTRQRHGSA-N 0.000 description 4
- GQZDDFRXSDGUNG-YVNDNENWSA-N Gln-Ile-Gln Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(N)=O)C(O)=O GQZDDFRXSDGUNG-YVNDNENWSA-N 0.000 description 4
- WLRYGVYQFXRJDA-DCAQKATOSA-N Gln-Pro-Pro Chemical compound NC(=O)CC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 WLRYGVYQFXRJDA-DCAQKATOSA-N 0.000 description 4
- ZFBBMCKQSNJZSN-AUTRQRHGSA-N Gln-Val-Gln Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O ZFBBMCKQSNJZSN-AUTRQRHGSA-N 0.000 description 4
- HUWSBFYAGXCXKC-CIUDSAMLSA-N Glu-Ala-Met Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCSC)C(O)=O HUWSBFYAGXCXKC-CIUDSAMLSA-N 0.000 description 4
- FYBSCGZLICNOBA-XQXXSGGOSA-N Glu-Ala-Thr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O FYBSCGZLICNOBA-XQXXSGGOSA-N 0.000 description 4
- GCYFUZJHAXJKKE-KKUMJFAQSA-N Glu-Arg-Tyr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O GCYFUZJHAXJKKE-KKUMJFAQSA-N 0.000 description 4
- WLIPTFCZLHCNFD-LPEHRKFASA-N Glu-Gln-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CCC(=O)O)N)C(=O)O WLIPTFCZLHCNFD-LPEHRKFASA-N 0.000 description 4
- HNVFSTLPVJWIDV-CIUDSAMLSA-N Glu-Glu-Gln Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O HNVFSTLPVJWIDV-CIUDSAMLSA-N 0.000 description 4
- MUSGDMDGNGXULI-DCAQKATOSA-N Glu-Glu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CCC(O)=O MUSGDMDGNGXULI-DCAQKATOSA-N 0.000 description 4
- INGJLBQKTRJLFO-UKJIMTQDSA-N Glu-Ile-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)CC)NC(=O)[C@@H](N)CCC(O)=O INGJLBQKTRJLFO-UKJIMTQDSA-N 0.000 description 4
- MWMJCGBSIORNCD-AVGNSLFASA-N Glu-Leu-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O MWMJCGBSIORNCD-AVGNSLFASA-N 0.000 description 4
- ILWHFUZZCFYSKT-AVGNSLFASA-N Glu-Lys-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O ILWHFUZZCFYSKT-AVGNSLFASA-N 0.000 description 4
- HQOGXFLBAKJUMH-CIUDSAMLSA-N Glu-Met-Ser Chemical compound CSCC[C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)[C@H](CCC(=O)O)N HQOGXFLBAKJUMH-CIUDSAMLSA-N 0.000 description 4
- UERORLSAFUHDGU-AVGNSLFASA-N Glu-Phe-Asn Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)O)NC(=O)[C@H](CCC(=O)O)N UERORLSAFUHDGU-AVGNSLFASA-N 0.000 description 4
- QCMVGXDELYMZET-GLLZPBPUSA-N Glu-Thr-Glu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(O)=O)C(O)=O QCMVGXDELYMZET-GLLZPBPUSA-N 0.000 description 4
- DTLLNDVORUEOTM-WDCWCFNPSA-N Glu-Thr-Lys Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCCN)C(O)=O DTLLNDVORUEOTM-WDCWCFNPSA-N 0.000 description 4
- HQTDNEZTGZUWSY-XVKPBYJWSA-N Glu-Val-Gly Chemical compound CC(C)[C@H](NC(=O)[C@@H](N)CCC(O)=O)C(=O)NCC(O)=O HQTDNEZTGZUWSY-XVKPBYJWSA-N 0.000 description 4
- SOYWRINXUSUWEQ-DLOVCJGASA-N Glu-Val-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](C(C)C)NC(=O)[C@@H](N)CCC(O)=O SOYWRINXUSUWEQ-DLOVCJGASA-N 0.000 description 4
- RLFSBAPJTYKSLG-WHFBIAKZSA-N Gly-Ala-Asp Chemical compound NCC(=O)N[C@@H](C)C(=O)N[C@@H](CC(O)=O)C(O)=O RLFSBAPJTYKSLG-WHFBIAKZSA-N 0.000 description 4
- OVSKVOOUFAKODB-UWVGGRQHSA-N Gly-Arg-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)CN)CCCN=C(N)N OVSKVOOUFAKODB-UWVGGRQHSA-N 0.000 description 4
- FZQLXNIMCPJVJE-YUMQZZPRSA-N Gly-Asp-Leu Chemical compound [H]NCC(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O FZQLXNIMCPJVJE-YUMQZZPRSA-N 0.000 description 4
- CQZDZKRHFWJXDF-WDSKDSINSA-N Gly-Gln-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](CCC(N)=O)NC(=O)CN CQZDZKRHFWJXDF-WDSKDSINSA-N 0.000 description 4
- YYPFZVIXAVDHIK-IUCAKERBSA-N Gly-Glu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)CN YYPFZVIXAVDHIK-IUCAKERBSA-N 0.000 description 4
- UFPXDFOYHVEIPI-BYPYZUCNSA-N Gly-Gly-Asp Chemical compound NCC(=O)NCC(=O)N[C@H](C(O)=O)CC(O)=O UFPXDFOYHVEIPI-BYPYZUCNSA-N 0.000 description 4
- QPTNELDXWKRIFX-YFKPBYRVSA-N Gly-Gly-Gln Chemical compound NCC(=O)NCC(=O)N[C@H](C(O)=O)CCC(N)=O QPTNELDXWKRIFX-YFKPBYRVSA-N 0.000 description 4
- ADZGCWWDPFDHCY-ZETCQYMHSA-N Gly-His-Gly Chemical compound OC(=O)CNC(=O)[C@@H](NC(=O)CN)CC1=CN=CN1 ADZGCWWDPFDHCY-ZETCQYMHSA-N 0.000 description 4
- LLZXNUUIBOALNY-QWRGUYRKSA-N Gly-Leu-Lys Chemical compound NCC(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCCCN LLZXNUUIBOALNY-QWRGUYRKSA-N 0.000 description 4
- MIIVFRCYJABHTQ-ONGXEEELSA-N Gly-Leu-Val Chemical compound [H]NCC(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C(C)C)C(O)=O MIIVFRCYJABHTQ-ONGXEEELSA-N 0.000 description 4
- ZLCLYFGMKFCDCN-XPUUQOCRSA-N Gly-Ser-Val Chemical compound CC(C)[C@H](NC(=O)[C@H](CO)NC(=O)CN)C(O)=O ZLCLYFGMKFCDCN-XPUUQOCRSA-N 0.000 description 4
- VSLXGYMEHVAJBH-DLOVCJGASA-N His-Ala-Leu Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(O)=O VSLXGYMEHVAJBH-DLOVCJGASA-N 0.000 description 4
- QQQHYJFKDLDUNK-CIUDSAMLSA-N His-Asp-Cys Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CS)C(=O)O)N QQQHYJFKDLDUNK-CIUDSAMLSA-N 0.000 description 4
- AKEDPWJFQULLPE-IUCAKERBSA-N His-Glu-Gly Chemical compound N[C@@H](Cc1cnc[nH]1)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O AKEDPWJFQULLPE-IUCAKERBSA-N 0.000 description 4
- QEYUCKCWTMIERU-SRVKXCTJSA-N His-Lys-Asp Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(=O)O)C(=O)O)N QEYUCKCWTMIERU-SRVKXCTJSA-N 0.000 description 4
- KFQDSSNYWKZFOO-LSJOCFKGSA-N His-Val-Ala Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C)C(O)=O KFQDSSNYWKZFOO-LSJOCFKGSA-N 0.000 description 4
- DMAPKBANYNZHNR-ULQDDVLXSA-N His-Val-Tyr Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)O)NC(=O)[C@H](CC2=CN=CN2)N DMAPKBANYNZHNR-ULQDDVLXSA-N 0.000 description 4
- TVSPLSZTKTUYLV-ZPFDUUQYSA-N Ile-Glu-Met Chemical compound N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CCSC)C(=O)O TVSPLSZTKTUYLV-ZPFDUUQYSA-N 0.000 description 4
- PDTMWFVVNZYWTR-NHCYSSNCSA-N Ile-Gly-Lys Chemical compound CC[C@H](C)[C@H](N)C(=O)NCC(=O)N[C@@H](CCCCN)C(O)=O PDTMWFVVNZYWTR-NHCYSSNCSA-N 0.000 description 4
- UWLHDGMRWXHFFY-HPCHECBXSA-N Ile-Ile-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H]([C@@H](C)CC)C(=O)N1CCC[C@@H]1C(=O)O)N UWLHDGMRWXHFFY-HPCHECBXSA-N 0.000 description 4
- PARSHQDZROHERM-NHCYSSNCSA-N Ile-Lys-Gly Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)NCC(=O)O)N PARSHQDZROHERM-NHCYSSNCSA-N 0.000 description 4
- XQLGNKLSPYCRMZ-HJWJTTGWSA-N Ile-Phe-Val Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](C(C)C)C(=O)O)N XQLGNKLSPYCRMZ-HJWJTTGWSA-N 0.000 description 4
- RQJUKVXWAKJDBW-SVSWQMSJSA-N Ile-Ser-Thr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(=O)O)N RQJUKVXWAKJDBW-SVSWQMSJSA-N 0.000 description 4
- IBMVEYRWAWIOTN-UHFFFAOYSA-N L-Leucyl-L-Arginyl-L-Proline Natural products CC(C)CC(N)C(=O)NC(CCCN=C(N)N)C(=O)N1CCCC1C(O)=O IBMVEYRWAWIOTN-UHFFFAOYSA-N 0.000 description 4
- RCFDOSNHHZGBOY-UHFFFAOYSA-N L-isoleucyl-L-alanine Natural products CCC(C)C(N)C(=O)NC(C)C(O)=O RCFDOSNHHZGBOY-UHFFFAOYSA-N 0.000 description 4
- SENJXOPIZNYLHU-UHFFFAOYSA-N L-leucyl-L-arginine Natural products CC(C)CC(N)C(=O)NC(C(O)=O)CCCN=C(N)N SENJXOPIZNYLHU-UHFFFAOYSA-N 0.000 description 4
- BQSLGJHIAGOZCD-CIUDSAMLSA-N Leu-Ala-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CO)C(O)=O BQSLGJHIAGOZCD-CIUDSAMLSA-N 0.000 description 4
- KSZCCRIGNVSHFH-UWVGGRQHSA-N Leu-Arg-Gly Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCNC(N)=N)C(=O)NCC(O)=O KSZCCRIGNVSHFH-UWVGGRQHSA-N 0.000 description 4
- IBMVEYRWAWIOTN-RWMBFGLXSA-N Leu-Arg-Pro Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N1CCC[C@@H]1C(O)=O IBMVEYRWAWIOTN-RWMBFGLXSA-N 0.000 description 4
- JKGHDYGZRDWHGA-SRVKXCTJSA-N Leu-Asn-Leu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O JKGHDYGZRDWHGA-SRVKXCTJSA-N 0.000 description 4
- VPKIQULSKFVCSM-SRVKXCTJSA-N Leu-Gln-Arg Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O VPKIQULSKFVCSM-SRVKXCTJSA-N 0.000 description 4
- DXYBNWJZJVSZAE-GUBZILKMSA-N Leu-Gln-Cys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CS)C(=O)O)N DXYBNWJZJVSZAE-GUBZILKMSA-N 0.000 description 4
- DPWGZWUMUUJQDT-IUCAKERBSA-N Leu-Gln-Gly Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)NCC(O)=O DPWGZWUMUUJQDT-IUCAKERBSA-N 0.000 description 4
- DZQMXBALGUHGJT-GUBZILKMSA-N Leu-Glu-Ala Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O DZQMXBALGUHGJT-GUBZILKMSA-N 0.000 description 4
- HFBCHNRFRYLZNV-GUBZILKMSA-N Leu-Glu-Asp Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O HFBCHNRFRYLZNV-GUBZILKMSA-N 0.000 description 4
- HPBCTWSUJOGJSH-MNXVOIDGSA-N Leu-Glu-Ile Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O HPBCTWSUJOGJSH-MNXVOIDGSA-N 0.000 description 4
- QVFGXCVIXXBFHO-AVGNSLFASA-N Leu-Glu-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O QVFGXCVIXXBFHO-AVGNSLFASA-N 0.000 description 4
- DBSLVQBXKVKDKJ-BJDJZHNGSA-N Leu-Ile-Ala Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](C)C(O)=O DBSLVQBXKVKDKJ-BJDJZHNGSA-N 0.000 description 4
- QNBVTHNJGCOVFA-AVGNSLFASA-N Leu-Leu-Glu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCC(O)=O QNBVTHNJGCOVFA-AVGNSLFASA-N 0.000 description 4
- KYIIALJHAOIAHF-KKUMJFAQSA-N Leu-Leu-His Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CC1=CN=CN1 KYIIALJHAOIAHF-KKUMJFAQSA-N 0.000 description 4
- FOBUGKUBUJOWAD-IHPCNDPISA-N Leu-Leu-Trp Chemical compound C1=CC=C2C(C[C@H](NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC(C)C)C(O)=O)=CNC2=C1 FOBUGKUBUJOWAD-IHPCNDPISA-N 0.000 description 4
- CHJKEDSZNSONPS-DCAQKATOSA-N Leu-Pro-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O CHJKEDSZNSONPS-DCAQKATOSA-N 0.000 description 4
- AKVBOOKXVAMKSS-GUBZILKMSA-N Leu-Ser-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(O)=O AKVBOOKXVAMKSS-GUBZILKMSA-N 0.000 description 4
- DAYQSYGBCUKVKT-VOAKCMCISA-N Leu-Thr-Lys Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCCN)C(O)=O DAYQSYGBCUKVKT-VOAKCMCISA-N 0.000 description 4
- WFCKERTZVCQXKH-KBPBESRZSA-N Leu-Tyr-Gly Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)NCC(O)=O WFCKERTZVCQXKH-KBPBESRZSA-N 0.000 description 4
- AXVIGSRGTMNSJU-YESZJQIVSA-N Leu-Tyr-Pro Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)N2CCC[C@@H]2C(=O)O)N AXVIGSRGTMNSJU-YESZJQIVSA-N 0.000 description 4
- FBNPMTNBFFAMMH-UHFFFAOYSA-N Leu-Val-Arg Natural products CC(C)CC(N)C(=O)NC(C(C)C)C(=O)NC(C(O)=O)CCCN=C(N)N FBNPMTNBFFAMMH-UHFFFAOYSA-N 0.000 description 4
- BTSXLXFPMZXVPR-DLOVCJGASA-N Lys-Ala-His Chemical compound C[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@H](CCCCN)N BTSXLXFPMZXVPR-DLOVCJGASA-N 0.000 description 4
- UWKNTTJNVSYXPC-CIUDSAMLSA-N Lys-Ala-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCCCN UWKNTTJNVSYXPC-CIUDSAMLSA-N 0.000 description 4
- 108010062166 Lys-Asn-Asp Proteins 0.000 description 4
- BYPMOIFBQPEWOH-CIUDSAMLSA-N Lys-Asn-Asp Chemical compound C(CCN)C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC(=O)O)C(=O)O)N BYPMOIFBQPEWOH-CIUDSAMLSA-N 0.000 description 4
- NRQRKMYZONPCTM-CIUDSAMLSA-N Lys-Asp-Ser Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O NRQRKMYZONPCTM-CIUDSAMLSA-N 0.000 description 4
- DFXQCCBKGUNYGG-GUBZILKMSA-N Lys-Gln-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](CCC(N)=O)NC(=O)[C@@H](N)CCCCN DFXQCCBKGUNYGG-GUBZILKMSA-N 0.000 description 4
- OWRUUFUVXFREBD-KKUMJFAQSA-N Lys-His-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(C)C)C(O)=O OWRUUFUVXFREBD-KKUMJFAQSA-N 0.000 description 4
- MXMDJEJWERYPMO-XUXIUFHCSA-N Lys-Ile-Arg Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O MXMDJEJWERYPMO-XUXIUFHCSA-N 0.000 description 4
- WRODMZBHNNPRLN-SRVKXCTJSA-N Lys-Leu-Ser Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O WRODMZBHNNPRLN-SRVKXCTJSA-N 0.000 description 4
- CAVRAQIDHUPECU-UVOCVTCTSA-N Lys-Thr-Thr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O CAVRAQIDHUPECU-UVOCVTCTSA-N 0.000 description 4
- LMMBAXJRYSXCOQ-ACRUOGEOSA-N Lys-Tyr-Phe Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](Cc1ccc(O)cc1)C(=O)N[C@@H](Cc1ccccc1)C(O)=O LMMBAXJRYSXCOQ-ACRUOGEOSA-N 0.000 description 4
- XABXVVSWUVCZST-GVXVVHGQSA-N Lys-Val-Gln Chemical compound NC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](C(C)C)NC(=O)[C@@H](N)CCCCN XABXVVSWUVCZST-GVXVVHGQSA-N 0.000 description 4
- KUQWVNFMZLHAPA-CIUDSAMLSA-N Met-Ala-Gln Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(O)=O KUQWVNFMZLHAPA-CIUDSAMLSA-N 0.000 description 4
- IYXDSYWCVVXSKB-CIUDSAMLSA-N Met-Asn-Glu Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O IYXDSYWCVVXSKB-CIUDSAMLSA-N 0.000 description 4
- OOSPRDCGTLQLBP-NHCYSSNCSA-N Met-Glu-Val Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C(C)C)C(O)=O OOSPRDCGTLQLBP-NHCYSSNCSA-N 0.000 description 4
- OXIWIYOJVNOKOV-SRVKXCTJSA-N Met-Met-Arg Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCSC)C(=O)N[C@H](C(O)=O)CCCNC(N)=N OXIWIYOJVNOKOV-SRVKXCTJSA-N 0.000 description 4
- WYDFQSJOARJAMM-GUBZILKMSA-N Met-Pro-Asp Chemical compound CSCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(O)=O)C(O)=O WYDFQSJOARJAMM-GUBZILKMSA-N 0.000 description 4
- GGXZOTSDJJTDGB-GUBZILKMSA-N Met-Ser-Val Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(O)=O GGXZOTSDJJTDGB-GUBZILKMSA-N 0.000 description 4
- 241001465754 Metazoa Species 0.000 description 4
- 241000283973 Oryctolagus cuniculus Species 0.000 description 4
- MPFGIYLYWUCSJG-AVGNSLFASA-N Phe-Glu-Asp Chemical compound OC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CC1=CC=CC=C1 MPFGIYLYWUCSJG-AVGNSLFASA-N 0.000 description 4
- UAYHMOIGIQZLFR-NHCYSSNCSA-N Pro-Gln-Val Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(O)=O UAYHMOIGIQZLFR-NHCYSSNCSA-N 0.000 description 4
- MGDFPGCFVJFITQ-CIUDSAMLSA-N Pro-Glu-Asp Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O MGDFPGCFVJFITQ-CIUDSAMLSA-N 0.000 description 4
- UEHYFUCOGHWASA-HJGDQZAQSA-N Pro-Glu-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H]1CCCN1 UEHYFUCOGHWASA-HJGDQZAQSA-N 0.000 description 4
- HAAQQNHQZBOWFO-LURJTMIESA-N Pro-Gly-Gly Chemical compound OC(=O)CNC(=O)CNC(=O)[C@@H]1CCCN1 HAAQQNHQZBOWFO-LURJTMIESA-N 0.000 description 4
- FEPSEIDIPBMIOS-QXEWZRGKSA-N Pro-Gly-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H]1CCCN1 FEPSEIDIPBMIOS-QXEWZRGKSA-N 0.000 description 4
- YXHYJEPDKSYPSQ-AVGNSLFASA-N Pro-Leu-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H]1CCCN1 YXHYJEPDKSYPSQ-AVGNSLFASA-N 0.000 description 4
- XYSXOCIWCPFOCG-IHRRRGAJSA-N Pro-Leu-Leu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O XYSXOCIWCPFOCG-IHRRRGAJSA-N 0.000 description 4
- SBVPYBFMIGDIDX-SRVKXCTJSA-N Pro-Pro-Pro Chemical compound OC(=O)[C@@H]1CCCN1C(=O)[C@H]1N(C(=O)[C@H]2NCCC2)CCC1 SBVPYBFMIGDIDX-SRVKXCTJSA-N 0.000 description 4
- POQFNPILEQEODH-FXQIFTODSA-N Pro-Ser-Ala Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O POQFNPILEQEODH-FXQIFTODSA-N 0.000 description 4
- SNGZLPOXVRTNMB-LPEHRKFASA-N Pro-Ser-Pro Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CO)C(=O)N2CCC[C@@H]2C(=O)O SNGZLPOXVRTNMB-LPEHRKFASA-N 0.000 description 4
- MKGIILKDUGDRRO-FXQIFTODSA-N Pro-Ser-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H]1CCCN1 MKGIILKDUGDRRO-FXQIFTODSA-N 0.000 description 4
- IMNVAOPEMFDAQD-NHCYSSNCSA-N Pro-Val-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O IMNVAOPEMFDAQD-NHCYSSNCSA-N 0.000 description 4
- 238000012300 Sequence Analysis Methods 0.000 description 4
- FIXILCYTSAUERA-FXQIFTODSA-N Ser-Ala-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O FIXILCYTSAUERA-FXQIFTODSA-N 0.000 description 4
- OYEDZGNMSBZCIM-XGEHTFHBSA-N Ser-Arg-Thr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)O)C(O)=O OYEDZGNMSBZCIM-XGEHTFHBSA-N 0.000 description 4
- CRZRTKAVUUGKEQ-ACZMJKKPSA-N Ser-Gln-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(O)=O CRZRTKAVUUGKEQ-ACZMJKKPSA-N 0.000 description 4
- MAWSJXHRLWVJEZ-ACZMJKKPSA-N Ser-Gln-Cys Chemical compound C(CC(=O)N)[C@@H](C(=O)N[C@@H](CS)C(=O)O)NC(=O)[C@H](CO)N MAWSJXHRLWVJEZ-ACZMJKKPSA-N 0.000 description 4
- UOLGINIHBRIECN-FXQIFTODSA-N Ser-Glu-Glu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O UOLGINIHBRIECN-FXQIFTODSA-N 0.000 description 4
- GJFYFGOEWLDQGW-GUBZILKMSA-N Ser-Leu-Gln Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CO)N GJFYFGOEWLDQGW-GUBZILKMSA-N 0.000 description 4
- XUDRHBPSPAPDJP-SRVKXCTJSA-N Ser-Lys-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CO XUDRHBPSPAPDJP-SRVKXCTJSA-N 0.000 description 4
- UPLYXVPQLJVWMM-KKUMJFAQSA-N Ser-Phe-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(C)C)C(O)=O UPLYXVPQLJVWMM-KKUMJFAQSA-N 0.000 description 4
- BMKNXTJLHFIAAH-CIUDSAMLSA-N Ser-Ser-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O BMKNXTJLHFIAAH-CIUDSAMLSA-N 0.000 description 4
- CUXJENOFJXOSOZ-BIIVOSGPSA-N Ser-Ser-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CO)NC(=O)[C@H](CO)N)C(=O)O CUXJENOFJXOSOZ-BIIVOSGPSA-N 0.000 description 4
- UIIMBOGNXHQVGW-UHFFFAOYSA-M Sodium bicarbonate Chemical compound [Na+].OC([O-])=O UIIMBOGNXHQVGW-UHFFFAOYSA-M 0.000 description 4
- ZUXQFMVPAYGPFJ-JXUBOQSCSA-N Thr-Ala-Lys Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCCCN ZUXQFMVPAYGPFJ-JXUBOQSCSA-N 0.000 description 4
- XYEXCEPTALHNEV-RCWTZXSCSA-N Thr-Arg-Arg Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O XYEXCEPTALHNEV-RCWTZXSCSA-N 0.000 description 4
- GKMYGVQDGVYCPC-IUKAMOBKSA-N Thr-Asp-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H]([C@@H](C)O)N GKMYGVQDGVYCPC-IUKAMOBKSA-N 0.000 description 4
- XDARBNMYXKUFOJ-GSSVUCPTSA-N Thr-Asp-Thr Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O XDARBNMYXKUFOJ-GSSVUCPTSA-N 0.000 description 4
- LHEZGZQRLDBSRR-WDCWCFNPSA-N Thr-Glu-Leu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O LHEZGZQRLDBSRR-WDCWCFNPSA-N 0.000 description 4
- MEJHFIOYJHTWMK-VOAKCMCISA-N Thr-Leu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)[C@@H](C)O MEJHFIOYJHTWMK-VOAKCMCISA-N 0.000 description 4
- YRJOLUDFVAUXLI-GSSVUCPTSA-N Thr-Thr-Asp Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@H](C(O)=O)CC(O)=O YRJOLUDFVAUXLI-GSSVUCPTSA-N 0.000 description 4
- ZMYCLHFLHRVOEA-HEIBUPTGSA-N Thr-Thr-Ser Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(O)=O ZMYCLHFLHRVOEA-HEIBUPTGSA-N 0.000 description 4
- JNKAYADBODLPMQ-HSHDSVGOSA-N Thr-Trp-Val Chemical compound C1=CC=C2C(C[C@@H](C(=O)N[C@@H](C(C)C)C(O)=O)NC(=O)[C@@H](N)[C@@H](C)O)=CNC2=C1 JNKAYADBODLPMQ-HSHDSVGOSA-N 0.000 description 4
- OGOYMQWIWHGTGH-KZVJFYERSA-N Thr-Val-Ala Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C)C(O)=O OGOYMQWIWHGTGH-KZVJFYERSA-N 0.000 description 4
- UPNRACRNHISCAF-SZMVWBNQSA-N Trp-Lys-Gln Chemical compound C1=CC=C2C(C[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(N)=O)C(O)=O)=CNC2=C1 UPNRACRNHISCAF-SZMVWBNQSA-N 0.000 description 4
- 108090000631 Trypsin Proteins 0.000 description 4
- 102000004142 Trypsin Human genes 0.000 description 4
- QUILOGWWLXMSAT-IHRRRGAJSA-N Tyr-Gln-Gln Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O QUILOGWWLXMSAT-IHRRRGAJSA-N 0.000 description 4
- QKXAEWMHAAVVGS-KKUMJFAQSA-N Tyr-Pro-Glu Chemical compound N[C@@H](Cc1ccc(O)cc1)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(O)=O)C(O)=O QKXAEWMHAAVVGS-KKUMJFAQSA-N 0.000 description 4
- KLQPIEVIKOQRAW-IZPVPAKOSA-N Tyr-Thr-Thr Chemical compound C[C@H]([C@@H](C(=O)N[C@@H]([C@@H](C)O)C(=O)O)NC(=O)[C@H](CC1=CC=C(C=C1)O)N)O KLQPIEVIKOQRAW-IZPVPAKOSA-N 0.000 description 4
- ASQFIHTXXMFENG-XPUUQOCRSA-N Val-Ala-Gly Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](C)C(=O)NCC(O)=O ASQFIHTXXMFENG-XPUUQOCRSA-N 0.000 description 4
- LABUITCFCAABSV-UHFFFAOYSA-N Val-Ala-Tyr Natural products CC(C)C(N)C(=O)NC(C)C(=O)NC(C(O)=O)CC1=CC=C(O)C=C1 LABUITCFCAABSV-UHFFFAOYSA-N 0.000 description 4
- NMANTMWGQZASQN-QXEWZRGKSA-N Val-Arg-Asp Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CC(=O)O)C(=O)O)N NMANTMWGQZASQN-QXEWZRGKSA-N 0.000 description 4
- KKHRWGYHBZORMQ-NHCYSSNCSA-N Val-Arg-Glu Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N KKHRWGYHBZORMQ-NHCYSSNCSA-N 0.000 description 4
- XGJLNBNZNMVJRS-NRPADANISA-N Val-Glu-Ala Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O XGJLNBNZNMVJRS-NRPADANISA-N 0.000 description 4
- LTTQCQRTSHJPPL-ZKWXMUAHSA-N Val-Ser-Asp Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(=O)O)C(=O)O)N LTTQCQRTSHJPPL-ZKWXMUAHSA-N 0.000 description 4
- 108010070944 alanylhistidine Proteins 0.000 description 4
- 229960000723 ampicillin Drugs 0.000 description 4
- AVKUERGKIZMTKX-NJBDSQKTSA-N ampicillin Chemical compound C1([C@@H](N)C(=O)N[C@H]2[C@H]3SC([C@@H](N3C2=O)C(O)=O)(C)C)=CC=CC=C1 AVKUERGKIZMTKX-NJBDSQKTSA-N 0.000 description 4
- 108010092854 aspartyllysine Proteins 0.000 description 4
- 238000006243 chemical reaction Methods 0.000 description 4
- 238000012790 confirmation Methods 0.000 description 4
- 238000010276 construction Methods 0.000 description 4
- 238000001962 electrophoresis Methods 0.000 description 4
- 239000000499 gel Substances 0.000 description 4
- 108010090037 glycyl-alanyl-isoleucine Proteins 0.000 description 4
- 108010033719 glycyl-histidyl-glycine Proteins 0.000 description 4
- 239000001963 growth medium Substances 0.000 description 4
- 108010085325 histidylproline Proteins 0.000 description 4
- 238000001727 in vivo Methods 0.000 description 4
- 230000005764 inhibitory process Effects 0.000 description 4
- 108010000761 leucylarginine Proteins 0.000 description 4
- 108010038320 lysylphenylalanine Proteins 0.000 description 4
- 238000005259 measurement Methods 0.000 description 4
- 210000004379 membrane Anatomy 0.000 description 4
- 239000012528 membrane Substances 0.000 description 4
- 125000004184 methoxymethyl group Chemical group [H]C([H])([H])OC([H])([H])* 0.000 description 4
- 125000001997 phenyl group Chemical group [H]C1=C([H])C([H])=C(*)C([H])=C1[H] 0.000 description 4
- 108010020755 prolyl-glycyl-glycine Proteins 0.000 description 4
- 108010014614 prolyl-glycyl-proline Proteins 0.000 description 4
- 210000002966 serum Anatomy 0.000 description 4
- 108010048397 seryl-lysyl-leucine Proteins 0.000 description 4
- DAEPDZWVDSPTHF-UHFFFAOYSA-M sodium pyruvate Chemical compound [Na+].CC(=O)C([O-])=O DAEPDZWVDSPTHF-UHFFFAOYSA-M 0.000 description 4
- 210000001685 thyroid gland Anatomy 0.000 description 4
- 238000002054 transplantation Methods 0.000 description 4
- 239000012588 trypsin Substances 0.000 description 4
- IBIDRSSEHFLGSD-UHFFFAOYSA-N valinyl-arginine Natural products CC(C)C(N)C(=O)NC(C(O)=O)CCCN=C(N)N IBIDRSSEHFLGSD-UHFFFAOYSA-N 0.000 description 4
- NTUPOKHATNSWCY-PMPSAXMXSA-N (2s)-2-[[(2s)-1-[(2r)-2-amino-3-phenylpropanoyl]pyrrolidine-2-carbonyl]amino]-5-(diaminomethylideneamino)pentanoic acid Chemical compound C([C@@H](N)C(=O)N1[C@@H](CCC1)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O)C1=CC=CC=C1 NTUPOKHATNSWCY-PMPSAXMXSA-N 0.000 description 3
- 125000006432 1-methyl cyclopropyl group Chemical group [H]C([H])([H])C1(*)C([H])([H])C1([H])[H] 0.000 description 3
- HHGYNJRJIINWAK-FXQIFTODSA-N Ala-Ala-Arg Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N HHGYNJRJIINWAK-FXQIFTODSA-N 0.000 description 3
- KIUYPHAMDKDICO-WHFBIAKZSA-N Ala-Asp-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)NCC(O)=O KIUYPHAMDKDICO-WHFBIAKZSA-N 0.000 description 3
- OYJCVIGKMXUVKB-GARJFASQSA-N Ala-Leu-Pro Chemical compound C[C@@H](C(=O)N[C@@H](CC(C)C)C(=O)N1CCC[C@@H]1C(=O)O)N OYJCVIGKMXUVKB-GARJFASQSA-N 0.000 description 3
- NINQYGGNRIBFSC-CIUDSAMLSA-N Ala-Lys-Ser Chemical compound NCCCC[C@H](NC(=O)[C@@H](N)C)C(=O)N[C@@H](CO)C(O)=O NINQYGGNRIBFSC-CIUDSAMLSA-N 0.000 description 3
- KQESEZXHYOUIIM-CQDKDKBSSA-N Ala-Lys-Tyr Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O KQESEZXHYOUIIM-CQDKDKBSSA-N 0.000 description 3
- WQKAQKZRDIZYNV-VZFHVOOUSA-N Ala-Ser-Thr Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(O)=O WQKAQKZRDIZYNV-VZFHVOOUSA-N 0.000 description 3
- IETUUAHKCHOQHP-KZVJFYERSA-N Ala-Thr-Val Chemical compound CC(C)[C@H](NC(=O)[C@@H](NC(=O)[C@H](C)N)[C@@H](C)O)C(O)=O IETUUAHKCHOQHP-KZVJFYERSA-N 0.000 description 3
- XCIGOVDXZULBBV-DCAQKATOSA-N Ala-Val-Lys Chemical compound CC(C)[C@H](NC(=O)[C@H](C)N)C(=O)N[C@@H](CCCCN)C(O)=O XCIGOVDXZULBBV-DCAQKATOSA-N 0.000 description 3
- VXXHDZKEQNGXNU-QXEWZRGKSA-N Arg-Asp-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H](N)CCCN=C(N)N VXXHDZKEQNGXNU-QXEWZRGKSA-N 0.000 description 3
- HAVKMRGWNXMCDR-STQMWFEESA-N Arg-Gly-Phe Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)NCC(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O HAVKMRGWNXMCDR-STQMWFEESA-N 0.000 description 3
- GNYUVVJYGJFKHN-RVMXOQNASA-N Arg-Ile-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N GNYUVVJYGJFKHN-RVMXOQNASA-N 0.000 description 3
- VIINVRPKMUZYOI-DCAQKATOSA-N Arg-Met-Glu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCC(O)=O)C(O)=O VIINVRPKMUZYOI-DCAQKATOSA-N 0.000 description 3
- OVQJAKFLFTZDNC-GUBZILKMSA-N Arg-Pro-Asp Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(O)=O)C(O)=O OVQJAKFLFTZDNC-GUBZILKMSA-N 0.000 description 3
- HGKHPCFTRQDHCU-IUCAKERBSA-N Arg-Pro-Gly Chemical compound NC(N)=NCCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O HGKHPCFTRQDHCU-IUCAKERBSA-N 0.000 description 3
- YNSUUAOAFCVINY-OSUNSFLBSA-N Arg-Thr-Ile Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O YNSUUAOAFCVINY-OSUNSFLBSA-N 0.000 description 3
- ZZXMOQIUIJJOKZ-ZLUOBGJFSA-N Asn-Asn-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CC(N)=O ZZXMOQIUIJJOKZ-ZLUOBGJFSA-N 0.000 description 3
- WPOLSNAQGVHROR-GUBZILKMSA-N Asn-Gln-Leu Chemical compound CC(C)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CC(=O)N)N WPOLSNAQGVHROR-GUBZILKMSA-N 0.000 description 3
- SGAUXNZEFIEAAI-GARJFASQSA-N Asn-His-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CN=CN2)NC(=O)[C@H](CC(=O)N)N)C(=O)O SGAUXNZEFIEAAI-GARJFASQSA-N 0.000 description 3
- NCFJQJRLQJEECD-NHCYSSNCSA-N Asn-Leu-Val Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C(C)C)C(O)=O NCFJQJRLQJEECD-NHCYSSNCSA-N 0.000 description 3
- SNYCNNPOFYBCEK-ZLUOBGJFSA-N Asn-Ser-Ser Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O SNYCNNPOFYBCEK-ZLUOBGJFSA-N 0.000 description 3
- WSOKZUVWBXVJHX-CIUDSAMLSA-N Asp-Arg-Glu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(O)=O WSOKZUVWBXVJHX-CIUDSAMLSA-N 0.000 description 3
- RQHLMGCXCZUOGT-ZPFDUUQYSA-N Asp-Leu-Ile Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O RQHLMGCXCZUOGT-ZPFDUUQYSA-N 0.000 description 3
- TZBJAXGYGSIUHQ-XUXIUFHCSA-N Asp-Leu-Leu-Ser Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O TZBJAXGYGSIUHQ-XUXIUFHCSA-N 0.000 description 3
- UZFHNLYQWMGUHU-DCAQKATOSA-N Asp-Lys-Arg Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O UZFHNLYQWMGUHU-DCAQKATOSA-N 0.000 description 3
- UAXIKORUDGGIGA-DCAQKATOSA-N Asp-Pro-Lys Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CC(=O)O)N)C(=O)N[C@@H](CCCCN)C(=O)O UAXIKORUDGGIGA-DCAQKATOSA-N 0.000 description 3
- XYPJXLLXNSAWHZ-SRVKXCTJSA-N Asp-Ser-Tyr Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O XYPJXLLXNSAWHZ-SRVKXCTJSA-N 0.000 description 3
- NWAHPBGBDIFUFD-KKUMJFAQSA-N Asp-Tyr-Leu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(C)C)C(O)=O NWAHPBGBDIFUFD-KKUMJFAQSA-N 0.000 description 3
- GYNUXDMCDILYIQ-QRTARXTBSA-N Asp-Val-Trp Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)O)NC(=O)[C@H](CC(=O)O)N GYNUXDMCDILYIQ-QRTARXTBSA-N 0.000 description 3
- 238000002965 ELISA Methods 0.000 description 3
- LGIKBBLQVSWUGK-DCAQKATOSA-N Gln-Leu-Gln Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O LGIKBBLQVSWUGK-DCAQKATOSA-N 0.000 description 3
- KHNJVFYHIKLUPD-SRVKXCTJSA-N Gln-Leu-Met Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCSC)C(=O)O)NC(=O)[C@H](CCC(=O)N)N KHNJVFYHIKLUPD-SRVKXCTJSA-N 0.000 description 3
- YPMDZWPZFOZYFG-GUBZILKMSA-N Gln-Leu-Ser Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O YPMDZWPZFOZYFG-GUBZILKMSA-N 0.000 description 3
- KPNWAJMEMRCLAL-GUBZILKMSA-N Gln-Ser-Lys Chemical compound C(CCN)C[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CCC(=O)N)N KPNWAJMEMRCLAL-GUBZILKMSA-N 0.000 description 3
- DYVMTEWCGAVKSE-HJGDQZAQSA-N Gln-Thr-Arg Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)NC(=O)[C@H](CCC(=O)N)N)O DYVMTEWCGAVKSE-HJGDQZAQSA-N 0.000 description 3
- AVZHGSCDKIQZPQ-CIUDSAMLSA-N Glu-Arg-Ala Chemical compound C[C@H](NC(=O)[C@H](CCCNC(N)=N)NC(=O)[C@@H](N)CCC(O)=O)C(O)=O AVZHGSCDKIQZPQ-CIUDSAMLSA-N 0.000 description 3
- AKJRHDMTEJXTPV-ACZMJKKPSA-N Glu-Asn-Ala Chemical compound C[C@H](NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CCC(O)=O)C(O)=O AKJRHDMTEJXTPV-ACZMJKKPSA-N 0.000 description 3
- SOEPMWQCTJITPZ-SRVKXCTJSA-N Glu-Met-Lys Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCC(=O)O)N SOEPMWQCTJITPZ-SRVKXCTJSA-N 0.000 description 3
- JDUKCSSHWNIQQZ-IHRRRGAJSA-N Glu-Phe-Glu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CCC(O)=O)C(O)=O JDUKCSSHWNIQQZ-IHRRRGAJSA-N 0.000 description 3
- IDEODOAVGCMUQV-GUBZILKMSA-N Glu-Ser-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O IDEODOAVGCMUQV-GUBZILKMSA-N 0.000 description 3
- VIPDPMHGICREIS-GVXVVHGQSA-N Glu-Val-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O VIPDPMHGICREIS-GVXVVHGQSA-N 0.000 description 3
- GQGAFTPXAPKSCF-WHFBIAKZSA-N Gly-Ala-Cys Chemical compound NCC(=O)N[C@@H](C)C(=O)N[C@@H](CS)C(=O)O GQGAFTPXAPKSCF-WHFBIAKZSA-N 0.000 description 3
- JXYMPBCYRKWJEE-BQBZGAKWSA-N Gly-Arg-Ala Chemical compound [H]NCC(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C)C(O)=O JXYMPBCYRKWJEE-BQBZGAKWSA-N 0.000 description 3
- OCQUNKSFDYDXBG-QXEWZRGKSA-N Gly-Arg-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)CN)CCCN=C(N)N OCQUNKSFDYDXBG-QXEWZRGKSA-N 0.000 description 3
- SOEATRRYCIPEHA-BQBZGAKWSA-N Gly-Glu-Glu Chemical compound [H]NCC(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O SOEATRRYCIPEHA-BQBZGAKWSA-N 0.000 description 3
- KMSGYZQRXPUKGI-BYPYZUCNSA-N Gly-Gly-Asn Chemical compound NCC(=O)NCC(=O)N[C@H](C(O)=O)CC(N)=O KMSGYZQRXPUKGI-BYPYZUCNSA-N 0.000 description 3
- BHPQOIPBLYJNAW-NGZCFLSTSA-N Gly-Ile-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)CN BHPQOIPBLYJNAW-NGZCFLSTSA-N 0.000 description 3
- YKJUITHASJAGHO-HOTGVXAUSA-N Gly-Lys-Trp Chemical compound C1=CC=C2C(=C1)C(=CN2)C[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)CN YKJUITHASJAGHO-HOTGVXAUSA-N 0.000 description 3
- IFHJOBKVXBESRE-YUMQZZPRSA-N Gly-Met-Gln Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)CN IFHJOBKVXBESRE-YUMQZZPRSA-N 0.000 description 3
- HFPVRZWORNJRRC-UWVGGRQHSA-N Gly-Pro-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)CN HFPVRZWORNJRRC-UWVGGRQHSA-N 0.000 description 3
- YOBGUCWZPXJHTN-BQBZGAKWSA-N Gly-Ser-Arg Chemical compound NCC(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCCN=C(N)N YOBGUCWZPXJHTN-BQBZGAKWSA-N 0.000 description 3
- SOEGEPHNZOISMT-BYPYZUCNSA-N Gly-Ser-Gly Chemical compound NCC(=O)N[C@@H](CO)C(=O)NCC(O)=O SOEGEPHNZOISMT-BYPYZUCNSA-N 0.000 description 3
- RIYIFUFFFBIOEU-KBPBESRZSA-N Gly-Tyr-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)CN)CC1=CC=C(O)C=C1 RIYIFUFFFBIOEU-KBPBESRZSA-N 0.000 description 3
- JBCLFWXMTIKCCB-UHFFFAOYSA-N H-Gly-Phe-OH Natural products NCC(=O)NC(C(O)=O)CC1=CC=CC=C1 JBCLFWXMTIKCCB-UHFFFAOYSA-N 0.000 description 3
- RVKIPWVMZANZLI-UHFFFAOYSA-N H-Lys-Trp-OH Natural products C1=CC=C2C(CC(NC(=O)C(N)CCCCN)C(O)=O)=CNC2=C1 RVKIPWVMZANZLI-UHFFFAOYSA-N 0.000 description 3
- TVRMJKNELJKNRS-GUBZILKMSA-N His-Glu-Asn Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CC(=O)N)C(=O)O)N TVRMJKNELJKNRS-GUBZILKMSA-N 0.000 description 3
- ORERHHPZDDEMSC-VGDYDELISA-N His-Ile-Ser Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)[C@H](CC1=CN=CN1)N ORERHHPZDDEMSC-VGDYDELISA-N 0.000 description 3
- RNMNYMDTESKEAJ-KKUMJFAQSA-N His-Leu-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC1=CN=CN1 RNMNYMDTESKEAJ-KKUMJFAQSA-N 0.000 description 3
- BZAQOPHNBFOOJS-DCAQKATOSA-N His-Pro-Asp Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(O)=O)C(O)=O BZAQOPHNBFOOJS-DCAQKATOSA-N 0.000 description 3
- QLBXWYXMLHAREM-PYJNHQTQSA-N His-Val-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](C(C)C)NC(=O)[C@H](CC1=CN=CN1)N QLBXWYXMLHAREM-PYJNHQTQSA-N 0.000 description 3
- BEWFWZRGBDVXRP-PEFMBERDSA-N Ile-Glu-Asn Chemical compound [H]N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O BEWFWZRGBDVXRP-PEFMBERDSA-N 0.000 description 3
- OUUCIIJSBIBCHB-ZPFDUUQYSA-N Ile-Leu-Asp Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O OUUCIIJSBIBCHB-ZPFDUUQYSA-N 0.000 description 3
- YCKPUHHMCFSUMD-IUKAMOBKSA-N Ile-Thr-Asp Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(=O)O)C(=O)O)N YCKPUHHMCFSUMD-IUKAMOBKSA-N 0.000 description 3
- NGKPIPCGMLWHBX-WZLNRYEVSA-N Ile-Tyr-Thr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)N[C@@H]([C@@H](C)O)C(=O)O)N NGKPIPCGMLWHBX-WZLNRYEVSA-N 0.000 description 3
- OGCQGUIWMSBHRZ-CIUDSAMLSA-N Leu-Asn-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(O)=O OGCQGUIWMSBHRZ-CIUDSAMLSA-N 0.000 description 3
- HYIFFZAQXPUEAU-QWRGUYRKSA-N Leu-Gly-Leu Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CC(C)C HYIFFZAQXPUEAU-QWRGUYRKSA-N 0.000 description 3
- APFJUBGRZGMQFF-QWRGUYRKSA-N Leu-Gly-Lys Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CCCCN APFJUBGRZGMQFF-QWRGUYRKSA-N 0.000 description 3
- JNDYEOUZBLOVOF-AVGNSLFASA-N Leu-Leu-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O JNDYEOUZBLOVOF-AVGNSLFASA-N 0.000 description 3
- FAELBUXXFQLUAX-AJNGGQMLSA-N Leu-Leu-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC(C)C FAELBUXXFQLUAX-AJNGGQMLSA-N 0.000 description 3
- RZXLZBIUTDQHJQ-SRVKXCTJSA-N Leu-Lys-Asp Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(O)=O RZXLZBIUTDQHJQ-SRVKXCTJSA-N 0.000 description 3
- LMDVGHQPPPLYAR-IHRRRGAJSA-N Leu-Val-His Chemical compound N[C@@H](CC(C)C)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC1=CNC=N1)C(=O)O LMDVGHQPPPLYAR-IHRRRGAJSA-N 0.000 description 3
- 206010058467 Lung neoplasm malignant Diseases 0.000 description 3
- 239000006142 Luria-Bertani Agar Substances 0.000 description 3
- CLBGMWIYPYAZPR-AVGNSLFASA-N Lys-Arg-Arg Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O CLBGMWIYPYAZPR-AVGNSLFASA-N 0.000 description 3
- WTZUSCUIVPVCRH-SRVKXCTJSA-N Lys-Gln-Arg Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@H](C(O)=O)CCCN=C(N)N WTZUSCUIVPVCRH-SRVKXCTJSA-N 0.000 description 3
- IMAKMJCBYCSMHM-AVGNSLFASA-N Lys-Glu-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CCCCN IMAKMJCBYCSMHM-AVGNSLFASA-N 0.000 description 3
- WGLAORUKDGRINI-WDCWCFNPSA-N Lys-Glu-Thr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O WGLAORUKDGRINI-WDCWCFNPSA-N 0.000 description 3
- QOJDBRUCOXQSSK-AJNGGQMLSA-N Lys-Ile-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCCCN)C(O)=O QOJDBRUCOXQSSK-AJNGGQMLSA-N 0.000 description 3
- MUXNCRWTWBMNHX-SRVKXCTJSA-N Lys-Leu-Asp Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O MUXNCRWTWBMNHX-SRVKXCTJSA-N 0.000 description 3
- SKRGVGLIRUGANF-AVGNSLFASA-N Lys-Leu-Glu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O SKRGVGLIRUGANF-AVGNSLFASA-N 0.000 description 3
- KQAREVUPVXMNNP-WDSOQIARSA-N Lys-Trp-Met Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CCSC)C(O)=O KQAREVUPVXMNNP-WDSOQIARSA-N 0.000 description 3
- QGQGAIBGTUJRBR-NAKRPEOUSA-N Met-Ala-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCSC QGQGAIBGTUJRBR-NAKRPEOUSA-N 0.000 description 3
- WDTLNWHPIPCMMP-AVGNSLFASA-N Met-Arg-Leu Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(O)=O WDTLNWHPIPCMMP-AVGNSLFASA-N 0.000 description 3
- KMSMNUFBNCHMII-IHRRRGAJSA-N Met-Leu-Lys Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCCCN KMSMNUFBNCHMII-IHRRRGAJSA-N 0.000 description 3
- XGIQKEAKUSPCBU-SRVKXCTJSA-N Met-Met-Val Chemical compound CC(C)[C@@H](C(=O)O)NC(=O)[C@H](CCSC)NC(=O)[C@H](CCSC)N XGIQKEAKUSPCBU-SRVKXCTJSA-N 0.000 description 3
- WUGMRIBZSVSJNP-UHFFFAOYSA-N N-L-alanyl-L-tryptophan Natural products C1=CC=C2C(CC(NC(=O)C(N)C)C(O)=O)=CNC2=C1 WUGMRIBZSVSJNP-UHFFFAOYSA-N 0.000 description 3
- PESQCPHRXOFIPX-UHFFFAOYSA-N N-L-methionyl-L-tyrosine Natural products CSCCC(N)C(=O)NC(C(O)=O)CC1=CC=C(O)C=C1 PESQCPHRXOFIPX-UHFFFAOYSA-N 0.000 description 3
- 108010087066 N2-tryptophyllysine Proteins 0.000 description 3
- VUYCNYVLKACHPA-KKUMJFAQSA-N Phe-Asp-His Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)N VUYCNYVLKACHPA-KKUMJFAQSA-N 0.000 description 3
- HBGFEEQFVBWYJQ-KBPBESRZSA-N Phe-Gly-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CC1=CC=CC=C1 HBGFEEQFVBWYJQ-KBPBESRZSA-N 0.000 description 3
- NJJBATPLUQHRBM-IHRRRGAJSA-N Phe-Pro-Ser Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CC2=CC=CC=C2)N)C(=O)N[C@@H](CO)C(=O)O NJJBATPLUQHRBM-IHRRRGAJSA-N 0.000 description 3
- KIZQGKLMXKGDIV-BQBZGAKWSA-N Pro-Ala-Gly Chemical compound OC(=O)CNC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1 KIZQGKLMXKGDIV-BQBZGAKWSA-N 0.000 description 3
- QSKCKTUQPICLSO-AVGNSLFASA-N Pro-Arg-Lys Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCCN)C(=O)O QSKCKTUQPICLSO-AVGNSLFASA-N 0.000 description 3
- PULPZRAHVFBVTO-DCAQKATOSA-N Pro-Glu-Arg Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O PULPZRAHVFBVTO-DCAQKATOSA-N 0.000 description 3
- KBUAPZAZPWNYSW-SRVKXCTJSA-N Pro-Pro-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@H]1NCCC1 KBUAPZAZPWNYSW-SRVKXCTJSA-N 0.000 description 3
- GMJDSFYVTAMIBF-FXQIFTODSA-N Pro-Ser-Asp Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O GMJDSFYVTAMIBF-FXQIFTODSA-N 0.000 description 3
- QUBVFEANYYWBTM-VEVYYDQMSA-N Pro-Thr-Asp Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(O)=O)C(O)=O QUBVFEANYYWBTM-VEVYYDQMSA-N 0.000 description 3
- IALSFJSONJZBKB-HRCADAONSA-N Pro-Tyr-Pro Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CC2=CC=C(C=C2)O)C(=O)N3CCC[C@@H]3C(=O)O IALSFJSONJZBKB-HRCADAONSA-N 0.000 description 3
- VDHGTOHMHHQSKG-JYJNAYRXSA-N Pro-Val-Phe Chemical compound CC(C)[C@H](NC(=O)[C@@H]1CCCN1)C(=O)N[C@@H](Cc1ccccc1)C(O)=O VDHGTOHMHHQSKG-JYJNAYRXSA-N 0.000 description 3
- IDQFQFVEWMWRQQ-DLOVCJGASA-N Ser-Ala-Phe Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O IDQFQFVEWMWRQQ-DLOVCJGASA-N 0.000 description 3
- QFBNNYNWKYKVJO-DCAQKATOSA-N Ser-Arg-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)CO)CCCN=C(N)N QFBNNYNWKYKVJO-DCAQKATOSA-N 0.000 description 3
- OLIJLNWFEQEFDM-SRVKXCTJSA-N Ser-Asp-Phe Chemical compound OC[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 OLIJLNWFEQEFDM-SRVKXCTJSA-N 0.000 description 3
- ULVMNZOKDBHKKI-ACZMJKKPSA-N Ser-Gln-Asp Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O ULVMNZOKDBHKKI-ACZMJKKPSA-N 0.000 description 3
- HMRAQFJFTOLDKW-GUBZILKMSA-N Ser-His-Glu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCC(O)=O)C(O)=O HMRAQFJFTOLDKW-GUBZILKMSA-N 0.000 description 3
- RIAKPZVSNBBNRE-BJDJZHNGSA-N Ser-Ile-Leu Chemical compound OC[C@H](N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(C)C)C(O)=O RIAKPZVSNBBNRE-BJDJZHNGSA-N 0.000 description 3
- HEUVHBXOVZONPU-BJDJZHNGSA-N Ser-Leu-Ile Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O HEUVHBXOVZONPU-BJDJZHNGSA-N 0.000 description 3
- PPNPDKGQRFSCAC-CIUDSAMLSA-N Ser-Lys-Asp Chemical compound NCCCC[C@H](NC(=O)[C@@H](N)CO)C(=O)N[C@@H](CC(O)=O)C(O)=O PPNPDKGQRFSCAC-CIUDSAMLSA-N 0.000 description 3
- XKFJENWJGHMDLI-QWRGUYRKSA-N Ser-Phe-Gly Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)NCC(O)=O XKFJENWJGHMDLI-QWRGUYRKSA-N 0.000 description 3
- ADJDNJCSPNFFPI-FXQIFTODSA-N Ser-Pro-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CO ADJDNJCSPNFFPI-FXQIFTODSA-N 0.000 description 3
- FKYWFUYPVKLJLP-DCAQKATOSA-N Ser-Pro-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CO FKYWFUYPVKLJLP-DCAQKATOSA-N 0.000 description 3
- QNBVFKZSSRYNFX-CUJWVEQBSA-N Ser-Thr-His Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@H](CO)N)O QNBVFKZSSRYNFX-CUJWVEQBSA-N 0.000 description 3
- OJFFAQFRCVPHNN-JYBASQMISA-N Ser-Thr-Trp Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC1=CNC2=C1C=CC=C2)C(O)=O OJFFAQFRCVPHNN-JYBASQMISA-N 0.000 description 3
- WFUAUEQXPVNAEF-ZJDVBMNYSA-N Thr-Arg-Thr Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@H](C(=O)N[C@@H]([C@@H](C)O)C(O)=O)CCCN=C(N)N WFUAUEQXPVNAEF-ZJDVBMNYSA-N 0.000 description 3
- PQLXHSACXPGWPD-GSSVUCPTSA-N Thr-Asn-Thr Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O PQLXHSACXPGWPD-GSSVUCPTSA-N 0.000 description 3
- YZUWGFXVVZQJEI-PMVVWTBXSA-N Thr-Gly-His Chemical compound C[C@H]([C@@H](C(=O)NCC(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N)O YZUWGFXVVZQJEI-PMVVWTBXSA-N 0.000 description 3
- YUPVPKZBKCLFLT-QTKMDUPCSA-N Thr-His-Val Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](C(C)C)C(=O)O)N)O YUPVPKZBKCLFLT-QTKMDUPCSA-N 0.000 description 3
- KRDSCBLRHORMRK-JXUBOQSCSA-N Thr-Lys-Ala Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(O)=O KRDSCBLRHORMRK-JXUBOQSCSA-N 0.000 description 3
- PCMDGXKXVMBIFP-VEVYYDQMSA-N Thr-Met-Asn Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(N)=O)C(O)=O PCMDGXKXVMBIFP-VEVYYDQMSA-N 0.000 description 3
- QHUWWSQZTFLXPQ-FJXKBIBVSA-N Thr-Met-Gly Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCSC)C(=O)NCC(O)=O QHUWWSQZTFLXPQ-FJXKBIBVSA-N 0.000 description 3
- WYLAVUAWOUVUCA-XVSYOHENSA-N Thr-Phe-Asp Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(O)=O)C(O)=O WYLAVUAWOUVUCA-XVSYOHENSA-N 0.000 description 3
- LECUEEHKUFYOOV-ZJDVBMNYSA-N Thr-Thr-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)O)NC(=O)[C@@H](N)[C@@H](C)O LECUEEHKUFYOOV-ZJDVBMNYSA-N 0.000 description 3
- OKAMOYTUQMIFJO-JBACZVJFSA-N Trp-Glu-Phe Chemical compound C([C@H](NC(=O)[C@H](CCC(O)=O)NC(=O)[C@H](CC=1C2=CC=CC=C2NC=1)N)C(O)=O)C1=CC=CC=C1 OKAMOYTUQMIFJO-JBACZVJFSA-N 0.000 description 3
- GAYLGYUVTDMLKC-UWJYBYFXSA-N Tyr-Asp-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 GAYLGYUVTDMLKC-UWJYBYFXSA-N 0.000 description 3
- BEIGSKUPTIFYRZ-SRVKXCTJSA-N Tyr-Asp-Asp Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CC(=O)O)C(=O)O)N)O BEIGSKUPTIFYRZ-SRVKXCTJSA-N 0.000 description 3
- WVRUKYLYMFGKAN-IHRRRGAJSA-N Tyr-Glu-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 WVRUKYLYMFGKAN-IHRRRGAJSA-N 0.000 description 3
- GULIUBBXCYPDJU-CQDKDKBSSA-N Tyr-Leu-Ala Chemical compound [O-]C(=O)[C@H](C)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H]([NH3+])CC1=CC=C(O)C=C1 GULIUBBXCYPDJU-CQDKDKBSSA-N 0.000 description 3
- KXUKIBHIVRYOIP-ZKWXMUAHSA-N Val-Asp-Cys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CS)C(=O)O)N KXUKIBHIVRYOIP-ZKWXMUAHSA-N 0.000 description 3
- XLDYBRXERHITNH-QSFUFRPTSA-N Val-Asp-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H](N)C(C)C XLDYBRXERHITNH-QSFUFRPTSA-N 0.000 description 3
- ZXAGTABZUOMUDO-GVXVVHGQSA-N Val-Glu-Lys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CCCCN)C(=O)O)N ZXAGTABZUOMUDO-GVXVVHGQSA-N 0.000 description 3
- PMXBARDFIAPBGK-DZKIICNBSA-N Val-Glu-Tyr Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 PMXBARDFIAPBGK-DZKIICNBSA-N 0.000 description 3
- YTPLVNUZZOBFFC-SCZZXKLOSA-N Val-Gly-Pro Chemical compound CC(C)[C@H](N)C(=O)NCC(=O)N1CCC[C@@H]1C(O)=O YTPLVNUZZOBFFC-SCZZXKLOSA-N 0.000 description 3
- XXROXFHCMVXETG-UWVGGRQHSA-N Val-Gly-Val Chemical compound CC(C)[C@H](N)C(=O)NCC(=O)N[C@@H](C(C)C)C(O)=O XXROXFHCMVXETG-UWVGGRQHSA-N 0.000 description 3
- RWOGENDAOGMHLX-DCAQKATOSA-N Val-Lys-Ala Chemical compound C[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](C(C)C)N RWOGENDAOGMHLX-DCAQKATOSA-N 0.000 description 3
- KTEZUXISLQTDDQ-NHCYSSNCSA-N Val-Lys-Asp Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(=O)O)C(=O)O)N KTEZUXISLQTDDQ-NHCYSSNCSA-N 0.000 description 3
- PHZGFLFMGLXCFG-FHWLQOOXSA-N Val-Lys-Trp Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)O)N PHZGFLFMGLXCFG-FHWLQOOXSA-N 0.000 description 3
- RYHUIHUOYRNNIE-NRPADANISA-N Val-Ser-Gln Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N RYHUIHUOYRNNIE-NRPADANISA-N 0.000 description 3
- 239000011543 agarose gel Substances 0.000 description 3
- 108010076324 alanyl-glycyl-glycine Proteins 0.000 description 3
- 108010077245 asparaginyl-proline Proteins 0.000 description 3
- 108010093581 aspartyl-proline Proteins 0.000 description 3
- 238000003556 assay Methods 0.000 description 3
- 244000309466 calf Species 0.000 description 3
- 108010054813 diprotin B Proteins 0.000 description 3
- 238000002474 experimental method Methods 0.000 description 3
- 239000000706 filtrate Substances 0.000 description 3
- 108010037850 glycylvaline Proteins 0.000 description 3
- 108010092114 histidylphenylalanine Proteins 0.000 description 3
- 238000002347 injection Methods 0.000 description 3
- 239000007924 injection Substances 0.000 description 3
- 201000005202 lung cancer Diseases 0.000 description 3
- 208000020816 lung neoplasm Diseases 0.000 description 3
- 230000013011 mating Effects 0.000 description 3
- 208000002154 non-small cell lung carcinoma Diseases 0.000 description 3
- 239000003002 pH adjusting agent Substances 0.000 description 3
- 239000000546 pharmaceutical excipient Substances 0.000 description 3
- 238000000746 purification Methods 0.000 description 3
- 230000035939 shock Effects 0.000 description 3
- 239000003381 stabilizer Substances 0.000 description 3
- 208000029729 tumor suppressor gene on chromosome 11 Diseases 0.000 description 3
- MTCFGRXMJLQNBG-REOHCLBHSA-N (2S)-2-Amino-3-hydroxypropansäure Chemical group OC[C@H](N)C(O)=O MTCFGRXMJLQNBG-REOHCLBHSA-N 0.000 description 2
- CWFMWBHMIMNZLN-NAKRPEOUSA-N (2s)-1-[(2s)-2-[[(2s,3s)-2-amino-3-methylpentanoyl]amino]propanoyl]pyrrolidine-2-carboxylic acid Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](C)C(=O)N1CCC[C@H]1C(O)=O CWFMWBHMIMNZLN-NAKRPEOUSA-N 0.000 description 2
- CBNVOPQSNMPGDO-UHFFFAOYSA-N 2-[[2-[[1-[2-amino-3-(4-hydroxyphenyl)propanoyl]pyrrolidine-2-carbonyl]amino]acetyl]amino]-3-methylpentanoic acid Chemical compound CCC(C)C(C(O)=O)NC(=O)CNC(=O)C1CCCN1C(=O)C(N)CC1=CC=C(O)C=C1 CBNVOPQSNMPGDO-UHFFFAOYSA-N 0.000 description 2
- QIBXPQVQBRFRIY-UHFFFAOYSA-N 4-amino-1-tert-butyl-N-(5-methyl-1H-pyrazol-3-yl)pyrazolo[3,4-d]pyrimidine-3-carboxamide Chemical compound NC1=C2C(=NC=N1)N(N=C2C(=O)NC1=NNC(=C1)C)C(C)(C)C QIBXPQVQBRFRIY-UHFFFAOYSA-N 0.000 description 2
- UFKINFUUYHPLQS-UHFFFAOYSA-N 4-amino-7-(1-fluoro-2-methylpropan-2-yl)-N-(5-methyl-1H-pyrazol-3-yl)pyrrolo[2,3-d]pyrimidine-5-carboxamide Chemical compound NC=1C2=C(N=CN=1)N(C=C2C(=O)NC1=NNC(=C1)C)C(CF)(C)C UFKINFUUYHPLQS-UHFFFAOYSA-N 0.000 description 2
- VKNSWNWSJODLGL-UHFFFAOYSA-N 4-amino-7-(1-methylcyclopropyl)-N-(5-methyl-1H-pyrazol-3-yl)pyrrolo[2,3-d]pyrimidine-5-carboxamide Chemical compound NC=1C2=C(N=CN=1)N(C=C2C(=O)NC1=NNC(=C1)C)C1(CC1)C VKNSWNWSJODLGL-UHFFFAOYSA-N 0.000 description 2
- ZJVPEIBBMXWONH-UHFFFAOYSA-N 4-amino-7-(2-cyclopropylpropan-2-yl)-N-(5-methyl-1H-pyrazol-3-yl)pyrrolo[2,3-d]pyrimidine-5-carboxamide Chemical compound NC=1C2=C(N=CN=1)N(C=C2C(=O)NC1=NNC(=C1)C)C(C)(C)C1CC1 ZJVPEIBBMXWONH-UHFFFAOYSA-N 0.000 description 2
- ZQADJVAMOQTKDB-UHFFFAOYSA-N 4-amino-7-tert-butyl-N-(5-methyl-1H-pyrazol-3-yl)pyrrolo[2,3-d]pyrimidine-5-carboxamide Chemical compound NC=1C2=C(N=CN=1)N(C=C2C(=O)NC1=NNC(=C1)C)C(C)(C)C ZQADJVAMOQTKDB-UHFFFAOYSA-N 0.000 description 2
- WJFXFPRKXLEDGY-UHFFFAOYSA-N 4-amino-N-[4-(methoxymethyl)phenyl]-7-(1-methylcyclopropyl)-6-[2-(oxan-4-yl)ethynyl]pyrrolo[2,3-d]pyrimidine-5-carboxamide Chemical compound NC=1C2=C(N=CN=1)N(C(=C2C(=O)NC1=CC=C(C=C1)COC)C#CC1CCOCC1)C1(CC1)C WJFXFPRKXLEDGY-UHFFFAOYSA-N 0.000 description 2
- IMIZPWSVYADSCN-UHFFFAOYSA-N 4-methyl-2-[[4-methyl-2-[[4-methyl-2-(pyrrolidine-2-carbonylamino)pentanoyl]amino]pentanoyl]amino]pentanoic acid Chemical compound CC(C)CC(C(O)=O)NC(=O)C(CC(C)C)NC(=O)C(CC(C)C)NC(=O)C1CCCN1 IMIZPWSVYADSCN-UHFFFAOYSA-N 0.000 description 2
- DVWVZSJAYIJZFI-FXQIFTODSA-N Ala-Arg-Asn Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(O)=O DVWVZSJAYIJZFI-FXQIFTODSA-N 0.000 description 2
- SVBXIUDNTRTKHE-CIUDSAMLSA-N Ala-Arg-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(O)=O SVBXIUDNTRTKHE-CIUDSAMLSA-N 0.000 description 2
- XEXJJJRVTFGWIC-FXQIFTODSA-N Ala-Asn-Arg Chemical compound C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N XEXJJJRVTFGWIC-FXQIFTODSA-N 0.000 description 2
- WDIYWDJLXOCGRW-ACZMJKKPSA-N Ala-Asp-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O WDIYWDJLXOCGRW-ACZMJKKPSA-N 0.000 description 2
- ZIWWTZWAKYBUOB-CIUDSAMLSA-N Ala-Asp-Leu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O ZIWWTZWAKYBUOB-CIUDSAMLSA-N 0.000 description 2
- OILNWMNBLIHXQK-ZLUOBGJFSA-N Ala-Cys-Ser Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CS)C(=O)N[C@@H](CO)C(O)=O OILNWMNBLIHXQK-ZLUOBGJFSA-N 0.000 description 2
- ZODMADSIQZZBSQ-FXQIFTODSA-N Ala-Gln-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O ZODMADSIQZZBSQ-FXQIFTODSA-N 0.000 description 2
- SFNFGFDRYJKZKN-XQXXSGGOSA-N Ala-Gln-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](C)N)O SFNFGFDRYJKZKN-XQXXSGGOSA-N 0.000 description 2
- YIGLXQRFQVWFEY-NRPADANISA-N Ala-Gln-Val Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(O)=O YIGLXQRFQVWFEY-NRPADANISA-N 0.000 description 2
- NJPMYXWVWQWCSR-ACZMJKKPSA-N Ala-Glu-Asn Chemical compound C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O NJPMYXWVWQWCSR-ACZMJKKPSA-N 0.000 description 2
- UHMQKOBNPRAZGB-CIUDSAMLSA-N Ala-Glu-Met Chemical compound C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CCSC)C(=O)O)N UHMQKOBNPRAZGB-CIUDSAMLSA-N 0.000 description 2
- PCIFXPRIFWKWLK-YUMQZZPRSA-N Ala-Gly-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)CNC(=O)[C@H](C)N PCIFXPRIFWKWLK-YUMQZZPRSA-N 0.000 description 2
- NIZKGBJVCMRDKO-KWQFWETISA-N Ala-Gly-Tyr Chemical compound C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 NIZKGBJVCMRDKO-KWQFWETISA-N 0.000 description 2
- JDIQCVUDDFENPU-ZKWXMUAHSA-N Ala-His-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@@H](NC(=O)[C@@H](N)C)CC1=CNC=N1 JDIQCVUDDFENPU-ZKWXMUAHSA-N 0.000 description 2
- MEFILNJXAVSUTO-JXUBOQSCSA-N Ala-Leu-Thr Chemical compound C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O MEFILNJXAVSUTO-JXUBOQSCSA-N 0.000 description 2
- SDZRIBWEVVRDQI-CIUDSAMLSA-N Ala-Lys-Asp Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(O)=O SDZRIBWEVVRDQI-CIUDSAMLSA-N 0.000 description 2
- NLOMBWNGESDVJU-GUBZILKMSA-N Ala-Met-Arg Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O NLOMBWNGESDVJU-GUBZILKMSA-N 0.000 description 2
- HYIDEIQUCBKIPL-CQDKDKBSSA-N Ala-Phe-His Chemical compound C[C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)N HYIDEIQUCBKIPL-CQDKDKBSSA-N 0.000 description 2
- WEZNQZHACPSMEF-QEJZJMRPSA-N Ala-Phe-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)C)CC1=CC=CC=C1 WEZNQZHACPSMEF-QEJZJMRPSA-N 0.000 description 2
- RNHKOQHGYMTHFR-UBHSHLNASA-N Ala-Phe-Met Chemical compound CSCC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@H](C)N)CC1=CC=CC=C1 RNHKOQHGYMTHFR-UBHSHLNASA-N 0.000 description 2
- MSWSRLGNLKHDEI-ACZMJKKPSA-N Ala-Ser-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(O)=O MSWSRLGNLKHDEI-ACZMJKKPSA-N 0.000 description 2
- NZGRHTKZFSVPAN-BIIVOSGPSA-N Ala-Ser-Pro Chemical compound C[C@@H](C(=O)N[C@@H](CO)C(=O)N1CCC[C@@H]1C(=O)O)N NZGRHTKZFSVPAN-BIIVOSGPSA-N 0.000 description 2
- JJHBEVZAZXZREW-LFSVMHDDSA-N Ala-Thr-Phe Chemical compound C[C@@H](O)[C@H](NC(=O)[C@H](C)N)C(=O)N[C@@H](Cc1ccccc1)C(O)=O JJHBEVZAZXZREW-LFSVMHDDSA-N 0.000 description 2
- IEAUDUOCWNPZBR-LKTVYLICSA-N Ala-Trp-Gln Chemical compound C[C@@H](C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N IEAUDUOCWNPZBR-LKTVYLICSA-N 0.000 description 2
- ANNKVZSFQJGVDY-XUXIUFHCSA-N Ala-Val-Pro-Pro Chemical compound C[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 ANNKVZSFQJGVDY-XUXIUFHCSA-N 0.000 description 2
- KWKQGHSSNHPGOW-BQBZGAKWSA-N Arg-Ala-Gly Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C)C(=O)NCC(O)=O KWKQGHSSNHPGOW-BQBZGAKWSA-N 0.000 description 2
- BIOCIVSVEDFKDJ-GUBZILKMSA-N Arg-Arg-Asp Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(O)=O BIOCIVSVEDFKDJ-GUBZILKMSA-N 0.000 description 2
- OVVUNXXROOFSIM-SDDRHHMPSA-N Arg-Arg-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCCN=C(N)N)N)C(=O)O OVVUNXXROOFSIM-SDDRHHMPSA-N 0.000 description 2
- QPOARHANPULOTM-GMOBBJLQSA-N Arg-Asn-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@H](CCCN=C(N)N)N QPOARHANPULOTM-GMOBBJLQSA-N 0.000 description 2
- BVBKBQRPOJFCQM-DCAQKATOSA-N Arg-Asn-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O BVBKBQRPOJFCQM-DCAQKATOSA-N 0.000 description 2
- IIABBYGHLYWVOS-FXQIFTODSA-N Arg-Asn-Ser Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(O)=O IIABBYGHLYWVOS-FXQIFTODSA-N 0.000 description 2
- FBLMOFHNVQBKRR-IHRRRGAJSA-N Arg-Asp-Tyr Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 FBLMOFHNVQBKRR-IHRRRGAJSA-N 0.000 description 2
- BJNUAWGXPSHQMJ-DCAQKATOSA-N Arg-Gln-Met Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCSC)C(O)=O BJNUAWGXPSHQMJ-DCAQKATOSA-N 0.000 description 2
- LLZXKVAAEWBUPB-KKUMJFAQSA-N Arg-Gln-Phe Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O LLZXKVAAEWBUPB-KKUMJFAQSA-N 0.000 description 2
- HPKSHFSEXICTLI-CIUDSAMLSA-N Arg-Glu-Ala Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O HPKSHFSEXICTLI-CIUDSAMLSA-N 0.000 description 2
- XLWSGICNBZGYTA-CIUDSAMLSA-N Arg-Glu-Asp Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O XLWSGICNBZGYTA-CIUDSAMLSA-N 0.000 description 2
- UFBURHXMKFQVLM-CIUDSAMLSA-N Arg-Glu-Ser Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(O)=O UFBURHXMKFQVLM-CIUDSAMLSA-N 0.000 description 2
- OQCWXQJLCDPRHV-UWVGGRQHSA-N Arg-Gly-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)NCC(=O)N[C@@H](CC(C)C)C(O)=O OQCWXQJLCDPRHV-UWVGGRQHSA-N 0.000 description 2
- AGVNTAUPLWIQEN-ZPFDUUQYSA-N Arg-Ile-Glu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N AGVNTAUPLWIQEN-ZPFDUUQYSA-N 0.000 description 2
- UHFUZWSZQKMDSX-DCAQKATOSA-N Arg-Leu-Asn Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N UHFUZWSZQKMDSX-DCAQKATOSA-N 0.000 description 2
- GMFAGHNRXPSSJS-SRVKXCTJSA-N Arg-Leu-Gln Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O GMFAGHNRXPSSJS-SRVKXCTJSA-N 0.000 description 2
- OTZMRMHZCMZOJZ-SRVKXCTJSA-N Arg-Leu-Glu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O OTZMRMHZCMZOJZ-SRVKXCTJSA-N 0.000 description 2
- WMEVEPXNCMKNGH-IHRRRGAJSA-N Arg-Leu-His Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N WMEVEPXNCMKNGH-IHRRRGAJSA-N 0.000 description 2
- COXMUHNBYCVVRG-DCAQKATOSA-N Arg-Leu-Ser Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O COXMUHNBYCVVRG-DCAQKATOSA-N 0.000 description 2
- SSZGOKWBHLOCHK-DCAQKATOSA-N Arg-Lys-Asn Chemical compound NC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CCCN=C(N)N SSZGOKWBHLOCHK-DCAQKATOSA-N 0.000 description 2
- CVXXSWQORBZAAA-SRVKXCTJSA-N Arg-Lys-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CCCN=C(N)N CVXXSWQORBZAAA-SRVKXCTJSA-N 0.000 description 2
- DIIGDGJKTMLQQW-IHRRRGAJSA-N Arg-Lys-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](CCCN=C(N)N)N DIIGDGJKTMLQQW-IHRRRGAJSA-N 0.000 description 2
- MTYLORHAQXVQOW-AVGNSLFASA-N Arg-Lys-Met Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCSC)C(O)=O MTYLORHAQXVQOW-AVGNSLFASA-N 0.000 description 2
- PAPSMOYMQDWIOR-AVGNSLFASA-N Arg-Lys-Val Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C(C)C)C(O)=O PAPSMOYMQDWIOR-AVGNSLFASA-N 0.000 description 2
- YCYXHLZRUSJITQ-SRVKXCTJSA-N Arg-Pro-Pro Chemical compound NC(=N)NCCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 YCYXHLZRUSJITQ-SRVKXCTJSA-N 0.000 description 2
- VUGWHBXPMAHEGZ-SRVKXCTJSA-N Arg-Pro-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CCCN=C(N)N VUGWHBXPMAHEGZ-SRVKXCTJSA-N 0.000 description 2
- VRTWYUYCJGNFES-CIUDSAMLSA-N Arg-Ser-Gln Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(O)=O VRTWYUYCJGNFES-CIUDSAMLSA-N 0.000 description 2
- INOIAEUXVVNJKA-XGEHTFHBSA-N Arg-Thr-Ser Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(O)=O INOIAEUXVVNJKA-XGEHTFHBSA-N 0.000 description 2
- FOWOZYAWODIRFZ-JYJNAYRXSA-N Arg-Tyr-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CC1=CC=C(C=C1)O)NC(=O)[C@H](CCCN=C(N)N)N FOWOZYAWODIRFZ-JYJNAYRXSA-N 0.000 description 2
- QCTOLCVIGRLMQS-HRCADAONSA-N Arg-Tyr-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CC=C(C=C2)O)NC(=O)[C@H](CCCN=C(N)N)N)C(=O)O QCTOLCVIGRLMQS-HRCADAONSA-N 0.000 description 2
- ISVACHFCVRKIDG-SRVKXCTJSA-N Arg-Val-Arg Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O ISVACHFCVRKIDG-SRVKXCTJSA-N 0.000 description 2
- NUHQMYUWLUSRJX-BIIVOSGPSA-N Asn-Ala-Pro Chemical compound C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CC(=O)N)N NUHQMYUWLUSRJX-BIIVOSGPSA-N 0.000 description 2
- XWGJDUSDTRPQRK-ZLUOBGJFSA-N Asn-Ala-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CC(N)=O XWGJDUSDTRPQRK-ZLUOBGJFSA-N 0.000 description 2
- VYLVOMUVLMGCRF-ZLUOBGJFSA-N Asn-Asp-Ser Chemical compound NC(=O)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O VYLVOMUVLMGCRF-ZLUOBGJFSA-N 0.000 description 2
- PQAIOUVVZCOLJK-FXQIFTODSA-N Asn-Gln-Gln Chemical compound C(CC(=O)N)[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CC(=O)N)N PQAIOUVVZCOLJK-FXQIFTODSA-N 0.000 description 2
- ASCGFDYEKSRNPL-CIUDSAMLSA-N Asn-Glu-Met Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCSC)C(O)=O ASCGFDYEKSRNPL-CIUDSAMLSA-N 0.000 description 2
- SUIJFTJDTJKSRK-IHRRRGAJSA-N Asn-Pro-Tyr Chemical compound NC(=O)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 SUIJFTJDTJKSRK-IHRRRGAJSA-N 0.000 description 2
- JWQWPRCDYWNVNM-ACZMJKKPSA-N Asn-Ser-Gln Chemical compound C(CC(=O)N)[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CC(=O)N)N JWQWPRCDYWNVNM-ACZMJKKPSA-N 0.000 description 2
- LUJQEUOZJUWRRX-BPUTZDHNSA-N Asn-Trp-Met Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CCSC)C(O)=O LUJQEUOZJUWRRX-BPUTZDHNSA-N 0.000 description 2
- ANRZCQXIXGDXLR-CWRNSKLLSA-N Asn-Trp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CNC3=CC=CC=C32)NC(=O)[C@H](CC(=O)N)N)C(=O)O ANRZCQXIXGDXLR-CWRNSKLLSA-N 0.000 description 2
- XBQSLMACWDXWLJ-GHCJXIJMSA-N Asp-Ala-Ile Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O XBQSLMACWDXWLJ-GHCJXIJMSA-N 0.000 description 2
- MUWDILPCTSMUHI-ZLUOBGJFSA-N Asp-Asn-Cys Chemical compound C([C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CS)C(=O)O)N)C(=O)O MUWDILPCTSMUHI-ZLUOBGJFSA-N 0.000 description 2
- FANQWNCPNFEPGZ-WHFBIAKZSA-N Asp-Asp-Gly Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(O)=O)C(=O)NCC(O)=O FANQWNCPNFEPGZ-WHFBIAKZSA-N 0.000 description 2
- ZRAOLTNMSCSCLN-ZLUOBGJFSA-N Asp-Cys-Asn Chemical compound C([C@@H](C(=O)N[C@@H](CS)C(=O)N[C@@H](CC(=O)N)C(=O)O)N)C(=O)O ZRAOLTNMSCSCLN-ZLUOBGJFSA-N 0.000 description 2
- DZQKLNLLWFQONU-LKXGYXEUSA-N Asp-Cys-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CS)NC(=O)[C@H](CC(=O)O)N)O DZQKLNLLWFQONU-LKXGYXEUSA-N 0.000 description 2
- IJHUZMGJRGNXIW-CIUDSAMLSA-N Asp-Glu-Arg Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O IJHUZMGJRGNXIW-CIUDSAMLSA-N 0.000 description 2
- XDGBFDYXZCMYEX-NUMRIWBASA-N Asp-Glu-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CC(=O)O)N)O XDGBFDYXZCMYEX-NUMRIWBASA-N 0.000 description 2
- YDJVIBMKAMQPPP-LAEOZQHASA-N Asp-Glu-Val Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C(C)C)C(O)=O YDJVIBMKAMQPPP-LAEOZQHASA-N 0.000 description 2
- QCVXMEHGFUMKCO-YUMQZZPRSA-N Asp-Gly-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CC(O)=O QCVXMEHGFUMKCO-YUMQZZPRSA-N 0.000 description 2
- POTCZYQVVNXUIG-BQBZGAKWSA-N Asp-Gly-Pro Chemical compound OC(=O)C[C@H](N)C(=O)NCC(=O)N1CCC[C@H]1C(O)=O POTCZYQVVNXUIG-BQBZGAKWSA-N 0.000 description 2
- TVIZQBFURPLQDV-DJFWLOJKSA-N Asp-His-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CC1=CN=CN1)NC(=O)[C@H](CC(=O)O)N TVIZQBFURPLQDV-DJFWLOJKSA-N 0.000 description 2
- UBPMOJLRVMGTOQ-GARJFASQSA-N Asp-His-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CN=CN2)NC(=O)[C@H](CC(=O)O)N)C(=O)O UBPMOJLRVMGTOQ-GARJFASQSA-N 0.000 description 2
- TZOZNVLBTAFJRW-UGYAYLCHSA-N Asp-Ile-Asp Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)O)NC(=O)[C@H](CC(=O)O)N TZOZNVLBTAFJRW-UGYAYLCHSA-N 0.000 description 2
- KLYPOCBLKMPBIQ-GHCJXIJMSA-N Asp-Ile-Ser Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)[C@H](CC(=O)O)N KLYPOCBLKMPBIQ-GHCJXIJMSA-N 0.000 description 2
- SPWXXPFDTMYTRI-IUKAMOBKSA-N Asp-Ile-Thr Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H]([C@@H](C)O)C(O)=O SPWXXPFDTMYTRI-IUKAMOBKSA-N 0.000 description 2
- GYWQGGUCMDCUJE-DLOVCJGASA-N Asp-Phe-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](C)C(O)=O GYWQGGUCMDCUJE-DLOVCJGASA-N 0.000 description 2
- QJHOOKBAHRJPPX-QWRGUYRKSA-N Asp-Phe-Gly Chemical compound OC(=O)C[C@H](N)C(=O)N[C@H](C(=O)NCC(O)=O)CC1=CC=CC=C1 QJHOOKBAHRJPPX-QWRGUYRKSA-N 0.000 description 2
- WMLFFCRUSPNENW-ZLUOBGJFSA-N Asp-Ser-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O WMLFFCRUSPNENW-ZLUOBGJFSA-N 0.000 description 2
- QSFHZPQUAAQHAQ-CIUDSAMLSA-N Asp-Ser-Leu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O QSFHZPQUAAQHAQ-CIUDSAMLSA-N 0.000 description 2
- MNQMTYSEKZHIDF-GCJQMDKQSA-N Asp-Thr-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(O)=O MNQMTYSEKZHIDF-GCJQMDKQSA-N 0.000 description 2
- GXHDGYOXPNQCKM-XVSYOHENSA-N Asp-Thr-Phe Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)O)NC(=O)[C@H](CC(=O)O)N)O GXHDGYOXPNQCKM-XVSYOHENSA-N 0.000 description 2
- ITGFVUYOLWBPQW-KKHAAJSZSA-N Asp-Thr-Val Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C(C)C)C(O)=O ITGFVUYOLWBPQW-KKHAAJSZSA-N 0.000 description 2
- IJGRMHOSHXDMSA-UHFFFAOYSA-N Atomic nitrogen Chemical compound N#N IJGRMHOSHXDMSA-UHFFFAOYSA-N 0.000 description 2
- 206010009944 Colon cancer Diseases 0.000 description 2
- KIQKJXYVGSYDFS-ZLUOBGJFSA-N Cys-Asn-Asn Chemical compound SC[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O KIQKJXYVGSYDFS-ZLUOBGJFSA-N 0.000 description 2
- ABLJDBFJPUWQQB-DCAQKATOSA-N Cys-Leu-Arg Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)NC(=O)[C@H](CS)N ABLJDBFJPUWQQB-DCAQKATOSA-N 0.000 description 2
- KGIHMGPYGXBYJJ-SRVKXCTJSA-N Cys-Lys-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CS KGIHMGPYGXBYJJ-SRVKXCTJSA-N 0.000 description 2
- ZGERHCJBLPQPGV-ACZMJKKPSA-N Cys-Ser-Gln Chemical compound C(CC(=O)N)[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CS)N ZGERHCJBLPQPGV-ACZMJKKPSA-N 0.000 description 2
- LKHMGNHQULEPFY-ACZMJKKPSA-N Cys-Ser-Glu Chemical compound SC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(O)=O LKHMGNHQULEPFY-ACZMJKKPSA-N 0.000 description 2
- NDNZRWUDUMTITL-FXQIFTODSA-N Cys-Ser-Val Chemical compound [H]N[C@@H](CS)C(=O)N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(O)=O NDNZRWUDUMTITL-FXQIFTODSA-N 0.000 description 2
- 102000053602 DNA Human genes 0.000 description 2
- 102000012410 DNA Ligases Human genes 0.000 description 2
- 108010061982 DNA Ligases Proteins 0.000 description 2
- 102000016928 DNA-directed DNA polymerase Human genes 0.000 description 2
- 108010014303 DNA-directed DNA polymerase Proteins 0.000 description 2
- 101710195747 Dynactin subunit 1 Proteins 0.000 description 2
- 241000196324 Embryophyta Species 0.000 description 2
- 108010067770 Endopeptidase K Proteins 0.000 description 2
- WSFSSNUMVMOOMR-UHFFFAOYSA-N Formaldehyde Chemical compound O=C WSFSSNUMVMOOMR-UHFFFAOYSA-N 0.000 description 2
- 241000233866 Fungi Species 0.000 description 2
- WUAYFMZULZDSLB-ACZMJKKPSA-N Gln-Ala-Asn Chemical compound NC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCC(N)=O WUAYFMZULZDSLB-ACZMJKKPSA-N 0.000 description 2
- XXLBHPPXDUWYAG-XQXXSGGOSA-N Gln-Ala-Thr Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O XXLBHPPXDUWYAG-XQXXSGGOSA-N 0.000 description 2
- LZRMPXRYLLTAJX-GUBZILKMSA-N Gln-Arg-Glu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(O)=O LZRMPXRYLLTAJX-GUBZILKMSA-N 0.000 description 2
- PRBLYKYHAJEABA-SRVKXCTJSA-N Gln-Arg-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(O)=O PRBLYKYHAJEABA-SRVKXCTJSA-N 0.000 description 2
- XOKGKOQWADCLFQ-GARJFASQSA-N Gln-Arg-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCC(=O)N)N)C(=O)O XOKGKOQWADCLFQ-GARJFASQSA-N 0.000 description 2
- INFBPLSHYFALDE-ACZMJKKPSA-N Gln-Asn-Ala Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](C)C(O)=O INFBPLSHYFALDE-ACZMJKKPSA-N 0.000 description 2
- WQWMZOIPXWSZNE-WDSKDSINSA-N Gln-Asp-Gly Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(O)=O)C(=O)NCC(O)=O WQWMZOIPXWSZNE-WDSKDSINSA-N 0.000 description 2
- MFLMFRZBAJSGHK-ACZMJKKPSA-N Gln-Cys-Ser Chemical compound C(CC(=O)N)[C@@H](C(=O)N[C@@H](CS)C(=O)N[C@@H](CO)C(=O)O)N MFLMFRZBAJSGHK-ACZMJKKPSA-N 0.000 description 2
- XEZWLWNGUMXAJI-QEJZJMRPSA-N Gln-Cys-Trp Chemical compound C1=CC=C2C(C[C@H](NC(=O)[C@H](CS)NC(=O)[C@H](CCC(N)=O)N)C(O)=O)=CNC2=C1 XEZWLWNGUMXAJI-QEJZJMRPSA-N 0.000 description 2
- AJDMYLOISOCHHC-YVNDNENWSA-N Gln-Gln-Ile Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O AJDMYLOISOCHHC-YVNDNENWSA-N 0.000 description 2
- LWDGZZGWDMHBOF-FXQIFTODSA-N Gln-Glu-Asn Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O LWDGZZGWDMHBOF-FXQIFTODSA-N 0.000 description 2
- LLRJEFPKIIBGJP-DCAQKATOSA-N Gln-Glu-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CCC(=O)N)N LLRJEFPKIIBGJP-DCAQKATOSA-N 0.000 description 2
- DRDSQGHKTLSNEA-GLLZPBPUSA-N Gln-Glu-Thr Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O DRDSQGHKTLSNEA-GLLZPBPUSA-N 0.000 description 2
- QQAPDATZKKTBIY-YUMQZZPRSA-N Gln-Gly-Met Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)NCC(=O)N[C@@H](CCSC)C(O)=O QQAPDATZKKTBIY-YUMQZZPRSA-N 0.000 description 2
- GLEGHWQNGPMKHO-DCAQKATOSA-N Gln-His-Glu Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)O)NC(=O)[C@H](CCC(=O)N)N GLEGHWQNGPMKHO-DCAQKATOSA-N 0.000 description 2
- IWUFOVSLWADEJC-AVGNSLFASA-N Gln-His-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(C)C)C(O)=O IWUFOVSLWADEJC-AVGNSLFASA-N 0.000 description 2
- JKGHMESJHRTHIC-SIUGBPQLSA-N Gln-Ile-Tyr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)O)NC(=O)[C@H](CCC(=O)N)N JKGHMESJHRTHIC-SIUGBPQLSA-N 0.000 description 2
- QBLMTCRYYTVUQY-GUBZILKMSA-N Gln-Leu-Asp Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O QBLMTCRYYTVUQY-GUBZILKMSA-N 0.000 description 2
- GURIQZQSTBBHRV-SRVKXCTJSA-N Gln-Lys-Arg Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O GURIQZQSTBBHRV-SRVKXCTJSA-N 0.000 description 2
- FKXCBKCOSVIGCT-AVGNSLFASA-N Gln-Lys-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O FKXCBKCOSVIGCT-AVGNSLFASA-N 0.000 description 2
- ZXGLLNZQSBLQLT-SRVKXCTJSA-N Gln-Met-Lys Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCC(=O)N)N ZXGLLNZQSBLQLT-SRVKXCTJSA-N 0.000 description 2
- UTOQQOMEJDPDMX-ACZMJKKPSA-N Gln-Ser-Asp Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O UTOQQOMEJDPDMX-ACZMJKKPSA-N 0.000 description 2
- MFHVAWMMKZBSRQ-ACZMJKKPSA-N Gln-Ser-Cys Chemical compound C(CC(=O)N)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CS)C(=O)O)N MFHVAWMMKZBSRQ-ACZMJKKPSA-N 0.000 description 2
- JKDBRTNMYXYLHO-JYJNAYRXSA-N Gln-Tyr-Leu Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CC(C)C)C(O)=O)CC1=CC=C(O)C=C1 JKDBRTNMYXYLHO-JYJNAYRXSA-N 0.000 description 2
- ZZLDMBMFKZFQMU-NRPADANISA-N Gln-Val-Ala Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C)C(O)=O ZZLDMBMFKZFQMU-NRPADANISA-N 0.000 description 2
- ICRKQMRFXYDYMK-LAEOZQHASA-N Gln-Val-Asn Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O ICRKQMRFXYDYMK-LAEOZQHASA-N 0.000 description 2
- RUFHOVYUYSNDNY-ACZMJKKPSA-N Glu-Ala-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCC(O)=O RUFHOVYUYSNDNY-ACZMJKKPSA-N 0.000 description 2
- SZXSSXUNOALWCH-ACZMJKKPSA-N Glu-Ala-Asn Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(N)=O)C(O)=O SZXSSXUNOALWCH-ACZMJKKPSA-N 0.000 description 2
- OGMQXTXGLDNBSS-FXQIFTODSA-N Glu-Ala-Gln Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(O)=O OGMQXTXGLDNBSS-FXQIFTODSA-N 0.000 description 2
- MXOODARRORARSU-ACZMJKKPSA-N Glu-Ala-Ser Chemical compound C[C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)[C@H](CCC(=O)O)N MXOODARRORARSU-ACZMJKKPSA-N 0.000 description 2
- WOSRKEJQESVHGA-CIUDSAMLSA-N Glu-Arg-Ser Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(O)=O WOSRKEJQESVHGA-CIUDSAMLSA-N 0.000 description 2
- RJONUNZIMUXUOI-GUBZILKMSA-N Glu-Asn-Lys Chemical compound C(CCN)C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@H](CCC(=O)O)N RJONUNZIMUXUOI-GUBZILKMSA-N 0.000 description 2
- NADWTMLCUDMDQI-ACZMJKKPSA-N Glu-Asp-Cys Chemical compound C(CC(=O)O)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CS)C(=O)O)N NADWTMLCUDMDQI-ACZMJKKPSA-N 0.000 description 2
- CKOFNWCLWRYUHK-XHNCKOQMSA-N Glu-Asp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)O)NC(=O)[C@H](CCC(=O)O)N)C(=O)O CKOFNWCLWRYUHK-XHNCKOQMSA-N 0.000 description 2
- JRCUFCXYZLPSDZ-ACZMJKKPSA-N Glu-Asp-Ser Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O JRCUFCXYZLPSDZ-ACZMJKKPSA-N 0.000 description 2
- XHWLNISLUFEWNS-CIUDSAMLSA-N Glu-Gln-Gln Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O XHWLNISLUFEWNS-CIUDSAMLSA-N 0.000 description 2
- ILGFBUGLBSAQQB-GUBZILKMSA-N Glu-Glu-Arg Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O ILGFBUGLBSAQQB-GUBZILKMSA-N 0.000 description 2
- NKLRYVLERDYDBI-FXQIFTODSA-N Glu-Glu-Asp Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O NKLRYVLERDYDBI-FXQIFTODSA-N 0.000 description 2
- QXDXIXFSFHUYAX-MNXVOIDGSA-N Glu-Ile-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)CC)NC(=O)[C@@H](N)CCC(O)=O QXDXIXFSFHUYAX-MNXVOIDGSA-N 0.000 description 2
- UGSVSNXPJJDJKL-SDDRHHMPSA-N Glu-Leu-Pro Chemical compound CC(C)C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCC(=O)O)N UGSVSNXPJJDJKL-SDDRHHMPSA-N 0.000 description 2
- NJCALAAIGREHDR-WDCWCFNPSA-N Glu-Leu-Thr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O NJCALAAIGREHDR-WDCWCFNPSA-N 0.000 description 2
- OQXDUSZKISQQSS-GUBZILKMSA-N Glu-Lys-Ala Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(O)=O OQXDUSZKISQQSS-GUBZILKMSA-N 0.000 description 2
- BCYGDJXHAGZNPQ-DCAQKATOSA-N Glu-Lys-Glu Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O BCYGDJXHAGZNPQ-DCAQKATOSA-N 0.000 description 2
- HRBYTAIBKPNZKQ-AVGNSLFASA-N Glu-Lys-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CCC(O)=O HRBYTAIBKPNZKQ-AVGNSLFASA-N 0.000 description 2
- ZGEJRLJEAMPEDV-SRVKXCTJSA-N Glu-Lys-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](CCC(=O)O)N ZGEJRLJEAMPEDV-SRVKXCTJSA-N 0.000 description 2
- MFNUFCFRAZPJFW-JYJNAYRXSA-N Glu-Lys-Phe Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 MFNUFCFRAZPJFW-JYJNAYRXSA-N 0.000 description 2
- NPMSEUWUMOSEFM-CIUDSAMLSA-N Glu-Met-Asn Chemical compound CSCC[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)O)NC(=O)[C@H](CCC(=O)O)N NPMSEUWUMOSEFM-CIUDSAMLSA-N 0.000 description 2
- ZTVGZOIBLRPQNR-KKUMJFAQSA-N Glu-Met-Tyr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O ZTVGZOIBLRPQNR-KKUMJFAQSA-N 0.000 description 2
- YRMZCZIRHYCNHX-RYUDHWBXSA-N Glu-Phe-Gly Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)NCC(O)=O YRMZCZIRHYCNHX-RYUDHWBXSA-N 0.000 description 2
- CBWKURKPYSLMJV-SOUVJXGZSA-N Glu-Phe-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CC=CC=C2)NC(=O)[C@H](CCC(=O)O)N)C(=O)O CBWKURKPYSLMJV-SOUVJXGZSA-N 0.000 description 2
- UDEPRBFQTWGLCW-CIUDSAMLSA-N Glu-Pro-Asp Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(O)=O)C(O)=O UDEPRBFQTWGLCW-CIUDSAMLSA-N 0.000 description 2
- WIKMTDVSCUJIPJ-CIUDSAMLSA-N Glu-Ser-Arg Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCCN=C(N)N WIKMTDVSCUJIPJ-CIUDSAMLSA-N 0.000 description 2
- JWNZHMSRZXXGTM-XKBZYTNZSA-N Glu-Ser-Thr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(O)=O JWNZHMSRZXXGTM-XKBZYTNZSA-N 0.000 description 2
- MWTGQXBHVRTCOR-GLLZPBPUSA-N Glu-Thr-Gln Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(N)=O)C(O)=O MWTGQXBHVRTCOR-GLLZPBPUSA-N 0.000 description 2
- UMZHHILWZBFPGL-LOKLDPHHSA-N Glu-Thr-Pro Chemical compound C[C@H]([C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCC(=O)O)N)O UMZHHILWZBFPGL-LOKLDPHHSA-N 0.000 description 2
- YOTHMZZSJKKEHZ-SZMVWBNQSA-N Glu-Trp-Lys Chemical compound C1=CC=C2C(C[C@@H](C(=O)N[C@@H](CCCCN)C(O)=O)NC(=O)[C@@H](N)CCC(O)=O)=CNC2=C1 YOTHMZZSJKKEHZ-SZMVWBNQSA-N 0.000 description 2
- HVKAAUOFFTUSAA-XDTLVQLUSA-N Glu-Tyr-Ala Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](C)C(O)=O HVKAAUOFFTUSAA-XDTLVQLUSA-N 0.000 description 2
- LZEUDRYSAZAJIO-AUTRQRHGSA-N Glu-Val-Glu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O LZEUDRYSAZAJIO-AUTRQRHGSA-N 0.000 description 2
- WQZGKKKJIJFFOK-GASJEMHNSA-N Glucose Natural products OC[C@H]1OC(O)[C@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-GASJEMHNSA-N 0.000 description 2
- VSVZIEVNUYDAFR-YUMQZZPRSA-N Gly-Ala-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)CN VSVZIEVNUYDAFR-YUMQZZPRSA-N 0.000 description 2
- QSDKBRMVXSWAQE-BFHQHQDPSA-N Gly-Ala-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)CN QSDKBRMVXSWAQE-BFHQHQDPSA-N 0.000 description 2
- XRTDOIOIBMAXCT-NKWVEPMBSA-N Gly-Asn-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)N)NC(=O)CN)C(=O)O XRTDOIOIBMAXCT-NKWVEPMBSA-N 0.000 description 2
- XTQFHTHIAKKCTM-YFKPBYRVSA-N Gly-Glu-Gly Chemical compound NCC(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O XTQFHTHIAKKCTM-YFKPBYRVSA-N 0.000 description 2
- LHRXAHLCRMQBGJ-RYUDHWBXSA-N Gly-Glu-Phe Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)CN LHRXAHLCRMQBGJ-RYUDHWBXSA-N 0.000 description 2
- QSVCIFZPGLOZGH-WDSKDSINSA-N Gly-Glu-Ser Chemical compound NCC(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(O)=O QSVCIFZPGLOZGH-WDSKDSINSA-N 0.000 description 2
- CUYLIWAAAYJKJH-RYUDHWBXSA-N Gly-Glu-Tyr Chemical compound NCC(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 CUYLIWAAAYJKJH-RYUDHWBXSA-N 0.000 description 2
- CCQOOWAONKGYKQ-BYPYZUCNSA-N Gly-Gly-Ala Chemical compound OC(=O)[C@H](C)NC(=O)CNC(=O)CN CCQOOWAONKGYKQ-BYPYZUCNSA-N 0.000 description 2
- YWAQATDNEKZFFK-BYPYZUCNSA-N Gly-Gly-Ser Chemical compound NCC(=O)NCC(=O)N[C@@H](CO)C(O)=O YWAQATDNEKZFFK-BYPYZUCNSA-N 0.000 description 2
- SWQALSGKVLYKDT-ZKWXMUAHSA-N Gly-Ile-Ala Chemical compound NCC(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](C)C(O)=O SWQALSGKVLYKDT-ZKWXMUAHSA-N 0.000 description 2
- FXGRXIATVXUAHO-WEDXCCLWSA-N Gly-Lys-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)CN)CCCCN FXGRXIATVXUAHO-WEDXCCLWSA-N 0.000 description 2
- OQQKUTVULYLCDG-ONGXEEELSA-N Gly-Lys-Val Chemical compound CC(C)[C@H](NC(=O)[C@H](CCCCN)NC(=O)CN)C(O)=O OQQKUTVULYLCDG-ONGXEEELSA-N 0.000 description 2
- JPVGHHQGKPQYIL-KBPBESRZSA-N Gly-Phe-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)CN)CC1=CC=CC=C1 JPVGHHQGKPQYIL-KBPBESRZSA-N 0.000 description 2
- IEGFSKKANYKBDU-QWHCGFSZSA-N Gly-Phe-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CC=CC=C2)NC(=O)CN)C(=O)O IEGFSKKANYKBDU-QWHCGFSZSA-N 0.000 description 2
- HAOUOFNNJJLVNS-BQBZGAKWSA-N Gly-Pro-Ser Chemical compound NCC(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O HAOUOFNNJJLVNS-BQBZGAKWSA-N 0.000 description 2
- IRJWAYCXIYUHQE-WHFBIAKZSA-N Gly-Ser-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](CO)NC(=O)CN IRJWAYCXIYUHQE-WHFBIAKZSA-N 0.000 description 2
- OHUKZZYSJBKFRR-WHFBIAKZSA-N Gly-Ser-Asp Chemical compound [H]NCC(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O OHUKZZYSJBKFRR-WHFBIAKZSA-N 0.000 description 2
- MYXNLWDWWOTERK-BHNWBGBOSA-N Gly-Thr-Pro Chemical compound C[C@H]([C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)CN)O MYXNLWDWWOTERK-BHNWBGBOSA-N 0.000 description 2
- GBYYQVBXFVDJPJ-WLTAIBSBSA-N Gly-Tyr-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CC1=CC=C(C=C1)O)NC(=O)CN)O GBYYQVBXFVDJPJ-WLTAIBSBSA-N 0.000 description 2
- BAYQNCWLXIDLHX-ONGXEEELSA-N Gly-Val-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](C(C)C)NC(=O)CN BAYQNCWLXIDLHX-ONGXEEELSA-N 0.000 description 2
- DHMQDGOQFOQNFH-UHFFFAOYSA-N Glycine Chemical compound NCC(O)=O DHMQDGOQFOQNFH-UHFFFAOYSA-N 0.000 description 2
- 241000238631 Hexapoda Species 0.000 description 2
- VCDNHBNNPCDBKV-DLOVCJGASA-N His-Ala-Lys Chemical compound C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CC1=CN=CN1)N VCDNHBNNPCDBKV-DLOVCJGASA-N 0.000 description 2
- SVHKVHBPTOMLTO-DCAQKATOSA-N His-Arg-Asp Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(O)=O SVHKVHBPTOMLTO-DCAQKATOSA-N 0.000 description 2
- UCDWNBFOZCZSNV-AVGNSLFASA-N His-Arg-Met Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCSC)C(O)=O UCDWNBFOZCZSNV-AVGNSLFASA-N 0.000 description 2
- WTJBVCUCLWFGAH-JUKXBJQTSA-N His-Ile-Tyr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)O)NC(=O)[C@H](CC2=CN=CN2)N WTJBVCUCLWFGAH-JUKXBJQTSA-N 0.000 description 2
- VYUXYMRNGALHEA-DLOVCJGASA-N His-Leu-Ala Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(O)=O VYUXYMRNGALHEA-DLOVCJGASA-N 0.000 description 2
- UROVZOUMHNXPLZ-AVGNSLFASA-N His-Leu-Gln Chemical compound NC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC1=CN=CN1 UROVZOUMHNXPLZ-AVGNSLFASA-N 0.000 description 2
- LVWIJITYHRZHBO-IXOXFDKPSA-N His-Leu-Thr Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O LVWIJITYHRZHBO-IXOXFDKPSA-N 0.000 description 2
- LDFWDDVELNOGII-MXAVVETBSA-N His-Lys-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](CC1=CN=CN1)N LDFWDDVELNOGII-MXAVVETBSA-N 0.000 description 2
- FJCGVRRVBKYYOU-DCAQKATOSA-N His-Met-Ser Chemical compound CSCC[C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)[C@H](CC1=CN=CN1)N FJCGVRRVBKYYOU-DCAQKATOSA-N 0.000 description 2
- XIGFLVCAVQQGNS-IHRRRGAJSA-N His-Pro-His Chemical compound C([C@H](N)C(=O)N1[C@@H](CCC1)C(=O)N[C@@H](CC=1NC=NC=1)C(O)=O)C1=CN=CN1 XIGFLVCAVQQGNS-IHRRRGAJSA-N 0.000 description 2
- IXQGOKWTQPCIQM-YJRXYDGGSA-N His-Thr-His Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@H](CC2=CN=CN2)N)O IXQGOKWTQPCIQM-YJRXYDGGSA-N 0.000 description 2
- DRKZDEFADVYTLU-AVGNSLFASA-N His-Val-Val Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C(C)C)C(O)=O DRKZDEFADVYTLU-AVGNSLFASA-N 0.000 description 2
- 241000282412 Homo Species 0.000 description 2
- 241000725303 Human immunodeficiency virus Species 0.000 description 2
- GRRNUXAQVGOGFE-UHFFFAOYSA-N Hygromycin-B Natural products OC1C(NC)CC(N)C(O)C1OC1C2OC3(C(C(O)C(O)C(C(N)CO)O3)O)OC2C(O)C(CO)O1 GRRNUXAQVGOGFE-UHFFFAOYSA-N 0.000 description 2
- HLYBGMZJVDHJEO-CYDGBPFRSA-N Ile-Arg-Arg Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N HLYBGMZJVDHJEO-CYDGBPFRSA-N 0.000 description 2
- YOTNPRLPIPHQSB-XUXIUFHCSA-N Ile-Arg-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCCN)C(=O)O)N YOTNPRLPIPHQSB-XUXIUFHCSA-N 0.000 description 2
- AWTDTFXPVCTHAK-BJDJZHNGSA-N Ile-Cys-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CS)C(=O)N[C@@H](CCCCN)C(=O)O)N AWTDTFXPVCTHAK-BJDJZHNGSA-N 0.000 description 2
- MTFVYKQRLXYAQN-LAEOZQHASA-N Ile-Glu-Gly Chemical compound [H]N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O MTFVYKQRLXYAQN-LAEOZQHASA-N 0.000 description 2
- TVYWVSJGSHQWMT-AJNGGQMLSA-N Ile-Leu-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)O)N TVYWVSJGSHQWMT-AJNGGQMLSA-N 0.000 description 2
- DSDPLOODKXISDT-XUXIUFHCSA-N Ile-Leu-Val Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C(C)C)C(O)=O DSDPLOODKXISDT-XUXIUFHCSA-N 0.000 description 2
- ADDYYRVQQZFIMW-MNXVOIDGSA-N Ile-Lys-Glu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N ADDYYRVQQZFIMW-MNXVOIDGSA-N 0.000 description 2
- YSGBJIQXTIVBHZ-AJNGGQMLSA-N Ile-Lys-Leu Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O YSGBJIQXTIVBHZ-AJNGGQMLSA-N 0.000 description 2
- GVNNAHIRSDRIII-AJNGGQMLSA-N Ile-Lys-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)O)N GVNNAHIRSDRIII-AJNGGQMLSA-N 0.000 description 2
- FFJQAEYLAQMGDL-MGHWNKPDSA-N Ile-Lys-Tyr Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 FFJQAEYLAQMGDL-MGHWNKPDSA-N 0.000 description 2
- KCTIFOCXAIUQQK-QXEWZRGKSA-N Ile-Pro-Gly Chemical compound CC[C@H](C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O KCTIFOCXAIUQQK-QXEWZRGKSA-N 0.000 description 2
- KTNGVMMGIQWIDV-OSUNSFLBSA-N Ile-Pro-Thr Chemical compound CC[C@H](C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(O)=O KTNGVMMGIQWIDV-OSUNSFLBSA-N 0.000 description 2
- MLSUZXHSNRBDCI-CYDGBPFRSA-N Ile-Pro-Val Chemical compound CC[C@H](C)[C@@H](C(=O)N1CCC[C@H]1C(=O)N[C@@H](C(C)C)C(=O)O)N MLSUZXHSNRBDCI-CYDGBPFRSA-N 0.000 description 2
- YKZAMJXNJUWFIK-JBDRJPRFSA-N Ile-Ser-Ala Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(=O)O)N YKZAMJXNJUWFIK-JBDRJPRFSA-N 0.000 description 2
- VGSPNSSCMOHRRR-BJDJZHNGSA-N Ile-Ser-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)O)N VGSPNSSCMOHRRR-BJDJZHNGSA-N 0.000 description 2
- ZDNNDIJTUHQCAM-MXAVVETBSA-N Ile-Ser-Phe Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)O)N ZDNNDIJTUHQCAM-MXAVVETBSA-N 0.000 description 2
- PZWBBXHHUSIGKH-OSUNSFLBSA-N Ile-Thr-Arg Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@H](C(O)=O)CCCN=C(N)N PZWBBXHHUSIGKH-OSUNSFLBSA-N 0.000 description 2
- OMDWJWGZGMCQND-CFMVVWHZSA-N Ile-Tyr-Asp Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)N[C@@H](CC(=O)O)C(=O)O)N OMDWJWGZGMCQND-CFMVVWHZSA-N 0.000 description 2
- YWCJXQKATPNPOE-UKJIMTQDSA-N Ile-Val-Glu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N YWCJXQKATPNPOE-UKJIMTQDSA-N 0.000 description 2
- APQYGMBHIVXFML-OSUNSFLBSA-N Ile-Val-Thr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](C(C)C)C(=O)N[C@@H]([C@@H](C)O)C(=O)O)N APQYGMBHIVXFML-OSUNSFLBSA-N 0.000 description 2
- WHUUTDBJXJRKMK-VKHMYHEASA-N L-glutamic acid Chemical group OC(=O)[C@@H](N)CCC(O)=O WHUUTDBJXJRKMK-VKHMYHEASA-N 0.000 description 2
- ZDXPYRJPNDTMRX-VKHMYHEASA-N L-glutamine Chemical compound OC(=O)[C@@H](N)CCC(N)=O ZDXPYRJPNDTMRX-VKHMYHEASA-N 0.000 description 2
- 229930182816 L-glutamine Natural products 0.000 description 2
- ROHFNLRQFUQHCH-YFKPBYRVSA-N L-leucine Chemical group CC(C)C[C@H](N)C(O)=O ROHFNLRQFUQHCH-YFKPBYRVSA-N 0.000 description 2
- FFEARJCKVFRZRR-BYPYZUCNSA-N L-methionine Chemical group CSCC[C@H](N)C(O)=O FFEARJCKVFRZRR-BYPYZUCNSA-N 0.000 description 2
- LZDNBBYBDGBADK-UHFFFAOYSA-N L-valyl-L-tryptophan Natural products C1=CC=C2C(CC(NC(=O)C(N)C(C)C)C(O)=O)=CNC2=C1 LZDNBBYBDGBADK-UHFFFAOYSA-N 0.000 description 2
- RFUBXQQFJFGJFV-GUBZILKMSA-N Leu-Asn-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O RFUBXQQFJFGJFV-GUBZILKMSA-N 0.000 description 2
- ZDSNOSQHMJBRQN-SRVKXCTJSA-N Leu-Asp-His Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N ZDSNOSQHMJBRQN-SRVKXCTJSA-N 0.000 description 2
- DLCOFDAHNMMQPP-SRVKXCTJSA-N Leu-Asp-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O DLCOFDAHNMMQPP-SRVKXCTJSA-N 0.000 description 2
- MYGQXVYRZMKRDB-SRVKXCTJSA-N Leu-Asp-Lys Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@H](C(O)=O)CCCCN MYGQXVYRZMKRDB-SRVKXCTJSA-N 0.000 description 2
- JQSXWJXBASFONF-KKUMJFAQSA-N Leu-Asp-Phe Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O JQSXWJXBASFONF-KKUMJFAQSA-N 0.000 description 2
- PVMPDMIKUVNOBD-CIUDSAMLSA-N Leu-Asp-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O PVMPDMIKUVNOBD-CIUDSAMLSA-N 0.000 description 2
- BOFAFKVZQUMTID-AVGNSLFASA-N Leu-Gln-His Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N BOFAFKVZQUMTID-AVGNSLFASA-N 0.000 description 2
- KGCLIYGPQXUNLO-IUCAKERBSA-N Leu-Gly-Glu Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CCC(O)=O KGCLIYGPQXUNLO-IUCAKERBSA-N 0.000 description 2
- VWHGTYCRDRBSFI-ZETCQYMHSA-N Leu-Gly-Gly Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)NCC(O)=O VWHGTYCRDRBSFI-ZETCQYMHSA-N 0.000 description 2
- VGPCJSXPPOQPBK-YUMQZZPRSA-N Leu-Gly-Ser Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)N[C@@H](CO)C(O)=O VGPCJSXPPOQPBK-YUMQZZPRSA-N 0.000 description 2
- VZBIUJURDLFFOE-IHRRRGAJSA-N Leu-His-Arg Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O VZBIUJURDLFFOE-IHRRRGAJSA-N 0.000 description 2
- XQXGNBFMAXWIGI-MXAVVETBSA-N Leu-His-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)CC(C)C)CC1=CN=CN1 XQXGNBFMAXWIGI-MXAVVETBSA-N 0.000 description 2
- SGIIOQQGLUUMDQ-IHRRRGAJSA-N Leu-His-Val Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](C(C)C)C(=O)O)N SGIIOQQGLUUMDQ-IHRRRGAJSA-N 0.000 description 2
- HGFGEMSVBMCFKK-MNXVOIDGSA-N Leu-Ile-Glu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(O)=O)C(O)=O HGFGEMSVBMCFKK-MNXVOIDGSA-N 0.000 description 2
- NRFGTHFONZYFNY-MGHWNKPDSA-N Leu-Ile-Tyr Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 NRFGTHFONZYFNY-MGHWNKPDSA-N 0.000 description 2
- JKSIBWITFMQTOA-XUXIUFHCSA-N Leu-Ile-Val Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](C(C)C)C(O)=O JKSIBWITFMQTOA-XUXIUFHCSA-N 0.000 description 2
- RXGLHDWAZQECBI-SRVKXCTJSA-N Leu-Leu-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O RXGLHDWAZQECBI-SRVKXCTJSA-N 0.000 description 2
- IEWBEPKLKUXQBU-VOAKCMCISA-N Leu-Leu-Thr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O IEWBEPKLKUXQBU-VOAKCMCISA-N 0.000 description 2
- KPYAOIVPJKPIOU-KKUMJFAQSA-N Leu-Lys-Lys Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(O)=O KPYAOIVPJKPIOU-KKUMJFAQSA-N 0.000 description 2
- VCHVSKNMTXWIIP-SRVKXCTJSA-N Leu-Lys-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(O)=O VCHVSKNMTXWIIP-SRVKXCTJSA-N 0.000 description 2
- DDVHDMSBLRAKNV-IHRRRGAJSA-N Leu-Met-Leu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(C)C)C(O)=O DDVHDMSBLRAKNV-IHRRRGAJSA-N 0.000 description 2
- ZAVCJRJOQKIOJW-KKUMJFAQSA-N Leu-Phe-Asp Chemical compound CC(C)C[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CC(O)=O)C(O)=O)CC1=CC=CC=C1 ZAVCJRJOQKIOJW-KKUMJFAQSA-N 0.000 description 2
- WMIOEVKKYIMVKI-DCAQKATOSA-N Leu-Pro-Ala Chemical compound [H]N[C@@H](CC(C)C)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C)C(O)=O WMIOEVKKYIMVKI-DCAQKATOSA-N 0.000 description 2
- QMKFDEUJGYNFMC-AVGNSLFASA-N Leu-Pro-Arg Chemical compound CC(C)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCCN=C(N)N)C(O)=O QMKFDEUJGYNFMC-AVGNSLFASA-N 0.000 description 2
- DPURXCQCHSQPAN-AVGNSLFASA-N Leu-Pro-Pro Chemical compound CC(C)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 DPURXCQCHSQPAN-AVGNSLFASA-N 0.000 description 2
- IDGZVZJLYFTXSL-DCAQKATOSA-N Leu-Ser-Arg Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCCN=C(N)N IDGZVZJLYFTXSL-DCAQKATOSA-N 0.000 description 2
- IZPVWNSAVUQBGP-CIUDSAMLSA-N Leu-Ser-Asp Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O IZPVWNSAVUQBGP-CIUDSAMLSA-N 0.000 description 2
- SBANPBVRHYIMRR-GARJFASQSA-N Leu-Ser-Pro Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CO)C(=O)N1CCC[C@@H]1C(=O)O)N SBANPBVRHYIMRR-GARJFASQSA-N 0.000 description 2
- SBANPBVRHYIMRR-UHFFFAOYSA-N Leu-Ser-Pro Natural products CC(C)CC(N)C(=O)NC(CO)C(=O)N1CCCC1C(O)=O SBANPBVRHYIMRR-UHFFFAOYSA-N 0.000 description 2
- BRTVHXHCUSXYRI-CIUDSAMLSA-N Leu-Ser-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O BRTVHXHCUSXYRI-CIUDSAMLSA-N 0.000 description 2
- PPGBXYKMUMHFBF-KATARQTJSA-N Leu-Ser-Thr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(O)=O PPGBXYKMUMHFBF-KATARQTJSA-N 0.000 description 2
- ZJZNLRVCZWUONM-JXUBOQSCSA-N Leu-Thr-Ala Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(O)=O ZJZNLRVCZWUONM-JXUBOQSCSA-N 0.000 description 2
- ZDJQVSIPFLMNOX-RHYQMDGZSA-N Leu-Thr-Arg Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@H](C(O)=O)CCCN=C(N)N ZDJQVSIPFLMNOX-RHYQMDGZSA-N 0.000 description 2
- ICYRCNICGBJLGM-HJGDQZAQSA-N Leu-Thr-Asp Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@H](C(O)=O)CC(O)=O ICYRCNICGBJLGM-HJGDQZAQSA-N 0.000 description 2
- URHJPNHRQMQGOZ-RHYQMDGZSA-N Leu-Thr-Met Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCSC)C(O)=O URHJPNHRQMQGOZ-RHYQMDGZSA-N 0.000 description 2
- GZRABTMNWJXFMH-UVOCVTCTSA-N Leu-Thr-Thr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O GZRABTMNWJXFMH-UVOCVTCTSA-N 0.000 description 2
- RNYLNYTYMXACRI-VFAJRCTISA-N Leu-Thr-Trp Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC1=CNC2=C1C=CC=C2)C(O)=O RNYLNYTYMXACRI-VFAJRCTISA-N 0.000 description 2
- WGAZVKFCPHXZLO-SZMVWBNQSA-N Leu-Trp-Glu Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N WGAZVKFCPHXZLO-SZMVWBNQSA-N 0.000 description 2
- FBNPMTNBFFAMMH-AVGNSLFASA-N Leu-Val-Arg Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N FBNPMTNBFFAMMH-AVGNSLFASA-N 0.000 description 2
- ROHFNLRQFUQHCH-UHFFFAOYSA-N Leucine Natural products CC(C)CC(N)C(O)=O ROHFNLRQFUQHCH-UHFFFAOYSA-N 0.000 description 2
- IXHKPDJKKCUKHS-GARJFASQSA-N Lys-Ala-Pro Chemical compound C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCCCN)N IXHKPDJKKCUKHS-GARJFASQSA-N 0.000 description 2
- KNKHAVVBVXKOGX-JXUBOQSCSA-N Lys-Ala-Thr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O KNKHAVVBVXKOGX-JXUBOQSCSA-N 0.000 description 2
- FUKDBQGFSJUXGX-RWMBFGLXSA-N Lys-Arg-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCCCN)N)C(=O)O FUKDBQGFSJUXGX-RWMBFGLXSA-N 0.000 description 2
- NQCJGQHHYZNUDK-DCAQKATOSA-N Lys-Arg-Ser Chemical compound NCCCC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CO)C(O)=O)CCCN=C(N)N NQCJGQHHYZNUDK-DCAQKATOSA-N 0.000 description 2
- SWWCDAGDQHTKIE-RHYQMDGZSA-N Lys-Arg-Thr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)O)C(O)=O SWWCDAGDQHTKIE-RHYQMDGZSA-N 0.000 description 2
- NTSPQIONFJUMJV-AVGNSLFASA-N Lys-Arg-Val Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C(C)C)C(O)=O NTSPQIONFJUMJV-AVGNSLFASA-N 0.000 description 2
- YKIRNDPUWONXQN-GUBZILKMSA-N Lys-Asn-Gln Chemical compound C(CCN)C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N YKIRNDPUWONXQN-GUBZILKMSA-N 0.000 description 2
- FACUGMGEFUEBTI-SRVKXCTJSA-N Lys-Asn-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CCCCN FACUGMGEFUEBTI-SRVKXCTJSA-N 0.000 description 2
- HIIZIQUUHIXUJY-GUBZILKMSA-N Lys-Asp-Gln Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O HIIZIQUUHIXUJY-GUBZILKMSA-N 0.000 description 2
- OVIVOCSURJYCTM-GUBZILKMSA-N Lys-Asp-Glu Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@H](C(O)=O)CCC(O)=O OVIVOCSURJYCTM-GUBZILKMSA-N 0.000 description 2
- IWWMPCPLFXFBAF-SRVKXCTJSA-N Lys-Asp-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O IWWMPCPLFXFBAF-SRVKXCTJSA-N 0.000 description 2
- GJJQCBVRWDGLMQ-GUBZILKMSA-N Lys-Glu-Ala Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O GJJQCBVRWDGLMQ-GUBZILKMSA-N 0.000 description 2
- ZXEUFAVXODIPHC-GUBZILKMSA-N Lys-Glu-Asn Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O ZXEUFAVXODIPHC-GUBZILKMSA-N 0.000 description 2
- GRADYHMSAUIKPS-DCAQKATOSA-N Lys-Glu-Gln Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O GRADYHMSAUIKPS-DCAQKATOSA-N 0.000 description 2
- DUTMKEAPLLUGNO-JYJNAYRXSA-N Lys-Glu-Phe Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O DUTMKEAPLLUGNO-JYJNAYRXSA-N 0.000 description 2
- ITWQLSZTLBKWJM-YUMQZZPRSA-N Lys-Gly-Ala Chemical compound OC(=O)[C@H](C)NC(=O)CNC(=O)[C@@H](N)CCCCN ITWQLSZTLBKWJM-YUMQZZPRSA-N 0.000 description 2
- GQZMPWBZQALKJO-UWVGGRQHSA-N Lys-Gly-Arg Chemical compound [H]N[C@@H](CCCCN)C(=O)NCC(=O)N[C@@H](CCCNC(N)=N)C(O)=O GQZMPWBZQALKJO-UWVGGRQHSA-N 0.000 description 2
- PRSBSVAVOQOAMI-BJDJZHNGSA-N Lys-Ile-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)CC)NC(=O)[C@@H](N)CCCCN PRSBSVAVOQOAMI-BJDJZHNGSA-N 0.000 description 2
- ORVFEGYUJITPGI-IHRRRGAJSA-N Lys-Leu-Met Chemical compound CSCC[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CCCCN ORVFEGYUJITPGI-IHRRRGAJSA-N 0.000 description 2
- LJADEBULDNKJNK-IHRRRGAJSA-N Lys-Leu-Val Chemical compound CC(C)C[C@H](NC(=O)[C@@H](N)CCCCN)C(=O)N[C@@H](C(C)C)C(O)=O LJADEBULDNKJNK-IHRRRGAJSA-N 0.000 description 2
- UQRZFMQQXXJTTF-AVGNSLFASA-N Lys-Lys-Glu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O UQRZFMQQXXJTTF-AVGNSLFASA-N 0.000 description 2
- HVAUKHLDSDDROB-KKUMJFAQSA-N Lys-Lys-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O HVAUKHLDSDDROB-KKUMJFAQSA-N 0.000 description 2
- QQPSCXKFDSORFT-IHRRRGAJSA-N Lys-Lys-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CCCCN QQPSCXKFDSORFT-IHRRRGAJSA-N 0.000 description 2
- WKUXWMWQTOYTFI-SRVKXCTJSA-N Lys-Met-Gln Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CCCCN)N WKUXWMWQTOYTFI-SRVKXCTJSA-N 0.000 description 2
- VSTNAUBHKQPVJX-IHRRRGAJSA-N Lys-Met-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(C)C)C(O)=O VSTNAUBHKQPVJX-IHRRRGAJSA-N 0.000 description 2
- INMBONMDMGPADT-AVGNSLFASA-N Lys-Met-Met Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCSC)C(=O)O)NC(=O)[C@H](CCCCN)N INMBONMDMGPADT-AVGNSLFASA-N 0.000 description 2
- ALEVUGKHINJNIF-QEJZJMRPSA-N Lys-Phe-Ala Chemical compound NCCCC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](C)C(O)=O)CC1=CC=CC=C1 ALEVUGKHINJNIF-QEJZJMRPSA-N 0.000 description 2
- ZJSZPXISKMDJKQ-JYJNAYRXSA-N Lys-Phe-Glu Chemical compound NCCCC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CCC(O)=O)C(O)=O)CC1=CC=CC=C1 ZJSZPXISKMDJKQ-JYJNAYRXSA-N 0.000 description 2
- LUTDBHBIHHREDC-IHRRRGAJSA-N Lys-Pro-Lys Chemical compound NCCCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCCCN)C(O)=O LUTDBHBIHHREDC-IHRRRGAJSA-N 0.000 description 2
- UQJOKDAYFULYIX-AVGNSLFASA-N Lys-Pro-Pro Chemical compound NCCCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 UQJOKDAYFULYIX-AVGNSLFASA-N 0.000 description 2
- WZVSHTFTCYOFPL-GARJFASQSA-N Lys-Ser-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CO)NC(=O)[C@H](CCCCN)N)C(=O)O WZVSHTFTCYOFPL-GARJFASQSA-N 0.000 description 2
- RPWTZTBIFGENIA-VOAKCMCISA-N Lys-Thr-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(O)=O RPWTZTBIFGENIA-VOAKCMCISA-N 0.000 description 2
- XGZDDOKIHSYHTO-SZMVWBNQSA-N Lys-Trp-Glu Chemical compound C1=CC=C2C(C[C@H](NC(=O)[C@@H](N)CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O)=CNC2=C1 XGZDDOKIHSYHTO-SZMVWBNQSA-N 0.000 description 2
- RQILLQOQXLZTCK-KBPBESRZSA-N Lys-Tyr-Gly Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)NCC(O)=O RQILLQOQXLZTCK-KBPBESRZSA-N 0.000 description 2
- FPQMQEOVSKMVMA-ACRUOGEOSA-N Lys-Tyr-Tyr Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CC2=CC=C(C=C2)O)C(=O)O)NC(=O)[C@H](CCCCN)N)O FPQMQEOVSKMVMA-ACRUOGEOSA-N 0.000 description 2
- OHXUUQDOBQKSNB-AVGNSLFASA-N Lys-Val-Arg Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O OHXUUQDOBQKSNB-AVGNSLFASA-N 0.000 description 2
- VKCPHIOZDWUFSW-ONGXEEELSA-N Lys-Val-Gly Chemical compound OC(=O)CNC(=O)[C@H](C(C)C)NC(=O)[C@@H](N)CCCCN VKCPHIOZDWUFSW-ONGXEEELSA-N 0.000 description 2
- IKXQOBUBZSOWDY-AVGNSLFASA-N Lys-Val-Val Chemical compound CC(C)[C@@H](C(=O)N[C@@H](C(C)C)C(=O)O)NC(=O)[C@H](CCCCN)N IKXQOBUBZSOWDY-AVGNSLFASA-N 0.000 description 2
- 241000829100 Macaca mulatta polyomavirus 1 Species 0.000 description 2
- GAELMDJMQDUDLJ-BQBZGAKWSA-N Met-Ala-Gly Chemical compound CSCC[C@H](N)C(=O)N[C@@H](C)C(=O)NCC(O)=O GAELMDJMQDUDLJ-BQBZGAKWSA-N 0.000 description 2
- CWFYZYQMUDWGTI-GUBZILKMSA-N Met-Arg-Asp Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(O)=O CWFYZYQMUDWGTI-GUBZILKMSA-N 0.000 description 2
- DRXODWRPPUFIAY-DCAQKATOSA-N Met-Asn-Lys Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@H](C(O)=O)CCCCN DRXODWRPPUFIAY-DCAQKATOSA-N 0.000 description 2
- UZVWDRPUTHXQAM-FXQIFTODSA-N Met-Asp-Ala Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](C)C(O)=O UZVWDRPUTHXQAM-FXQIFTODSA-N 0.000 description 2
- FWTBMGAKKPSTBT-GUBZILKMSA-N Met-Gln-Glu Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O FWTBMGAKKPSTBT-GUBZILKMSA-N 0.000 description 2
- VOOINLQYUZOREH-SRVKXCTJSA-N Met-Gln-Leu Chemical compound CC(C)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CCSC)N VOOINLQYUZOREH-SRVKXCTJSA-N 0.000 description 2
- QMIXOTQHYHOUJP-KKUMJFAQSA-N Met-Gln-Tyr Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)O)N QMIXOTQHYHOUJP-KKUMJFAQSA-N 0.000 description 2
- KQBJYJXPZBNEIK-DCAQKATOSA-N Met-Glu-Arg Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CCCNC(N)=N KQBJYJXPZBNEIK-DCAQKATOSA-N 0.000 description 2
- DJDFBVNNDAUPRW-GUBZILKMSA-N Met-Glu-Gln Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O DJDFBVNNDAUPRW-GUBZILKMSA-N 0.000 description 2
- SJDQOYTYNGZZJX-SRVKXCTJSA-N Met-Glu-Leu Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O SJDQOYTYNGZZJX-SRVKXCTJSA-N 0.000 description 2
- VZBXCMCHIHEPBL-SRVKXCTJSA-N Met-Glu-Lys Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CCCCN VZBXCMCHIHEPBL-SRVKXCTJSA-N 0.000 description 2
- DGNZGCQSVGGYJS-BQBZGAKWSA-N Met-Gly-Asp Chemical compound CSCC[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CC(O)=O DGNZGCQSVGGYJS-BQBZGAKWSA-N 0.000 description 2
- HSJIGJRZYUADSS-IHRRRGAJSA-N Met-Lys-Leu Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O HSJIGJRZYUADSS-IHRRRGAJSA-N 0.000 description 2
- VSJAPSMRFYUOKS-IUCAKERBSA-N Met-Pro-Gly Chemical compound CSCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O VSJAPSMRFYUOKS-IUCAKERBSA-N 0.000 description 2
- CIDICGYKRUTYLE-FXQIFTODSA-N Met-Ser-Ala Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O CIDICGYKRUTYLE-FXQIFTODSA-N 0.000 description 2
- VOAKKHOIAFKOQZ-JYJNAYRXSA-N Met-Tyr-Arg Chemical compound NC(=N)NCCC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)CCSC)CC1=CC=C(O)C=C1 VOAKKHOIAFKOQZ-JYJNAYRXSA-N 0.000 description 2
- LPNWWHBFXPNHJG-AVGNSLFASA-N Met-Val-Lys Chemical compound CSCC[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@H](C(O)=O)CCCCN LPNWWHBFXPNHJG-AVGNSLFASA-N 0.000 description 2
- 241000699666 Mus <mouse, genus> Species 0.000 description 2
- 108010002311 N-glycylglutamic acid Proteins 0.000 description 2
- 108010066427 N-valyltryptophan Proteins 0.000 description 2
- 239000012124 Opti-MEM Substances 0.000 description 2
- LSXGADJXBDFXQU-DLOVCJGASA-N Phe-Ala-Asp Chemical compound OC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CC1=CC=CC=C1 LSXGADJXBDFXQU-DLOVCJGASA-N 0.000 description 2
- NEHSHYOUIWBYSA-DCPHZVHLSA-N Phe-Ala-Trp Chemical compound C[C@@H](C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)O)NC(=O)[C@H](CC3=CC=CC=C3)N NEHSHYOUIWBYSA-DCPHZVHLSA-N 0.000 description 2
- JIYJYFIXQTYDNF-YDHLFZDLSA-N Phe-Asn-Val Chemical compound CC(C)[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@H](CC1=CC=CC=C1)N JIYJYFIXQTYDNF-YDHLFZDLSA-N 0.000 description 2
- KYYMILWEGJYPQZ-IHRRRGAJSA-N Phe-Glu-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CC1=CC=CC=C1 KYYMILWEGJYPQZ-IHRRRGAJSA-N 0.000 description 2
- APJPXSFJBMMOLW-KBPBESRZSA-N Phe-Gly-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CC1=CC=CC=C1 APJPXSFJBMMOLW-KBPBESRZSA-N 0.000 description 2
- HNFUGJUZJRYUHN-JSGCOSHPSA-N Phe-Gly-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CC1=CC=CC=C1 HNFUGJUZJRYUHN-JSGCOSHPSA-N 0.000 description 2
- VADLTGVIOIOKGM-BZSNNMDCSA-N Phe-His-Leu Chemical compound C([C@@H](C(=O)N[C@@H](CC(C)C)C(O)=O)NC(=O)[C@@H](N)CC=1C=CC=CC=1)C1=CN=CN1 VADLTGVIOIOKGM-BZSNNMDCSA-N 0.000 description 2
- ZUQACJLOHYRVPJ-DKIMLUQUSA-N Phe-Lys-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CC1=CC=CC=C1 ZUQACJLOHYRVPJ-DKIMLUQUSA-N 0.000 description 2
- BELBBZDIHDAJOR-UHFFFAOYSA-N Phenolsulfonephthalein Chemical compound C1=CC(O)=CC=C1C1(C=2C=CC(O)=CC=2)C2=CC=CC=C2S(=O)(=O)O1 BELBBZDIHDAJOR-UHFFFAOYSA-N 0.000 description 2
- XQLBWXHVZVBNJM-FXQIFTODSA-N Pro-Ala-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1 XQLBWXHVZVBNJM-FXQIFTODSA-N 0.000 description 2
- NHDVNAKDACFHPX-GUBZILKMSA-N Pro-Arg-Ala Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C)C(O)=O NHDVNAKDACFHPX-GUBZILKMSA-N 0.000 description 2
- VCYJKOLZYPYGJV-AVGNSLFASA-N Pro-Arg-Leu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(O)=O VCYJKOLZYPYGJV-AVGNSLFASA-N 0.000 description 2
- CJZTUKSFZUSNCC-FXQIFTODSA-N Pro-Asp-Asn Chemical compound NC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H]1CCCN1 CJZTUKSFZUSNCC-FXQIFTODSA-N 0.000 description 2
- VJLJGKQAOQJXJG-CIUDSAMLSA-N Pro-Asp-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O VJLJGKQAOQJXJG-CIUDSAMLSA-N 0.000 description 2
- XKHCJJPNXFBADI-DCAQKATOSA-N Pro-Asp-Lys Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CCCCN)C(=O)O XKHCJJPNXFBADI-DCAQKATOSA-N 0.000 description 2
- CLNJSLSHKJECME-BQBZGAKWSA-N Pro-Gly-Ala Chemical compound OC(=O)[C@H](C)NC(=O)CNC(=O)[C@@H]1CCCN1 CLNJSLSHKJECME-BQBZGAKWSA-N 0.000 description 2
- QEWBZBLXDKIQPS-STQMWFEESA-N Pro-Gly-Tyr Chemical compound [H]N1CCC[C@H]1C(=O)NCC(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O QEWBZBLXDKIQPS-STQMWFEESA-N 0.000 description 2
- LPGSNRSLPHRNBW-AVGNSLFASA-N Pro-His-Val Chemical compound C([C@@H](C(=O)N[C@@H](C(C)C)C([O-])=O)NC(=O)[C@H]1[NH2+]CCC1)C1=CN=CN1 LPGSNRSLPHRNBW-AVGNSLFASA-N 0.000 description 2
- FMLRRBDLBJLJIK-DCAQKATOSA-N Pro-Leu-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H]1CCCN1 FMLRRBDLBJLJIK-DCAQKATOSA-N 0.000 description 2
- VTFXTWDFPTWNJY-RHYQMDGZSA-N Pro-Leu-Thr Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O VTFXTWDFPTWNJY-RHYQMDGZSA-N 0.000 description 2
- SUENWIFTSTWUKD-AVGNSLFASA-N Pro-Leu-Val Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C(C)C)C(O)=O SUENWIFTSTWUKD-AVGNSLFASA-N 0.000 description 2
- IQAGKQWXVHTPOT-FHWLQOOXSA-N Pro-Lys-Trp Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC2=CNC3=CC=CC=C32)C(=O)O IQAGKQWXVHTPOT-FHWLQOOXSA-N 0.000 description 2
- FDMKYQQYJKYCLV-GUBZILKMSA-N Pro-Pro-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@H]1NCCC1 FDMKYQQYJKYCLV-GUBZILKMSA-N 0.000 description 2
- FNGOXVQBBCMFKV-CIUDSAMLSA-N Pro-Ser-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(O)=O FNGOXVQBBCMFKV-CIUDSAMLSA-N 0.000 description 2
- SXJOPONICMGFCR-DCAQKATOSA-N Pro-Ser-Lys Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)O SXJOPONICMGFCR-DCAQKATOSA-N 0.000 description 2
- PKHDJFHFMGQMPS-RCWTZXSCSA-N Pro-Thr-Arg Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O PKHDJFHFMGQMPS-RCWTZXSCSA-N 0.000 description 2
- AIOWVDNPESPXRB-YTWAJWBKSA-N Pro-Thr-Pro Chemical compound C[C@H]([C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@@H]2CCCN2)O AIOWVDNPESPXRB-YTWAJWBKSA-N 0.000 description 2
- FZXSYIPVAFVYBH-KKUMJFAQSA-N Pro-Tyr-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCC(O)=O)C(O)=O FZXSYIPVAFVYBH-KKUMJFAQSA-N 0.000 description 2
- ZMLRZBWCXPQADC-TUAOUCFPSA-N Pro-Val-Pro Chemical compound CC(C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@@H]2CCCN2 ZMLRZBWCXPQADC-TUAOUCFPSA-N 0.000 description 2
- 108090000412 Protein-Tyrosine Kinases Proteins 0.000 description 2
- 102000004022 Protein-Tyrosine Kinases Human genes 0.000 description 2
- ZUGXSSFMTXKHJS-ZLUOBGJFSA-N Ser-Ala-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C)C(=O)N[C@@H](C)C(O)=O ZUGXSSFMTXKHJS-ZLUOBGJFSA-N 0.000 description 2
- MMGJPDWSIOAGTH-ACZMJKKPSA-N Ser-Ala-Gln Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(O)=O MMGJPDWSIOAGTH-ACZMJKKPSA-N 0.000 description 2
- BTKUIVBNGBFTTP-WHFBIAKZSA-N Ser-Ala-Gly Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C)C(=O)NCC(O)=O BTKUIVBNGBFTTP-WHFBIAKZSA-N 0.000 description 2
- DWUIECHTAMYEFL-XVYDVKMFSA-N Ser-Ala-His Chemical compound OC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CC1=CN=CN1 DWUIECHTAMYEFL-XVYDVKMFSA-N 0.000 description 2
- IYCBDVBJWDXQRR-FXQIFTODSA-N Ser-Ala-Met Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C)C(=O)N[C@@H](CCSC)C(O)=O IYCBDVBJWDXQRR-FXQIFTODSA-N 0.000 description 2
- FCRMLGJMPXCAHD-FXQIFTODSA-N Ser-Arg-Asn Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(O)=O FCRMLGJMPXCAHD-FXQIFTODSA-N 0.000 description 2
- QEDMOZUJTGEIBF-FXQIFTODSA-N Ser-Arg-Asp Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(O)=O QEDMOZUJTGEIBF-FXQIFTODSA-N 0.000 description 2
- QPFJSHSJFIYDJZ-GHCJXIJMSA-N Ser-Asp-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H](N)CO QPFJSHSJFIYDJZ-GHCJXIJMSA-N 0.000 description 2
- GHPQVUYZQQGEDA-BIIVOSGPSA-N Ser-Asp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)O)NC(=O)[C@H](CO)N)C(=O)O GHPQVUYZQQGEDA-BIIVOSGPSA-N 0.000 description 2
- SWSRFJZZMNLMLY-ZKWXMUAHSA-N Ser-Asp-Val Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](C(C)C)C(O)=O SWSRFJZZMNLMLY-ZKWXMUAHSA-N 0.000 description 2
- RNMRYWZYFHHOEV-CIUDSAMLSA-N Ser-Gln-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O RNMRYWZYFHHOEV-CIUDSAMLSA-N 0.000 description 2
- OJPHFSOMBZKQKQ-GUBZILKMSA-N Ser-Gln-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(N)=O)NC(=O)[C@@H](N)CO OJPHFSOMBZKQKQ-GUBZILKMSA-N 0.000 description 2
- FMDHKPRACUXATF-ACZMJKKPSA-N Ser-Gln-Ser Chemical compound OC[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(O)=O FMDHKPRACUXATF-ACZMJKKPSA-N 0.000 description 2
- QKQDTEYDEIJPNK-GUBZILKMSA-N Ser-Glu-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CO QKQDTEYDEIJPNK-GUBZILKMSA-N 0.000 description 2
- UFKPDBLKLOBMRH-XHNCKOQMSA-N Ser-Glu-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CO)N)C(=O)O UFKPDBLKLOBMRH-XHNCKOQMSA-N 0.000 description 2
- YMTLKLXDFCSCNX-BYPYZUCNSA-N Ser-Gly-Gly Chemical compound OC[C@H](N)C(=O)NCC(=O)NCC(O)=O YMTLKLXDFCSCNX-BYPYZUCNSA-N 0.000 description 2
- IOVHBRCQOGWAQH-ZKWXMUAHSA-N Ser-Gly-Ile Chemical compound [H]N[C@@H](CO)C(=O)NCC(=O)N[C@@H]([C@@H](C)CC)C(O)=O IOVHBRCQOGWAQH-ZKWXMUAHSA-N 0.000 description 2
- QYSFWUIXDFJUDW-DCAQKATOSA-N Ser-Leu-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O QYSFWUIXDFJUDW-DCAQKATOSA-N 0.000 description 2
- UBRMZSHOOIVJPW-SRVKXCTJSA-N Ser-Leu-Lys Chemical compound OC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(O)=O UBRMZSHOOIVJPW-SRVKXCTJSA-N 0.000 description 2
- VZQRNAYURWAEFE-KKUMJFAQSA-N Ser-Leu-Phe Chemical compound OC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 VZQRNAYURWAEFE-KKUMJFAQSA-N 0.000 description 2
- YUJLIIRMIAGMCQ-CIUDSAMLSA-N Ser-Leu-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O YUJLIIRMIAGMCQ-CIUDSAMLSA-N 0.000 description 2
- RHAPJNVNWDBFQI-BQBZGAKWSA-N Ser-Pro-Gly Chemical compound OC[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O RHAPJNVNWDBFQI-BQBZGAKWSA-N 0.000 description 2
- XGQKSRGHEZNWIS-IHRRRGAJSA-N Ser-Pro-Tyr Chemical compound N[C@@H](CO)C(=O)N1CCC[C@H]1C(=O)N[C@@H](Cc1ccc(O)cc1)C(O)=O XGQKSRGHEZNWIS-IHRRRGAJSA-N 0.000 description 2
- HHJFMHQYEAAOBM-ZLUOBGJFSA-N Ser-Ser-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O HHJFMHQYEAAOBM-ZLUOBGJFSA-N 0.000 description 2
- OLKICIBQRVSQMA-SRVKXCTJSA-N Ser-Ser-Tyr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O OLKICIBQRVSQMA-SRVKXCTJSA-N 0.000 description 2
- FLMYSKVSDVHLEW-SVSWQMSJSA-N Ser-Thr-Ile Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O FLMYSKVSDVHLEW-SVSWQMSJSA-N 0.000 description 2
- ZSDXEKUKQAKZFE-XAVMHZPKSA-N Ser-Thr-Pro Chemical compound C[C@H]([C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CO)N)O ZSDXEKUKQAKZFE-XAVMHZPKSA-N 0.000 description 2
- VEVYMLNYMULSMS-AVGNSLFASA-N Ser-Tyr-Gln Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCC(N)=O)C(O)=O VEVYMLNYMULSMS-AVGNSLFASA-N 0.000 description 2
- HAYADTTXNZFUDM-IHRRRGAJSA-N Ser-Tyr-Val Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](C(C)C)C(O)=O HAYADTTXNZFUDM-IHRRRGAJSA-N 0.000 description 2
- MTCFGRXMJLQNBG-UHFFFAOYSA-N Serine Chemical group OCC(N)C(O)=O MTCFGRXMJLQNBG-UHFFFAOYSA-N 0.000 description 2
- 206010041067 Small cell lung cancer Diseases 0.000 description 2
- FAPWRFPIFSIZLT-UHFFFAOYSA-M Sodium chloride Chemical compound [Na+].[Cl-] FAPWRFPIFSIZLT-UHFFFAOYSA-M 0.000 description 2
- VFEHSAJCWWHDBH-RHYQMDGZSA-N Thr-Arg-Leu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(O)=O VFEHSAJCWWHDBH-RHYQMDGZSA-N 0.000 description 2
- NLSNVZAREYQMGR-HJGDQZAQSA-N Thr-Asp-Leu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O NLSNVZAREYQMGR-HJGDQZAQSA-N 0.000 description 2
- KRPKYGOFYUNIGM-XVSYOHENSA-N Thr-Asp-Phe Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)O)N)O KRPKYGOFYUNIGM-XVSYOHENSA-N 0.000 description 2
- JKGGPMOUIAAJAA-YEPSODPASA-N Thr-Gly-Val Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)NCC(=O)N[C@@H](C(C)C)C(O)=O JKGGPMOUIAAJAA-YEPSODPASA-N 0.000 description 2
- SXAGUVRFGJSFKC-ZEILLAHLSA-N Thr-His-Thr Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H]([C@@H](C)O)C(O)=O SXAGUVRFGJSFKC-ZEILLAHLSA-N 0.000 description 2
- XTCNBOBTROGWMW-RWRJDSDZSA-N Thr-Ile-Glu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)O)NC(=O)[C@H]([C@@H](C)O)N XTCNBOBTROGWMW-RWRJDSDZSA-N 0.000 description 2
- AHOLTQCAVBSUDP-PPCPHDFISA-N Thr-Ile-Lys Chemical compound CC[C@H](C)[C@H](NC(=O)[C@@H](N)[C@@H](C)O)C(=O)N[C@@H](CCCCN)C(O)=O AHOLTQCAVBSUDP-PPCPHDFISA-N 0.000 description 2
- IMDMLDSVUSMAEJ-HJGDQZAQSA-N Thr-Leu-Asn Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O IMDMLDSVUSMAEJ-HJGDQZAQSA-N 0.000 description 2
- RRRRCRYTLZVCEN-HJGDQZAQSA-N Thr-Leu-Asp Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O RRRRCRYTLZVCEN-HJGDQZAQSA-N 0.000 description 2
- JMBRNXUOLJFURW-BEAPCOKYSA-N Thr-Phe-Pro Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N2CCC[C@@H]2C(=O)O)N)O JMBRNXUOLJFURW-BEAPCOKYSA-N 0.000 description 2
- NBIIPOKZPUGATB-BWBBJGPYSA-N Thr-Ser-Cys Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CS)C(=O)O)N)O NBIIPOKZPUGATB-BWBBJGPYSA-N 0.000 description 2
- DOBIBIXIHJKVJF-XKBZYTNZSA-N Thr-Ser-Gln Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(N)=O DOBIBIXIHJKVJF-XKBZYTNZSA-N 0.000 description 2
- MFMGPEKYBXFIRF-SUSMZKCASA-N Thr-Thr-Gln Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(N)=O)C(O)=O MFMGPEKYBXFIRF-SUSMZKCASA-N 0.000 description 2
- COYHRQWNJDJCNA-NUJDXYNKSA-N Thr-Thr-Thr Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O COYHRQWNJDJCNA-NUJDXYNKSA-N 0.000 description 2
- GJOBRAHDRIDAPT-NGTWOADLSA-N Thr-Trp-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CC1=CNC2=CC=CC=C21)NC(=O)[C@H]([C@@H](C)O)N GJOBRAHDRIDAPT-NGTWOADLSA-N 0.000 description 2
- KPMIQCXJDVKWKO-IFFSRLJSSA-N Thr-Val-Glu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O KPMIQCXJDVKWKO-IFFSRLJSSA-N 0.000 description 2
- QNXZCKMXHPULME-ZNSHCXBVSA-N Thr-Val-Pro Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](C(C)C)C(=O)N1CCC[C@@H]1C(=O)O)N)O QNXZCKMXHPULME-ZNSHCXBVSA-N 0.000 description 2
- MNYNCKZAEIAONY-XGEHTFHBSA-N Thr-Val-Ser Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CO)C(O)=O MNYNCKZAEIAONY-XGEHTFHBSA-N 0.000 description 2
- LGEPIBQBGZTBHL-SXNHZJKMSA-N Trp-Gln-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CC1=CNC2=CC=CC=C21)N LGEPIBQBGZTBHL-SXNHZJKMSA-N 0.000 description 2
- YTCNLMSUXPCFBW-SXNHZJKMSA-N Trp-Ile-Glu Chemical compound [H]N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(O)=O)C(O)=O YTCNLMSUXPCFBW-SXNHZJKMSA-N 0.000 description 2
- ULHASJWZGUEUNN-XIRDDKMYSA-N Trp-Lys-Ser Chemical compound [H]N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(O)=O ULHASJWZGUEUNN-XIRDDKMYSA-N 0.000 description 2
- GBEAUNVBIMLWIB-IHPCNDPISA-N Trp-Ser-Phe Chemical compound C([C@H](NC(=O)[C@H](CO)NC(=O)[C@H](CC=1C2=CC=CC=C2NC=1)N)C(O)=O)C1=CC=CC=C1 GBEAUNVBIMLWIB-IHPCNDPISA-N 0.000 description 2
- MXKUGFHWYYKVDV-SZMVWBNQSA-N Trp-Val-Val Chemical compound CC(C)[C@H](NC(=O)[C@@H](NC(=O)[C@@H](N)Cc1c[nH]c2ccccc12)C(C)C)C(O)=O MXKUGFHWYYKVDV-SZMVWBNQSA-N 0.000 description 2
- ZWZOCUWOXSDYFZ-CQDKDKBSSA-N Tyr-Ala-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 ZWZOCUWOXSDYFZ-CQDKDKBSSA-N 0.000 description 2
- HKIUVWMZYFBIHG-KKUMJFAQSA-N Tyr-Arg-Gln Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N)O HKIUVWMZYFBIHG-KKUMJFAQSA-N 0.000 description 2
- GFZQWWDXJVGEMW-ULQDDVLXSA-N Tyr-Arg-Lys Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCCN)C(=O)O)N)O GFZQWWDXJVGEMW-ULQDDVLXSA-N 0.000 description 2
- LMLBOGIOLHZXOT-JYJNAYRXSA-N Tyr-Glu-His Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)N)O LMLBOGIOLHZXOT-JYJNAYRXSA-N 0.000 description 2
- CNLKDWSAORJEMW-KWQFWETISA-N Tyr-Gly-Ala Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)NCC(=O)N[C@@H](C)C(O)=O CNLKDWSAORJEMW-KWQFWETISA-N 0.000 description 2
- OLWFDNLLBWQWCP-STQMWFEESA-N Tyr-Gly-Met Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)NCC(=O)N[C@@H](CCSC)C(O)=O OLWFDNLLBWQWCP-STQMWFEESA-N 0.000 description 2
- AZGZDDNKFFUDEH-QWRGUYRKSA-N Tyr-Gly-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CC1=CC=C(O)C=C1 AZGZDDNKFFUDEH-QWRGUYRKSA-N 0.000 description 2
- BSCBBPKDVOZICB-KKUMJFAQSA-N Tyr-Leu-Asp Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O BSCBBPKDVOZICB-KKUMJFAQSA-N 0.000 description 2
- DWAMXBFJNZIHMC-KBPBESRZSA-N Tyr-Leu-Gly Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(C)C)C(=O)NCC(O)=O DWAMXBFJNZIHMC-KBPBESRZSA-N 0.000 description 2
- XDGPTBVOSHKDFT-KKUMJFAQSA-N Tyr-Met-Glu Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCC(O)=O)C(O)=O XDGPTBVOSHKDFT-KKUMJFAQSA-N 0.000 description 2
- QPOUERMDWKKZEG-HJPIBITLSA-N Tyr-Ser-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 QPOUERMDWKKZEG-HJPIBITLSA-N 0.000 description 2
- GOPQNCQSXBJAII-ULQDDVLXSA-N Tyr-Val-Lys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CC1=CC=C(C=C1)O)N GOPQNCQSXBJAII-ULQDDVLXSA-N 0.000 description 2
- 108091023045 Untranslated Region Proteins 0.000 description 2
- FZSPNKUFROZBSG-ZKWXMUAHSA-N Val-Ala-Asp Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CC(O)=O FZSPNKUFROZBSG-ZKWXMUAHSA-N 0.000 description 2
- IZFVRRYRMQFVGX-NRPADANISA-N Val-Ala-Gln Chemical compound C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](C(C)C)N IZFVRRYRMQFVGX-NRPADANISA-N 0.000 description 2
- JIODCDXKCJRMEH-NHCYSSNCSA-N Val-Arg-Gln Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N JIODCDXKCJRMEH-NHCYSSNCSA-N 0.000 description 2
- XPYNXORPPVTVQK-SRVKXCTJSA-N Val-Arg-Met Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCSC)C(=O)O)N XPYNXORPPVTVQK-SRVKXCTJSA-N 0.000 description 2
- GNWUWQAVVJQREM-NHCYSSNCSA-N Val-Asn-His Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N GNWUWQAVVJQREM-NHCYSSNCSA-N 0.000 description 2
- DBOXBUDEAJVKRE-LSJOCFKGSA-N Val-Asn-Val Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](C(C)C)C(=O)O)N DBOXBUDEAJVKRE-LSJOCFKGSA-N 0.000 description 2
- ISERLACIZUGCDX-ZKWXMUAHSA-N Val-Asp-Ala Chemical compound C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](C(C)C)N ISERLACIZUGCDX-ZKWXMUAHSA-N 0.000 description 2
- VLOYGOZDPGYWFO-LAEOZQHASA-N Val-Asp-Glu Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O VLOYGOZDPGYWFO-LAEOZQHASA-N 0.000 description 2
- BRPKEERLGYNCNC-NHCYSSNCSA-N Val-Glu-Arg Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CCCN=C(N)N BRPKEERLGYNCNC-NHCYSSNCSA-N 0.000 description 2
- MHAHQDBEIDPFQS-NHCYSSNCSA-N Val-Glu-Met Chemical compound CSCC[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)C(C)C MHAHQDBEIDPFQS-NHCYSSNCSA-N 0.000 description 2
- OPGWZDIYEYJVRX-AVGNSLFASA-N Val-His-Arg Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N OPGWZDIYEYJVRX-AVGNSLFASA-N 0.000 description 2
- UMPVMAYCLYMYGA-ONGXEEELSA-N Val-Leu-Gly Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)NCC(O)=O UMPVMAYCLYMYGA-ONGXEEELSA-N 0.000 description 2
- SYSWVVCYSXBVJG-RHYQMDGZSA-N Val-Leu-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CC(C)C)NC(=O)[C@H](C(C)C)N)O SYSWVVCYSXBVJG-RHYQMDGZSA-N 0.000 description 2
- ZRSZTKTVPNSUNA-IHRRRGAJSA-N Val-Lys-Leu Chemical compound CC(C)C[C@H](NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)C(C)C)C(O)=O ZRSZTKTVPNSUNA-IHRRRGAJSA-N 0.000 description 2
- IEBGHUMBJXIXHM-AVGNSLFASA-N Val-Lys-Met Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCSC)C(=O)O)N IEBGHUMBJXIXHM-AVGNSLFASA-N 0.000 description 2
- LJSZPMSUYKKKCP-UBHSHLNASA-N Val-Phe-Ala Chemical compound CC(C)[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](C)C(O)=O)CC1=CC=CC=C1 LJSZPMSUYKKKCP-UBHSHLNASA-N 0.000 description 2
- SJRUJQFQVLMZFW-WPRPVWTQSA-N Val-Pro-Gly Chemical compound CC(C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O SJRUJQFQVLMZFW-WPRPVWTQSA-N 0.000 description 2
- MIKHIIQMRFYVOR-RCWTZXSCSA-N Val-Pro-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@@H]1CCCN1C(=O)[C@H](C(C)C)N)O MIKHIIQMRFYVOR-RCWTZXSCSA-N 0.000 description 2
- UGFMVXRXULGLNO-XPUUQOCRSA-N Val-Ser-Gly Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CO)C(=O)NCC(O)=O UGFMVXRXULGLNO-XPUUQOCRSA-N 0.000 description 2
- SVLAAUGFIHSJPK-JYJNAYRXSA-N Val-Trp-Ser Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)N[C@@H](CO)C(=O)O)N SVLAAUGFIHSJPK-JYJNAYRXSA-N 0.000 description 2
- QPJSIBAOZBVELU-BPNCWPANSA-N Val-Tyr-Ala Chemical compound C[C@@H](C(=O)O)NC(=O)[C@H](CC1=CC=C(C=C1)O)NC(=O)[C@H](C(C)C)N QPJSIBAOZBVELU-BPNCWPANSA-N 0.000 description 2
- PFMSJVIPEZMKSC-DZKIICNBSA-N Val-Tyr-Glu Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N PFMSJVIPEZMKSC-DZKIICNBSA-N 0.000 description 2
- RTJPAGFXOWEBAI-SRVKXCTJSA-N Val-Val-Arg Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N RTJPAGFXOWEBAI-SRVKXCTJSA-N 0.000 description 2
- ZLNYBMWGPOKSLW-LSJOCFKGSA-N Val-Val-Asp Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O ZLNYBMWGPOKSLW-LSJOCFKGSA-N 0.000 description 2
- AOILQMZPNLUXCM-AVGNSLFASA-N Val-Val-Lys Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@H](C(O)=O)CCCCN AOILQMZPNLUXCM-AVGNSLFASA-N 0.000 description 2
- 241000700605 Viruses Species 0.000 description 2
- 235000004279 alanine Nutrition 0.000 description 2
- 125000003295 alanine group Chemical group N[C@@H](C)C(=O)* 0.000 description 2
- 108010011559 alanylphenylalanine Proteins 0.000 description 2
- 229960001611 alectinib Drugs 0.000 description 2
- KDGFLJKFZUIJMX-UHFFFAOYSA-N alectinib Chemical compound CCC1=CC=2C(=O)C(C3=CC=C(C=C3N3)C#N)=C3C(C)(C)C=2C=C1N(CC1)CCC1N1CCOCC1 KDGFLJKFZUIJMX-UHFFFAOYSA-N 0.000 description 2
- 108010040443 aspartyl-aspartic acid Proteins 0.000 description 2
- 108010068265 aspartyltyrosine Proteins 0.000 description 2
- 239000011230 binding agent Substances 0.000 description 2
- 230000000903 blocking effect Effects 0.000 description 2
- 239000008280 blood Substances 0.000 description 2
- 238000010805 cDNA synthesis kit Methods 0.000 description 2
- 239000013592 cell lysate Substances 0.000 description 2
- 238000012054 celltiter-glo Methods 0.000 description 2
- 208000029742 colonic neoplasm Diseases 0.000 description 2
- 239000003086 colorant Substances 0.000 description 2
- 230000007423 decrease Effects 0.000 description 2
- 238000011161 development Methods 0.000 description 2
- 230000001079 digestive effect Effects 0.000 description 2
- 239000012895 dilution Substances 0.000 description 2
- 238000010790 dilution Methods 0.000 description 2
- 239000007884 disintegrant Substances 0.000 description 2
- 239000002552 dosage form Substances 0.000 description 2
- 230000037437 driver mutation Effects 0.000 description 2
- 239000003937 drug carrier Substances 0.000 description 2
- 238000011156 evaluation Methods 0.000 description 2
- 239000012530 fluid Substances 0.000 description 2
- 238000002866 fluorescence resonance energy transfer Methods 0.000 description 2
- 230000003325 follicular Effects 0.000 description 2
- 239000012634 fragment Substances 0.000 description 2
- 230000005861 gene abnormality Effects 0.000 description 2
- 238000007429 general method Methods 0.000 description 2
- 239000008103 glucose Substances 0.000 description 2
- 235000013922 glutamic acid Nutrition 0.000 description 2
- 239000004220 glutamic acid Chemical group 0.000 description 2
- 229960002743 glutamine Drugs 0.000 description 2
- 108010013768 glutamyl-aspartyl-proline Proteins 0.000 description 2
- 108010015792 glycyllysine Proteins 0.000 description 2
- 108010081551 glycylphenylalanine Proteins 0.000 description 2
- 108010087823 glycyltyrosine Proteins 0.000 description 2
- 108010040030 histidinoalanine Proteins 0.000 description 2
- 102000050427 human RET Human genes 0.000 description 2
- GRRNUXAQVGOGFE-NZSRVPFOSA-N hygromycin B Chemical compound O[C@@H]1[C@@H](NC)C[C@@H](N)[C@H](O)[C@H]1O[C@H]1[C@H]2O[C@@]3([C@@H]([C@@H](O)[C@@H](O)[C@@H](C(N)CO)O3)O)O[C@H]2[C@@H](O)[C@@H](CO)O1 GRRNUXAQVGOGFE-NZSRVPFOSA-N 0.000 description 2
- 229940097277 hygromycin b Drugs 0.000 description 2
- 238000011532 immunohistochemical staining Methods 0.000 description 2
- 238000011065 in-situ storage Methods 0.000 description 2
- 239000003112 inhibitor Substances 0.000 description 2
- 230000003834 intracellular effect Effects 0.000 description 2
- 108010044374 isoleucyl-tyrosine Proteins 0.000 description 2
- 239000007951 isotonicity adjuster Substances 0.000 description 2
- 229930027917 kanamycin Natural products 0.000 description 2
- 229960000318 kanamycin Drugs 0.000 description 2
- SBUJHOSQTJFQJX-NOAMYHISSA-N kanamycin Chemical compound O[C@@H]1[C@@H](O)[C@H](O)[C@@H](CN)O[C@@H]1O[C@H]1[C@H](O)[C@@H](O[C@@H]2[C@@H]([C@@H](N)[C@H](O)[C@@H](CO)O2)O)[C@H](N)C[C@@H]1N SBUJHOSQTJFQJX-NOAMYHISSA-N 0.000 description 2
- 229930182823 kanamycin A Natural products 0.000 description 2
- 239000012669 liquid formulation Substances 0.000 description 2
- 239000000314 lubricant Substances 0.000 description 2
- 238000004020 luminiscence type Methods 0.000 description 2
- 108020004999 messenger RNA Proteins 0.000 description 2
- 229930182817 methionine Chemical group 0.000 description 2
- 108010005942 methionylglycine Proteins 0.000 description 2
- 108010085203 methionylmethionine Proteins 0.000 description 2
- 239000007758 minimum essential medium Substances 0.000 description 2
- 102000039446 nucleic acids Human genes 0.000 description 2
- 108020004707 nucleic acids Proteins 0.000 description 2
- 150000007523 nucleic acids Chemical class 0.000 description 2
- 210000000056 organ Anatomy 0.000 description 2
- 229960003531 phenolsulfonphthalein Drugs 0.000 description 2
- 150000003230 pyrimidines Chemical class 0.000 description 2
- 239000011541 reaction mixture Substances 0.000 description 2
- 238000005070 sampling Methods 0.000 description 2
- 235000004400 serine Nutrition 0.000 description 2
- 208000000587 small cell lung carcinoma Diseases 0.000 description 2
- 235000017557 sodium bicarbonate Nutrition 0.000 description 2
- 229910000030 sodium bicarbonate Inorganic materials 0.000 description 2
- 238000002415 sodium dodecyl sulfate polyacrylamide gel electrophoresis Methods 0.000 description 2
- 229940054269 sodium pyruvate Drugs 0.000 description 2
- 239000007787 solid Substances 0.000 description 2
- 238000007920 subcutaneous administration Methods 0.000 description 2
- 229940124597 therapeutic agent Drugs 0.000 description 2
- 239000012096 transfection reagent Substances 0.000 description 2
- 238000012546 transfer Methods 0.000 description 2
- 230000009466 transformation Effects 0.000 description 2
- 108010051110 tyrosyl-lysine Proteins 0.000 description 2
- 108010003137 tyrosyltyrosine Proteins 0.000 description 2
- 239000004474 valine Substances 0.000 description 2
- 239000013603 viral vector Substances 0.000 description 2
- 238000005406 washing Methods 0.000 description 2
- NHBKXEKEPDILRR-UHFFFAOYSA-N 2,3-bis(butanoylsulfanyl)propyl butanoate Chemical compound CCCC(=O)OCC(SC(=O)CCC)CSC(=O)CCC NHBKXEKEPDILRR-UHFFFAOYSA-N 0.000 description 1
- GVRVHSKQODOXTL-UHFFFAOYSA-N 4-amino-N-[4-(methoxymethyl)phenyl]-7-(1-methylcyclopropyl)-6-[2-(1-methylpyrazol-4-yl)ethynyl]pyrrolo[2,3-d]pyrimidine-5-carboxamide Chemical compound NC=1C2=C(N=CN=1)N(C(=C2C(=O)NC1=CC=C(C=C1)COC)C#CC=1C=NN(C=1)C)C1(CC1)C GVRVHSKQODOXTL-UHFFFAOYSA-N 0.000 description 1
- 239000013607 AAV vector Substances 0.000 description 1
- HRPVXLWXLXDGHG-UHFFFAOYSA-N Acrylamide Chemical compound NC(=O)C=C HRPVXLWXLXDGHG-UHFFFAOYSA-N 0.000 description 1
- MBWYUTNBYSSUIQ-HERUPUMHSA-N Ala-Asn-Trp Chemical compound C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)O)N MBWYUTNBYSSUIQ-HERUPUMHSA-N 0.000 description 1
- BEMGNWZECGIJOI-WDSKDSINSA-N Ala-Gly-Glu Chemical compound [H]N[C@@H](C)C(=O)NCC(=O)N[C@@H](CCC(O)=O)C(O)=O BEMGNWZECGIJOI-WDSKDSINSA-N 0.000 description 1
- CYBJZLQSUJEMAS-LFSVMHDDSA-N Ala-Phe-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CC1=CC=CC=C1)NC(=O)[C@H](C)N)O CYBJZLQSUJEMAS-LFSVMHDDSA-N 0.000 description 1
- WNHNMKOFKCHKKD-BFHQHQDPSA-N Ala-Thr-Gly Chemical compound [H]N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(=O)NCC(O)=O WNHNMKOFKCHKKD-BFHQHQDPSA-N 0.000 description 1
- 244000303258 Annona diversifolia Species 0.000 description 1
- 235000002198 Annona diversifolia Nutrition 0.000 description 1
- 108091023037 Aptamer Proteins 0.000 description 1
- GXXWTNKNFFKTJB-NAKRPEOUSA-N Arg-Ile-Ser Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CO)C(O)=O GXXWTNKNFFKTJB-NAKRPEOUSA-N 0.000 description 1
- FMNBYVSGRCXWEK-FOHZUACHSA-N Asn-Thr-Gly Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)NCC(O)=O FMNBYVSGRCXWEK-FOHZUACHSA-N 0.000 description 1
- CBHVAFXKOYAHOY-NHCYSSNCSA-N Asn-Val-Leu Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O CBHVAFXKOYAHOY-NHCYSSNCSA-N 0.000 description 1
- SVABRQFIHCSNCI-FOHZUACHSA-N Asp-Gly-Thr Chemical compound [H]N[C@@H](CC(O)=O)C(=O)NCC(=O)N[C@@H]([C@@H](C)O)C(O)=O SVABRQFIHCSNCI-FOHZUACHSA-N 0.000 description 1
- GBSUGIXJAAKZOW-GMOBBJLQSA-N Asp-Ile-Arg Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O GBSUGIXJAAKZOW-GMOBBJLQSA-N 0.000 description 1
- BWJZSLQJNBSUPM-FXQIFTODSA-N Asp-Pro-Asn Chemical compound OC(=O)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(O)=O BWJZSLQJNBSUPM-FXQIFTODSA-N 0.000 description 1
- DWBZEJHQQIURML-IMJSIDKUSA-N Asp-Ser Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CO)C(O)=O DWBZEJHQQIURML-IMJSIDKUSA-N 0.000 description 1
- JSHWXQIZOCVWIA-ZKWXMUAHSA-N Asp-Ser-Val Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(O)=O JSHWXQIZOCVWIA-ZKWXMUAHSA-N 0.000 description 1
- DCXYFEDJOCDNAF-UHFFFAOYSA-N Asparagine Chemical group OC(=O)C(N)CC(N)=O DCXYFEDJOCDNAF-UHFFFAOYSA-N 0.000 description 1
- MLDQJTXFUGDVEO-UHFFFAOYSA-N BAY-43-9006 Chemical compound C1=NC(C(=O)NC)=CC(OC=2C=CC(NC(=O)NC=3C=C(C(Cl)=CC=3)C(F)(F)F)=CC=2)=C1 MLDQJTXFUGDVEO-UHFFFAOYSA-N 0.000 description 1
- 241000193830 Bacillus <bacterium> Species 0.000 description 1
- 241000894006 Bacteria Species 0.000 description 1
- 206010004593 Bile duct cancer Diseases 0.000 description 1
- 206010005003 Bladder cancer Diseases 0.000 description 1
- 241000255789 Bombyx mori Species 0.000 description 1
- 206010006187 Breast cancer Diseases 0.000 description 1
- 208000026310 Breast neoplasm Diseases 0.000 description 1
- OKTJSMMVPCPJKN-UHFFFAOYSA-N Carbon Chemical compound [C] OKTJSMMVPCPJKN-UHFFFAOYSA-N 0.000 description 1
- 206010008342 Cervix carcinoma Diseases 0.000 description 1
- PXEGEYISOXISDV-XIRDDKMYSA-N Cys-Trp-Lys Chemical compound C1=CC=C2C(C[C@@H](C(=O)N[C@@H](CCCCN)C(O)=O)NC(=O)[C@@H](N)CS)=CNC2=C1 PXEGEYISOXISDV-XIRDDKMYSA-N 0.000 description 1
- 102000004127 Cytokines Human genes 0.000 description 1
- 108090000695 Cytokines Proteins 0.000 description 1
- 238000000018 DNA microarray Methods 0.000 description 1
- 241000702421 Dependoparvovirus Species 0.000 description 1
- 206010061825 Duodenal neoplasm Diseases 0.000 description 1
- 241001269524 Dura Species 0.000 description 1
- 108010012830 Dynactin Complex Proteins 0.000 description 1
- 102000019205 Dynactin Complex Human genes 0.000 description 1
- 241000991587 Enterovirus C Species 0.000 description 1
- 241000620209 Escherichia coli DH5[alpha] Species 0.000 description 1
- 208000000461 Esophageal Neoplasms Diseases 0.000 description 1
- 241000287828 Gallus gallus Species 0.000 description 1
- 206010051066 Gastrointestinal stromal tumour Diseases 0.000 description 1
- VOLVNCMGXWDDQY-LPEHRKFASA-N Gln-Glu-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CCC(=O)N)N)C(=O)O VOLVNCMGXWDDQY-LPEHRKFASA-N 0.000 description 1
- DSRVQBZAMPGEKU-AVGNSLFASA-N Gln-Phe-Cys Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CS)C(=O)O)NC(=O)[C@H](CCC(=O)N)N DSRVQBZAMPGEKU-AVGNSLFASA-N 0.000 description 1
- SYWCGQOIIARSIX-SRVKXCTJSA-N Glu-Pro-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(C)C)C(O)=O SYWCGQOIIARSIX-SRVKXCTJSA-N 0.000 description 1
- HMJULNMJWOZNFI-XHNCKOQMSA-N Glu-Ser-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CO)NC(=O)[C@H](CCC(=O)O)N)C(=O)O HMJULNMJWOZNFI-XHNCKOQMSA-N 0.000 description 1
- MXJYXYDREQWUMS-XKBZYTNZSA-N Glu-Thr-Ser Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(O)=O MXJYXYDREQWUMS-XKBZYTNZSA-N 0.000 description 1
- GZUKEVBTYNNUQF-WDSKDSINSA-N Gly-Ala-Gln Chemical compound NCC(=O)N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(O)=O GZUKEVBTYNNUQF-WDSKDSINSA-N 0.000 description 1
- YMUFWNJHVPQNQD-ZKWXMUAHSA-N Gly-Ala-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)CN YMUFWNJHVPQNQD-ZKWXMUAHSA-N 0.000 description 1
- OMOZPGCHVWOXHN-BQBZGAKWSA-N Gly-Met-Ser Chemical compound CSCC[C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)CN OMOZPGCHVWOXHN-BQBZGAKWSA-N 0.000 description 1
- ZVXMEWXHFBYJPI-LSJOCFKGSA-N Gly-Val-Ile Chemical compound [H]NCC(=O)N[C@@H](C(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O ZVXMEWXHFBYJPI-LSJOCFKGSA-N 0.000 description 1
- 239000004471 Glycine Substances 0.000 description 1
- FLUVGKKRRMLNPU-CQDKDKBSSA-N His-Ala-Phe Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](C)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O FLUVGKKRRMLNPU-CQDKDKBSSA-N 0.000 description 1
- 101000984753 Homo sapiens Serine/threonine-protein kinase B-raf Proteins 0.000 description 1
- SPQWWEZBHXHUJN-KBIXCLLPSA-N Ile-Glu-Ser Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(O)=O SPQWWEZBHXHUJN-KBIXCLLPSA-N 0.000 description 1
- SHVFUCSSACPBTF-VGDYDELISA-N Ile-Ser-His Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N SHVFUCSSACPBTF-VGDYDELISA-N 0.000 description 1
- 108060003951 Immunoglobulin Proteins 0.000 description 1
- 108010054477 Immunoglobulin Fab Fragments Proteins 0.000 description 1
- 102000001706 Immunoglobulin Fab Fragments Human genes 0.000 description 1
- 108010021625 Immunoglobulin Fragments Proteins 0.000 description 1
- 102000008394 Immunoglobulin Fragments Human genes 0.000 description 1
- 102100034343 Integrase Human genes 0.000 description 1
- 108010061833 Integrases Proteins 0.000 description 1
- 208000008839 Kidney Neoplasms Diseases 0.000 description 1
- XUJNEKJLAYXESH-REOHCLBHSA-N L-Cysteine Chemical group SC[C@H](N)C(O)=O XUJNEKJLAYXESH-REOHCLBHSA-N 0.000 description 1
- QNAYBMKLOCPYGJ-REOHCLBHSA-N L-alanine Chemical compound C[C@H](N)C(O)=O QNAYBMKLOCPYGJ-REOHCLBHSA-N 0.000 description 1
- HNDVDQJCIGZPNO-YFKPBYRVSA-N L-histidine Chemical group OC(=O)[C@@H](N)CC1=CN=CN1 HNDVDQJCIGZPNO-YFKPBYRVSA-N 0.000 description 1
- KZSNJWFQEVHDMF-BYPYZUCNSA-N L-valine Chemical compound CC(C)[C@H](N)C(O)=O KZSNJWFQEVHDMF-BYPYZUCNSA-N 0.000 description 1
- 239000002147 L01XE04 - Sunitinib Substances 0.000 description 1
- 239000005511 L01XE05 - Sorafenib Substances 0.000 description 1
- WXZOHBVPVKABQN-DCAQKATOSA-N Leu-Met-Asp Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(=O)O)C(=O)O)N WXZOHBVPVKABQN-DCAQKATOSA-N 0.000 description 1
- 206010025323 Lymphomas Diseases 0.000 description 1
- NDORZBUHCOJQDO-GVXVVHGQSA-N Lys-Gln-Val Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(O)=O NDORZBUHCOJQDO-GVXVVHGQSA-N 0.000 description 1
- ZFNYWKHYUMEZDZ-WDSOQIARSA-N Lys-Trp-Val Chemical compound CC(C)[C@@H](C(=O)O)NC(=O)[C@H](CC1=CNC2=CC=CC=C21)NC(=O)[C@H](CCCCN)N ZFNYWKHYUMEZDZ-WDSOQIARSA-N 0.000 description 1
- MCNGIXXCMJAURZ-VEVYYDQMSA-N Met-Asp-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CCSC)N)O MCNGIXXCMJAURZ-VEVYYDQMSA-N 0.000 description 1
- DBXMFHGGHMXYHY-DCAQKATOSA-N Met-Leu-Ser Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O DBXMFHGGHMXYHY-DCAQKATOSA-N 0.000 description 1
- UNPGTBHYKJOCCZ-DCAQKATOSA-N Met-Lys-Ala Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(O)=O UNPGTBHYKJOCCZ-DCAQKATOSA-N 0.000 description 1
- RDLSEGZJMYGFNS-FXQIFTODSA-N Met-Ser-Asp Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O RDLSEGZJMYGFNS-FXQIFTODSA-N 0.000 description 1
- 241000711408 Murine respirovirus Species 0.000 description 1
- 241000699660 Mus musculus Species 0.000 description 1
- ZOFYNKHSCZTTIE-QGZVFWFLSA-N NC=1C2=C(N=CN=1)N(C(=C2C(=O)NC1=CC=C(C=C1)COC)OC[C@@H]1OCCC1)C1(CC1)C Chemical compound NC=1C2=C(N=CN=1)N(C(=C2C(=O)NC1=CC=C(C=C1)COC)OC[C@@H]1OCCC1)C1(CC1)C ZOFYNKHSCZTTIE-QGZVFWFLSA-N 0.000 description 1
- 238000000636 Northern blotting Methods 0.000 description 1
- 108091028043 Nucleic acid sequence Proteins 0.000 description 1
- 206010030155 Oesophageal carcinoma Diseases 0.000 description 1
- 108700020796 Oncogene Proteins 0.000 description 1
- 206010033128 Ovarian cancer Diseases 0.000 description 1
- 206010061535 Ovarian neoplasm Diseases 0.000 description 1
- 206010061902 Pancreatic neoplasm Diseases 0.000 description 1
- BJEYSVHMGIJORT-NHCYSSNCSA-N Phe-Ala-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](C)NC(=O)[C@@H](N)CC1=CC=CC=C1 BJEYSVHMGIJORT-NHCYSSNCSA-N 0.000 description 1
- OJUMUUXGSXUZJZ-SRVKXCTJSA-N Phe-Asp-Ser Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O OJUMUUXGSXUZJZ-SRVKXCTJSA-N 0.000 description 1
- LTAWNJXSRUCFAN-UNQGMJICSA-N Phe-Thr-Arg Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O LTAWNJXSRUCFAN-UNQGMJICSA-N 0.000 description 1
- 229940122907 Phosphatase inhibitor Drugs 0.000 description 1
- OAICVXFJPJFONN-UHFFFAOYSA-N Phosphorus Chemical compound [P] OAICVXFJPJFONN-UHFFFAOYSA-N 0.000 description 1
- OYEUSRAZOGIDBY-JYJNAYRXSA-N Pro-Arg-Tyr Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O OYEUSRAZOGIDBY-JYJNAYRXSA-N 0.000 description 1
- SMCHPSMKAFIERP-FXQIFTODSA-N Pro-Asn-Asp Chemical compound OC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@@H]1CCCN1 SMCHPSMKAFIERP-FXQIFTODSA-N 0.000 description 1
- RETPETNFPLNLRV-JYJNAYRXSA-N Pro-Asn-Trp Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC2=CNC3=CC=CC=C32)C(=O)O RETPETNFPLNLRV-JYJNAYRXSA-N 0.000 description 1
- JMVQDLDPDBXAAX-YUMQZZPRSA-N Pro-Gly-Gln Chemical compound NC(=O)CC[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H]1CCCN1 JMVQDLDPDBXAAX-YUMQZZPRSA-N 0.000 description 1
- VYWNORHENYEQDW-YUMQZZPRSA-N Pro-Gly-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H]1CCCN1 VYWNORHENYEQDW-YUMQZZPRSA-N 0.000 description 1
- FYPGHGXAOZTOBO-IHRRRGAJSA-N Pro-Leu-His Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@@H]2CCCN2 FYPGHGXAOZTOBO-IHRRRGAJSA-N 0.000 description 1
- IIRBTQHFVNGPMQ-AVGNSLFASA-N Pro-Val-Lys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@@H]1CCCN1 IIRBTQHFVNGPMQ-AVGNSLFASA-N 0.000 description 1
- 206010060862 Prostate cancer Diseases 0.000 description 1
- 208000000236 Prostatic Neoplasms Diseases 0.000 description 1
- 229940124158 Protease/peptidase inhibitor Drugs 0.000 description 1
- 108010029485 Protein Isoforms Proteins 0.000 description 1
- 102000001708 Protein Isoforms Human genes 0.000 description 1
- 108010001648 Proto-Oncogene Proteins c-ret Proteins 0.000 description 1
- 102000000813 Proto-Oncogene Proteins c-ret Human genes 0.000 description 1
- 238000012181 QIAquick gel extraction kit Methods 0.000 description 1
- 238000011530 RNeasy Mini Kit Methods 0.000 description 1
- 208000015634 Rectal Neoplasms Diseases 0.000 description 1
- 206010038389 Renal cancer Diseases 0.000 description 1
- 240000004808 Saccharomyces cerevisiae Species 0.000 description 1
- MOVJSUIKUNCVMG-ZLUOBGJFSA-N Ser-Cys-Ser Chemical compound C([C@@H](C(=O)N[C@@H](CS)C(=O)N[C@@H](CO)C(=O)O)N)O MOVJSUIKUNCVMG-ZLUOBGJFSA-N 0.000 description 1
- FYUIFUJFNCLUIX-XVYDVKMFSA-N Ser-His-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](C)C(O)=O FYUIFUJFNCLUIX-XVYDVKMFSA-N 0.000 description 1
- FUMGHWDRRFCKEP-CIUDSAMLSA-N Ser-Leu-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(O)=O FUMGHWDRRFCKEP-CIUDSAMLSA-N 0.000 description 1
- KCGIREHVWRXNDH-GARJFASQSA-N Ser-Leu-Pro Chemical compound CC(C)C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CO)N KCGIREHVWRXNDH-GARJFASQSA-N 0.000 description 1
- CKDXFSPMIDSMGV-GUBZILKMSA-N Ser-Pro-Val Chemical compound [H]N[C@@H](CO)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C(C)C)C(O)=O CKDXFSPMIDSMGV-GUBZILKMSA-N 0.000 description 1
- ANOQEBQWIAYIMV-AEJSXWLSSA-N Ser-Val-Pro Chemical compound CC(C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CO)N ANOQEBQWIAYIMV-AEJSXWLSSA-N 0.000 description 1
- ODRUTDLAONAVDV-IHRRRGAJSA-N Ser-Val-Tyr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O ODRUTDLAONAVDV-IHRRRGAJSA-N 0.000 description 1
- 102100027103 Serine/threonine-protein kinase B-raf Human genes 0.000 description 1
- 241000710960 Sindbis virus Species 0.000 description 1
- 108010003723 Single-Domain Antibodies Proteins 0.000 description 1
- 108020004682 Single-Stranded DNA Proteins 0.000 description 1
- 208000000453 Skin Neoplasms Diseases 0.000 description 1
- 206010054184 Small intestine carcinoma Diseases 0.000 description 1
- 238000002105 Southern blotting Methods 0.000 description 1
- 208000005718 Stomach Neoplasms Diseases 0.000 description 1
- CAGTXGDOIFXLPC-KZVJFYERSA-N Thr-Arg-Ala Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](C)C(O)=O)CCCN=C(N)N CAGTXGDOIFXLPC-KZVJFYERSA-N 0.000 description 1
- UTSWGQNAQRIHAI-UNQGMJICSA-N Thr-Arg-Phe Chemical compound NC(N)=NCCC[C@H](NC(=O)[C@@H](N)[C@H](O)C)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 UTSWGQNAQRIHAI-UNQGMJICSA-N 0.000 description 1
- PRNGXSILMXSWQQ-OEAJRASXSA-N Thr-Leu-Phe Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O PRNGXSILMXSWQQ-OEAJRASXSA-N 0.000 description 1
- AYFVYJQAPQTCCC-UHFFFAOYSA-N Threonine Natural products CC(O)C(N)C(O)=O AYFVYJQAPQTCCC-UHFFFAOYSA-N 0.000 description 1
- 239000004473 Threonine Substances 0.000 description 1
- 239000007984 Tris EDTA buffer Substances 0.000 description 1
- 239000007983 Tris buffer Substances 0.000 description 1
- 241000209140 Triticum Species 0.000 description 1
- 235000021307 Triticum Nutrition 0.000 description 1
- WCTYCXZYBNKEIV-SXNHZJKMSA-N Trp-Glu-Ile Chemical compound C1=CC=C2C(C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O)=CNC2=C1 WCTYCXZYBNKEIV-SXNHZJKMSA-N 0.000 description 1
- OFTGYORHQMSPAI-PJODQICGSA-N Trp-Met-Ala Chemical compound [H]N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](C)C(O)=O OFTGYORHQMSPAI-PJODQICGSA-N 0.000 description 1
- RQLNEFOBQAVGSY-WDSOQIARSA-N Trp-Met-Leu Chemical compound [H]N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(C)C)C(O)=O RQLNEFOBQAVGSY-WDSOQIARSA-N 0.000 description 1
- WMIUTJPFHMMUGY-ZFWWWQNUSA-N Trp-Pro-Gly Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CC2=CNC3=CC=CC=C32)N)C(=O)NCC(=O)O WMIUTJPFHMMUGY-ZFWWWQNUSA-N 0.000 description 1
- JONPRIHUYSPIMA-UWJYBYFXSA-N Tyr-Ala-Asn Chemical compound NC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 JONPRIHUYSPIMA-UWJYBYFXSA-N 0.000 description 1
- PMDWYLVWHRTJIW-STQMWFEESA-N Tyr-Gly-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CC1=CC=C(O)C=C1 PMDWYLVWHRTJIW-STQMWFEESA-N 0.000 description 1
- PYJKETPLFITNKS-IHRRRGAJSA-N Tyr-Pro-Asn Chemical compound N[C@@H](Cc1ccc(O)cc1)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(O)=O PYJKETPLFITNKS-IHRRRGAJSA-N 0.000 description 1
- 208000007097 Urinary Bladder Neoplasms Diseases 0.000 description 1
- 208000006105 Uterine Cervical Neoplasms Diseases 0.000 description 1
- 208000002495 Uterine Neoplasms Diseases 0.000 description 1
- 241000700618 Vaccinia virus Species 0.000 description 1
- NHXZRXLFOBFMDM-AVGNSLFASA-N Val-Pro-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)C(C)C NHXZRXLFOBFMDM-AVGNSLFASA-N 0.000 description 1
- LCHZBEUVGAVMKS-RHYQMDGZSA-N Val-Thr-Leu Chemical compound CC(C)C[C@H](NC(=O)[C@@H](NC(=O)[C@@H](N)C(C)C)[C@@H](C)O)C(O)=O LCHZBEUVGAVMKS-RHYQMDGZSA-N 0.000 description 1
- 230000005856 abnormality Effects 0.000 description 1
- 230000004913 activation Effects 0.000 description 1
- 239000000654 additive Substances 0.000 description 1
- 210000004102 animal cell Anatomy 0.000 description 1
- 230000003466 anti-cipated effect Effects 0.000 description 1
- 239000003963 antioxidant agent Substances 0.000 description 1
- 239000007864 aqueous solution Substances 0.000 description 1
- 235000009582 asparagine Nutrition 0.000 description 1
- 229960001230 asparagine Drugs 0.000 description 1
- 125000000613 asparagine group Chemical group N[C@@H](CC(N)=O)C(=O)* 0.000 description 1
- 235000003704 aspartic acid Nutrition 0.000 description 1
- CKLJMWTZIZZHCS-REOHCLBHSA-N aspartic acid group Chemical group N[C@@H](CC(=O)O)C(=O)O CKLJMWTZIZZHCS-REOHCLBHSA-N 0.000 description 1
- 210000002469 basement membrane Anatomy 0.000 description 1
- OQFSQFPPLPISGP-UHFFFAOYSA-N beta-carboxyaspartic acid Natural products OC(=O)C(N)C(C(O)=O)C(O)=O OQFSQFPPLPISGP-UHFFFAOYSA-N 0.000 description 1
- 208000026900 bile duct neoplasm Diseases 0.000 description 1
- 201000009036 biliary tract cancer Diseases 0.000 description 1
- 208000020790 biliary tract neoplasm Diseases 0.000 description 1
- 210000004369 blood Anatomy 0.000 description 1
- 210000001124 body fluid Anatomy 0.000 description 1
- 239000010839 body fluid Substances 0.000 description 1
- 239000006172 buffering agent Substances 0.000 description 1
- 239000002775 capsule Substances 0.000 description 1
- 229910052799 carbon Inorganic materials 0.000 description 1
- 239000012876 carrier material Substances 0.000 description 1
- 238000010370 cell cloning Methods 0.000 description 1
- 238000004113 cell culture Methods 0.000 description 1
- 238000005119 centrifugation Methods 0.000 description 1
- 201000010881 cervical cancer Diseases 0.000 description 1
- 210000000038 chest Anatomy 0.000 description 1
- 208000006990 cholangiocarcinoma Diseases 0.000 description 1
- 238000010367 cloning Methods 0.000 description 1
- 239000013599 cloning vector Substances 0.000 description 1
- 239000011248 coating agent Substances 0.000 description 1
- 239000013065 commercial product Substances 0.000 description 1
- 239000012141 concentrate Substances 0.000 description 1
- 235000008504 concentrate Nutrition 0.000 description 1
- 210000004748 cultured cell Anatomy 0.000 description 1
- 238000012258 culturing Methods 0.000 description 1
- XUJNEKJLAYXESH-UHFFFAOYSA-N cysteine Natural products SCC(N)C(O)=O XUJNEKJLAYXESH-UHFFFAOYSA-N 0.000 description 1
- 235000018417 cysteine Nutrition 0.000 description 1
- 238000003745 diagnosis Methods 0.000 description 1
- 238000007847 digital PCR Methods 0.000 description 1
- 239000003085 diluting agent Substances 0.000 description 1
- 238000004090 dissolution Methods 0.000 description 1
- 239000012153 distilled water Substances 0.000 description 1
- 230000000857 drug effect Effects 0.000 description 1
- 241001493065 dsRNA viruses Species 0.000 description 1
- 201000000312 duodenum cancer Diseases 0.000 description 1
- 238000013399 early diagnosis Methods 0.000 description 1
- 238000004520 electroporation Methods 0.000 description 1
- 102000052116 epidermal growth factor receptor activity proteins Human genes 0.000 description 1
- 108700015053 epidermal growth factor receptor activity proteins Proteins 0.000 description 1
- 201000004101 esophageal cancer Diseases 0.000 description 1
- 238000000605 extraction Methods 0.000 description 1
- 230000001605 fetal effect Effects 0.000 description 1
- 210000002950 fibroblast Anatomy 0.000 description 1
- 238000000684 flow cytometry Methods 0.000 description 1
- 239000007850 fluorescent dye Substances 0.000 description 1
- 238000009472 formulation Methods 0.000 description 1
- 239000003205 fragrance Substances 0.000 description 1
- 230000008014 freezing Effects 0.000 description 1
- 238000007710 freezing Methods 0.000 description 1
- 210000000232 gallbladder Anatomy 0.000 description 1
- 206010017758 gastric cancer Diseases 0.000 description 1
- 201000011243 gastrointestinal stromal tumor Diseases 0.000 description 1
- 239000008187 granular material Substances 0.000 description 1
- 229940094991 herring sperm dna Drugs 0.000 description 1
- HNDVDQJCIGZPNO-UHFFFAOYSA-N histidine Chemical group OC(=O)C(N)CC1=CN=CN1 HNDVDQJCIGZPNO-UHFFFAOYSA-N 0.000 description 1
- 235000014304 histidine Nutrition 0.000 description 1
- 102000045233 human DCTN1 Human genes 0.000 description 1
- 210000005260 human cell Anatomy 0.000 description 1
- 238000009396 hybridization Methods 0.000 description 1
- 210000004408 hybridoma Anatomy 0.000 description 1
- 210000000987 immune system Anatomy 0.000 description 1
- 102000018358 immunoglobulin Human genes 0.000 description 1
- 229940072221 immunoglobulins Drugs 0.000 description 1
- 238000003364 immunohistochemistry Methods 0.000 description 1
- 238000001114 immunoprecipitation Methods 0.000 description 1
- 230000001976 improved effect Effects 0.000 description 1
- 230000006872 improvement Effects 0.000 description 1
- 238000000338 in vitro Methods 0.000 description 1
- 208000015181 infectious disease Diseases 0.000 description 1
- 238000010255 intramuscular injection Methods 0.000 description 1
- 238000001990 intravenous administration Methods 0.000 description 1
- 238000010253 intravenous injection Methods 0.000 description 1
- 201000010982 kidney cancer Diseases 0.000 description 1
- 125000001909 leucine group Chemical group [H]N(*)C(C(*)=O)C([H])([H])C(C([H])([H])[H])C([H])([H])[H] 0.000 description 1
- 108010091871 leucylmethionine Proteins 0.000 description 1
- 238000007169 ligase reaction Methods 0.000 description 1
- 210000004185 liver Anatomy 0.000 description 1
- 201000007270 liver cancer Diseases 0.000 description 1
- 208000014018 liver neoplasm Diseases 0.000 description 1
- 239000003589 local anesthetic agent Substances 0.000 description 1
- 238000000504 luminescence detection Methods 0.000 description 1
- 230000001926 lymphatic effect Effects 0.000 description 1
- 208000015486 malignant pancreatic neoplasm Diseases 0.000 description 1
- 239000000463 material Substances 0.000 description 1
- 108010082117 matrigel Proteins 0.000 description 1
- 239000011159 matrix material Substances 0.000 description 1
- 229910052751 metal Inorganic materials 0.000 description 1
- 239000002184 metal Substances 0.000 description 1
- 108091070501 miRNA Proteins 0.000 description 1
- 239000002679 microRNA Substances 0.000 description 1
- 239000011259 mixed solution Substances 0.000 description 1
- 238000010369 molecular cloning Methods 0.000 description 1
- RAHBGWKEPAQNFF-UHFFFAOYSA-N motesanib Chemical compound C=1C=C2C(C)(C)CNC2=CC=1NC(=O)C1=CC=CN=C1NCC1=CC=NC=C1 RAHBGWKEPAQNFF-UHFFFAOYSA-N 0.000 description 1
- 229950003968 motesanib Drugs 0.000 description 1
- 229940124303 multikinase inhibitor Drugs 0.000 description 1
- YOHYSYJDKVYCJI-UHFFFAOYSA-N n-[3-[[6-[3-(trifluoromethyl)anilino]pyrimidin-4-yl]amino]phenyl]cyclopropanecarboxamide Chemical compound FC(F)(F)C1=CC=CC(NC=2N=CN=C(NC=3C=C(NC(=O)C4CC4)C=CC=3)C=2)=C1 YOHYSYJDKVYCJI-UHFFFAOYSA-N 0.000 description 1
- 230000035407 negative regulation of cell proliferation Effects 0.000 description 1
- 230000027405 negative regulation of phosphorylation Effects 0.000 description 1
- 229910052757 nitrogen Inorganic materials 0.000 description 1
- 239000002674 ointment Substances 0.000 description 1
- 201000002528 pancreatic cancer Diseases 0.000 description 1
- 208000008443 pancreatic carcinoma Diseases 0.000 description 1
- 239000012188 paraffin wax Substances 0.000 description 1
- 239000000137 peptide hydrolase inhibitor Substances 0.000 description 1
- 238000002823 phage display Methods 0.000 description 1
- COLNVLDHVKWLRT-UHFFFAOYSA-N phenylalanine Natural products OC(=O)C(N)CC1=CC=CC=C1 COLNVLDHVKWLRT-UHFFFAOYSA-N 0.000 description 1
- COLNVLDHVKWLRT-QMMMGPOBSA-N phenylalanine group Chemical group N[C@@H](CC1=CC=CC=C1)C(=O)O COLNVLDHVKWLRT-QMMMGPOBSA-N 0.000 description 1
- 229910052698 phosphorus Inorganic materials 0.000 description 1
- 239000011574 phosphorus Substances 0.000 description 1
- 238000010837 poor prognosis Methods 0.000 description 1
- 239000000843 powder Substances 0.000 description 1
- 239000003755 preservative agent Substances 0.000 description 1
- 230000002062 proliferating effect Effects 0.000 description 1
- 238000001243 protein synthesis Methods 0.000 description 1
- 210000001938 protoplast Anatomy 0.000 description 1
- QDXGKRWDQCEABB-UHFFFAOYSA-N pyrimidine-5-carboxamide Chemical compound NC(=O)C1=CN=CN=C1 QDXGKRWDQCEABB-UHFFFAOYSA-N 0.000 description 1
- 238000011002 quantification Methods 0.000 description 1
- 239000011535 reaction buffer Substances 0.000 description 1
- 238000003753 real-time PCR Methods 0.000 description 1
- 206010038038 rectal cancer Diseases 0.000 description 1
- 201000001275 rectum cancer Diseases 0.000 description 1
- 108091008146 restriction endonucleases Proteins 0.000 description 1
- 210000001995 reticulocyte Anatomy 0.000 description 1
- 238000003757 reverse transcription PCR Methods 0.000 description 1
- 150000003839 salts Chemical class 0.000 description 1
- 239000012723 sample buffer Substances 0.000 description 1
- 238000000926 separation method Methods 0.000 description 1
- 238000012163 sequencing technique Methods 0.000 description 1
- 230000003584 silencer Effects 0.000 description 1
- 201000000849 skin cancer Diseases 0.000 description 1
- 201000002314 small intestine cancer Diseases 0.000 description 1
- 239000011780 sodium chloride Substances 0.000 description 1
- 239000001509 sodium citrate Substances 0.000 description 1
- NLJMYIDDQXHKNR-UHFFFAOYSA-K sodium citrate Chemical compound O.O.[Na+].[Na+].[Na+].[O-]C(=O)CC(O)(CC([O-])=O)C([O-])=O NLJMYIDDQXHKNR-UHFFFAOYSA-K 0.000 description 1
- 239000002904 solvent Substances 0.000 description 1
- 229960003787 sorafenib Drugs 0.000 description 1
- 201000011549 stomach cancer Diseases 0.000 description 1
- 238000010254 subcutaneous injection Methods 0.000 description 1
- 239000000126 substance Substances 0.000 description 1
- 239000000758 substrate Substances 0.000 description 1
- 229960001796 sunitinib Drugs 0.000 description 1
- WINHZLLDWRZWRT-ATVHPVEESA-N sunitinib Chemical compound CCN(CC)CCNC(=O)C1=C(C)NC(\C=C/2C3=CC(F)=CC=C3NC\2=O)=C1C WINHZLLDWRZWRT-ATVHPVEESA-N 0.000 description 1
- 239000000829 suppository Substances 0.000 description 1
- 230000004083 survival effect Effects 0.000 description 1
- 239000000375 suspending agent Substances 0.000 description 1
- 208000024891 symptom Diseases 0.000 description 1
- 238000003786 synthesis reaction Methods 0.000 description 1
- 230000002194 synthesizing effect Effects 0.000 description 1
- 239000006188 syrup Substances 0.000 description 1
- 235000020357 syrup Nutrition 0.000 description 1
- 239000003826 tablet Substances 0.000 description 1
- 230000001225 therapeutic effect Effects 0.000 description 1
- 125000000341 threoninyl group Chemical group [H]OC([H])(C([H])([H])[H])C([H])(N([H])[H])C(*)=O 0.000 description 1
- 208000013077 thyroid gland carcinoma Diseases 0.000 description 1
- 238000013518 transcription Methods 0.000 description 1
- 230000035897 transcription Effects 0.000 description 1
- 230000014616 translation Effects 0.000 description 1
- LENZDBCJOHFCAS-UHFFFAOYSA-N tris Chemical compound OCC(N)(CO)CO LENZDBCJOHFCAS-UHFFFAOYSA-N 0.000 description 1
- 108010084932 tryptophyl-proline Proteins 0.000 description 1
- 239000000107 tumor biomarker Substances 0.000 description 1
- 210000004881 tumor cell Anatomy 0.000 description 1
- 108010020532 tyrosyl-proline Proteins 0.000 description 1
- 241000701161 unidentified adenovirus Species 0.000 description 1
- 241000701447 unidentified baculovirus Species 0.000 description 1
- 241001529453 unidentified herpesvirus Species 0.000 description 1
- 241001430294 unidentified retrovirus Species 0.000 description 1
- 201000005112 urinary bladder cancer Diseases 0.000 description 1
- 210000002700 urine Anatomy 0.000 description 1
- 206010046766 uterine cancer Diseases 0.000 description 1
- 150000003679 valine derivatives Chemical class 0.000 description 1
- 125000002987 valine group Chemical compound [H]N([H])C([H])(C(*)=O)C([H])(C([H])([H])[H])C([H])([H])[H] 0.000 description 1
- 108010009962 valyltyrosine Proteins 0.000 description 1
- 239000011782 vitamin Substances 0.000 description 1
- 235000013343 vitamin Nutrition 0.000 description 1
- 229940088594 vitamin Drugs 0.000 description 1
- 229930003231 vitamin Natural products 0.000 description 1
- 150000003722 vitamin derivatives Chemical class 0.000 description 1
Images
Classifications
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N33/00—Investigating or analysing materials by specific methods not covered by groups G01N1/00 - G01N31/00
- G01N33/48—Biological material, e.g. blood, urine; Haemocytometers
- G01N33/50—Chemical analysis of biological material, e.g. blood, urine; Testing involving biospecific ligand binding methods; Immunological testing
- G01N33/53—Immunoassay; Biospecific binding assay; Materials therefor
- G01N33/574—Immunoassay; Biospecific binding assay; Materials therefor for cancer
- G01N33/57407—Specifically defined cancers
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/82—Translation products from oncogenes
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/32—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against translation products of oncogenes
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K19/00—Hybrid peptides, i.e. peptides covalently bound to nucleic acids, or non-covalently bound protein-protein complexes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/10—Transferases (2.)
- C12N9/12—Transferases (2.) transferring phosphorus containing groups, e.g. kinases (2.7)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/02—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving viable microorganisms
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6806—Preparing nucleic acids for analysis, e.g. for polymerase chain reaction [PCR] assay
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6813—Hybridisation assays
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N33/00—Investigating or analysing materials by specific methods not covered by groups G01N1/00 - G01N31/00
- G01N33/15—Medicinal preparations ; Physical properties thereof, e.g. dissolubility
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N33/00—Investigating or analysing materials by specific methods not covered by groups G01N1/00 - G01N31/00
- G01N33/48—Biological material, e.g. blood, urine; Haemocytometers
- G01N33/50—Chemical analysis of biological material, e.g. blood, urine; Testing involving biospecific ligand binding methods; Immunological testing
- G01N33/53—Immunoassay; Biospecific binding assay; Materials therefor
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/33—Heterocyclic compounds
- A61K31/395—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins
- A61K31/435—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins having six-membered rings with one nitrogen as the only ring hetero atom
- A61K31/47—Quinolines; Isoquinolines
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/33—Heterocyclic compounds
- A61K31/395—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins
- A61K31/495—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins having six-membered rings with two or more nitrogen atoms as the only ring heteroatoms, e.g. piperazine or tetrazines
- A61K31/505—Pyrimidines; Hydrogenated pyrimidines, e.g. trimethoprim
- A61K31/517—Pyrimidines; Hydrogenated pyrimidines, e.g. trimethoprim ortho- or peri-condensed with carbocyclic ring systems, e.g. quinazoline, perimidine
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/33—Heterocyclic compounds
- A61K31/395—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins
- A61K31/495—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins having six-membered rings with two or more nitrogen atoms as the only ring heteroatoms, e.g. piperazine or tetrazines
- A61K31/505—Pyrimidines; Hydrogenated pyrimidines, e.g. trimethoprim
- A61K31/519—Pyrimidines; Hydrogenated pyrimidines, e.g. trimethoprim ortho- or peri-condensed with heterocyclic rings
- A61K31/52—Purines, e.g. adenine
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/33—Heterocyclic compounds
- A61K31/395—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins
- A61K31/535—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins having six-membered rings with at least one nitrogen and one oxygen as the ring hetero atoms, e.g. 1,2-oxazines
- A61K31/5375—1,4-Oxazines, e.g. morpholine
- A61K31/5377—1,4-Oxazines, e.g. morpholine not condensed and containing further heterocyclic rings, e.g. timolol
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K45/00—Medicinal preparations containing active ingredients not provided for in groups A61K31/00 - A61K41/00
- A61K45/06—Mixtures of active ingredients without chemical characterisation, e.g. antiphlogistics and cardiaca
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/40—Fusion polypeptide containing a tag for immunodetection, or an epitope for immunisation
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
- C12N15/1135—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing against oncogenes or tumor suppressor genes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/14—Type of nucleic acid interfering N.A.
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/158—Expression markers
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2800/00—Detection or diagnosis of diseases
- G01N2800/52—Predicting or monitoring the response to treatment, e.g. for selection of therapy based on assay results in personalised medicine; Prognosis
Landscapes
- Chemical & Material Sciences (AREA)
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Genetics & Genomics (AREA)
- Zoology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Molecular Biology (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Wood Science & Technology (AREA)
- Biophysics (AREA)
- Immunology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biotechnology (AREA)
- Medicinal Chemistry (AREA)
- Analytical Chemistry (AREA)
- General Engineering & Computer Science (AREA)
- Microbiology (AREA)
- Physics & Mathematics (AREA)
- Biomedical Technology (AREA)
- Oncology (AREA)
- Gastroenterology & Hepatology (AREA)
- Pathology (AREA)
- Hematology (AREA)
- Urology & Nephrology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Toxicology (AREA)
- General Physics & Mathematics (AREA)
- Food Science & Technology (AREA)
- Plant Pathology (AREA)
- Pharmacology & Pharmacy (AREA)
- Cell Biology (AREA)
- Hospice & Palliative Care (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- General Chemical & Material Sciences (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
Abstract
DCTN1 단백질의 일부와 RET 단백질의 일부가 융합한 신규 폴리펩티드, 상기 폴리펩티드를 코딩하는 폴리뉴클레오티드, 상기 폴리뉴클레오티드 또는 폴리펩티드의 검출 방법, 상기 폴리뉴클레오티드의 발현 또는 폴리펩티드의 발현 및/또는 활성을 억제하는 화합물을 스크리닝하는 방법, 및 RET를 저해하는 화합물을 유효 성분으로 하는 의약 조성물.
Description
본 발명은 DCTN1 단백질과 RET 단백질과의 융합 단백질인 폴리펩티드, 당해 폴리펩티드를 코딩하는 폴리뉴클레오티드, 당해 폴리펩티드 또는 폴리뉴클레오티드의 검출 방법, 당해 폴리펩티드 또는 폴리뉴클레오티드를 표적으로 한 화합물, 당해 화합물을 스크리닝하는 방법에 관한 것이다.
암은 일본에 있어서의 사인 제1위인 질환이며, 그 치료법의 개선이 요구되고 있다. 갑상선암의 이환자수는 증가하고 있지만, 많은 증례에서 진행이 느려 초기 단계에 적절한 치료를 하는 것으로 높은 생존율을 나타낸다. 반면에 자각 증상이 거의 없기 때문에 적절한 치료에는 조기의 진단이 불가결하다.
갑상선암은 조직형에 따라 유두암, 여포암, 수양암, 미분화암, 악성 림프종으로 나뉜다. 유두암은 갑상선암의 80% 정도를 차지하고 있고, 또한 미분화암은 저빈도이지만 예후가 매우 나쁜 것이 알려져 있다(비특허문헌 1).
유두암에 있어서, 대부분이 암 유전자의 활성화에 의해 발생하는 것이 알려져 있으며 BRAF 변이 유전자(50 내지 60%), RAS 변이 유전자(10 내지 20%), RET 융합 유전자(5 내지 10%) 등의 유전자 이상이 상호 배타적으로 발생하는 것이 명확해지고 있다. 또한 비소세포 폐암에 있어서도, RET 융합 유전자가 1 내지 2%의 빈도로 EGFR 변이 유전자 등의 다른 드라이버 변이 유전자와 상호 배타적으로 존재하는 것이 보고되어 있다(비특허문헌 2 내지 5).
진행한 갑상선암에 있어서는 약물 치료가 주체가 되어, 다양한 멀티키나아제 저해제가 승인되었지만, 드라이버 변이 유전자 특이적으로 효과를 나타내는 약제는 아직 승인되지 않았다. 한편, 폐암에 있어서는 RET 융합 유전자 양성인 환자에서의 RET를 저해함으로써 효과를 나타내는 것이 보고되어 있으며(비특허문헌 6), 갑상선암에 있어서도 변이 유전자나 융합 유전자와 같은 약제 효과의 지표가 될 수 있는 유전자 이상을 동정할 필요가 있다.
암에 있어서, 드라이버가 되는 변이 유전자(변이 단백질)나 융합 유전자(융합 단백질) 등을 동정하는 것은, 암의 성질을 명백히 함과 함께, 이들 변이 유전자나 융합 유전자를 표적으로 한 새로운 암 치료약이나 검사 방법의 개발에 크게 기여하기 때문에, 강하게 요망되고 있다.
그러나, 암 발생의 드라이버가 될 수 있는 변이 유전자나 융합 유전자 등은 충분히 해명되지 않았으며, 약제의 치료 효과에 관련될 수 있는 유전자 이상을 동정하는 것은 매우 의미가 있다.
Cancer, 115(16), pp3801-7(2009)
Oncogene, 22(29), pp4578-80(2003)
Cell, 159(3), pp676-90(2014)
Cancer Discov., 3(6), pp630-5(2013)
Nature, 511(7511), pp543-50(2014)
Lancet Respir Med., 5(1), pp42-50(2017)
본 발명은, RET 단백질의 적어도 일부를 포함하는 융합 단백질인 신규 폴리펩티드, 또는 당해 폴리펩티드를 코딩하는 폴리뉴클레오티드, 당해 폴리펩티드 또는 폴리뉴클레오티드의 검출 방법, 당해 폴리펩티드 또는 폴리뉴클레오티드를 표적으로 한 화합물, 당해 화합물을 스크리닝하는 방법을 제공하는 것을 과제로 한다.
본 발명자들은, 상기 과제를 해결하기 위해 예의 검토한 결과, 갑상선암 환자 유래의 세포에 있어서, DCTN1 단백질의 일부와 RET 단백질의 일부가 융합한 신규 폴리펩티드 및 당해 폴리펩티드를 코딩하는 폴리뉴클레오티드를 동정했다. 또한, 암 세포에 있어서의 본 발명의 폴리뉴클레오티드 또는 폴리펩티드의 검출 방법, 상기 폴리뉴클레오티드의 발현 또는 폴리펩티드의 발현 및/또는 활성을 억제하는 화합물을 스크리닝하는 방법을 발견했다. 다종다양하게 있는 단백질 중, DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분과의 조합을 갖는 융합 단백질이 세포 내에서 자연스럽게 발생하는 것, 또한 DCTN1과 RET와의 융합 유전자가 암 드라이버로서 작용하기 때문에, 상기 융합 단백질이, 암 진단에 유효한 것은 신규의 지견이며 또한 종래 기술로부터 예상할 수 없는 사항이다. 또한 당해 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 암 환자를 치료하기 위한 RET를 저해하는 화합물을 유효 성분으로 하는 의약 조성물을 발견하고, 본 발명을 완성하기에 이르렀다.
즉, 본 발명은, 이하의 양태를 제공하는 것이다.
항 1. DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드.
항 2. 이하의 (a) 내지 (c)에서 선택되는 항 1에 기재된 폴리펩티드.
(a) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열을 포함하는 폴리펩티드.
(b) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실, 또는 부가된 아미노산 서열을 포함하는 폴리펩티드.
(c) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열과 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드.
항 3. 항 1 또는 2에 기재된 폴리펩티드를 코딩하는 폴리뉴클레오티드.
항 4. 이하의 (d) 내지 (f)에서 선택되는 항 3에 기재된 폴리뉴클레오티드.
(d) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(e) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실, 또는 부가된 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(f) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열과 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
항 5. 이하의 (g) 내지 (i)에서 선택되는 항 3에 기재된 폴리뉴클레오티드.
(g) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열을 포함하는 폴리뉴클레오티드.
(h) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열과 상보적인 염기 서열을 포함하는 폴리뉴클레오티드와 엄격한 조건 하에서 하이브리다이징하는 폴리뉴클레오티드.
(i) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열과 90% 이상의 동일성을 갖는 폴리뉴클레오티드.
항 6. 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드를 포함하는 발현 벡터.
항 7. 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드를 도입한 세포.
항 8. 항 1 또는 2에 기재된 폴리펩티드에 특이적으로 결합하는 항체.
항 9. 시료 중 항 1 또는 2에 기재된 폴리펩티드의 존재를 검출하는 방법.
항 10. 시료 중 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하기 위한 프라이머 또는 프로브로서, 당해 프라이머 또는 프로브가 이하의 (j) 내지 (l)에서 선택되는 폴리뉴클레오티드.
(j) DCTN1 단백질을 코딩하는 폴리뉴클레오티드에 하이브리다이징하는 프로브 및 RET 단백질을 코딩하는 폴리뉴클레오티드에 하이브리다이징하는 프로브로 이루어지는 군에서 선택되는 적어도 하나의 프로브인 폴리뉴클레오티드.
(k) DCTN1 단백질을 코딩하는 폴리뉴클레오티드와 RET 단백질을 코딩하는 폴리뉴클레오티드와의 융합점에 하이브리다이징하는 프로브인 폴리뉴클레오티드.
(l) DCTN1 단백질을 코딩하는 폴리뉴클레오티드와 RET 단백질을 코딩하는 폴리뉴클레오티드와의 융합점을 끼워넣도록 설계된 센스 프라이머와 안티센스 프라이머의 세트인 폴리뉴클레오티드.
항 11. 시료 중 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하는 방법.
항 12. 항 9 또는 11에 기재된 검출 방법에 있어서, 시료 중 항 1 또는 2에 기재된 폴리펩티드 또는 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출한 경우에, 시료의 유래가 되는 환자가 암이라고 판정하는 방법.
항 13. RET를 저해하는 화합물을 유효 성분으로 하는, DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 치료용 의약 조성물.
항 14. 이하의 (1) 및 (2)의 공정을 포함하는 항 1 또는 2에 기재된 폴리펩티드의 발현 및/또는 활성, 또는 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 발현을 억제하는 화합물을 스크리닝하는 방법.
(1) 항 1 또는 2에 기재된 폴리펩티드, 항 1 또는 2에 기재된 폴리펩티드 또는 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드를 발현하고 있는 세포, 또는 항 7에 기재된 세포에 시험 화합물을 접촉하는 공정.
(2) 상기 공정 (1)에 있어서, 항 1 또는 2에 기재된 폴리펩티드의 발현 및/또는 활성, 또는 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 발현이 억제되는지 측정하는 공정, 또는 상기 공정 (1)에 기재된 세포의 증식이 억제되는지 측정하는 공정.
항 15. 항 1 또는 2에 기재된 폴리펩티드 또는 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드를, RET를 저해하는 화합물을 사용한 화학 요법이 유효한지 여부의 지표로 하는 방법으로서, 항 9에 기재된 검출 방법에 의해 시료 중에서 항 1 또는 2에 기재된 폴리펩티드를 검출한 경우, 및/또는 항 11에 기재된 검출 방법에 의해 시료 중에서 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출한 경우에, RET를 저해하는 화합물을 사용한 화학 요법이 유효하다고 판정하는 방법.
항 16. DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드 및 당해 폴리펩티드를 코딩하는 폴리뉴클레오티드로 이루어지는 군에서 선택되는 적어도 1종을 포함하는 암을 검출하기 위한 바이오마커.
항 17. DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 환자에게, RET를 저해하는 화합물을 사용한 화학 요법을 행하는 공정을 포함하는, 암의 치료 방법.
항 18. 피험자 유래의 시료 중에서 항 1 또는 2에 기재된 폴리펩티드의 존재를 검출하는 것 및/또는 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하는 것을 행하는 공정, 그리고
항 1 또는 2에 기재된 폴리펩티드의 존재가 검출되고/되거나 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재가 검출된 경우에, 당해 피험자에 대하여, RET를 저해하는 화합물을 사용한 화학 요법을 행하는 공정을 포함하는, 암의 치료 방법.
항 19. DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 환자를 치료하기 위한, RET를 저해하는 화합물.
항 20. DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 환자를 치료하기 위한 암 치료용 의약 조성물을 제조하기 위한, RET를 저해하는 화합물의 사용.
항 21. 시료 중 항 1 또는 2에 기재된 폴리펩티드의 존재를 검출하기 위한 수단 및/또는 시료 중 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하기 위한 수단의, RET를 저해하는 화합물을 사용한 화학 요법이 유효한지 여부의 판정약의 제조 방법.
항 22. 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하기 위한 항DCTN1 항체 및 항RET 항체의 조합.
항 23. 항 1 또는 2에 기재된 폴리펩티드 또는 항 3 내지 5 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하기 위한 검출약을 제조하기 위한, 항 8에 기재된 항체, 항 22에 기재된 항체의 조합 또는 항 10에 기재된 프라이머 또는 프로브의 사용.
본 발명에 따르면, 본 발명의 폴리뉴클레오티드 및/또는 폴리펩티드는 암 세포에서 특이적으로 발현하고 있는 것이 나타났다. 또한, 본 발명의 폴리뉴클레오티드, 폴리펩티드 및 당해 폴리뉴클레오티드 및/또는 폴리펩티드를 발현하고 있는 세포는, 본 발명의 폴리뉴클레오티드의 발현 또는 폴리펩티드의 발현 및/또는 활성을 억제하는 화합물을 스크리닝하는 방법에 사용할 수 있다. 또한, 본 발명의 폴리뉴클레오티드 및/또는 폴리펩티드의 존재를 지표로 함으로써, DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 대상 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성 대상의 검출을 행하는 것이 가능하다. 또한, RET를 저해하는 화합물은, DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 치료용 의약 조성물로서 유용하다.
도 1은 액적 디지털(Droplet Digital) PCR(ddPCR)을 사용한 갑상선암 조직 중의 DCTN1-RET 융합 유전자 및 GAPDH의 발현 확인.
도 2는 액적 디지털 PCR(ddPCR)을 사용한 정상 갑상선 조직 중의 DCTN1-RET 융합 유전자 및 GAPDH의 발현 확인.
도 3은 정상 갑상선 조직 및 갑상선암 조직 중의 DCTN1-RET 융합 유전자 전장의 발현 확인.
도 4는 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서의 DCTN1-RET 융합 단백질의 발현 확인. a) 항 인산화 RET 항체를 사용한 DCTN1-RET 융합 단백질의 검출, b) 항RET 항체를 사용한 DCTN1-RET 융합 단백질의 검출, c) 항DCTN1 항체를 사용한 DCTN1-RET 융합 단백질의 검출.
도 5는 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 3차원 배양에 있어서의 증식 확인. N=3, 평균+SD.
도 6은 생체 내(in vivo)에 있어서의 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 종양 형성성 확인.
도 7은 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서의 RET siRNA에 의한 인산화 RET의 발현 억제의 확인.
도 8은 RET siRNA에 의한 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 증식 억제 효과의 확인.
도 9는 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서의 RET를 저해하는 화합물에 의한 인산화 RET의 발현 억제의 확인.
도 2는 액적 디지털 PCR(ddPCR)을 사용한 정상 갑상선 조직 중의 DCTN1-RET 융합 유전자 및 GAPDH의 발현 확인.
도 3은 정상 갑상선 조직 및 갑상선암 조직 중의 DCTN1-RET 융합 유전자 전장의 발현 확인.
도 4는 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서의 DCTN1-RET 융합 단백질의 발현 확인. a) 항 인산화 RET 항체를 사용한 DCTN1-RET 융합 단백질의 검출, b) 항RET 항체를 사용한 DCTN1-RET 융합 단백질의 검출, c) 항DCTN1 항체를 사용한 DCTN1-RET 융합 단백질의 검출.
도 5는 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 3차원 배양에 있어서의 증식 확인. N=3, 평균+SD.
도 6은 생체 내(in vivo)에 있어서의 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 종양 형성성 확인.
도 7은 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서의 RET siRNA에 의한 인산화 RET의 발현 억제의 확인.
도 8은 RET siRNA에 의한 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 증식 억제 효과의 확인.
도 9는 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서의 RET를 저해하는 화합물에 의한 인산화 RET의 발현 억제의 확인.
본 발명은, 신규 폴리뉴클레오티드 또는 폴리펩티드, 당해 폴리뉴클레오티드 또는 폴리펩티드의 검출 방법, 당해 폴리뉴클레오티드 또는 폴리펩티드를 표적으로 한 화합물, 당해 화합물을 스크리닝하는 방법에 관한 것이다.
본 발명은, DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드(이하, 「본 발명의 폴리펩티드」라고도 칭한다)를 제공한다. 또한, 본 발명은, 당해 폴리펩티드를 코딩하는 폴리뉴클레오티드(이하, 「본 발명의 폴리뉴클레오티드」라고도 칭한다)를 제공한다.
본 발명에 관한 「DCTN1(다이낙틴 서브유닛(Dynactin Subunit) 1) 단백질」은, 150 kDa 디네인-연관 폴리펩티드(dynein-associated polypeptide) 단백질, 또는 DAP-150 단백질이라고도 칭해지는 단백질이며, 인간 또는 비인간 포유 동물의 DCTN1 단백질을 포함하고, 바람직하게는 인간 DCTN1 단백질이다. 인간에 있어서 2p13.1에 좌승해 있는 유전자에 코딩되어 있는 단백질이다. 본 발명에 있어서, 「DCTN1 단백질」은, 그의 스플라이스 변이체인 이소형을 포함하고, 인간 유래의 것이면, 예를 들어 GenPept 액세션 번호 NP_004073, NP_075408, NP_001128512, NP_001128513, NP_001177765, 또는 NP_001177766으로 나타나는 아미노산 서열을 포함하는 폴리펩티드를 들 수 있다. 또한, 보다 구체적으로는, 예를 들어 서열번호 25, 서열번호 26, 서열번호 27, 서열번호 28, 서열번호 29, 또는 서열번호 30으로 나타나는 아미노산 서열을 포함하는 폴리펩티드를 들 수 있다. 또한, 본 발명에 있어서, 「DCTN1 단백질의 N 말단 부분」이란, 상기 DCTN1 단백질의 N 말단측에 있는 코일드-코일 도메인의 일부 또는 전부를 포함하는 폴리펩티드이고, 바람직하게는 DCTN1 단백질의 N 말단측에 있는 코일드-코일 도메인의 전부를 포함하는 폴리펩티드이다.
본 발명에 관한 「RET 단백질」은, Ret 원종양유전자 단백질, RET 수용체 티로신 키나아제(Receptor Tyrosine Kinase) 단백질, 또는 형질감염 중 재배열(Rearranged During Transfection) 단백질이라고도 칭해지는 단백질이며, 인간 또는 비인간 포유 동물의 RET 단백질을 포함하고, 바람직하게는 인간 RET 단백질이다. 인간에 있어서 10q11.2에 좌승해 있는 유전자에 코딩되어 있는 단백질이다. 본 발명에 있어서, 「RET 단백질」은, 그의 스플라이스 변이체인 이소형을 포함하고, 인간 유래의 것이면, 예를 들어 GenPept 액세션 번호 NP_066124, 또는 NP_065681로 나타나는 아미노산 서열을 포함하는 폴리펩티드를 들 수 있다. 또한, 보다 구체적으로는, 예를 들어 서열번호 31, 또는 서열번호 32로 나타나는 아미노산 서열을 포함하는 폴리펩티드를 들 수 있다. 또한, 본 발명에 있어서, 「RET 단백질의 C 말단 부분」이란, 상기 RET 단백질의 C 말단측에 있는 키나아제 도메인을 포함하는 폴리펩티드이다.
또한, 본 발명의 「DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드」로서는, 상기 DCTN1 단백질의 N 말단측에 있는 코일드-코일 도메인의 일부 또는 전부를 포함하는 폴리펩티드와, 상기 RET 단백질의 C 말단측에 있는 키나아제 도메인을 포함하는 폴리펩티드가 융합되어 있는 폴리펩티드이고, 바람직하게는 상기 DCTN1 단백질의 N 말단측에 있는 코일드-코일 도메인의 전부를 포함하는 폴리펩티드와, 상기 RET 단백질의 C 말단측에 있는 키나아제 도메인을 포함하는 폴리펩티드가 융합되어 있는 폴리펩티드이고, 보다 바람직하게는 이하의 (a) 내지 (c)에서 선택되는 폴리펩티드이다. 이들 폴리펩티드는 키나아제 활성 및/또는 세포 증식 효과를 갖는 것이 바람직하다.
(a) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열을 포함하는 폴리펩티드.
(b) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실, 또는 부가된 아미노산 서열을 포함하는 폴리펩티드.
(c) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열과 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드.
보다 바람직하게는 이하의 (a) 내지 (c)에서 선택되는 폴리펩티드이다. 이들 폴리펩티드는 키나아제 활성 또는 세포 증식 효과를 갖는 것이 바람직하다.
(a) 서열번호 18로 나타나는 아미노산 서열을 포함하는 폴리펩티드.
(b) 서열번호 18로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실, 또는 부가된 아미노산 서열을 포함하는 폴리펩티드.
(c) 서열번호 18로 나타나는 아미노산 서열과 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드.
본 발명의 「DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드」에는, 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실 또는 부가된 아미노산 서열을 포함하는 폴리펩티드(상기 (b))가 포함된다. 이러한 아미노산 서열을 포함하는 DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드로서는, 예를 들어 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열을 갖는 DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드의 이소형을 들 수 있다. 이들 폴리펩티드는 키나아제 활성 또는 세포 증식 효과를 갖는 것이 바람직하다. 여기서, 결실, 치환 또는 부가되는 수개의 아미노산이란, 예를 들어 1 내지 10개가 바람직하고, 보다 바람직하게는 1 내지 5개의 아미노산이다. 또한, 상기 부가에는, N 말단 또는 C 말단으로의 1 내지 수개의 아미노산의 부가, 또는 양 말단으로의 1 내지 수개의 아미노산의 부가가 포함된다.
상기 폴리펩티드의 아미노산이 치환된 폴리펩티드로서는, 예를 들어 GenPept 액세션 번호: NP_066124(서열번호 31) 또는 NP_065681(서열번호 32)로 나타나는 아미노산 서열을 갖는 RET 단백질의 게이트키퍼 부위인 804번째(서열번호 2 및 서열번호 4에서는 1325번째, 서열번호 6 및 서열번호 8에서는 1191번째, 서열번호 10 및 서열번호 12에서는 1300번째, 서열번호 14 및 서열번호 16에서는 1186번째, 서열번호 18 및 서열번호 20에서는 1283번째, 서열번호 22 및 서열번호 24에서는 1318번째)의 발린이 류신, 메티오닌, 또는 글루탐산으로 치환된 폴리펩티드, 또는 806번째(서열번호 2 및 서열번호 4에서는 1327번째, 서열번호 6 및 서열번호 8에서는 1193번째, 서열번호 10 및 서열번호 12에서는 1302번째, 서열번호 14 및 서열번호 16에서는 1188번째, 서열번호 18 및 서열번호 20에서는 1285번째, 서열번호 22 및 서열번호 24에서는 1320번째)의 티로신이 시스테인, 글루탐산, 세린, 히스티딘, 또는 아스파라긴으로 치환된 폴리펩티드를 들 수 있다.
또한, 게이트키퍼 부위 이외의 아미노산인 768번째(서열번호 2 및 서열번호 4에서는 1289번째, 서열번호 6 및 서열번호 8에서는 1155번째, 서열번호 10 및 서열번호 12에서는 1264번째, 서열번호 14 및 서열번호 16에서는 1150번째, 서열번호 18 및 서열번호 20에서는 1247번째, 서열번호 22 및 서열번호 24에서는 1282번째)의 글루탐산이 아스파르트산으로 치환된 폴리펩티드, 883번째(서열번호 2 및 서열번호 4에서는 1404번째, 서열번호 6 및 서열번호 8에서는 1270번째, 서열번호 10 및 서열번호 12에서는 1379번째, 서열번호 14 및 서열번호 16에서는 1265번째, 서열번호 18 및 서열번호 20에서는 1362번째, 서열번호 22 및 서열번호 24에서는 1397번째)의 알라닌이 페닐알라닌, 또는 세린으로 치환된 폴리펩티드, 884번째(서열번호 2 및 서열번호 4에서는 1405번째, 서열번호 6 및 서열번호 8에서는 1271번째, 서열번호 10 및 서열번호 12에서는 1380번째, 서열번호 14 및 서열번호 16에서는 1266번째, 서열번호 18 및 서열번호 20에서는 1363번째, 서열번호 22 및 서열번호 24에서는 1398번째)의 글루탐산이 발린으로 치환된 폴리펩티드, 891번째(서열번호 2 및 서열번호 4에서는 1412번째, 서열번호 6 및 서열번호 8에서는 1278번째, 서열번호 10 및 서열번호 12에서는 1387번째, 서열번호 14 및 서열번호 16에서는 1273번째, 서열번호 18 및 서열번호 20에서는 1370번째, 서열번호 22 및 서열번호 24에서는 1405번째)의 세린이 알라닌, 또는 류신으로 치환된 폴리펩티드, 또는 918번째(서열번호 2 및 서열번호 4에서는 1439번째, 서열번호 6 및 서열번호 8에서는 1305번째, 서열번호 10 및 서열번호 12에서는 1414번째, 서열번호 14 및 서열번호 16에서는 1300번째, 서열번호 18 및 서열번호 20에서는 1397번째, 서열번호 22 및 서열번호 24에서는 1432번째)의 메티오닌이 트레오닌으로 치환된 폴리펩티드를 들 수 있지만, 이들에 한정되지 않는다.
또한, 본 발명의 DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드에는, 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열에 있어서 상당하는 서열을 적절하게 얼라인먼트했을 때, 상기 서열번호 중 어느 것에서 나타나는 아미노산 서열의 1개와 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드(상기 (c))가 포함된다. 이들 폴리펩티드는 키나아제 활성 또는 세포 증식 효과를 갖는 것이 바람직하다.
여기서, 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열과의 동일성은, 바람직하게는 90% 이상이고, 보다 바람직하게는 95% 이상이고, 더욱 바람직하게는 98% 이상이다. 당해 아미노산 서열의 동일성은, 통상 관용의 방법으로 계산할 수 있다.
본 발명의 폴리펩티드는, 본 발명에 관한 폴리펩티드를 구성하는 아미노산 서열 이외에, 단백질 태그를 구성하는 아미노산을 갖고 있어도 된다. 태그의 예로서는, 발현 효율을 향상시키는 태그나, 정제 효율을 향상시키는 태그 등, 당업자에게 주지인 태그를 사용할 수 있고, His 태그, Myc 태그, FLAG 태그 등을 들 수 있다.
본 발명의 폴리뉴클레오티드는, 상기 DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드를 코딩하는 폴리뉴클레오티드이고, 바람직하게는 이하의 (d) 내지 (i)에서 선택되는 폴리뉴클레오티드이다. 이들 폴리뉴클레오티드는, 키나아제 활성 또는 세포 증식 효과를 갖는 폴리펩티드를 코딩하는 폴리뉴클레오티드인 것이 바람직하다.
(d) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(e) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실, 또는 부가된 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(f) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열과 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(g) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열을 포함하는 폴리뉴클레오티드.
(h) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열과 상보적인 염기 서열을 포함하는 폴리뉴클레오티드와 엄격한 조건 하에서 하이브리다이징하는 폴리뉴클레오티드.
(i) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열과 90% 이상의 동일성을 갖는 폴리뉴클레오티드.
보다 바람직하게는 이하의 (d) 내지 (i)에서 선택되는 폴리뉴클레오티드이다. 이들 폴리뉴클레오티드는, 키나아제 활성 또는 세포 증식 효과를 갖는 폴리펩티드를 코딩하는 폴리뉴클레오티드인 것이 바람직하다.
(d) 서열번호 18로 나타나는 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(e) 서열번호 18로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실, 또는 부가된 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(f) 서열번호 18로 나타나는 아미노산 서열과 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(g) 서열번호 17로 나타나는 염기 서열을 포함하는 폴리뉴클레오티드.
(h) 서열번호 17로 나타나는 염기 서열과 상보적인 염기 서열을 포함하는 폴리뉴클레오티드와 엄격한 조건 하에서 하이브리다이징하는 폴리뉴클레오티드.
(i) 서열번호 17로 나타나는 염기 서열과 90% 이상의 동일성을 갖는 폴리뉴클레오티드.
본 발명의 폴리뉴클레오티드는, 이중쇄 DNA 뿐만 아니라, 그것을 구성하는 센스쇄 및 안티센스쇄와 같은 각종 단일쇄 DNA나 RNA도 포함한다. 안티센스쇄는, 프로브 등으로서 이용 가능하다. DNA에는, 예를 들어 클로닝이나 화학 합성 기술 또는 그들의 조합으로 얻어지는 cDNA나 게놈 DNA가 포함된다. 또한, 본 발명에 관한 폴리뉴클레오티드에 대하여, 본 발명에 관한 폴리펩티드를 코딩하는 염기 서열 이외에, 비번역 영역(UTR)의 서열 등의 염기 서열이 부가되어 있어도 된다.
여기서, 엄격한 조건이란, 예를 들어 Molecular Cloning: A Laboratory Manual(Second Edition, J.Sambrook et.al, 1989)에 기재된 조건을 들 수 있다. 즉, 6×SSC(1×SSC의 조성: 0.15M 염화나트륨, 0.015M 시트르산나트륨, pH7.0), 0.5% SDS, 5×덴하르트 및 100㎎/mL 청어 정자 DNA를 포함하는 용액에 프로브와 함께 65℃에서 8 내지 16시간 항온하고, 하이브리다이징시키는 조건 등을 들 수 있다.
또한, 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열과의 동일성은, 바람직하게는 90% 이상, 보다 바람직하게는 95% 이상, 더욱 바람직하게는 98% 이상이다. 당해 염기 서열의 동일성은, 통상 관용의 방법으로 계산할 수 있다.
본 명세서 중에 있어서, 「키나아제 활성 또는 세포 증식 효과를 갖는다」의 「키나아제 활성을 갖는다」란, 티로신을 인산화하는 효소로서의 활성을 갖는 것을 의미한다. 또한, 「키나아제 활성 또는 세포 증식 효과를 갖는다」의 「세포 증식 효과를 갖는다」란, 본 발명의 폴리뉴클레오티드 및/또는 폴리펩티드를 세포에 도입함으로써, 도입하지 않은 세포에 비하여, 도입한 세포의 증식능이 향상되는 효과이다. 이러한 효과는, 예를 들어 사이토카인 의존적으로 증식하는 세포주에 폴리뉴클레오티드 및/또는 폴리펩티드를 도입했을 때, 사이토카인 비의존적으로 증식했을 때, 세포 증식 효과를 갖는다고 확인할 수 있다.
본 발명의 폴리뉴클레오티드는, 예를 들어 DCTN1 유전자와 RET 유전자와의 융합 유전자를 유지하는 갑상선암 등으로부터 조정한 cDNA 라이브러리나 게놈 DNA 라이브러리를 사용하여, 본 발명의 폴리뉴클레오티드의 염기 서열의 일부와 특이적으로 하이브리다이징하는 프라이머를 사용하여, 추출할 수 있다. 이러한 프라이머로서는, 본 발명에 관한 폴리뉴클레오티드 또는 그의 안티센스쇄의 적어도 일부에 특이적으로 하이브리다이징하는 프라이머이면, 어떠한 서열 및 길이의 것을 사용해도 된다. 또한 인공적으로 폴리뉴클레오티드를 합성하는 방법을 들 수 있다(Nat. Methods, 11: 499-507, 2014).
본 발명의 발현 벡터는, 본 발명의 폴리뉴클레오티드를 포함하고, 본 발명의 폴리펩티드를 발현시키는 한, 특별히 한정되는 것은 아니다. 예를 들어, 사용하는 숙주에 따라서 적절히 선택한 공지된 발현 벡터에, 본 발명의 폴리뉴클레오티드를 삽입함으로써 얻어지는 발현 벡터를 들 수 있다.
숙주로서는, 형질 전환이 가능한 생세포이면 특별히 한정되지 않고, 예를 들어, 대장균, 고초균 등의 세균, 효모나 사상균 등의 진균류, Sf9 세포 등의 곤충 세포, 누에 등의 곤충, 동물 세포, 식물 또는 식물 유래 세포를 들 수 있다.
본 발명의 폴리뉴클레오티드를 삽입하기 위한 벡터는, 숙주 안에서 복제 가능한 것이면 특별히 한정되지 않고, 도입하는 숙주의 종류, 도입 방법 등에 따라서 적절히 선택할 수 있다. 예를 들어, 플라스미드 DNA, 파지 DNA, 바이러스 벡터를 들 수 있다. 발현 벡터의 구축에 사용되는 벡터 DNA는, 널리 보급되어 입수가 용이한 것이 사용된다. 예를 들어, pUC19(타카라 바이오), pTV118N(타카라 바이오), pMAMneo(클론 테크), pGEX(GE 헬스케어), pET160(Invitrogen), pDEST(Invitrogen), pIEx(머크 밀리포아), pBacPAK(클론 테크)를 들 수 있다. 또한, 바이러스 벡터로서는, 예를 들어 바큘로바이러스 벡터, 레트로바이러스 벡터, 인간 면역 부전증 바이러스(HIV) 등의 렌티바이러스 벡터, 아데노바이러스 벡터, 아데노 수반 바이러스 벡터(AAV 벡터), 헤르페스 바이러스, 백시니아 바이러스, 폭스 바이러스, 폴리오 바이러스, 신드비스 바이러스, 센다이 바이러스, 시미안 바이러스-40(SV-40) 등의 DNA 바이러스나 RNA 바이러스를 들 수 있다.
당해 발현 벡터를 사용해서 숙주를 형질 전환하기 위해서는, 프로토플라스트 법, 적격 세포법, 일렉트로포레이션법 등을 사용해서 행할 수 있다. 얻어진 형질 전환체는, 자화할 수 있는 탄소원, 질소원, 금속염, 비타민 등을 포함하는 배지를 사용해서 적당한 조건 하에서 배양하면 된다.
본 발명의 폴리뉴클레오티드를 도입한 세포는, 예를 들어 상기한 본 발명의 발현 벡터로 형질 전환된 세포, 게놈 편집에 의해 본 발명의 폴리뉴클레오티드를 도입한 세포를 들 수 있다. 여기서 사용할 수 있는 세포로서는, 상기한 숙주 세포를 들 수 있다. 세포가, 발현 벡터로 형질 전환된 세포인지 확인하는 방법으로서는, 예를 들어, 본 발명의 폴리펩티드를 검출하기 위한 방법, 또는 본 발명의 폴리뉴클레오티드를 검출하기 위한 방법을 들 수 있다.
「게놈 편집에 의해 본 발명의 폴리뉴클레오티드를 도입한 세포」로서, 바람직하게는 각각 단독으로 존재하는 DCTN1 유전자와 RET 유전자를 게놈 편집에 의해 융합시킨 유전자를 갖는 세포이고, 보다 바람직하게는 각각 단독으로 존재하는 DCTN1 유전자와 RET 유전자의 DCTN1의 엑손 27과 RET의 엑손 12를 게놈 편집에 의해 융합시킨 유전자를 갖는 세포이다. 이러한 세포는 통상 관용의 방법으로 제작 가능하며, 예를 들어 Cell Rep., 9(4), pp1219-1227(2014), Nat.Commun., 5, 3728(2014)에 기재된 방법을 들 수 있다. 세포가, 게놈 편집에 의해 본 발명의 폴리뉴클레오티드를 도입한 세포인지 확인하는 방법으로서는, 예를 들어, 본 발명의 폴리펩티드를 검출하기 위한 방법, 또는 본 발명의 폴리뉴클레오티드를 검출하기 위한 방법을 들 수 있다.
본 발명의 폴리펩티드는, 본 발명의 발현 벡터로 형질 전환된 세포를, 세포 배양에 적합한 배지를 사용하여, 적당한 조건 하에서 배양함으로써 얻어진 배양액 및/또는 세포로부터, 일반적인 방법에 의해 단백질의 채취, 정제를 행함으로써, 얻을 수 있다. 또한, 본 발명의 폴리펩티드는, 본 발명의 폴리뉴클레오티드를 갖는 발현 벡터, 또는 본 발명의 폴리뉴클레오티드를 코딩하는 주형 RNA 또는 주형 DNA를 무세포 단백질 합성계(예를 들어, 인간 세포주 유래의 세포 추출액, 토끼 망상적혈구 추출액, 소맥 배아 추출액, 대장균 추출액)에 도입하고, 적당한 조건 하에서 인큐베이션함으로써 얻어진 반응액으로부터, 일반적인 방법에 의해 단백질의 채취, 정제를 행함으로써, 얻을 수 있다.
본 발명에 있어서, 본 발명의 폴리펩티드에 특이적으로 결합하는 항체는, DCTN1 단백질의 N 말단 부분과 RET 단백질의 C 말단 부분의 융합점에 특이적으로 결합하는 항체를 들 수 있다. 당해 항체는, DCTN1 단백질의 N 말단 부분과 RET 단백질의 C 말단 부분의 융합점에 특이적으로 결합하지만, 야생형의 DCTN1, 또는 야생형의 RET 단백질 중 어디에도 결합하지 않는 항체를 의미한다.
본 발명에 있어서, 「DCTN1 단백질의 N 말단 부분과 RET 단백질의 C 말단 부분의 융합점」에 있어서의 「융합점」이란, DCTN1 단백질의 N 말단 부분 유래의 폴리펩티드와 RET 단백질의 C 말단 부분 유래의 폴리펩티드가 융합한 점을 의미한다. 서열번호 2에 있어서의 융합점은, 서열번호 2에 있어서의 1-1233번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 2에 있어서의 1234-1635번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 4에 있어서의 융합점은, 서열번호 4에 있어서의 1-1233번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 4에 있어서의 1234-1593번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 6에 있어서의 융합점은, 서열번호 6에 있어서의 1-1099번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 6에 있어서의 1100-1501번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 8에 있어서의 융합점은, 서열번호 8에 있어서의 1-1099번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 8에 있어서의 1100-1459번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 10에 있어서의 융합점은, 서열번호 10에 있어서의 1-1208번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 10에 있어서의 1209-1610번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 12에 있어서의 융합점은, 서열번호 12에 있어서의 1-1208번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 12에 있어서의 1209-1568번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 14에 있어서의 융합점은, 서열번호 14에 있어서의 1-1094번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 14에 있어서의 1095-1496번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 16에 있어서의 융합점은, 서열번호 16에 있어서의 1-1094번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 16에 있어서의 1095-1454번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 18에 있어서의 융합점은, 서열번호 18에 있어서의 1-1191번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 18에 있어서의 1192-1593번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 20에 있어서의 융합점은, 서열번호 20에 있어서의 1-1191번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 20에 있어서의 1192-1551번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 22에 있어서의 융합점은, 서열번호 22에 있어서의 1-1226번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 22에 있어서의 1227-1628번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다. 서열번호 24에 있어서의 융합점은, 서열번호 24에 있어서의 1-1226번째의 DCTN1의 N 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드와 서열번호 24에 있어서의 1227-1586번째의 RET의 C 말단 부분 유래의 아미노산 서열을 갖는 폴리펩티드가 융합한 점이다.
상기 항체는, 예를 들어 면역 글로불린(IgA, IgD, IgE, IgG, IgM, IgY 등), Fab 프래그먼트, F(ab')2 프래그먼트, 단일쇄 항체 프래그먼트(scFv), 싱글 도메인 항체, 디아바디(Diabody) 등(Nat.Rev.I㎜unol., 6:343-357, 2006)을 들 수 있고, 이들은 인간 항체, 인간화 항체, 키메라 항체, 마우스 항체, 라마 항체, 닭 항체 등의 모노클로날 항체 또는 폴리클로날 항체를 들 수 있지만 이들에 한정되는 것은 아니다.
상기 항체는, 다양한 공지된 방법을 사용해서 제작할 수 있고, 제작 방법은 특별히 한정되는 것이 아니다. 이러한 공지된 방법으로서는, 당해 발명의 폴리펩티드, DCTN1 단백질의 N 말단 부분과 RET 단백질의 C 말단 부분의 융합점을 포함하는 폴리펩티드 단편 등을 면역 동물에 접종하고, 당해 동물의 면역계를 활성화시킨 후, 당해 동물의 혈청을 회수하고, 폴리클로날 항체로서 얻는 방법, 또는 하이브리도마법, 파지 디스플레이법 등에 의해 모노클로날 항체를 얻는 방법 등을 들 수 있다.
본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제하는 화합물을 스크리닝하는 방법은, 이하의 (1) 및 (2)의 공정을 포함하는 방법에 의해 행할 수 있다.
즉, 본 발명의 스크리닝 방법은,
(1) 본 발명의 폴리펩티드, 또는 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포에 시험 화합물을 접촉하는 공정.
(2) 상기 공정 (1)에 있어서, 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현이 억제되는지 측정하는 공정, 또는 상기 공정 (1)에 기재된 세포의 증식이 억제되는지 측정하는 공정.
을 포함하는 방법에 의해 행할 수 있다.
보다 바람직하게는, 이하의 (1) 및 (2)의 공정을 포함하는 방법이다.
(1) 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포에 시험 화합물을 접촉하는 공정.
(2) 상기 공정 (1)에 기재된 세포의 증식이 억제되는지 측정하는 공정.
또한, 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제하는 화합물을 스크리닝하는 방법은, 이하의 (1) 내지 (3)의 공정을 포함하는 방법에 의해 행할 수 있다.
즉, 본 발명의 스크리닝 방법은,
(1) 본 발명의 폴리펩티드, 또는 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포에 시험 화합물을 접촉하는 공정.
(2) 상기 공정 (1)에 있어서, 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현이 억제되는지 측정하는 공정, 또는 상기 공정 (1)에 기재된 세포의 증식이 억제되는지 측정하는 공정.
(3) 상기 공정 (2)에 있어서, 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현이 억제된 경우, 또는 상기 공정 (1)에 기재된 세포의 증식이 억제된 경우, 시험 화합물이 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제한다고 판정하는 공정.
을 포함하는 방법에 의해 행할 수 있다.
보다 바람직하게는, 이하의 (1) 내지 (3)의 공정을 포함하는 방법이다.
(1) 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포에 시험 화합물을 접촉하는 공정.
(2) 상기 공정 (1)에 기재된 세포의 증식이 억제되는지 측정하는 공정.
(3) 상기 공정 (2)에 있어서, 상기 공정 (1)에 기재된 세포의 증식이 억제된 경우, 시험 화합물이 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제한다고 판정하는 공정.
「본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포」는, 본 발명의 발현 벡터로 형질 전환된 세포, 게놈 편집에 의해 본 발명의 폴리뉴클레오티드를 도입한 세포, 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 초대 배양 세포, 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 주화 세포, 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 암 환자 유래의 세포 등을 들 수 있다. 세포가, 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포인지 확인하는 방법으로서는, 예를 들어, 본 발명의 폴리펩티드를 검출하기 위한 방법, 또는 본 발명의 폴리뉴클레오티드를 검출하기 위한 방법을 들 수 있다.
본 발명에 있어서, 「본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현이 억제된다」 중에서, 「본 발명의 폴리펩티드의 발현, 또는 본 발명의 폴리뉴클레오티드의 발현이 억제된다」란, 예를 들어 본 발명의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포에 시험 화합물을 접촉시킨 후, 당해 세포에 있어서의 본 발명의 폴리펩티드 또는 폴리뉴클레오티드의 발현량을, 본 발명의 폴리펩티드 또는 폴리뉴클레오티드의 존재를 검출하는 방법을 사용해서 평가했을 때, 시험 화합물을 접촉시키지 않은 세포와 비교하여, 시험 화합물을 접촉시킨 세포에 있어서, 본 발명의 폴리펩티드 또는 폴리뉴클레오티드의 발현량이 통계학상 유의미하게 저하되었을 때, 본 발명의 폴리펩티드 또는 폴리뉴클레오티드의 발현이 억제되었다고 판단할 수 있다.
또한, 「본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현이 억제된다」 중에서, 「본 발명의 폴리펩티드의 활성이 억제된다」란, 예를 들어 본 발명의 폴리펩티드, 또는 본 발명의 폴리펩티드를 발현하고 있는 세포를 사용하여, 시험 화합물을 접촉시키지 않은 경우와 비교하여, 시험 화합물을 접촉시킨 경우에, 티로신의 인산화의 비율이 통계학상 유의미하게 저하되었을 때, 본 발명의 폴리펩티드 활성이 억제되었다고 판단할 수 있다.
또한, 본 발명의 폴리펩티드를 발현하고 있는 세포를 사용하여, 시험 화합물을 접촉시키지 않은 경우와 비교하여, 시험 화합물을 접촉시킨 경우에, 세포 증식이 통계학상 유의미하게 억제되었을 때, 본 발명의 폴리펩티드의 활성이 억제되었다고 판단할 수 있다.
본 발명에 있어서, 「티로신의 인산화」란, RET 단백질(다른 단백질과 융합되어 있는 RET 단백질도 포함한다)의 티로신의 인산화뿐만 아니라, RET 하류 시그널 상의 단백질의 티로신의 인산화도 포함된다. RET 하류 시그널 상의 단백질로서는, 예를 들어 STAT, AKT, ERK를 들 수 있다. 바람직하게는, RET 단백질(다른 단백질과 융합되어 있는 RET 단백질도 포함한다)의 티로신의 인산화이다.
또한, 「티로신의 인산화의 비율」은, 예를 들어 인산화 RET 특이적 항체를 사용하여, 웨스턴 블로팅, 면역 침강, 면역 조직 화학, ELISA, 플로우 사이토메트리에 의해 측정 가능하다.
본 발명에 있어서 「시료」란, 생체 시료(예를 들어, 세포, 조직, 장기, 체액(혈액, 림프액 등), 소화액, 오줌)뿐만 아니라, 이들 생체 시료로부터 얻어지는 핵산 추출물(게놈 DNA 추출물, mRNA 추출물, mRNA 추출물로부터 제조된 cDNA 제조물이나 cRNA 제조물 등)이나 단백질 추출물도 포함한다. 또한, 상기 시료는, 포르말린 고정 처리, 알코올 고정 처리, 동결 처리 또는 파라핀 포매 처리가 실시되어 있는 것이어도 된다. 상기 생체 시료로서는, 생체로부터 채취한 것을 사용할 수 있다. 바람직하게는 암 환자 유래의 시료이고, 보다 바람직하게는 종양 세포를 포함하는 시료이다. 또한, 생체 시료의 채취 방법은, 생체 시료의 종류에 따라 적절히 선택할 수 있다.
본 발명에는, 시료 중 본 발명의 폴리펩티드의 존재를 검출하는 방법이 포함된다.
본 발명에 있어서, 시료 중 본 발명의 폴리펩티드의 존재를 검출하는 방법은, 본 발명의 폴리펩티드에 특이적으로 결합하는 항체를 사용한 ELISA법, 웨스턴 블로팅법, 또는 면역 조직 화학 염색법, 또는 DCTN1 단백질에 특이적으로 결합하는 항체 및 RET 단백질에 특이적으로 결합하는 항체를 사용한 FRET(Fluorescence Resonance Energy Transfer; 형광 공명 에너지 전달)법 등, 통상 관용의 검출법에 의해 검출하는 방법을 들 수 있다. 바람직하게는, 본 발명의 폴리펩티드에 특이적으로 결합하는 항체를 사용한 ELISA법, 웨스턴 블로팅법, 또는 면역 조직 화학 염색법이다.
DCTN1 단백질에 특이적으로 결합하는 항체 및 RET 단백질에 특이적으로 결합하는 항체로서는, DCTN1 단백질의 상기 융합점으로부터 N 말단 부분에 결합하는 항체 및 RET 단백질의 상기 융합점으로부터 C 말단 부분에 결합하는 항체가 바람직하고, 이들 항체는, 시판품을 사용하는 것, 또는 통상 공지된 방법으로 제작하는 것이 가능하다.
본 발명에 있어서, 시료 중 본 발명의 폴리펩티드의 존재를 검출하는 방법으로서, 바람직하게는 본 발명의 폴리펩티드에 특이적으로 결합하는 항체 또는 DCTN1 단백질에 특이적으로 결합하는 항체 및 RET 단백질에 특이적으로 결합하는 항체를 사용해서 본 발명의 폴리펩티드를 검출하는 공정을 포함하는 것을 특징으로 하는, 본 발명의 폴리펩티드의 존재를 검출하는 방법이고, 보다 바람직하게는 본 발명의 폴리펩티드에 특이적으로 결합하는 항체를 사용해서 본 발명의 폴리펩티드를 검출하는 공정을 포함하는 것을 특징으로 하는, 본 발명의 폴리펩티드의 존재를 검출하는 방법이다. 따라서, 본 발명의 폴리펩티드의 존재를 검출하기 위한 수단으로서는, 특별히 한정되지 않지만, 예를 들어 DCTN1 단백질에 특이적으로 결합하는 항체 및 RET 단백질에 특이적으로 결합하는 항체의 조합; 본 발명의 폴리펩티드에 특이적으로 결합하는 항체 등을 들 수 있다.
본 발명에는, 시료 중 본 발명의 폴리뉴클레오티드의 존재를 검출하기 위한 프라이머 또는 프로브가 포함된다. 본 발명에 있어서, 본 발명의 폴리펩티드의 존재를 검출하기 위한 수단으로서는, 특별히 한정되지 않지만, 예를 들어 본 발명의 폴리뉴클레오티드의 존재를 검출하기 위한 프라이머 또는 프로브 등을 들 수 있다.
당해 프라이머 또는 프로브로서는, (j) 내지 (l)에서 선택되는 폴리뉴클레오티드;
(j) DCTN1 단백질을 코딩하는 폴리뉴클레오티드에 하이브리다이징하는 프로브 및 RET 단백질을 코딩하는 폴리뉴클레오티드에 하이브리다이징하는 프로브로 이루어지는 군에서 선택되는 적어도 하나의 프로브인 폴리뉴클레오티드.
(k) DCTN1 단백질을 코딩하는 폴리뉴클레오티드와 RET 단백질을 코딩하는 폴리뉴클레오티드와의 융합점에 하이브리다이징하는 프로브인 폴리뉴클레오티드.
(l) DCTN1 단백질을 코딩하는 폴리뉴클레오티드와 RET 단백질을 코딩하는 폴리뉴클레오티드와의 융합점을 끼워넣도록 설계된 센스 프라이머와 안티센스 프라이머의 세트인 폴리뉴클레오티드.
를 들 수 있다.
본 발명에 있어서, 「DCTN1 단백질을 코딩하는 폴리뉴클레오티드와 RET 단백질을 코딩하는 폴리뉴클레오티드와의 융합점」에 있어서의 「융합점」이란, DCTN1 단백질을 코딩하는 폴리뉴클레오티드와 RET 단백질을 코딩하는 폴리뉴클레오티드가 융합한 점을 의미한다. 서열번호 1에 있어서의 융합점은, 서열번호 1에 있어서의 1-3699번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 1에 있어서의 3700-4905번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 3에 있어서의 융합점은, 서열번호 3에 있어서의 1-3699번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 3에 있어서의 3700-4779번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 5에 있어서의 융합점은, 서열번호 5에 있어서의 1-3297번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 5에 있어서의 3298-4503번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 7에 있어서의 융합점은, 서열번호 7에 있어서의 1-3297번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 7에 있어서의 3298-4377번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 9에 있어서의 융합점은, 서열번호 9에 있어서의 1-3624번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 9에 있어서의 3625-4830번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 11에 있어서의 융합점은, 서열번호 11에 있어서의 1-3624번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 11에 있어서의 3625-4704번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 13에 있어서의 융합점은, 서열번호 13에 있어서의 1-3282번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 13에 있어서의 3283-4488번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 15에 있어서의 융합점은, 서열번호 15에 있어서의 1-3282번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 15에 있어서의 3283-4362번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 17에 있어서의 융합점은, 서열번호 17에 있어서의 1-3573번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 17에 있어서의 3574-4779번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 19에 있어서의 융합점은, 서열번호 19에 있어서의 1-3573번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 19에 있어서의 3574-4653번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 21에 있어서의 융합점은, 서열번호 21에 있어서의 1-3678번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 21에 있어서의 3679-4884번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다. 서열번호 23에 있어서의 융합점은, 서열번호 23에 있어서의 1-3678번째의 DCTN1을 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드와 서열번호 23에 있어서의 3679-4758번째의 RET를 코딩하는 폴리뉴클레오티드 유래의 염기 서열을 갖는 폴리뉴클레오티드가 융합한 점이다.
본 발명에 있어서, 프라이머 또는 프로브는, 본 발명의 폴리뉴클레오티드 서열 정보에 기초하여, 본 발명의 폴리뉴클레오티드와 특이적으로 하이브리다이징하는 폴리뉴클레오티드로서, 통상 공지된 방법에 의해 제작된다. 당해 프라이머 또는 프로브의 염기수는, 10 내지 50 염기, 바람직하게는 15 내지 50 염기, 보다 바람직하게는 18 내지 35 염기이다.
당해 프라이머 또는 프로브는, 본 발명의 폴리뉴클레오티드와 특이적으로 하이브리다이징하는 것이면, 완전히 상보적일 필요는 없다. 이러한 프라이머 또는 프로브는, 대응하는 염기 서열과 비교하여, 70% 이상, 바람직하게는 80% 이상, 보다 바람직하게는 90% 이상, 보다 바람직하게는 95% 이상, 보다 바람직하게는 98% 이상의 동일성을 갖는 폴리뉴클레오티드이다.
본 발명의 프라이머 또는 프로브로서는, 바람직하게는 (i) 서열번호 69, (ii) 서열번호 70, 또는 (iii) 서열번호 71로 표현되는 폴리뉴클레오티드이고, 보다 바람직하게는 (iv) 서열번호 69와 서열번호 70으로 표현되는 센스 프라이머와 안티센스 프라이머의 세트인 폴리뉴클레오티드이고, 보다 바람직하게는 (v) 서열번호 69, 서열번호 70 및 서열번호 71로 표현되는 센스 프라이머, 안티센스 프라이머 및 프로브의 세트인 폴리뉴클레오티드이다.
본 발명에는, 시료 중 본 발명의 폴리뉴클레오티드의 존재를 검출하는 방법이 포함된다.
본 발명에 있어서, 시료 중 본 발명의 폴리뉴클레오티드의 존재를 검출하는 방법은, 노던 블로팅법, 서던 블로팅법, RT-PCR법, 리얼타임 PCR법, 디지털 PCR법, DNA 마이크로어레이법, 계내(in situ) 하이브리다이제이션법, 시퀀스 해석법 등, 통상 관용의 검출법에 의해 검출하는 방법이다.
본 발명에 있어서, 시료 중 본 발명의 폴리뉴클레오티드의 존재를 검출하는 방법에는, 본 발명의 폴리뉴클레오티드를 포함하는 RET 융합 유전자의 폴리뉴클레오티드의 존재를 검출하기 위한 방법도 포함된다. 당해 방법으로서는, RET 단백질을 코딩하는 폴리뉴클레오티드에 하이브리다이징하는 프라이머(예를 들어, RET 키나아제 도메인 이후의 3'측의 서열에 하이브리다이징하는 프라이머)를 사용하여, 5'RACE법에 의해 증폭시킨 PCR 산물을 시퀀스 해석하는 방법 등을 들 수 있다.
본 발명에 있어서, 시료 중 본 발명의 폴리뉴클레오티드의 존재를 검출하는 방법으로서, 바람직하게는 본 발명의 프라이머 또는 프로브를 사용해서 본 발명의 폴리뉴클레오티드를 검출하는 공정을 포함하는 것을 특징으로 하는, 본 발명의 폴리뉴클레오티드의 존재를 검출하는 방법이다.
본 발명에는, RET를 저해하는 화합물을 유효 성분으로 하는, DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 치료용 의약 조성물이 포함된다.
보다 바람직하게는, 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제하는 화합물을 유효 성분으로 하는, DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 치료용 의약 조성물이 포함된다.
본 발명에 있어서, 「DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암」에 있어서의 「DCTN1 유전자와 RET 유전자와의 융합 유전자 양성인 암」이란, 본 발명의 폴리뉴클레오티드가 발현하고 있는 암이고, 바람직하게는 본 발명의 폴리뉴클레오티드의 존재를 검출하는 방법을 사용하여, 본 발명의 폴리뉴클레오티드가 검출된 암이다.
또한, 본 발명에 있어서, 「DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암」에 있어서의 「DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암」이란, 본 발명의 폴리펩티드가 발현하고 있는 암이고, 바람직하게는 본 발명의 폴리펩티드의 존재를 검출하는 방법을 사용하여, 본 발명의 폴리펩티드가 검출된 암이다.
본 발명의 암 치료용 의약 조성물에 있어서의 유효 성분으로서는, RET를 저해하는 화합물이고, 보다 바람직하게는 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제하는 화합물이다. 또한, 본 발명의 스크리닝 방법에 의해 선택한 화합물을 유효 성분으로서 사용할 수 있다. 예를 들어, 공지된 RET를 저해하는 화합물을 본 발명의 의약 조성물에 있어서의 유효 성분으로서 사용할 수 있다. RET를 저해하는 화합물로서는, RET의 발현 및/또는 활성을 저해할 수 있는 화합물이면, 다른 티로신 키나아제의 발현 및/또는 활성을 저해하는 화합물이어도 되고, 보다 바람직하게는 RET의 활성을 저해할 수 있는 화합물이며, 다른 티로신 키나아제의 발현 및/또는 활성을 저해하는 화합물이어도 된다. 이러한 화합물로서, 예를 들어, 반데타닙, 소라페닙, 수니티닙, 모테사닙, 카보잔티닙, 렌바티닙, 국제공개 제2016/127074호 팸플릿, 국제공개 제2017/043550호 팸플릿, 국제공개 제2017/011776호 팸플릿, 국제공개 제2017/146116호 팸플릿에 기재된 화합물을 들 수 있다.
DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 치료용 의약 조성물의 유효 성분으로서는, RET를 저해하는 화합물이고, 보다 바람직하게는 반데타닙, 카보잔티닙, 렌바티닙, 국제공개 제2017/043550호 팸플릿에 기재된 일반식 (1)로 나타나는 축합 피리미딘 화합물 또는 국제공개 제2017/146116호 팸플릿에 기재된 일반식 (1)로 나타나는 축합 피리미딘 화합물이고, 보다 바람직하게는, 반데타닙, 카보잔티닙, 렌바티닙, 국제공개 제2017/043550호 팸플릿에 기재된 실시예 화합물 1 내지 90 또는 국제공개 제2017/146116호 팸플릿에 기재된 실시예 화합물 1 내지 207이고, 더욱 바람직하게는 반데타닙, 카보잔티닙, 렌바티닙, 4-아미노-1-(tert-부틸)-N-(5-메틸-1H-피라졸-3-일)-1H-피라졸로[3,4-d]피리미딘-3-카르복사미드, 4-아미노-7-(tert-부틸)-N-(5-메틸-1H-피라졸-3-일)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-7-(1-플루오로-2-메틸프로판-2-일)-N-(5-메틸-1H-피라졸-3-일)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-N-(5-메틸-1H-피라졸-3-일)-7-(1-메틸시클로프로필)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-7-(2-시클로프로필프로판-2-일)-N-(5-메틸-1H-피라졸-3-일)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-N-[4-(메톡시메틸)페닐]-7-(1-메틸시클로프로필)-6-(3-모르폴리노프로-1-핀-1-일)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-N-[4-(메톡시메틸)페닐]-7-(1-메틸시클로프로필)-6-((테트라히드로-2H-피란-4-일)에티닐)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, (R)-4-아미노-N-[4-(메톡시메틸)페닐]-7-(1-메틸시클로프로필)-6-((테트라히드로푸란-2-일)메톡시)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드 또는 4-아미노-N-[4-(메톡시메틸)페닐]-6-((1-메틸-1H-피라졸-4-일)에티닐)-7-(1-메틸시클로프로필)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드이며, 특히 바람직하게는 4-아미노-1-(tert-부틸)-N-(5-메틸-1H-피라졸-3-일)-1H-피라졸로[3,4-d]피리미딘-3-카르복사미드, 4-아미노-7-(tert-부틸)-N-(5-메틸-1H-피라졸-3-일)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-7-(1-플루오로-2-메틸프로판-2-일)-N-(5-메틸-1H-피라졸-3-일)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-N-(5-메틸-1H-피라졸-3-일)-7-(1-메틸시클로프로필)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-7-(2-시클로프로필프로판-2-일)-N-(5-메틸-1H-피라졸-3-일)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-N-[4-(메톡시메틸)페닐]-7-(1-메틸시클로프로필)-6-(3-모르폴리노프로-1-핀-1-일)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, 4-아미노-N-[4-(메톡시메틸)페닐]-7-(1-메틸시클로프로필)-6-((테트라히드로-2H-피란-4-일)에티닐)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드, (R)-4-아미노-N-[4-(메톡시메틸)페닐]-7-(1-메틸시클로프로필)-6-((테트라히드로푸란-2-일)메톡시)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드 또는 4-아미노-N-[4-(메톡시메틸)페닐]-6-((1-메틸-1H-피라졸-4-일)에티닐)-7-(1-메틸시클로프로필)-7H-피롤로[2,3-d]피리미딘-5-카르복사미드이다.
본 발명에 있어서, 「RET의 발현 및/또는 활성을 저해할 수 있는 화합물」 중에서, 「RET의 발현을 저해할 수 있다」란, 예를 들어 RET의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포에 시험 화합물을 접촉시킨 후, 당해 세포에 있어서의 RET의 폴리펩티드 또는 폴리뉴클레오티드의 발현량을 검출했을 때, 시험 화합물을 접촉시키지 않은 세포와 비교하여, 시험 화합물을 접촉시킨 세포에 있어서, RET의 폴리펩티드 또는 폴리뉴클레오티드의 발현량이 저하되었을 때, RET의 발현이 억제되었다고 판단할 수 있다. 이러한 화합물로서는, 상기와 같은 화합물, siRNA, miRNA, 핵산(DNA, RNA) 앱타머 등을 들 수 있다. siRNA로서는, 예를 들어 CACAUGUCAUCAAAUUGUATT(서열번호 74), GGAUUGAAAACAAACUCUATT(서열번호 75), GCUUGUCCCGAGAUGUUUATT(서열번호 76)을 들 수 있으며, 바람직하게는CACAUGUCAUCAAAUUGUATT(서열번호 74) 또는 GGAUUGAAAACAAACUCUATT(서열번호 75)이다.
또한, 「RET의 발현 및/또는 활성을 저해할 수 있는 화합물」 중에서, 「RET의 활성을 저해할 수 있다」란, 예를 들어 티로신의 인산화를 지표로 판단할 수 있다. 티로신의 인산화를 측정하는 방법으로서는, 예를 들어, 국제공개 제2017/043550호 팸플릿의 시험예 1에 기재된 방법을 들 수 있다.
또한, RET의 폴리펩티드 및/또는 폴리뉴클레오티드를 발현하고 있는 세포를 사용하여, 세포 증식 억제 효과를 지표로 하여, 「RET의 활성을 저해할 수 있다」라고 판단할 수 있다. 세포 증식 억제 효과는, 예를 들어 국제공개 제2017/043550호 팸플릿의 시험예 3 및 시험예 4에 기재된 방법을 들 수 있다.
본 발명의 의약 조성물의 대상이 되는 암은, 본 발명의 폴리뉴클레오티드 및/또는 폴리펩티드를 발현하고 있는 한 특별히 제한되지 않지만, 예를 들어 두경부암, 갑상선암, 소화기암(식도암, 위암, 십이지장암, 간암, 담도암(담낭·담관암 등), 췌장암, 소장암, 대장암(결장직장암, 결장암, 직장암 등), 소화관 간질 종양 등), 폐암(비소세포 폐암, 소세포 폐암), 유방암, 난소암, 자궁암(자궁경부암, 자궁체암 등), 신장암, 방광암, 전립선암, 피부암 등을 들 수 있고, 바람직하게는 갑상선암, 또는 폐암(비소세포 폐암, 소세포 폐암)이다. 부언하면, 여기에서 암에는, 원발소 뿐만 아니라, 다른 장기(간장 등)로 전이한 암도 포함한다.
본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제하는 화합물을 유효 성분으로 하는 제제는, 약학적 담체를 배합하여, 다양한 투여 형태에 따른 의약 조성물로서 제조할 수 있다. 당해 형태로서는, 예를 들어 경구제, 주사제, 좌제, 연고제, 첩부제를 들 수 있다. 이들 투여 형태는, 각각 당업자에게 공지 관용인 제제 방법에 의해 제조할 수 있다.
약학적 담체로서는, 제제 소재로서 관용인 각종 유기 또는 무기 담체 물질이 사용되고, 고형 제제에 있어서의 부형제, 결합제, 붕괴제, 활택제, 코팅제 등, 액상 제제에 있어서의 용제, 용해 보조제, 현탁화제, 등장화제, pH 조절제·완충제, 무통화제 등으로서 배합된다. 또한, 필요에 따라 방부제, 항산화제, 착색제, 교미·교취제, 안정화제 등의 제제 첨가물을 사용할 수도 있다.
경구용 고형 제제를 제조하는 경우에는, 본 발명 화합물에 부형제, 필요에 따라 부형제, 결합제, 붕괴제, 활택제, 착색제, 교미·교취제 등을 첨가한 후, 통상의 방법에 의해 정제, 피복 정제, 과립제, 산제, 캡슐제 등을 제조할 수 있다.
경구용 액체 제제를 제조하는 경우에는, 본 발명 화합물에 pH 조절제·완충제, 안정화제, 교미·교취제 등을 첨가해서 통상의 방법에 의해 내복 액제, 시럽제, 엘릭시르제 등을 제조할 수 있다.
주사제를 제조하는 경우에는, 본 발명 화합물에 pH 조절제·완충제, 안정화제, 등장화제, 국소 마취제 등을 첨가하고, 통상의 방법에 의해 피하, 근육내 및 정맥내용 주사제를 제조할 수 있다.
본 발명에는, 본 발명의 폴리펩티드를 검출하기 위한 방법 또는 본 발명의 폴리뉴클레오티드를 검출하기 위한 방법에 있어서, 시료 중 본 발명의 폴리펩티드 또는 본 발명의 폴리뉴클레오티드의 존재를 검출한 경우에, 암이라고 판정하는 방법이 포함된다. 본 발명에 의해 판정할 수 있는 암으로서는, 본 발명의 의약 조성물의 대상이 되는 암으로서 열거한 것 등을 들 수 있다. 상기한 바와 같이 본 발명의 폴리펩티드 또는 폴리뉴클레오티드를 사용함으로써, 암의 판정을 할 수 있다. 따라서, 본 발명의 폴리펩티드, 폴리뉴클레오티드는, 암을 검출하기 위한 바이오마커로서 사용할 수도 있다.
본 발명의 폴리펩티드 또는 본 발명의 폴리뉴클레오티드를, RET를 저해하는 화합물을 사용한 화학 요법이 유효한지 여부의 지표로 하는 방법으로서, 본 발명의 검출 방법에 의해 시료 중에서 본 발명의 폴리펩티드를 검출한 경우, 및/또는 본 발명의 검출 방법에 의해 시료 중에서 본 발명의 폴리뉴클레오티드의 존재를 검출한 경우에, RET를 저해하는 화합물을 사용한 화학 요법이 유효하다고 판정하는 방법이 포함된다.
보다 바람직하게는, 본 발명의 폴리펩티드 또는 본 발명의 폴리뉴클레오티드를, 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제하는 화합물을 사용한 화학 요법이 유효한지 여부의 지표로 하는 방법으로서, 본 발명의 검출 방법에 의해 시료 중에서 본 발명의 폴리펩티드를 검출한 경우, 및/또는 본 발명의 검출 방법에 의해 시료 중에서 본 발명의 폴리뉴클레오티드의 존재를 검출한 경우에, 본 발명의 폴리펩티드의 발현 및/또는 활성, 또는 본 발명의 폴리뉴클레오티드의 발현을 억제하는 화합물을 사용한 화학 요법이 유효하다고 판정하는 방법이 포함된다.
보다 바람직하게는, 본 발명의 폴리펩티드 또는 본 발명의 폴리뉴클레오티드를, 본 발명의 스크리닝 방법에 의해 얻어진 화합물을 사용한 화학 요법이 유효한지 여부의 지표로 하는 방법으로서, 본 발명의 검출 방법에 의해 시료 중에서 본 발명의 폴리펩티드를 검출한 경우, 및/또는 본 발명의 검출 방법에 의해 시료 중에서 본 발명의 폴리뉴클레오티드의 존재를 검출한 경우에, 본 발명의 스크리닝 방법에 의해 얻어진 화합물을 사용한 화학 요법이 유효하다고 판정하는 방법이 포함된다.
이하, 실시예에 의해 본 발명을 구체적으로 설명하지만, 본 발명은 이들 실시예에 의해 한정되는 것은 아니다.
실시예
실시예 1 DCTN1 유전자와 RET 유전자와의 융합 유전자(DCTN1-RET 융합 유전자)의 취득
<1-1 임상 검체 유래 RNA의 추출>
Asterand Bioscience에서 구입한 인간 갑상선암 조직으로부터, RNeasy 미니 키트(Qiagen)를 사용해서 이하의 방법에 의해 RNA를 추출했다. 갑상선암 조직에 완충제 RLT를 600μL 첨가하고, QIAshredder 스핀 칼럼에 어플라이한 후 원심하고 (16,000rpm, 2분, 실온), 여액을 회수했다. 회수한 여액에 동량의 70% 에탄올 수용액을 첨가하고 혼화한 후에, RNeasy 미니 칼럼에 어플라이하고 원심하였다(10,000rpm, 15초, 실온). RNeasy 미니 칼럼에 700μL의 완충제 RW1을 첨가하고 원심하였다(10,000rpm, 15초, 실온). 추가로 500μL의 완충제 RPE를 첨가하고 원심하였다(10,000rpm, 15초, 실온). 마찬가지로 다시 500μL의 완충제 RPE를 첨가하고 원심하였다(10,000rpm, 2분, 실온). 다시 RNeasy 미니 칼럼을 원심(16,000rpm, 1분, 실온)하고, 남은 완충제를 제거했다. RNeasy 미니 칼럼에 RNase 무함유 물을 40μL 어플라이한 후, 원심(10,000rpm, 1분, 실온)하고, 여액을 총 RNA(total RNA)로서 회수했다.
<1-2 임상 검체 유래 cDNA의 제작>
상기 1-1에서 얻어진 총 RNA로부터, 슈퍼스크립트(SuperScript) VILO cDNA 합성 키트(invitorgen)를 사용하여, 이하의 방법에 의해 cDNA를 합성했다. 500ng의 총 RNA를 용량이 14μL가 되도록 RNase 무함유 물로 제조하고, 4μL의 5×VILO 반응 믹스(Reaction Mix) 및 2μL의 10×슈퍼스크립트 효소 믹스(Enzyme Mix)를 첨가하고 혼화했다. 25℃에서 10분간 보온하고 그 후 42℃에서 60분 보온했다. 반응을 정지시키기 위해서 마지막으로 85℃에서 5분간 인큐베이트하여, cDNA를 얻었다.
<1-3 클로닝 벡터의 제작 및 정제>
DCTN1-RET 융합 유전자를 증폭시키기 위해서, 표 1에 나타낸 바와 같이, 센스 프라이머로서 프라이머 1(서열번호 33)과 안티센스 프라이머로서 프라이머 2(서열번호 34), 추가로 네스티드(nested) PCR에 사용하는 센스 프라이머로서 프라이머 3(서열번호 35)과 안티센스 프라이머로서 프라이머 4(서열번호 36)를 설계했다.
이들 프라이머를 사용해서 상기 1-2에서 합성한 cDNA를 주형으로 하여, KOD-Plus-Neo(TOYOBO)를 사용하고, 이하의 방법으로 DCTN1-RET 융합 유전자를 증폭시켰다. 2μL의 cDNA, 5μL의 10×KOD-Plus-Neo용 PCR 완충제, 5μL의 2mM dNTPs, 3μL의 25mM MgSO4, 1μL의 KOD-Plus-Neo, 1.5μL의 프라이머 1(10μM), 1.5μL의 프라이머 2(10μM) 및 31μL의 재증류수(DDW)를 혼화하여, PCR을 행하였다. 이어서 얻어진 PCR 산물을 100배로 희석하고, 2μL의 희석한 PCR 산물, 5μL의 10×KOD-Plus-Neo용 PCR 완충제, 5μL의 2mM dNTPs, 3μL의 25mM MgSO4, 1μL의 KOD-Plus-Neo, 1.5μL의 프라이머 3(10μM), 1.5μL의 프라이머 4(10μM) 및 31μL의 DDW를 혼화하여, 네스티드 PCR을 행하였다.
네스티드 PCR 산물을 1% 아가로오스 겔(나카라이)을 사용한 전기 영동에 의해 분리하고, QIAquick 겔 추출 키트(Qiagen)를 사용하여, 겔로부터 PCR 산물을 정제했다.
제한 효소 SmaI(NEB)로 절단한 pUC18 DNA(타카라 바이오)와 정제한 PCR 산물과 T4 DNA 리가아제(ligase)(NEB)와 T4 DNA 리가아제 반응 완충제(NEB)를 혼화하고, 하룻밤 16℃에서 인큐베이트했다. 라이게이션 산물을 SmaI(NEB)로 처리하고, 이하의 방법으로 적격 세포로의 형질전환을 행하였다. 50μL의 대장균 DH5α 적격 세포(타카라 바이오)에 SmaI 처리를 한 라이게이션 산물을 첨가하여 30분 빙상에서 정치했다. 그 후 42℃에서 30초간 히트 쇼크를 가하고, 2분간 빙상에서 정치했다. SOC 배지(타카라 바이오)를 첨가하여, 37℃에서 1시간 진탕 배양한 후, 암피실린 함유 LB 한천 배지 플레이트(UNITECH)에 배양액을 도포하고, 37℃에서 하룻밤 정치했다. 대장균 콜로니를 암피실린 함유 LB 배지(InvivoGen)에 현탁하고 37℃에서 하룻밤 진탕 배양했다. 증식한 대장균으로부터 QIAquick 스핀 미니프렙 키트(Qiagen)를 사용하여, 첨부의 프로토콜에 준해서 DCTN1-RET 융합 유전자가 삽입된 플라스미드 DNA를 정제했다.
<1-4 서열의 결정>
상기 1-3에서 얻어진 플라스미드 DNA를 주형으로 해서, 표 2에 나타낸 시퀀스용 프라이머의 프라이머 5 내지 프라이머 36을 사용하여, BigDye 터미네이터 v3.1 사이클 시퀀싱 키트를 사용하여 시퀀스 반응을 행하고, Applied Biosystems 3730xl DNA 애널라이저를 사용해서 시퀀스 해석을 실시했다. 시퀀스 해석의 결과, DCTN1-RET 융합 유전자는, DCTN1 변이체 5(GenBank 액세션 번호: NM_001190836)의 엑손 1 내지 엑손 27의 3'측의 하류에 RET 변이체 2(GenBank 액세션 번호: NM_020975)의 엑손 12 내지 엑손 20이 융합한 유전자(서열번호 17)였다.
실시예 2 DCTN1-RET 융합 유전자의 검출
Asterand Bioscience에서 구입한 인간 정상 갑상선 조직 유래 RNA와 상기 1-1에서 얻어진 인간 갑상선암 조직 유래 RNA로부터 슈퍼스크립트 VILO cDNA 합성 키트(invitorgen)를 사용하여, 이하의 방법에 의해 cDNA를 합성했다. 280ng의 총 RNA를 용량이 14μL가 되도록 RNase 무함유 물로 제조하고, 5×VILO 반응 믹스를 4μL, 10×슈퍼스크립트 효소 믹스를 2μL 각각 첨가하고 혼화했다. 25℃에서 10분간 보온하고 그 후 42℃에서 60분 보온했다. 반응을 정지시키기 위해서 마지막으로 85℃에서 5분간 인큐베이트했다.
DCTN1-RET 융합 유전자의 검출을 위해서, 표 3에 나타낸 바와 같이, DCTN-RET 융합 유전자 검출용 센스 프라이머로서 프라이머 37(서열번호 69)과 DCTN-RET 융합 유전자 검출용 안티센스 프라이머로서 프라이머 38(서열번호 70), DCTN1-RET 융합 유전자 검출용 프로브(프로브는 TaqMan MGB 프로브, 형광 색소는 FAM(Thermo Fisher Scientific))으로서 프라이머 39(서열번호 71)를 설계했다.
얻어진 cDNA를 10배 희석하고 1.1μL을 주형으로서 사용하고, 11μL의 프로브용 ddPCR 슈퍼믹스(Bio-Rad), 2μL의 프라이머 37(10μM), 2μL의 프라이머 38(10μM), 0.6μL의 프라이머 39(10μM), 1.1μL의 GAPDH를 검출하는 20×HEX 검정(PrimePCR ddPCR 발현 프로브 검정(Expression Probe Assay): GAPDH, 인간, Bio-Rad)를 혼화시켜서, 자동화 액적 제너레이터(Bio-Rad)로 액적을 제작했다. 제작한 액적을 사용해서 PCR을 행하고, 액적 리더(Bio-Rad)로 DCTN1-RET 및 GAPDH 양성인 액적을 카운트했다. 결과를 도 1 및 도 2에 도시한다.
또한, 상기에서 합성한 cDNA를 주형으로 하여, KOD-Plus-Neo(TOYOBO)를 사용하여 이하의 방법으로 DCTN1-RET 융합 유전자를 증폭시켰다. 2μL의 cDNA, 5μL의 10×KOD-Plus-Neo용 PCR 완충제, 5μL의 2mM dNTPs, 3μL의 25mM MgSO4, 1μL의 KOD-Plus-Neo, 1.5μL의 프라이머 1(10μM), 1.5μL의 프라이머 2(10μM) 및 31μL의 DDW를 혼화하여, PCR을 행하였다. 이어서 얻어진 PCR 산물을 100배로 희석하고, 2μL의 희석한 PCR 산물, 5μL의 10×KOD-Plus-Neo용 PCR 완충제, 5μL의 2mM dNTPs, 3μL의 25mM MgSO4, 1μL의 KOD-Plus-Neo, 1.5μL의 프라이머 3(10μM), 1.5μL의 프라이머 4(10μM) 및 31μL의 DDW를 혼화하여, 네스티드 PCR을 행하였다. 네스티드 PCR 산물을 1% 아가로오스 겔(나카라이)을 사용한 전기 영동에 의해 분리하고 촬영을 행하였다. 결과를 도 3에 도시한다.
도 1, 도 2 및 도 3에 도시한 바와 같이, 인간 갑상선암 조직 유래의 RNA로부터 합성한 cDNA에 있어서는, DCTN1-RET 융합 유전자가 검출된 데 반해, 인간 정상 갑상선 조직 유래 RNA로부터 합성한 cDNA에 있어서는, DCTN1-RET 융합 유전자가 검출되지 않았다. 이상의 결과로부터, DCTN1-RET 융합 유전자는 암 바이오 마커로서 유용한 것이 나타났다.
실시예 3 DCTN1-RET 융합 유전자의 발현 벡터의 구축
발현 벡터 구축을 위해, 표 4에 나타낸 바와 같이, 센스 프라이머로서 프라이머 40(서열번호 72)과 안티센스 프라이머로서 프라이머 41(서열번호 73)을 설계했다.
이들 프라이머를 사용해서 상기 1-2에서 합성한 cDNA를 주형으로 하여, Prime STAR Max DNA 폴리머라아제(Polymerase)(TaKaRa)를 사용하여 이하의 방법으로 DCTN1-RET 융합 유전자를 증폭시켰다. 1μL의 cDNA, 25μL의 2×Prime STAR Max DNA 폴리머라아제, 1μL의 프라이머 40(10μM), 1μL의 프라이머 41(10μM) 및 22μL의 재증류수(DDW)를 혼화하여, PCR을 행하였다. 얻어진 PCR 산물을 1% 아가로오스 겔(나카라이)을 사용한 전기 영동에 의해 분리하고, GFX PCR DNA 및 겔 밴드 정제 키트(GE Healthcare)를 사용하여, 겔로부터 PCR 산물을 정제했다. 이어서 얻어진 정제 PCR 산물을 Gateway BP Clonase II 효소 믹스(ThermoFisher)를 사용해서 이하의 방법으로 Gateway pDONR221 벡터에 삽입하고, 엔트리 벡터를 제작했다. 구체적으로는, 5.0μL의 정제 PCR 산물, 3.5μL의 pDONR221(85ng/μL), 4.0μL의 BP Clonase II 효소 믹스, 7.5μL의 TE를 혼화하고, 25℃에서 90분간 인큐베이트했다. 인큐베이트 후에 프로테이나아제(Proteinase) K(2㎎/mL)를 1μL 첨가하고, 37℃에서 10분간 인큐베이트함으로써 엔트리 벡터를 얻었다.
얻어진 엔트리 벡터를 50μL의 대장균 DH5α 적격 세포(타카라 바이오)에 첨가하여 30분 빙상에서 정치했다. 그 후 37℃에서 20초간 히트 쇼크를 가하여, 2분간 빙상에서 정치했다. SOC 배지(타카라 바이오)를 첨가하여, 37℃에서 1시간 진탕 배양한 후, 카나마이신 함유 LB 한천 배지 플레이트에 배양액을 도포하고, 37℃에서 하룻밤 정치했다. 대장균 콜로니를 카나마이신 함유 LB 배지에 현탁하고, 37℃에서 하룻밤 진탕 배양했다. 증식한 대장균으로부터 DCTN1-RET 융합 유전자가 삽입된 플라스미드 DNA(엔트리벡터 크론)를 DNA 자동 분리 장치 GENE PREP STAR PI-480(구라보)에서 정제했다.
얻어진 플라스미드와 Gateway LR Clonase II 효소 믹스(ThermoFisher)를 사용하여, 이하의 방법으로, pJTI Fast DEST 벡터에 DCTN1-RET 융합 유전자를 삽입하여, 발현 벡터로 했다. 150ng의 엔트리 벡터 크론, 1μL의 pJTI Fast DEST 벡터(150ng/μL), 2μL의 LR Clonase II 효소 믹스 및 TE 완충제를 혼화하고, 전량을 10μL로 하고, 25℃에서 90분간 인큐베이트했다. 인큐베이트 후에 프로테이나아제 K(2㎎/mL)를 1μL 첨가하고, 37℃에서 10분간 인큐베이트함으로써, DCTN1-RET 융합 유전자가 삽입된 pJTI Fast DEST 벡터(DCTN1-RET 융합 유전자 발현 벡터)를 얻었다. 얻어진 DCTN1-RET 융합 유전자 발현 벡터를 50μL의 대장균 DH5α 적격 세포(타카라 바이오)에 첨가하여 30분 빙상에서 정치했다. 그 후 37℃에서 20초간 히트 쇼크를 가하여, 2분간 빙상에서 정치했다. SOC 배지(타카라 바이오)를 첨가하여, 37℃에서 1시간 진탕 배양한 후, 암피실린 함유 LB 한천 배지 플레이트에 배양액을 도포하고, 37℃에서 하룻밤 정치했다. 대장균 콜로니를 암피실린 함유 LB 배지에 현탁하고 37℃에서 하룻밤 진탕 배양했다. 증식한 대장균으로부터 DCTN1-RET 융합 유전자가 삽입된 플라스미드 DNA(DCTN1-RET 융합 유전자 발현 벡터)를 플라스미드 플러스 맥시 키트(QIAGEN)를 사용하여 정제했다.
실시예 4 DCTN1-RET 융합 유전자 발현 세포의 수립
<4-1 세포의 수립>
DCTN1-RET 융합 유전자 발현 세포의 수립을 위한 숙주 세포로서, 마우스 태아 섬유아 세포 NIH/3T3 세포(American Type Culture Collection)를 선택하고, 상기에서 제작한 DCTN1-RET 융합 유전자가 삽입된 발현 벡터를 형질감염함으로써, DCTN1-RET 융합 유전자 발현 세포를 수립했다. 상세한 방법은 이하에 나타내는 바와 같이 실시했다. NIH/3T3 세포는, 통상의 배양(2차원 배양)을 위해서, D-MEM(고글루코오스)(L-글루타민, 페놀레드, 피루브산나트륨, 1500㎎/L 탄산수소나트륨 함유)(WAKO)에 10%가 되도록 초생 송아지 혈청(Newborn Calf Serum; NBCS)(GIBCO)을 첨가한 것을 2차원 배양용 배지로서 사용하고, 37℃, 5% CO2로 배양한 것을 사용했다. 형질감염을 행하기 전날에, NIH/3T3 세포를 1.5×105cells/2mL에서, 6웰 플레이트(IWAKI)에 파종하고, 37℃, 5% CO2로, 하룻밤 인큐베이트했다. 1.5μg의 DCTN1-RET 융합 유전자 발현 벡터, 1.5μg의 pJTI phiC31 인테그라아제 벡터를 혼합한 혼합액에, 그 혼합액의 6배량의 ViaFect 형질감염 시약을 첨가한 후, 전량이 300μL가 되도록 Opti-MEM을 첨가하고, 실온에서 5분간 인큐베이트함으로써 형질감염 용액을 제작했다. NIH/3T3 세포가 파종되어 있는 웰로부터 배지를 300μL 제거하고, 상기에서 제작한 형질감염 용액을 300μL 첨가하고, 37℃, 5% CO2로, 하룻밤 인큐베이트했다. 다음날, 형질감염 용액을 제거하기 위해서, 배지를 교환했다. 배지를 교환 할 때, 새로운 배지에는, 하이그로마이신 B(나카라이테스크)를 500μg/mL가 되도록 첨가했다. 하이그로마이신 B에 의해, DCTN1-RET 융합 유전자 도입 발현 벡터가 도입되어 있지 않은 세포를 제거했다.
형질감염 후, 1주일에 2회 정도의 배지 교환을 하면서, 세포가 증식할 때까지, 배양을 행하였다. 형질감염 22일 후에, 세포를 트립신으로 회수하고, 이하의 방법으로 단세포 클로닝을 행하였다. 회수한 세포의 세포수를 측정하여, 1cell/200μL가 되도록 배지를 첨가했다. 96웰 플레이트(ThermoFisher)에 200μL/1웰이 되도록 세포를 파종했다. 파종 후, 매일 관찰을 행하여, 단세포로부터 증식해 온 세포를 취득하고, DCTN1-RET 융합 유전자 발현 세포(DCTN1-RET 융합 유전자 발현 NIH/3T3 세포)로 했다.
<4-2 목적 단백질의 발현 확인>
얻어진 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서의 DCTN1-RET 융합 단백질의 발현을, 웨스턴 블로트법으로 확인했다. 즉, 배양 플라스크로부터 배지를 제거하고, PBS로 한번 세정했다. 배양 플라스크 내에 포스파타아제 저해제(phosphatase inhibitor)(로슈)와 프로테아제 저해제(protease inhibitor)(로슈)를 함유한 샘플 희석 농축물(Sample Diluent Concentrate) 2(R&D SYSTEMS)를 첨가하고, 스크레이퍼로 세포 용해액을 회수했다. 회수한 세포 용해액으로부터 원심 분리에 의해, 단백질 샘플을 얻었다. 단백질 샘플은, 단백질 정량을 행하여, 단백질 농도를 일정하게 했다. 일정 농도의 단백질 샘플에 대하여, SDS-PAGE용 환원 시약 함유 샘플 완충 용액(6x)(나카라이테스크)을 첨가하여, 95℃에서 5분간 인큐베이트함으로써, 단백질을 변성시켜서, 웨스턴 블로팅용 샘플을 얻었다. 또한, 네거티브 컨트롤용 샘플로서, 친주인 NIH/3T3 세포를 사용하여, 마찬가지 방법으로 웨스턴 블로팅용 샘플을 얻었다. 상기 샘플을 사용하여, 이하에 나타내는 방법으로, 단백질의 발현을 확인했다. 4 내지 15퍼센트 아크릴아미드 겔(BIO-RAD) 및 1×Tris/글리신/SDS 완충제를 사용하여, SDS-PAGE 전기 영동(200V에서 30분)에서, 단백질을 분리했다. Trans-Blot Turbo RTA 미디 PVDF 트랜스퍼 키트(BIO-RAD)와 Transblot Turbo 전사 시스템(BIO-RAD)을 사용해서 PVDF막에 단백질을 전사하고, PVDF막을 블로킹 원-피(Blocking One-P)에 1시간 침지했다. 블로킹 원-피가 10%가 되도록 TBS-T로 희석한 용액으로 1차 항체(포스포-Ret(Tyr905) 항체(CST), Ret(C31B4) 토끼 mAb(CST) 및 항-Dctn1 항체(ATLAS ANTIBODIES))를 1/1000 농도가 되도록 희석하고, PVDF막을 침지시켜서, 4℃에서 하룻밤 인큐베이트했다. TBS-T로 세정 후, 항-토끼 IgG, HRP-연결 항체(CST)를 1/2000 농도가 되도록 TBS-T로 희석한 2차 항체 희석액으로, PVDF막을 침지시켜서, 실온에서 1시간 인큐베이트했다. TBS-T로 세정 후, SuperSignal 웨스트 듀라 연장 기간 기질(West Dura Extended Duration Substrate)(ThermoFisher) 및 루미노·이미지 애널라이저 Amersham Imager 600(GE 헬스케어)을 사용해서 단백질의 검출을 행하였다. 또한, 검출 단백질의 분자량은, 프리시전 플러스 프로테인 칼레이도스코프 스탠다드(BIORAD)에 의해 확인했다.
그 결과, 도 4의 a) 및 b)에 나타낸 바와 같이, 항pRET 항체와 항RET 항체를 사용한바 내재성의 RET(150 및 175kDa)는 검출되지 않았다. 한편, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서만, 175kDa 부근에 DCTN1-RET 융합 단백질이라고 추측되는 밴드를 확인할 수 있었다.
또한, 도 4의 c)에 나타낸 바와 같이 항DCTN1 항체를 사용한바, 150kDa 부근에 내재성의 DCTN1이 검출되었다. 한편, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서만, 내재성의 150kDa의 DCTN1의 밴드 상의 175kDa 부근에 밴드가 검출되었다. 즉, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 있어서만, RET에 대한 항체 및 DCTN1에 대한 항체의 양쪽에 175kDa 부근의 밴드가 검출된 점에서, 제작한 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에서는, DCTN1과 RET가 융합한 단백질이 발현하고 있는 것이 명확해졌다.
실시예 5 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 3차원 배양에 의한 증식 확인
NIH/3T3 세포는, 2차원 배양 조건에서는 양호한 증식을 나타내지만, 3차원 배양 조건에서는, 대부분 증식하지 않는다. 그 반면, NIH/3T3 세포에 암 유전자를 발현시킴으로써, 3차원 배양 조건에서도 증식하는 것이 알려져 있다. 그 성질을 이용하여, DCTN1-RET 융합 유전자가 암 유전자인지의 확인을 행하였다. 37℃, 5% CO2로, 2차원 배양으로 배양한 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포 및 NIH/3T3 세포를 트립신으로 회수하고, 세포수를 측정했다. 3차원 배양을 행하기 위해서, FCeM 시리즈 제조 키트(닛산 가가쿠 고교 가부시키가이샤)와 D-MEM(고글루코오스)(L-글루타민, 페놀레드, 피루브산나트륨, 1500㎎/L 탄산수소나트륨 함유)(WAKO)과 초생 송아지 혈청(NBCS)(GIBCO)을 사용하여, 3차원 배양용 배지를 제작했다. 제작한 3차원 배양용 배지에, 세포를 1000cells/90μL가 되도록 현탁하고, 96웰 투명 검정 둥근 바닥의 타원 마이크로플레이트(Corning)에 90μL/1well이 되도록 파종하고, 37℃, 5% CO2로 인큐베이트했다. 파종 다음날(Day1) 및 파종 8일 후(Day8)에 세포 내 ATP 발광 검출 시약인 Celltiter-Glo 2.0 시약(Progema) 및 루미노미터(EnSpire, PerkinElmer)를 사용해서 발광량(counts per second: cps)을 측정하여, 생세포수의 지표로 했다. Day1에서의 측정 결과와 Day8에서의 측정 결과로부터 각 세포의 증식율을 산출했다(N=3).
그 결과, 도 5에 도시한 바와 같이, NIH/3T3 세포로는, Day8의 세포수는 Day1의 세포수와 비교해서 2.4배였던 데 비해, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에서는, 20.9배였다. 또한, NIH/3T3 세포는, 3차원 배양에 있어서 세포의 응집 덩어리는 형성되지 않지만, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에서는, 3차원 배양에 의해 세포의 응집 덩어리가 형성되어 있는 것이 확인되었다.
즉, DCTN1-RET 융합 유전자를 도입함으로써, 세포의 증식이 항진한 것이 명확해지고, DCTN1-RET 융합 유전자가 암 유전자인 것이 시사되었다.
실시예 6 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 생체 내에 있어서의 종양 형성성 확인
DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 생체 내에 있어서의 종양 형성성을 확인하기 위해서, 누드마우스를 사용한 이식 실험을 행하였다. 부언하면, 친주인 NIH/3T3 세포는, 누드마우스의 피하에서는 증식하지 않는 것이, 일반적으로 알려져 있고, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포를 누드마우스의 피하에 이식함으로써, DCTN1-RET 융합 유전자가 종양 형성성에 기여하는지, 즉 암 유전자인지의 확인을 할 수 있다. 피이식 동물로서는, 누드마우스(BALB/cAJcl-nu/nu, 니혼 클레아)를 사용했다. DCTN1-RET 융합 유전자 발현 NIH/3T3 세포를 트립신으로 회수하고, 최종적으로 1×108cells/mL가 되도록 PBS에 현탁하고, 동량의 마트리겔 기저막 매트릭스(Corning)를 첨가하고, 5×107cells/mL로 한 것을 이식용 세포액으로 했다. 25G 주사 바늘과 1mL 시린지를 사용하여, 이식용 세포액을 누드마우스(N=10)의 우측 흉부의 피하에 0.1mL씩 이식했다. 전자 노기스(미츠토요)를 사용하여, 이식 후, 10, 13, 17일째에, 1마리씩 종양의 긴 직경 및 짧은 직경을 측정하고, 이하의 식을 사용해서 종양 체적을 산출했다.
종양 체적(㎣)=(긴 직경, ㎜)×(짧은 직경, ㎜)×(짧은 직경, ㎜)/2
종양 체적의 측정 결과를 도 6에 나타낸다. 그 결과, 누드마우스의 피하에 이식된 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포는, 종양을 형성하고, 양호하게 증식하는 것이 확인되고, 생체 내 실험에 있어서도 DCTN1-RET 융합 유전자가 암 유전자인 것이 시사되었다.
실시예 7 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포를 사용한 siRNA에 의한 DCTN1-RET 융합 단백질의 억제와 세포 증식 억제 효과의 확인
DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 대한 siRNA 처리에 의한 영향을 확인했다. 사용한 siRNA는, 하기의 표 5에 나타낸 3종류의 RET siRNA와 네거티브 컨트롤로서 Silencer 셀렉트 네거티브 컨트롤 #1 siRNA(Ambion)를 사용했다. 또한, 3종류의 RET siRNA는, 모두 인간 RET를 표적으로 하는 siRNA이지만, RET siRNA1 및 RET siRNA2는 DCTN1-RET 융합 유전자 내의 RET 부분에 결합하는 서열을 포함하고, RET siRNA3은 DCTN1-RET 융합 유전자 내에 결합하는 서열을 포함하지 않는다. 즉, RET siRNA1 및 RET siRNA2는 DCTN1-RET 융합 유전자의 발현을 억제하지만, RET siRNA3은 DCTN1-RET 융합 유전자의 발현을 억제하지 않는 것이 상정되었다. 이하에 siRNA를 사용한 실험의 방법에 대해서 기재했다.
DCTN1-RET 융합 유전자 발현 NIH/3T3 세포는, 2차원 배양용 배지를 사용하고, 37℃, 5% CO2에서 배양한 것을 사용했다. siRNA 처리를 행하기 전날에, 각 세포를 3×105cells/2mL로, 6웰 플레이트(IWAKI)에 파종하고, 37℃, 5% CO2로, 하룻밤 인큐베이트했다. 사전에 물을 사용해서 20μM으로 제조한 각 siRNA를 12μL, 4μL의 리포펙타민(Lipofectamin) RNAiMAX 형질감염 시약(ThermoFisher) 및 384μL의 Opti-MEM을 혼합하고, 실온에서 15분 인큐베이트함으로써 siRNA 용액을 제작했다. 각 세포가 파종되어 있는 웰에 siRNA 용액을 400μL 첨가하고, 37℃, 5% CO2로, 하룻밤 인큐베이트했다.
다음날, 일부는, 단백질 발현 해석용으로 샘플링을 행하고, 일부에 대해서는, 세포 증식 확인용으로 재파종을 행하였다. 단백질 발현 해석용 샘플링 및 단백질 발현 해석은, 1차 항체로서, 포스포-Ret(Tyr905) 항체(CST), Ret(C31B4) 토끼 mAb(CST) 및 GAPDH(D16H11) XP 토끼 mAb(CST)를 사용한 것 이외에는, 상기 <4-2 목적 단백질의 발현 확인>과 마찬가지 방법으로 실시했다. 그 결과, 도 7에 나타낸 바와 같이, siRNA 처리하지 않은 세포(무처리)에 비해, 네거티브 컨트롤 siRNA(NC)를 처리한 세포에 있어서, DCTN1-RET 융합 단백질의 발현은 억제되어 있지 않은 것을 확인할 수 있었다. 한편, RET siRNA1 및 RET siRNA2를 처리한 경우, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 DCTN1-RET 융합 단백질의 발현이 억제되는 것이 확인되고, RET siRNA3에서는 억제되지 않는 것이 명확해졌다.
다음에 세포 증식 억제 효과의 확인을 위해, 무처리 또는 siRNA 처리된 웰로부터 트립신에 의해 세포를 회수하고, 세포수를 측정했다. 상기 실시예 5와 마찬가지 방법으로 3차원 배양을 행하고, 파종 당일(Day0) 및 파종 4일 후(Day4)에 실시예 5와 마찬가지 방법으로 생세포수를 측정했다. Day0에서의 측정 결과와 Day4에서의 측정 결과로부터 각 세포에 있어서의 증식율을 산출했다.
그 결과, 도 8에 나타낸 바와 같이, siRNA를 처리하지 않은 세포(무처리) 및 네거티브 컨트롤 siRNA(NC)를 처리한 세포에서는, Day4의 세포수는, Day0의 세포수와 비교하여, 4.9배 및 3.6배였던 데 비해, RET siRNA1 및 RET siRNA2로 처리한 세포에서는, 2.0배 및 2.4배 정도의 증식으로, 현저하게 증식율이 저하되었다. 한편, RET siRNA3로 처리한 세포에서는, 3.7배의 증식으로, 네거티브 컨트롤 siRNA와 동일 정도의 증식율이며, 증식율의 저하는 보이지 않았다. 이들 결과로부터, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 증식은, siRNA에 의해 RET의 발현을 저해한 경우에도 억제되는 것이 명확해졌다.
실시예 8 DCTN1-RET 융합 유전자 발현 NIH/3T3 세포를 사용한 세포 증식 억제 효과
DCTN1-RET 융합 유전자 발현 NIH/3T3 세포에 대한 시험관 내(in vitro) 세포 증식 시험을 행하였다. 상기 실시예 5와 마찬가지 방법으로, 3차원 배양 및 파종을 행하였다. 파종 후 37℃, 5% CO2로 하룻밤 인큐베이트하였다(Day0). RET를 저해한다고 보고되어 있는 카보잔티닙, 반데타닙, 알렉티닙, 렌바티닙, 축합 피리미딘 화합물(화합물 1 내지 9; 표 6에 나타낸다)을 디메틸술폭시드로 10mmol/L의 농도로 용해하고, 추가로 3차원 배양용 배지를 사용하여, 이들 화합물의 최종 농도가 각각 1000, 333, 111, 37.0, 12.3, 4.12, 1.37, 0.457nmol/L가 되도록 희석을 행하였다. 이것을 먼저 설명한 세포가 파종된 플레이트의 각 웰에 0.01mL씩 첨가하여(Day1), 37℃, 5% CO2로 7일간 인큐베이트했다. 배양 후(Day8), 모든 웰에 세포 내 ATP 발광 검출 시약인 Celltiter-Glo 2.0 시약(Progema)를 첨가하고, 루미노미터(EnSpire, PerkinElmer)를 사용해서 발광량(counts per second: cps)을 측정했다. Tday8과 Cday1의 값의 크기에 따라, 이하의 식으로부터 화합물의 각 농도에 있어서의 Day1로부터의 증식율을 산출하고, 세포 증식을 50% 억제하는 피검 화합물의 농도(GI50(nM))를 구했다.
1) Tday8≥Cday1인 경우
증식율(%)=(Tday8-Cday1)/(Cday8-Cday1)×100
T: 피검 화합물을 첨가한 웰의 cps
C: 피검 화합물을 첨가하지 않은 웰의 cps
Day1: 피검 화합물을 첨가한 날
Day8: 평가일
2) Tday8<Cday1인 경우
증식율(%)=(Tday8-Cday1)/(Cday1)×100
T: 피검 화합물을 첨가한 웰의 cps
C: 피검 화합물을 첨가하지 않은 웰의 cps
Day1: 피검 화합물을 첨가한 날
Day8: 평가일
그 결과, 표 7에 나타낸 바와 같이, 카보잔티닙, 반데타닙, 렌바티닙 및 축합 피리미딘 화합물(화합물 1 내지 9)에 있어서, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 증식이 억제되었다.
이상의 결과로부터, 상기 RET 저해제는, DCTN1-RET 융합 유전자가 검출된 암에 대한 치료약으로서 유용할 가능성이 시사되었다. 또한, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포를 사용함으로써, DCTN1-RET를 억제하는 화합물의 스크리닝이 가능한 것이 시사되었다.
실시예 9 DCTN1-RET 융합 유전자 발현 세포를 사용한 RET의 인산화 저해
DCTN1-RET 융합 유전자 발현 세포에 있어서의 RET의 인산화가, RET를 저해한다고 보고되어 있는 기존의 약제로 저해되는지에 대해서 이하의 방법으로 검토했다.
DCTN1-RET 융합 유전자 발현 NIH/3T3 세포는, 2차원 배양용 배지를 사용하고, 37℃, 5% CO2로 배양한 것을 사용했다. 약제 처리를 행하기 전날에, 각 세포를 3×105cells/2mL로, 6웰 플레이트(IWAKI)에 파종하고, 37℃, 5% CO2로, 하룻밤 인큐베이트했다. 카보잔티닙, 반데타닙, 알렉티닙, 렌바티닙을 디메틸술폭시드로 10mmol/L의 농도로 용해하고, 추가로 PBS를 사용하여, 이들 화합물의 최종 농도가 각각 1000, 100, 10nmol/L가 되도록 희석을 행하였다. 이것을 먼저 설명한 세포가 파종된 플레이트의 각 웰에 20μL씩 첨가하여(Day1), 37℃, 5% CO2로 1시간 인큐베이트했다. 인큐베이트 후, 상기 실시예 7에 기재되어 있는 방법과 마찬가지 방법으로 단백질 발현 해석용 샘플링을 행하여, 단백질 발현 해석을 실시했다.
그 결과, 도 9에 나타낸 바와 같이, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 인산화 RET 레벨이 카보잔티닙 및 렌바티닙에 의해 현저하게 감소하는 것이 확인되었다. 또한, 상기와 마찬가지 방법으로, 축합 피리미딘 화합물을 사용하여, RET의 인산화 저해를 평가한 결과, 축합 피리미딘 화합물에 있어서도 RET의 인산화가 현저하게 감소하는 것을 확인할 수 있었다.
이상의 결과로부터, 인산화 RET 레벨을 현저하게 감소시키는 약제는, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 증식을 억제할 수 있는 화합물이며, DCTN1-RET 융합 유전자가 검출된 암에 대한 치료약으로서 유용할 가능성이 시사되었다. 또한, DCTN1-RET 융합 유전자 발현 NIH/3T3 세포의 인산화 RET 레벨을 사용함으로써 RET 저해제의 스크리닝이 가능한 것이 시사되었다.
서열번호 1은 DCTN1 변이체 1(v1)[서열번호 25의 일부]과 RET 변이체 2(v2)[서열번호 31의 일부]와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 2는 DCTN1 v1과 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 3은 DCTN1 v1과 RET 변이체 4(v4)[서열번호 32의 일부]와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 4는 DCTN1 v1과 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 5는 DCTN1 변이체 2(v2)[서열번호 26의 일부]와 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 6은 DCTN1 v2와 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 7은 DCTN1 v2와 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 8은 DCTN1 v2와 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 9는 DCTN1 변이체 3(v3)[서열번호 27의 일부]과 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 10은 DCTN1 v3과 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 11은 DCTN1 v3과 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 12는 DCTN1 v3과 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 13은 DCTN1 변이체 4(v4)[서열번호 28의 일부]와 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 14는 DCTN1 v4와 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 15는 DCTN1 v4와 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 16은 DCTN1 v4와 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 17은 DCTN1 변이체 5(v5)[서열번호 29의 일부]와 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 18은 DCTN1 v5와 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 19는 DCTN1 v5와 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 20은 DCTN1 v5와 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 21은 DCTN1 v6과 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 22는 DCTN1 v6과 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 23은 DCTN1 v6과 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 24는 DCTN1 v6과 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 33은 프라이머의 염기 서열을 나타낸다.
서열번호 34는 프라이머의 염기 서열을 나타낸다.
서열번호 35는 프라이머의 염기 서열을 나타낸다.
서열번호 36은 프라이머의 염기 서열을 나타낸다.
서열번호 37은 프라이머의 염기 서열을 나타낸다.
서열번호 38은 프라이머의 염기 서열을 나타낸다.
서열번호 39는 프라이머의 염기 서열을 나타낸다.
서열번호 40은 프라이머의 염기 서열을 나타낸다.
서열번호 41은 프라이머의 염기 서열을 나타낸다.
서열번호 42는 프라이머의 염기 서열을 나타낸다.
서열번호 43은 프라이머의 염기 서열을 나타낸다.
서열번호 44는 프라이머의 염기 서열을 나타낸다.
서열번호 45는 프라이머의 염기 서열을 나타낸다.
서열번호 46은 프라이머의 염기 서열을 나타낸다.
서열번호 47은 프라이머의 염기 서열을 나타낸다.
서열번호 48은 프라이머의 염기 서열을 나타낸다.
서열번호 49는 프라이머의 염기 서열을 나타낸다.
서열번호 50은 프라이머의 염기 서열을 나타낸다.
서열번호 51은 프라이머의 염기 서열을 나타낸다.
서열번호 52는 프라이머의 염기 서열을 나타낸다.
서열번호 53은 프라이머의 염기 서열을 나타낸다.
서열번호 54는 프라이머의 염기 서열을 나타낸다.
서열번호 55는 프라이머의 염기 서열을 나타낸다.
서열번호 56은 프라이머의 염기 서열을 나타낸다.
서열번호 57은 프라이머의 염기 서열을 나타낸다.
서열번호 58은 프라이머의 염기 서열을 나타낸다.
서열번호 59는 프라이머의 염기 서열을 나타낸다.
서열번호 60은 프라이머의 염기 서열을 나타낸다.
서열번호 61은 프라이머의 염기 서열을 나타낸다.
서열번호 62는 프라이머의 염기 서열을 나타낸다.
서열번호 63은 프라이머의 염기 서열을 나타낸다.
서열번호 64는 프라이머의 염기 서열을 나타낸다.
서열번호 65는 프라이머의 염기 서열을 나타낸다.
서열번호 66은 프라이머의 염기 서열을 나타낸다.
서열번호 67은 프라이머의 염기 서열을 나타낸다.
서열번호 68은 프라이머의 염기 서열을 나타낸다.
서열번호 69는 프라이머의 염기 서열을 나타낸다.
서열번호 70은 프라이머의 염기 서열을 나타낸다.
서열번호 71은 프라이머의 염기 서열을 나타낸다.
서열번호 72는 프라이머의 염기 서열을 나타낸다.
서열번호 73은 프라이머의 염기 서열을 나타낸다.
서열번호 74는 RET siRNA의 염기 서열을 나타낸다.
서열번호 75는 RET siRNA의 염기 서열을 나타낸다.
서열번호 76은 RET siRNA의 염기 서열을 나타낸다.
서열번호 77은 RET siRNA의 염기 서열을 나타낸다.
서열번호 2는 DCTN1 v1과 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 3은 DCTN1 v1과 RET 변이체 4(v4)[서열번호 32의 일부]와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 4는 DCTN1 v1과 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 5는 DCTN1 변이체 2(v2)[서열번호 26의 일부]와 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 6은 DCTN1 v2와 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 7은 DCTN1 v2와 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 8은 DCTN1 v2와 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 9는 DCTN1 변이체 3(v3)[서열번호 27의 일부]과 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 10은 DCTN1 v3과 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 11은 DCTN1 v3과 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 12는 DCTN1 v3과 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 13은 DCTN1 변이체 4(v4)[서열번호 28의 일부]와 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 14는 DCTN1 v4와 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 15는 DCTN1 v4와 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 16은 DCTN1 v4와 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 17은 DCTN1 변이체 5(v5)[서열번호 29의 일부]와 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 18은 DCTN1 v5와 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 19는 DCTN1 v5와 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 20은 DCTN1 v5와 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 21은 DCTN1 v6과 RET v2와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 22는 DCTN1 v6과 RET v2와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 23은 DCTN1 v6과 RET v4와의 융합 펩티드를 코딩하는 폴리뉴클레오티드의 염기 서열을 나타낸다.
서열번호 24는 DCTN1 v6과 RET v4와의 융합 펩티드의 아미노산 서열을 나타낸다.
서열번호 33은 프라이머의 염기 서열을 나타낸다.
서열번호 34는 프라이머의 염기 서열을 나타낸다.
서열번호 35는 프라이머의 염기 서열을 나타낸다.
서열번호 36은 프라이머의 염기 서열을 나타낸다.
서열번호 37은 프라이머의 염기 서열을 나타낸다.
서열번호 38은 프라이머의 염기 서열을 나타낸다.
서열번호 39는 프라이머의 염기 서열을 나타낸다.
서열번호 40은 프라이머의 염기 서열을 나타낸다.
서열번호 41은 프라이머의 염기 서열을 나타낸다.
서열번호 42는 프라이머의 염기 서열을 나타낸다.
서열번호 43은 프라이머의 염기 서열을 나타낸다.
서열번호 44는 프라이머의 염기 서열을 나타낸다.
서열번호 45는 프라이머의 염기 서열을 나타낸다.
서열번호 46은 프라이머의 염기 서열을 나타낸다.
서열번호 47은 프라이머의 염기 서열을 나타낸다.
서열번호 48은 프라이머의 염기 서열을 나타낸다.
서열번호 49는 프라이머의 염기 서열을 나타낸다.
서열번호 50은 프라이머의 염기 서열을 나타낸다.
서열번호 51은 프라이머의 염기 서열을 나타낸다.
서열번호 52는 프라이머의 염기 서열을 나타낸다.
서열번호 53은 프라이머의 염기 서열을 나타낸다.
서열번호 54는 프라이머의 염기 서열을 나타낸다.
서열번호 55는 프라이머의 염기 서열을 나타낸다.
서열번호 56은 프라이머의 염기 서열을 나타낸다.
서열번호 57은 프라이머의 염기 서열을 나타낸다.
서열번호 58은 프라이머의 염기 서열을 나타낸다.
서열번호 59는 프라이머의 염기 서열을 나타낸다.
서열번호 60은 프라이머의 염기 서열을 나타낸다.
서열번호 61은 프라이머의 염기 서열을 나타낸다.
서열번호 62는 프라이머의 염기 서열을 나타낸다.
서열번호 63은 프라이머의 염기 서열을 나타낸다.
서열번호 64는 프라이머의 염기 서열을 나타낸다.
서열번호 65는 프라이머의 염기 서열을 나타낸다.
서열번호 66은 프라이머의 염기 서열을 나타낸다.
서열번호 67은 프라이머의 염기 서열을 나타낸다.
서열번호 68은 프라이머의 염기 서열을 나타낸다.
서열번호 69는 프라이머의 염기 서열을 나타낸다.
서열번호 70은 프라이머의 염기 서열을 나타낸다.
서열번호 71은 프라이머의 염기 서열을 나타낸다.
서열번호 72는 프라이머의 염기 서열을 나타낸다.
서열번호 73은 프라이머의 염기 서열을 나타낸다.
서열번호 74는 RET siRNA의 염기 서열을 나타낸다.
서열번호 75는 RET siRNA의 염기 서열을 나타낸다.
서열번호 76은 RET siRNA의 염기 서열을 나타낸다.
서열번호 77은 RET siRNA의 염기 서열을 나타낸다.
SEQUENCE LISTING
<110> TAIHO PHARMACEUTICAL CO., LTD.
<120> Fusion peptide of DCTN1 protein and RET protein
<130> P17-158WO
<150> JP 2017-158796
<151> 2017-08-21
<160> 77
<170> PatentIn version 3.5
<210> 1
<211> 4908
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 1
atggcacaga gcaagaggca cgtgtacagc cggacgccca gcggcagcag gatgagtgcg 60
gaggcaagcg cccggcctct gcgggtgggc tcccgtgtag aggtgattgg aaaaggccac 120
cgaggcactg tggcctatgt tggagccaca ctgtttgcca ctggcaaatg ggtaggcgtg 180
attctggatg aagcaaaggg caaaaatgat ggaactgttc aaggcaggaa gtacttcact 240
tgtgatgaag ggcatggcat ctttgtgcgc cagtcccaga tccaggtatt tgaagatgga 300
gcagatacta cttccccaga gacacctgat tcttctgctt caaaagtcct caaaagagag 360
ggaactgata caactgcaaa gactagcaaa ctgcggggac tgaagcctaa gaaggcaccg 420
acagcccgaa agaccacaac tcggcgaccc aagcccacgc gcccagccag tactggggtg 480
gctggggcca gtagctccct gggcccctct ggctcagcgt cagcaggtga gctgagcagc 540
agtgagccca gcaccccggc tcagactccg ctggcagcac ccatcatccc cacgccggtc 600
ctcacctctc ctggagcagt ccccccgctt ccttccccat ccaaggagga ggagggacta 660
agggctcagg tgcgggacct ggaggagaaa ctagagaccc tgagactgaa acgggcagaa 720
gacaaagcaa agctaaaaga gctggagaaa cacaaaatcc agctggagca ggtgcaggaa 780
tggaagagca aaatgcagga gcagcaggcc gacctgcagc ggcgcctcaa ggaggcgaga 840
aaggaagcca aggaggcgct ggaggcaaag gaacgctata tggaggagat ggctgatact 900
gctgatgcca ttgagatggc cactttggac aaggagatgg ctgaagagcg ggctgagtcc 960
ctgcagcagg aggtggaggc actgaaggag cgggtggacg agctcactac tgacttagag 1020
atcctcaagg ctgagattga agagaagggc tcagatggcg ctgcatccag ttatcagctc 1080
aagcagcttg aggagcagaa tgcccgcctg aaggatgccc tggtgaggat gcgggatctt 1140
tcttcctcag agaagcagga gcatgtgaag ctccagaagc tcatggaaaa gaagaaccaa 1200
gagctggaag ttgtgaggca acagcgggag cgtctgcagg aggagctaag ccaggcagag 1260
agcaccattg atgagctcaa ggagcaggtg gatgctgctc tgggtgctga ggagatggtg 1320
gagatgctga cagatcggaa cctgaatctg gaagagaaag tgcgcgagtt gagggagact 1380
gtgggagact tggaagcgat gaatgagatg aacgatgagc tgcaggagaa tgcacgtgag 1440
acagaactgg agctgcggga gcagctggac atggcaggcg cgcgggttcg tgaggcccag 1500
aagcgtgtgg aggcagccca ggagacggtt gcagactacc agcagaccat caagaagtac 1560
cgccagctga ccgcccatct acaggatgtg aatcgggaac tgacaaacca gcaggaagca 1620
tctgtggaga ggcaacagca gccacctcca gagacctttg acttcaaaat caagtttgct 1680
gagactaagg cccatgccaa ggcaattgag atggaattga ggcagatgga ggtggcccag 1740
gccaatcgac acatgtccct gctgacagcc ttcatgcctg acagcttcct tcggccaggt 1800
ggggaccatg actgcgttct ggtgctgttg ctcatgcctc gtctcatttg caaggcagag 1860
ctgatccgga agcaggccca ggagaagttt gaactaagtg agaactgttc agagcggcct 1920
gggctgcgag gagctgctgg ggagcaactc agctttgctg ctggactggt gtactcgctg 1980
agcctgctgc aggccacgct acaccgctat gagcatgccc tctctcagtg cagtgtggat 2040
gtgtataaga aagtgggcag cctgtaccct gagatgagtg cccatgagcg ctccttggat 2100
ttcctcattg aactgctgca caaggatcag ctggatgaga ctgtcaatgt ggagcctctc 2160
accaaggcca tcaagtacta tcagcatctg tacagcatcc accttgccga acagcctgag 2220
gactgtacta tgcagctggc tgaccacatt aagttcacgc agagtgctct ggactgcatg 2280
agtgtggagg taggacggct gcgtgccttc ttgcagggtg ggcaggaggc tacagatatt 2340
gccctcctgc tccgggatct ggaaacttca tgcagtgaca tccgccagtt ctgcaagaag 2400
atccgaaggc gaatgccagg gacagatgct cctgggatcc cagctgcact ggcctttgga 2460
ccacaggtat ctgacacgct cctagactgc aggaaacact tgacgtgggt cgtggctgtg 2520
ctgcaggagg tggcagctgc tgctgcccag ctcattgccc cactggcaga gaatgagggg 2580
ctacttgtgg ctgctctgga ggaactggct ttcaaagcaa gcgagcagat ctatgggacc 2640
ccctccagca gcccctatga gtgtctgcgc cagtcatgca acatcctcat cagtaccatg 2700
aacaagctgg ccacagccat gcaggagggg gagtatgatg cagagcggcc ccccagcaag 2760
cctccaccgg ttgaactgcg ggctgctgcc cttcgtgcag agatcacaga tgctgaaggc 2820
ctgggtttga agctcgaaga tcgagagaca gttattaagg agttgaagaa gtcactcaag 2880
attaagggag aggagctaag tgaggccaat gtgcggctga gcctcctgga gaagaagttg 2940
gacagtgctg ccaaggatgc agatgagcgc atcgagaaag tccagactcg gctggaggag 3000
acccaggcac tgctgcgaaa gaaggagaaa gagtttgagg agacaatgga tgcactccag 3060
gctgacatcg accagctgga ggcagagaag gcagaactaa agcagcgtct gaacagccag 3120
tccaaacgca cgattgaggg actccggggc cctcctcctt caggcattgc tactctggtc 3180
tctggcattg ctggtgaaga acagcagcga ggagccatcc ctgggcaggc tccagggtct 3240
gtgccaggcc cagggctggt gaaggactca ccactgctgc ttcagcagat ctctgccatg 3300
aggctgcaca tctcccagct ccagcatgag aacagcatcc tcaagggagc ccagatgaag 3360
gcatccttgg catccctgcc ccctctgcat gttgcaaagc tatcccatga gggccctggc 3420
agtgagttac cagctggagc gctgtatcgt aagaccagcc agctgctgga gacattgaat 3480
caattgagca cacacacgca cgtagtagac atcactcgca ccagccctgc tgccaagagc 3540
ccgtcggccc aacttatgga gcaagtggct cagcttaagt ccctgagtga caccgtcgag 3600
aagctcaagg atgaggtcct caaggagaca gtatctcagc gccctggagc cacagtaccc 3660
actgactttg ccaccttccc ttcatcagcc ttcctcaggg aggatccaaa gtgggaattc 3720
cctcggaaga acttggttct tggaaaaact ctaggagaag gcgaatttgg aaaagtggtc 3780
aaggcaacgg ccttccatct gaaaggcaga gcagggtaca ccacggtggc cgtgaagatg 3840
ctgaaagaga acgcctcccc gagtgagctt cgagacctgc tgtcagagtt caacgtcctg 3900
aagcaggtca accacccaca tgtcatcaaa ttgtatgggg cctgcagcca ggatggcccg 3960
ctcctcctca tcgtggagta cgccaaatac ggctccctgc ggggcttcct ccgcgagagc 4020
cgcaaagtgg ggcctggcta cctgggcagt ggaggcagcc gcaactccag ctccctggac 4080
cacccggatg agcgggccct caccatgggc gacctcatct catttgcctg gcagatctca 4140
caggggatgc agtatctggc cgagatgaag ctcgttcatc gggacttggc agccagaaac 4200
atcctggtag ctgaggggcg gaagatgaag atttcggatt tcggcttgtc ccgagatgtt 4260
tatgaagagg attcctacgt gaagaggagc cagggtcgga ttccagttaa atggatggca 4320
attgaatccc tttttgatca tatctacacc acgcaaagtg atgtatggtc ttttggtgtc 4380
ctgctgtggg agatcgtgac cctaggggga aacccctatc ctgggattcc tcctgagcgg 4440
ctcttcaacc ttctgaagac cggccaccgg atggagaggc cagacaactg cagcgaggag 4500
atgtaccgcc tgatgctgca atgctggaag caggagccgg acaaaaggcc ggtgtttgcg 4560
gacatcagca aagacctgga gaagatgatg gttaagagga gagactactt ggaccttgcg 4620
gcgtccactc catctgactc cctgatttat gacgacggcc tctcagagga ggagacaccg 4680
ctggtggact gtaataatgc ccccctccct cgagccctcc cttccacatg gattgaaaac 4740
aaactctatg gcatgtcaga cccgaactgg cctggagaga gtcctgtacc actcacgaga 4800
gctgatggca ctaacactgg gtttccaaga tatccaaatg atagtgtata tgctaactgg 4860
atgctttcac cctcagcggc aaaattaatg gacacgtttg atagttaa 4908
<210> 2
<211> 1635
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 2
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Arg Gly Leu Lys Pro Lys Lys Ala Pro Thr Ala Arg Lys
130 135 140
Thr Thr Thr Arg Arg Pro Lys Pro Thr Arg Pro Ala Ser Thr Gly Val
145 150 155 160
Ala Gly Ala Ser Ser Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly
165 170 175
Glu Leu Ser Ser Ser Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala
180 185 190
Ala Pro Ile Ile Pro Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro
195 200 205
Pro Leu Pro Ser Pro Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val
210 215 220
Arg Asp Leu Glu Glu Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu
225 230 235 240
Asp Lys Ala Lys Leu Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu
245 250 255
Gln Val Gln Glu Trp Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu
260 265 270
Gln Arg Arg Leu Lys Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu
275 280 285
Ala Lys Glu Arg Tyr Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile
290 295 300
Glu Met Ala Thr Leu Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser
305 310 315 320
Leu Gln Gln Glu Val Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr
325 330 335
Thr Asp Leu Glu Ile Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp
340 345 350
Gly Ala Ala Ser Ser Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala
355 360 365
Arg Leu Lys Asp Ala Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu
370 375 380
Lys Gln Glu His Val Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln
385 390 395 400
Glu Leu Glu Val Val Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu
405 410 415
Ser Gln Ala Glu Ser Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala
420 425 430
Ala Leu Gly Ala Glu Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu
435 440 445
Asn Leu Glu Glu Lys Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu
450 455 460
Glu Ala Met Asn Glu Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu
465 470 475 480
Thr Glu Leu Glu Leu Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val
485 490 495
Arg Glu Ala Gln Lys Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp
500 505 510
Tyr Gln Gln Thr Ile Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln
515 520 525
Asp Val Asn Arg Glu Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg
530 535 540
Gln Gln Gln Pro Pro Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala
545 550 555 560
Glu Thr Lys Ala His Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met
565 570 575
Glu Val Ala Gln Ala Asn Arg His Met Ser Leu Leu Thr Ala Phe Met
580 585 590
Pro Asp Ser Phe Leu Arg Pro Gly Gly Asp His Asp Cys Val Leu Val
595 600 605
Leu Leu Leu Met Pro Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys
610 615 620
Gln Ala Gln Glu Lys Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro
625 630 635 640
Gly Leu Arg Gly Ala Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu
645 650 655
Val Tyr Ser Leu Ser Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His
660 665 670
Ala Leu Ser Gln Cys Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu
675 680 685
Tyr Pro Glu Met Ser Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu
690 695 700
Leu Leu His Lys Asp Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu
705 710 715 720
Thr Lys Ala Ile Lys Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala
725 730 735
Glu Gln Pro Glu Asp Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe
740 745 750
Thr Gln Ser Ala Leu Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg
755 760 765
Ala Phe Leu Gln Gly Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu
770 775 780
Arg Asp Leu Glu Thr Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys
785 790 795 800
Ile Arg Arg Arg Met Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala
805 810 815
Leu Ala Phe Gly Pro Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys
820 825 830
His Leu Thr Trp Val Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala
835 840 845
Ala Gln Leu Ile Ala Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala
850 855 860
Ala Leu Glu Glu Leu Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr
865 870 875 880
Pro Ser Ser Ser Pro Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu
885 890 895
Ile Ser Thr Met Asn Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr
900 905 910
Asp Ala Glu Arg Pro Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala
915 920 925
Ala Ala Leu Arg Ala Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys
930 935 940
Leu Glu Asp Arg Glu Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys
945 950 955 960
Ile Lys Gly Glu Glu Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu
965 970 975
Glu Lys Lys Leu Asp Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu
980 985 990
Lys Val Gln Thr Arg Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys
995 1000 1005
Glu Lys Glu Phe Glu Glu Thr Met Asp Ala Leu Gln Ala Asp Ile
1010 1015 1020
Asp Gln Leu Glu Ala Glu Lys Ala Glu Leu Lys Gln Arg Leu Asn
1025 1030 1035
Ser Gln Ser Lys Arg Thr Ile Glu Gly Leu Arg Gly Pro Pro Pro
1040 1045 1050
Ser Gly Ile Ala Thr Leu Val Ser Gly Ile Ala Gly Glu Glu Gln
1055 1060 1065
Gln Arg Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser Val Pro Gly
1070 1075 1080
Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln Gln Ile Ser
1085 1090 1095
Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu Asn Ser Ile
1100 1105 1110
Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu Pro Pro
1115 1120 1125
Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu Leu
1130 1135 1140
Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu Thr
1145 1150 1155
Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp Ile Thr Arg
1160 1165 1170
Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu Met Glu Gln
1175 1180 1185
Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu Lys Leu Lys
1190 1195 1200
Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro Gly Ala Thr
1205 1210 1215
Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu Arg
1220 1225 1230
Glu Asp Pro Lys Trp Glu Phe Pro Arg Lys Asn Leu Val Leu Gly
1235 1240 1245
Lys Thr Leu Gly Glu Gly Glu Phe Gly Lys Val Val Lys Ala Thr
1250 1255 1260
Ala Phe His Leu Lys Gly Arg Ala Gly Tyr Thr Thr Val Ala Val
1265 1270 1275
Lys Met Leu Lys Glu Asn Ala Ser Pro Ser Glu Leu Arg Asp Leu
1280 1285 1290
Leu Ser Glu Phe Asn Val Leu Lys Gln Val Asn His Pro His Val
1295 1300 1305
Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp Gly Pro Leu Leu Leu
1310 1315 1320
Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu Arg Gly Phe Leu Arg
1325 1330 1335
Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu Gly Ser Gly Gly Ser
1340 1345 1350
Arg Asn Ser Ser Ser Leu Asp His Pro Asp Glu Arg Ala Leu Thr
1355 1360 1365
Met Gly Asp Leu Ile Ser Phe Ala Trp Gln Ile Ser Gln Gly Met
1370 1375 1380
Gln Tyr Leu Ala Glu Met Lys Leu Val His Arg Asp Leu Ala Ala
1385 1390 1395
Arg Asn Ile Leu Val Ala Glu Gly Arg Lys Met Lys Ile Ser Asp
1400 1405 1410
Phe Gly Leu Ser Arg Asp Val Tyr Glu Glu Asp Ser Tyr Val Lys
1415 1420 1425
Arg Ser Gln Gly Arg Ile Pro Val Lys Trp Met Ala Ile Glu Ser
1430 1435 1440
Leu Phe Asp His Ile Tyr Thr Thr Gln Ser Asp Val Trp Ser Phe
1445 1450 1455
Gly Val Leu Leu Trp Glu Ile Val Thr Leu Gly Gly Asn Pro Tyr
1460 1465 1470
Pro Gly Ile Pro Pro Glu Arg Leu Phe Asn Leu Leu Lys Thr Gly
1475 1480 1485
His Arg Met Glu Arg Pro Asp Asn Cys Ser Glu Glu Met Tyr Arg
1490 1495 1500
Leu Met Leu Gln Cys Trp Lys Gln Glu Pro Asp Lys Arg Pro Val
1505 1510 1515
Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys Met Met Val Lys Arg
1520 1525 1530
Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr Pro Ser Asp Ser Leu
1535 1540 1545
Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu Thr Pro Leu Val Asp
1550 1555 1560
Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu Pro Ser Thr Trp Ile
1565 1570 1575
Glu Asn Lys Leu Tyr Gly Met Ser Asp Pro Asn Trp Pro Gly Glu
1580 1585 1590
Ser Pro Val Pro Leu Thr Arg Ala Asp Gly Thr Asn Thr Gly Phe
1595 1600 1605
Pro Arg Tyr Pro Asn Asp Ser Val Tyr Ala Asn Trp Met Leu Ser
1610 1615 1620
Pro Ser Ala Ala Lys Leu Met Asp Thr Phe Asp Ser
1625 1630 1635
<210> 3
<211> 4782
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 3
atggcacaga gcaagaggca cgtgtacagc cggacgccca gcggcagcag gatgagtgcg 60
gaggcaagcg cccggcctct gcgggtgggc tcccgtgtag aggtgattgg aaaaggccac 120
cgaggcactg tggcctatgt tggagccaca ctgtttgcca ctggcaaatg ggtaggcgtg 180
attctggatg aagcaaaggg caaaaatgat ggaactgttc aaggcaggaa gtacttcact 240
tgtgatgaag ggcatggcat ctttgtgcgc cagtcccaga tccaggtatt tgaagatgga 300
gcagatacta cttccccaga gacacctgat tcttctgctt caaaagtcct caaaagagag 360
ggaactgata caactgcaaa gactagcaaa ctgcggggac tgaagcctaa gaaggcaccg 420
acagcccgaa agaccacaac tcggcgaccc aagcccacgc gcccagccag tactggggtg 480
gctggggcca gtagctccct gggcccctct ggctcagcgt cagcaggtga gctgagcagc 540
agtgagccca gcaccccggc tcagactccg ctggcagcac ccatcatccc cacgccggtc 600
ctcacctctc ctggagcagt ccccccgctt ccttccccat ccaaggagga ggagggacta 660
agggctcagg tgcgggacct ggaggagaaa ctagagaccc tgagactgaa acgggcagaa 720
gacaaagcaa agctaaaaga gctggagaaa cacaaaatcc agctggagca ggtgcaggaa 780
tggaagagca aaatgcagga gcagcaggcc gacctgcagc ggcgcctcaa ggaggcgaga 840
aaggaagcca aggaggcgct ggaggcaaag gaacgctata tggaggagat ggctgatact 900
gctgatgcca ttgagatggc cactttggac aaggagatgg ctgaagagcg ggctgagtcc 960
ctgcagcagg aggtggaggc actgaaggag cgggtggacg agctcactac tgacttagag 1020
atcctcaagg ctgagattga agagaagggc tcagatggcg ctgcatccag ttatcagctc 1080
aagcagcttg aggagcagaa tgcccgcctg aaggatgccc tggtgaggat gcgggatctt 1140
tcttcctcag agaagcagga gcatgtgaag ctccagaagc tcatggaaaa gaagaaccaa 1200
gagctggaag ttgtgaggca acagcgggag cgtctgcagg aggagctaag ccaggcagag 1260
agcaccattg atgagctcaa ggagcaggtg gatgctgctc tgggtgctga ggagatggtg 1320
gagatgctga cagatcggaa cctgaatctg gaagagaaag tgcgcgagtt gagggagact 1380
gtgggagact tggaagcgat gaatgagatg aacgatgagc tgcaggagaa tgcacgtgag 1440
acagaactgg agctgcggga gcagctggac atggcaggcg cgcgggttcg tgaggcccag 1500
aagcgtgtgg aggcagccca ggagacggtt gcagactacc agcagaccat caagaagtac 1560
cgccagctga ccgcccatct acaggatgtg aatcgggaac tgacaaacca gcaggaagca 1620
tctgtggaga ggcaacagca gccacctcca gagacctttg acttcaaaat caagtttgct 1680
gagactaagg cccatgccaa ggcaattgag atggaattga ggcagatgga ggtggcccag 1740
gccaatcgac acatgtccct gctgacagcc ttcatgcctg acagcttcct tcggccaggt 1800
ggggaccatg actgcgttct ggtgctgttg ctcatgcctc gtctcatttg caaggcagag 1860
ctgatccgga agcaggccca ggagaagttt gaactaagtg agaactgttc agagcggcct 1920
gggctgcgag gagctgctgg ggagcaactc agctttgctg ctggactggt gtactcgctg 1980
agcctgctgc aggccacgct acaccgctat gagcatgccc tctctcagtg cagtgtggat 2040
gtgtataaga aagtgggcag cctgtaccct gagatgagtg cccatgagcg ctccttggat 2100
ttcctcattg aactgctgca caaggatcag ctggatgaga ctgtcaatgt ggagcctctc 2160
accaaggcca tcaagtacta tcagcatctg tacagcatcc accttgccga acagcctgag 2220
gactgtacta tgcagctggc tgaccacatt aagttcacgc agagtgctct ggactgcatg 2280
agtgtggagg taggacggct gcgtgccttc ttgcagggtg ggcaggaggc tacagatatt 2340
gccctcctgc tccgggatct ggaaacttca tgcagtgaca tccgccagtt ctgcaagaag 2400
atccgaaggc gaatgccagg gacagatgct cctgggatcc cagctgcact ggcctttgga 2460
ccacaggtat ctgacacgct cctagactgc aggaaacact tgacgtgggt cgtggctgtg 2520
ctgcaggagg tggcagctgc tgctgcccag ctcattgccc cactggcaga gaatgagggg 2580
ctacttgtgg ctgctctgga ggaactggct ttcaaagcaa gcgagcagat ctatgggacc 2640
ccctccagca gcccctatga gtgtctgcgc cagtcatgca acatcctcat cagtaccatg 2700
aacaagctgg ccacagccat gcaggagggg gagtatgatg cagagcggcc ccccagcaag 2760
cctccaccgg ttgaactgcg ggctgctgcc cttcgtgcag agatcacaga tgctgaaggc 2820
ctgggtttga agctcgaaga tcgagagaca gttattaagg agttgaagaa gtcactcaag 2880
attaagggag aggagctaag tgaggccaat gtgcggctga gcctcctgga gaagaagttg 2940
gacagtgctg ccaaggatgc agatgagcgc atcgagaaag tccagactcg gctggaggag 3000
acccaggcac tgctgcgaaa gaaggagaaa gagtttgagg agacaatgga tgcactccag 3060
gctgacatcg accagctgga ggcagagaag gcagaactaa agcagcgtct gaacagccag 3120
tccaaacgca cgattgaggg actccggggc cctcctcctt caggcattgc tactctggtc 3180
tctggcattg ctggtgaaga acagcagcga ggagccatcc ctgggcaggc tccagggtct 3240
gtgccaggcc cagggctggt gaaggactca ccactgctgc ttcagcagat ctctgccatg 3300
aggctgcaca tctcccagct ccagcatgag aacagcatcc tcaagggagc ccagatgaag 3360
gcatccttgg catccctgcc ccctctgcat gttgcaaagc tatcccatga gggccctggc 3420
agtgagttac cagctggagc gctgtatcgt aagaccagcc agctgctgga gacattgaat 3480
caattgagca cacacacgca cgtagtagac atcactcgca ccagccctgc tgccaagagc 3540
ccgtcggccc aacttatgga gcaagtggct cagcttaagt ccctgagtga caccgtcgag 3600
aagctcaagg atgaggtcct caaggagaca gtatctcagc gccctggagc cacagtaccc 3660
actgactttg ccaccttccc ttcatcagcc ttcctcaggg aggatccaaa gtgggaattc 3720
cctcggaaga acttggttct tggaaaaact ctaggagaag gcgaatttgg aaaagtggtc 3780
aaggcaacgg ccttccatct gaaaggcaga gcagggtaca ccacggtggc cgtgaagatg 3840
ctgaaagaga acgcctcccc gagtgagctt cgagacctgc tgtcagagtt caacgtcctg 3900
aagcaggtca accacccaca tgtcatcaaa ttgtatgggg cctgcagcca ggatggcccg 3960
ctcctcctca tcgtggagta cgccaaatac ggctccctgc ggggcttcct ccgcgagagc 4020
cgcaaagtgg ggcctggcta cctgggcagt ggaggcagcc gcaactccag ctccctggac 4080
cacccggatg agcgggccct caccatgggc gacctcatct catttgcctg gcagatctca 4140
caggggatgc agtatctggc cgagatgaag ctcgttcatc gggacttggc agccagaaac 4200
atcctggtag ctgaggggcg gaagatgaag atttcggatt tcggcttgtc ccgagatgtt 4260
tatgaagagg attcctacgt gaagaggagc cagggtcgga ttccagttaa atggatggca 4320
attgaatccc tttttgatca tatctacacc acgcaaagtg atgtatggtc ttttggtgtc 4380
ctgctgtggg agatcgtgac cctaggggga aacccctatc ctgggattcc tcctgagcgg 4440
ctcttcaacc ttctgaagac cggccaccgg atggagaggc cagacaactg cagcgaggag 4500
atgtaccgcc tgatgctgca atgctggaag caggagccgg acaaaaggcc ggtgtttgcg 4560
gacatcagca aagacctgga gaagatgatg gttaagagga gagactactt ggaccttgcg 4620
gcgtccactc catctgactc cctgatttat gacgacggcc tctcagagga ggagacaccg 4680
ctggtggact gtaataatgc ccccctccct cgagccctcc cttccacatg gattgaaaac 4740
aaactctatg gtagaatttc ccatgcattt actagattct ag 4782
<210> 4
<211> 1593
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 4
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Arg Gly Leu Lys Pro Lys Lys Ala Pro Thr Ala Arg Lys
130 135 140
Thr Thr Thr Arg Arg Pro Lys Pro Thr Arg Pro Ala Ser Thr Gly Val
145 150 155 160
Ala Gly Ala Ser Ser Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly
165 170 175
Glu Leu Ser Ser Ser Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala
180 185 190
Ala Pro Ile Ile Pro Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro
195 200 205
Pro Leu Pro Ser Pro Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val
210 215 220
Arg Asp Leu Glu Glu Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu
225 230 235 240
Asp Lys Ala Lys Leu Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu
245 250 255
Gln Val Gln Glu Trp Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu
260 265 270
Gln Arg Arg Leu Lys Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu
275 280 285
Ala Lys Glu Arg Tyr Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile
290 295 300
Glu Met Ala Thr Leu Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser
305 310 315 320
Leu Gln Gln Glu Val Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr
325 330 335
Thr Asp Leu Glu Ile Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp
340 345 350
Gly Ala Ala Ser Ser Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala
355 360 365
Arg Leu Lys Asp Ala Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu
370 375 380
Lys Gln Glu His Val Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln
385 390 395 400
Glu Leu Glu Val Val Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu
405 410 415
Ser Gln Ala Glu Ser Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala
420 425 430
Ala Leu Gly Ala Glu Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu
435 440 445
Asn Leu Glu Glu Lys Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu
450 455 460
Glu Ala Met Asn Glu Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu
465 470 475 480
Thr Glu Leu Glu Leu Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val
485 490 495
Arg Glu Ala Gln Lys Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp
500 505 510
Tyr Gln Gln Thr Ile Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln
515 520 525
Asp Val Asn Arg Glu Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg
530 535 540
Gln Gln Gln Pro Pro Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala
545 550 555 560
Glu Thr Lys Ala His Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met
565 570 575
Glu Val Ala Gln Ala Asn Arg His Met Ser Leu Leu Thr Ala Phe Met
580 585 590
Pro Asp Ser Phe Leu Arg Pro Gly Gly Asp His Asp Cys Val Leu Val
595 600 605
Leu Leu Leu Met Pro Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys
610 615 620
Gln Ala Gln Glu Lys Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro
625 630 635 640
Gly Leu Arg Gly Ala Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu
645 650 655
Val Tyr Ser Leu Ser Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His
660 665 670
Ala Leu Ser Gln Cys Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu
675 680 685
Tyr Pro Glu Met Ser Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu
690 695 700
Leu Leu His Lys Asp Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu
705 710 715 720
Thr Lys Ala Ile Lys Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala
725 730 735
Glu Gln Pro Glu Asp Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe
740 745 750
Thr Gln Ser Ala Leu Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg
755 760 765
Ala Phe Leu Gln Gly Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu
770 775 780
Arg Asp Leu Glu Thr Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys
785 790 795 800
Ile Arg Arg Arg Met Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala
805 810 815
Leu Ala Phe Gly Pro Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys
820 825 830
His Leu Thr Trp Val Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala
835 840 845
Ala Gln Leu Ile Ala Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala
850 855 860
Ala Leu Glu Glu Leu Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr
865 870 875 880
Pro Ser Ser Ser Pro Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu
885 890 895
Ile Ser Thr Met Asn Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr
900 905 910
Asp Ala Glu Arg Pro Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala
915 920 925
Ala Ala Leu Arg Ala Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys
930 935 940
Leu Glu Asp Arg Glu Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys
945 950 955 960
Ile Lys Gly Glu Glu Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu
965 970 975
Glu Lys Lys Leu Asp Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu
980 985 990
Lys Val Gln Thr Arg Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys
995 1000 1005
Glu Lys Glu Phe Glu Glu Thr Met Asp Ala Leu Gln Ala Asp Ile
1010 1015 1020
Asp Gln Leu Glu Ala Glu Lys Ala Glu Leu Lys Gln Arg Leu Asn
1025 1030 1035
Ser Gln Ser Lys Arg Thr Ile Glu Gly Leu Arg Gly Pro Pro Pro
1040 1045 1050
Ser Gly Ile Ala Thr Leu Val Ser Gly Ile Ala Gly Glu Glu Gln
1055 1060 1065
Gln Arg Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser Val Pro Gly
1070 1075 1080
Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln Gln Ile Ser
1085 1090 1095
Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu Asn Ser Ile
1100 1105 1110
Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu Pro Pro
1115 1120 1125
Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu Leu
1130 1135 1140
Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu Thr
1145 1150 1155
Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp Ile Thr Arg
1160 1165 1170
Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu Met Glu Gln
1175 1180 1185
Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu Lys Leu Lys
1190 1195 1200
Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro Gly Ala Thr
1205 1210 1215
Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu Arg
1220 1225 1230
Glu Asp Pro Lys Trp Glu Phe Pro Arg Lys Asn Leu Val Leu Gly
1235 1240 1245
Lys Thr Leu Gly Glu Gly Glu Phe Gly Lys Val Val Lys Ala Thr
1250 1255 1260
Ala Phe His Leu Lys Gly Arg Ala Gly Tyr Thr Thr Val Ala Val
1265 1270 1275
Lys Met Leu Lys Glu Asn Ala Ser Pro Ser Glu Leu Arg Asp Leu
1280 1285 1290
Leu Ser Glu Phe Asn Val Leu Lys Gln Val Asn His Pro His Val
1295 1300 1305
Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp Gly Pro Leu Leu Leu
1310 1315 1320
Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu Arg Gly Phe Leu Arg
1325 1330 1335
Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu Gly Ser Gly Gly Ser
1340 1345 1350
Arg Asn Ser Ser Ser Leu Asp His Pro Asp Glu Arg Ala Leu Thr
1355 1360 1365
Met Gly Asp Leu Ile Ser Phe Ala Trp Gln Ile Ser Gln Gly Met
1370 1375 1380
Gln Tyr Leu Ala Glu Met Lys Leu Val His Arg Asp Leu Ala Ala
1385 1390 1395
Arg Asn Ile Leu Val Ala Glu Gly Arg Lys Met Lys Ile Ser Asp
1400 1405 1410
Phe Gly Leu Ser Arg Asp Val Tyr Glu Glu Asp Ser Tyr Val Lys
1415 1420 1425
Arg Ser Gln Gly Arg Ile Pro Val Lys Trp Met Ala Ile Glu Ser
1430 1435 1440
Leu Phe Asp His Ile Tyr Thr Thr Gln Ser Asp Val Trp Ser Phe
1445 1450 1455
Gly Val Leu Leu Trp Glu Ile Val Thr Leu Gly Gly Asn Pro Tyr
1460 1465 1470
Pro Gly Ile Pro Pro Glu Arg Leu Phe Asn Leu Leu Lys Thr Gly
1475 1480 1485
His Arg Met Glu Arg Pro Asp Asn Cys Ser Glu Glu Met Tyr Arg
1490 1495 1500
Leu Met Leu Gln Cys Trp Lys Gln Glu Pro Asp Lys Arg Pro Val
1505 1510 1515
Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys Met Met Val Lys Arg
1520 1525 1530
Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr Pro Ser Asp Ser Leu
1535 1540 1545
Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu Thr Pro Leu Val Asp
1550 1555 1560
Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu Pro Ser Thr Trp Ile
1565 1570 1575
Glu Asn Lys Leu Tyr Gly Arg Ile Ser His Ala Phe Thr Arg Phe
1580 1585 1590
<210> 5
<211> 4506
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 5
atgatgagac aggcaccgac agcccgaaag accacaactc ggcgacccaa gcccacgcgc 60
ccagccagta ctggggtggc tggggccagt agctccctgg gcccctctgg ctcagcgtca 120
gcaggtgagc tgagcagcag tgagcccagc accccggctc agactccgct ggcagcaccc 180
atcatcccca cgccggtcct cacctctcct ggagcagtcc ccccgcttcc ttccccatcc 240
aaggaggagg agggactaag ggctcaggtg cgggacctgg aggagaaact agagaccctg 300
agactgaaac gggcagaaga caaagcaaag ctaaaagagc tggagaaaca caaaatccag 360
ctggagcagg tgcaggaatg gaagagcaaa atgcaggagc agcaggccga cctgcagcgg 420
cgcctcaagg aggcgagaaa ggaagccaag gaggcgctgg aggcaaagga acgctatatg 480
gaggagatgg ctgatactgc tgatgccatt gagatggcca ctttggacaa ggagatggct 540
gaagagcggg ctgagtccct gcagcaggag gtggaggcac tgaaggagcg ggtggacgag 600
ctcactactg acttagagat cctcaaggct gagattgaag agaagggctc agatggcgct 660
gcatccagtt atcagctcaa gcagcttgag gagcagaatg cccgcctgaa ggatgccctg 720
gtgaggatgc gggatctttc ttcctcagag aagcaggagc atgtgaagct ccagaagctc 780
atggaaaaga agaaccaaga gctggaagtt gtgaggcaac agcgggagcg tctgcaggag 840
gagctaagcc aggcagagag caccattgat gagctcaagg agcaggtgga tgctgctctg 900
ggtgctgagg agatggtgga gatgctgaca gatcggaacc tgaatctgga agagaaagtg 960
cgcgagttga gggagactgt gggagacttg gaagcgatga atgagatgaa cgatgagctg 1020
caggagaatg cacgtgagac agaactggag ctgcgggagc agctggacat ggcaggcgcg 1080
cgggttcgtg aggcccagaa gcgtgtggag gcagcccagg agacggttgc agactaccag 1140
cagaccatca agaagtaccg ccagctgacc gcccatctac aggatgtgaa tcgggaactg 1200
acaaaccagc aggaagcatc tgtggagagg caacagcagc cacctccaga gacctttgac 1260
ttcaaaatca agtttgctga gactaaggcc catgccaagg caattgagat ggaattgagg 1320
cagatggagg tggcccaggc caatcgacac atgtccctgc tgacagcctt catgcctgac 1380
agcttccttc ggccaggtgg ggaccatgac tgcgttctgg tgctgttgct catgcctcgt 1440
ctcatttgca aggcagagct gatccggaag caggcccagg agaagtttga actaagtgag 1500
aactgttcag agcggcctgg gctgcgagga gctgctgggg agcaactcag ctttgctgct 1560
ggactggtgt actcgctgag cctgctgcag gccacgctac accgctatga gcatgccctc 1620
tctcagtgca gtgtggatgt gtataagaaa gtgggcagcc tgtaccctga gatgagtgcc 1680
catgagcgct ccttggattt cctcattgaa ctgctgcaca aggatcagct ggatgagact 1740
gtcaatgtgg agcctctcac caaggccatc aagtactatc agcatctgta cagcatccac 1800
cttgccgaac agcctgagga ctgtactatg cagctggctg accacattaa gttcacgcag 1860
agtgctctgg actgcatgag tgtggaggta ggacggctgc gtgccttctt gcagggtggg 1920
caggaggcta cagatattgc cctcctgctc cgggatctgg aaacttcatg cagtgacatc 1980
cgccagttct gcaagaagat ccgaaggcga atgccaggga cagatgctcc tgggatccca 2040
gctgcactgg cctttggacc acaggtatct gacacgctcc tagactgcag gaaacacttg 2100
acgtgggtcg tggctgtgct gcaggaggtg gcagctgctg ctgcccagct cattgcccca 2160
ctggcagaga atgaggggct acttgtggct gctctggagg aactggcttt caaagcaagc 2220
gagcagatct atgggacccc ctccagcagc ccctatgagt gtctgcgcca gtcatgcaac 2280
atcctcatca gtaccatgaa caagctggcc acagccatgc aggaggggga gtatgatgca 2340
gagcggcccc ccagcaagcc tccaccggtt gaactgcggg ctgctgccct tcgtgcagag 2400
atcacagatg ctgaaggcct gggtttgaag ctcgaagatc gagagacagt tattaaggag 2460
ttgaagaagt cactcaagat taagggagag gagctaagtg aggccaatgt gcggctgagc 2520
ctcctggaga agaagttgga cagtgctgcc aaggatgcag atgagcgcat cgagaaagtc 2580
cagactcggc tggaggagac ccaggcactg ctgcgaaaga aggagaaaga gtttgaggag 2640
acaatggatg cactccaggc tgacatcgac cagctggagg cagagaaggc agaactaaag 2700
cagcgtctga acagccagtc caaacgcacg attgagggac tccggggccc tcctccttca 2760
ggcattgcta ctctggtctc tggcattgct ggtgaagaac agcagcgagg agccatccct 2820
gggcaggctc cagggtctgt gccaggccca gggctggtga aggactcacc actgctgctt 2880
cagcagatct ctgccatgag gctgcacatc tcccagctcc agcatgagaa cagcatcctc 2940
aagggagccc agatgaaggc atccttggca tccctgcccc ctctgcatgt tgcaaagcta 3000
tcccatgagg gccctggcag tgagttacca gctggagcgc tgtatcgtaa gaccagccag 3060
ctgctggaga cattgaatca attgagcaca cacacgcacg tagtagacat cactcgcacc 3120
agccctgctg ccaagagccc gtcggcccaa cttatggagc aagtggctca gcttaagtcc 3180
ctgagtgaca ccgtcgagaa gctcaaggat gaggtcctca aggagacagt atctcagcgc 3240
cctggagcca cagtacccac tgactttgcc accttccctt catcagcctt cctcagggag 3300
gatccaaagt gggaattccc tcggaagaac ttggttcttg gaaaaactct aggagaaggc 3360
gaatttggaa aagtggtcaa ggcaacggcc ttccatctga aaggcagagc agggtacacc 3420
acggtggccg tgaagatgct gaaagagaac gcctccccga gtgagcttcg agacctgctg 3480
tcagagttca acgtcctgaa gcaggtcaac cacccacatg tcatcaaatt gtatggggcc 3540
tgcagccagg atggcccgct cctcctcatc gtggagtacg ccaaatacgg ctccctgcgg 3600
ggcttcctcc gcgagagccg caaagtgggg cctggctacc tgggcagtgg aggcagccgc 3660
aactccagct ccctggacca cccggatgag cgggccctca ccatgggcga cctcatctca 3720
tttgcctggc agatctcaca ggggatgcag tatctggccg agatgaagct cgttcatcgg 3780
gacttggcag ccagaaacat cctggtagct gaggggcgga agatgaagat ttcggatttc 3840
ggcttgtccc gagatgttta tgaagaggat tcctacgtga agaggagcca gggtcggatt 3900
ccagttaaat ggatggcaat tgaatccctt tttgatcata tctacaccac gcaaagtgat 3960
gtatggtctt ttggtgtcct gctgtgggag atcgtgaccc tagggggaaa cccctatcct 4020
gggattcctc ctgagcggct cttcaacctt ctgaagaccg gccaccggat ggagaggcca 4080
gacaactgca gcgaggagat gtaccgcctg atgctgcaat gctggaagca ggagccggac 4140
aaaaggccgg tgtttgcgga catcagcaaa gacctggaga agatgatggt taagaggaga 4200
gactacttgg accttgcggc gtccactcca tctgactccc tgatttatga cgacggcctc 4260
tcagaggagg agacaccgct ggtggactgt aataatgccc ccctccctcg agccctccct 4320
tccacatgga ttgaaaacaa actctatggc atgtcagacc cgaactggcc tggagagagt 4380
cctgtaccac tcacgagagc tgatggcact aacactgggt ttccaagata tccaaatgat 4440
agtgtatatg ctaactggat gctttcaccc tcagcggcaa aattaatgga cacgtttgat 4500
agttaa 4506
<210> 6
<211> 1501
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 6
Met Met Arg Gln Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro
1 5 10 15
Lys Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser
20 25 30
Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu
35 40 45
Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr
50 55 60
Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser
65 70 75 80
Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys
85 90 95
Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys
100 105 110
Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys
115 120 125
Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu
130 135 140
Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met
145 150 155 160
Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp
165 170 175
Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu
180 185 190
Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu
195 200 205
Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr
210 215 220
Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu
225 230 235 240
Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys
245 250 255
Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg
260 265 270
Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr
275 280 285
Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu
290 295 300
Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val
305 310 315 320
Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met
325 330 335
Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg
340 345 350
Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg
355 360 365
Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys
370 375 380
Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu
385 390 395 400
Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro
405 410 415
Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala
420 425 430
Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn
435 440 445
Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg
450 455 460
Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg
465 470 475 480
Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe
485 490 495
Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala
500 505 510
Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu
515 520 525
Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser
530 535 540
Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala
545 550 555 560
His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln
565 570 575
Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr
580 585 590
Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys
595 600 605
Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp
610 615 620
Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly
625 630 635 640
Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser
645 650 655
Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro
660 665 670
Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln
675 680 685
Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val
690 695 700
Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro
705 710 715 720
Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala
725 730 735
Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr
740 745 750
Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys
755 760 765
Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro
770 775 780
Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu
785 790 795 800
Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr
805 810 815
Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu
820 825 830
Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser
835 840 845
Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu
850 855 860
Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu
865 870 875 880
Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
885 890 895
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile Glu
900 905 910
Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val Ser Gly
915 920 925
Ile Ala Gly Glu Glu Gln Gln Arg Gly Ala Ile Pro Gly Gln Ala Pro
930 935 940
Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu
945 950 955 960
Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu
965 970 975
Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu
980 985 990
Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu
995 1000 1005
Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu
1010 1015 1020
Thr Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp Ile Thr
1025 1030 1035
Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu Met Glu
1040 1045 1050
Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu Lys Leu
1055 1060 1065
Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro Gly Ala
1070 1075 1080
Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu
1085 1090 1095
Arg Glu Asp Pro Lys Trp Glu Phe Pro Arg Lys Asn Leu Val Leu
1100 1105 1110
Gly Lys Thr Leu Gly Glu Gly Glu Phe Gly Lys Val Val Lys Ala
1115 1120 1125
Thr Ala Phe His Leu Lys Gly Arg Ala Gly Tyr Thr Thr Val Ala
1130 1135 1140
Val Lys Met Leu Lys Glu Asn Ala Ser Pro Ser Glu Leu Arg Asp
1145 1150 1155
Leu Leu Ser Glu Phe Asn Val Leu Lys Gln Val Asn His Pro His
1160 1165 1170
Val Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp Gly Pro Leu Leu
1175 1180 1185
Leu Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu Arg Gly Phe Leu
1190 1195 1200
Arg Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu Gly Ser Gly Gly
1205 1210 1215
Ser Arg Asn Ser Ser Ser Leu Asp His Pro Asp Glu Arg Ala Leu
1220 1225 1230
Thr Met Gly Asp Leu Ile Ser Phe Ala Trp Gln Ile Ser Gln Gly
1235 1240 1245
Met Gln Tyr Leu Ala Glu Met Lys Leu Val His Arg Asp Leu Ala
1250 1255 1260
Ala Arg Asn Ile Leu Val Ala Glu Gly Arg Lys Met Lys Ile Ser
1265 1270 1275
Asp Phe Gly Leu Ser Arg Asp Val Tyr Glu Glu Asp Ser Tyr Val
1280 1285 1290
Lys Arg Ser Gln Gly Arg Ile Pro Val Lys Trp Met Ala Ile Glu
1295 1300 1305
Ser Leu Phe Asp His Ile Tyr Thr Thr Gln Ser Asp Val Trp Ser
1310 1315 1320
Phe Gly Val Leu Leu Trp Glu Ile Val Thr Leu Gly Gly Asn Pro
1325 1330 1335
Tyr Pro Gly Ile Pro Pro Glu Arg Leu Phe Asn Leu Leu Lys Thr
1340 1345 1350
Gly His Arg Met Glu Arg Pro Asp Asn Cys Ser Glu Glu Met Tyr
1355 1360 1365
Arg Leu Met Leu Gln Cys Trp Lys Gln Glu Pro Asp Lys Arg Pro
1370 1375 1380
Val Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys Met Met Val Lys
1385 1390 1395
Arg Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr Pro Ser Asp Ser
1400 1405 1410
Leu Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu Thr Pro Leu Val
1415 1420 1425
Asp Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu Pro Ser Thr Trp
1430 1435 1440
Ile Glu Asn Lys Leu Tyr Gly Met Ser Asp Pro Asn Trp Pro Gly
1445 1450 1455
Glu Ser Pro Val Pro Leu Thr Arg Ala Asp Gly Thr Asn Thr Gly
1460 1465 1470
Phe Pro Arg Tyr Pro Asn Asp Ser Val Tyr Ala Asn Trp Met Leu
1475 1480 1485
Ser Pro Ser Ala Ala Lys Leu Met Asp Thr Phe Asp Ser
1490 1495 1500
<210> 7
<211> 4380
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 7
atgatgagac aggcaccgac agcccgaaag accacaactc ggcgacccaa gcccacgcgc 60
ccagccagta ctggggtggc tggggccagt agctccctgg gcccctctgg ctcagcgtca 120
gcaggtgagc tgagcagcag tgagcccagc accccggctc agactccgct ggcagcaccc 180
atcatcccca cgccggtcct cacctctcct ggagcagtcc ccccgcttcc ttccccatcc 240
aaggaggagg agggactaag ggctcaggtg cgggacctgg aggagaaact agagaccctg 300
agactgaaac gggcagaaga caaagcaaag ctaaaagagc tggagaaaca caaaatccag 360
ctggagcagg tgcaggaatg gaagagcaaa atgcaggagc agcaggccga cctgcagcgg 420
cgcctcaagg aggcgagaaa ggaagccaag gaggcgctgg aggcaaagga acgctatatg 480
gaggagatgg ctgatactgc tgatgccatt gagatggcca ctttggacaa ggagatggct 540
gaagagcggg ctgagtccct gcagcaggag gtggaggcac tgaaggagcg ggtggacgag 600
ctcactactg acttagagat cctcaaggct gagattgaag agaagggctc agatggcgct 660
gcatccagtt atcagctcaa gcagcttgag gagcagaatg cccgcctgaa ggatgccctg 720
gtgaggatgc gggatctttc ttcctcagag aagcaggagc atgtgaagct ccagaagctc 780
atggaaaaga agaaccaaga gctggaagtt gtgaggcaac agcgggagcg tctgcaggag 840
gagctaagcc aggcagagag caccattgat gagctcaagg agcaggtgga tgctgctctg 900
ggtgctgagg agatggtgga gatgctgaca gatcggaacc tgaatctgga agagaaagtg 960
cgcgagttga gggagactgt gggagacttg gaagcgatga atgagatgaa cgatgagctg 1020
caggagaatg cacgtgagac agaactggag ctgcgggagc agctggacat ggcaggcgcg 1080
cgggttcgtg aggcccagaa gcgtgtggag gcagcccagg agacggttgc agactaccag 1140
cagaccatca agaagtaccg ccagctgacc gcccatctac aggatgtgaa tcgggaactg 1200
acaaaccagc aggaagcatc tgtggagagg caacagcagc cacctccaga gacctttgac 1260
ttcaaaatca agtttgctga gactaaggcc catgccaagg caattgagat ggaattgagg 1320
cagatggagg tggcccaggc caatcgacac atgtccctgc tgacagcctt catgcctgac 1380
agcttccttc ggccaggtgg ggaccatgac tgcgttctgg tgctgttgct catgcctcgt 1440
ctcatttgca aggcagagct gatccggaag caggcccagg agaagtttga actaagtgag 1500
aactgttcag agcggcctgg gctgcgagga gctgctgggg agcaactcag ctttgctgct 1560
ggactggtgt actcgctgag cctgctgcag gccacgctac accgctatga gcatgccctc 1620
tctcagtgca gtgtggatgt gtataagaaa gtgggcagcc tgtaccctga gatgagtgcc 1680
catgagcgct ccttggattt cctcattgaa ctgctgcaca aggatcagct ggatgagact 1740
gtcaatgtgg agcctctcac caaggccatc aagtactatc agcatctgta cagcatccac 1800
cttgccgaac agcctgagga ctgtactatg cagctggctg accacattaa gttcacgcag 1860
agtgctctgg actgcatgag tgtggaggta ggacggctgc gtgccttctt gcagggtggg 1920
caggaggcta cagatattgc cctcctgctc cgggatctgg aaacttcatg cagtgacatc 1980
cgccagttct gcaagaagat ccgaaggcga atgccaggga cagatgctcc tgggatccca 2040
gctgcactgg cctttggacc acaggtatct gacacgctcc tagactgcag gaaacacttg 2100
acgtgggtcg tggctgtgct gcaggaggtg gcagctgctg ctgcccagct cattgcccca 2160
ctggcagaga atgaggggct acttgtggct gctctggagg aactggcttt caaagcaagc 2220
gagcagatct atgggacccc ctccagcagc ccctatgagt gtctgcgcca gtcatgcaac 2280
atcctcatca gtaccatgaa caagctggcc acagccatgc aggaggggga gtatgatgca 2340
gagcggcccc ccagcaagcc tccaccggtt gaactgcggg ctgctgccct tcgtgcagag 2400
atcacagatg ctgaaggcct gggtttgaag ctcgaagatc gagagacagt tattaaggag 2460
ttgaagaagt cactcaagat taagggagag gagctaagtg aggccaatgt gcggctgagc 2520
ctcctggaga agaagttgga cagtgctgcc aaggatgcag atgagcgcat cgagaaagtc 2580
cagactcggc tggaggagac ccaggcactg ctgcgaaaga aggagaaaga gtttgaggag 2640
acaatggatg cactccaggc tgacatcgac cagctggagg cagagaaggc agaactaaag 2700
cagcgtctga acagccagtc caaacgcacg attgagggac tccggggccc tcctccttca 2760
ggcattgcta ctctggtctc tggcattgct ggtgaagaac agcagcgagg agccatccct 2820
gggcaggctc cagggtctgt gccaggccca gggctggtga aggactcacc actgctgctt 2880
cagcagatct ctgccatgag gctgcacatc tcccagctcc agcatgagaa cagcatcctc 2940
aagggagccc agatgaaggc atccttggca tccctgcccc ctctgcatgt tgcaaagcta 3000
tcccatgagg gccctggcag tgagttacca gctggagcgc tgtatcgtaa gaccagccag 3060
ctgctggaga cattgaatca attgagcaca cacacgcacg tagtagacat cactcgcacc 3120
agccctgctg ccaagagccc gtcggcccaa cttatggagc aagtggctca gcttaagtcc 3180
ctgagtgaca ccgtcgagaa gctcaaggat gaggtcctca aggagacagt atctcagcgc 3240
cctggagcca cagtacccac tgactttgcc accttccctt catcagcctt cctcagggag 3300
gatccaaagt gggaattccc tcggaagaac ttggttcttg gaaaaactct aggagaaggc 3360
gaatttggaa aagtggtcaa ggcaacggcc ttccatctga aaggcagagc agggtacacc 3420
acggtggccg tgaagatgct gaaagagaac gcctccccga gtgagcttcg agacctgctg 3480
tcagagttca acgtcctgaa gcaggtcaac cacccacatg tcatcaaatt gtatggggcc 3540
tgcagccagg atggcccgct cctcctcatc gtggagtacg ccaaatacgg ctccctgcgg 3600
ggcttcctcc gcgagagccg caaagtgggg cctggctacc tgggcagtgg aggcagccgc 3660
aactccagct ccctggacca cccggatgag cgggccctca ccatgggcga cctcatctca 3720
tttgcctggc agatctcaca ggggatgcag tatctggccg agatgaagct cgttcatcgg 3780
gacttggcag ccagaaacat cctggtagct gaggggcgga agatgaagat ttcggatttc 3840
ggcttgtccc gagatgttta tgaagaggat tcctacgtga agaggagcca gggtcggatt 3900
ccagttaaat ggatggcaat tgaatccctt tttgatcata tctacaccac gcaaagtgat 3960
gtatggtctt ttggtgtcct gctgtgggag atcgtgaccc tagggggaaa cccctatcct 4020
gggattcctc ctgagcggct cttcaacctt ctgaagaccg gccaccggat ggagaggcca 4080
gacaactgca gcgaggagat gtaccgcctg atgctgcaat gctggaagca ggagccggac 4140
aaaaggccgg tgtttgcgga catcagcaaa gacctggaga agatgatggt taagaggaga 4200
gactacttgg accttgcggc gtccactcca tctgactccc tgatttatga cgacggcctc 4260
tcagaggagg agacaccgct ggtggactgt aataatgccc ccctccctcg agccctccct 4320
tccacatgga ttgaaaacaa actctatggt agaatttccc atgcatttac tagattctag 4380
<210> 8
<211> 1459
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 8
Met Met Arg Gln Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro
1 5 10 15
Lys Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser
20 25 30
Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu
35 40 45
Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr
50 55 60
Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser
65 70 75 80
Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys
85 90 95
Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys
100 105 110
Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys
115 120 125
Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu
130 135 140
Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met
145 150 155 160
Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp
165 170 175
Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu
180 185 190
Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu
195 200 205
Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr
210 215 220
Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu
225 230 235 240
Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys
245 250 255
Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg
260 265 270
Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr
275 280 285
Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu
290 295 300
Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val
305 310 315 320
Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met
325 330 335
Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg
340 345 350
Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg
355 360 365
Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys
370 375 380
Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu
385 390 395 400
Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro
405 410 415
Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala
420 425 430
Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn
435 440 445
Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg
450 455 460
Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg
465 470 475 480
Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe
485 490 495
Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala
500 505 510
Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu
515 520 525
Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser
530 535 540
Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala
545 550 555 560
His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln
565 570 575
Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr
580 585 590
Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys
595 600 605
Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp
610 615 620
Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly
625 630 635 640
Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser
645 650 655
Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro
660 665 670
Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln
675 680 685
Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val
690 695 700
Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro
705 710 715 720
Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala
725 730 735
Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr
740 745 750
Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys
755 760 765
Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro
770 775 780
Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu
785 790 795 800
Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr
805 810 815
Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu
820 825 830
Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser
835 840 845
Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu
850 855 860
Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu
865 870 875 880
Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
885 890 895
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile Glu
900 905 910
Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val Ser Gly
915 920 925
Ile Ala Gly Glu Glu Gln Gln Arg Gly Ala Ile Pro Gly Gln Ala Pro
930 935 940
Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu
945 950 955 960
Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu
965 970 975
Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu
980 985 990
Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu
995 1000 1005
Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu
1010 1015 1020
Thr Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp Ile Thr
1025 1030 1035
Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu Met Glu
1040 1045 1050
Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu Lys Leu
1055 1060 1065
Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro Gly Ala
1070 1075 1080
Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu
1085 1090 1095
Arg Glu Asp Pro Lys Trp Glu Phe Pro Arg Lys Asn Leu Val Leu
1100 1105 1110
Gly Lys Thr Leu Gly Glu Gly Glu Phe Gly Lys Val Val Lys Ala
1115 1120 1125
Thr Ala Phe His Leu Lys Gly Arg Ala Gly Tyr Thr Thr Val Ala
1130 1135 1140
Val Lys Met Leu Lys Glu Asn Ala Ser Pro Ser Glu Leu Arg Asp
1145 1150 1155
Leu Leu Ser Glu Phe Asn Val Leu Lys Gln Val Asn His Pro His
1160 1165 1170
Val Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp Gly Pro Leu Leu
1175 1180 1185
Leu Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu Arg Gly Phe Leu
1190 1195 1200
Arg Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu Gly Ser Gly Gly
1205 1210 1215
Ser Arg Asn Ser Ser Ser Leu Asp His Pro Asp Glu Arg Ala Leu
1220 1225 1230
Thr Met Gly Asp Leu Ile Ser Phe Ala Trp Gln Ile Ser Gln Gly
1235 1240 1245
Met Gln Tyr Leu Ala Glu Met Lys Leu Val His Arg Asp Leu Ala
1250 1255 1260
Ala Arg Asn Ile Leu Val Ala Glu Gly Arg Lys Met Lys Ile Ser
1265 1270 1275
Asp Phe Gly Leu Ser Arg Asp Val Tyr Glu Glu Asp Ser Tyr Val
1280 1285 1290
Lys Arg Ser Gln Gly Arg Ile Pro Val Lys Trp Met Ala Ile Glu
1295 1300 1305
Ser Leu Phe Asp His Ile Tyr Thr Thr Gln Ser Asp Val Trp Ser
1310 1315 1320
Phe Gly Val Leu Leu Trp Glu Ile Val Thr Leu Gly Gly Asn Pro
1325 1330 1335
Tyr Pro Gly Ile Pro Pro Glu Arg Leu Phe Asn Leu Leu Lys Thr
1340 1345 1350
Gly His Arg Met Glu Arg Pro Asp Asn Cys Ser Glu Glu Met Tyr
1355 1360 1365
Arg Leu Met Leu Gln Cys Trp Lys Gln Glu Pro Asp Lys Arg Pro
1370 1375 1380
Val Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys Met Met Val Lys
1385 1390 1395
Arg Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr Pro Ser Asp Ser
1400 1405 1410
Leu Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu Thr Pro Leu Val
1415 1420 1425
Asp Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu Pro Ser Thr Trp
1430 1435 1440
Ile Glu Asn Lys Leu Tyr Gly Arg Ile Ser His Ala Phe Thr Arg
1445 1450 1455
Phe
<210> 9
<211> 4833
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 9
atggcacaga gcaagaggca cgtgtacagc cggacgccca gcggcagcag gatgagtgcg 60
gaggcaagcg cccggcctct gcgggtgggc tcccgtgtag aggtgattgg aaaaggccac 120
cgaggcactg tggcctatgt tggagccaca ctgtttgcca ctggcaaatg ggtaggcgtg 180
attctggatg aagcaaaggg caaaaatgat ggaactgttc aaggcaggaa gtacttcact 240
tgtgatgaag ggcatggcat ctttgtgcgc cagtcccaga tccaggtatt tgaagatgga 300
gcagatacta cttccccaga gacacctgat tcttctgctt caaaagtcct caaaagagag 360
ggaactgata caactgcaaa gactagcaaa ctgcccacgc gcccagccag tactggggtg 420
gctggggcca gtagctccct gggcccctct ggctcagcgt cagcaggtga gctgagcagc 480
agtgagccca gcaccccggc tcagactccg ctggcagcac ccatcatccc cacgccggtc 540
ctcacctctc ctggagcagt ccccccgctt ccttccccat ccaaggagga ggagggacta 600
agggctcagg tgcgggacct ggaggagaaa ctagagaccc tgagactgaa acgggcagaa 660
gacaaagcaa agctaaaaga gctggagaaa cacaaaatcc agctggagca ggtgcaggaa 720
tggaagagca aaatgcagga gcagcaggcc gacctgcagc ggcgcctcaa ggaggcgaga 780
aaggaagcca aggaggcgct ggaggcaaag gaacgctata tggaggagat ggctgatact 840
gctgatgcca ttgagatggc cactttggac aaggagatgg ctgaagagcg ggctgagtcc 900
ctgcagcagg aggtggaggc actgaaggag cgggtggacg agctcactac tgacttagag 960
atcctcaagg ctgagattga agagaagggc tcagatggcg ctgcatccag ttatcagctc 1020
aagcagcttg aggagcagaa tgcccgcctg aaggatgccc tggtgaggat gcgggatctt 1080
tcttcctcag agaagcagga gcatgtgaag ctccagaagc tcatggaaaa gaagaaccaa 1140
gagctggaag ttgtgaggca acagcgggag cgtctgcagg aggagctaag ccaggcagag 1200
agcaccattg atgagctcaa ggagcaggtg gatgctgctc tgggtgctga ggagatggtg 1260
gagatgctga cagatcggaa cctgaatctg gaagagaaag tgcgcgagtt gagggagact 1320
gtgggagact tggaagcgat gaatgagatg aacgatgagc tgcaggagaa tgcacgtgag 1380
acagaactgg agctgcggga gcagctggac atggcaggcg cgcgggttcg tgaggcccag 1440
aagcgtgtgg aggcagccca ggagacggtt gcagactacc agcagaccat caagaagtac 1500
cgccagctga ccgcccatct acaggatgtg aatcgggaac tgacaaacca gcaggaagca 1560
tctgtggaga ggcaacagca gccacctcca gagacctttg acttcaaaat caagtttgct 1620
gagactaagg cccatgccaa ggcaattgag atggaattga ggcagatgga ggtggcccag 1680
gccaatcgac acatgtccct gctgacagcc ttcatgcctg acagcttcct tcggccaggt 1740
ggggaccatg actgcgttct ggtgctgttg ctcatgcctc gtctcatttg caaggcagag 1800
ctgatccgga agcaggccca ggagaagttt gaactaagtg agaactgttc agagcggcct 1860
gggctgcgag gagctgctgg ggagcaactc agctttgctg ctggactggt gtactcgctg 1920
agcctgctgc aggccacgct acaccgctat gagcatgccc tctctcagtg cagtgtggat 1980
gtgtataaga aagtgggcag cctgtaccct gagatgagtg cccatgagcg ctccttggat 2040
ttcctcattg aactgctgca caaggatcag ctggatgaga ctgtcaatgt ggagcctctc 2100
accaaggcca tcaagtacta tcagcatctg tacagcatcc accttgccga acagcctgag 2160
gactgtacta tgcagctggc tgaccacatt aagttcacgc agagtgctct ggactgcatg 2220
agtgtggagg taggacggct gcgtgccttc ttgcagggtg ggcaggaggc tacagatatt 2280
gccctcctgc tccgggatct ggaaacttca tgcagtgaca tccgccagtt ctgcaagaag 2340
atccgaaggc gaatgccagg gacagatgct cctgggatcc cagctgcact ggcctttgga 2400
ccacaggtat ctgacacgct cctagactgc aggaaacact tgacgtgggt cgtggctgtg 2460
ctgcaggagg tggcagctgc tgctgcccag ctcattgccc cactggcaga gaatgagggg 2520
ctacttgtgg ctgctctgga ggaactggct ttcaaagcaa gcgagcagat ctatgggacc 2580
ccctccagca gcccctatga gtgtctgcgc cagtcatgca acatcctcat cagtaccatg 2640
aacaagctgg ccacagccat gcaggagggg gagtatgatg cagagcggcc ccccagcaag 2700
cctccaccgg ttgaactgcg ggctgctgcc cttcgtgcag agatcacaga tgctgaaggc 2760
ctgggtttga agctcgaaga tcgagagaca gttattaagg agttgaagaa gtcactcaag 2820
attaagggag aggagctaag tgaggccaat gtgcggctga gcctcctgga gaagaagttg 2880
gacagtgctg ccaaggatgc agatgagcgc atcgagaaag tccagactcg gctggaggag 2940
acccaggcac tgctgcgaaa gaaggagaaa gagtttgagg agacaatgga tgcactccag 3000
gctgacatcg accagctgga ggcagagaag gcagaactaa agcagcgtct gaacagccag 3060
tccaaacgca cgattgaggg actccggggc cctcctcctt caggcattgc tactctggtc 3120
tctggcattg ctggtggagc catccctggg caggctccag ggtctgtgcc aggcccaggg 3180
ctggtgaagg actcaccact gctgcttcag cagatctctg ccatgaggct gcacatctcc 3240
cagctccagc atgagaacag catcctcaag ggagcccaga tgaaggcatc cttggcatcc 3300
ctgccccctc tgcatgttgc aaagctatcc catgagggcc ctggcagtga gttaccagct 3360
ggagcgctgt atcgtaagac cagccagctg ctggagacat tgaatcaatt gagcacacac 3420
acgcacgtag tagacatcac tcgcaccagc cctgctgcca agagcccgtc ggcccaactt 3480
atggagcaag tggctcagct taagtccctg agtgacaccg tcgagaagct caaggatgag 3540
gtcctcaagg agacagtatc tcagcgccct ggagccacag tacccactga ctttgccacc 3600
ttcccttcat cagccttcct cagggaggat ccaaagtggg aattccctcg gaagaacttg 3660
gttcttggaa aaactctagg agaaggcgaa tttggaaaag tggtcaaggc aacggccttc 3720
catctgaaag gcagagcagg gtacaccacg gtggccgtga agatgctgaa agagaacgcc 3780
tccccgagtg agcttcgaga cctgctgtca gagttcaacg tcctgaagca ggtcaaccac 3840
ccacatgtca tcaaattgta tggggcctgc agccaggatg gcccgctcct cctcatcgtg 3900
gagtacgcca aatacggctc cctgcggggc ttcctccgcg agagccgcaa agtggggcct 3960
ggctacctgg gcagtggagg cagccgcaac tccagctccc tggaccaccc ggatgagcgg 4020
gccctcacca tgggcgacct catctcattt gcctggcaga tctcacaggg gatgcagtat 4080
ctggccgaga tgaagctcgt tcatcgggac ttggcagcca gaaacatcct ggtagctgag 4140
gggcggaaga tgaagatttc ggatttcggc ttgtcccgag atgtttatga agaggattcc 4200
tacgtgaaga ggagccaggg tcggattcca gttaaatgga tggcaattga atcccttttt 4260
gatcatatct acaccacgca aagtgatgta tggtcttttg gtgtcctgct gtgggagatc 4320
gtgaccctag ggggaaaccc ctatcctggg attcctcctg agcggctctt caaccttctg 4380
aagaccggcc accggatgga gaggccagac aactgcagcg aggagatgta ccgcctgatg 4440
ctgcaatgct ggaagcagga gccggacaaa aggccggtgt ttgcggacat cagcaaagac 4500
ctggagaaga tgatggttaa gaggagagac tacttggacc ttgcggcgtc cactccatct 4560
gactccctga tttatgacga cggcctctca gaggaggaga caccgctggt ggactgtaat 4620
aatgcccccc tccctcgagc cctcccttcc acatggattg aaaacaaact ctatggcatg 4680
tcagacccga actggcctgg agagagtcct gtaccactca cgagagctga tggcactaac 4740
actgggtttc caagatatcc aaatgatagt gtatatgcta actggatgct ttcaccctca 4800
gcggcaaaat taatggacac gtttgatagt taa 4833
<210> 10
<211> 1610
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 10
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser
130 135 140
Ser Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser
145 150 155 160
Ser Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile
165 170 175
Pro Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser
180 185 190
Pro Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu
195 200 205
Glu Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys
210 215 220
Leu Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu
225 230 235 240
Trp Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu
245 250 255
Lys Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg
260 265 270
Tyr Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr
275 280 285
Leu Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu
290 295 300
Val Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu
305 310 315 320
Ile Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser
325 330 335
Ser Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp
340 345 350
Ala Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His
355 360 365
Val Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val
370 375 380
Val Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu
385 390 395 400
Ser Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala
405 410 415
Glu Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu
420 425 430
Lys Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn
435 440 445
Glu Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu
450 455 460
Leu Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln
465 470 475 480
Lys Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr
485 490 495
Ile Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg
500 505 510
Glu Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro
515 520 525
Pro Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala
530 535 540
His Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln
545 550 555 560
Ala Asn Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe
565 570 575
Leu Arg Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met
580 585 590
Pro Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu
595 600 605
Lys Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly
610 615 620
Ala Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu
625 630 635 640
Ser Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln
645 650 655
Cys Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met
660 665 670
Ser Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys
675 680 685
Asp Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile
690 695 700
Lys Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu
705 710 715 720
Asp Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala
725 730 735
Leu Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln
740 745 750
Gly Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu
755 760 765
Thr Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg
770 775 780
Met Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly
785 790 795 800
Pro Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp
805 810 815
Val Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile
820 825 830
Ala Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu
835 840 845
Leu Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser
850 855 860
Pro Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met
865 870 875 880
Asn Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg
885 890 895
Pro Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg
900 905 910
Ala Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg
915 920 925
Glu Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu
930 935 940
Glu Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu
945 950 955 960
Asp Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr
965 970 975
Arg Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe
980 985 990
Glu Glu Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala
995 1000 1005
Glu Lys Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg
1010 1015 1020
Thr Ile Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr
1025 1030 1035
Leu Val Ser Gly Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro
1040 1045 1050
Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu
1055 1060 1065
Leu Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln
1070 1075 1080
His Glu Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu
1085 1090 1095
Ala Ser Leu Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly
1100 1105 1110
Pro Gly Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser
1115 1120 1125
Gln Leu Leu Glu Thr Leu Asn Gln Leu Ser Thr His Thr His Val
1130 1135 1140
Val Asp Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala
1145 1150 1155
Gln Leu Met Glu Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr
1160 1165 1170
Val Glu Lys Leu Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln
1175 1180 1185
Arg Pro Gly Ala Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser
1190 1195 1200
Ser Ala Phe Leu Arg Glu Asp Pro Lys Trp Glu Phe Pro Arg Lys
1205 1210 1215
Asn Leu Val Leu Gly Lys Thr Leu Gly Glu Gly Glu Phe Gly Lys
1220 1225 1230
Val Val Lys Ala Thr Ala Phe His Leu Lys Gly Arg Ala Gly Tyr
1235 1240 1245
Thr Thr Val Ala Val Lys Met Leu Lys Glu Asn Ala Ser Pro Ser
1250 1255 1260
Glu Leu Arg Asp Leu Leu Ser Glu Phe Asn Val Leu Lys Gln Val
1265 1270 1275
Asn His Pro His Val Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp
1280 1285 1290
Gly Pro Leu Leu Leu Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu
1295 1300 1305
Arg Gly Phe Leu Arg Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu
1310 1315 1320
Gly Ser Gly Gly Ser Arg Asn Ser Ser Ser Leu Asp His Pro Asp
1325 1330 1335
Glu Arg Ala Leu Thr Met Gly Asp Leu Ile Ser Phe Ala Trp Gln
1340 1345 1350
Ile Ser Gln Gly Met Gln Tyr Leu Ala Glu Met Lys Leu Val His
1355 1360 1365
Arg Asp Leu Ala Ala Arg Asn Ile Leu Val Ala Glu Gly Arg Lys
1370 1375 1380
Met Lys Ile Ser Asp Phe Gly Leu Ser Arg Asp Val Tyr Glu Glu
1385 1390 1395
Asp Ser Tyr Val Lys Arg Ser Gln Gly Arg Ile Pro Val Lys Trp
1400 1405 1410
Met Ala Ile Glu Ser Leu Phe Asp His Ile Tyr Thr Thr Gln Ser
1415 1420 1425
Asp Val Trp Ser Phe Gly Val Leu Leu Trp Glu Ile Val Thr Leu
1430 1435 1440
Gly Gly Asn Pro Tyr Pro Gly Ile Pro Pro Glu Arg Leu Phe Asn
1445 1450 1455
Leu Leu Lys Thr Gly His Arg Met Glu Arg Pro Asp Asn Cys Ser
1460 1465 1470
Glu Glu Met Tyr Arg Leu Met Leu Gln Cys Trp Lys Gln Glu Pro
1475 1480 1485
Asp Lys Arg Pro Val Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys
1490 1495 1500
Met Met Val Lys Arg Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr
1505 1510 1515
Pro Ser Asp Ser Leu Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu
1520 1525 1530
Thr Pro Leu Val Asp Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu
1535 1540 1545
Pro Ser Thr Trp Ile Glu Asn Lys Leu Tyr Gly Met Ser Asp Pro
1550 1555 1560
Asn Trp Pro Gly Glu Ser Pro Val Pro Leu Thr Arg Ala Asp Gly
1565 1570 1575
Thr Asn Thr Gly Phe Pro Arg Tyr Pro Asn Asp Ser Val Tyr Ala
1580 1585 1590
Asn Trp Met Leu Ser Pro Ser Ala Ala Lys Leu Met Asp Thr Phe
1595 1600 1605
Asp Ser
1610
<210> 11
<211> 4707
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 11
atggcacaga gcaagaggca cgtgtacagc cggacgccca gcggcagcag gatgagtgcg 60
gaggcaagcg cccggcctct gcgggtgggc tcccgtgtag aggtgattgg aaaaggccac 120
cgaggcactg tggcctatgt tggagccaca ctgtttgcca ctggcaaatg ggtaggcgtg 180
attctggatg aagcaaaggg caaaaatgat ggaactgttc aaggcaggaa gtacttcact 240
tgtgatgaag ggcatggcat ctttgtgcgc cagtcccaga tccaggtatt tgaagatgga 300
gcagatacta cttccccaga gacacctgat tcttctgctt caaaagtcct caaaagagag 360
ggaactgata caactgcaaa gactagcaaa ctgcccacgc gcccagccag tactggggtg 420
gctggggcca gtagctccct gggcccctct ggctcagcgt cagcaggtga gctgagcagc 480
agtgagccca gcaccccggc tcagactccg ctggcagcac ccatcatccc cacgccggtc 540
ctcacctctc ctggagcagt ccccccgctt ccttccccat ccaaggagga ggagggacta 600
agggctcagg tgcgggacct ggaggagaaa ctagagaccc tgagactgaa acgggcagaa 660
gacaaagcaa agctaaaaga gctggagaaa cacaaaatcc agctggagca ggtgcaggaa 720
tggaagagca aaatgcagga gcagcaggcc gacctgcagc ggcgcctcaa ggaggcgaga 780
aaggaagcca aggaggcgct ggaggcaaag gaacgctata tggaggagat ggctgatact 840
gctgatgcca ttgagatggc cactttggac aaggagatgg ctgaagagcg ggctgagtcc 900
ctgcagcagg aggtggaggc actgaaggag cgggtggacg agctcactac tgacttagag 960
atcctcaagg ctgagattga agagaagggc tcagatggcg ctgcatccag ttatcagctc 1020
aagcagcttg aggagcagaa tgcccgcctg aaggatgccc tggtgaggat gcgggatctt 1080
tcttcctcag agaagcagga gcatgtgaag ctccagaagc tcatggaaaa gaagaaccaa 1140
gagctggaag ttgtgaggca acagcgggag cgtctgcagg aggagctaag ccaggcagag 1200
agcaccattg atgagctcaa ggagcaggtg gatgctgctc tgggtgctga ggagatggtg 1260
gagatgctga cagatcggaa cctgaatctg gaagagaaag tgcgcgagtt gagggagact 1320
gtgggagact tggaagcgat gaatgagatg aacgatgagc tgcaggagaa tgcacgtgag 1380
acagaactgg agctgcggga gcagctggac atggcaggcg cgcgggttcg tgaggcccag 1440
aagcgtgtgg aggcagccca ggagacggtt gcagactacc agcagaccat caagaagtac 1500
cgccagctga ccgcccatct acaggatgtg aatcgggaac tgacaaacca gcaggaagca 1560
tctgtggaga ggcaacagca gccacctcca gagacctttg acttcaaaat caagtttgct 1620
gagactaagg cccatgccaa ggcaattgag atggaattga ggcagatgga ggtggcccag 1680
gccaatcgac acatgtccct gctgacagcc ttcatgcctg acagcttcct tcggccaggt 1740
ggggaccatg actgcgttct ggtgctgttg ctcatgcctc gtctcatttg caaggcagag 1800
ctgatccgga agcaggccca ggagaagttt gaactaagtg agaactgttc agagcggcct 1860
gggctgcgag gagctgctgg ggagcaactc agctttgctg ctggactggt gtactcgctg 1920
agcctgctgc aggccacgct acaccgctat gagcatgccc tctctcagtg cagtgtggat 1980
gtgtataaga aagtgggcag cctgtaccct gagatgagtg cccatgagcg ctccttggat 2040
ttcctcattg aactgctgca caaggatcag ctggatgaga ctgtcaatgt ggagcctctc 2100
accaaggcca tcaagtacta tcagcatctg tacagcatcc accttgccga acagcctgag 2160
gactgtacta tgcagctggc tgaccacatt aagttcacgc agagtgctct ggactgcatg 2220
agtgtggagg taggacggct gcgtgccttc ttgcagggtg ggcaggaggc tacagatatt 2280
gccctcctgc tccgggatct ggaaacttca tgcagtgaca tccgccagtt ctgcaagaag 2340
atccgaaggc gaatgccagg gacagatgct cctgggatcc cagctgcact ggcctttgga 2400
ccacaggtat ctgacacgct cctagactgc aggaaacact tgacgtgggt cgtggctgtg 2460
ctgcaggagg tggcagctgc tgctgcccag ctcattgccc cactggcaga gaatgagggg 2520
ctacttgtgg ctgctctgga ggaactggct ttcaaagcaa gcgagcagat ctatgggacc 2580
ccctccagca gcccctatga gtgtctgcgc cagtcatgca acatcctcat cagtaccatg 2640
aacaagctgg ccacagccat gcaggagggg gagtatgatg cagagcggcc ccccagcaag 2700
cctccaccgg ttgaactgcg ggctgctgcc cttcgtgcag agatcacaga tgctgaaggc 2760
ctgggtttga agctcgaaga tcgagagaca gttattaagg agttgaagaa gtcactcaag 2820
attaagggag aggagctaag tgaggccaat gtgcggctga gcctcctgga gaagaagttg 2880
gacagtgctg ccaaggatgc agatgagcgc atcgagaaag tccagactcg gctggaggag 2940
acccaggcac tgctgcgaaa gaaggagaaa gagtttgagg agacaatgga tgcactccag 3000
gctgacatcg accagctgga ggcagagaag gcagaactaa agcagcgtct gaacagccag 3060
tccaaacgca cgattgaggg actccggggc cctcctcctt caggcattgc tactctggtc 3120
tctggcattg ctggtggagc catccctggg caggctccag ggtctgtgcc aggcccaggg 3180
ctggtgaagg actcaccact gctgcttcag cagatctctg ccatgaggct gcacatctcc 3240
cagctccagc atgagaacag catcctcaag ggagcccaga tgaaggcatc cttggcatcc 3300
ctgccccctc tgcatgttgc aaagctatcc catgagggcc ctggcagtga gttaccagct 3360
ggagcgctgt atcgtaagac cagccagctg ctggagacat tgaatcaatt gagcacacac 3420
acgcacgtag tagacatcac tcgcaccagc cctgctgcca agagcccgtc ggcccaactt 3480
atggagcaag tggctcagct taagtccctg agtgacaccg tcgagaagct caaggatgag 3540
gtcctcaagg agacagtatc tcagcgccct ggagccacag tacccactga ctttgccacc 3600
ttcccttcat cagccttcct cagggaggat ccaaagtggg aattccctcg gaagaacttg 3660
gttcttggaa aaactctagg agaaggcgaa tttggaaaag tggtcaaggc aacggccttc 3720
catctgaaag gcagagcagg gtacaccacg gtggccgtga agatgctgaa agagaacgcc 3780
tccccgagtg agcttcgaga cctgctgtca gagttcaacg tcctgaagca ggtcaaccac 3840
ccacatgtca tcaaattgta tggggcctgc agccaggatg gcccgctcct cctcatcgtg 3900
gagtacgcca aatacggctc cctgcggggc ttcctccgcg agagccgcaa agtggggcct 3960
ggctacctgg gcagtggagg cagccgcaac tccagctccc tggaccaccc ggatgagcgg 4020
gccctcacca tgggcgacct catctcattt gcctggcaga tctcacaggg gatgcagtat 4080
ctggccgaga tgaagctcgt tcatcgggac ttggcagcca gaaacatcct ggtagctgag 4140
gggcggaaga tgaagatttc ggatttcggc ttgtcccgag atgtttatga agaggattcc 4200
tacgtgaaga ggagccaggg tcggattcca gttaaatgga tggcaattga atcccttttt 4260
gatcatatct acaccacgca aagtgatgta tggtcttttg gtgtcctgct gtgggagatc 4320
gtgaccctag ggggaaaccc ctatcctggg attcctcctg agcggctctt caaccttctg 4380
aagaccggcc accggatgga gaggccagac aactgcagcg aggagatgta ccgcctgatg 4440
ctgcaatgct ggaagcagga gccggacaaa aggccggtgt ttgcggacat cagcaaagac 4500
ctggagaaga tgatggttaa gaggagagac tacttggacc ttgcggcgtc cactccatct 4560
gactccctga tttatgacga cggcctctca gaggaggaga caccgctggt ggactgtaat 4620
aatgcccccc tccctcgagc cctcccttcc acatggattg aaaacaaact ctatggtaga 4680
atttcccatg catttactag attctag 4707
<210> 12
<211> 1568
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 12
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser
130 135 140
Ser Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser
145 150 155 160
Ser Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile
165 170 175
Pro Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser
180 185 190
Pro Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu
195 200 205
Glu Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys
210 215 220
Leu Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu
225 230 235 240
Trp Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu
245 250 255
Lys Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg
260 265 270
Tyr Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr
275 280 285
Leu Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu
290 295 300
Val Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu
305 310 315 320
Ile Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser
325 330 335
Ser Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp
340 345 350
Ala Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His
355 360 365
Val Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val
370 375 380
Val Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu
385 390 395 400
Ser Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala
405 410 415
Glu Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu
420 425 430
Lys Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn
435 440 445
Glu Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu
450 455 460
Leu Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln
465 470 475 480
Lys Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr
485 490 495
Ile Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg
500 505 510
Glu Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro
515 520 525
Pro Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala
530 535 540
His Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln
545 550 555 560
Ala Asn Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe
565 570 575
Leu Arg Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met
580 585 590
Pro Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu
595 600 605
Lys Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly
610 615 620
Ala Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu
625 630 635 640
Ser Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln
645 650 655
Cys Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met
660 665 670
Ser Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys
675 680 685
Asp Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile
690 695 700
Lys Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu
705 710 715 720
Asp Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala
725 730 735
Leu Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln
740 745 750
Gly Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu
755 760 765
Thr Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg
770 775 780
Met Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly
785 790 795 800
Pro Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp
805 810 815
Val Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile
820 825 830
Ala Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu
835 840 845
Leu Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser
850 855 860
Pro Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met
865 870 875 880
Asn Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg
885 890 895
Pro Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg
900 905 910
Ala Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg
915 920 925
Glu Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu
930 935 940
Glu Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu
945 950 955 960
Asp Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr
965 970 975
Arg Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe
980 985 990
Glu Glu Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala
995 1000 1005
Glu Lys Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg
1010 1015 1020
Thr Ile Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr
1025 1030 1035
Leu Val Ser Gly Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro
1040 1045 1050
Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu
1055 1060 1065
Leu Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln
1070 1075 1080
His Glu Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu
1085 1090 1095
Ala Ser Leu Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly
1100 1105 1110
Pro Gly Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser
1115 1120 1125
Gln Leu Leu Glu Thr Leu Asn Gln Leu Ser Thr His Thr His Val
1130 1135 1140
Val Asp Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala
1145 1150 1155
Gln Leu Met Glu Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr
1160 1165 1170
Val Glu Lys Leu Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln
1175 1180 1185
Arg Pro Gly Ala Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser
1190 1195 1200
Ser Ala Phe Leu Arg Glu Asp Pro Lys Trp Glu Phe Pro Arg Lys
1205 1210 1215
Asn Leu Val Leu Gly Lys Thr Leu Gly Glu Gly Glu Phe Gly Lys
1220 1225 1230
Val Val Lys Ala Thr Ala Phe His Leu Lys Gly Arg Ala Gly Tyr
1235 1240 1245
Thr Thr Val Ala Val Lys Met Leu Lys Glu Asn Ala Ser Pro Ser
1250 1255 1260
Glu Leu Arg Asp Leu Leu Ser Glu Phe Asn Val Leu Lys Gln Val
1265 1270 1275
Asn His Pro His Val Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp
1280 1285 1290
Gly Pro Leu Leu Leu Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu
1295 1300 1305
Arg Gly Phe Leu Arg Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu
1310 1315 1320
Gly Ser Gly Gly Ser Arg Asn Ser Ser Ser Leu Asp His Pro Asp
1325 1330 1335
Glu Arg Ala Leu Thr Met Gly Asp Leu Ile Ser Phe Ala Trp Gln
1340 1345 1350
Ile Ser Gln Gly Met Gln Tyr Leu Ala Glu Met Lys Leu Val His
1355 1360 1365
Arg Asp Leu Ala Ala Arg Asn Ile Leu Val Ala Glu Gly Arg Lys
1370 1375 1380
Met Lys Ile Ser Asp Phe Gly Leu Ser Arg Asp Val Tyr Glu Glu
1385 1390 1395
Asp Ser Tyr Val Lys Arg Ser Gln Gly Arg Ile Pro Val Lys Trp
1400 1405 1410
Met Ala Ile Glu Ser Leu Phe Asp His Ile Tyr Thr Thr Gln Ser
1415 1420 1425
Asp Val Trp Ser Phe Gly Val Leu Leu Trp Glu Ile Val Thr Leu
1430 1435 1440
Gly Gly Asn Pro Tyr Pro Gly Ile Pro Pro Glu Arg Leu Phe Asn
1445 1450 1455
Leu Leu Lys Thr Gly His Arg Met Glu Arg Pro Asp Asn Cys Ser
1460 1465 1470
Glu Glu Met Tyr Arg Leu Met Leu Gln Cys Trp Lys Gln Glu Pro
1475 1480 1485
Asp Lys Arg Pro Val Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys
1490 1495 1500
Met Met Val Lys Arg Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr
1505 1510 1515
Pro Ser Asp Ser Leu Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu
1520 1525 1530
Thr Pro Leu Val Asp Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu
1535 1540 1545
Pro Ser Thr Trp Ile Glu Asn Lys Leu Tyr Gly Arg Ile Ser His
1550 1555 1560
Ala Phe Thr Arg Phe
1565
<210> 13
<211> 4491
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 13
atgatgagac aggcaccgac agcccgaaag accacaactc ggcgacccaa gcccacgcgc 60
ccagccagta ctggggtggc tggggccagt agctccctgg gcccctctgg ctcagcgtca 120
gcaggtgagc tgagcagcag tgagcccagc accccggctc agactccgct ggcagcaccc 180
atcatcccca cgccggtcct cacctctcct ggagcagtcc ccccgcttcc ttccccatcc 240
aaggaggagg agggactaag ggctcaggtg cgggacctgg aggagaaact agagaccctg 300
agactgaaac gggcagaaga caaagcaaag ctaaaagagc tggagaaaca caaaatccag 360
ctggagcagg tgcaggaatg gaagagcaaa atgcaggagc agcaggccga cctgcagcgg 420
cgcctcaagg aggcgagaaa ggaagccaag gaggcgctgg aggcaaagga acgctatatg 480
gaggagatgg ctgatactgc tgatgccatt gagatggcca ctttggacaa ggagatggct 540
gaagagcggg ctgagtccct gcagcaggag gtggaggcac tgaaggagcg ggtggacgag 600
ctcactactg acttagagat cctcaaggct gagattgaag agaagggctc agatggcgct 660
gcatccagtt atcagctcaa gcagcttgag gagcagaatg cccgcctgaa ggatgccctg 720
gtgaggatgc gggatctttc ttcctcagag aagcaggagc atgtgaagct ccagaagctc 780
atggaaaaga agaaccaaga gctggaagtt gtgaggcaac agcgggagcg tctgcaggag 840
gagctaagcc aggcagagag caccattgat gagctcaagg agcaggtgga tgctgctctg 900
ggtgctgagg agatggtgga gatgctgaca gatcggaacc tgaatctgga agagaaagtg 960
cgcgagttga gggagactgt gggagacttg gaagcgatga atgagatgaa cgatgagctg 1020
caggagaatg cacgtgagac agaactggag ctgcgggagc agctggacat ggcaggcgcg 1080
cgggttcgtg aggcccagaa gcgtgtggag gcagcccagg agacggttgc agactaccag 1140
cagaccatca agaagtaccg ccagctgacc gcccatctac aggatgtgaa tcgggaactg 1200
acaaaccagc aggaagcatc tgtggagagg caacagcagc cacctccaga gacctttgac 1260
ttcaaaatca agtttgctga gactaaggcc catgccaagg caattgagat ggaattgagg 1320
cagatggagg tggcccaggc caatcgacac atgtccctgc tgacagcctt catgcctgac 1380
agcttccttc ggccaggtgg ggaccatgac tgcgttctgg tgctgttgct catgcctcgt 1440
ctcatttgca aggcagagct gatccggaag caggcccagg agaagtttga actaagtgag 1500
aactgttcag agcggcctgg gctgcgagga gctgctgggg agcaactcag ctttgctgct 1560
ggactggtgt actcgctgag cctgctgcag gccacgctac accgctatga gcatgccctc 1620
tctcagtgca gtgtggatgt gtataagaaa gtgggcagcc tgtaccctga gatgagtgcc 1680
catgagcgct ccttggattt cctcattgaa ctgctgcaca aggatcagct ggatgagact 1740
gtcaatgtgg agcctctcac caaggccatc aagtactatc agcatctgta cagcatccac 1800
cttgccgaac agcctgagga ctgtactatg cagctggctg accacattaa gttcacgcag 1860
agtgctctgg actgcatgag tgtggaggta ggacggctgc gtgccttctt gcagggtggg 1920
caggaggcta cagatattgc cctcctgctc cgggatctgg aaacttcatg cagtgacatc 1980
cgccagttct gcaagaagat ccgaaggcga atgccaggga cagatgctcc tgggatccca 2040
gctgcactgg cctttggacc acaggtatct gacacgctcc tagactgcag gaaacacttg 2100
acgtgggtcg tggctgtgct gcaggaggtg gcagctgctg ctgcccagct cattgcccca 2160
ctggcagaga atgaggggct acttgtggct gctctggagg aactggcttt caaagcaagc 2220
gagcagatct atgggacccc ctccagcagc ccctatgagt gtctgcgcca gtcatgcaac 2280
atcctcatca gtaccatgaa caagctggcc acagccatgc aggaggggga gtatgatgca 2340
gagcggcccc ccagcaagcc tccaccggtt gaactgcggg ctgctgccct tcgtgcagag 2400
atcacagatg ctgaaggcct gggtttgaag ctcgaagatc gagagacagt tattaaggag 2460
ttgaagaagt cactcaagat taagggagag gagctaagtg aggccaatgt gcggctgagc 2520
ctcctggaga agaagttgga cagtgctgcc aaggatgcag atgagcgcat cgagaaagtc 2580
cagactcggc tggaggagac ccaggcactg ctgcgaaaga aggagaaaga gtttgaggag 2640
acaatggatg cactccaggc tgacatcgac cagctggagg cagagaaggc agaactaaag 2700
cagcgtctga acagccagtc caaacgcacg attgagggac tccggggccc tcctccttca 2760
ggcattgcta ctctggtctc tggcattgct ggtggagcca tccctgggca ggctccaggg 2820
tctgtgccag gcccagggct ggtgaaggac tcaccactgc tgcttcagca gatctctgcc 2880
atgaggctgc acatctccca gctccagcat gagaacagca tcctcaaggg agcccagatg 2940
aaggcatcct tggcatccct gccccctctg catgttgcaa agctatccca tgagggccct 3000
ggcagtgagt taccagctgg agcgctgtat cgtaagacca gccagctgct ggagacattg 3060
aatcaattga gcacacacac gcacgtagta gacatcactc gcaccagccc tgctgccaag 3120
agcccgtcgg cccaacttat ggagcaagtg gctcagctta agtccctgag tgacaccgtc 3180
gagaagctca aggatgaggt cctcaaggag acagtatctc agcgccctgg agccacagta 3240
cccactgact ttgccacctt cccttcatca gccttcctca gggaggatcc aaagtgggaa 3300
ttccctcgga agaacttggt tcttggaaaa actctaggag aaggcgaatt tggaaaagtg 3360
gtcaaggcaa cggccttcca tctgaaaggc agagcagggt acaccacggt ggccgtgaag 3420
atgctgaaag agaacgcctc cccgagtgag cttcgagacc tgctgtcaga gttcaacgtc 3480
ctgaagcagg tcaaccaccc acatgtcatc aaattgtatg gggcctgcag ccaggatggc 3540
ccgctcctcc tcatcgtgga gtacgccaaa tacggctccc tgcggggctt cctccgcgag 3600
agccgcaaag tggggcctgg ctacctgggc agtggaggca gccgcaactc cagctccctg 3660
gaccacccgg atgagcgggc cctcaccatg ggcgacctca tctcatttgc ctggcagatc 3720
tcacagggga tgcagtatct ggccgagatg aagctcgttc atcgggactt ggcagccaga 3780
aacatcctgg tagctgaggg gcggaagatg aagatttcgg atttcggctt gtcccgagat 3840
gtttatgaag aggattccta cgtgaagagg agccagggtc ggattccagt taaatggatg 3900
gcaattgaat ccctttttga tcatatctac accacgcaaa gtgatgtatg gtcttttggt 3960
gtcctgctgt gggagatcgt gaccctaggg ggaaacccct atcctgggat tcctcctgag 4020
cggctcttca accttctgaa gaccggccac cggatggaga ggccagacaa ctgcagcgag 4080
gagatgtacc gcctgatgct gcaatgctgg aagcaggagc cggacaaaag gccggtgttt 4140
gcggacatca gcaaagacct ggagaagatg atggttaaga ggagagacta cttggacctt 4200
gcggcgtcca ctccatctga ctccctgatt tatgacgacg gcctctcaga ggaggagaca 4260
ccgctggtgg actgtaataa tgcccccctc cctcgagccc tcccttccac atggattgaa 4320
aacaaactct atggcatgtc agacccgaac tggcctggag agagtcctgt accactcacg 4380
agagctgatg gcactaacac tgggtttcca agatatccaa atgatagtgt atatgctaac 4440
tggatgcttt caccctcagc ggcaaaatta atggacacgt ttgatagtta a 4491
<210> 14
<211> 1496
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 14
Met Met Arg Gln Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro
1 5 10 15
Lys Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser
20 25 30
Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu
35 40 45
Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr
50 55 60
Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser
65 70 75 80
Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys
85 90 95
Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys
100 105 110
Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys
115 120 125
Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu
130 135 140
Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met
145 150 155 160
Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp
165 170 175
Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu
180 185 190
Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu
195 200 205
Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr
210 215 220
Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu
225 230 235 240
Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys
245 250 255
Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg
260 265 270
Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr
275 280 285
Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu
290 295 300
Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val
305 310 315 320
Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met
325 330 335
Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg
340 345 350
Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg
355 360 365
Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys
370 375 380
Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu
385 390 395 400
Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro
405 410 415
Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala
420 425 430
Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn
435 440 445
Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg
450 455 460
Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg
465 470 475 480
Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe
485 490 495
Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala
500 505 510
Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu
515 520 525
Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser
530 535 540
Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala
545 550 555 560
His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln
565 570 575
Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr
580 585 590
Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys
595 600 605
Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp
610 615 620
Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly
625 630 635 640
Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser
645 650 655
Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro
660 665 670
Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln
675 680 685
Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val
690 695 700
Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro
705 710 715 720
Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala
725 730 735
Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr
740 745 750
Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys
755 760 765
Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro
770 775 780
Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu
785 790 795 800
Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr
805 810 815
Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu
820 825 830
Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser
835 840 845
Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu
850 855 860
Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu
865 870 875 880
Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
885 890 895
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile Glu
900 905 910
Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val Ser Gly
915 920 925
Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser Val Pro Gly
930 935 940
Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln Gln Ile Ser Ala
945 950 955 960
Met Arg Leu His Ile Ser Gln Leu Gln His Glu Asn Ser Ile Leu Lys
965 970 975
Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu Pro Pro Leu His Val
980 985 990
Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu Leu Pro Ala Gly Ala
995 1000 1005
Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu Thr Leu Asn Gln Leu
1010 1015 1020
Ser Thr His Thr His Val Val Asp Ile Thr Arg Thr Ser Pro Ala
1025 1030 1035
Ala Lys Ser Pro Ser Ala Gln Leu Met Glu Gln Val Ala Gln Leu
1040 1045 1050
Lys Ser Leu Ser Asp Thr Val Glu Lys Leu Lys Asp Glu Val Leu
1055 1060 1065
Lys Glu Thr Val Ser Gln Arg Pro Gly Ala Thr Val Pro Thr Asp
1070 1075 1080
Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu Arg Glu Asp Pro Lys
1085 1090 1095
Trp Glu Phe Pro Arg Lys Asn Leu Val Leu Gly Lys Thr Leu Gly
1100 1105 1110
Glu Gly Glu Phe Gly Lys Val Val Lys Ala Thr Ala Phe His Leu
1115 1120 1125
Lys Gly Arg Ala Gly Tyr Thr Thr Val Ala Val Lys Met Leu Lys
1130 1135 1140
Glu Asn Ala Ser Pro Ser Glu Leu Arg Asp Leu Leu Ser Glu Phe
1145 1150 1155
Asn Val Leu Lys Gln Val Asn His Pro His Val Ile Lys Leu Tyr
1160 1165 1170
Gly Ala Cys Ser Gln Asp Gly Pro Leu Leu Leu Ile Val Glu Tyr
1175 1180 1185
Ala Lys Tyr Gly Ser Leu Arg Gly Phe Leu Arg Glu Ser Arg Lys
1190 1195 1200
Val Gly Pro Gly Tyr Leu Gly Ser Gly Gly Ser Arg Asn Ser Ser
1205 1210 1215
Ser Leu Asp His Pro Asp Glu Arg Ala Leu Thr Met Gly Asp Leu
1220 1225 1230
Ile Ser Phe Ala Trp Gln Ile Ser Gln Gly Met Gln Tyr Leu Ala
1235 1240 1245
Glu Met Lys Leu Val His Arg Asp Leu Ala Ala Arg Asn Ile Leu
1250 1255 1260
Val Ala Glu Gly Arg Lys Met Lys Ile Ser Asp Phe Gly Leu Ser
1265 1270 1275
Arg Asp Val Tyr Glu Glu Asp Ser Tyr Val Lys Arg Ser Gln Gly
1280 1285 1290
Arg Ile Pro Val Lys Trp Met Ala Ile Glu Ser Leu Phe Asp His
1295 1300 1305
Ile Tyr Thr Thr Gln Ser Asp Val Trp Ser Phe Gly Val Leu Leu
1310 1315 1320
Trp Glu Ile Val Thr Leu Gly Gly Asn Pro Tyr Pro Gly Ile Pro
1325 1330 1335
Pro Glu Arg Leu Phe Asn Leu Leu Lys Thr Gly His Arg Met Glu
1340 1345 1350
Arg Pro Asp Asn Cys Ser Glu Glu Met Tyr Arg Leu Met Leu Gln
1355 1360 1365
Cys Trp Lys Gln Glu Pro Asp Lys Arg Pro Val Phe Ala Asp Ile
1370 1375 1380
Ser Lys Asp Leu Glu Lys Met Met Val Lys Arg Arg Asp Tyr Leu
1385 1390 1395
Asp Leu Ala Ala Ser Thr Pro Ser Asp Ser Leu Ile Tyr Asp Asp
1400 1405 1410
Gly Leu Ser Glu Glu Glu Thr Pro Leu Val Asp Cys Asn Asn Ala
1415 1420 1425
Pro Leu Pro Arg Ala Leu Pro Ser Thr Trp Ile Glu Asn Lys Leu
1430 1435 1440
Tyr Gly Met Ser Asp Pro Asn Trp Pro Gly Glu Ser Pro Val Pro
1445 1450 1455
Leu Thr Arg Ala Asp Gly Thr Asn Thr Gly Phe Pro Arg Tyr Pro
1460 1465 1470
Asn Asp Ser Val Tyr Ala Asn Trp Met Leu Ser Pro Ser Ala Ala
1475 1480 1485
Lys Leu Met Asp Thr Phe Asp Ser
1490 1495
<210> 15
<211> 4365
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 15
atgatgagac aggcaccgac agcccgaaag accacaactc ggcgacccaa gcccacgcgc 60
ccagccagta ctggggtggc tggggccagt agctccctgg gcccctctgg ctcagcgtca 120
gcaggtgagc tgagcagcag tgagcccagc accccggctc agactccgct ggcagcaccc 180
atcatcccca cgccggtcct cacctctcct ggagcagtcc ccccgcttcc ttccccatcc 240
aaggaggagg agggactaag ggctcaggtg cgggacctgg aggagaaact agagaccctg 300
agactgaaac gggcagaaga caaagcaaag ctaaaagagc tggagaaaca caaaatccag 360
ctggagcagg tgcaggaatg gaagagcaaa atgcaggagc agcaggccga cctgcagcgg 420
cgcctcaagg aggcgagaaa ggaagccaag gaggcgctgg aggcaaagga acgctatatg 480
gaggagatgg ctgatactgc tgatgccatt gagatggcca ctttggacaa ggagatggct 540
gaagagcggg ctgagtccct gcagcaggag gtggaggcac tgaaggagcg ggtggacgag 600
ctcactactg acttagagat cctcaaggct gagattgaag agaagggctc agatggcgct 660
gcatccagtt atcagctcaa gcagcttgag gagcagaatg cccgcctgaa ggatgccctg 720
gtgaggatgc gggatctttc ttcctcagag aagcaggagc atgtgaagct ccagaagctc 780
atggaaaaga agaaccaaga gctggaagtt gtgaggcaac agcgggagcg tctgcaggag 840
gagctaagcc aggcagagag caccattgat gagctcaagg agcaggtgga tgctgctctg 900
ggtgctgagg agatggtgga gatgctgaca gatcggaacc tgaatctgga agagaaagtg 960
cgcgagttga gggagactgt gggagacttg gaagcgatga atgagatgaa cgatgagctg 1020
caggagaatg cacgtgagac agaactggag ctgcgggagc agctggacat ggcaggcgcg 1080
cgggttcgtg aggcccagaa gcgtgtggag gcagcccagg agacggttgc agactaccag 1140
cagaccatca agaagtaccg ccagctgacc gcccatctac aggatgtgaa tcgggaactg 1200
acaaaccagc aggaagcatc tgtggagagg caacagcagc cacctccaga gacctttgac 1260
ttcaaaatca agtttgctga gactaaggcc catgccaagg caattgagat ggaattgagg 1320
cagatggagg tggcccaggc caatcgacac atgtccctgc tgacagcctt catgcctgac 1380
agcttccttc ggccaggtgg ggaccatgac tgcgttctgg tgctgttgct catgcctcgt 1440
ctcatttgca aggcagagct gatccggaag caggcccagg agaagtttga actaagtgag 1500
aactgttcag agcggcctgg gctgcgagga gctgctgggg agcaactcag ctttgctgct 1560
ggactggtgt actcgctgag cctgctgcag gccacgctac accgctatga gcatgccctc 1620
tctcagtgca gtgtggatgt gtataagaaa gtgggcagcc tgtaccctga gatgagtgcc 1680
catgagcgct ccttggattt cctcattgaa ctgctgcaca aggatcagct ggatgagact 1740
gtcaatgtgg agcctctcac caaggccatc aagtactatc agcatctgta cagcatccac 1800
cttgccgaac agcctgagga ctgtactatg cagctggctg accacattaa gttcacgcag 1860
agtgctctgg actgcatgag tgtggaggta ggacggctgc gtgccttctt gcagggtggg 1920
caggaggcta cagatattgc cctcctgctc cgggatctgg aaacttcatg cagtgacatc 1980
cgccagttct gcaagaagat ccgaaggcga atgccaggga cagatgctcc tgggatccca 2040
gctgcactgg cctttggacc acaggtatct gacacgctcc tagactgcag gaaacacttg 2100
acgtgggtcg tggctgtgct gcaggaggtg gcagctgctg ctgcccagct cattgcccca 2160
ctggcagaga atgaggggct acttgtggct gctctggagg aactggcttt caaagcaagc 2220
gagcagatct atgggacccc ctccagcagc ccctatgagt gtctgcgcca gtcatgcaac 2280
atcctcatca gtaccatgaa caagctggcc acagccatgc aggaggggga gtatgatgca 2340
gagcggcccc ccagcaagcc tccaccggtt gaactgcggg ctgctgccct tcgtgcagag 2400
atcacagatg ctgaaggcct gggtttgaag ctcgaagatc gagagacagt tattaaggag 2460
ttgaagaagt cactcaagat taagggagag gagctaagtg aggccaatgt gcggctgagc 2520
ctcctggaga agaagttgga cagtgctgcc aaggatgcag atgagcgcat cgagaaagtc 2580
cagactcggc tggaggagac ccaggcactg ctgcgaaaga aggagaaaga gtttgaggag 2640
acaatggatg cactccaggc tgacatcgac cagctggagg cagagaaggc agaactaaag 2700
cagcgtctga acagccagtc caaacgcacg attgagggac tccggggccc tcctccttca 2760
ggcattgcta ctctggtctc tggcattgct ggtggagcca tccctgggca ggctccaggg 2820
tctgtgccag gcccagggct ggtgaaggac tcaccactgc tgcttcagca gatctctgcc 2880
atgaggctgc acatctccca gctccagcat gagaacagca tcctcaaggg agcccagatg 2940
aaggcatcct tggcatccct gccccctctg catgttgcaa agctatccca tgagggccct 3000
ggcagtgagt taccagctgg agcgctgtat cgtaagacca gccagctgct ggagacattg 3060
aatcaattga gcacacacac gcacgtagta gacatcactc gcaccagccc tgctgccaag 3120
agcccgtcgg cccaacttat ggagcaagtg gctcagctta agtccctgag tgacaccgtc 3180
gagaagctca aggatgaggt cctcaaggag acagtatctc agcgccctgg agccacagta 3240
cccactgact ttgccacctt cccttcatca gccttcctca gggaggatcc aaagtgggaa 3300
ttccctcgga agaacttggt tcttggaaaa actctaggag aaggcgaatt tggaaaagtg 3360
gtcaaggcaa cggccttcca tctgaaaggc agagcagggt acaccacggt ggccgtgaag 3420
atgctgaaag agaacgcctc cccgagtgag cttcgagacc tgctgtcaga gttcaacgtc 3480
ctgaagcagg tcaaccaccc acatgtcatc aaattgtatg gggcctgcag ccaggatggc 3540
ccgctcctcc tcatcgtgga gtacgccaaa tacggctccc tgcggggctt cctccgcgag 3600
agccgcaaag tggggcctgg ctacctgggc agtggaggca gccgcaactc cagctccctg 3660
gaccacccgg atgagcgggc cctcaccatg ggcgacctca tctcatttgc ctggcagatc 3720
tcacagggga tgcagtatct ggccgagatg aagctcgttc atcgggactt ggcagccaga 3780
aacatcctgg tagctgaggg gcggaagatg aagatttcgg atttcggctt gtcccgagat 3840
gtttatgaag aggattccta cgtgaagagg agccagggtc ggattccagt taaatggatg 3900
gcaattgaat ccctttttga tcatatctac accacgcaaa gtgatgtatg gtcttttggt 3960
gtcctgctgt gggagatcgt gaccctaggg ggaaacccct atcctgggat tcctcctgag 4020
cggctcttca accttctgaa gaccggccac cggatggaga ggccagacaa ctgcagcgag 4080
gagatgtacc gcctgatgct gcaatgctgg aagcaggagc cggacaaaag gccggtgttt 4140
gcggacatca gcaaagacct ggagaagatg atggttaaga ggagagacta cttggacctt 4200
gcggcgtcca ctccatctga ctccctgatt tatgacgacg gcctctcaga ggaggagaca 4260
ccgctggtgg actgtaataa tgcccccctc cctcgagccc tcccttccac atggattgaa 4320
aacaaactct atggtagaat ttcccatgca tttactagat tctag 4365
<210> 16
<211> 1454
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 16
Met Met Arg Gln Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro
1 5 10 15
Lys Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser
20 25 30
Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu
35 40 45
Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr
50 55 60
Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser
65 70 75 80
Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys
85 90 95
Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys
100 105 110
Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys
115 120 125
Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu
130 135 140
Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met
145 150 155 160
Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp
165 170 175
Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu
180 185 190
Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu
195 200 205
Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr
210 215 220
Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu
225 230 235 240
Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys
245 250 255
Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg
260 265 270
Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr
275 280 285
Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu
290 295 300
Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val
305 310 315 320
Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met
325 330 335
Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg
340 345 350
Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg
355 360 365
Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys
370 375 380
Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu
385 390 395 400
Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro
405 410 415
Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala
420 425 430
Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn
435 440 445
Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg
450 455 460
Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg
465 470 475 480
Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe
485 490 495
Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala
500 505 510
Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu
515 520 525
Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser
530 535 540
Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala
545 550 555 560
His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln
565 570 575
Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr
580 585 590
Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys
595 600 605
Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp
610 615 620
Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly
625 630 635 640
Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser
645 650 655
Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro
660 665 670
Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln
675 680 685
Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val
690 695 700
Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro
705 710 715 720
Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala
725 730 735
Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr
740 745 750
Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys
755 760 765
Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro
770 775 780
Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu
785 790 795 800
Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr
805 810 815
Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu
820 825 830
Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser
835 840 845
Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu
850 855 860
Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu
865 870 875 880
Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
885 890 895
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile Glu
900 905 910
Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val Ser Gly
915 920 925
Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser Val Pro Gly
930 935 940
Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln Gln Ile Ser Ala
945 950 955 960
Met Arg Leu His Ile Ser Gln Leu Gln His Glu Asn Ser Ile Leu Lys
965 970 975
Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu Pro Pro Leu His Val
980 985 990
Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu Leu Pro Ala Gly Ala
995 1000 1005
Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu Thr Leu Asn Gln Leu
1010 1015 1020
Ser Thr His Thr His Val Val Asp Ile Thr Arg Thr Ser Pro Ala
1025 1030 1035
Ala Lys Ser Pro Ser Ala Gln Leu Met Glu Gln Val Ala Gln Leu
1040 1045 1050
Lys Ser Leu Ser Asp Thr Val Glu Lys Leu Lys Asp Glu Val Leu
1055 1060 1065
Lys Glu Thr Val Ser Gln Arg Pro Gly Ala Thr Val Pro Thr Asp
1070 1075 1080
Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu Arg Glu Asp Pro Lys
1085 1090 1095
Trp Glu Phe Pro Arg Lys Asn Leu Val Leu Gly Lys Thr Leu Gly
1100 1105 1110
Glu Gly Glu Phe Gly Lys Val Val Lys Ala Thr Ala Phe His Leu
1115 1120 1125
Lys Gly Arg Ala Gly Tyr Thr Thr Val Ala Val Lys Met Leu Lys
1130 1135 1140
Glu Asn Ala Ser Pro Ser Glu Leu Arg Asp Leu Leu Ser Glu Phe
1145 1150 1155
Asn Val Leu Lys Gln Val Asn His Pro His Val Ile Lys Leu Tyr
1160 1165 1170
Gly Ala Cys Ser Gln Asp Gly Pro Leu Leu Leu Ile Val Glu Tyr
1175 1180 1185
Ala Lys Tyr Gly Ser Leu Arg Gly Phe Leu Arg Glu Ser Arg Lys
1190 1195 1200
Val Gly Pro Gly Tyr Leu Gly Ser Gly Gly Ser Arg Asn Ser Ser
1205 1210 1215
Ser Leu Asp His Pro Asp Glu Arg Ala Leu Thr Met Gly Asp Leu
1220 1225 1230
Ile Ser Phe Ala Trp Gln Ile Ser Gln Gly Met Gln Tyr Leu Ala
1235 1240 1245
Glu Met Lys Leu Val His Arg Asp Leu Ala Ala Arg Asn Ile Leu
1250 1255 1260
Val Ala Glu Gly Arg Lys Met Lys Ile Ser Asp Phe Gly Leu Ser
1265 1270 1275
Arg Asp Val Tyr Glu Glu Asp Ser Tyr Val Lys Arg Ser Gln Gly
1280 1285 1290
Arg Ile Pro Val Lys Trp Met Ala Ile Glu Ser Leu Phe Asp His
1295 1300 1305
Ile Tyr Thr Thr Gln Ser Asp Val Trp Ser Phe Gly Val Leu Leu
1310 1315 1320
Trp Glu Ile Val Thr Leu Gly Gly Asn Pro Tyr Pro Gly Ile Pro
1325 1330 1335
Pro Glu Arg Leu Phe Asn Leu Leu Lys Thr Gly His Arg Met Glu
1340 1345 1350
Arg Pro Asp Asn Cys Ser Glu Glu Met Tyr Arg Leu Met Leu Gln
1355 1360 1365
Cys Trp Lys Gln Glu Pro Asp Lys Arg Pro Val Phe Ala Asp Ile
1370 1375 1380
Ser Lys Asp Leu Glu Lys Met Met Val Lys Arg Arg Asp Tyr Leu
1385 1390 1395
Asp Leu Ala Ala Ser Thr Pro Ser Asp Ser Leu Ile Tyr Asp Asp
1400 1405 1410
Gly Leu Ser Glu Glu Glu Thr Pro Leu Val Asp Cys Asn Asn Ala
1415 1420 1425
Pro Leu Pro Arg Ala Leu Pro Ser Thr Trp Ile Glu Asn Lys Leu
1430 1435 1440
Tyr Gly Arg Ile Ser His Ala Phe Thr Arg Phe
1445 1450
<210> 17
<211> 4782
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 17
atgagtgcgg aggcaagcgc ccggcctctg cgggtgggct cccgtgtaga ggtgattgga 60
aaaggccacc gaggcactgt ggcctatgtt ggagccacac tgtttgccac tggcaaatgg 120
gtaggcgtga ttctggatga agcaaagggc aaaaatgatg gaactgttca aggcaggaag 180
tacttcactt gtgatgaagg gcatggcatc tttgtgcgcc agtcccagat ccaggtattt 240
gaagatggag cagatactac ttccccagag acacctgatt cttctgcttc aaaagtcctc 300
aaaagagagg gaactgatac aactgcaaag actagcaaac tgcccacgcg cccagccagt 360
actggggtgg ctggggccag tagctccctg ggcccctctg gctcagcgtc agcaggtgag 420
ctgagcagca gtgagcccag caccccggct cagactccgc tggcagcacc catcatcccc 480
acgccggtcc tcacctctcc tggagcagtc cccccgcttc cttccccatc caaggaggag 540
gagggactaa gggctcaggt gcgggacctg gaggagaaac tagagaccct gagactgaaa 600
cgggcagaag acaaagcaaa gctaaaagag ctggagaaac acaaaatcca gctggagcag 660
gtgcaggaat ggaagagcaa aatgcaggag cagcaggccg acctgcagcg gcgcctcaag 720
gaggcgagaa aggaagccaa ggaggcgctg gaggcaaagg aacgctatat ggaggagatg 780
gctgatactg ctgatgccat tgagatggcc actttggaca aggagatggc tgaagagcgg 840
gctgagtccc tgcagcagga ggtggaggca ctgaaggagc gggtggacga gctcactact 900
gacttagaga tcctcaaggc tgagattgaa gagaagggct cagatggcgc tgcatccagt 960
tatcagctca agcagcttga ggagcagaat gcccgcctga aggatgccct ggtgaggatg 1020
cgggatcttt cttcctcaga gaagcaggag catgtgaagc tccagaagct catggaaaag 1080
aagaaccaag agctggaagt tgtgaggcaa cagcgggagc gtctgcagga ggagctaagc 1140
caggcagaga gcaccattga tgagctcaag gagcaggtgg atgctgctct gggtgctgag 1200
gagatggtgg agatgctgac agatcggaac ctgaatctgg aagagaaagt gcgcgagttg 1260
agggagactg tgggagactt ggaagcgatg aatgagatga acgatgagct gcaggagaat 1320
gcacgtgaga cagaactgga gctgcgggag cagctggaca tggcaggcgc gcgggttcgt 1380
gaggcccaga agcgtgtgga ggcagcccag gagacggttg cagactacca gcagaccatc 1440
aagaagtacc gccagctgac cgcccatcta caggatgtga atcgggaact gacaaaccag 1500
caggaagcat ctgtggagag gcaacagcag ccacctccag agacctttga cttcaaaatc 1560
aagtttgctg agactaaggc ccatgccaag gcaattgaga tggaattgag gcagatggag 1620
gtggcccagg ccaatcgaca catgtccctg ctgacagcct tcatgcctga cagcttcctt 1680
cggccaggtg gggaccatga ctgcgttctg gtgctgttgc tcatgcctcg tctcatttgc 1740
aaggcagagc tgatccggaa gcaggcccag gagaagtttg aactaagtga gaactgttca 1800
gagcggcctg ggctgcgagg agctgctggg gagcaactca gctttgctgc tggactggtg 1860
tactcgctga gcctgctgca ggccacgcta caccgctatg agcatgccct ctctcagtgc 1920
agtgtggatg tgtataagaa agtgggcagc ctgtaccctg agatgagtgc ccatgagcgc 1980
tccttggatt tcctcattga actgctgcac aaggatcagc tggatgagac tgtcaatgtg 2040
gagcctctca ccaaggccat caagtactat cagcatctgt acagcatcca ccttgccgaa 2100
cagcctgagg actgtactat gcagctggct gaccacatta agttcacgca gagtgctctg 2160
gactgcatga gtgtggaggt aggacggctg cgtgccttct tgcagggtgg gcaggaggct 2220
acagatattg ccctcctgct ccgggatctg gaaacttcat gcagtgacat ccgccagttc 2280
tgcaagaaga tccgaaggcg aatgccaggg acagatgctc ctgggatccc agctgcactg 2340
gcctttggac cacaggtatc tgacacgctc ctagactgca ggaaacactt gacgtgggtc 2400
gtggctgtgc tgcaggaggt ggcagctgct gctgcccagc tcattgcccc actggcagag 2460
aatgaggggc tacttgtggc tgctctggag gaactggctt tcaaagcaag cgagcagatc 2520
tatgggaccc cctccagcag cccctatgag tgtctgcgcc agtcatgcaa catcctcatc 2580
agtaccatga acaagctggc cacagccatg caggaggggg agtatgatgc agagcggccc 2640
cccagcaagc ctccaccggt tgaactgcgg gctgctgccc ttcgtgcaga gatcacagat 2700
gctgaaggcc tgggtttgaa gctcgaagat cgagagacag ttattaagga gttgaagaag 2760
tcactcaaga ttaagggaga ggagctaagt gaggccaatg tgcggctgag cctcctggag 2820
aagaagttgg acagtgctgc caaggatgca gatgagcgca tcgagaaagt ccagactcgg 2880
ctggaggaga cccaggcact gctgcgaaag aaggagaaag agtttgagga gacaatggat 2940
gcactccagg ctgacatcga ccagctggag gcagagaagg cagaactaaa gcagcgtctg 3000
aacagccagt ccaaacgcac gattgaggga ctccggggcc ctcctccttc aggcattgct 3060
actctggtct ctggcattgc tggtggagcc atccctgggc aggctccagg gtctgtgcca 3120
ggcccagggc tggtgaagga ctcaccactg ctgcttcagc agatctctgc catgaggctg 3180
cacatctccc agctccagca tgagaacagc atcctcaagg gagcccagat gaaggcatcc 3240
ttggcatccc tgccccctct gcatgttgca aagctatccc atgagggccc tggcagtgag 3300
ttaccagctg gagcgctgta tcgtaagacc agccagctgc tggagacatt gaatcaattg 3360
agcacacaca cgcacgtagt agacatcact cgcaccagcc ctgctgccaa gagcccgtcg 3420
gcccaactta tggagcaagt ggctcagctt aagtccctga gtgacaccgt cgagaagctc 3480
aaggatgagg tcctcaagga gacagtatct cagcgccctg gagccacagt acccactgac 3540
tttgccacct tcccttcatc agccttcctc agggaggatc caaagtggga attccctcgg 3600
aagaacttgg ttcttggaaa aactctagga gaaggcgaat ttggaaaagt ggtcaaggca 3660
acggccttcc atctgaaagg cagagcaggg tacaccacgg tggccgtgaa gatgctgaaa 3720
gagaacgcct ccccgagtga gcttcgagac ctgctgtcag agttcaacgt cctgaagcag 3780
gtcaaccacc cacatgtcat caaattgtat ggggcctgca gccaggatgg cccgctcctc 3840
ctcatcgtgg agtacgccaa atacggctcc ctgcggggct tcctccgcga gagccgcaaa 3900
gtggggcctg gctacctggg cagtggaggc agccgcaact ccagctccct ggaccacccg 3960
gatgagcggg ccctcaccat gggcgacctc atctcatttg cctggcagat ctcacagggg 4020
atgcagtatc tggccgagat gaagctcgtt catcgggact tggcagccag aaacatcctg 4080
gtagctgagg ggcggaagat gaagatttcg gatttcggct tgtcccgaga tgtttatgaa 4140
gaggattcct acgtgaagag gagccagggt cggattccag ttaaatggat ggcaattgaa 4200
tccctttttg atcatatcta caccacgcaa agtgatgtat ggtcttttgg tgtcctgctg 4260
tgggagatcg tgaccctagg gggaaacccc tatcctggga ttcctcctga gcggctcttc 4320
aaccttctga agaccggcca ccggatggag aggccagaca actgcagcga ggagatgtac 4380
cgcctgatgc tgcaatgctg gaagcaggag ccggacaaaa ggccggtgtt tgcggacatc 4440
agcaaagacc tggagaagat gatggttaag aggagagact acttggacct tgcggcgtcc 4500
actccatctg actccctgat ttatgacgac ggcctctcag aggaggagac accgctggtg 4560
gactgtaata atgcccccct ccctcgagcc ctcccttcca catggattga aaacaaactc 4620
tatggcatgt cagacccgaa ctggcctgga gagagtcctg taccactcac gagagctgat 4680
ggcactaaca ctgggtttcc aagatatcca aatgatagtg tatatgctaa ctggatgctt 4740
tcaccctcag cggcaaaatt aatggacacg tttgatagtt aa 4782
<210> 18
<211> 1593
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 18
Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg Val
1 5 10 15
Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly Ala
20 25 30
Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu Ala
35 40 45
Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr Cys
50 55 60
Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val Phe
65 70 75 80
Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser Ala
85 90 95
Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr Ser
100 105 110
Lys Leu Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser
115 120 125
Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser
130 135 140
Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro
145 150 155 160
Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro
165 170 175
Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu
180 185 190
Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu
195 200 205
Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp
210 215 220
Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys
225 230 235 240
Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr
245 250 255
Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu
260 265 270
Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val
275 280 285
Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile
290 295 300
Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser
305 310 315 320
Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala
325 330 335
Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val
340 345 350
Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val
355 360 365
Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser
370 375 380
Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu
385 390 395 400
Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys
405 410 415
Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu
420 425 430
Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu
435 440 445
Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys
450 455 460
Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile
465 470 475 480
Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu
485 490 495
Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro
500 505 510
Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His
515 520 525
Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala
530 535 540
Asn Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu
545 550 555 560
Arg Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro
565 570 575
Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys
580 585 590
Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala
595 600 605
Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser
610 615 620
Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys
625 630 635 640
Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser
645 650 655
Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp
660 665 670
Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys
675 680 685
Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp
690 695 700
Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu
705 710 715 720
Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly
725 730 735
Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr
740 745 750
Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met
755 760 765
Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro
770 775 780
Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val
785 790 795 800
Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala
805 810 815
Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu
820 825 830
Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro
835 840 845
Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn
850 855 860
Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro
865 870 875 880
Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala
885 890 895
Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu
900 905 910
Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu
915 920 925
Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp
930 935 940
Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg
945 950 955 960
Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu
965 970 975
Glu Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu
980 985 990
Lys Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile
995 1000 1005
Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val
1010 1015 1020
Ser Gly Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser
1025 1030 1035
Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln
1040 1045 1050
Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu
1055 1060 1065
Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser
1070 1075 1080
Leu Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly
1085 1090 1095
Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu
1100 1105 1110
Leu Glu Thr Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp
1115 1120 1125
Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu
1130 1135 1140
Met Glu Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu
1145 1150 1155
Lys Leu Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro
1160 1165 1170
Gly Ala Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala
1175 1180 1185
Phe Leu Arg Glu Asp Pro Lys Trp Glu Phe Pro Arg Lys Asn Leu
1190 1195 1200
Val Leu Gly Lys Thr Leu Gly Glu Gly Glu Phe Gly Lys Val Val
1205 1210 1215
Lys Ala Thr Ala Phe His Leu Lys Gly Arg Ala Gly Tyr Thr Thr
1220 1225 1230
Val Ala Val Lys Met Leu Lys Glu Asn Ala Ser Pro Ser Glu Leu
1235 1240 1245
Arg Asp Leu Leu Ser Glu Phe Asn Val Leu Lys Gln Val Asn His
1250 1255 1260
Pro His Val Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp Gly Pro
1265 1270 1275
Leu Leu Leu Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu Arg Gly
1280 1285 1290
Phe Leu Arg Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu Gly Ser
1295 1300 1305
Gly Gly Ser Arg Asn Ser Ser Ser Leu Asp His Pro Asp Glu Arg
1310 1315 1320
Ala Leu Thr Met Gly Asp Leu Ile Ser Phe Ala Trp Gln Ile Ser
1325 1330 1335
Gln Gly Met Gln Tyr Leu Ala Glu Met Lys Leu Val His Arg Asp
1340 1345 1350
Leu Ala Ala Arg Asn Ile Leu Val Ala Glu Gly Arg Lys Met Lys
1355 1360 1365
Ile Ser Asp Phe Gly Leu Ser Arg Asp Val Tyr Glu Glu Asp Ser
1370 1375 1380
Tyr Val Lys Arg Ser Gln Gly Arg Ile Pro Val Lys Trp Met Ala
1385 1390 1395
Ile Glu Ser Leu Phe Asp His Ile Tyr Thr Thr Gln Ser Asp Val
1400 1405 1410
Trp Ser Phe Gly Val Leu Leu Trp Glu Ile Val Thr Leu Gly Gly
1415 1420 1425
Asn Pro Tyr Pro Gly Ile Pro Pro Glu Arg Leu Phe Asn Leu Leu
1430 1435 1440
Lys Thr Gly His Arg Met Glu Arg Pro Asp Asn Cys Ser Glu Glu
1445 1450 1455
Met Tyr Arg Leu Met Leu Gln Cys Trp Lys Gln Glu Pro Asp Lys
1460 1465 1470
Arg Pro Val Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys Met Met
1475 1480 1485
Val Lys Arg Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr Pro Ser
1490 1495 1500
Asp Ser Leu Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu Thr Pro
1505 1510 1515
Leu Val Asp Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu Pro Ser
1520 1525 1530
Thr Trp Ile Glu Asn Lys Leu Tyr Gly Met Ser Asp Pro Asn Trp
1535 1540 1545
Pro Gly Glu Ser Pro Val Pro Leu Thr Arg Ala Asp Gly Thr Asn
1550 1555 1560
Thr Gly Phe Pro Arg Tyr Pro Asn Asp Ser Val Tyr Ala Asn Trp
1565 1570 1575
Met Leu Ser Pro Ser Ala Ala Lys Leu Met Asp Thr Phe Asp Ser
1580 1585 1590
<210> 19
<211> 4656
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 19
atgagtgcgg aggcaagcgc ccggcctctg cgggtgggct cccgtgtaga ggtgattgga 60
aaaggccacc gaggcactgt ggcctatgtt ggagccacac tgtttgccac tggcaaatgg 120
gtaggcgtga ttctggatga agcaaagggc aaaaatgatg gaactgttca aggcaggaag 180
tacttcactt gtgatgaagg gcatggcatc tttgtgcgcc agtcccagat ccaggtattt 240
gaagatggag cagatactac ttccccagag acacctgatt cttctgcttc aaaagtcctc 300
aaaagagagg gaactgatac aactgcaaag actagcaaac tgcccacgcg cccagccagt 360
actggggtgg ctggggccag tagctccctg ggcccctctg gctcagcgtc agcaggtgag 420
ctgagcagca gtgagcccag caccccggct cagactccgc tggcagcacc catcatcccc 480
acgccggtcc tcacctctcc tggagcagtc cccccgcttc cttccccatc caaggaggag 540
gagggactaa gggctcaggt gcgggacctg gaggagaaac tagagaccct gagactgaaa 600
cgggcagaag acaaagcaaa gctaaaagag ctggagaaac acaaaatcca gctggagcag 660
gtgcaggaat ggaagagcaa aatgcaggag cagcaggccg acctgcagcg gcgcctcaag 720
gaggcgagaa aggaagccaa ggaggcgctg gaggcaaagg aacgctatat ggaggagatg 780
gctgatactg ctgatgccat tgagatggcc actttggaca aggagatggc tgaagagcgg 840
gctgagtccc tgcagcagga ggtggaggca ctgaaggagc gggtggacga gctcactact 900
gacttagaga tcctcaaggc tgagattgaa gagaagggct cagatggcgc tgcatccagt 960
tatcagctca agcagcttga ggagcagaat gcccgcctga aggatgccct ggtgaggatg 1020
cgggatcttt cttcctcaga gaagcaggag catgtgaagc tccagaagct catggaaaag 1080
aagaaccaag agctggaagt tgtgaggcaa cagcgggagc gtctgcagga ggagctaagc 1140
caggcagaga gcaccattga tgagctcaag gagcaggtgg atgctgctct gggtgctgag 1200
gagatggtgg agatgctgac agatcggaac ctgaatctgg aagagaaagt gcgcgagttg 1260
agggagactg tgggagactt ggaagcgatg aatgagatga acgatgagct gcaggagaat 1320
gcacgtgaga cagaactgga gctgcgggag cagctggaca tggcaggcgc gcgggttcgt 1380
gaggcccaga agcgtgtgga ggcagcccag gagacggttg cagactacca gcagaccatc 1440
aagaagtacc gccagctgac cgcccatcta caggatgtga atcgggaact gacaaaccag 1500
caggaagcat ctgtggagag gcaacagcag ccacctccag agacctttga cttcaaaatc 1560
aagtttgctg agactaaggc ccatgccaag gcaattgaga tggaattgag gcagatggag 1620
gtggcccagg ccaatcgaca catgtccctg ctgacagcct tcatgcctga cagcttcctt 1680
cggccaggtg gggaccatga ctgcgttctg gtgctgttgc tcatgcctcg tctcatttgc 1740
aaggcagagc tgatccggaa gcaggcccag gagaagtttg aactaagtga gaactgttca 1800
gagcggcctg ggctgcgagg agctgctggg gagcaactca gctttgctgc tggactggtg 1860
tactcgctga gcctgctgca ggccacgcta caccgctatg agcatgccct ctctcagtgc 1920
agtgtggatg tgtataagaa agtgggcagc ctgtaccctg agatgagtgc ccatgagcgc 1980
tccttggatt tcctcattga actgctgcac aaggatcagc tggatgagac tgtcaatgtg 2040
gagcctctca ccaaggccat caagtactat cagcatctgt acagcatcca ccttgccgaa 2100
cagcctgagg actgtactat gcagctggct gaccacatta agttcacgca gagtgctctg 2160
gactgcatga gtgtggaggt aggacggctg cgtgccttct tgcagggtgg gcaggaggct 2220
acagatattg ccctcctgct ccgggatctg gaaacttcat gcagtgacat ccgccagttc 2280
tgcaagaaga tccgaaggcg aatgccaggg acagatgctc ctgggatccc agctgcactg 2340
gcctttggac cacaggtatc tgacacgctc ctagactgca ggaaacactt gacgtgggtc 2400
gtggctgtgc tgcaggaggt ggcagctgct gctgcccagc tcattgcccc actggcagag 2460
aatgaggggc tacttgtggc tgctctggag gaactggctt tcaaagcaag cgagcagatc 2520
tatgggaccc cctccagcag cccctatgag tgtctgcgcc agtcatgcaa catcctcatc 2580
agtaccatga acaagctggc cacagccatg caggaggggg agtatgatgc agagcggccc 2640
cccagcaagc ctccaccggt tgaactgcgg gctgctgccc ttcgtgcaga gatcacagat 2700
gctgaaggcc tgggtttgaa gctcgaagat cgagagacag ttattaagga gttgaagaag 2760
tcactcaaga ttaagggaga ggagctaagt gaggccaatg tgcggctgag cctcctggag 2820
aagaagttgg acagtgctgc caaggatgca gatgagcgca tcgagaaagt ccagactcgg 2880
ctggaggaga cccaggcact gctgcgaaag aaggagaaag agtttgagga gacaatggat 2940
gcactccagg ctgacatcga ccagctggag gcagagaagg cagaactaaa gcagcgtctg 3000
aacagccagt ccaaacgcac gattgaggga ctccggggcc ctcctccttc aggcattgct 3060
actctggtct ctggcattgc tggtggagcc atccctgggc aggctccagg gtctgtgcca 3120
ggcccagggc tggtgaagga ctcaccactg ctgcttcagc agatctctgc catgaggctg 3180
cacatctccc agctccagca tgagaacagc atcctcaagg gagcccagat gaaggcatcc 3240
ttggcatccc tgccccctct gcatgttgca aagctatccc atgagggccc tggcagtgag 3300
ttaccagctg gagcgctgta tcgtaagacc agccagctgc tggagacatt gaatcaattg 3360
agcacacaca cgcacgtagt agacatcact cgcaccagcc ctgctgccaa gagcccgtcg 3420
gcccaactta tggagcaagt ggctcagctt aagtccctga gtgacaccgt cgagaagctc 3480
aaggatgagg tcctcaagga gacagtatct cagcgccctg gagccacagt acccactgac 3540
tttgccacct tcccttcatc agccttcctc agggaggatc caaagtggga attccctcgg 3600
aagaacttgg ttcttggaaa aactctagga gaaggcgaat ttggaaaagt ggtcaaggca 3660
acggccttcc atctgaaagg cagagcaggg tacaccacgg tggccgtgaa gatgctgaaa 3720
gagaacgcct ccccgagtga gcttcgagac ctgctgtcag agttcaacgt cctgaagcag 3780
gtcaaccacc cacatgtcat caaattgtat ggggcctgca gccaggatgg cccgctcctc 3840
ctcatcgtgg agtacgccaa atacggctcc ctgcggggct tcctccgcga gagccgcaaa 3900
gtggggcctg gctacctggg cagtggaggc agccgcaact ccagctccct ggaccacccg 3960
gatgagcggg ccctcaccat gggcgacctc atctcatttg cctggcagat ctcacagggg 4020
atgcagtatc tggccgagat gaagctcgtt catcgggact tggcagccag aaacatcctg 4080
gtagctgagg ggcggaagat gaagatttcg gatttcggct tgtcccgaga tgtttatgaa 4140
gaggattcct acgtgaagag gagccagggt cggattccag ttaaatggat ggcaattgaa 4200
tccctttttg atcatatcta caccacgcaa agtgatgtat ggtcttttgg tgtcctgctg 4260
tgggagatcg tgaccctagg gggaaacccc tatcctggga ttcctcctga gcggctcttc 4320
aaccttctga agaccggcca ccggatggag aggccagaca actgcagcga ggagatgtac 4380
cgcctgatgc tgcaatgctg gaagcaggag ccggacaaaa ggccggtgtt tgcggacatc 4440
agcaaagacc tggagaagat gatggttaag aggagagact acttggacct tgcggcgtcc 4500
actccatctg actccctgat ttatgacgac ggcctctcag aggaggagac accgctggtg 4560
gactgtaata atgcccccct ccctcgagcc ctcccttcca catggattga aaacaaactc 4620
tatggtagaa tttcccatgc atttactaga ttctag 4656
<210> 20
<211> 1551
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 20
Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg Val
1 5 10 15
Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly Ala
20 25 30
Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu Ala
35 40 45
Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr Cys
50 55 60
Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val Phe
65 70 75 80
Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser Ala
85 90 95
Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr Ser
100 105 110
Lys Leu Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser
115 120 125
Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser
130 135 140
Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro
145 150 155 160
Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro
165 170 175
Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu
180 185 190
Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu
195 200 205
Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp
210 215 220
Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys
225 230 235 240
Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr
245 250 255
Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu
260 265 270
Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val
275 280 285
Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile
290 295 300
Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser
305 310 315 320
Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala
325 330 335
Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val
340 345 350
Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val
355 360 365
Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser
370 375 380
Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu
385 390 395 400
Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys
405 410 415
Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu
420 425 430
Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu
435 440 445
Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys
450 455 460
Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile
465 470 475 480
Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu
485 490 495
Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro
500 505 510
Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His
515 520 525
Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala
530 535 540
Asn Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu
545 550 555 560
Arg Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro
565 570 575
Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys
580 585 590
Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala
595 600 605
Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser
610 615 620
Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys
625 630 635 640
Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser
645 650 655
Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp
660 665 670
Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys
675 680 685
Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp
690 695 700
Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu
705 710 715 720
Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly
725 730 735
Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr
740 745 750
Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met
755 760 765
Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro
770 775 780
Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val
785 790 795 800
Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala
805 810 815
Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu
820 825 830
Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro
835 840 845
Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn
850 855 860
Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro
865 870 875 880
Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala
885 890 895
Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu
900 905 910
Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu
915 920 925
Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp
930 935 940
Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg
945 950 955 960
Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu
965 970 975
Glu Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu
980 985 990
Lys Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile
995 1000 1005
Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val
1010 1015 1020
Ser Gly Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser
1025 1030 1035
Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln
1040 1045 1050
Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu
1055 1060 1065
Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser
1070 1075 1080
Leu Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly
1085 1090 1095
Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu
1100 1105 1110
Leu Glu Thr Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp
1115 1120 1125
Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu
1130 1135 1140
Met Glu Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu
1145 1150 1155
Lys Leu Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro
1160 1165 1170
Gly Ala Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala
1175 1180 1185
Phe Leu Arg Glu Asp Pro Lys Trp Glu Phe Pro Arg Lys Asn Leu
1190 1195 1200
Val Leu Gly Lys Thr Leu Gly Glu Gly Glu Phe Gly Lys Val Val
1205 1210 1215
Lys Ala Thr Ala Phe His Leu Lys Gly Arg Ala Gly Tyr Thr Thr
1220 1225 1230
Val Ala Val Lys Met Leu Lys Glu Asn Ala Ser Pro Ser Glu Leu
1235 1240 1245
Arg Asp Leu Leu Ser Glu Phe Asn Val Leu Lys Gln Val Asn His
1250 1255 1260
Pro His Val Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp Gly Pro
1265 1270 1275
Leu Leu Leu Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu Arg Gly
1280 1285 1290
Phe Leu Arg Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu Gly Ser
1295 1300 1305
Gly Gly Ser Arg Asn Ser Ser Ser Leu Asp His Pro Asp Glu Arg
1310 1315 1320
Ala Leu Thr Met Gly Asp Leu Ile Ser Phe Ala Trp Gln Ile Ser
1325 1330 1335
Gln Gly Met Gln Tyr Leu Ala Glu Met Lys Leu Val His Arg Asp
1340 1345 1350
Leu Ala Ala Arg Asn Ile Leu Val Ala Glu Gly Arg Lys Met Lys
1355 1360 1365
Ile Ser Asp Phe Gly Leu Ser Arg Asp Val Tyr Glu Glu Asp Ser
1370 1375 1380
Tyr Val Lys Arg Ser Gln Gly Arg Ile Pro Val Lys Trp Met Ala
1385 1390 1395
Ile Glu Ser Leu Phe Asp His Ile Tyr Thr Thr Gln Ser Asp Val
1400 1405 1410
Trp Ser Phe Gly Val Leu Leu Trp Glu Ile Val Thr Leu Gly Gly
1415 1420 1425
Asn Pro Tyr Pro Gly Ile Pro Pro Glu Arg Leu Phe Asn Leu Leu
1430 1435 1440
Lys Thr Gly His Arg Met Glu Arg Pro Asp Asn Cys Ser Glu Glu
1445 1450 1455
Met Tyr Arg Leu Met Leu Gln Cys Trp Lys Gln Glu Pro Asp Lys
1460 1465 1470
Arg Pro Val Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys Met Met
1475 1480 1485
Val Lys Arg Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr Pro Ser
1490 1495 1500
Asp Ser Leu Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu Thr Pro
1505 1510 1515
Leu Val Asp Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu Pro Ser
1520 1525 1530
Thr Trp Ile Glu Asn Lys Leu Tyr Gly Arg Ile Ser His Ala Phe
1535 1540 1545
Thr Arg Phe
1550
<210> 21
<211> 4887
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 21
atggcacaga gcaagaggca cgtgtacagc cggacgccca gcggcagcag gatgagtgcg 60
gaggcaagcg cccggcctct gcgggtgggc tcccgtgtag aggtgattgg aaaaggccac 120
cgaggcactg tggcctatgt tggagccaca ctgtttgcca ctggcaaatg ggtaggcgtg 180
attctggatg aagcaaaggg caaaaatgat ggaactgttc aaggcaggaa gtacttcact 240
tgtgatgaag ggcatggcat ctttgtgcgc cagtcccaga tccaggtatt tgaagatgga 300
gcagatacta cttccccaga gacacctgat tcttctgctt caaaagtcct caaaagagag 360
ggaactgata caactgcaaa gactagcaaa ctggcaccga cagcccgaaa gaccacaact 420
cggcgaccca agcccacgcg cccagccagt actggggtgg ctggggccag tagctccctg 480
ggcccctctg gctcagcgtc agcaggtgag ctgagcagca gtgagcccag caccccggct 540
cagactccgc tggcagcacc catcatcccc acgccggtcc tcacctctcc tggagcagtc 600
cccccgcttc cttccccatc caaggaggag gagggactaa gggctcaggt gcgggacctg 660
gaggagaaac tagagaccct gagactgaaa cgggcagaag acaaagcaaa gctaaaagag 720
ctggagaaac acaaaatcca gctggagcag gtgcaggaat ggaagagcaa aatgcaggag 780
cagcaggccg acctgcagcg gcgcctcaag gaggcgagaa aggaagccaa ggaggcgctg 840
gaggcaaagg aacgctatat ggaggagatg gctgatactg ctgatgccat tgagatggcc 900
actttggaca aggagatggc tgaagagcgg gctgagtccc tgcagcagga ggtggaggca 960
ctgaaggagc gggtggacga gctcactact gacttagaga tcctcaaggc tgagattgaa 1020
gagaagggct cagatggcgc tgcatccagt tatcagctca agcagcttga ggagcagaat 1080
gcccgcctga aggatgccct ggtgaggatg cgggatcttt cttcctcaga gaagcaggag 1140
catgtgaagc tccagaagct catggaaaag aagaaccaag agctggaagt tgtgaggcaa 1200
cagcgggagc gtctgcagga ggagctaagc caggcagaga gcaccattga tgagctcaag 1260
gagcaggtgg atgctgctct gggtgctgag gagatggtgg agatgctgac agatcggaac 1320
ctgaatctgg aagagaaagt gcgcgagttg agggagactg tgggagactt ggaagcgatg 1380
aatgagatga acgatgagct gcaggagaat gcacgtgaga cagaactgga gctgcgggag 1440
cagctggaca tggcaggcgc gcgggttcgt gaggcccaga agcgtgtgga ggcagcccag 1500
gagacggttg cagactacca gcagaccatc aagaagtacc gccagctgac cgcccatcta 1560
caggatgtga atcgggaact gacaaaccag caggaagcat ctgtggagag gcaacagcag 1620
ccacctccag agacctttga cttcaaaatc aagtttgctg agactaaggc ccatgccaag 1680
gcaattgaga tggaattgag gcagatggag gtggcccagg ccaatcgaca catgtccctg 1740
ctgacagcct tcatgcctga cagcttcctt cggccaggtg gggaccatga ctgcgttctg 1800
gtgctgttgc tcatgcctcg tctcatttgc aaggcagagc tgatccggaa gcaggcccag 1860
gagaagtttg aactaagtga gaactgttca gagcggcctg ggctgcgagg agctgctggg 1920
gagcaactca gctttgctgc tggactggtg tactcgctga gcctgctgca ggccacgcta 1980
caccgctatg agcatgccct ctctcagtgc agtgtggatg tgtataagaa agtgggcagc 2040
ctgtaccctg agatgagtgc ccatgagcgc tccttggatt tcctcattga actgctgcac 2100
aaggatcagc tggatgagac tgtcaatgtg gagcctctca ccaaggccat caagtactat 2160
cagcatctgt acagcatcca ccttgccgaa cagcctgagg actgtactat gcagctggct 2220
gaccacatta agttcacgca gagtgctctg gactgcatga gtgtggaggt aggacggctg 2280
cgtgccttct tgcagggtgg gcaggaggct acagatattg ccctcctgct ccgggatctg 2340
gaaacttcat gcagtgacat ccgccagttc tgcaagaaga tccgaaggcg aatgccaggg 2400
acagatgctc ctgggatccc agctgcactg gcctttggac cacaggtatc tgacacgctc 2460
ctagactgca ggaaacactt gacgtgggtc gtggctgtgc tgcaggaggt ggcagctgct 2520
gctgcccagc tcattgcccc actggcagag aatgaggggc tacttgtggc tgctctggag 2580
gaactggctt tcaaagcaag cgagcagatc tatgggaccc cctccagcag cccctatgag 2640
tgtctgcgcc agtcatgcaa catcctcatc agtaccatga acaagctggc cacagccatg 2700
caggaggggg agtatgatgc agagcggccc cccagcaagc ctccaccggt tgaactgcgg 2760
gctgctgccc ttcgtgcaga gatcacagat gctgaaggcc tgggtttgaa gctcgaagat 2820
cgagagacag ttattaagga gttgaagaag tcactcaaga ttaagggaga ggagctaagt 2880
gaggccaatg tgcggctgag cctcctggag aagaagttgg acagtgctgc caaggatgca 2940
gatgagcgca tcgagaaagt ccagactcgg ctggaggaga cccaggcact gctgcgaaag 3000
aaggagaaag agtttgagga gacaatggat gcactccagg ctgacatcga ccagctggag 3060
gcagagaagg cagaactaaa gcagcgtctg aacagccagt ccaaacgcac gattgaggga 3120
ctccggggcc ctcctccttc aggcattgct actctggtct ctggcattgc tggtgaagaa 3180
cagcagcgag gagccatccc tgggcaggct ccagggtctg tgccaggccc agggctggtg 3240
aaggactcac cactgctgct tcagcagatc tctgccatga ggctgcacat ctcccagctc 3300
cagcatgaga acagcatcct caagggagcc cagatgaagg catccttggc atccctgccc 3360
cctctgcatg ttgcaaagct atcccatgag ggccctggca gtgagttacc agctggagcg 3420
ctgtatcgta agaccagcca gctgctggag acattgaatc aattgagcac acacacgcac 3480
gtagtagaca tcactcgcac cagccctgct gccaagagcc cgtcggccca acttatggag 3540
caagtggctc agcttaagtc cctgagtgac accgtcgaga agctcaagga tgaggtcctc 3600
aaggagacag tatctcagcg ccctggagcc acagtaccca ctgactttgc caccttccct 3660
tcatcagcct tcctcaggga ggatccaaag tgggaattcc ctcggaagaa cttggttctt 3720
ggaaaaactc taggagaagg cgaatttgga aaagtggtca aggcaacggc cttccatctg 3780
aaaggcagag cagggtacac cacggtggcc gtgaagatgc tgaaagagaa cgcctccccg 3840
agtgagcttc gagacctgct gtcagagttc aacgtcctga agcaggtcaa ccacccacat 3900
gtcatcaaat tgtatggggc ctgcagccag gatggcccgc tcctcctcat cgtggagtac 3960
gccaaatacg gctccctgcg gggcttcctc cgcgagagcc gcaaagtggg gcctggctac 4020
ctgggcagtg gaggcagccg caactccagc tccctggacc acccggatga gcgggccctc 4080
accatgggcg acctcatctc atttgcctgg cagatctcac aggggatgca gtatctggcc 4140
gagatgaagc tcgttcatcg ggacttggca gccagaaaca tcctggtagc tgaggggcgg 4200
aagatgaaga tttcggattt cggcttgtcc cgagatgttt atgaagagga ttcctacgtg 4260
aagaggagcc agggtcggat tccagttaaa tggatggcaa ttgaatccct ttttgatcat 4320
atctacacca cgcaaagtga tgtatggtct tttggtgtcc tgctgtggga gatcgtgacc 4380
ctagggggaa acccctatcc tgggattcct cctgagcggc tcttcaacct tctgaagacc 4440
ggccaccgga tggagaggcc agacaactgc agcgaggaga tgtaccgcct gatgctgcaa 4500
tgctggaagc aggagccgga caaaaggccg gtgtttgcgg acatcagcaa agacctggag 4560
aagatgatgg ttaagaggag agactacttg gaccttgcgg cgtccactcc atctgactcc 4620
ctgatttatg acgacggcct ctcagaggag gagacaccgc tggtggactg taataatgcc 4680
cccctccctc gagccctccc ttccacatgg attgaaaaca aactctatgg catgtcagac 4740
ccgaactggc ctggagagag tcctgtacca ctcacgagag ctgatggcac taacactggg 4800
tttccaagat atccaaatga tagtgtatat gctaactgga tgctttcacc ctcagcggca 4860
aaattaatgg acacgtttga tagttaa 4887
<210> 22
<211> 1628
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 22
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro Lys
130 135 140
Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser Leu
145 150 155 160
Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu Pro
165 170 175
Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr Pro
180 185 190
Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser Lys
195 200 205
Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys Leu
210 215 220
Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys Glu
225 230 235 240
Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys Ser
245 250 255
Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu Ala
260 265 270
Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met Glu
275 280 285
Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp Lys
290 295 300
Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu Ala
305 310 315 320
Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu Lys
325 330 335
Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr Gln
340 345 350
Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu Val
355 360 365
Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys Leu
370 375 380
Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg Gln
385 390 395 400
Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr Ile
405 410 415
Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu Met
420 425 430
Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val Arg
435 440 445
Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met Asn
450 455 460
Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg Glu
465 470 475 480
Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg Val
485 490 495
Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys Lys
500 505 510
Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu Thr
515 520 525
Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro Glu
530 535 540
Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala Lys
545 550 555 560
Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn Arg
565 570 575
His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg Pro
580 585 590
Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg Leu
595 600 605
Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe Glu
610 615 620
Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala Gly
625 630 635 640
Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu Leu
645 650 655
Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser Val
660 665 670
Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala His
675 680 685
Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln Leu
690 695 700
Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr Tyr
705 710 715 720
Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys Thr
725 730 735
Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp Cys
740 745 750
Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly Gln
755 760 765
Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser Cys
770 775 780
Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro Gly
785 790 795 800
Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln Val
805 810 815
Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val Ala
820 825 830
Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro Leu
835 840 845
Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala Phe
850 855 860
Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr Glu
865 870 875 880
Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys Leu
885 890 895
Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro Ser
900 905 910
Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu Ile
915 920 925
Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr Val
930 935 940
Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu Ser
945 950 955 960
Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser Ala
965 970 975
Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu Glu
980 985 990
Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu Thr
995 1000 1005
Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
1010 1015 1020
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile
1025 1030 1035
Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val
1040 1045 1050
Ser Gly Ile Ala Gly Glu Glu Gln Gln Arg Gly Ala Ile Pro Gly
1055 1060 1065
Gln Ala Pro Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser
1070 1075 1080
Pro Leu Leu Leu Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser
1085 1090 1095
Gln Leu Gln His Glu Asn Ser Ile Leu Lys Gly Ala Gln Met Lys
1100 1105 1110
Ala Ser Leu Ala Ser Leu Pro Pro Leu His Val Ala Lys Leu Ser
1115 1120 1125
His Glu Gly Pro Gly Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg
1130 1135 1140
Lys Thr Ser Gln Leu Leu Glu Thr Leu Asn Gln Leu Ser Thr His
1145 1150 1155
Thr His Val Val Asp Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser
1160 1165 1170
Pro Ser Ala Gln Leu Met Glu Gln Val Ala Gln Leu Lys Ser Leu
1175 1180 1185
Ser Asp Thr Val Glu Lys Leu Lys Asp Glu Val Leu Lys Glu Thr
1190 1195 1200
Val Ser Gln Arg Pro Gly Ala Thr Val Pro Thr Asp Phe Ala Thr
1205 1210 1215
Phe Pro Ser Ser Ala Phe Leu Arg Glu Asp Pro Lys Trp Glu Phe
1220 1225 1230
Pro Arg Lys Asn Leu Val Leu Gly Lys Thr Leu Gly Glu Gly Glu
1235 1240 1245
Phe Gly Lys Val Val Lys Ala Thr Ala Phe His Leu Lys Gly Arg
1250 1255 1260
Ala Gly Tyr Thr Thr Val Ala Val Lys Met Leu Lys Glu Asn Ala
1265 1270 1275
Ser Pro Ser Glu Leu Arg Asp Leu Leu Ser Glu Phe Asn Val Leu
1280 1285 1290
Lys Gln Val Asn His Pro His Val Ile Lys Leu Tyr Gly Ala Cys
1295 1300 1305
Ser Gln Asp Gly Pro Leu Leu Leu Ile Val Glu Tyr Ala Lys Tyr
1310 1315 1320
Gly Ser Leu Arg Gly Phe Leu Arg Glu Ser Arg Lys Val Gly Pro
1325 1330 1335
Gly Tyr Leu Gly Ser Gly Gly Ser Arg Asn Ser Ser Ser Leu Asp
1340 1345 1350
His Pro Asp Glu Arg Ala Leu Thr Met Gly Asp Leu Ile Ser Phe
1355 1360 1365
Ala Trp Gln Ile Ser Gln Gly Met Gln Tyr Leu Ala Glu Met Lys
1370 1375 1380
Leu Val His Arg Asp Leu Ala Ala Arg Asn Ile Leu Val Ala Glu
1385 1390 1395
Gly Arg Lys Met Lys Ile Ser Asp Phe Gly Leu Ser Arg Asp Val
1400 1405 1410
Tyr Glu Glu Asp Ser Tyr Val Lys Arg Ser Gln Gly Arg Ile Pro
1415 1420 1425
Val Lys Trp Met Ala Ile Glu Ser Leu Phe Asp His Ile Tyr Thr
1430 1435 1440
Thr Gln Ser Asp Val Trp Ser Phe Gly Val Leu Leu Trp Glu Ile
1445 1450 1455
Val Thr Leu Gly Gly Asn Pro Tyr Pro Gly Ile Pro Pro Glu Arg
1460 1465 1470
Leu Phe Asn Leu Leu Lys Thr Gly His Arg Met Glu Arg Pro Asp
1475 1480 1485
Asn Cys Ser Glu Glu Met Tyr Arg Leu Met Leu Gln Cys Trp Lys
1490 1495 1500
Gln Glu Pro Asp Lys Arg Pro Val Phe Ala Asp Ile Ser Lys Asp
1505 1510 1515
Leu Glu Lys Met Met Val Lys Arg Arg Asp Tyr Leu Asp Leu Ala
1520 1525 1530
Ala Ser Thr Pro Ser Asp Ser Leu Ile Tyr Asp Asp Gly Leu Ser
1535 1540 1545
Glu Glu Glu Thr Pro Leu Val Asp Cys Asn Asn Ala Pro Leu Pro
1550 1555 1560
Arg Ala Leu Pro Ser Thr Trp Ile Glu Asn Lys Leu Tyr Gly Met
1565 1570 1575
Ser Asp Pro Asn Trp Pro Gly Glu Ser Pro Val Pro Leu Thr Arg
1580 1585 1590
Ala Asp Gly Thr Asn Thr Gly Phe Pro Arg Tyr Pro Asn Asp Ser
1595 1600 1605
Val Tyr Ala Asn Trp Met Leu Ser Pro Ser Ala Ala Lys Leu Met
1610 1615 1620
Asp Thr Phe Asp Ser
1625
<210> 23
<211> 4761
<212> DNA
<213> Artificial Sequence
<220>
<223> coding sequence of fusion peptide
<400> 23
atggcacaga gcaagaggca cgtgtacagc cggacgccca gcggcagcag gatgagtgcg 60
gaggcaagcg cccggcctct gcgggtgggc tcccgtgtag aggtgattgg aaaaggccac 120
cgaggcactg tggcctatgt tggagccaca ctgtttgcca ctggcaaatg ggtaggcgtg 180
attctggatg aagcaaaggg caaaaatgat ggaactgttc aaggcaggaa gtacttcact 240
tgtgatgaag ggcatggcat ctttgtgcgc cagtcccaga tccaggtatt tgaagatgga 300
gcagatacta cttccccaga gacacctgat tcttctgctt caaaagtcct caaaagagag 360
ggaactgata caactgcaaa gactagcaaa ctggcaccga cagcccgaaa gaccacaact 420
cggcgaccca agcccacgcg cccagccagt actggggtgg ctggggccag tagctccctg 480
ggcccctctg gctcagcgtc agcaggtgag ctgagcagca gtgagcccag caccccggct 540
cagactccgc tggcagcacc catcatcccc acgccggtcc tcacctctcc tggagcagtc 600
cccccgcttc cttccccatc caaggaggag gagggactaa gggctcaggt gcgggacctg 660
gaggagaaac tagagaccct gagactgaaa cgggcagaag acaaagcaaa gctaaaagag 720
ctggagaaac acaaaatcca gctggagcag gtgcaggaat ggaagagcaa aatgcaggag 780
cagcaggccg acctgcagcg gcgcctcaag gaggcgagaa aggaagccaa ggaggcgctg 840
gaggcaaagg aacgctatat ggaggagatg gctgatactg ctgatgccat tgagatggcc 900
actttggaca aggagatggc tgaagagcgg gctgagtccc tgcagcagga ggtggaggca 960
ctgaaggagc gggtggacga gctcactact gacttagaga tcctcaaggc tgagattgaa 1020
gagaagggct cagatggcgc tgcatccagt tatcagctca agcagcttga ggagcagaat 1080
gcccgcctga aggatgccct ggtgaggatg cgggatcttt cttcctcaga gaagcaggag 1140
catgtgaagc tccagaagct catggaaaag aagaaccaag agctggaagt tgtgaggcaa 1200
cagcgggagc gtctgcagga ggagctaagc caggcagaga gcaccattga tgagctcaag 1260
gagcaggtgg atgctgctct gggtgctgag gagatggtgg agatgctgac agatcggaac 1320
ctgaatctgg aagagaaagt gcgcgagttg agggagactg tgggagactt ggaagcgatg 1380
aatgagatga acgatgagct gcaggagaat gcacgtgaga cagaactgga gctgcgggag 1440
cagctggaca tggcaggcgc gcgggttcgt gaggcccaga agcgtgtgga ggcagcccag 1500
gagacggttg cagactacca gcagaccatc aagaagtacc gccagctgac cgcccatcta 1560
caggatgtga atcgggaact gacaaaccag caggaagcat ctgtggagag gcaacagcag 1620
ccacctccag agacctttga cttcaaaatc aagtttgctg agactaaggc ccatgccaag 1680
gcaattgaga tggaattgag gcagatggag gtggcccagg ccaatcgaca catgtccctg 1740
ctgacagcct tcatgcctga cagcttcctt cggccaggtg gggaccatga ctgcgttctg 1800
gtgctgttgc tcatgcctcg tctcatttgc aaggcagagc tgatccggaa gcaggcccag 1860
gagaagtttg aactaagtga gaactgttca gagcggcctg ggctgcgagg agctgctggg 1920
gagcaactca gctttgctgc tggactggtg tactcgctga gcctgctgca ggccacgcta 1980
caccgctatg agcatgccct ctctcagtgc agtgtggatg tgtataagaa agtgggcagc 2040
ctgtaccctg agatgagtgc ccatgagcgc tccttggatt tcctcattga actgctgcac 2100
aaggatcagc tggatgagac tgtcaatgtg gagcctctca ccaaggccat caagtactat 2160
cagcatctgt acagcatcca ccttgccgaa cagcctgagg actgtactat gcagctggct 2220
gaccacatta agttcacgca gagtgctctg gactgcatga gtgtggaggt aggacggctg 2280
cgtgccttct tgcagggtgg gcaggaggct acagatattg ccctcctgct ccgggatctg 2340
gaaacttcat gcagtgacat ccgccagttc tgcaagaaga tccgaaggcg aatgccaggg 2400
acagatgctc ctgggatccc agctgcactg gcctttggac cacaggtatc tgacacgctc 2460
ctagactgca ggaaacactt gacgtgggtc gtggctgtgc tgcaggaggt ggcagctgct 2520
gctgcccagc tcattgcccc actggcagag aatgaggggc tacttgtggc tgctctggag 2580
gaactggctt tcaaagcaag cgagcagatc tatgggaccc cctccagcag cccctatgag 2640
tgtctgcgcc agtcatgcaa catcctcatc agtaccatga acaagctggc cacagccatg 2700
caggaggggg agtatgatgc agagcggccc cccagcaagc ctccaccggt tgaactgcgg 2760
gctgctgccc ttcgtgcaga gatcacagat gctgaaggcc tgggtttgaa gctcgaagat 2820
cgagagacag ttattaagga gttgaagaag tcactcaaga ttaagggaga ggagctaagt 2880
gaggccaatg tgcggctgag cctcctggag aagaagttgg acagtgctgc caaggatgca 2940
gatgagcgca tcgagaaagt ccagactcgg ctggaggaga cccaggcact gctgcgaaag 3000
aaggagaaag agtttgagga gacaatggat gcactccagg ctgacatcga ccagctggag 3060
gcagagaagg cagaactaaa gcagcgtctg aacagccagt ccaaacgcac gattgaggga 3120
ctccggggcc ctcctccttc aggcattgct actctggtct ctggcattgc tggtgaagaa 3180
cagcagcgag gagccatccc tgggcaggct ccagggtctg tgccaggccc agggctggtg 3240
aaggactcac cactgctgct tcagcagatc tctgccatga ggctgcacat ctcccagctc 3300
cagcatgaga acagcatcct caagggagcc cagatgaagg catccttggc atccctgccc 3360
cctctgcatg ttgcaaagct atcccatgag ggccctggca gtgagttacc agctggagcg 3420
ctgtatcgta agaccagcca gctgctggag acattgaatc aattgagcac acacacgcac 3480
gtagtagaca tcactcgcac cagccctgct gccaagagcc cgtcggccca acttatggag 3540
caagtggctc agcttaagtc cctgagtgac accgtcgaga agctcaagga tgaggtcctc 3600
aaggagacag tatctcagcg ccctggagcc acagtaccca ctgactttgc caccttccct 3660
tcatcagcct tcctcaggga ggatccaaag tgggaattcc ctcggaagaa cttggttctt 3720
ggaaaaactc taggagaagg cgaatttgga aaagtggtca aggcaacggc cttccatctg 3780
aaaggcagag cagggtacac cacggtggcc gtgaagatgc tgaaagagaa cgcctccccg 3840
agtgagcttc gagacctgct gtcagagttc aacgtcctga agcaggtcaa ccacccacat 3900
gtcatcaaat tgtatggggc ctgcagccag gatggcccgc tcctcctcat cgtggagtac 3960
gccaaatacg gctccctgcg gggcttcctc cgcgagagcc gcaaagtggg gcctggctac 4020
ctgggcagtg gaggcagccg caactccagc tccctggacc acccggatga gcgggccctc 4080
accatgggcg acctcatctc atttgcctgg cagatctcac aggggatgca gtatctggcc 4140
gagatgaagc tcgttcatcg ggacttggca gccagaaaca tcctggtagc tgaggggcgg 4200
aagatgaaga tttcggattt cggcttgtcc cgagatgttt atgaagagga ttcctacgtg 4260
aagaggagcc agggtcggat tccagttaaa tggatggcaa ttgaatccct ttttgatcat 4320
atctacacca cgcaaagtga tgtatggtct tttggtgtcc tgctgtggga gatcgtgacc 4380
ctagggggaa acccctatcc tgggattcct cctgagcggc tcttcaacct tctgaagacc 4440
ggccaccgga tggagaggcc agacaactgc agcgaggaga tgtaccgcct gatgctgcaa 4500
tgctggaagc aggagccgga caaaaggccg gtgtttgcgg acatcagcaa agacctggag 4560
aagatgatgg ttaagaggag agactacttg gaccttgcgg cgtccactcc atctgactcc 4620
ctgatttatg acgacggcct ctcagaggag gagacaccgc tggtggactg taataatgcc 4680
cccctccctc gagccctccc ttccacatgg attgaaaaca aactctatgg tagaatttcc 4740
catgcattta ctagattcta g 4761
<210> 24
<211> 1586
<212> PRT
<213> Artificial Sequence
<220>
<223> fusion peptide
<400> 24
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro Lys
130 135 140
Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser Leu
145 150 155 160
Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu Pro
165 170 175
Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr Pro
180 185 190
Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser Lys
195 200 205
Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys Leu
210 215 220
Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys Glu
225 230 235 240
Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys Ser
245 250 255
Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu Ala
260 265 270
Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met Glu
275 280 285
Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp Lys
290 295 300
Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu Ala
305 310 315 320
Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu Lys
325 330 335
Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr Gln
340 345 350
Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu Val
355 360 365
Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys Leu
370 375 380
Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg Gln
385 390 395 400
Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr Ile
405 410 415
Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu Met
420 425 430
Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val Arg
435 440 445
Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met Asn
450 455 460
Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg Glu
465 470 475 480
Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg Val
485 490 495
Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys Lys
500 505 510
Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu Thr
515 520 525
Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro Glu
530 535 540
Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala Lys
545 550 555 560
Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn Arg
565 570 575
His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg Pro
580 585 590
Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg Leu
595 600 605
Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe Glu
610 615 620
Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala Gly
625 630 635 640
Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu Leu
645 650 655
Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser Val
660 665 670
Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala His
675 680 685
Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln Leu
690 695 700
Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr Tyr
705 710 715 720
Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys Thr
725 730 735
Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp Cys
740 745 750
Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly Gln
755 760 765
Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser Cys
770 775 780
Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro Gly
785 790 795 800
Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln Val
805 810 815
Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val Ala
820 825 830
Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro Leu
835 840 845
Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala Phe
850 855 860
Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr Glu
865 870 875 880
Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys Leu
885 890 895
Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro Ser
900 905 910
Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu Ile
915 920 925
Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr Val
930 935 940
Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu Ser
945 950 955 960
Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser Ala
965 970 975
Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu Glu
980 985 990
Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu Thr
995 1000 1005
Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
1010 1015 1020
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile
1025 1030 1035
Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val
1040 1045 1050
Ser Gly Ile Ala Gly Glu Glu Gln Gln Arg Gly Ala Ile Pro Gly
1055 1060 1065
Gln Ala Pro Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser
1070 1075 1080
Pro Leu Leu Leu Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser
1085 1090 1095
Gln Leu Gln His Glu Asn Ser Ile Leu Lys Gly Ala Gln Met Lys
1100 1105 1110
Ala Ser Leu Ala Ser Leu Pro Pro Leu His Val Ala Lys Leu Ser
1115 1120 1125
His Glu Gly Pro Gly Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg
1130 1135 1140
Lys Thr Ser Gln Leu Leu Glu Thr Leu Asn Gln Leu Ser Thr His
1145 1150 1155
Thr His Val Val Asp Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser
1160 1165 1170
Pro Ser Ala Gln Leu Met Glu Gln Val Ala Gln Leu Lys Ser Leu
1175 1180 1185
Ser Asp Thr Val Glu Lys Leu Lys Asp Glu Val Leu Lys Glu Thr
1190 1195 1200
Val Ser Gln Arg Pro Gly Ala Thr Val Pro Thr Asp Phe Ala Thr
1205 1210 1215
Phe Pro Ser Ser Ala Phe Leu Arg Glu Asp Pro Lys Trp Glu Phe
1220 1225 1230
Pro Arg Lys Asn Leu Val Leu Gly Lys Thr Leu Gly Glu Gly Glu
1235 1240 1245
Phe Gly Lys Val Val Lys Ala Thr Ala Phe His Leu Lys Gly Arg
1250 1255 1260
Ala Gly Tyr Thr Thr Val Ala Val Lys Met Leu Lys Glu Asn Ala
1265 1270 1275
Ser Pro Ser Glu Leu Arg Asp Leu Leu Ser Glu Phe Asn Val Leu
1280 1285 1290
Lys Gln Val Asn His Pro His Val Ile Lys Leu Tyr Gly Ala Cys
1295 1300 1305
Ser Gln Asp Gly Pro Leu Leu Leu Ile Val Glu Tyr Ala Lys Tyr
1310 1315 1320
Gly Ser Leu Arg Gly Phe Leu Arg Glu Ser Arg Lys Val Gly Pro
1325 1330 1335
Gly Tyr Leu Gly Ser Gly Gly Ser Arg Asn Ser Ser Ser Leu Asp
1340 1345 1350
His Pro Asp Glu Arg Ala Leu Thr Met Gly Asp Leu Ile Ser Phe
1355 1360 1365
Ala Trp Gln Ile Ser Gln Gly Met Gln Tyr Leu Ala Glu Met Lys
1370 1375 1380
Leu Val His Arg Asp Leu Ala Ala Arg Asn Ile Leu Val Ala Glu
1385 1390 1395
Gly Arg Lys Met Lys Ile Ser Asp Phe Gly Leu Ser Arg Asp Val
1400 1405 1410
Tyr Glu Glu Asp Ser Tyr Val Lys Arg Ser Gln Gly Arg Ile Pro
1415 1420 1425
Val Lys Trp Met Ala Ile Glu Ser Leu Phe Asp His Ile Tyr Thr
1430 1435 1440
Thr Gln Ser Asp Val Trp Ser Phe Gly Val Leu Leu Trp Glu Ile
1445 1450 1455
Val Thr Leu Gly Gly Asn Pro Tyr Pro Gly Ile Pro Pro Glu Arg
1460 1465 1470
Leu Phe Asn Leu Leu Lys Thr Gly His Arg Met Glu Arg Pro Asp
1475 1480 1485
Asn Cys Ser Glu Glu Met Tyr Arg Leu Met Leu Gln Cys Trp Lys
1490 1495 1500
Gln Glu Pro Asp Lys Arg Pro Val Phe Ala Asp Ile Ser Lys Asp
1505 1510 1515
Leu Glu Lys Met Met Val Lys Arg Arg Asp Tyr Leu Asp Leu Ala
1520 1525 1530
Ala Ser Thr Pro Ser Asp Ser Leu Ile Tyr Asp Asp Gly Leu Ser
1535 1540 1545
Glu Glu Glu Thr Pro Leu Val Asp Cys Asn Asn Ala Pro Leu Pro
1550 1555 1560
Arg Ala Leu Pro Ser Thr Trp Ile Glu Asn Lys Leu Tyr Gly Arg
1565 1570 1575
Ile Ser His Ala Phe Thr Arg Phe
1580 1585
<210> 25
<211> 1278
<212> PRT
<213> Homo sapiens
<400> 25
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Arg Gly Leu Lys Pro Lys Lys Ala Pro Thr Ala Arg Lys
130 135 140
Thr Thr Thr Arg Arg Pro Lys Pro Thr Arg Pro Ala Ser Thr Gly Val
145 150 155 160
Ala Gly Ala Ser Ser Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly
165 170 175
Glu Leu Ser Ser Ser Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala
180 185 190
Ala Pro Ile Ile Pro Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro
195 200 205
Pro Leu Pro Ser Pro Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val
210 215 220
Arg Asp Leu Glu Glu Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu
225 230 235 240
Asp Lys Ala Lys Leu Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu
245 250 255
Gln Val Gln Glu Trp Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu
260 265 270
Gln Arg Arg Leu Lys Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu
275 280 285
Ala Lys Glu Arg Tyr Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile
290 295 300
Glu Met Ala Thr Leu Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser
305 310 315 320
Leu Gln Gln Glu Val Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr
325 330 335
Thr Asp Leu Glu Ile Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp
340 345 350
Gly Ala Ala Ser Ser Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala
355 360 365
Arg Leu Lys Asp Ala Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu
370 375 380
Lys Gln Glu His Val Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln
385 390 395 400
Glu Leu Glu Val Val Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu
405 410 415
Ser Gln Ala Glu Ser Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala
420 425 430
Ala Leu Gly Ala Glu Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu
435 440 445
Asn Leu Glu Glu Lys Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu
450 455 460
Glu Ala Met Asn Glu Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu
465 470 475 480
Thr Glu Leu Glu Leu Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val
485 490 495
Arg Glu Ala Gln Lys Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp
500 505 510
Tyr Gln Gln Thr Ile Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln
515 520 525
Asp Val Asn Arg Glu Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg
530 535 540
Gln Gln Gln Pro Pro Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala
545 550 555 560
Glu Thr Lys Ala His Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met
565 570 575
Glu Val Ala Gln Ala Asn Arg His Met Ser Leu Leu Thr Ala Phe Met
580 585 590
Pro Asp Ser Phe Leu Arg Pro Gly Gly Asp His Asp Cys Val Leu Val
595 600 605
Leu Leu Leu Met Pro Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys
610 615 620
Gln Ala Gln Glu Lys Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro
625 630 635 640
Gly Leu Arg Gly Ala Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu
645 650 655
Val Tyr Ser Leu Ser Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His
660 665 670
Ala Leu Ser Gln Cys Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu
675 680 685
Tyr Pro Glu Met Ser Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu
690 695 700
Leu Leu His Lys Asp Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu
705 710 715 720
Thr Lys Ala Ile Lys Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala
725 730 735
Glu Gln Pro Glu Asp Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe
740 745 750
Thr Gln Ser Ala Leu Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg
755 760 765
Ala Phe Leu Gln Gly Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu
770 775 780
Arg Asp Leu Glu Thr Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys
785 790 795 800
Ile Arg Arg Arg Met Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala
805 810 815
Leu Ala Phe Gly Pro Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys
820 825 830
His Leu Thr Trp Val Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala
835 840 845
Ala Gln Leu Ile Ala Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala
850 855 860
Ala Leu Glu Glu Leu Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr
865 870 875 880
Pro Ser Ser Ser Pro Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu
885 890 895
Ile Ser Thr Met Asn Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr
900 905 910
Asp Ala Glu Arg Pro Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala
915 920 925
Ala Ala Leu Arg Ala Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys
930 935 940
Leu Glu Asp Arg Glu Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys
945 950 955 960
Ile Lys Gly Glu Glu Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu
965 970 975
Glu Lys Lys Leu Asp Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu
980 985 990
Lys Val Gln Thr Arg Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys
995 1000 1005
Glu Lys Glu Phe Glu Glu Thr Met Asp Ala Leu Gln Ala Asp Ile
1010 1015 1020
Asp Gln Leu Glu Ala Glu Lys Ala Glu Leu Lys Gln Arg Leu Asn
1025 1030 1035
Ser Gln Ser Lys Arg Thr Ile Glu Gly Leu Arg Gly Pro Pro Pro
1040 1045 1050
Ser Gly Ile Ala Thr Leu Val Ser Gly Ile Ala Gly Glu Glu Gln
1055 1060 1065
Gln Arg Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser Val Pro Gly
1070 1075 1080
Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln Gln Ile Ser
1085 1090 1095
Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu Asn Ser Ile
1100 1105 1110
Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu Pro Pro
1115 1120 1125
Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu Leu
1130 1135 1140
Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu Thr
1145 1150 1155
Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp Ile Thr Arg
1160 1165 1170
Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu Met Glu Gln
1175 1180 1185
Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu Lys Leu Lys
1190 1195 1200
Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro Gly Ala Thr
1205 1210 1215
Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu Arg
1220 1225 1230
Ala Lys Glu Glu Gln Gln Asp Asp Thr Val Tyr Met Gly Lys Val
1235 1240 1245
Thr Phe Ser Cys Ala Ala Gly Phe Gly Gln Arg His Arg Leu Val
1250 1255 1260
Leu Thr Gln Glu Gln Leu His Gln Leu His Ser Arg Leu Ile Ser
1265 1270 1275
<210> 26
<211> 1144
<212> PRT
<213> Homo sapiens
<400> 26
Met Met Arg Gln Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro
1 5 10 15
Lys Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser
20 25 30
Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu
35 40 45
Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr
50 55 60
Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser
65 70 75 80
Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys
85 90 95
Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys
100 105 110
Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys
115 120 125
Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu
130 135 140
Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met
145 150 155 160
Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp
165 170 175
Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu
180 185 190
Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu
195 200 205
Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr
210 215 220
Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu
225 230 235 240
Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys
245 250 255
Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg
260 265 270
Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr
275 280 285
Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu
290 295 300
Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val
305 310 315 320
Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met
325 330 335
Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg
340 345 350
Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg
355 360 365
Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys
370 375 380
Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu
385 390 395 400
Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro
405 410 415
Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala
420 425 430
Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn
435 440 445
Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg
450 455 460
Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg
465 470 475 480
Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe
485 490 495
Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala
500 505 510
Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu
515 520 525
Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser
530 535 540
Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala
545 550 555 560
His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln
565 570 575
Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr
580 585 590
Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys
595 600 605
Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp
610 615 620
Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly
625 630 635 640
Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser
645 650 655
Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro
660 665 670
Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln
675 680 685
Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val
690 695 700
Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro
705 710 715 720
Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala
725 730 735
Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr
740 745 750
Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys
755 760 765
Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro
770 775 780
Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu
785 790 795 800
Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr
805 810 815
Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu
820 825 830
Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser
835 840 845
Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu
850 855 860
Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu
865 870 875 880
Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
885 890 895
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile Glu
900 905 910
Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val Ser Gly
915 920 925
Ile Ala Gly Glu Glu Gln Gln Arg Gly Ala Ile Pro Gly Gln Ala Pro
930 935 940
Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu
945 950 955 960
Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu
965 970 975
Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu
980 985 990
Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu
995 1000 1005
Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu
1010 1015 1020
Thr Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp Ile Thr
1025 1030 1035
Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu Met Glu
1040 1045 1050
Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu Lys Leu
1055 1060 1065
Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro Gly Ala
1070 1075 1080
Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu
1085 1090 1095
Arg Ala Lys Glu Glu Gln Gln Asp Asp Thr Val Tyr Met Gly Lys
1100 1105 1110
Val Thr Phe Ser Cys Ala Ala Gly Phe Gly Gln Arg His Arg Leu
1115 1120 1125
Val Leu Thr Gln Glu Gln Leu His Gln Leu His Ser Arg Leu Ile
1130 1135 1140
Ser
<210> 27
<211> 1253
<212> PRT
<213> Homo sapiens
<400> 27
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser
130 135 140
Ser Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser
145 150 155 160
Ser Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile
165 170 175
Pro Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser
180 185 190
Pro Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu
195 200 205
Glu Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys
210 215 220
Leu Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu
225 230 235 240
Trp Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu
245 250 255
Lys Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg
260 265 270
Tyr Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr
275 280 285
Leu Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu
290 295 300
Val Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu
305 310 315 320
Ile Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser
325 330 335
Ser Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp
340 345 350
Ala Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His
355 360 365
Val Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val
370 375 380
Val Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu
385 390 395 400
Ser Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala
405 410 415
Glu Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu
420 425 430
Lys Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn
435 440 445
Glu Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu
450 455 460
Leu Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln
465 470 475 480
Lys Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr
485 490 495
Ile Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg
500 505 510
Glu Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro
515 520 525
Pro Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala
530 535 540
His Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln
545 550 555 560
Ala Asn Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe
565 570 575
Leu Arg Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met
580 585 590
Pro Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu
595 600 605
Lys Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly
610 615 620
Ala Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu
625 630 635 640
Ser Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln
645 650 655
Cys Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met
660 665 670
Ser Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys
675 680 685
Asp Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile
690 695 700
Lys Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu
705 710 715 720
Asp Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala
725 730 735
Leu Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln
740 745 750
Gly Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu
755 760 765
Thr Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg
770 775 780
Met Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly
785 790 795 800
Pro Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp
805 810 815
Val Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile
820 825 830
Ala Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu
835 840 845
Leu Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser
850 855 860
Pro Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met
865 870 875 880
Asn Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg
885 890 895
Pro Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg
900 905 910
Ala Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg
915 920 925
Glu Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu
930 935 940
Glu Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu
945 950 955 960
Asp Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr
965 970 975
Arg Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe
980 985 990
Glu Glu Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala
995 1000 1005
Glu Lys Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg
1010 1015 1020
Thr Ile Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr
1025 1030 1035
Leu Val Ser Gly Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro
1040 1045 1050
Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu
1055 1060 1065
Leu Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln
1070 1075 1080
His Glu Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu
1085 1090 1095
Ala Ser Leu Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly
1100 1105 1110
Pro Gly Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser
1115 1120 1125
Gln Leu Leu Glu Thr Leu Asn Gln Leu Ser Thr His Thr His Val
1130 1135 1140
Val Asp Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala
1145 1150 1155
Gln Leu Met Glu Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr
1160 1165 1170
Val Glu Lys Leu Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln
1175 1180 1185
Arg Pro Gly Ala Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser
1190 1195 1200
Ser Ala Phe Leu Arg Ala Lys Glu Glu Gln Gln Asp Asp Thr Val
1205 1210 1215
Tyr Met Gly Lys Val Thr Phe Ser Cys Ala Ala Gly Phe Gly Gln
1220 1225 1230
Arg His Arg Leu Val Leu Thr Gln Glu Gln Leu His Gln Leu His
1235 1240 1245
Ser Arg Leu Ile Ser
1250
<210> 28
<211> 1139
<212> PRT
<213> Homo sapiens
<400> 28
Met Met Arg Gln Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro
1 5 10 15
Lys Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser
20 25 30
Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu
35 40 45
Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr
50 55 60
Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser
65 70 75 80
Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys
85 90 95
Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys
100 105 110
Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys
115 120 125
Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu
130 135 140
Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met
145 150 155 160
Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp
165 170 175
Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu
180 185 190
Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu
195 200 205
Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr
210 215 220
Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu
225 230 235 240
Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys
245 250 255
Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg
260 265 270
Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr
275 280 285
Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu
290 295 300
Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val
305 310 315 320
Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met
325 330 335
Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg
340 345 350
Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg
355 360 365
Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys
370 375 380
Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu
385 390 395 400
Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro
405 410 415
Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala
420 425 430
Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn
435 440 445
Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg
450 455 460
Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg
465 470 475 480
Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe
485 490 495
Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala
500 505 510
Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu
515 520 525
Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser
530 535 540
Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala
545 550 555 560
His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln
565 570 575
Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr
580 585 590
Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys
595 600 605
Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp
610 615 620
Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly
625 630 635 640
Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser
645 650 655
Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro
660 665 670
Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln
675 680 685
Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val
690 695 700
Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro
705 710 715 720
Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala
725 730 735
Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr
740 745 750
Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys
755 760 765
Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro
770 775 780
Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu
785 790 795 800
Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr
805 810 815
Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu
820 825 830
Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser
835 840 845
Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu
850 855 860
Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu
865 870 875 880
Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
885 890 895
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile Glu
900 905 910
Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val Ser Gly
915 920 925
Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser Val Pro Gly
930 935 940
Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln Gln Ile Ser Ala
945 950 955 960
Met Arg Leu His Ile Ser Gln Leu Gln His Glu Asn Ser Ile Leu Lys
965 970 975
Gly Ala Gln Met Lys Ala Ser Leu Ala Ser Leu Pro Pro Leu His Val
980 985 990
Ala Lys Leu Ser His Glu Gly Pro Gly Ser Glu Leu Pro Ala Gly Ala
995 1000 1005
Leu Tyr Arg Lys Thr Ser Gln Leu Leu Glu Thr Leu Asn Gln Leu
1010 1015 1020
Ser Thr His Thr His Val Val Asp Ile Thr Arg Thr Ser Pro Ala
1025 1030 1035
Ala Lys Ser Pro Ser Ala Gln Leu Met Glu Gln Val Ala Gln Leu
1040 1045 1050
Lys Ser Leu Ser Asp Thr Val Glu Lys Leu Lys Asp Glu Val Leu
1055 1060 1065
Lys Glu Thr Val Ser Gln Arg Pro Gly Ala Thr Val Pro Thr Asp
1070 1075 1080
Phe Ala Thr Phe Pro Ser Ser Ala Phe Leu Arg Ala Lys Glu Glu
1085 1090 1095
Gln Gln Asp Asp Thr Val Tyr Met Gly Lys Val Thr Phe Ser Cys
1100 1105 1110
Ala Ala Gly Phe Gly Gln Arg His Arg Leu Val Leu Thr Gln Glu
1115 1120 1125
Gln Leu His Gln Leu His Ser Arg Leu Ile Ser
1130 1135
<210> 29
<211> 1236
<212> PRT
<213> Homo sapiens
<400> 29
Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg Val
1 5 10 15
Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly Ala
20 25 30
Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu Ala
35 40 45
Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr Cys
50 55 60
Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val Phe
65 70 75 80
Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser Ala
85 90 95
Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr Ser
100 105 110
Lys Leu Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser
115 120 125
Ser Leu Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser
130 135 140
Glu Pro Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro
145 150 155 160
Thr Pro Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro
165 170 175
Ser Lys Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu
180 185 190
Lys Leu Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu
195 200 205
Lys Glu Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp
210 215 220
Lys Ser Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys
225 230 235 240
Glu Ala Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr
245 250 255
Met Glu Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu
260 265 270
Asp Lys Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val
275 280 285
Glu Ala Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile
290 295 300
Leu Lys Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser
305 310 315 320
Tyr Gln Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala
325 330 335
Leu Val Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val
340 345 350
Lys Leu Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val
355 360 365
Arg Gln Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser
370 375 380
Thr Ile Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu
385 390 395 400
Glu Met Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys
405 410 415
Val Arg Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu
420 425 430
Met Asn Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu
435 440 445
Arg Glu Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys
450 455 460
Arg Val Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile
465 470 475 480
Lys Lys Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu
485 490 495
Leu Thr Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro
500 505 510
Pro Glu Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His
515 520 525
Ala Lys Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala
530 535 540
Asn Arg His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu
545 550 555 560
Arg Pro Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro
565 570 575
Arg Leu Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys
580 585 590
Phe Glu Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala
595 600 605
Ala Gly Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser
610 615 620
Leu Leu Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys
625 630 635 640
Ser Val Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser
645 650 655
Ala His Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp
660 665 670
Gln Leu Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys
675 680 685
Tyr Tyr Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp
690 695 700
Cys Thr Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu
705 710 715 720
Asp Cys Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly
725 730 735
Gly Gln Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr
740 745 750
Ser Cys Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met
755 760 765
Pro Gly Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro
770 775 780
Gln Val Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val
785 790 795 800
Val Ala Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala
805 810 815
Pro Leu Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu
820 825 830
Ala Phe Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro
835 840 845
Tyr Glu Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn
850 855 860
Lys Leu Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro
865 870 875 880
Pro Ser Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala
885 890 895
Glu Ile Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu
900 905 910
Thr Val Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu
915 920 925
Leu Ser Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp
930 935 940
Ser Ala Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg
945 950 955 960
Leu Glu Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu
965 970 975
Glu Thr Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu
980 985 990
Lys Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile
995 1000 1005
Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val
1010 1015 1020
Ser Gly Ile Ala Gly Gly Ala Ile Pro Gly Gln Ala Pro Gly Ser
1025 1030 1035
Val Pro Gly Pro Gly Leu Val Lys Asp Ser Pro Leu Leu Leu Gln
1040 1045 1050
Gln Ile Ser Ala Met Arg Leu His Ile Ser Gln Leu Gln His Glu
1055 1060 1065
Asn Ser Ile Leu Lys Gly Ala Gln Met Lys Ala Ser Leu Ala Ser
1070 1075 1080
Leu Pro Pro Leu His Val Ala Lys Leu Ser His Glu Gly Pro Gly
1085 1090 1095
Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg Lys Thr Ser Gln Leu
1100 1105 1110
Leu Glu Thr Leu Asn Gln Leu Ser Thr His Thr His Val Val Asp
1115 1120 1125
Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser Pro Ser Ala Gln Leu
1130 1135 1140
Met Glu Gln Val Ala Gln Leu Lys Ser Leu Ser Asp Thr Val Glu
1145 1150 1155
Lys Leu Lys Asp Glu Val Leu Lys Glu Thr Val Ser Gln Arg Pro
1160 1165 1170
Gly Ala Thr Val Pro Thr Asp Phe Ala Thr Phe Pro Ser Ser Ala
1175 1180 1185
Phe Leu Arg Ala Lys Glu Glu Gln Gln Asp Asp Thr Val Tyr Met
1190 1195 1200
Gly Lys Val Thr Phe Ser Cys Ala Ala Gly Phe Gly Gln Arg His
1205 1210 1215
Arg Leu Val Leu Thr Gln Glu Gln Leu His Gln Leu His Ser Arg
1220 1225 1230
Leu Ile Ser
1235
<210> 30
<211> 1271
<212> PRT
<213> Homo sapiens
<400> 30
Met Ala Gln Ser Lys Arg His Val Tyr Ser Arg Thr Pro Ser Gly Ser
1 5 10 15
Arg Met Ser Ala Glu Ala Ser Ala Arg Pro Leu Arg Val Gly Ser Arg
20 25 30
Val Glu Val Ile Gly Lys Gly His Arg Gly Thr Val Ala Tyr Val Gly
35 40 45
Ala Thr Leu Phe Ala Thr Gly Lys Trp Val Gly Val Ile Leu Asp Glu
50 55 60
Ala Lys Gly Lys Asn Asp Gly Thr Val Gln Gly Arg Lys Tyr Phe Thr
65 70 75 80
Cys Asp Glu Gly His Gly Ile Phe Val Arg Gln Ser Gln Ile Gln Val
85 90 95
Phe Glu Asp Gly Ala Asp Thr Thr Ser Pro Glu Thr Pro Asp Ser Ser
100 105 110
Ala Ser Lys Val Leu Lys Arg Glu Gly Thr Asp Thr Thr Ala Lys Thr
115 120 125
Ser Lys Leu Ala Pro Thr Ala Arg Lys Thr Thr Thr Arg Arg Pro Lys
130 135 140
Pro Thr Arg Pro Ala Ser Thr Gly Val Ala Gly Ala Ser Ser Ser Leu
145 150 155 160
Gly Pro Ser Gly Ser Ala Ser Ala Gly Glu Leu Ser Ser Ser Glu Pro
165 170 175
Ser Thr Pro Ala Gln Thr Pro Leu Ala Ala Pro Ile Ile Pro Thr Pro
180 185 190
Val Leu Thr Ser Pro Gly Ala Val Pro Pro Leu Pro Ser Pro Ser Lys
195 200 205
Glu Glu Glu Gly Leu Arg Ala Gln Val Arg Asp Leu Glu Glu Lys Leu
210 215 220
Glu Thr Leu Arg Leu Lys Arg Ala Glu Asp Lys Ala Lys Leu Lys Glu
225 230 235 240
Leu Glu Lys His Lys Ile Gln Leu Glu Gln Val Gln Glu Trp Lys Ser
245 250 255
Lys Met Gln Glu Gln Gln Ala Asp Leu Gln Arg Arg Leu Lys Glu Ala
260 265 270
Arg Lys Glu Ala Lys Glu Ala Leu Glu Ala Lys Glu Arg Tyr Met Glu
275 280 285
Glu Met Ala Asp Thr Ala Asp Ala Ile Glu Met Ala Thr Leu Asp Lys
290 295 300
Glu Met Ala Glu Glu Arg Ala Glu Ser Leu Gln Gln Glu Val Glu Ala
305 310 315 320
Leu Lys Glu Arg Val Asp Glu Leu Thr Thr Asp Leu Glu Ile Leu Lys
325 330 335
Ala Glu Ile Glu Glu Lys Gly Ser Asp Gly Ala Ala Ser Ser Tyr Gln
340 345 350
Leu Lys Gln Leu Glu Glu Gln Asn Ala Arg Leu Lys Asp Ala Leu Val
355 360 365
Arg Met Arg Asp Leu Ser Ser Ser Glu Lys Gln Glu His Val Lys Leu
370 375 380
Gln Lys Leu Met Glu Lys Lys Asn Gln Glu Leu Glu Val Val Arg Gln
385 390 395 400
Gln Arg Glu Arg Leu Gln Glu Glu Leu Ser Gln Ala Glu Ser Thr Ile
405 410 415
Asp Glu Leu Lys Glu Gln Val Asp Ala Ala Leu Gly Ala Glu Glu Met
420 425 430
Val Glu Met Leu Thr Asp Arg Asn Leu Asn Leu Glu Glu Lys Val Arg
435 440 445
Glu Leu Arg Glu Thr Val Gly Asp Leu Glu Ala Met Asn Glu Met Asn
450 455 460
Asp Glu Leu Gln Glu Asn Ala Arg Glu Thr Glu Leu Glu Leu Arg Glu
465 470 475 480
Gln Leu Asp Met Ala Gly Ala Arg Val Arg Glu Ala Gln Lys Arg Val
485 490 495
Glu Ala Ala Gln Glu Thr Val Ala Asp Tyr Gln Gln Thr Ile Lys Lys
500 505 510
Tyr Arg Gln Leu Thr Ala His Leu Gln Asp Val Asn Arg Glu Leu Thr
515 520 525
Asn Gln Gln Glu Ala Ser Val Glu Arg Gln Gln Gln Pro Pro Pro Glu
530 535 540
Thr Phe Asp Phe Lys Ile Lys Phe Ala Glu Thr Lys Ala His Ala Lys
545 550 555 560
Ala Ile Glu Met Glu Leu Arg Gln Met Glu Val Ala Gln Ala Asn Arg
565 570 575
His Met Ser Leu Leu Thr Ala Phe Met Pro Asp Ser Phe Leu Arg Pro
580 585 590
Gly Gly Asp His Asp Cys Val Leu Val Leu Leu Leu Met Pro Arg Leu
595 600 605
Ile Cys Lys Ala Glu Leu Ile Arg Lys Gln Ala Gln Glu Lys Phe Glu
610 615 620
Leu Ser Glu Asn Cys Ser Glu Arg Pro Gly Leu Arg Gly Ala Ala Gly
625 630 635 640
Glu Gln Leu Ser Phe Ala Ala Gly Leu Val Tyr Ser Leu Ser Leu Leu
645 650 655
Gln Ala Thr Leu His Arg Tyr Glu His Ala Leu Ser Gln Cys Ser Val
660 665 670
Asp Val Tyr Lys Lys Val Gly Ser Leu Tyr Pro Glu Met Ser Ala His
675 680 685
Glu Arg Ser Leu Asp Phe Leu Ile Glu Leu Leu His Lys Asp Gln Leu
690 695 700
Asp Glu Thr Val Asn Val Glu Pro Leu Thr Lys Ala Ile Lys Tyr Tyr
705 710 715 720
Gln His Leu Tyr Ser Ile His Leu Ala Glu Gln Pro Glu Asp Cys Thr
725 730 735
Met Gln Leu Ala Asp His Ile Lys Phe Thr Gln Ser Ala Leu Asp Cys
740 745 750
Met Ser Val Glu Val Gly Arg Leu Arg Ala Phe Leu Gln Gly Gly Gln
755 760 765
Glu Ala Thr Asp Ile Ala Leu Leu Leu Arg Asp Leu Glu Thr Ser Cys
770 775 780
Ser Asp Ile Arg Gln Phe Cys Lys Lys Ile Arg Arg Arg Met Pro Gly
785 790 795 800
Thr Asp Ala Pro Gly Ile Pro Ala Ala Leu Ala Phe Gly Pro Gln Val
805 810 815
Ser Asp Thr Leu Leu Asp Cys Arg Lys His Leu Thr Trp Val Val Ala
820 825 830
Val Leu Gln Glu Val Ala Ala Ala Ala Ala Gln Leu Ile Ala Pro Leu
835 840 845
Ala Glu Asn Glu Gly Leu Leu Val Ala Ala Leu Glu Glu Leu Ala Phe
850 855 860
Lys Ala Ser Glu Gln Ile Tyr Gly Thr Pro Ser Ser Ser Pro Tyr Glu
865 870 875 880
Cys Leu Arg Gln Ser Cys Asn Ile Leu Ile Ser Thr Met Asn Lys Leu
885 890 895
Ala Thr Ala Met Gln Glu Gly Glu Tyr Asp Ala Glu Arg Pro Pro Ser
900 905 910
Lys Pro Pro Pro Val Glu Leu Arg Ala Ala Ala Leu Arg Ala Glu Ile
915 920 925
Thr Asp Ala Glu Gly Leu Gly Leu Lys Leu Glu Asp Arg Glu Thr Val
930 935 940
Ile Lys Glu Leu Lys Lys Ser Leu Lys Ile Lys Gly Glu Glu Leu Ser
945 950 955 960
Glu Ala Asn Val Arg Leu Ser Leu Leu Glu Lys Lys Leu Asp Ser Ala
965 970 975
Ala Lys Asp Ala Asp Glu Arg Ile Glu Lys Val Gln Thr Arg Leu Glu
980 985 990
Glu Thr Gln Ala Leu Leu Arg Lys Lys Glu Lys Glu Phe Glu Glu Thr
995 1000 1005
Met Asp Ala Leu Gln Ala Asp Ile Asp Gln Leu Glu Ala Glu Lys
1010 1015 1020
Ala Glu Leu Lys Gln Arg Leu Asn Ser Gln Ser Lys Arg Thr Ile
1025 1030 1035
Glu Gly Leu Arg Gly Pro Pro Pro Ser Gly Ile Ala Thr Leu Val
1040 1045 1050
Ser Gly Ile Ala Gly Glu Glu Gln Gln Arg Gly Ala Ile Pro Gly
1055 1060 1065
Gln Ala Pro Gly Ser Val Pro Gly Pro Gly Leu Val Lys Asp Ser
1070 1075 1080
Pro Leu Leu Leu Gln Gln Ile Ser Ala Met Arg Leu His Ile Ser
1085 1090 1095
Gln Leu Gln His Glu Asn Ser Ile Leu Lys Gly Ala Gln Met Lys
1100 1105 1110
Ala Ser Leu Ala Ser Leu Pro Pro Leu His Val Ala Lys Leu Ser
1115 1120 1125
His Glu Gly Pro Gly Ser Glu Leu Pro Ala Gly Ala Leu Tyr Arg
1130 1135 1140
Lys Thr Ser Gln Leu Leu Glu Thr Leu Asn Gln Leu Ser Thr His
1145 1150 1155
Thr His Val Val Asp Ile Thr Arg Thr Ser Pro Ala Ala Lys Ser
1160 1165 1170
Pro Ser Ala Gln Leu Met Glu Gln Val Ala Gln Leu Lys Ser Leu
1175 1180 1185
Ser Asp Thr Val Glu Lys Leu Lys Asp Glu Val Leu Lys Glu Thr
1190 1195 1200
Val Ser Gln Arg Pro Gly Ala Thr Val Pro Thr Asp Phe Ala Thr
1205 1210 1215
Phe Pro Ser Ser Ala Phe Leu Arg Ala Lys Glu Glu Gln Gln Asp
1220 1225 1230
Asp Thr Val Tyr Met Gly Lys Val Thr Phe Ser Cys Ala Ala Gly
1235 1240 1245
Phe Gly Gln Arg His Arg Leu Val Leu Thr Gln Glu Gln Leu His
1250 1255 1260
Gln Leu His Ser Arg Leu Ile Ser
1265 1270
<210> 31
<211> 1114
<212> PRT
<213> Homo sapiens
<400> 31
Met Ala Lys Ala Thr Ser Gly Ala Ala Gly Leu Arg Leu Leu Leu Leu
1 5 10 15
Leu Leu Leu Pro Leu Leu Gly Lys Val Ala Leu Gly Leu Tyr Phe Ser
20 25 30
Arg Asp Ala Tyr Trp Glu Lys Leu Tyr Val Asp Gln Ala Ala Gly Thr
35 40 45
Pro Leu Leu Tyr Val His Ala Leu Arg Asp Ala Pro Glu Glu Val Pro
50 55 60
Ser Phe Arg Leu Gly Gln His Leu Tyr Gly Thr Tyr Arg Thr Arg Leu
65 70 75 80
His Glu Asn Asn Trp Ile Cys Ile Gln Glu Asp Thr Gly Leu Leu Tyr
85 90 95
Leu Asn Arg Ser Leu Asp His Ser Ser Trp Glu Lys Leu Ser Val Arg
100 105 110
Asn Arg Gly Phe Pro Leu Leu Thr Val Tyr Leu Lys Val Phe Leu Ser
115 120 125
Pro Thr Ser Leu Arg Glu Gly Glu Cys Gln Trp Pro Gly Cys Ala Arg
130 135 140
Val Tyr Phe Ser Phe Phe Asn Thr Ser Phe Pro Ala Cys Ser Ser Leu
145 150 155 160
Lys Pro Arg Glu Leu Cys Phe Pro Glu Thr Arg Pro Ser Phe Arg Ile
165 170 175
Arg Glu Asn Arg Pro Pro Gly Thr Phe His Gln Phe Arg Leu Leu Pro
180 185 190
Val Gln Phe Leu Cys Pro Asn Ile Ser Val Ala Tyr Arg Leu Leu Glu
195 200 205
Gly Glu Gly Leu Pro Phe Arg Cys Ala Pro Asp Ser Leu Glu Val Ser
210 215 220
Thr Arg Trp Ala Leu Asp Arg Glu Gln Arg Glu Lys Tyr Glu Leu Val
225 230 235 240
Ala Val Cys Thr Val His Ala Gly Ala Arg Glu Glu Val Val Met Val
245 250 255
Pro Phe Pro Val Thr Val Tyr Asp Glu Asp Asp Ser Ala Pro Thr Phe
260 265 270
Pro Ala Gly Val Asp Thr Ala Ser Ala Val Val Glu Phe Lys Arg Lys
275 280 285
Glu Asp Thr Val Val Ala Thr Leu Arg Val Phe Asp Ala Asp Val Val
290 295 300
Pro Ala Ser Gly Glu Leu Val Arg Arg Tyr Thr Ser Thr Leu Leu Pro
305 310 315 320
Gly Asp Thr Trp Ala Gln Gln Thr Phe Arg Val Glu His Trp Pro Asn
325 330 335
Glu Thr Ser Val Gln Ala Asn Gly Ser Phe Val Arg Ala Thr Val His
340 345 350
Asp Tyr Arg Leu Val Leu Asn Arg Asn Leu Ser Ile Ser Glu Asn Arg
355 360 365
Thr Met Gln Leu Ala Val Leu Val Asn Asp Ser Asp Phe Gln Gly Pro
370 375 380
Gly Ala Gly Val Leu Leu Leu His Phe Asn Val Ser Val Leu Pro Val
385 390 395 400
Ser Leu His Leu Pro Ser Thr Tyr Ser Leu Ser Val Ser Arg Arg Ala
405 410 415
Arg Arg Phe Ala Gln Ile Gly Lys Val Cys Val Glu Asn Cys Gln Ala
420 425 430
Phe Ser Gly Ile Asn Val Gln Tyr Lys Leu His Ser Ser Gly Ala Asn
435 440 445
Cys Ser Thr Leu Gly Val Val Thr Ser Ala Glu Asp Thr Ser Gly Ile
450 455 460
Leu Phe Val Asn Asp Thr Lys Ala Leu Arg Arg Pro Lys Cys Ala Glu
465 470 475 480
Leu His Tyr Met Val Val Ala Thr Asp Gln Gln Thr Ser Arg Gln Ala
485 490 495
Gln Ala Gln Leu Leu Val Thr Val Glu Gly Ser Tyr Val Ala Glu Glu
500 505 510
Ala Gly Cys Pro Leu Ser Cys Ala Val Ser Lys Arg Arg Leu Glu Cys
515 520 525
Glu Glu Cys Gly Gly Leu Gly Ser Pro Thr Gly Arg Cys Glu Trp Arg
530 535 540
Gln Gly Asp Gly Lys Gly Ile Thr Arg Asn Phe Ser Thr Cys Ser Pro
545 550 555 560
Ser Thr Lys Thr Cys Pro Asp Gly His Cys Asp Val Val Glu Thr Gln
565 570 575
Asp Ile Asn Ile Cys Pro Gln Asp Cys Leu Arg Gly Ser Ile Val Gly
580 585 590
Gly His Glu Pro Gly Glu Pro Arg Gly Ile Lys Ala Gly Tyr Gly Thr
595 600 605
Cys Asn Cys Phe Pro Glu Glu Glu Lys Cys Phe Cys Glu Pro Glu Asp
610 615 620
Ile Gln Asp Pro Leu Cys Asp Glu Leu Cys Arg Thr Val Ile Ala Ala
625 630 635 640
Ala Val Leu Phe Ser Phe Ile Val Ser Val Leu Leu Ser Ala Phe Cys
645 650 655
Ile His Cys Tyr His Lys Phe Ala His Lys Pro Pro Ile Ser Ser Ala
660 665 670
Glu Met Thr Phe Arg Arg Pro Ala Gln Ala Phe Pro Val Ser Tyr Ser
675 680 685
Ser Ser Gly Ala Arg Arg Pro Ser Leu Asp Ser Met Glu Asn Gln Val
690 695 700
Ser Val Asp Ala Phe Lys Ile Leu Glu Asp Pro Lys Trp Glu Phe Pro
705 710 715 720
Arg Lys Asn Leu Val Leu Gly Lys Thr Leu Gly Glu Gly Glu Phe Gly
725 730 735
Lys Val Val Lys Ala Thr Ala Phe His Leu Lys Gly Arg Ala Gly Tyr
740 745 750
Thr Thr Val Ala Val Lys Met Leu Lys Glu Asn Ala Ser Pro Ser Glu
755 760 765
Leu Arg Asp Leu Leu Ser Glu Phe Asn Val Leu Lys Gln Val Asn His
770 775 780
Pro His Val Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp Gly Pro Leu
785 790 795 800
Leu Leu Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu Arg Gly Phe Leu
805 810 815
Arg Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu Gly Ser Gly Gly Ser
820 825 830
Arg Asn Ser Ser Ser Leu Asp His Pro Asp Glu Arg Ala Leu Thr Met
835 840 845
Gly Asp Leu Ile Ser Phe Ala Trp Gln Ile Ser Gln Gly Met Gln Tyr
850 855 860
Leu Ala Glu Met Lys Leu Val His Arg Asp Leu Ala Ala Arg Asn Ile
865 870 875 880
Leu Val Ala Glu Gly Arg Lys Met Lys Ile Ser Asp Phe Gly Leu Ser
885 890 895
Arg Asp Val Tyr Glu Glu Asp Ser Tyr Val Lys Arg Ser Gln Gly Arg
900 905 910
Ile Pro Val Lys Trp Met Ala Ile Glu Ser Leu Phe Asp His Ile Tyr
915 920 925
Thr Thr Gln Ser Asp Val Trp Ser Phe Gly Val Leu Leu Trp Glu Ile
930 935 940
Val Thr Leu Gly Gly Asn Pro Tyr Pro Gly Ile Pro Pro Glu Arg Leu
945 950 955 960
Phe Asn Leu Leu Lys Thr Gly His Arg Met Glu Arg Pro Asp Asn Cys
965 970 975
Ser Glu Glu Met Tyr Arg Leu Met Leu Gln Cys Trp Lys Gln Glu Pro
980 985 990
Asp Lys Arg Pro Val Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys Met
995 1000 1005
Met Val Lys Arg Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr Pro
1010 1015 1020
Ser Asp Ser Leu Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu Thr
1025 1030 1035
Pro Leu Val Asp Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu Pro
1040 1045 1050
Ser Thr Trp Ile Glu Asn Lys Leu Tyr Gly Met Ser Asp Pro Asn
1055 1060 1065
Trp Pro Gly Glu Ser Pro Val Pro Leu Thr Arg Ala Asp Gly Thr
1070 1075 1080
Asn Thr Gly Phe Pro Arg Tyr Pro Asn Asp Ser Val Tyr Ala Asn
1085 1090 1095
Trp Met Leu Ser Pro Ser Ala Ala Lys Leu Met Asp Thr Phe Asp
1100 1105 1110
Ser
<210> 32
<211> 1072
<212> PRT
<213> Homo sapiens
<400> 32
Met Ala Lys Ala Thr Ser Gly Ala Ala Gly Leu Arg Leu Leu Leu Leu
1 5 10 15
Leu Leu Leu Pro Leu Leu Gly Lys Val Ala Leu Gly Leu Tyr Phe Ser
20 25 30
Arg Asp Ala Tyr Trp Glu Lys Leu Tyr Val Asp Gln Ala Ala Gly Thr
35 40 45
Pro Leu Leu Tyr Val His Ala Leu Arg Asp Ala Pro Glu Glu Val Pro
50 55 60
Ser Phe Arg Leu Gly Gln His Leu Tyr Gly Thr Tyr Arg Thr Arg Leu
65 70 75 80
His Glu Asn Asn Trp Ile Cys Ile Gln Glu Asp Thr Gly Leu Leu Tyr
85 90 95
Leu Asn Arg Ser Leu Asp His Ser Ser Trp Glu Lys Leu Ser Val Arg
100 105 110
Asn Arg Gly Phe Pro Leu Leu Thr Val Tyr Leu Lys Val Phe Leu Ser
115 120 125
Pro Thr Ser Leu Arg Glu Gly Glu Cys Gln Trp Pro Gly Cys Ala Arg
130 135 140
Val Tyr Phe Ser Phe Phe Asn Thr Ser Phe Pro Ala Cys Ser Ser Leu
145 150 155 160
Lys Pro Arg Glu Leu Cys Phe Pro Glu Thr Arg Pro Ser Phe Arg Ile
165 170 175
Arg Glu Asn Arg Pro Pro Gly Thr Phe His Gln Phe Arg Leu Leu Pro
180 185 190
Val Gln Phe Leu Cys Pro Asn Ile Ser Val Ala Tyr Arg Leu Leu Glu
195 200 205
Gly Glu Gly Leu Pro Phe Arg Cys Ala Pro Asp Ser Leu Glu Val Ser
210 215 220
Thr Arg Trp Ala Leu Asp Arg Glu Gln Arg Glu Lys Tyr Glu Leu Val
225 230 235 240
Ala Val Cys Thr Val His Ala Gly Ala Arg Glu Glu Val Val Met Val
245 250 255
Pro Phe Pro Val Thr Val Tyr Asp Glu Asp Asp Ser Ala Pro Thr Phe
260 265 270
Pro Ala Gly Val Asp Thr Ala Ser Ala Val Val Glu Phe Lys Arg Lys
275 280 285
Glu Asp Thr Val Val Ala Thr Leu Arg Val Phe Asp Ala Asp Val Val
290 295 300
Pro Ala Ser Gly Glu Leu Val Arg Arg Tyr Thr Ser Thr Leu Leu Pro
305 310 315 320
Gly Asp Thr Trp Ala Gln Gln Thr Phe Arg Val Glu His Trp Pro Asn
325 330 335
Glu Thr Ser Val Gln Ala Asn Gly Ser Phe Val Arg Ala Thr Val His
340 345 350
Asp Tyr Arg Leu Val Leu Asn Arg Asn Leu Ser Ile Ser Glu Asn Arg
355 360 365
Thr Met Gln Leu Ala Val Leu Val Asn Asp Ser Asp Phe Gln Gly Pro
370 375 380
Gly Ala Gly Val Leu Leu Leu His Phe Asn Val Ser Val Leu Pro Val
385 390 395 400
Ser Leu His Leu Pro Ser Thr Tyr Ser Leu Ser Val Ser Arg Arg Ala
405 410 415
Arg Arg Phe Ala Gln Ile Gly Lys Val Cys Val Glu Asn Cys Gln Ala
420 425 430
Phe Ser Gly Ile Asn Val Gln Tyr Lys Leu His Ser Ser Gly Ala Asn
435 440 445
Cys Ser Thr Leu Gly Val Val Thr Ser Ala Glu Asp Thr Ser Gly Ile
450 455 460
Leu Phe Val Asn Asp Thr Lys Ala Leu Arg Arg Pro Lys Cys Ala Glu
465 470 475 480
Leu His Tyr Met Val Val Ala Thr Asp Gln Gln Thr Ser Arg Gln Ala
485 490 495
Gln Ala Gln Leu Leu Val Thr Val Glu Gly Ser Tyr Val Ala Glu Glu
500 505 510
Ala Gly Cys Pro Leu Ser Cys Ala Val Ser Lys Arg Arg Leu Glu Cys
515 520 525
Glu Glu Cys Gly Gly Leu Gly Ser Pro Thr Gly Arg Cys Glu Trp Arg
530 535 540
Gln Gly Asp Gly Lys Gly Ile Thr Arg Asn Phe Ser Thr Cys Ser Pro
545 550 555 560
Ser Thr Lys Thr Cys Pro Asp Gly His Cys Asp Val Val Glu Thr Gln
565 570 575
Asp Ile Asn Ile Cys Pro Gln Asp Cys Leu Arg Gly Ser Ile Val Gly
580 585 590
Gly His Glu Pro Gly Glu Pro Arg Gly Ile Lys Ala Gly Tyr Gly Thr
595 600 605
Cys Asn Cys Phe Pro Glu Glu Glu Lys Cys Phe Cys Glu Pro Glu Asp
610 615 620
Ile Gln Asp Pro Leu Cys Asp Glu Leu Cys Arg Thr Val Ile Ala Ala
625 630 635 640
Ala Val Leu Phe Ser Phe Ile Val Ser Val Leu Leu Ser Ala Phe Cys
645 650 655
Ile His Cys Tyr His Lys Phe Ala His Lys Pro Pro Ile Ser Ser Ala
660 665 670
Glu Met Thr Phe Arg Arg Pro Ala Gln Ala Phe Pro Val Ser Tyr Ser
675 680 685
Ser Ser Gly Ala Arg Arg Pro Ser Leu Asp Ser Met Glu Asn Gln Val
690 695 700
Ser Val Asp Ala Phe Lys Ile Leu Glu Asp Pro Lys Trp Glu Phe Pro
705 710 715 720
Arg Lys Asn Leu Val Leu Gly Lys Thr Leu Gly Glu Gly Glu Phe Gly
725 730 735
Lys Val Val Lys Ala Thr Ala Phe His Leu Lys Gly Arg Ala Gly Tyr
740 745 750
Thr Thr Val Ala Val Lys Met Leu Lys Glu Asn Ala Ser Pro Ser Glu
755 760 765
Leu Arg Asp Leu Leu Ser Glu Phe Asn Val Leu Lys Gln Val Asn His
770 775 780
Pro His Val Ile Lys Leu Tyr Gly Ala Cys Ser Gln Asp Gly Pro Leu
785 790 795 800
Leu Leu Ile Val Glu Tyr Ala Lys Tyr Gly Ser Leu Arg Gly Phe Leu
805 810 815
Arg Glu Ser Arg Lys Val Gly Pro Gly Tyr Leu Gly Ser Gly Gly Ser
820 825 830
Arg Asn Ser Ser Ser Leu Asp His Pro Asp Glu Arg Ala Leu Thr Met
835 840 845
Gly Asp Leu Ile Ser Phe Ala Trp Gln Ile Ser Gln Gly Met Gln Tyr
850 855 860
Leu Ala Glu Met Lys Leu Val His Arg Asp Leu Ala Ala Arg Asn Ile
865 870 875 880
Leu Val Ala Glu Gly Arg Lys Met Lys Ile Ser Asp Phe Gly Leu Ser
885 890 895
Arg Asp Val Tyr Glu Glu Asp Ser Tyr Val Lys Arg Ser Gln Gly Arg
900 905 910
Ile Pro Val Lys Trp Met Ala Ile Glu Ser Leu Phe Asp His Ile Tyr
915 920 925
Thr Thr Gln Ser Asp Val Trp Ser Phe Gly Val Leu Leu Trp Glu Ile
930 935 940
Val Thr Leu Gly Gly Asn Pro Tyr Pro Gly Ile Pro Pro Glu Arg Leu
945 950 955 960
Phe Asn Leu Leu Lys Thr Gly His Arg Met Glu Arg Pro Asp Asn Cys
965 970 975
Ser Glu Glu Met Tyr Arg Leu Met Leu Gln Cys Trp Lys Gln Glu Pro
980 985 990
Asp Lys Arg Pro Val Phe Ala Asp Ile Ser Lys Asp Leu Glu Lys Met
995 1000 1005
Met Val Lys Arg Arg Asp Tyr Leu Asp Leu Ala Ala Ser Thr Pro
1010 1015 1020
Ser Asp Ser Leu Ile Tyr Asp Asp Gly Leu Ser Glu Glu Glu Thr
1025 1030 1035
Pro Leu Val Asp Cys Asn Asn Ala Pro Leu Pro Arg Ala Leu Pro
1040 1045 1050
Ser Thr Trp Ile Glu Asn Lys Leu Tyr Gly Arg Ile Ser His Ala
1055 1060 1065
Phe Thr Arg Phe
1070
<210> 33
<211> 26
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 33
tgtccagctt tgtgcctgat tgatgt 26
<210> 34
<211> 26
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 34
gctgggcact gaagagaaag gaatgc 26
<210> 35
<211> 24
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 35
agcaggatga gtgcggaggc aagc 24
<210> 36
<211> 33
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 36
ttaactatca aacgtgtcca ttaattttgc cgc 33
<210> 37
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 37
agtactgggg tggctggg 18
<210> 38
<211> 19
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 38
cactttggac aaggagatg 19
<210> 39
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 39
acagaactgg agctgcgg 18
<210> 40
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 40
ggactggtgt actcgctg 18
<210> 41
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 41
tcctagactg caggaaacac 20
<210> 42
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 42
catcgagaaa gtccagac 18
<210> 43
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 43
gctgctggag acattgaa 18
<210> 44
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 44
tcactgctgc tcagctca 18
<210> 45
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 45
gaggatccaa agtgggaatt 20
<210> 46
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 46
agtatctggc cgagatgaag 20
<210> 47
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 47
gcaaagacct ggagaagatg 20
<210> 48
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 48
aggacgttga actctgacag 20
<210> 49
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 49
cctttgcttc atccagaatc 20
<210> 50
<211> 22
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 50
gattttgtgt ttctccagct ct 22
<210> 51
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 51
cctgcttctc tgaggaagaa 20
<210> 52
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 52
gggccttagt ctcagcaaac 20
<210> 53
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 53
gagcactctg cgtgaactta 20
<210> 54
<211> 21
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 54
cagcttgttc atggtactga t 21
<210> 55
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 55
tggtgagtcc ttcaccag 18
<210> 56
<211> 22
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 56
cctagagttt ttccaagaac ca 22
<210> 57
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 57
catttaactg gaatccgacc 20
<210> 58
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 58
gactctctcc aggccagttc 20
<210> 59
<211> 22
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 59
ggctatcaga agtaaaacca cc 22
<210> 60
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 60
cgagagctga tggcacta 18
<210> 61
<211> 24
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 61
cttcatcaca agtgaagtac ttcc 24
<210> 62
<211> 19
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 62
cgtactccac gatgaggag 19
<210> 63
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 63
gattctggat gaagcaaagg 20
<210> 64
<211> 24
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 64
ggaagtactt cacttgtgat gaag 24
<210> 65
<211> 18
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 65
cccagccacc ccagtact 18
<210> 66
<211> 17
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 66
gtaaaacgac ggccagt 17
<210> 67
<211> 17
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 67
gttttcccag tcacgac 17
<210> 68
<211> 17
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 68
caggaaacag ctatgac 17
<210> 69
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 69
ctggagccac agtacccact 20
<210> 70
<211> 20
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 70
tccaaattcg ccttctccta 20
<210> 71
<211> 25
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 71
ttcatcagcc ttcctcaggg aggat 25
<210> 72
<211> 41
<212> DNA
<213> Artificial Sequence
<220>
<223> Primer
<400> 72
ggggacaagt ttgtacaaaa aagcaggctt cgccaccagc a 41
<210> 73
<211> 42
<212> DNA
<213> Artificial Sequence
<220>
<223> primer
<400> 73
ggggaccact ttgtacaaga aagctgggtt ttaactatca aa 42
<210> 74
<211> 21
<212> RNA
<213> Artificial Sequence
<220>
<223> RET siRNA1
<220>
<221> misc_feature
<223> n stand for thymine
<220>
<221> misc_feature
<222> (20)..(21)
<223> n stand for thymine
<400> 74
cacaugucau caaauuguan n 21
<210> 75
<211> 21
<212> RNA
<213> Artificial Sequence
<220>
<223> RET siRNA2
<220>
<221> misc_feature
<222> (20)..(21)
<223> n stand for thymine
<400> 75
ggauugaaaa caaacucuan n 21
<210> 76
<211> 21
<212> RNA
<213> Artificial Sequence
<220>
<223> RET siRNA
<220>
<221> misc_feature
<222> (20)..(21)
<223> n stand for thymine
<400> 76
gcuugucccg agauguuuan n 21
<210> 77
<211> 21
<212> RNA
<213> Artificial Sequence
<220>
<223> RET siRNA3
<220>
<221> misc_feature
<222> (20)..(21)
<223> n stand for thymine
<400> 77
ccacugcuac cacaaguuun n 21
Claims (23)
- DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드.
- 제1항에 있어서, 이하의 (a) 내지 (c)에서 선택되는 폴리펩티드.
(a) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열을 포함하는 폴리펩티드.
(b) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실, 또는 부가된 아미노산 서열을 포함하는 폴리펩티드.
(c) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열과 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드. - 제1항 또는 제2항에 기재된 폴리펩티드를 코딩하는 폴리뉴클레오티드.
- 제3항에 있어서, 이하의 (d) 내지 (f)에서 선택되는 폴리뉴클레오티드.
(d) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(e) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열에 있어서, 1개 또는 수개의 아미노산이 치환, 결실, 또는 부가된 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드.
(f) 서열번호 2, 서열번호 4, 서열번호 6, 서열번호 8, 서열번호 10, 서열번호 12, 서열번호 14, 서열번호 16, 서열번호 18, 서열번호 20, 서열번호 22, 또는 서열번호 24로 나타나는 아미노산 서열과 90% 이상의 동일성을 갖는 아미노산 서열을 포함하는 폴리펩티드를 코딩하는 폴리뉴클레오티드. - 제3항에 있어서, 이하의 (g) 내지 (i)에서 선택되는 폴리뉴클레오티드.
(g) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열을 포함하는 폴리뉴클레오티드.
(h) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열과 상보적인 염기 서열을 포함하는 폴리뉴클레오티드와 엄격한 조건 하에서 하이브리다이징하는 폴리뉴클레오티드.
(i) 서열번호 1, 서열번호 3, 서열번호 5, 서열번호 7, 서열번호 9, 서열번호 11, 서열번호 13, 서열번호 15, 서열번호 17, 서열번호 19, 서열번호 21, 또는 서열번호 23으로 나타나는 염기 서열과 90% 이상의 동일성을 갖는 폴리뉴클레오티드. - 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드를 포함하는 발현 벡터.
- 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드를 도입한 세포.
- 제1항 또는 제2항에 기재된 폴리펩티드에 특이적으로 결합하는 항체.
- 시료 중 제1항 또는 제2항에 기재된 폴리펩티드의 존재를 검출하는 방법.
- 시료 중 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하기 위한 프라이머 또는 프로브로서, 당해 프라이머 또는 프로브가 이하의 (j) 내지 (l)에서 선택되는 폴리뉴클레오티드.
(j) DCTN1 단백질을 코딩하는 폴리뉴클레오티드에 하이브리다이징하는 프로브 및 RET 단백질을 코딩하는 폴리뉴클레오티드에 하이브리다이징하는 프로브로 이루어지는 군에서 선택되는 적어도 하나의 프로브인 폴리뉴클레오티드.
(k) DCTN1 단백질을 코딩하는 폴리뉴클레오티드와 RET 단백질을 코딩하는 폴리뉴클레오티드와의 융합점에 하이브리다이징하는 프로브인 폴리뉴클레오티드.
(l) DCTN1 단백질을 코딩하는 폴리뉴클레오티드와 RET 단백질을 코딩하는 폴리뉴클레오티드와의 융합점을 끼워넣도록 설계된 센스 프라이머와 안티센스 프라이머의 세트인 폴리뉴클레오티드. - 시료 중 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하는 방법.
- 제9항 또는 제11항에 기재된 검출 방법에 있어서, 시료 중 제1항 또는 제2항에 기재된 폴리펩티드 또는 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출한 경우에, 시료의 유래가 되는 환자가 암이라고 판정하는 방법.
- RET를 저해하는 화합물을 유효 성분으로 하는, DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 치료용 의약 조성물.
- 이하의 (1) 및 (2)의 공정을 포함하는 제1항 또는 제2항에 기재된 폴리펩티드의 발현 및/또는 활성, 또는 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 발현을 억제하는 화합물을 스크리닝하는 방법.
(1) 제1항 또는 제2항에 기재된 폴리펩티드, 제1항 또는 제2항에 기재된 폴리펩티드 또는 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드를 발현하고 있는 세포, 또는 제7항의 세포에 시험 화합물을 접촉하는 공정.
(2) 상기 공정 (1)에 있어서, 제1항 또는 제2항에 기재된 폴리펩티드의 발현 및/또는 활성, 또는 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 발현이 억제되는지 측정하는 공정, 또는 상기 공정 (1)에 기재된 세포의 증식이 억제되는지 측정하는 공정. - 제1항 또는 제2항에 기재된 폴리펩티드 또는 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드를, RET를 저해하는 화합물을 사용한 화학 요법이 유효한지 여부의 지표로 하는 방법으로서, 제9항에 기재된 검출 방법에 의해 시료 중에서 제1항 또는 제2항에 기재된 폴리펩티드를 검출한 경우, 및/또는 제11항에 기재된 검출 방법에 의해 시료 중에서 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출한 경우에, RET를 저해하는 화합물을 사용한 화학 요법이 유효하다고 판정하는 방법.
- DCTN1 단백질의 N 말단 부분과, RET 단백질의 C 말단 부분이 융합되어 있는 폴리펩티드 및 당해 폴리펩티드를 코딩하는 폴리뉴클레오티드로 이루어지는 군에서 선택되는 적어도 1종을 포함하는 암을 검출하기 위한 바이오마커.
- DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 환자에게, RET를 저해하는 화합물을 사용한 화학 요법을 행하는 공정을 포함하는, 암의 치료 방법.
- 피험자 유래의 시료 중에서 제1항 또는 제2항에 기재된 폴리펩티드의 존재를 검출하는 것 및/또는 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하는 것을 행하는 공정, 그리고
제1항 또는 제2항에 기재된 폴리펩티드의 존재가 검출되고/되거나 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재가 검출된 경우에, 당해 피험자에 대하여, RET를 저해하는 화합물을 사용한 화학 요법을 행하는 공정을 포함하는, 암의 치료 방법. - DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 환자를 치료하기 위한, RET를 저해하는 화합물.
- DCTN1 유전자와 RET 유전자와의 융합 유전자 양성 및/또는 DCTN1 단백질과 RET 단백질과의 융합 단백질 양성인 암 환자를 치료하기 위한 암 치료용 의약 조성물을 제조하기 위한, RET를 저해하는 화합물의 사용.
- 시료 중 제1항 또는 제2항에 기재된 폴리펩티드의 존재를 검출하기 위한 수단 및/또는 시료 중 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하기 위한 수단의, RET를 저해하는 화합물을 사용한 화학 요법이 유효한지 여부의 판정약의 제조 방법.
- 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하기 위한 항DCTN1 항체 및 항RET 항체의 조합.
- 제1항 또는 제2항에 기재된 폴리펩티드 또는 제3항 내지 제5항 중 어느 한 항에 기재된 폴리뉴클레오티드의 존재를 검출하기 위한 검출약을 제조하기 위한, 제8항에 기재된 항체, 제22항에 기재된 항체의 조합 또는 제10항에 기재된 프라이머 또는 프로브의 사용.
Applications Claiming Priority (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JPJP-P-2017-158796 | 2017-08-21 | ||
| JP2017158796 | 2017-08-21 | ||
| PCT/JP2018/030688 WO2019039439A1 (ja) | 2017-08-21 | 2018-08-20 | Dctn1タンパク質とretタンパク質との融合タンパク質 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| KR20200038528A true KR20200038528A (ko) | 2020-04-13 |
Family
ID=65439881
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| KR1020207007893A KR20200038528A (ko) | 2017-08-21 | 2018-08-20 | Dctn1 단백질과 ret 단백질과의 융합 단백질 |
Country Status (15)
| Country | Link |
|---|---|
| US (1) | US20200190154A1 (ko) |
| EP (1) | EP3674325A4 (ko) |
| JP (1) | JP7033143B2 (ko) |
| KR (1) | KR20200038528A (ko) |
| CN (1) | CN111032696B (ko) |
| AU (1) | AU2018322286B2 (ko) |
| BR (1) | BR112020003006A2 (ko) |
| CA (1) | CA3073375A1 (ko) |
| MA (1) | MA49988A (ko) |
| MX (1) | MX2020002002A (ko) |
| MY (1) | MY201941A (ko) |
| PH (1) | PH12020500269A1 (ko) |
| SG (1) | SG11202001212SA (ko) |
| TW (1) | TW201915027A (ko) |
| WO (1) | WO2019039439A1 (ko) |
Families Citing this family (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2024500628A (ja) * | 2020-11-20 | 2024-01-10 | ヘルシン ヘルスケア ソシエテ アノニム | 腫瘍治療のための4-アミノ-n-[4-(メトキシメチル)フェニル]-7-(1-メチルシクロプロピル)-6-(3-モルホリノプロパ-1-イン-1-イル)-7h-ピロロ[2,3-d]ピリミジン-5-カルボキサミドの使用法 |
Family Cites Families (11)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN100478355C (zh) * | 2002-08-07 | 2009-04-15 | 上海新世界基因技术开发有限公司 | 具有促进小鼠nih/3t3细胞转化功能的新的人蛋白及其编码序列 |
| WO2013018882A1 (ja) * | 2011-08-04 | 2013-02-07 | 独立行政法人国立がん研究センター | Kif5b遺伝子とret遺伝子との融合遺伝子、並びに該融合遺伝子を標的としたがん治療の有効性を判定する方法 |
| WO2013059740A1 (en) * | 2011-10-21 | 2013-04-25 | Foundation Medicine, Inc. | Novel alk and ntrk1 fusion molecules and uses thereof |
| US10023855B2 (en) * | 2011-10-31 | 2018-07-17 | Macrogen, Inc. | Fusion protein comprising C-terminal domain of RET protein and use thereof as a diagnosing marker |
| US20160010068A1 (en) * | 2013-02-22 | 2016-01-14 | Boris C. Bastian | Fusion polynucleotides and fusion polypeptides associated with cancer and particularly melanoma and their uses as therapeutic and diagnostic targets |
| WO2015117040A1 (en) * | 2014-01-31 | 2015-08-06 | Swift Biosciences, Inc. | Improved methods for processing dna substrates |
| US10202365B2 (en) | 2015-02-06 | 2019-02-12 | Blueprint Medicines Corporation | 2-(pyridin-3-yl)-pyrimidine derivatives as RET inhibitors |
| HUE053067T2 (hu) | 2015-07-16 | 2021-06-28 | Array Biopharma Inc | Helyettesített pirazolo[1,5-A]piridin vegyületek, mint RET kináz inhibitorok |
| MA41559A (fr) | 2015-09-08 | 2017-12-26 | Taiho Pharmaceutical Co Ltd | Composé de pyrimidine condensé ou un sel de celui-ci |
| AU2016362876B2 (en) * | 2015-12-01 | 2023-02-09 | LGC Clinical Diagnostics, Inc. | Multiplex cellular reference materials |
| BR112018016724B1 (pt) | 2016-02-23 | 2024-02-20 | Taiho Pharmaceutical Co., Ltd | Composto de pirimidina condensado ou sal do mesmo, seus usos, inibidor de ret, agente antitumor e composição farmacêutica |
-
2018
- 2018-08-20 MX MX2020002002A patent/MX2020002002A/es unknown
- 2018-08-20 CN CN201880054583.2A patent/CN111032696B/zh active Active
- 2018-08-20 JP JP2019537622A patent/JP7033143B2/ja active Active
- 2018-08-20 AU AU2018322286A patent/AU2018322286B2/en active Active
- 2018-08-20 BR BR112020003006-6A patent/BR112020003006A2/pt unknown
- 2018-08-20 MA MA049988A patent/MA49988A/fr unknown
- 2018-08-20 EP EP18849160.9A patent/EP3674325A4/en active Pending
- 2018-08-20 KR KR1020207007893A patent/KR20200038528A/ko not_active Application Discontinuation
- 2018-08-20 SG SG11202001212SA patent/SG11202001212SA/en unknown
- 2018-08-20 US US16/640,955 patent/US20200190154A1/en active Pending
- 2018-08-20 MY MYPI2020000799A patent/MY201941A/en unknown
- 2018-08-20 TW TW107129031A patent/TW201915027A/zh unknown
- 2018-08-20 CA CA3073375A patent/CA3073375A1/en active Pending
- 2018-08-20 WO PCT/JP2018/030688 patent/WO2019039439A1/ja unknown
-
2020
- 2020-02-05 PH PH12020500269A patent/PH12020500269A1/en unknown
Non-Patent Citations (6)
| Title |
|---|
| Cancer Discov., 3(6), pp630-5(2013) |
| Cancer, 115(16), pp3801-7(2009) |
| Cell, 159(3), pp676-90(2014) |
| Lancet Respir Med., 5(1), pp42-50(2017) |
| Nature, 511(7511), pp543-50(2014) |
| Oncogene, 22(29), pp4578-80(2003) |
Also Published As
| Publication number | Publication date |
|---|---|
| MY201941A (en) | 2024-03-25 |
| AU2018322286A1 (en) | 2020-04-02 |
| EP3674325A4 (en) | 2021-11-03 |
| CN111032696A (zh) | 2020-04-17 |
| JP7033143B2 (ja) | 2022-03-09 |
| SG11202001212SA (en) | 2020-03-30 |
| RU2020111214A (ru) | 2021-09-24 |
| EP3674325A1 (en) | 2020-07-01 |
| MA49988A (fr) | 2020-07-01 |
| CN111032696B (zh) | 2023-11-03 |
| PH12020500269A1 (en) | 2020-09-21 |
| CA3073375A1 (en) | 2019-02-28 |
| MX2020002002A (es) | 2020-07-20 |
| WO2019039439A1 (ja) | 2019-02-28 |
| RU2020111214A3 (ko) | 2021-12-17 |
| TW201915027A (zh) | 2019-04-16 |
| US20200190154A1 (en) | 2020-06-18 |
| JPWO2019039439A1 (ja) | 2020-08-13 |
| BR112020003006A2 (pt) | 2020-08-11 |
| AU2018322286B2 (en) | 2022-03-10 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP6493681B2 (ja) | 肺がんで見出された新規融合遺伝子 | |
| EP1692281B8 (fr) | Identification d'une mutation de jak2 impliquee dans la polyglobulie de vaquez | |
| EP1730303B1 (en) | Mutations of the pik3ca gene in human cancers | |
| JP2006500946A (ja) | 精巣精上皮腫の診断方法 | |
| US20130137111A1 (en) | Detection method of novel ret fusion | |
| WO2016084883A1 (ja) | 胆道がんにおける新規治療標的融合遺伝子 | |
| TW202018090A (zh) | 癌促進因子表現抑制劑、其有效成分的篩選方法、對於該方法有用的表現匣、診斷藥及診斷方法 | |
| WO2015093557A1 (ja) | 胃がんの責任因子としての新規融合遺伝子 | |
| CN111032696B (zh) | Dctn1蛋白质与ret蛋白质的融合蛋白 | |
| RU2813996C2 (ru) | Слитый белок из белка dctn1 с белком ret | |
| KR101496745B1 (ko) | 인간 암세포의 불사화에 관련되는 유전자 및 그 용도 | |
| KR101897322B1 (ko) | 원발성 갑상선암 전이의 진단방법 및 이를 이용한 진단키트 | |
| WO2022244807A1 (ja) | Ltk融合遺伝子 | |
| KR101683961B1 (ko) | 방광암 재발 진단 마커 | |
| KR101104105B1 (ko) | 대장암 진단용 조성물 및 그 용도 | |
| US20040072192A1 (en) | Cancer-linked genes as targets for chemotherapy | |
| KR20110036559A (ko) | 대장암 진단용 조성물 및 그 용도 | |
| KR20110036560A (ko) | 대장암 진단용 조성물 및 그 용도 | |
| KR20110036561A (ko) | 대장암 진단용 조성물 및 그 용도 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| A201 | Request for examination | ||
| E902 | Notification of reason for refusal |