Disclosure of Invention
The invention provides a new application of a SlCYP707A gene as a negative regulatory factor in improving the resistance of tomato at a sub-low temperature, and provides a basis for cultivating a tomato variety resistant to the sub-low temperature.
The specific technical scheme is as follows:
the invention provides an application of a SlCYP707A gene as a negative regulatory factor in improving the resistance of tomato at a sub-low temperature, wherein the SlCYP707A gene is at least one of a SlCYP707A1 gene and a SlCYP707A2 gene; the nucleotide sequence of the SlCYP707A1 gene is shown in SEQ ID No.1, and the nucleotide sequence of the SlCYP707A2 gene is shown in SEQ ID No. 2.
The amino acid sequence of the SlCYP707A1 gene is shown in SEQ ID No. 3; the amino acid sequence of the SlCYP707A2 gene is shown in SEQ ID No. 4.
The knockout of the SlCYP707A1 or SlCYP707A2 gene has no obvious influence on the growth phenotype of the tomato at normal temperature (22 ℃/20 ℃), but the root growth of a single mutant of the two genes is obviously stronger than that of a control group in the environment of sub-low temperature (16 ℃/12 ℃), and the root development is more vigorous after the two genes are jointly mutated; meanwhile, the biomass and root cap ratio of the single mutant and the double mutant of the SlCYP707A1 gene and the SlCYP707A2 gene on the overground part at the sub-low temperature are also obviously increased; after the treatment at the sub-low temperature, the resistance of plants knocked out by the SlCYP707A1 and SlCYP707A2 genes is obviously enhanced, mainly expressed by that the maximum photochemical efficiency (Fv/Fm) of leaves is improved, and the relative electrolyte permeability of the leaves is reduced.
Furthermore, the application approach is to obtain a single mutant by knocking out the SlCYP707A1 gene or the SlCYP707A2 gene, or to obtain a homozygous double mutant by hybridizing the single mutant of the SlCYP707A1 gene and the single mutant of the SlCYP707A2 gene, so that the sub-low temperature resistance of the tomato mutant is improved.
Furthermore, the application approach is to obtain a single mutant by knocking out the SlCYP707A1 gene or the SlCYP707A2 gene, or to obtain a homozygous double mutant by hybridizing the single mutant of the SlCYP707A1 gene and the single mutant of the SlCYP707A2 gene, so that the relative electrolyte permeability and/or the maximum photochemical efficiency of leaves are reduced, and the sub-low temperature resistance of the tomato mutant is improved.
Further, the means of the knockout is CRISPR/Cas9 gene editing technology.
According to experimental observation, the plant height, stem thickness, aboveground part biomass and underground part biomass at normal temperature do not change obviously after single mutation and double mutation of SlCYP707A1 and SlCYP707A2 genes, but after sub-low temperature treatment, compared with a control group, the root system of the mutant material grows better, the aboveground part biomass, the underground part biomass and the root cap ratio are all increased obviously, and the double mutant grows better than the single mutant.
The resistance phenotype analysis also proves that the plant resistance of the SlCYP707A1 and SlCYP707A2 genes after single mutation and double mutation is obviously improved, and the results show that compared with a control group, the mutant has the advantages that the maximum photochemical efficiency (Fv/Fm) of leaves is improved, the relative electrolyte permeability of the leaves is reduced, and the resistance of the double mutant is stronger than that of the single mutant. The discovery provides a new idea for creating the low-temperature resistant germplasm of the tomato by degrading the key gene CYP707As through ABA.
Further, the application comprises the steps of:
(1) designing a target sequence of the SlCYP707A gene, and constructing a CRISPR/Cas9 vector;
the nucleotide sequence of the target sequence sgRNA1 is shown in SEQ ID NO.5 or SEQ ID NO. 6;
(2) constructing agrobacterium gene engineering bacteria containing the CRISPR/Cas9 vector in the step (1);
(3) transforming the genetically engineered bacteria in the step (2) into tomato cotyledons to obtain a homozygous single mutant strain which does not contain exogenous Cas9 protein and is stably inherited.
Further, the method also comprises the step (4): crossing the single mutant lacking the SlCYP707A1 gene and the single mutant lacking the SlCYP707A2 gene to obtain an F1 generation, selfing to obtain an F2 generation, and finding out a homozygous double-mutant plant lacking the SlCYP707A1 and the SlCYP707A2 gene from an F2 generation segregating population by a sequencing technology.
In view of the above findings, a double mutant strain of genes SlCYP707A1 and SlCYP707A2 can be used as a male parent, hybridized with a variety with excellent properties but low-temperature resistance to be improved as a female parent, backcrossed with the female parent for multiple generations, and then selfed, and a variety with more excellent comprehensive properties can be screened in a later generation group.
Compared with the prior art, the invention has the following beneficial effects:
(1) the invention utilizes CRISPR/Cas9 gene editing technology to respectively carry out site-directed knockout on SlCYP707A1 and SlCYP707A2 genes to construct single mutants, the two single mutants are hybridized, and after selfing for one generation, homozygous SlCYP707A1 and SlCYP707A2 double-mutant materials are separated from F2 generation, and the mutant materials have stronger growth and resistance in a sub-low temperature environment.
(2) The invention utilizes gene editing technology to obtain single mutants and double mutants of SlCYP707A1 and SlCYP707A2 genes, and a series of growth phenotype observation and resistance experiments prove that after sub-low temperature treatment, compared with a control group of tomatoes, the biomass of the overground parts and the underground parts of the single mutants and the double mutants of the SlCYP707A1 and SlCYP707A2 genes are obviously improved, the root cap ratio is increased, and the resistance is enhanced. The method is helpful for deeply understanding the regulation and control functions and action mechanisms of ABA on plant growth and development under adversity stress, and can provide a new way for cultivating plant growth reinforced tomato germplasm at a sub-low temperature by combining hybrid breeding.
(3) The application method provided by the invention can be applied to low-temperature resistance breeding of common tomatoes and has the advantages of high efficiency, rapidness, simplicity, convenience and feasibility.
Detailed Description
The present invention will be further described with reference to the following specific examples, which are only illustrative of the present invention, but the scope of the present invention is not limited thereto.
Example 1 qRT-PCR analysis of expression of SlCYP707A1 and SlCYP707A2 genes in different tissues and organs of tomato
The expression patterns of SlCYP707A1 and SlCYP707A2 genes were investigated by qRT-PCR.
The specific method comprises the following steps:
by using
Detection was performed by 480II fluorescent quantitative PCR instrument (Roche, Swiss). The reaction system is described in detail in 2 × SYBR Green Supermix (Vazyme) instructions.
The specific primers of the SlCYP707A1 gene are (SlCYP707A 1-F: 5'-GTCCAGGGAATGAACTTGCC-3'; SlCYP707A 1-R: 5'-TTGCAGCTAGTCCACCCAAT-3'), and the specific primers of the SlCYP707A2 gene are (SlCYP707A 2-F: 5'-CTCGGGTCATTCATGGGAGA-3'; SlCYP707A 2-R: 5'-AGCAGTAGTGTCTCGAGCTG-3') and utilize 2-ΔΔCtThe relative expression level of the gene is calculated.
As a result, it was found that: the genes SlCYP707A1 and SlCYP707A2 are expressed in different tissues, the expression level of SlCYP707A1 is the highest in leaves, and the expression level of SlCYP707A1 is the lowest in roots. SlCYP707A2 expresses the highest amount in roots, while the expression amounts in terminal buds, lateral buds, leaves and stems are relatively low.
Example 2
First, acquisition of single mutant and double mutant of SlCYP707A1 and SlCYP707A2 genes
In order to determine the influence of SlCYP707A1 and SlCYP707A2 gene deletion on tomato plant growth and low-temperature resistance, target sequences sgRNA1 and sgRNA2 of SlCYP707A1 and SlCYP707A2 are respectively designed, and pCAMBIA1301-U6-26-sgRNA1-SlCYP707A1/A2-35S-Cas9SK vectors are constructed by enzyme digestion and connection. The vector is introduced into tomato plants through genetic transformation, and functions of the tomato plants are researched by knocking out SlCYP707A1 and SlCYP707A2 respectively by using a CRISPR/Cas9 technology.
The method comprises the following specific steps:
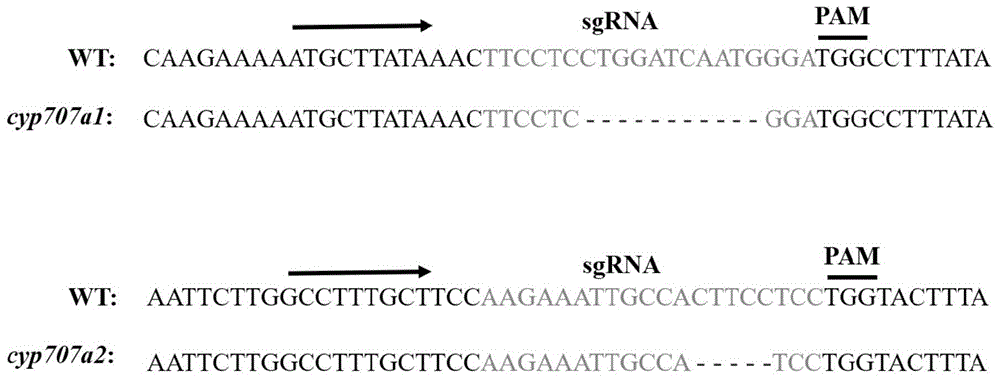
the CRISPR-P website (http:// cbi. hzau. edu. cn/cgi-bin/CRISPR) was used to design the target sequence sgRNA1 of the SlCYP707A1 gene: 5'-TTCCTCCTGGATCAATGGGA-3' and the target sequence sgRNA2 of the SlCYP707a2 gene: 5'-AAGAAATTGCCACTTCCTCC-3' are provided.
Annealing the synthesized sgRNA1 and sgRNA2 sequences (single strands) to form a double-stranded sgRNA, wherein both ends of the sgRNA have Bbs I restriction endonuclease sites. The formed sgRNA is respectively connected with AtU6-26SK vectors cut by Bbs I restriction endonuclease, positive plasmids are extracted for later use, and the names are U6-26-sgRNA1-SlCYP707A 1-SK and U6-26-sgRNA1-SlCYP707A 2-SK. The U6-26-sgRNA1-SlCYP707A1/A2-SK and 35S-Cas9SK vectors are subjected to double enzyme digestion by Kpn I and Sal I restriction enzymes, and the respective enzyme digestion products are recovered and connected. PCR detection of bacterial liquid, sequencing verification of positive clone, extraction of positive plasmid for later use, and designation of U6-26-sgRNA1-SlCYP707A1/A2-35S-cas9 SK.
U6-26-sgRNA1-SlCYP707A1/A2-35S-cas9SK and pCAMBIA1301 vectors are subjected to double enzyme digestion by using Kpn I and Xba I restriction endonucleases, and a band of about 6kb is recovered from U6-26-sgRNA1-SlCYP707A1/A2-35S-cas9SK and is connected to the digested pCAMBIA1301 vector. The ligation product was transformed into E.coli DH 5. alpha. competent cells, and a single colony was picked up and cultured overnight in liquid LB medium containing 50mg/L kanamycin (Kan) at 37 ℃ with shaking at 200 rpm. Primers were designed at the 5' end of the pCAMBIA1301 vector for PCR detection of bacterial suspension (about 550 bp). Sequencing and verifying positive clones, extracting positive plasmids, and naming the plasmids as pCAMBIA1301-U6-26-sgRNA1-SlCYP707A1-35S-cas9SK and pCAMBIA1301-U6-26-sgRNA1-SlCYP707A2-35S-cas9 SK.
Transferring the vector into GV3101 agrobacterium to infect common tomato cotyledons, obtaining resistant buds of a transformation pCAMBIA1301-U6-26-sgRNA1-SlCYP707A1/A2-35S-cas9SK knockout vector by a plant tissue culture technology, transplanting after rooting culture, separating and identifying a SlCYP707A1 and SlCYP707A2 gene homozygous mutant plant after selfing for one generation.
And hybridizing the SlCYP707A1 and SlCYP707A2 gene homozygous mutant plants to obtain an F1 generation, selfing to obtain an F2 generation, and reserving the plants which are simultaneously subjected to SlCYP707A1 and SlCYP707A2 gene homozygous mutation as double mutants for subsequent experimental study by sequencing verification from a segregation population.
Secondly, observing growth phenotype and measuring growth indexes of SlCYP707A1 and SlCYP707A2 gene mutants at normal temperature and subambient temperature
Observing the growth phenotype of the SlCYP707A1 and SlCYP707A2 gene mutants at normal temperature and sub-low temperature, and measuring growth indexes;
the specific operation is as follows:
common (wild type, WT) tomatoes (Solanum lycopersicum cv. condine Red) and SlCYP707A1 and SlCYP707A2 gene single mutants and double mutants were used as experimental materials. Will be provided withSoaking the seeds in warm water at 50 ℃ for 15min, then placing the seeds in a constant speed shaking table (200rpm/min) at 28 ℃ for about 2 days, changing water once a day, and sowing the seeds in a container filled with grass peat when the length of the embryo roots of the seeds reaches about 1 cm: vermiculite 2: 1, 72-hole cell plate. Placing the plug in a plant growth chamber, wherein the growth conditions are as follows: the light period is 12h/12h, the ambient temperature is 22 ℃/20 ℃, the relative humidity is about 75 percent, and the average light intensity is 200 mu mol m- 2s-1. The nutrient solution is 1/2Hoagland nutrient solution. When the seedlings grow to have three leaves and one heart, transplanting the single seedlings into a plastic pot for culture under the same culture conditions, and evenly dividing the removed tomato seedlings into two parts after one week. One batch of seedlings is transferred to an artificial climate box according to the original culture conditions, the conditions such as illumination, water and fertilizer are unchanged, the environmental temperature is 16 ℃/12 ℃, and after treatment is carried out for 10 ℃, the tomato seedlings treated at normal temperature and sub-low temperature are observed and counted.
The method for observing and measuring the growth indexes comprises the following steps: selecting materials with consistent growth vigor from each material for statistics, wherein the plant height statistics starts from the cotyledon of the plant to the plant height of the terminal bud; stem thickness statistics the diameter of a two-internode stem is measured with a vernier caliper; the biomass statistics takes the root-stem junction as a dividing point, and the mass of the overground part and the underground part is weighed; taking root length statistics, wherein a root-tuber junction is taken as an initial position, bunching root systems, and taking the tail end as an end point to measure the length; each treatment contained 6 biological replicates.
As a result: the growth phenotype of the SlCYP707A1 and SlCYP707A2 knockout tomato mutant at normal temperature is not obviously different from that of a control group (WT), but the mutant shows that the mutant has the following characteristics after being treated at the sub-low temperature: the ratio of the biomass of the underground part of the plant, the biomass of the overground part and the root cap is increased.
Thirdly, observing and measuring resistance indexes of SlCYP707A1 and SlCYP707A2 gene mutants after sub-low temperature treatment
In order to explore the influence of the sub-low temperature environment on the photosynthetic efficiency and leaf electrolyte permeability of tomato SlCYP707A1 and SlCYP707A2 knockout mutants, Wild Type (WT) and mutant plants were subjected to sub-low temperature treatment at 16 ℃/12 ℃.
Determination of relative electrolyte permeability: cutting the main leaf vein parallel to the leaf into 0.5cm long strips, weighing 0.2g placed in a container containing 20mL of ddH2O in a 50ml centrifuge tube, incubated at 28 ℃ for 2 hours with a shaker (200rpm/min), and the conductivity was measured as EC1 with a digital conductivity meter (DDS-11A, Otorion instruments, Ltd., Hangzhou). Then placing the centrifuge tube with the sample in a water bath kettle at 100 ℃ for half an hour, cooling and then measuring the conductivity again, wherein the measured value is EC 2; finally, the relative electrolyte permeability EC (%) ═ EC1/EC2 × 100% was calculated.
Determination of maximum photochemical efficiency of the blade: and (3) carrying out dark treatment on the tomato plant for half an hour, selecting the second and third leaf sections from top to bottom, and carrying out chlorophyll fluorescence measurement by using an Imaging PAM fluorescence Imaging system (IMAG-MAX/L, Germany) to obtain chlorophyll fluorescence parameters such as maximum photochemical efficiency (Fv/Fm).
As a result: compared with a control plant, the resistance of the SlCYP707A1 and SlCYP707A2 knockout mutants is obviously improved, the relative electrolyte permeability of leaves is reduced, the maximum photochemical efficiency (Fv/Fm) is improved, and the resistance of double mutants is better than that of single mutants.
Sequence listing
<110> Anqing city Ming triangle future industry research institute
ZHEJIANG University
Application of <120> SlCYP707A gene as negative regulatory factor in promotion of tomato sub-low temperature resistance
<160> 10
<170> SIPOSequenceListing 1.0
<210> 1
<211> 1431
<212> DNA
<213> tomato (Solanum lycopersicum L.)
<400> 1
atggttaatt actttgaaat atttctctat atctctatgt ttgttcttgg ttacttgtct 60
tattattttt gttttggcaa aaataacaat tcatcatcca agaaaaatgc ttataaactt 120
cctcctggat caatgggatg gccttatatt ggtgaaactc ttcaacttta ctctcaagac 180
cctaatgcct tcttcattaa taggcaaaga aggtttggtg aaatttttaa gacaaaaatt 240
ctaggttgtc catgtgtgat gttggcaagt ccagaagctg ctagatttgt actagtgaac 300
caagcaaatt tgtttaagcc aacttatcct aaaagtaaag agaatttaat tggtcaatct 360
gcaatttttt ttcatcaagg agattatcat aatcacctta gaaaacttgt tcaagctcct 420
ttaaatccag aatccattcg aaatcaaatc ccttatatcg aagaattatc gatttctgca 480
ttgaattcct gggttggagg acatgttgtc aacacttatc atgagatgaa aaagttttct 540
tttgaagtag gtatacttgc tatatttgga catttggatg gtcatgttaa ggaagaattg 600
aagaagaatt atagcatagt tgataaaggt tataattcat ttccaataaa tttgccaggg 660
actctttaca gaaaagccct tcaagcaaga aagaaacttg ggaagatttt gagtgaaata 720
ataagagaaa tgaaggagaa gaagacacta gaaaaagggt tgttgagttg tttcttgaat 780
gctaaagaag aaaaaggatt tttggtatta aatgaagatc aaattgctga taatataatt 840
ggagttttat ttgctgcaca agatacaaca gcaagtgttt taacttggat tataaagtat 900
cttcatgata atccaaagct tcttgagtgt gtcaaggctg aacagaaagt tatttggcag 960
tcaaatgaac aagaaaatca tgggttgaca tggacacaaa caagaaagat gcctatcact 1020
agcagggttg ttttggagac actgagaatg gctagtatca tatcttttgc atttagagag 1080
gctgtagctg atgtggaata caaaggatac ctaattccaa aaggatggaa ggtgatgcct 1140
ttgttcagaa atattcatca caacccagag ttctttcctg atccacaaaa atttgatcct 1200
tcaagatttg agaatgcgcc gaaacccaat acatttatgc catttggcag tggtgtacat 1260
gcttgtccag ggaatgaact tgccaagctg gaaattctca ttatgacaca tcatctagtc 1320
actaaattca ggtgggaagt ggtaggatct ggtagtggca ttcaatatgg accattccca 1380
gtcccattgg gtggactagc tgcaagattt tggaaaacta catcaaccta a 1431
<210> 2
<211> 1410
<212> DNA
<213> tomato (Solanum lycopersicum L.)
<400> 2
atggaatttg tttctatgtt gtgtttgttt actttcattt ctttaactct tcttctaatc 60
cattctatct tcaaattctt ggcctttgct tccaagaaat tgccacttcc tcctggtact 120
ttaggtttac cttatattgg tgaaaccttc caactctact cacaaaatcc caatgttttc 180
tttgcttcca aagtcaaaaa gtatggttca attttcaaga cttacatatt gggttgtcct 240
tgtgtaatga tatcaagtcc agaggcagct aaacaagttt tggtcacaaa ggctaatttg 300
tttaagccta catttcctgc tagcaaagaa agaatgctgg gaaaacaagc aattttcttt 360
catcaaggtg attatcatgc caaattgaga aaattagtcc tccaggcttt caagcccgat 420
tctatcagaa acatcatccc cgacattgaa tccatcgcga taacatcact cgaatcattt 480
caaggaagat tgatcaacac ttatcaagaa atgaagacat atacattcaa tgtggcattg 540
atttcgatat ttggtaaaga tgaatttcta tacagagagg agctcaagaa atgttactac 600
attctcgaaa aaggatacaa ttcgatgcca attaatctcc ccggtacact cttcaacaaa 660
gcaatgaaag cgaggaaaga gctagctaaa atcgttgcca aaatcatctc gactagacga 720
gaaatgaaga ttgatcatgg cgatttgctc gggtcattca tgggagataa agaaggactc 780
actgacgaac aaattgcaga taatgtaatc ggagtcatct ttgcagctcg agacactact 840
gctagtgttc ttacatggat cctcaaatac cttggagaaa atcccagtgt cctacaagct 900
gtcacagaag agcaagagaa cataatgaga aaaaaagagg tgaatggtga agaaaaagtt 960
ttaaattggc aagatacaag acaaatgcca atgacaacaa gagttattca agaaacactt 1020
agagttgctt caattttatc attcacattt agagaagctg ttgaagatgt tgaatttgaa 1080
ggatatttaa tacctaaagg atggaaagta ttaccactct ttaggaatat tcatcatagt 1140
ccagacaatt ttcctgaacc agagaaattt gatccttcaa gatttgaggt gtcaccaaaa 1200
cccaatacat tcatgccatt tggcaatggg gtccactcat gtccagggaa tgacttagcc 1260
aagctggaga ttttgatcct tgtacatcat ctgaccacaa agtacaggtg gtctatggtg 1320
ggcccacaaa atggaattca gtatgggcca tttgctcttc cccaaaatgg tttacccatt 1380
aaactctctc tcaaaacatc atcaacataa 1410
<210> 3
<211> 476
<212> PRT
<213> tomato (Solanum lycopersicum L.)
<400> 3
Met Val Asn Tyr Phe Glu Ile Phe Leu Tyr Ile Ser Met Phe Val Leu
1 5 10 15
Gly Tyr Leu Ser Tyr Tyr Phe Cys Phe Gly Lys Asn Asn Asn Ser Ser
20 25 30
Ser Lys Lys Asn Ala Tyr Lys Leu Pro Pro Gly Ser Met Gly Trp Pro
35 40 45
Tyr Ile Gly Glu Thr Leu Gln Leu Tyr Ser Gln Asp Pro Asn Ala Phe
50 55 60
Phe Ile Asn Arg Gln Arg Arg Phe Gly Glu Ile Phe Lys Thr Lys Ile
65 70 75 80
Leu Gly Cys Pro Cys Val Met Leu Ala Ser Pro Glu Ala Ala Arg Phe
85 90 95
Val Leu Val Asn Gln Ala Asn Leu Phe Lys Pro Thr Tyr Pro Lys Ser
100 105 110
Lys Glu Asn Leu Ile Gly Gln Ser Ala Ile Phe Phe His Gln Gly Asp
115 120 125
Tyr His Asn His Leu Arg Lys Leu Val Gln Ala Pro Leu Asn Pro Glu
130 135 140
Ser Ile Arg Asn Gln Ile Pro Tyr Ile Glu Glu Leu Ser Ile Ser Ala
145 150 155 160
Leu Asn Ser Trp Val Gly Gly His Val Val Asn Thr Tyr His Glu Met
165 170 175
Lys Lys Phe Ser Phe Glu Val Gly Ile Leu Ala Ile Phe Gly His Leu
180 185 190
Asp Gly His Val Lys Glu Glu Leu Lys Lys Asn Tyr Ser Ile Val Asp
195 200 205
Lys Gly Tyr Asn Ser Phe Pro Ile Asn Leu Pro Gly Thr Leu Tyr Arg
210 215 220
Lys Ala Leu Gln Ala Arg Lys Lys Leu Gly Lys Ile Leu Ser Glu Ile
225 230 235 240
Ile Arg Glu Met Lys Glu Lys Lys Thr Leu Glu Lys Gly Leu Leu Ser
245 250 255
Cys Phe Leu Asn Ala Lys Glu Glu Lys Gly Phe Leu Val Leu Asn Glu
260 265 270
Asp Gln Ile Ala Asp Asn Ile Ile Gly Val Leu Phe Ala Ala Gln Asp
275 280 285
Thr Thr Ala Ser Val Leu Thr Trp Ile Ile Lys Tyr Leu His Asp Asn
290 295 300
Pro Lys Leu Leu Glu Cys Val Lys Ala Glu Gln Lys Val Ile Trp Gln
305 310 315 320
Ser Asn Glu Gln Glu Asn His Gly Leu Thr Trp Thr Gln Thr Arg Lys
325 330 335
Met Pro Ile Thr Ser Arg Val Val Leu Glu Thr Leu Arg Met Ala Ser
340 345 350
Ile Ile Ser Phe Ala Phe Arg Glu Ala Val Ala Asp Val Glu Tyr Lys
355 360 365
Gly Tyr Leu Ile Pro Lys Gly Trp Lys Val Met Pro Leu Phe Arg Asn
370 375 380
Ile His His Asn Pro Glu Phe Phe Pro Asp Pro Gln Lys Phe Asp Pro
385 390 395 400
Ser Arg Phe Glu Asn Ala Pro Lys Pro Asn Thr Phe Met Pro Phe Gly
405 410 415
Ser Gly Val His Ala Cys Pro Gly Asn Glu Leu Ala Lys Leu Glu Ile
420 425 430
Leu Ile Met Thr His His Leu Val Thr Lys Phe Arg Trp Glu Val Val
435 440 445
Gly Ser Gly Ser Gly Ile Gln Tyr Gly Pro Phe Pro Val Pro Leu Gly
450 455 460
Gly Leu Ala Ala Arg Phe Trp Lys Thr Thr Ser Thr
465 470 475
<210> 4
<211> 469
<212> PRT
<213> tomato (Solanum lycopersicum L.)
<400> 4
Met Glu Phe Val Ser Met Leu Cys Leu Phe Thr Phe Ile Ser Leu Thr
1 5 10 15
Leu Leu Leu Ile His Ser Ile Phe Lys Phe Leu Ala Phe Ala Ser Lys
20 25 30
Lys Leu Pro Leu Pro Pro Gly Thr Leu Gly Leu Pro Tyr Ile Gly Glu
35 40 45
Thr Phe Gln Leu Tyr Ser Gln Asn Pro Asn Val Phe Phe Ala Ser Lys
50 55 60
Val Lys Lys Tyr Gly Ser Ile Phe Lys Thr Tyr Ile Leu Gly Cys Pro
65 70 75 80
Cys Val Met Ile Ser Ser Pro Glu Ala Ala Lys Gln Val Leu Val Thr
85 90 95
Lys Ala Asn Leu Phe Lys Pro Thr Phe Pro Ala Ser Lys Glu Arg Met
100 105 110
Leu Gly Lys Gln Ala Ile Phe Phe His Gln Gly Asp Tyr His Ala Lys
115 120 125
Leu Arg Lys Leu Val Leu Gln Ala Phe Lys Pro Asp Ser Ile Arg Asn
130 135 140
Ile Ile Pro Asp Ile Glu Ser Ile Ala Ile Thr Ser Leu Glu Ser Phe
145 150 155 160
Gln Gly Arg Leu Ile Asn Thr Tyr Gln Glu Met Lys Thr Tyr Thr Phe
165 170 175
Asn Val Ala Leu Ile Ser Ile Phe Gly Lys Asp Glu Phe Leu Tyr Arg
180 185 190
Glu Glu Leu Lys Lys Cys Tyr Tyr Ile Leu Glu Lys Gly Tyr Asn Ser
195 200 205
Met Pro Ile Asn Leu Pro Gly Thr Leu Phe Asn Lys Ala Met Lys Ala
210 215 220
Arg Lys Glu Leu Ala Lys Ile Val Ala Lys Ile Ile Ser Thr Arg Arg
225 230 235 240
Glu Met Lys Ile Asp His Gly Asp Leu Leu Gly Ser Phe Met Gly Asp
245 250 255
Lys Glu Gly Leu Thr Asp Glu Gln Ile Ala Asp Asn Val Ile Gly Val
260 265 270
Ile Phe Ala Ala Arg Asp Thr Thr Ala Ser Val Leu Thr Trp Ile Leu
275 280 285
Lys Tyr Leu Gly Glu Asn Pro Ser Val Leu Gln Ala Val Thr Glu Glu
290 295 300
Gln Glu Asn Ile Met Arg Lys Lys Glu Val Asn Gly Glu Glu Lys Val
305 310 315 320
Leu Asn Trp Gln Asp Thr Arg Gln Met Pro Met Thr Thr Arg Val Ile
325 330 335
Gln Glu Thr Leu Arg Val Ala Ser Ile Leu Ser Phe Thr Phe Arg Glu
340 345 350
Ala Val Glu Asp Val Glu Phe Glu Gly Tyr Leu Ile Pro Lys Gly Trp
355 360 365
Lys Val Leu Pro Leu Phe Arg Asn Ile His His Ser Pro Asp Asn Phe
370 375 380
Pro Glu Pro Glu Lys Phe Asp Pro Ser Arg Phe Glu Val Ser Pro Lys
385 390 395 400
Pro Asn Thr Phe Met Pro Phe Gly Asn Gly Val His Ser Cys Pro Gly
405 410 415
Asn Asp Leu Ala Lys Leu Glu Ile Leu Ile Leu Val His His Leu Thr
420 425 430
Thr Lys Tyr Arg Trp Ser Met Val Gly Pro Gln Asn Gly Ile Gln Tyr
435 440 445
Gly Pro Phe Ala Leu Pro Gln Asn Gly Leu Pro Ile Lys Leu Ser Leu
450 455 460
Lys Thr Ser Ser Thr
465
<210> 5
<211> 20
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400> 5
ttcctcctgg atcaatggga 20
<210> 6
<211> 20
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400> 6
aagaaattgc cacttcctcc 20
<210> 7
<211> 20
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400> 7
gtccagggaa tgaacttgcc 20
<210> 8
<211> 20
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400> 8
ttgcagctag tccacccaat 20
<210> 9
<211> 20
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400> 9
ctcgggtcat tcatgggaga 20
<210> 10
<211> 20
<212> DNA
<213> Artificial Sequence (Artificial Sequence)
<400> 10
agcagtagtg tctcgagctg 20