WO2022145178A1 - フェニルアラニンアンモニアリアーゼを用いた鎖状の不飽和カルボン酸化合物の製造方法 - Google Patents
フェニルアラニンアンモニアリアーゼを用いた鎖状の不飽和カルボン酸化合物の製造方法 Download PDFInfo
- Publication number
- WO2022145178A1 WO2022145178A1 PCT/JP2021/044890 JP2021044890W WO2022145178A1 WO 2022145178 A1 WO2022145178 A1 WO 2022145178A1 JP 2021044890 W JP2021044890 W JP 2021044890W WO 2022145178 A1 WO2022145178 A1 WO 2022145178A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- amino acid
- seq
- site
- acid sequence
- group
- Prior art date
Links
- 108700023158 Phenylalanine ammonia-lyases Proteins 0.000 title claims abstract description 118
- -1 carboxylic acid compound Chemical class 0.000 title claims abstract description 116
- 238000004519 manufacturing process Methods 0.000 title claims abstract description 66
- 238000000034 method Methods 0.000 claims abstract description 60
- 239000011203 carbon fibre reinforced carbon Substances 0.000 claims abstract description 34
- 125000003277 amino group Chemical group 0.000 claims abstract description 22
- 150000001413 amino acids Chemical class 0.000 claims description 195
- 235000001014 amino acid Nutrition 0.000 claims description 108
- 239000013598 vector Substances 0.000 claims description 96
- 229940024606 amino acid Drugs 0.000 claims description 87
- 102000004190 Enzymes Human genes 0.000 claims description 71
- 108090000790 Enzymes Proteins 0.000 claims description 71
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 claims description 60
- 108010056979 phenylacrylic acid decarboxylase Proteins 0.000 claims description 55
- 150000002430 hydrocarbons Chemical group 0.000 claims description 52
- 125000004432 carbon atom Chemical group C* 0.000 claims description 44
- 229930195735 unsaturated hydrocarbon Natural products 0.000 claims description 41
- 108090000623 proteins and genes Proteins 0.000 claims description 38
- COLNVLDHVKWLRT-QMMMGPOBSA-N L-phenylalanine Chemical compound OC(=O)[C@@H](N)CC1=CC=CC=C1 COLNVLDHVKWLRT-QMMMGPOBSA-N 0.000 claims description 24
- COLNVLDHVKWLRT-UHFFFAOYSA-N phenylalanine Natural products OC(=O)C(N)CC1=CC=CC=C1 COLNVLDHVKWLRT-UHFFFAOYSA-N 0.000 claims description 24
- 150000001875 compounds Chemical class 0.000 claims description 23
- 229960000310 isoleucine Drugs 0.000 claims description 22
- AGPKZVBTJJNPAG-UHFFFAOYSA-N isoleucine Natural products CCC(C)C(N)C(O)=O AGPKZVBTJJNPAG-UHFFFAOYSA-N 0.000 claims description 22
- 235000018102 proteins Nutrition 0.000 claims description 22
- 102000004169 proteins and genes Human genes 0.000 claims description 22
- 238000006467 substitution reaction Methods 0.000 claims description 21
- 125000000741 isoleucyl group Chemical group [H]N([H])C(C(C([H])([H])[H])C([H])([H])C([H])([H])[H])C(=O)O* 0.000 claims description 14
- QIVBCDIJIAJPQS-UHFFFAOYSA-N Tryptophan Natural products C1=CC=C2C(CC(N)C(O)=O)=CNC2=C1 QIVBCDIJIAJPQS-UHFFFAOYSA-N 0.000 claims description 13
- 229930182817 methionine Natural products 0.000 claims description 13
- KZSNJWFQEVHDMF-BYPYZUCNSA-N L-valine Chemical compound CC(C)[C@H](N)C(O)=O KZSNJWFQEVHDMF-BYPYZUCNSA-N 0.000 claims description 12
- 125000004435 hydrogen atom Chemical group [H]* 0.000 claims description 12
- KZSNJWFQEVHDMF-UHFFFAOYSA-N Valine Natural products CC(C)C(N)C(O)=O KZSNJWFQEVHDMF-UHFFFAOYSA-N 0.000 claims description 11
- 235000004279 alanine Nutrition 0.000 claims description 11
- 239000004474 valine Substances 0.000 claims description 11
- QIVBCDIJIAJPQS-VIFPVBQESA-N L-tryptophane Chemical compound C1=CC=C2C(C[C@H](N)C(O)=O)=CNC2=C1 QIVBCDIJIAJPQS-VIFPVBQESA-N 0.000 claims description 10
- 238000012258 culturing Methods 0.000 claims description 10
- 125000003545 alkoxy group Chemical group 0.000 claims description 9
- 125000000217 alkyl group Chemical group 0.000 claims description 9
- AGPKZVBTJJNPAG-WHFBIAKZSA-N L-isoleucine Chemical compound CC[C@H](C)[C@H](N)C(O)=O AGPKZVBTJJNPAG-WHFBIAKZSA-N 0.000 claims description 8
- 125000002887 hydroxy group Chemical group [H]O* 0.000 claims description 8
- 125000003295 alanine group Chemical group N[C@@H](C)C(=O)* 0.000 claims description 7
- 125000003178 carboxy group Chemical group [H]OC(*)=O 0.000 claims description 7
- 125000001360 methionine group Chemical group N[C@@H](CCSC)C(=O)* 0.000 claims description 7
- FFEARJCKVFRZRR-BYPYZUCNSA-N L-methionine Chemical compound CSCC[C@H](N)C(O)=O FFEARJCKVFRZRR-BYPYZUCNSA-N 0.000 claims description 6
- QNAYBMKLOCPYGJ-REOHCLBHSA-N L-alanine Chemical compound C[C@H](N)C(O)=O QNAYBMKLOCPYGJ-REOHCLBHSA-N 0.000 claims description 4
- 125000000430 tryptophan group Chemical group [H]N([H])C(C(=O)O*)C([H])([H])C1=C([H])N([H])C2=C([H])C([H])=C([H])C([H])=C12 0.000 claims description 4
- 150000001336 alkenes Chemical group 0.000 claims description 3
- QKHXKQJMUZVCJM-QRPNPIFTSA-N (2s)-2-amino-3-phenylpropanoic acid;azane Chemical class N.OC(=O)[C@@H](N)CC1=CC=CC=C1 QKHXKQJMUZVCJM-QRPNPIFTSA-N 0.000 claims description 2
- 125000001570 methylene group Chemical group [H]C([H])([*:1])[*:2] 0.000 claims description 2
- 150000001732 carboxylic acid derivatives Chemical class 0.000 claims 1
- 108020004414 DNA Proteins 0.000 description 114
- KAKZBPTYRLMSJV-UHFFFAOYSA-N Butadiene Chemical compound C=CC=C KAKZBPTYRLMSJV-UHFFFAOYSA-N 0.000 description 84
- 210000004027 cell Anatomy 0.000 description 76
- 238000006243 chemical reaction Methods 0.000 description 42
- 241000588724 Escherichia coli Species 0.000 description 24
- KDXKERNSBIXSRK-YFKPBYRVSA-N L-lysine Chemical compound NCCCC[C@H](N)C(O)=O KDXKERNSBIXSRK-YFKPBYRVSA-N 0.000 description 22
- 229960005190 phenylalanine Drugs 0.000 description 21
- 230000015572 biosynthetic process Effects 0.000 description 17
- 239000003795 chemical substances by application Substances 0.000 description 16
- 230000003197 catalytic effect Effects 0.000 description 15
- 239000000243 solution Substances 0.000 description 14
- 235000019766 L-Lysine Nutrition 0.000 description 11
- 239000004472 Lysine Substances 0.000 description 11
- 230000000694 effects Effects 0.000 description 11
- 239000000758 substrate Substances 0.000 description 11
- 230000035772 mutation Effects 0.000 description 10
- 125000003729 nucleotide group Chemical group 0.000 description 10
- 239000000126 substance Substances 0.000 description 10
- ROHFNLRQFUQHCH-UHFFFAOYSA-N Leucine Natural products CC(C)CC(N)C(O)=O ROHFNLRQFUQHCH-UHFFFAOYSA-N 0.000 description 9
- 108091028043 Nucleic acid sequence Proteins 0.000 description 9
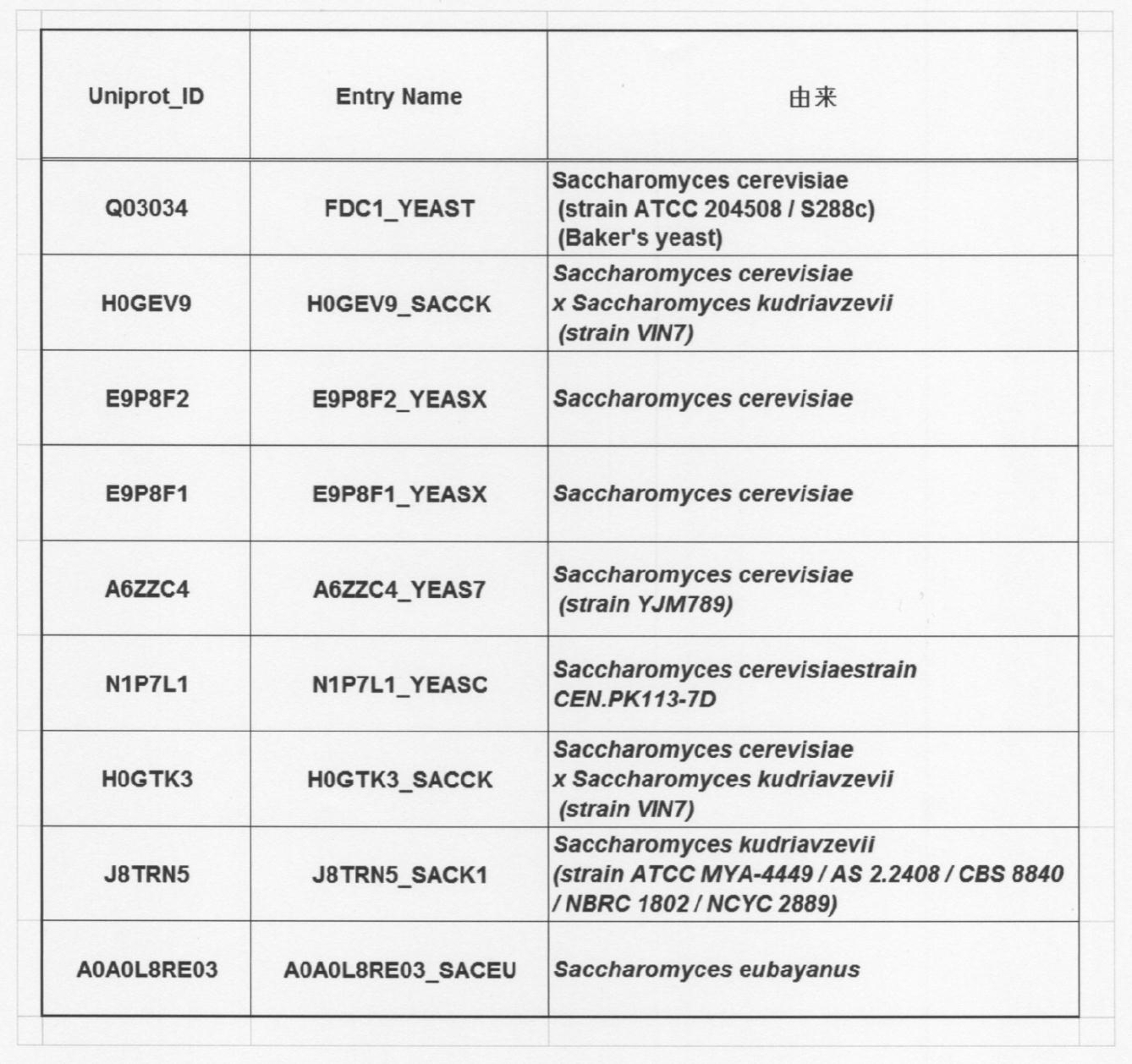
- 241000235070 Saccharomyces Species 0.000 description 9
- 125000001909 leucine group Chemical group [H]N(*)C(C(*)=O)C([H])([H])C(C([H])([H])[H])C([H])([H])[H] 0.000 description 9
- 230000001737 promoting effect Effects 0.000 description 9
- 238000000746 purification Methods 0.000 description 8
- 239000002994 raw material Substances 0.000 description 8
- 240000004808 Saccharomyces cerevisiae Species 0.000 description 7
- 235000014680 Saccharomyces cerevisiae Nutrition 0.000 description 7
- 239000002773 nucleotide Substances 0.000 description 7
- 239000013612 plasmid Substances 0.000 description 7
- 239000013600 plasmid vector Substances 0.000 description 7
- PEDCQBHIVMGVHV-UHFFFAOYSA-N Glycerine Chemical compound OCC(O)CO PEDCQBHIVMGVHV-UHFFFAOYSA-N 0.000 description 6
- WCUXLLCKKVVCTQ-UHFFFAOYSA-M Potassium chloride Chemical compound [Cl-].[K+] WCUXLLCKKVVCTQ-UHFFFAOYSA-M 0.000 description 6
- TXXHDPDFNKHHGW-UHFFFAOYSA-N muconic acid Chemical compound OC(=O)C=CC=CC(O)=O TXXHDPDFNKHHGW-UHFFFAOYSA-N 0.000 description 6
- 238000002360 preparation method Methods 0.000 description 6
- 230000035484 reaction time Effects 0.000 description 6
- WNNNWFKQCKFSDK-BYPYZUCNSA-N (2s)-2-aminopent-4-enoic acid Chemical compound OC(=O)[C@@H](N)CC=C WNNNWFKQCKFSDK-BYPYZUCNSA-N 0.000 description 5
- 241000238631 Hexapoda Species 0.000 description 5
- 241000078013 Trichormus variabilis Species 0.000 description 5
- AVKUERGKIZMTKX-NJBDSQKTSA-N ampicillin Chemical compound C1([C@@H](N)C(=O)N[C@H]2[C@H]3SC([C@@H](N3C2=O)C(O)=O)(C)C)=CC=CC=C1 AVKUERGKIZMTKX-NJBDSQKTSA-N 0.000 description 5
- 229960000723 ampicillin Drugs 0.000 description 5
- 239000000872 buffer Substances 0.000 description 5
- 238000004587 chromatography analysis Methods 0.000 description 5
- 244000005700 microbiome Species 0.000 description 5
- 239000000203 mixture Substances 0.000 description 5
- 239000000047 product Substances 0.000 description 5
- 238000011084 recovery Methods 0.000 description 5
- 239000007858 starting material Substances 0.000 description 5
- SDVVLIIVFBKBMG-ONEGZZNKSA-N (E)-penta-2,4-dienoic acid Chemical compound OC(=O)\C=C\C=C SDVVLIIVFBKBMG-ONEGZZNKSA-N 0.000 description 4
- 241000219195 Arabidopsis thaliana Species 0.000 description 4
- 108020004705 Codon Proteins 0.000 description 4
- SRBFZHDQGSBBOR-IOVATXLUSA-N D-xylopyranose Chemical compound O[C@@H]1COC(O)[C@H](O)[C@H]1O SRBFZHDQGSBBOR-IOVATXLUSA-N 0.000 description 4
- DHMQDGOQFOQNFH-UHFFFAOYSA-N Glycine Chemical compound NCC(O)=O DHMQDGOQFOQNFH-UHFFFAOYSA-N 0.000 description 4
- PPBRXRYQALVLMV-UHFFFAOYSA-N Styrene Chemical compound C=CC1=CC=CC=C1 PPBRXRYQALVLMV-UHFFFAOYSA-N 0.000 description 4
- 239000007853 buffer solution Substances 0.000 description 4
- 239000002299 complementary DNA Substances 0.000 description 4
- 238000004821 distillation Methods 0.000 description 4
- 238000006911 enzymatic reaction Methods 0.000 description 4
- 238000009396 hybridization Methods 0.000 description 4
- 238000005259 measurement Methods 0.000 description 4
- 230000000717 retained effect Effects 0.000 description 4
- 230000035939 shock Effects 0.000 description 4
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Chemical compound O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 4
- DCXYFEDJOCDNAF-UHFFFAOYSA-N Asparagine Natural products OC(=O)C(N)CC(N)=O DCXYFEDJOCDNAF-UHFFFAOYSA-N 0.000 description 3
- 241000894006 Bacteria Species 0.000 description 3
- 108090000489 Carboxy-Lyases Proteins 0.000 description 3
- 102000004031 Carboxy-Lyases Human genes 0.000 description 3
- GUBGYTABKSRVRQ-QKKXKWKRSA-N Lactose Natural products OC[C@H]1O[C@@H](O[C@H]2[C@H](O)[C@@H](O)C(O)O[C@@H]2CO)[C@H](O)[C@@H](O)[C@H]1O GUBGYTABKSRVRQ-QKKXKWKRSA-N 0.000 description 3
- 241001465754 Metazoa Species 0.000 description 3
- TXXHDPDFNKHHGW-CCAGOZQPSA-N Muconic acid Natural products OC(=O)\C=C/C=C\C(O)=O TXXHDPDFNKHHGW-CCAGOZQPSA-N 0.000 description 3
- 241000700605 Viruses Species 0.000 description 3
- WNNNWFKQCKFSDK-UHFFFAOYSA-N allylglycine Chemical compound OC(=O)C(N)CC=C WNNNWFKQCKFSDK-UHFFFAOYSA-N 0.000 description 3
- 229960001230 asparagine Drugs 0.000 description 3
- 235000009582 asparagine Nutrition 0.000 description 3
- 125000000613 asparagine group Chemical group N[C@@H](CC(N)=O)C(=O)* 0.000 description 3
- 229940041514 candida albicans extract Drugs 0.000 description 3
- 239000013592 cell lysate Substances 0.000 description 3
- 238000013461 design Methods 0.000 description 3
- 238000001514 detection method Methods 0.000 description 3
- 238000004520 electroporation Methods 0.000 description 3
- FVTCRASFADXXNN-SCRDCRAPSA-N flavin mononucleotide Chemical class OP(=O)(O)OC[C@@H](O)[C@@H](O)[C@@H](O)CN1C=2C=C(C)C(C)=CC=2N=C2C1=NC(=O)NC2=O FVTCRASFADXXNN-SCRDCRAPSA-N 0.000 description 3
- 238000002290 gas chromatography-mass spectrometry Methods 0.000 description 3
- BPHPUYQFMNQIOC-NXRLNHOXSA-N isopropyl beta-D-thiogalactopyranoside Chemical compound CC(C)S[C@@H]1O[C@H](CO)[C@H](O)[C@H](O)[C@H]1O BPHPUYQFMNQIOC-NXRLNHOXSA-N 0.000 description 3
- 229930027917 kanamycin Natural products 0.000 description 3
- 229960000318 kanamycin Drugs 0.000 description 3
- SBUJHOSQTJFQJX-NOAMYHISSA-N kanamycin Chemical compound O[C@@H]1[C@@H](O)[C@H](O)[C@@H](CN)O[C@@H]1O[C@H]1[C@H](O)[C@@H](O[C@@H]2[C@@H]([C@@H](N)[C@H](O)[C@@H](CO)O2)O)[C@H](N)C[C@@H]1N SBUJHOSQTJFQJX-NOAMYHISSA-N 0.000 description 3
- 229930182823 kanamycin A Natural products 0.000 description 3
- 239000008101 lactose Substances 0.000 description 3
- 150000002632 lipids Chemical class 0.000 description 3
- 125000002496 methyl group Chemical group [H]C([H])([H])* 0.000 description 3
- 239000011259 mixed solution Substances 0.000 description 3
- 230000001590 oxidative effect Effects 0.000 description 3
- 239000002504 physiological saline solution Substances 0.000 description 3
- 239000001103 potassium chloride Substances 0.000 description 3
- 235000011164 potassium chloride Nutrition 0.000 description 3
- 239000003755 preservative agent Substances 0.000 description 3
- 125000006308 propyl amino group Chemical group 0.000 description 3
- 229920006395 saturated elastomer Polymers 0.000 description 3
- 239000001488 sodium phosphate Substances 0.000 description 3
- 229910000162 sodium phosphate Inorganic materials 0.000 description 3
- 239000002904 solvent Substances 0.000 description 3
- 239000008223 sterile water Substances 0.000 description 3
- 239000004094 surface-active agent Substances 0.000 description 3
- 238000003786 synthesis reaction Methods 0.000 description 3
- RYFMWSXOAZQYPI-UHFFFAOYSA-K trisodium phosphate Chemical compound [Na+].[Na+].[Na+].[O-]P([O-])([O-])=O RYFMWSXOAZQYPI-UHFFFAOYSA-K 0.000 description 3
- 235000015112 vegetable and seed oil Nutrition 0.000 description 3
- 239000008158 vegetable oil Substances 0.000 description 3
- 239000012138 yeast extract Substances 0.000 description 3
- KSEBMYQBYZTDHS-HWKANZROSA-M (E)-Ferulic acid Natural products COC1=CC(\C=C\C([O-])=O)=CC=C1O KSEBMYQBYZTDHS-HWKANZROSA-M 0.000 description 2
- WBYWAXJHAXSJNI-VOTSOKGWSA-M .beta-Phenylacrylic acid Natural products [O-]C(=O)\C=C\C1=CC=CC=C1 WBYWAXJHAXSJNI-VOTSOKGWSA-M 0.000 description 2
- FAMLQSCGIQGDPV-UHFFFAOYSA-N 3,4-dihydroxy-6-[4-hydroxy-5-(hydroxymethyl)-2-(sulfooxymethyl)oxolan-3-yl]oxy-5-sulfooxyoxane-2-carboxylic acid Chemical compound OC1C(CO)OC(COS(O)(=O)=O)C1OC1C(OS(O)(=O)=O)C(O)C(O)C(C(O)=O)O1 FAMLQSCGIQGDPV-UHFFFAOYSA-N 0.000 description 2
- QGZKDVFQNNGYKY-UHFFFAOYSA-N Ammonia Chemical compound N QGZKDVFQNNGYKY-UHFFFAOYSA-N 0.000 description 2
- 108090000673 Ammonia-Lyases Proteins 0.000 description 2
- 102000004118 Ammonia-Lyases Human genes 0.000 description 2
- IJGRMHOSHXDMSA-UHFFFAOYSA-N Atomic nitrogen Chemical compound N#N IJGRMHOSHXDMSA-UHFFFAOYSA-N 0.000 description 2
- CURLTUGMZLYLDI-UHFFFAOYSA-N Carbon dioxide Chemical compound O=C=O CURLTUGMZLYLDI-UHFFFAOYSA-N 0.000 description 2
- WBYWAXJHAXSJNI-SREVYHEPSA-N Cinnamic acid Chemical compound OC(=O)\C=C/C1=CC=CC=C1 WBYWAXJHAXSJNI-SREVYHEPSA-N 0.000 description 2
- 102000016911 Deoxyribonucleases Human genes 0.000 description 2
- 108010053770 Deoxyribonucleases Proteins 0.000 description 2
- 206010059866 Drug resistance Diseases 0.000 description 2
- 241000196324 Embryophyta Species 0.000 description 2
- 108030006742 Flavin prenyltransferases Proteins 0.000 description 2
- 108010070675 Glutathione transferase Proteins 0.000 description 2
- 102000005720 Glutathione transferase Human genes 0.000 description 2
- 239000004471 Glycine Substances 0.000 description 2
- RRHGJUQNOFWUDK-UHFFFAOYSA-N Isoprene Chemical compound CC(=C)C=C RRHGJUQNOFWUDK-UHFFFAOYSA-N 0.000 description 2
- 241000519545 Plagiochasma Species 0.000 description 2
- 241000026988 Plagiochasma appendiculatum Species 0.000 description 2
- 241000589540 Pseudomonas fluorescens Species 0.000 description 2
- 241000198071 Saccharomyces cariocanus Species 0.000 description 2
- 241001123227 Saccharomyces pastorianus Species 0.000 description 2
- 108010052982 Tyrosine 2,3-aminomutase Proteins 0.000 description 2
- 238000004458 analytical method Methods 0.000 description 2
- 210000004102 animal cell Anatomy 0.000 description 2
- PYMYPHUHKUWMLA-UHFFFAOYSA-N arabinose Natural products OCC(O)C(O)C(O)C=O PYMYPHUHKUWMLA-UHFFFAOYSA-N 0.000 description 2
- SRBFZHDQGSBBOR-UHFFFAOYSA-N beta-D-Pyranose-Lyxose Natural products OC1COC(O)C(O)C1O SRBFZHDQGSBBOR-UHFFFAOYSA-N 0.000 description 2
- 235000013985 cinnamic acid Nutrition 0.000 description 2
- 229930016911 cinnamic acid Natural products 0.000 description 2
- 238000007796 conventional method Methods 0.000 description 2
- 230000009089 cytolysis Effects 0.000 description 2
- 230000001461 cytolytic effect Effects 0.000 description 2
- 230000000911 decarboxylating effect Effects 0.000 description 2
- 238000006114 decarboxylation reaction Methods 0.000 description 2
- 238000003795 desorption Methods 0.000 description 2
- ZPWVASYFFYYZEW-UHFFFAOYSA-L dipotassium hydrogen phosphate Chemical compound [K+].[K+].OP([O-])([O-])=O ZPWVASYFFYYZEW-UHFFFAOYSA-L 0.000 description 2
- 230000002255 enzymatic effect Effects 0.000 description 2
- 238000002474 experimental method Methods 0.000 description 2
- 239000013604 expression vector Substances 0.000 description 2
- KSEBMYQBYZTDHS-HWKANZROSA-N ferulic acid Chemical compound COC1=CC(\C=C\C(O)=O)=CC=C1O KSEBMYQBYZTDHS-HWKANZROSA-N 0.000 description 2
- 235000001785 ferulic acid Nutrition 0.000 description 2
- 229940114124 ferulic acid Drugs 0.000 description 2
- KSEBMYQBYZTDHS-UHFFFAOYSA-N ferulic acid Natural products COC1=CC(C=CC(O)=O)=CC=C1O KSEBMYQBYZTDHS-UHFFFAOYSA-N 0.000 description 2
- 229940013640 flavin mononucleotide Drugs 0.000 description 2
- FVTCRASFADXXNN-UHFFFAOYSA-N flavin mononucleotide Natural products OP(=O)(O)OCC(O)C(O)C(O)CN1C=2C=C(C)C(C)=CC=2N=C2C1=NC(=O)NC2=O FVTCRASFADXXNN-UHFFFAOYSA-N 0.000 description 2
- 239000011768 flavin mononucleotide Substances 0.000 description 2
- 239000007789 gas Substances 0.000 description 2
- 238000004817 gas chromatography Methods 0.000 description 2
- 239000003112 inhibitor Substances 0.000 description 2
- 239000003550 marker Substances 0.000 description 2
- WBYWAXJHAXSJNI-UHFFFAOYSA-N methyl p-hydroxycinnamate Natural products OC(=O)C=CC1=CC=CC=C1 WBYWAXJHAXSJNI-UHFFFAOYSA-N 0.000 description 2
- 229910000402 monopotassium phosphate Inorganic materials 0.000 description 2
- 235000019796 monopotassium phosphate Nutrition 0.000 description 2
- 125000003935 n-pentoxy group Chemical group [H]C([H])([H])C([H])([H])C([H])([H])C([H])([H])C([H])([H])O* 0.000 description 2
- 150000007523 nucleic acids Chemical class 0.000 description 2
- 239000000137 peptide hydrolase inhibitor Substances 0.000 description 2
- 239000003208 petroleum Substances 0.000 description 2
- PJNZPQUBCPKICU-UHFFFAOYSA-N phosphoric acid;potassium Chemical compound [K].OP(O)(O)=O PJNZPQUBCPKICU-UHFFFAOYSA-N 0.000 description 2
- 229920000642 polymer Polymers 0.000 description 2
- 238000001556 precipitation Methods 0.000 description 2
- 108090000765 processed proteins & peptides Proteins 0.000 description 2
- 230000035755 proliferation Effects 0.000 description 2
- 238000001243 protein synthesis Methods 0.000 description 2
- 230000010076 replication Effects 0.000 description 2
- 108091008146 restriction endonucleases Proteins 0.000 description 2
- 235000019231 riboflavin-5'-phosphate Nutrition 0.000 description 2
- 229930195734 saturated hydrocarbon Natural products 0.000 description 2
- 239000007790 solid phase Substances 0.000 description 2
- 241000894007 species Species 0.000 description 2
- 125000001424 substituent group Chemical group 0.000 description 2
- 229920003051 synthetic elastomer Polymers 0.000 description 2
- 239000005061 synthetic rubber Substances 0.000 description 2
- QURCVMIEKCOAJU-UHFFFAOYSA-N trans-isoferulic acid Natural products COC1=CC=C(C=CC(O)=O)C=C1O QURCVMIEKCOAJU-UHFFFAOYSA-N 0.000 description 2
- 230000014616 translation Effects 0.000 description 2
- 241000701447 unidentified baculovirus Species 0.000 description 2
- FSPHIBLXJRNTSN-GDVGLLTNSA-N (2s)-2,6-diamino-3-methylhexanoic acid Chemical compound OC(=O)[C@@H](N)C(C)CCCN FSPHIBLXJRNTSN-GDVGLLTNSA-N 0.000 description 1
- NNIFTGRSDICEMZ-GDVGLLTNSA-N (2s)-2,6-diamino-4-methylhexanoic acid Chemical compound NCCC(C)C[C@H](N)C(O)=O NNIFTGRSDICEMZ-GDVGLLTNSA-N 0.000 description 1
- 125000005923 1,2-dimethylpropyloxy group Chemical group 0.000 description 1
- WIIZWVCIJKGZOK-IUCAKERBSA-N 2,2-dichloro-n-[(1s,2s)-1,3-dihydroxy-1-(4-nitrophenyl)propan-2-yl]acetamide Chemical compound ClC(Cl)C(=O)N[C@@H](CO)[C@@H](O)C1=CC=C([N+]([O-])=O)C=C1 WIIZWVCIJKGZOK-IUCAKERBSA-N 0.000 description 1
- WCASXYBKJHWFMY-NSCUHMNNSA-N 2-Buten-1-ol Chemical compound C\C=C\CO WCASXYBKJHWFMY-NSCUHMNNSA-N 0.000 description 1
- YMQIIPLNNNGNEB-UHFFFAOYSA-N 3-methylpenta-2,4-dienoic acid Chemical compound C=CC(C)=CC(O)=O YMQIIPLNNNGNEB-UHFFFAOYSA-N 0.000 description 1
- VVHRGZKMNAOETC-UHFFFAOYSA-N 4-methylpenta-2,4-dienoic acid Chemical compound CC(=C)C=CC(O)=O VVHRGZKMNAOETC-UHFFFAOYSA-N 0.000 description 1
- ZCYVEMRRCGMTRW-UHFFFAOYSA-N 7553-56-2 Chemical compound [I] ZCYVEMRRCGMTRW-UHFFFAOYSA-N 0.000 description 1
- 241000589158 Agrobacterium Species 0.000 description 1
- 239000004475 Arginine Substances 0.000 description 1
- 241000228212 Aspergillus Species 0.000 description 1
- 241000193830 Bacillus <bacterium> Species 0.000 description 1
- 244000063299 Bacillus subtilis Species 0.000 description 1
- 235000014469 Bacillus subtilis Nutrition 0.000 description 1
- 244000289276 Bambusa oldhamii Species 0.000 description 1
- 235000004270 Bambusa oldhamii Nutrition 0.000 description 1
- 239000002028 Biomass Substances 0.000 description 1
- WKBOTKDWSSQWDR-UHFFFAOYSA-N Bromine atom Chemical compound [Br] WKBOTKDWSSQWDR-UHFFFAOYSA-N 0.000 description 1
- OKTJSMMVPCPJKN-UHFFFAOYSA-N Carbon Chemical compound [C] OKTJSMMVPCPJKN-UHFFFAOYSA-N 0.000 description 1
- ZAMOUSCENKQFHK-UHFFFAOYSA-N Chlorine atom Chemical compound [Cl] ZAMOUSCENKQFHK-UHFFFAOYSA-N 0.000 description 1
- 241000701959 Escherichia virus Lambda Species 0.000 description 1
- VGGSQFUCUMXWEO-UHFFFAOYSA-N Ethene Chemical compound C=C VGGSQFUCUMXWEO-UHFFFAOYSA-N 0.000 description 1
- 239000005977 Ethylene Substances 0.000 description 1
- PXGOKWXKJXAPGV-UHFFFAOYSA-N Fluorine Chemical compound FF PXGOKWXKJXAPGV-UHFFFAOYSA-N 0.000 description 1
- 208000033962 Fontaine progeroid syndrome Diseases 0.000 description 1
- 241000233866 Fungi Species 0.000 description 1
- 241001159781 Kazachstania spencerorum Species 0.000 description 1
- 241000039979 Kazachstania turicensis Species 0.000 description 1
- 241001123232 Kazachstania unispora Species 0.000 description 1
- 238000012218 Kunkel's method Methods 0.000 description 1
- ODKSFYDXXFIFQN-BYPYZUCNSA-P L-argininium(2+) Chemical compound NC(=[NH2+])NCCC[C@H]([NH3+])C(O)=O ODKSFYDXXFIFQN-BYPYZUCNSA-P 0.000 description 1
- RBWXRFBKVDBXEG-DMTCNVIQSA-N L-beta-ethynylserine zwitterion Chemical compound OC(=O)[C@@H](N)[C@H](O)C#C RBWXRFBKVDBXEG-DMTCNVIQSA-N 0.000 description 1
- HNDVDQJCIGZPNO-YFKPBYRVSA-N L-histidine Chemical compound OC(=O)[C@@H](N)CC1=CN=CN1 HNDVDQJCIGZPNO-YFKPBYRVSA-N 0.000 description 1
- 241000235087 Lachancea kluyveri Species 0.000 description 1
- 108060001084 Luciferase Proteins 0.000 description 1
- 239000005089 Luciferase Substances 0.000 description 1
- 108010047357 Luminescent Proteins Proteins 0.000 description 1
- 102000006830 Luminescent Proteins Human genes 0.000 description 1
- 241001123224 Naumovozyma dairenensis Species 0.000 description 1
- 244000061176 Nicotiana tabacum Species 0.000 description 1
- 235000002637 Nicotiana tabacum Nutrition 0.000 description 1
- 229920000459 Nitrile rubber Polymers 0.000 description 1
- 229920002302 Nylon 6,6 Polymers 0.000 description 1
- 241000283973 Oryctolagus cuniculus Species 0.000 description 1
- 240000007594 Oryza sativa Species 0.000 description 1
- 235000007164 Oryza sativa Nutrition 0.000 description 1
- 239000001888 Peptone Substances 0.000 description 1
- 108010080698 Peptones Proteins 0.000 description 1
- 239000005062 Polybutadiene Substances 0.000 description 1
- 229940124158 Protease/peptidase inhibitor Drugs 0.000 description 1
- 241000589516 Pseudomonas Species 0.000 description 1
- 241000223253 Rhodotorula glutinis Species 0.000 description 1
- 241000589953 Rubinisphaera brasiliensis Species 0.000 description 1
- 241000235072 Saccharomyces bayanus Species 0.000 description 1
- 241000877399 Saccharomyces chevalieri Species 0.000 description 1
- 241000877401 Saccharomyces ellipsoideus Species 0.000 description 1
- 241001063879 Saccharomyces eubayanus Species 0.000 description 1
- 241000198063 Saccharomyces kudriavzevii Species 0.000 description 1
- 241001407717 Saccharomyces norbensis Species 0.000 description 1
- 241001123228 Saccharomyces paradoxus Species 0.000 description 1
- 241000582914 Saccharomyces uvarum Species 0.000 description 1
- 241000304195 Salvia miltiorrhiza Species 0.000 description 1
- 235000011135 Salvia miltiorrhiza Nutrition 0.000 description 1
- 235000002634 Solanum Nutrition 0.000 description 1
- 241000207763 Solanum Species 0.000 description 1
- 241001147855 Streptomyces cattleya Species 0.000 description 1
- 241000201081 Streptomyces maritimus Species 0.000 description 1
- QAOWNCQODCNURD-UHFFFAOYSA-L Sulfate Chemical compound [O-]S([O-])(=O)=O QAOWNCQODCNURD-UHFFFAOYSA-L 0.000 description 1
- NINIDFKCEFEMDL-UHFFFAOYSA-N Sulfur Chemical compound [S] NINIDFKCEFEMDL-UHFFFAOYSA-N 0.000 description 1
- 206010042602 Supraventricular extrasystoles Diseases 0.000 description 1
- 241001149649 Taxus wallichiana var. chinensis Species 0.000 description 1
- 241000209140 Triticum Species 0.000 description 1
- 235000021307 Triticum Nutrition 0.000 description 1
- 229920000122 acrylonitrile butadiene styrene Polymers 0.000 description 1
- 238000001042 affinity chromatography Methods 0.000 description 1
- 125000002521 alkyl halide group Chemical group 0.000 description 1
- 150000001350 alkyl halides Chemical class 0.000 description 1
- 229910021529 ammonia Inorganic materials 0.000 description 1
- 239000003242 anti bacterial agent Substances 0.000 description 1
- ODKSFYDXXFIFQN-UHFFFAOYSA-N arginine Natural products OC(=O)C(N)CCCNC(N)=N ODKSFYDXXFIFQN-UHFFFAOYSA-N 0.000 description 1
- QVGXLLKOCUKJST-UHFFFAOYSA-N atomic oxygen Chemical compound [O] QVGXLLKOCUKJST-UHFFFAOYSA-N 0.000 description 1
- 230000001580 bacterial effect Effects 0.000 description 1
- 230000003115 biocidal effect Effects 0.000 description 1
- GDTBXPJZTBHREO-UHFFFAOYSA-N bromine Substances BrBr GDTBXPJZTBHREO-UHFFFAOYSA-N 0.000 description 1
- 229910052794 bromium Inorganic materials 0.000 description 1
- OWBTYPJTUOEWEK-UHFFFAOYSA-N butane-2,3-diol Chemical compound CC(O)C(C)O OWBTYPJTUOEWEK-UHFFFAOYSA-N 0.000 description 1
- 239000006227 byproduct Substances 0.000 description 1
- 239000001506 calcium phosphate Substances 0.000 description 1
- 229910000389 calcium phosphate Inorganic materials 0.000 description 1
- 235000011010 calcium phosphates Nutrition 0.000 description 1
- 229910052799 carbon Inorganic materials 0.000 description 1
- CREMABGTGYGIQB-UHFFFAOYSA-N carbon carbon Chemical compound C.C CREMABGTGYGIQB-UHFFFAOYSA-N 0.000 description 1
- 239000001569 carbon dioxide Substances 0.000 description 1
- 229910002092 carbon dioxide Inorganic materials 0.000 description 1
- 239000005018 casein Substances 0.000 description 1
- BECPQYXYKAMYBN-UHFFFAOYSA-N casein, tech. Chemical compound NCCCCC(C(O)=O)N=C(O)C(CC(O)=O)N=C(O)C(CCC(O)=N)N=C(O)C(CC(C)C)N=C(O)C(CCC(O)=O)N=C(O)C(CC(O)=O)N=C(O)C(CCC(O)=O)N=C(O)C(C(C)O)N=C(O)C(CCC(O)=N)N=C(O)C(CCC(O)=N)N=C(O)C(CCC(O)=N)N=C(O)C(CCC(O)=O)N=C(O)C(CCC(O)=O)N=C(O)C(COP(O)(O)=O)N=C(O)C(CCC(O)=N)N=C(O)C(N)CC1=CC=CC=C1 BECPQYXYKAMYBN-UHFFFAOYSA-N 0.000 description 1
- 235000021240 caseins Nutrition 0.000 description 1
- 238000004113 cell culture Methods 0.000 description 1
- 230000006037 cell lysis Effects 0.000 description 1
- 238000005119 centrifugation Methods 0.000 description 1
- 239000003153 chemical reaction reagent Substances 0.000 description 1
- 229960005091 chloramphenicol Drugs 0.000 description 1
- WIIZWVCIJKGZOK-RKDXNWHRSA-N chloramphenicol Chemical compound ClC(Cl)C(=O)N[C@H](CO)[C@H](O)C1=CC=C([N+]([O-])=O)C=C1 WIIZWVCIJKGZOK-RKDXNWHRSA-N 0.000 description 1
- 239000000460 chlorine Substances 0.000 description 1
- 229910052801 chlorine Inorganic materials 0.000 description 1
- 230000002759 chromosomal effect Effects 0.000 description 1
- 210000000349 chromosome Anatomy 0.000 description 1
- 239000012141 concentrate Substances 0.000 description 1
- 125000004093 cyano group Chemical group *C#N 0.000 description 1
- 230000009615 deamination Effects 0.000 description 1
- 238000006481 deamination reaction Methods 0.000 description 1
- 230000007547 defect Effects 0.000 description 1
- 238000010790 dilution Methods 0.000 description 1
- 239000012895 dilution Substances 0.000 description 1
- 229940042399 direct acting antivirals protease inhibitors Drugs 0.000 description 1
- 238000004090 dissolution Methods 0.000 description 1
- 239000003623 enhancer Substances 0.000 description 1
- 230000007613 environmental effect Effects 0.000 description 1
- 125000001301 ethoxy group Chemical group [H]C([H])([H])C([H])([H])O* 0.000 description 1
- 125000001495 ethyl group Chemical group [H]C([H])([H])C([H])([H])* 0.000 description 1
- 239000000284 extract Substances 0.000 description 1
- 238000000605 extraction Methods 0.000 description 1
- 238000000855 fermentation Methods 0.000 description 1
- 230000004151 fermentation Effects 0.000 description 1
- 238000001914 filtration Methods 0.000 description 1
- 239000012467 final product Substances 0.000 description 1
- 239000007850 fluorescent dye Substances 0.000 description 1
- 108091006047 fluorescent proteins Proteins 0.000 description 1
- 102000034287 fluorescent proteins Human genes 0.000 description 1
- 229910052731 fluorine Inorganic materials 0.000 description 1
- 239000011737 fluorine Substances 0.000 description 1
- 125000002485 formyl group Chemical group [H]C(*)=O 0.000 description 1
- 239000002803 fossil fuel Substances 0.000 description 1
- WCASXYBKJHWFMY-UHFFFAOYSA-N gamma-methylallyl alcohol Natural products CC=CCO WCASXYBKJHWFMY-UHFFFAOYSA-N 0.000 description 1
- 238000005227 gel permeation chromatography Methods 0.000 description 1
- 230000002068 genetic effect Effects 0.000 description 1
- 238000010353 genetic engineering Methods 0.000 description 1
- 239000005431 greenhouse gas Substances 0.000 description 1
- 238000000227 grinding Methods 0.000 description 1
- 125000005843 halogen group Chemical group 0.000 description 1
- HNDVDQJCIGZPNO-UHFFFAOYSA-N histidine Natural products OC(=O)C(N)CC1=CN=CN1 HNDVDQJCIGZPNO-UHFFFAOYSA-N 0.000 description 1
- 238000002744 homologous recombination Methods 0.000 description 1
- 230000006801 homologous recombination Effects 0.000 description 1
- 229910052739 hydrogen Inorganic materials 0.000 description 1
- 239000001257 hydrogen Substances 0.000 description 1
- 230000001939 inductive effect Effects 0.000 description 1
- 239000011630 iodine Substances 0.000 description 1
- 229910052740 iodine Inorganic materials 0.000 description 1
- 238000004255 ion exchange chromatography Methods 0.000 description 1
- 229910000358 iron sulfate Inorganic materials 0.000 description 1
- BAUYGSIQEAFULO-UHFFFAOYSA-L iron(2+) sulfate (anhydrous) Chemical compound [Fe+2].[O-]S([O-])(=O)=O BAUYGSIQEAFULO-UHFFFAOYSA-L 0.000 description 1
- 229910000462 iron(III) oxide hydroxide Inorganic materials 0.000 description 1
- 125000000959 isobutyl group Chemical group [H]C([H])([H])C([H])(C([H])([H])[H])C([H])([H])* 0.000 description 1
- 125000001449 isopropyl group Chemical group [H]C([H])([H])C([H])(*)C([H])([H])[H] 0.000 description 1
- 150000004658 ketimines Chemical class 0.000 description 1
- 238000001638 lipofection Methods 0.000 description 1
- 239000007788 liquid Substances 0.000 description 1
- XIXADJRWDQXREU-UHFFFAOYSA-M lithium acetate Chemical compound [Li+].CC([O-])=O XIXADJRWDQXREU-UHFFFAOYSA-M 0.000 description 1
- 125000005439 maleimidyl group Chemical group C1(C=CC(N1*)=O)=O 0.000 description 1
- 238000004949 mass spectrometry Methods 0.000 description 1
- 235000013372 meat Nutrition 0.000 description 1
- 108020004999 messenger RNA Proteins 0.000 description 1
- 239000002207 metabolite Substances 0.000 description 1
- 229910052751 metal Inorganic materials 0.000 description 1
- 239000002184 metal Substances 0.000 description 1
- 150000002739 metals Chemical class 0.000 description 1
- 125000005394 methallyl group Chemical group 0.000 description 1
- 125000000956 methoxy group Chemical group [H]C([H])([H])O* 0.000 description 1
- 208000024191 minimally invasive lung adenocarcinoma Diseases 0.000 description 1
- 230000004048 modification Effects 0.000 description 1
- 238000012986 modification Methods 0.000 description 1
- 238000002703 mutagenesis Methods 0.000 description 1
- 231100000350 mutagenesis Toxicity 0.000 description 1
- 125000006606 n-butoxy group Chemical group 0.000 description 1
- 125000004108 n-butyl group Chemical group [H]C([H])([H])C([H])([H])C([H])([H])C([H])([H])* 0.000 description 1
- 125000000740 n-pentyl group Chemical group [H]C([H])([H])C([H])([H])C([H])([H])C([H])([H])C([H])([H])* 0.000 description 1
- 125000003506 n-propoxy group Chemical group [H]C([H])([H])C([H])([H])C([H])([H])O* 0.000 description 1
- 125000004123 n-propyl group Chemical group [H]C([H])([H])C([H])([H])C([H])([H])* 0.000 description 1
- 125000000449 nitro group Chemical group [O-][N+](*)=O 0.000 description 1
- 229910052757 nitrogen Inorganic materials 0.000 description 1
- 108020004707 nucleic acids Proteins 0.000 description 1
- 102000039446 nucleic acids Human genes 0.000 description 1
- 235000015097 nutrients Nutrition 0.000 description 1
- JRZJOMJEPLMPRA-UHFFFAOYSA-N olefin Natural products CCCCCCCC=C JRZJOMJEPLMPRA-UHFFFAOYSA-N 0.000 description 1
- 150000002894 organic compounds Chemical class 0.000 description 1
- 229910052760 oxygen Inorganic materials 0.000 description 1
- 239000001301 oxygen Substances 0.000 description 1
- 239000002245 particle Substances 0.000 description 1
- 239000008188 pellet Substances 0.000 description 1
- SDVVLIIVFBKBMG-UHFFFAOYSA-N penta-2,4-dienoic acid Chemical compound OC(=O)C=CC=C SDVVLIIVFBKBMG-UHFFFAOYSA-N 0.000 description 1
- 235000019319 peptone Nutrition 0.000 description 1
- 238000005373 pervaporation Methods 0.000 description 1
- 150000002994 phenylalanines Chemical class 0.000 description 1
- 239000008363 phosphate buffer Substances 0.000 description 1
- 229910052698 phosphorus Inorganic materials 0.000 description 1
- 239000011574 phosphorus Substances 0.000 description 1
- 229920002857 polybutadiene Polymers 0.000 description 1
- 229920002704 polyhistidine Polymers 0.000 description 1
- 239000002952 polymeric resin Substances 0.000 description 1
- 230000013823 prenylation Effects 0.000 description 1
- 230000002335 preservative effect Effects 0.000 description 1
- 125000002924 primary amino group Chemical group [H]N([H])* 0.000 description 1
- QQONPFPTGQHPMA-UHFFFAOYSA-N propylene Natural products CC=C QQONPFPTGQHPMA-UHFFFAOYSA-N 0.000 description 1
- 125000004805 propylene group Chemical group [H]C([H])([H])C([H])([*:1])C([H])([H])[*:2] 0.000 description 1
- 238000000751 protein extraction Methods 0.000 description 1
- 239000012429 reaction media Substances 0.000 description 1
- 238000011160 research Methods 0.000 description 1
- 210000001995 reticulocyte Anatomy 0.000 description 1
- PYWVYCXTNDRMGF-UHFFFAOYSA-N rhodamine B Chemical compound [Cl-].C=12C=CC(=[N+](CC)CC)C=C2OC2=CC(N(CC)CC)=CC=C2C=1C1=CC=CC=C1C(O)=O PYWVYCXTNDRMGF-UHFFFAOYSA-N 0.000 description 1
- 229960002181 saccharomyces boulardii Drugs 0.000 description 1
- 238000009938 salting Methods 0.000 description 1
- 150000003839 salts Chemical class 0.000 description 1
- 239000000523 sample Substances 0.000 description 1
- 238000005070 sampling Methods 0.000 description 1
- 230000003248 secreting effect Effects 0.000 description 1
- 210000002966 serum Anatomy 0.000 description 1
- 238000002741 site-directed mutagenesis Methods 0.000 description 1
- 238000001179 sorption measurement Methods 0.000 description 1
- 229920003048 styrene butadiene rubber Polymers 0.000 description 1
- WPLOVIFNBMNBPD-ATHMIXSHSA-N subtilin Chemical compound CC1SCC(NC2=O)C(=O)NC(CC(N)=O)C(=O)NC(C(=O)NC(CCCCN)C(=O)NC(C(C)CC)C(=O)NC(=C)C(=O)NC(CCCCN)C(O)=O)CSC(C)C2NC(=O)C(CC(C)C)NC(=O)C1NC(=O)C(CCC(N)=O)NC(=O)C(CC(C)C)NC(=O)C(NC(=O)C1NC(=O)C(=C/C)/NC(=O)C(CCC(N)=O)NC(=O)C(CC(C)C)NC(=O)C(C)NC(=O)CNC(=O)C(NC(=O)C(NC(=O)C2NC(=O)CNC(=O)C3CCCN3C(=O)C(NC(=O)C3NC(=O)C(CC(C)C)NC(=O)C(=C)NC(=O)C(CCC(O)=O)NC(=O)C(NC(=O)C(CCCCN)NC(=O)C(N)CC=4C5=CC=CC=C5NC=4)CSC3)C(C)SC2)C(C)C)C(C)SC1)CC1=CC=CC=C1 WPLOVIFNBMNBPD-ATHMIXSHSA-N 0.000 description 1
- 229910052717 sulfur Inorganic materials 0.000 description 1
- 239000011593 sulfur Substances 0.000 description 1
- 229910021653 sulphate ion Inorganic materials 0.000 description 1
- 239000006228 supernatant Substances 0.000 description 1
- 238000001308 synthesis method Methods 0.000 description 1
- 230000002194 synthesizing effect Effects 0.000 description 1
- 229920001059 synthetic polymer Polymers 0.000 description 1
- 229920003002 synthetic resin Polymers 0.000 description 1
- 125000000999 tert-butyl group Chemical group [H]C([H])([H])C(*)(C([H])([H])[H])C([H])([H])[H] 0.000 description 1
- 125000003396 thiol group Chemical group [H]S* 0.000 description 1
- 150000003573 thiols Chemical class 0.000 description 1
- 231100000331 toxic Toxicity 0.000 description 1
- 230000002588 toxic effect Effects 0.000 description 1
- 230000002103 transcriptional effect Effects 0.000 description 1
- QORWJWZARLRLPR-UHFFFAOYSA-H tricalcium bis(phosphate) Chemical compound [Ca+2].[Ca+2].[Ca+2].[O-]P([O-])([O-])=O.[O-]P([O-])([O-])=O QORWJWZARLRLPR-UHFFFAOYSA-H 0.000 description 1
- 229910021642 ultra pure water Inorganic materials 0.000 description 1
- 238000000108 ultra-filtration Methods 0.000 description 1
- 239000012498 ultrapure water Substances 0.000 description 1
- 241001446247 uncultured actinomycete Species 0.000 description 1
- 241000701161 unidentified adenovirus Species 0.000 description 1
- 241001430294 unidentified retrovirus Species 0.000 description 1
- 238000010792 warming Methods 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/88—Lyases (4.)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/70—Vectors or expression systems specially adapted for E. coli
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/0004—Oxidoreductases (1.)
- C12N9/0071—Oxidoreductases (1.) acting on paired donors with incorporation of molecular oxygen (1.14)
- C12N9/0083—Miscellaneous (1.14.99)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/04—Alpha- or beta- amino acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P5/00—Preparation of hydrocarbons or halogenated hydrocarbons
- C12P5/007—Preparation of hydrocarbons or halogenated hydrocarbons containing one or more isoprene units, i.e. terpenes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P5/00—Preparation of hydrocarbons or halogenated hydrocarbons
- C12P5/02—Preparation of hydrocarbons or halogenated hydrocarbons acyclic
- C12P5/026—Unsaturated compounds, i.e. alkenes, alkynes or allenes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/40—Preparation of oxygen-containing organic compounds containing a carboxyl group including Peroxycarboxylic acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y401/00—Carbon-carbon lyases (4.1)
- C12Y401/01—Carboxy-lyases (4.1.1)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y403/00—Carbon-nitrogen lyases (4.3)
- C12Y403/01—Ammonia-lyases (4.3.1)
- C12Y403/01024—Phenylalanine ammonia-lyase (4.3.1.24)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12R—INDEXING SCHEME ASSOCIATED WITH SUBCLASSES C12C - C12Q, RELATING TO MICROORGANISMS
- C12R2001/00—Microorganisms ; Processes using microorganisms
- C12R2001/01—Bacteria or Actinomycetales ; using bacteria or Actinomycetales
- C12R2001/185—Escherichia
- C12R2001/19—Escherichia coli
Definitions
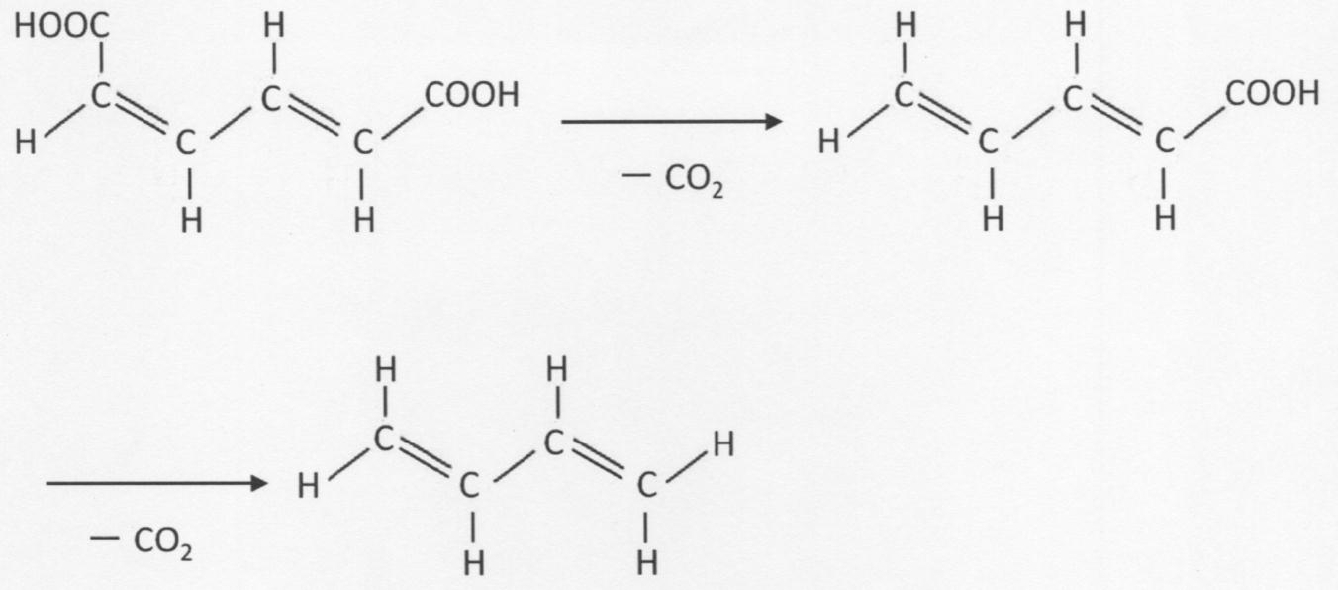
- a carbon-carbon double bond is further introduced into a second chain unsaturated carboxylic acid compound having a carbon-carbon double bond at the terminal, and a third carbon-carbon double bond is introduced.
- the present invention relates to a method for producing a chain unsaturated carboxylic acid compound. Further, the present invention comprises producing a second chain unsaturated carboxylic acid compound from a first chain unsaturated carboxylic acid compound having an amino group at the terminal in the presence of the terminal alkene-producing enzyme BesC.
- the present invention relates to a method for producing a third chain unsaturated carboxylic acid compound in the presence of phenylalanine ammonia lyase. Further, the present invention thus produces a third chain unsaturated carboxylic acid compound, which is then carbon-carbon at both ends from the unsaturated carboxylic acid compound in the presence of a ferulic acid decarboxylase.
- the present invention relates to a method for producing a chain unsaturated hydrocarbon compound having a double bond.
- the present invention also relates to a phenylalanine ammonia-lyase variant, a DNA encoding the variant, a vector into which the DNA is inserted, and a host cell into which the DNA or the vector has been introduced, which can be used in these production methods. Furthermore, the present invention relates to a method for producing the variant using the host cell.
- Butadiene (1,3-butadiene) is used as a raw material for various polymer compounds such as various synthetic rubbers (butadiene rubber, styrene-butadiene rubber, acrylonitrile-butadiene rubber, etc.) and polymer resins (ABS resin, nylon 66, etc.). Therefore, it can be said that it is an extremely important organic compound in the chemical industry. Further, these polymer compounds made from butadiene as a raw material are widely used not only for industrial products such as automobile tires but also for daily goods such as clothing. As a result, the demand for butadiene is increasing year by year, with annual demand reaching 13 million tons and a market size of $ 15 billion.
- Butadiene has been produced mainly by purifying the C4 fraction produced as a by-product in the production of ethylene and propylene from petroleum.
- environmental problems such as the depletion of fossil fuels such as petroleum and global warming due to greenhouse gas emissions
- a method for producing butadiene from a substance derived from a biomass resource, which is a renewable resource, using an enzyme is being actively developed.
- Patent Document 1 discloses a method for producing butadiene using xylose as a raw material and a microorganism having an enzymatic activity capable of converting it into crotyl alcohol or the like.
- Patent Document 2 discloses a method for producing butadiene using xylose as a raw material and a microorganism having an enzymatic activity capable of converting it into 2,3-butanediol.
- many attempts have been made to produce unsaturated hydrocarbon compounds such as butadiene using enzymes.
- the present invention has been made in view of the above-mentioned problems of the prior art, and an object of the present invention is to provide a method for producing an unsaturated compound having at least two carbon-carbon double bonds using an enzyme. do.
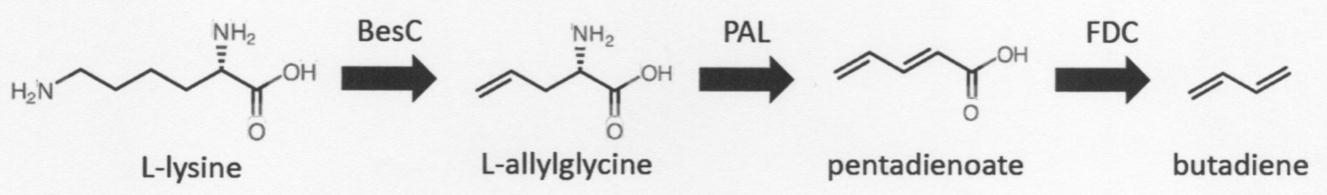
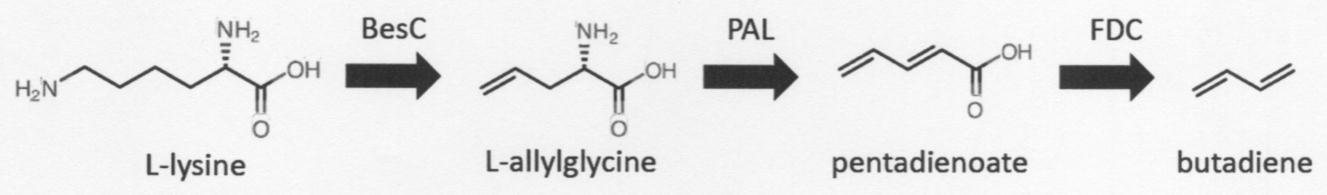
- the present inventors have conceived the following reaction scheme leading to the production of butadiene, using L-lysine as a starting material instead of muconic acid. Since L-lysine can be obtained at a relatively low cost, it is possible to reduce the cost in producing butadiene.
- BesC an enzyme disclosed in Non-Patent Document 2
- the enzyme has the activity of catalyzing a reaction that cleaves the carbon-carbon bond in the group by oxidizing the terminal propylamino group and forms a carbon-carbon double bond at the terminal.
- butadiene butadiene
- pentadienoate it was conceived that the FDC clarified by the present inventors can be used as described above (Patent Document 3).
- PAL phenylalanine ammonia-lyase
- a carbon-carbon double bond is further introduced into a second chain unsaturated carboxylic acid compound having a carbon-carbon double bond at the terminal, and a third carbon-carbon double bond is introduced.
- the present invention relates to a method for producing a chain unsaturated carboxylic acid compound. Further, the present invention produces a second chain unsaturated carboxylic acid compound from a first chain unsaturated carboxylic acid compound having an amino group at the terminal in the presence of BesC, and further phenylalanine from the compound.
- the present invention relates to a method for producing a third chain unsaturated carboxylic acid compound in the presence of ammonia lyase. Furthermore, the present invention thus produces a third chain unsaturated carboxylic acid compound, from which carbon-carbon double bonds are formed at both ends in the presence of PDC.
- the present invention relates to a method for producing a chain unsaturated hydrocarbon compound having.
- the present invention also relates to a PAL variant, a DNA encoding the variant, a vector into which the DNA is inserted, and a host cell into which the DNA or the vector has been introduced, which can be used in these production methods. Furthermore, the present invention relates to a method for producing the mutant using the host cell.
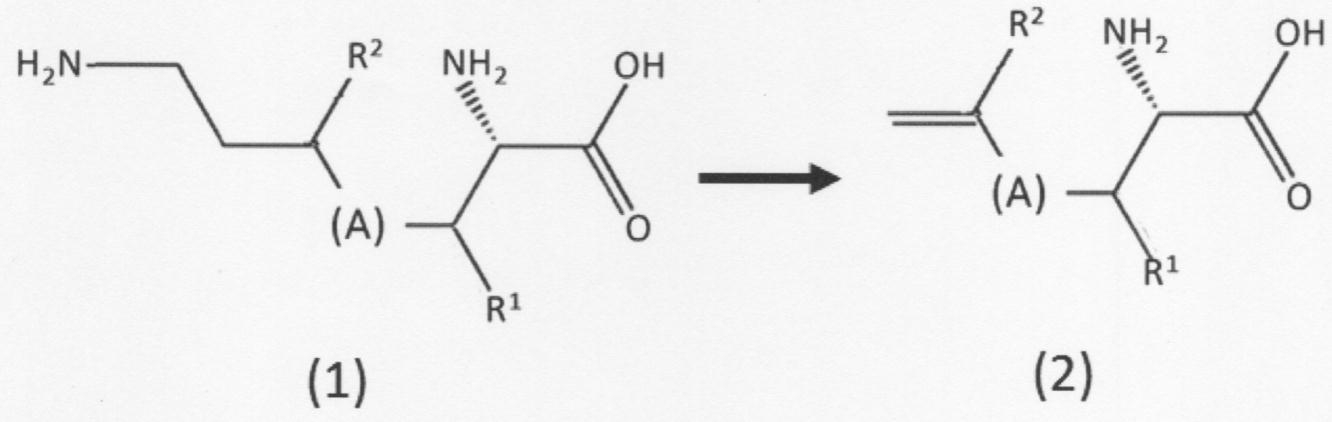
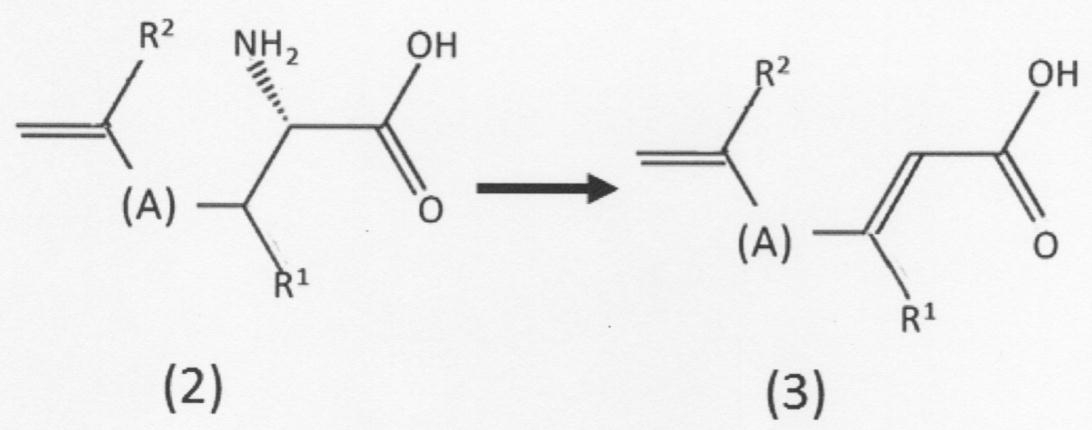
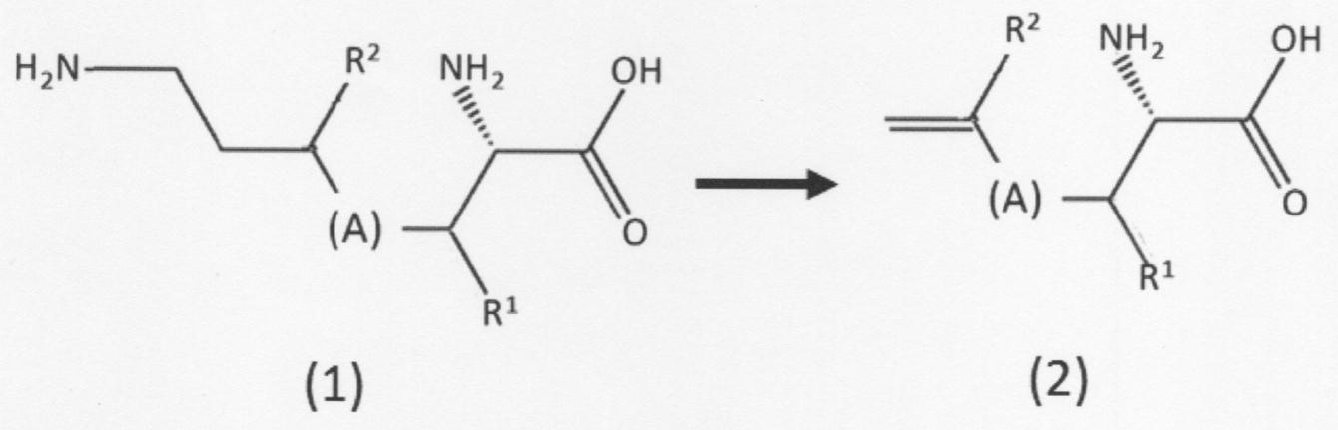
- the present invention provides the following: [1] A chain unsaturated carboxylic acid compound represented by the following formula (2) or having a first amino group and a first carbon-carbon double bond at the terminal in the presence of phenylalanine ammonia lyase.
- a chain unsaturated carboxylic acid compound represented by the following formula (3) which comprises a step of removing the first amino group from the geometric isomer to form a second carbon-carbon double bond. Or a method for producing its geometric isomer
- [(A) in the above formula indicates a linear hydrocarbon group having 0 to 5 carbon atoms which may be substituted, and when the number of carbon atoms is 2 to 5, a double bond is formed between adjacent carbon atoms. It may be formed.
- Each of R 1 and R 2 independently has a hydrogen atom, a linear or branched alkyl group having 1 to 5 carbon atoms, a linear or branched alkoxy group having 1 to 5 carbon atoms, or a hydroxyl group. Show].
- [(A) in the above formula indicates a linear hydrocarbon group having 0 to 5 carbon atoms which may be substituted, and when the number of carbon atoms is 2 to 5, a double bond is formed between adjacent carbon atoms. It may be formed.
- Each of R 1 and R 2 independently has a hydrogen atom, a linear or branched alkyl group having 1 to 5 carbon atoms, a linear or branched alkoxy group having 1 to 5 carbon atoms, or a hydroxyl group. Show].
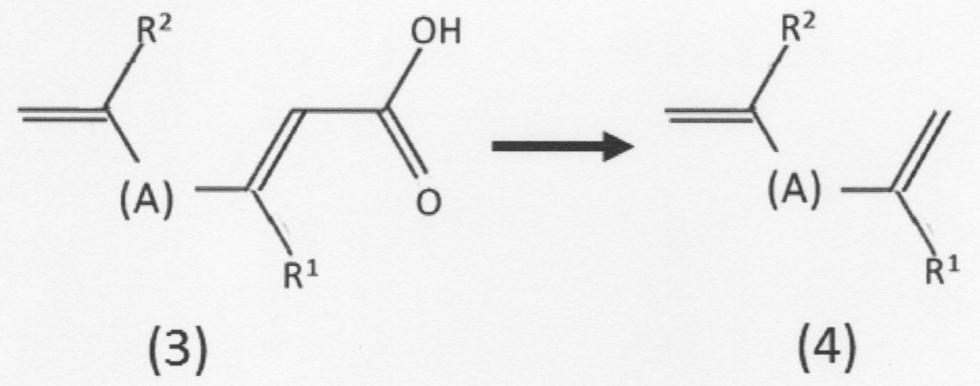
- [3] By the method according to [1] or [2], a chain unsaturated carboxylic acid compound represented by the above formula (3) or a geometric isomer thereof is produced, and in the presence of ferulic acid decarboxylase.
- [(A) in the above formula indicates a linear hydrocarbon group having 0 to 5 carbon atoms which may be substituted, and when the number of carbon atoms is 2 to 5, a double bond is formed between adjacent carbon atoms. It may be formed.
- Each of R 1 and R 2 independently has a hydrogen atom, a linear or branched alkyl group having 1 to 5 carbon atoms, a linear or branched alkoxy group having 1 to 5 carbon atoms, or a hydroxyl group. Show].
- [4] The method according to any one of [1] to [3], wherein the phenylalanine ammonia-lyase is a phenylalanine ammonia-lyase having at least one of the following characteristics (1) to (5).
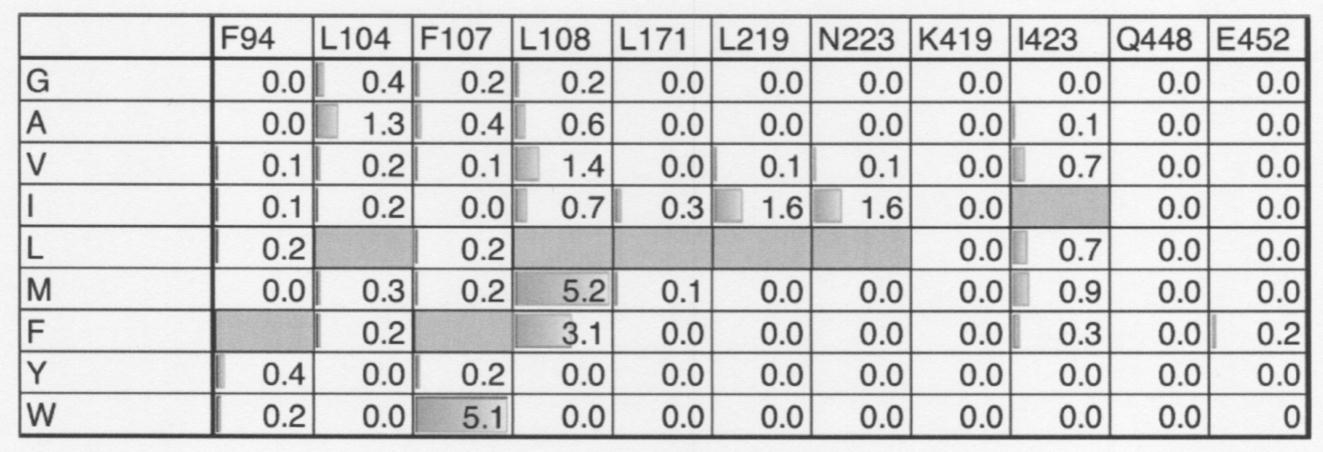
- the amino acid at position 108 of the amino acid sequence shown in SEQ ID NO: 2 or the amino acid corresponding to the site is methionine, phenylalanine or valine.
- the amino acid corresponding to the 107th position or the site of the amino acid sequence shown in SEQ ID NO: 2 is tryptophan.
- the amino acid corresponding to the 219th position or the site of the amino acid sequence shown in SEQ ID NO: 2 is isoleucine.
- the amino acid corresponding to the 223rd position or the site of the amino acid sequence shown in SEQ ID NO: 2 is isoleucine.
- the amino acid corresponding to the 104th position or the site of the amino acid sequence shown in SEQ ID NO: 2 is alanine.
- Phenylalanine ammonia-lyase variant in which at least one amino acid substitution from the following (1) to (5) has been introduced (1) Corresponds to position 108 or the site of the amino acid sequence set forth in SEQ ID NO: 2. Amino acid is replaced with methionine, phenylalanine or valine, (2) Amino acid at position 107 of the amino acid sequence shown in SEQ ID NO: 2 or the amino acid corresponding to the site is replaced with tryptophan. (3) The amino acid at position 219 of the amino acid sequence shown in SEQ ID NO: 2 or the amino acid corresponding to the site is replaced with isoleucine.
- a method for producing a phenylalanine ammonia-lyase variant which comprises a step of culturing the host cell according to [8] and collecting the protein expressed in the host cell.
- a method for producing a modified phenylalanine ammonia lyase which comprises a step of introducing an amino acid substitution of at least one of the following (1) to (5) in phenylalanine ammonia lyase.
- Method (1) SEQ ID NO:: Substitute the amino acid at position 108 of the amino acid sequence described in 2 or the amino acid corresponding to the site with methionine, phenylalanine or valine.
- the present invention it is possible to provide a method for producing an unsaturated compound having at least two carbon-carbon double bonds using an enzyme.
- the present invention also makes it possible to produce butadiene using relatively inexpensive L-lysine as a starting material.
- Phenylalanine ammonia-lyase (AtPAL, AvPAL, or PaPAL) derived from Arabidopsis thaliana, Anabaena variabilis, or Plagiochasma appendiculutum was added to a mixed solution of ferulic acid decarboxylase, and phenylalanine was added to the mixed solution.
- Graph It is a graph which shows the result of having added allylglycine to the mixed solution of AtPAL, AvPAL, or PaPAL and ferulic acid decarboxylase, and measuring the amount of butadiene produced.
- PAL phenylalanine ammonia-lyase
- the present invention relates to a method for producing a chain unsaturated compound having at least two carbon-carbon double bonds, which comprises the following reaction steps.
- the present invention has a first amino group and a first carbon-carbon double bond at the terminal in the presence of phenylalanine ammonia lyase, and is a chain-like defect represented by the above formula (2). It comprises a step of desorbing a first amino group from a saturated carboxylic acid compound or a geometric isomer thereof (second chain unsaturated carboxylic acid compound) to form a second carbon-carbon double bond.
- the present invention provides a method for producing a chain-shaped unsaturated carboxylic acid compound represented by the above formula (3) or a geometric isomer thereof (a third chain-shaped unsaturated carboxylic acid compound).
- the "third chain unsaturated carboxylic acid compound" produced in the reaction has at least two carbon-carbon double bonds including a terminal carbon-carbon double bond.
- (A) in each chemical formula indicates a linear hydrocarbon group having 0 to 5 carbon atoms which may be substituted.
- the "linear hydrocarbon group having 0 carbon atoms” means that the compounds represented by each chemical formula and the carbon atoms bonded via (A) in their geometric isomers are (A). It means that they are directly connected without going through. Further, when the linear hydrocarbon group which may be substituted has 2 to 5 carbon atoms, at least one double bond may be formed between adjacent carbon atoms. Further, the substituent that the hydrocarbon group may have in (A) is, for example, a linear or branched alkyl group having 1 to 5 carbon atoms, a linear group having 1 to 5 carbon atoms, or the substituent.
- Examples thereof include a branched alkoxy group, a hydroxyl group, a halogen atom (for example, fluorine, chlorine, bromine, iodine), a nitro group, a cyano group, an amino group, a carboxyl group, and a formyl group.
- a branched alkoxy group for example, a hydroxyl group, a halogen atom (for example, fluorine, chlorine, bromine, iodine), a nitro group, a cyano group, an amino group, a carboxyl group, and a formyl group.
- a halogen atom for example, fluorine, chlorine, bromine, iodine
- R 1 and R 2 in each chemical formula are independently hydrogen atoms, linear or branched alkyl groups having 1 to 5 carbon atoms, linear or linear groups having 1 to 5 carbon atoms, respectively. It shows a branched alkoxy group or hydroxyl group.
- Examples of the "linear or branched alkyl group having 1 to 5 carbon atoms" include a methyl group, an ethyl group, an n-propyl group, an i-propyl group, an n-butyl group, an i-butyl group, and s.
- -Butyl group, t-butyl group, n-pentyl group, i-pentyl group can be mentioned.
- Examples of the "linear or branched alkoxy group having 1 to 5 carbon atoms" include a methoxy group, an ethoxy group, an n-propoxy group, an i-propoxy group, an n-butoxy group, an i-butoxy group, and s.
- -Butoxy group, t-butoxy group, n-pentyloxy group, i-pentyloxy group, n-pentyloxy group, 1,2-dimethyl-propoxy group can be mentioned.
- R 1 in each chemical formula it is preferably a hydrogen atom.
- R2 is preferably a hydrogen atom or a methyl group.
- the combination of (A), R 1 and R 2 in each chemical formula is preferably a linear hydrocarbon group having 0 carbon atoms, a hydrogen group and a hydrogen group, or a linear hydrocarbon group having 0 carbon atoms, respectively. , Hydrogen group and methyl group.
- the "third chain unsaturated carboxylic acid compound” is preferably pentadienoic acid, 4-methylpentadienoic acid, or 3-methylpentadienoic acid.
- the deamination is promoted and the third chain unsaturated carboxylic acid compound is produced.
- Any condition can be used, and a person skilled in the art can appropriately adjust and set the composition of the reaction solution, the pH of the reaction solution, the reaction temperature, the reaction time, and the like.
- reaction solution to which the phenylalanine ammonia-lyase according to the present invention and the second chain unsaturated carboxylic acid compound as a substrate thereof are added is not particularly limited as long as the reaction is not hindered, but is preferable.
- a buffer solution having a pH of 6 to 8 is mentioned, and more preferably, a buffer solution containing potassium chloride and sodium phosphate having a pH of 6 to 7 is mentioned.
- the reaction temperature is not particularly limited as long as it does not interfere with the reaction, but is usually 20 to 40 ° C, preferably 25 to 37 ° C. Further, the reaction time may be any time as long as the unsaturated hydrocarbon compound can be produced, and is not particularly limited, but is usually 30 minutes to 7 days, preferably 12 hours to 2 days.
- the produced third chain unsaturated carboxylic acid compound can be collected by appropriately using a known recovery and purification method (distillation, chromatography, etc.). Further, these methods may be carried out alone or in combination as appropriate and may be carried out in multiple steps.
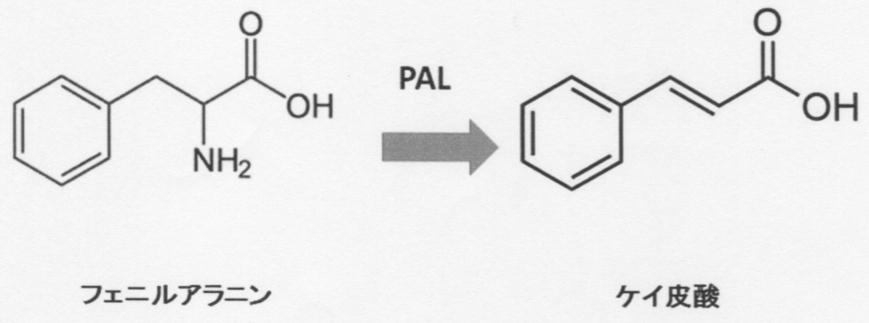
- Phenylalanine ammonia-lyase is an enzyme registered as EC number: 4.3.1.24, and means an enzyme that uses phenylalanine as a substrate and catalyzes a reaction that produces cinnamic acid and ammonia. It is also an enzyme also called PAL, tylase, phenylalanine deaminase, tyrosine ammonia-lyase, L-tyrosine ammonia-lyase, phenylalanine ammonium-lyase, and L-phenylalanine ammonia-lyase.
- the phenylalanine ammonia-lyase according to the present invention is not particularly limited, and those derived from various organisms can be used.
- phenylalanine ammonia-lyase derived from Anabaena variabilis ⁇ Arabidopsis thaliana ⁇ Plagiochasma appendiculatum ⁇ Rhodotorula glutinis ⁇ Planctomyces brasiliensis ⁇ Oryza sativa ⁇ Bambusa oldhamii ⁇ Taxus chinensis ⁇ Nicotiana tabacum ⁇ Streptomyces maritimus ⁇ Salvia miltiorrhiza ⁇ Solanum lycopersicum ⁇ Among these, as shown in Examples described later, from the viewpoint of higher catalytic activity for producing a third chain unsaturated carboxylic acid compound, phenylalanine ammonia-lyase derived from Anabaena variabilis ⁇ Arabidopsis thaliana ⁇ Plagiochasma append
- the phenylalanine ammonia-lyase according to the present invention has an identity of 15% or more (for example, 16% or more, 17% or more, 18% or more, 19% or more) with the amino acid sequence shown in SEQ ID NO: 2. It is preferably 20% or more (for example, 30% or more, 40% or more), more preferably 50% or more (for example, 60% or more, 70% or more), and further preferably 80% or more (for example, 60% or more, 70% or more). For example, it is more preferably 85% or more, 86% or more, 87% or more, 88% or more, 89% or more, and 90% or more (for example, 91% or more, 92% or more, 93% or more, 94% or more).
- the "identity" with the amino acid sequence set forth in SEQ ID NO: 2 is the amino acid sequence set forth in the phenylalanine ammonia lyase according to the present invention and the amino acid sequence set forth in SEQ ID NO: 2 with respect to the total number of amino acids of the phenylalanine ammonia lyase according to the present invention. It means the ratio (%) of the number of matching amino acids.
- the phenylalanine ammonia-lyase according to the present invention may have a natural or non-natural (artificial) mutation introduced into the amino acid sequence set forth in SEQ ID NO: 2. That is, one or more amino acids were substituted, deleted, added, and / or inserted into the phenylalanine ammonia-lyase according to the present invention in the amino acid sequence of phenylalanine ammonia-lyase (amino acid sequence shown in SEQ ID NO: 2, etc.). It also includes proteins consisting of amino acid sequences.
- the term "plurality” is not particularly limited, but is usually 2 to 200, preferably 2 to 150, more preferably 2 to 100, still more preferably 2 to 70, and more preferably 2 to 50. , More preferably 2 to 30, more preferably 2 to 20, and even more preferably 2 to 10 (eg, 2 to 9, 2 to 8, 2 to 7, 2 to 6, 2 to 5). 2 to 4 pieces, 2 to 3 pieces, 2 pieces).
- the phenylalanine ammonia-lyase according to the present invention is preferably phenylalanine ammonia-lyase having at least one of the following characteristics (1) to (5).
- the amino acid corresponding to the 108th position or the site of the amino acid sequence shown in SEQ ID NO: 2 is methionine, phenylalanine or valine.
- the amino acid corresponding to the 107th position or the site of the amino acid sequence shown in SEQ ID NO: 2 is tryptophan.
- the amino acid corresponding to the 219th position or the site of the amino acid sequence shown in SEQ ID NO: 2 isoleucine.
- the amino acid corresponding to the 223rd position or the site of the amino acid sequence shown in SEQ ID NO: 2 is isoleucine.
- the amino acid corresponding to the 104th position or the site of the amino acid sequence shown in SEQ ID NO: 2 is alanine.
- the "corresponding site" of phenylalanine ammonia lyase refers to nucleotide and amino acid sequence analysis software (GENETYX-MAC, Sequencer, etc.) and BLAST (http://blast.ncbi.nlm.nih.gov/).
- the amino acid at position 108 of the amino acid sequence set forth in SEQ ID NO: 2 or the amino acid corresponding to the site is methionine, and the amino acid at position 107 of the amino acid sequence set forth in SEQ ID NO: 2 or the same.
- the amino acid corresponding to the site is preferably tryptophan, and the amino acid at position 108 of the amino acid sequence set forth in SEQ ID NO: 2 or the amino acid corresponding to the site is preferably phenylalanine ammoniamariase having at least one characteristic of phenylalanine.
- the amino acid at position 108 of the amino acid sequence set forth in No .: 2 or the amino acid corresponding to the site is methionine, and the amino acid corresponding to position 107 or the site of the amino acid sequence set forth in SEQ ID NO: 2 is characterized by at least one of tryptophan. It is more preferable to have phenylalanine ammonia lyase.
- the phenylalanine ammonia-lyase having a specific amino acid at each such site may be a wild-type phenylalanine ammonia-lyase, and may have at least one of the above-mentioned characteristics (1) to (5). It may be a variant of phenylalanine ammonia-lyase into which an amino acid substitution has been introduced.
- any of the following (a) to (c) can be mentioned as such a mutant.
- SEQ ID NO: 2 which comprises an amino acid sequence in which at least one of the following (1) to (5) has been introduced with an amino acid substitution
- SEQ ID NO: The amino acid at position 108 of the amino acid sequence described in 2 or the amino acid corresponding to the site is replaced with methionine, phenylalanine or valine.
- Amino acid at position 107 of the amino acid sequence shown in SEQ ID NO: 2 or the amino acid corresponding to the site is replaced with tryptophan.
- the "identity" of the "plurality” of amino acids substituted, deleted, added, and / or inserted with the amino acid sequence set forth in SEQ ID NO: 2, including their preferred embodiments (range), is described above. That's right.
- the phenylalanine ammonia-lyase variant may be naturally or unnaturally (artificially) generated. That is, a phenylalanine ammonia-lyase variant having an artificially introduced amino acid substitution or the like is also included.
- phenylalanine ammonia-lyase or a natural or non-natural variant thereof has catalytic activity for producing a third chain unsaturated carboxylic acid compound is a method known to those skilled in the art. It can be determined by directly measuring the amount of the unsaturated carboxylic acid compound produced (eg, chromatographic mass analysis).
- the phenylalanine ammonia-lyase according to the present invention may be directly or indirectly added with another compound.
- the addition is not particularly limited, and may be an addition at the gene level or a chemical addition.
- the site to be added is not particularly limited, and may be either the amino terminal or the carboxyl terminal of the phenylalanine ammonia-lyase according to the present invention, or both of them. Addition at the gene level is achieved by using a DNA encoding phenylalanine ammonia-lyase according to the present invention to which a reading frame of DNA encoding another protein is added.
- the "other protein” added in this manner is not particularly limited, and for the purpose of facilitating the purification of the phenylalanine ammonia lyase according to the present invention, a polyhistidine (His-) tag (tag) protein, Purification tag proteins such as FLAG-tag protein (registered trademark, Sigma-Aldrich) and glutathione-S-transferase (GST) are preferably used, and for the purpose of facilitating the detection of phenylalanine ammonia lyase according to the present invention.
- a tag protein for detection such as a fluorescent protein such as GFP and a chemically luminescent protein such as luciferase is preferably used.
- the chemical addition may be covalent or non-covalent.
- the "covalent bond” is not particularly limited, and is, for example, an amide bond between an amino group and a carboxyl group, an alkylamine bond between an amino group and an alkyl halide group, a disulfide bond between thiols, a thiol group and a maleimide group or an alkyl halide.
- a thioether bond with a group can be mentioned.
- Examples of the "non-covalent bond” include a biotin-avidin bond.
- fluorescent dyes such as Cy3 and rhodamine are suitable. Used for.
- phenylalanine ammonia-lyase according to the present invention may be mixed with other components and used.
- the other components are not particularly limited, and examples thereof include sterile water, physiological saline, vegetable oils, surfactants, lipids, lysis aids, buffers, protease inhibitors, and preservatives.
- the present invention can also take an aspect of a method for producing a third chain unsaturated carboxylic acid compound through the production of a second chain unsaturated carboxylic acid compound.
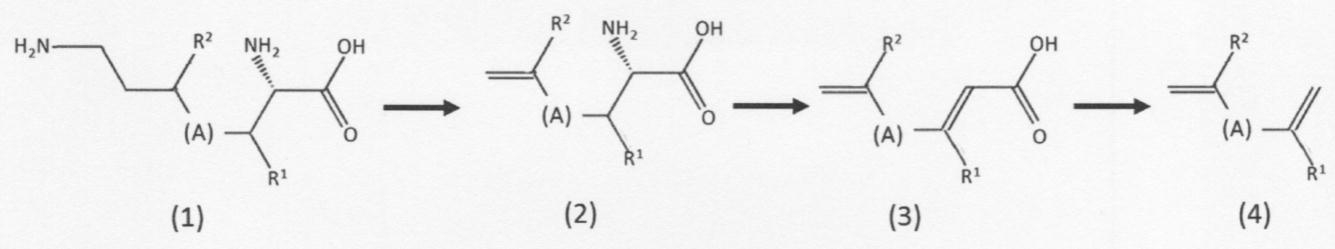
- the "second chain unsaturated carboxylic acid compound” is represented by the above formula (2) having a first amino group and a first carbon-carbon double bond at the terminal. It means a chain unsaturated carboxylic acid compound or a geometric isomer thereof. Further, the "first chain unsaturated carboxylic acid compound” used as a raw material for producing the same has a first amino group and a second amino group at the end according to the above formula (1). It means a chain unsaturated carboxylic acid compound represented or a geometric isomer thereof.
- (A) and R 1 are as described above, including their preferred embodiments, but the "second chain unsaturated carboxylic acid compound” is preferably L-allylglycine. L- (2-methylallyl) glycine and L- (3-methylallyl) glycine.
- the "first chain unsaturated carboxylic acid compound” is preferably L-lysine, 4-methyllysine, or 3-methyllysine.
- the condition for forming a carbon-carbon double bond at the terminal of the first chain unsaturated carboxylic acid compound in the presence of BesC according to the present invention the formation of the double bond is promoted, and the second
- the conditions may be such that a chain unsaturated carboxylic acid compound is produced, and those skilled in the art can appropriately adjust and set the composition of the reaction solution, the pH of the reaction solution, the reaction temperature, the reaction time, and the like. ..
- the reaction solution to which BesC according to the present invention and the first chain unsaturated carboxylic acid compound as a substrate thereof are added is not particularly limited as long as the reaction is not hindered, but is preferably pH 6 to. 8 buffers, more preferably buffers containing potassium chloride and sodium phosphate at pH 6-7. Further, iron sulfate is preferably contained from the viewpoint of facilitating the reaction.
- the reaction temperature is not particularly limited as long as it does not interfere with the reaction, but is usually 20 to 40 ° C, preferably 25 to 37 ° C. Further, the reaction time may be any time as long as the unsaturated hydrocarbon compound can be produced, and is not particularly limited, but is usually 30 minutes to 7 days, preferably 12 hours to 2 days.
- the produced second chain unsaturated carboxylic acid compound can be collected by appropriately using a known recovery and purification method (distillation, chromatography, etc.). Further, these methods may be carried out alone or in combination as appropriate and may be carried out in multiple steps.
- the first chain unsaturated carboxylic acid compound as a raw material can be purchased as a commercially available product as shown in Examples described later. Further, those skilled in the art can also synthesize by appropriately considering a known synthesis method (for example, the method described in the method for producing L-lysine by a fermentation method (JPH0530985A)).
- BesC is one enzyme related to ⁇ -ethynylserine biosynthesis (Bes).
- the enzyme has an activity of catalyzing a reaction that cleaves a carbon-carbon bond in the group by oxidizing a propylamino group at the terminal and forms a carbon-carbon double bond at the terminal. It is an alkene-producing enzyme (Non-Patent Document 2).
- the origin of BesC is not particularly limited as long as it has a catalytic activity that promotes the formation of a second chain unsaturated carboxylic acid compound, and various biological sources can be used.
- the origin of BesC includes Pseudomonas fluororescens and Streptomyces cattleya.
- BesC derived from Pseudomonas fluorescens (BesC consisting of the amino acid sequence shown in SEQ ID NO: 11) is preferable.
- the BesC according to the present invention has an identity of 15% or more (for example, 16% or more, 17% or more, 18% or more, 19% or more) with the amino acid sequence shown in SEQ ID NO: 11. It is more preferably 20% or more (for example, 30% or more, 40% or more), further preferably 50% or more (for example, 60% or more, 70% or more), and even more preferably 80% or more (for example, 60% or more). It is more preferably 85% or more, 86% or more, 87% or more, 88% or more, 89% or more, and 90% or more (for example, 91% or more, 92% or more, 93% or more, 94% or more, 95).
- identity with the amino acid sequence shown in SEQ ID NO: 11 means the number of amino acids matching the total number of BesC amino acids according to the present invention with the BesC according to the present invention and the amino acid sequence shown in SEQ ID NO: 11. Means the percentage of.
- the BesC according to the present invention may have a natural or non-natural (artificial) mutation introduced into the amino acid sequence set forth in SEQ ID NO: 11. That is, the BesC according to the present invention comprises an amino acid sequence in which one or more amino acids are substituted, deleted, added, and / or inserted in the amino acid sequence of BesC (amino acid sequence shown in SEQ ID NO: 11 or the like). It also contains proteins.
- the term "plurality” is not particularly limited, but is usually 2 to 80, preferably 2 to 70, more preferably 2 to 60, still more preferably 2 to 50, and more preferably 2 to 40. , More preferably 2 to 30, more preferably 2 to 20, and even more preferably 2 to 10 (eg, 2 to 9, 2 to 8, 2 to 7, 2 to 6, 2 to 5). 2 to 4 pieces, 2 to 3 pieces, 2 pieces).
- BesC or a natural or non-natural variant thereof has a catalytic activity for producing a second chain unsaturated carboxylic acid compound is known by those skilled in the art (for example, a method known to those skilled in the art). , Chromatographic mass analysis), it can be determined by directly measuring the amount of the unsaturated carboxylic acid compound produced.
- BesC according to the present invention may be directly or indirectly added with another compound, similarly to the above-mentioned phenylalanine ammonia-lyase according to the present invention. Furthermore, BesC according to the present invention may be mixed with other components and used in the same manner as phenylalanine ammonia-lyase.
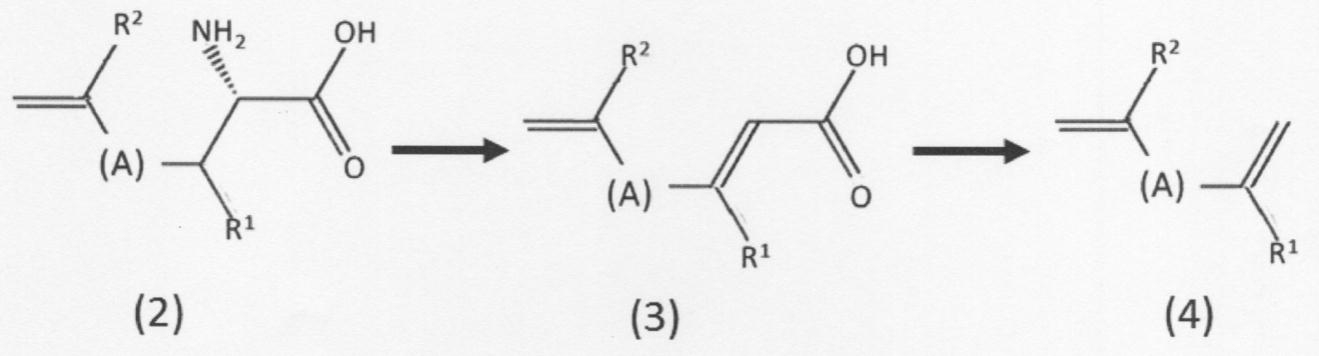
- the second chain unsaturated carboxylic acid compound represented by the following formula (2) is replaced with the third chain represented by the following formula (3).
- the chain represented by the following formula (4) is formed.
- a mode of the production method shown below may also be taken, which comprises a step of producing an unsaturated hydrocarbon compound or a geometric isomer thereof (hereinafter, also simply referred to as “chain unsaturated hydrocarbon compound”).
- the "chain unsaturated hydrocarbon compound” produced in the reaction has a carbon-carbon double bond at both ends and is represented by the above formula (4). It means a hydrocarbon compound or a geometric isomer thereof.
- (A) and R 1 are as described above, including their preferred embodiments, but the "chain unsaturated hydrocarbon compound” is preferably butadiene or isoprene.
- the decarboxylation is promoted and the unsaturated hydrocarbon compound is produced.
- Any person skilled in the art can appropriately adjust and set the composition of the reaction solution, the pH of the reaction solution, the reaction temperature, the reaction time, and the like.
- the reaction solution to which the ferulic acid decarboxylase according to the present invention and the unsaturated hydrocarbon dicarboxylic acid compound as a substrate thereof are added is not particularly limited as long as the reaction is not hindered, but is preferably pH 6 to 8.
- the above-mentioned buffer solution is mentioned, and more preferably, a buffer solution containing potassium chloride and sodium phosphate having a pH of 6 to 7 is mentioned.
- the prenylated flavin mononucleotide (prFMN) or its isomer prFMN ketimine , prFMN iminiu , these prFMN and its isomer are referred to in Non-Patent Document 1). Is preferably contained.
- the reaction temperature is not particularly limited as long as it does not interfere with the reaction, but is usually 20 to 40 ° C, preferably 25 to 37 ° C. Further, the reaction time may be any time as long as the unsaturated hydrocarbon compound can be produced, and is not particularly limited, but is usually 30 minutes to 7 days, preferably 12 hours to 2 days.
- the chain unsaturated hydrocarbon compound produced under such conditions is generally easily vaporized, it can be collected by a known recovery and purification method for volatile gas.
- sampling methods include gas stripping, distillation, adsorption, desorption, pervaporation, desorption of unsaturated hydrocarbon compounds adsorbed on the solid phase from the solid phase by heat or vacuum, extraction with a solvent, or chromatography. (For example, gas chromatography) and the like.
- the unsaturated hydrocarbon compound to be produced is a liquid, it can be collected by appropriately using a known recovery / purification method (distillation, chromatography, etc.). Further, these methods may be carried out alone or in combination as appropriate and may be carried out in multiple steps.
- Ferulic acid decarboxylase is an enzyme registered as EC number: 4.1.1.102, and is usually a reaction of decarboxylating ferulic acid to produce 4-vinylguaiyacol (4VG). Means an enzyme that catalyzes.
- the "ferulic acid decarboxylase” has an activity of catalyzing a reaction of desorbing a carboxyl group from a third chain unsaturated carboxylic acid compound to form a chain unsaturated hydrocarbon compound.
- a thing there is no particular limitation, and a thing derived from various organisms can be used.
- Tables 1 to 6 of Patent Document 3 a protein corresponding to "Ferric acid decarboxylase” on UNIPROT can be mentioned.
- examples of the ferulic acid decarboxylase according to the present invention include ferulic acid decarboxylase derived from saccharomyces and ferulic acid decarboxylase derived from Aspergillus (UNIPROT ID: A2QHE5, etc.), and more preferably ferulic acid decarboxylase derived from saccharomyces.
- Carboxylase can be mentioned. ⁇ Saccharomyces( ⁇ ) ⁇ Saccharomyces cerevisiae ⁇ Saccharomyces kudriavzevii ⁇ Saccharomyces eubayanus ⁇ Saccharomyces bayanus ⁇ Saccharomyces boulardii ⁇ Saccharomyces bulderi ⁇ Saccharomyces cariocanus ⁇ Saccharomyces cariocus ⁇ Saccharomyces chevalieri ⁇ Saccharomyces dairenensis ⁇ Saccharomyces ellipsoideus ⁇ Saccharomyces florentinus ⁇ Saccharomyces kluyveri ⁇ Saccharomyces martiniae ⁇ Saccharomyces monacensis ⁇ Saccharomyces norbensis ⁇ Saccharomyces paradoxus ⁇ Saccharomyces pastorianus ⁇ Saccharomyces spencerorum ⁇ Saccharomyces turic
- ferulic acid decarboxylase derived from Saccharomyces for example, FDC (UNIPROT ID: Q03034, SEQ ID NO: 13) derived from FDC (UNIPROT ID: Q03034, SEQ ID NO: 13) derived from Saccharomyces cerevisiae (ATCC 204508 / S288c strain) (Pan yeast).
- FDC UNIPROT ID: Q03034, SEQ ID NO: 13
- Saccharomyces cerevisiae ATCC 204508 / S288c strain
- a protein derived from Saccharomyces corresponding to "Ferulic acid decarboxylase" on UNIPROT can be mentioned, and more specifically, ferulic acid decarboxylase shown in Table 1 below can be mentioned. It should be understood that mutations in the nucleotide sequence in nature can result in changes in the amino acid sequence of the protein.
- the ferulic acid decarboxylase according to the present invention has an identity of 15% or more (for example, 16% or more, 17% or more, 18% or more, 19% or more) with the amino acid sequence shown in SEQ ID NO: 13. It is preferably 20% or more (for example, 30% or more, 40% or more), more preferably 50% or more (for example, 60% or more, 70% or more), and further preferably 80% or more. (For example, 85% or more, 86% or more, 87% or more, 88% or more, 89% or more) is more preferable, and 90% or more (for example, 91% or more, 92% or more, 93% or more, 94%).
- identity with the amino acid sequence shown in SEQ ID NO: 13 means the amino acid decarboxylase according to the present invention and the amino acid shown in SEQ ID NO: 13 with respect to the total number of amino acids of the ferulic acid decarboxylase according to the present invention. It means the percentage of amino acids that match the sequence.
- the ferulic acid decarboxylase according to the present invention may have a mutation introduced into the amino acid sequence set forth in SEQ ID NO: 13. That is, in the ferulic acid decarboxylase according to the present invention, one or more amino acids are substituted, deleted, added, and / or inserted in the amino acid sequence of the ferulic acid decarboxylase (amino acid sequence shown in SEQ ID NO: 13, etc.). Also included is a protein consisting of the amino acid sequence.
- the term "plurality” is not particularly limited, but is usually 2 to 150, preferably 2 to 100, more preferably 2 to 80, still more preferably 2 to 50, and more preferably 2 to 30.
- such ferulic acid decarboxylase mutants may be naturally or unnaturally (artificially) generated. That is, a ferulic acid decarboxylase variant into which an amino acid substitution or the like is artificially introduced is also included.
- ferulic acid decarboxylase or a natural or non-natural variant thereof has a catalytic activity for producing a chain unsaturated hydrocarbon compound can be determined by those skilled in the art, for example, as described below. As shown in the example, it can be determined by directly measuring the amount of unsaturated hydrocarbon compound by gas chromatography-mass spectrometry (GC-MS).
- GC-MS gas chromatography-mass spectrometry
- the ferulic acid decarboxylase according to the present invention may be directly or indirectly added with another compound, similarly to the above-mentioned phenylalanine ammonia-lyase according to the present invention. Furthermore, the ferulic acid decarboxylase according to the present invention may be used in combination with other components in the same manner as phenylalanine ammonia-lyase.
- DNA encoding the enzyme according to the present invention phenylalanine ammonia-lyase according to the present invention, BesC according to the present invention, ferulic acid decarboxylase according to the present invention
- the traits of the host cell can be transformed to produce various enzymes according to the present invention in the cell, and thus a third chain of unsaturated carboxylic acid compound or chain of unsaturated hydrocarbons can be produced. It becomes possible to produce a hydrogen compound.
- the DNA according to the present invention may be a natural DNA or a DNA in which a mutation is artificially introduced into the natural DNA as long as it encodes the above-mentioned enzyme according to the present invention, and may be artificially introduced. It may be a DNA consisting of a designed nucleotide sequence. Further, the morphology is not particularly limited, and genomic DNA and chemically synthesized DNA are included in addition to cDNA. Preparation of these DNAs can be carried out by those skilled in the art using conventional means. For genomic DNA, for example, genomic DNA is extracted from various bacteria to prepare a genomic library (plasmids, phages, cosmids, BACs, PACs, etc. can be used as vectors), and the present invention is developed.
- genomic DNA for example, genomic DNA is extracted from various bacteria to prepare a genomic library (plasmids, phages, cosmids, BACs, PACs, etc. can be used as vectors), and the present invention is developed.
- cDNA for example, cDNA is synthesized based on mRNA extracted from various bacteria, and this is inserted into a vector such as ⁇ ZAP to prepare a cDNA library, which is expanded to develop colony high in the same manner as above. It can be prepared by performing hybridization or plaque hybridization, or by performing PCR.
- a mutation encoding the above-mentioned amino acid substitution can be introduced into the DNA thus prepared by a person skilled in the art by using a known part-specific mutation introduction method.
- the part-specific mutagenesis method include the Kunkel method (Kunkel, TA, Proc Natl Acad Sci USA, 1985, Vol. 82, No. 2, pp. 488-492), SOE (splicing-by-overlap).
- SOE splicing-by-overlap
- -Extension) -PCR method Ho, S.N., Hunt, HD, Horton, RM, Pullen, JK, and Peace, LR, Gene, 1989, Vol. 77 , Pages 51-59).
- nucleotide sequence encoding the protein into which the above-mentioned amino acid substitution has been introduced, and based on the sequence information, use an automatic nucleic acid synthesizer to chemically synthesize the DNA according to the present invention. It can also be synthesized as a target.
- the DNA according to the present invention is an enzyme according to the present invention in which codons are optimized according to the type of the host cell from the viewpoint of further improving the expression efficiency of the decarboxylase according to the present invention to be encoded.
- the aspect of the DNA encoding the above can also be taken.
- the vector in which the DNA is inserted can also be taken so that the above-mentioned DNA can be replicated in the host cell.
- the "vector” exists as a self-replicating vector, that is, an extrachromosomal independent entity, and its replication does not depend on chromosomal replication, for example, it can be constructed on the basis of a plasmid.
- the vector may also be one that, when introduced into a host cell, integrates into the genome of the host cell and replicates with the chromosome into which it has been integrated.
- Examples of such a vector include a plasmid and phage DNA.
- the plasmids include Escherichia coli-derived plasmids (pET22, pBR322, pBR325, pUC118, pUC119, pUC18, pUC19, etc.), yeast-derived plasmids (YEp13, YEp24, YCp50, etc.), Bacillus subtilis-derived plasmids (pUB110, pTP5, etc.).
- Examples of the phage DNA include ⁇ phage (Charon4A, Charon21A, EMBL3, EMBL4, ⁇ gt10, ⁇ gt11, ⁇ ZAP, etc.).
- the present invention also comprises an insect virus vector such as baculovirus if the host cell is derived from an insect, T-DNA or the like if the host cell is derived from an animal, and an animal virus vector such as a retrovirus or an adenovirus vector if the host cell is derived from an animal. It can also be used as a vector for the virus.
- an insect virus vector such as baculovirus if the host cell is derived from an insect, T-DNA or the like if the host cell is derived from an animal
- an animal virus vector such as a retrovirus or an adenovirus vector if the host cell is derived from an animal. It can also be used as a vector for the virus.
- the procedure and method for constructing the vector according to the present invention those commonly used in the field of genetic engineering can be used. For example, in order to insert the DNA according to the present invention into a vector, first, the purified DNA is cleaved with an appropriate restriction enzyme, inserted into a restriction enzyme site or a
- the vector according to the present invention may be in the form of an expression vector containing the enzyme according to the present invention encoded by the DNA in a state capable of being expressed in a host cell.
- a DNA sequence that controls the expression and a transformed host cell are used. It is desirable to include a genetic marker or the like for selection.
- the DNA sequence that controls expression includes a promoter, an enhancer, a splicing signal, a poly A addition signal, a ribosome binding sequence (SD sequence), a terminator, and the like.
- the promoter is not particularly limited as long as it exhibits transcriptional activity in the host cell, and can be obtained as a DNA sequence that controls the expression of a gene encoding a protein of the same species or heterologous to the host cell.
- a DNA sequence that induces the expression may be included.
- the expression of the gene located downstream is induced by adding isopropyl- ⁇ -D-thiogalactopyranoside (IPTG).
- IPTG isopropyl- ⁇ -D-thiogalactopyranoside
- IPTG isopropyl- ⁇ -D-thiogalactopyranoside
- lactose operon There is a lactose operon that can be used.
- the gene marker in the present invention may be appropriately selected depending on the method for selecting the transformed host cell, and for example, a gene encoding drug resistance or a gene complementing auxotrophy can be used.
- DNA encoding the enzyme according to the present invention may be inserted into the vector, or a plurality of types of DNA may be inserted into one vector.
- a plurality of types of DNA When inserting a plurality of types of DNA into a vector, it is preferable that these DNAs form an operon when a plurality of types of DNA are inserted into one vector.
- the "operon" is a nucleic acid sequence unit composed of one or a plurality of genes transcribed under the control of the same promoter.
- a vector into which a DNA encoding phenylalanine ammonia-lyase according to the present invention is inserted A combination of a vector in which a DNA encoding BesC according to the present invention is inserted and a vector in which a DNA encoding phenylalanine ammonia-lyase according to the present invention is inserted, A combination of a vector in which a DNA encoding phenylalanine ammonia-lyase according to the present invention is inserted and a vector in which a DNA encoding a ferulic acid decarboxylase according to the present invention is inserted, or a BesC according to the present invention is encoded.
- the combination of the vector in which the DNA is inserted, the vector in which the DNA encoding the phenylalanine ammonia-lyase according to the present invention is inserted, and the vector in which the DNA encoding the ferulic acid decarboxylase according to the present invention is inserted is , Can be mentioned. Further, a vector in which a DNA encoding BesC according to the present invention and a DNA encoding phenylalanine ammonia-lyase according to the present invention are inserted.
- a vector in which the DNA to be used and the ferulic acid decarboxylase according to the present invention are inserted can also be mentioned.
- DNA or vector according to the present invention may be mixed with other components and used.
- the other components are not particularly limited, and examples thereof include sterile water, physiological saline, vegetable oils, surfactants, lipids, solubilizing agents, buffers, DNase inhibitors, and preservatives.
- ⁇ An agent for promoting the formation of a third chain unsaturated carboxylic acid compound or a chain unsaturated hydrocarbon compound As described above, by using the enzyme according to the present invention, the DNA encoding the enzyme, or the vector into which the DNA is inserted, a third chain unsaturated carboxylic acid compound or a chain unsaturated hydrocarbon is used. It is possible to promote the formation of compounds.
- the present invention is a third chain unsaturated carboxylic acid compound or chain unsaturated hydrocarbon comprising the enzyme according to the invention, the DNA encoding the enzyme or the vector into which the DNA is inserted.
- An agent for promoting the formation of a compound is provided.
- the DNA encoding the enzyme or the vector into which the DNA is inserted which comprises the following aspects.
- An agent for promoting the formation of a third chain unsaturated carboxylic acid compound Includes BesC according to the present invention, DNA encoding the enzyme or a vector into which the DNA is inserted, and phenylalanine ammonia-lyase according to the present invention, DNA encoding the enzyme or a vector into which the DNA is inserted.
- An agent for promoting the formation of a third chain unsaturated carboxylic acid compound from a first chain unsaturated carboxylic acid compound A third chain of unsaturated carboxylic acid compounds from a first chain containing a vector into which a DNA encoding BesC according to the present invention and a DNA encoding phenylalanine ammonia-lyase according to the present invention are inserted.
- An agent for promoting the formation of a chain unsaturated hydrocarbon compound from a second chain unsaturated carboxylic acid compound which comprises. Chained from a second chain of unsaturated carboxylic acid compounds, comprising a vector into which the DNA encoding phenylalanine ammonia-lyase according to the present invention and the DNA encoding ferulic acid decarboxylase according to the present invention are inserted.
- An agent for promoting the formation of unsaturated hydrocarbon compounds, The present invention includes BesC according to the present invention, DNA encoding the enzyme or a vector into which the DNA is inserted, phenylalanine ammonia lyase according to the present invention, DNA encoding the enzyme or a vector into which the DNA is inserted.
- a first strand comprising a vector into which a DNA encoding BesC according to the present invention, a DNA encoding phenylalanine ammonia-lyase according to the present invention, and a DNA encoding ferulic acid decarboxylase according to the present invention are inserted.
- an agent it may be any one containing the enzyme or the like according to the present invention, but it may be mixed with other components and used.
- the other components are not particularly limited, and examples thereof include sterile water, physiological saline, vegetable oil, surfactant, lipid, lysis aid, buffer, protease inhibitor, DNase inhibitor, and preservative.
- the present invention can also provide a kit containing such an agent.
- the above-mentioned agent may be contained in the mode of a host cell described later, to which the DNA or the like according to the present invention has been introduced and transformed.
- a host cell for introducing various substrates first chain-like unsaturated carboxylic acid compound, second chain-like unsaturated carboxylic acid compound
- a medium for culturing the host cells, instructions for use thereof, and the like may be included in the kit of the present invention.
- Such an instruction manual is an instruction for using the agent or the like of the present invention in the method for producing the above-mentioned third chain unsaturated carboxylic acid compound or chain unsaturated hydrocarbon compound. ..
- the description describes, for example, information on the experimental method and conditions of the production method of the present invention, information on the agent of the present invention, etc. (for example, information such as a vector map showing a vector sequence of a vector, etc., an enzyme according to the present invention. Information on the sequence of the host cell, the origin and properties of the host cell, information on the culture conditions of the host cell, etc.) can be included.
- the host cell into which the DNA or vector according to the present invention is introduced is not particularly limited, and is, for example, a microorganism (Escherichia coli, sprouting yeast, dividing yeast, bacillus, actinomycete, filamentous fungus, etc.), plant cell, insect cell, animal cell, etc.
- a microorganism Esscherichia coli, sprouting yeast, dividing yeast, bacillus, actinomycete, filamentous fungus, etc.
- a third chain unsaturated carboxylic acid compound or a chain unsaturated hydrocarbon compound which exhibits high growth in a short time and is highly productive, can be used. From the viewpoint of being able to contribute to production, it is preferable to use a microorganism as a host cell, and it is more preferable to use Escherichia coli.
- the host cell into which the DNA or vector according to the present invention is introduced contains prFMN or an isomer thereof that induces prenylation of flavin mononucleotide (FMN) and contributes to the improvement of productivity of chain unsaturated hydrocarbon compounds. From the viewpoint of production, cells carrying flavin prenyl transferase are preferable.
- the host cell into which the DNA or vector according to the present invention is introduced is preferably Escherichia coli C41 (DE3) cell from the viewpoint that it is easy to handle and can express a toxic protein.
- the introduction of the DNA or vector according to the present invention can also be carried out according to a method commonly used in this field.
- examples of the method of introduction into microorganisms such as Escherichia coli include a heat shock method, an electroporation method, a spheroplast method, and a lithium acetate method
- examples of the method of introduction into plant cells include a method using agrobacterium.
- the particle gun method can be mentioned, and the method of introduction into insect cells includes a method using baculovirus and an electroporation method, and the method of introduction into animal cells includes a calcium phosphate method, a lipofection method, and an electroporation method. Can be mentioned.
- the DNA or the like introduced into the host cell in this manner may be retained in the host cell by being randomly inserted into the genomic DNA, may be retained by homologous recombination, or may be a vector. For example, it can be replicated and retained as an independent body outside its genomic DNA.
- a host cell into which a DNA or vector encoding the enzyme according to the present invention has been introduced is cultured, and a third chain unsaturated product produced in the host cell and / or the culture thereof is saturated. Also provided is a method for producing a third chain unsaturated carboxylic acid compound or chain unsaturated hydrocarbon compound, comprising the step of collecting a carboxylic acid compound or a chain unsaturated hydrocarbon compound.
- the "host cell transformed to express the enzyme according to the present invention” is as described above. Further, the combination of the enzyme according to the present invention, the DNA encoding the enzyme, and the vector into which the DNA is inserted, which is expressed or retained in such a host cell, is the above-mentioned ⁇ DNA encoding the enzyme according to the present invention, and
- the starting material based on the examples shown in the vector having the DNA> and ⁇ an agent for promoting the formation of a third chain unsaturated carboxylic acid compound or a chain unsaturated hydrocarbon compound> ( In consideration of the combination of the substrate) and the final product, those skilled in the art can appropriately change the design.
- the cell culture conditions are as described below, but various substrates (first chain unsaturated carboxylic acid compound, second chain unsaturated carboxylic acid compound) are added to the medium. It is preferable to have.
- the culture temperature may be appropriately changed in design according to the type of host cell used, but is usually 20 to 40 ° C, preferably 25 to 37 ° C.
- the "culture" is a medium containing a proliferated host cell, a secretory product of the host cell, a metabolite of the host cell, etc., which is obtained by culturing the host cell in a medium. , Includes their dilutions and concentrates.
- the collection of the third chain unsaturated carboxylic acid compound or the chain unsaturated hydrocarbon compound from such host cells and / or the culture is also not particularly limited, and the above-mentioned known recovery and purification are not particularly limited. It can be done using the method.
- the collection time may be any time as long as it is appropriately adjusted according to the type of host cell to be used and a third chain unsaturated carboxylic acid compound or chain unsaturated hydrocarbon compound can be produced. It is usually 30 minutes to 7 days, preferably 12 hours to 2 days.
- ⁇ Method for producing a modified phenylalanine ammonia-lyase according to the present invention As shown in Examples described later, by culturing a host cell into which a DNA or the like encoding a phenylalanine ammonia-lyase variant according to the present invention has been introduced, the variant can be produced in the host cell.
- the present invention comprises a step of culturing a host cell into which a DNA encoding a phenylalanine ammonia-lyase variant according to the present invention or a vector containing the DNA is introduced, and collecting a protein expressed in the host cell.
- a method for producing a modified ammonia-lyase can also be provided.
- the condition of "culturing the host cell” may be any condition as long as the host cell can produce the phenylalanine ammonia lyase variant according to the present invention.
- the temperature, the presence or absence of addition of air, the concentration of oxygen, the concentration of carbon dioxide, the pH of the medium, the culture temperature, the culture time, the humidity and the like can be appropriately adjusted and set according to the above.
- the medium may contain a medium that can be assimilated by the host cell, and may contain carbon sources, nitrogen sources, sulfur sources, inorganic salts, metals, peptone, yeast extract, meat extract, casein hydrolyzate, serum and the like. It is mentioned as an inclusion.
- an IPTG for inducing the expression of the DNA encoding the phenylalanine ammonia lyase variant according to the present invention and an antibiotic corresponding to a drug resistance gene that can be encoded by the vector according to the present invention (for example, ampicillin) or a nutrient corresponding to a gene that complements the auxotrophy that can be encoded by the vector according to the present invention (eg, arginine, histidine) may be added.
- an antibiotic corresponding to a drug resistance gene that can be encoded by the vector according to the present invention
- an antibiotic corresponding to a drug resistance gene that can be encoded by the vector according to the present invention for example, ampicillin
- a nutrient corresponding to a gene that complements the auxotrophy that can be encoded by the vector according to the present invention eg, arginine, histidine
- the host cell is recovered from the medium by filtration, centrifugation or the like, and the recovered host cell is used as a cell. It is treated by dissolution, grinding treatment, pressure crushing, etc., and further, solvent precipitation such as ultrafiltration treatment, salting, sulphate precipitation, chromatography (for example, gel chromatography, ion exchange chromatography, affinity chromatography), etc.
- solvent precipitation such as ultrafiltration treatment, salting, sulphate precipitation, chromatography (for example, gel chromatography, ion exchange chromatography, affinity chromatography), etc.
- a method for purifying and concentrating a protein expressed in a host cell can be mentioned.
- the above-mentioned purified tag protein when added to the phenylalanine ammonia-lyase variant according to the present invention, it can be purified and collected using a substrate to which the tag protein is adsorbed. Further, these purification and concentration methods may be carried out alone or in combination as appropriate and may be carried out in multiple steps.
- the phenylalanine ammonia-lyase variant according to the present invention is not limited to the above biological synthesis, and can also be produced by using the DNA or the like and the cell-free protein synthesis system according to the present invention.
- the cell-free protein synthesis system is not particularly limited, and examples thereof include wheat germ-derived, Escherichia coli-derived, rabbit reticulocyte-derived, and insect cell-derived synthetic systems.
- those skilled in the art can chemically synthesize the phenylalanine ammonia-lyase variant according to the present invention using a commercially available peptide synthesizer or the like.
- the present invention is a method for producing a phenylalanine ammonia-lyase variant having enhanced catalytic activity for producing a third chain unsaturated carboxylic acid compound from a second chain unsaturated carboxylic acid compound.
- a production method comprising the step of introducing an amino acid substitution of at least one of the following (1) to (5) in phenylalanine ammonia-lyase. (1) Substitute the amino acid at position 108 of the amino acid sequence shown in SEQ ID NO: 2 or the amino acid corresponding to the site with methionine, phenylalanine or valine.
- the "phenylalanine ammonia-lyase variant having enhanced catalytic activity for producing a third chain unsaturated carboxylic acid compound from a second chain unsaturated carboxylic acid compound” means that the amino acid substitution is introduced. This means phenylalanine ammonia-lyase having a higher catalytic activity to produce a third chain unsaturated carboxylic acid compound as compared with that before the introduction, and the comparison target is usually phenylalanine ammonia-lyase derived from the above-mentioned various organisms and the above-mentioned phenylalanine ammonia-lyase. It is its natural variant.
- “Introduction of amino acid substitution” in phenylalanine ammonia-lyase can be performed by modifying the encoding DNA. As described above, such "modification of DNA” is appropriately carried out by a method known to those skilled in the art, for example, a site-directed mutagenesis method or a chemical synthesis method of DNA based on the modified sequence information. It is possible to do. Further, as described above, "introduction of amino acid substitution” can also be performed by using a method for chemically synthesizing a peptide.
- the phenylalanine ammonia-lyase variant thus produced has a catalytic activity for producing a third chain unsaturated carboxylic acid compound with respect to the phenylalanine ammonia-lyase having the amino acid sequence set forth in SEQ ID NO: 2.
- 1.2 times or more (1.3 times or more, 1.4 times or more, 1.5 times or more, 1.6 times or more, 1.7 times or more, 1.8 times or more, 1.9 times or more) It is preferably 2 times or more, more preferably 3 times or more, further preferably 4 times or more, still more preferably 5 times or more.
- the present inventors conceived the following reaction scheme in which butadiene is produced using L-lysine as a starting material in a three-step reaction using three kinds of enzymes (BesC, PAL, FDC).
- Example 1 Preparation of plasmid vector> First, in order to efficiently express FDC derived from Pseudomonas fluorescens, PAL derived from Arabidopsis thaliana, and FDC derived from Saccharomyces cerevisiae in Escherichia coli, a nucleotide sequence modified in consideration of the frequency of codon use in Escherichia coli was designed. Then, the DNA consisting of such a modified nucleotide sequence was chemically synthesized according to a conventional method.
- the DNA thus prepared and the pET22b (+) vector are ligated by the Gibson Assembly method (registered by the Gibson Associates kit NEBliderHiFi DNA Secembury Master Mix (registered trademark).
- the various wild-type genes were prepared as plasmid vectors (BesC vector, PAL vector, FDC vector 1) that can be expressed in Escherichia coli.
- a DNA obtained by amplifying a gene (SEQ ID NO: 15) encoding a flavinprenyl transferase (hereinafter, also referred to as “UbiX”) from an Escherichia coli (K-12) strain by a PCR method and a pColADute vector (manufactured by Novagen) were obtained.
- a plasmid vector (UbiX vector) capable of expressing the wild type UbiX in Escherichia coli was prepared.
- the vector prepared as described above (5 ⁇ g BesC vector or 5 ⁇ g PAL vector) was introduced into Escherichia coli C41 (DE3) strain (Lucien Corporation, 100 ⁇ L) by the heat shock method to obtain wild-type BesC or PAL.
- a transformant to be expressed was prepared. Then, each of these transformants was cultured in LB medium supplemented with ampicillin for 6 hours. By culturing for 6 hours (preculture), the proliferation of these transformants has reached a plateau. Therefore, the amount of cells at the start of the enzyme reaction described later becomes uniform among these transformants.
- the prepared vectors (5 ⁇ g FDC vector 1 and 5 ⁇ g UbiX vector) were introduced into Escherichia coli C41 (DE3) strain (Lucien Corporation, 100 ⁇ L) by the heat shock method, and wild-type FDC and UbiX were introduced.
- a transformant co-expressing with was prepared.
- the cells were cultured in LB medium supplemented with ampicillin and kanamycin for 6 hours.
- the above-mentioned enzyme reaction medium to which 50 mg / L kanamycin was added was prepared, and a cytolytic solution was prepared in the same manner.
- Example 2 As shown in Example 1, the production of butadiene from L-lysine was observed by the combination of various wild-type enzymes. Therefore, in order to further improve this catalytic activity, PALs derived from other hosts were examined for PALs to be subjected to the second stage reaction.
- plasmid vectors (various PAL vectors) capable of expressing the various field genes in Escherichia coli were prepared in the same manner as described above.
- the prepared vector (5 ⁇ g PAL vector, 5 ⁇ g UbiX vector and 5 ⁇ g FDC vector 2) was introduced into Escherichia coli C41 (DE3) strain (Lucien Corporation, 100 ⁇ L) by the heat shock method, and various PALs and wild type were introduced. A transformant co-expressing FDC and UbiX was prepared.
- each of these transformants was cultured for 6 hours in LB medium supplemented with ampicillin, kanamycin, and chloramphenicol. By culturing for 6 hours (preculture), the proliferation of these transformants has reached a plateau. Therefore, the amount of cells at the start of the enzyme reaction described later becomes uniform among these transformants.
- butadiene was also produced from all PALs even when allylglycine was added as a substrate.
- allylglycine was added as a substrate.
- Example 3 As shown in Example 2, high butadiene formation catalytic activity was observed in wild-type AvPAL. Therefore, in order to further improve this high catalytic activity, a mutation was introduced in which the enzyme active site was replaced with another amino acid. Then, the catalytic activity of the obtained mutant introducer was evaluated.
- a primer that encodes the amino acid sequence into which each mutation was introduced. Then, using the vector encoding the wild-type AvPAL prepared in Example 2 as a template and using the above-mentioned primers, a plasmid vector into which each mutation is introduced can be expressed in Escherichia coli according to the protocol of the Gibson Assembly method. (PAL variant vector) was prepared. Then, in the above ⁇ Preparation of medium for measuring enzyme activity and measurement of enzyme activity>, a PAL variant vector was introduced instead of the wild-type PAL vector, a transformant was prepared, and the enzyme activity was measured. The butadiene production amount when the wild-type PAL is used is 1.0, and the butadiene production amount when the PAL variant is used is shown in Table 3 below.
- the present invention it is possible to provide a method for producing an unsaturated compound having at least two carbon-carbon double bonds using an enzyme.
- the present invention also makes it possible to produce butadiene using relatively inexpensive L-lysine as a starting material.
- an unsaturated compound can be produced by biosynthesis without using chemical synthesis, so that the burden on the environment is small. Therefore, the present invention is extremely useful in the production of raw materials for various synthetic polymers such as synthetic rubber such as butadiene.
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Zoology (AREA)
- Health & Medical Sciences (AREA)
- Wood Science & Technology (AREA)
- Genetics & Genomics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Biotechnology (AREA)
- Microbiology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Biomedical Technology (AREA)
- General Chemical & Material Sciences (AREA)
- Molecular Biology (AREA)
- Medicinal Chemistry (AREA)
- Biophysics (AREA)
- Plant Pathology (AREA)
- Physics & Mathematics (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
- Enzymes And Modification Thereof (AREA)
Abstract
Description
(1)108位のロイシンを、メチオニン、フェニルアラニン又はバリンに置換、
(2)107位のフェニルアラニンを、トリプトファンに置換、
(3)219位のロイシンを、イソロイシンに置換、
(4)223位のアスパラギンを、イソロイシンに置換、
(5)104位のロイシンを、アラニンに置換。
[1] フェニルアラニンアンモニアリアーゼの存在下、第1のアミノ基と末端に第1の炭素-炭素間二重結合とを有する、下記式(2)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体から、第1のアミノ基を脱離させ、第2の炭素-炭素間二重結合を形成させる工程を含む、下記式(3)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体を製造する方法
[2] 末端アルケン生成酵素 BesCの存在下、第1のアミノ基と末端に第2のアミノ基とを有する、下記式(1)で表される鎖状のカルボン酸化合物又はその幾何異性体から、第2のアミノ基及びメチレン基を脱離させ、前記式(2)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体を製造する工程を含み、当該化合物又はその幾何異性体から、前記式(3)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体を製造する、[1]に記載の方法
[3] [1]又は[2]に記載の方法により、前記式(3)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体を製造し、フェルラ酸デカルボキシラーゼの存在下、前記不飽和カルボン酸化合物又はその幾何異性体から、カルボキシル基を脱離させる工程を含む、下記式(4)で表される鎖状の不飽和炭化水素化合物又はその幾何異性体を製造する方法
[4] 前記フェニルアラニンアンモニアリアーゼが、下記(1)~(5)のうちの少なくとも1の特徴を有するフェニルアラニンアンモニアリアーゼである、[1]~[3]のうちのいずれか一項に記載の方法
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸が、メチオニン、フェニルアラニン若しくはバリンである、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸が、トリプトファンである、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸が、イソロイシンである、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸が、イソロイシンである、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸が、アラニンである。
[5] 下記(1)~(5)のうちの少なくとも1のアミノ酸置換が導入されている、フェニルアラニンアンモニアリアーゼ改変体
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸が、メチオニン、フェニルアラニン若しくはバリンに置換、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸が、トリプトファンに置換、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸が、イソロイシンに置換、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸が、イソロイシンに置換、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸が、アラニンに置換。
[6] [5]に記載のフェニルアラニンアンモニアリアーゼ改変体をコードするDNA。
[7] [6]に記載のDNAを含むベクター。
[8] [6]に記載のDNA又は[7]に記載のベクターが導入された宿主細胞。
[9] [8]に記載の宿主細胞を培養し、該宿主細胞に発現したタンパク質を採取する工程を含む、フェニルアラニンアンモニアリアーゼ改変体の製造方法。
[10] フェニルアラニンアンモニアリアーゼ改変体の製造方法であって、フェニルアラニンアンモニアリアーゼにおいて、下記(1)~(5)のうちの少なくとも1のアミノ酸置換を導入する工程を含む、方法
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸を、メチオニン、フェニルアラニン若しくはバリンに置換、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸を、トリプトファンに置換、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸を、イソロイシンに置換、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸を、イソロイシンに置換、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸を、アラニンに置換。
前記のとおり、本発明は、フェニルアラニンアンモニアリアーゼの存在下、第1のアミノ基と末端に第1の炭素-炭素間二重結合とを有する、前記式(2)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体(第2の鎖状の不飽和カルボン酸化合物)から、第1のアミノ基を脱離させ、第2の炭素-炭素間二重結合を形成させる工程を含む、前記式(3)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体(第3の鎖状の不飽和カルボン酸化合物)を製造する方法を、提供するものである。
「フェニルアラニンアンモニアリアーゼ」とは、EC番号:4.3.1.24として登録されている酵素であり、フェニルアラニンを基質とし、ケイ皮酸とアンモニアを生成する反応を、触媒する酵素を意味する。また、PAL、チラーゼ、フェニルアラニンデアミナーゼ、チロシンアンモニアリアーゼ、L-チロシンアンモニアリアーゼ、フェニルアラニンアンモニウムリアーゼ、L-フェニルアラニンアンモニアリアーゼとも称される酵素である。
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸が、メチオニン、フェニルアラニン又はバリンである、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸が、トリプトファンである、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸が、イソロイシンである、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸が、イソロイシンである、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸が、アラニンである。
(a)配列番号:2に記載のアミノ酸配列において、下記(1)~(5)のうちの少なくとも1のアミノ酸置換が導入されているアミノ酸配列を含む、フェニルアラニンアンモニアリアーゼ変異体
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸が、メチオニン、フェニルアラニン若しくはバリンに置換、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸が、トリプトファンに置換、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸が、イソロイシンに置換、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸が、イソロイシンに置換、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸が、アラニンに置換。
(b)配列番号:2に記載のアミノ酸配列において、前記(1)~(5)のうちの少なくとも1のアミノ酸置換が導入され、更に、前記部位以外の1若しくは数個の部位にて、1又は複数のアミノ酸が置換、欠失、付加、及び/又は挿入されているアミノ酸配列を含む、フェニルアラニンアンモニアリアーゼ変異体。
(c)配列番号:2に記載のアミノ酸配列と少なくとも15%の同一性を有するアミノ酸配列であって、前記(1)~(5)のうちの少なくとも1のアミノ酸置換が導入されているアミノ酸配列を含む、フェニルアラニンアンモニアリアーゼ変異体。
後述の実施例に示すとおり、本発明者らは、上述の第3の鎖状の不飽和カルボン酸化合物製造の原料となる、第2の鎖状の不飽和カルボン酸化合物も、下記反応に示すとおり、生成できることを見出している。
「BesC」とは、β-エチニルセリン生合成(Bes)に関する1の酵素である。当該酵素は、末端のプロピルアミノ基を酸化することにより、当該基中の炭素-炭素間結合を開裂し、末端に炭素-炭素間二重結合を形成させる反応を、触媒する活性を有する、末端アルケン生成酵素である(非特許文献2)。
後述の実施例に示すとおり、本発明者らは、上述の第3の鎖状の不飽和カルボン酸化合物から、両末端に炭素-炭素間二重結合を有する鎖状の不飽和炭化水素化合物を、生成できることも見出している。
「フェルラ酸デカルボキシラーゼ」とは、EC番号:4.1.1.102として登録されている酵素であり、通常、フェルラ酸を脱炭酸して4-ビニルグアイヤコール(4VG)を生成する反応を、触媒する酵素を意味する。
次に、本発明にかかる酵素(本発明にかかるフェニルアラニンアンモニアリアーゼ、本発明にかかるBesC、本発明にかかるフェルラ酸デカルボキシラーゼ)をコードするDNA等について説明する。かかるDNAを導入することによって、宿主細胞の形質を転換し、本発明にかかる各種酵素を当該細胞において製造させること、ひいては第3の鎖状の不飽和カルボン酸化合物、又は鎖状の不飽和炭化水素化合物を製造させることが可能となる。
本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAが挿入されているベクター、
本発明にかかるBesCをコードするDNAが挿入されているベクターと、本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAが挿入されているベクターとの組み合わせ、
本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAが挿入されているベクターと、本発明にかかるフェルラ酸デカルボキシラーゼをコードするDNAが挿入されているベクターとの組み合わせ、又は
本発明にかかるBesCをコードするDNAが挿入されているベクターと、本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAが挿入されているベクターと、本発明にかかるフェルラ酸デカルボキシラーゼをコードするDNAが挿入されているベクターとの組み合わせが、挙げられる。
また、本発明にかかるBesCをコードするDNAと、本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAとが挿入されているベクター、
本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAと、本発明にかかるフェルラ酸デカルボキシラーゼとが挿入されているベクター、又は
本発明にかかるBesCをコードするDNAと、本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAと、本発明にかかるフェルラ酸デカルボキシラーゼとが挿入されているベクターも、挙げることができる。
上述のとおり、本発明にかかる酵素、該酵素をコードするDNA又は該DNAが挿入されているベクターを用いることにより、第3の鎖状の不飽和カルボン酸化合物、又は鎖状の不飽和炭化水素化合物の生成を促進することが可能となる。
本発明にかかるフェニルアラニンアンモニアリアーゼ、該酵素をコードするDNA又は該DNAが挿入されているベクターを含む、第2の鎖状の不飽和カルボン酸化合物から第3の鎖状の不飽和カルボン酸化合物の生成を促進するための剤、
本発明にかかるBesC、該酵素をコードするDNA又は該DNAが挿入されているベクターと、本発明にかかるフェニルアラニンアンモニアリアーゼ、該酵素をコードするDNA又は該DNAが挿入されているベクターとを含む、第1の鎖状の不飽和カルボン酸化合物から第3の鎖状の不飽和カルボン酸化合物の生成を促進するための剤、
本発明にかかるBesCをコードするDNAと、本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAとが挿入されているベクターを含む、第1の鎖状の不飽和カルボン酸化合物から第3の鎖状の不飽和カルボン酸化合物の生成を促進するための剤、
本発明にかかるフェニルアラニンアンモニアリアーゼ、該酵素をコードするDNA又は該DNAが挿入されているベクターと、本発明にかかるフェルラ酸デカルボキシラーゼ、該酵素をコードするDNA又は該DNAが挿入されているベクターとを含む、第2の鎖状の不飽和カルボン酸化合物から鎖状の不飽和炭化水素化合物の生成を促進するための剤、
本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAと、本発明にかかるフェルラ酸デカルボキシラーゼをコードするDNAとが挿入されているベクターを含む、第2の鎖状の不飽和カルボン酸化合物から鎖状の不飽和炭化水素化合物の生成を促進するための剤、
本発明にかかるBesC、該酵素をコードするDNA又は該DNAが挿入されているベクターと、本発明にかかるフェニルアラニンアンモニアリアーゼ、該酵素をコードするDNA又は該DNAが挿入されているベクターと、本発明にかかるフェルラ酸デカルボキシラーゼ、該酵素をコードするDNA又は該DNAが挿入されているベクターとを含む、第1の鎖状の不飽和カルボン酸化合物から鎖状の不飽和炭化水素化合物の生成を促進するための剤、
本発明にかかるBesCをコードするDNAと、本発明にかかるフェニルアラニンアンモニアリアーゼをコードするDNAと、本発明にかかるフェルラ酸デカルボキシラーゼをコードするDNAとが挿入されているベクターを含む、第1の鎖状の不飽和カルボン酸化合物から鎖状の不飽和炭化水素化合物の生成を促進するための剤。
次に、本発明にかかるDNA又はベクターが導入された宿主細胞について説明する。前述のDNA又はベクターの導入によって形質転換された宿主細胞を用いれば、本発明にかかる酵素を製造することが可能となり、ひいては、第3の鎖状の不飽和カルボン酸化合物、又は鎖状の不飽和炭化水素化合物を製造させることも可能となる。
上述のとおり、本発明にかかる酵素を発現するように形質転換された宿主細胞を、培養することにより、第3の鎖状の不飽和カルボン酸化合物又は鎖状の不飽和炭化水素化合物を製造することができる。したがって、本発明においては、本発明にかかる酵素をコードするDNA又はベクターが導入された宿主細胞を培養し、該宿主細胞及び/又はその培養物において生成された、第3の鎖状の不飽和カルボン酸化合物又は鎖状の不飽和炭化水素化合物を採取する工程を含む、第3の鎖状の不飽和カルボン酸化合物又は鎖状の不飽和炭化水素化合物の製造方法も提供される。
後述の実施例に示す通り、本発明にかかるフェニルアラニンアンモニアリアーゼ改変体をコードするDNA等が導入された宿主細胞を培養することにより、該宿主細胞内にて当該改変体を製造することができる。
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸を、メチオニン、フェニルアラニン若しくはバリンに置換、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸を、トリプトファンに置換、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸を、イソロイシンに置換、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸を、イソロイシンに置換、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸を、アラニンに置換。
<プラスミドベクターの調製>
先ず、Pseudomonas fluorescens由来BesC、Arabidopsis thaliana由来のPAL、そしてSaccharomyces cerevisiae由来のFDCを大腸菌にて効率良く発現させるために、大腸菌におけるコドンの使用頻度を考慮して改変したヌクレオチド配列を設計した。次いで、かかる改変ヌクレオチド配列からなるDNAを常法に沿って化学合成した。そして、このようにして調製したDNAとpET22b(+)ベクター(Novagen社製)を、Gibson Assembly法(New England Biolabs社のキットNEBuilder HiFi DNA Assembly Master Mix(登録商標)を使用)により連結することによって、当該各種野生型遺伝子を大腸菌において各々発現可能なプラスミドベクター(BesCベクター、PALベクター、FDCベクター1)として調製した。同様にして、大腸菌(K-12)株からフラビンプレニルトランスフェラーゼ(以下「UbiX」とも称する)をコードする遺伝子(配列番号:15)をPCR法により増幅したDNAとpColADuetベクター(Novagen社製)を、Gibson Assembly法により連結することにより、当該野生型のUbiXを大腸菌において発現可能なプラスミドベクター(UbiXベクター)を調製した。
前記のとおり調製したベクター(5μgのBesCベクター、又は5μgのPALベクター)を、大腸菌C41(DE3)株(Lucigen Corporation社製、100μL)に、ヒートショック法により導入し、野生型のBesC又はPALを発現する形質転換体を調製した。そして、これら形質転換体を各々、アンピシリンを添加したLB培地にて6時間培養した。なお、かかる6時間の培養(前培養)により、これら形質転換体の増殖は頭打ちとなる。そのため、後述の酵素反応開始時点での菌体量は、これら形質転換体間において均一となる。
実施例1に示したように、各種野生型酵素の組み合わせによって、L-リジンからのブタジエンの生成が認められた。そこで、この触媒活性をより向上させるべく、二段階目の反応を行うPALについて、他の宿主由来のPALを検討した。
Anabaena variabilis又はPlagiochasma appendiculatum由来のPALを大腸菌にて効率良く発現させるために、大腸菌におけるコドンの使用頻度を考慮して改変したヌクレオチド配列を設計した。次いで、かかる改変ヌクレオチド配列からなるDNAを常法に沿って化学合成し、前述と同様に当該各種野遺伝子を、大腸菌において発現可能なプラスミドベクター(各種PALベクター)を各々調製した。
調整したベクター(5μgのPALベクター、5μgのUbiXベクターと5μgのFDCベクター2)を、大腸菌C41(DE3)株(Lucigen Corporation社製、100μL)に、ヒートショック法により導入し、各種PALと野生型のFDCとUbiXとを共発現する形質転換体を調製した。
実施例2に示したように、野生型のAvPALにおいて、高いブタジエンの生成触媒活性が認められた。そこで、この高い触媒活性をより向上させるべく、酵素活性部位を各々他のアミノ酸に置換する変異を導入した。そして、得られた変異導入体について、前記触媒活性を評価した。
(1)108位のロイシンが、メチオニン、フェニルアラニン又はバリンに置換、
(2)107位のフェニルアラニンが、トリプトファンに置換、
(3)219位のロイシンが、イソロイシンに置換、
(4)223位のアスパラギンが、イソロイシンに置換、
(5)104位のロイシンが、アラニンに置換。
Claims (10)
- フェニルアラニンアンモニアリアーゼの存在下、第1のアミノ基と末端に第1の炭素-炭素間二重結合とを有する、下記式(2)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体から、第1のアミノ基を脱離させ、第2の炭素-炭素間二重結合を形成させる工程を含む、下記式(3)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体を製造する方法
- 末端アルケン生成酵素 BesCの存在下、第1のアミノ基と末端に第2のアミノ基とを有する、下記式(1)で表される鎖状のカルボン酸化合物又はその幾何異性体から、第2のアミノ基及びメチレン基を脱離させ、前記式(2)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体を製造する工程を含み、当該化合物又はその幾何異性体から、前記式(3)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体を製造する、請求項1に記載の方法
- 請求項1又は2に記載の方法により、前記式(3)で表される鎖状の不飽和カルボン酸化合物又はその幾何異性体を製造し、フェルラ酸デカルボキシラーゼの存在下、前記不飽和カルボン酸化合物又はその幾何異性体から、カルボキシル基を脱離させる工程を含む、下記式(4)で表される鎖状の不飽和炭化水素化合物又はその幾何異性体を製造する方法
- 前記フェニルアラニンアンモニアリアーゼが、下記(1)~(5)のうちの少なくとも1の特徴を有するフェニルアラニンアンモニアリアーゼである、請求項1~3のうちのいずれか一項に記載の方法
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸が、メチオニン、フェニルアラニン若しくはバリンである、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸が、トリプトファンである、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸が、イソロイシンである、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸が、イソロイシンである、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸が、アラニンである。 - 下記(1)~(5)のうちの少なくとも1のアミノ酸置換が導入されている、フェニルアラニンアンモニアリアーゼ改変体
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸が、メチオニン、フェニルアラニン若しくはバリンに置換、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸が、トリプトファンに置換、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸が、イソロイシンに置換、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸が、イソロイシンに置換、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸が、アラニンに置換。 - 請求項5に記載のフェニルアラニンアンモニアリアーゼ改変体をコードするDNA。
- 請求項6に記載のDNAを含むベクター。
- 請求項6に記載のDNA又は請求項7に記載のベクターが導入された宿主細胞。
- 請求項8に記載の宿主細胞を培養し、該宿主細胞に発現したタンパク質を採取する工程を含む、フェニルアラニンアンモニアリアーゼ改変体の製造方法。
- フェニルアラニンアンモニアリアーゼ改変体の製造方法であって、フェニルアラニンアンモニアリアーゼにおいて、下記(1)~(5)のうちの少なくとも1のアミノ酸置換を導入する工程を含む、方法
(1)配列番号:2に記載のアミノ酸配列の108位又は該部位に対応するアミノ酸を、メチオニン、フェニルアラニン若しくはバリンに置換、
(2)配列番号:2に記載のアミノ酸配列の107位又は該部位に対応するアミノ酸を、トリプトファンに置換、
(3)配列番号:2に記載のアミノ酸配列の219位又は該部位に対応するアミノ酸を、イソロイシンに置換、
(4)配列番号:2に記載のアミノ酸配列の223位又は該部位に対応するアミノ酸を、イソロイシンに置換、
(5)配列番号:2に記載のアミノ酸配列の104位又は該部位に対応するアミノ酸を、アラニンに置換。
Priority Applications (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP21915039.8A EP4269593A1 (en) | 2020-12-28 | 2021-12-07 | Method for producing chain unsaturated carboxylic acid compound using phenylalanine ammonia lyase |
| JP2022572955A JPWO2022145178A1 (ja) | 2020-12-28 | 2021-12-07 | |
| CN202180087826.4A CN116710549A (zh) | 2020-12-28 | 2021-12-07 | 使用苯丙氨酸氨裂合酶的链状不饱和羧酸化合物的制造方法 |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2020-219017 | 2020-12-28 | ||
| JP2020219017 | 2020-12-28 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2022145178A1 true WO2022145178A1 (ja) | 2022-07-07 |
Family
ID=82260391
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2021/044890 WO2022145178A1 (ja) | 2020-12-28 | 2021-12-07 | フェニルアラニンアンモニアリアーゼを用いた鎖状の不飽和カルボン酸化合物の製造方法 |
Country Status (4)
| Country | Link |
|---|---|
| EP (1) | EP4269593A1 (ja) |
| JP (1) | JPWO2022145178A1 (ja) |
| CN (1) | CN116710549A (ja) |
| WO (1) | WO2022145178A1 (ja) |
Citations (8)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20130005012A1 (en) * | 2011-06-23 | 2013-01-03 | Phytogene, Inc. | Enzymatic system for monomer synthesis |
| JP2014030376A (ja) | 2012-08-02 | 2014-02-20 | Mitsubishi Chemicals Corp | ブタジエンの製造方法 |
| US20150337336A1 (en) * | 2012-06-22 | 2015-11-26 | Phytogene, Inc. | Enzymes and methods for styrene synthesis |
| JP2015228804A (ja) | 2014-06-03 | 2015-12-21 | 三菱化学株式会社 | 2,3−ブタンジオール生産能を有する微生物およびそれを用いた2,3−ブタンジオールの製造方法、1,3−ブタジエンの製造方法 |
| US20160230194A1 (en) * | 2013-09-18 | 2016-08-11 | Arizona Board Of Regents On Behalf Of Arizona State University | Enzymes and Microorganisms for the Production of 1,3-Butadiene and Other Dienes |
| WO2019022083A1 (ja) | 2017-07-24 | 2019-01-31 | 国立研究開発法人理化学研究所 | デカルボキシラーゼ、及びそれを用いた不飽和炭化水素化合物の製造方法 |
| WO2020013951A1 (en) * | 2018-07-12 | 2020-01-16 | Codexis, Inc. | Engineered phenylalanine ammonia lyase polypeptides |
| US20200048674A1 (en) * | 2017-04-24 | 2020-02-13 | The Regents Of The University Of California | Biosynthesis of bioorthogonal amino acids |
-
2021
- 2021-12-07 JP JP2022572955A patent/JPWO2022145178A1/ja active Pending
- 2021-12-07 CN CN202180087826.4A patent/CN116710549A/zh active Pending
- 2021-12-07 WO PCT/JP2021/044890 patent/WO2022145178A1/ja active Application Filing
- 2021-12-07 EP EP21915039.8A patent/EP4269593A1/en active Pending
Patent Citations (8)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20130005012A1 (en) * | 2011-06-23 | 2013-01-03 | Phytogene, Inc. | Enzymatic system for monomer synthesis |
| US20150337336A1 (en) * | 2012-06-22 | 2015-11-26 | Phytogene, Inc. | Enzymes and methods for styrene synthesis |
| JP2014030376A (ja) | 2012-08-02 | 2014-02-20 | Mitsubishi Chemicals Corp | ブタジエンの製造方法 |
| US20160230194A1 (en) * | 2013-09-18 | 2016-08-11 | Arizona Board Of Regents On Behalf Of Arizona State University | Enzymes and Microorganisms for the Production of 1,3-Butadiene and Other Dienes |
| JP2015228804A (ja) | 2014-06-03 | 2015-12-21 | 三菱化学株式会社 | 2,3−ブタンジオール生産能を有する微生物およびそれを用いた2,3−ブタンジオールの製造方法、1,3−ブタジエンの製造方法 |
| US20200048674A1 (en) * | 2017-04-24 | 2020-02-13 | The Regents Of The University Of California | Biosynthesis of bioorthogonal amino acids |
| WO2019022083A1 (ja) | 2017-07-24 | 2019-01-31 | 国立研究開発法人理化学研究所 | デカルボキシラーゼ、及びそれを用いた不飽和炭化水素化合物の製造方法 |
| WO2020013951A1 (en) * | 2018-07-12 | 2020-01-16 | Codexis, Inc. | Engineered phenylalanine ammonia lyase polypeptides |
Non-Patent Citations (5)
| Title |
|---|
| "UNIPROT", Database accession no. A2QHE5 |
| HO, S. N.HUNT, H. D.HORTON, R. M.PULLEN, J. K.PEASE, L. R., GENE, vol. 77, 1989, pages 51 - 59 |
| J A MARCHAND ET AL., NATURE, vol. 567, 2019 |
| KARL A. P. PAYNE ET AL., NATURE, vol. 522, 2015, pages 497 - 501 |
| KUNKEL, T. A., PROC NATL ACAD SCI, vol. 82, 1985, pages 488 - 492 |
Also Published As
| Publication number | Publication date |
|---|---|
| EP4269593A1 (en) | 2023-11-01 |
| JPWO2022145178A1 (ja) | 2022-07-07 |
| CN116710549A (zh) | 2023-09-05 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11142778B2 (en) | Decarboxylase and method for producing unsaturated hydrocarbon compound using same | |
| JP5590649B2 (ja) | 変異体ピロリジル−tRNA合成酵素及びこれを用いる非天然アミノ酸組み込みタンパク質の製造方法 | |
| US20220348974A1 (en) | Biotin synthases for efficient production of biotin | |
| JP2023120401A (ja) | N-アシル-アミノ基含有化合物の製造方法 | |
| WO2023190564A1 (ja) | メタクリル酸の製造方法 | |
| WO2022145178A1 (ja) | フェニルアラニンアンモニアリアーゼを用いた鎖状の不飽和カルボン酸化合物の製造方法 | |
| WO2021054441A1 (ja) | サッカロミケス由来フェルラ酸デカルボキシラーゼ変異体、及びそれを用いた不飽和炭化水素化合物の製造方法 | |
| Wang et al. | Bioconversion of recombinantly produced precursor peptide pqqA into pyrroloquinoline quinone (PQQ) using a cell-free in vitro system | |
| JP6803047B2 (ja) | ジホスホメバロン酸デカルボキシラーゼ変異体、及びそれを用いたオレフィン化合物の製造方法 | |
| CN110234761B (zh) | 二磷酸甲羟戊酸脱羧酶变异体、和利用了该变异体的烯烃化合物的制造方法 | |
| JP6995315B2 (ja) | ジホスホメバロン酸デカルボキシラーゼ変異体、及びそれを用いたオレフィン化合物の製造方法 | |
| WO2024090440A1 (ja) | フェルラ酸デカルボキシラーゼ、及びそれを用いた不飽和炭化水素化合物の製造方法 | |
| JP6635535B1 (ja) | Efpタンパク質を発現する大腸菌およびそれを用いたフラボノイド化合物製造方法 | |
| JP2024517485A (ja) | フェニルプロパノイド化合物の生合成 | |
| Zhou et al. | Exploring, Cloning, and Characterization of Aryl-alcohol Dehydrogenase for β-Phenylethanol Production in Paocai | |
| WO2023150533A1 (en) | Microorganisms and methods for producing menaquinone-7 | |
| JP2023527107A (ja) | シスチンを生体触媒還元するための酵素的触媒酸化還元系のコア構成成分としての生体触媒 | |
| JP2009131254A (ja) | 4−ヒドロキシイソロイシン又は2−アミノ−3−メチル−4−ケトペンタン酸の製造法 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 21915039 Country of ref document: EP Kind code of ref document: A1 |
|
| ENP | Entry into the national phase |
Ref document number: 2022572955 Country of ref document: JP Kind code of ref document: A |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 202180087826.4 Country of ref document: CN |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2021915039 Country of ref document: EP Effective date: 20230728 |