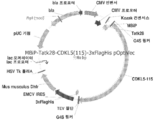
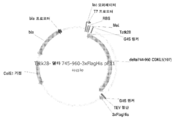
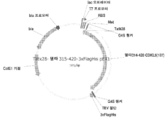
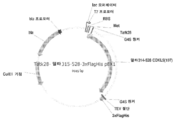
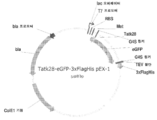
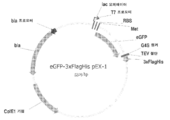
KR20200090889A - Cdkl5 발현 변이체 및 cdkl5 융합 단백질 - Google Patents
Cdkl5 발현 변이체 및 cdkl5 융합 단백질 Download PDFInfo
- Publication number
- KR20200090889A KR20200090889A KR1020207018589A KR20207018589A KR20200090889A KR 20200090889 A KR20200090889 A KR 20200090889A KR 1020207018589 A KR1020207018589 A KR 1020207018589A KR 20207018589 A KR20207018589 A KR 20207018589A KR 20200090889 A KR20200090889 A KR 20200090889A
- Authority
- KR
- South Korea
- Prior art keywords
- ser
- leu
- lys
- pro
- glu
- Prior art date
Links
- 102100034746 Cyclin-dependent kinase-like 5 Human genes 0.000 title claims abstract description 164
- 108020001507 fusion proteins Proteins 0.000 title claims abstract description 144
- 102000037865 fusion proteins Human genes 0.000 title claims abstract description 143
- 101000945692 Homo sapiens Cyclin-dependent kinase-like 5 Proteins 0.000 title claims abstract description 63
- 230000014509 gene expression Effects 0.000 title description 14
- 108090000765 processed proteins & peptides Proteins 0.000 claims abstract description 113
- 102000004196 processed proteins & peptides Human genes 0.000 claims abstract description 109
- 229920001184 polypeptide Polymers 0.000 claims abstract description 103
- 238000000034 method Methods 0.000 claims abstract description 32
- 239000008194 pharmaceutical composition Substances 0.000 claims abstract description 6
- 208000006289 Rett Syndrome Diseases 0.000 claims description 25
- 241000588724 Escherichia coli Species 0.000 claims description 24
- 230000007812 deficiency Effects 0.000 claims description 23
- 239000000203 mixture Substances 0.000 claims description 23
- 230000035772 mutation Effects 0.000 claims description 21
- 241000282414 Homo sapiens Species 0.000 claims description 14
- 238000009472 formulation Methods 0.000 claims description 13
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 claims description 10
- 208000005849 atypical Rett syndrome Diseases 0.000 claims description 10
- 108091033319 polynucleotide Proteins 0.000 claims description 10
- 102000040430 polynucleotide Human genes 0.000 claims description 10
- 239000002157 polynucleotide Substances 0.000 claims description 10
- 208000012902 Nervous system disease Diseases 0.000 claims description 9
- 230000001404 mediated effect Effects 0.000 claims description 8
- 239000013598 vector Substances 0.000 claims description 7
- 239000003937 drug carrier Substances 0.000 claims description 5
- 238000004519 manufacturing process Methods 0.000 claims description 4
- 241000699802 Cricetulus griseus Species 0.000 claims description 3
- 238000007913 intrathecal administration Methods 0.000 claims description 3
- 210000003734 kidney Anatomy 0.000 claims description 3
- 210000001672 ovary Anatomy 0.000 claims description 3
- 108010008281 Recombinant Fusion Proteins Proteins 0.000 abstract description 19
- 102000007056 Recombinant Fusion Proteins Human genes 0.000 abstract description 19
- 238000011282 treatment Methods 0.000 abstract description 12
- 101710178912 Cyclin-dependent kinase-like 5 Proteins 0.000 description 151
- 108010034529 leucyl-lysine Proteins 0.000 description 73
- KDXKERNSBIXSRK-UHFFFAOYSA-N Lysine Natural products NCCCCC(N)C(O)=O KDXKERNSBIXSRK-UHFFFAOYSA-N 0.000 description 69
- 108010050848 glycylleucine Proteins 0.000 description 58
- SBANPBVRHYIMRR-UHFFFAOYSA-N Leu-Ser-Pro Natural products CC(C)CC(N)C(=O)NC(CO)C(=O)N1CCCC1C(O)=O SBANPBVRHYIMRR-UHFFFAOYSA-N 0.000 description 57
- 210000004027 cell Anatomy 0.000 description 55
- VPZXBVLAVMBEQI-UHFFFAOYSA-N glycyl-DL-alpha-alanine Natural products OC(=O)C(C)NC(=O)CN VPZXBVLAVMBEQI-UHFFFAOYSA-N 0.000 description 49
- 108010026333 seryl-proline Proteins 0.000 description 47
- YBAFDPFAUTYYRW-UHFFFAOYSA-N N-L-alpha-glutamyl-L-leucine Natural products CC(C)CC(C(O)=O)NC(=O)C(N)CCC(O)=O YBAFDPFAUTYYRW-UHFFFAOYSA-N 0.000 description 44
- 108010093581 aspartyl-proline Proteins 0.000 description 43
- 241000880493 Leptailurus serval Species 0.000 description 40
- 239000013612 plasmid Substances 0.000 description 39
- 108010018006 histidylserine Proteins 0.000 description 35
- WGLAORUKDGRINI-WDCWCFNPSA-N Lys-Glu-Thr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O WGLAORUKDGRINI-WDCWCFNPSA-N 0.000 description 34
- KOSRFJWDECSPRO-UHFFFAOYSA-N alpha-L-glutamyl-L-glutamic acid Natural products OC(=O)CCC(N)C(=O)NC(CCC(O)=O)C(O)=O KOSRFJWDECSPRO-UHFFFAOYSA-N 0.000 description 32
- 108090000623 proteins and genes Proteins 0.000 description 32
- 108010085325 histidylproline Proteins 0.000 description 30
- 102000004169 proteins and genes Human genes 0.000 description 30
- NTBOEZICHOSJEE-YUMQZZPRSA-N Gly-Lys-Ser Chemical compound [H]NCC(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(O)=O NTBOEZICHOSJEE-YUMQZZPRSA-N 0.000 description 28
- AEMPCGRFEZTWIF-IHRRRGAJSA-N Val-Leu-Lys Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(O)=O AEMPCGRFEZTWIF-IHRRRGAJSA-N 0.000 description 28
- 108010051109 Cell-Penetrating Peptides Proteins 0.000 description 27
- 102000020313 Cell-Penetrating Peptides Human genes 0.000 description 27
- QVOGDCQNGLBNCR-FXQIFTODSA-N Ser-Arg-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(O)=O QVOGDCQNGLBNCR-FXQIFTODSA-N 0.000 description 27
- YRBGKVIWMNEVCZ-WDSKDSINSA-N Ser-Glu-Gly Chemical compound OC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O YRBGKVIWMNEVCZ-WDSKDSINSA-N 0.000 description 26
- 108010045023 alanyl-prolyl-tyrosine Proteins 0.000 description 26
- 108010047857 aspartylglycine Proteins 0.000 description 26
- 108010055341 glutamyl-glutamic acid Proteins 0.000 description 26
- 108010089804 glycyl-threonine Proteins 0.000 description 26
- 108010003700 lysyl aspartic acid Proteins 0.000 description 26
- 108010031719 prolyl-serine Proteins 0.000 description 26
- 108010090461 DFG peptide Proteins 0.000 description 24
- IVGJYOOGJLFKQE-AVGNSLFASA-N Glu-Leu-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCC(=O)O)N IVGJYOOGJLFKQE-AVGNSLFASA-N 0.000 description 24
- KSOBNUBCYHGUKH-UWVGGRQHSA-N Gly-Val-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](C(C)C)NC(=O)CN KSOBNUBCYHGUKH-UWVGGRQHSA-N 0.000 description 24
- GLYJPWIRLBAIJH-UHFFFAOYSA-N Ile-Lys-Pro Natural products CCC(C)C(N)C(=O)NC(CCCCN)C(=O)N1CCCC1C(O)=O GLYJPWIRLBAIJH-UHFFFAOYSA-N 0.000 description 24
- FNGOXVQBBCMFKV-CIUDSAMLSA-N Pro-Ser-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(O)=O FNGOXVQBBCMFKV-CIUDSAMLSA-N 0.000 description 24
- 108010024654 phenylalanyl-prolyl-alanine Proteins 0.000 description 24
- CQJHFKKGZXKZBC-BPNCWPANSA-N Ala-Pro-Tyr Chemical compound C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 CQJHFKKGZXKZBC-BPNCWPANSA-N 0.000 description 23
- UUYBFNKHOCJCHT-VHSXEESVSA-N Gly-Leu-Pro Chemical compound CC(C)C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)CN UUYBFNKHOCJCHT-VHSXEESVSA-N 0.000 description 23
- YOKVEHGYYQEQOP-QWRGUYRKSA-N Leu-Leu-Gly Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)NCC(O)=O YOKVEHGYYQEQOP-QWRGUYRKSA-N 0.000 description 23
- 108010008355 arginyl-glutamine Proteins 0.000 description 23
- DVWVZSJAYIJZFI-FXQIFTODSA-N Ala-Arg-Asn Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(O)=O DVWVZSJAYIJZFI-FXQIFTODSA-N 0.000 description 22
- CRCCTGPNZUCAHE-DCAQKATOSA-N Arg-His-Ser Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CO)C(O)=O)CC1=CN=CN1 CRCCTGPNZUCAHE-DCAQKATOSA-N 0.000 description 22
- GLWFAWNYGWBMOC-SRVKXCTJSA-N Asn-Leu-Leu Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O GLWFAWNYGWBMOC-SRVKXCTJSA-N 0.000 description 22
- 101100342977 Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) leu-1 gene Proteins 0.000 description 22
- IGFJVXOATGZTHD-UHFFFAOYSA-N Arg-Phe-His Natural products NC(CCNC(=N)N)C(=O)NC(Cc1ccccc1)C(=O)NC(Cc2c[nH]cn2)C(=O)O IGFJVXOATGZTHD-UHFFFAOYSA-N 0.000 description 21
- 102000001708 Protein Isoforms Human genes 0.000 description 21
- 108010029485 Protein Isoforms Proteins 0.000 description 21
- 108010087924 alanylproline Proteins 0.000 description 21
- 102000054946 human CDKL5 Human genes 0.000 description 21
- 108010048818 seryl-histidine Proteins 0.000 description 21
- 108010090894 prolylleucine Proteins 0.000 description 20
- HHGYNJRJIINWAK-FXQIFTODSA-N Ala-Ala-Arg Chemical compound C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N HHGYNJRJIINWAK-FXQIFTODSA-N 0.000 description 19
- OGCQGUIWMSBHRZ-CIUDSAMLSA-N Leu-Asn-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(O)=O OGCQGUIWMSBHRZ-CIUDSAMLSA-N 0.000 description 19
- RXGLHDWAZQECBI-SRVKXCTJSA-N Leu-Leu-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O RXGLHDWAZQECBI-SRVKXCTJSA-N 0.000 description 19
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 19
- UVTGNSWSRSCPLP-UHFFFAOYSA-N Arg-Tyr Natural products NC(CCNC(=N)N)C(=O)NC(Cc1ccc(O)cc1)C(=O)O UVTGNSWSRSCPLP-UHFFFAOYSA-N 0.000 description 18
- CHJKEDSZNSONPS-DCAQKATOSA-N Leu-Pro-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O CHJKEDSZNSONPS-DCAQKATOSA-N 0.000 description 18
- GJJQCBVRWDGLMQ-GUBZILKMSA-N Lys-Glu-Ala Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O GJJQCBVRWDGLMQ-GUBZILKMSA-N 0.000 description 18
- VOZIBWWZSBIXQN-SRVKXCTJSA-N Pro-Glu-Lys Chemical compound NCCCC[C@H](NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H]1CCCN1)C(O)=O VOZIBWWZSBIXQN-SRVKXCTJSA-N 0.000 description 18
- 108010062796 arginyllysine Proteins 0.000 description 18
- 108010004914 prolylarginine Proteins 0.000 description 18
- RYQSYXFGFOTJDJ-RHYQMDGZSA-N Arg-Thr-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(O)=O RYQSYXFGFOTJDJ-RHYQMDGZSA-N 0.000 description 17
- VYWNORHENYEQDW-YUMQZZPRSA-N Pro-Gly-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H]1CCCN1 VYWNORHENYEQDW-YUMQZZPRSA-N 0.000 description 17
- 229940024606 amino acid Drugs 0.000 description 17
- 150000001413 amino acids Chemical class 0.000 description 17
- 108010069117 seryl-lysyl-aspartic acid Proteins 0.000 description 17
- 108010072986 threonyl-seryl-lysine Proteins 0.000 description 17
- OMNVOTCFQQLEQU-CIUDSAMLSA-N His-Asn-Asp Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC(=O)O)C(=O)O)N OMNVOTCFQQLEQU-CIUDSAMLSA-N 0.000 description 16
- SITWEMZOJNKJCH-UHFFFAOYSA-N L-alanine-L-arginine Natural products CC(N)C(=O)NC(C(O)=O)CCCNC(N)=N SITWEMZOJNKJCH-UHFFFAOYSA-N 0.000 description 16
- HRTRLSRYZZKPCO-BJDJZHNGSA-N Leu-Ile-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CO)C(O)=O HRTRLSRYZZKPCO-BJDJZHNGSA-N 0.000 description 16
- IBSGMIPRBMPMHE-IHRRRGAJSA-N Leu-Met-Lys Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCCCN)C(O)=O IBSGMIPRBMPMHE-IHRRRGAJSA-N 0.000 description 16
- SBANPBVRHYIMRR-GARJFASQSA-N Leu-Ser-Pro Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CO)C(=O)N1CCC[C@@H]1C(=O)O)N SBANPBVRHYIMRR-GARJFASQSA-N 0.000 description 16
- BSXKBOUZDAZXHE-CIUDSAMLSA-N Ser-Pro-Glu Chemical compound [H]N[C@@H](CO)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(O)=O)C(O)=O BSXKBOUZDAZXHE-CIUDSAMLSA-N 0.000 description 16
- HOVLHEKTGVIKAP-WDCWCFNPSA-N Thr-Leu-Gln Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O HOVLHEKTGVIKAP-WDCWCFNPSA-N 0.000 description 16
- 108010060035 arginylproline Proteins 0.000 description 16
- 108010015792 glycyllysine Proteins 0.000 description 16
- 108010073969 valyllysine Proteins 0.000 description 16
- WDIYWDJLXOCGRW-ACZMJKKPSA-N Ala-Asp-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O WDIYWDJLXOCGRW-ACZMJKKPSA-N 0.000 description 15
- LXMKTIZAGIBQRX-HRCADAONSA-N Arg-Phe-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CC=CC=C2)NC(=O)[C@H](CCCN=C(N)N)N)C(=O)O LXMKTIZAGIBQRX-HRCADAONSA-N 0.000 description 15
- ICRHGPYYXMWHIE-LPEHRKFASA-N Arg-Ser-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CO)NC(=O)[C@H](CCCN=C(N)N)N)C(=O)O ICRHGPYYXMWHIE-LPEHRKFASA-N 0.000 description 15
- CGWVCWFQGXOUSJ-ULQDDVLXSA-N Arg-Tyr-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(C)C)C(O)=O CGWVCWFQGXOUSJ-ULQDDVLXSA-N 0.000 description 15
- 206010010904 Convulsion Diseases 0.000 description 15
- YRHZWVKUFWCEPW-GLLZPBPUSA-N Gln-Thr-Gln Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CCC(=O)N)N)O YRHZWVKUFWCEPW-GLLZPBPUSA-N 0.000 description 15
- AFODTOLGSZQDSL-PEFMBERDSA-N Glu-Asn-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@H](CCC(=O)O)N AFODTOLGSZQDSL-PEFMBERDSA-N 0.000 description 15
- VNCNWQPIQYAMAK-ACZMJKKPSA-N Glu-Ser-Ser Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O VNCNWQPIQYAMAK-ACZMJKKPSA-N 0.000 description 15
- CQGSYZCULZMEDE-UHFFFAOYSA-N Leu-Gln-Pro Natural products CC(C)CC(N)C(=O)NC(CCC(N)=O)C(=O)N1CCCC1C(O)=O CQGSYZCULZMEDE-UHFFFAOYSA-N 0.000 description 15
- KIPIKSXPPLABPN-CIUDSAMLSA-N Pro-Glu-Asn Chemical compound NC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H]1CCCN1 KIPIKSXPPLABPN-CIUDSAMLSA-N 0.000 description 15
- BRGQQXQKPUCUJQ-KBIXCLLPSA-N Ser-Glu-Ile Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O BRGQQXQKPUCUJQ-KBIXCLLPSA-N 0.000 description 15
- CAOYHZOWXFFAIR-CIUDSAMLSA-N Ser-His-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CO)C(O)=O CAOYHZOWXFFAIR-CIUDSAMLSA-N 0.000 description 15
- 108010000434 glycyl-alanyl-leucine Proteins 0.000 description 15
- PXKLCFFSVLKOJM-ACZMJKKPSA-N Ala-Asn-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O PXKLCFFSVLKOJM-ACZMJKKPSA-N 0.000 description 14
- RZZMZYZXNJRPOJ-BJDJZHNGSA-N Ala-Ile-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](C)N RZZMZYZXNJRPOJ-BJDJZHNGSA-N 0.000 description 14
- VRTOMXFZHGWHIJ-KZVJFYERSA-N Ala-Thr-Arg Chemical compound [H]N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O VRTOMXFZHGWHIJ-KZVJFYERSA-N 0.000 description 14
- XSLGWYYNOSUMRM-ZKWXMUAHSA-N Ala-Val-Asn Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O XSLGWYYNOSUMRM-ZKWXMUAHSA-N 0.000 description 14
- IASNWHAGGYTEKX-IUCAKERBSA-N Arg-Arg-Gly Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)NCC(O)=O IASNWHAGGYTEKX-IUCAKERBSA-N 0.000 description 14
- ZWASIOHRQWRWAS-UGYAYLCHSA-N Asn-Asp-Ile Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O ZWASIOHRQWRWAS-UGYAYLCHSA-N 0.000 description 14
- QXOPPIDJKPEKCW-GUBZILKMSA-N Asn-Pro-Arg Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CC(=O)N)N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O QXOPPIDJKPEKCW-GUBZILKMSA-N 0.000 description 14
- RNAQPBOOJRDICC-BPUTZDHNSA-N Asp-Met-Trp Chemical compound CSCC[C@@H](C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)O)NC(=O)[C@H](CC(=O)O)N RNAQPBOOJRDICC-BPUTZDHNSA-N 0.000 description 14
- BRRPVTUFESPTCP-ACZMJKKPSA-N Asp-Ser-Glu Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(O)=O BRRPVTUFESPTCP-ACZMJKKPSA-N 0.000 description 14
- LRZPRGJXAZFXCR-DCAQKATOSA-N Cys-Arg-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CS)N LRZPRGJXAZFXCR-DCAQKATOSA-N 0.000 description 14
- 102000004190 Enzymes Human genes 0.000 description 14
- 108090000790 Enzymes Proteins 0.000 description 14
- XQDGOJPVMSWZSO-SRVKXCTJSA-N Gln-Pro-Leu Chemical compound CC(C)C[C@@H](C(=O)O)NC(=O)[C@@H]1CCCN1C(=O)[C@H](CCC(=O)N)N XQDGOJPVMSWZSO-SRVKXCTJSA-N 0.000 description 14
- YYOBUPFZLKQUAX-FXQIFTODSA-N Glu-Asn-Glu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O YYOBUPFZLKQUAX-FXQIFTODSA-N 0.000 description 14
- XMVLTPMCUJTJQP-FXQIFTODSA-N Glu-Gln-Cys Chemical compound C(CC(=O)O)[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CS)C(=O)O)N XMVLTPMCUJTJQP-FXQIFTODSA-N 0.000 description 14
- XJFITURPHAKKAI-SRVKXCTJSA-N His-Pro-Gln Chemical compound C([C@H](N)C(=O)N1[C@@H](CCC1)C(=O)N[C@@H](CCC(N)=O)C(O)=O)C1=CN=CN1 XJFITURPHAKKAI-SRVKXCTJSA-N 0.000 description 14
- RRSLQOLASISYTB-CIUDSAMLSA-N Leu-Cys-Asp Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CS)C(=O)N[C@@H](CC(O)=O)C(O)=O RRSLQOLASISYTB-CIUDSAMLSA-N 0.000 description 14
- VVQJGYPTIYOFBR-IHRRRGAJSA-N Leu-Lys-Met Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCSC)C(=O)O)N VVQJGYPTIYOFBR-IHRRRGAJSA-N 0.000 description 14
- MVVSHHJKJRZVNY-ACRUOGEOSA-N Leu-Phe-Tyr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O MVVSHHJKJRZVNY-ACRUOGEOSA-N 0.000 description 14
- IZPVWNSAVUQBGP-CIUDSAMLSA-N Leu-Ser-Asp Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O IZPVWNSAVUQBGP-CIUDSAMLSA-N 0.000 description 14
- AZOFEHCPMBRNFD-BZSNNMDCSA-N Lys-Phe-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CCCCN)C(O)=O)CC1=CC=CC=C1 AZOFEHCPMBRNFD-BZSNNMDCSA-N 0.000 description 14
- YRNRVKTYDSLKMD-KKUMJFAQSA-N Lys-Ser-Tyr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O YRNRVKTYDSLKMD-KKUMJFAQSA-N 0.000 description 14
- DRRXXZBXDMLGFC-IHRRRGAJSA-N Lys-Val-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](C(C)C)NC(=O)[C@@H](N)CCCCN DRRXXZBXDMLGFC-IHRRRGAJSA-N 0.000 description 14
- MPCKIRSXNKACRF-GUBZILKMSA-N Met-Pro-Asn Chemical compound CSCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(O)=O MPCKIRSXNKACRF-GUBZILKMSA-N 0.000 description 14
- MGECUMGTSHYHEJ-QEWYBTABSA-N Phe-Glu-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CC1=CC=CC=C1 MGECUMGTSHYHEJ-QEWYBTABSA-N 0.000 description 14
- NHCKESBLOMHIIE-IRXDYDNUSA-N Phe-Gly-Phe Chemical compound C([C@H](N)C(=O)NCC(=O)N[C@@H](CC=1C=CC=CC=1)C(O)=O)C1=CC=CC=C1 NHCKESBLOMHIIE-IRXDYDNUSA-N 0.000 description 14
- FGWUALWGCZJQDJ-URLPEUOOSA-N Phe-Thr-Ile Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O FGWUALWGCZJQDJ-URLPEUOOSA-N 0.000 description 14
- DRKAXLDECUGLFE-ULQDDVLXSA-N Pro-Leu-Phe Chemical compound CC(C)C[C@H](NC(=O)[C@@H]1CCCN1)C(=O)N[C@@H](Cc1ccccc1)C(O)=O DRKAXLDECUGLFE-ULQDDVLXSA-N 0.000 description 14
- BNFVPSRLHHPQKS-WHFBIAKZSA-N Ser-Asp-Gly Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(=O)NCC(O)=O BNFVPSRLHHPQKS-WHFBIAKZSA-N 0.000 description 14
- ZIFYDQAFEMIZII-GUBZILKMSA-N Ser-Leu-Glu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O ZIFYDQAFEMIZII-GUBZILKMSA-N 0.000 description 14
- UPLYXVPQLJVWMM-KKUMJFAQSA-N Ser-Phe-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(C)C)C(O)=O UPLYXVPQLJVWMM-KKUMJFAQSA-N 0.000 description 14
- JZRYFUGREMECBH-XPUUQOCRSA-N Ser-Val-Gly Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(=O)NCC(O)=O JZRYFUGREMECBH-XPUUQOCRSA-N 0.000 description 14
- GZTKZDGIEBKZAH-XIRDDKMYSA-N Trp-Cys-His Chemical compound C1=CC=C2C(=C1)C(=CN2)C[C@@H](C(=O)N[C@@H](CS)C(=O)N[C@@H](CC3=CN=CN3)C(=O)O)N GZTKZDGIEBKZAH-XIRDDKMYSA-N 0.000 description 14
- IIJWXEUNETVJPV-IHRRRGAJSA-N Tyr-Arg-Ser Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CO)C(=O)O)N)O IIJWXEUNETVJPV-IHRRRGAJSA-N 0.000 description 14
- YLRAFVVWZRSZQC-DZKIICNBSA-N Val-Phe-Glu Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N YLRAFVVWZRSZQC-DZKIICNBSA-N 0.000 description 14
- 108010013835 arginine glutamate Proteins 0.000 description 14
- 210000004978 chinese hamster ovary cell Anatomy 0.000 description 14
- 201000010099 disease Diseases 0.000 description 14
- XKUKSGPZAADMRA-UHFFFAOYSA-N glycyl-glycyl-glycine Natural products NCC(=O)NCC(=O)NCC(O)=O XKUKSGPZAADMRA-UHFFFAOYSA-N 0.000 description 14
- 108010037850 glycylvaline Proteins 0.000 description 14
- 108010073472 leucyl-prolyl-proline Proteins 0.000 description 14
- 108010054155 lysyllysine Proteins 0.000 description 14
- 108010017391 lysylvaline Proteins 0.000 description 14
- UGLPMYSCWHTZQU-AUTRQRHGSA-N Ala-Ala-Tyr Chemical compound C[C@H]([NH3+])C(=O)N[C@@H](C)C(=O)N[C@H](C([O-])=O)CC1=CC=C(O)C=C1 UGLPMYSCWHTZQU-AUTRQRHGSA-N 0.000 description 13
- LWUWMHIOBPTZBA-DCAQKATOSA-N Ala-Arg-Lys Chemical compound NC(=N)NCCC[C@H](NC(=O)[C@@H](N)C)C(=O)N[C@@H](CCCCN)C(O)=O LWUWMHIOBPTZBA-DCAQKATOSA-N 0.000 description 13
- NINQYGGNRIBFSC-CIUDSAMLSA-N Ala-Lys-Ser Chemical compound NCCCC[C@H](NC(=O)[C@@H](N)C)C(=O)N[C@@H](CO)C(O)=O NINQYGGNRIBFSC-CIUDSAMLSA-N 0.000 description 13
- DJIMLSXHXKWADV-CIUDSAMLSA-N Asn-Leu-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC(N)=O DJIMLSXHXKWADV-CIUDSAMLSA-N 0.000 description 13
- INKFLNZBTSNFON-CIUDSAMLSA-N Gln-Ala-Arg Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O INKFLNZBTSNFON-CIUDSAMLSA-N 0.000 description 13
- SHERTACNJPYHAR-ACZMJKKPSA-N Gln-Ala-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCC(N)=O SHERTACNJPYHAR-ACZMJKKPSA-N 0.000 description 13
- ATRHMOJQJWPVBQ-DRZSPHRISA-N Glu-Ala-Phe Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O ATRHMOJQJWPVBQ-DRZSPHRISA-N 0.000 description 13
- QVFGXCVIXXBFHO-AVGNSLFASA-N Leu-Glu-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O QVFGXCVIXXBFHO-AVGNSLFASA-N 0.000 description 13
- CCQLQKZTXZBXTN-NHCYSSNCSA-N Leu-Gly-Ile Chemical compound [H]N[C@@H](CC(C)C)C(=O)NCC(=O)N[C@@H]([C@@H](C)CC)C(O)=O CCQLQKZTXZBXTN-NHCYSSNCSA-N 0.000 description 13
- BGZCJDGBBUUBHA-KKUMJFAQSA-N Leu-Lys-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O BGZCJDGBBUUBHA-KKUMJFAQSA-N 0.000 description 13
- HGZHSNBZDOLMLH-DCAQKATOSA-N Lys-Asn-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@H](CCCCN)N HGZHSNBZDOLMLH-DCAQKATOSA-N 0.000 description 13
- MUXNCRWTWBMNHX-SRVKXCTJSA-N Lys-Leu-Asp Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O MUXNCRWTWBMNHX-SRVKXCTJSA-N 0.000 description 13
- APKRGYLBSCWJJP-FXQIFTODSA-N Pro-Ala-Asp Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](C)C(=O)N[C@@H](CC(O)=O)C(O)=O APKRGYLBSCWJJP-FXQIFTODSA-N 0.000 description 13
- YUJLIIRMIAGMCQ-CIUDSAMLSA-N Ser-Leu-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O YUJLIIRMIAGMCQ-CIUDSAMLSA-N 0.000 description 13
- VUXIQSUQQYNLJP-XAVMHZPKSA-N Thr-Ser-Pro Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CO)C(=O)N1CCC[C@@H]1C(=O)O)N)O VUXIQSUQQYNLJP-XAVMHZPKSA-N 0.000 description 13
- PQPWEALFTLKSEB-DZKIICNBSA-N Tyr-Val-Glu Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(O)=O)C(O)=O PQPWEALFTLKSEB-DZKIICNBSA-N 0.000 description 13
- 108010045350 alanyl-tyrosyl-alanine Proteins 0.000 description 13
- 108010006664 gamma-glutamyl-glycyl-glycine Proteins 0.000 description 13
- 108010074027 glycyl-seryl-phenylalanine Proteins 0.000 description 13
- 108010009298 lysylglutamic acid Proteins 0.000 description 13
- 108010029020 prolylglycine Proteins 0.000 description 13
- NMXKFWOEASXOGB-QSFUFRPTSA-N Ala-Ile-His Chemical compound C[C@H](N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@H](C(O)=O)CC1=CN=CN1 NMXKFWOEASXOGB-QSFUFRPTSA-N 0.000 description 12
- LDLSENBXQNDTPB-DCAQKATOSA-N Ala-Lys-Arg Chemical compound NCCCC[C@H](NC(=O)[C@@H](N)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N LDLSENBXQNDTPB-DCAQKATOSA-N 0.000 description 12
- PGNNQOJOEGFAOR-KWQFWETISA-N Ala-Tyr-Gly Chemical compound OC(=O)CNC(=O)[C@@H](NC(=O)[C@@H](N)C)CC1=CC=C(O)C=C1 PGNNQOJOEGFAOR-KWQFWETISA-N 0.000 description 12
- NABSCJGZKWSNHX-RCWTZXSCSA-N Arg-Arg-Thr Chemical compound NC(N)=NCCC[C@@H](C(=O)N[C@@H]([C@H](O)C)C(O)=O)NC(=O)[C@@H](N)CCCN=C(N)N NABSCJGZKWSNHX-RCWTZXSCSA-N 0.000 description 12
- YUIGJDNAGKJLDO-JYJNAYRXSA-N Arg-Arg-Tyr Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O YUIGJDNAGKJLDO-JYJNAYRXSA-N 0.000 description 12
- ZZXMOQIUIJJOKZ-ZLUOBGJFSA-N Asn-Asn-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CC(N)=O ZZXMOQIUIJJOKZ-ZLUOBGJFSA-N 0.000 description 12
- XEGZSHSPQNDNRH-JRQIVUDYSA-N Asn-Tyr-Thr Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H]([C@@H](C)O)C(O)=O XEGZSHSPQNDNRH-JRQIVUDYSA-N 0.000 description 12
- HRGGPWBIMIQANI-GUBZILKMSA-N Asp-Gln-Leu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O HRGGPWBIMIQANI-GUBZILKMSA-N 0.000 description 12
- YFSLJHLQOALGSY-ZPFDUUQYSA-N Asp-Ile-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CC(=O)O)N YFSLJHLQOALGSY-ZPFDUUQYSA-N 0.000 description 12
- OXFOKRAFNYSREH-BJDJZHNGSA-N Cys-Ile-Leu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(C)C)C(=O)O)NC(=O)[C@H](CS)N OXFOKRAFNYSREH-BJDJZHNGSA-N 0.000 description 12
- ZXGLLNZQSBLQLT-SRVKXCTJSA-N Gln-Met-Lys Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCC(=O)N)N ZXGLLNZQSBLQLT-SRVKXCTJSA-N 0.000 description 12
- PVBBEKPHARMPHX-DCAQKATOSA-N Glu-Gln-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(N)=O)NC(=O)[C@@H](N)CCC(O)=O PVBBEKPHARMPHX-DCAQKATOSA-N 0.000 description 12
- CAQXJMUDOLSBPF-SUSMZKCASA-N Glu-Thr-Thr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O CAQXJMUDOLSBPF-SUSMZKCASA-N 0.000 description 12
- HBMRTXJZQDVRFT-DZKIICNBSA-N Glu-Tyr-Val Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](C(C)C)C(O)=O HBMRTXJZQDVRFT-DZKIICNBSA-N 0.000 description 12
- ZYRXTRTUCAVNBQ-GVXVVHGQSA-N Glu-Val-Lys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCC(=O)O)N ZYRXTRTUCAVNBQ-GVXVVHGQSA-N 0.000 description 12
- GRIRDMVMJJDZKV-RCOVLWMOSA-N Gly-Asn-Val Chemical compound [H]NCC(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](C(C)C)C(O)=O GRIRDMVMJJDZKV-RCOVLWMOSA-N 0.000 description 12
- XTQFHTHIAKKCTM-YFKPBYRVSA-N Gly-Glu-Gly Chemical compound NCC(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O XTQFHTHIAKKCTM-YFKPBYRVSA-N 0.000 description 12
- YYPFZVIXAVDHIK-IUCAKERBSA-N Gly-Glu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)CN YYPFZVIXAVDHIK-IUCAKERBSA-N 0.000 description 12
- HFPVRZWORNJRRC-UWVGGRQHSA-N Gly-Pro-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)CN HFPVRZWORNJRRC-UWVGGRQHSA-N 0.000 description 12
- YGHSQRJSHKYUJY-SCZZXKLOSA-N Gly-Val-Pro Chemical compound CC(C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)CN YGHSQRJSHKYUJY-SCZZXKLOSA-N 0.000 description 12
- JBCLFWXMTIKCCB-UHFFFAOYSA-N H-Gly-Phe-OH Natural products NCC(=O)NC(C(O)=O)CC1=CC=CC=C1 JBCLFWXMTIKCCB-UHFFFAOYSA-N 0.000 description 12
- HQKADFMLECZIQJ-HVTMNAMFSA-N His-Glu-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CC1=CN=CN1)N HQKADFMLECZIQJ-HVTMNAMFSA-N 0.000 description 12
- GNXGAVNTVNOCLL-SIUGBPQLSA-N Ile-Tyr-Gln Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N GNXGAVNTVNOCLL-SIUGBPQLSA-N 0.000 description 12
- HASRFYOMVPJRPU-SRVKXCTJSA-N Leu-Arg-Glu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCC(O)=O)C(O)=O HASRFYOMVPJRPU-SRVKXCTJSA-N 0.000 description 12
- STAVRDQLZOTNKJ-RHYQMDGZSA-N Leu-Arg-Thr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)O)C(O)=O STAVRDQLZOTNKJ-RHYQMDGZSA-N 0.000 description 12
- VIWUBXKCYJGNCL-SRVKXCTJSA-N Leu-Asn-His Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@H](C(O)=O)CC1=CN=CN1 VIWUBXKCYJGNCL-SRVKXCTJSA-N 0.000 description 12
- WIDZHJTYKYBLSR-DCAQKATOSA-N Leu-Glu-Glu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O WIDZHJTYKYBLSR-DCAQKATOSA-N 0.000 description 12
- HNDWYLYAYNBWMP-AJNGGQMLSA-N Leu-Ile-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CC(C)C)N HNDWYLYAYNBWMP-AJNGGQMLSA-N 0.000 description 12
- HVHRPWQEQHIQJF-AVGNSLFASA-N Leu-Lys-Glu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O HVHRPWQEQHIQJF-AVGNSLFASA-N 0.000 description 12
- JIHDFWWRYHSAQB-GUBZILKMSA-N Leu-Ser-Glu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(O)=O JIHDFWWRYHSAQB-GUBZILKMSA-N 0.000 description 12
- 108010062166 Lys-Asn-Asp Proteins 0.000 description 12
- VUTWYNQUSJWBHO-BZSNNMDCSA-N Lys-Leu-Tyr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O VUTWYNQUSJWBHO-BZSNNMDCSA-N 0.000 description 12
- DRXODWRPPUFIAY-DCAQKATOSA-N Met-Asn-Lys Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@H](C(O)=O)CCCCN DRXODWRPPUFIAY-DCAQKATOSA-N 0.000 description 12
- HOZNVKDCKZPRER-XUXIUFHCSA-N Met-Lys-Ile Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O HOZNVKDCKZPRER-XUXIUFHCSA-N 0.000 description 12
- 108010079364 N-glycylalanine Proteins 0.000 description 12
- SFKOEHXABNPLRT-KBPBESRZSA-N Phe-His-Gly Chemical compound N[C@@H](Cc1ccccc1)C(=O)N[C@@H](Cc1cnc[nH]1)C(=O)NCC(O)=O SFKOEHXABNPLRT-KBPBESRZSA-N 0.000 description 12
- XWYXZPHPYKRYPA-GMOBBJLQSA-N Pro-Asn-Ile Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O XWYXZPHPYKRYPA-GMOBBJLQSA-N 0.000 description 12
- NXEYSLRNNPWCRN-SRVKXCTJSA-N Pro-Glu-Leu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O NXEYSLRNNPWCRN-SRVKXCTJSA-N 0.000 description 12
- JDJMFMVVJHLWDP-UNQGMJICSA-N Pro-Thr-Phe Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O JDJMFMVVJHLWDP-UNQGMJICSA-N 0.000 description 12
- PPNPDKGQRFSCAC-CIUDSAMLSA-N Ser-Lys-Asp Chemical compound NCCCC[C@H](NC(=O)[C@@H](N)CO)C(=O)N[C@@H](CC(O)=O)C(O)=O PPNPDKGQRFSCAC-CIUDSAMLSA-N 0.000 description 12
- FZXOPYUEQGDGMS-ACZMJKKPSA-N Ser-Ser-Gln Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(O)=O FZXOPYUEQGDGMS-ACZMJKKPSA-N 0.000 description 12
- GKWNLDNXMMLRMC-GLLZPBPUSA-N Thr-Glu-Gln Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N)O GKWNLDNXMMLRMC-GLLZPBPUSA-N 0.000 description 12
- OPGWZDIYEYJVRX-AVGNSLFASA-N Val-His-Arg Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N OPGWZDIYEYJVRX-AVGNSLFASA-N 0.000 description 12
- 108010076324 alanyl-glycyl-glycine Proteins 0.000 description 12
- 108010068265 aspartyltyrosine Proteins 0.000 description 12
- 108010078274 isoleucylvaline Proteins 0.000 description 12
- 108010057821 leucylproline Proteins 0.000 description 12
- 108010038320 lysylphenylalanine Proteins 0.000 description 12
- 108010091617 pentalysine Proteins 0.000 description 12
- 108010018625 phenylalanylarginine Proteins 0.000 description 12
- 108010077112 prolyl-proline Proteins 0.000 description 12
- BUANFPRKJKJSRR-ACZMJKKPSA-N Ala-Ala-Gln Chemical compound C[C@H]([NH3+])C(=O)N[C@@H](C)C(=O)N[C@H](C([O-])=O)CCC(N)=O BUANFPRKJKJSRR-ACZMJKKPSA-N 0.000 description 11
- GXCSUJQOECMKPV-CIUDSAMLSA-N Arg-Ala-Gln Chemical compound C[C@H](NC(=O)[C@@H](N)CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(O)=O GXCSUJQOECMKPV-CIUDSAMLSA-N 0.000 description 11
- VBFJESQBIWCWRL-DCAQKATOSA-N Arg-Ala-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCCNC(N)=N VBFJESQBIWCWRL-DCAQKATOSA-N 0.000 description 11
- VYZBPPBKFCHCIS-WPRPVWTQSA-N Arg-Val-Gly Chemical compound OC(=O)CNC(=O)[C@H](C(C)C)NC(=O)[C@@H](N)CCCN=C(N)N VYZBPPBKFCHCIS-WPRPVWTQSA-N 0.000 description 11
- SUMJNGAMIQSNGX-TUAOUCFPSA-N Arg-Val-Pro Chemical compound CC(C)[C@H](NC(=O)[C@@H](N)CCCNC(N)=N)C(=O)N1CCC[C@@H]1C(O)=O SUMJNGAMIQSNGX-TUAOUCFPSA-N 0.000 description 11
- SPCONPVIDFMDJI-QSFUFRPTSA-N Asn-Ile-Val Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](C(C)C)C(O)=O SPCONPVIDFMDJI-QSFUFRPTSA-N 0.000 description 11
- KKBWDNZXYLGJEY-UHFFFAOYSA-N Gly-Arg-Pro Natural products NCC(=O)NC(CCNC(=N)N)C(=O)N1CCCC1C(=O)O KKBWDNZXYLGJEY-UHFFFAOYSA-N 0.000 description 11
- POJJAZJHBGXEGM-YUMQZZPRSA-N Gly-Ser-Lys Chemical compound C(CCN)C[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)CN POJJAZJHBGXEGM-YUMQZZPRSA-N 0.000 description 11
- PMGDADKJMCOXHX-UHFFFAOYSA-N L-Arginyl-L-glutamin-acetat Natural products NC(=N)NCCCC(N)C(=O)NC(CCC(N)=O)C(O)=O PMGDADKJMCOXHX-UHFFFAOYSA-N 0.000 description 11
- LHSGPCFBGJHPCY-UHFFFAOYSA-N L-leucine-L-tyrosine Natural products CC(C)CC(N)C(=O)NC(C(O)=O)CC1=CC=C(O)C=C1 LHSGPCFBGJHPCY-UHFFFAOYSA-N 0.000 description 11
- HWMZUBUEOYAQSC-DCAQKATOSA-N Lys-Gln-Glu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O HWMZUBUEOYAQSC-DCAQKATOSA-N 0.000 description 11
- KZNQNBZMBZJQJO-UHFFFAOYSA-N N-glycyl-L-proline Natural products NCC(=O)N1CCCC1C(O)=O KZNQNBZMBZJQJO-UHFFFAOYSA-N 0.000 description 11
- 108091000080 Phosphotransferase Proteins 0.000 description 11
- VXCHGLYSIOOZIS-GUBZILKMSA-N Pro-Ala-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1 VXCHGLYSIOOZIS-GUBZILKMSA-N 0.000 description 11
- RFWXYTJSVDUBBZ-DCAQKATOSA-N Pro-Pro-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@H]1NCCC1 RFWXYTJSVDUBBZ-DCAQKATOSA-N 0.000 description 11
- MKGIILKDUGDRRO-FXQIFTODSA-N Pro-Ser-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H]1CCCN1 MKGIILKDUGDRRO-FXQIFTODSA-N 0.000 description 11
- UGDMQJSXSSZUKL-IHRRRGAJSA-N Pro-Ser-Tyr Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC2=CC=C(C=C2)O)C(=O)O UGDMQJSXSSZUKL-IHRRRGAJSA-N 0.000 description 11
- BQWCDDAISCPDQV-XHNCKOQMSA-N Ser-Gln-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CO)N)C(=O)O BQWCDDAISCPDQV-XHNCKOQMSA-N 0.000 description 11
- ZKBKUWQVDWWSRI-BZSNNMDCSA-N Ser-Phe-Tyr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O ZKBKUWQVDWWSRI-BZSNNMDCSA-N 0.000 description 11
- HSBZWINKRYZCSQ-KKUMJFAQSA-N Tyr-Lys-Asp Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(O)=O HSBZWINKRYZCSQ-KKUMJFAQSA-N 0.000 description 11
- ROLGIBMFNMZANA-GVXVVHGQSA-N Val-Glu-Leu Chemical compound CC(C)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](C(C)C)N ROLGIBMFNMZANA-GVXVVHGQSA-N 0.000 description 11
- PZTZYZUTCPZWJH-FXQIFTODSA-N Val-Ser-Ser Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)O)N PZTZYZUTCPZWJH-FXQIFTODSA-N 0.000 description 11
- 108010086434 alanyl-seryl-glycine Proteins 0.000 description 11
- 125000003275 alpha amino acid group Chemical group 0.000 description 11
- 210000000170 cell membrane Anatomy 0.000 description 11
- 230000000694 effects Effects 0.000 description 11
- 108010010147 glycylglutamine Proteins 0.000 description 11
- 108010077515 glycylproline Proteins 0.000 description 11
- 108010012058 leucyltyrosine Proteins 0.000 description 11
- 108010076718 lysyl-glutamyl-tryptophan Proteins 0.000 description 11
- 102000020233 phosphotransferase Human genes 0.000 description 11
- 108010079317 prolyl-tyrosine Proteins 0.000 description 11
- 208000024891 symptom Diseases 0.000 description 11
- XQJAFSDFQZPYCU-UWJYBYFXSA-N Ala-Asn-Tyr Chemical compound C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)O)N XQJAFSDFQZPYCU-UWJYBYFXSA-N 0.000 description 10
- VGPWRRFOPXVGOH-BYPYZUCNSA-N Ala-Gly-Gly Chemical compound C[C@H](N)C(=O)NCC(=O)NCC(O)=O VGPWRRFOPXVGOH-BYPYZUCNSA-N 0.000 description 10
- XPSGESXVBSQZPL-SRVKXCTJSA-N Arg-Arg-Arg Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O XPSGESXVBSQZPL-SRVKXCTJSA-N 0.000 description 10
- JSHVMZANPXCDTL-GMOBBJLQSA-N Arg-Asp-Ile Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O JSHVMZANPXCDTL-GMOBBJLQSA-N 0.000 description 10
- OOWSBIOUKIUWLO-RCOVLWMOSA-N Asn-Gly-Val Chemical compound [H]N[C@@H](CC(N)=O)C(=O)NCC(=O)N[C@@H](C(C)C)C(O)=O OOWSBIOUKIUWLO-RCOVLWMOSA-N 0.000 description 10
- SSNJZBGOMNLSLA-CIUDSAMLSA-N Cys-Leu-Asn Chemical compound [H]N[C@@H](CS)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O SSNJZBGOMNLSLA-CIUDSAMLSA-N 0.000 description 10
- PSERKXGRRADTKA-MNXVOIDGSA-N Gln-Leu-Ile Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O PSERKXGRRADTKA-MNXVOIDGSA-N 0.000 description 10
- QKWBEMCLYTYBNI-GVXVVHGQSA-N Gln-Lys-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](N)CCC(N)=O QKWBEMCLYTYBNI-GVXVVHGQSA-N 0.000 description 10
- LPIKVBWNNVFHCQ-GUBZILKMSA-N Gln-Ser-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O LPIKVBWNNVFHCQ-GUBZILKMSA-N 0.000 description 10
- WOMUDRVDJMHTCV-DCAQKATOSA-N Glu-Arg-Arg Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O WOMUDRVDJMHTCV-DCAQKATOSA-N 0.000 description 10
- GYCPQVFKCPPRQB-GUBZILKMSA-N Glu-Gln-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CCC(=O)O)N GYCPQVFKCPPRQB-GUBZILKMSA-N 0.000 description 10
- QQLBPVKLJBAXBS-FXQIFTODSA-N Glu-Glu-Asn Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O QQLBPVKLJBAXBS-FXQIFTODSA-N 0.000 description 10
- QJCKNLPMTPXXEM-AUTRQRHGSA-N Glu-Glu-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CCC(O)=O QJCKNLPMTPXXEM-AUTRQRHGSA-N 0.000 description 10
- UMHRCVCZUPBBQW-GARJFASQSA-N Glu-Met-Pro Chemical compound CSCC[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCC(=O)O)N UMHRCVCZUPBBQW-GARJFASQSA-N 0.000 description 10
- GMVCSRBOSIUTFC-FXQIFTODSA-N Glu-Ser-Glu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(O)=O GMVCSRBOSIUTFC-FXQIFTODSA-N 0.000 description 10
- QIZJOTQTCAGKPU-KWQFWETISA-N Gly-Ala-Tyr Chemical compound [NH3+]CC(=O)N[C@@H](C)C(=O)N[C@H](C([O-])=O)CC1=CC=C(O)C=C1 QIZJOTQTCAGKPU-KWQFWETISA-N 0.000 description 10
- CIMULJZTTOBOPN-WHFBIAKZSA-N Gly-Asn-Asn Chemical compound NCC(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O CIMULJZTTOBOPN-WHFBIAKZSA-N 0.000 description 10
- LEGMTEAZGRRIMY-ZKWXMUAHSA-N Gly-Cys-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CS)NC(=O)CN LEGMTEAZGRRIMY-ZKWXMUAHSA-N 0.000 description 10
- MHXKHKWHPNETGG-QWRGUYRKSA-N Gly-Lys-Leu Chemical compound [H]NCC(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O MHXKHKWHPNETGG-QWRGUYRKSA-N 0.000 description 10
- JUIOPCXACJLRJK-AVGNSLFASA-N His-Lys-Glu Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N JUIOPCXACJLRJK-AVGNSLFASA-N 0.000 description 10
- OWYIDJCNRWRSJY-QTKMDUPCSA-N His-Pro-Thr Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(O)=O OWYIDJCNRWRSJY-QTKMDUPCSA-N 0.000 description 10
- UOYGZBIPZYKGSH-SRVKXCTJSA-N His-Ser-Lys Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)O)N UOYGZBIPZYKGSH-SRVKXCTJSA-N 0.000 description 10
- UDLAWRKOVFDKFL-PEFMBERDSA-N Ile-Asp-Gln Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N UDLAWRKOVFDKFL-PEFMBERDSA-N 0.000 description 10
- GVNNAHIRSDRIII-AJNGGQMLSA-N Ile-Lys-Lys Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)O)N GVNNAHIRSDRIII-AJNGGQMLSA-N 0.000 description 10
- YJRSIJZUIUANHO-NAKRPEOUSA-N Ile-Val-Ala Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C)C(=O)O)N YJRSIJZUIUANHO-NAKRPEOUSA-N 0.000 description 10
- JCGMFFQQHJQASB-PYJNHQTQSA-N Ile-Val-His Chemical compound N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC1=CNC=N1)C(=O)O JCGMFFQQHJQASB-PYJNHQTQSA-N 0.000 description 10
- KGCLIYGPQXUNLO-IUCAKERBSA-N Leu-Gly-Glu Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CCC(O)=O KGCLIYGPQXUNLO-IUCAKERBSA-N 0.000 description 10
- YFBBUHJJUXXZOF-UWVGGRQHSA-N Leu-Gly-Pro Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)N1CCC[C@H]1C(O)=O YFBBUHJJUXXZOF-UWVGGRQHSA-N 0.000 description 10
- POZULHZYLPGXMR-ONGXEEELSA-N Leu-Gly-Val Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)N[C@@H](C(C)C)C(O)=O POZULHZYLPGXMR-ONGXEEELSA-N 0.000 description 10
- QNBVTHNJGCOVFA-AVGNSLFASA-N Leu-Leu-Glu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCC(O)=O QNBVTHNJGCOVFA-AVGNSLFASA-N 0.000 description 10
- ZGUMORRUBUCXEH-AVGNSLFASA-N Leu-Lys-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(N)=O)C(O)=O ZGUMORRUBUCXEH-AVGNSLFASA-N 0.000 description 10
- NFLFJGGKOHYZJF-BJDJZHNGSA-N Lys-Ala-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCCCN NFLFJGGKOHYZJF-BJDJZHNGSA-N 0.000 description 10
- BYPMOIFBQPEWOH-CIUDSAMLSA-N Lys-Asn-Asp Chemical compound C(CCN)C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC(=O)O)C(=O)O)N BYPMOIFBQPEWOH-CIUDSAMLSA-N 0.000 description 10
- CNGOEHJCLVCJHN-SRVKXCTJSA-N Lys-Pro-Glu Chemical compound NCCCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(O)=O)C(O)=O CNGOEHJCLVCJHN-SRVKXCTJSA-N 0.000 description 10
- MEQLGHAMAUPOSJ-DCAQKATOSA-N Lys-Ser-Val Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(O)=O MEQLGHAMAUPOSJ-DCAQKATOSA-N 0.000 description 10
- HGAJNEWOUHDUMZ-SRVKXCTJSA-N Met-Leu-Glu Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCC(O)=O HGAJNEWOUHDUMZ-SRVKXCTJSA-N 0.000 description 10
- NKLDZIPTGKBDBB-HTUGSXCWSA-N Phe-Gln-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CC1=CC=CC=C1)N)O NKLDZIPTGKBDBB-HTUGSXCWSA-N 0.000 description 10
- UAMFZRNCIFFMLE-FHWLQOOXSA-N Phe-Glu-Tyr Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CC2=CC=C(C=C2)O)C(=O)O)N UAMFZRNCIFFMLE-FHWLQOOXSA-N 0.000 description 10
- MMJJFXWMCMJMQA-STQMWFEESA-N Phe-Pro-Gly Chemical compound C([C@H](N)C(=O)N1[C@@H](CCC1)C(=O)NCC(O)=O)C1=CC=CC=C1 MMJJFXWMCMJMQA-STQMWFEESA-N 0.000 description 10
- XQPHBAKJJJZOBX-SRVKXCTJSA-N Pro-Lys-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O XQPHBAKJJJZOBX-SRVKXCTJSA-N 0.000 description 10
- DKKGAAJTDKHWOD-BIIVOSGPSA-N Ser-Asn-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)N)NC(=O)[C@H](CO)N)C(=O)O DKKGAAJTDKHWOD-BIIVOSGPSA-N 0.000 description 10
- YQQKYAZABFEYAF-FXQIFTODSA-N Ser-Glu-Gln Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O YQQKYAZABFEYAF-FXQIFTODSA-N 0.000 description 10
- LRZLZIUXQBIWTB-KATARQTJSA-N Ser-Lys-Thr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)O)C(O)=O LRZLZIUXQBIWTB-KATARQTJSA-N 0.000 description 10
- FLONGDPORFIVQW-XGEHTFHBSA-N Ser-Pro-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CO FLONGDPORFIVQW-XGEHTFHBSA-N 0.000 description 10
- SNXUIBACCONSOH-BWBBJGPYSA-N Ser-Thr-Ser Chemical compound OC[C@H](N)C(=O)N[C@@H]([C@H](O)C)C(=O)N[C@@H](CO)C(O)=O SNXUIBACCONSOH-BWBBJGPYSA-N 0.000 description 10
- BSNZTJXVDOINSR-JXUBOQSCSA-N Thr-Ala-Leu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(O)=O BSNZTJXVDOINSR-JXUBOQSCSA-N 0.000 description 10
- LKEKWDJCJSPXNI-IRIUXVKKSA-N Thr-Glu-Tyr Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 LKEKWDJCJSPXNI-IRIUXVKKSA-N 0.000 description 10
- WPSDXXQRIVKBAY-NKIYYHGXSA-N Thr-His-Glu Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N)O WPSDXXQRIVKBAY-NKIYYHGXSA-N 0.000 description 10
- AMXMBCAXAZUCFA-RHYQMDGZSA-N Thr-Leu-Arg Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O AMXMBCAXAZUCFA-RHYQMDGZSA-N 0.000 description 10
- PRTHQBSMXILLPC-XGEHTFHBSA-N Thr-Ser-Arg Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O PRTHQBSMXILLPC-XGEHTFHBSA-N 0.000 description 10
- HIZDHWHVOLUGOX-BPUTZDHNSA-N Trp-Ser-Val Chemical compound [H]N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(O)=O HIZDHWHVOLUGOX-BPUTZDHNSA-N 0.000 description 10
- UIDJDMVRDUANDL-BVSLBCMMSA-N Trp-Tyr-Arg Chemical compound [H]N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O UIDJDMVRDUANDL-BVSLBCMMSA-N 0.000 description 10
- YMUQBRQQCPQEQN-CXTHYWKRSA-N Tyr-Ile-Tyr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)O)NC(=O)[C@H](CC2=CC=C(C=C2)O)N YMUQBRQQCPQEQN-CXTHYWKRSA-N 0.000 description 10
- CDKZJGMPZHPAJC-ULQDDVLXSA-N Tyr-Leu-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 CDKZJGMPZHPAJC-ULQDDVLXSA-N 0.000 description 10
- ZSZFTYVFQLUWBF-QXEWZRGKSA-N Val-Asp-Met Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CCSC)C(=O)O)N ZSZFTYVFQLUWBF-QXEWZRGKSA-N 0.000 description 10
- WFENBJPLZMPVAX-XVKPBYJWSA-N Val-Gly-Glu Chemical compound CC(C)[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CCC(O)=O WFENBJPLZMPVAX-XVKPBYJWSA-N 0.000 description 10
- JAKHAONCJJZVHT-DCAQKATOSA-N Val-Lys-Ser Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(=O)O)N JAKHAONCJJZVHT-DCAQKATOSA-N 0.000 description 10
- 239000003814 drug Substances 0.000 description 10
- 108010064235 lysylglycine Proteins 0.000 description 10
- 108010056582 methionylglutamic acid Proteins 0.000 description 10
- 108010034507 methionyltryptophan Proteins 0.000 description 10
- BLIMFWGRQKRCGT-YUMQZZPRSA-N Ala-Gly-Lys Chemical compound C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CCCCN BLIMFWGRQKRCGT-YUMQZZPRSA-N 0.000 description 9
- HOVPGJUNRLMIOZ-CIUDSAMLSA-N Ala-Ser-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@H](C)N HOVPGJUNRLMIOZ-CIUDSAMLSA-N 0.000 description 9
- CPSHGRGUPZBMOK-CIUDSAMLSA-N Arg-Asn-Gln Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O CPSHGRGUPZBMOK-CIUDSAMLSA-N 0.000 description 9
- FIQKRDXFTANIEJ-ULQDDVLXSA-N Arg-Phe-His Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N FIQKRDXFTANIEJ-ULQDDVLXSA-N 0.000 description 9
- IOTKDTZEEBZNCM-UGYAYLCHSA-N Asn-Asn-Ile Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O IOTKDTZEEBZNCM-UGYAYLCHSA-N 0.000 description 9
- DXVMJJNAOVECBA-WHFBIAKZSA-N Asn-Gly-Asn Chemical compound NC(=O)C[C@H](N)C(=O)NCC(=O)N[C@@H](CC(N)=O)C(O)=O DXVMJJNAOVECBA-WHFBIAKZSA-N 0.000 description 9
- TZFQICWZWFNIKU-KKUMJFAQSA-N Asn-Leu-Tyr Chemical compound NC(=O)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 TZFQICWZWFNIKU-KKUMJFAQSA-N 0.000 description 9
- UGXYFDQFLVCDFC-CIUDSAMLSA-N Asn-Ser-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H](N)CC(N)=O UGXYFDQFLVCDFC-CIUDSAMLSA-N 0.000 description 9
- QNFRBNZGVVKBNJ-PEFMBERDSA-N Asp-Ile-Gln Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CC(=O)O)N QNFRBNZGVVKBNJ-PEFMBERDSA-N 0.000 description 9
- BJDHEININLSZOT-KKUMJFAQSA-N Asp-Tyr-Lys Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCCCN)C(O)=O BJDHEININLSZOT-KKUMJFAQSA-N 0.000 description 9
- ZBKUIQNCRIYVGH-SDDRHHMPSA-N Gln-Leu-Pro Chemical compound CC(C)C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCC(=O)N)N ZBKUIQNCRIYVGH-SDDRHHMPSA-N 0.000 description 9
- FNAJNWPDTIXYJN-CIUDSAMLSA-N Gln-Pro-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CCC(N)=O FNAJNWPDTIXYJN-CIUDSAMLSA-N 0.000 description 9
- KVQOVQVGVKDZNW-GUBZILKMSA-N Gln-Ser-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CCC(=O)N)N KVQOVQVGVKDZNW-GUBZILKMSA-N 0.000 description 9
- LHIPZASLKPYDPI-AVGNSLFASA-N Glu-Phe-Asp Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(O)=O)C(O)=O LHIPZASLKPYDPI-AVGNSLFASA-N 0.000 description 9
- NSTUFLGQJCOCDL-UWVGGRQHSA-N Gly-Leu-Arg Chemical compound NCC(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N NSTUFLGQJCOCDL-UWVGGRQHSA-N 0.000 description 9
- FGPLUIQCSKGLTI-WDSKDSINSA-N Gly-Ser-Glu Chemical compound NCC(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(O)=O FGPLUIQCSKGLTI-WDSKDSINSA-N 0.000 description 9
- ZZWUYQXMIFTIIY-WEDXCCLWSA-N Gly-Thr-Leu Chemical compound [H]NCC(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(O)=O ZZWUYQXMIFTIIY-WEDXCCLWSA-N 0.000 description 9
- DHMQDGOQFOQNFH-UHFFFAOYSA-N Glycine Natural products NCC(O)=O DHMQDGOQFOQNFH-UHFFFAOYSA-N 0.000 description 9
- ZPVJJPAIUZLSNE-DCAQKATOSA-N His-Arg-Ser Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(O)=O ZPVJJPAIUZLSNE-DCAQKATOSA-N 0.000 description 9
- 108010093488 His-His-His-His-His-His Proteins 0.000 description 9
- BFOGZWSSGMLYKV-DCAQKATOSA-N His-Ser-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CC1=CN=CN1)N BFOGZWSSGMLYKV-DCAQKATOSA-N 0.000 description 9
- SLQVFYWBGNNOTK-BYULHYEWSA-N Ile-Gly-Asn Chemical compound CC[C@H](C)[C@@H](C(=O)NCC(=O)N[C@@H](CC(=O)N)C(=O)O)N SLQVFYWBGNNOTK-BYULHYEWSA-N 0.000 description 9
- TWYOYAKMLHWMOJ-ZPFDUUQYSA-N Ile-Leu-Asn Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O TWYOYAKMLHWMOJ-ZPFDUUQYSA-N 0.000 description 9
- XHBYEMIUENPZLY-GMOBBJLQSA-N Ile-Pro-Asn Chemical compound CC[C@H](C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(O)=O XHBYEMIUENPZLY-GMOBBJLQSA-N 0.000 description 9
- WNGVUZWBXZKQES-YUMQZZPRSA-N Leu-Ala-Gly Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](C)C(=O)NCC(O)=O WNGVUZWBXZKQES-YUMQZZPRSA-N 0.000 description 9
- KZZCOWMDDXDKSS-CIUDSAMLSA-N Leu-Ser-Asn Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(N)=O)C(O)=O KZZCOWMDDXDKSS-CIUDSAMLSA-N 0.000 description 9
- IBQMEXQYZMVIFU-SRVKXCTJSA-N Lys-Asp-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CCCCN)N IBQMEXQYZMVIFU-SRVKXCTJSA-N 0.000 description 9
- IMAKMJCBYCSMHM-AVGNSLFASA-N Lys-Glu-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CCCCN IMAKMJCBYCSMHM-AVGNSLFASA-N 0.000 description 9
- DGNZGCQSVGGYJS-BQBZGAKWSA-N Met-Gly-Asp Chemical compound CSCC[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CC(O)=O DGNZGCQSVGGYJS-BQBZGAKWSA-N 0.000 description 9
- PESQCPHRXOFIPX-UHFFFAOYSA-N N-L-methionyl-L-tyrosine Natural products CSCCC(N)C(=O)NC(C(O)=O)CC1=CC=C(O)C=C1 PESQCPHRXOFIPX-UHFFFAOYSA-N 0.000 description 9
- JLLJTMHNXQTMCK-UBHSHLNASA-N Phe-Pro-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CC1=CC=CC=C1 JLLJTMHNXQTMCK-UBHSHLNASA-N 0.000 description 9
- CJZTUKSFZUSNCC-FXQIFTODSA-N Pro-Asp-Asn Chemical compound NC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H]1CCCN1 CJZTUKSFZUSNCC-FXQIFTODSA-N 0.000 description 9
- DXTOOBDIIAJZBJ-BQBZGAKWSA-N Pro-Gly-Ser Chemical compound [H]N1CCC[C@H]1C(=O)NCC(=O)N[C@@H](CO)C(O)=O DXTOOBDIIAJZBJ-BQBZGAKWSA-N 0.000 description 9
- STASJMBVVHNWCG-IHRRRGAJSA-N Pro-His-Leu Chemical compound C([C@@H](C(=O)N[C@@H](CC(C)C)C([O-])=O)NC(=O)[C@H]1[NH2+]CCC1)C1=CN=CN1 STASJMBVVHNWCG-IHRRRGAJSA-N 0.000 description 9
- MMAPOBOTRUVNKJ-ZLUOBGJFSA-N Ser-Asp-Ser Chemical compound C([C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)[C@H](CO)N)C(=O)O MMAPOBOTRUVNKJ-ZLUOBGJFSA-N 0.000 description 9
- FKYWFUYPVKLJLP-DCAQKATOSA-N Ser-Pro-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CO FKYWFUYPVKLJLP-DCAQKATOSA-N 0.000 description 9
- PYTKULIABVRXSC-BWBBJGPYSA-N Ser-Ser-Thr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(O)=O PYTKULIABVRXSC-BWBBJGPYSA-N 0.000 description 9
- IQPWNQRRAJHOKV-KATARQTJSA-N Thr-Ser-Lys Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCCCN IQPWNQRRAJHOKV-KATARQTJSA-N 0.000 description 9
- QHEGAOPHISYNDF-XDTLVQLUSA-N Tyr-Gln-Ala Chemical compound C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CC1=CC=C(C=C1)O)N QHEGAOPHISYNDF-XDTLVQLUSA-N 0.000 description 9
- DWAMXBFJNZIHMC-KBPBESRZSA-N Tyr-Leu-Gly Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(C)C)C(=O)NCC(O)=O DWAMXBFJNZIHMC-KBPBESRZSA-N 0.000 description 9
- ZXAGTABZUOMUDO-GVXVVHGQSA-N Val-Glu-Lys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CCCCN)C(=O)O)N ZXAGTABZUOMUDO-GVXVVHGQSA-N 0.000 description 9
- 108010069205 aspartyl-phenylalanine Proteins 0.000 description 9
- 108010053037 kyotorphin Proteins 0.000 description 9
- 108010051242 phenylalanylserine Proteins 0.000 description 9
- USNSOPDIZILSJP-FXQIFTODSA-N Arg-Asn-Asn Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O USNSOPDIZILSJP-FXQIFTODSA-N 0.000 description 8
- SQKPKIJVWHAWNF-DCAQKATOSA-N Arg-Asp-Lys Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CCCCN)C(O)=O SQKPKIJVWHAWNF-DCAQKATOSA-N 0.000 description 8
- KXOPYFNQLVUOAQ-FXQIFTODSA-N Arg-Ser-Ala Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O KXOPYFNQLVUOAQ-FXQIFTODSA-N 0.000 description 8
- DDBMKOCQWNFDBH-RHYQMDGZSA-N Arg-Thr-Lys Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N)O DDBMKOCQWNFDBH-RHYQMDGZSA-N 0.000 description 8
- LFWOQHSQNCKXRU-UFYCRDLUSA-N Arg-Tyr-Phe Chemical compound C([C@H](NC(=O)[C@H](CCCN=C(N)N)N)C(=O)N[C@@H](CC=1C=CC=CC=1)C(O)=O)C1=CC=C(O)C=C1 LFWOQHSQNCKXRU-UFYCRDLUSA-N 0.000 description 8
- LXTGAOAXPSJWOU-DCAQKATOSA-N Asn-Arg-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CC(=O)N)N LXTGAOAXPSJWOU-DCAQKATOSA-N 0.000 description 8
- LTZIRYMWOJHRCH-GUDRVLHUSA-N Asn-Ile-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CC(=O)N)N LTZIRYMWOJHRCH-GUDRVLHUSA-N 0.000 description 8
- FAEIQWHBRBWUBN-FXQIFTODSA-N Asp-Arg-Ser Chemical compound C(C[C@@H](C(=O)N[C@@H](CO)C(=O)O)NC(=O)[C@H](CC(=O)O)N)CN=C(N)N FAEIQWHBRBWUBN-FXQIFTODSA-N 0.000 description 8
- ILJQISGMGXRZQQ-IHRRRGAJSA-N Asp-Arg-Tyr Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O ILJQISGMGXRZQQ-IHRRRGAJSA-N 0.000 description 8
- OGTCOKZFOJIZFG-CIUDSAMLSA-N Asp-His-Asp Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(O)=O)C(O)=O OGTCOKZFOJIZFG-CIUDSAMLSA-N 0.000 description 8
- KFAFUJMGHVVYRC-DCAQKATOSA-N Asp-Leu-Met Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCSC)C(O)=O KFAFUJMGHVVYRC-DCAQKATOSA-N 0.000 description 8
- QJHOOKBAHRJPPX-QWRGUYRKSA-N Asp-Phe-Gly Chemical compound OC(=O)C[C@H](N)C(=O)N[C@H](C(=O)NCC(O)=O)CC1=CC=CC=C1 QJHOOKBAHRJPPX-QWRGUYRKSA-N 0.000 description 8
- KPSHWSWFPUDEGF-FXQIFTODSA-N Asp-Pro-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CC(O)=O KPSHWSWFPUDEGF-FXQIFTODSA-N 0.000 description 8
- UAXIKORUDGGIGA-DCAQKATOSA-N Asp-Pro-Lys Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CC(=O)O)N)C(=O)N[C@@H](CCCCN)C(=O)O UAXIKORUDGGIGA-DCAQKATOSA-N 0.000 description 8
- GIKOVDMXBAFXDF-NHCYSSNCSA-N Asp-Val-Leu Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O GIKOVDMXBAFXDF-NHCYSSNCSA-N 0.000 description 8
- BLGNLNRBABWDST-CIUDSAMLSA-N Cys-Leu-Asp Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)O)NC(=O)[C@H](CS)N BLGNLNRBABWDST-CIUDSAMLSA-N 0.000 description 8
- PRBLYKYHAJEABA-SRVKXCTJSA-N Gln-Arg-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(C)C)C(O)=O PRBLYKYHAJEABA-SRVKXCTJSA-N 0.000 description 8
- CITDWMLWXNUQKD-FXQIFTODSA-N Gln-Gln-Asn Chemical compound C(CC(=O)N)[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CC(=O)N)C(=O)O)N CITDWMLWXNUQKD-FXQIFTODSA-N 0.000 description 8
- ZEEPYMXTJWIMSN-GUBZILKMSA-N Gln-Lys-Ser Chemical compound NCCCC[C@@H](C(=O)N[C@@H](CO)C(O)=O)NC(=O)[C@@H](N)CCC(N)=O ZEEPYMXTJWIMSN-GUBZILKMSA-N 0.000 description 8
- NYCVMJGIJYQWDO-CIUDSAMLSA-N Gln-Ser-Arg Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O NYCVMJGIJYQWDO-CIUDSAMLSA-N 0.000 description 8
- GLWXKFRTOHKGIT-ACZMJKKPSA-N Glu-Asn-Asn Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O GLWXKFRTOHKGIT-ACZMJKKPSA-N 0.000 description 8
- JVWPPCWUDRJGAE-YUMQZZPRSA-N Gly-Asn-Leu Chemical compound [H]NCC(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O JVWPPCWUDRJGAE-YUMQZZPRSA-N 0.000 description 8
- QGZSAHIZRQHCEQ-QWRGUYRKSA-N Gly-Asp-Tyr Chemical compound NCC(=O)N[C@@H](CC(O)=O)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 QGZSAHIZRQHCEQ-QWRGUYRKSA-N 0.000 description 8
- YTSVAIMKVLZUDU-YUMQZZPRSA-N Gly-Leu-Asp Chemical compound [H]NCC(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O YTSVAIMKVLZUDU-YUMQZZPRSA-N 0.000 description 8
- CCBIBMKQNXHNIN-ZETCQYMHSA-N Gly-Leu-Gly Chemical compound NCC(=O)N[C@@H](CC(C)C)C(=O)NCC(O)=O CCBIBMKQNXHNIN-ZETCQYMHSA-N 0.000 description 8
- WNGHUXFWEWTKAO-YUMQZZPRSA-N Gly-Ser-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)CN WNGHUXFWEWTKAO-YUMQZZPRSA-N 0.000 description 8
- LCRDMSSAKLTKBU-ZDLURKLDSA-N Gly-Ser-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)CN LCRDMSSAKLTKBU-ZDLURKLDSA-N 0.000 description 8
- 102000019058 Glycogen Synthase Kinase 3 beta Human genes 0.000 description 8
- 108010051975 Glycogen Synthase Kinase 3 beta Proteins 0.000 description 8
- YADRBUZBKHHDAO-XPUUQOCRSA-N His-Gly-Ala Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)NCC(=O)N[C@@H](C)C(O)=O YADRBUZBKHHDAO-XPUUQOCRSA-N 0.000 description 8
- XDIVYNSPYBLSME-DCAQKATOSA-N His-Met-Asp Chemical compound CSCC[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)O)NC(=O)[C@H](CC1=CN=CN1)N XDIVYNSPYBLSME-DCAQKATOSA-N 0.000 description 8
- FFKJUTZARGRVTH-KKUMJFAQSA-N His-Ser-Tyr Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O FFKJUTZARGRVTH-KKUMJFAQSA-N 0.000 description 8
- DGVYSZUCRYXKOJ-XIRDDKMYSA-N His-Trp-Cys Chemical compound C1=CC=C2C(=C1)C(=CN2)C[C@@H](C(=O)N[C@@H](CS)C(=O)O)NC(=O)[C@H](CC3=CN=CN3)N DGVYSZUCRYXKOJ-XIRDDKMYSA-N 0.000 description 8
- SYPULFZAGBBIOM-GVXVVHGQSA-N His-Val-Glu Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)O)NC(=O)[C@H](CC1=CN=CN1)N SYPULFZAGBBIOM-GVXVVHGQSA-N 0.000 description 8
- JRHFQUPIZOYKQP-KBIXCLLPSA-N Ile-Ala-Glu Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCC(O)=O JRHFQUPIZOYKQP-KBIXCLLPSA-N 0.000 description 8
- GYAFMRQGWHXMII-IUKAMOBKSA-N Ile-Asp-Thr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H]([C@@H](C)O)C(=O)O)N GYAFMRQGWHXMII-IUKAMOBKSA-N 0.000 description 8
- BATWGBRIZANGPN-ZPFDUUQYSA-N Ile-Pro-Gln Chemical compound CC[C@H](C)[C@@H](C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(=O)N)C(=O)O)N BATWGBRIZANGPN-ZPFDUUQYSA-N 0.000 description 8
- 108010065920 Insulin Lispro Proteins 0.000 description 8
- DCGXHWINSHEPIR-SRVKXCTJSA-N Leu-Lys-Cys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CS)C(=O)O)N DCGXHWINSHEPIR-SRVKXCTJSA-N 0.000 description 8
- YWKNKRAKOCLOLH-OEAJRASXSA-N Leu-Phe-Thr Chemical compound CC(C)C[C@H](N)C(=O)N[C@H](C(=O)N[C@@H]([C@@H](C)O)C(O)=O)CC1=CC=CC=C1 YWKNKRAKOCLOLH-OEAJRASXSA-N 0.000 description 8
- QMKFDEUJGYNFMC-AVGNSLFASA-N Leu-Pro-Arg Chemical compound CC(C)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCCN=C(N)N)C(O)=O QMKFDEUJGYNFMC-AVGNSLFASA-N 0.000 description 8
- SVBJIZVVYJYGLA-DCAQKATOSA-N Leu-Ser-Val Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(O)=O SVBJIZVVYJYGLA-DCAQKATOSA-N 0.000 description 8
- AEDWWMMHUGYIFD-HJGDQZAQSA-N Leu-Thr-Asn Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(N)=O)C(O)=O AEDWWMMHUGYIFD-HJGDQZAQSA-N 0.000 description 8
- XFIHDSBIPWEYJJ-YUMQZZPRSA-N Lys-Ala-Gly Chemical compound OC(=O)CNC(=O)[C@H](C)NC(=O)[C@@H](N)CCCCN XFIHDSBIPWEYJJ-YUMQZZPRSA-N 0.000 description 8
- FACUGMGEFUEBTI-SRVKXCTJSA-N Lys-Asn-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CCCCN FACUGMGEFUEBTI-SRVKXCTJSA-N 0.000 description 8
- PINHPJWGVBKQII-SRVKXCTJSA-N Lys-Leu-Cys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CS)C(=O)O)NC(=O)[C@H](CCCCN)N PINHPJWGVBKQII-SRVKXCTJSA-N 0.000 description 8
- XIZQPFCRXLUNMK-BZSNNMDCSA-N Lys-Leu-Phe Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)O)NC(=O)[C@H](CCCCN)N XIZQPFCRXLUNMK-BZSNNMDCSA-N 0.000 description 8
- GHKXHCMRAUYLBS-CIUDSAMLSA-N Lys-Ser-Asn Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(N)=O)C(O)=O GHKXHCMRAUYLBS-CIUDSAMLSA-N 0.000 description 8
- MIMXMVDLMDMOJD-BZSNNMDCSA-N Lys-Tyr-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(C)C)C(O)=O MIMXMVDLMDMOJD-BZSNNMDCSA-N 0.000 description 8
- HLQWFLJOJRFXHO-CIUDSAMLSA-N Met-Glu-Ser Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(O)=O HLQWFLJOJRFXHO-CIUDSAMLSA-N 0.000 description 8
- 102100039124 Methyl-CpG-binding protein 2 Human genes 0.000 description 8
- 241001529936 Murinae Species 0.000 description 8
- KIEPQOIQHFKQLK-PCBIJLKTSA-N Phe-Asn-Ile Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O KIEPQOIQHFKQLK-PCBIJLKTSA-N 0.000 description 8
- DJPXNKUDJKGQEE-BZSNNMDCSA-N Phe-Asp-Phe Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O DJPXNKUDJKGQEE-BZSNNMDCSA-N 0.000 description 8
- BVHFFNYBKRTSIU-MEYUZBJRSA-N Phe-His-Thr Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H]([C@@H](C)O)C(O)=O BVHFFNYBKRTSIU-MEYUZBJRSA-N 0.000 description 8
- OQTDZEJJWWAGJT-KKUMJFAQSA-N Phe-Lys-Asp Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(O)=O OQTDZEJJWWAGJT-KKUMJFAQSA-N 0.000 description 8
- XNMYNGDKJNOKHH-BZSNNMDCSA-N Phe-Ser-Tyr Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O XNMYNGDKJNOKHH-BZSNNMDCSA-N 0.000 description 8
- LCUOTSLIVGSGAU-AVGNSLFASA-N Pro-His-Arg Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O LCUOTSLIVGSGAU-AVGNSLFASA-N 0.000 description 8
- GOMUXSCOIWIJFP-GUBZILKMSA-N Pro-Ser-Arg Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O GOMUXSCOIWIJFP-GUBZILKMSA-N 0.000 description 8
- BGWKULMLUIUPKY-BQBZGAKWSA-N Pro-Ser-Gly Chemical compound OC(=O)CNC(=O)[C@H](CO)NC(=O)[C@@H]1CCCN1 BGWKULMLUIUPKY-BQBZGAKWSA-N 0.000 description 8
- ZMLRZBWCXPQADC-TUAOUCFPSA-N Pro-Val-Pro Chemical compound CC(C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@@H]2CCCN2 ZMLRZBWCXPQADC-TUAOUCFPSA-N 0.000 description 8
- WTUJZHKANPDPIN-CIUDSAMLSA-N Ser-Ala-Lys Chemical compound C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CO)N WTUJZHKANPDPIN-CIUDSAMLSA-N 0.000 description 8
- VQBLHWSPVYYZTB-DCAQKATOSA-N Ser-Arg-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CO)N VQBLHWSPVYYZTB-DCAQKATOSA-N 0.000 description 8
- FIDMVVBUOCMMJG-CIUDSAMLSA-N Ser-Asn-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CO FIDMVVBUOCMMJG-CIUDSAMLSA-N 0.000 description 8
- RDFQNDHEHVSONI-ZLUOBGJFSA-N Ser-Asn-Ser Chemical compound OC[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(O)=O RDFQNDHEHVSONI-ZLUOBGJFSA-N 0.000 description 8
- MESDJCNHLZBMEP-ZLUOBGJFSA-N Ser-Asp-Asp Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O MESDJCNHLZBMEP-ZLUOBGJFSA-N 0.000 description 8
- FMDHKPRACUXATF-ACZMJKKPSA-N Ser-Gln-Ser Chemical compound OC[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(O)=O FMDHKPRACUXATF-ACZMJKKPSA-N 0.000 description 8
- SMIDBHKWSYUBRZ-ACZMJKKPSA-N Ser-Glu-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(O)=O SMIDBHKWSYUBRZ-ACZMJKKPSA-N 0.000 description 8
- RJHJPZQOMKCSTP-CIUDSAMLSA-N Ser-His-Asn Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(N)=O)C(O)=O RJHJPZQOMKCSTP-CIUDSAMLSA-N 0.000 description 8
- IOVBCLGAJJXOHK-SRVKXCTJSA-N Ser-His-His Chemical compound C([C@H](NC(=O)[C@H](CO)N)C(=O)N[C@@H](CC=1NC=NC=1)C(O)=O)C1=CN=CN1 IOVBCLGAJJXOHK-SRVKXCTJSA-N 0.000 description 8
- GJFYFGOEWLDQGW-GUBZILKMSA-N Ser-Leu-Gln Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CO)N GJFYFGOEWLDQGW-GUBZILKMSA-N 0.000 description 8
- WOJYIMBIKTWKJO-KKUMJFAQSA-N Ser-Phe-His Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)NC(=O)[C@H](CO)N WOJYIMBIKTWKJO-KKUMJFAQSA-N 0.000 description 8
- NUEHQDHDLDXCRU-GUBZILKMSA-N Ser-Pro-Arg Chemical compound OC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCCN=C(N)N)C(O)=O NUEHQDHDLDXCRU-GUBZILKMSA-N 0.000 description 8
- KQNDIKOYWZTZIX-FXQIFTODSA-N Ser-Ser-Arg Chemical compound OC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCCNC(N)=N KQNDIKOYWZTZIX-FXQIFTODSA-N 0.000 description 8
- NVNPWELENFJOHH-CIUDSAMLSA-N Ser-Ser-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CO)N NVNPWELENFJOHH-CIUDSAMLSA-N 0.000 description 8
- NADLKBTYNKUJEP-KATARQTJSA-N Ser-Thr-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(O)=O NADLKBTYNKUJEP-KATARQTJSA-N 0.000 description 8
- FGBLCMLXHRPVOF-IHRRRGAJSA-N Ser-Tyr-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O FGBLCMLXHRPVOF-IHRRRGAJSA-N 0.000 description 8
- JGUWRQWULDWNCM-FXQIFTODSA-N Ser-Val-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CO)C(O)=O JGUWRQWULDWNCM-FXQIFTODSA-N 0.000 description 8
- JWQNAFHCXKVZKZ-UVOCVTCTSA-N Thr-Lys-Thr Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)O)C(O)=O JWQNAFHCXKVZKZ-UVOCVTCTSA-N 0.000 description 8
- GVMXJJAJLIEASL-ZJDVBMNYSA-N Thr-Pro-Thr Chemical compound C[C@@H](O)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(O)=O GVMXJJAJLIEASL-ZJDVBMNYSA-N 0.000 description 8
- COYHRQWNJDJCNA-NUJDXYNKSA-N Thr-Thr-Thr Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O COYHRQWNJDJCNA-NUJDXYNKSA-N 0.000 description 8
- PRONOHBTMLNXCZ-BZSNNMDCSA-N Tyr-Leu-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 PRONOHBTMLNXCZ-BZSNNMDCSA-N 0.000 description 8
- ABSXSJZNRAQDDI-KJEVXHAQSA-N Tyr-Val-Thr Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O ABSXSJZNRAQDDI-KJEVXHAQSA-N 0.000 description 8
- DDRBQONWVBDQOY-GUBZILKMSA-N Val-Ala-Arg Chemical compound CC(C)[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O DDRBQONWVBDQOY-GUBZILKMSA-N 0.000 description 8
- SLLKXDSRVAOREO-KZVJFYERSA-N Val-Ala-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](C)NC(=O)[C@H](C(C)C)N)O SLLKXDSRVAOREO-KZVJFYERSA-N 0.000 description 8
- GNWUWQAVVJQREM-NHCYSSNCSA-N Val-Asn-His Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N GNWUWQAVVJQREM-NHCYSSNCSA-N 0.000 description 8
- 108010047495 alanylglycine Proteins 0.000 description 8
- 108010068380 arginylarginine Proteins 0.000 description 8
- 238000012217 deletion Methods 0.000 description 8
- 230000037430 deletion Effects 0.000 description 8
- XBGGUPMXALFZOT-UHFFFAOYSA-N glycyl-L-tyrosine hemihydrate Natural products NCC(=O)NC(C(O)=O)CC1=CC=C(O)C=C1 XBGGUPMXALFZOT-UHFFFAOYSA-N 0.000 description 8
- 108010087823 glycyltyrosine Proteins 0.000 description 8
- 108010025306 histidylleucine Proteins 0.000 description 8
- 108010092114 histidylphenylalanine Proteins 0.000 description 8
- VBRDBGCROKWTPV-XHNCKOQMSA-N Ala-Glu-Pro Chemical compound C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N1CCC[C@@H]1C(=O)O)N VBRDBGCROKWTPV-XHNCKOQMSA-N 0.000 description 7
- BFMIRJBURUXDRG-DLOVCJGASA-N Ala-Phe-Asp Chemical compound OC(=O)C[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)C)CC1=CC=CC=C1 BFMIRJBURUXDRG-DLOVCJGASA-N 0.000 description 7
- RVDVDRUZWZIBJQ-CIUDSAMLSA-N Arg-Asn-Glu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O RVDVDRUZWZIBJQ-CIUDSAMLSA-N 0.000 description 7
- OBFTYSPXDRROQO-SRVKXCTJSA-N Arg-Gln-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CCC(N)=O)NC(=O)[C@@H](N)CCCN=C(N)N OBFTYSPXDRROQO-SRVKXCTJSA-N 0.000 description 7
- GRRXPUAICOGISM-RWMBFGLXSA-N Arg-Lys-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCCN)NC(=O)[C@H](CCCN=C(N)N)N)C(=O)O GRRXPUAICOGISM-RWMBFGLXSA-N 0.000 description 7
- FAEFJTCTNZTPHX-ACZMJKKPSA-N Asn-Gln-Ala Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(O)=O FAEFJTCTNZTPHX-ACZMJKKPSA-N 0.000 description 7
- OLGCWMNDJTWQAG-GUBZILKMSA-N Asn-Glu-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CC(N)=O OLGCWMNDJTWQAG-GUBZILKMSA-N 0.000 description 7
- HPNDKUOLNRVRAY-BIIVOSGPSA-N Asn-Ser-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CO)NC(=O)[C@H](CC(=O)N)N)C(=O)O HPNDKUOLNRVRAY-BIIVOSGPSA-N 0.000 description 7
- DTNUIAJCPRMNBT-WHFBIAKZSA-N Asp-Gly-Ala Chemical compound [H]N[C@@H](CC(O)=O)C(=O)NCC(=O)N[C@@H](C)C(O)=O DTNUIAJCPRMNBT-WHFBIAKZSA-N 0.000 description 7
- ORRJQLIATJDMQM-HJGDQZAQSA-N Asp-Leu-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC(O)=O ORRJQLIATJDMQM-HJGDQZAQSA-N 0.000 description 7
- QTIZKMMLNUMHHU-DCAQKATOSA-N Asp-Pro-His Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CC(=O)O)N)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O QTIZKMMLNUMHHU-DCAQKATOSA-N 0.000 description 7
- MJJIHRWNWSQTOI-VEVYYDQMSA-N Asp-Thr-Arg Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H]([C@H](O)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O MJJIHRWNWSQTOI-VEVYYDQMSA-N 0.000 description 7
- 102100036279 DNA (cytosine-5)-methyltransferase 1 Human genes 0.000 description 7
- NKCZYEDZTKOFBG-GUBZILKMSA-N Gln-Gln-Arg Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O NKCZYEDZTKOFBG-GUBZILKMSA-N 0.000 description 7
- QQAPDATZKKTBIY-YUMQZZPRSA-N Gln-Gly-Met Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)NCC(=O)N[C@@H](CCSC)C(O)=O QQAPDATZKKTBIY-YUMQZZPRSA-N 0.000 description 7
- MUSGDMDGNGXULI-DCAQKATOSA-N Glu-Glu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CCC(O)=O MUSGDMDGNGXULI-DCAQKATOSA-N 0.000 description 7
- UJMNFCAHLYKWOZ-DCAQKATOSA-N Glu-Lys-Gln Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(N)=O)C(O)=O UJMNFCAHLYKWOZ-DCAQKATOSA-N 0.000 description 7
- YHOJJFFTSMWVGR-HJGDQZAQSA-N Glu-Met-Thr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCSC)C(=O)N[C@@H]([C@@H](C)O)C(O)=O YHOJJFFTSMWVGR-HJGDQZAQSA-N 0.000 description 7
- FFJQHWKSGAWSTJ-BFHQHQDPSA-N Gly-Thr-Ala Chemical compound [H]NCC(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(O)=O FFJQHWKSGAWSTJ-BFHQHQDPSA-N 0.000 description 7
- KNNSUUOHFVVJOP-GUBZILKMSA-N His-Glu-Ser Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CO)C(=O)O)N KNNSUUOHFVVJOP-GUBZILKMSA-N 0.000 description 7
- IAYPZSHNZQHQNO-KKUMJFAQSA-N His-Ser-Phe Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CC2=CN=CN2)N IAYPZSHNZQHQNO-KKUMJFAQSA-N 0.000 description 7
- JUCZDDVZBMPKRT-IXOXFDKPSA-N His-Thr-Lys Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CC1=CN=CN1)N)O JUCZDDVZBMPKRT-IXOXFDKPSA-N 0.000 description 7
- ULXYQAJWJGLCNR-YUMQZZPRSA-N Leu-Asp-Gly Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)NCC(O)=O ULXYQAJWJGLCNR-YUMQZZPRSA-N 0.000 description 7
- VULJUQZPSOASBZ-SRVKXCTJSA-N Leu-Pro-Glu Chemical compound [H]N[C@@H](CC(C)C)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(O)=O)C(O)=O VULJUQZPSOASBZ-SRVKXCTJSA-N 0.000 description 7
- IRMLZWSRWSGTOP-CIUDSAMLSA-N Leu-Ser-Ala Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O IRMLZWSRWSGTOP-CIUDSAMLSA-N 0.000 description 7
- GHOIOYHDDKXIDX-SZMVWBNQSA-N Lys-Glu-Trp Chemical compound C1=CC=C2C(C[C@H](NC(=O)[C@H](CCC(O)=O)NC(=O)[C@@H](N)CCCCN)C(O)=O)=CNC2=C1 GHOIOYHDDKXIDX-SZMVWBNQSA-N 0.000 description 7
- ZJSZPXISKMDJKQ-JYJNAYRXSA-N Lys-Phe-Glu Chemical compound NCCCC[C@H](N)C(=O)N[C@H](C(=O)N[C@@H](CCC(O)=O)C(O)=O)CC1=CC=CC=C1 ZJSZPXISKMDJKQ-JYJNAYRXSA-N 0.000 description 7
- ZUGVARDEGWMMLK-SRVKXCTJSA-N Lys-Ser-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCCCN ZUGVARDEGWMMLK-SRVKXCTJSA-N 0.000 description 7
- WAAZECNCPVGPIV-RHYQMDGZSA-N Lys-Thr-Met Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCSC)C(O)=O WAAZECNCPVGPIV-RHYQMDGZSA-N 0.000 description 7
- TZLYIHDABYBOCJ-FXQIFTODSA-N Met-Asp-Ser Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O TZLYIHDABYBOCJ-FXQIFTODSA-N 0.000 description 7
- LLGTYVHITPVGKR-RYUDHWBXSA-N Phe-Gln-Gly Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CCC(N)=O)C(=O)NCC(O)=O LLGTYVHITPVGKR-RYUDHWBXSA-N 0.000 description 7
- SKICPQLTOXGWGO-GARJFASQSA-N Pro-Gln-Pro Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CCC(=O)N)C(=O)N2CCC[C@@H]2C(=O)O SKICPQLTOXGWGO-GARJFASQSA-N 0.000 description 7
- AFXCXDQNRXTSBD-FJXKBIBVSA-N Pro-Gly-Thr Chemical compound [H]N1CCC[C@H]1C(=O)NCC(=O)N[C@@H]([C@@H](C)O)C(O)=O AFXCXDQNRXTSBD-FJXKBIBVSA-N 0.000 description 7
- JUJGNDZIKKQMDJ-IHRRRGAJSA-N Pro-His-His Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC1=CNC=N1)C(O)=O JUJGNDZIKKQMDJ-IHRRRGAJSA-N 0.000 description 7
- MRYUJHGPZQNOAD-IHRRRGAJSA-N Pro-Leu-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@@H]1CCCN1 MRYUJHGPZQNOAD-IHRRRGAJSA-N 0.000 description 7
- GFHXZNVJIKMAGO-IHRRRGAJSA-N Pro-Phe-Ser Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CO)C(O)=O GFHXZNVJIKMAGO-IHRRRGAJSA-N 0.000 description 7
- NLQUOHDCLSFABG-GUBZILKMSA-N Ser-Arg-Arg Chemical compound NC(N)=NCCC[C@H](NC(=O)[C@H](CO)N)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O NLQUOHDCLSFABG-GUBZILKMSA-N 0.000 description 7
- IXUGADGDCQDLSA-FXQIFTODSA-N Ser-Gln-Gln Chemical compound C(CC(=O)N)[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CO)N IXUGADGDCQDLSA-FXQIFTODSA-N 0.000 description 7
- AEGUWTFAQQWVLC-BQBZGAKWSA-N Ser-Gly-Arg Chemical compound [H]N[C@@H](CO)C(=O)NCC(=O)N[C@@H](CCCNC(N)=N)C(O)=O AEGUWTFAQQWVLC-BQBZGAKWSA-N 0.000 description 7
- IFPBAGJBHSNYPR-ZKWXMUAHSA-N Ser-Ile-Gly Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)CC)C(=O)NCC(O)=O IFPBAGJBHSNYPR-ZKWXMUAHSA-N 0.000 description 7
- VGQVAVQWKJLIRM-FXQIFTODSA-N Ser-Ser-Val Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(O)=O VGQVAVQWKJLIRM-FXQIFTODSA-N 0.000 description 7
- XJDMUQCLVSCRSJ-VZFHVOOUSA-N Ser-Thr-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(O)=O XJDMUQCLVSCRSJ-VZFHVOOUSA-N 0.000 description 7
- FHXGMDRKJHKLKW-QWRGUYRKSA-N Ser-Tyr-Gly Chemical compound OC[C@H](N)C(=O)N[C@H](C(=O)NCC(O)=O)CC1=CC=C(O)C=C1 FHXGMDRKJHKLKW-QWRGUYRKSA-N 0.000 description 7
- YEDSOSIKVUMIJE-DCAQKATOSA-N Ser-Val-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O YEDSOSIKVUMIJE-DCAQKATOSA-N 0.000 description 7
- WPSKTVVMQCXPRO-BWBBJGPYSA-N Thr-Ser-Ser Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O WPSKTVVMQCXPRO-BWBBJGPYSA-N 0.000 description 7
- QYDKSNXSBXZPFK-ZJDVBMNYSA-N Thr-Thr-Arg Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O QYDKSNXSBXZPFK-ZJDVBMNYSA-N 0.000 description 7
- OEVJGIHPQOXYFE-SRVKXCTJSA-N Tyr-Asn-Asp Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O OEVJGIHPQOXYFE-SRVKXCTJSA-N 0.000 description 7
- RIFVTNDKUMSSMN-ULQDDVLXSA-N Tyr-His-Val Chemical compound CC(C)[C@H](NC(=O)[C@H](Cc1c[nH]cn1)NC(=O)[C@@H](N)Cc1ccc(O)cc1)C(O)=O RIFVTNDKUMSSMN-ULQDDVLXSA-N 0.000 description 7
- WQOHKVRQDLNDIL-YJRXYDGGSA-N Tyr-Thr-Ser Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(O)=O WQOHKVRQDLNDIL-YJRXYDGGSA-N 0.000 description 7
- SVFRYKBZHUGKLP-QXEWZRGKSA-N Val-Met-Asn Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(=O)N)C(=O)O)N SVFRYKBZHUGKLP-QXEWZRGKSA-N 0.000 description 7
- MNSSBIHFEUUXNW-RCWTZXSCSA-N Val-Thr-Arg Chemical compound CC(C)[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@H](C(O)=O)CCCN=C(N)N MNSSBIHFEUUXNW-RCWTZXSCSA-N 0.000 description 7
- 238000011161 development Methods 0.000 description 7
- 230000018109 developmental process Effects 0.000 description 7
- 108010073025 phenylalanylphenylalanine Proteins 0.000 description 7
- ZVFVBBGVOILKPO-WHFBIAKZSA-N Ala-Gly-Ala Chemical compound C[C@H](N)C(=O)NCC(=O)N[C@@H](C)C(O)=O ZVFVBBGVOILKPO-WHFBIAKZSA-N 0.000 description 6
- OBVSBEYOMDWLRJ-BFHQHQDPSA-N Ala-Gly-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)CNC(=O)[C@H](C)N OBVSBEYOMDWLRJ-BFHQHQDPSA-N 0.000 description 6
- SUMYEVXWCAYLLJ-GUBZILKMSA-N Ala-Leu-Gln Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O SUMYEVXWCAYLLJ-GUBZILKMSA-N 0.000 description 6
- BTRULDJUUVGRNE-DCAQKATOSA-N Ala-Pro-Lys Chemical compound C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCCCN)C(O)=O BTRULDJUUVGRNE-DCAQKATOSA-N 0.000 description 6
- KLALXKYLOMZDQT-ZLUOBGJFSA-N Ala-Ser-Asn Chemical compound C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CC(N)=O KLALXKYLOMZDQT-ZLUOBGJFSA-N 0.000 description 6
- ZEAYJGRKRUBDOB-GARJFASQSA-N Arg-Gln-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CCCN=C(N)N)N)C(=O)O ZEAYJGRKRUBDOB-GARJFASQSA-N 0.000 description 6
- PBSOQGZLPFVXPU-YUMQZZPRSA-N Arg-Glu-Gly Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O PBSOQGZLPFVXPU-YUMQZZPRSA-N 0.000 description 6
- NVCIXQYNWYTLDO-IHRRRGAJSA-N Arg-His-Leu Chemical compound CC(C)C[C@@H](C(=O)O)NC(=O)[C@H](CC1=CN=CN1)NC(=O)[C@H](CCCN=C(N)N)N NVCIXQYNWYTLDO-IHRRRGAJSA-N 0.000 description 6
- XSPKAHFVDKRGRL-DCAQKATOSA-N Arg-Pro-Glu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(O)=O)C(O)=O XSPKAHFVDKRGRL-DCAQKATOSA-N 0.000 description 6
- AMIQZQAAYGYKOP-FXQIFTODSA-N Arg-Ser-Asn Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(N)=O)C(O)=O AMIQZQAAYGYKOP-FXQIFTODSA-N 0.000 description 6
- VRTWYUYCJGNFES-CIUDSAMLSA-N Arg-Ser-Gln Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(O)=O VRTWYUYCJGNFES-CIUDSAMLSA-N 0.000 description 6
- ZDOQDYFZNGASEY-BIIVOSGPSA-N Asn-Asp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)O)NC(=O)[C@H](CC(=O)N)N)C(=O)O ZDOQDYFZNGASEY-BIIVOSGPSA-N 0.000 description 6
- GNKVBRYFXYWXAB-WDSKDSINSA-N Asn-Glu-Gly Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O GNKVBRYFXYWXAB-WDSKDSINSA-N 0.000 description 6
- UJGRZQYSNYTCAX-SRVKXCTJSA-N Asp-Leu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC(O)=O UJGRZQYSNYTCAX-SRVKXCTJSA-N 0.000 description 6
- HRVQDZOWMLFAOD-BIIVOSGPSA-N Asp-Ser-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CO)NC(=O)[C@H](CC(=O)O)N)C(=O)O HRVQDZOWMLFAOD-BIIVOSGPSA-N 0.000 description 6
- KBJVTFWQWXCYCQ-IUKAMOBKSA-N Asp-Thr-Ile Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O KBJVTFWQWXCYCQ-IUKAMOBKSA-N 0.000 description 6
- 108020004414 DNA Proteins 0.000 description 6
- VGTDBGYFVWOQTI-RYUDHWBXSA-N Gln-Gly-Phe Chemical compound NC(=O)CC[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 VGTDBGYFVWOQTI-RYUDHWBXSA-N 0.000 description 6
- XFAUJGNLHIGXET-AVGNSLFASA-N Gln-Leu-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O XFAUJGNLHIGXET-AVGNSLFASA-N 0.000 description 6
- LGWNISYVKDNJRP-FXQIFTODSA-N Gln-Ser-Gln Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(O)=O LGWNISYVKDNJRP-FXQIFTODSA-N 0.000 description 6
- KPNWAJMEMRCLAL-GUBZILKMSA-N Gln-Ser-Lys Chemical compound C(CCN)C[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CCC(=O)N)N KPNWAJMEMRCLAL-GUBZILKMSA-N 0.000 description 6
- HLRLXVPRJJITSK-IFFSRLJSSA-N Gln-Thr-Val Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C(C)C)C(O)=O HLRLXVPRJJITSK-IFFSRLJSSA-N 0.000 description 6
- FHPXTPQBODWBIY-CIUDSAMLSA-N Glu-Ala-Arg Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O FHPXTPQBODWBIY-CIUDSAMLSA-N 0.000 description 6
- JJKKWYQVHRUSDG-GUBZILKMSA-N Glu-Ala-Lys Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCCN)C(O)=O JJKKWYQVHRUSDG-GUBZILKMSA-N 0.000 description 6
- VXKCPBPQEKKERH-IUCAKERBSA-N Gly-Arg-Pro Chemical compound NC(N)=NCCC[C@H](NC(=O)CN)C(=O)N1CCC[C@H]1C(O)=O VXKCPBPQEKKERH-IUCAKERBSA-N 0.000 description 6
- HFXJIZNEXNIZIJ-BQBZGAKWSA-N Gly-Glu-Gln Chemical compound NCC(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O HFXJIZNEXNIZIJ-BQBZGAKWSA-N 0.000 description 6
- NSVOVKWEKGEOQB-LURJTMIESA-N Gly-Pro-Gly Chemical compound NCC(=O)N1CCC[C@H]1C(=O)NCC(O)=O NSVOVKWEKGEOQB-LURJTMIESA-N 0.000 description 6
- 239000004471 Glycine Substances 0.000 description 6
- YAALVYQFVJNXIV-KKUMJFAQSA-N His-Leu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC1=CN=CN1 YAALVYQFVJNXIV-KKUMJFAQSA-N 0.000 description 6
- ZVKDCQVQTGYBQT-LSJOCFKGSA-N His-Pro-Ala Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C)C(O)=O ZVKDCQVQTGYBQT-LSJOCFKGSA-N 0.000 description 6
- KAXZXLSXFWSNNZ-XVYDVKMFSA-N His-Ser-Ala Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O KAXZXLSXFWSNNZ-XVYDVKMFSA-N 0.000 description 6
- STGQSBKUYSPPIG-CIUDSAMLSA-N His-Ser-Asp Chemical compound OC(=O)C[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H](N)CC1=CN=CN1 STGQSBKUYSPPIG-CIUDSAMLSA-N 0.000 description 6
- PZAJPILZRFPYJJ-SRVKXCTJSA-N His-Ser-Leu Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O PZAJPILZRFPYJJ-SRVKXCTJSA-N 0.000 description 6
- KMBPQYKVZBMRMH-PEFMBERDSA-N Ile-Gln-Asn Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O KMBPQYKVZBMRMH-PEFMBERDSA-N 0.000 description 6
- IGJWJGIHUFQANP-LAEOZQHASA-N Ile-Gly-Gln Chemical compound CC[C@H](C)[C@@H](C(=O)NCC(=O)N[C@@H](CCC(=O)N)C(=O)O)N IGJWJGIHUFQANP-LAEOZQHASA-N 0.000 description 6
- DPURXCQCHSQPAN-AVGNSLFASA-N Leu-Pro-Pro Chemical compound CC(C)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 DPURXCQCHSQPAN-AVGNSLFASA-N 0.000 description 6
- PRSBSVAVOQOAMI-BJDJZHNGSA-N Lys-Ile-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)CC)NC(=O)[C@@H](N)CCCCN PRSBSVAVOQOAMI-BJDJZHNGSA-N 0.000 description 6
- AIRZWUMAHCDDHR-KKUMJFAQSA-N Lys-Leu-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O AIRZWUMAHCDDHR-KKUMJFAQSA-N 0.000 description 6
- YDDDRTIPNTWGIG-SRVKXCTJSA-N Lys-Lys-Ser Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(O)=O YDDDRTIPNTWGIG-SRVKXCTJSA-N 0.000 description 6
- MIROMRNASYKZNL-ULQDDVLXSA-N Lys-Pro-Tyr Chemical compound NCCCC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 MIROMRNASYKZNL-ULQDDVLXSA-N 0.000 description 6
- LKDXINHHSWFFJC-SRVKXCTJSA-N Lys-Ser-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CCCCN)N LKDXINHHSWFFJC-SRVKXCTJSA-N 0.000 description 6
- JOSAKOKSPXROGQ-BJDJZHNGSA-N Lys-Ser-Ile Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O JOSAKOKSPXROGQ-BJDJZHNGSA-N 0.000 description 6
- ZBLSZPYQQRIHQU-RCWTZXSCSA-N Met-Thr-Val Chemical compound CSCC[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C(C)C)C(O)=O ZBLSZPYQQRIHQU-RCWTZXSCSA-N 0.000 description 6
- 108010072388 Methyl-CpG-Binding Protein 2 Proteins 0.000 description 6
- 241000699670 Mus sp. Species 0.000 description 6
- OBVCYFIHIIYIQF-CIUDSAMLSA-N Pro-Asn-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O OBVCYFIHIIYIQF-CIUDSAMLSA-N 0.000 description 6
- FUVBEZJCRMHWEM-FXQIFTODSA-N Pro-Asn-Ser Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(O)=O FUVBEZJCRMHWEM-FXQIFTODSA-N 0.000 description 6
- AQSMZTIEJMZQEC-DCAQKATOSA-N Pro-His-Ser Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CC2=CN=CN2)C(=O)N[C@@H](CO)C(=O)O AQSMZTIEJMZQEC-DCAQKATOSA-N 0.000 description 6
- FYPGHGXAOZTOBO-IHRRRGAJSA-N Pro-Leu-His Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@@H]2CCCN2 FYPGHGXAOZTOBO-IHRRRGAJSA-N 0.000 description 6
- ULWBBFKQBDNGOY-RWMBFGLXSA-N Pro-Lys-Pro Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CCCCN)C(=O)N2CCC[C@@H]2C(=O)O ULWBBFKQBDNGOY-RWMBFGLXSA-N 0.000 description 6
- FYXCBXDAMPEHIQ-FHWLQOOXSA-N Pro-Trp-Lys Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CC2=CNC3=CC=CC=C32)C(=O)N[C@@H](CCCCN)C(=O)O FYXCBXDAMPEHIQ-FHWLQOOXSA-N 0.000 description 6
- NRCJWSGXMAPYQX-LPEHRKFASA-N Ser-Arg-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CO)N)C(=O)O NRCJWSGXMAPYQX-LPEHRKFASA-N 0.000 description 6
- OBXVZEAMXFSGPU-FXQIFTODSA-N Ser-Asn-Arg Chemical compound C(C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@H](CO)N)CN=C(N)N OBXVZEAMXFSGPU-FXQIFTODSA-N 0.000 description 6
- YMTLKLXDFCSCNX-BYPYZUCNSA-N Ser-Gly-Gly Chemical compound OC[C@H](N)C(=O)NCC(=O)NCC(O)=O YMTLKLXDFCSCNX-BYPYZUCNSA-N 0.000 description 6
- SFTZWNJFZYOLBD-ZDLURKLDSA-N Ser-Gly-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CO SFTZWNJFZYOLBD-ZDLURKLDSA-N 0.000 description 6
- QYSFWUIXDFJUDW-DCAQKATOSA-N Ser-Leu-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O QYSFWUIXDFJUDW-DCAQKATOSA-N 0.000 description 6
- NMZXJDSKEGFDLJ-DCAQKATOSA-N Ser-Pro-Lys Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CO)N)C(=O)N[C@@H](CCCCN)C(=O)O NMZXJDSKEGFDLJ-DCAQKATOSA-N 0.000 description 6
- VFWQQZMRKFOGLE-ZLUOBGJFSA-N Ser-Ser-Cys Chemical compound C([C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CS)C(=O)O)N)O VFWQQZMRKFOGLE-ZLUOBGJFSA-N 0.000 description 6
- BMKNXTJLHFIAAH-CIUDSAMLSA-N Ser-Ser-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O BMKNXTJLHFIAAH-CIUDSAMLSA-N 0.000 description 6
- ILZAUMFXKSIUEF-SRVKXCTJSA-N Ser-Ser-Phe Chemical compound OC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 ILZAUMFXKSIUEF-SRVKXCTJSA-N 0.000 description 6
- UBTNVMGPMYDYIU-HJPIBITLSA-N Ser-Tyr-Ile Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O UBTNVMGPMYDYIU-HJPIBITLSA-N 0.000 description 6
- KRDSCBLRHORMRK-JXUBOQSCSA-N Thr-Lys-Ala Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(O)=O KRDSCBLRHORMRK-JXUBOQSCSA-N 0.000 description 6
- BDGBHYCAZJPLHX-HJGDQZAQSA-N Thr-Lys-Asn Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(N)=O)C(O)=O BDGBHYCAZJPLHX-HJGDQZAQSA-N 0.000 description 6
- SPVHQURZJCUDQC-VOAKCMCISA-N Thr-Lys-Leu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O SPVHQURZJCUDQC-VOAKCMCISA-N 0.000 description 6
- MCDVZTRGHNXTGK-HJGDQZAQSA-N Thr-Met-Glu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCC(O)=O)C(O)=O MCDVZTRGHNXTGK-HJGDQZAQSA-N 0.000 description 6
- WURLIFOWSMBUAR-SLFFLAALSA-N Tyr-Phe-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CC=CC=C2)NC(=O)[C@H](CC3=CC=C(C=C3)O)N)C(=O)O WURLIFOWSMBUAR-SLFFLAALSA-N 0.000 description 6
- UJMCYJKPDFQLHX-XGEHTFHBSA-N Val-Ser-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](C(C)C)N)O UJMCYJKPDFQLHX-XGEHTFHBSA-N 0.000 description 6
- JPBGMZDTPVGGMQ-ULQDDVLXSA-N Val-Tyr-His Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)N JPBGMZDTPVGGMQ-ULQDDVLXSA-N 0.000 description 6
- 108010044940 alanylglutamine Proteins 0.000 description 6
- 210000000805 cytoplasm Anatomy 0.000 description 6
- 238000006366 phosphorylation reaction Methods 0.000 description 6
- RXTBLQVXNIECFP-FXQIFTODSA-N Ala-Gln-Gln Chemical compound C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(O)=O RXTBLQVXNIECFP-FXQIFTODSA-N 0.000 description 5
- FVSOUJZKYWEFOB-KBIXCLLPSA-N Ala-Gln-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CCC(N)=O)NC(=O)[C@H](C)N FVSOUJZKYWEFOB-KBIXCLLPSA-N 0.000 description 5
- XAXHGSOBFPIRFG-LSJOCFKGSA-N Ala-Pro-His Chemical compound C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](Cc1cnc[nH]1)C(O)=O XAXHGSOBFPIRFG-LSJOCFKGSA-N 0.000 description 5
- 102100033830 Amphiphysin Human genes 0.000 description 5
- DFCIPNHFKOQAME-FXQIFTODSA-N Arg-Ala-Asn Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(N)=O)C(O)=O DFCIPNHFKOQAME-FXQIFTODSA-N 0.000 description 5
- FLYANDHDFRGGTM-PYJNHQTQSA-N Arg-Ile-His Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N FLYANDHDFRGGTM-PYJNHQTQSA-N 0.000 description 5
- FVBZXNSRIDVYJS-AVGNSLFASA-N Arg-Pro-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CCCN=C(N)N FVBZXNSRIDVYJS-AVGNSLFASA-N 0.000 description 5
- JQHASVQBAKRJKD-GUBZILKMSA-N Arg-Ser-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CCCN=C(N)N)N JQHASVQBAKRJKD-GUBZILKMSA-N 0.000 description 5
- HAJWYALLJIATCX-FXQIFTODSA-N Asn-Asn-Arg Chemical compound C(C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@H](CC(=O)N)N)CN=C(N)N HAJWYALLJIATCX-FXQIFTODSA-N 0.000 description 5
- UGXVKHRDGLYFKR-CIUDSAMLSA-N Asn-Asp-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H](N)CC(N)=O UGXVKHRDGLYFKR-CIUDSAMLSA-N 0.000 description 5
- SGAUXNZEFIEAAI-GARJFASQSA-N Asn-His-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CN=CN2)NC(=O)[C@H](CC(=O)N)N)C(=O)O SGAUXNZEFIEAAI-GARJFASQSA-N 0.000 description 5
- ANPFQTJEPONRPL-UGYAYLCHSA-N Asn-Ile-Asp Chemical compound NC(=O)C[C@H](N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(O)=O)C(O)=O ANPFQTJEPONRPL-UGYAYLCHSA-N 0.000 description 5
- VWADICJNCPFKJS-ZLUOBGJFSA-N Asn-Ser-Asp Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O VWADICJNCPFKJS-ZLUOBGJFSA-N 0.000 description 5
- MKJBPDLENBUHQU-CIUDSAMLSA-N Asn-Ser-Leu Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O MKJBPDLENBUHQU-CIUDSAMLSA-N 0.000 description 5
- VLDRQOHCMKCXLY-SRVKXCTJSA-N Asn-Ser-Phe Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O VLDRQOHCMKCXLY-SRVKXCTJSA-N 0.000 description 5
- TZOZNVLBTAFJRW-UGYAYLCHSA-N Asp-Ile-Asp Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)O)NC(=O)[C@H](CC(=O)O)N TZOZNVLBTAFJRW-UGYAYLCHSA-N 0.000 description 5
- ZBYLEBZCVKLPCY-FXQIFTODSA-N Asp-Ser-Arg Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O ZBYLEBZCVKLPCY-FXQIFTODSA-N 0.000 description 5
- 108010009540 DNA (Cytosine-5-)-Methyltransferase 1 Proteins 0.000 description 5
- OETQLUYCMBARHJ-CIUDSAMLSA-N Gln-Asn-Arg Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O OETQLUYCMBARHJ-CIUDSAMLSA-N 0.000 description 5
- NPTGGVQJYRSMCM-GLLZPBPUSA-N Gln-Gln-Thr Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O NPTGGVQJYRSMCM-GLLZPBPUSA-N 0.000 description 5
- VOLVNCMGXWDDQY-LPEHRKFASA-N Gln-Glu-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CCC(=O)N)N)C(=O)O VOLVNCMGXWDDQY-LPEHRKFASA-N 0.000 description 5
- OPAINBJQDQTGJY-JGVFFNPUSA-N Glu-Gly-Pro Chemical compound C1C[C@@H](N(C1)C(=O)CNC(=O)[C@H](CCC(=O)O)N)C(=O)O OPAINBJQDQTGJY-JGVFFNPUSA-N 0.000 description 5
- HPJLZFTUUJKWAJ-JHEQGTHGSA-N Glu-Gly-Thr Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)NCC(=O)N[C@@H]([C@@H](C)O)C(O)=O HPJLZFTUUJKWAJ-JHEQGTHGSA-N 0.000 description 5
- QIQABBIDHGQXGA-ZPFDUUQYSA-N Glu-Ile-Arg Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O QIQABBIDHGQXGA-ZPFDUUQYSA-N 0.000 description 5
- VSVZIEVNUYDAFR-YUMQZZPRSA-N Gly-Ala-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)CN VSVZIEVNUYDAFR-YUMQZZPRSA-N 0.000 description 5
- JBRBACJPBZNFMF-YUMQZZPRSA-N Gly-Ala-Lys Chemical compound NCC(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CCCCN JBRBACJPBZNFMF-YUMQZZPRSA-N 0.000 description 5
- DBJYVKDPGIFXFO-BQBZGAKWSA-N Gly-Met-Ala Chemical compound [H]NCC(=O)N[C@@H](CCSC)C(=O)N[C@@H](C)C(O)=O DBJYVKDPGIFXFO-BQBZGAKWSA-N 0.000 description 5
- IZVICCORZOSGPT-JSGCOSHPSA-N Gly-Val-Tyr Chemical compound [H]NCC(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O IZVICCORZOSGPT-JSGCOSHPSA-N 0.000 description 5
- IDNNYVGVSZMQTK-IHRRRGAJSA-N His-Arg-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)N IDNNYVGVSZMQTK-IHRRRGAJSA-N 0.000 description 5
- ZJSMFRTVYSLKQU-DJFWLOJKSA-N His-Asp-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CC1=CN=CN1)N ZJSMFRTVYSLKQU-DJFWLOJKSA-N 0.000 description 5
- ASCFJMSGKUIRDU-ZPFDUUQYSA-N Ile-Arg-Gln Chemical compound CC[C@H](C)[C@H](N)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(O)=O ASCFJMSGKUIRDU-ZPFDUUQYSA-N 0.000 description 5
- SHVFUCSSACPBTF-VGDYDELISA-N Ile-Ser-His Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N SHVFUCSSACPBTF-VGDYDELISA-N 0.000 description 5
- DLCOFDAHNMMQPP-SRVKXCTJSA-N Leu-Asp-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O DLCOFDAHNMMQPP-SRVKXCTJSA-N 0.000 description 5
- CQGSYZCULZMEDE-SRVKXCTJSA-N Leu-Gln-Pro Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N1CCC[C@H]1C(O)=O CQGSYZCULZMEDE-SRVKXCTJSA-N 0.000 description 5
- CIVKXGPFXDIQBV-WDCWCFNPSA-N Leu-Gln-Thr Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O CIVKXGPFXDIQBV-WDCWCFNPSA-N 0.000 description 5
- UCDHVOALNXENLC-KBPBESRZSA-N Leu-Gly-Tyr Chemical compound CC(C)C[C@H]([NH3+])C(=O)NCC(=O)N[C@H](C([O-])=O)CC1=CC=C(O)C=C1 UCDHVOALNXENLC-KBPBESRZSA-N 0.000 description 5
- IEWBEPKLKUXQBU-VOAKCMCISA-N Leu-Leu-Thr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O IEWBEPKLKUXQBU-VOAKCMCISA-N 0.000 description 5
- BRTVHXHCUSXYRI-CIUDSAMLSA-N Leu-Ser-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O BRTVHXHCUSXYRI-CIUDSAMLSA-N 0.000 description 5
- YPLVCBKEPJPBDQ-MELADBBJSA-N Lys-Leu-Pro Chemical compound CC(C)C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCCCN)N YPLVCBKEPJPBDQ-MELADBBJSA-N 0.000 description 5
- DNWBUCHHMRQWCZ-GUBZILKMSA-N Lys-Ser-Gln Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(N)=O DNWBUCHHMRQWCZ-GUBZILKMSA-N 0.000 description 5
- IDUCUXTUHHIQIP-SOUVJXGZSA-N Phe-Gln-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CC2=CC=CC=C2)N)C(=O)O IDUCUXTUHHIQIP-SOUVJXGZSA-N 0.000 description 5
- MCIXMYKSPQUMJG-SRVKXCTJSA-N Phe-Ser-Ser Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O MCIXMYKSPQUMJG-SRVKXCTJSA-N 0.000 description 5
- IFMDQWDAJUMMJC-DCAQKATOSA-N Pro-Ala-Leu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(O)=O IFMDQWDAJUMMJC-DCAQKATOSA-N 0.000 description 5
- KDIIENQUNVNWHR-JYJNAYRXSA-N Pro-Arg-Phe Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O KDIIENQUNVNWHR-JYJNAYRXSA-N 0.000 description 5
- ICTZKEXYDDZZFP-SRVKXCTJSA-N Pro-Arg-Pro Chemical compound N([C@@H](CCCN=C(N)N)C(=O)N1[C@@H](CCC1)C(O)=O)C(=O)[C@@H]1CCCN1 ICTZKEXYDDZZFP-SRVKXCTJSA-N 0.000 description 5
- DIFXZGPHVCIVSQ-CIUDSAMLSA-N Pro-Gln-Ser Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(O)=O DIFXZGPHVCIVSQ-CIUDSAMLSA-N 0.000 description 5
- AJCRQOHDLCBHFA-SRVKXCTJSA-N Pro-His-Glu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCC(O)=O)C(O)=O AJCRQOHDLCBHFA-SRVKXCTJSA-N 0.000 description 5
- PKHDJFHFMGQMPS-RCWTZXSCSA-N Pro-Thr-Arg Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O PKHDJFHFMGQMPS-RCWTZXSCSA-N 0.000 description 5
- BRKHVZNDAOMAHX-BIIVOSGPSA-N Ser-Ala-Pro Chemical compound C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CO)N BRKHVZNDAOMAHX-BIIVOSGPSA-N 0.000 description 5
- YQHZVYJAGWMHES-ZLUOBGJFSA-N Ser-Ala-Ser Chemical compound OC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CO)C(O)=O YQHZVYJAGWMHES-ZLUOBGJFSA-N 0.000 description 5
- RZUOXAKGNHXZTB-GUBZILKMSA-N Ser-Arg-Met Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCSC)C(O)=O RZUOXAKGNHXZTB-GUBZILKMSA-N 0.000 description 5
- WXWDPFVKQRVJBJ-CIUDSAMLSA-N Ser-Asn-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@H](CO)N WXWDPFVKQRVJBJ-CIUDSAMLSA-N 0.000 description 5
- CRZRTKAVUUGKEQ-ACZMJKKPSA-N Ser-Gln-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(O)=O CRZRTKAVUUGKEQ-ACZMJKKPSA-N 0.000 description 5
- KDGARKCAKHBEDB-NKWVEPMBSA-N Ser-Gly-Pro Chemical compound C1C[C@@H](N(C1)C(=O)CNC(=O)[C@H](CO)N)C(=O)O KDGARKCAKHBEDB-NKWVEPMBSA-N 0.000 description 5
- MQUZANJDFOQOBX-SRVKXCTJSA-N Ser-Phe-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CO)C(O)=O MQUZANJDFOQOBX-SRVKXCTJSA-N 0.000 description 5
- PJIQEIFXZPCWOJ-FXQIFTODSA-N Ser-Pro-Asp Chemical compound [H]N[C@@H](CO)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(O)=O)C(O)=O PJIQEIFXZPCWOJ-FXQIFTODSA-N 0.000 description 5
- WNDUPCKKKGSKIQ-CIUDSAMLSA-N Ser-Pro-Gln Chemical compound OC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(N)=O)C(O)=O WNDUPCKKKGSKIQ-CIUDSAMLSA-N 0.000 description 5
- WLJPJRGQRNCIQS-ZLUOBGJFSA-N Ser-Ser-Asn Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(N)=O)C(O)=O WLJPJRGQRNCIQS-ZLUOBGJFSA-N 0.000 description 5
- PPCZVWHJWJFTFN-ZLUOBGJFSA-N Ser-Ser-Asp Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(O)=O PPCZVWHJWJFTFN-ZLUOBGJFSA-N 0.000 description 5
- RXUOAOOZIWABBW-XGEHTFHBSA-N Ser-Thr-Arg Chemical compound OC[C@H](N)C(=O)N[C@@H]([C@H](O)C)C(=O)N[C@H](C(O)=O)CCCN=C(N)N RXUOAOOZIWABBW-XGEHTFHBSA-N 0.000 description 5
- LGIMRDKGABDMBN-DCAQKATOSA-N Ser-Val-Lys Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CO)N LGIMRDKGABDMBN-DCAQKATOSA-N 0.000 description 5
- SIEBDTCABMZCLF-XGEHTFHBSA-N Ser-Val-Thr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O SIEBDTCABMZCLF-XGEHTFHBSA-N 0.000 description 5
- LVHHEVGYAZGXDE-KDXUFGMBSA-N Thr-Ala-Pro Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](C)C(=O)N1CCC[C@@H]1C(=O)O)N)O LVHHEVGYAZGXDE-KDXUFGMBSA-N 0.000 description 5
- WFUAUEQXPVNAEF-ZJDVBMNYSA-N Thr-Arg-Thr Chemical compound C[C@@H](O)[C@H](N)C(=O)N[C@H](C(=O)N[C@@H]([C@@H](C)O)C(O)=O)CCCN=C(N)N WFUAUEQXPVNAEF-ZJDVBMNYSA-N 0.000 description 5
- RRRRCRYTLZVCEN-HJGDQZAQSA-N Thr-Leu-Asp Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O RRRRCRYTLZVCEN-HJGDQZAQSA-N 0.000 description 5
- MEJHFIOYJHTWMK-VOAKCMCISA-N Thr-Leu-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)[C@@H](C)O MEJHFIOYJHTWMK-VOAKCMCISA-N 0.000 description 5
- KERCOYANYUPLHJ-XGEHTFHBSA-N Thr-Pro-Ser Chemical compound C[C@@H](O)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O KERCOYANYUPLHJ-XGEHTFHBSA-N 0.000 description 5
- QNXZCKMXHPULME-ZNSHCXBVSA-N Thr-Val-Pro Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](C(C)C)C(=O)N1CCC[C@@H]1C(=O)O)N)O QNXZCKMXHPULME-ZNSHCXBVSA-N 0.000 description 5
- IJGPOONOTBNTFS-GVXVVHGQSA-N Val-Lys-Glu Chemical compound [H]N[C@@H](C(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O IJGPOONOTBNTFS-GVXVVHGQSA-N 0.000 description 5
- 210000004556 brain Anatomy 0.000 description 5
- 230000008859 change Effects 0.000 description 5
- 208000035475 disorder Diseases 0.000 description 5
- 230000006870 function Effects 0.000 description 5
- 230000009368 gene silencing by RNA Effects 0.000 description 5
- 108010078144 glutaminyl-glycine Proteins 0.000 description 5
- 210000004940 nucleus Anatomy 0.000 description 5
- 230000026731 phosphorylation Effects 0.000 description 5
- 108010071207 serylmethionine Proteins 0.000 description 5
- IYLGMFKRTLBESI-ATIWLJMLSA-N (2s)-2-[[(2s)-2-[[(2s)-2-[[(2s)-2-[[(2s)-2-amino-5-(diaminomethylideneamino)pentanoyl]amino]-5-(diaminomethylideneamino)pentanoyl]amino]propanoyl]amino]-5-(diaminomethylideneamino)pentanoyl]amino]-5-(diaminomethylideneamino)pentanoic acid Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O IYLGMFKRTLBESI-ATIWLJMLSA-N 0.000 description 4
- VBDMWOKJZDCFJM-FXQIFTODSA-N Ala-Ala-Met Chemical compound CSCC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@H](C)N VBDMWOKJZDCFJM-FXQIFTODSA-N 0.000 description 4
- ODWSTKXGQGYHSH-FXQIFTODSA-N Ala-Arg-Ala Chemical compound C[C@H](N)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C)C(O)=O ODWSTKXGQGYHSH-FXQIFTODSA-N 0.000 description 4
- TTXMOJWKNRJWQJ-FXQIFTODSA-N Ala-Arg-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)C)CCCN=C(N)N TTXMOJWKNRJWQJ-FXQIFTODSA-N 0.000 description 4
- SUHLZMHFRALVSY-YUMQZZPRSA-N Ala-Lys-Gly Chemical compound NCCCC[C@H](NC(=O)[C@@H](N)C)C(=O)NCC(O)=O SUHLZMHFRALVSY-YUMQZZPRSA-N 0.000 description 4
- WQKAQKZRDIZYNV-VZFHVOOUSA-N Ala-Ser-Thr Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(O)=O WQKAQKZRDIZYNV-VZFHVOOUSA-N 0.000 description 4
- 108700031308 Antennapedia Homeodomain Proteins 0.000 description 4
- XHFXZQHTLJVZBN-FXQIFTODSA-N Asn-Arg-Asn Chemical compound C(C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)O)NC(=O)[C@H](CC(=O)N)N)CN=C(N)N XHFXZQHTLJVZBN-FXQIFTODSA-N 0.000 description 4
- POOCJCRBHHMAOS-FXQIFTODSA-N Asn-Arg-Ser Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(O)=O POOCJCRBHHMAOS-FXQIFTODSA-N 0.000 description 4
- GWNMUVANAWDZTI-YUMQZZPRSA-N Asn-Gly-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)CNC(=O)[C@H](CC(=O)N)N GWNMUVANAWDZTI-YUMQZZPRSA-N 0.000 description 4
- AITGTTNYKAWKDR-CIUDSAMLSA-N Asn-His-Ser Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CO)C(O)=O AITGTTNYKAWKDR-CIUDSAMLSA-N 0.000 description 4
- NVWJMQNYLYWVNQ-BYULHYEWSA-N Asn-Ile-Gly Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)NCC(O)=O NVWJMQNYLYWVNQ-BYULHYEWSA-N 0.000 description 4
- SEKBHZJLARBNPB-GHCJXIJMSA-N Asn-Ile-Ser Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CO)C(O)=O SEKBHZJLARBNPB-GHCJXIJMSA-N 0.000 description 4
- ZMUQQMGITUJQTI-CIUDSAMLSA-N Asn-Leu-Asn Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O ZMUQQMGITUJQTI-CIUDSAMLSA-N 0.000 description 4
- SYZWMVSXBZCOBZ-QXEWZRGKSA-N Asn-Val-Met Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCSC)C(=O)O)NC(=O)[C@H](CC(=O)N)N SYZWMVSXBZCOBZ-QXEWZRGKSA-N 0.000 description 4
- FANQWNCPNFEPGZ-WHFBIAKZSA-N Asp-Asp-Gly Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(O)=O)C(=O)NCC(O)=O FANQWNCPNFEPGZ-WHFBIAKZSA-N 0.000 description 4
- CELPEWWLSXMVPH-CIUDSAMLSA-N Asp-Asp-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H](N)CC(O)=O CELPEWWLSXMVPH-CIUDSAMLSA-N 0.000 description 4
- WBDWQKRLTVCDSY-WHFBIAKZSA-N Asp-Gly-Asp Chemical compound OC(=O)C[C@H](N)C(=O)NCC(=O)N[C@@H](CC(O)=O)C(O)=O WBDWQKRLTVCDSY-WHFBIAKZSA-N 0.000 description 4
- XLILXFRAKOYEJX-GUBZILKMSA-N Asp-Leu-Gln Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O XLILXFRAKOYEJX-GUBZILKMSA-N 0.000 description 4
- CAXXTYYGFYTBPV-IUCAKERBSA-N Gln-Leu-Gly Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)NCC(O)=O CAXXTYYGFYTBPV-IUCAKERBSA-N 0.000 description 4
- FQCILXROGNOZON-YUMQZZPRSA-N Gln-Pro-Gly Chemical compound NC(=O)CC[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O FQCILXROGNOZON-YUMQZZPRSA-N 0.000 description 4
- OAGVHWYIBZMWLA-YFKPBYRVSA-N Glu-Gly-Gly Chemical compound OC(=O)CC[C@H](N)C(=O)NCC(=O)NCC(O)=O OAGVHWYIBZMWLA-YFKPBYRVSA-N 0.000 description 4
- BXSZPACYCMNKLS-AVGNSLFASA-N Glu-Ser-Phe Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O BXSZPACYCMNKLS-AVGNSLFASA-N 0.000 description 4
- MXJYXYDREQWUMS-XKBZYTNZSA-N Glu-Thr-Ser Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(O)=O MXJYXYDREQWUMS-XKBZYTNZSA-N 0.000 description 4
- YZPVGIVFMZLQMM-YUMQZZPRSA-N Gly-Gln-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)CN YZPVGIVFMZLQMM-YUMQZZPRSA-N 0.000 description 4
- SCJJPCQUJYPHRZ-BQBZGAKWSA-N Gly-Pro-Asn Chemical compound NCC(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(O)=O SCJJPCQUJYPHRZ-BQBZGAKWSA-N 0.000 description 4
- HAOUOFNNJJLVNS-BQBZGAKWSA-N Gly-Pro-Ser Chemical compound NCC(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O HAOUOFNNJJLVNS-BQBZGAKWSA-N 0.000 description 4
- LVXFNTIIGOQBMD-SRVKXCTJSA-N His-Leu-Ser Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O LVXFNTIIGOQBMD-SRVKXCTJSA-N 0.000 description 4
- VCBWXASUBZIFLQ-IHRRRGAJSA-N His-Pro-Leu Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(C)C)C(O)=O VCBWXASUBZIFLQ-IHRRRGAJSA-N 0.000 description 4
- CHIAUHSHDARFBD-ULQDDVLXSA-N His-Pro-Tyr Chemical compound C([C@H](N)C(=O)N1[C@@H](CCC1)C(=O)N[C@@H](CC=1C=CC(O)=CC=1)C(O)=O)C1=CN=CN1 CHIAUHSHDARFBD-ULQDDVLXSA-N 0.000 description 4
- FCPSGEVYIVXPPO-QTKMDUPCSA-N His-Thr-Arg Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O FCPSGEVYIVXPPO-QTKMDUPCSA-N 0.000 description 4
- XGBVLRJLHUVCNK-DCAQKATOSA-N His-Val-Ser Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CO)C(O)=O XGBVLRJLHUVCNK-DCAQKATOSA-N 0.000 description 4
- YOZCKMXHBYKOMQ-IHRRRGAJSA-N Leu-Arg-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCCN)C(=O)O)N YOZCKMXHBYKOMQ-IHRRRGAJSA-N 0.000 description 4
- YKNBJXOJTURHCU-DCAQKATOSA-N Leu-Asp-Arg Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@H](C(O)=O)CCCN=C(N)N YKNBJXOJTURHCU-DCAQKATOSA-N 0.000 description 4
- CSFVADKICPDRRF-KKUMJFAQSA-N Leu-His-Leu Chemical compound CC(C)C[C@H]([NH3+])C(=O)N[C@H](C(=O)N[C@@H](CC(C)C)C([O-])=O)CC1=CN=CN1 CSFVADKICPDRRF-KKUMJFAQSA-N 0.000 description 4
- PDQDCFBVYXEFSD-SRVKXCTJSA-N Leu-Leu-Asp Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(O)=O)C(O)=O PDQDCFBVYXEFSD-SRVKXCTJSA-N 0.000 description 4
- FOBUGKUBUJOWAD-IHPCNDPISA-N Leu-Leu-Trp Chemical compound C1=CC=C2C(C[C@H](NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](N)CC(C)C)C(O)=O)=CNC2=C1 FOBUGKUBUJOWAD-IHPCNDPISA-N 0.000 description 4
- UCBPDSYUVAAHCD-UWVGGRQHSA-N Leu-Pro-Gly Chemical compound CC(C)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O UCBPDSYUVAAHCD-UWVGGRQHSA-N 0.000 description 4
- ZJZNLRVCZWUONM-JXUBOQSCSA-N Leu-Thr-Ala Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(O)=O ZJZNLRVCZWUONM-JXUBOQSCSA-N 0.000 description 4
- 102100022187 Leucine-rich repeat-containing protein 4C Human genes 0.000 description 4
- 101710084185 Leucine-rich repeat-containing protein 4C Proteins 0.000 description 4
- SJNZALDHDUYDBU-IHRRRGAJSA-N Lys-Arg-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCCN)C(O)=O SJNZALDHDUYDBU-IHRRRGAJSA-N 0.000 description 4
- ZXEUFAVXODIPHC-GUBZILKMSA-N Lys-Glu-Asn Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O ZXEUFAVXODIPHC-GUBZILKMSA-N 0.000 description 4
- GQZMPWBZQALKJO-UWVGGRQHSA-N Lys-Gly-Arg Chemical compound [H]N[C@@H](CCCCN)C(=O)NCC(=O)N[C@@H](CCCNC(N)=N)C(O)=O GQZMPWBZQALKJO-UWVGGRQHSA-N 0.000 description 4
- WZVSHTFTCYOFPL-GARJFASQSA-N Lys-Ser-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CO)NC(=O)[C@H](CCCCN)N)C(=O)O WZVSHTFTCYOFPL-GARJFASQSA-N 0.000 description 4
- OHXUUQDOBQKSNB-AVGNSLFASA-N Lys-Val-Arg Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O OHXUUQDOBQKSNB-AVGNSLFASA-N 0.000 description 4
- DNDVVILEHVMWIS-LPEHRKFASA-N Met-Asp-Pro Chemical compound CSCC[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N1CCC[C@@H]1C(=O)O)N DNDVVILEHVMWIS-LPEHRKFASA-N 0.000 description 4
- HSJIGJRZYUADSS-IHRRRGAJSA-N Met-Lys-Leu Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O HSJIGJRZYUADSS-IHRRRGAJSA-N 0.000 description 4
- JEGFCFLCRSJCMA-IHRRRGAJSA-N Phe-Arg-Ser Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CO)C(=O)O)N JEGFCFLCRSJCMA-IHRRRGAJSA-N 0.000 description 4
- ISYSEOWLRQKQEQ-JYJNAYRXSA-N Phe-His-Glu Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCC(O)=O)C(O)=O ISYSEOWLRQKQEQ-JYJNAYRXSA-N 0.000 description 4
- OXKJSGGTHFMGDT-UFYCRDLUSA-N Phe-Phe-Arg Chemical compound C([C@H](N)C(=O)N[C@@H](CC=1C=CC=CC=1)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O)C1=CC=CC=C1 OXKJSGGTHFMGDT-UFYCRDLUSA-N 0.000 description 4
- BONHGTUEEPIMPM-AVGNSLFASA-N Phe-Ser-Glu Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(O)=O BONHGTUEEPIMPM-AVGNSLFASA-N 0.000 description 4
- CGBYDGAJHSOGFQ-LPEHRKFASA-N Pro-Ala-Pro Chemical compound C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@@H]2CCCN2 CGBYDGAJHSOGFQ-LPEHRKFASA-N 0.000 description 4
- XROLYVMNVIKVEM-BQBZGAKWSA-N Pro-Asn-Gly Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(=O)NCC(O)=O XROLYVMNVIKVEM-BQBZGAKWSA-N 0.000 description 4
- AMBLXEMWFARNNQ-DCAQKATOSA-N Pro-Asn-Leu Chemical compound CC(C)C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)N)NC(=O)[C@@H]1CCCN1 AMBLXEMWFARNNQ-DCAQKATOSA-N 0.000 description 4
- FKYKZHOKDOPHSA-DCAQKATOSA-N Pro-Leu-Ser Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O FKYKZHOKDOPHSA-DCAQKATOSA-N 0.000 description 4
- VTFXTWDFPTWNJY-RHYQMDGZSA-N Pro-Leu-Thr Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O VTFXTWDFPTWNJY-RHYQMDGZSA-N 0.000 description 4
- 101710149951 Protein Tat Proteins 0.000 description 4
- 108091030071 RNAI Proteins 0.000 description 4
- JPIDMRXXNMIVKY-VZFHVOOUSA-N Ser-Ala-Thr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O JPIDMRXXNMIVKY-VZFHVOOUSA-N 0.000 description 4
- YUSRGTQIPCJNHQ-CIUDSAMLSA-N Ser-Arg-Glu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(O)=O YUSRGTQIPCJNHQ-CIUDSAMLSA-N 0.000 description 4
- YMEXHZTVKDAKIY-GHCJXIJMSA-N Ser-Asn-Ile Chemical compound CC[C@H](C)[C@H](NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CO)C(O)=O YMEXHZTVKDAKIY-GHCJXIJMSA-N 0.000 description 4
- BGOWRLSWJCVYAQ-CIUDSAMLSA-N Ser-Asp-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O BGOWRLSWJCVYAQ-CIUDSAMLSA-N 0.000 description 4
- GHPQVUYZQQGEDA-BIIVOSGPSA-N Ser-Asp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)O)NC(=O)[C@H](CO)N)C(=O)O GHPQVUYZQQGEDA-BIIVOSGPSA-N 0.000 description 4
- BEAFYHFQTOTVFS-VGDYDELISA-N Ser-Ile-His Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@H](CO)N BEAFYHFQTOTVFS-VGDYDELISA-N 0.000 description 4
- AZWNCEBQZXELEZ-FXQIFTODSA-N Ser-Pro-Ser Chemical compound OC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O AZWNCEBQZXELEZ-FXQIFTODSA-N 0.000 description 4
- XQJCEKXQUJQNNK-ZLUOBGJFSA-N Ser-Ser-Ser Chemical compound OC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O XQJCEKXQUJQNNK-ZLUOBGJFSA-N 0.000 description 4
- ZSDXEKUKQAKZFE-XAVMHZPKSA-N Ser-Thr-Pro Chemical compound C[C@H]([C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CO)N)O ZSDXEKUKQAKZFE-XAVMHZPKSA-N 0.000 description 4
- IRKWVRSEQFTGGV-VEVYYDQMSA-N Thr-Asn-Arg Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O IRKWVRSEQFTGGV-VEVYYDQMSA-N 0.000 description 4
- PAOYNIKMYOGBMR-PBCZWWQYSA-N Thr-Asn-His Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N)O PAOYNIKMYOGBMR-PBCZWWQYSA-N 0.000 description 4
- SHOMROOOQBDGRL-JHEQGTHGSA-N Thr-Glu-Gly Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O SHOMROOOQBDGRL-JHEQGTHGSA-N 0.000 description 4
- ULHASJWZGUEUNN-XIRDDKMYSA-N Trp-Lys-Ser Chemical compound [H]N[C@@H](CC1=CNC2=C1C=CC=C2)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(O)=O ULHASJWZGUEUNN-XIRDDKMYSA-N 0.000 description 4
- 108010040443 aspartyl-aspartic acid Proteins 0.000 description 4
- 108010092854 aspartyllysine Proteins 0.000 description 4
- 229940079593 drug Drugs 0.000 description 4
- 108010048367 enhanced green fluorescent protein Proteins 0.000 description 4
- 108010049041 glutamylalanine Proteins 0.000 description 4
- 108010001064 glycyl-glycyl-glycyl-glycine Proteins 0.000 description 4
- 230000003834 intracellular effect Effects 0.000 description 4
- 108010073093 leucyl-glycyl-glycyl-glycine Proteins 0.000 description 4
- 230000004048 modification Effects 0.000 description 4
- 238000012986 modification Methods 0.000 description 4
- 210000002569 neuron Anatomy 0.000 description 4
- 108010082795 phenylalanyl-arginyl-arginine Proteins 0.000 description 4
- 230000028327 secretion Effects 0.000 description 4
- 238000006467 substitution reaction Methods 0.000 description 4
- 238000013518 transcription Methods 0.000 description 4
- 230000035897 transcription Effects 0.000 description 4
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 4
- HKZAAJSTFUZYTO-LURJTMIESA-N (2s)-2-[[2-[[2-[[2-[(2-aminoacetyl)amino]acetyl]amino]acetyl]amino]acetyl]amino]-3-hydroxypropanoic acid Chemical compound NCC(=O)NCC(=O)NCC(=O)NCC(=O)N[C@@H](CO)C(O)=O HKZAAJSTFUZYTO-LURJTMIESA-N 0.000 description 3
- XWTNPSHCJMZAHQ-QMMMGPOBSA-N 2-[[2-[[2-[[(2s)-2-amino-4-methylpentanoyl]amino]acetyl]amino]acetyl]amino]acetic acid Chemical compound CC(C)C[C@H](N)C(=O)NCC(=O)NCC(=O)NCC(O)=O XWTNPSHCJMZAHQ-QMMMGPOBSA-N 0.000 description 3
- PBAMJJXWDQXOJA-FXQIFTODSA-N Ala-Asp-Arg Chemical compound C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@H](C(O)=O)CCCN=C(N)N PBAMJJXWDQXOJA-FXQIFTODSA-N 0.000 description 3
- SOBIAADAMRHGKH-CIUDSAMLSA-N Ala-Leu-Ser Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O SOBIAADAMRHGKH-CIUDSAMLSA-N 0.000 description 3
- NCQMBSJGJMYKCK-ZLUOBGJFSA-N Ala-Ser-Ser Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O NCQMBSJGJMYKCK-ZLUOBGJFSA-N 0.000 description 3
- CLICCYPMVFGUOF-IHRRRGAJSA-N Arg-Lys-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(C)C)C(O)=O CLICCYPMVFGUOF-IHRRRGAJSA-N 0.000 description 3
- LCBSSOCDWUTQQV-SDDRHHMPSA-N Arg-Met-Pro Chemical compound CSCC[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N LCBSSOCDWUTQQV-SDDRHHMPSA-N 0.000 description 3
- GOVUDFOGXOONFT-VEVYYDQMSA-N Asn-Arg-Thr Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)O)C(O)=O GOVUDFOGXOONFT-VEVYYDQMSA-N 0.000 description 3
- QHBMKQWOIYJYMI-BYULHYEWSA-N Asn-Asn-Val Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](C(C)C)C(O)=O QHBMKQWOIYJYMI-BYULHYEWSA-N 0.000 description 3
- OLISTMZJGQUOGS-GMOBBJLQSA-N Asn-Ile-Arg Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)NC(=O)[C@H](CC(=O)N)N OLISTMZJGQUOGS-GMOBBJLQSA-N 0.000 description 3
- WXVGISRWSYGEDK-KKUMJFAQSA-N Asn-Lys-Phe Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)O)NC(=O)[C@H](CCCCN)NC(=O)[C@H](CC(=O)N)N WXVGISRWSYGEDK-KKUMJFAQSA-N 0.000 description 3
- PQKSVQSMTHPRIB-ZKWXMUAHSA-N Asn-Val-Ser Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CO)C(O)=O PQKSVQSMTHPRIB-ZKWXMUAHSA-N 0.000 description 3
- KNMRXHIAVXHCLW-ZLUOBGJFSA-N Asp-Asn-Ser Chemical compound C([C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CO)C(=O)O)N)C(=O)O KNMRXHIAVXHCLW-ZLUOBGJFSA-N 0.000 description 3
- CLUMZOKVGUWUFD-CIUDSAMLSA-N Asp-Leu-Asn Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O CLUMZOKVGUWUFD-CIUDSAMLSA-N 0.000 description 3
- CTWCFPWFIGRAEP-CIUDSAMLSA-N Asp-Lys-Asp Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(O)=O CTWCFPWFIGRAEP-CIUDSAMLSA-N 0.000 description 3
- MYLZFUMPZCPJCJ-NHCYSSNCSA-N Asp-Lys-Val Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C(C)C)C(O)=O MYLZFUMPZCPJCJ-NHCYSSNCSA-N 0.000 description 3
- ZKAOJVJQGVUIIU-GUBZILKMSA-N Asp-Pro-Arg Chemical compound OC(=O)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCCNC(N)=N)C(O)=O ZKAOJVJQGVUIIU-GUBZILKMSA-N 0.000 description 3
- HICVMZCGVFKTPM-BQBZGAKWSA-N Asp-Pro-Gly Chemical compound OC(=O)C[C@H](N)C(=O)N1CCC[C@H]1C(=O)NCC(O)=O HICVMZCGVFKTPM-BQBZGAKWSA-N 0.000 description 3
- SXLCDCZHNCLFGZ-BPUTZDHNSA-N Asp-Pro-Trp Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC1=CNC2=C1C=CC=C2)C(O)=O SXLCDCZHNCLFGZ-BPUTZDHNSA-N 0.000 description 3
- VNXQRBXEQXLERQ-CIUDSAMLSA-N Asp-Ser-Lys Chemical compound C(CCN)C[C@@H](C(=O)O)NC(=O)[C@H](CO)NC(=O)[C@H](CC(=O)O)N VNXQRBXEQXLERQ-CIUDSAMLSA-N 0.000 description 3
- 108700024394 Exon Proteins 0.000 description 3
- XOKGKOQWADCLFQ-GARJFASQSA-N Gln-Arg-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCC(=O)N)N)C(=O)O XOKGKOQWADCLFQ-GARJFASQSA-N 0.000 description 3
- PIUPHASDUFSHTF-CIUDSAMLSA-N Gln-Pro-Asn Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CCC(=O)N)N)C(=O)N[C@@H](CC(=O)N)C(=O)O PIUPHASDUFSHTF-CIUDSAMLSA-N 0.000 description 3
- JILRMFFFCHUUTJ-ACZMJKKPSA-N Gln-Ser-Ser Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O JILRMFFFCHUUTJ-ACZMJKKPSA-N 0.000 description 3
- FLLRAEJOLZPSMN-CIUDSAMLSA-N Glu-Asn-Arg Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@H](C(O)=O)CCCN=C(N)N FLLRAEJOLZPSMN-CIUDSAMLSA-N 0.000 description 3
- ZOXBSICWUDAOHX-GUBZILKMSA-N Glu-Asn-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CCC(O)=O ZOXBSICWUDAOHX-GUBZILKMSA-N 0.000 description 3
- MWMJCGBSIORNCD-AVGNSLFASA-N Glu-Leu-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O MWMJCGBSIORNCD-AVGNSLFASA-N 0.000 description 3
- BCYGDJXHAGZNPQ-DCAQKATOSA-N Glu-Lys-Glu Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCC(O)=O)C(O)=O BCYGDJXHAGZNPQ-DCAQKATOSA-N 0.000 description 3
- YKBUCXNNBYZYAY-MNXVOIDGSA-N Glu-Lys-Ile Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O YKBUCXNNBYZYAY-MNXVOIDGSA-N 0.000 description 3
- QJVZSVUYZFYLFQ-CIUDSAMLSA-N Glu-Pro-Ala Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C)C(O)=O QJVZSVUYZFYLFQ-CIUDSAMLSA-N 0.000 description 3
- HZISRJBYZAODRV-XQXXSGGOSA-N Glu-Thr-Ala Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(O)=O HZISRJBYZAODRV-XQXXSGGOSA-N 0.000 description 3
- JDAYMLXPUJRSDJ-XIRDDKMYSA-N Glu-Trp-Arg Chemical compound C1=CC=C2C(C[C@H](NC(=O)[C@H](CCC(O)=O)N)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O)=CNC2=C1 JDAYMLXPUJRSDJ-XIRDDKMYSA-N 0.000 description 3
- YWAQATDNEKZFFK-BYPYZUCNSA-N Gly-Gly-Ser Chemical compound NCC(=O)NCC(=O)N[C@@H](CO)C(O)=O YWAQATDNEKZFFK-BYPYZUCNSA-N 0.000 description 3
- OLPPXYMMIARYAL-QMMMGPOBSA-N Gly-Gly-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)CNC(=O)CN OLPPXYMMIARYAL-QMMMGPOBSA-N 0.000 description 3
- FEUPVVCGQLNXNP-IRXDYDNUSA-N Gly-Phe-Phe Chemical compound C([C@H](NC(=O)CN)C(=O)N[C@@H](CC=1C=CC=CC=1)C(O)=O)C1=CC=CC=C1 FEUPVVCGQLNXNP-IRXDYDNUSA-N 0.000 description 3
- WCORRBXVISTKQL-WHFBIAKZSA-N Gly-Ser-Ser Chemical compound NCC(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O WCORRBXVISTKQL-WHFBIAKZSA-N 0.000 description 3
- FKESCSGWBPUTPN-FOHZUACHSA-N Gly-Thr-Asn Chemical compound [H]NCC(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(N)=O)C(O)=O FKESCSGWBPUTPN-FOHZUACHSA-N 0.000 description 3
- XHVONGZZVUUORG-WEDXCCLWSA-N Gly-Thr-Lys Chemical compound NCC(=O)N[C@@H]([C@H](O)C)C(=O)N[C@H](C(O)=O)CCCCN XHVONGZZVUUORG-WEDXCCLWSA-N 0.000 description 3
- RIUZKUJUPVFAGY-HOTGVXAUSA-N Gly-Trp-His Chemical compound C1=CC=C2C(=C1)C(=CN2)C[C@@H](C(=O)N[C@@H](CC3=CN=CN3)C(=O)O)NC(=O)CN RIUZKUJUPVFAGY-HOTGVXAUSA-N 0.000 description 3
- RGPWUJOMKFYFSR-QWRGUYRKSA-N His-Gly-Leu Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)NCC(=O)N[C@@H](CC(C)C)C(O)=O RGPWUJOMKFYFSR-QWRGUYRKSA-N 0.000 description 3
- KHUFDBQXGLEIHC-BZSNNMDCSA-N His-Leu-Tyr Chemical compound C([C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC=1C=CC(O)=CC=1)C(O)=O)C1=CN=CN1 KHUFDBQXGLEIHC-BZSNNMDCSA-N 0.000 description 3
- 241000713772 Human immunodeficiency virus 1 Species 0.000 description 3
- 108700000788 Human immunodeficiency virus 1 tat peptide (47-57) Proteins 0.000 description 3
- LNJLOZYNZFGJMM-DEQVHRJGSA-N Ile-His-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N2CCC[C@@H]2C(=O)O)N LNJLOZYNZFGJMM-DEQVHRJGSA-N 0.000 description 3
- KEKTTYCXKGBAAL-VGDYDELISA-N Ile-His-Ser Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](CO)C(=O)O)N KEKTTYCXKGBAAL-VGDYDELISA-N 0.000 description 3
- JZNVOBUNTWNZPW-GHCJXIJMSA-N Ile-Ser-Asp Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(=O)O)C(=O)O)N JZNVOBUNTWNZPW-GHCJXIJMSA-N 0.000 description 3
- MMEDVBWCMGRKKC-GARJFASQSA-N Leu-Asp-Pro Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N1CCC[C@@H]1C(=O)O)N MMEDVBWCMGRKKC-GARJFASQSA-N 0.000 description 3
- PVMPDMIKUVNOBD-CIUDSAMLSA-N Leu-Asp-Ser Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O PVMPDMIKUVNOBD-CIUDSAMLSA-N 0.000 description 3
- LOLUPZNNADDTAA-AVGNSLFASA-N Leu-Gln-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O LOLUPZNNADDTAA-AVGNSLFASA-N 0.000 description 3
- FQZPTCNSNPWHLJ-AVGNSLFASA-N Leu-Gln-Lys Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCCN)C(O)=O FQZPTCNSNPWHLJ-AVGNSLFASA-N 0.000 description 3
- OXRLYTYUXAQTHP-YUMQZZPRSA-N Leu-Gly-Ala Chemical compound [H]N[C@@H](CC(C)C)C(=O)NCC(=O)N[C@@H](C)C(O)=O OXRLYTYUXAQTHP-YUMQZZPRSA-N 0.000 description 3
- KYIIALJHAOIAHF-KKUMJFAQSA-N Leu-Leu-His Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@H](C(O)=O)CC1=CN=CN1 KYIIALJHAOIAHF-KKUMJFAQSA-N 0.000 description 3
- VCHVSKNMTXWIIP-SRVKXCTJSA-N Leu-Lys-Ser Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(O)=O VCHVSKNMTXWIIP-SRVKXCTJSA-N 0.000 description 3
- DGWXCIORNLWGGG-CIUDSAMLSA-N Lys-Asn-Ser Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(O)=O DGWXCIORNLWGGG-CIUDSAMLSA-N 0.000 description 3
- VSRXPEHZMHSFKU-IUCAKERBSA-N Lys-Gln-Gly Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)NCC(O)=O VSRXPEHZMHSFKU-IUCAKERBSA-N 0.000 description 3
- SBQDRNOLGSYHQA-YUMQZZPRSA-N Lys-Ser-Gly Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(=O)NCC(O)=O SBQDRNOLGSYHQA-YUMQZZPRSA-N 0.000 description 3
- IOQWIOPSKJOEKI-SRVKXCTJSA-N Lys-Ser-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(O)=O IOQWIOPSKJOEKI-SRVKXCTJSA-N 0.000 description 3
- 239000004472 Lysine Substances 0.000 description 3
- 101150083522 MECP2 gene Proteins 0.000 description 3
- YRAWWKUTNBILNT-FXQIFTODSA-N Met-Ala-Ala Chemical compound CSCC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](C)C(O)=O YRAWWKUTNBILNT-FXQIFTODSA-N 0.000 description 3
- MUDYEFAKNSTFAI-JYJNAYRXSA-N Met-Tyr-Val Chemical compound [H]N[C@@H](CCSC)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](C(C)C)C(O)=O MUDYEFAKNSTFAI-JYJNAYRXSA-N 0.000 description 3
- AJHCSUXXECOXOY-UHFFFAOYSA-N N-glycyl-L-tryptophan Natural products C1=CC=C2C(CC(NC(=O)CN)C(O)=O)=CNC2=C1 AJHCSUXXECOXOY-UHFFFAOYSA-N 0.000 description 3
- 125000001429 N-terminal alpha-amino-acid group Chemical group 0.000 description 3
- MQVFHOPCKNTHGT-MELADBBJSA-N Phe-Asp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)O)NC(=O)[C@H](CC2=CC=CC=C2)N)C(=O)O MQVFHOPCKNTHGT-MELADBBJSA-N 0.000 description 3
- VFDRDMOMHBJGKD-UFYCRDLUSA-N Phe-Tyr-Arg Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CC2=CC=C(C=C2)O)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N VFDRDMOMHBJGKD-UFYCRDLUSA-N 0.000 description 3
- LNLNHXIQPGKRJQ-SRVKXCTJSA-N Pro-Arg-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@@H]1CCCN1 LNLNHXIQPGKRJQ-SRVKXCTJSA-N 0.000 description 3
- VPFGPKIWSDVTOY-SRVKXCTJSA-N Pro-Glu-His Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O VPFGPKIWSDVTOY-SRVKXCTJSA-N 0.000 description 3
- FFSLAIOXRMOFIZ-GJZGRUSLSA-N Pro-Gly-Trp Chemical compound N([C@@H](CC=1C2=CC=CC=C2NC=1)C(=O)O)C(=O)CNC(=O)[C@@H]1CCCN1 FFSLAIOXRMOFIZ-GJZGRUSLSA-N 0.000 description 3
- QHSSUIHLAIWXEE-IHRRRGAJSA-N Pro-Tyr-Asn Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(N)=O)C(O)=O QHSSUIHLAIWXEE-IHRRRGAJSA-N 0.000 description 3
- 241000700159 Rattus Species 0.000 description 3
- NNFMANHDYSVNIO-DCAQKATOSA-N Ser-Lys-Arg Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O NNFMANHDYSVNIO-DCAQKATOSA-N 0.000 description 3
- IFLVBVIYADZIQO-DCAQKATOSA-N Ser-Met-Lys Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CO)N IFLVBVIYADZIQO-DCAQKATOSA-N 0.000 description 3
- AXOHAHIUJHCLQR-IHRRRGAJSA-N Ser-Met-Tyr Chemical compound CSCC[C@@H](C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)O)NC(=O)[C@H](CO)N AXOHAHIUJHCLQR-IHRRRGAJSA-N 0.000 description 3
- TVPQRPNBYCRRLL-IHRRRGAJSA-N Ser-Phe-Met Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CCSC)C(O)=O TVPQRPNBYCRRLL-IHRRRGAJSA-N 0.000 description 3
- HHJFMHQYEAAOBM-ZLUOBGJFSA-N Ser-Ser-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CO)C(=O)N[C@@H](C)C(O)=O HHJFMHQYEAAOBM-ZLUOBGJFSA-N 0.000 description 3
- VVKVHAOOUGNDPJ-SRVKXCTJSA-N Ser-Tyr-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CO)C(O)=O VVKVHAOOUGNDPJ-SRVKXCTJSA-N 0.000 description 3
- 102100029462 Sodium-dependent lysophosphatidylcholine symporter 1 Human genes 0.000 description 3
- 101710185583 Sodium-dependent lysophosphatidylcholine symporter 1 Proteins 0.000 description 3
- 102000019197 Superoxide Dismutase Human genes 0.000 description 3
- 108010012715 Superoxide dismutase Proteins 0.000 description 3
- GLQFKOVWXPPFTP-VEVYYDQMSA-N Thr-Arg-Asp Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(O)=O GLQFKOVWXPPFTP-VEVYYDQMSA-N 0.000 description 3
- JNQZPAWOPBZGIX-RCWTZXSCSA-N Thr-Arg-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)[C@@H](C)O)CCCN=C(N)N JNQZPAWOPBZGIX-RCWTZXSCSA-N 0.000 description 3
- VIWQOOBRKCGSDK-RYQLBKOJSA-N Trp-Arg-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CC2=CNC3=CC=CC=C32)N)C(=O)O VIWQOOBRKCGSDK-RYQLBKOJSA-N 0.000 description 3
- CKKFTIQYURNSEI-IHRRRGAJSA-N Tyr-Asn-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@H](CC(N)=O)NC(=O)[C@@H](N)CC1=CC=C(O)C=C1 CKKFTIQYURNSEI-IHRRRGAJSA-N 0.000 description 3
- FIRUOPRJKCBLST-KKUMJFAQSA-N Tyr-His-Asp Chemical compound C1=CC(=CC=C1C[C@@H](C(=O)N[C@@H](CC2=CN=CN2)C(=O)N[C@@H](CC(=O)O)C(=O)O)N)O FIRUOPRJKCBLST-KKUMJFAQSA-N 0.000 description 3
- DMWNPLOERDAHSY-MEYUZBJRSA-N Tyr-Leu-Thr Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O DMWNPLOERDAHSY-MEYUZBJRSA-N 0.000 description 3
- QFXVAFIHVWXXBJ-AVGNSLFASA-N Tyr-Ser-Glu Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(O)=O QFXVAFIHVWXXBJ-AVGNSLFASA-N 0.000 description 3
- COYSIHFOCOMGCF-UHFFFAOYSA-N Val-Arg-Gly Natural products CC(C)C(N)C(=O)NC(C(=O)NCC(O)=O)CCCN=C(N)N COYSIHFOCOMGCF-UHFFFAOYSA-N 0.000 description 3
- LAYSXAOGWHKNED-XPUUQOCRSA-N Val-Gly-Ser Chemical compound CC(C)[C@H](N)C(=O)NCC(=O)N[C@@H](CO)C(O)=O LAYSXAOGWHKNED-XPUUQOCRSA-N 0.000 description 3
- YTNGABPUXFEOGU-SRVKXCTJSA-N Val-Pro-Arg Chemical compound CC(C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCCN=C(N)N)C(O)=O YTNGABPUXFEOGU-SRVKXCTJSA-N 0.000 description 3
- LGXUZJIQCGXKGZ-QXEWZRGKSA-N Val-Pro-Asn Chemical compound CC(C)[C@@H](C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(=O)N)C(=O)O)N LGXUZJIQCGXKGZ-QXEWZRGKSA-N 0.000 description 3
- SSYBNWFXCFNRFN-GUBZILKMSA-N Val-Pro-Ser Chemical compound CC(C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O SSYBNWFXCFNRFN-GUBZILKMSA-N 0.000 description 3
- 108010011559 alanylphenylalanine Proteins 0.000 description 3
- AVKUERGKIZMTKX-NJBDSQKTSA-N ampicillin Chemical compound C1([C@@H](N)C(=O)N[C@H]2[C@H]3SC([C@@H](N3C2=O)C(O)=O)(C)C)=CC=CC=C1 AVKUERGKIZMTKX-NJBDSQKTSA-N 0.000 description 3
- 230000008901 benefit Effects 0.000 description 3
- 230000008499 blood brain barrier function Effects 0.000 description 3
- 210000001218 blood-brain barrier Anatomy 0.000 description 3
- 238000010790 dilution Methods 0.000 description 3
- 239000012895 dilution Substances 0.000 description 3
- 230000002068 genetic effect Effects 0.000 description 3
- 108010079547 glutamylmethionine Proteins 0.000 description 3
- 125000003630 glycyl group Chemical group [H]N([H])C([H])([H])C(*)=O 0.000 description 3
- 108010084389 glycyltryptophan Proteins 0.000 description 3
- 108010028295 histidylhistidine Proteins 0.000 description 3
- 230000001771 impaired effect Effects 0.000 description 3
- 238000000338 in vitro Methods 0.000 description 3
- 238000002347 injection Methods 0.000 description 3
- 239000007924 injection Substances 0.000 description 3
- 238000003780 insertion Methods 0.000 description 3
- 230000037431 insertion Effects 0.000 description 3
- 210000004962 mammalian cell Anatomy 0.000 description 3
- 230000007246 mechanism Effects 0.000 description 3
- 229930182817 methionine Natural products 0.000 description 3
- 230000011987 methylation Effects 0.000 description 3
- 238000007069 methylation reaction Methods 0.000 description 3
- 210000002241 neurite Anatomy 0.000 description 3
- 230000001717 pathogenic effect Effects 0.000 description 3
- 230000035515 penetration Effects 0.000 description 3
- 239000000546 pharmaceutical excipient Substances 0.000 description 3
- 230000004797 therapeutic response Effects 0.000 description 3
- 210000001519 tissue Anatomy 0.000 description 3
- 238000010361 transduction Methods 0.000 description 3
- 230000026683 transduction Effects 0.000 description 3
- 238000001890 transfection Methods 0.000 description 3
- 230000005945 translocation Effects 0.000 description 3
- 238000001262 western blot Methods 0.000 description 3
- MPLOSMWGDNJSEV-WHFBIAKZSA-N Ala-Gly-Asp Chemical compound [H]N[C@@H](C)C(=O)NCC(=O)N[C@@H](CC(O)=O)C(O)=O MPLOSMWGDNJSEV-WHFBIAKZSA-N 0.000 description 2
- DPNZTBKGAUAZQU-DLOVCJGASA-N Ala-Leu-His Chemical compound C[C@@H](C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N DPNZTBKGAUAZQU-DLOVCJGASA-N 0.000 description 2
- QPBSRMDNJOTFAL-AICCOOGYSA-N Ala-Leu-Leu-Thr Chemical compound C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H]([C@@H](C)O)C(O)=O QPBSRMDNJOTFAL-AICCOOGYSA-N 0.000 description 2
- PEIBBAXIKUAYGN-UBHSHLNASA-N Ala-Phe-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)C)CC1=CC=CC=C1 PEIBBAXIKUAYGN-UBHSHLNASA-N 0.000 description 2
- LSMDIAAALJJLRO-XQXXSGGOSA-N Ala-Thr-Glu Chemical compound [H]N[C@@H](C)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(O)=O)C(O)=O LSMDIAAALJJLRO-XQXXSGGOSA-N 0.000 description 2
- KWKQGHSSNHPGOW-BQBZGAKWSA-N Arg-Ala-Gly Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C)C(=O)NCC(O)=O KWKQGHSSNHPGOW-BQBZGAKWSA-N 0.000 description 2
- BVBKBQRPOJFCQM-DCAQKATOSA-N Arg-Asn-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O BVBKBQRPOJFCQM-DCAQKATOSA-N 0.000 description 2
- FEZJJKXNPSEYEV-CIUDSAMLSA-N Arg-Gln-Ala Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C)C(O)=O FEZJJKXNPSEYEV-CIUDSAMLSA-N 0.000 description 2
- VDBKFYYIBLXEIF-GUBZILKMSA-N Arg-Gln-Glu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O VDBKFYYIBLXEIF-GUBZILKMSA-N 0.000 description 2
- NKBQZKVMKJJDLX-SRVKXCTJSA-N Arg-Glu-Leu Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O NKBQZKVMKJJDLX-SRVKXCTJSA-N 0.000 description 2
- VEAIMHJZTIDCIH-KKUMJFAQSA-N Arg-Phe-Gln Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CCC(N)=O)C(O)=O VEAIMHJZTIDCIH-KKUMJFAQSA-N 0.000 description 2
- FRBAHXABMQXSJQ-FXQIFTODSA-N Arg-Ser-Ser Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CO)C(O)=O FRBAHXABMQXSJQ-FXQIFTODSA-N 0.000 description 2
- SYFHFLGAROUHNT-VEVYYDQMSA-N Arg-Thr-Asn Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(N)=O)C(O)=O SYFHFLGAROUHNT-VEVYYDQMSA-N 0.000 description 2
- 239000004475 Arginine Substances 0.000 description 2
- YNDLOUMBVDVALC-ZLUOBGJFSA-N Asn-Ala-Ala Chemical compound C[C@@H](C(=O)N[C@@H](C)C(=O)O)NC(=O)[C@H](CC(=O)N)N YNDLOUMBVDVALC-ZLUOBGJFSA-N 0.000 description 2
- SWLOHUMCUDRTCL-ZLUOBGJFSA-N Asn-Ala-Asn Chemical compound C[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)O)NC(=O)[C@H](CC(=O)N)N SWLOHUMCUDRTCL-ZLUOBGJFSA-N 0.000 description 2
- UBKOVSLDWIHYSY-ACZMJKKPSA-N Asn-Glu-Ser Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(O)=O UBKOVSLDWIHYSY-ACZMJKKPSA-N 0.000 description 2
- OOXUBGLNDRGOKT-FXQIFTODSA-N Asn-Ser-Arg Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O OOXUBGLNDRGOKT-FXQIFTODSA-N 0.000 description 2
- NCXTYSVDWLAQGZ-ZKWXMUAHSA-N Asn-Ser-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H](N)CC(N)=O NCXTYSVDWLAQGZ-ZKWXMUAHSA-N 0.000 description 2
- KRXIWXCXOARFNT-ZLUOBGJFSA-N Asp-Ala-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](C)NC(=O)[C@@H](N)CC(O)=O KRXIWXCXOARFNT-ZLUOBGJFSA-N 0.000 description 2
- VFUXXFVCYZPOQG-WDSKDSINSA-N Asp-Glu-Gly Chemical compound [H]N[C@@H](CC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(O)=O VFUXXFVCYZPOQG-WDSKDSINSA-N 0.000 description 2
- HAFCJCDJGIOYPW-WDSKDSINSA-N Asp-Gly-Gln Chemical compound OC(=O)C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CCC(N)=O HAFCJCDJGIOYPW-WDSKDSINSA-N 0.000 description 2
- SVABRQFIHCSNCI-FOHZUACHSA-N Asp-Gly-Thr Chemical compound [H]N[C@@H](CC(O)=O)C(=O)NCC(=O)N[C@@H]([C@@H](C)O)C(O)=O SVABRQFIHCSNCI-FOHZUACHSA-N 0.000 description 2
- CJUKAWUWBZCTDQ-SRVKXCTJSA-N Asp-Leu-Lys Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(O)=O CJUKAWUWBZCTDQ-SRVKXCTJSA-N 0.000 description 2
- JSNWZMFSLIWAHS-HJGDQZAQSA-N Asp-Thr-Leu Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC(C)C)C(=O)O)NC(=O)[C@H](CC(=O)O)N)O JSNWZMFSLIWAHS-HJGDQZAQSA-N 0.000 description 2
- 102100026596 Bcl-2-like protein 1 Human genes 0.000 description 2
- 208000014644 Brain disease Diseases 0.000 description 2
- 125000001433 C-terminal amino-acid group Chemical group 0.000 description 2
- 108091026890 Coding region Proteins 0.000 description 2
- 108020004705 Codon Proteins 0.000 description 2
- IIGHQOPGMGKDMT-SRVKXCTJSA-N Cys-Asp-Phe Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CS)N IIGHQOPGMGKDMT-SRVKXCTJSA-N 0.000 description 2
- HAYVLBZZBDCKRA-SRVKXCTJSA-N Cys-His-Lys Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CS)N HAYVLBZZBDCKRA-SRVKXCTJSA-N 0.000 description 2
- 230000007067 DNA methylation Effects 0.000 description 2
- 208000032274 Encephalopathy Diseases 0.000 description 2
- XZWYTXMRWQJBGX-VXBMVYAYSA-N FLAG peptide Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H](NC(=O)[C@@H](N)CC(O)=O)CC1=CC=C(O)C=C1 XZWYTXMRWQJBGX-VXBMVYAYSA-N 0.000 description 2
- HHWQMFIGMMOVFK-WDSKDSINSA-N Gln-Ala-Gly Chemical compound OC(=O)CNC(=O)[C@H](C)NC(=O)[C@@H](N)CCC(N)=O HHWQMFIGMMOVFK-WDSKDSINSA-N 0.000 description 2
- AAOBFSKXAVIORT-GUBZILKMSA-N Gln-Asn-Leu Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(O)=O AAOBFSKXAVIORT-GUBZILKMSA-N 0.000 description 2
- LWDGZZGWDMHBOF-FXQIFTODSA-N Gln-Glu-Asn Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O LWDGZZGWDMHBOF-FXQIFTODSA-N 0.000 description 2
- NSNUZSPSADIMJQ-WDSKDSINSA-N Gln-Gly-Asp Chemical compound NC(=O)CC[C@H](N)C(=O)NCC(=O)N[C@@H](CC(O)=O)C(O)=O NSNUZSPSADIMJQ-WDSKDSINSA-N 0.000 description 2
- SHAUZYVSXAMYAZ-JYJNAYRXSA-N Gln-Leu-Phe Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)O)NC(=O)[C@H](CCC(=O)N)N SHAUZYVSXAMYAZ-JYJNAYRXSA-N 0.000 description 2
- XBWGJWXGUNSZAT-CIUDSAMLSA-N Gln-Met-Asp Chemical compound CSCC[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)O)NC(=O)[C@H](CCC(=O)N)N XBWGJWXGUNSZAT-CIUDSAMLSA-N 0.000 description 2
- OACPJRQRAHMQEQ-NHCYSSNCSA-N Gln-Val-Arg Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O OACPJRQRAHMQEQ-NHCYSSNCSA-N 0.000 description 2
- KASDBWKLWJKTLJ-GUBZILKMSA-N Glu-Glu-Met Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCSC)C(O)=O KASDBWKLWJKTLJ-GUBZILKMSA-N 0.000 description 2
- AIGROOHQXCACHL-WDSKDSINSA-N Glu-Gly-Ala Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)NCC(=O)N[C@@H](C)C(O)=O AIGROOHQXCACHL-WDSKDSINSA-N 0.000 description 2
- LGYCLOCORAEQSZ-PEFMBERDSA-N Glu-Ile-Asp Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(O)=O)C(O)=O LGYCLOCORAEQSZ-PEFMBERDSA-N 0.000 description 2
- QXDXIXFSFHUYAX-MNXVOIDGSA-N Glu-Ile-Leu Chemical compound CC(C)C[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)CC)NC(=O)[C@@H](N)CCC(O)=O QXDXIXFSFHUYAX-MNXVOIDGSA-N 0.000 description 2
- INGJLBQKTRJLFO-UKJIMTQDSA-N Glu-Ile-Val Chemical compound CC(C)[C@@H](C(O)=O)NC(=O)[C@H]([C@@H](C)CC)NC(=O)[C@@H](N)CCC(O)=O INGJLBQKTRJLFO-UKJIMTQDSA-N 0.000 description 2
- FBEJIDRSQCGFJI-GUBZILKMSA-N Glu-Leu-Ser Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(O)=O FBEJIDRSQCGFJI-GUBZILKMSA-N 0.000 description 2
- SWRVAQHFBRZVNX-GUBZILKMSA-N Glu-Lys-Asn Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CC(N)=O)C(O)=O SWRVAQHFBRZVNX-GUBZILKMSA-N 0.000 description 2
- SUIAHERNFYRBDZ-GVXVVHGQSA-N Glu-Lys-Val Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C(C)C)C(O)=O SUIAHERNFYRBDZ-GVXVVHGQSA-N 0.000 description 2
- 102000005720 Glutathione transferase Human genes 0.000 description 2
- 108010070675 Glutathione transferase Proteins 0.000 description 2
- QXPRJQPCFXMCIY-NKWVEPMBSA-N Gly-Ala-Pro Chemical compound C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)CN QXPRJQPCFXMCIY-NKWVEPMBSA-N 0.000 description 2
- LJPIRKICOISLKN-WHFBIAKZSA-N Gly-Ala-Ser Chemical compound NCC(=O)N[C@@H](C)C(=O)N[C@@H](CO)C(O)=O LJPIRKICOISLKN-WHFBIAKZSA-N 0.000 description 2
- BYYNJRSNDARRBX-YFKPBYRVSA-N Gly-Gln-Gly Chemical compound NCC(=O)N[C@@H](CCC(N)=O)C(=O)NCC(O)=O BYYNJRSNDARRBX-YFKPBYRVSA-N 0.000 description 2
- QSVCIFZPGLOZGH-WDSKDSINSA-N Gly-Glu-Ser Chemical compound NCC(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(O)=O QSVCIFZPGLOZGH-WDSKDSINSA-N 0.000 description 2
- UFPXDFOYHVEIPI-BYPYZUCNSA-N Gly-Gly-Asp Chemical compound NCC(=O)NCC(=O)N[C@H](C(O)=O)CC(O)=O UFPXDFOYHVEIPI-BYPYZUCNSA-N 0.000 description 2
- YFGONBOFGGWKKY-VHSXEESVSA-N Gly-His-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC2=CN=CN2)NC(=O)CN)C(=O)O YFGONBOFGGWKKY-VHSXEESVSA-N 0.000 description 2
- UESJMAMHDLEHGM-NHCYSSNCSA-N Gly-Ile-Leu Chemical compound NCC(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(C)C)C(O)=O UESJMAMHDLEHGM-NHCYSSNCSA-N 0.000 description 2
- WMGHDYWNHNLGBV-ONGXEEELSA-N Gly-Phe-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@@H](NC(=O)CN)CC1=CC=CC=C1 WMGHDYWNHNLGBV-ONGXEEELSA-N 0.000 description 2
- FFALDIDGPLUDKV-ZDLURKLDSA-N Gly-Thr-Ser Chemical compound [H]NCC(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CO)C(O)=O FFALDIDGPLUDKV-ZDLURKLDSA-N 0.000 description 2
- GBYYQVBXFVDJPJ-WLTAIBSBSA-N Gly-Tyr-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CC1=CC=C(C=C1)O)NC(=O)CN)O GBYYQVBXFVDJPJ-WLTAIBSBSA-N 0.000 description 2
- 208000031886 HIV Infections Diseases 0.000 description 2
- SVHKVHBPTOMLTO-DCAQKATOSA-N His-Arg-Asp Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CC(O)=O)C(O)=O SVHKVHBPTOMLTO-DCAQKATOSA-N 0.000 description 2
- LSQHWKPPOFDHHZ-YUMQZZPRSA-N His-Asp-Gly Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)NCC(=O)O)N LSQHWKPPOFDHHZ-YUMQZZPRSA-N 0.000 description 2
- TVRMJKNELJKNRS-GUBZILKMSA-N His-Glu-Asn Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CC(=O)N)C(=O)O)N TVRMJKNELJKNRS-GUBZILKMSA-N 0.000 description 2
- JMSONHOUHFDOJH-GUBZILKMSA-N His-Ser-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H](N)CC1=CN=CN1 JMSONHOUHFDOJH-GUBZILKMSA-N 0.000 description 2
- DGLAHESNTJWGDO-SRVKXCTJSA-N His-Ser-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)N[C@@H](CO)C(=O)N[C@@H](CC2=CN=CN2)C(=O)O)N DGLAHESNTJWGDO-SRVKXCTJSA-N 0.000 description 2
- 102100021454 Histone deacetylase 4 Human genes 0.000 description 2
- 101710177324 Histone deacetylase 4 Proteins 0.000 description 2
- 241000282412 Homo Species 0.000 description 2
- 101000590272 Homo sapiens 26S proteasome non-ATPase regulatory subunit 2 Proteins 0.000 description 2
- 101000931098 Homo sapiens DNA (cytosine-5)-methyltransferase 1 Proteins 0.000 description 2
- ATXGFMOBVKSOMK-PEDHHIEDSA-N Ile-Arg-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H]([C@@H](C)CC)C(=O)O)N ATXGFMOBVKSOMK-PEDHHIEDSA-N 0.000 description 2
- NPROWIBAWYMPAZ-GUDRVLHUSA-N Ile-Asp-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N1CCC[C@@H]1C(=O)O)N NPROWIBAWYMPAZ-GUDRVLHUSA-N 0.000 description 2
- AQTWDZDISVGCAC-CFMVVWHZSA-N Ile-Asp-Tyr Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(=O)O)C(=O)N[C@@H](CC1=CC=C(C=C1)O)C(=O)O)N AQTWDZDISVGCAC-CFMVVWHZSA-N 0.000 description 2
- GTSAALPQZASLPW-KJYZGMDISA-N Ile-His-Trp Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](CC2=CNC3=CC=CC=C32)C(=O)O)N GTSAALPQZASLPW-KJYZGMDISA-N 0.000 description 2
- FZWVCYCYWCLQDH-NHCYSSNCSA-N Ile-Leu-Gly Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC(C)C)C(=O)NCC(=O)O)N FZWVCYCYWCLQDH-NHCYSSNCSA-N 0.000 description 2
- UIEZQYNXCYHMQS-BJDJZHNGSA-N Ile-Lys-Ala Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(=O)O)N UIEZQYNXCYHMQS-BJDJZHNGSA-N 0.000 description 2
- GLYJPWIRLBAIJH-FQUUOJAGSA-N Ile-Lys-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N1CCC[C@@H]1C(=O)O)N GLYJPWIRLBAIJH-FQUUOJAGSA-N 0.000 description 2
- YWCJXQKATPNPOE-UKJIMTQDSA-N Ile-Val-Glu Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(=O)O)C(=O)O)N YWCJXQKATPNPOE-UKJIMTQDSA-N 0.000 description 2
- PWWVAXIEGOYWEE-UHFFFAOYSA-N Isophenergan Chemical compound C1=CC=C2N(CC(C)N(C)C)C3=CC=CC=C3SC2=C1 PWWVAXIEGOYWEE-UHFFFAOYSA-N 0.000 description 2
- KFKWRHQBZQICHA-STQMWFEESA-N L-leucyl-L-phenylalanine Natural products CC(C)C[C@H](N)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 KFKWRHQBZQICHA-STQMWFEESA-N 0.000 description 2
- FFEARJCKVFRZRR-BYPYZUCNSA-N L-methionine Chemical compound CSCC[C@H](N)C(O)=O FFEARJCKVFRZRR-BYPYZUCNSA-N 0.000 description 2
- TYYLDKGBCJGJGW-UHFFFAOYSA-N L-tryptophan-L-tyrosine Natural products C=1NC2=CC=CC=C2C=1CC(N)C(=O)NC(C(O)=O)CC1=CC=C(O)C=C1 TYYLDKGBCJGJGW-UHFFFAOYSA-N 0.000 description 2
- OHZIZVWQXJPBJS-IXOXFDKPSA-N Leu-His-Thr Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H]([C@@H](C)O)C(O)=O OHZIZVWQXJPBJS-IXOXFDKPSA-N 0.000 description 2
- LXKNSJLSGPNHSK-KKUMJFAQSA-N Leu-Leu-Lys Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)O)N LXKNSJLSGPNHSK-KKUMJFAQSA-N 0.000 description 2
- WMIOEVKKYIMVKI-DCAQKATOSA-N Leu-Pro-Ala Chemical compound [H]N[C@@H](CC(C)C)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C)C(O)=O WMIOEVKKYIMVKI-DCAQKATOSA-N 0.000 description 2
- AKVBOOKXVAMKSS-GUBZILKMSA-N Leu-Ser-Gln Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(O)=O AKVBOOKXVAMKSS-GUBZILKMSA-N 0.000 description 2
- MVHXGBZUJLWZOH-BJDJZHNGSA-N Leu-Ser-Ile Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)CC)C(O)=O MVHXGBZUJLWZOH-BJDJZHNGSA-N 0.000 description 2
- JGKHAFUAPZCCDU-BZSNNMDCSA-N Leu-Tyr-Leu Chemical compound CC(C)C[C@H]([NH3+])C(=O)N[C@H](C(=O)N[C@@H](CC(C)C)C([O-])=O)CC1=CC=C(O)C=C1 JGKHAFUAPZCCDU-BZSNNMDCSA-N 0.000 description 2
- RDFIVFHPOSOXMW-ACRUOGEOSA-N Leu-Tyr-Phe Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O RDFIVFHPOSOXMW-ACRUOGEOSA-N 0.000 description 2
- FZIJIFCXUCZHOL-CIUDSAMLSA-N Lys-Ala-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCCCN FZIJIFCXUCZHOL-CIUDSAMLSA-N 0.000 description 2
- WSXTWLJHTLRFLW-SRVKXCTJSA-N Lys-Ala-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCCN)C(O)=O WSXTWLJHTLRFLW-SRVKXCTJSA-N 0.000 description 2
- NTEVEUCLFMWSND-SRVKXCTJSA-N Lys-Arg-Gln Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(O)=O NTEVEUCLFMWSND-SRVKXCTJSA-N 0.000 description 2
- WGCKDDHUFPQSMZ-ZPFDUUQYSA-N Lys-Asp-Ile Chemical compound CC[C@H](C)[C@@H](C(O)=O)NC(=O)[C@H](CC(O)=O)NC(=O)[C@@H](N)CCCCN WGCKDDHUFPQSMZ-ZPFDUUQYSA-N 0.000 description 2
- IWWMPCPLFXFBAF-SRVKXCTJSA-N Lys-Asp-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(O)=O IWWMPCPLFXFBAF-SRVKXCTJSA-N 0.000 description 2
- NRQRKMYZONPCTM-CIUDSAMLSA-N Lys-Asp-Ser Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CO)C(O)=O NRQRKMYZONPCTM-CIUDSAMLSA-N 0.000 description 2
- SVJRVFPSHPGWFF-DCAQKATOSA-N Lys-Cys-Arg Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CS)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O SVJRVFPSHPGWFF-DCAQKATOSA-N 0.000 description 2
- IZJGPPIGYTVXLB-FQUUOJAGSA-N Lys-Ile-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CCCCN)N IZJGPPIGYTVXLB-FQUUOJAGSA-N 0.000 description 2
- VSTNAUBHKQPVJX-IHRRRGAJSA-N Lys-Met-Leu Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(C)C)C(O)=O VSTNAUBHKQPVJX-IHRRRGAJSA-N 0.000 description 2
- YTJFXEDRUOQGSP-DCAQKATOSA-N Lys-Pro-Ser Chemical compound [H]N[C@@H](CCCCN)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O YTJFXEDRUOQGSP-DCAQKATOSA-N 0.000 description 2
- JMNRXRPBHFGXQX-GUBZILKMSA-N Lys-Ser-Glu Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@H](C(O)=O)CCC(O)=O JMNRXRPBHFGXQX-GUBZILKMSA-N 0.000 description 2
- MIFFFXHMAHFACR-KATARQTJSA-N Lys-Ser-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)[C@H](CO)NC(=O)[C@@H](N)CCCCN MIFFFXHMAHFACR-KATARQTJSA-N 0.000 description 2
- QVTDVTONTRSQMF-WDCWCFNPSA-N Lys-Thr-Glu Chemical compound OC(=O)CC[C@@H](C(O)=O)NC(=O)[C@H]([C@H](O)C)NC(=O)[C@@H](N)CCCCN QVTDVTONTRSQMF-WDCWCFNPSA-N 0.000 description 2
- YFQSSOAGMZGXFT-MEYUZBJRSA-N Lys-Thr-Tyr Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(O)=O YFQSSOAGMZGXFT-MEYUZBJRSA-N 0.000 description 2
- 101710175625 Maltose/maltodextrin-binding periplasmic protein Proteins 0.000 description 2
- GPAHWYRSHCKICP-GUBZILKMSA-N Met-Glu-Glu Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CCC(O)=O)C(O)=O GPAHWYRSHCKICP-GUBZILKMSA-N 0.000 description 2
- JPCHYAUKOUGOIB-HJGDQZAQSA-N Met-Glu-Thr Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O JPCHYAUKOUGOIB-HJGDQZAQSA-N 0.000 description 2
- HAQLBBVZAGMESV-IHRRRGAJSA-N Met-Lys-Lys Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(O)=O HAQLBBVZAGMESV-IHRRRGAJSA-N 0.000 description 2
- 241000699666 Mus <mouse, genus> Species 0.000 description 2
- XMBSYZWANAQXEV-UHFFFAOYSA-N N-alpha-L-glutamyl-L-phenylalanine Natural products OC(=O)CCC(N)C(=O)NC(C(O)=O)CC1=CC=CC=C1 XMBSYZWANAQXEV-UHFFFAOYSA-N 0.000 description 2
- 208000025966 Neurological disease Diseases 0.000 description 2
- 239000000020 Nitrocellulose Substances 0.000 description 2
- 241000283973 Oryctolagus cuniculus Species 0.000 description 2
- KBVJZCVLQWCJQN-KKUMJFAQSA-N Phe-Leu-Asn Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(O)=O KBVJZCVLQWCJQN-KKUMJFAQSA-N 0.000 description 2
- NJJBATPLUQHRBM-IHRRRGAJSA-N Phe-Pro-Ser Chemical compound C1C[C@H](N(C1)C(=O)[C@H](CC2=CC=CC=C2)N)C(=O)N[C@@H](CO)C(=O)O NJJBATPLUQHRBM-IHRRRGAJSA-N 0.000 description 2
- APMXLWHMIVWLLR-BZSNNMDCSA-N Phe-Tyr-Ser Chemical compound C([C@H](N)C(=O)N[C@@H](CC=1C=CC(O)=CC=1)C(=O)N[C@@H](CO)C(O)=O)C1=CC=CC=C1 APMXLWHMIVWLLR-BZSNNMDCSA-N 0.000 description 2
- XQLBWXHVZVBNJM-FXQIFTODSA-N Pro-Ala-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1 XQLBWXHVZVBNJM-FXQIFTODSA-N 0.000 description 2
- KPDRZQUWJKTMBP-DCAQKATOSA-N Pro-Asp-Leu Chemical compound CC(C)C[C@@H](C(=O)O)NC(=O)[C@H](CC(=O)O)NC(=O)[C@@H]1CCCN1 KPDRZQUWJKTMBP-DCAQKATOSA-N 0.000 description 2
- JMVQDLDPDBXAAX-YUMQZZPRSA-N Pro-Gly-Gln Chemical compound NC(=O)CC[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H]1CCCN1 JMVQDLDPDBXAAX-YUMQZZPRSA-N 0.000 description 2
- MCWHYUWXVNRXFV-RWMBFGLXSA-N Pro-Leu-Pro Chemical compound CC(C)C[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@@H]2CCCN2 MCWHYUWXVNRXFV-RWMBFGLXSA-N 0.000 description 2
- ABSSTGUCBCDKMU-UWVGGRQHSA-N Pro-Lys-Gly Chemical compound NCCCC[C@@H](C(=O)NCC(O)=O)NC(=O)[C@@H]1CCCN1 ABSSTGUCBCDKMU-UWVGGRQHSA-N 0.000 description 2
- KWMZPPWYBVZIER-XGEHTFHBSA-N Pro-Ser-Thr Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CO)C(=O)N[C@@H]([C@@H](C)O)C(O)=O KWMZPPWYBVZIER-XGEHTFHBSA-N 0.000 description 2
- LZHHZYDPMZEMRX-STQMWFEESA-N Pro-Tyr-Gly Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)NCC(O)=O LZHHZYDPMZEMRX-STQMWFEESA-N 0.000 description 2
- SHTKRJHDMNSKRM-ULQDDVLXSA-N Pro-Tyr-His Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CC2=CC=C(C=C2)O)C(=O)N[C@@H](CC3=CN=CN3)C(=O)O SHTKRJHDMNSKRM-ULQDDVLXSA-N 0.000 description 2
- WXUBSIDKNMFAGS-IHRRRGAJSA-N Ser-Arg-Tyr Chemical compound NC(N)=NCCC[C@H](NC(=O)[C@H](CO)N)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 WXUBSIDKNMFAGS-IHRRRGAJSA-N 0.000 description 2
- BTPAWKABYQMKKN-LKXGYXEUSA-N Ser-Asp-Thr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O BTPAWKABYQMKKN-LKXGYXEUSA-N 0.000 description 2
- DSSOYPJWSWFOLK-CIUDSAMLSA-N Ser-Cys-Leu Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CS)C(=O)N[C@@H](CC(C)C)C(O)=O DSSOYPJWSWFOLK-CIUDSAMLSA-N 0.000 description 2
- KJMOINFQVCCSDX-XKBZYTNZSA-N Ser-Gln-Thr Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H]([C@@H](C)O)C(O)=O KJMOINFQVCCSDX-XKBZYTNZSA-N 0.000 description 2
- VQBCMLMPEWPUTB-ACZMJKKPSA-N Ser-Glu-Ser Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CO)C(O)=O VQBCMLMPEWPUTB-ACZMJKKPSA-N 0.000 description 2
- QGAHMVHBORDHDC-YUMQZZPRSA-N Ser-His-Gly Chemical compound OC[C@H](N)C(=O)N[C@H](C(=O)NCC(O)=O)CC1=CN=CN1 QGAHMVHBORDHDC-YUMQZZPRSA-N 0.000 description 2
- IXZHZUGGKLRHJD-DCAQKATOSA-N Ser-Leu-Val Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C(C)C)C(O)=O IXZHZUGGKLRHJD-DCAQKATOSA-N 0.000 description 2
- GZSZPKSBVAOGIE-CIUDSAMLSA-N Ser-Lys-Ala Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(O)=O GZSZPKSBVAOGIE-CIUDSAMLSA-N 0.000 description 2
- QUGRFWPMPVIAPW-IHRRRGAJSA-N Ser-Pro-Phe Chemical compound OC[C@H](N)C(=O)N1CCC[C@H]1C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 QUGRFWPMPVIAPW-IHRRRGAJSA-N 0.000 description 2
- SRSPTFBENMJHMR-WHFBIAKZSA-N Ser-Ser-Gly Chemical compound OC[C@H](N)C(=O)N[C@@H](CO)C(=O)NCC(O)=O SRSPTFBENMJHMR-WHFBIAKZSA-N 0.000 description 2
- FAPWRFPIFSIZLT-UHFFFAOYSA-M Sodium chloride Chemical compound [Na+].[Cl-] FAPWRFPIFSIZLT-UHFFFAOYSA-M 0.000 description 2
- DDPVJPIGACCMEH-XQXXSGGOSA-N Thr-Ala-Gln Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(O)=O DDPVJPIGACCMEH-XQXXSGGOSA-N 0.000 description 2
- LHUBVKCLOVALIA-HJGDQZAQSA-N Thr-Arg-Gln Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(N)=O)C(O)=O LHUBVKCLOVALIA-HJGDQZAQSA-N 0.000 description 2
- PKXHGEXFMIZSER-QTKMDUPCSA-N Thr-Arg-His Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N)O PKXHGEXFMIZSER-QTKMDUPCSA-N 0.000 description 2
- CEXFELBFVHLYDZ-XGEHTFHBSA-N Thr-Arg-Ser Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CO)C(O)=O CEXFELBFVHLYDZ-XGEHTFHBSA-N 0.000 description 2
- VOGXLRKCWFLJBY-HSHDSVGOSA-N Thr-Arg-Trp Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)O)N)O VOGXLRKCWFLJBY-HSHDSVGOSA-N 0.000 description 2
- VIBXMCZWVUOZLA-OLHMAJIHSA-N Thr-Asn-Asn Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CC(=O)N)C(=O)O)N)O VIBXMCZWVUOZLA-OLHMAJIHSA-N 0.000 description 2
- QILPDQCTQZDHFM-HJGDQZAQSA-N Thr-Gln-Arg Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O QILPDQCTQZDHFM-HJGDQZAQSA-N 0.000 description 2
- LIXBDERDAGNVAV-XKBZYTNZSA-N Thr-Gln-Ser Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(O)=O LIXBDERDAGNVAV-XKBZYTNZSA-N 0.000 description 2
- ZBKDBZUTTXINIX-RWRJDSDZSA-N Thr-Ile-Gln Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CCC(N)=O)C(O)=O ZBKDBZUTTXINIX-RWRJDSDZSA-N 0.000 description 2
- YJCVECXVYHZOBK-KNZXXDILSA-N Thr-Ile-Pro Chemical compound CC[C@H](C)[C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H]([C@@H](C)O)N YJCVECXVYHZOBK-KNZXXDILSA-N 0.000 description 2
- PZSDPRBZINDEJV-HTUGSXCWSA-N Thr-Phe-Gln Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CCC(N)=O)C(O)=O PZSDPRBZINDEJV-HTUGSXCWSA-N 0.000 description 2
- BBPCSGKKPJUYRB-UVOCVTCTSA-N Thr-Thr-Leu Chemical compound [H]N[C@@H]([C@@H](C)O)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(O)=O BBPCSGKKPJUYRB-UVOCVTCTSA-N 0.000 description 2
- QEJHHFFFCUDPDV-WDSOQIARSA-N Trp-His-Val Chemical compound CC(C)[C@@H](C(=O)O)NC(=O)[C@H](CC1=CN=CN1)NC(=O)[C@H](CC2=CNC3=CC=CC=C32)N QEJHHFFFCUDPDV-WDSOQIARSA-N 0.000 description 2
- TWAVEIJGFCBWCG-JYJNAYRXSA-N Tyr-Gln-Leu Chemical compound CC(C)C[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CC1=CC=C(C=C1)O)N TWAVEIJGFCBWCG-JYJNAYRXSA-N 0.000 description 2
- CTDPLKMBVALCGN-JSGCOSHPSA-N Tyr-Gly-Val Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)NCC(=O)N[C@@H](C(C)C)C(O)=O CTDPLKMBVALCGN-JSGCOSHPSA-N 0.000 description 2
- GFJXBLSZOFWHAW-JYJNAYRXSA-N Tyr-His-Glu Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CCC(O)=O)C(O)=O GFJXBLSZOFWHAW-JYJNAYRXSA-N 0.000 description 2
- KIJLSRYAUGGZIN-CFMVVWHZSA-N Tyr-Ile-Asp Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(O)=O)C(O)=O KIJLSRYAUGGZIN-CFMVVWHZSA-N 0.000 description 2
- LVFZXRQQQDTBQH-IRIUXVKKSA-N Tyr-Thr-Glu Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(O)=O)C(O)=O LVFZXRQQQDTBQH-IRIUXVKKSA-N 0.000 description 2
- RGJZPXFZIUUQDN-BPNCWPANSA-N Tyr-Val-Ala Chemical compound [H]N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](C)C(O)=O RGJZPXFZIUUQDN-BPNCWPANSA-N 0.000 description 2
- OQWNEUXPKHIEJO-NRPADANISA-N Val-Glu-Ser Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CO)C(=O)O)N OQWNEUXPKHIEJO-NRPADANISA-N 0.000 description 2
- OXGVAUFVTOPFFA-XPUUQOCRSA-N Val-Gly-Cys Chemical compound CC(C)[C@@H](C(=O)NCC(=O)N[C@@H](CS)C(=O)O)N OXGVAUFVTOPFFA-XPUUQOCRSA-N 0.000 description 2
- DOFAQXCYFQKSHT-SRVKXCTJSA-N Val-Pro-Pro Chemical compound CC(C)[C@H](N)C(=O)N1CCC[C@H]1C(=O)N1[C@H](C(O)=O)CCC1 DOFAQXCYFQKSHT-SRVKXCTJSA-N 0.000 description 2
- NIJJYAXOARWZEE-UHFFFAOYSA-N Valproic acid Chemical compound CCCC(C(O)=O)CCC NIJJYAXOARWZEE-UHFFFAOYSA-N 0.000 description 2
- 125000000539 amino acid group Chemical group 0.000 description 2
- 239000003708 ampul Substances 0.000 description 2
- ODKSFYDXXFIFQN-UHFFFAOYSA-N arginine Natural products OC(=O)C(N)CCCNC(N)=N ODKSFYDXXFIFQN-UHFFFAOYSA-N 0.000 description 2
- 230000001580 bacterial effect Effects 0.000 description 2
- 230000008827 biological function Effects 0.000 description 2
- OWMVSZAMULFTJU-UHFFFAOYSA-N bis-tris Chemical compound OCCN(CCO)C(CO)(CO)CO OWMVSZAMULFTJU-UHFFFAOYSA-N 0.000 description 2
- 210000004899 c-terminal region Anatomy 0.000 description 2
- 206010008118 cerebral infarction Diseases 0.000 description 2
- 238000003776 cleavage reaction Methods 0.000 description 2
- 150000001875 compounds Chemical class 0.000 description 2
- 210000003618 cortical neuron Anatomy 0.000 description 2
- 230000001086 cytosolic effect Effects 0.000 description 2
- 230000007547 defect Effects 0.000 description 2
- 210000001787 dendrite Anatomy 0.000 description 2
- 231100000673 dose–response relationship Toxicity 0.000 description 2
- 230000029142 excretion Effects 0.000 description 2
- 238000002474 experimental method Methods 0.000 description 2
- 108010067216 glycyl-glycyl-glycine Proteins 0.000 description 2
- 108010020688 glycylhistidine Proteins 0.000 description 2
- 230000027984 hippocampus development Effects 0.000 description 2
- 238000001727 in vivo Methods 0.000 description 2
- 238000011534 incubation Methods 0.000 description 2
- 238000001802 infusion Methods 0.000 description 2
- 239000004615 ingredient Substances 0.000 description 2
- 230000005764 inhibitory process Effects 0.000 description 2
- 230000003993 interaction Effects 0.000 description 2
- 238000001990 intravenous administration Methods 0.000 description 2
- 238000000021 kinase assay Methods 0.000 description 2
- 108010044056 leucyl-phenylalanine Proteins 0.000 description 2
- 239000006166 lysate Substances 0.000 description 2
- 125000003588 lysine group Chemical group [H]N([H])C([H])([H])C([H])([H])C([H])([H])C([H])([H])C([H])(N([H])[H])C(*)=O 0.000 description 2
- 108010025153 lysyl-alanyl-alanine Proteins 0.000 description 2
- 230000014759 maintenance of location Effects 0.000 description 2
- 239000000463 material Substances 0.000 description 2
- 235000013336 milk Nutrition 0.000 description 2
- 239000008267 milk Substances 0.000 description 2
- 210000004080 milk Anatomy 0.000 description 2
- 230000001537 neural effect Effects 0.000 description 2
- 230000007472 neurodevelopment Effects 0.000 description 2
- 229920001220 nitrocellulos Polymers 0.000 description 2
- 230000002018 overexpression Effects 0.000 description 2
- 230000004853 protein function Effects 0.000 description 2
- 230000004044 response Effects 0.000 description 2
- 230000007017 scission Effects 0.000 description 2
- 239000011780 sodium chloride Substances 0.000 description 2
- 238000002415 sodium dodecyl sulfate polyacrylamide gel electrophoresis Methods 0.000 description 2
- 239000000758 substrate Substances 0.000 description 2
- 210000000225 synapse Anatomy 0.000 description 2
- 230000001225 therapeutic effect Effects 0.000 description 2
- 108010061238 threonyl-glycine Proteins 0.000 description 2
- 230000014616 translation Effects 0.000 description 2
- 108010062760 transportan Proteins 0.000 description 2
- PBKWZFANFUTEPS-CWUSWOHSSA-N transportan Chemical compound C([C@@H](C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CC(C)C)C(=O)NCC(=O)N[C@@H](CCCCN)C(=O)N[C@H](C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(C)C)C(N)=O)[C@@H](C)CC)NC(=O)CNC(=O)[C@H](C)NC(=O)[C@H](CO)NC(=O)[C@H](CC(N)=O)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](NC(=O)[C@H](CC=1C2=CC=CC=C2NC=1)NC(=O)CN)[C@@H](C)O)C1=CC=C(O)C=C1 PBKWZFANFUTEPS-CWUSWOHSSA-N 0.000 description 2
- 108010060175 trypsinogen activation peptide Proteins 0.000 description 2
- 108010044292 tryptophyltyrosine Proteins 0.000 description 2
- 230000003612 virological effect Effects 0.000 description 2
- 238000005406 washing Methods 0.000 description 2
- IGXNPQWXIRIGBF-KEOOTSPTSA-N (2s)-2-[[(2s)-2-[[(2s)-2-[[(2s)-2-[[(2s)-2-amino-3-(1h-imidazol-5-yl)propanoyl]amino]-3-(1h-imidazol-5-yl)propanoyl]amino]-3-(1h-imidazol-5-yl)propanoyl]amino]-3-(1h-imidazol-5-yl)propanoyl]amino]-3-(1h-imidazol-5-yl)propanoic acid Chemical compound C([C@H](N)C(=O)N[C@@H](CC=1NC=NC=1)C(=O)N[C@@H](CC=1NC=NC=1)C(=O)N[C@@H](CC=1NC=NC=1)C(=O)N[C@@H](CC=1NC=NC=1)C(O)=O)C1=CN=CN1 IGXNPQWXIRIGBF-KEOOTSPTSA-N 0.000 description 1
- RAVVEEJGALCVIN-AGVBWZICSA-N (2s)-2-[[(2s)-2-[[(2s)-2-[[(2s)-5-amino-2-[[(2s)-2-[[(2s)-2-[[(2s)-6-amino-2-[[(2s)-6-amino-2-[[(2s)-2-[[2-[[(2s)-2-amino-3-(4-hydroxyphenyl)propanoyl]amino]acetyl]amino]-5-(diaminomethylideneamino)pentanoyl]amino]hexanoyl]amino]hexanoyl]amino]-5-(diamino Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCC(N)=O)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCCCN)NC(=O)[C@H](CCCCN)NC(=O)[C@H](CCCN=C(N)N)NC(=O)CNC(=O)[C@@H](N)CC1=CC=C(O)C=C1 RAVVEEJGALCVIN-AGVBWZICSA-N 0.000 description 1
- AUXMWYRZQPIXCC-KNIFDHDWSA-N (2s)-2-amino-4-methylpentanoic acid;(2s)-2-aminopropanoic acid Chemical compound C[C@H](N)C(O)=O.CC(C)C[C@H](N)C(O)=O AUXMWYRZQPIXCC-KNIFDHDWSA-N 0.000 description 1
- RRBGTUQJDFBWNN-MUGJNUQGSA-N (2s)-6-amino-2-[[(2s)-6-amino-2-[[(2s)-6-amino-2-[[(2s)-2,6-diaminohexanoyl]amino]hexanoyl]amino]hexanoyl]amino]hexanoic acid Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(O)=O RRBGTUQJDFBWNN-MUGJNUQGSA-N 0.000 description 1
- KDCGOANMDULRCW-UHFFFAOYSA-N 7H-purine Chemical class N1=CNC2=NC=NC2=C1 KDCGOANMDULRCW-UHFFFAOYSA-N 0.000 description 1
- FXKNPWNXPQZLES-ZLUOBGJFSA-N Ala-Asn-Ser Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CO)C(O)=O FXKNPWNXPQZLES-ZLUOBGJFSA-N 0.000 description 1
- YIGLXQRFQVWFEY-NRPADANISA-N Ala-Gln-Val Chemical compound [H]N[C@@H](C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(O)=O YIGLXQRFQVWFEY-NRPADANISA-N 0.000 description 1
- NIZKGBJVCMRDKO-KWQFWETISA-N Ala-Gly-Tyr Chemical compound C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 NIZKGBJVCMRDKO-KWQFWETISA-N 0.000 description 1
- MNZHHDPWDWQJCQ-YUMQZZPRSA-N Ala-Leu-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)NCC(O)=O MNZHHDPWDWQJCQ-YUMQZZPRSA-N 0.000 description 1
- VCSABYLVNWQYQE-SRVKXCTJSA-N Ala-Lys-Lys Chemical compound NCCCC[C@H](NC(=O)[C@@H](N)C)C(=O)N[C@@H](CCCCN)C(O)=O VCSABYLVNWQYQE-SRVKXCTJSA-N 0.000 description 1
- VCSABYLVNWQYQE-UHFFFAOYSA-N Ala-Lys-Lys Natural products NCCCCC(NC(=O)C(N)C)C(=O)NC(CCCCN)C(O)=O VCSABYLVNWQYQE-UHFFFAOYSA-N 0.000 description 1
- YCRAFFCYWOUEOF-DLOVCJGASA-N Ala-Phe-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@@H](NC(=O)[C@@H](N)C)CC1=CC=CC=C1 YCRAFFCYWOUEOF-DLOVCJGASA-N 0.000 description 1
- RTZCUEHYUQZIDE-WHFBIAKZSA-N Ala-Ser-Gly Chemical compound C[C@H](N)C(=O)N[C@@H](CO)C(=O)NCC(O)=O RTZCUEHYUQZIDE-WHFBIAKZSA-N 0.000 description 1
- 101150019028 Antp gene Proteins 0.000 description 1
- GIVWETPOBCRTND-DCAQKATOSA-N Arg-Gln-Arg Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O GIVWETPOBCRTND-DCAQKATOSA-N 0.000 description 1
- JCAISGGAOQXEHJ-ZPFDUUQYSA-N Arg-Gln-Ile Chemical compound CC[C@H](C)[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)N)NC(=O)[C@H](CCCN=C(N)N)N JCAISGGAOQXEHJ-ZPFDUUQYSA-N 0.000 description 1
- KRQSPVKUISQQFS-FJXKBIBVSA-N Arg-Gly-Thr Chemical compound C[C@@H](O)[C@@H](C(O)=O)NC(=O)CNC(=O)[C@@H](N)CCCN=C(N)N KRQSPVKUISQQFS-FJXKBIBVSA-N 0.000 description 1
- DGFXIWKPTDKBLF-AVGNSLFASA-N Arg-His-Val Chemical compound CC(C)[C@@H](C(=O)O)NC(=O)[C@H](CC1=CN=CN1)NC(=O)[C@H](CCCN=C(N)N)N DGFXIWKPTDKBLF-AVGNSLFASA-N 0.000 description 1
- NGTYEHIRESTSRX-UWVGGRQHSA-N Arg-Lys-Gly Chemical compound NCCCC[C@@H](C(=O)NCC(O)=O)NC(=O)[C@@H](N)CCCN=C(N)N NGTYEHIRESTSRX-UWVGGRQHSA-N 0.000 description 1
- NYDIVDKTULRINZ-AVGNSLFASA-N Arg-Met-Lys Chemical compound CSCC[C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCCN=C(N)N)N NYDIVDKTULRINZ-AVGNSLFASA-N 0.000 description 1
- RATVAFHGEFAWDH-JYJNAYRXSA-N Arg-Phe-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CC1=CC=CC=C1)NC(=O)[C@H](CCCN=C(N)N)N RATVAFHGEFAWDH-JYJNAYRXSA-N 0.000 description 1
- BSYKSCBTTQKOJG-GUBZILKMSA-N Arg-Pro-Ala Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N1CCC[C@H]1C(=O)N[C@@H](C)C(O)=O BSYKSCBTTQKOJG-GUBZILKMSA-N 0.000 description 1
- UULLJGQFCDXVTQ-CYDGBPFRSA-N Arg-Pro-Ile Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)CC)C(O)=O UULLJGQFCDXVTQ-CYDGBPFRSA-N 0.000 description 1
- VENMDXUVHSKEIN-GUBZILKMSA-N Arg-Ser-Arg Chemical compound NC(N)=NCCC[C@H](N)C(=O)N[C@@H](CO)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O VENMDXUVHSKEIN-GUBZILKMSA-N 0.000 description 1
- CPTXATAOUQJQRO-GUBZILKMSA-N Arg-Val-Ser Chemical compound [H]N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CO)C(O)=O CPTXATAOUQJQRO-GUBZILKMSA-N 0.000 description 1
- AEZCCDMZZJOGII-DCAQKATOSA-N Asn-Met-Leu Chemical compound [H]N[C@@H](CC(N)=O)C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CC(C)C)C(O)=O AEZCCDMZZJOGII-DCAQKATOSA-N 0.000 description 1
- NECWUSYTYSIFNC-DLOVCJGASA-N Asp-Ala-Phe Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@H](C(O)=O)CC1=CC=CC=C1 NECWUSYTYSIFNC-DLOVCJGASA-N 0.000 description 1
- GVPSCJQLUGIKAM-GUBZILKMSA-N Asp-Arg-Arg Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCN=C(N)N)C(O)=O GVPSCJQLUGIKAM-GUBZILKMSA-N 0.000 description 1
- VPSHHQXIWLGVDD-ZLUOBGJFSA-N Asp-Asp-Asp Chemical compound OC(=O)C[C@H](N)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O VPSHHQXIWLGVDD-ZLUOBGJFSA-N 0.000 description 1
- YNCHFVRXEQFPBY-BQBZGAKWSA-N Asp-Gly-Arg Chemical compound OC(=O)C[C@H](N)C(=O)NCC(=O)N[C@H](C(O)=O)CCCN=C(N)N YNCHFVRXEQFPBY-BQBZGAKWSA-N 0.000 description 1
- 206010003805 Autism Diseases 0.000 description 1
- 208000020706 Autistic disease Diseases 0.000 description 1
- 201000006474 Brain Ischemia Diseases 0.000 description 1
- 208000013576 CDKL5 disease Diseases 0.000 description 1
- 101150032457 CDKL5 gene Proteins 0.000 description 1
- 208000027412 CDKL5-deficiency disease Diseases 0.000 description 1
- 101100512078 Caenorhabditis elegans lys-1 gene Proteins 0.000 description 1
- 102000000584 Calmodulin Human genes 0.000 description 1
- 108010041952 Calmodulin Proteins 0.000 description 1
- 206010008120 Cerebral ischaemia Diseases 0.000 description 1
- 208000028698 Cognitive impairment Diseases 0.000 description 1
- GUKYYUFHWYRMEU-WHFBIAKZSA-N Cys-Gly-Asp Chemical compound [H]N[C@@H](CS)C(=O)NCC(=O)N[C@@H](CC(O)=O)C(O)=O GUKYYUFHWYRMEU-WHFBIAKZSA-N 0.000 description 1
- 102000053602 DNA Human genes 0.000 description 1
- 230000008836 DNA modification Effects 0.000 description 1
- 230000009946 DNA mutation Effects 0.000 description 1
- 230000004543 DNA replication Effects 0.000 description 1
- 102000047174 Disks Large Homolog 4 Human genes 0.000 description 1
- 108700019745 Disks Large Homolog 4 Proteins 0.000 description 1
- 101100401560 Drosophila melanogaster mib1 gene Proteins 0.000 description 1
- 208000012661 Dyskinesia Diseases 0.000 description 1
- 208000018522 Gastrointestinal disease Diseases 0.000 description 1
- RGRMOYQUIJVQQD-SRVKXCTJSA-N Gln-Arg-His Chemical compound C1=C(NC=N1)C[C@@H](C(=O)O)NC(=O)[C@H](CCCN=C(N)N)NC(=O)[C@H](CCC(=O)N)N RGRMOYQUIJVQQD-SRVKXCTJSA-N 0.000 description 1
- BTSPOOHJBYJRKO-CIUDSAMLSA-N Gln-Asp-Arg Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O BTSPOOHJBYJRKO-CIUDSAMLSA-N 0.000 description 1
- MFJAPSYJQJCQDN-BQBZGAKWSA-N Gln-Gly-Glu Chemical compound NC(=O)CC[C@H](N)C(=O)NCC(=O)N[C@@H](CCC(O)=O)C(O)=O MFJAPSYJQJCQDN-BQBZGAKWSA-N 0.000 description 1
- HDUDGCZEOZEFOA-KBIXCLLPSA-N Gln-Ile-Ala Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](C)C(=O)O)NC(=O)[C@H](CCC(=O)N)N HDUDGCZEOZEFOA-KBIXCLLPSA-N 0.000 description 1
- IULKWYSYZSURJK-AVGNSLFASA-N Gln-Leu-Lys Chemical compound NC(=O)CC[C@H](N)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(O)=O IULKWYSYZSURJK-AVGNSLFASA-N 0.000 description 1
- OSCLNNWLKKIQJM-WDSKDSINSA-N Gln-Ser-Gly Chemical compound [H]N[C@@H](CCC(N)=O)C(=O)N[C@@H](CO)C(=O)NCC(O)=O OSCLNNWLKKIQJM-WDSKDSINSA-N 0.000 description 1
- OUBUHIODTNUUTC-WDCWCFNPSA-N Gln-Thr-Lys Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCCCN)C(=O)O)NC(=O)[C@H](CCC(=O)N)N)O OUBUHIODTNUUTC-WDCWCFNPSA-N 0.000 description 1
- ITYRYNUZHPNCIK-GUBZILKMSA-N Glu-Ala-Leu Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(O)=O ITYRYNUZHPNCIK-GUBZILKMSA-N 0.000 description 1
- BIYNPVYAZOUVFQ-CIUDSAMLSA-N Glu-Pro-Ser Chemical compound [H]N[C@@H](CCC(O)=O)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CO)C(O)=O BIYNPVYAZOUVFQ-CIUDSAMLSA-N 0.000 description 1
- WQZGKKKJIJFFOK-GASJEMHNSA-N Glucose Natural products OC[C@H]1OC(O)[C@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-GASJEMHNSA-N 0.000 description 1
- UPOJUWHGMDJUQZ-IUCAKERBSA-N Gly-Arg-Arg Chemical compound NC(=N)NCCC[C@H](NC(=O)CN)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O UPOJUWHGMDJUQZ-IUCAKERBSA-N 0.000 description 1
- LXXLEUBUOMCAMR-NKWVEPMBSA-N Gly-Asp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)O)NC(=O)CN)C(=O)O LXXLEUBUOMCAMR-NKWVEPMBSA-N 0.000 description 1
- GZBZACMXFIPIDX-WHFBIAKZSA-N Gly-Cys-Asp Chemical compound C([C@@H](C(=O)O)NC(=O)[C@H](CS)NC(=O)CN)C(=O)O GZBZACMXFIPIDX-WHFBIAKZSA-N 0.000 description 1
- CUYLIWAAAYJKJH-RYUDHWBXSA-N Gly-Glu-Tyr Chemical compound NCC(=O)N[C@@H](CCC(O)=O)C(=O)N[C@H](C(O)=O)CC1=CC=C(O)C=C1 CUYLIWAAAYJKJH-RYUDHWBXSA-N 0.000 description 1
- IDOGEHIWMJMAHT-BYPYZUCNSA-N Gly-Gly-Cys Chemical compound NCC(=O)NCC(=O)N[C@@H](CS)C(O)=O IDOGEHIWMJMAHT-BYPYZUCNSA-N 0.000 description 1
- PDUHNKAFQXQNLH-ZETCQYMHSA-N Gly-Lys-Gly Chemical compound NCCCC[C@H](NC(=O)CN)C(=O)NCC(O)=O PDUHNKAFQXQNLH-ZETCQYMHSA-N 0.000 description 1
- HJARVELKOSZUEW-YUMQZZPRSA-N Gly-Pro-Gln Chemical compound [H]NCC(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(N)=O)C(O)=O HJARVELKOSZUEW-YUMQZZPRSA-N 0.000 description 1
- NVTPVQLIZCOJFK-FOHZUACHSA-N Gly-Thr-Asp Chemical compound [H]NCC(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(O)=O)C(O)=O NVTPVQLIZCOJFK-FOHZUACHSA-N 0.000 description 1
- NIKBMHGRNAPJFW-IUCAKERBSA-N His-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@@H](N)CC1=CN=CN1 NIKBMHGRNAPJFW-IUCAKERBSA-N 0.000 description 1
- LMMPTUVWHCFTOT-GARJFASQSA-N His-Asp-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CC(=O)O)NC(=O)[C@H](CC2=CN=CN2)N)C(=O)O LMMPTUVWHCFTOT-GARJFASQSA-N 0.000 description 1
- BZKDJRSZWLPJNI-SRVKXCTJSA-N His-His-Ser Chemical compound [H]N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CC1=CNC=N1)C(=O)N[C@@H](CO)C(O)=O BZKDJRSZWLPJNI-SRVKXCTJSA-N 0.000 description 1
- QCBYAHHNOHBXIH-UWVGGRQHSA-N His-Pro-Gly Chemical compound C([C@H](N)C(=O)N1[C@@H](CCC1)C(=O)NCC(O)=O)C1=CN=CN1 QCBYAHHNOHBXIH-UWVGGRQHSA-N 0.000 description 1
- UPJODPVSKKWGDQ-KLHWPWHYSA-N His-Thr-Pro Chemical compound C[C@H]([C@@H](C(=O)N1CCC[C@@H]1C(=O)O)NC(=O)[C@H](CC2=CN=CN2)N)O UPJODPVSKKWGDQ-KLHWPWHYSA-N 0.000 description 1
- 108010033040 Histones Proteins 0.000 description 1
- 101100439048 Homo sapiens CDKL5 gene Proteins 0.000 description 1
- 101000899240 Homo sapiens Endoplasmic reticulum chaperone BiP Proteins 0.000 description 1
- 101001059454 Homo sapiens Serine/threonine-protein kinase MARK2 Proteins 0.000 description 1
- 241000725303 Human immunodeficiency virus Species 0.000 description 1
- 206010021143 Hypoxia Diseases 0.000 description 1
- AREBLHSMLMRICD-PYJNHQTQSA-N Ile-His-Arg Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N AREBLHSMLMRICD-PYJNHQTQSA-N 0.000 description 1
- 206010021750 Infantile Spasms Diseases 0.000 description 1
- VCSBGUACOYUIGD-CIUDSAMLSA-N Leu-Asn-Asp Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O VCSBGUACOYUIGD-CIUDSAMLSA-N 0.000 description 1
- JFSGIJSCJFQGSZ-MXAVVETBSA-N Leu-Ile-His Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)NC(=O)[C@H](CC(C)C)N JFSGIJSCJFQGSZ-MXAVVETBSA-N 0.000 description 1
- ZRHDPZAAWLXXIR-SRVKXCTJSA-N Leu-Lys-Ala Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](C)C(O)=O ZRHDPZAAWLXXIR-SRVKXCTJSA-N 0.000 description 1
- BMVFXOQHDQZAQU-DCAQKATOSA-N Leu-Pro-Asp Chemical compound CC(C)C[C@@H](C(=O)N1CCC[C@H]1C(=O)N[C@@H](CC(=O)O)C(=O)O)N BMVFXOQHDQZAQU-DCAQKATOSA-N 0.000 description 1
- VDIARPPNADFEAV-WEDXCCLWSA-N Leu-Thr-Gly Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)NCC(O)=O VDIARPPNADFEAV-WEDXCCLWSA-N 0.000 description 1
- QWWPYKKLXWOITQ-VOAKCMCISA-N Leu-Thr-Leu Chemical compound CC(C)C[C@H](N)C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@H](C(O)=O)CC(C)C QWWPYKKLXWOITQ-VOAKCMCISA-N 0.000 description 1
- WUHBLPVELFTPQK-KKUMJFAQSA-N Leu-Tyr-Asn Chemical compound [H]N[C@@H](CC(C)C)C(=O)N[C@@H](CC1=CC=C(O)C=C1)C(=O)N[C@@H](CC(N)=O)C(O)=O WUHBLPVELFTPQK-KKUMJFAQSA-N 0.000 description 1
- RVOMPSJXSRPFJT-DCAQKATOSA-N Lys-Ala-Arg Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](C)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O RVOMPSJXSRPFJT-DCAQKATOSA-N 0.000 description 1
- UWKNTTJNVSYXPC-CIUDSAMLSA-N Lys-Ala-Ser Chemical compound OC[C@@H](C(O)=O)NC(=O)[C@H](C)NC(=O)[C@@H](N)CCCCN UWKNTTJNVSYXPC-CIUDSAMLSA-N 0.000 description 1
- QUYCUALODHJQLK-CIUDSAMLSA-N Lys-Asp-Asp Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O QUYCUALODHJQLK-CIUDSAMLSA-N 0.000 description 1
- AAORVPFVUIHEAB-YUMQZZPRSA-N Lys-Asp-Gly Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H](CC(O)=O)C(=O)NCC(O)=O AAORVPFVUIHEAB-YUMQZZPRSA-N 0.000 description 1
- ISHNZELVUVPCHY-ZETCQYMHSA-N Lys-Gly-Gly Chemical compound NCCCC[C@H](N)C(=O)NCC(=O)NCC(O)=O ISHNZELVUVPCHY-ZETCQYMHSA-N 0.000 description 1
- KYNNSEJZFVCDIV-ZPFDUUQYSA-N Lys-Ile-Asn Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(N)=O)C(O)=O KYNNSEJZFVCDIV-ZPFDUUQYSA-N 0.000 description 1
- IUWMQCZOTYRXPL-ZPFDUUQYSA-N Lys-Ile-Asp Chemical compound [H]N[C@@H](CCCCN)C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CC(O)=O)C(O)=O IUWMQCZOTYRXPL-ZPFDUUQYSA-N 0.000 description 1
- GFWLIJDQILOEPP-HSCHXYMDSA-N Lys-Ile-Trp Chemical compound CC[C@H](C)[C@@H](C(=O)N[C@@H](CC1=CNC2=CC=CC=C21)C(=O)O)NC(=O)[C@H](CCCCN)N GFWLIJDQILOEPP-HSCHXYMDSA-N 0.000 description 1
- WBSCNDJQPKSPII-KKUMJFAQSA-N Lys-Lys-Lys Chemical compound NCCCC[C@H](N)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(O)=O WBSCNDJQPKSPII-KKUMJFAQSA-N 0.000 description 1
- YXPJCVNIDDKGOE-MELADBBJSA-N Lys-Lys-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CCCCN)NC(=O)[C@H](CCCCN)N)C(=O)O YXPJCVNIDDKGOE-MELADBBJSA-N 0.000 description 1
- 241000124008 Mammalia Species 0.000 description 1
- 208000036626 Mental retardation Diseases 0.000 description 1
- CTVJSFRHUOSCQQ-DCAQKATOSA-N Met-Arg-Glu Chemical compound CSCC[C@H](N)C(=O)N[C@@H](CCCNC(N)=N)C(=O)N[C@@H](CCC(O)=O)C(O)=O CTVJSFRHUOSCQQ-DCAQKATOSA-N 0.000 description 1
- WVTYEEPGEUSFGQ-LPEHRKFASA-N Met-Cys-Pro Chemical compound CSCC[C@@H](C(=O)N[C@@H](CS)C(=O)N1CCC[C@@H]1C(=O)O)N WVTYEEPGEUSFGQ-LPEHRKFASA-N 0.000 description 1
- 102000029749 Microtubule Human genes 0.000 description 1
- 108091022875 Microtubule Proteins 0.000 description 1
- 108010006519 Molecular Chaperones Proteins 0.000 description 1
- 206010060860 Neurological symptom Diseases 0.000 description 1
- 206010029333 Neurosis Diseases 0.000 description 1
- 108020004485 Nonsense Codon Proteins 0.000 description 1
- 102000007999 Nuclear Proteins Human genes 0.000 description 1
- 108010089610 Nuclear Proteins Proteins 0.000 description 1
- 108091034117 Oligonucleotide Proteins 0.000 description 1
- 108010033276 Peptide Fragments Proteins 0.000 description 1
- 102000007079 Peptide Fragments Human genes 0.000 description 1
- HHOOEUSPFGPZFP-QWRGUYRKSA-N Phe-Asn-Gly Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC(N)=O)C(=O)NCC(O)=O HHOOEUSPFGPZFP-QWRGUYRKSA-N 0.000 description 1
- AEEQKUDWJGOFQI-SRVKXCTJSA-N Phe-Cys-Cys Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CS)C(=O)N[C@@H](CS)C(=O)O)N AEEQKUDWJGOFQI-SRVKXCTJSA-N 0.000 description 1
- UMKYAYXCMYYNHI-AVGNSLFASA-N Phe-Gln-Asn Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CC(=O)N)C(=O)O)N UMKYAYXCMYYNHI-AVGNSLFASA-N 0.000 description 1
- GRVMHFCZUIYNKQ-UFYCRDLUSA-N Phe-Phe-Val Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](C(C)C)C(O)=O GRVMHFCZUIYNKQ-UFYCRDLUSA-N 0.000 description 1
- GLJZDMZJHFXJQG-BZSNNMDCSA-N Phe-Ser-Phe Chemical compound [H]N[C@@H](CC1=CC=CC=C1)C(=O)N[C@@H](CO)C(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O GLJZDMZJHFXJQG-BZSNNMDCSA-N 0.000 description 1
- DBALDZKOTNSBFM-FXQIFTODSA-N Pro-Ala-Asn Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](C)C(=O)N[C@@H](CC(N)=O)C(O)=O DBALDZKOTNSBFM-FXQIFTODSA-N 0.000 description 1
- CQZNGNCAIXMAIQ-UBHSHLNASA-N Pro-Ala-Phe Chemical compound C[C@H](NC(=O)[C@@H]1CCCN1)C(=O)N[C@@H](Cc1ccccc1)C(O)=O CQZNGNCAIXMAIQ-UBHSHLNASA-N 0.000 description 1
- HMNSRTLZAJHSIK-YUMQZZPRSA-N Pro-Arg Chemical compound NC(=N)NCCC[C@@H](C(O)=O)NC(=O)[C@@H]1CCCN1 HMNSRTLZAJHSIK-YUMQZZPRSA-N 0.000 description 1
- VPVHXWGPALPDGP-GUBZILKMSA-N Pro-Asn-Arg Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CC(N)=O)C(=O)N[C@@H](CCCNC(N)=N)C(O)=O VPVHXWGPALPDGP-GUBZILKMSA-N 0.000 description 1
- SGCZFWSQERRKBD-BQBZGAKWSA-N Pro-Asp-Gly Chemical compound OC(=O)CNC(=O)[C@H](CC(O)=O)NC(=O)[C@@H]1CCCN1 SGCZFWSQERRKBD-BQBZGAKWSA-N 0.000 description 1
- OZAPWFHRPINHND-GUBZILKMSA-N Pro-Cys-Val Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](CS)C(=O)N[C@@H](C(C)C)C(O)=O OZAPWFHRPINHND-GUBZILKMSA-N 0.000 description 1
- PTLOFJZJADCNCD-DCAQKATOSA-N Pro-Glu-Met Chemical compound CSCC[C@@H](C(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@@H]1CCCN1 PTLOFJZJADCNCD-DCAQKATOSA-N 0.000 description 1
- BODDREDDDRZUCF-QTKMDUPCSA-N Pro-His-Thr Chemical compound C[C@H]([C@@H](C(=O)O)NC(=O)[C@H](CC1=CN=CN1)NC(=O)[C@@H]2CCCN2)O BODDREDDDRZUCF-QTKMDUPCSA-N 0.000 description 1
- UREQLMJCKFLLHM-NAKRPEOUSA-N Pro-Ile-Ser Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)CC)C(=O)N[C@@H](CO)C(O)=O UREQLMJCKFLLHM-NAKRPEOUSA-N 0.000 description 1
- DWGFLKQSGRUQTI-IHRRRGAJSA-N Pro-Lys-Lys Chemical compound NCCCC[C@@H](C(O)=O)NC(=O)[C@H](CCCCN)NC(=O)[C@@H]1CCCN1 DWGFLKQSGRUQTI-IHRRRGAJSA-N 0.000 description 1
- HRIXMVRZRGFKNQ-HJGDQZAQSA-N Pro-Thr-Gln Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CCC(N)=O)C(O)=O HRIXMVRZRGFKNQ-HJGDQZAQSA-N 0.000 description 1
- FDMCIBSQRKFSTJ-RHYQMDGZSA-N Pro-Thr-Leu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H]([C@@H](C)O)C(=O)N[C@@H](CC(C)C)C(O)=O FDMCIBSQRKFSTJ-RHYQMDGZSA-N 0.000 description 1
- IALSFJSONJZBKB-HRCADAONSA-N Pro-Tyr-Pro Chemical compound C1C[C@H](NC1)C(=O)N[C@@H](CC2=CC=C(C=C2)O)C(=O)N3CCC[C@@H]3C(=O)O IALSFJSONJZBKB-HRCADAONSA-N 0.000 description 1
- KHRLUIPIMIQFGT-AVGNSLFASA-N Pro-Val-Leu Chemical compound [H]N1CCC[C@H]1C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CC(C)C)C(O)=O KHRLUIPIMIQFGT-AVGNSLFASA-N 0.000 description 1
- 101710118538 Protease Proteins 0.000 description 1
- 241000589516 Pseudomonas Species 0.000 description 1
- 101150058540 RAC1 gene Proteins 0.000 description 1
- 238000012228 RNA interference-mediated gene silencing Methods 0.000 description 1
- 102100022122 Ras-related C3 botulinum toxin substrate 1 Human genes 0.000 description 1
- 238000012300 Sequence Analysis Methods 0.000 description 1
- RFBKULCUBJAQFT-BIIVOSGPSA-N Ser-Cys-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CS)NC(=O)[C@H](CO)N)C(=O)O RFBKULCUBJAQFT-BIIVOSGPSA-N 0.000 description 1
- SQBLRDDJTUJDMV-ACZMJKKPSA-N Ser-Glu-Asn Chemical compound [H]N[C@@H](CO)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(N)=O)C(O)=O SQBLRDDJTUJDMV-ACZMJKKPSA-N 0.000 description 1
- JFWDJFULOLKQFY-QWRGUYRKSA-N Ser-Gly-Phe Chemical compound [H]N[C@@H](CO)C(=O)NCC(=O)N[C@@H](CC1=CC=CC=C1)C(O)=O JFWDJFULOLKQFY-QWRGUYRKSA-N 0.000 description 1
- AXVNLRQLPLSIPQ-FXQIFTODSA-N Ser-Met-Cys Chemical compound CSCC[C@@H](C(=O)N[C@@H](CS)C(=O)O)NC(=O)[C@H](CO)N AXVNLRQLPLSIPQ-FXQIFTODSA-N 0.000 description 1
- ADJDNJCSPNFFPI-FXQIFTODSA-N Ser-Pro-Ala Chemical compound OC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1C(=O)[C@@H](N)CO ADJDNJCSPNFFPI-FXQIFTODSA-N 0.000 description 1
- CUXJENOFJXOSOZ-BIIVOSGPSA-N Ser-Ser-Pro Chemical compound C1C[C@@H](N(C1)C(=O)[C@H](CO)NC(=O)[C@H](CO)N)C(=O)O CUXJENOFJXOSOZ-BIIVOSGPSA-N 0.000 description 1
- 102100028904 Serine/threonine-protein kinase MARK2 Human genes 0.000 description 1
- ZQUKYJOKQBRBCS-GLLZPBPUSA-N Thr-Gln-Gln Chemical compound C[C@H]([C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N)O ZQUKYJOKQBRBCS-GLLZPBPUSA-N 0.000 description 1
- XPNSAQMEAVSQRD-FBCQKBJTSA-N Thr-Gly-Gly Chemical compound C[C@@H](O)[C@H](N)C(=O)NCC(=O)NCC(O)=O XPNSAQMEAVSQRD-FBCQKBJTSA-N 0.000 description 1
- KJADKKWYZYXHBB-XBWDGYHZSA-N Topiramic acid Chemical compound C1O[C@@]2(COS(N)(=O)=O)OC(C)(C)O[C@H]2[C@@H]2OC(C)(C)O[C@@H]21 KJADKKWYZYXHBB-XBWDGYHZSA-N 0.000 description 1
- UUIYFDAWNBSWPG-IHPCNDPISA-N Trp-Lys-Lys Chemical compound C1=CC=C2C(=C1)C(=CN2)C[C@@H](C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](CCCCN)C(=O)O)N UUIYFDAWNBSWPG-IHPCNDPISA-N 0.000 description 1
- UPODKYBYUBTWSV-BZSNNMDCSA-N Tyr-Phe-Cys Chemical compound C([C@H](N)C(=O)N[C@@H](CC=1C=CC=CC=1)C(=O)N[C@@H](CS)C(O)=O)C1=CC=C(O)C=C1 UPODKYBYUBTWSV-BZSNNMDCSA-N 0.000 description 1
- FDKDGFGTHGJKNV-FHWLQOOXSA-N Tyr-Phe-Gln Chemical compound C1=CC=C(C=C1)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](CC2=CC=C(C=C2)O)N FDKDGFGTHGJKNV-FHWLQOOXSA-N 0.000 description 1
- COYSIHFOCOMGCF-WPRPVWTQSA-N Val-Arg-Gly Chemical compound CC(C)[C@H](N)C(=O)N[C@H](C(=O)NCC(O)=O)CCCN=C(N)N COYSIHFOCOMGCF-WPRPVWTQSA-N 0.000 description 1
- HNWQUBBOBKSFQV-AVGNSLFASA-N Val-Arg-His Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCCN=C(N)N)C(=O)N[C@@H](CC1=CN=CN1)C(=O)O)N HNWQUBBOBKSFQV-AVGNSLFASA-N 0.000 description 1
- GXAZTLJYINLMJL-LAEOZQHASA-N Val-Asn-Gln Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CC(=O)N)C(=O)N[C@@H](CCC(=O)N)C(=O)O)N GXAZTLJYINLMJL-LAEOZQHASA-N 0.000 description 1
- HGJRMXOWUWVUOA-GVXVVHGQSA-N Val-Leu-Gln Chemical compound CC(C)C[C@@H](C(=O)N[C@@H](CCC(=O)N)C(=O)O)NC(=O)[C@H](C(C)C)N HGJRMXOWUWVUOA-GVXVVHGQSA-N 0.000 description 1
- OJPRSVJGNCAKQX-SRVKXCTJSA-N Val-Met-Arg Chemical compound CC(C)[C@@H](C(=O)N[C@@H](CCSC)C(=O)N[C@@H](CCCN=C(N)N)C(=O)O)N OJPRSVJGNCAKQX-SRVKXCTJSA-N 0.000 description 1
- 201000006791 West syndrome Diseases 0.000 description 1
- JLCPHMBAVCMARE-UHFFFAOYSA-N [3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-hydroxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methyl [5-(6-aminopurin-9-yl)-2-(hydroxymethyl)oxolan-3-yl] hydrogen phosphate Polymers Cc1cn(C2CC(OP(O)(=O)OCC3OC(CC3OP(O)(=O)OCC3OC(CC3O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c3nc(N)[nH]c4=O)C(COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3CO)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cc(C)c(=O)[nH]c3=O)n3cc(C)c(=O)[nH]c3=O)n3ccc(N)nc3=O)n3cc(C)c(=O)[nH]c3=O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)O2)c(=O)[nH]c1=O JLCPHMBAVCMARE-UHFFFAOYSA-N 0.000 description 1
- 238000009825 accumulation Methods 0.000 description 1
- 230000004913 activation Effects 0.000 description 1
- 239000012190 activator Substances 0.000 description 1
- 239000013543 active substance Substances 0.000 description 1
- 239000002671 adjuvant Substances 0.000 description 1
- 108010005233 alanylglutamic acid Proteins 0.000 description 1
- 102000004111 amphiphysin Human genes 0.000 description 1
- 108090000686 amphiphysin Proteins 0.000 description 1
- 238000004458 analytical method Methods 0.000 description 1
- 230000019552 anatomical structure morphogenesis Effects 0.000 description 1
- 235000020244 animal milk Nutrition 0.000 description 1
- 238000010171 animal model Methods 0.000 description 1
- 239000001961 anticonvulsive agent Substances 0.000 description 1
- 239000012062 aqueous buffer Substances 0.000 description 1
- CKLJMWTZIZZHCS-REOHCLBHSA-L aspartate group Chemical group N[C@@H](CC(=O)[O-])C(=O)[O-] CKLJMWTZIZZHCS-REOHCLBHSA-L 0.000 description 1
- 230000002567 autonomic effect Effects 0.000 description 1
- 238000011888 autopsy Methods 0.000 description 1
- 230000003542 behavioural effect Effects 0.000 description 1
- WQZGKKKJIJFFOK-VFUOTHLCSA-N beta-D-glucose Chemical compound OC[C@H]1O[C@@H](O)[C@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-VFUOTHLCSA-N 0.000 description 1
- 239000003124 biologic agent Substances 0.000 description 1
- 230000015572 biosynthetic process Effects 0.000 description 1
- 210000004958 brain cell Anatomy 0.000 description 1
- 208000029028 brain injury Diseases 0.000 description 1
- 230000015556 catabolic process Effects 0.000 description 1
- 239000006143 cell culture medium Substances 0.000 description 1
- 239000013592 cell lysate Substances 0.000 description 1
- 210000003855 cell nucleus Anatomy 0.000 description 1
- 230000001413 cellular effect Effects 0.000 description 1
- 208000026106 cerebrovascular disease Diseases 0.000 description 1
- 238000010367 cloning Methods 0.000 description 1
- 208000010877 cognitive disease Diseases 0.000 description 1
- 238000004891 communication Methods 0.000 description 1
- 239000012141 concentrate Substances 0.000 description 1
- 238000010276 construction Methods 0.000 description 1
- 210000004748 cultured cell Anatomy 0.000 description 1
- 108010004073 cysteinylcysteine Proteins 0.000 description 1
- 108010060199 cysteinylproline Proteins 0.000 description 1
- 230000006378 damage Effects 0.000 description 1
- 230000002950 deficient Effects 0.000 description 1
- 230000006735 deficit Effects 0.000 description 1
- 238000006731 degradation reaction Methods 0.000 description 1
- 230000001934 delay Effects 0.000 description 1
- 238000013461 design Methods 0.000 description 1
- 208000025384 developmental and epileptic encephalopathy, 1 Diseases 0.000 description 1
- 239000008121 dextrose Substances 0.000 description 1
- 235000005911 diet Nutrition 0.000 description 1
- 230000037213 diet Effects 0.000 description 1
- 239000003085 diluting agent Substances 0.000 description 1
- 101150069842 dlg4 gene Proteins 0.000 description 1
- 239000002552 dosage form Substances 0.000 description 1
- 230000003828 downregulation Effects 0.000 description 1
- 238000002651 drug therapy Methods 0.000 description 1
- 230000008482 dysregulation Effects 0.000 description 1
- 230000012202 endocytosis Effects 0.000 description 1
- 238000002641 enzyme replacement therapy Methods 0.000 description 1
- 230000001973 epigenetic effect Effects 0.000 description 1
- 230000007608 epigenetic mechanism Effects 0.000 description 1
- 206010015037 epilepsy Diseases 0.000 description 1
- 230000002964 excitative effect Effects 0.000 description 1
- 230000001747 exhibiting effect Effects 0.000 description 1
- 230000006390 fear memory Effects 0.000 description 1
- 230000014061 fear response Effects 0.000 description 1
- 239000012530 fluid Substances 0.000 description 1
- 239000012634 fragment Substances 0.000 description 1
- 210000005153 frontal cortex Anatomy 0.000 description 1
- 230000005714 functional activity Effects 0.000 description 1
- 238000003197 gene knockdown Methods 0.000 description 1
- 108010081985 glycyl-cystinyl-aspartic acid Proteins 0.000 description 1
- 108010078326 glycyl-glycyl-valine Proteins 0.000 description 1
- 108010081551 glycylphenylalanine Proteins 0.000 description 1
- 239000005090 green fluorescent protein Substances 0.000 description 1
- 230000012010 growth Effects 0.000 description 1
- 239000001963 growth medium Substances 0.000 description 1
- 230000000971 hippocampal effect Effects 0.000 description 1
- 210000004295 hippocampal neuron Anatomy 0.000 description 1
- 230000007954 hypoxia Effects 0.000 description 1
- 230000006872 improvement Effects 0.000 description 1
- 238000000099 in vitro assay Methods 0.000 description 1
- 238000010874 in vitro model Methods 0.000 description 1
- 238000005462 in vivo assay Methods 0.000 description 1
- 230000002779 inactivation Effects 0.000 description 1
- 239000003112 inhibitor Substances 0.000 description 1
- 238000007918 intramuscular administration Methods 0.000 description 1
- 238000007914 intraventricular administration Methods 0.000 description 1
- 230000000302 ischemic effect Effects 0.000 description 1
- 150000002576 ketones Chemical class 0.000 description 1
- 238000011813 knockout mouse model Methods 0.000 description 1
- 150000002605 large molecules Chemical class 0.000 description 1
- 210000004185 liver Anatomy 0.000 description 1
- 239000003589 local anesthetic agent Substances 0.000 description 1
- 229960005015 local anesthetics Drugs 0.000 description 1
- 230000033001 locomotion Effects 0.000 description 1
- 230000004777 loss-of-function mutation Effects 0.000 description 1
- 210000004072 lung Anatomy 0.000 description 1
- 239000008176 lyophilized powder Substances 0.000 description 1
- 108010043322 lysyl-tryptophyl-alpha-lysine Proteins 0.000 description 1
- 229920002521 macromolecule Polymers 0.000 description 1
- 238000012423 maintenance Methods 0.000 description 1
- 239000002609 medium Substances 0.000 description 1
- 239000012528 membrane Substances 0.000 description 1
- 230000015654 memory Effects 0.000 description 1
- 206010027175 memory impairment Diseases 0.000 description 1
- 108020004999 messenger RNA Proteins 0.000 description 1
- 125000001360 methionine group Chemical group N[C@@H](CCSC)C(=O)* 0.000 description 1
- 210000004688 microtubule Anatomy 0.000 description 1
- 230000004973 motor coordination Effects 0.000 description 1
- 210000000337 motor cortex Anatomy 0.000 description 1
- 238000010172 mouse model Methods 0.000 description 1
- 210000004165 myocardium Anatomy 0.000 description 1
- 230000007383 nerve stimulation Effects 0.000 description 1
- 210000000653 nervous system Anatomy 0.000 description 1
- 230000000926 neurological effect Effects 0.000 description 1
- 230000014511 neuron projection development Effects 0.000 description 1
- 230000009223 neuronal apoptosis Effects 0.000 description 1
- 230000009207 neuronal maturation Effects 0.000 description 1
- 230000006576 neuronal survival Effects 0.000 description 1
- 230000000324 neuroprotective effect Effects 0.000 description 1
- 208000015238 neurotic disease Diseases 0.000 description 1
- 230000037434 nonsense mutation Effects 0.000 description 1
- 230000001818 nuclear effect Effects 0.000 description 1
- 238000005457 optimization Methods 0.000 description 1
- 210000003463 organelle Anatomy 0.000 description 1
- 230000036542 oxidative stress Effects 0.000 description 1
- 238000004806 packaging method and process Methods 0.000 description 1
- 239000002245 particle Substances 0.000 description 1
- 231100000255 pathogenic effect Toxicity 0.000 description 1
- 230000007903 penetration ability Effects 0.000 description 1
- 239000008177 pharmaceutical agent Substances 0.000 description 1
- 210000002826 placenta Anatomy 0.000 description 1
- 108010011110 polyarginine Proteins 0.000 description 1
- 229920002704 polyhistidine Polymers 0.000 description 1
- 230000001242 postsynaptic effect Effects 0.000 description 1
- 239000002244 precipitate Substances 0.000 description 1
- 230000002028 premature Effects 0.000 description 1
- 230000008569 process Effects 0.000 description 1
- 239000000047 product Substances 0.000 description 1
- 108010053725 prolylvaline Proteins 0.000 description 1
- 230000007398 protein translocation Effects 0.000 description 1
- 239000003642 reactive oxygen metabolite Substances 0.000 description 1
- 238000003259 recombinant expression Methods 0.000 description 1
- 230000001105 regulatory effect Effects 0.000 description 1
- 230000003014 reinforcing effect Effects 0.000 description 1
- 230000010076 replication Effects 0.000 description 1
- 230000002441 reversible effect Effects 0.000 description 1
- 230000011664 signaling Effects 0.000 description 1
- 210000002027 skeletal muscle Anatomy 0.000 description 1
- 230000007958 sleep Effects 0.000 description 1
- 208000019116 sleep disease Diseases 0.000 description 1
- 230000003997 social interaction Effects 0.000 description 1
- 239000000243 solution Substances 0.000 description 1
- 239000002904 solvent Substances 0.000 description 1
- 238000010561 standard procedure Methods 0.000 description 1
- 239000008227 sterile water for injection Substances 0.000 description 1
- 238000001356 surgical procedure Methods 0.000 description 1
- 238000012360 testing method Methods 0.000 description 1
- 238000002560 therapeutic procedure Methods 0.000 description 1
- 229960004394 topiramate Drugs 0.000 description 1
- 230000009466 transformation Effects 0.000 description 1
- 238000013519 translation Methods 0.000 description 1
- 108010029384 tryptophyl-histidine Proteins 0.000 description 1
- 108010078580 tyrosylleucine Proteins 0.000 description 1
- 230000009452 underexpressoin Effects 0.000 description 1
- 210000001186 vagus nerve Anatomy 0.000 description 1
- 229960000604 valproic acid Drugs 0.000 description 1
- 239000003981 vehicle Substances 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/10—Transferases (2.)
- C12N9/12—Transferases (2.) transferring phosphorus containing groups, e.g. kinases (2.7)
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K9/00—Medicinal preparations characterised by special physical form
- A61K9/0012—Galenical forms characterised by the site of application
- A61K9/0019—Injectable compositions; Intramuscular, intravenous, arterial, subcutaneous administration; Compositions to be administered through the skin in an invasive manner
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/62—DNA sequences coding for fusion proteins
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y207/00—Transferases transferring phosphorus-containing groups (2.7)
- C12Y207/11—Protein-serine/threonine kinases (2.7.11)
- C12Y207/11022—Cyclin-dependent kinase (2.7.11.22)
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
- C07K2319/02—Fusion polypeptide containing a localisation/targetting motif containing a signal sequence
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
- C07K2319/09—Fusion polypeptide containing a localisation/targetting motif containing a nuclear localisation signal
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
- C07K2319/10—Fusion polypeptide containing a localisation/targetting motif containing a tag for extracellular membrane crossing, e.g. TAT or VP22
Abstract
신규한 CDKL5 효소 변이체뿐만 아니라 전장 CDKL5 폴리펩티드 또는 CDKL5 변이체를 포함하는 융합 단백질이 제공된다. 그러한 융합 단백질은 세포-침투 폴리펩티드를 포함할 수 있으며 선택적으로 리더 신호 폴리펩티드 및/또는 태그를 포함한다. 또한, 그러한 CDKL5 변이체 및 융합 단백질의 제조 방법뿐만 아니라 약학 조성물, 치료 방법, 및 그러한 재조합 단백질의 용도가 제공된다.
Description
본 발명은 전반적으로 키나제 결핍 장애의 치료에 관한 것이고, 특히 CDKL5의 결핍과 관련된 장애의 치료를 위한 신규한 재조합 단백질에 관한 것이다.
CDKL5는 세린/트레오닌 키나제이며, 과거에 STK9로 알려져 있었다. 이 유전자의 돌연변이는 최근 정신 지체, 의사 소통 및 운동 능력의 상실, 유아 경련 및 발작, 비정형 레트 증후군(Rett Syndrome), 및 X-연관 웨스트 증후군(West Syndrome)과 같은 다수의 신경계 장애와 관련되어 왔다. X-연관 유전자 사이클린-의존성 키나제-유사 5(CDKL5)의 돌연변이 또는 결실은 조기 발생 중증 신경학적 손상 및 난치성 발작을 동반하는 간질성 뇌병증을 유발하는 것으로 밝혀졌다.
현재, CDKL5 결핍을 갖는 의료 문헌에 기재된 알려진 최고령자들은 41세가 되었다. 많은 다른 사람들은 20대 및 10대이지만, 이 질병은 지난 15년 동안 확인되었을 뿐이기 때문에 새로 진단된 대다수는 걸음마 단계의 유아(toddler) 또는 유아(infant)이다. CDKL5 결핍 장애로 진단된 개체는 일반적으로 신경 발달의 지연을 겪고 발작 위험이 높으며, 발병 연령 중앙값은 6주이다. 111명의 참가자로 이루어진 한 연구에 따르면, 개체의 85.6%가 발작이 매일 발생하는 간질을 가졌으며 발작은 하루 평균 6건이었다.
현재의 치료법은 발작 약물 치료로부터 케톤생성 식이요법, 미주 신경 자극, 및 수술에 이르기까지 다양하다. 일반적으로 투여되는 항-간질 약물에는 클로바잠, 발프로산 및 토피라메이트가 포함되고, 많은 경우에 둘 이상의 약물 요법이 동시에 사용된다. 개체는 새로운 유형의 약물 치료를 시작한 후 일정 기간 동안 발작이 없는 "허니문 기간(honeymoon period)"을 갖는 것으로 보였지만, 궁극적으로 발작의 재발이 있게 된다. 관찰된 허니문의 지속 기간은 2개월 내지 7년이며, 중앙값은 6개월이다. 예를 들어, 연구에 따르면, 111명의 참가자 중 16명은 현재 발작이 없었고, 1명의 개체는 발작을 일으킨 적이 없었다.
병원성 발현에 대한 정확한 메카니즘은 여전히 명확하지 않다. 일부 실험 데이터는 C-말단에서의 특정 넌센스(non-sense) 돌연변이가 단백질이 핵에 항시적으로 위치하게 하는 한편, 다른 미스센스 돌연변이가 세포질에서 높게 표현됨을 시사한다. 핵 위치 신호 및 핵 배출 신호가 모두 단백질의 C-말단에서 확인되었다.
일부 돌연변이 효소 변이체는 인산화 기능의 부분적 또는 완전한 상실을 가져오는 한편, 다른 돌연변이 및 트렁케이션(truncation)은 인산화 능력의 증가를 가져오며, 이는 기능 상실 및 획득 둘 모두가 병원성일 수 있음을 시사한다. 효소 활성 상실/기능 획득 및 효소 핵 위치 대 세포질 내 체류에 기인하는 상호 작용 및 병원성 효과는 여전히 불명확하다. 광범위한 CDKL5 돌연변이를 가지며 임상 증상을 나타내는 환자의 분석은 임상 증상을 유발하는 돌연변이가 C-말단 또는 키나제 활성 도메인에서 발견될 가능성이 높음을 시사하고, 이는 CDKL5의 키나제 활성 및 단백질 전위 능력 둘 모두가 증상의 임상 발현에 영향을 줄 수 있음을 시사한다.
따라서, 본 발명의 다양한 양태는 신규한 CDKL5 변이체 및 CDKL5 융합 단백질에 관한 것이며, 이들은 CDKL5 결핍 또는 CDKL5 돌연변이 또는 결핍에 의해 야기된 비정형 레트 증후군과 같은 CDKL5-매개 신경계 장애를 치료하는 데 사용될 수 있다. 본 발명의 다른 양태는 그러한 CDKL5 변이체 및 융합 단백질의 제조 방법뿐만 아니라 약학 조성물, 치료 방법, 및 그러한 재조합 단백질의 용도에 관한 것이다.
본 발명의 일 양태는 본원에 기재된 바와 같은 CDKL5 폴리펩티드에 관한 것이다. 하나 이상의 구현예에서, CDKL5 폴리펩티드는 SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 대해 적어도 98%의 서열 동일성을 갖는 서열을 포함한다. 하나 이상의 구현예에서, CDKL5 폴리펩티드는 SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 대해 적어도 99%의 서열 동일성을 갖는 서열을 포함한다. 하나 이상의 구현예에서, CDKL5 폴리펩티드는 SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 대해 100%의 서열 동일성을 갖는 서열을 포함한다.
본 발명의 다른 양태는 핵 배출 신호(NES)를 결여한 CDKL5 폴리펩티드에 관한 것이다. 하나 이상의 구현예에서, CDKL5 폴리펩티드는 핵 위치 신호(NLS)를 함유한다.
본 발명의 다른 양태는 핵 위치 신호(NLS)를 결여하고 핵 배출 신호(NES)를 함유하는 CDKL5 폴리펩티드에 관한 것이다.
본 발명의 다른 양태는 본원에 기재된 바와 같은 CDKL5 폴리펩티드 및 세포-침투 폴리펩티드를 포함하는 융합 단백질에 관한 것이다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 90%의 서열 동일성을 갖는다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 95%의 서열 동일성을 갖는다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 100%의 서열 동일성을 갖는다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 적어도 90%의 서열 동일성을 갖는다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 적어도 95%의 서열 동일성을 갖는다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 적어도 100%의 서열 동일성을 갖는다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 적어도 90%의 서열 동일성을 갖는다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 적어도 95%의 서열 동일성을 갖는다. 하나 이상의 구현예에서, 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 적어도 100%의 서열 동일성을 갖는다. 다양한 구현예에서, CDKL5 폴리펩티드는 (예를 들어, SEQ ID NO. 1 또는 SEQ ID NO: 47에 제시된 바와 같은) 전장 CDKL5 폴리펩티드이다. 다른 구현예에서, CDKL5 폴리펩티드는 (예를 들어, SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 제시된 바와 같은) 본원에 기재된 바와 같은 변이체이다.
본 발명의 다른 양태는 본원에 기재된 바와 같은 CDKL5 폴리펩티드 또는 본원에 기재된 바와 같은 융합 단백질, 및 약학적으로 허용되는 담체를 포함하는 약학 제형에 관한 것이다.
본 발명의 다른 양태는 본원에 기재된 바와 같은 CDKL5 폴리펩티드 또는 본원에 기재된 바와 같은 융합 단백질; 및 약학적으로 허용되는 담체를 포함하는 제형을 투여하는 단계를 포함하는, CDKL5-매개 신경계 장애를 치료하는 방법에 관한 것이다. 하나 이상의 구현예에서, 제형은 척수강내 투여된다. 하나 이상의 구현예에서, 제형은 정맥내 투여된다. 하나 이상의 구현예에서, 제형은 수조내 투여된다. 하나 이상의 구현예에서, 제형은 뇌실내 투여된다. 하나 이상의 구현예에서, 제형은 실질내 투여된다. 하나 이상의 구현예에서, CDKL5-매개 신경계 장애는 CDKL5 결핍, 또는 CDKL5 돌연변이 또는 결핍에 의해 야기된 비정형 레트 증후군 중 하나 이상이다.
본 발명의 다른 양태는 본원에 기재된 바와 같은 CDKL5 폴리펩티드 또는 본원에 기재된 바와 같은 융합 단백질의 제조 방법에 관한 것이다. 하나 이상의 구현예에서, 방법은 CDKL5 폴리펩티드 또는 융합 단백질을 발현시키는 단계; 및 CDKL5 폴리펩티드 또는 융합 단백질을 정제하는 단계를 포함한다. 하나 이상의 구현예에서, CDKL5 폴리펩티드 또는 융합 단백질은 차이니즈 햄스터 난소(CHO) 세포, HeLa 세포, 인간 배아 신장(HEK) 세포 또는 대장균 세포에서 발현된다.
본 발명의 다른 양태는 본원에 기재된 바와 같은 CDKL5 폴리펩티드 또는 본원에 기재된 바와 같은 융합 단백질을 인코딩하는 폴리뉴클레오티드에 관한 것이다. 본 발명의 다른 양태는 그러한 폴리뉴클레오티드를 포함하는 벡터에 관한 것이다.
도 1a는 CDKL5107의 폴리펩티드 맵(map)을 나타낸다. 맵은 ATP 결합 부위, 키나제 도메인 및 키나제 활성 부위, 2개의 핵 위치 신호, 및 핵 배출 신호를 비롯한 폴리펩티드의 중요한 특징을 확인해준다.
도 1b 및 도 1c는 합성된 CDKL5 작제물 변이체를 도시한 그래프를 나타내고(도 1b), 범례는 작제물이 어떻게 합성되었는지를 설명하기 위해 관련 아미노산 결실 정보와 함께 폴리펩티드의 길이를 기술한다(도 1c).
도 2a 내지 도 2ad는 CHO 세포 또는 대장균 세포와 같은 세포에서 다양한 융합 단백질을 발현하기 위한 예시적인 플라스미드를 나타낸다.
도 1b 및 도 1c는 합성된 CDKL5 작제물 변이체를 도시한 그래프를 나타내고(도 1b), 범례는 작제물이 어떻게 합성되었는지를 설명하기 위해 관련 아미노산 결실 정보와 함께 폴리펩티드의 길이를 기술한다(도 1c).
도 2a 내지 도 2ad는 CHO 세포 또는 대장균 세포와 같은 세포에서 다양한 융합 단백질을 발현하기 위한 예시적인 플라스미드를 나타낸다.
본 발명의 몇몇 예시적인 구현예를 설명하기 전에, 본 발명은 다음의 설명에 제시된 구성 또는 공정 단계의 세부 사항으로 제한되지 않음을 이해해야 한다. 본 발명은 다른 구현예가 가능하며 다양한 방식으로 실시되거나 수행될 수 있다.
본 발명의 다양한 양태는 신규한 CDKL5 변이체 및 CDKL5 융합 단백질에 관한 것이다. 본 발명의 다른 양태는 그러한 CDKL5 변이체 및 융합 단백질의 제조 방법뿐만 아니라 약학 조성물, 치료 방법, 및 그러한 재조합 단백질의 용도에 관한 것이다.
임의의 특정 이론에 구속시키고자 함이 없이, 기능적 활성을 유지하는 짧은 CDKL5 변이체는, 특히 CDKL5 폴리펩티드를 포함하는 융합 단백질에 통합될 때, 전장 CDKL5 폴리펩티드에 비해 이점을 제공할 수 있는 것으로 여겨진다. 하나 이상의 구현예에서, 그러한 이점은 단백질 생성 동안 숙주 세포로부터의 분비 개선, 용해도 개선, 혈액-뇌 장벽(BBB)을 횡단하는 능력 향상, 및/또는 표적 세포에 침투하는 능력 향상을 포함할 수 있다.
정의
본원에 사용되는 바와 같이, "CDKL5-매개 신경계 장애"는 CDKL5 단백질의 발현 또는 과발현에 의해 치료될 수 있는 임의의 질병 또는 장애를 지칭한다.
본원에 사용되는 바와 같이, "CDKL5 결핍"은 단백질의 생물학적 기능에서의 임의의 결핍을 지칭한다. 결핍은 단백질을 코딩하는 DNA 또는 DNA 관련 조절 영역에서의 임의의 DNA 돌연변이, 또는 DNA 메틸화 또는 히스톤 변형을 포함하지만 이로 제한되지 않는 후성학적 DNA 변형에서의 임의의 변화, CDKL5 단백질의 2차, 3차, 또는 4차 구조의 임의의 변화, 또는 야생형 또는 정상 대상체와 비교되는 생물학적 기능을 수행하는 CDKL5 단백질의 능력의 임의의 변화로 인한 단백질 기능의 임의의 변화에 기인할 수 있다. 결핍은 CDKL5 단백질의 결여, 예컨대 완전히 기능하는 단백질의 널(null) 돌연변이 또는 저발현을 또한 포함할 수 있다.
본원에 사용되는 바와 같이, "CDKL5 돌연변이 또는 결핍에 의해 야기된 비정형 레트 증후군"은 레트 증후군과 유사한 임상 징후를 갖는 비정형 형태의 레트 증후군을 지칭하지만 CDKL5 돌연변이 또는 결핍에 의해 야기된다.
CDKL5 결핍, 레트 증후군, 또는 비정형 레트 증후군의 증상 또는 마커는 발작, 인지 장애, 저산소증뿐만 아니라 자율신경, 수면, 및 위장 장애를 포함하지만 이로 제한되지는 않는다.
본원에 사용되는 바와 같이, 용어 "담체"는 화합물과 함께 투여되는 희석제, 애주번트, 부형제, 또는 비히클을 지칭하는 것으로 의도된다. 적합한 약학적 담체는 당업계에 공지되어 있으며, 적어도 하나의 구현예에서, 문헌["Remington's Pharmaceutical Sciences" by E. W. Martin](제18 판) 또는 이 문헌의 다른 판본에 기재되어 있다.
본원에 사용되는 바와 같이, 용어 "효소 대체 요법" 또는 "ERT"는 정제된 외인성 효소가 그러한 효소의 결핍을 갖는 개체로 도입되는 것을 지칭하는 것으로 의도된다. 투여된 단백질은 천연 공급원으로부터 얻거나 재조합 발현에 의해 얻을 수 있다. 이 용어는 또한 정제된 효소의 투여를 달리 필요로 하거나 이로부터 이익을 얻는 개체로의 정제된 효소의 도입을 지칭한다. 적어도 하나의 구현예에서, 그러한 개체는 효소 부족을 겪는다. 도입된 효소는 시험관내에서 생성된 정제된 재조합 효소, 또는 예를 들어 태반 또는 동물 젖과 같은 분리된 조직 또는 유체로부터 정제되거나 식물로부터 정제된 단백질일 수 있다.
본원에 사용되는 바와 같이, 용어 "대상체" 또는 "환자"는 인간 또는 비-인간 동물을 지칭하는 것으로 의도된다. 적어도 하나의 구현예에서, 대상체는 포유 동물이다. 적어도 하나의 구현예에서, 대상체는 인간이다.
본원에 사용되는 바와 같이, "치료적 유효량" 및 "유효량"은 대상체에서 치료 반응을 가져오기에 충분한 재조합 단백질(예를 들어, CDKL5 변이체 또는 융합 단백질)의 양을 지칭하는 것으로 의도된다. 치료 반응은, 본원에 기재되고 당업계에 공지된 임의의 대용 임상 마커 또는 증상을 비롯한, 사용자(예를 들어, 임상의)가 요법에 대한 효과적인 반응으로 인식할 임의의 반응일 수 있다. 따라서, 적어도 하나의 구현예에서, 치료 반응은 당업계에 공지된 것들과 같은 CDKL5 결핍, 레트 증후군, 또는 비정형 레트 증후군의 하나 이상의 증상 또는 마커의 개선 또는 억제일 수 있다.
CDKL5 단백질의 기능
인간 CDKL5 유전자는 24개의 엑손으로 구성되며, 이들 중 처음 3개(엑손 1, 엑손 1a 및 엑손 1b)는 번역되지 않는다.
원래 발견된 인간 CDKL5 변이체는 분자량이 115 kDa인 1030개의 아미노산(CDKL5115)이었다. 다른 두드러진 변이체인 CDKL5107은 변경된 C-말단 영역을 함유하는데, 이는 선택적 스플라이싱이 CDKL5115 변이체의 경우와는 상이한 엑손들을 조합하기 때문이다. CDKL5107(107 kDa)은 더 짧은데, 이는 그것이 엑손 19의 대안적 버전(alternate version)을 보유하며 CDKL5115 변이체에 존재하는 엑손 20 내지 엑손 21을 함유하지 않기 때문이다. hCDKL5107 mRNA는 hCDKL5115 전사체보다 인간 뇌에서 37배 더 풍부한 것으로 밝혀졌고, 뮤린 CDKL5107은 뮤린 뇌에서 뮤린 CDKL5105 변이체보다 160배 더 풍부한 것으로 밝혀졌다. 인간 및 뮤린 CDKL5107 아이소형 둘 모두는 인간 CDKL5115 변이체와 비교하여 더 긴 반감기 및 분해 내성을 나타냈다.
CDKL5 녹아웃 마우스 모델은 Lox-Cre 재조합 시스템을 사용하여 생성되었고, 이들 마우스는 사회적 상호 작용에서의 자폐-유사 결함, 운동 조절 장애, 및 공포 기억의 상실의 증상을 나타낸다(문헌[Wang et al., Proc Natl Acad Sci U.S.A, 109(52), 21516-21521]). 예를 들어, 녹아웃 CDKL5 마우스는 운동 협응 감소의 증상을 가지며 자극에 반복적으로 노출될 때 기억력 및 공포 반응 장애를 나타낸다. 이러한 변화로 인해 과학자들은 CDKL5 키나제 활성의 상실이 뉴런 네트워크 발달 장애를 초래한다는 가설을 세웠다. 이전 데이터는 CDKL5가 메틸-CpG 결합 단백질 2(MeCP2)를 인산화하고, MeCP2에서의 독립적인 기능 상실 돌연변이가 레트 증후군 표현형을 가져온다는 것을 시사하였다. CDKL5의 다른 기질은 네트린(Netrin) G1 리간드(NGL-1), 슈틴(Shootin)1(SHTN1), 마인드밤(Mindbomb) 1(MIB1), DNA (시토신-5)-메틸트랜스퍼라제 1(DNMT1), 암피피신(Amphiphysin) 1(AMPH1), 말단-결합 단백질 EB2, 미세소관 관련 단백질 1S(MAP1S) 및 히스톤 데아세틸라제 4(HDAC4)를 포함한다. CDKL5의 정확한 역할이 아직 밝혀지지 않았지만, 이들 데이터는 CDKL5가 MeCP2를 비롯한 올바른 뉴런 발달에 중요한 다운스트림 표적의 인산화에서 역할을 한다는 것을 시사한다. 인간에서, CDKL5의 돌연변이는 레트 증후군과 중첩되며 추가로 조기 발생 발작을 나타내는 표현형과 관련된다. CDKL5 KO 마우스는 임의의 조기 발생 발작 증상을 나타내지 않았지만, 운동 결함, 사교성 감소, 및 학습 및 기억력 장애를 나타냈다(문헌[Chen et al. CDKL5, a protein associated with Rett Syndrome, regulates neuronal morphogenesis via Rac1 signaling, J Neurosci 30: 12777-12786]).
2개의 CDKL5 아이소형이 래트에서 발견되는데, 하나는 CDKL5a로 일컬어지고 다른 하나는 CDKL5b로 일컬어진다(문헌[Chen et al.]). 일반적으로, C-말단 근처의 마지막 100개 내지 150개 아미노산을 제외하면, 인간, 래트, 및 마우스 종에 걸쳐 CDKL5 유전자에서 높은 수준의 서열 보존이 존재한다. 웨스턴 블롯 데이터에 따르면, 래트 발달 동안 두 변이체 모두가 존재하지만 성체는 단일 변이체를 우세하게 발현하는 것으로 보인다. 또한, CDKL5는 뇌, 간, 및 폐에 확인 가능한 양으로 존재한다.
CDKL5는 핵에서 기능하지만 배양된 뉴런의 수상 돌기에서도 발견되며, 이는 가능한 대안적 세포질 역할을 시사한다. 배양된 피질 뉴런에서의 RNAi(RNA 간섭)에 의한 CDKL5 발현의 하향 조절은 신경 돌기 성장 및 수상 돌기 분지(dendritic arborization)(분지형성(branching))를 억제하였고, CDKL5의 과발현은 반대 효과를 가졌다(문헌[Chen et al.]). CDKL5의 핵 및 세포질 효과 둘 모두를 특성규명하기 위해, 핵 배출 서열(NES)을 갖는 CDKL5a의 변이체를 배양된 피질 뉴런 RNAi 모델에서 발현시켰다. 이 NES-CDKL5a 변이체는 야생형 유전자 발현을 침묵시키는 데 사용된 RNAi에 내성이 있으므로, 세포질에서만 발현될 때 CDKL5a를 모델링하는 데 사용되었다. 이 CDKL5 변이체가 오로지 세포질에 존재한다는 것을 확인하기 위해 GFP 태그를 사용한 후, 신경 돌기 길이 및 신경 돌기 분지 수의 증가가 관찰되었다. 내인성 CDKL5 발현을 녹다운시키기 위해 RNAi가 사용될 때 관찰되는 질병 표현형을 부분적으로 구제하는 NES-GFP-CDKL5a의 능력은 세포질에서의 CDKL5의 발현이 신경 돌기의 발달 및 성장에서 중요한 인자임을 시사한다.
CDKL5의 인간 돌연변이는 레트 증후군과 유사한 표현형과 관련이 있으며, CDKL5 돌연변이를 갖는 개체는 또한 조기 발생 발작을 나타낸다. 이러한 발작의 발생은 레트 증상의 발생 전에 초기 정상 발달 기간이 존재하는 고전적인 레트 증후군 표현형과 다르다. 고전적인 레트 증후군(RTT)을 갖는 환자는 6개월령 내지 18개월령까지 정상적으로 발달하는 것으로 보이고, 이어서 이들 환자는 언어 및 운동 상실을 비롯한 신경학적 증상을 나타내기 시작한다. RTT 뇌의 부검은 운동 및 전두 피질에서 더 짧은 수상 돌기를 갖는 더 작고 더 조밀하게 팩킹된 뉴런을 나타내며, 이는 뉴런 발달이 손상되어 있음을 시사한다. 대부분의 고전적 RTT 사례는 MECP2 유전자의 돌연변이에 기인하며, 이 유전자는 포유 동물 게놈에서 CpG 디뉴클레오티드에 선택적으로 결합하고 복합체의 동원을 통해 전사를 조절하는 핵 단백질을 인코딩하는 X-연관 유전자이다. 불충분하게 이해되어 있지만, MECP2의 돌연변이에 의해 유발된 유전자 발현의 조절 이상이 레트 증후군의 근본 원인인 것으로 일반적으로 생각된다. 고전적 레트 증후군 사례의 대략 20% 및 다른 레트 증후군 변이체의 60% 내지 80%는 MECP2에 돌연변이를 지니지 않으며, 이는 병인에 대한 대안적인 유전적 원인을 시사한다. 최근에, 일부 CDKL5 돌연변이가 RTT의 특정 변이체 및 다른 중증 뇌병증을 갖는 환자에서 확인되었으며, CDKL5는 생체내 및 시험관내 모두에서 MeCP2와 상호 작용하는 것으로 밝혀졌다. MeCP2 이외에, CDKL5는 NGL-1을 비롯한 다수의 다운스트림 표적과 상호 작용하고 이를 인산화하는 것으로 밝혀졌다. 인산화될 때, NGL-1은 PSD95와 상호 작용하고 수상돌기 가시 및 시냅스 형성의 올바른 발생 및 발달에 중요하다(문헌[Ricciardi S, et al. "CDKL5 ensures excitatory synapse stability by reinforcing NGL-1-PSD95 interaction in the postsynaptic compartment and is impaired in patient iPSC-derived neurons." Nat Cell Biol 14(9):911-923]).
CDKL5는 또한 단백질 DNA 메틸트랜스퍼라제 1(DNMT1)을 인산화하는 것으로 밝혀졌다(문헌[Kameshita I, et al. "Cyclin-dependent kinase-like 5 binds and phosphorylates DNA methyltransferase 1." Biochem Biophys Res Commun 377:1162-1167]). 이 인산화는 DNMT1의 활성화를 가져오며, DNMT1는 헤미메틸화된(hemimethylated) DNA를 우선적으로 메틸화하는 유지형(maintenance-type) 메틸화 단백질이다. 이 공정은 DNA 복제 동안 DNA 메틸화 패턴의 유지에 유용하여, 새로 합성된 딸(daughter) DNA 가닥이 그것이 대체한 모 가닥의 메틸화 패턴을 유지할 수 있게 한다. DNA의 메틸화가 일반적으로 유전자 발현을 침묵시키는 후성학적 메커니즘인 것으로 생각됨에 따라, DNMT1의 이러한 유지 기능은 세포 세대에 걸쳐 유전자 발현 패턴을 보존하는 데 중요하다.
현재의 모델은 CDKL5 키나제 도메인이 GSK-3β를 인산화하고, GSK-3β의 인산화가 그의 비활성화를 가져온다는 것을 시사한다. 그에 따라 CDKL5 활성이 결핍된 개체는 증가된 GSK-3β 활성을 나타내는 것으로 보인다. 이전의 연구에 따르면, GSK-3β는 해마 신경발생을 조절하고, GSK-3β의 증가된 활성이 신생아 해마 뉴런의 수상돌기 형태를 심각하게 손상시키는 것으로 밝혀졌다. 또한, GSK-3β는 뉴런 생존 및 성숙과 같은 주요 발달 사건의 음성 조절 인자로서 작용하는 것으로 보인다. CDKL5 KO 마우스를 사용하여 수행된 연구는 GSK-3β 억제제에 의한 처리가 CDKL5 활성이 결핍된 마우스에서 해마 발달 및 행동 결함을 거의 완전히 구제할 수 있음을 입증했다(문헌[Fuchs et al. "Inhibition of GSK3β Rescues Hippocampal Development and Learning in a Mouse Model of CDKL5 Disorder." Neurobiology of Disease 82: 298-310]). 이 발달 구제는 또한 치료 이후에도 지속되는 것으로 보였다.
CDKL5
107
폴리펩티드 작제물
도 1a는 CDKL5107의 폴리펩티드 맵을 나타낸다. 야생형 전장 인간 CDKL5107 아이소형의 아미노산 서열은 SEQ ID NO: 1에 제공된다. CDKL5107 단백질은 960개의 아미노산으로 구성되고, 키나제 도메인은 처음 약 300개의 아미노산에 포함된다. 960개 중 잔기 42는 인산화 반응 동안 ATP 결합에 참여하는 키나제 도메인 내에 위치한 주요 리신 잔기이고, 이 잔기의 돌연변이는 일반적으로 키나제 활성의 상실("키나제 사멸")을 가져온다. 또한, 2개의 핵 위치 신호가 스패닝 잔기 312-315(NLS1) 및 스패닝 잔기 784-789(NLS2)에 존재하고, 핵 배출 신호(NES)가 스패닝 잔기 836-845에 존재한다. 잔기 905 내지 960에 걸쳐 있는 C-말단의 아미노산은 CDKL5107에 고유하며 CDKL5115에는 존재하지 않는다. 아미노산 잔기 1-904는 CDKL5115와 CDKL5107 사이에 동일하다. 야생형 전장 인간 CDKL5115 아이소형의 아미노산 서열은 SEQ ID NO: 47에 제공된다.
본 발명의 다양한 구현예는 신규한 CDKL5 변이체를 제공한다. 도 1b 및 도 1c는 전장 인간 CDKL5107 아이소형(작제물 1) 및 신규한 CDKL5 작제물(작제물 2 내지 작제물 12로 표시됨)의 폴리펩티드를 나타낸다. 이들 CDKL5 작제물은 일반적으로 두 가지 범주에 속한다: C-말단에서 몇 개의 아미노산이 결여된 것(작제물 2 내지 작제물 7) 및 폴리펩티드 사슬의 중간에서 몇 개의 아미노산이 결여된 것(작제물 8 내지 작제물 12). 더욱이, CDKL5가 추가의 N-말단 아미노산 서열에 C-말단적으로 융합된 작제물에서, CDKL5의 개시 메티오닌이 제거된다. 이들 작제물에서, CDKL5 폴리펩티드는 제2 아미노산인 리신으로 시작한다. 작제물 1은 전장 인간 CDKL5107 아이소형의 모든 960개의 아미노산을 함유한다. 전체 960개 아미노산 사슬 중 처음 851개 아미노산을 함유하는 작제물 2는, CDKL5107과 CDKL5115 사이에 상이한 꼬리 서열이 제거되지만 키나제 도메인, 핵 위치 신호(NLS1 및 NLS2), 및 핵 배출 신호(NES)는 온전히 유지되는 단축된 CDKL5 폴리펩티드를 나타낸다. 작제물 3은 추가로 단축되며, 여기서 핵 위치 신호(NLS2) 및 핵 배출 신호(NES)가 추가로 제거된다. 도 1b 및 도 1c에 나타낸 바와 같이, 작제물 4 내지 작제물 7은 훨씬 더 단축된다. 작제물 2 내지 작제물 7은 모두 활성 키나제 도메인을 함유하는 한편, 작제물 3 내지 작제물 7은 NLS2 또는 NES 서열을 함유하지 않는다. 작제물 7은 NLS1 서열까지 추가로 단축된다. 나머지 작제물(작제물 8 내지 작제물 12)은 모두 CDKL5107에 고유한 C-말단 아미노산을 보유하면서 폴리펩티드 사슬의 중간 부분에서 결실을 갖는다. 이들 작제물 중, 작제물 12는 NES 및 NLS2 서열을 결여하고 있다. 작제물 1 내지 작제물 12의 아미노산 서열은 각각 SEQ ID NO: 1 내지 SEQ ID NO: 12에 제공된다.
하나 이상의 구현예에서, CDKL5 폴리펩티드는 SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 대해 적어도 98%, 적어도 98.5%, 적어도 99% 또는 적어도 99.5%의 서열 동일성을 갖는다. CDKL5 폴리펩티드는, SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12로 기재된 아미노산 서열에 대해 1개, 2개, 3개, 4개, 5개, 6개, 7 개, 8개, 9개, 10개, 11개, 12개, 13개, 14개, 15개 또는 그 초과의 결실, 치환 및/또는 삽입을 갖는 것과 같은, SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 대한 결실, 치환 및/또는 삽입을 함유할 수 있다.
하나 이상의 구현예에서, CDKL5 폴리펩티드는 SEQ ID NO: 1 또는 SEQ ID NO: 47에 대해 적어도 98%, 적어도 98.5%, 적어도 99% 또는 적어도 99.5%의 서열 동일성을 갖는다. CDKL5 폴리펩티드는, SEQ ID NO: 1 또는 SEQ ID NO: 47로 기재된 아미노산 서열에 대해 1개, 2개, 3개, 4개, 5개, 6개, 7 개, 8개, 9개, 10개, 11개, 12개, 13개, 14개, 15개 또는 그 초과의 결실, 치환 및/또는 삽입을 갖는 것과 같은, SEQ ID NO: 1 또는 SEQ ID NO: 47에 대한 결실, 치환 및/또는 삽입을 함유할 수 있다.
GCG 서열 분석 패키지(미국 위스콘신주 매디슨 소재의 University of Wisconsin)의 일부로서 이용 가능하며, 예를 들어, 초기 설정으로 사용될 수 있는 FASTA 또는 BLAST를 비롯한 다양한 정렬 알고리즘 및/또는 프로그램이 2개의 서열 사이의 동일성을 계산하기 위해 사용될 수 있다. 예를 들어, 본원에 기재된 특정 폴리펩티드에 대해 적어도 98%, 98.5%, 99% 또는 99.5%의 동일성을 가지며 바람직하게는 실질적으로 동일한 기능을 나타내는 폴리펩티드뿐만 아니라 그러한 폴리펩티드를 인코딩하는 폴리뉴클레오티드가 고려된다. 달리 지시되지 않는 한, 유사성 점수는 BLOSUM62의 사용에 기초할 것이다. BLASTP가 사용될 때, 유사성 퍼센트는 BLASTP 양성 스코어에 기초하고, 서열 동일성 퍼센트는 BLASTP 동일성 스코어에 기초한다. BLASTP "동일성"은 높은 스코어링 서열 쌍에서의 동일한 총 잔기의 수 및 분율을 나타내고; BLASTP "양성"은 정렬 스코어가 양의 값을 가지며 서로 유사한 잔기의 수 및 분율을 나타낸다. 본원에 개시된 아미노산 서열과 이러한 정도의 동일성 또는 유사성 또는 임의의 중간 정도의 동일성 또는 유사성을 갖는 아미노산 서열이 본 개시에 의해 고려되고 포함된다. 유사한 폴리펩티드의 폴리뉴클레오티드 서열은 유전자 코드를 사용하여 추론되고, 통상적인 수단, 특히 유전자 코드를 사용하여 그의 아미노산 서열을 역 번역함으로써 얻을 수 있다.
당업자는 특정 폴리펩티드 서열을 인코딩하는 폴리뉴클레오티드 서열을 용이하게 유도할 수 있다. 그러한 폴리뉴클레오티드 서열은, OptimumGeneTM 코돈 최적화 도구(미국 뉴저지주 피스카터웨이 소재의 GenScript)를 사용하는 것과 같이, 시판되는 제품을 사용하여 표적 세포에서의 발현을 위해 코돈 최적화될 수 있다.
세포-침투 펩티드(CPP)
다양한 바이러스 및 세포 단백질은 세포막을 가로지르는 전위를 매개하는 기본 폴리펩티드 서열을 보유한다. 세포막을 가로질러 전위하는 능력은 막을 가로지르는 고분자량 폴리펩티드의 전달을 위한 중요한 도구가 되었다. "단백질 전달 도메인(protein transduction domain)"(PTD) 및 "세포-침투 펩티드"(CPP)라는 문구는 전부는 아니지만 다수의 포유 동물 세포의 원형질 막을 통과할 수 있는 짧은 펩티드(30개 미만의 아미노산)를 지칭하기 위해 일반적으로 사용된다. 이들이 원형질막을 집합적으로 횡단하게 하는 도메인의 특정 특성을 확인하기 위한 연구 후, 연구자들은 이들 도메인이 리신 및 아르기닌과 같은 다수의 염기성 아미노산 잔기를 함유한다는 것을 관찰하였다. 따라서, 세포-침투 펩티드는 두 가지 부류로 분류되는데, 첫 번째 부류는 양전하에 기여하는 리신 잔기를 함유하는 양친매성 나선형 펩티드로 구성되는 한편, 두 번째 부류는 아르기닌이 풍부한 펩티드를 포함한다. 이들 펩티드는 세포내 표적에 전달하기 어려운 다른 단백질과 함께 사용되는 경우 치료 가능성을 가질 수 있다. PTD의 가장 빈번한 실험 용도는 TAT, 안테나페디아(Antennapedia)(Antp), 및 기타 폴리-아르기닌 펩티드이다.
지금까지, TAT는 PTD의 가장 잘 특성규명된 것이며, 짧은 펩티드 및 올리고뉴클레오티드와 같은 작은 카고(cargo)를 세포간 표적에 성공적으로 전달하는 데 사용되어 왔다. HIV-TAT(HIV 전사 활성화제)는 인간 면역 결핍 바이러스 타입 1(HIV-1)의 복제에 관여하는 86개 아미노산 단백질이며, 많은 연구에 의하면 TAT는 바이러스 게놈의 전사를 활성화시키기 위해 원형질막을 통해 전위하여 핵에 도달할 수 있는 것으로 밝혀졌다. 연구에 의하면 TAT는 몇몇 상이한 단백질에 커플링될 때 그의 침투 특성을 유지하는 것으로 또한 밝혀졌다. TAT 단백질의 어느 영역이 전위 특성에 중요한지를 이해하기 위해, TAT의 다양한 길이의 펩티드 단편이 합성되고 그들의 침투 능력이 평가되는 실험이 수행되었다(문헌[Lebleu et al. "A Truncated HIV-1 TAT Protein Basic Domain Rapidly Translocates through the Plasma Membrane and Accumulates in the Cell Nucleus." J. Biol. Chem. 1997, 272:16010-16017]). 염기성 아미노산의 영역은 이러한 침투 특성을 유지하는 TAT의 양태로서 확인되었으며, 이러한 염기성 아미노산 클러스터가 없는 TAT 단백질이 세포 원형질막을 침투할 수 없는 실험이 수행되었다. 일부 경우에, 더 짧은 서열의 세포-침투 펩티드는 퓨린(furin)과 같은 엔도프로테아제 효소에 의한 분비 동안의 절단을 방지하도록 변형되었다. 이러한 변형은 단축된 세포-침투 TAT 아미노산 서열을 YGRKKRRQRRR에서 YARKAARQARA로 변화시키며, 이 짧은 펩티드는 TATκ로 지칭된다.
TAT가 원형질막을 가로질러 전위할 수 있는 정확한 메커니즘은 여전히 불확실하다. 최근의 연구는 특별한 유형의 세포내이입이 TAT 흡수와 관련될 가능성을 탐구하였고, TAT 침투에 내성인 것으로 보이는 몇몇 세포주가 확인되었다. TAT에 의해 전달될 특정 카고는 전달의 효율에서 또한 역할을 할 수 있다. 이전의 연구 데이터는 TAT 융합 단백질이 변성 조건에서 제조될 때 세포 흡수가 더 우수하다는 것을 시사하였는데, 이는 올바르게 폴딩된 단백질 카고가 구조적 제약으로 인해 원형질막을 횡단하는 데 훨씬 더 많은 에너지(델타-G)를 필요로 할 가능성이 있기 때문이다.
TAT 카고를 리폴딩하는 세포내 단백질 샤페론(chaperone)의 능력은 리폴딩될 단백질 카고의 아이덴티티(identity) 및 크기에 따라 달라질 가능성이 있다. 일부 경우에, TAT-융합 단백질은 수성 환경에 놓일 때 침전되므로, 변성되는 방식으로 제조될 수도 없고 본래의 입체 형태로 매우 오랫동안 안정적으로 유지될 수도 없다. TAT-융합 단백질의 설계는 또한 전달될 특정 카고에 맞추어져야 한다. 카고 단백질이 N-말단에서 밀접하게 관련되고 TAT 도메인이 N-말단에서 또한 발견되는 경우, TAT 전위 도메인은 카고 단백질에 묻힐 수 있고 전달이 불량할 수 있다.
다수의 TAT-카고 변이체는 1차 배양 세포, 형질 전환된 세포, 및 마우스 조직에 존재하는 세포를 비롯한 다양한 세포 유형 내로 성공적으로 전달되었다. 배양 시에, TAT-융합 단백질은 일반적으로 세포 내외로 쉽게 확산되어, 균일한 농도의 매우 빠른 확립을 가져온다.
효소, 항체, 다른 단백질, 또는 심지어 약물 로딩된 담체 입자와 같은 많은 약학 제제는 세포질, 핵, 또는 다른 특정 소기관 내에서 치료 작용을 발휘하기 위해 세포내로 전달될 필요가 있다. 따라서, 이들 상이한 유형의 큰 분자의 전달은 생물학적 제제의 개발에서 중요한 도전을 나타낸다. 현재 데이터는 TAT가 하나 초과의 메커니즘을 통해 원형질막을 횡단할 수 있음을 시사한다.
TAT 전달 도메인은 효소인 슈퍼옥사이드 디스뮤타제(SOD)에 또한 융합되었다(문헌[Torchilin, "Intracellular delivery of protein and peptide therapeutics." Protein Therapeutics. 2008. 5(2-3):e95-e103]). 이 융합 단백질은 그것이 세포내 환경으로 SOD 효소를 전달하기 위해 세포막을 가로질러 전위할 수 있음을 입증하기 위해 사용되었고, 그에 따라 여기서 융합 단백질은 숙주 세포에 대한 반응성 산소 종 및 산화 스트레스의 더 높은 축적을 가져오는 효소 결핍 장애를 치료하는 데 있어서 치료 가능성을 갖는다.
TAT 융합 단백질은 혈액 뇌 장벽을 가로질러 전달되는 것으로 또한 밝혀졌다. 신경보호 단백질인 Bcl-xL에 융합된 TAT 도메인은 배양 시에 세포에 빠르게 침투할 수 있었고, 뇌 허혈을 겪고 있는 마우스에 투여될 때, 융합 단백질은 1시간 내지 2시간 내에 뇌 세포에 전달되었다. 전달 후, 뇌경색은 용량-의존적 방식으로 크기가 감소되었다(문헌[Cao, G. et al., "In Vivo Delivery of a Bcl-xL Fusion Protein Containing the TAT Protein Transduction Domain Protects against Ischemic Brain Injury and Neuronal Apoptosis." J. Neurosci. 22, 5423, 2002]).
다양한 구현예에서, 본원에 기재된 CDKL5 변이체는 TAT, 변형된 TAT(TATκ), 트랜스포탄(Transportan), 안테나페디아 또는 P97과 같은 CPP에 작동적으로 연결된다. 본원에 사용되는 바와 같이, TAT는 11개 아미노산을 갖는 원래의 TAT 펩티드(TAT11로 표시됨)를 지칭할 수 있거나, 클로닝에 사용되는 플라스미드의 폴리링커로부터 유래된 추가의 16개 N-말단 아미노산을 갖는 TAT 펩티드(TAT28으로 표시됨)를 지칭할 수 있다. 유사하게, TATκ는 TAT11의 변형된 버전(TATκ11로 표시됨) 또는 TAT28의 변형된 버전(TATκ28로 표시됨)을 지칭할 수 있다. CPP인 TAT28, TATκ28, TAT11, TATκ11, 트랜스포탄, 안테나피디아 및 P97의 아미노산 서열은 각각 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 및 SEQ ID NO: 50에 제공된다.
일부 구현예에서, CPP는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 90%의 서열 동일성을 갖는다. 일부 구현예에서, CPP는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 95%의 서열 동일성을 갖는다. 일부 구현예에서, CPP는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 100%의 서열 동일성을 갖는다. 일부 구현예에서, CPP는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 적어도 90%의 서열 동일성을 갖는다. 일부 구현예에서, CPP는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 적어도 95%의 서열 동일성을 갖는다. 일부 구현예에서, CPP는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17 또는 SEQ ID NO: 18에 대해 100%의 서열 동일성을 갖는다. 다양한 구현예에서, CPP는 SEQ ID NO: 16의 서열을 갖지 않는다.
다양한 구현예에서, CPP에는 N-말단 글리신이 첨가될 수 있다. 예를 들어, TATκ28 및 TAT28은 낮은 안정성을 갖는 N-말단 아스파테이트 잔기를 달리 가질 것이다. 서열에 N-말단 글리신을 첨가하면 N-말단 규칙을 통해 단백질 안정성을 증가시킬 수 있다. 따라서, 일부 구현예에서, 리더 신호 폴리펩티드를 갖는 융합 단백질들 중 임의의 것에는 리더 신호 폴리펩티드의 C-말단부에 글리신이 첨가될 수 있어서, 리더 신호 폴리펩티드의 절단 시에 융합 단백질의 새로운 N-말단은 글리신으로 시작할 것이다. 유사한 방식으로, 리더 신호 폴리펩티드를 결여한 융합 단백질에는 N-말단 메티오닌과 융합 단백질의 나머지 사이에 글리신이 또한 첨가될 수 있다. 또한 유사한 방식으로, TAT28 또는 TATκ28 이외의 CPP를 갖는 융합 단백질에는 리더 신호 폴리펩티드와 CPP 사이에 글리신이 또한 첨가될 수 있다.
CDKL5 변이체를 포함하는 융합 단백질
전술한 바와 같이, CDKL5 변이체는 CPP를 또한 함유하는 단백질과 같은 융합 단백질에 사용될 수 있다. 단백질 분비를 향상시키기 위한 리더 신호 폴리펩티드 또는 융합 단백질을 검출하고/검출하거나 정제하기 위한 태그뿐만 아니라 기능성 폴리펩티드들을 연결하는 데 사용될 수 있는 링커 폴리펩티드와 같은 다른 폴리펩티드가 그러한 융합 단백질에 또한 통합될 수 있다.
리더 신호 폴리펩티드의 예에는 인간 면역 글로불린 중쇄 결합 단백질의 변형된 단편(변형된 BiP, 예를 들어, SEQ ID NO: 48, SEQ ID NO: 51, SEQ ID NO: 52 또는 SEQ ID NO: 53) 또는 뮤린 Igκ 사슬 리더 폴리펩티드(SEQ ID NO: 49, 예를 들어, ThermoFisher 벡터로부터의 pSecTag2)가 포함되지만 이로 제한되지 않는다. 변형된 BiP 신호 폴리펩티드의 예에는 전문이 본원에 참조로 포함된 미국 특허 제9,279,007호에 기재된 것들이 포함된다.
융합 단백질에 첨가될 수 있는 태그의 예에는 에피토프 태그(예를 들어, MYC, HA, V5, NE), 글루타티온 S-트랜스퍼라제(GST), 말토스-결합 단백질(MBP), 칼모둘린-결합 펩티드(CBP), FLAG®, 3xFLAG® 및 폴리히스티딘이 포함되지만 이로 제한되지 않는다.
제형, 치료 방법 및 용도
재조합 단백질(예를 들어, CDKL5 변이체 또는 융합 단백질)은 일상적인 절차에 따라 인간에게 투여하기에 적합한 약학 조성물로서 제형화될 수 있다. 예를 들어, 하나 이상의 구현예에서, 정맥내 투여용 조성물은 멸균 등장성 수성 완충제 중의 용액이다. 필요한 경우, 조성물은 주사 부위에서의 통증을 완화시키기 위해 가용화제 및 국소 마취제를 또한 포함할 수 있다. 일반적으로, 성분들은 단위 투여 형태로, 예를 들어, 활성 제제의 양을 나타내는 앰풀 또는 사세(sachet)와 같은 기밀 밀봉된 용기 중의 건식 동결 건조된 분말 또는 수분 비함유 농축물로서, 개별적으로 또는 함께 혼합되어 공급된다. 조성물이 주입에 의해 투여되는 경우, 이는 멸균 의약 등급의 물, 식염수 또는 덱스트로스/물을 함유하는 주입 병으로 분배될 수 있다. 조성물이 주사에 의해 투여되는 경우, 성분들이 투여 전에 혼합될 수 있도록 주사용 멸균수 또는 식염수의 앰풀이 제공될 수 있다.
재조합 단백질(예를 들어, CDKL5 변이체 또는 융합 단백질)(또는 재조합 단백질을 함유하는 조성물 또는 의약)은 적절한 경로로 투여된다. 하나 이상의 구현예에서, 재조합 단백질은 정맥내 투여된다. 다른 구현예에서, 재조합 단백질은 표적 조직, 예컨대 심장 또는 골격근(예를 들어, 근육내; 심실내), 또는 신경계(예를 들어, 뇌 내로의 직접 주사; 척수강내)로의 직접 투여에 의해 투여된다. 원하는 경우, 하나 초과의 경로가 동시에 사용될 수 있다.
재조합 단백질(예를 들어, CDKL5 변이체 또는 융합 단백질)(또는 재조합 단백질을 함유하는 조성물 또는 의약)은 치료적 유효량(예를 들어, 규칙적인 간격으로 투여될 때, 질병과 관련된 증상을 개선하고, 질병의 발병을 예방하거나 지연시키고/지연시키거나, 질병의 중증도 또는 질병 증상의 빈도를 감소시키는 것과 같이 질병을 치료하기에 충분한 투여량)으로 투여된다. 질병의 치료에 치료적으로 유효한 양은 질병의 영향의 본질 및 정도에 좌우될 것이다. 또한, 최적의 투여량 범위를 확인하는 데 도움을 주기 위해 시험관내 또는 생체내 검정이 선택적으로 사용될 수 있다. 사용되는 정확한 용량은 또한 투여 경로 및 질병의 심각성에 따라 달라질 것이며, 의사의 판단 및 각 환자의 상황에 따라 결정되어야 한다. 시험관내 또는 동물 모델 시험 시스템으로부터 유도된 용량-반응 곡선으로부터 유효량이 추정될 수 있다.
치료적 유효량의 재조합 단백질(예를 들어, CDKL5 변이체 또는 융합 단백질)(또는 재조합 단백질을 함유하는 조성물 또는 의약)은, 질병의 영향의 본질 및 정도에 따라 및/또는 지속적으로, 규칙적인 간격으로 투여될 수 있다. 본원에 사용되는 바와 같은 "규칙적인 간격"으로 투여는 치료적 유효량이 (일회 용량과 구별되는 바와 같이) 주기적으로 투여됨을 나타낸다. 단일 개체에 대한 투여 간격은 고정된 간격일 필요는 없으며, 개체의 필요에 따라 시간이 지나면서 변할 수 있다.
재조합 단백질(예를 들어, CDKL5 변이체 또는 융합 단백질)은 추후 사용을 위해, 예컨대 단위 용량 바이알 또는 주사기, 또는 정맥내 투여용 병 또는 백에 넣어져 제조될 수 있다. 재조합 단백질(예를 들어, CDKL5 변이체 또는 융합 단백질)뿐만 아니라 선택적인 부형제 또는 다른 약물과 같은 다른 활성 성분을 함유하는 키트는 포장재에 봉입될 수 있고, CDKL5 결핍, 레트 증후군, 또는 레트 증후군 변종을 갖는 환자와 같은 치료가 필요한 대상체를 치료하기 위한 재구성, 희석 또는 투여에 대한 설명서가 딸려 있을 수 있다.
제조 방법
재조합 단백질(예를 들어, CDKL5 변이체 또는 융합 단백질)은 적절한 벡터를 사용하여 숙주 세포에서 발현되고 그로부터 분비될 수 있다. 예를 들어, 포유 동물 세포(예를 들어, CHO, HeLa 또는 HEK 세포) 또는 세균 세포(예를 들어, 대장균 또는 슈도모나스 할로플랑크티스(P. haloplanktis) TAC 125 세포)가 사용될 수 있다. 예시적인 플라스미드는 하기 실시예에 기재되어 있고, 도 2a 내지 도 2ad에 도시되어 있다. 당업자는 본원에 기재된 CDKL5 변이체 및 융합 단백질을 생성하기 위해 세포의 형질전환, 형질감염, 또는 형질도입에 적합한 대안적인 벡터를 선택할 수 있다.
발현 및 분비 후, 재조합 단백질은 표준 기법을 사용하여 주위의 세포 배양 배지로부터 회수되고 정제될 수 있다. 대안적으로, 재조합 단백질은 배지보다는 세포로부터 직접 단리되고 정제될 수 있다.
실시예
실시예 1 - CDKL5 융합 단백질
도 2a 내지 도 2ad는 포유 동물 세포(예를 들어, CHO 세포) 또는 세균 세포(예를 들어, 대장균 세포)와 같은 적합한 세포에서 융합 단백질을 발현하기 위한 플라스미드를 도시한다. 이들 단백질은 SEQ ID NO: 19 내지 SEQ ID NO: 46에 제시된 아미노산 서열을 갖는다. 결실 또는 트렁케이션의 번호 매김은 전장 CDKL5107 폴리펩티드(1-960)에 대한 것이다. CDKL5가 추가의 N-말단 아미노산 서열에 C-말단적으로 융합된 작제물에서, CDKL5의 개시 메티오닌(아미노산 1)이 제거된다. 이들 작제물에서, CDKL5 폴리펩티드는 제2 아미노산인 리신으로 시작한다. 도 2a 내지 도 2ad 및 SEQ ID NO: 19 내지 SEQ ID NO: 46과 SEQ ID NO: 54 내지 SEQ ID NO: 55에 사용된 약어는 하기 표 1에 요약되어 있다:
[표 1]
도 2a는 CHO 세포에서 SEQ ID NO: 19의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 변형된 BiP 리더 신호 폴리펩티드, TATκ28 및 전장 인간 CDKL5107 아이소형을 포함한다.
도 2b는 CHO 세포에서 SEQ ID NO: 20의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 뮤린 Igκ 사슬 리더 폴리펩티드, TATκ28 및 전장 인간 CDKL5107 아이소형을 포함한다.
도 2c는 CHO 세포에서 SEQ ID NO: 21의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 변형된 BiP 리더 신호 폴리펩티드, TATκ28 및 전장 인간 CDKL5115 아이소형을 포함한다.
도 2d는 CHO 세포에서 SEQ ID NO: 22의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 뮤린 Igκ 사슬 리더 폴리펩티드, TATκ28 및 전장 인간 CDKL5115 아이소형을 포함한다.
도 2e는 CHO 세포에서 SEQ ID NO: 23의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 전장 인간 CDKL5107 아이소형을 포함한다.
도 2f는 대장균 세포에서 SEQ ID NO: 24의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 전장 인간 CDKL5107 아이소형을 포함한다.
도 2g는 대장균 세포에서 SEQ ID NO: 25의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 2의 CDKL5107 변이체를 포함한다.
도 2h는 대장균 세포에서 SEQ ID NO: 26의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 3의 CDKL5107 변이체를 포함한다.
도 2i는 대장균 세포에서 SEQ ID NO: 27의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 4의 CDKL5107 변이체를 포함한다.
도 2j는 대장균 세포에서 SEQ ID NO: 28의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 5의 CDKL5107 변이체를 포함한다.
도 2k는 대장균 세포에서 SEQ ID NO: 29의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 6의 CDKL5107 변이체를 포함한다.
도 2l은 대장균 세포에서 SEQ ID NO: 30의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 7의 CDKL5107 변이체를 포함한다.
도 2m은 대장균 세포에서 SEQ ID NO: 31의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 8의 CDKL5107 변이체를 포함한다.
도 2n은 대장균 세포에서 SEQ ID NO: 32의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 9의 CDKL5107 변이체를 포함한다.
도 2o는 대장균 세포에서 SEQ ID NO: 33의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 10의 CDKL5107 변이체를 포함한다.
도 2p는 대장균 세포에서 SEQ ID NO: 34의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 11의 CDKL5107 변이체를 포함한다.
도 2q는 대장균 세포에서 SEQ ID NO: 35의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 작제물 12의 CDKL5107 변이체를 포함한다.
도 2r은 대장균 세포에서 SEQ ID NO: 36의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TAT28 및 전장 인간 CDKL5107 아이소형을 포함한다.
도 2s는 대장균 세포에서 SEQ ID NO: 37의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ28 및 강화 녹색 형광 단백질(eGFP)을 포함한다.
도 2t는 대장균 세포에서 SEQ ID NO: 38의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 CPP가 없는 eGFP를 포함한다.
도 2u는 대장균 세포에서 SEQ ID NO: 39의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 인간 암피피신 1(AMPH1)을 포함한다.
도 2v는 CHO 세포에서 SEQ ID NO: 40의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 인간 암피피신 1(AMPH1)을 포함한다.
도 2w는 CHO 세포에서 SEQ ID NO: 41의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 변형된 BiP 리더 신호 폴리펩티드, TATκ11 및 전장 인간 CDKL5107 아이소형을 포함한다.
도 2x는 CHO 세포에서 SEQ ID NO: 42의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 뮤린 Igκ 사슬 리더 폴리펩티드, TATκ11 및 전장 인간 CDKL5107 아이소형을 포함한다.
도 2y는 CHO 세포에서 SEQ ID NO: 43의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ11, 및 리더 신호 폴리펩티드가 없는 전장 인간 CDKL5107 아이소형을 포함한다.
도 2z는 대장균 세포에서 SEQ ID NO: 44의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TATκ11, 및 리더 신호 폴리펩티드가 없는 전장 인간 CDKL5107 아이소형을 포함한다.
도 2aa는 대장균 세포에서 SEQ ID NO: 45의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TAT11, 및 리더 신호 폴리펩티드가 없는 전장 인간 CDKL5107 아이소형을 포함한다.
도 2ab는 CHO 세포에서 SEQ ID NO: 46의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 TAT11, 및 리더 신호 폴리펩티드가 없는 전장 인간 CDKL5107 아이소형을 포함한다.
도 2ac는 CHO 세포에서 SEQ ID NO: 54의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 안테나페디아 CPP, 및 리더 신호 폴리펩티드가 없는 전장 인간 CDKL5107 아이소형을 포함한다.
도 2ad는 CHO 세포에서 SEQ ID NO: 55의 융합 단백질을 발현하기 위한 예시적인 플라스미드를 도시한다. 이 융합 단백질은 트랜스포탄 CPP, 및 리더 신호 폴리펩티드가 없는 전장 인간 CDKL5107 아이소형을 포함한다.
SEQ ID NO: 19 내지 SEQ ID NO: 36 및 SEQ ID NO: 41 내지 SEQ ID NO: 46의 CDKL5 융합 단백질은 각각 도 2a 내지 도 2r 및 도 2w 내지 도 2ab의 플라스미드를 사용하여 발현되고 활성이 평가될 것이다. 인간 암피피신 1(AMPH1)은 CDKL5 키나제 검정에서 기질이 될 것이다. 도 2u 및 도 2v의 플라스미드는 CDKL5 키나제 검정을 위해 친화성-태깅된 AMPH1(SEQ ID NO: 39 및 SEQ ID NO: 40)을 발현시키는 데 사용될 것이다. 친화성-태깅된 eGFP 단독(SEQ ID NO: 38)뿐만 아니라 친화성-태깅된 TATk28-eGFP(SEQ ID NO: 37)는 CDKL5 융합 단백질에 대한 대조군으로서 작용할 것이며, 이들은 각각 도 2s 및 도 2t의 플라스미드를 사용하여 발현될 것이다.
다양한 CDKL5 융합 단백질을 CHO 및 HEK 세포에서뿐만 아니라 HeLa 세포 용해물과 함께 시험관내 전사/번역을 사용하여 발현시켰다. 간략하게 말하면, CHO-S 세포(20x106개 세포)를 Maxcyte STX를 사용하여 8개의 플라스미드로 전기 천공하였다: (1) pOptiVec 빈(empty) 벡터; 2) TATk28-CDKL5-107-3xFlagHis; 3) TATk11-CDKL5-107-3xFlagHis; 4) TAT11-CDKL5-107-3xFlagHis; 5) TAT28-CDKL5-107-3xFlagHis; 6) ANTP-CDKL5-107-3xFlagHis; 7) TRANSP-CDKL5-107-3xFlagHis 및 8) MBiP-TATK28-CDKL5-107-3xFlagHis(코딩 서열은 CHO 코돈-최적화됨). 세포를 배양 배지에서 회수하고, 하루 동안 배양하였다. 세포를 수거하고 용해시켰다. 각각의 형질감염을 위해, 20 ㎍의 용해물을 4% 내지 12% BisTris SDS-PAGE에 적용하고, iBlot2 시스템을 사용하여 니트로셀룰로오스 블롯으로 옮겼다. 블롯을 1xTBS-T 중의 5% 우유로 차단시켰다. 토끼 항-His 항체의 1:2000 희석물과 함께 밤새 인큐베이션함으로써 블롯을 웨스턴 블롯에 적용하였다. 일련의 세척 후, 블롯을 1:10000 항-토끼 IgG DyaLight 680 2차 항체와 함께 인큐베이션하였다. 추가 세척을 수행하였다. 블롯을 Licor Odyssey 스캐너에서 이미지화하였다. 블롯은 CDKL5 융합 단백질의 발현을 확인해주었다.
HEK293F 세포(8x106개 세포)를 FuGeneHD(24 μl의 FuGeneHD : 8 μg의 DNA 비) 및 7개의 플라스미드로 형질감염시켰다: 1) 빈 pOptiVec; 2) TATk11-CDKL5_107-3xFlagHis; 3) TAT11-CDKL5_1-FH; 4) TAT28-CDKL5_1-FH; 5) ANTP-CDKL5_107-3xFlagHis; 6) TRANSP-CDKL5_107-3xFlagHis 및 7) TATk28-CDKL5_107-3xFlagHis(코딩 서열은 인간 코돈-최적화됨). 세포를 인큐베이션하고, 형질감염 후 2일차에 수거하였다. 세포를 용해시키고, 20 ㎍의 용해물을 4% 내지 12% BisTris SDS-PAGE에 적용하고, iBlot2 시스템을 사용하여 니트로셀룰로스 블롯으로 옮겼다. 블롯을 1xTBS-T 중의 5% 우유로 차단시켰다. 토끼 항-His 항체의 1:2000 희석물과 함께 밤새 인큐베이션함으로써 블롯을 웨스턴 블롯에 적용하였다. 일련의 세척 후, 블롯을 1:10000 항-토끼 IgG DyaLight 680 2차 항체와 함께 인큐베이션하였다. 추가 세척을 수행하였다. 블롯을 Licor Odyssey 스캐너에서 이미지화하였다. 블롯은 CDKL5 융합 단백질의 발현을 확인해주었다.
본 명세서 전체에 걸쳐 "일 구현예", "특정 구현예", "다양한 구현예", "하나 이상의 구현예" 또는 "구현예"에 대한 언급은 구현예와 관련하여 기재된 특정한 특징, 구조, 재료, 또는 특성이 본 개시의 적어도 하나의 구현예에 포함됨을 의미한다. 따라서, 본 명세서 전체에 걸쳐 여러 곳에서 "하나 이상의 구현예에서", "특정 구현예에서", "다양한 구현예에서", "일 구현예에서" 또는 "구현예에서"와 같은 문구의 출현은 반드시 본 개시의 동일한 구현예를 지칭하는 것은 아니다. 또한, 특정한 특징, 구조, 재료, 또는 특성은 하나 이상의 구현예에서 임의의 적합한 방식으로 조합될 수 있다.
본원의 개시는 특정한 구현예를 참조로 설명을 제공했지만, 이들 구현예는 단지 본 개시의 원리 및 응용을 예시하는 것으로 이해해야 한다. 본 개시의 사상 및 범주를 벗어나지 않고 본 개시에 대해 다양한 수정 및 변형이 이루어질 수 있음이 당업자에게 명백할 것이다. 따라서, 본 개시는 첨부된 청구범위 및 그 균등물의 범주 내에 있는 수정 및 변형을 포함하는 것으로 의도된다.
<110> Amicus Therapeutics, Inc.
<120> CDKL5 EXPRESSION VARIANTS AND CDKL5 FUSION PROTEINS
<130> AT17-013
<160> 55
<170> PatentIn version 3.5
<210> 1
<211> 960
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 isoform polypeptide 1-960 (full-length)
<400> 1
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser
305 310 315 320
Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln
325 330 335
Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly
340 345 350
Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn
355 360 365
Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr
370 375 380
Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn
385 390 395 400
Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu
405 410 415
Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys
420 425 430
Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met
435 440 445
Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln
450 455 460
Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro
465 470 475 480
Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys
485 490 495
Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser
500 505 510
Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr
515 520 525
Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro
530 535 540
Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr
545 550 555 560
Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His
565 570 575
Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser Phe
580 585 590
Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro
595 600 605
His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly
610 615 620
Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn
625 630 635 640
Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu
645 650 655
Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr
660 665 670
Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp
675 680 685
Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr
690 695 700
Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser
705 710 715 720
Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val
725 730 735
Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg
740 745 750
Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser
755 760 765
Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys
770 775 780
Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp
785 790 795 800
Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser
805 810 815
Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln
820 825 830
Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala
835 840 845
Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala
850 855 860
Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser
865 870 875 880
Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys
885 890 895
Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp His
900 905 910
Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser Glu
915 920 925
Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg Thr
930 935 940
Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala Leu
945 950 955 960
<210> 2
<211> 852
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?853-960
<400> 2
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser
305 310 315 320
Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln
325 330 335
Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly
340 345 350
Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn
355 360 365
Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr
370 375 380
Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn
385 390 395 400
Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu
405 410 415
Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys
420 425 430
Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met
435 440 445
Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln
450 455 460
Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro
465 470 475 480
Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys
485 490 495
Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser
500 505 510
Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr
515 520 525
Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro
530 535 540
Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr
545 550 555 560
Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His
565 570 575
Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser Phe
580 585 590
Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro
595 600 605
His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly
610 615 620
Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn
625 630 635 640
Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu
645 650 655
Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr
660 665 670
Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp
675 680 685
Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr
690 695 700
Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser
705 710 715 720
Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val
725 730 735
Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg
740 745 750
Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser
755 760 765
Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys
770 775 780
Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp
785 790 795 800
Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser
805 810 815
Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln
820 825 830
Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala
835 840 845
Ser Asn His Pro
850
<210> 3
<211> 744
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?745-960
<400> 3
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser
305 310 315 320
Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln
325 330 335
Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly
340 345 350
Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn
355 360 365
Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr
370 375 380
Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn
385 390 395 400
Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu
405 410 415
Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys
420 425 430
Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met
435 440 445
Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln
450 455 460
Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro
465 470 475 480
Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys
485 490 495
Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser
500 505 510
Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr
515 520 525
Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro
530 535 540
Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr
545 550 555 560
Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His
565 570 575
Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser Phe
580 585 590
Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro
595 600 605
His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly
610 615 620
Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn
625 630 635 640
Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu
645 650 655
Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr
660 665 670
Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp
675 680 685
Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr
690 695 700
Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser
705 710 715 720
Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val
725 730 735
Ser Ser Leu Pro Ser Glu Ser Ser
740
<210> 4
<211> 636
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?637-960
<400> 4
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser
305 310 315 320
Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln
325 330 335
Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly
340 345 350
Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn
355 360 365
Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr
370 375 380
Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn
385 390 395 400
Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu
405 410 415
Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys
420 425 430
Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met
435 440 445
Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln
450 455 460
Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro
465 470 475 480
Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys
485 490 495
Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser
500 505 510
Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr
515 520 525
Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro
530 535 540
Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr
545 550 555 560
Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His
565 570 575
Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser Phe
580 585 590
Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro
595 600 605
His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly
610 615 620
Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala
625 630 635
<210> 5
<211> 528
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?529-960
<400> 5
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser
305 310 315 320
Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln
325 330 335
Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly
340 345 350
Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn
355 360 365
Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr
370 375 380
Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn
385 390 395 400
Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu
405 410 415
Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys
420 425 430
Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met
435 440 445
Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln
450 455 460
Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro
465 470 475 480
Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys
485 490 495
Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser
500 505 510
Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr
515 520 525
<210> 6
<211> 420
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?421-960
<400> 6
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser
305 310 315 320
Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln
325 330 335
Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly
340 345 350
Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn
355 360 365
Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr
370 375 380
Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn
385 390 395 400
Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu
405 410 415
Phe Asp Phe Asn
420
<210> 7
<211> 314
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?315-960
<400> 7
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys
305 310
<210> 8
<211> 854
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?315-420
<400> 8
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ile Asp Pro Lys Pro Ser
305 310 315 320
Glu Gly Pro Gly Thr Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln
325 330 335
Asn Arg His Ser Phe Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu
340 345 350
Gln Pro Asn Glu Lys Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro
355 360 365
Gln Ser Ser Arg Ser Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly
370 375 380
Ala Leu Ser Asp Ser Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala
385 390 395 400
Gln Ile Ala Glu Pro Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu
405 410 415
Asp Leu Asn Ser Pro Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr
420 425 430
Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr
435 440 445
Leu Asp Ser Arg Arg Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu
450 455 460
Leu Lys Leu Pro Glu His Met Asp Ser Ser His Ser His Ser Leu Ser
465 470 475 480
Ala Pro His Glu Ser Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe
485 490 495
Ser Ser Gln Gln Arg Pro His Arg His Ser Met Tyr Val Thr Arg Asp
500 505 510
Lys Val Arg Ala Lys Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly
515 520 525
Met Ala Ala Arg Ala Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly
530 535 540
Glu Gln Leu Pro Pro Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu
545 550 555 560
Thr Ser Arg Glu Gly Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu
565 570 575
Gly Gly Val Tyr His Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys
580 585 590
Glu Asn Arg His Leu Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser
595 600 605
Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn
610 615 620
Asn Val Ser Thr Arg Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly
625 630 635 640
Thr Asn His Ser Lys Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro
645 650 655
Glu Asn Ile Ser His Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly
660 665 670
Phe Phe Arg Ser Met Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro
675 680 685
Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser
690 695 700
Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile
705 710 715 720
Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu
725 730 735
Leu His Leu Ser Ser Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg
740 745 750
Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile
755 760 765
Arg Ile His Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg
770 775 780
Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln
785 790 795 800
Met Asp Pro Gly Trp His Val Ser Ser Val Thr Arg Ser Ala Thr Glu
805 810 815
Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly
820 825 830
His Pro Tyr Asn Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp
835 840 845
Leu Lys Glu Thr Ala Leu
850
<210> 9
<211> 746
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?315-528
<400> 9
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ser Pro Thr Pro Thr Arg
305 310 315 320
His Ser Asp Thr Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn Asn Arg
325 330 335
Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr Thr Thr Arg His Ser Lys
340 345 350
Thr Met Glu Glu Leu Lys Leu Pro Glu His Met Asp Ser Ser His Ser
355 360 365
His Ser Leu Ser Ala Pro His Glu Ser Phe Ser Tyr Gly Leu Gly Tyr
370 375 380
Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro His Arg His Ser Met Tyr
385 390 395 400
Val Thr Arg Asp Lys Val Arg Ala Lys Gly Leu Asp Gly Ser Leu Ser
405 410 415
Ile Gly Gln Gly Met Ala Ala Arg Ala Asn Ser Leu Gln Leu Leu Ser
420 425 430
Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu Met Thr Val Ala Arg Ser
435 440 445
Ser Val Lys Glu Thr Ser Arg Glu Gly Thr Ser Ser Phe His Thr Arg
450 455 460
Gln Lys Ser Glu Gly Gly Val Tyr His Asp Pro His Ser Asp Asp Gly
465 470 475 480
Thr Ala Pro Lys Glu Asn Arg His Leu Tyr Asn Asp Pro Val Pro Arg
485 490 495
Arg Val Gly Ser Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp Asn Ser
500 505 510
Phe His Glu Asn Asn Val Ser Thr Arg Val Ser Ser Leu Pro Ser Glu
515 520 525
Ser Ser Ser Gly Thr Asn His Ser Lys Arg Gln Pro Ala Phe Asp Pro
530 535 540
Trp Lys Ser Pro Glu Asn Ile Ser His Ser Glu Gln Leu Lys Glu Lys
545 550 555 560
Glu Lys Gln Gly Phe Phe Arg Ser Met Lys Lys Lys Lys Lys Lys Ser
565 570 575
Gln Thr Val Pro Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu Gln Lys
580 585 590
Ser Ile His Ser Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu Trp Arg
595 600 605
Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu Lys Ser
610 615 620
Leu Arg Lys Leu Leu His Leu Ser Ser Ala Ser Asn His Pro Ala Ser
625 630 635 640
Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser
645 650 655
Phe Ser Glu Ile Arg Ile His Pro Leu Ser Gln Ala Ser Gly Gly Ser
660 665 670
Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln
675 680 685
Leu Pro Gly Gln Met Asp Pro Gly Trp His Val Ser Ser Val Thr Arg
690 695 700
Ser Ala Thr Glu Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala Lys Ser
705 710 715 720
Gly Pro Asn Gly His Pro Tyr Asn Arg Thr Asn Arg Ser Arg Met Pro
725 730 735
Asn Leu Asn Asp Leu Lys Glu Thr Ala Leu
740 745
<210> 10
<211> 638
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?315-636
<400> 10
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ala Arg Ala Asn Ser Leu
305 310 315 320
Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu Met Thr
325 330 335
Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr Ser Ser
340 345 350
Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp Pro His
355 360 365
Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr Asn Asp
370 375 380
Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser Pro Arg
385 390 395 400
Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val Ser Ser
405 410 415
Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg Gln Pro
420 425 430
Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser Glu Gln
435 440 445
Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys Lys Lys
450 455 460
Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp Leu Leu
465 470 475 480
Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser Arg Pro
485 490 495
Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln Ser Gln
500 505 510
Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala Ser Asn
515 520 525
His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala Gln Gln
530 535 540
Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser Gln Ala
545 550 555 560
Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys Gly Arg
565 570 575
Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp His Val Ser
580 585 590
Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser Glu Gln Leu
595 600 605
Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg Thr Asn Arg
610 615 620
Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala Leu
625 630 635
<210> 11
<211> 530
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?315-744
<400> 11
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ser Gly Thr Asn His Ser
305 310 315 320
Lys Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser
325 330 335
His Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser
340 345 350
Met Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser
355 360 365
Pro Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro
370 375 380
Ser Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln
385 390 395 400
Thr Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser
405 410 415
Ser Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu
420 425 430
Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro
435 440 445
Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala
450 455 460
Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly
465 470 475 480
Trp His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr
485 490 495
Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn
500 505 510
Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr
515 520 525
Ala Leu
530
<210> 12
<211> 422
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5107 Variant ?315-852
<400> 12
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ala Ser Ser Asp Pro Arg
305 310 315 320
Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile
325 330 335
Arg Ile His Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg
340 345 350
Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln
355 360 365
Met Asp Pro Gly Trp His Val Ser Ser Val Thr Arg Ser Ala Thr Glu
370 375 380
Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly
385 390 395 400
His Pro Tyr Asn Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp
405 410 415
Leu Lys Glu Thr Ala Leu
420
<210> 13
<211> 27
<212> PRT
<213> Artificial Sequence
<220>
<223> TAT28 CPP
<400> 13
Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu Ala Ala
1 5 10 15
Tyr Gly Arg Lys Lys Arg Arg Gln Arg Arg Arg
20 25
<210> 14
<211> 27
<212> PRT
<213> Artificial Sequence
<220>
<223> TAT?28 CPP
<400> 14
Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu Ala Ala
1 5 10 15
Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala
20 25
<210> 15
<211> 11
<212> PRT
<213> Artificial Sequence
<220>
<223> TAT11 CPP
<400> 15
Tyr Gly Arg Lys Lys Arg Arg Gln Arg Arg Arg
1 5 10
<210> 16
<211> 11
<212> PRT
<213> Artificial Sequence
<220>
<223> TAT?11 CPP
<400> 16
Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala
1 5 10
<210> 17
<211> 21
<212> PRT
<213> Artificial Sequence
<220>
<223> Transportan CPP
<400> 17
Ala Gly Tyr Leu Leu Gly Lys Ile Asn Leu Lys Ala Leu Ala Ala Leu
1 5 10 15
Ala Lys Lys Ile Leu
20
<210> 18
<211> 16
<212> PRT
<213> Artificial Sequence
<220>
<223> Antennapedia CPP
<400> 18
Arg Gln Ile Lys Ile Trp Phe Gln Asn Arg Arg Met Lys Trp Lys Lys
1 5 10 15
<210> 19
<211> 1064
<212> PRT
<213> Artificial Sequence
<220>
<223> >MBip_Tk28p_107_3xFlagHis_cho-opt in pOptiVec
<400> 19
Met Lys Leu Ser Leu Val Ala Ala Met Leu Leu Leu Leu Ser Leu Val
1 5 10 15
Ala Ala Met Leu Leu Leu Leu Ser Ala Ala Arg Ala Gly Asp Ala Ala
20 25 30
Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu Ala Ala Tyr Ala Arg
35 40 45
Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly Gly Ser Lys Ile Pro
50 55 60
Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu Gly Val Val Gly
65 70 75 80
Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His Lys Glu Thr His
85 90 95
Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu Glu Asn Glu Glu
100 105 110
Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu Arg Thr Leu Lys
115 120 125
Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg Arg Arg Gly Lys
130 135 140
Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met Leu Glu Leu Leu
145 150 155 160
Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val Lys Ser Tyr Ile
165 170 175
Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys Asn Asp Ile Val
180 185 190
His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser His Asn Asp Val
195 200 205
Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu Ser Glu Gly Asn
210 215 220
Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp Tyr Arg Ser Pro
225 230 235 240
Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val Asp Met Trp Ser
245 250 255
Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln Pro Leu Phe Pro
260 265 270
Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln Lys Val Leu Gly
275 280 285
Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser Asn Pro Arg Phe
290 295 300
His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln Ser Leu Glu Arg
305 310 315 320
Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp Leu Met Lys Asn
325 330 335
Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr Glu Gln Cys Leu
340 345 350
Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp Arg Ser Pro Ser
355 360 365
Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser Ser Thr Leu Ser
370 375 380
Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln Ser His His Arg
385 390 395 400
Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly Leu Pro Arg Ala
405 410 415
Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn Gly Asn Leu Ala
420 425 430
Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr Gln Ala Ser Ser
435 440 445
Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn Asn Ile Pro His
450 455 460
Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu Phe Asp Phe Asn
465 470 475 480
Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys Tyr Leu Lys Ser
485 490 495
Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met Glu Ser Ser Gln
500 505 510
Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln Ser Arg His Ser
515 520 525
Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro Ser Tyr Arg Thr
530 535 540
Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys Ser Val Ser Asn
545 550 555 560
Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser Thr Ser Arg Tyr
565 570 575
Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr Ser Pro Thr Pro
580 585 590
Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn
595 600 605
Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr Thr Thr Arg His
610 615 620
Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His Met Asp Ser Ser
625 630 635 640
His Ser His Ser Leu Ser Ala Pro His Glu Ser Phe Ser Tyr Gly Leu
645 650 655
Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro His Arg His Ser
660 665 670
Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly Leu Asp Gly Ser
675 680 685
Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn Ser Leu Gln Leu
690 695 700
Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu Met Thr Val Ala
705 710 715 720
Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr Ser Ser Phe His
725 730 735
Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp Pro His Ser Asp
740 745 750
Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr Asn Asp Pro Val
755 760 765
Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp
770 775 780
Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val Ser Ser Leu Pro
785 790 795 800
Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg Gln Pro Ala Phe
805 810 815
Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser Glu Gln Leu Lys
820 825 830
Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys Lys Lys Lys Lys
835 840 845
Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu
850 855 860
Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu
865 870 875 880
Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu
885 890 895
Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala Ser Asn His Pro
900 905 910
Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys
915 920 925
Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser Gln Ala Ser Gly
930 935 940
Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala
945 950 955 960
Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp His Val Ser Ser Val
965 970 975
Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala
980 985 990
Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg Thr Asn Arg Ser Arg
995 1000 1005
Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala Leu Gly Gly Gly Gly
1010 1015 1020
Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp
1025 1030 1035 1040
Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly
1045 1050 1055
Ala Pro His His His His His His
1060
<210> 20
<211> 1056
<212> PRT
<213> Artificial Sequence
<220>
<223> >IgK_Tk28p_107_3xFlagHis_cho-opt in pOptiVec
<400> 20
Met Glu Thr Asp Thr Leu Leu Leu Trp Val Leu Leu Leu Trp Val Pro
1 5 10 15
Gly Ser Thr Gly Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg
20 25 30
Thr Lys Leu Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala
35 40 45
Gly Gly Gly Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys
50 55 60
Phe Glu Ile Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu
65 70 75 80
Lys Cys Arg His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe
85 90 95
Lys Asp Ser Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu
100 105 110
Leu Lys Met Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys
115 120 125
Glu Ala Phe Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val
130 135 140
Glu Lys Asn Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro
145 150 155 160
Pro Glu Lys Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His
165 170 175
Trp Cys His Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn
180 185 190
Leu Leu Ile Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe
195 200 205
Ala Arg Asn Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val
210 215 220
Ala Thr Arg Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr
225 230 235 240
Gly Lys Ser Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu
245 250 255
Ser Asp Gly Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu
260 265 270
Phe Thr Ile Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys
275 280 285
Leu Phe Tyr Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val
290 295 300
Asn His Pro Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser
305 310 315 320
Val Leu Leu Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp
325 330 335
Arg Tyr Leu Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln
340 345 350
Arg Leu Leu Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr
355 360 365
His Val Glu Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser
370 375 380
Thr Ala Leu Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn
385 390 395 400
Leu Ser Val Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu
405 410 415
Ser Phe Leu Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His
420 425 430
Thr Lys Thr Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp
435 440 445
Leu Thr Asn Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys
450 455 460
Ser Lys Thr Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly
465 470 475 480
Pro Gly Thr Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg
485 490 495
His Ser Phe Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro
500 505 510
Asn Glu Lys Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser
515 520 525
Ser Arg Ser Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu
530 535 540
Ser Asp Ser Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile
545 550 555 560
Ala Glu Pro Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu
565 570 575
Asn Ser Pro Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr
580 585 590
Leu Leu Ser Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp
595 600 605
Ser Arg Arg Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys
610 615 620
Leu Pro Glu His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro
625 630 635 640
His Glu Ser Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser
645 650 655
Gln Gln Arg Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val
660 665 670
Arg Ala Lys Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala
675 680 685
Ala Arg Ala Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln
690 695 700
Leu Pro Pro Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser
705 710 715 720
Arg Glu Gly Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly
725 730 735
Val Tyr His Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn
740 745 750
Arg His Leu Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr
755 760 765
Arg Val Pro Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val
770 775 780
Ser Thr Arg Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn
785 790 795 800
His Ser Lys Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn
805 810 815
Ile Ser His Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe
820 825 830
Arg Ser Met Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser
835 840 845
Asp Ser Pro Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser
850 855 860
Thr Pro Ser Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp
865 870 875 880
Leu Gln Thr Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His
885 890 895
Leu Ser Ser Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln
900 905 910
Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile
915 920 925
His Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu
930 935 940
Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp
945 950 955 960
Pro Gly Trp His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro
965 970 975
Ser Tyr Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro
980 985 990
Tyr Asn Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys
995 1000 1005
Glu Thr Ala Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly
1010 1015 1020
Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr
1025 1030 1035 1040
Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His His His His
1045 1050 1055
<210> 21
<211> 1134
<212> PRT
<213> Artificial Sequence
<220>
<223> >MBiP_Tk28p_115_3xFlagHis_cho-opt in pOptiVec
<400> 21
Met Lys Leu Ser Leu Val Ala Ala Met Leu Leu Leu Leu Ser Leu Val
1 5 10 15
Ala Ala Met Leu Leu Leu Leu Ser Ala Ala Arg Ala Gly Asp Ala Ala
20 25 30
Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu Ala Ala Tyr Ala Arg
35 40 45
Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly Gly Ser Lys Ile Pro
50 55 60
Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu Gly Val Val Gly
65 70 75 80
Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His Lys Glu Thr His
85 90 95
Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu Glu Asn Glu Glu
100 105 110
Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu Arg Thr Leu Lys
115 120 125
Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg Arg Arg Gly Lys
130 135 140
Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met Leu Glu Leu Leu
145 150 155 160
Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val Lys Ser Tyr Ile
165 170 175
Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys Asn Asp Ile Val
180 185 190
His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser His Asn Asp Val
195 200 205
Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu Ser Glu Gly Asn
210 215 220
Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp Tyr Arg Ser Pro
225 230 235 240
Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val Asp Met Trp Ser
245 250 255
Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln Pro Leu Phe Pro
260 265 270
Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln Lys Val Leu Gly
275 280 285
Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser Asn Pro Arg Phe
290 295 300
His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln Ser Leu Glu Arg
305 310 315 320
Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp Leu Met Lys Asn
325 330 335
Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr Glu Gln Cys Leu
340 345 350
Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp Arg Ser Pro Ser
355 360 365
Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser Ser Thr Leu Ser
370 375 380
Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln Ser His His Arg
385 390 395 400
Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly Leu Pro Arg Ala
405 410 415
Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn Gly Asn Leu Ala
420 425 430
Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr Gln Ala Ser Ser
435 440 445
Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn Asn Ile Pro His
450 455 460
Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu Phe Asp Phe Asn
465 470 475 480
Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys Tyr Leu Lys Ser
485 490 495
Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met Glu Ser Ser Gln
500 505 510
Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln Ser Arg His Ser
515 520 525
Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro Ser Tyr Arg Thr
530 535 540
Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys Ser Val Ser Asn
545 550 555 560
Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser Thr Ser Arg Tyr
565 570 575
Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr Ser Pro Thr Pro
580 585 590
Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn
595 600 605
Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr Thr Thr Arg His
610 615 620
Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His Met Asp Ser Ser
625 630 635 640
His Ser His Ser Leu Ser Ala Pro His Glu Ser Phe Ser Tyr Gly Leu
645 650 655
Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro His Arg His Ser
660 665 670
Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly Leu Asp Gly Ser
675 680 685
Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn Ser Leu Gln Leu
690 695 700
Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu Met Thr Val Ala
705 710 715 720
Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr Ser Ser Phe His
725 730 735
Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp Pro His Ser Asp
740 745 750
Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr Asn Asp Pro Val
755 760 765
Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp
770 775 780
Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val Ser Ser Leu Pro
785 790 795 800
Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg Gln Pro Ala Phe
805 810 815
Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser Glu Gln Leu Lys
820 825 830
Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys Lys Lys Lys Lys
835 840 845
Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu
850 855 860
Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu
865 870 875 880
Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu
885 890 895
Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala Ser Asn His Pro
900 905 910
Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys
915 920 925
Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser Gln Ala Ser Gly
930 935 940
Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala
945 950 955 960
Leu Gln Leu Pro Asp Gly Gly Cys Asp Gly Arg Arg Gln Arg His His
965 970 975
Ser Gly Pro Gln Asp Arg Arg Phe Met Leu Arg Thr Thr Glu Gln Gln
980 985 990
Gly Glu Tyr Phe Cys Cys Gly Asp Pro Lys Lys Pro His Thr Pro Cys
995 1000 1005
Val Pro Asn Arg Ala Leu His Arg Pro Ile Ser Ser Pro Ala Pro Tyr
1010 1015 1020
Pro Val Leu Gln Val Arg Gly Thr Ser Met Cys Pro Thr Leu Gln Val
1025 1030 1035 1040
Arg Gly Thr Asp Ala Phe Ser Cys Pro Thr Gln Gln Ser Gly Phe Ser
1045 1050 1055
Phe Phe Val Arg His Val Met Arg Glu Ala Leu Ile His Arg Ala Gln
1060 1065 1070
Val Asn Gln Ala Ala Leu Leu Thr Tyr His Glu Asn Ala Ala Leu Thr
1075 1080 1085
Gly Lys Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr
1090 1095 1100
Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp
1105 1110 1115 1120
Asp Asp Asp Lys Asp Gly Ala Pro His His His His His His
1125 1130
<210> 22
<211> 1126
<212> PRT
<213> Artificial Sequence
<220>
<223> >IgK_Tk28p_115_3xFlagHis_cho-opt in pOptiVec
<400> 22
Met Glu Thr Asp Thr Leu Leu Leu Trp Val Leu Leu Leu Trp Val Pro
1 5 10 15
Gly Ser Thr Gly Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg
20 25 30
Thr Lys Leu Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala
35 40 45
Gly Gly Gly Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys
50 55 60
Phe Glu Ile Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu
65 70 75 80
Lys Cys Arg His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe
85 90 95
Lys Asp Ser Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu
100 105 110
Leu Lys Met Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys
115 120 125
Glu Ala Phe Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val
130 135 140
Glu Lys Asn Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro
145 150 155 160
Pro Glu Lys Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His
165 170 175
Trp Cys His Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn
180 185 190
Leu Leu Ile Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe
195 200 205
Ala Arg Asn Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val
210 215 220
Ala Thr Arg Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr
225 230 235 240
Gly Lys Ser Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu
245 250 255
Ser Asp Gly Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu
260 265 270
Phe Thr Ile Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys
275 280 285
Leu Phe Tyr Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val
290 295 300
Asn His Pro Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser
305 310 315 320
Val Leu Leu Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp
325 330 335
Arg Tyr Leu Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln
340 345 350
Arg Leu Leu Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr
355 360 365
His Val Glu Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser
370 375 380
Thr Ala Leu Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn
385 390 395 400
Leu Ser Val Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu
405 410 415
Ser Phe Leu Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His
420 425 430
Thr Lys Thr Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp
435 440 445
Leu Thr Asn Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys
450 455 460
Ser Lys Thr Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly
465 470 475 480
Pro Gly Thr Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg
485 490 495
His Ser Phe Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro
500 505 510
Asn Glu Lys Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser
515 520 525
Ser Arg Ser Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu
530 535 540
Ser Asp Ser Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile
545 550 555 560
Ala Glu Pro Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu
565 570 575
Asn Ser Pro Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr
580 585 590
Leu Leu Ser Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp
595 600 605
Ser Arg Arg Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys
610 615 620
Leu Pro Glu His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro
625 630 635 640
His Glu Ser Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser
645 650 655
Gln Gln Arg Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val
660 665 670
Arg Ala Lys Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala
675 680 685
Ala Arg Ala Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln
690 695 700
Leu Pro Pro Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser
705 710 715 720
Arg Glu Gly Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly
725 730 735
Val Tyr His Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn
740 745 750
Arg His Leu Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr
755 760 765
Arg Val Pro Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val
770 775 780
Ser Thr Arg Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn
785 790 795 800
His Ser Lys Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn
805 810 815
Ile Ser His Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe
820 825 830
Arg Ser Met Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser
835 840 845
Asp Ser Pro Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser
850 855 860
Thr Pro Ser Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp
865 870 875 880
Leu Gln Thr Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His
885 890 895
Leu Ser Ser Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln
900 905 910
Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile
915 920 925
His Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu
930 935 940
Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Asp Gly Gly Cys
945 950 955 960
Asp Gly Arg Arg Gln Arg His His Ser Gly Pro Gln Asp Arg Arg Phe
965 970 975
Met Leu Arg Thr Thr Glu Gln Gln Gly Glu Tyr Phe Cys Cys Gly Asp
980 985 990
Pro Lys Lys Pro His Thr Pro Cys Val Pro Asn Arg Ala Leu His Arg
995 1000 1005
Pro Ile Ser Ser Pro Ala Pro Tyr Pro Val Leu Gln Val Arg Gly Thr
1010 1015 1020
Ser Met Cys Pro Thr Leu Gln Val Arg Gly Thr Asp Ala Phe Ser Cys
1025 1030 1035 1040
Pro Thr Gln Gln Ser Gly Phe Ser Phe Phe Val Arg His Val Met Arg
1045 1050 1055
Glu Ala Leu Ile His Arg Ala Gln Val Asn Gln Ala Ala Leu Leu Thr
1060 1065 1070
Tyr His Glu Asn Ala Ala Leu Thr Gly Lys Gly Gly Gly Gly Ser Glu
1075 1080 1085
Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys
1090 1095 1100
Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro
1105 1110 1115 1120
His His His His His His
1125
<210> 23
<211> 1037
<212> PRT
<213> Artificial Sequence
<220>
<223> >Tk28p_107_3xFlagHis_cho-opt in pOptiVec
<400> 23
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
340 345 350
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
355 360 365
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
370 375 380
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
385 390 395 400
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
405 410 415
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
420 425 430
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
435 440 445
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
450 455 460
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
465 470 475 480
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
485 490 495
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
500 505 510
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
515 520 525
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
530 535 540
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
545 550 555 560
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
565 570 575
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
580 585 590
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
595 600 605
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
610 615 620
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
625 630 635 640
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
645 650 655
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
660 665 670
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
675 680 685
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
690 695 700
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
705 710 715 720
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
725 730 735
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
740 745 750
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
755 760 765
Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys
770 775 780
Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His
785 790 795 800
Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met
805 810 815
Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro
820 825 830
Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser
835 840 845
Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr
850 855 860
Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser
865 870 875 880
Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr
885 890 895
Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu
900 905 910
Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro
915 920 925
Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp
930 935 940
His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser
945 950 955 960
Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg
965 970 975
Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala
980 985 990
Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys
995 1000 1005
Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp
1010 1015 1020
Asp Asp Lys Asp Gly Ala Pro His His His His His His
1025 1030 1035
<210> 24
<211> 1037
<212> PRT
<213> Artificial Sequence
<220>
<223> >Tk28p_107_3xFlagHis_ecoli-opt in pEX-1
<400> 24
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
340 345 350
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
355 360 365
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
370 375 380
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
385 390 395 400
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
405 410 415
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
420 425 430
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
435 440 445
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
450 455 460
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
465 470 475 480
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
485 490 495
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
500 505 510
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
515 520 525
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
530 535 540
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
545 550 555 560
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
565 570 575
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
580 585 590
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
595 600 605
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
610 615 620
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
625 630 635 640
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
645 650 655
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
660 665 670
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
675 680 685
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
690 695 700
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
705 710 715 720
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
725 730 735
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
740 745 750
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
755 760 765
Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys
770 775 780
Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His
785 790 795 800
Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met
805 810 815
Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro
820 825 830
Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser
835 840 845
Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr
850 855 860
Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser
865 870 875 880
Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr
885 890 895
Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu
900 905 910
Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro
915 920 925
Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp
930 935 940
His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser
945 950 955 960
Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg
965 970 975
Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala
980 985 990
Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys
995 1000 1005
Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp
1010 1015 1020
Asp Asp Lys Asp Gly Ala Pro His His His His His His
1025 1030 1035
<210> 25
<211> 929
<212> PRT
<213> Artificial Sequence
<220>
<223> >?853-960 in pEX-1
<400> 25
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
340 345 350
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
355 360 365
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
370 375 380
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
385 390 395 400
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
405 410 415
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
420 425 430
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
435 440 445
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
450 455 460
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
465 470 475 480
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
485 490 495
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
500 505 510
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
515 520 525
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
530 535 540
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
545 550 555 560
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
565 570 575
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
580 585 590
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
595 600 605
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
610 615 620
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
625 630 635 640
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
645 650 655
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
660 665 670
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
675 680 685
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
690 695 700
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
705 710 715 720
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
725 730 735
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
740 745 750
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
755 760 765
Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys
770 775 780
Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His
785 790 795 800
Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met
805 810 815
Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro
820 825 830
Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser
835 840 845
Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr
850 855 860
Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser
865 870 875 880
Ala Ser Asn His Pro Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln
885 890 895
Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp
900 905 910
Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His His His
915 920 925
His
<210> 26
<211> 821
<212> PRT
<213> Artificial Sequence
<220>
<223> >?745-960 in pEX-1
<400> 26
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
340 345 350
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
355 360 365
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
370 375 380
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
385 390 395 400
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
405 410 415
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
420 425 430
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
435 440 445
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
450 455 460
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
465 470 475 480
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
485 490 495
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
500 505 510
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
515 520 525
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
530 535 540
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
545 550 555 560
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
565 570 575
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
580 585 590
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
595 600 605
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
610 615 620
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
625 630 635 640
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
645 650 655
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
660 665 670
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
675 680 685
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
690 695 700
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
705 710 715 720
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
725 730 735
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
740 745 750
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
755 760 765
Val Ser Ser Leu Pro Ser Glu Ser Ser Gly Gly Gly Gly Ser Glu Asn
770 775 780
Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp
785 790 795 800
His Asp Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His
805 810 815
His His His His His
820
<210> 27
<211> 713
<212> PRT
<213> Artificial Sequence
<220>
<223> >?637-960 in pEX-1
<400> 27
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
340 345 350
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
355 360 365
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
370 375 380
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
385 390 395 400
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
405 410 415
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
420 425 430
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
435 440 445
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
450 455 460
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
465 470 475 480
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
485 490 495
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
500 505 510
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
515 520 525
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
530 535 540
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
545 550 555 560
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
565 570 575
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
580 585 590
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
595 600 605
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
610 615 620
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
625 630 635 640
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
645 650 655
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Gly Gly Gly
660 665 670
Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His Asp Gly
675 680 685
Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp
690 695 700
Gly Ala Pro His His His His His His
705 710
<210> 28
<211> 605
<212> PRT
<213> Artificial Sequence
<220>
<223> >?529-960 in pEX-1
<400> 28
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
340 345 350
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
355 360 365
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
370 375 380
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
385 390 395 400
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
405 410 415
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
420 425 430
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
435 440 445
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
450 455 460
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
465 470 475 480
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
485 490 495
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
500 505 510
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
515 520 525
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
530 535 540
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
545 550 555 560
Thr Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys
565 570 575
Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp
580 585 590
Asp Asp Lys Asp Gly Ala Pro His His His His His His
595 600 605
<210> 29
<211> 497
<212> PRT
<213> Artificial Sequence
<220>
<223> >?421-960 in pEX-1
<400> 29
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
340 345 350
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
355 360 365
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
370 375 380
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
385 390 395 400
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
405 410 415
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
420 425 430
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
435 440 445
Glu Phe Asp Phe Asn Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln
450 455 460
Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp
465 470 475 480
Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His His His
485 490 495
His
<210> 30
<211> 391
<212> PRT
<213> Artificial Sequence
<220>
<223> >?315-960 in pEX-1
<400> 30
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Gly Gly Gly Gly Ser
340 345 350
Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr
355 360 365
Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala
370 375 380
Pro His His His His His His
385 390
<210> 31
<211> 931
<212> PRT
<213> Artificial Sequence
<220>
<223> >?315-420 in pEX-1
<400> 31
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ile Asp Pro Lys Pro
340 345 350
Ser Glu Gly Pro Gly Thr Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln
355 360 365
Gln Asn Arg His Ser Phe Met Glu Ser Ser Gln Ser Lys Ala Gly Thr
370 375 380
Leu Gln Pro Asn Glu Lys Gln Ser Arg His Ser Tyr Ile Asp Thr Ile
385 390 395 400
Pro Gln Ser Ser Arg Ser Pro Ser Tyr Arg Thr Lys Ala Lys Ser His
405 410 415
Gly Ala Leu Ser Asp Ser Lys Ser Val Ser Asn Leu Ser Glu Ala Arg
420 425 430
Ala Gln Ile Ala Glu Pro Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys
435 440 445
Leu Asp Leu Asn Ser Pro Thr Ser Pro Thr Pro Thr Arg His Ser Asp
450 455 460
Thr Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly
465 470 475 480
Thr Leu Asp Ser Arg Arg Thr Thr Thr Arg His Ser Lys Thr Met Glu
485 490 495
Glu Leu Lys Leu Pro Glu His Met Asp Ser Ser His Ser His Ser Leu
500 505 510
Ser Ala Pro His Glu Ser Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro
515 520 525
Phe Ser Ser Gln Gln Arg Pro His Arg His Ser Met Tyr Val Thr Arg
530 535 540
Asp Lys Val Arg Ala Lys Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln
545 550 555 560
Gly Met Ala Ala Arg Ala Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro
565 570 575
Gly Glu Gln Leu Pro Pro Glu Met Thr Val Ala Arg Ser Ser Val Lys
580 585 590
Glu Thr Ser Arg Glu Gly Thr Ser Ser Phe His Thr Arg Gln Lys Ser
595 600 605
Glu Gly Gly Val Tyr His Asp Pro His Ser Asp Asp Gly Thr Ala Pro
610 615 620
Lys Glu Asn Arg His Leu Tyr Asn Asp Pro Val Pro Arg Arg Val Gly
625 630 635 640
Ser Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp Asn Ser Phe His Glu
645 650 655
Asn Asn Val Ser Thr Arg Val Ser Ser Leu Pro Ser Glu Ser Ser Ser
660 665 670
Gly Thr Asn His Ser Lys Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser
675 680 685
Pro Glu Asn Ile Ser His Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln
690 695 700
Gly Phe Phe Arg Ser Met Lys Lys Lys Lys Lys Lys Ser Gln Thr Val
705 710 715 720
Pro Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu Gln Lys Ser Ile His
725 730 735
Ser Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys
740 745 750
Ile Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys
755 760 765
Leu Leu His Leu Ser Ser Ala Ser Asn His Pro Ala Ser Ser Asp Pro
770 775 780
Arg Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu
785 790 795 800
Ile Arg Ile His Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile
805 810 815
Arg Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly
820 825 830
Gln Met Asp Pro Gly Trp His Val Ser Ser Val Thr Arg Ser Ala Thr
835 840 845
Glu Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn
850 855 860
Gly His Pro Tyr Asn Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn
865 870 875 880
Asp Leu Lys Glu Thr Ala Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr
885 890 895
Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp
900 905 910
Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His
915 920 925
His His His
930
<210> 32
<211> 823
<212> PRT
<213> Artificial Sequence
<220>
<223> >?315-528 in pEX-1
<400> 32
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ser Pro Thr Pro Thr
340 345 350
Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn Asn
355 360 365
Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr Thr Thr Arg His Ser
370 375 380
Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His Met Asp Ser Ser His
385 390 395 400
Ser His Ser Leu Ser Ala Pro His Glu Ser Phe Ser Tyr Gly Leu Gly
405 410 415
Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro His Arg His Ser Met
420 425 430
Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly Leu Asp Gly Ser Leu
435 440 445
Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn Ser Leu Gln Leu Leu
450 455 460
Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu Met Thr Val Ala Arg
465 470 475 480
Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr Ser Ser Phe His Thr
485 490 495
Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp Pro His Ser Asp Asp
500 505 510
Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr Asn Asp Pro Val Pro
515 520 525
Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp Asn
530 535 540
Ser Phe His Glu Asn Asn Val Ser Thr Arg Val Ser Ser Leu Pro Ser
545 550 555 560
Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg Gln Pro Ala Phe Asp
565 570 575
Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser Glu Gln Leu Lys Glu
580 585 590
Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys Lys Lys Lys Lys Lys
595 600 605
Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu Gln
610 615 620
Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu Trp
625 630 635 640
Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu Lys
645 650 655
Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala Ser Asn His Pro Ala
660 665 670
Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys Asn
675 680 685
Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser Gln Ala Ser Gly Gly
690 695 700
Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala Leu
705 710 715 720
Gln Leu Pro Gly Gln Met Asp Pro Gly Trp His Val Ser Ser Val Thr
725 730 735
Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala Lys
740 745 750
Ser Gly Pro Asn Gly His Pro Tyr Asn Arg Thr Asn Arg Ser Arg Met
755 760 765
Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala Leu Gly Gly Gly Gly Ser
770 775 780
Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr
785 790 795 800
Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala
805 810 815
Pro His His His His His His
820
<210> 33
<211> 715
<212> PRT
<213> Artificial Sequence
<220>
<223> >?315-636 in pEX-1
<400> 33
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ala Arg Ala Asn Ser
340 345 350
Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu Met
355 360 365
Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr Ser
370 375 380
Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp Pro
385 390 395 400
His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr Asn
405 410 415
Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser Pro
420 425 430
Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val Ser
435 440 445
Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg Gln
450 455 460
Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser Glu
465 470 475 480
Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys Lys
485 490 495
Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp Leu
500 505 510
Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser Arg
515 520 525
Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln Ser
530 535 540
Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala Ser
545 550 555 560
Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala Gln
565 570 575
Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser Gln
580 585 590
Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys Gly
595 600 605
Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp His Val
610 615 620
Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser Glu Gln
625 630 635 640
Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg Thr Asn
645 650 655
Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala Leu Gly
660 665 670
Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His
675 680 685
Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp
690 695 700
Lys Asp Gly Ala Pro His His His His His His
705 710 715
<210> 34
<211> 607
<212> PRT
<213> Artificial Sequence
<220>
<223> >?315-744 in pEX-1
<400> 34
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ser Gly Thr Asn His
340 345 350
Ser Lys Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile
355 360 365
Ser His Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg
370 375 380
Ser Met Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp
385 390 395 400
Ser Pro Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr
405 410 415
Pro Ser Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu
420 425 430
Gln Thr Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu
435 440 445
Ser Ser Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro
450 455 460
Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His
465 470 475 480
Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro
485 490 495
Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro
500 505 510
Gly Trp His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser
515 520 525
Tyr Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr
530 535 540
Asn Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu
545 550 555 560
Thr Ala Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp
565 570 575
Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys
580 585 590
Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His His His His
595 600 605
<210> 35
<211> 499
<212> PRT
<213> Artificial Sequence
<220>
<223> >?315-852 in pEX-1
<400> 35
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Ala Ser Ser Asp Pro
340 345 350
Arg Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu
355 360 365
Ile Arg Ile His Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile
370 375 380
Arg Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly
385 390 395 400
Gln Met Asp Pro Gly Trp His Val Ser Ser Val Thr Arg Ser Ala Thr
405 410 415
Glu Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn
420 425 430
Gly His Pro Tyr Asn Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn
435 440 445
Asp Leu Lys Glu Thr Ala Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr
450 455 460
Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp
465 470 475 480
Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His
485 490 495
His His His
<210> 36
<211> 1037
<212> PRT
<213> Artificial Sequence
<220>
<223> >Tt28p_107_3xFlagHis_ecoli-opt in pEX-1
<400> 36
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Gly Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Gly Gly
20 25 30
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
35 40 45
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
50 55 60
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
65 70 75 80
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
85 90 95
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
100 105 110
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
115 120 125
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
130 135 140
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
145 150 155 160
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
165 170 175
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
180 185 190
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
195 200 205
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
210 215 220
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
225 230 235 240
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
245 250 255
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
260 265 270
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
275 280 285
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
290 295 300
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
305 310 315 320
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
325 330 335
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
340 345 350
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
355 360 365
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
370 375 380
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
385 390 395 400
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
405 410 415
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
420 425 430
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
435 440 445
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
450 455 460
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
465 470 475 480
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
485 490 495
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
500 505 510
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
515 520 525
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
530 535 540
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
545 550 555 560
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
565 570 575
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
580 585 590
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
595 600 605
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
610 615 620
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
625 630 635 640
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
645 650 655
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
660 665 670
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
675 680 685
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
690 695 700
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
705 710 715 720
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
725 730 735
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
740 745 750
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
755 760 765
Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys
770 775 780
Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His
785 790 795 800
Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met
805 810 815
Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro
820 825 830
Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser
835 840 845
Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr
850 855 860
Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser
865 870 875 880
Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr
885 890 895
Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu
900 905 910
Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro
915 920 925
Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp
930 935 940
His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser
945 950 955 960
Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg
965 970 975
Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala
980 985 990
Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys
995 1000 1005
Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp
1010 1015 1020
Asp Asp Lys Asp Gly Ala Pro His His His His His His
1025 1030 1035
<210> 37
<211> 316
<212> PRT
<213> Artificial Sequence
<220>
<223> >Tk28p_eGFP_ecoli-opt_3xFlagHis in pEX-1
<400> 37
Met Gly Asp Ala Ala Gln Pro Ala Arg Arg Ala Arg Arg Thr Lys Leu
1 5 10 15
Ala Ala Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
20 25 30
Gly Ser Val Ser Lys Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile
35 40 45
Leu Val Glu Leu Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser
50 55 60
Gly Glu Gly Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu Lys Phe
65 70 75 80
Ile Cys Thr Thr Gly Lys Leu Pro Val Pro Trp Pro Thr Leu Val Thr
85 90 95
Thr Leu Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr Pro Asp His Met
100 105 110
Lys Gln His Asp Phe Phe Lys Ser Ala Met Pro Glu Gly Tyr Val Gln
115 120 125
Glu Arg Thr Ile Phe Phe Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala
130 135 140
Glu Val Lys Phe Glu Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys
145 150 155 160
Gly Ile Asp Phe Lys Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu
165 170 175
Tyr Asn Tyr Asn Ser His Asn Val Tyr Ile Met Ala Asp Lys Gln Lys
180 185 190
Asn Gly Ile Lys Val Asn Phe Lys Ile Arg His Asn Ile Glu Asp Gly
195 200 205
Ser Val Gln Leu Ala Asp His Tyr Gln Gln Asn Thr Pro Ile Gly Asp
210 215 220
Gly Pro Val Leu Leu Pro Asp Asn His Tyr Leu Ser Thr Gln Ser Ala
225 230 235 240
Leu Ser Lys Asp Pro Asn Glu Lys Arg Asp His Met Val Leu Leu Glu
245 250 255
Phe Val Thr Ala Ala Gly Ile Thr Leu Gly Met Asp Glu Leu Tyr Lys
260 265 270
Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp
275 280 285
His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp
290 295 300
Asp Lys Asp Gly Ala Pro His His His His His His
305 310 315
<210> 38
<211> 283
<212> PRT
<213> Artificial Sequence
<220>
<223> >eGFP_3xFlagHis_ecoli-opt in pEX-1
<400> 38
Met Val Ser Lys Gly Glu Glu Leu Phe Thr Gly Val Val Pro Ile Leu
1 5 10 15
Val Glu Leu Asp Gly Asp Val Asn Gly His Lys Phe Ser Val Ser Gly
20 25 30
Glu Gly Glu Gly Asp Ala Thr Tyr Gly Lys Leu Thr Leu Lys Phe Ile
35 40 45
Cys Thr Thr Gly Lys Leu Pro Val Pro Trp Pro Thr Leu Val Thr Thr
50 55 60
Leu Thr Tyr Gly Val Gln Cys Phe Ser Arg Tyr Pro Asp His Met Lys
65 70 75 80
Gln His Asp Phe Phe Lys Ser Ala Met Pro Glu Gly Tyr Val Gln Glu
85 90 95
Arg Thr Ile Phe Phe Lys Asp Asp Gly Asn Tyr Lys Thr Arg Ala Glu
100 105 110
Val Lys Phe Glu Gly Asp Thr Leu Val Asn Arg Ile Glu Leu Lys Gly
115 120 125
Ile Asp Phe Lys Glu Asp Gly Asn Ile Leu Gly His Lys Leu Glu Tyr
130 135 140
Asn Tyr Asn Ser His Asn Val Tyr Ile Met Ala Asp Lys Gln Lys Asn
145 150 155 160
Gly Ile Lys Val Asn Phe Lys Ile Arg His Asn Ile Glu Asp Gly Ser
165 170 175
Val Gln Leu Ala Asp His Tyr Gln Gln Asn Thr Pro Ile Gly Asp Gly
180 185 190
Pro Val Leu Leu Pro Asp Asn His Tyr Leu Ser Thr Gln Ser Ala Leu
195 200 205
Ser Lys Asp Pro Asn Glu Lys Arg Asp His Met Val Leu Leu Glu Phe
210 215 220
Val Thr Ala Ala Gly Ile Thr Leu Gly Met Asp Glu Leu Tyr Lys Gly
225 230 235 240
Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His
245 250 255
Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp
260 265 270
Lys Asp Gly Ala Pro His His His His His His
275 280
<210> 39
<211> 739
<212> PRT
<213> Artificial Sequence
<220>
<223> >AMPH1-3xFlagHis in pEX-1 (ecoli-opt)
<400> 39
Met Ala Asp Ile Lys Thr Gly Ile Phe Ala Lys Asn Val Gln Lys Arg
1 5 10 15
Leu Asn Arg Ala Gln Glu Lys Val Leu Gln Lys Leu Gly Lys Ala Asp
20 25 30
Glu Thr Lys Asp Glu Gln Phe Glu Glu Tyr Val Gln Asn Phe Lys Arg
35 40 45
Gln Glu Ala Glu Gly Thr Arg Leu Gln Arg Glu Leu Arg Gly Tyr Leu
50 55 60
Ala Ala Ile Lys Gly Met Gln Glu Ala Ser Met Lys Leu Thr Glu Ser
65 70 75 80
Leu His Glu Val Tyr Glu Pro Asp Trp Tyr Gly Arg Glu Asp Val Lys
85 90 95
Met Val Gly Glu Lys Cys Asp Val Leu Trp Glu Asp Phe His Gln Lys
100 105 110
Leu Val Asp Gly Ser Leu Leu Thr Leu Asp Thr Tyr Leu Gly Gln Phe
115 120 125
Pro Asp Ile Lys Asn Arg Ile Ala Lys Arg Ser Arg Lys Leu Val Asp
130 135 140
Tyr Asp Ser Ala Arg His His Leu Glu Ala Leu Gln Ser Ser Lys Arg
145 150 155 160
Lys Asp Glu Ser Arg Ile Ser Lys Ala Glu Glu Glu Phe Gln Lys Ala
165 170 175
Gln Lys Val Phe Glu Glu Phe Asn Val Asp Leu Gln Glu Glu Leu Pro
180 185 190
Ser Leu Trp Ser Arg Arg Val Gly Phe Tyr Val Asn Thr Phe Lys Asn
195 200 205
Val Ser Ser Leu Glu Ala Lys Phe His Lys Glu Ile Ala Val Leu Cys
210 215 220
His Lys Leu Tyr Glu Val Met Thr Lys Leu Gly Asp Gln His Ala Asp
225 230 235 240
Lys Ala Phe Thr Ile Gln Gly Ala Pro Ser Asp Ser Gly Pro Leu Arg
245 250 255
Ile Ala Lys Thr Pro Ser Pro Pro Glu Glu Pro Ser Pro Leu Pro Ser
260 265 270
Pro Thr Ala Ser Pro Asn His Thr Leu Ala Pro Ala Ser Pro Ala Pro
275 280 285
Ala Arg Pro Arg Ser Pro Ser Gln Thr Arg Lys Gly Pro Pro Val Pro
290 295 300
Pro Leu Pro Lys Val Thr Pro Thr Lys Glu Leu Gln Gln Glu Asn Ile
305 310 315 320
Ile Ser Phe Phe Glu Asp Asn Phe Val Pro Glu Ile Ser Val Thr Thr
325 330 335
Pro Ser Gln Asn Glu Val Pro Glu Val Lys Lys Glu Glu Thr Leu Leu
340 345 350
Asp Leu Asp Phe Asp Pro Phe Lys Pro Glu Val Thr Pro Ala Gly Ser
355 360 365
Ala Gly Val Thr His Ser Pro Met Ser Gln Thr Leu Pro Trp Asp Leu
370 375 380
Trp Thr Thr Ser Thr Asp Leu Val Gln Pro Ala Ser Gly Gly Ser Phe
385 390 395 400
Asn Gly Phe Thr Gln Pro Gln Asp Thr Ser Leu Phe Thr Met Gln Thr
405 410 415
Asp Gln Ser Met Ile Cys Asn Leu Ala Glu Ser Glu Gln Ala Pro Pro
420 425 430
Thr Glu Pro Lys Ala Glu Glu Pro Leu Ala Ala Val Thr Pro Ala Val
435 440 445
Gly Leu Asp Leu Gly Met Asp Thr Arg Ala Glu Glu Pro Val Glu Glu
450 455 460
Ala Val Ile Ile Pro Gly Ala Asp Ala Asp Ala Ala Val Gly Thr Leu
465 470 475 480
Val Ser Ala Ala Glu Gly Ala Pro Gly Glu Glu Ala Glu Ala Glu Lys
485 490 495
Ala Thr Val Pro Ala Gly Glu Gly Val Ser Leu Glu Glu Ala Lys Ile
500 505 510
Gly Thr Glu Thr Thr Glu Gly Ala Glu Ser Ala Gln Pro Glu Ala Glu
515 520 525
Glu Leu Glu Ala Thr Val Pro Gln Glu Lys Val Ile Pro Ser Val Val
530 535 540
Ile Glu Pro Ala Ser Asn His Glu Glu Glu Gly Glu Asn Glu Ile Thr
545 550 555 560
Ile Gly Ala Glu Pro Lys Glu Thr Thr Glu Asp Ala Ala Pro Pro Gly
565 570 575
Pro Thr Ser Glu Thr Pro Glu Leu Ala Thr Glu Gln Lys Pro Ile Gln
580 585 590
Asp Pro Gln Pro Thr Pro Ser Ala Pro Ala Met Gly Ala Ala Asp Gln
595 600 605
Leu Ala Ser Ala Arg Glu Ala Ser Gln Glu Leu Pro Pro Gly Phe Leu
610 615 620
Tyr Lys Val Glu Thr Leu His Asp Phe Glu Ala Ala Asn Ser Asp Glu
625 630 635 640
Leu Thr Leu Gln Arg Gly Asp Val Val Leu Val Val Pro Ser Asp Ser
645 650 655
Glu Ala Asp Gln Asp Ala Gly Trp Leu Val Gly Val Lys Glu Ser Asp
660 665 670
Trp Leu Gln Tyr Arg Asp Leu Ala Thr Tyr Lys Gly Leu Phe Pro Glu
675 680 685
Asn Phe Thr Arg Arg Leu Asp Glu Asn Leu Tyr Phe Gln Gly Gly Gly
690 695 700
Gly Gly Ser Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp
705 710 715 720
Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His
725 730 735
His His His
<210> 40
<211> 739
<212> PRT
<213> Artificial Sequence
<220>
<223> >AMPH1-3xFlagHis cho-opt in pOptiVec
<400> 40
Met Ala Asp Ile Lys Thr Gly Ile Phe Ala Lys Asn Val Gln Lys Arg
1 5 10 15
Leu Asn Arg Ala Gln Glu Lys Val Leu Gln Lys Leu Gly Lys Ala Asp
20 25 30
Glu Thr Lys Asp Glu Gln Phe Glu Glu Tyr Val Gln Asn Phe Lys Arg
35 40 45
Gln Glu Ala Glu Gly Thr Arg Leu Gln Arg Glu Leu Arg Gly Tyr Leu
50 55 60
Ala Ala Ile Lys Gly Met Gln Glu Ala Ser Met Lys Leu Thr Glu Ser
65 70 75 80
Leu His Glu Val Tyr Glu Pro Asp Trp Tyr Gly Arg Glu Asp Val Lys
85 90 95
Met Val Gly Glu Lys Cys Asp Val Leu Trp Glu Asp Phe His Gln Lys
100 105 110
Leu Val Asp Gly Ser Leu Leu Thr Leu Asp Thr Tyr Leu Gly Gln Phe
115 120 125
Pro Asp Ile Lys Asn Arg Ile Ala Lys Arg Ser Arg Lys Leu Val Asp
130 135 140
Tyr Asp Ser Ala Arg His His Leu Glu Ala Leu Gln Ser Ser Lys Arg
145 150 155 160
Lys Asp Glu Ser Arg Ile Ser Lys Ala Glu Glu Glu Phe Gln Lys Ala
165 170 175
Gln Lys Val Phe Glu Glu Phe Asn Val Asp Leu Gln Glu Glu Leu Pro
180 185 190
Ser Leu Trp Ser Arg Arg Val Gly Phe Tyr Val Asn Thr Phe Lys Asn
195 200 205
Val Ser Ser Leu Glu Ala Lys Phe His Lys Glu Ile Ala Val Leu Cys
210 215 220
His Lys Leu Tyr Glu Val Met Thr Lys Leu Gly Asp Gln His Ala Asp
225 230 235 240
Lys Ala Phe Thr Ile Gln Gly Ala Pro Ser Asp Ser Gly Pro Leu Arg
245 250 255
Ile Ala Lys Thr Pro Ser Pro Pro Glu Glu Pro Ser Pro Leu Pro Ser
260 265 270
Pro Thr Ala Ser Pro Asn His Thr Leu Ala Pro Ala Ser Pro Ala Pro
275 280 285
Ala Arg Pro Arg Ser Pro Ser Gln Thr Arg Lys Gly Pro Pro Val Pro
290 295 300
Pro Leu Pro Lys Val Thr Pro Thr Lys Glu Leu Gln Gln Glu Asn Ile
305 310 315 320
Ile Ser Phe Phe Glu Asp Asn Phe Val Pro Glu Ile Ser Val Thr Thr
325 330 335
Pro Ser Gln Asn Glu Val Pro Glu Val Lys Lys Glu Glu Thr Leu Leu
340 345 350
Asp Leu Asp Phe Asp Pro Phe Lys Pro Glu Val Thr Pro Ala Gly Ser
355 360 365
Ala Gly Val Thr His Ser Pro Met Ser Gln Thr Leu Pro Trp Asp Leu
370 375 380
Trp Thr Thr Ser Thr Asp Leu Val Gln Pro Ala Ser Gly Gly Ser Phe
385 390 395 400
Asn Gly Phe Thr Gln Pro Gln Asp Thr Ser Leu Phe Thr Met Gln Thr
405 410 415
Asp Gln Ser Met Ile Cys Asn Leu Ala Glu Ser Glu Gln Ala Pro Pro
420 425 430
Thr Glu Pro Lys Ala Glu Glu Pro Leu Ala Ala Val Thr Pro Ala Val
435 440 445
Gly Leu Asp Leu Gly Met Asp Thr Arg Ala Glu Glu Pro Val Glu Glu
450 455 460
Ala Val Ile Ile Pro Gly Ala Asp Ala Asp Ala Ala Val Gly Thr Leu
465 470 475 480
Val Ser Ala Ala Glu Gly Ala Pro Gly Glu Glu Ala Glu Ala Glu Lys
485 490 495
Ala Thr Val Pro Ala Gly Glu Gly Val Ser Leu Glu Glu Ala Lys Ile
500 505 510
Gly Thr Glu Thr Thr Glu Gly Ala Glu Ser Ala Gln Pro Glu Ala Glu
515 520 525
Glu Leu Glu Ala Thr Val Pro Gln Glu Lys Val Ile Pro Ser Val Val
530 535 540
Ile Glu Pro Ala Ser Asn His Glu Glu Glu Gly Glu Asn Glu Ile Thr
545 550 555 560
Ile Gly Ala Glu Pro Lys Glu Thr Thr Glu Asp Ala Ala Pro Pro Gly
565 570 575
Pro Thr Ser Glu Thr Pro Glu Leu Ala Thr Glu Gln Lys Pro Ile Gln
580 585 590
Asp Pro Gln Pro Thr Pro Ser Ala Pro Ala Met Gly Ala Ala Asp Gln
595 600 605
Leu Ala Ser Ala Arg Glu Ala Ser Gln Glu Leu Pro Pro Gly Phe Leu
610 615 620
Tyr Lys Val Glu Thr Leu His Asp Phe Glu Ala Ala Asn Ser Asp Glu
625 630 635 640
Leu Thr Leu Gln Arg Gly Asp Val Val Leu Val Val Pro Ser Asp Ser
645 650 655
Glu Ala Asp Gln Asp Ala Gly Trp Leu Val Gly Val Lys Glu Ser Asp
660 665 670
Trp Leu Gln Tyr Arg Asp Leu Ala Thr Tyr Lys Gly Leu Phe Pro Glu
675 680 685
Asn Phe Thr Arg Arg Leu Asp Glu Asn Leu Tyr Phe Gln Gly Gly Gly
690 695 700
Gly Gly Ser Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp
705 710 715 720
Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His
725 730 735
His His His
<210> 41
<211> 1048
<212> PRT
<213> Artificial Sequence
<220>
<223> >MBip_Tatk11_107_3xFlagHis_cho-opt in pOptiVec
<400> 41
Met Lys Leu Ser Leu Val Ala Ala Met Leu Leu Leu Leu Ser Leu Val
1 5 10 15
Ala Ala Met Leu Leu Leu Leu Ser Ala Ala Arg Ala Gly Tyr Ala Arg
20 25 30
Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly Gly Ser Lys Ile Pro
35 40 45
Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu Gly Val Val Gly
50 55 60
Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His Lys Glu Thr His
65 70 75 80
Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu Glu Asn Glu Glu
85 90 95
Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu Arg Thr Leu Lys
100 105 110
Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg Arg Arg Gly Lys
115 120 125
Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met Leu Glu Leu Leu
130 135 140
Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val Lys Ser Tyr Ile
145 150 155 160
Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys Asn Asp Ile Val
165 170 175
His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser His Asn Asp Val
180 185 190
Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu Ser Glu Gly Asn
195 200 205
Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp Tyr Arg Ser Pro
210 215 220
Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val Asp Met Trp Ser
225 230 235 240
Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln Pro Leu Phe Pro
245 250 255
Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln Lys Val Leu Gly
260 265 270
Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser Asn Pro Arg Phe
275 280 285
His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln Ser Leu Glu Arg
290 295 300
Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp Leu Met Lys Asn
305 310 315 320
Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr Glu Gln Cys Leu
325 330 335
Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp Arg Ser Pro Ser
340 345 350
Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser Ser Thr Leu Ser
355 360 365
Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln Ser His His Arg
370 375 380
Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly Leu Pro Arg Ala
385 390 395 400
Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn Gly Asn Leu Ala
405 410 415
Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr Gln Ala Ser Ser
420 425 430
Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn Asn Ile Pro His
435 440 445
Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu Phe Asp Phe Asn
450 455 460
Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys Tyr Leu Lys Ser
465 470 475 480
Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met Glu Ser Ser Gln
485 490 495
Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln Ser Arg His Ser
500 505 510
Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro Ser Tyr Arg Thr
515 520 525
Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys Ser Val Ser Asn
530 535 540
Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser Thr Ser Arg Tyr
545 550 555 560
Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr Ser Pro Thr Pro
565 570 575
Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn
580 585 590
Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr Thr Thr Arg His
595 600 605
Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His Met Asp Ser Ser
610 615 620
His Ser His Ser Leu Ser Ala Pro His Glu Ser Phe Ser Tyr Gly Leu
625 630 635 640
Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro His Arg His Ser
645 650 655
Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly Leu Asp Gly Ser
660 665 670
Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn Ser Leu Gln Leu
675 680 685
Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu Met Thr Val Ala
690 695 700
Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr Ser Ser Phe His
705 710 715 720
Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp Pro His Ser Asp
725 730 735
Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr Asn Asp Pro Val
740 745 750
Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp
755 760 765
Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val Ser Ser Leu Pro
770 775 780
Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg Gln Pro Ala Phe
785 790 795 800
Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser Glu Gln Leu Lys
805 810 815
Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys Lys Lys Lys Lys
820 825 830
Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu
835 840 845
Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu
850 855 860
Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu
865 870 875 880
Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala Ser Asn His Pro
885 890 895
Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys
900 905 910
Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser Gln Ala Ser Gly
915 920 925
Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala
930 935 940
Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp His Val Ser Ser Val
945 950 955 960
Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala
965 970 975
Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg Thr Asn Arg Ser Arg
980 985 990
Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala Leu Gly Gly Gly Gly
995 1000 1005
Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp
1010 1015 1020
Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly
1025 1030 1035 1040
Ala Pro His His His His His His
1045
<210> 42
<211> 1040
<212> PRT
<213> Artificial Sequence
<220>
<223> >IgK_Tatk11_107_3xFlagHis_cho-opt in pOptiVec
<400> 42
Met Glu Thr Asp Thr Leu Leu Leu Trp Val Leu Leu Leu Trp Val Pro
1 5 10 15
Gly Ser Thr Gly Gly Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala
20 25 30
Gly Gly Gly Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys
35 40 45
Phe Glu Ile Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu
50 55 60
Lys Cys Arg His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe
65 70 75 80
Lys Asp Ser Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu
85 90 95
Leu Lys Met Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys
100 105 110
Glu Ala Phe Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val
115 120 125
Glu Lys Asn Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro
130 135 140
Pro Glu Lys Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His
145 150 155 160
Trp Cys His Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn
165 170 175
Leu Leu Ile Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe
180 185 190
Ala Arg Asn Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val
195 200 205
Ala Thr Arg Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr
210 215 220
Gly Lys Ser Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu
225 230 235 240
Ser Asp Gly Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu
245 250 255
Phe Thr Ile Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys
260 265 270
Leu Phe Tyr Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val
275 280 285
Asn His Pro Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser
290 295 300
Val Leu Leu Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp
305 310 315 320
Arg Tyr Leu Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln
325 330 335
Arg Leu Leu Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr
340 345 350
His Val Glu Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser
355 360 365
Thr Ala Leu Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn
370 375 380
Leu Ser Val Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu
385 390 395 400
Ser Phe Leu Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His
405 410 415
Thr Lys Thr Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp
420 425 430
Leu Thr Asn Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys
435 440 445
Ser Lys Thr Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly
450 455 460
Pro Gly Thr Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg
465 470 475 480
His Ser Phe Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro
485 490 495
Asn Glu Lys Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser
500 505 510
Ser Arg Ser Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu
515 520 525
Ser Asp Ser Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile
530 535 540
Ala Glu Pro Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu
545 550 555 560
Asn Ser Pro Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr
565 570 575
Leu Leu Ser Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp
580 585 590
Ser Arg Arg Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys
595 600 605
Leu Pro Glu His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro
610 615 620
His Glu Ser Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser
625 630 635 640
Gln Gln Arg Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val
645 650 655
Arg Ala Lys Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala
660 665 670
Ala Arg Ala Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln
675 680 685
Leu Pro Pro Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser
690 695 700
Arg Glu Gly Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly
705 710 715 720
Val Tyr His Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn
725 730 735
Arg His Leu Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr
740 745 750
Arg Val Pro Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val
755 760 765
Ser Thr Arg Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn
770 775 780
His Ser Lys Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn
785 790 795 800
Ile Ser His Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe
805 810 815
Arg Ser Met Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser
820 825 830
Asp Ser Pro Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser
835 840 845
Thr Pro Ser Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp
850 855 860
Leu Gln Thr Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His
865 870 875 880
Leu Ser Ser Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln
885 890 895
Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile
900 905 910
His Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu
915 920 925
Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp
930 935 940
Pro Gly Trp His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro
945 950 955 960
Ser Tyr Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro
965 970 975
Tyr Asn Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys
980 985 990
Glu Thr Ala Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly
995 1000 1005
Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr
1010 1015 1020
Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His His His His
1025 1030 1035 1040
<210> 43
<211> 1021
<212> PRT
<213> Artificial Sequence
<220>
<223> >Tatk11_107_3xFlagHis_cho-opt in pOptiVec (leaderless)
<400> 43
Met Gly Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
1 5 10 15
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
20 25 30
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
35 40 45
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
50 55 60
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
65 70 75 80
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
85 90 95
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
100 105 110
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
115 120 125
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
130 135 140
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
145 150 155 160
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
165 170 175
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
180 185 190
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
195 200 205
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
210 215 220
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
225 230 235 240
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
245 250 255
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
260 265 270
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
275 280 285
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
290 295 300
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
305 310 315 320
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
325 330 335
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
340 345 350
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
355 360 365
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
370 375 380
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
385 390 395 400
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
405 410 415
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
420 425 430
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
435 440 445
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
450 455 460
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
465 470 475 480
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
485 490 495
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
500 505 510
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
515 520 525
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
530 535 540
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
545 550 555 560
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
565 570 575
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
580 585 590
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
595 600 605
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
610 615 620
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
625 630 635 640
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
645 650 655
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
660 665 670
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
675 680 685
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
690 695 700
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
705 710 715 720
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
725 730 735
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
740 745 750
Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys
755 760 765
Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His
770 775 780
Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met
785 790 795 800
Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro
805 810 815
Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser
820 825 830
Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr
835 840 845
Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser
850 855 860
Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr
865 870 875 880
Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu
885 890 895
Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro
900 905 910
Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp
915 920 925
His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser
930 935 940
Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg
945 950 955 960
Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala
965 970 975
Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys
980 985 990
Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp
995 1000 1005
Asp Asp Lys Asp Gly Ala Pro His His His His His His
1010 1015 1020
<210> 44
<211> 1021
<212> PRT
<213> Artificial Sequence
<220>
<223> >Tatk11_107_3xFlagHis_ecoli-opt in pEX-1
<400> 44
Met Gly Tyr Ala Arg Lys Ala Ala Arg Gln Ala Arg Ala Gly Gly Gly
1 5 10 15
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
20 25 30
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
35 40 45
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
50 55 60
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
65 70 75 80
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
85 90 95
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
100 105 110
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
115 120 125
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
130 135 140
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
145 150 155 160
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
165 170 175
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
180 185 190
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
195 200 205
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
210 215 220
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
225 230 235 240
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
245 250 255
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
260 265 270
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
275 280 285
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
290 295 300
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
305 310 315 320
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
325 330 335
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
340 345 350
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
355 360 365
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
370 375 380
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
385 390 395 400
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
405 410 415
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
420 425 430
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
435 440 445
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
450 455 460
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
465 470 475 480
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
485 490 495
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
500 505 510
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
515 520 525
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
530 535 540
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
545 550 555 560
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
565 570 575
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
580 585 590
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
595 600 605
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
610 615 620
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
625 630 635 640
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
645 650 655
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
660 665 670
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
675 680 685
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
690 695 700
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
705 710 715 720
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
725 730 735
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
740 745 750
Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys
755 760 765
Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His
770 775 780
Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met
785 790 795 800
Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro
805 810 815
Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser
820 825 830
Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr
835 840 845
Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser
850 855 860
Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr
865 870 875 880
Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu
885 890 895
Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro
900 905 910
Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp
915 920 925
His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser
930 935 940
Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg
945 950 955 960
Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala
965 970 975
Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys
980 985 990
Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp
995 1000 1005
Asp Asp Lys Asp Gly Ala Pro His His His His His His
1010 1015 1020
<210> 45
<211> 1021
<212> PRT
<213> Artificial Sequence
<220>
<223> >Tat11_107_3xFlagHis_ecoli-opt in pEX-1
<400> 45
Met Gly Tyr Gly Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Gly Gly
1 5 10 15
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
20 25 30
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
35 40 45
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
50 55 60
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
65 70 75 80
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
85 90 95
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
100 105 110
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
115 120 125
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
130 135 140
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
145 150 155 160
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
165 170 175
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
180 185 190
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
195 200 205
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
210 215 220
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
225 230 235 240
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
245 250 255
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
260 265 270
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
275 280 285
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
290 295 300
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
305 310 315 320
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
325 330 335
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
340 345 350
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
355 360 365
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
370 375 380
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
385 390 395 400
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
405 410 415
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
420 425 430
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
435 440 445
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
450 455 460
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
465 470 475 480
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
485 490 495
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
500 505 510
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
515 520 525
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
530 535 540
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
545 550 555 560
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
565 570 575
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
580 585 590
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
595 600 605
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
610 615 620
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
625 630 635 640
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
645 650 655
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
660 665 670
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
675 680 685
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
690 695 700
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
705 710 715 720
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
725 730 735
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
740 745 750
Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys
755 760 765
Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His
770 775 780
Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met
785 790 795 800
Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro
805 810 815
Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser
820 825 830
Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr
835 840 845
Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser
850 855 860
Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr
865 870 875 880
Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu
885 890 895
Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro
900 905 910
Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp
915 920 925
His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser
930 935 940
Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg
945 950 955 960
Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala
965 970 975
Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys
980 985 990
Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp
995 1000 1005
Asp Asp Lys Asp Gly Ala Pro His His His His His His
1010 1015 1020
<210> 46
<211> 1021
<212> PRT
<213> Artificial Sequence
<220>
<223> >Tat11_107_3xFlagHis_cho-opt in pOptiVec (leaderless)
<400> 46
Met Gly Tyr Gly Arg Lys Lys Arg Arg Gln Arg Arg Arg Gly Gly Gly
1 5 10 15
Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile
20 25 30
Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg
35 40 45
His Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser
50 55 60
Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met
65 70 75 80
Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe
85 90 95
Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn
100 105 110
Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys
115 120 125
Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His
130 135 140
Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile
145 150 155 160
Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn
165 170 175
Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg
180 185 190
Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser
195 200 205
Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly
210 215 220
Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile
225 230 235 240
Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr
245 250 255
Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro
260 265 270
Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu
275 280 285
Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu
290 295 300
Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu
305 310 315 320
Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu
325 330 335
Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu
340 345 350
Gln Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val
355 360 365
Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu
370 375 380
Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr
385 390 395 400
Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn
405 410 415
Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr
420 425 430
Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr
435 440 445
Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe
450 455 460
Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys
465 470 475 480
Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser
485 490 495
Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser
500 505 510
Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro
515 520 525
Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro
530 535 540
Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser
545 550 555 560
Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg
565 570 575
Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu
580 585 590
His Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser
595 600 605
Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg
610 615 620
Pro His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys
625 630 635 640
Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala
645 650 655
Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro
660 665 670
Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly
675 680 685
Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His
690 695 700
Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu
705 710 715 720
Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro
725 730 735
Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg
740 745 750
Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys
755 760 765
Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His
770 775 780
Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met
785 790 795 800
Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro
805 810 815
Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser
820 825 830
Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr
835 840 845
Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser
850 855 860
Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr
865 870 875 880
Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu
885 890 895
Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro
900 905 910
Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln Met Asp Pro Gly Trp
915 920 925
His Val Ser Ser Val Thr Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser
930 935 940
Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly His Pro Tyr Asn Arg
945 950 955 960
Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala
965 970 975
Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys
980 985 990
Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile Asp Tyr Lys Asp Asp
995 1000 1005
Asp Asp Lys Asp Gly Ala Pro His His His His His His
1010 1015 1020
<210> 47
<211> 1030
<212> PRT
<213> Artificial Sequence
<220>
<223> CDKL5115 isoform polypeptide 1-1030 (full-length)
<400> 47
Met Lys Ile Pro Asn Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu
1 5 10 15
Gly Val Val Gly Glu Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His
20 25 30
Lys Glu Thr His Glu Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu
35 40 45
Glu Asn Glu Glu Val Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu
50 55 60
Arg Thr Leu Lys Gln Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg
65 70 75 80
Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met
85 90 95
Leu Glu Leu Leu Glu Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val
100 105 110
Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala Ile His Trp Cys His Lys
115 120 125
Asn Asp Ile Val His Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser
130 135 140
His Asn Asp Val Leu Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu
145 150 155 160
Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp
165 170 175
Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val
180 185 190
Asp Met Trp Ser Val Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln
195 200 205
Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln
210 215 220
Lys Val Leu Gly Pro Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser
225 230 235 240
Asn Pro Arg Phe His Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln
245 250 255
Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp
260 265 270
Leu Met Lys Asn Leu Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr
275 280 285
Glu Gln Cys Leu Asn His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp
290 295 300
Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser
305 310 315 320
Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln
325 330 335
Ser His His Arg Ser Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly
340 345 350
Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn
355 360 365
Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr
370 375 380
Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn
385 390 395 400
Asn Ile Pro His Leu Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu
405 410 415
Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys
420 425 430
Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met
435 440 445
Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln
450 455 460
Ser Arg His Ser Tyr Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro
465 470 475 480
Ser Tyr Arg Thr Lys Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys
485 490 495
Ser Val Ser Asn Leu Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser
500 505 510
Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr
515 520 525
Ser Pro Thr Pro Thr Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro
530 535 540
Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr
545 550 555 560
Thr Thr Arg His Ser Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His
565 570 575
Met Asp Ser Ser His Ser His Ser Leu Ser Ala Pro His Glu Ser Phe
580 585 590
Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro
595 600 605
His Arg His Ser Met Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly
610 615 620
Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn
625 630 635 640
Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu
645 650 655
Met Thr Val Ala Arg Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr
660 665 670
Ser Ser Phe His Thr Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp
675 680 685
Pro His Ser Asp Asp Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr
690 695 700
Asn Asp Pro Val Pro Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser
705 710 715 720
Pro Arg Pro Asp Asn Ser Phe His Glu Asn Asn Val Ser Thr Arg Val
725 730 735
Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg
740 745 750
Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser
755 760 765
Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys
770 775 780
Lys Lys Lys Lys Lys Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp
785 790 795 800
Leu Leu Thr Leu Gln Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser
805 810 815
Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln
820 825 830
Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala
835 840 845
Ser Asn His Pro Ala Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala
850 855 860
Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser
865 870 875 880
Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys
885 890 895
Gly Arg Pro Ala Leu Gln Leu Pro Asp Gly Gly Cys Asp Gly Arg Arg
900 905 910
Gln Arg His His Ser Gly Pro Gln Asp Arg Arg Phe Met Leu Arg Thr
915 920 925
Thr Glu Gln Gln Gly Glu Tyr Phe Cys Cys Gly Asp Pro Lys Lys Pro
930 935 940
His Thr Pro Cys Val Pro Asn Arg Ala Leu His Arg Pro Ile Ser Ser
945 950 955 960
Pro Ala Pro Tyr Pro Val Leu Gln Val Arg Gly Thr Ser Met Cys Pro
965 970 975
Thr Leu Gln Val Arg Gly Thr Asp Ala Phe Ser Cys Pro Thr Gln Gln
980 985 990
Ser Gly Phe Ser Phe Phe Val Arg His Val Met Arg Glu Ala Leu Ile
995 1000 1005
His Arg Ala Gln Val Asn Gln Ala Ala Leu Leu Thr Tyr His Glu Asn
1010 1015 1020
Ala Ala Leu Thr Gly Lys
1025 1030
<210> 48
<211> 28
<212> PRT
<213> Artificial Sequence
<220>
<223> MBiP
<400> 48
Met Lys Leu Ser Leu Val Ala Ala Met Leu Leu Leu Leu Ser Leu Val
1 5 10 15
Ala Ala Met Leu Leu Leu Leu Ser Ala Ala Arg Ala
20 25
<210> 49
<211> 20
<212> PRT
<213> Artificial Sequence
<220>
<223> Murine Ig?
<400> 49
Met Glu Thr Asp Thr Leu Leu Leu Trp Val Leu Leu Leu Trp Val Pro
1 5 10 15
Gly Ser Thr Gly
20
<210> 50
<211> 12
<212> PRT
<213> Artificial Sequence
<220>
<223> P97
<400> 50
Asp Ser Ser His Ala Phe Thr Leu Asp Glu Leu Arg
1 5 10
<210> 51
<211> 25
<212> PRT
<213> Artificial Sequence
<220>
<223> MBiP2
<400> 51
Met Lys Leu Ser Leu Val Ala Ala Met Leu Leu Leu Leu Trp Val Ala
1 5 10 15
Leu Leu Leu Leu Ser Ala Ala Arg Ala
20 25
<210> 52
<211> 26
<212> PRT
<213> Artificial Sequence
<220>
<223> MBiP3
<400> 52
Met Lys Leu Ser Leu Val Ala Ala Met Leu Leu Leu Leu Ser Leu Val
1 5 10 15
Ala Leu Leu Leu Leu Ser Ala Ala Arg Ala
20 25
<210> 53
<211> 26
<212> PRT
<213> Artificial Sequence
<220>
<223> MBiP4
<400> 53
Met Lys Leu Ser Leu Val Ala Ala Met Leu Leu Leu Leu Ala Leu Val
1 5 10 15
Ala Leu Leu Leu Leu Ser Ala Ala Arg Ala
20 25
<210> 54
<211> 1026
<212> PRT
<213> Artificial Sequence
<220>
<223> >ANTP_107_3xFlagHis_cho-opt in pOptiVec
<400> 54
Met Gly Arg Gln Ile Lys Ile Trp Phe Gln Asn Arg Arg Met Lys Trp
1 5 10 15
Lys Lys Gly Gly Gly Gly Ser Lys Ile Pro Asn Ile Gly Asn Val Met
20 25 30
Asn Lys Phe Glu Ile Leu Gly Val Val Gly Glu Gly Ala Tyr Gly Val
35 40 45
Val Leu Lys Cys Arg His Lys Glu Thr His Glu Ile Val Ala Ile Lys
50 55 60
Lys Phe Lys Asp Ser Glu Glu Asn Glu Glu Val Lys Glu Thr Thr Leu
65 70 75 80
Arg Glu Leu Lys Met Leu Arg Thr Leu Lys Gln Glu Asn Ile Val Glu
85 90 95
Leu Lys Glu Ala Phe Arg Arg Arg Gly Lys Leu Tyr Leu Val Phe Glu
100 105 110
Tyr Val Glu Lys Asn Met Leu Glu Leu Leu Glu Glu Met Pro Asn Gly
115 120 125
Val Pro Pro Glu Lys Val Lys Ser Tyr Ile Tyr Gln Leu Ile Lys Ala
130 135 140
Ile His Trp Cys His Lys Asn Asp Ile Val His Arg Asp Ile Lys Pro
145 150 155 160
Glu Asn Leu Leu Ile Ser His Asn Asp Val Leu Lys Leu Cys Asp Phe
165 170 175
Gly Phe Ala Arg Asn Leu Ser Glu Gly Asn Asn Ala Asn Tyr Thr Glu
180 185 190
Tyr Val Ala Thr Arg Trp Tyr Arg Ser Pro Glu Leu Leu Leu Gly Ala
195 200 205
Pro Tyr Gly Lys Ser Val Asp Met Trp Ser Val Gly Cys Ile Leu Gly
210 215 220
Glu Leu Ser Asp Gly Gln Pro Leu Phe Pro Gly Glu Ser Glu Ile Asp
225 230 235 240
Gln Leu Phe Thr Ile Gln Lys Val Leu Gly Pro Leu Pro Ser Glu Gln
245 250 255
Met Lys Leu Phe Tyr Ser Asn Pro Arg Phe His Gly Leu Arg Phe Pro
260 265 270
Ala Val Asn His Pro Gln Ser Leu Glu Arg Arg Tyr Leu Gly Ile Leu
275 280 285
Asn Ser Val Leu Leu Asp Leu Met Lys Asn Leu Leu Lys Leu Asp Pro
290 295 300
Ala Asp Arg Tyr Leu Thr Glu Gln Cys Leu Asn His Pro Thr Phe Gln
305 310 315 320
Thr Gln Arg Leu Leu Asp Arg Ser Pro Ser Arg Ser Ala Lys Arg Lys
325 330 335
Pro Tyr His Val Glu Ser Ser Thr Leu Ser Asn Arg Asn Gln Ala Gly
340 345 350
Lys Ser Thr Ala Leu Gln Ser His His Arg Ser Asn Ser Lys Asp Ile
355 360 365
Gln Asn Leu Ser Val Gly Leu Pro Arg Ala Asp Glu Gly Leu Pro Ala
370 375 380
Asn Glu Ser Phe Leu Asn Gly Asn Leu Ala Gly Ala Ser Leu Ser Pro
385 390 395 400
Leu His Thr Lys Thr Tyr Gln Ala Ser Ser Gln Pro Gly Ser Thr Ser
405 410 415
Lys Asp Leu Thr Asn Asn Asn Ile Pro His Leu Leu Ser Pro Lys Glu
420 425 430
Ala Lys Ser Lys Thr Glu Phe Asp Phe Asn Ile Asp Pro Lys Pro Ser
435 440 445
Glu Gly Pro Gly Thr Lys Tyr Leu Lys Ser Asn Ser Arg Ser Gln Gln
450 455 460
Asn Arg His Ser Phe Met Glu Ser Ser Gln Ser Lys Ala Gly Thr Leu
465 470 475 480
Gln Pro Asn Glu Lys Gln Ser Arg His Ser Tyr Ile Asp Thr Ile Pro
485 490 495
Gln Ser Ser Arg Ser Pro Ser Tyr Arg Thr Lys Ala Lys Ser His Gly
500 505 510
Ala Leu Ser Asp Ser Lys Ser Val Ser Asn Leu Ser Glu Ala Arg Ala
515 520 525
Gln Ile Ala Glu Pro Ser Thr Ser Arg Tyr Phe Pro Ser Ser Cys Leu
530 535 540
Asp Leu Asn Ser Pro Thr Ser Pro Thr Pro Thr Arg His Ser Asp Thr
545 550 555 560
Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn Asn Arg Asn Glu Gly Thr
565 570 575
Leu Asp Ser Arg Arg Thr Thr Thr Arg His Ser Lys Thr Met Glu Glu
580 585 590
Leu Lys Leu Pro Glu His Met Asp Ser Ser His Ser His Ser Leu Ser
595 600 605
Ala Pro His Glu Ser Phe Ser Tyr Gly Leu Gly Tyr Thr Ser Pro Phe
610 615 620
Ser Ser Gln Gln Arg Pro His Arg His Ser Met Tyr Val Thr Arg Asp
625 630 635 640
Lys Val Arg Ala Lys Gly Leu Asp Gly Ser Leu Ser Ile Gly Gln Gly
645 650 655
Met Ala Ala Arg Ala Asn Ser Leu Gln Leu Leu Ser Pro Gln Pro Gly
660 665 670
Glu Gln Leu Pro Pro Glu Met Thr Val Ala Arg Ser Ser Val Lys Glu
675 680 685
Thr Ser Arg Glu Gly Thr Ser Ser Phe His Thr Arg Gln Lys Ser Glu
690 695 700
Gly Gly Val Tyr His Asp Pro His Ser Asp Asp Gly Thr Ala Pro Lys
705 710 715 720
Glu Asn Arg His Leu Tyr Asn Asp Pro Val Pro Arg Arg Val Gly Ser
725 730 735
Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp Asn Ser Phe His Glu Asn
740 745 750
Asn Val Ser Thr Arg Val Ser Ser Leu Pro Ser Glu Ser Ser Ser Gly
755 760 765
Thr Asn His Ser Lys Arg Gln Pro Ala Phe Asp Pro Trp Lys Ser Pro
770 775 780
Glu Asn Ile Ser His Ser Glu Gln Leu Lys Glu Lys Glu Lys Gln Gly
785 790 795 800
Phe Phe Arg Ser Met Lys Lys Lys Lys Lys Lys Ser Gln Thr Val Pro
805 810 815
Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu Gln Lys Ser Ile His Ser
820 825 830
Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu Trp Arg Pro Glu Lys Ile
835 840 845
Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu Lys Ser Leu Arg Lys Leu
850 855 860
Leu His Leu Ser Ser Ala Ser Asn His Pro Ala Ser Ser Asp Pro Arg
865 870 875 880
Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys Asn Ser Phe Ser Glu Ile
885 890 895
Arg Ile His Pro Leu Ser Gln Ala Ser Gly Gly Ser Ser Asn Ile Arg
900 905 910
Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala Leu Gln Leu Pro Gly Gln
915 920 925
Met Asp Pro Gly Trp His Val Ser Ser Val Thr Arg Ser Ala Thr Glu
930 935 940
Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala Lys Ser Gly Pro Asn Gly
945 950 955 960
His Pro Tyr Asn Arg Thr Asn Arg Ser Arg Met Pro Asn Leu Asn Asp
965 970 975
Leu Lys Glu Thr Ala Leu Gly Gly Gly Gly Ser Glu Asn Leu Tyr Phe
980 985 990
Gln Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr Lys Asp His Asp Ile
995 1000 1005
Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala Pro His His His His
1010 1015 1020
His His
1025
<210> 55
<211> 1031
<212> PRT
<213> Artificial Sequence
<220>
<223> >TRANSP_107_3xFlagHis_cho-opt in pOptiVec
<400> 55
Met Gly Ala Gly Tyr Leu Leu Gly Lys Ile Asn Leu Lys Ala Leu Ala
1 5 10 15
Ala Leu Ala Lys Lys Ile Leu Gly Gly Gly Gly Ser Lys Ile Pro Asn
20 25 30
Ile Gly Asn Val Met Asn Lys Phe Glu Ile Leu Gly Val Val Gly Glu
35 40 45
Gly Ala Tyr Gly Val Val Leu Lys Cys Arg His Lys Glu Thr His Glu
50 55 60
Ile Val Ala Ile Lys Lys Phe Lys Asp Ser Glu Glu Asn Glu Glu Val
65 70 75 80
Lys Glu Thr Thr Leu Arg Glu Leu Lys Met Leu Arg Thr Leu Lys Gln
85 90 95
Glu Asn Ile Val Glu Leu Lys Glu Ala Phe Arg Arg Arg Gly Lys Leu
100 105 110
Tyr Leu Val Phe Glu Tyr Val Glu Lys Asn Met Leu Glu Leu Leu Glu
115 120 125
Glu Met Pro Asn Gly Val Pro Pro Glu Lys Val Lys Ser Tyr Ile Tyr
130 135 140
Gln Leu Ile Lys Ala Ile His Trp Cys His Lys Asn Asp Ile Val His
145 150 155 160
Arg Asp Ile Lys Pro Glu Asn Leu Leu Ile Ser His Asn Asp Val Leu
165 170 175
Lys Leu Cys Asp Phe Gly Phe Ala Arg Asn Leu Ser Glu Gly Asn Asn
180 185 190
Ala Asn Tyr Thr Glu Tyr Val Ala Thr Arg Trp Tyr Arg Ser Pro Glu
195 200 205
Leu Leu Leu Gly Ala Pro Tyr Gly Lys Ser Val Asp Met Trp Ser Val
210 215 220
Gly Cys Ile Leu Gly Glu Leu Ser Asp Gly Gln Pro Leu Phe Pro Gly
225 230 235 240
Glu Ser Glu Ile Asp Gln Leu Phe Thr Ile Gln Lys Val Leu Gly Pro
245 250 255
Leu Pro Ser Glu Gln Met Lys Leu Phe Tyr Ser Asn Pro Arg Phe His
260 265 270
Gly Leu Arg Phe Pro Ala Val Asn His Pro Gln Ser Leu Glu Arg Arg
275 280 285
Tyr Leu Gly Ile Leu Asn Ser Val Leu Leu Asp Leu Met Lys Asn Leu
290 295 300
Leu Lys Leu Asp Pro Ala Asp Arg Tyr Leu Thr Glu Gln Cys Leu Asn
305 310 315 320
His Pro Thr Phe Gln Thr Gln Arg Leu Leu Asp Arg Ser Pro Ser Arg
325 330 335
Ser Ala Lys Arg Lys Pro Tyr His Val Glu Ser Ser Thr Leu Ser Asn
340 345 350
Arg Asn Gln Ala Gly Lys Ser Thr Ala Leu Gln Ser His His Arg Ser
355 360 365
Asn Ser Lys Asp Ile Gln Asn Leu Ser Val Gly Leu Pro Arg Ala Asp
370 375 380
Glu Gly Leu Pro Ala Asn Glu Ser Phe Leu Asn Gly Asn Leu Ala Gly
385 390 395 400
Ala Ser Leu Ser Pro Leu His Thr Lys Thr Tyr Gln Ala Ser Ser Gln
405 410 415
Pro Gly Ser Thr Ser Lys Asp Leu Thr Asn Asn Asn Ile Pro His Leu
420 425 430
Leu Ser Pro Lys Glu Ala Lys Ser Lys Thr Glu Phe Asp Phe Asn Ile
435 440 445
Asp Pro Lys Pro Ser Glu Gly Pro Gly Thr Lys Tyr Leu Lys Ser Asn
450 455 460
Ser Arg Ser Gln Gln Asn Arg His Ser Phe Met Glu Ser Ser Gln Ser
465 470 475 480
Lys Ala Gly Thr Leu Gln Pro Asn Glu Lys Gln Ser Arg His Ser Tyr
485 490 495
Ile Asp Thr Ile Pro Gln Ser Ser Arg Ser Pro Ser Tyr Arg Thr Lys
500 505 510
Ala Lys Ser His Gly Ala Leu Ser Asp Ser Lys Ser Val Ser Asn Leu
515 520 525
Ser Glu Ala Arg Ala Gln Ile Ala Glu Pro Ser Thr Ser Arg Tyr Phe
530 535 540
Pro Ser Ser Cys Leu Asp Leu Asn Ser Pro Thr Ser Pro Thr Pro Thr
545 550 555 560
Arg His Ser Asp Thr Arg Thr Leu Leu Ser Pro Ser Gly Arg Asn Asn
565 570 575
Arg Asn Glu Gly Thr Leu Asp Ser Arg Arg Thr Thr Thr Arg His Ser
580 585 590
Lys Thr Met Glu Glu Leu Lys Leu Pro Glu His Met Asp Ser Ser His
595 600 605
Ser His Ser Leu Ser Ala Pro His Glu Ser Phe Ser Tyr Gly Leu Gly
610 615 620
Tyr Thr Ser Pro Phe Ser Ser Gln Gln Arg Pro His Arg His Ser Met
625 630 635 640
Tyr Val Thr Arg Asp Lys Val Arg Ala Lys Gly Leu Asp Gly Ser Leu
645 650 655
Ser Ile Gly Gln Gly Met Ala Ala Arg Ala Asn Ser Leu Gln Leu Leu
660 665 670
Ser Pro Gln Pro Gly Glu Gln Leu Pro Pro Glu Met Thr Val Ala Arg
675 680 685
Ser Ser Val Lys Glu Thr Ser Arg Glu Gly Thr Ser Ser Phe His Thr
690 695 700
Arg Gln Lys Ser Glu Gly Gly Val Tyr His Asp Pro His Ser Asp Asp
705 710 715 720
Gly Thr Ala Pro Lys Glu Asn Arg His Leu Tyr Asn Asp Pro Val Pro
725 730 735
Arg Arg Val Gly Ser Phe Tyr Arg Val Pro Ser Pro Arg Pro Asp Asn
740 745 750
Ser Phe His Glu Asn Asn Val Ser Thr Arg Val Ser Ser Leu Pro Ser
755 760 765
Glu Ser Ser Ser Gly Thr Asn His Ser Lys Arg Gln Pro Ala Phe Asp
770 775 780
Pro Trp Lys Ser Pro Glu Asn Ile Ser His Ser Glu Gln Leu Lys Glu
785 790 795 800
Lys Glu Lys Gln Gly Phe Phe Arg Ser Met Lys Lys Lys Lys Lys Lys
805 810 815
Ser Gln Thr Val Pro Asn Ser Asp Ser Pro Asp Leu Leu Thr Leu Gln
820 825 830
Lys Ser Ile His Ser Ala Ser Thr Pro Ser Ser Arg Pro Lys Glu Trp
835 840 845
Arg Pro Glu Lys Ile Ser Asp Leu Gln Thr Gln Ser Gln Pro Leu Lys
850 855 860
Ser Leu Arg Lys Leu Leu His Leu Ser Ser Ala Ser Asn His Pro Ala
865 870 875 880
Ser Ser Asp Pro Arg Phe Gln Pro Leu Thr Ala Gln Gln Thr Lys Asn
885 890 895
Ser Phe Ser Glu Ile Arg Ile His Pro Leu Ser Gln Ala Ser Gly Gly
900 905 910
Ser Ser Asn Ile Arg Gln Glu Pro Ala Pro Lys Gly Arg Pro Ala Leu
915 920 925
Gln Leu Pro Gly Gln Met Asp Pro Gly Trp His Val Ser Ser Val Thr
930 935 940
Arg Ser Ala Thr Glu Gly Pro Ser Tyr Ser Glu Gln Leu Gly Ala Lys
945 950 955 960
Ser Gly Pro Asn Gly His Pro Tyr Asn Arg Thr Asn Arg Ser Arg Met
965 970 975
Pro Asn Leu Asn Asp Leu Lys Glu Thr Ala Leu Gly Gly Gly Gly Ser
980 985 990
Glu Asn Leu Tyr Phe Gln Gly Asp Tyr Lys Asp His Asp Gly Asp Tyr
995 1000 1005
Lys Asp His Asp Ile Asp Tyr Lys Asp Asp Asp Asp Lys Asp Gly Ala
1010 1015 1020
Pro His His His His His His
1025 1030
Claims (44)
- CDKL5 폴리펩티드로서, SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 대해 적어도 98%의 서열 동일성을 갖는 서열을 포함하는, CDKL5 폴리펩티드.
- 제1항에 있어서, 상기 CDKL5 폴리펩티드는 SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 대해 적어도 99%의 서열 동일성을 갖는 서열을 포함하는, CDKL5 폴리펩티드.
- 제1항에 있어서, 상기 CDKL5 폴리펩티드는 SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 또는 SEQ ID NO: 12에 대해 100%의 서열 동일성을 갖는 서열을 포함하는, CDKL5 폴리펩티드.
- 핵 배출 신호(NES)를 결여한 CDKL5 폴리펩티드.
- 제4항에 있어서, 상기 CDKL5 폴리펩티드는 핵 위치 신호(NLS)를 함유하는, CDKL5 폴리펩티드.
- 제4항에 있어서, 상기 CDKL5 폴리펩티드는 핵 위치 신호(NLS)를 함유하지 않는, CDKL5 폴리펩티드.
- 핵 위치 신호(NLS)를 결여하며 핵 배출 신호(NES)를 함유하는, CDKL5 폴리펩티드.
- 제1항 내지 제7항 중 어느 한 항의 CDKL5 폴리펩티드 및 상기 CDKL5 폴리펩티드에 작동적으로 연결된 리더 신호 폴리펩티드를 포함하는, 융합 단백질.
- 제1항 내지 제7항 중 어느 한 항의 CDKL5 폴리펩티드 및 상기 CDKL5 폴리펩티드에 작동적으로 연결된 세포-침투 폴리펩티드를 포함하는, 융합 단백질.
- 제9항에 있어서, 상기 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 90%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제9항에 있어서, 상기 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 100%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제9항 내지 제11항 중 어느 한 항에 있어서, 제9항 내지 제11항 중 어느 한 항의 융합 단백질에 작동적으로 연결된 리더 신호 폴리펩티드를 추가로 포함하는, 융합 단백질.
- 제8항 또는 제12항에 있어서, 상기 리더 신호 폴리펩티드는 SEQ ID NO: 48, SEQ ID NO: 49, SEQ ID NO: 51, SEQ ID NO: 52 또는 SEQ ID NO: 53에 대해 적어도 90%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제8항 또는 제12항에 있어서, 상기 리더 신호 폴리펩티드는 SEQ ID NO: 48, SEQ ID NO: 49, SEQ ID NO: 51, SEQ ID NO: 52 또는 SEQ ID NO: 53에 대해 100%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제1항 내지 제7항 중 어느 한 항의 CDKL5 폴리펩티드 또는 제8항 내지 제14항 중 어느 한 항의 융합 단백질; 및
약학적으로 허용되는 담체를 포함하는, 약학 제형. - 제15항의 제형을 이를 필요로 하는 환자에게 투여하는 단계를 포함하는, CDKL5-매개 신경계 장애의 치료 방법.
- 제16항에 있어서, 상기 제형은 척수강내, 정맥내, 수조내, 뇌실내 또는 실질내 투여되는, 방법.
- 제16항 또는 제17항에 있어서, 상기 제형은 척수강내 또는 정맥내 투여되는, 방법.
- 제16항 내지 제18항 중 어느 한 항에 있어서, 상기 CDKL5-매개 신경계 장애는 CDKL5 결핍, 또는 CDKL5 돌연변이 또는 결핍에 의해 야기된 비정형 레트 증후군(Rett Syndrome) 중 하나 이상인, 방법.
- 제1항 내지 제7항 중 어느 한 항의 CDKL5 폴리펩티드 또는 제8항 내지 제14항 중 어느 한 항의 융합 단백질을 제조하는 방법으로서,
상기 CDKL5 폴리펩티드 또는 상기 융합 단백질을 발현시키는 단계; 및
상기 CDKL5 폴리펩티드 또는 상기 융합 단백질을 정제하는 단계를 포함하는, 방법. - 제20항에 있어서, 상기 CDKL5 폴리펩티드 또는 상기 융합 단백질은 차이니즈 햄스터 난소(CHO) 세포, HeLa 세포, 인간 배아 신장(HEK) 세포 또는 대장균 세포에서 발현되는, 방법.
- 제1항 내지 제7항 중 어느 한 항의 CDKL5 폴리펩티드 또는 제8항 내지 제14항 중 어느 한 항의 융합 단백질을 인코딩하는, 폴리뉴클레오티드.
- 제22항의 폴리뉴클레오티드를 포함하는 벡터.
- CDKL5 폴리펩티드 및 함께 작동적으로 연결된 세포-침투 폴리펩티드를 포함하는 융합 단백질로서, 상기 CDKL5 폴리펩티드는 SEQ ID NO: 1 또는 SEQ ID NO: 47에 대해 적어도 98%의 서열 동일성을 갖는 서열을 포함하고, 상기 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 90%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제24항에 있어서, 상기 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 95%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제25항에 있어서, 상기 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 100%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제24항 내지 제26항 중 어느 한 항에 있어서, 리더 신호 폴리펩티드를 추가로 포함하는, 융합 단백질.
- 제27항에 있어서, 상기 리더 신호 폴리펩티드는 SEQ ID NO: 48, SEQ ID NO: 49, SEQ ID NO: 51, SEQ ID NO: 52 또는 SEQ ID NO: 53에 대해 적어도 90%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제28항에 있어서, 상기 리더 신호 폴리펩티드는 SEQ ID NO: 48, SEQ ID NO: 49, SEQ ID NO: 51, SEQ ID NO: 52 또는 SEQ ID NO: 53에 대해 100%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- CDKL5 폴리펩티드 및 함께 작동적으로 연결된 리더 신호 폴리펩티드를 포함하는 융합 단백질로서, 상기 CDKL5 폴리펩티드는 SEQ ID NO: 1 또는 SEQ ID NO: 47에 대해 적어도 98%의 서열 동일성을 갖는 서열을 포함하고, 상기 리더 신호 폴리펩티드는 SEQ ID NO: 48, SEQ ID NO: 51, SEQ ID NO: 52 또는 SEQ ID NO: 53에 대해 적어도 90%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제30항에 있어서, 상기 리더 신호 폴리펩티드는 SEQ ID NO: 48, SEQ ID NO: 51, SEQ ID NO: 52 또는 SEQ ID NO: 53에 대해 100%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제30항 또는 제31항에 있어서, 세포-침투 폴리펩티드를 추가로 포함하는, 융합 단백질.
- 제32항에 있어서, 상기 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 90%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제33항에 있어서, 상기 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 적어도 95%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제34항에 있어서, 상기 세포-침투 폴리펩티드는 SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18 또는 SEQ ID NO: 50에 대해 100%의 서열 동일성을 갖는 서열을 포함하는, 융합 단백질.
- 제24항 내지 제35항 중 어느 한 항의 융합 단백질; 및 약학적으로 허용되는 담체를 포함하는, 약학 제형.
- 제36항의 제형을 이를 필요로 하는 환자에게 투여하는 단계를 포함하는, CDKL5-매개 신경계 장애의 치료 방법.
- 제37항에 있어서, 상기 제형은 척수강내, 정맥내, 수조내, 뇌실내 또는 실질내 투여되는, 방법.
- 제37항 또는 제38항에 있어서, 상기 제형은 척수강내 또는 정맥내 투여되는, 방법.
- 제37항 내지 제39항 중 어느 한 항에 있어서, 상기 CDKL5-매개 신경계 장애는 CDKL5 결핍, 또는 CDKL5 돌연변이 또는 결핍에 의해 야기된 비정형 레트 증후군 중 하나 이상인, 방법.
- 제24항 내지 제35항 중 어느 한 항에 따른 융합 단백질을 제조하는 방법으로서,
상기 융합 단백질을 발현시키는 단계; 및
상기 융합 단백질을 정제하는 단계를 포함하는, 방법. - 제41항에 있어서, 상기 융합 단백질은 차이니즈 햄스터 난소(CHO) 세포, HeLa 세포, 인간 배아 신장(HEK) 세포 또는 대장균 세포에서 발현되는, 방법.
- 제24항 내지 제35항 중 어느 한 항의 융합 단백질을 인코딩하는 폴리뉴클레오티드.
- 제43항의 폴리뉴클레오티드를 포함하는 벡터.
Applications Claiming Priority (5)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US201762592936P | 2017-11-30 | 2017-11-30 | |
| US201762592944P | 2017-11-30 | 2017-11-30 | |
| US62/592,936 | 2017-11-30 | ||
| US62/592,944 | 2017-11-30 | ||
| PCT/US2018/063294 WO2019108924A2 (en) | 2017-11-30 | 2018-11-30 | Cdkl5 expression variants and cdkl5 fusion proteins |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| KR20200090889A true KR20200090889A (ko) | 2020-07-29 |
Family
ID=65003456
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| KR1020207018589A KR20200090889A (ko) | 2017-11-30 | 2018-11-30 | Cdkl5 발현 변이체 및 cdkl5 융합 단백질 |
Country Status (10)
| Country | Link |
|---|---|
| US (1) | US20200299654A1 (ko) |
| EP (1) | EP3717642A2 (ko) |
| JP (1) | JP2021505135A (ko) |
| KR (1) | KR20200090889A (ko) |
| CN (1) | CN111936624A (ko) |
| AU (1) | AU2018375754A1 (ko) |
| CA (1) | CA3083951A1 (ko) |
| MX (1) | MX2020005670A (ko) |
| TW (1) | TW201927825A (ko) |
| WO (1) | WO2019108924A2 (ko) |
Families Citing this family (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20230043046A1 (en) * | 2019-10-30 | 2023-02-09 | Amicus Therapeutics, Inc. | Recombinant CDKL5 Proteins, Gene Therapy and Production Methods |
| CA3200192A1 (en) | 2020-12-01 | 2022-06-09 | Justin PERCIVAL | Compositions and uses thereof for treatment of angelman syndrome |
| CN114716569B (zh) * | 2022-04-13 | 2023-11-10 | 浙江大学 | 一种携带目标蛋白自主进入真核细胞的重组蛋白、重组表达载体和重组菌及应用 |
| CN116377050A (zh) * | 2022-12-16 | 2023-07-04 | 湖南家辉生物技术有限公司 | 一种发育性癫痫性脑病2型致病基因cdkl5突变位点的应用及其检测试剂和应用 |
Family Cites Families (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2005072470A2 (en) * | 2004-01-28 | 2005-08-11 | Exelixis, Inc | Mbms as modifiers of branching morphogenesis and methods of use |
| BR112013012671B1 (pt) * | 2010-11-22 | 2022-03-03 | Amicus Therapeutics, Inc | Sequência sinal de polipeptídio, proteína de fusão e vetor de expressão de proteína |
| WO2015128746A2 (en) * | 2014-02-28 | 2015-09-03 | Alma Mater Studiorum-Universita Di Bologna | Tatk-cdkl5 fusion proteins, compositions, formulations, and use thereof |
-
2018
- 2018-11-30 US US16/768,511 patent/US20200299654A1/en active Pending
- 2018-11-30 AU AU2018375754A patent/AU2018375754A1/en active Pending
- 2018-11-30 WO PCT/US2018/063294 patent/WO2019108924A2/en unknown
- 2018-11-30 CN CN201880085312.3A patent/CN111936624A/zh active Pending
- 2018-11-30 MX MX2020005670A patent/MX2020005670A/es unknown
- 2018-11-30 JP JP2020529748A patent/JP2021505135A/ja active Pending
- 2018-11-30 CA CA3083951A patent/CA3083951A1/en active Pending
- 2018-11-30 EP EP18830554.4A patent/EP3717642A2/en active Pending
- 2018-11-30 TW TW107142931A patent/TW201927825A/zh unknown
- 2018-11-30 KR KR1020207018589A patent/KR20200090889A/ko not_active Application Discontinuation
Also Published As
| Publication number | Publication date |
|---|---|
| MX2020005670A (es) | 2020-11-24 |
| AU2018375754A1 (en) | 2020-07-16 |
| TW201927825A (zh) | 2019-07-16 |
| CA3083951A1 (en) | 2019-06-06 |
| JP2021505135A (ja) | 2021-02-18 |
| CN111936624A (zh) | 2020-11-13 |
| US20200299654A1 (en) | 2020-09-24 |
| WO2019108924A3 (en) | 2019-08-15 |
| WO2019108924A2 (en) | 2019-06-06 |
| EP3717642A2 (en) | 2020-10-07 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US9932377B2 (en) | Mitochondrial targeting and therapeutic use thereof | |
| KR20200090889A (ko) | Cdkl5 발현 변이체 및 cdkl5 융합 단백질 | |
| US8735341B2 (en) | Non-viral delivery of compounds to mitochondria | |
| Nakayama et al. | Fibroblast growth factor-12 (FGF12) translocation into intestinal epithelial cells is dependent on a novel cell-penetrating peptide domain: Involvement of internalization in the in vivo role of exogenous FGF12 | |
| CA2855223A1 (en) | Use of cell-permeable peptide inhibitors of the jnk signal transduction pathway for the treatment of dry eye syndrome | |
| JP2022554267A (ja) | 組換えcdkl5タンパク質、遺伝子療法、及び製造方法 | |
| AU2017248353A1 (en) | TDP-43 mitochondrial localization inhibitor for the treatment of neurodegenerative disease | |
| US20180170983A1 (en) | New Use of Cell-Permeable Peptide Inhibitors of the JNK Signal Transduction Pathway for the Treatment of Mild Cognitive Impairment | |
| US10550151B2 (en) | Cell-penetrating compositions and methods using same | |
| AU2017296189A1 (en) | Ocular delivery of cell permeant therapeutics for the treatment of retinal edema | |
| US10036016B2 (en) | Methods for inducing glucose uptake | |
| JP2012511309A (ja) | Ec−sodのカルボキシル末端のアポプチンタンパク質導入ドメインの融合蛋白質 | |
| US20220033450A1 (en) | Virally expressed inhibitors of pdz domains, such as pick1 and uses thereof | |
| US10385113B2 (en) | Engineered FGF compositions and methods of use thereof | |
| KR101131512B1 (ko) | 퇴행성신경질환 예방 또는 치료용 약제학적 조성물 | |
| US20140303093A1 (en) | Micro-utrophin polypeptides and methods | |
| WO2014161095A1 (en) | Crp40 fragments for the treatment of neurological disorders | |
| Risco Quiroz | Development of a Novel Strategy to Treat Spinal Muscular Atrophy | |
| EP2785363A1 (en) | Use of cell-permeable peptide inhibitors of the jnk signal transduction pathway for the treatment of dry eye syndrome |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| A201 | Request for examination | ||
| E902 | Notification of reason for refusal |