WO2023051802A1 - 直接转分化治疗神经系统疾病 - Google Patents
直接转分化治疗神经系统疾病 Download PDFInfo
- Publication number
- WO2023051802A1 WO2023051802A1 PCT/CN2022/123409 CN2022123409W WO2023051802A1 WO 2023051802 A1 WO2023051802 A1 WO 2023051802A1 CN 2022123409 W CN2022123409 W CN 2022123409W WO 2023051802 A1 WO2023051802 A1 WO 2023051802A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- rest
- cells
- protein
- neurons
- variant
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Images
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K45/00—Medicinal preparations containing active ingredients not provided for in groups A61K31/00 - A61K41/00
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
- C07K14/4701—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals not used
- C07K14/4702—Regulators; Modulating activity
- C07K14/4703—Inhibitors; Suppressors
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/70—Carbohydrates; Sugars; Derivatives thereof
- A61K31/7088—Compounds having three or more nucleosides or nucleotides
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/70—Carbohydrates; Sugars; Derivatives thereof
- A61K31/7088—Compounds having three or more nucleosides or nucleotides
- A61K31/713—Double-stranded nucleic acids or oligonucleotides
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K48/00—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy
- A61K48/0008—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy characterised by an aspect of the 'non-active' part of the composition delivered, e.g. wherein such 'non-active' part is not delivered simultaneously with the 'active' part of the composition
- A61K48/0025—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy characterised by an aspect of the 'non-active' part of the composition delivered, e.g. wherein such 'non-active' part is not delivered simultaneously with the 'active' part of the composition wherein the non-active part clearly interacts with the delivered nucleic acid
- A61K48/0033—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy characterised by an aspect of the 'non-active' part of the composition delivered, e.g. wherein such 'non-active' part is not delivered simultaneously with the 'active' part of the composition wherein the non-active part clearly interacts with the delivered nucleic acid the non-active part being non-polymeric
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K48/00—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy
- A61K48/005—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy characterised by an aspect of the 'active' part of the composition delivered, i.e. the nucleic acid delivered
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K48/00—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy
- A61K48/005—Medicinal preparations containing genetic material which is inserted into cells of the living body to treat genetic diseases; Gene therapy characterised by an aspect of the 'active' part of the composition delivered, i.e. the nucleic acid delivered
- A61K48/0058—Nucleic acids adapted for tissue specific expression, e.g. having tissue specific promoters as part of a contruct
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P21/00—Drugs for disorders of the muscular or neuromuscular system
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
- A61P25/08—Antiepileptics; Anticonvulsants
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
- A61P25/14—Drugs for disorders of the nervous system for treating abnormal movements, e.g. chorea, dyskinesia
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
- A61P25/14—Drugs for disorders of the nervous system for treating abnormal movements, e.g. chorea, dyskinesia
- A61P25/16—Anti-Parkinson drugs
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
- A61P25/18—Antipsychotics, i.e. neuroleptics; Drugs for mania or schizophrenia
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
- A61P25/20—Hypnotics; Sedatives
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
- A61P25/28—Drugs for disorders of the nervous system for treating neurodegenerative disorders of the central nervous system, e.g. nootropic agents, cognition enhancers, drugs for treating Alzheimer's disease or other forms of dementia
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P27/00—Drugs for disorders of the senses
- A61P27/02—Ophthalmic agents
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P27/00—Drugs for disorders of the senses
- A61P27/02—Ophthalmic agents
- A61P27/06—Antiglaucoma agents or miotics
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P27/00—Drugs for disorders of the senses
- A61P27/16—Otologicals
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P9/00—Drugs for disorders of the cardiovascular system
- A61P9/10—Drugs for disorders of the cardiovascular system for treating ischaemic or atherosclerotic diseases, e.g. antianginal drugs, coronary vasodilators, drugs for myocardial infarction, retinopathy, cerebrovascula insufficiency, renal arteriosclerosis
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
- C12N15/1137—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing against enzymes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/85—Vectors or expression systems specially adapted for eukaryotic hosts for animal cells
- C12N15/86—Viral vectors
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N5/00—Undifferentiated human, animal or plant cells, e.g. cell lines; Tissues; Cultivation or maintenance thereof; Culture media therefor
- C12N5/06—Animal cells or tissues; Human cells or tissues
- C12N5/0602—Vertebrate cells
- C12N5/0618—Cells of the nervous system
- C12N5/0619—Neurons
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N5/00—Undifferentiated human, animal or plant cells, e.g. cell lines; Tissues; Cultivation or maintenance thereof; Culture media therefor
- C12N5/06—Animal cells or tissues; Human cells or tissues
- C12N5/0602—Vertebrate cells
- C12N5/0618—Cells of the nervous system
- C12N5/062—Sensory transducers, e.g. photoreceptors; Sensory neurons, e.g. for hearing, taste, smell, pH, touch, temperature, pain
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y301/00—Hydrolases acting on ester bonds (3.1)
- C12Y301/03—Phosphoric monoester hydrolases (3.1.3)
- C12Y301/03016—Phosphoprotein phosphatase (3.1.3.16), i.e. calcineurin
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/14—Type of nucleic acid interfering nucleic acids [NA]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/14—Type of nucleic acid interfering nucleic acids [NA]
- C12N2310/141—MicroRNAs, miRNAs
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/20—Type of nucleic acid involving clustered regularly interspaced short palindromic repeats [CRISPR]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2320/00—Applications; Uses
- C12N2320/30—Special therapeutic applications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2506/00—Differentiation of animal cells from one lineage to another; Differentiation of pluripotent cells
- C12N2506/08—Differentiation of animal cells from one lineage to another; Differentiation of pluripotent cells from cells of the nervous system
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2750/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA ssDNA viruses
- C12N2750/00011—Details
- C12N2750/14011—Parvoviridae
- C12N2750/14111—Dependovirus, e.g. adenoassociated viruses
- C12N2750/14141—Use of virus, viral particle or viral elements as a vector
- C12N2750/14143—Use of virus, viral particle or viral elements as a vector viral genome or elements thereof as genetic vector
Definitions
- the present disclosure relates to the field of biomedicine. More specifically, the present disclosure relates to RE1/NRSE elements as targets for transdifferentiation of non-neuronal cells into neuronal cells; and RE1/NRSE blockers in preventing and/or treating neuronal function loss or death associated with use in diseases.
- Repressor element 1/neuron-restrictive silencer element is a specific DNA sequence, about 21bp in length (ranging from 20-23bp), mainly related to REST (RE1 silencing Transcription factor (RE1-silencing transcription factor), which is also known as neuron-restrictive silencer factor (neuron-restrictive silencer factor, NRSF) binding, regulates the expression of genes related to neuronal development and maturation.
- RE1 is a negative regulatory element related to neuron maturation, which was first discovered at the 5' end of the promoters of NaV1.2 and SCG10, and regulates the expression of these genes.
- the RE1 site is bound by a silencing complex composed of histone deacetylases and methylases to inhibit the expression of neuron-related genes.
- a silencing complex composed of histone deacetylases and methylases to inhibit the expression of neuron-related genes.
- CRISPR-mediated gene regulation and epigenetic modification technology has very high precision and can precisely regulate the expression of specific genes, but it is difficult to regulate the expression of genes regulated by RE1 in this way.
- Parkinson's disease is a disease associated with loss of neuronal function or death and is characterized by the loss of dopamine neurons in the substantia nigra of the midbrain.
- the treatment methods for Parkinson's disease are mainly small molecule drugs represented by dopamine analogues such as levodopa.
- the surgical treatment developed in recent years can also improve the symptoms of the disease to a certain extent, such as deep brain electrode stimulation.
- these methods can only partially alleviate the disease, and cannot prevent the development of the disease, nor can it slow down the death of dopamine neurons.
- the concept of transdifferentiation therapy has brought hope to the regeneration of dopamine neurons.
- glial cells By overexpressing some genes in glial cells, or gene editing of glial cells, glial cells can be transdifferentiated into dopamine neurons to supplement missing or dead dopamine neurons.
- Müller glia MG is the main glial cell in retinal tissue.
- Retinal ganglion cell RRC is a nerve cell located in the innermost layer of the retina. Its dendrites mainly establish synaptic connections with bipolar cells, and its axons extend to the optic nerve head to form the optic nerve, extending to brain.
- Retinal ganglion cell (RGC) damage or degeneration is a major cause of permanent blindness.
- RGCs are the only output neurons in the retina, so RGC damage or degeneration will lead to permanent blindness. Reconstructing functional RGC is currently the only way to help blind patients restore vision, and RGC cells are very difficult to regenerate. Therefore, exploring how to regenerate RGC cells will bring hope to the majority of blind patients caused by RGC cell death.
- Photoreceptor cell death is the main cause of blindness. Photoreceptor cells are divided into rod cells and cone cells. There are many reasons for photoreceptor cell death, including hereditary blindness, acquired blindness and senile degenerative blindness. Common genetic blindnesses, such as retinitis pigmentosa and amaurosis, result from the death of photoreceptor cells in the retina. In the visual system, photoreceptor cells are the only cells that convert light signals into neural electrical signals. The photoreceptor cells establish a connection with downstream bipolar cells, and transmit the neural electrical signals to bipolar cells. Long-distance connections are established between cells, which continue to transmit neural signals downstream.
- Inner ear spiral ganglion cells are located in the inner ear spiral ganglion, one end of which is connected to the inner ear hair cells, and the other end is connected to the central nervous system. It is the only channel for transmitting auditory signals to the central nervous system. Inner ear spiral ganglion cells are a very special type of neurons, and their gene expression profiles are very different from ordinary neurons. Permanent deafness caused by the death of spiral ganglion cells in the inner ear, no matter whether it is hereditary or non-hereditary, there is currently no drug available for treatment. Due to the special structure of the inner ear spiral ganglion, stem cell transplantation technology has not yet made progress.
- NgN2 and NeuroD1 can induce neurons in vivo, they can only produce ordinary glutamatergic neurons, but cannot induce neurons with special functions, such as: dopamine neurons, serotonergic neurons, Cholinergic neurons, retinal ganglion cells, photoreceptor cells and cochlear spiral ganglion cells, etc.
- Ptbp1 can induce glial cells to transdifferentiate into a special type of neuron in vivo, and its type is dopamine neuron.
- the application provides a method for blocking RE1/NRSE elements to regulate the expression of neuron-related genes in non-neuronal cells, which includes reducing the combination of REST and RE1/NRSE elements, or reducing the amount or activity of REST .
- the amount of REST is reduced by methods such as gene editing, small RNA interference, or accelerated protein degradation.
- the amount of REST is reduced by methods such as gene editing, antisense oligonucleotides (ASOs), small RNA interference, miRNA technology, small molecule compounds, or accelerated protein degradation.
- ASOs antisense oligonucleotides
- small RNA interference small RNA interference
- miRNA technology small molecule compounds
- small molecule compounds small molecule compounds
- the REST inhibitory active region is removed by gene editing or the REST activity is reduced by an inhibitor of REST activity.
- the binding of REST to the RE1/NRSE element is blocked by binding of a REST-binding agent, such as a REST antibody, to REST.
- a REST-binding agent such as a REST antibody
- binding of REST to a RE1/NRSE element is blocked by binding of a RE1/NRSE element blocker to the RE1/NRSE element.
- the RE1/NRSE element blocking agent is a REST competitive binding protein, short peptide or gene editing protein or its encoding nucleic acid, or nucleic acid and nucleic acid analogs, or a small molecule RE1/NRSE element blocking agent agent.
- the RE1/NRSE element blocker is a REST variant or nucleic acid encoding it.
- the REST variant is the DNA binding domain of REST, which lacks the N-terminal and C-terminal repression domains of REST, preferably amino acids 155-420 of REST.
- the DNA binding domain of the REST is fused to an activation domain.
- the activation domain is selected from: epigenetic modification proteins or gene activation regulatory elements, such as VP64, P65-HSF1, VP16, RTA, Suntag, P300, CBP or combinations thereof, preferably VP64 or P65- HSF1.
- the REST variant has the amino acid sequence of SEQ ID NO: 1, 3, 5 and 9 or the nucleotide sequence of SEQ ID NO: 2, 4, 6 and 10, or at least 70% thereof , 60%, 50% percent identity sequences.
- the non-neuronal cells include, for example, glial cells, fibroblasts, stem cells, neural precursor cells, neural stem cells, wherein the glial cells are selected from the group consisting of astrocytes, oligodendrocytes, Ependymal cells, Schwann cells, NG2 cells, satellite cells, Müller glial cells, inner ear glial cells or combinations thereof, preferably astrocytes, Müller glial cells and cochlear glial cells.
- glial cells are selected from the group consisting of astrocytes, oligodendrocytes, Ependymal cells, Schwann cells, NG2 cells, satellite cells, Müller glial cells, inner ear glial cells or combinations thereof, preferably astrocytes, Müller glial cells and cochlear glial cells.
- the glial cells are derived from the brain, spinal cord, eyes or ears, wherein the glial cells in the brain are derived from the striatum, the substantia nigra, the ventral tegmental area of the midbrain, the spinal cord, the hypothalamus, the dorsal Lateral midbrain or cerebral cortex, preferably striatum and substantia nigra.
- neuronal cells are mammalian neurons, wherein preferably Dopamine neurons, GABA neurons, 5-HT neurons, glutamatergic neurons, ChAT neurons, NE neurons, motor neurons, spinal cord neurons, spinal motor neurons, spinal cord sensory neurons, photoreceptor cells ( rods and cones), bipolar cells, horizontal cells, amacrine cells, retinal ganglion cells (RGCs), cochlear neurons (cochlear spiral ganglion cells and vestibular neurons), pyramidal neurons, intermediary Neurons, medium spiny neurons (MSN), Purkinje cells, granule cells, olfactory sensory neurons, peribulbar cells or combinations thereof, more preferably dopamine neurons, retinal ganglion cells, photoreceptor cells and cochlear spiral ganglion cell.
- GABA neurons GABA neurons
- 5-HT neurons glutamatergic neurons
- ChAT neurons NE neurons
- motor neurons spinal cord neurons
- spinal motor neurons spinal motor neurons
- spinal cord sensory neurons spinal cord sensory neurons
- photoreceptor cells rods and cone
- the non-neuronal and/or neuronal cells are from, for example, humans, non-human primates, rats and mice, preferably humans.
- the present application provides a RE1/NRSE element blocker for the preparation of medicines for the prevention and/or treatment of neuron function loss or death-related diseases, wherein the RE1/NRSE element blocker
- the blocking agent reduces the binding of RE1/NRSE endogenous binding factors to RE1/NRSE elements, wherein the RE1/NRSE endogenous binding factors include zinc finger proteins such as REST.
- the RE1/NRSE element blocking agent binds to the RE1/NRSE element thereby blocking the binding of the RE1/NRSE endogenous binding factor to the RE1/NRSE element.
- the RE1/NRSE element blocking agent is a REST competitive binding protein, short peptide or gene editing protein or its encoding nucleic acid, or nucleic acid and nucleic acid analogs, or a small molecule RE1/NRSE element blocking agent agent.
- the RE1/NRSE element blocker is a REST variant or nucleic acid encoding it.
- the REST variant is the DNA binding domain of REST, which lacks the N-terminal and C-terminal repression domains of REST, preferably amino acids 155-420 of REST.
- the DNA binding domain of the REST is fused to an activation domain.
- the activation domain is selected from: epigenetic modification proteins or gene activation regulatory elements, such as VP64, P65-HSF1, VP16, RTA, Suntag, P300, CBP or combinations thereof, preferably VP64 or P65- HSF1.
- the REST variant has the amino acid sequence of SEQ ID NO: 1, 3, 5 and 9 or the nucleotide sequence of SEQ ID NO: 2, 4, 6 and 10, or at least 70% thereof , 60%, 50% percent identity sequences.
- the disease associated with loss of neuron function or death is selected from: Parkinson's disease, Alzheimer's disease, stroke (stroke), schizophrenia, Huntington's disease, depression, motor neuron Yuan disease, amyotrophic lateral sclerosis, spinal muscular atrophy, Pick disease, sleep disorders, epilepsy, ataxia, visual impairment due to RGC cell death, glaucoma, age-related RGC lesions, optic nerve damage, retinal ischemia or hemorrhage, Leber hereditary optic neuropathy, degeneration or death of photoreceptor cells due to damage or degeneration, macular degeneration, retinitis pigmentosa, diabetic-related blindness, night blindness, color blindness, hereditary blindness, amaurosis congenita, spiral nerve Deafness or hearing loss due to ganglion cell death, or a combination thereof.
- Parkinson's disease Alzheimer's disease
- stroke stroke
- schizophrenia Huntington's disease
- depression depression
- motor neuron Yuan disease amyotrophic lateral sclerosis
- the present application provides a RE1/NRSE element blocking agent, which is a REST variant or its encoding nucleic acid, or an artificially designed analog of the DNA binding domain of REST.
- the REST variant is the DNA binding domain of REST, which lacks the N-terminal and C-terminal repression domains of REST, preferably amino acids 155-420 of REST.
- the DNA binding domain of the REST is fused to an activation domain.
- the activation domain is selected from: epigenetic modification proteins or gene activation regulatory elements, such as VP64, P65-HSF1, VP16, RTA, Suntag, P300, CBP or combinations thereof, preferably VP64 or P65- HSF1.
- the REST variant has the amino acid sequence of SEQ ID NO: 1, 3, 5 and 9 or the nucleotide sequence of SEQ ID NO: 2, 4, 6 and 10, or at least 70% thereof , 60%, 50% percent identity sequences.
- the REST variant or the DNA binding domain of REST is from, for example, human, non-human primate, rat and mouse, preferably human.
- the present application provides a pharmaceutical composition or kit or kit, which comprises the above-mentioned blocking agent.
- the pharmaceutical composition or kit or kit is formulated for injection, intracranial administration, intraocular administration, intraaural administration, inhalation, parenteral administration, intravenous administration, intramuscular administration administration, intradermal administration, topical administration or oral administration.
- the pharmaceutical composition or kit or kit further comprises a carrier or carrier for delivering the RE1/NRSE element blocking agent, wherein the carrier or carrier is a viral vector, a lipid body, nanoparticle, exosome, virus-like particle, among which viral vectors include recombinant adeno-associated viral vectors (rAAV), adeno-associated viral (AAV) vectors, adenoviral vectors, lentiviral vectors, retroviral vectors, and poxviral vectors , herpes virus, SV40 virus vector, and combinations thereof, wherein AAV and rAAV are preferred.
- rAAV recombinant adeno-associated viral vectors
- AAV adeno-associated viral vectors
- AAV adenoviral vectors
- lentiviral vectors lentiviral vectors
- retroviral vectors retroviral vectors
- poxviral vectors herpes virus, SV40 virus vector, and combinations thereof, wherein AAV and rA
- the pharmaceutical composition or kit or kit comprises an expression vector for expressing a REST variant, wherein the expression vector comprises a nucleotide sequence encoding a REST variant in combination with the expression vector causing its expression
- the promoter is operably linked.
- the pharmaceutical composition or kit or kit is for topical administration to at least one of: 1) glial cells in the striatum; ii) glial cells in the substantia nigra of the brain; iii) ) glial cells in the retina; iv) glial cells in the inner ear; v) glial cells in the spinal cord; vi) glial cells in the prefrontal cortex; vii) glial cells in the motor cortex; viii) hypothalamus and ix) glial cells in the ventral tegmental area (VTA).
- VTA ventral tegmental area
- the pharmaceutical composition or kit or kit further comprises i) one or more dopamine neuron-related factors, or ii) expression of one or more dopamine neuron-related factors in Müller glial cells A retinal ganglion cell-associated factor,
- said one or more dopamine neuron related factors are selected from: FoxA2, Lmx1a, Lmx1b, Nurr1, Pbx1a, Pitx3, Gata2, Gata3, FGF8, BMP, En1, En2, PET1, Pax family protein (Pax3 , Pax6, etc.), SHH, Wnt family proteins and TGF- ⁇ family proteins, or combinations thereof;
- retinal ganglion cell-related factors include: ⁇ -catenin, Oct4, Sox2, Klf4, Crx, aCamKII, Brn3a, Brn3b, Brn3C, Math5, Otx2, Ngn2, Ngn1, AscL1, miRNA9 , miRNA-124, Nr2e3 and Nrl and other factors.
- the promoter is a glial cell-specific promoter or a Muller glial cell (MG) cell-specific promoter
- the glial cell-specific promoter is selected from the group consisting of GFAP promoter, ALDH1L1 promoter promoter, EAAT1/GLAST promoter, glutamine synthetase promoter, S100 ⁇ promoter EAAT2/GLT-1 promoter and Rlbp1 promoter, preferably GFAP promoter.
- the glial cells have a transdifferentiation efficiency of at least 1%, or at least 10%, 20%, 30%, 40%, or 50%.
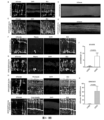
- FIG. 1 Schematic diagram of endogenous zinc finger proteins and suppression system design.
- A Schematic diagram of the REST protein structure.
- the REST protein includes an N-terminal repression domain, a DNA-binding structure in the middle responsible for binding to RE1, and a C-terminal transcription repression domain.
- RZFD-V1 represents the first design of RZFD, including 8 zinc finger domains (RZFD, REST Zinc Figure domain) in the middle of the human REST protein responsible for binding to RE1 DNA.
- RZFD-V2 and RZFD-V3 we fused RZFD with two different activators, and named them RZFD-V2 and RZFD-V3, respectively.
- RZFD-V2 is obtained by fusing the C-terminus of RZFD with VP64, and RZFD-V3 is composed of a P65 and a transcriptional activation domain of HSF1 fused to the C-terminus of RZFD.
- RZFDmax optimizes the codon of RZFD to increase its expression in mammals, and at the same time adds BPNLS sequences at both ends of RZFD to enhance its efficiency in nuclear entry and function.
- B In glial cells, REST binds to RE1, and the C-terminus and N-terminus of REST respectively recruit some transcriptional repressors, so that the neuron-related genes regulated by RE1 cannot be expressed.
- RZFD-V1 we imagined that after expressing RZFD-V1 in glial cells, RZFD would compete with RE1 for binding, preventing the binding of REST to RE1, so that the REST silencing complex could not be formed and the genes regulated by RE1 could not be inhibited, including neural Meta-associated gene expression.
- RZFD-V2 competitively binds to RE1 through the RZFD domain, preventing the binding of REST/NRSF to RE1, and its VP64 domain can recruit transcriptional activators to enhance the expression of neuron-related genes regulated by RE1.
- FIG. 1 Schematic diagram of dCas9-Krab or dCas9-3xKrab inhibiting the expression of target genes under the action of gRNA.
- I-K Schematic diagram of N-dCas9 or 3xN-dCas9 or 3xN-dCas9-3xC inhibiting target gene expression under the guidance of gRNA, wherein N represents the N-terminal inhibitory domain of REST, and C represents the C-terminal inhibitory domain of REST.
- L Transfection of HEK293T cells with various suppression systems inhibits the expression efficiency of Ptbp1 gene.
- the inhibition efficiency of dCas9-Krab, N-dCas9 and 3xN-dCas9 is relatively low, and the inhibition efficiency of dCas9-3xKrab and 3xN-dCas9-3xC is relatively high.
- FIG. 1 Schematic diagram of AAV vector design.
- Vector 1 is a vector schematic diagram of GFAP-driven expression of mCherry.
- GFAP is a promoter specifically expressed in glial cells.
- mCherry is a red fluorescent protein used to label glial cells.
- Vector 2 is a schematic diagram of the human RZFD expression vector, and the expression of RZFD is driven by the astrocyte-specific promoter GFAP.
- B Schematic diagram of injection and sample analysis. The day of AAV injection was recorded as day 0, and the results of transdifferentiation of glial cells into neurons were analyzed 2 weeks after injection, and the transdifferentiation of glial cells was analyzed at 1.5 months after injection.
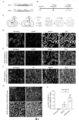
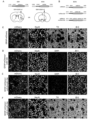
- FIG. 1 Schematic diagram of AAV virus injection and transdifferentiation. Inject GFAP-mCherry alone in the striatum or substantia nigra of mice, or inject a mixed AAV of GFAP-mCherry and GFAP-RZFD. GFAP-mCherry will label glial cells as Red, while GFAP-RZFD transdifferentiates glial cells into neuronal neurons.
- FIG. 1 Shows indicate neurons expressing both mCherry and TH.
- FIG. 1 About 1.5 months after injection of mixed AAV of GFAP-mCherry and GFAP-RZFDmax in the striatum of mice, GFAP-mCherry+GFAP-RZFDmax promoted the transdifferentiation of astrocytes in the striatum of mice is a dopamine neuron, and TH is a specific marker of dopamine. Scale bar is 50 microns.
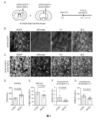
- FIG. 3 RZFD transdifferentiates astrocytes into dopamine neurons in DAT-Cre:Ai9 mice modeled with 6-OHDA.
- A Schematic diagram of AAV injection.
- GFAP-EGFP is the labeled AAV, which specifically labels astrocytes under the GFAP promoter.
- GFAP-EGFP mixed with GFAP-RZFD was injected into the striatum or substantia nigra of mice to induce the transdifferentiation of astrocytes into dopamine neurons.
- the green fluorescent signal is the astrocytes specifically labeled by GFAP-EGFP, which still presents a typical astrocyte morphology, and there is no tdTomato signal that specifically labels dopamine neurons, and There are no TH-positive cells, and the yellow arrows point to EGFP-positive astrocytes.
- C GFAP-EGFP and GFAP-RZFD mixed injection group, almost all green fluorescent labeled cells have neuron morphology, and tdTomato signal appears in striatum, and tdTomato signal is co-labeled with TH signal, the white arrow points to TH Positive tdTomato red cells. Scale bar is 50 microns.
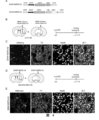
- FIG. 4 Transdifferentiation of glial cells into neurons mediated by RZFD-VP64 or RZFD-P65-HSF1.
- A Schematic diagram of the design of GFAP-RZFD-V2 and GFAP-RZFD-V3 expression vectors.
- RZFD-V2 is a fusion protein of RZFD and VP64, and its expression is driven by the glial cell-specific promoter GFAP.
- RZFD-V3 is composed of RZFD and P65-HSF1 activation domain.
- FIG. 1 Representative diagram of GFAP-RZFD-V2 transdifferentiation of glial cells into neurons or dopamine neurons in the striatum
- mCherry is the cell labeled by GFAP-mCherry
- TH is the specific marker of dopamine neurons
- NeuN is a neuron-specific marker. Arrows indicate neurons expressing both mCherry and TH.
- FIG. D Schematic diagram of injecting GFAP-RZFD-V3 in DAT-Cre:Ai9 mice, GFAP-RZFD-V3 was injected into the striatum or substantia nigra of DAT-Cre:Ai9 mice, and the samples were analyzed after 1.5 months, Dat -Cre is Cre inserted behind the Dat endogenous promoter, Ai9 is Rosa26-CAG-LSL-tdTomato-WPRE mice, DAT-Cre: only mature dopamine neurons in the brain of Ai9 mice can be labeled by tdTomato red fluorescent signal superior.
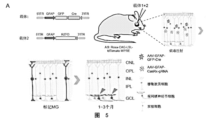
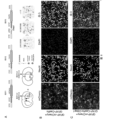
- FIG. 5 Overexpression of GFAP-RZFD in the retina transdifferentiates Müller glia cells into photoreceptor cells or retinal ganglion cells.
- A Schematic diagram of the experimental process of overexpressing GFAP-RZFD to transdifferentiate Müller glial cells into photoreceptor cells or retinal ganglion cells, in which the carrier 1 is GFAP-EGFP-2A-Cre, which is injected into the retina of Ai9 mice and can be used for Specifically label muller glial cells in mice, and the labeled cells will be labeled with tdTomato.
- B After the control virus GFAP-EGFP-2A-Cre was injected into the retina of Ai9 mice, the red fluorescent signal was specifically expressed in Müller glial cells.
- FIG. Overexpression of miRNA can transdifferentiate mouse astrocytes into neurons but not dopamine neurons.
- A Schematic illustration of astrocyte-to-neuron transition achieved by overexpressing miRNAs.
- Vector 1 is AAV-GFAP-mCherry, and vector 2 (AAV-GFAP-miRNA) is driven by GFAP to overexpress miRNA.
- Vector 1 was injected alone as the control group, and vector 1+2 was injected as the experimental group.
- B Corresponds to carrier 2 in Figure A.
- miR-124 (comprising miR-124-5p and miR-124-3p), miR-9 (comprising miR-9-5p and miR-9-3p) or miR-9-miR-124 driven by GFAP promoter, respectively Schematic diagram of the AAV expression vector.
- C Injection of GFAP-mCherry in the striatum of mice, mCherry is specifically expressed in glial cells, and there are no TH positive cells in the striatum.
- GFAP-mCherry+GFAP-miR-9+miR-124 was injected into the striatum of mice, TH is a specific marker of dopamine neurons, the white arrow points to the marked cells, and the overlay shows red fluorescence and TH signals are non-overlapping, and the bar is 50 ⁇ m.
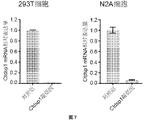
- CasRx can knock down the mRNA expression of Ctdsp1 in vitro.
- Expression of CasRx and a gRNA targeting Ctdsp1 mRNA in human 293T and mouse N2A cell lines in vitro achieved mRNA knockdown of Ctdsp1.
- the full name of Ctdsp1 is Carboxy-terminal domain RNA polymerase II polypeptide A small phosphatase 1 (Ctdsp1).
- FIG. 8 Knockdown of Ctdsp1 can transdifferentiate mouse astrocytes into neurons but not dopamine neurons.
- A Schematic representation of astrocyte-to-neuron conversion via gene knockdown.
- Vector 1 AAV-GFAP-mCherry
- vector 2 AAV-GFAP-CasRx
- Vector 3 AAV-GFAP-CasRx-Ctdsp1 encodes CasRx and a gRNA targeting Ctdsp1.
- AAV-GFAP-CasRx-Ctdsp1 or control vectors AAV-GFAP-CasRx and AAV-GFAP-mCherry without gRNA were injected into the striatum or substantia nigra of C57BL/6 mice.
- AAV-GFAP-mCherry co-injection was used to specifically label astrocytes or neurons transformed from astrocytes. The occurrence of transformation was checked 1-2 months after the injection.
- Carrier 1+2 is the control group
- carrier 1+3 is the test group.
- FIG. 9 Knockdown of Ctdsp1 or overexpression of miRNA fails to generate retinal ganglion cells, photoreceptor cells, or cochlear spiral ganglion cells.
- vector 1 GFAP-EGFP-2A-Cre
- vector 2 GFAP-CasRx
- Vector 3 GFAP-CasRx-Ctdsp1 encodes CasRx and a gRNA targeting Ctdsp1.
- FIG. 1 Schematic representation of miRNA overexpression in retina.
- Vector 1 is GFAP-EGFP-2A-Cre, and vector 2 (GFAP-miRNA) is driven by GFAP to overexpress miRNA.
- Vector 1 was injected alone as the control group, and vector 1+2 was injected as the experimental group.
- D Corresponds to vector 2 in Figure C. Schematic diagram of AAV expression vectors driven by GFAP promoter to express miR-124, miR-9 or miR-9-miR-124, respectively.
- the present application provides compositions related to RE1/NRSE regulation, biologically active molecules modified based on different domains of endogenous RE1/NRSE binding protein REST and applications thereof.
- this application relates to the modification of the endogenous RE1-binding protein REST, so as to utilize the function of different regions in the REST protein to regulate gene expression.
- Repressor element 1/neuron-restrictive silencer element is a specific DNA sequence, about 21bp in length (ranging from 20-23bp), present in many nerve-related genes the promoter region.
- RE1/NRSE and RE1 are used interchangeably.
- RE1 is a negative regulatory element related to neuron maturation, which was first discovered at the 5' end of the promoters of NaV1.2 and SCG10, and regulates the expression of these genes. In non-neuronal cells, the RE1 site is bound by a silencing complex composed of histone deacetylases and methylases to inhibit the expression of neuron-related genes. There are more than 1800 RE1 elements in mice and humans.
- RE1 mainly binds to REST (RE1-silencing transcription factor) to regulate the expression of genes related to neuronal development and maturation.
- REST also known as neuron-restrictive silencer factor (NRSF)
- NRSF neuron-restrictive silencer factor
- human REST protein No. 159-412 contains 8 zinc finger protein domains (Zinc Finger Domain, ZFD) ( Figure 1A), in its N- The 1-83 position of the terminal is its N-terminal inhibitory region, which mainly binds to proteins such as Sin3a and Sin3b, and the 1008-1097th position is its C-terminal inhibitory domain and a zinc finger domain, which mainly binds to proteins such as RCOR1.
- ZFD Zinc Finger Domain
- the zinc finger domain in the REST protein may be related to its binding to RE1, thereby allowing the transcriptional repression domain in the REST protein (thought to exist at the N-terminal or C-terminal of the REST protein) to regulate the transcription of RE1, thereby inhibiting the RE1 target gene expression.
- Previous studies have shown that deleting positions 1-83 and 1008-1097 of the REST protein does not affect the binding of REST to RE1, but cannot function normally.
- the present application provides methods for transdifferentiating non-neuronal cells into functional neuronal cells in an individual by modulating RE1.
- by blocking REST binding to RE1/NRSE elements at a site of interest in vivo eg, a site affected by a disease
- transdifferentiation of non-neuronal cells into functional neuronal cells can be achieved at a site of interest. Yuan.
- the inventors of the present application found that the N-terminal and C-terminal of the REST protein can recruit various epigenetic regulatory elements, and then negatively regulate the genes they act on.
- the DNA-binding protein is combined or fused with the N-terminal and/or C-terminal of the REST protein, negative regulation can be achieved on the target gene region to which the DNA-binding protein binds.
- the present application provides a method of transdifferentiating non-neuronal cells into functional neurons in an individual.
- the present application provides a method of preventing and/or treating a disease associated with loss of neuronal function or death in an individual in need thereof, the method comprising displacing non-neuronal cells at a site affected by the disease Transdifferentiated into functional neurons.
- a functional neuron refers to a neuron cell capable of specific functions, such as dopamine neurons, retinal ganglion cells, photoreceptor cells and other neurons with specific functions. .
- a functional neuron has at least one morphological characteristic of a neuron, eg, has synapses, eg, axons.
- the functional neuron expresses at least one marker of a mature neuron, such as a NeuN gene expression product.
- a functional neuron has electrophysiological properties.
- Functional neurons can have different functions.
- the functional neurons include dopamine neurons, retinal ganglion cells, photoreceptor cells and cochlear spiral ganglion cells, GABA neurons, 5-HT neurons, glutamatergic neurons, ChAT neurons neurons, NE neurons, motor neurons, spinal cord neurons, spinal motor neurons, spinal cord sensory neurons, bipolar cells, horizontal cells, amacrine cells, pyramidal neurons, interneurons, medium spiny neurons (MSN), Purkinje cells, granule cells, olfactory sensory neurons, periglobular cells, or any combination thereof.
- the functional neuron expresses the NeuN gene.
- the NeuN gene is a known specific marker of mature neurons. Detection of NeuN gene expression products (such as NeuN protein) in non-neuronal cells indicates that non-neuronal cells are transdifferentiated into functional neurons.
- the functional neuron has an axon.
- the axons of neurons can be observed through a microscope.
- the functional neurons comprise dopamine neurons, retinal ganglion cells, photoreceptor cells, or cochlear spiral ganglion cells.
- the functional neurons comprise dopamine neurons.
- dopamine neuron and dopaminergic neuron are used interchangeably.
- Dopaminergic neurons are neurons that contain and release dopamine (DA) as a neurotransmitter.
- Dopaminergic neurons are the major source of dopamine in the central nervous system.
- Dopamine belongs to catecholamine neurotransmitters, which can affect neural functions such as emotion and reward, and play an important biological role in the central nervous system.
- Dopaminergic neurons in the brain are mainly concentrated in the substantria nigra pars compacta (SNc), ventral tegmental area (VTA), hypothalamus, and periventricular areas of the midbrain.
- SNc substantria nigra pars compacta
- VTA ventral tegmental area
- hypothalamus hypothalamus
- periventricular areas of the midbrain periventricular areas of the midbrain.
- the dopamine neuron expresses one or more markers selected from the group consisting of tyrosine hydroxylase (TH), FoxA2, Nurrl, Pitx3, Vmat2, and DAT.
- a "marker” in this application may refer to an expression product of a gene, such as mRNA or protein. Detection of the expression of one or more of these markers in functional neuron cells indicates that the functional neuron cells are dopamine neurons. Exemplary gene sequences and protein sequences of these markers are well known in the art, and can be queried through public databases (such as the gene database and protein database of the National Center for Bioinformatics (NCBI) under the National Institutes of Health), in this application are listed in Table A.
- NCBI National Center for Bioinformatics
- Tyrosine hydroxylase is an enzyme responsible for catalyzing the conversion of the amino acid L-tyrosine into dihydroxyphenylalanine (dopa), and is an enzyme involved in the synthesis and metabolism of dopamine in dopaminergic neurons.
- Other markers of dopamine neurons include FoxA2, Nurr1, Vmat2 and DAT, etc.
- the dopamine neuron expresses NeuN, TH, and DAT.
- the functional neurons comprise retinal ganglion cells.
- Retinal ganglion cells are a type of neuron located near the inner surface of the retina (the ganglion cell layer) that receive visual information from photoreceptors via two types of interneurons, bipolar cells and amacrine cells. Its dendrites mainly establish synaptic connections with bipolar cells, and its axons extend to the optic nerve head, forming the optic nerve, which extends to the brain.
- the retinal ganglion cells express one or more markers selected from the group consisting of RBPMS, Pax6, Brn3a, Brn3b, Brn3c, and Map2.
- RBPMS is a specific marker of RGCs. If the expression of RBPMS is detected in functional neuron cells, it is suggested that the functional neuron cells are RGCs. Exemplary gene sequences and protein sequences of RBPMS are well known in the art, can be queried through public databases, and are listed in Table A in this application.
- the retinal ganglion cells (RGCs) express NeuN and RBPMS.
- the functional neurons comprise photoreceptor cells.
- Photoreceptor cells are specialized neuroepithelial cells found in the retina that have the function of sensing light and performing phototransduction functions. It can be processed by bipolar cells and ganglion cells to convert light signals into electrical signals and transmit them to the brain. Photoreceptor cells include rods and cones.
- the photoreceptor cells express one or more markers selected from the group consisting of Rhodopsin, mCAR, m-opsin, and S-opsin.
- Rhodopsin, mCAR, m-opsin, and S-opsin are all specific markers of RGCs. Detecting the expression of Rhodopsin, mCAR, m-opsi and/or S-opsin in functional neuron cells indicates that the functional neuron cells are photoreceptor cells.
- Exemplary gene sequences and protein sequences of Rhodopsin, mCAR, m-opsi and S-opsin are well known in the art, can be queried through public databases, and are listed in Table A in this application.
- the photoreceptor cells express NeuN, Rhodopsin and/or mCAR.
- the functional neurons comprise cochlear spiral ganglion cells.
- the cochlear spiral ganglion cell is a kind of bipolar ganglion cell, which is the first-order neuron of the auditory conduction pathway. Its peripheral process is connected with hair cells, and its central process participates in the formation of the auditory nerve. Spiral ganglion cells play an important role in the transmission and encoding of sound signals.
- the cochlear spiral ganglion cells express one or more markers selected from the group consisting of NeuN, Prox1, Tuj-1, and Map2.
- the expression of Prox1 and Map2 detected in functional neuron cells indicates that the functional neuron cells are cochlear spiral ganglion cells.
- Exemplary gene sequences and protein sequences of Prox1, Tuj-1, and Map2 are well known in the art, can be queried through public databases, and are listed in Table A in this application.
- the cochlear spiral ganglion cells express NeuN, Prox1, Tuj-1, and Map2.
- the non-neuronal cells include glial cells (eg, glial cells), fibroblasts, stem cells, neural precursor cells, or neural stem cells. In some embodiments, the non-neuronal cells comprise glial cells (eg, glial cells).
- the glial cells are selected from astrocytes, oligodendrocytes, ependymal cells, Schwann cells, NG2 cells, satellite cells, Müller glia cells, inner ear glia cells and any combination thereof.
- Müller glia MG is the main glial cell in retinal tissue.
- the glial cells are located in the brain, spinal cord, eye or ear. In some embodiments, the glial cells are located in the striatum, substantia nigra, ventral tegmental area of the midbrain, medulla oblongata, hypothalamus, dorsal midbrain, or cerebral cortex of the brain.
- the active substance is administered topically to glial cells in one or more of the following sites in the subject: 1) glial cells in the striatum; ii) glial cells in the substantia nigra of the brain iii) glial cells in the retina; iv) glial cells in the inner ear; v) glial cells in the spinal cord; vi) glial cells in the prefrontal cortex; vii) glial cells in the motor cortex; viii ) glial cells in the hypothalamus; and ix) glial cells in the ventral tegmental area (VTA).
- VTA ventral tegmental area
- the microenvironment of glial cells in the body helps to promote the transdifferentiation induced by the active substances described in the application, and promotes the transdifferentiation of glial cells into functional cells in vivo. Neurons.
- the glial cells are selected from astrocytes, Müller glia, and cochlear glia.
- the glial cells comprise astrocytes and the functional neurons comprise dopamine neurons.
- the methods provided herein relate to methods of transdifferentiating astrocytes into dopamine neuronal cells in an individual.
- the astrocytes are located in the striatum and/or the substantia nigra.
- the method comprises administering an active substance provided herein to the individual's striatum and/or substantia nigra.
- the glial cells comprise Muller glial cells and the functional neuronal cells comprise retinal ganglion cells (RGCs) and/or photoreceptor cells.
- the methods provided herein relate to methods of transdifferentiating Muller glial cells into retinal ganglion cells (RGCs) and/or photoreceptor cells in an individual.
- the Müller glial cells are located under the retina or in the vitreous cavity.
- the method comprises administering an active substance provided herein to the subretinal or vitreous cavity of the individual.
- the glial cells comprise cochlear glial cells and the functional neuronal cells comprise cochlear spiral ganglion cells.
- the methods provided herein relate to methods of transdifferentiating cochlear glial cells into cochlear spiral ganglion cells in an individual.
- the cochlear glial cells are located in the inner ear.
- the method comprises administering an active agent provided herein to the inner ear of the individual.
- the transdifferentiation efficiency of the glial cells into functional neurons achieved after the administration of the active substance is at least 1%, or at least 10%, 20%, 30%, 40% or 50%. %.
- Transdifferentiation efficiency can be detected and calculated by methods known to those skilled in the art.
- fluorescent proteins can be used to label the initial cells of transdifferentiation (such as glial cells), such as GFAP-mCherry, GFAP-tdTomato, GFAP-EGFP, etc.
- Ai9 transgenic mice can also be used, in which the glial cells have fluorescent labeling.
- the transdifferentiation efficiency can be calculated by calculating the percentage of the number of transdifferentiated cells to the number of initially labeled cells.
- the transdifferentiation efficiency can also be calculated as a percentage of the number of cells produced by transdifferentiation compared to the number of cells of this type at the site of administration, for example, in the substantia nigra, the percentage of newly generated dopamine neurons to dopamine neurons in the substantia nigra .
- the method provided herein for transdifferentiating non-neuronal cells into functional neurons in an individual comprises administering to the individual an amount capable of reducing the binding of REST to RE1/NRSE elements, or reducing the amount of REST or active active substances.
- the method for preventing and/or treating diseases related to neuronal function loss or death in an individual in need thereof comprising administering a therapeutically effective A dose of an active substance capable of reducing the binding of REST to the RE1/NRSE element, or reducing the amount or activity of REST, to transdifferentiate non-neuronal cells into functional neurons at the site affected by the disease.
- the active substance is capable of reducing the amount or activity of REST. Any active substance that reduces the amount or activity of REST can be used.
- the amount of REST is reduced by methods such as gene editing, small RNA interference, or accelerated protein degradation.
- the amount of REST is reduced by methods such as gene editing, antisense oligonucleotides (Antisense Oligonucleotide, ASO), small RNA interference, miRNA technology, small molecular compounds, or accelerated protein degradation.
- the REST inhibitory active region is removed by gene editing or the REST activity is reduced by an inhibitor of REST activity.
- the active agent is capable of reducing REST binding to the RE1/NRSE element.
- the binding of REST to the RE1/NRSE element is blocked by binding of a REST-binding agent, such as a REST antibody, to REST.
- the active substance includes a RE1/NRSE element blocking agent, which can bind to the RE1/NRSE element to block the binding of REST to the RE1/NRSE element.
- the RE1/NRSE element blocker comprises a small molecule compound, nucleic acid, or nucleic acid analog that competes with REST for binding to RE1.
- the RE1/NRSE element blocker comprises a protein or a nucleic acid encoding the protein that competes with REST for binding to RE1.
- the protein that competes with REST for binding to RE1 comprises a REST variant.
- "Variant” in this application refers to a derivative sequence having one or more substitutions (including but not limited to conservative substitutions), additions, deletions, insertions or truncations, or any combination thereof, of one or more amino acid residues compared to the parent sequence .
- a REST variant may comprise a fragment of a REST protein, or a fusion protein of a fragment of a REST protein with another protein.
- the REST variant comprises the DNA binding domain of REST but lacks the N-terminal and/or C-terminal repression domain of REST.
- Native REST proteins contain an N-terminal repression domain, an intermediate DNA-binding domain responsible for binding to DNA, and a C-terminal transcription repression domain.
- the DNA binding domain of REST may be the 8 zinc finger domains in the REST protein or fragments thereof capable of binding DNA (for example, zinc finger domains with less than 8 zinc finger domains).
- the REST variant comprises amino acids 155-420 of REST (especially human REST), but lacks the N-terminal and/or C-terminal inhibitory domain of REST. In some embodiments, the REST variant comprises a RE1 binding fragment in amino acids 155-420 of REST (especially human REST), but lacks the N-terminal and/or C-terminal inhibitory domain of REST.
- RE1-binding fragment refers to a protein fragment capable of binding RE1 element in the present application.
- the REST variant has the amino acid sequence of SEQ ID NO: 1, 3, 5 or 9, or a sequence having at least 70%, 60%, or 50% percent identity thereto.
- the RE1/NRSE element blocker comprises nucleic acid encoding the REST variant, the nucleic acid encoding the REST variant has a nucleoside of SEQ ID NO: 2, 4, 6 or 10 acid sequence, or a sequence having at least 70%, 60%, or 50% identity thereto.
- percentage (%) of sequence identity with respect to amino acid sequences (or nucleic acid sequences) is defined as the number of identical amino acids (or nucleic acids) in a candidate sequence with a reference sequence after aligning the sequences and introducing gaps, if necessary, to achieve the maximum number of identical amino acids (or nucleic acids). The percentage of amino acid (or nucleic acid) residues in which the amino acid (or nucleic acid) residues are identical.
- the percent sequence identity (%) of an amino acid sequence (or nucleic acid sequence) can be calculated by dividing the number of amino acid residues (or bases) that are identical to the reference sequence to which it is compared by the number of amino acid residues (or bases) in the candidate sequence or reference sequence The total number of residues (or bases), whichever is shorter. Conservative substitutions of amino acid residues may or may not be considered identical residues. Alignment for the purpose of determining percent amino acid (or nucleic acid) sequence identity can be achieved, for example, using publicly available tools such as BLASTN, BLASTp (found at the U.S. National Center for Biotechnology Information ; NCBI), see also Altschul S.F.
- the nucleic acid encoding the REST variant is codon-optimized, optionally having the nucleotide sequence of SEQ ID NO: 15, or at least 70%, 60%, or 50% thereof Sequences with percent identity. Without being bound by theory, it is believed that the nucleic acid sequence of the codon-optimized REST variant is capable of higher expression in human cells.
- the REST variant further comprises an activation domain fused to the REST DNA binding domain.
- activation domain refers to a domain capable of interacting with the regulatory sequence of a target gene and activating or increasing the expression of the target gene.
- the activation domain comprises an epigenetic modification protein or a gene activation regulatory element, optionally, the activation domain comprises VP64, P65-HSF1, VP16, RTA, Suntag, P300, CBP or its In any combination, optionally, the activation domain includes VP64 or P65-HSF1.
- the REST variant is fused to one or more nuclear localization signal sequences (ie, NLS).
- NLS nuclear localization signal sequences
- the nuclear localization signal sequence can promote the REST variant to enter the nucleus, thereby better regulating gene expression and cell transdifferentiation. Any suitable nuclear localization signal sequence can be used.
- nuclear localization signal sequences include, but are not limited to, BPNLS (e.g., the amino acid sequence shown in SEQ ID NO: 13), the NLS of the SV40 viral large T antigen, with the amino acid sequence PKKKRKV (SEQ ID NO: 41); from NLS of a nucleoplasmic protein (e.g., a nucleoplasmic protein bipartite NLS with the sequence KRPAATKKAGQAKKKK) (SEQ ID NO: 42); a c-myc NLS with the amino acid sequence PAAKRVKLD (SEQ ID NO: 43) or RQRRNELKRSP (SEQ ID NO: 44 ); hRNPA1M9NLS with sequence NQSSNFGPMKGGNFGGRSSGPYGGGGQYFAKPRNQGGY (SEQ ID NO: 45); sequence RMRIZFKNKGKDTAELRRRRVEVSVELRKAKKDEQILKRRNV (SEQ ID NO: 46) from the IBB domain of importin-
- At least one of said nuclear localization signal sequences is fused to the N-terminus of said REST variant.
- At least one of said nuclear localization signal sequences is fused to the C-terminus of said REST variant.
- At least one of said nuclear localization signal sequences is fused to the N-terminus and C-terminus of said REST variant, respectively.
- the nuclear localization signal sequence comprises an amino acid sequence sequence selected from the group consisting of SEQ ID Nos: 13, and 41-58.
- the inventors performed a series of analyzes and modifications on the endogenous zinc finger domain of REST by using protein structure prediction and genetic engineering technology.
- the inventors found that overexpressing amino acids 155-420 of REST (containing only 8 zinc finger domains, called RZFD: REST Zinc Finger Domain), can block the combination of REST and RE1.
- RZFD REST Zinc Finger Domain
- Overexpression of RZFD in mouse striatal astrocytes can transdifferentiate glial cells into functional neurons by AAV-mediated gene delivery technology.
- RZFD-P65-HSF1 transdifferentiation domains of P65 and HSF1
- AAV-mediated in vivo transdifferentiation technology we found that RZFD-P65-HSF1 can also transdifferentiate glial cells into functional neurons in the striatum.
- REST which binds to a DNA sequence in the genome called RE1
- RE1 is a type of sequence that cannot be targeted by CRISPR technology.
- RZFD human endogenous zinc finger structure
- RZFD-V1 activation domains
- RZFD-V3 P65-HSF1
- RZFD, RZFD-VP64 and RZFD-P65-HSF1 could transdifferentiate Müller cells into retinal ganglion cells, and some photoreceptor cells were observed.
- Retinal ganglion cells are the only cells in the visual pathway that carry visual signals to the brain, and their absence or death can lead to permanent blindness.
- RZFD RZFD-VP64 and RZFD-P65-HSF1 can transdifferentiate glial cells into dopamine neurons
- RZFD-VP64 and RZFD-P65-HSF1 can transdifferentiate Müller cells into retinal ganglion cells and photoreceptor cells.
- the individual is a human or an animal.
- the animal is a non-human primate (eg, monkey), rat or mouse.
- diseases related to loss of function or death of neurons mainly include diseases related to loss of function or death of dopamine neurons, and visual impairment related to loss or death of optic ganglion or photoreceptor cells.
- the disease associated with loss of neuron function or death is selected from: Parkinson's disease, Alzheimer's disease, stroke (stroke), schizophrenia, Huntington's disease, depression, motor neuron Yuan disease, amyotrophic lateral sclerosis, spinal muscular atrophy, Pick disease, sleep disorders, epilepsy, ataxia, visual impairment due to RGC cell death, glaucoma, age-related RGC lesions, optic nerve damage, retinal ischemia or hemorrhage, Leber hereditary optic neuropathy, degeneration or death of photoreceptor cells due to damage or degeneration, macular degeneration, retinitis pigmentosa, diabetic-related blindness, night blindness, color blindness, hereditary blindness, amaurosis congenita, spiral nerve Deafness or hearing loss due
- the present application provides a method for preventing and/or treating diseases related to loss of neuron function or death in an individual in need thereof, comprising administering a therapeutically effective dose to the striatum and/or substantia nigra of the individual
- the active substances provided by the present application are used to transdifferentiate the astrocytes of the striatum and/or substantia nigra into dopamine neurons, wherein the diseases related to the loss of neuron function or death are selected from the following Group: Parkinson's disease depression and Alzheimer's disease.
- the present application provides a method for preventing and/or treating diseases related to neuron function loss or death in an individual in need thereof, comprising administering a therapeutically effective dose of the present application to the subretinal or vitreous cavity of the individual.
- RGC retinal ganglion cells
- the present application provides a method for preventing and/or treating diseases related to neuron function loss or death in an individual in need thereof, comprising administering a therapeutically effective dose of the active substance provided by the application to the inner ear of the individual , to transdifferentiate the cochlear glial cells of the inner ear into cochlear spiral ganglion cells, wherein the disease associated with loss of neuron function or death is selected from the group consisting of: deafness or hearing loss caused by spiral ganglion cell death .
- the application provides a REST variant comprising the DNA binding domain of REST but lacking the N-terminal and/or C-terminal repression domain of REST.
- REST in this application refers to native or endogenous REST molecules, including REST of any species.
- the REST variant comprises amino acids 155-420 of REST, but lacks the N-terminal and/or C-terminal repression domain of REST. In some embodiments, the REST variant comprises a RE1 binding fragment in amino acids 155-420 of REST (especially human REST), but lacks the N-terminal and/or C-terminal inhibitory domain of REST.
- the REST variant has the amino acid sequence of SEQ ID NO: 1, 3, 5 or 9, or a sequence having at least 70%, 60%, or 50% percent identity thereto.
- the RE1/NRSE element blocker comprises nucleic acid encoding the REST variant, the nucleic acid encoding the REST variant has a nucleoside of SEQ ID NO: 2, 4, 6 or 10 acid sequence, or a sequence having a percent identity thereto of at least 70%, 60%, or 50%.
- the REST variant further comprises an activation domain fused to the REST DNA binding domain.
- the activation domain comprises an epigenetic modification protein or a gene activation regulatory element, optionally, the activation domain comprises VP64, P65-HSF1, VP16, RTA, Suntag, P300, CBP or its In any combination, optionally, the activation domain includes VP64 or P65-HSF1.
- the REST variant is fused to one or more nuclear localization signal sequences.
- At least one of said nuclear localization signal sequences is fused to the N-terminus of said REST variant.
- At least one of said nuclear localization signal sequences is fused to the C-terminus of said REST variant.
- At least one of said nuclear localization signal sequences is fused to the N-terminus and C-terminus of said REST variant, respectively.
- the nuclear localization signal sequence comprises the amino acid sequence shown in SEQ ID NO: 13.
- the present application provides a polynucleotide comprising a nucleic acid sequence encoding a REST variant as described in the present application.
- the present application provides an expression vector comprising a polynucleotide encoding a REST variant, and optionally further comprising a promoter operably linked to the polynucleotide.
- the promoter is a glial cell-specific promoter.
- the glial cell-specific promoter is an astrocyte-specific promoter or a Muller glia (MG) cell-specific promoter.
- the glial cell-specific promoter is selected from GFAP promoter, ALDH1L1 promoter, EAAT1/GLAST promoter, glutamine synthetase promoter, S100 ⁇ promoter EAAT2/GLT-1 promoter and Rlbp1 promoter, preferably GFAP promoter.
- the glial cell-specific promoter is a cochlear glial cell-specific promoter.
- the cochlear glial cell-specific promoter is selected from the group consisting of: GFAP promoter (e.g., see SEQ ID NO: 39 or 40), ALDH1L1 promoter, EAAT1/GLAST promoter, and Plp1 promoter .
- the application provides a pharmaceutical composition comprising a REST variant as described in the application, or a polynucleotide encoding the REST variant, or a polynucleotide comprising a REST variant encoding Expression vectors, and pharmaceutically acceptable vectors.
- the pharmaceutical composition further comprises a carrier for delivering the polynucleotide, wherein the carrier comprises a viral vector, liposome, nanoparticle, exosome, or virus-like particle.
- the viral vectors include recombinant adeno-associated viral vectors (rAAV), adeno-associated viral (AAV) vectors, adenoviral vectors, lentiviral vectors, retroviral vectors, poxvirus vectors, herpesviruses, SV40 viruses vector, or any combination thereof, among which AAV or rAAV is preferred.
- rAAV recombinant adeno-associated viral vectors
- AAV adeno-associated viral vectors
- AAV adeno-associated viral vectors
- adenoviral vectors adenoviral vectors
- lentiviral vectors lentiviral vectors
- retroviral vectors retroviral vectors
- poxvirus vectors poxvirus vectors
- herpesviruses herpesviruses
- SV40 viruses vector or any combination thereof, among which AAV or rAAV is preferred.
- the pharmaceutical composition is suitable for local administration to glial cells in one or more of the following sites: 1) glial cells in the striatum; ii) glial cells in the substantia nigra of the brain; iii) glial cells in the retina; iv) glial cells in the inner ear; v) glial cells in the spinal cord; vi) glial cells in the prefrontal cortex; vii) glial cells in the motor cortex; viii) lower Glial cells in the thalamus; and ix) Glial cells in the ventral tegmental area (VTA).
- VTA ventral tegmental area
- the pharmaceutical composition is suitable for intracranial or intraocular administration.
- the pharmaceutical composition further comprises i) one or more dopamine neuron-related factors, or ii) expression of one or more retinal ganglion cell-related factors in Müller glial cells factor,
- the one or more dopamine neuron-related factors are selected from the group consisting of: FoxA2, Lmx1a, Lmx1b, Nurr1, Pbx1a, Pitx3, Gata2, Gata3, FGF8, BMP, En1, En2, PET1, Pax family proteins (Pax3, Pax6, etc.), SHH, Wnt family proteins, TGF- ⁇ family proteins, and any combination thereof;
- retinal ganglion cell-related factors include: ⁇ -catenin, Oct4, Sox2, Klf4, Crx, aCamKII, Brn3a, Brn3b, Brn3C, Math5, Otx2, Ngn2, Ngn1, AscL1, miRNA9, miRNA-124, Nr2e3, Nrl, and any combination thereof.
- the present application provides a kit or kit, which comprises the pharmaceutical composition provided in the present application.
- the present application provides a fusion protein comprising a DNA binding protein fused to one or more REST inhibitory domains.
- the DNA binding protein can bind to the target DNA to be regulated, and allow the REST inhibitory domain fused with it to negatively regulate the target DNA to be regulated, so as to inhibit the transcriptional activity of the target DNA to be regulated.
- DNA-binding proteins and one or more REST inhibitory domains can also be non-covalently bound or complexed to form protein complexes.
- the present application also provides a composition comprising a) a DNA binding protein and b) a protein comprising one or more REST inhibitory domains, wherein said a) and b) are capable of combining to form a protein complex.
- protein complex refers to a complex formed by two protein molecules bound by non-covalent bonds.
- the DNA-binding protein may additionally have a binding moiety (such as an antibody or an antigen-binding fragment thereof) capable of binding to the protein comprising one or more REST inhibitory domains.
- a protein comprising one or more REST inhibitory domains may additionally have a binding moiety (e.g., an antibody or an antigen-binding fragment thereof) capable of binding the DNA-binding protein.
- the binding part may not affect the binding of the DNA binding protein to the target DNA, nor the binding of the protein comprising one or more REST inhibitory domains to RE1.
- the formed protein complex still has at least part of the function of binding the target DNA and at least part of the function of binding RE1.
- a non-covalently bound protein complex can be obtained, which contains both the DNA binding protein, which can bind the target DNA to be regulated, and the inhibitory domain of REST, which can inhibit the target DNA to be regulated transcriptional activity.
- the a) and b) are respectively linked to a pair of self-assembled assemblies, and the self-assembled assemblies can be combined with each other.
- self-assembling assembly refers to two ligands that can spontaneously bind to each other. Since the two ligands are capable of spontaneous association, they are also called a pair of self-assembling assemblies.
- the pair of self-assembling assemblies may be selected from the group consisting of: i) two protein domains that associate with each other; and ii) an RNA splice donor and a splice acceptor. Any two protein domains that can bind to each other can be used as the self-assembling assembly described in this application, for example: antigen and antibody; antigen-binding fragments in antigen and antibody; receptor and ligand; kind of protein.
- the pair of self-assembling assemblies includes: KFBP and FRB; or PYL and ABI. Any RNA cleavage donor and cleavage acceptor can be used as the self-assembly assembly described in the present application, for example, a split intein (Split Intein) can be used.
- the two ligands of the pair of self-assembling assemblies are respectively linked to: a) a DNA binding protein and b) a protein comprising one or more repression domains of REST.
- the inhibitory domain of REST can be linked to an antigen
- the CRISPR-Cas protein can be linked to an antibody bound to it, thereby self-assembling the protein complex described in this application through the interaction between the antigen and the antibody.
- the fusion proteins, compositions and protein complexes provided in this application all have one or more REST inhibitory domains.
- the "REST inhibitory domain” in the present application refers to a domain in the REST protein that has the function of inhibiting gene expression. Without wishing to be bound by any theory, it is believed that there are different REST inhibitory domains in REST proteins, including but not limited to, the N-terminal region of the REST protein and the C-terminal region of the REST protein.
- the "one or more REST inhibitory domains” may refer to one or more identical REST inhibitory domains, such as multiple repetitions of the same inhibitory domain, or one or more Different REST inhibitory domains.
- the one or more REST inhibitory domains comprise a REST N-terminal inhibitory domain and/or a REST C-terminal inhibitory domain.
- the one or more REST inhibitory domains are derived from human REST protein or animal REST protein.
- the animal REST protein may include non-human primate REST protein (such as monkey), rodent REST protein (such as mouse, rat), poultry (such as chicken, duck, goose etc.), farm animals (e.g. cattle, sheep, pigs, etc.).
- the N-terminal inhibitory region of REST comprises amino acids 1-83 of REST or a fragment thereof having transcriptional inhibitory activity.
- transcriptional repression activity means that when the repression domain of REST interacts with the transcriptional regulatory sequence of a target gene, the repression domain of REST can reduce or reduce the transcription of the target gene.
- a fragment having transcriptional repressive activity refers to a fragment capable of providing at least part of the transcriptional repressive activity in the amino acid sequence of REST.
- the N-terminal inhibitory region of the REST comprises the sequence shown in SEQ ID NO: 16 or a fragment thereof having transcriptional inhibitory activity.
- the C-terminal inhibitory region of REST comprises amino acids 1008-1097 of REST or a fragment thereof having transcriptional inhibitory activity.
- the C-terminal inhibitory region of the REST comprises the sequence shown in SEQ ID NO: 18 or a fragment thereof having transcriptional inhibitory activity.
- the fragment with transcriptional repression activity comprises a fragment of at least 20 consecutive amino acids, 30 consecutive amino acids, 40 consecutive amino acids, or 50 consecutive amino acids on the REST protein.
- the DNA binding protein can be targeted to bind to a specific target DNA sequence.
- the DNA binding protein can be a transcription activator-like effector nuclease (TALEN), a zinc finger ribozyme (ZFN), a sequence-guided DNA binding protein such as a CRISPR-Cas protein, or a transcription factor. DNA binding moiety. ZFNs and TALENs achieve specific DNA binding through protein-DNA interactions, and CRISPR-Cas proteins (such as Cas9, Cas12) are guided by short guide RNAs that base-pair with the target DNA to bind to specific DNA sequences.
- the DNA binding portion of the transcription factor is not the naturally occurring DNA binding domain of REST.
- the DNA binding protein is selected from sequence guided DNA binding proteins.
- Sequence-guided DNA-binding protein refers to a DNA-binding protein that can bind to a specific nucleotide sequence under the guidance of a targeting moiety. Sequence-guided DNA binding proteins include, but are not limited to, CRISPR-Cas proteins.
- the sequence-guided DNA binding protein is a CRISPR-Cas protein or a variant thereof.
- CRISPR-Cas protein refers to clustered regularly interspaced short palindromic repeat-related protein, which is a type of nuclease derived from the adaptive immune system of bacteria or archaea.
- CRISPR-Cas protein can be used in targeted Guided by guide RNA (gRNA), it targets and cuts specific nucleic acid sequences.
- CRISPR-Cas proteins capable of binding DNA include, but are not limited to, Cas9 and Cas12.
- the CRISPR-Cas protein variant does not have nuclease activity.
- the sequence-guided DNA binding protein is a Cas9 protein or a Cas9 variant that does not have nuclease activity.
- Cas9 variants that do not have nuclease activity include, nCas9 or dCas9.
- the Cas9 protein can be derived from a variety of bacteria, including, but not limited to, Cas9 from Streptococcus pyogenes (SpCas9), Cas9 from Staphylococcus aureus (SaCas9), Cas9 from Streptococcus thermophilus thermophilus) Cas9 (StCas9), etc.
- the Cas9 protein can be engineered to contain one or more mutations that reduce or eliminate nuclease activity. Mutations in the Cas9 protein can render it inactive at cutting double-stranded DNA, or only able to cut one single-stranded DNA.
- the Asp residue at position 10 can be changed to Ala residue (ie D10A mutant), or the His residue at position 840 can be changed to Ala residue (H840A mutant), thus A mutant that can only cut single-stranded DNA was obtained, also known as nCas9.
- D10A and H840A double mutations can be introduced at the same time, so that Cas9 loses the activity of cutting DNA double strands.
- Such a Cas9 mutant is also called dCas9.
- the dCas9 has an amino acid sequence as shown in SEQ ID NO: 21 or 22.
- sequence-guided DNA-binding protein is Cas12 protein or a Cas12 variant that does not have nuclease activity.
- Cas12 protein also known as Cpf1
- Cpf1 can be derived from a variety of bacteria, including but not limited to Cpf1 (LbCpf1) of Lachnospiraceae bacterium, Cpf1 (AsCpf1) of Acidaminococcus sp, Cpf1 of Francisella novicida (FnCpf1), etc.
- the Cas12 protein can be engineered to comprise one or more mutations capable of reducing or eliminating nuclease activity.
- At least one N-terminal inhibitory domain of the REST protein is linked to the N-terminus or C-terminus of the DNA-binding protein.
- At least one of the C-terminal inhibitory regions of the REST protein is linked to the N- or C-terminus of the DNA-binding protein.
- said fusion protein comprises at least one N-terminal inhibitory domain of said REST linked in tandem to either the N-terminus or the C-terminus of said DNA binding protein.
- said fusion protein comprises at least one C-terminal inhibitory domain of said REST linked in tandem to either the N-terminus or the C-terminus of said DNA binding protein.
- the fusion protein comprises the amino acid sequence shown in SEQ ID NO: 25-27.
- the application provides a polynucleotide encoding the fusion protein or composition or protein complex as described in the application.
- the first polynucleotide fragment and the second polynucleotide fragment are linked by a self-cleavable third nucleotide fragment.
- the application provides a method for inhibiting the expression of a target gene in a cell, comprising delivering a fusion protein as described in the application or a polynucleotide encoding the fusion protein to the cell, wherein the The DNA binding protein can bind the target gene or its regulatory sequence, and inhibit the expression of the target gene.
- the application provides a method for inhibiting the expression of a target gene in an individual, comprising delivering a fusion protein as described in the application or a polynucleotide encoding the fusion protein to the individual, wherein the The DNA binding protein can bind the target gene or its regulatory sequence, and inhibit the expression of the target gene.
- the DNA binding protein is a CRISPR-Cas protein or a variant thereof.
- the CRISPR-Cas protein variant does not have nuclease activity.
- the sequence-guided DNA binding protein is a Cas9 protein or a Cas9 variant that does not have nuclease activity.
- the method further comprises delivering to the cell or the individual a guide RNA comprising a targeting sequence complementary to a target region of the gene of interest or its regulatory sequence, and a DNA binding protein binding sequence.
- the plasmids in this study were all constructed by our laboratory.
- the AAV backbone vector was digested with restriction endonucleases, and agarose gel electrophoresis was performed to recover the backbone vector.
- the inserted DNA fragment is carried out PCR with cell eDNA as a template, and the PCR fragment is recovered after agarose gel electrophoresis.
- the ClonExpress MultiS One Step Cloning Kit (Vazyme, C113-02) from Novizyme Biotechnology Co., Ltd. was used to connect the backbone vector and fragments. After connection, transform into DH5a Escherichia coli and plate. On the second day, single clones were picked for identification, positive clones were sequenced, and clones with completely correct sequencing were expanded for culture and plasmid extraction.
- the AAV serotype used in this study was AAV8.
- Stereotaxic injections were performed using the Revoid Stereotaxic Injection System (C57BL/6 or Dat-Cre:Ai9 mice, older than two months).
- the titers of AAV-GFAP-RZFD-V1, AAV-GFAP-RZFD-V2 and AAV-GFAP-RZFD-V3 were greater than 5 ⁇ 10 12 vg/ml (1-3 ⁇ l per injection).
- AAV was injected into striatum (AP+0.8mm, ML ⁇ 1.6mm and DV-2.8mm) or substantia nigra (AP-3.0mm, ML ⁇ 1.25mm and DV-4.5mm).
- the primary antibodies used in this study include: guinea pig anti-NeuN (1:500, ABN90, Millipore), rabbit anti-TH (1:500, AB152, Millipore), rat anti-DAT (1:100, MAB369, Millipore), rabbit Anti-RBPMS (Proteintech, Cat#15187-1-AP) and mouse anti-Flag (1:2000, F3165, Sigma).
- the secondary antibodies used in this study were: Cy5-AffiniPure Donkey Anti-Guinea Pig IgG(H+L) (1:500, 706-175-148, Jackson ImmunoResearch), Alexa Fluora-488AffiniPure Donkey Anti-Rabbit IgG(H+ L) (1:500, 711-545-152, Jackson ImmunoResearch), Alexa Fluora-488 AffiniPure Donkey Anti-Mouse IgG (H+L) (1:500, 715-545-150, Jackson ImmunoResearch), and Cy5 AffiniPure Donkey Anti-Rabbit IgG (H+L) (1:500, 711-175-152, Jackson ImmunoResearch).
- mice used in this experiment were adult C57BL/6 mice (7-10 weeks) mice.
- 25 mg/kg desipramine hydrochloride (D3900, Sigma-Aldrich) was intraperitoneally injected.
- mice were injected subcutaneously with 1 ml of 4% glucose-saline solution.
- mice were injected intraperitoneally with 0.5 mg/kg apomorphine (A4393, Sigma-Aldrich) 10 minutes before the test. During the test, the mice were each placed in an opaque cylinder (diameter 30 cm), and the behavior was recorded by a video camera above it for 20 minutes. Rotation was defined as a full-body turn with one hind paw as the center and no change in head orientation. Count the number of rotations on the injected side and the injected contralateral side, and quantify the data as the number of contralateral reversals within 20 min.
- apomorphine A4393, Sigma-Aldrich
- mice were gently placed in a glass beaker (1000 ml) and their behavior was recorded for 10 minutes with a video camera in front of them.
- the number of wall touches on the injected side and the contralateral forelimb were counted separately, and the data were quantified as the ratio of the number of wall touches on the ipsilateral side to the total number of wall touches.
- mice were first trained for 2 days, and behavioral tests were performed on the 3rd day. On day 1, the mice were trained 4 times with the rotating rod at a fixed speed of 4 turns/min for 300 s each time. On day 2, train the mice 4 times at an accelerated pace ranging from 4 to 40 laps/min. On the third day, the behavioral test was performed at 4-40 laps/minute, and the test was performed 4 times. The time the mouse stayed on the pole before falling off was recorded as the dwell period and the average of the 3 longest dwell periods was used for analysis.
- NMDA solution 200 mM NMDA solution was prepared in PBS, and then 1.5 ⁇ l NMDA solution was injected into 4-8 week-old Ai9 mice or 5-6 week-old Ai9 mice by intravitreal injection. In the eyes of C57BL/6 mice (for VEP and black and white scene preference tests). 2–3 weeks after NMDA injection, GFAP-GFP-Cre was co-delivered to the retina by subretinal injection together with GFAP-CasRx-REST or GFAP-CasRx.
- mice C57BL/6 were injected with NMDA to induce retinal damage and subretinal injection 2-3 weeks after injection GFAP-mCherry (0.1 ⁇ l) mixed with PBS (0.9 ⁇ l) or GFAP-RZFD-V1 (0.9 ⁇ l)/GFAP-RZFD-V2 (0.9 ⁇ l)/GFAP-RZFD-V3 (0.9 ⁇ l) mixture.
- Eyes were injected with high titer (>1 ⁇ 10 13 vg/ml) AAV with a Hamilton syringe (32G needle) under an Olympus microscope (Olympus, Tokyo, Japan).
- a total of 1 ⁇ l of GFAP-GFP-Cre (0.1ul) + pbs (0.9ul) was injected by subretinal injection (Ai9 and C57BL/6 mice, 4 weeks to 12 months old), or GFAP-GFP-Cre (0.1 ⁇ l) and GFAP-RZFD-V1 (0.9 ⁇ l)/GFAP-RZFD-V2 (0.9 ⁇ l)/GFAP-RZFD-V3 (0.9 ⁇ l) were injected subretinal.
- the eyes, optic nerve and brain tissue were collected, fixed with 4% paraformaldehyde (PFA) for 2 hours (eye and optic nerve) or 24 hours (brain), and then dehydrated in 30% sucrose solution After 2h (eyes) or 24 (brain) hours, the optic nerve was directly washed with PBS and mounted for observation. Then embedded with OCT, the eyes and brain were sliced at a thickness of 30 ⁇ m.
- PFA paraformaldehyde
- the secondary antibodies were: Cy TM 5AffiniPure Donkey mouse anti-IgG (H+L) (1:500, 715-175-150, Jackson ImmunoResearch) and Cy TM 5AffiniPure Donkey rabbit anti-IgG (H+L) (1:500, 711 -175-152, Jackson ImmunoResearch).
- the primary antibody was incubated overnight at 4°C, and then washed 3 times with PBS, 10 minutes each time.
- the secondary antibody was incubated at room temperature for 2-3 hours, and then washed 3 times with PBS, 10 minutes each time. Finally, slides were mounted with anti-fade reagent (Life technology) and imaged using an Olympus FV3000 microscope.
- REST Zinc Finger Domain REST Zinc Finger Domain
- VP64 SEQ ID NO: 7
- P65-HSF1 SEQ ID NO: 11
- VP64 and P65-HSF can recruit transcription factors and histone acetylation proteins to the vicinity of RE1, regulate the chromosome structure near RE1, and allow the genes controlled by them to express.
- RZFD codon of RZFD (SEQ ID NO: 15) to increase the expression of RZFD in mammalian cells.
- RZFDmax bpNLS sequences at both ends of RZFD to enhance For its nuclear efficiency, this version was named RZFDmax (Fig. 1A).
- RZFD-VP64 and RZFD-P65-HSF1 can not only competitively bind to RE1, but also further change the chromosomal state of RE1 attachment, and promote the expression of nerve-related genes regulated by it.
- the N-terminal (SEQ ID NO: 16) and C-terminal (SEQ ID NO: 18) of the REST protein have the function of recruiting various epigenetic regulatory elements, thereby realizing the regulation of the expression of REST target genes.
- N-dCas9 SEQ ID NO: 25
- 3x N -dCas9 SEQ ID NO: 26
- 3x N-dCas9-3xC SEQ ID NO: 27
- an AAV expression vector In order to realize the transdifferentiation of glial cells into neurons in mice, we constructed an AAV expression vector. In order to specifically label astrocytes, in the AAV expression vector, we used the astrocyte-specific promoter GFAP to drive the expression of the fluorescent protein mCherry (GFAP-mCherry), and the human RZFD was also driven by the GFAP promoter (GFAP - RZFD, wherein the amino acid sequence of RZFD is SEQ ID NO: 1, and the nucleic acid sequence is SEQ ID NO: 2), which is specifically expressed in astrocytes ( FIG. 2A ).
- TH neuron-specific markers NeuN and dopamine neuron-specific marker tyrosine hydroxylase staining ( FIG. 2B ).
- AAV expressing GFAP-mCherry+GFAP-RZFD was injected into the striatum or substantia nigra of 8-week-old C57 mice as the experimental group, or AAV expressing GFAP-mCherry alone was injected as the control group (Fig. 2C).
- Example 4 RZFD transdifferentiates astrocytes into dopamine neurons and alleviates disease symptoms in a Parkinson's mouse model
- Dat-Cre:Ai9 can specifically label dopamine neurons, and only dopamine neurons will be labeled as tdTomato positive.
- mice in the control group could maintain about 120s, while the treated mice could reach about 180s (Fig. 3D).
- the balance of the mice in the treatment group was significantly better than that in the control group (Fig. 3E).
- the drug-induced rotatory behavior test the circling behavior was significantly reduced in the treatment group relative to the control group (Fig. 3F and 3G).
- Example 5 RZFD-VP64 and RZFD-P65-HSF1 transdifferentiate astrocytes into dopamine neurons
- RZFD-VP64 annotated as GFAP-RZFD-V2, amino acid sequence SEQ ID NO: 5, nucleic acid sequence SEQ ID NO: 6) and RZFD-P65-HSF1 (annotated as GFAP-RZFD-V3, amino acid sequence SEQ ID NO: 9, nucleic acid sequence SEQ ID NO: 10) AAV expression vector (Fig. 4A).
- RZFD-VP64 and RZFD-P65-HSF1 were expressed.
- Example 6 RZFD transdifferentiates Müller glial cells into photoreceptor cells and retinal ganglion cells
- AAV expressing NLS-RZFD-V1 SEQ ID NO: 3
- AAV expressing GFAP-EGFP-2A-Cre was delivered into the eyes of mice by subretinal injection.
- GFAP-EGFP-2A-Cre is used to label Müller glial cells.
- Cre is specifically expressed in Müller glial cells, and the LSL sequence of Ai9 mice is unwrapped, so as to achieve the purpose of labeling Ai9 mice. Effect of murine Müller glial cells (Fig.
- Rhodopsin and Cone arrestin Staining with photoreceptor cell-specific protein markers Rhodopsin and Cone arrestin revealed that tdTomato-positive cells located in the outer granular layer were co-labeled with Rhodopsin or Cone arrestin (ie, MCAR) ( Figure 5H and 5I). This suggests that overexpression of RZFD in Müller glial cells can transdifferentiate them into photoreceptor cells. The number of photoreceptor cells was counted, and it was found that the average number of photoreceptor cells in each visual field was about 8, while almost no tdTomato-positive photoreceptor cells were observed in the control group (Fig. 5J).
- tissue-specific promoter GFAP tissue-specific promoter GFAP to promote the expression of miR-124pri-miRNA (as shown in SEQ ID NO: 34)
- pri-miRNA will be processed into pre-miRNA (as shown in SEQ ID NO: 34) in the cell after expression : 33), and finally processed into mature miRNA (SEQ ID NO: 31 and SEQ ID NO: 32), thereby achieving tissue-specific expression of miR-124.
- glial cells we used the same method in glial cells to use similar AAV vectors to promote the overexpression of miR-124 with the tissue-specific promoter GFAP to verify whether it can transdifferentiate glial cells into neurons.
- Analysis of injected mouse brain slices revealed that overexpression of miR-124 did not transdifferentiate glial cells into neurons (Fig. 6A-D).
- the red fluorescent signal mCherry was expressed in glial cells, and it was not co-labeled with the neuron-specific marker NeuN, and there were no TH-positive neurons in the striatum (Fig. 6C). Therefore, upregulation of miR-124 alone is not sufficient to achieve transdifferentiation of glial cells into dopamine neurons.
- miR-9pri-miRNA (as shown in SEQ ID NO: 38) specifically Expressed in glial cells, miR-9pri-miRNA will be processed into pre-miRNA (as shown in SEQ ID NO: 37) in the cell after expression, and finally processed into mature miRNA (SEQ ID NO: 35 and SEQ ID NO: 35 and SEQ ID NO: 36), so as to achieve tissue-specific expression of miR-9.
- GFAP-mCherry was used to label astrocytes in the striatum.
- miR-9 could not promote the transdifferentiation of glial cells into neurons. Similar to miR-124, overexpression of either miR-9 or miR-124 failed to transdifferentiate astrocytes into neurons or dopaminergic neurons, although some glial cells underwent morphological changes.
- Example 8 Co-overexpression of miR-9 and miR124 can transdifferentiate glial cells into neurons
- Ctdsp1 amino acid sequence SEQ ID NO: 29, nucleic acid sequence SEQ ID NO: 30
- shRNA knockdown was used.
- CasRx-mediated RNA editing technology is a new generation of RNA knockdown technology with higher efficiency and stronger specificity.
- a gRNA targeting Ctdsp1 was first designed, and the gRNA targeting region was the 318-347th position of the CDS sequence of the Ctdsp1 gene, and was tested in cultured human 293T and mouse N2A cells Their inhibition efficiencies were compared.
- the plasmids transfected in the control group were CAG-CasRx-P2A-GFP and U6-nontarget-CMV-mCherry
- the plasmids transfected in the experimental group were CAG-CasRx-P2A-GFP and U6-gRNA(Ctdsp1)-CMV-mCherry.
- Example 10 Knockdown of Ctdsp1 in astrocytes transdifferentiates glial cells into neurons but fails to generate dopamine neurons
- NLS-RZFD-V1 amino acid sequence (SEQ ID NO: 3)
- NLS-RZFD-V1 nucleotide sequence (SEQ ID NO: 4)
- Amino acid sequence (SEQ ID NO: 5) of NLS-RZFD-V2
- VP64 nucleotide sequence (SEQ ID NO: 8)
- NLS-RZFD-V3 amino acid sequence (SEQ ID NO: 9)
- NLS-RZFD-V3 nucleotide sequence (SEQ ID NO: 10)
- the sequence of expressing human miRNA-9Pri-miRNA is (SEQ ID NO: 38)
- Short GFAP promoter (SEQ ID NO: 40)
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Health & Medical Sciences (AREA)
- Biomedical Technology (AREA)
- Medicinal Chemistry (AREA)
- Pharmacology & Pharmacy (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- Animal Behavior & Ethology (AREA)
- Genetics & Genomics (AREA)
- General Chemical & Material Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Neurology (AREA)
- Neurosurgery (AREA)
- Molecular Biology (AREA)
- Zoology (AREA)
- Biochemistry (AREA)
- Biotechnology (AREA)
- Wood Science & Technology (AREA)
- General Engineering & Computer Science (AREA)
- Epidemiology (AREA)
- Biophysics (AREA)
- Microbiology (AREA)
- Ophthalmology & Optometry (AREA)
- Physics & Mathematics (AREA)
- Toxicology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Gastroenterology & Hepatology (AREA)
- Plant Pathology (AREA)
- Psychiatry (AREA)
- Psychology (AREA)
- Anesthesiology (AREA)
- Orthopedic Medicine & Surgery (AREA)
- Vascular Medicine (AREA)
- Cardiology (AREA)
Priority Applications (7)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202280063875.9A CN118201623A (zh) | 2021-09-30 | 2022-09-30 | 直接转分化治疗神经系统疾病 |
| EP22875183.0A EP4410296A4 (en) | 2021-09-30 | 2022-09-30 | DIRECT TRANSDIFFERENTIATION FOR THE TREATMENT OF NEUROLOGICAL DISEASES |
| KR1020247013258A KR20240082366A (ko) | 2021-09-30 | 2022-09-30 | 신경계 질환 치료를 위한 직접 전환분화 |
| AU2022357103A AU2022357103A1 (en) | 2021-09-30 | 2022-09-30 | Direct transdifferentiation for treatment of neurological disease |
| US18/697,216 US20250115650A1 (en) | 2021-09-30 | 2022-09-30 | Direct transdifferentiation for treatment of neurological disease |
| CA3233534A CA3233534A1 (en) | 2021-09-30 | 2022-09-30 | Direct transdifferentiation for treatment of neurological disease |
| JP2024519557A JP2024534667A (ja) | 2021-09-30 | 2022-09-30 | 神経系疾患の直接分化転換治療 |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202111158620.4 | 2021-09-30 | ||
| CN202111158620.4A CN115887655B (zh) | 2021-09-30 | 2021-09-30 | 直接转分化治疗神经系统疾病 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2023051802A1 true WO2023051802A1 (zh) | 2023-04-06 |
Family
ID=85750328
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/CN2022/123409 Ceased WO2023051802A1 (zh) | 2021-09-30 | 2022-09-30 | 直接转分化治疗神经系统疾病 |
Country Status (8)
| Country | Link |
|---|---|
| US (1) | US20250115650A1 (enExample) |
| EP (1) | EP4410296A4 (enExample) |
| JP (1) | JP2024534667A (enExample) |
| KR (1) | KR20240082366A (enExample) |
| CN (3) | CN115887655B (enExample) |
| AU (1) | AU2022357103A1 (enExample) |
| CA (1) | CA3233534A1 (enExample) |
| WO (1) | WO2023051802A1 (enExample) |
Cited By (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2024199511A1 (zh) * | 2023-03-30 | 2024-10-03 | 上海鲸奇生物科技有限公司 | 治疗神经系统疾病的表达载体和组合物 |
| WO2025173411A1 (ja) * | 2024-02-14 | 2025-08-21 | 国立大学法人筑波大学 | 網膜神経節細胞分化誘導用キット、医薬組成物及び緑内障モデル |
Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20150299698A1 (en) * | 2012-11-01 | 2015-10-22 | The Regents Of The University Of California | Methods for engineering non-neuronal cells into neurons and using newly engineered neurons to treat neurodegenerative diseases |
| US20160031958A1 (en) * | 2014-08-01 | 2016-02-04 | Oregon Health & Science University | Methods and compositions useful in manipulating the stability of re1 silencing transcription factor |
| CN112566640A (zh) * | 2018-06-18 | 2021-03-26 | 罗切斯特大学 | 治疗精神分裂症和其他神经精神病症的方法 |
| WO2021102345A1 (en) * | 2019-11-22 | 2021-05-27 | Alcamena Stem Cell Therapeutics, Llc | Compositions and methods for derepressing re1 silencing transcription factor target genes |
Family Cites Families (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| FR2774698B1 (fr) * | 1998-02-12 | 2002-03-01 | Rhone Poulenc Rorer Sa | Utilisation d'elements de regulation negative pour l'expression neurospecifique de transgenes |
| ITVA20060041A1 (it) * | 2006-07-05 | 2008-01-06 | Dialectica Srl | Uso di composti derivati amminotiazolici, di loro composizioni farmaceutiche, nel trattamento di malattie caratterizzate dalla anormale repressione della trascrizione genica, particolarmente il morbo di huntington |
| WO2009058014A2 (en) * | 2007-11-02 | 2009-05-07 | Vereniging Voor Christelijk Hoger Onderwijs, Wetenschappelijk Onderzoek En Patientenzorg | Polypeptides involved in neuronal regeneration-associated gene expression |
| JP5850321B2 (ja) * | 2010-02-10 | 2016-02-03 | 公立大学法人横浜市立大学 | 神経選択的転写抑制因子NRSFに特異的に結合するmSin3Bに結合する化合物の利用 |
| EP3622057A1 (en) * | 2017-05-12 | 2020-03-18 | New York Stem Cell Foundation, Inc. | Systems |
| CA3096691A1 (en) * | 2018-04-11 | 2019-10-17 | The Regents Of The University Of California | Reprogramming of non-neuronal cells into neurons and methods and compositions to treat neurodegenerative diseases and disorders |
-
2021
- 2021-09-30 CN CN202111158620.4A patent/CN115887655B/zh active Active
- 2021-09-30 CN CN202510816959.0A patent/CN120754249A/zh active Pending
-
2022
- 2022-09-30 WO PCT/CN2022/123409 patent/WO2023051802A1/zh not_active Ceased
- 2022-09-30 EP EP22875183.0A patent/EP4410296A4/en active Pending
- 2022-09-30 CA CA3233534A patent/CA3233534A1/en active Pending
- 2022-09-30 US US18/697,216 patent/US20250115650A1/en active Pending
- 2022-09-30 JP JP2024519557A patent/JP2024534667A/ja active Pending
- 2022-09-30 KR KR1020247013258A patent/KR20240082366A/ko active Pending
- 2022-09-30 AU AU2022357103A patent/AU2022357103A1/en active Pending
- 2022-09-30 CN CN202280063875.9A patent/CN118201623A/zh active Pending
Patent Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20150299698A1 (en) * | 2012-11-01 | 2015-10-22 | The Regents Of The University Of California | Methods for engineering non-neuronal cells into neurons and using newly engineered neurons to treat neurodegenerative diseases |
| US20160031958A1 (en) * | 2014-08-01 | 2016-02-04 | Oregon Health & Science University | Methods and compositions useful in manipulating the stability of re1 silencing transcription factor |
| CN112566640A (zh) * | 2018-06-18 | 2021-03-26 | 罗切斯特大学 | 治疗精神分裂症和其他神经精神病症的方法 |
| WO2021102345A1 (en) * | 2019-11-22 | 2021-05-27 | Alcamena Stem Cell Therapeutics, Llc | Compositions and methods for derepressing re1 silencing transcription factor target genes |
Non-Patent Citations (8)
| Title |
|---|
| ALTSCHUL S.F ET AL., J. MOLECULAR BIOLOGY, vol. 215, 1990, pages 403 - 410 |
| HIGGINS D.G. ET AL., METHODS IN ENZYMOLOGY, vol. 266, 1996, pages 383 - 402 |
| HWANG JEE-YEON, ZUKIN R SUZANNE: "REST, a master transcriptional regulator in neurodegenerative disease", CURRENT OPINION IN NEUROBIOLOGY, vol. 48, 1 February 2018 (2018-02-01), GB , pages 193 - 200, XP093052784, ISSN: 0959-4388, DOI: 10.1016/j.conb.2017.12.008 * |
| KONG QING-RAN, XIE BING-TENG, ZHANG HENG, LI JING-YU, HUANG TIAN-QING, WEI REN-YUE, LIU ZHONG-HUA: "RE1-silencing Transcription Factor (REST) Is Required for Nuclear Reprogramming by Inhibiting Transforming Growth Factor β Signaling Pathway", JOURNAL OF BIOLOGICAL CHEMISTRY, vol. 291, no. 53, 1 December 2016 (2016-12-01), US , pages 27334 - 27342, XP093052786, ISSN: 0021-9258, DOI: 10.1074/jbc.M116.743849 * |
| LARKIN M.A ET AL., BIOINFORMATICS (OXFORD, ENGLAND, vol. 23, no. 21, 2007, pages 2947 - 8 |
| See also references of EP4410296A4 |
| SHIMOJO MASAHITO, LEE JEONG-HEON, HERSH LOUIS B.: "Role of Zinc Finger Domains of the Transcription Factor Neuron-restrictive Silencer Factor/Repressor Element-1 Silencing Transcription Factor in DNA Binding and Nuclear Localization", JOURNAL OF BIOLOGICAL CHEMISTRY, vol. 276, no. 16, 1 April 2001 (2001-04-01), US , pages 13121 - 13126, XP093052785, ISSN: 0021-9258, DOI: 10.1074/jbc.M011193200 * |
| STEPHEN F. ET AL., NUCLEIC ACIDS RES., vol. 25, 1997, pages 3389 - 3402 |
Cited By (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2024199511A1 (zh) * | 2023-03-30 | 2024-10-03 | 上海鲸奇生物科技有限公司 | 治疗神经系统疾病的表达载体和组合物 |
| WO2025173411A1 (ja) * | 2024-02-14 | 2025-08-21 | 国立大学法人筑波大学 | 網膜神経節細胞分化誘導用キット、医薬組成物及び緑内障モデル |
Also Published As
| Publication number | Publication date |
|---|---|
| CN118201623A (zh) | 2024-06-14 |
| KR20240082366A (ko) | 2024-06-10 |
| CN115887655A (zh) | 2023-04-04 |
| US20250115650A1 (en) | 2025-04-10 |
| CN115887655B (zh) | 2025-06-17 |
| EP4410296A1 (en) | 2024-08-07 |
| CA3233534A1 (en) | 2023-04-06 |
| CN120754249A (zh) | 2025-10-10 |
| JP2024534667A (ja) | 2024-09-20 |
| EP4410296A4 (en) | 2025-10-15 |
| AU2022357103A1 (en) | 2024-05-02 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| IL241539B2 (en) | Methods for using zscan4 to rejuvenate human cells | |
| WO2021031810A1 (zh) | Ptbp1抑制剂在预防和/或治疗功能性神经元死亡相关的神经系统疾病中的应用 | |
| CN112386699A (zh) | Ptbp1抑制剂在预防和/或治疗功能性神经元死亡相关的神经系统疾病中的应用 | |
| US20250115650A1 (en) | Direct transdifferentiation for treatment of neurological disease | |
| CN112566640A (zh) | 治疗精神分裂症和其他神经精神病症的方法 | |
| JP7776157B2 (ja) | 神経疾患の治療 | |
| WO2021032068A1 (zh) | Ptbp1抑制剂在预防和/或治疗功能性神经元死亡相关的神经系统疾病中的应用 | |
| CA3197321A1 (en) | Neurod1 vector | |
| US20250281648A1 (en) | Use of transdifferentiation of glial cells into neurons in prevention or treatment of diseases associated with neuron loss-of-function or death | |
| JP7680072B2 (ja) | リプログラミング用機能性断片、組み合わせ、およびその用途 | |
| IL301750A (en) | Neurod1 and dlx2 vector | |
| WO2021031025A1 (zh) | Ptbp1抑制剂在预防和/或治疗神经退行性疾病中的应用 | |
| US20240400984A1 (en) | Treatment of Neuronal Absence Disease by Transdifferentiating Treatment | |
| JP2023524900A (ja) | 機能性ニューロンへのグリア細胞の分化転換を誘導するための方法、及びその用途 | |
| WO2024046393A1 (zh) | 非神经元细胞转分化为神经元的方法及应用 | |
| JPWO2023051802A5 (enExample) | ||
| WO2024199511A1 (zh) | 治疗神经系统疾病的表达载体和组合物 | |
| KR20250057176A (ko) | Arid4b 유전자 발현을 통한 신경세포로의 직접교차분화 유도 방법 | |
| WO2025038955A9 (en) | Systems and methods for improving safety of split intein aav mediated gene therapy | |
| KR20230123926A (ko) | Dlx2 벡터 | |
| JP2024038021A (ja) | 機能的ミエリン産生を増進させるための方法および組成物 | |
| CN116670160A (zh) | Dlx2载体 | |
| CN116761812A (zh) | Neurod1和dlx2载体 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 22875183 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 202280063875.9 Country of ref document: CN |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 3233534 Country of ref document: CA |
|
| ENP | Entry into the national phase |
Ref document number: 2024519557 Country of ref document: JP Kind code of ref document: A |
|
| WWE | Wipo information: entry into national phase |
Ref document number: AU2022357103 Country of ref document: AU |
|
| ENP | Entry into the national phase |
Ref document number: 20247013258 Country of ref document: KR Kind code of ref document: A |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2022357103 Country of ref document: AU Date of ref document: 20220930 Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 2022875183 Country of ref document: EP Effective date: 20240430 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 11202402178T Country of ref document: SG |
|
| WWP | Wipo information: published in national office |
Ref document number: 18697216 Country of ref document: US |