WO2021241728A1 - 抗ウイルス剤 - Google Patents
抗ウイルス剤 Download PDFInfo
- Publication number
- WO2021241728A1 WO2021241728A1 PCT/JP2021/020382 JP2021020382W WO2021241728A1 WO 2021241728 A1 WO2021241728 A1 WO 2021241728A1 JP 2021020382 W JP2021020382 W JP 2021020382W WO 2021241728 A1 WO2021241728 A1 WO 2021241728A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- strain
- lactobacillus
- antiviral
- receipt number
- number nite
- Prior art date
Links
- 239000003443 antiviral agent Substances 0.000 title claims abstract description 36
- 230000014509 gene expression Effects 0.000 claims abstract description 157
- 241000186660 Lactobacillus Species 0.000 claims abstract description 152
- 229940039696 lactobacillus Drugs 0.000 claims abstract description 142
- 230000000840 anti-viral effect Effects 0.000 claims abstract description 134
- 241000700605 Viruses Species 0.000 claims abstract description 65
- 108090000623 proteins and genes Proteins 0.000 claims abstract description 60
- 230000001580 bacterial effect Effects 0.000 claims abstract description 57
- 239000002773 nucleotide Substances 0.000 claims abstract description 41
- 125000003729 nucleotide group Chemical group 0.000 claims abstract description 41
- 108020004465 16S ribosomal RNA Proteins 0.000 claims abstract description 29
- 230000002708 enhancing effect Effects 0.000 claims abstract description 15
- 230000001603 reducing effect Effects 0.000 claims abstract description 12
- 230000000694 effects Effects 0.000 claims description 43
- 108091032973 (ribonucleotides)n+m Proteins 0.000 claims description 36
- 240000006024 Lactobacillus plantarum Species 0.000 claims description 31
- 235000013965 Lactobacillus plantarum Nutrition 0.000 claims description 31
- 229940072205 lactobacillus plantarum Drugs 0.000 claims description 31
- 241001261506 Undaria pinnatifida Species 0.000 claims description 28
- 241000186869 Lactobacillus salivarius Species 0.000 claims description 25
- 241001474374 Blennius Species 0.000 claims description 19
- 102100034170 Interferon-induced, double-stranded RNA-activated protein kinase Human genes 0.000 claims description 19
- 101710089751 Interferon-induced, double-stranded RNA-activated protein kinase Proteins 0.000 claims description 19
- 101100198353 Mus musculus Rnasel gene Proteins 0.000 claims description 19
- 108020003175 receptors Proteins 0.000 claims description 18
- 230000002222 downregulating effect Effects 0.000 claims description 17
- 102000005962 receptors Human genes 0.000 claims description 16
- 102100037435 Antiviral innate immune response receptor RIG-I Human genes 0.000 claims description 15
- 102000014150 Interferons Human genes 0.000 claims description 14
- 108010050904 Interferons Proteins 0.000 claims description 14
- 229940079322 interferon Drugs 0.000 claims description 14
- 102000040650 (ribonucleotides)n+m Human genes 0.000 claims description 13
- 101000952099 Homo sapiens Antiviral innate immune response receptor RIG-I Proteins 0.000 claims description 12
- 244000144972 livestock Species 0.000 claims description 12
- 101000669447 Homo sapiens Toll-like receptor 4 Proteins 0.000 claims description 9
- 102100039360 Toll-like receptor 4 Human genes 0.000 claims description 9
- 241001493065 dsRNA viruses Species 0.000 claims description 9
- 235000013305 food Nutrition 0.000 claims description 8
- 102000002689 Toll-like receptor Human genes 0.000 claims description 6
- 108020000411 Toll-like receptor Proteins 0.000 claims description 6
- 101001128393 Homo sapiens Interferon-induced GTP-binding protein Mx1 Proteins 0.000 claims description 4
- 102100031802 Interferon-induced GTP-binding protein Mx1 Human genes 0.000 claims description 4
- 102100024652 Toll-interacting protein Human genes 0.000 claims description 4
- 101710182709 Toll-interacting protein Proteins 0.000 claims description 4
- 238000006384 oligomerization reaction Methods 0.000 claims description 3
- SIIZPVYVXNXXQG-KGXOGWRBSA-N [(2r,3r,4r,5r)-5-(6-aminopurin-9-yl)-4-[[(3s,4r)-5-(6-aminopurin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-3-hydroxyoxolan-2-yl]methyl [(2r,4r,5r)-2-(6-aminopurin-9-yl)-4-hydroxy-5-(phosphonooxymethyl)oxolan-3-yl] hydrogen phosphate Polymers C1=NC2=C(N)N=CN=C2N1[C@@H]1O[C@H](COP(O)(=O)OC2[C@@H](O[C@H](COP(O)(O)=O)[C@H]2O)N2C3=NC=NC(N)=C3N=C2)[C@@H](O)[C@H]1OP(O)(=O)OCC([C@@H](O)[C@H]1O)OC1N1C(N=CN=C2N)=C2N=C1 SIIZPVYVXNXXQG-KGXOGWRBSA-N 0.000 claims description 2
- 208000015181 infectious disease Diseases 0.000 abstract description 28
- 244000052616 bacterial pathogen Species 0.000 abstract description 12
- 239000006041 probiotic Substances 0.000 abstract description 12
- 235000018291 probiotics Nutrition 0.000 abstract description 12
- 239000003795 chemical substances by application Substances 0.000 abstract description 5
- 239000004480 active ingredient Substances 0.000 abstract description 3
- 210000004027 cell Anatomy 0.000 description 204
- 239000002609 medium Substances 0.000 description 100
- 238000000034 method Methods 0.000 description 87
- 241000702670 Rotavirus Species 0.000 description 70
- 239000007788 liquid Substances 0.000 description 70
- 108091036414 Polyinosinic:polycytidylic acid Proteins 0.000 description 64
- 239000000243 solution Substances 0.000 description 64
- 239000006144 Dulbecco’s modified Eagle's medium Substances 0.000 description 47
- 108020004999 messenger RNA Proteins 0.000 description 46
- 230000000638 stimulation Effects 0.000 description 39
- 108090000467 Interferon-beta Proteins 0.000 description 38
- 102100026720 Interferon beta Human genes 0.000 description 33
- 101150112867 MX1 gene Proteins 0.000 description 32
- 238000004458 analytical method Methods 0.000 description 28
- 230000012010 growth Effects 0.000 description 28
- SIKJAQJRHWYJAI-UHFFFAOYSA-N Indole Chemical compound C1=CC=C2NC=CC2=C1 SIKJAQJRHWYJAI-UHFFFAOYSA-N 0.000 description 26
- 230000006799 invasive growth in response to glucose limitation Effects 0.000 description 26
- 101150062121 tollip gene Proteins 0.000 description 22
- 230000009385 viral infection Effects 0.000 description 21
- 229920001817 Agar Polymers 0.000 description 20
- 239000008272 agar Substances 0.000 description 20
- 238000011156 evaluation Methods 0.000 description 19
- 229910052760 oxygen Inorganic materials 0.000 description 18
- 102000004127 Cytokines Human genes 0.000 description 17
- 108090000695 Cytokines Proteins 0.000 description 17
- 244000005700 microbiome Species 0.000 description 17
- 108010085238 Actins Proteins 0.000 description 16
- 238000012360 testing method Methods 0.000 description 15
- 206010067470 Rotavirus infection Diseases 0.000 description 14
- LFYJSSARVMHQJB-QIXNEVBVSA-N bakuchiol Chemical compound CC(C)=CCC[C@@](C)(C=C)\C=C\C1=CC=C(O)C=C1 LFYJSSARVMHQJB-QIXNEVBVSA-N 0.000 description 14
- 238000004519 manufacturing process Methods 0.000 description 14
- 102100027769 2'-5'-oligoadenylate synthase 1 Human genes 0.000 description 13
- 241000193830 Bacillus <bacterium> Species 0.000 description 13
- 229920001342 Bakelite® Polymers 0.000 description 13
- 102000016938 Catalase Human genes 0.000 description 13
- 108010053835 Catalase Proteins 0.000 description 13
- 102000012422 Collagen Type I Human genes 0.000 description 13
- 108010022452 Collagen Type I Proteins 0.000 description 13
- 101001008907 Homo sapiens 2'-5'-oligoadenylate synthase 1 Proteins 0.000 description 13
- QVGXLLKOCUKJST-UHFFFAOYSA-N atomic oxygen Chemical compound [O] QVGXLLKOCUKJST-UHFFFAOYSA-N 0.000 description 13
- 239000004637 bakelite Substances 0.000 description 13
- 230000015572 biosynthetic process Effects 0.000 description 13
- 239000007789 gas Substances 0.000 description 13
- PZOUSPYUWWUPPK-UHFFFAOYSA-N indole Natural products CC1=CC=CC2=C1C=CN2 PZOUSPYUWWUPPK-UHFFFAOYSA-N 0.000 description 13
- RKJUIXBNRJVNHR-UHFFFAOYSA-N indolenine Natural products C1=CC=C2CC=NC2=C1 RKJUIXBNRJVNHR-UHFFFAOYSA-N 0.000 description 13
- 230000004899 motility Effects 0.000 description 13
- 239000001301 oxygen Substances 0.000 description 13
- 238000010195 expression analysis Methods 0.000 description 12
- 238000002360 preparation method Methods 0.000 description 12
- 230000002519 immonomodulatory effect Effects 0.000 description 11
- 230000000968 intestinal effect Effects 0.000 description 11
- 230000002829 reductive effect Effects 0.000 description 10
- 208000036142 Viral infection Diseases 0.000 description 9
- 210000004962 mammalian cell Anatomy 0.000 description 9
- 239000000203 mixture Substances 0.000 description 9
- 241000894006 Bacteria Species 0.000 description 8
- 108020004414 DNA Proteins 0.000 description 8
- 241000588724 Escherichia coli Species 0.000 description 8
- -1 antibody] Proteins 0.000 description 8
- 238000010166 immunofluorescence Methods 0.000 description 8
- JVTAAEKCZFNVCJ-UHFFFAOYSA-N lactic acid Chemical compound CC(O)C(O)=O JVTAAEKCZFNVCJ-UHFFFAOYSA-N 0.000 description 8
- 238000011084 recovery Methods 0.000 description 8
- PEDCQBHIVMGVHV-UHFFFAOYSA-N Glycerine Chemical compound OCC(O)CO PEDCQBHIVMGVHV-UHFFFAOYSA-N 0.000 description 7
- 241000124008 Mammalia Species 0.000 description 7
- 241001465754 Metazoa Species 0.000 description 7
- 230000002155 anti-virotic effect Effects 0.000 description 7
- 239000002299 complementary DNA Substances 0.000 description 7
- 230000036541 health Effects 0.000 description 7
- 230000002265 prevention Effects 0.000 description 7
- 238000005406 washing Methods 0.000 description 7
- CSCPPACGZOOCGX-UHFFFAOYSA-N Acetone Chemical compound CC(C)=O CSCPPACGZOOCGX-UHFFFAOYSA-N 0.000 description 6
- 101000831567 Homo sapiens Toll-like receptor 2 Proteins 0.000 description 6
- 102100037850 Interferon gamma Human genes 0.000 description 6
- 108010074328 Interferon-gamma Proteins 0.000 description 6
- 102100024333 Toll-like receptor 2 Human genes 0.000 description 6
- 239000000463 material Substances 0.000 description 6
- 108091069025 single-strand RNA Proteins 0.000 description 6
- 230000000052 comparative effect Effects 0.000 description 5
- 238000012258 culturing Methods 0.000 description 5
- 230000028993 immune response Effects 0.000 description 5
- 230000001717 pathogenic effect Effects 0.000 description 5
- 239000000047 product Substances 0.000 description 5
- 238000003753 real-time PCR Methods 0.000 description 5
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 5
- 208000035143 Bacterial infection Diseases 0.000 description 4
- 241000282412 Homo Species 0.000 description 4
- 101000831496 Homo sapiens Toll-like receptor 3 Proteins 0.000 description 4
- 102100024324 Toll-like receptor 3 Human genes 0.000 description 4
- 102000004142 Trypsin Human genes 0.000 description 4
- 108090000631 Trypsin Proteins 0.000 description 4
- 230000009471 action Effects 0.000 description 4
- 239000000654 additive Substances 0.000 description 4
- 208000022362 bacterial infectious disease Diseases 0.000 description 4
- 239000003153 chemical reaction reagent Substances 0.000 description 4
- 235000013376 functional food Nutrition 0.000 description 4
- 238000011534 incubation Methods 0.000 description 4
- 210000002490 intestinal epithelial cell Anatomy 0.000 description 4
- 235000014655 lactic acid Nutrition 0.000 description 4
- 239000004310 lactic acid Substances 0.000 description 4
- BASFCYQUMIYNBI-UHFFFAOYSA-N platinum Chemical compound [Pt] BASFCYQUMIYNBI-UHFFFAOYSA-N 0.000 description 4
- 239000000843 powder Substances 0.000 description 4
- 235000013406 prebiotics Nutrition 0.000 description 4
- 230000009467 reduction Effects 0.000 description 4
- 238000011160 research Methods 0.000 description 4
- DAEPDZWVDSPTHF-UHFFFAOYSA-M sodium pyruvate Chemical compound [Na+].CC(=O)C([O-])=O DAEPDZWVDSPTHF-UHFFFAOYSA-M 0.000 description 4
- 230000004936 stimulating effect Effects 0.000 description 4
- 239000000126 substance Substances 0.000 description 4
- 239000012588 trypsin Substances 0.000 description 4
- 241000712461 unidentified influenza virus Species 0.000 description 4
- 101710127675 Antiviral innate immune response receptor RIG-I Proteins 0.000 description 3
- 241000283690 Bos taurus Species 0.000 description 3
- 208000035473 Communicable disease Diseases 0.000 description 3
- 241000711573 Coronaviridae Species 0.000 description 3
- 102100038284 Cytospin-B Human genes 0.000 description 3
- 102000004190 Enzymes Human genes 0.000 description 3
- 108090000790 Enzymes Proteins 0.000 description 3
- 101001125032 Homo sapiens Nucleotide-binding oligomerization domain-containing protein 1 Proteins 0.000 description 3
- 101150113091 NSP5 gene Proteins 0.000 description 3
- 101710144121 Non-structural protein 5 Proteins 0.000 description 3
- 102100029424 Nucleotide-binding oligomerization domain-containing protein 1 Human genes 0.000 description 3
- 241001137860 Rotavirus A Species 0.000 description 3
- 241000282887 Suidae Species 0.000 description 3
- 108010047933 Tumor Necrosis Factor alpha-Induced Protein 3 Proteins 0.000 description 3
- 102000007150 Tumor Necrosis Factor alpha-Induced Protein 3 Human genes 0.000 description 3
- 230000002953 anti-rotaviral effect Effects 0.000 description 3
- 230000007416 antiviral immune response Effects 0.000 description 3
- 238000012790 confirmation Methods 0.000 description 3
- 230000003247 decreasing effect Effects 0.000 description 3
- 210000000286 epitheliocyte Anatomy 0.000 description 3
- 210000001035 gastrointestinal tract Anatomy 0.000 description 3
- 235000011187 glycerol Nutrition 0.000 description 3
- 150000002632 lipids Chemical class 0.000 description 3
- 229940115272 polyinosinic:polycytidylic acid Drugs 0.000 description 3
- 102000004169 proteins and genes Human genes 0.000 description 3
- 239000002904 solvent Substances 0.000 description 3
- IJGRMHOSHXDMSA-UHFFFAOYSA-N Atomic nitrogen Chemical compound N#N IJGRMHOSHXDMSA-UHFFFAOYSA-N 0.000 description 2
- 108091003079 Bovine Serum Albumin Proteins 0.000 description 2
- 102100021943 C-C motif chemokine 2 Human genes 0.000 description 2
- OKTJSMMVPCPJKN-UHFFFAOYSA-N Carbon Chemical compound [C] OKTJSMMVPCPJKN-UHFFFAOYSA-N 0.000 description 2
- 241000195493 Cryptophyta Species 0.000 description 2
- 102000053602 DNA Human genes 0.000 description 2
- 102100034121 E3 ubiquitin-protein ligase RNF125 Human genes 0.000 description 2
- 108091092584 GDNA Proteins 0.000 description 2
- WQZGKKKJIJFFOK-GASJEMHNSA-N Glucose Natural products OC[C@H]1OC(O)[C@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-GASJEMHNSA-N 0.000 description 2
- 101001082073 Homo sapiens Interferon-induced helicase C domain-containing protein 1 Proteins 0.000 description 2
- 101000831708 Homo sapiens Ubiquitin carboxyl-terminal hydrolase CYLD Proteins 0.000 description 2
- 108010032038 Interferon Regulatory Factor-3 Proteins 0.000 description 2
- 102100029843 Interferon regulatory factor 3 Human genes 0.000 description 2
- 102100027353 Interferon-induced helicase C domain-containing protein 1 Human genes 0.000 description 2
- ZDXPYRJPNDTMRX-VKHMYHEASA-N L-glutamine Chemical compound OC(=O)[C@@H](N)CCC(N)=O ZDXPYRJPNDTMRX-VKHMYHEASA-N 0.000 description 2
- 229930182816 L-glutamine Natural products 0.000 description 2
- 241000914114 Lactobacillus hayakitensis Species 0.000 description 2
- 241000127282 Middle East respiratory syndrome-related coronavirus Species 0.000 description 2
- 241001494479 Pecora Species 0.000 description 2
- 238000010806 PrimeScriptTM RT Reagent kit Methods 0.000 description 2
- 238000011529 RT qPCR Methods 0.000 description 2
- 241000206572 Rhodophyta Species 0.000 description 2
- FAPWRFPIFSIZLT-UHFFFAOYSA-M Sodium chloride Chemical compound [Na+].[Cl-] FAPWRFPIFSIZLT-UHFFFAOYSA-M 0.000 description 2
- 102100024250 Ubiquitin carboxyl-terminal hydrolase CYLD Human genes 0.000 description 2
- 230000000996 additive effect Effects 0.000 description 2
- 230000000844 anti-bacterial effect Effects 0.000 description 2
- 230000000692 anti-sense effect Effects 0.000 description 2
- WQZGKKKJIJFFOK-VFUOTHLCSA-N beta-D-glucose Chemical compound OC[C@H]1O[C@@H](O)[C@H](O)[C@@H](O)[C@@H]1O WQZGKKKJIJFFOK-VFUOTHLCSA-N 0.000 description 2
- 230000037396 body weight Effects 0.000 description 2
- 239000000872 buffer Substances 0.000 description 2
- 150000001720 carbohydrates Chemical class 0.000 description 2
- 229910052799 carbon Inorganic materials 0.000 description 2
- 238000004113 cell culture Methods 0.000 description 2
- 210000002421 cell wall Anatomy 0.000 description 2
- 238000006243 chemical reaction Methods 0.000 description 2
- 238000007796 conventional method Methods 0.000 description 2
- 239000012228 culture supernatant Substances 0.000 description 2
- 230000001086 cytosolic effect Effects 0.000 description 2
- 235000015872 dietary supplement Nutrition 0.000 description 2
- 239000003814 drug Substances 0.000 description 2
- 239000012091 fetal bovine serum Substances 0.000 description 2
- 239000008103 glucose Substances 0.000 description 2
- 244000005709 gut microbiome Species 0.000 description 2
- 235000013402 health food Nutrition 0.000 description 2
- 230000003308 immunostimulating effect Effects 0.000 description 2
- 230000006872 improvement Effects 0.000 description 2
- 230000001939 inductive effect Effects 0.000 description 2
- 210000000936 intestine Anatomy 0.000 description 2
- 239000000314 lubricant Substances 0.000 description 2
- HQKMJHAJHXVSDF-UHFFFAOYSA-L magnesium stearate Chemical compound [Mg+2].CCCCCCCCCCCCCCCCCC([O-])=O.CCCCCCCCCCCCCCCCCC([O-])=O HQKMJHAJHXVSDF-UHFFFAOYSA-L 0.000 description 2
- 239000002207 metabolite Substances 0.000 description 2
- 238000002493 microarray Methods 0.000 description 2
- 235000016709 nutrition Nutrition 0.000 description 2
- 239000000546 pharmaceutical excipient Substances 0.000 description 2
- 239000002504 physiological saline solution Substances 0.000 description 2
- 229910052697 platinum Inorganic materials 0.000 description 2
- 102000040430 polynucleotide Human genes 0.000 description 2
- 108091033319 polynucleotide Proteins 0.000 description 2
- 239000002157 polynucleotide Substances 0.000 description 2
- 229920001184 polypeptide Polymers 0.000 description 2
- 102000004196 processed proteins & peptides Human genes 0.000 description 2
- 108090000765 processed proteins & peptides Proteins 0.000 description 2
- 239000000523 sample Substances 0.000 description 2
- 210000000813 small intestine Anatomy 0.000 description 2
- 229940054269 sodium pyruvate Drugs 0.000 description 2
- UCSJYZPVAKXKNQ-HZYVHMACSA-N streptomycin Chemical compound CN[C@H]1[C@H](O)[C@@H](O)[C@H](CO)O[C@H]1O[C@@H]1[C@](C=O)(O)[C@H](C)O[C@H]1O[C@@H]1[C@@H](NC(N)=N)[C@H](O)[C@@H](NC(N)=N)[C@H](O)[C@H]1O UCSJYZPVAKXKNQ-HZYVHMACSA-N 0.000 description 2
- 239000006228 supernatant Substances 0.000 description 2
- 230000001629 suppression Effects 0.000 description 2
- HDTRYLNUVZCQOY-UHFFFAOYSA-N α-D-glucopyranosyl-α-D-glucopyranoside Natural products OC1C(O)C(O)C(CO)OC1OC1C(O)C(O)C(O)C(CO)O1 HDTRYLNUVZCQOY-UHFFFAOYSA-N 0.000 description 1
- 239000001707 (E,7R,11R)-3,7,11,15-tetramethylhexadec-2-en-1-ol Substances 0.000 description 1
- MWZTVLNYXAKUKY-LBEKAKSKSA-N 4-hydroxy-N-[2-[(1R,13S)-3-methyl-8-oxo-11-azatetracyclo[8.4.0.01,13.02,7]tetradeca-2,4,6,9-tetraene-11-carbonyl]imidazo[1,2-a]pyridin-6-yl]benzamide Chemical compound C=1([C@]23C[C@@H]3C3)C(C)=CC=CC=1C(=O)C=C2N3C(=O)C(N=C1C=C2)=CN1C=C2NC(=O)C1=CC=C(O)C=C1 MWZTVLNYXAKUKY-LBEKAKSKSA-N 0.000 description 1
- 102100022900 Actin, cytoplasmic 1 Human genes 0.000 description 1
- 235000019737 Animal fat Nutrition 0.000 description 1
- 102000052666 B-Cell Lymphoma 3 Human genes 0.000 description 1
- 241000193749 Bacillus coagulans Species 0.000 description 1
- 101710155857 C-C motif chemokine 2 Proteins 0.000 description 1
- 208000025721 COVID-19 Diseases 0.000 description 1
- 241001678559 COVID-19 virus Species 0.000 description 1
- 241000282472 Canis lupus familiaris Species 0.000 description 1
- 241000283707 Capra Species 0.000 description 1
- 241000700199 Cavia porcellus Species 0.000 description 1
- 241000282693 Cercopithecidae Species 0.000 description 1
- 241000282994 Cervidae Species 0.000 description 1
- 108010093668 Deubiquitinating Enzymes Proteins 0.000 description 1
- 102000001477 Deubiquitinating Enzymes Human genes 0.000 description 1
- 229920002307 Dextran Polymers 0.000 description 1
- IAZDPXIOMUYVGZ-UHFFFAOYSA-N Dimethylsulphoxide Chemical compound CS(C)=O IAZDPXIOMUYVGZ-UHFFFAOYSA-N 0.000 description 1
- 208000010772 Dog disease Diseases 0.000 description 1
- 102100034428 Dual specificity protein phosphatase 1 Human genes 0.000 description 1
- 101710132784 Dual specificity protein phosphatase 1 Proteins 0.000 description 1
- 101710162555 E3 ubiquitin-protein ligase RNF125 Proteins 0.000 description 1
- KCXVZYZYPLLWCC-UHFFFAOYSA-N EDTA Chemical compound OC(=O)CN(CC(O)=O)CCN(CC(O)=O)CC(O)=O KCXVZYZYPLLWCC-UHFFFAOYSA-N 0.000 description 1
- 238000002965 ELISA Methods 0.000 description 1
- 241000194033 Enterococcus Species 0.000 description 1
- 241000194032 Enterococcus faecalis Species 0.000 description 1
- 241000283086 Equidae Species 0.000 description 1
- 241000282326 Felis catus Species 0.000 description 1
- 241000192125 Firmicutes Species 0.000 description 1
- 241000233866 Fungi Species 0.000 description 1
- 108010010803 Gelatin Proteins 0.000 description 1
- 241000711549 Hepacivirus C Species 0.000 description 1
- 241000700721 Hepatitis B virus Species 0.000 description 1
- 241000724675 Hepatitis E virus Species 0.000 description 1
- 241000709721 Hepatovirus A Species 0.000 description 1
- 101000711567 Homo sapiens E3 ubiquitin-protein ligase RNF125 Proteins 0.000 description 1
- 101001055222 Homo sapiens Interleukin-8 Proteins 0.000 description 1
- 101000652846 Homo sapiens Single Ig IL-1-related receptor Proteins 0.000 description 1
- 241000725303 Human immunodeficiency virus Species 0.000 description 1
- 229920002153 Hydroxypropyl cellulose Polymers 0.000 description 1
- 102100040019 Interferon alpha-1/13 Human genes 0.000 description 1
- 102000003996 Interferon-beta Human genes 0.000 description 1
- 102100036342 Interleukin-1 receptor-associated kinase 1 Human genes 0.000 description 1
- 101710199015 Interleukin-1 receptor-associated kinase 1 Proteins 0.000 description 1
- 101710199012 Interleukin-1 receptor-associated kinase 3 Proteins 0.000 description 1
- 102100023530 Interleukin-1 receptor-associated kinase 3 Human genes 0.000 description 1
- 108090001005 Interleukin-6 Proteins 0.000 description 1
- 108090001007 Interleukin-8 Proteins 0.000 description 1
- 102000004890 Interleukin-8 Human genes 0.000 description 1
- 102100026236 Interleukin-8 Human genes 0.000 description 1
- 238000007397 LAMP assay Methods 0.000 description 1
- 241000186716 Lactobacillus agilis Species 0.000 description 1
- 241000186711 Lactobacillus aviarius Species 0.000 description 1
- 240000001929 Lactobacillus brevis Species 0.000 description 1
- 235000013957 Lactobacillus brevis Nutrition 0.000 description 1
- 241000057444 Lactobacillus brevis subsp. coagulans Species 0.000 description 1
- 241000186840 Lactobacillus fermentum Species 0.000 description 1
- 241000186606 Lactobacillus gasseri Species 0.000 description 1
- 241000186605 Lactobacillus paracasei Species 0.000 description 1
- GUBGYTABKSRVRQ-QKKXKWKRSA-N Lactose Natural products OC[C@H]1O[C@@H](O[C@H]2[C@H](O)[C@@H](O)C(O)O[C@@H]2CO)[C@H](O)[C@@H](O)[C@H]1O GUBGYTABKSRVRQ-QKKXKWKRSA-N 0.000 description 1
- 241000192132 Leuconostoc Species 0.000 description 1
- 241000712079 Measles morbillivirus Species 0.000 description 1
- 208000025370 Middle East respiratory syndrome Diseases 0.000 description 1
- 101000971179 Mus musculus B-cell lymphoma 3 protein homolog Proteins 0.000 description 1
- 241000699670 Mus sp. Species 0.000 description 1
- 241000204031 Mycoplasma Species 0.000 description 1
- 241001263478 Norovirus Species 0.000 description 1
- 238000000636 Northern blotting Methods 0.000 description 1
- 101710089460 OTU domain-containing protein 5 Proteins 0.000 description 1
- 102100025194 OTU domain-containing protein 5 Human genes 0.000 description 1
- 101710090551 OTU domain-containing protein 5-A Proteins 0.000 description 1
- 241000283973 Oryctolagus cuniculus Species 0.000 description 1
- 240000007594 Oryza sativa Species 0.000 description 1
- 235000007164 Oryza sativa Nutrition 0.000 description 1
- 241000192001 Pediococcus Species 0.000 description 1
- 229930182555 Penicillin Natural products 0.000 description 1
- JGSARLDLIJGVTE-MBNYWOFBSA-N Penicillin G Chemical compound N([C@H]1[C@H]2SC([C@@H](N2C1=O)C(O)=O)(C)C)C(=O)CC1=CC=CC=C1 JGSARLDLIJGVTE-MBNYWOFBSA-N 0.000 description 1
- BLUHKGOSFDHHGX-UHFFFAOYSA-N Phytol Natural products CC(C)CCCC(C)CCCC(C)CCCC(C)C=CO BLUHKGOSFDHHGX-UHFFFAOYSA-N 0.000 description 1
- 241000255969 Pieris brassicae Species 0.000 description 1
- 239000004372 Polyvinyl alcohol Substances 0.000 description 1
- 208000009341 RNA Virus Infections Diseases 0.000 description 1
- 241000700159 Rattus Species 0.000 description 1
- 241000315672 SARS coronavirus Species 0.000 description 1
- 201000003176 Severe Acute Respiratory Syndrome Diseases 0.000 description 1
- 102100030929 Single Ig IL-1-related receptor Human genes 0.000 description 1
- 108020004682 Single-Stranded DNA Proteins 0.000 description 1
- 238000002105 Southern blotting Methods 0.000 description 1
- 229920002472 Starch Polymers 0.000 description 1
- 229930006000 Sucrose Natural products 0.000 description 1
- CZMRCDWAGMRECN-UGDNZRGBSA-N Sucrose Chemical compound O[C@H]1[C@H](O)[C@@H](CO)O[C@@]1(CO)O[C@@H]1[C@H](O)[C@@H](O)[C@H](O)[C@@H](CO)O1 CZMRCDWAGMRECN-UGDNZRGBSA-N 0.000 description 1
- HNZBNQYXWOLKBA-UHFFFAOYSA-N Tetrahydrofarnesol Natural products CC(C)CCCC(C)CCCC(C)=CCO HNZBNQYXWOLKBA-UHFFFAOYSA-N 0.000 description 1
- HDTRYLNUVZCQOY-WSWWMNSNSA-N Trehalose Natural products O[C@@H]1[C@@H](O)[C@@H](O)[C@@H](CO)O[C@@H]1O[C@@H]1[C@H](O)[C@@H](O)[C@@H](O)[C@@H](CO)O1 HDTRYLNUVZCQOY-WSWWMNSNSA-N 0.000 description 1
- 108060008682 Tumor Necrosis Factor Proteins 0.000 description 1
- 102000000852 Tumor Necrosis Factor-alpha Human genes 0.000 description 1
- 230000003213 activating effect Effects 0.000 description 1
- BOTWFXYSPFMFNR-OALUTQOASA-N all-rac-phytol Natural products CC(C)CCC[C@H](C)CCC[C@H](C)CCCC(C)=CCO BOTWFXYSPFMFNR-OALUTQOASA-N 0.000 description 1
- HDTRYLNUVZCQOY-LIZSDCNHSA-N alpha,alpha-trehalose Chemical compound O[C@@H]1[C@@H](O)[C@H](O)[C@@H](CO)O[C@@H]1O[C@@H]1[C@H](O)[C@@H](O)[C@H](O)[C@@H](CO)O1 HDTRYLNUVZCQOY-LIZSDCNHSA-N 0.000 description 1
- 239000003708 ampul Substances 0.000 description 1
- 239000003242 anti bacterial agent Substances 0.000 description 1
- 230000002429 anti-coagulating effect Effects 0.000 description 1
- 229940088710 antibiotic agent Drugs 0.000 description 1
- 239000003146 anticoagulant agent Substances 0.000 description 1
- 229940127219 anticoagulant drug Drugs 0.000 description 1
- 230000003078 antioxidant effect Effects 0.000 description 1
- 239000007864 aqueous solution Substances 0.000 description 1
- 229940054340 bacillus coagulans Drugs 0.000 description 1
- 230000008901 benefit Effects 0.000 description 1
- 239000011230 binding agent Substances 0.000 description 1
- 239000012620 biological material Substances 0.000 description 1
- 230000033228 biological regulation Effects 0.000 description 1
- 210000004369 blood Anatomy 0.000 description 1
- 239000008280 blood Substances 0.000 description 1
- 238000004422 calculation algorithm Methods 0.000 description 1
- 229940041514 candida albicans extract Drugs 0.000 description 1
- 239000013592 cell lysate Substances 0.000 description 1
- 210000000170 cell membrane Anatomy 0.000 description 1
- 239000001913 cellulose Substances 0.000 description 1
- 229920002678 cellulose Polymers 0.000 description 1
- 238000005119 centrifugation Methods 0.000 description 1
- 230000001886 ciliary effect Effects 0.000 description 1
- 239000011248 coating agent Substances 0.000 description 1
- 230000001332 colony forming effect Effects 0.000 description 1
- 230000002860 competitive effect Effects 0.000 description 1
- 238000013329 compounding Methods 0.000 description 1
- 230000001351 cycling effect Effects 0.000 description 1
- 230000001461 cytolytic effect Effects 0.000 description 1
- 238000001514 detection method Methods 0.000 description 1
- 238000010586 diagram Methods 0.000 description 1
- 239000003085 diluting agent Substances 0.000 description 1
- 201000010099 disease Diseases 0.000 description 1
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 1
- 239000007884 disintegrant Substances 0.000 description 1
- BNIILDVGGAEEIG-UHFFFAOYSA-L disodium hydrogen phosphate Chemical compound [Na+].[Na+].OP([O-])([O-])=O BNIILDVGGAEEIG-UHFFFAOYSA-L 0.000 description 1
- 239000003937 drug carrier Substances 0.000 description 1
- 235000005686 eating Nutrition 0.000 description 1
- 235000006694 eating habits Nutrition 0.000 description 1
- 230000000531 effect on virus Effects 0.000 description 1
- 229940032049 enterococcus faecalis Drugs 0.000 description 1
- 230000000688 enterotoxigenic effect Effects 0.000 description 1
- LYCAIKOWRPUZTN-UHFFFAOYSA-N ethylene glycol Natural products OCCO LYCAIKOWRPUZTN-UHFFFAOYSA-N 0.000 description 1
- 230000001747 exhibiting effect Effects 0.000 description 1
- 239000000284 extract Substances 0.000 description 1
- 238000000855 fermentation Methods 0.000 description 1
- 230000004151 fermentation Effects 0.000 description 1
- 238000011049 filling Methods 0.000 description 1
- 239000000796 flavoring agent Substances 0.000 description 1
- 238000000684 flow cytometry Methods 0.000 description 1
- 238000002637 fluid replacement therapy Methods 0.000 description 1
- 235000012041 food component Nutrition 0.000 description 1
- 235000013355 food flavoring agent Nutrition 0.000 description 1
- 235000003599 food sweetener Nutrition 0.000 description 1
- 238000009472 formulation Methods 0.000 description 1
- 235000014106 fortified food Nutrition 0.000 description 1
- 238000004108 freeze drying Methods 0.000 description 1
- 238000007710 freezing Methods 0.000 description 1
- 230000008014 freezing Effects 0.000 description 1
- 239000008273 gelatin Substances 0.000 description 1
- 229920000159 gelatin Polymers 0.000 description 1
- 235000019322 gelatine Nutrition 0.000 description 1
- 235000011852 gelatine desserts Nutrition 0.000 description 1
- 239000003349 gelling agent Substances 0.000 description 1
- 210000000003 hoof Anatomy 0.000 description 1
- WGCNASOHLSPBMP-UHFFFAOYSA-N hydroxyacetaldehyde Natural products OCC=O WGCNASOHLSPBMP-UHFFFAOYSA-N 0.000 description 1
- 239000001863 hydroxypropyl cellulose Substances 0.000 description 1
- 235000010977 hydroxypropyl cellulose Nutrition 0.000 description 1
- 230000008975 immunomodulatory function Effects 0.000 description 1
- 230000004957 immunoregulator effect Effects 0.000 description 1
- 230000000415 inactivating effect Effects 0.000 description 1
- 230000002458 infectious effect Effects 0.000 description 1
- 239000004615 ingredient Substances 0.000 description 1
- 239000000138 intercalating agent Substances 0.000 description 1
- 229960001388 interferon-beta Drugs 0.000 description 1
- 230000011542 interferon-beta production Effects 0.000 description 1
- 210000004347 intestinal mucosa Anatomy 0.000 description 1
- 238000001990 intravenous administration Methods 0.000 description 1
- FZWBNHMXJMCXLU-BLAUPYHCSA-N isomaltotriose Chemical compound O[C@@H]1[C@@H](O)[C@H](O)[C@@H](CO)O[C@@H]1OC[C@@H]1[C@@H](O)[C@H](O)[C@@H](O)[C@@H](OC[C@@H](O)[C@@H](O)[C@H](O)[C@@H](O)C=O)O1 FZWBNHMXJMCXLU-BLAUPYHCSA-N 0.000 description 1
- 239000007951 isotonicity adjuster Substances 0.000 description 1
- 229940012969 lactobacillus fermentum Drugs 0.000 description 1
- 239000008101 lactose Substances 0.000 description 1
- 239000003446 ligand Substances 0.000 description 1
- 230000002101 lytic effect Effects 0.000 description 1
- 235000019359 magnesium stearate Nutrition 0.000 description 1
- 230000007246 mechanism Effects 0.000 description 1
- 210000004379 membrane Anatomy 0.000 description 1
- 239000012528 membrane Substances 0.000 description 1
- 230000002503 metabolic effect Effects 0.000 description 1
- 230000000813 microbial effect Effects 0.000 description 1
- 238000002156 mixing Methods 0.000 description 1
- 210000003097 mucus Anatomy 0.000 description 1
- 229910052757 nitrogen Inorganic materials 0.000 description 1
- 102000039446 nucleic acids Human genes 0.000 description 1
- 108020004707 nucleic acids Proteins 0.000 description 1
- 150000007523 nucleic acids Chemical class 0.000 description 1
- 235000015097 nutrients Nutrition 0.000 description 1
- 239000003921 oil Substances 0.000 description 1
- 235000019198 oils Nutrition 0.000 description 1
- 150000007524 organic acids Chemical class 0.000 description 1
- 235000005985 organic acids Nutrition 0.000 description 1
- 239000006174 pH buffer Substances 0.000 description 1
- 238000007911 parenteral administration Methods 0.000 description 1
- 229940049954 penicillin Drugs 0.000 description 1
- 239000008194 pharmaceutical composition Substances 0.000 description 1
- 239000000825 pharmaceutical preparation Substances 0.000 description 1
- 229940127557 pharmaceutical product Drugs 0.000 description 1
- 230000026731 phosphorylation Effects 0.000 description 1
- 238000006366 phosphorylation reaction Methods 0.000 description 1
- BOTWFXYSPFMFNR-PYDDKJGSSA-N phytol Chemical compound CC(C)CCC[C@@H](C)CCC[C@@H](C)CCC\C(C)=C\CO BOTWFXYSPFMFNR-PYDDKJGSSA-N 0.000 description 1
- 229920002451 polyvinyl alcohol Polymers 0.000 description 1
- 230000003449 preventive effect Effects 0.000 description 1
- 230000000529 probiotic effect Effects 0.000 description 1
- 230000008569 process Effects 0.000 description 1
- 230000035755 proliferation Effects 0.000 description 1
- 230000000069 prophylactic effect Effects 0.000 description 1
- 239000003223 protective agent Substances 0.000 description 1
- 230000009257 reactivity Effects 0.000 description 1
- 238000010839 reverse transcription Methods 0.000 description 1
- 235000009566 rice Nutrition 0.000 description 1
- 230000035807 sensation Effects 0.000 description 1
- 230000035945 sensitivity Effects 0.000 description 1
- 238000012163 sequencing technique Methods 0.000 description 1
- 210000002966 serum Anatomy 0.000 description 1
- 230000001568 sexual effect Effects 0.000 description 1
- 235000015170 shellfish Nutrition 0.000 description 1
- 230000011664 signaling Effects 0.000 description 1
- 239000011780 sodium chloride Substances 0.000 description 1
- 241000894007 species Species 0.000 description 1
- 239000003381 stabilizer Substances 0.000 description 1
- 239000008107 starch Substances 0.000 description 1
- 235000019698 starch Nutrition 0.000 description 1
- 230000001954 sterilising effect Effects 0.000 description 1
- 238000004659 sterilization and disinfection Methods 0.000 description 1
- 229960005322 streptomycin Drugs 0.000 description 1
- 239000000758 substrate Substances 0.000 description 1
- 239000005720 sucrose Substances 0.000 description 1
- 235000000346 sugar Nutrition 0.000 description 1
- 150000008163 sugars Chemical class 0.000 description 1
- 239000013589 supplement Substances 0.000 description 1
- 239000003765 sweetening agent Substances 0.000 description 1
- 235000019722 synbiotics Nutrition 0.000 description 1
- 239000000454 talc Substances 0.000 description 1
- 229910052623 talc Inorganic materials 0.000 description 1
- 235000012222 talc Nutrition 0.000 description 1
- UEUXEKPTXMALOB-UHFFFAOYSA-J tetrasodium;2-[2-[bis(carboxylatomethyl)amino]ethyl-(carboxylatomethyl)amino]acetate Chemical compound [Na+].[Na+].[Na+].[Na+].[O-]C(=O)CN(CC([O-])=O)CCN(CC([O-])=O)CC([O-])=O UEUXEKPTXMALOB-UHFFFAOYSA-J 0.000 description 1
- 229940124597 therapeutic agent Drugs 0.000 description 1
- 210000001519 tissue Anatomy 0.000 description 1
- 231100000033 toxigenic Toxicity 0.000 description 1
- 230000001551 toxigenic effect Effects 0.000 description 1
- 239000007195 tryptone soya broth Substances 0.000 description 1
- 241000701161 unidentified adenovirus Species 0.000 description 1
- 241001529453 unidentified herpesvirus Species 0.000 description 1
- 229960005486 vaccine Drugs 0.000 description 1
- 235000015112 vegetable and seed oil Nutrition 0.000 description 1
- 239000008158 vegetable oil Substances 0.000 description 1
- 230000003612 virological effect Effects 0.000 description 1
- 238000001262 western blot Methods 0.000 description 1
- 239000012138 yeast extract Substances 0.000 description 1
Images
Classifications
-
- A—HUMAN NECESSITIES
- A23—FOODS OR FOODSTUFFS; TREATMENT THEREOF, NOT COVERED BY OTHER CLASSES
- A23L—FOODS, FOODSTUFFS, OR NON-ALCOHOLIC BEVERAGES, NOT COVERED BY SUBCLASSES A21D OR A23B-A23J; THEIR PREPARATION OR TREATMENT, e.g. COOKING, MODIFICATION OF NUTRITIVE QUALITIES, PHYSICAL TREATMENT; PRESERVATION OF FOODS OR FOODSTUFFS, IN GENERAL
- A23L33/00—Modifying nutritive qualities of foods; Dietetic products; Preparation or treatment thereof
- A23L33/10—Modifying nutritive qualities of foods; Dietetic products; Preparation or treatment thereof using additives
- A23L33/135—Bacteria or derivatives thereof, e.g. probiotics
-
- A—HUMAN NECESSITIES
- A23—FOODS OR FOODSTUFFS; TREATMENT THEREOF, NOT COVERED BY OTHER CLASSES
- A23K—FODDER
- A23K10/00—Animal feeding-stuffs
- A23K10/10—Animal feeding-stuffs obtained by microbiological or biochemical processes
- A23K10/16—Addition of microorganisms or extracts thereof, e.g. single-cell proteins, to feeding-stuff compositions
- A23K10/18—Addition of microorganisms or extracts thereof, e.g. single-cell proteins, to feeding-stuff compositions of live microorganisms
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K35/00—Medicinal preparations containing materials or reaction products thereof with undetermined constitution
- A61K35/66—Microorganisms or materials therefrom
- A61K35/74—Bacteria
- A61K35/741—Probiotics
- A61K35/744—Lactic acid bacteria, e.g. enterococci, pediococci, lactococci, streptococci or leuconostocs
- A61K35/747—Lactobacilli, e.g. L. acidophilus or L. brevis
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K9/00—Medicinal preparations characterised by special physical form
- A61K9/0012—Galenical forms characterised by the site of application
- A61K9/0053—Mouth and digestive tract, i.e. intraoral and peroral administration
- A61K9/0056—Mouth soluble or dispersible forms; Suckable, eatable, chewable coherent forms; Forms rapidly disintegrating in the mouth; Lozenges; Lollipops; Bite capsules; Baked products; Baits or other oral forms for animals
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P31/00—Antiinfectives, i.e. antibiotics, antiseptics, chemotherapeutics
- A61P31/12—Antivirals
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P31/00—Antiinfectives, i.e. antibiotics, antiseptics, chemotherapeutics
- A61P31/12—Antivirals
- A61P31/14—Antivirals for RNA viruses
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N1/00—Microorganisms, e.g. protozoa; Compositions thereof; Processes of propagating, maintaining or preserving microorganisms or compositions thereof; Processes of preparing or isolating a composition containing a microorganism; Culture media therefor
- C12N1/20—Bacteria; Culture media therefor
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N1/00—Microorganisms, e.g. protozoa; Compositions thereof; Processes of propagating, maintaining or preserving microorganisms or compositions thereof; Processes of preparing or isolating a composition containing a microorganism; Culture media therefor
- C12N1/20—Bacteria; Culture media therefor
- C12N1/205—Bacterial isolates
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- A—HUMAN NECESSITIES
- A23—FOODS OR FOODSTUFFS; TREATMENT THEREOF, NOT COVERED BY OTHER CLASSES
- A23V—INDEXING SCHEME RELATING TO FOODS, FOODSTUFFS OR NON-ALCOHOLIC BEVERAGES AND LACTIC OR PROPIONIC ACID BACTERIA USED IN FOODSTUFFS OR FOOD PREPARATION
- A23V2002/00—Food compositions, function of food ingredients or processes for food or foodstuffs
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K35/00—Medicinal preparations containing materials or reaction products thereof with undetermined constitution
- A61K2035/11—Medicinal preparations comprising living procariotic cells
- A61K2035/115—Probiotics
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12R—INDEXING SCHEME ASSOCIATED WITH SUBCLASSES C12C - C12Q, RELATING TO MICROORGANISMS
- C12R2001/00—Microorganisms ; Processes using microorganisms
- C12R2001/01—Bacteria or Actinomycetales ; using bacteria or Actinomycetales
- C12R2001/225—Lactobacillus
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12R—INDEXING SCHEME ASSOCIATED WITH SUBCLASSES C12C - C12Q, RELATING TO MICROORGANISMS
- C12R2001/00—Microorganisms ; Processes using microorganisms
- C12R2001/01—Bacteria or Actinomycetales ; using bacteria or Actinomycetales
- C12R2001/225—Lactobacillus
- C12R2001/25—Lactobacillus plantarum
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y02—TECHNOLOGIES OR APPLICATIONS FOR MITIGATION OR ADAPTATION AGAINST CLIMATE CHANGE
- Y02A—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE
- Y02A50/00—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE in human health protection, e.g. against extreme weather
- Y02A50/30—Against vector-borne diseases, e.g. mosquito-borne, fly-borne, tick-borne or waterborne diseases whose impact is exacerbated by climate change
Definitions
- the present invention relates to an antiviral agent containing antiviral probiotics (preferably immunobiotics [that is, probiotics having a mucosal immunomodulatory function such as intestinal mucosa] as an active ingredient].
- antiviral probiotics preferably immunobiotics [that is, probiotics having a mucosal immunomodulatory function such as intestinal mucosa] as an active ingredient].
- Probiotics are living microorganisms that act on the normal bacterial flora in the body of the host mammal (especially in the intestinal tract) and improve the balance to benefit the body of the mammal.
- Probiotics induce the production of bactericidal substances, competitive intake of nutritional components, competition of attachment sites, promotion / suppression of metabolic enzyme activity, etc. against infectious diseases of viruses and pathogenic bacteria, and have bactericidal effects. It is known to exert an immunostimulatory effect.
- probiotics include lactic acid bacteria such as Lactobacillus, Biffidobacterium, Enterococcus, Leuconostoc, and Pediococcus.
- lactic acid bacteria such as Lactobacillus, Biffidobacterium, Enterococcus, Leuconostoc, and Pediococcus.
- an immunostimulatory composition containing a cytoplasmic fraction of a lactic acid bacterium cell or a cytoplasmic fraction has been reported (Patent Document 1).
- Coagulans has an interferon (IFN) - ⁇ and IFN- ⁇ expression enhancing action and a natural killer (NK) cell activating action.
- IFN interferon
- NK natural killer
- lactic acid bacteria such as Bacillus coagulans have an antiviral effect against infectious diseases of sensation virus and influenza virus
- lactic acid bacteria such as Lactobacillus plantarum, Lactobacillus ramnosus, Lactobacillus fermentum, Lactobacillus paracasei, and Lactobacillus gasseri can be used to prevent or treat virus infection. It has been reported that it can be used (Patent Document 4). Further, it has been reported that the combined use of Labre bacterium and Enterococcus faecalis exerts an excellent preventive effect against influenza virus infectious diseases (Patent Document 5). However, no probiotics that effectively exert antiviral effects in complex infections of viruses and pathogenic bacteria have been known so far.
- An object of the present invention is to provide an antiviral agent containing probiotics that effectively exert an antiviral effect even in a complex infection of a virus and a pathogenic bacterium as an active ingredient.
- the present inventors are continuing diligent research to solve the above problems.
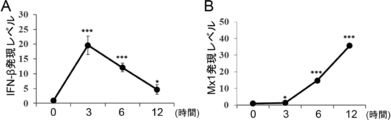
- two kinds of factors IFN- ⁇ and Mx1 were identified as an index for evaluating the antiviral property of probiotics.
- the expression levels of the above two types of factors were used as indicators to show antiviral properties.
- a genus bacterial strain was selected.
- the selected antiviral Lactobacillus bacterial strain has an effect of enhancing the expression of antiviral factors and an effect of reducing the expression of down-regulatory factors of antiviral factors, and also in the combined infection of viruses and pathogenic bacteria.
- a specific Lactobacillus bacterial strain exhibiting wakame assimilation has a 16S rRNA gene having the same nucleotide sequence as the selected antiviral Lactobacillus bacterial strain, and has an effect of enhancing the expression of antiviral factors. Confirmed to have. Furthermore, it is a species different from Lactobacillus salivarius and has at least 90% identity with the nucleotide sequence of the 16S rRNA gene of Lactobacillus salivalius (that is, the nucleotide sequence of SEQ ID NO: 1) that exerted an antiviral effect. It was confirmed that the Lactobacillus plantarum (also referred to as "Lactiplantibacillus plantarum”) strain having the 16S rRNA gene also has an antiviral effect. The present invention has been completed based on these findings.
- the present invention is as follows. [1] Expression of a 16S rRNA gene having at least 90% identity with the nucleotide sequence shown in SEQ ID NO: 1 and an antiviral factor expression-enhancing effect and / or an antiviral factor down-regulatory factor An antiviral agent comprising one or two or more strains of the genus Lactobacillus having a reducing action. [2] Antiviral factors are interferon (IFN) - ⁇ , IFN- ⁇ , MX1 (MX dynamin like GTPase 1), OAS1 (2'-5'-oligoadenylate synthetase 1), RNaseL, PKR (Protein kinase R).
- IFN interferon
- MX1 MX dynamin like GTPase 1
- OAS1 (2'-5'-oligoadenylate synthetase 1
- RNaseL RNaseL
- PKR Protein kinase R
- antiviral factors selected from RIG-I Retinoic acid inducible gene-I
- the down-regulatory factor of the antiviral factor is selected from A20 and Toll-interacting protein.
- the antiviral agent according to the above [1] which is a downward control factor of the antiviral factor of 1 or 2.
- the lactobacillus strain has an action of enhancing the expression of one or more receptors selected from TLR (Toll-like receptor) 2, TLR4, and NOD2 (Nucleotide binding oligomerization domain-like receptor 2).
- TLR Toll-like receptor
- NOD2 Nucleotide binding oligomerization domain-like receptor 2
- Lactobacillus bacterium strain is deposited as Receipt No. NITE ABP-03218; Lactobacillus salivalius strain deposited as Receipt No. NITE ABP-03219; Deposited as Receipt No. NITE ABP-03221.
- Lactobacillus salivalius strain Lactobacillus plantaram strain deposited under receipt number NITE ABP-03474; Lactobacillus plantalum strain deposited under receipt number NITE ABP-03467; deposited as receipt number NITE ABP-03468 Lactobacillus plantalum strains; Lactobacillus plantalum strains deposited as receipt number NITE ABP-03466; Lactobacillus plantalum strains deposited as receipt number NITE ABP-03471; deposits as receipt number NITE ABP-03469 Lactobacillus plantarum strains deposited; Lactobacillus plantaram strains deposited as receipt number NITE ABP-03470; Lactobacillus plantaram strains deposited as receipt number NITE ABP-03472; and receipt number NITE ABP-
- the antiviral agent according to any one of the above [1] to [4], which is one or more selected from the Lactobacillus plantarum strain deposited as 03473; [6]
- One or two or more of the lactobacillus bacterium strains having an expression-reducing effect (hereinafter, may be referred to as "the lactobacillus genus bacterial strain") are infected with a virus (preferably a virus infection in a combined infection of a virus and a pathogenic bacterium).
- Methods of preventing and / or treating a viral infection including the step of administering to a subject (patient) in need of prevention and / or treatment of the disease; One or more Lactobacillus strains for use as antiviral agents; One or more Lactobacillus strains for use in anti-virus; or one or more Lactobacillus strains for use in the prevention and / or treatment of viral infections; , Use of one or more Lactobacillus strains to produce antiviral agents; Use of one or more Lactobacillus bacterial strains to produce prophylactic and / or therapeutic agents for viral infections; Can be mentioned.
- the Lactobacillus bacterial strain has an effect of enhancing the expression of antiviral factors and an effect of reducing the expression of down-regulatory factors of antiviral factors, and effectively exerts an antiviral effect even in a complex infection of a virus and a pathogenic bacterium. Demonstrate.
- the lactobacillus bacterium strain is a bacterial strain (that is, probiotics) that coexists in the living body (particularly, in the intestinal tract) of the host mammal, unlike vaccines and antibiotics, it has side effects. It is possible to effectively prevent / improve (treat) viral infections (preferably viral infections in complex infections of viruses and pathogenic bacteria) in humans and non-human mammals (particularly domestic animals) with little occurrence. ..
- the Lactobacillus bacterium strain has wakame assimilation property
- the Lactobacillus genus bacterial strain preferably antiviral immunobiotics
- wakame (prebiotics) immunophores having wakame assimilation. Since synbiotics promote the growth of the bacterial strain, the antiviral effect of the Lactobacillus bacterial strain can be more effectively exerted.
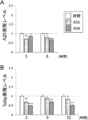
- FIG. 7A-C show the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 35” in the figure) before stimulation with PolyI: C for 6 hours, 3 hours, and 12 hours, respectively. It is a figure which shows the result of having analyzed the A20 expression level (FIG. 7A) and the Tollip expression level (FIGS. 7B and C) in the PIE cell line stimulated by # 58 "). “**” and “***” in the figure are statistically significantly different from the results of stimulation with PolyI: C alone (“Poly (I: C) +” in the figure). (P ⁇ 0.01 and p ⁇ 0.001).
- FIG. 7A shows the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 35” in the figure) before stimulation with PolyI: C for 6 hours, 3 hours, and 12 hours, respectively.
- FIG. 7A shows the result of having analyzed the A20 expression level (FIG. 7A) and the Tollip expression level (FIGS. 7B and C
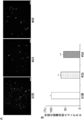
- FIG. 8A shows that the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) is stimulated or these prior to incubation with the activated rotavirus solution.
- 6 is a fluorescent image analyzed by an indirect immunofluorescent antibody method using an anti-rotavirus antibody for a PIE cell line that was not stimulated by the strain (“control” in the figure).
- FIG. 8B is a diagram showing the results of analysis of the proportion of rotavirus-infected cells based on the results of FIG. 8A. The proportion of rotavirus-infected cells is shown as a relative value when the control is 100.
- FIG. 10A is stimulated with the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) before incubating with the activated rotavirus solution for 3 or 6 hours. It is a figure which shows the result of having analyzed the A20 expression level for the PIE cell line which did or did not stimulate with these strains (the "control" in the figure).
- FIG. 10A is stimulated with the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) before incubating with the activated rotavirus solution for 3 or 6 hours. It is a figure which shows the result of having analyzed the A20 expression level for the PIE cell line which did or did not stimulate with these strains (the "control" in the figure).
- FIG. 10A is stimulated with the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) before incubating with the activated rotavirus solution for 3 or 6 hours. It is
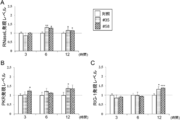
- 10B shows the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) before incubating with the activated rotavirus solution for 3 hours, 6 hours, or 12 hours. It is a figure which shows the result of having analyzed the Tollip expression level of the PIE cell line which was stimulated with (") or was not stimulated by these strains ("control "in the figure). The A20 expression level and the Tollip expression level are shown as relative values when the control is 1. "*”, "**”, and "***” in the figure are statistically significantly different from the control (p ⁇ 0.05, p ⁇ 0.01, and p ⁇ , respectively. 0.001) is shown. FIG.
- FIG. 11A shows the PIE cell line when incubated with activated rotavirus solution alone (“virus” in the figure) or with activated rotavirus solution and ETEC-containing solution (“virus + ETEC” in the figure). It is a figure which shows the result of having analyzed the ratio of the rotavirus-infected cell of "). The proportion of rotavirus-infected cells is shown as a relative value when "virus + ETEC” is set to 100.
- FIG. 11B shows whether to stimulate with the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) before incubating with the activated rotavirus solution and the ETEC-containing solution.
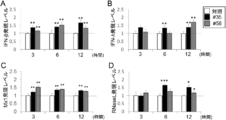
- control For PIE cell lines stimulated with or not stimulated with these strains (“control” in the figure), three antiviral factors (IFN- ⁇ [FIG. 12A], IFN- ⁇ [ It is a figure which shows the result of having analyzed the expression level of FIG. 12B] and Mx1 [FIG. 12C]).
- the expression levels of these antiviral factors are shown as relative values when the control is 1. "*”, "**”, and "***” in the figure are statistically significantly different from the control (p ⁇ 0.05, p ⁇ 0.01, and p ⁇ , respectively. 0.001) is shown.
- 3 types of antiviral factors RNaseL [FIG. 13A], PKR [FIG. 13B], for PIE cell lines stimulated with or not stimulated with these strains (“control” in the figure).
- RIG-1 [FIG. 13C] it is a figure which shows the result of having analyzed the expression level. The expression levels of these antiviral factors are shown as relative values when the control is 1.
- FIG. 14A shows the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) prior to incubation with the activated rotavirus solution and the ETEC-containing solution for 3 or 6 hours. It is a figure which shows the result of having analyzed the A20 expression level about the PIE cell line which was stimulated with (") or was not stimulated by these strains ("control "in the figure).
- FIG. 14A shows the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) prior to incubation with the activated rotavirus solution and the ETEC-containing solution for 3 or 6 hours. It is a figure which shows the result of having analyzed the A20 expression level about the PIE cell line which was stimulated with (") or was not stimulated by these strains ("control "in the figure).
- FIG. 14A shows the # 35 strain (“# 35” in the figure) or the # 58 strain (“# 58” in the figure) prior to incubation with
- the antiviral agent of the present invention comprises a 16S rRNA gene having at least 90% identity with the lactobacillus strain (ie, the nucleotide sequence set forth in SEQ ID NO: 1) identified for use as "for antivirus".
- An agent containing one or more hereinafter, “anti-virus” It is sometimes called a “virus agent”).
- the 16S rRNA gene is usually contained in the genomic DNA of the Lactobacillus bacterial strain.
- the Lactobacillus bacterial strain which is an antiviral probiotic
- a feed composition for livestock, a food or drink composition or a pharmaceutical composition examples include health foods (functional foods, nutritional supplements, health supplements, nutritionally fortified foods, nutritionally adjusted foods, supplements, etc.), health functional foods (specified health foods, nutritional functional foods, functional foods). Labeled foods, etc.) can be mentioned.
- livestock feed and food and drink are preferable.
- anti-virus means prevention and / or improvement of viral infections by suppressing the growth of the virus, inactivating the virus, reducing the susceptibility to the virus, and the like. It means to (treat).
- the antiviral factor may be a factor that causes an antiviral effect in mammalian cells (eg, polypeptide, protein [specifically, cytokine, antibody], polynucleotide, saccharide, lipid), for example.
- the above-mentioned "downregulatory factor of antiviral factor” may be a factor that causes a decrease in the expression of antiviral factor (for example, polypeptide, protein, polynucleotide, saccharide, lipid), and for example, A20 (TNFAIP3 [ Tumor Necrosis Factor Alpha-Induced Protein 3], Tolllip (Toll-interacting protein), RNF125 (Ring Finger Protein 125), DUBA (Deubiquitinase A), CYLD (CYLD Lysine 63 Deubiquitinase), etc. Since the effect has been demonstrated in this example, a downregulator of one or two antiviral factors selected from A20 and Tollip can be preferably exemplified.
- the Lactobacillus bacterial strain has one or more receptors selected from TLR (Toll-like receptor) 2, TLR4, and NOD2 (Nucleotide binding oligomerization domain-like receptor 2) (hereinafter, "antiviral factor receptor”). It can be characterized by the expression-enhancing action of the body).
- TLR Toll-like receptor
- NOD2 Nucleotide binding oligomerization domain-like receptor 2
- antiviral factors down-regulatory factors of antiviral factors, and antiviral factor receptors (hereinafter, these may be collectively referred to as "antiviral factors, etc.")
- antiviral factors etc.
- the expression level of the antiviral factor and the like can be detected by using a method such as Western blotting method, indirect immunofluorescence method, flow cytometry, ELISA method, EIA method, RIA method and the like.
- transcripts of genes such as antiviral factors can directly detect mRNA of genes such as antiviral factors, and cDNA synthesized using mRNA of genes such as antiviral factors as a template. Can also be detected indirectly.
- the method for detecting mRNA of a gene such as an antiviral factor include (Reverse Transcription) -PCR method; Northern blotting method, microarray method, ISH method and the like.
- a method for detecting cDNA synthesized using mRNA of a gene such as an antiviral factor as a template for example, LAMP method, PCR method (for example, real-time PCR method [intercalator method, 5'-nuclease method, cycling probe). Methods], ddPCR method); LCR method; sequencing method using next-generation sequencer; Southern hybridization method using the probe for cDNA detection, microarray method, ISH method; and the like.
- antiviral factors in the present specification, “enhancement of expression of antiviral factors”, “reduction of expression of downregulatory factors of antiviral factors”, and “enhancement of expression of antiviral factor receptors” are used to refer to mammalian cell lines.
- the expression level of the antiviral factor was higher, respectively. It means “increased”, “decreased expression level of downregulatory factor of antiviral factor”, and “increased expression level of antiviral factor receptor”.
- the above-mentioned mammalian cell line is a target virus of the antiviral agent, a substance that induces virus pseudo-infection (for example, Polyinosinic-polycytidylic acid [PolyI]. : C]) and / or may be stimulated with pathogenic bacteria.
- An arbitrary threshold (cutoff value) can be set as to whether the expression level of the antiviral factor or the like has increased or decreased, and the threshold can be set as, for example, the antivirus in the comparative control.
- Mean value of expression level of sex factors and the like mean value + standard deviation (SD); mean value + 2SD; mean value + 3SD; median; interquartile range and the like can be mentioned.
- the thresholds are sensitivity (ratio at which a mammalian cell line cultured in the presence of the Lactobacillus bacterium strain can be correctly determined to be positive) and specificity (mammalian cells cultured in the absence of the Lactobacillus genus bacterial strain).
- mammalian cell line examples include a human intestinal epithelial cell line (Cac Cincinnati-2), a porcine intestinal epithelial cell line (PIE [Porcine Intestinal Epitheliocyte] cell line), and a bovine intestinal epithelial cell line (BIE [Bovine Intestinal Epitheliocyte] cell line). ) Stocks can be mentioned.
- the target virus of the antiviral agent is not particularly limited, and is, for example, a DNA virus (double-stranded DNA virus, single-stranded DNA virus, etc.), RNA virus (double-stranded RNA virus, single-stranded RNA (+)).
- a double-stranded RNA virus is preferable because the effect on the double-stranded RNA model virus has been demonstrated in this example described later.
- the target virus includes, for example, an influenza virus (single-strand RNA (-)-strand virus), a norovirus (single-strand RNA (+)-strand virus), a rotavirus (double-strand RNA virus), and the like.
- Rave virus (single-strand RNA (+)-strand virus), measles virus (positive-strand RNA (+)-strand virus), RS virus (positive-strand RNA (-)-strand virus), herpes virus (double-strand DNA virus) ), Hepatitis A virus (positive-strand RNA (+)-strand virus), hepatitis B virus (double-strand DNA virus), hepatitis C virus (positive-strand RNA (+)-strand virus), hepatitis E virus (Single-strand RNA (+)-strand virus), adenovirus (double-strand DNA virus), hoof virus (positive-strand RNA (+)-strand virus), mad dog disease virus (positive-strand RNA (-)-strand virus), Human immunodeficiency virus (single-strand RNA (+)-strand virus), corona virus (single-strand RNA (+)-strand virus), etc.
- rotavirus can be preferably exemplified.
- Influenza virus includes type A, type B, type C, triinfluenza virus and various subtypes thereof, and coronavirus includes a general coronavirus that causes colds and a new type of corona.
- Viruses eg, Severe Acute Respiratory Syndrome Coronavirus [SARS and SARS-CoV-2], Middle East Respiratory Syndrome Coronavirus [MERS], COVID-19
- SARS and SARS-CoV-2 Severe Acute Respiratory Syndrome Coronavirus [SARS and SARS-CoV-2], Middle East Respiratory Syndrome Coronavirus [MERS], COVID-19
- the target of application of the antiviral agent may be a mammal that requires prevention and / or improvement (treatment) of a viral infection, and in this example described later, it is also effective in a complex infection of a virus and a pathogenic bacterium.
- Preferably exemplify mammals in need to prevent and / or ameliorate (treat) viral infections in a combined viral and pathogenic bacterial infection, as they have been demonstrated to exert antiviral effects. be able to.
- pathogenic bacteria include, for example, mycoplasma, enterohemorrhagic Escherichia coli (eg, enterohemorrhagic Escherichia coli [EPEC], enterohemorrhagic tissue invading Escherichia coli [EIEC], toxinogenic Escherichia coli [ETEC], intestinal aggregate adhesion.
- enterohemorrhagic Escherichia coli eg, enterohemorrhagic Escherichia coli [EPEC], enterohemorrhagic tissue invading Escherichia coli [EIEC], toxinogenic Escherichia coli [ETEC], intestinal aggregate adhesion.
- EAggEC enterohemorrhagic Escherichia coli
- EHEC enterohemorrhagic Escherichia coli
- lytic bacteria and the like can be mentioned.
- the mammals include humans and non-human mammals (eg, monkeys, mice, rats, dogs, cats, livestock [eg, rabbits, pigs, horses, cows, sheep, goats, deer]) and the like. It can be mentioned, and humans and livestock can be preferably exemplified.
- non-human mammals eg, monkeys, mice, rats, dogs, cats, livestock [eg, rabbits, pigs, horses, cows, sheep, goats, deer]
- livestock eg, rabbits, pigs, horses, cows, sheep, goats, deer
- the lactobacillus bacterium strain has a 16S rRNA gene having at least 90% identity with the nucleotide sequence of SEQ ID NO: 1, and has an antiviral factor expression enhancing action and / or an antiviral factor downward control.
- it is a bacterial strain of the genus Lactobacillus having an action of reducing the expression of the factor, it may be a living bacterial strain or a dead bacterial strain.
- the Lactobacillus genus bacterial strain is different from the Lactobacillus salivarius strain or the Lactobacillus plantaram strain, or the Lactobacillus salivalius strain or the Lactobacillus plantalum strain whose effect has been demonstrated in the present Examples described later (for example).
- Lactobacillus hayakitensis [Lactobacillus hayakitensis], Lactobacillus agilis, Lactobacillus aviarius subsp. , Has a 16S rRNA gene having at least 90% identity with the nucleotide sequence of SEQ ID NO: 1, and has an effect of enhancing the expression of antiviral factors and / or an effect of reducing the expression of down-regulatory factors of antiviral factors. It also includes Lactobacillus strains.
- the Lactobacillus genus bacterial strain is specifically deposited as the Lactobacillus salivarius strain (# 35 strain) deposited under the international deposit receipt number NITE ABP-03218; the international deposit receipt number NITE ABP-03219.
- Lactobacillus salivalius strain (# 58 shares); deposited as international deposit receipt number NITE ABP-03221; and Lactobacillus salivalius strain (# 131 shares); and deposited as international deposit receipt number NITE ABP-03220. Lactobacillus salivalius shares (# 71 shares); and Lactobacillus plantaram shares (# 16 shares) deposited under the international deposit receipt number NITE ABP-03474; deposited under the international deposit receipt number NITE ABP-03467.
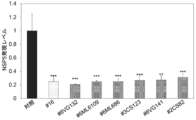
- Lactobacillus plantaram stock (# 6VG132 stock); Lactobacillus plantaram stock (# 6ML6109 stock) deposited as international deposit receipt number NITE ABP-03468; deposited as international deposit receipt number NITE ABP-03466 Lactobacillus plantaram stock (# 6ML686 shares); Lactobacillus plantaram stock (# 3CS123 shares) deposited as international deposit receipt number NITE ABP-03471; deposited as international deposit receipt number NITE ABP-03469 Lactobacillus plantaram stock (# 6VG141 stock); Lactobacillus plantaram stock (# 2CS82 stock) deposited as international deposit receipt number NITE ABP-03472; deposited as international deposit receipt number NITE ABP-03472 Lactobacillus plantalum strain (# 1FeB18 strain); and Lactobacillus plantalum strain (# 4FeB195 strain) deposited under the international deposit receipt number NITE ABP-03473; Lactobacillus salivarius strain (
- Lactobacillus salivalius stock (# 1) deposited under the international deposit receipt number NITE ABP-03221 31 shares); Lactobacillus plantarum shares deposited under the international deposit receipt number NITE ABP-03474 (# 16 shares); Lactobacillus plantaram shares deposited under the international deposit receipt number NITE ABP-03467 (the # 16 shares) 6VG132 shares); Lactobacillus plantarum shares deposited under the international deposit receipt number NITE ABP-03468 (# 6ML6109 shares); Lactobacillus plantaram shares deposited under the international deposit receipt number NITE ABP-03466 (the # #).
- 6ML686 shares Lactobacillus plantarum shares deposited under the international deposit receipt number NITE ABP-03471 (# 3CS123 shares); Lactobacillus plantaram shares deposited under the international deposit receipt number NITE ABP-03469 (the # 3). 6VG141 shares); Lactobacillus plantarum shares deposited under the international deposit receipt number NITE ABP-03470 (# 2CS82 shares); Lactobacillus plantaram shares deposited under the international deposit receipt number NITE ABP-03472 (the # 2). 1FeB18 strain); and one or more selected from the Lactobacillus plantarum strain (the # 4FeB195 strain) deposited under the international deposit receipt number NITE ABP-03473; can be preferably exemplified.
- nucleotide sequence shown in SEQ ID NO: 1 means that one or several nucleotides in the nucleotide sequence of SEQ ID NO: 1 are substituted, deleted, inserted, added or inverted. , 90% or more of the entire nucleotide sequence of SEQ ID NO: 1 is identical.
- a nucleotide sequence in which one or several nucleotides are substituted, deleted, inserted, added or inverted is defined as, for example, in the range of 1 to 149, preferably in the range of 1 to 100.
- the "at least 90% identity” is preferably 91% or more, more preferably 92% or more, still more preferably 93% or more, still more preferably 94% or more, and particularly preferably 95% or more.
- the identity is particularly more preferably 96% or more, particularly still more preferably 97% or more, particularly even more preferably 98% or more, and most preferably 99% or more (about 100%).
- Nucleotide sequence identity is called BLASTX or BLASTP based on the algorithm BLAST (Proc. Natl. Acad. Sci. USA 87: 2264-2268, 1990, Proc Natl Acad Sci USA 90: 5873, 1993) by Carlin and Altschul.
- BLASTN Altschul SF, et al: J Mol Biol 215: 403, 1990
- the Lactobacillus bacterial strain further includes those having wakame assimilation.
- “wakame assimilation” means the ability to synthesize substances necessary for Lactobacillus bacteria such as proteins, nucleic acids, sugars, and lipids using wakame as a carbon source or nitrogen source.
- the antiviral agent is roughly classified into a liquid type and a non-liquid type.
- the liquid type antiviral agent the culture solution of the Lactobacillus genus bacterial strain is purified, and if necessary, appropriate physiological saline or fluid replacement or pharmaceutical additives are added and filled in ampoules or vials.
- the non-liquid type antiviral agent is an ampoule, vial, etc. by adding an appropriate freeze-protecting agent (for example, glycerol, dimethylsulfoxide [DMSO], trehalose, dextran) to the liquid type antiviral agent. It can be produced by freezing or freeze-drying after filling in.
- DMSO dimethylsulfoxide
- the method of applying (administering) the antiviral agent can be either an oral application method (oral administration) or a parenteral application method (parenteral administration), and a parenteral application method (non-oral administration).
- oral administration include intravenous administration and local administration.
- the additives include conventional pharmaceutically acceptable carriers, binders, stabilizers, excipients, diluents, pH buffers, disintegrants, isotonic agents, additives, and coating agents.
- Solubilizers, lubricants, gliding agents, solubilizing agents, lubricants, flavoring agents, sweeteners, solvents, gelling agents, nutritional agents and the like can be exemplified.
- Specific examples of such compounding components include water, wakame component, physiological saline, animal fat and oil, vegetable oil, lactose, starch, gelatin, crystalline cellulose, gum, talc, magnesium stearate, hydroxypropyl cellulose, and poly.
- the antiviral agent contains a wakame component (prebiotic) which is a nutrient source of the Lactobacillus bacterial strain as an additive, and the Lactobacillus is contained.
- Bacterial strains of the genus preferably antiviral immunobiotics
- those comprising immunosin biotics by prebiotics can be preferably exemplified.
- the seaweed component may be a seaweed powder obtained by crushing a dried seaweed product, or a seaweed component extract obtained by further extracting the seaweed powder with water or the like. ..
- the applicable (administration) amount of the Lactobacillus bacterial strain contained in the antiviral agent varies depending on the sex, age, body weight, physical condition, etc. of the subject (mammal) to be ingested, and therefore cannot be unconditionally specified. It is 10 4 to 10 12 cfu (Colony Forming Unit) per 1 kg of body weight per day, preferably 10 6 to 10 10 cfu. It should be noted that such an amount may be ingested once or may be ingested in several divided doses.
- the amount of the Lactobacillus genus bacterial strain contained in the livestock feed composition is, for example, 10 4 to 10 12 cfu / g per 1 g of the livestock feed composition. It is preferably 10 6 to 10 10 cfu.
- Example 1 Search for antiviral factors We searched for factors that can be used as indicators for evaluating the antiviral properties of probiotics.
- Test cells For the PIE cell line, cells cloned from the small intestine of a primary piglet of a ternary hybrid (LWD; landrace ⁇ large white ⁇ durock) were used.
- the PIE cell line is a DMEM liquid medium (high glucose, L-glutamine) containing 10% FCS (fetal bovine serum) and 1% streptomycin / penicillin using a 250 mL flask (manufactured by Sumitomo Bakelite) coated with type I collagen. , Containing sodium pyruvate; manufactured by GIBCO) (hereinafter, simply referred to as "DMEM liquid medium").
- FCS fetal bovine serum
- the epithelial buffer was removed and incubated in PBS containing 0.25% trypsin and 0.02% EDTA for 5 minutes at 37 ° C.
- DMEM liquid medium was added, and the detached cells were collected and then centrifuged (12000 rpm ⁇ 5 minutes) to remove the culture supernatant.
- DMEM liquid medium was newly added, and the cells were collected. After counting the number of cells, cells were seeded such that 1 ⁇ 10 6 cells per flask. After culturing for 24 hours, the culture supernatant was removed by suction with an aspirator, and DMEM liquid medium was newly added for culturing.
- the cells were stored at ⁇ 80 ° C. for each generation using a cell bunker (registered trademark) (manufactured by Nippon Zenyaku Kogyo Co., Ltd.). In addition, cell culture was performed under the conditions of 5% CO 2 /20% O 2 and 37 ° C.
- the PIE cell line was stimulated with Poly I: C, which is a double-stranded RNA virus model that induces virus pseudo-infection, and the expression of cytokine-related factors was analyzed. Specifically, the analysis was performed according to the following procedures [1] to [5]. [1] The PIE cell line was inoculated on a 12-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / well, and cultured in DMEM liquid medium for 5 days.
- RNA concentration and purity were measured with a NanoDrop ND-1000 spectrophotometer (manufactured by Thermo Fisher Scientific).
- cDNA was synthesized from the obtained total RNA using the Prime Script RT reagent Kit with gDNA Eraser (Perfect Real Time) (manufactured by Takara) according to the protocol attached to the product.
- cytokine-related factors IFN- ⁇ , IFN- ⁇ , Mx1, OAS1, RNaseL, PKR, RIG-I, TLR2, TLR3, TLR4, NOD1, NOD2, MCP-1 [Also called CCL2], IL-6, IL-8 [also called CXCL8], IL-12, IL-18, TNF ⁇ , A20, BCL-3, Tollip, IRAK-M, MKP-1, and SIGIRR) and ⁇ .
- the primer set (sense primer and antisense primer) shown in Table 1 below, Platinum SYBR Green qPCR Super Mix-UDG with ROX (manufactured by Invitrogen), and ABI PRISM Quantitative PCR analysis using a 7300 real-time PCR system (manufactured by Applied Biosystem) was performed according to the protocol attached to the product.
- the expression level of various cytokine-related factors by Poly I: C stimulation is the formula (Poly I: C stimulation [expression level of mRNA of various cytokine-related factor genes / expression level of mRNA of ⁇ -actin gene] / Poly I: C unstimulated. [Expression level of mRNA of various cytokine-related factor genes / Expression level of mRNA of ⁇ -actin gene]).
- the Lactobacillus bacterial strain was selected using the IFN- ⁇ expression level 3 hours after PolyI: C stimulation and the Mx1 expression level 12 hours after PolyI: C stimulation as indicators. I decided to use it.
- Example 2 Selection of the Lactobacillus bacterial strain using antiviral property as an index
- the Lactobacillus bacterial strain was selected using the expression levels of two antiviral factors (IFN- ⁇ and Mx1) as indicators.
- the PIE cell line was stimulated with the prepared 116 kinds of Lactobacillus salivalius strain-containing solution, and then stimulated with Poly I: C to express two kinds of antiviral factors (IFN- ⁇ and Mx1).
- the Lactobacillus bacterial strain was selected using the level as an index. Specifically, the analysis was performed according to the following procedures [1] to [4]. [1] The PIE cell line was inoculated on a 12-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / well, and cultured in DMEM liquid medium for 3 days.
- the PIE cell line was similarly cultured in DMEM liquid medium free of the Lactobacillus salivalius strain.
- the cells were replaced with DMEM liquid medium containing 50 ng / mL Poly I: C (catalog number P9582, manufactured by SIGMA), and cultured for 3 hours and 12 hours. By doing so, Poly I: C stimulation was performed.
- the IFN- ⁇ expression level and Mx1 expression level stimulated by various lactobacillus salivarius strains are the formulas (before stimulating with Poly I: C, when stimulated with various lactobacillus salivarius strains [IFN- ⁇ gene or Mx1 gene mRNA]. Expression level / ⁇ -actin gene mRNA expression level]) / Based on [IFN- ⁇ gene or Mx1 gene mRNA expression level / ⁇ -actin gene mRNA expression level]) when stimulated with Poly I: C (See the vertical axis in FIG. 2).
- the # 35 strain is a Lactobacillus salivarius strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 1 and has the following characteristics.
- the # 35 strain is the NITE Patent Microorganisms Depositary (NPMD) (Address: Kazusa, Kisarazu City, Chiba Prefecture, 292-0818). Kazusakamatari Room 2-5-8 122 [# 122, 2-5-8 Kazusakamatari, Kisarazu-shi, Chiba 292-0818, Japan]) as international deposit receipt number NITE ABP-03218 on May 19, 2020. It has been deposited internationally.
- NPMD NITE Patent Microorganisms Depositary
- A Cell morphology: bacillus, spore formation: (-), motility: (-)
- B Colony morphology (Observation of colony morphology smeared on MRS agar medium and aerobically cultured at 37 ° C.
- the # 58 strain is a Lactobacillus salivarius strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 1 and has the following characteristics.
- # 58 strain was transferred to the National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: Room 2-5-8 122, Kazusakamatari, Kisarazu City, Chiba Prefecture, 292-0818). It has been internationally deposited under the international deposit receipt number NITE ABP-03219 on May 19, 2020.
- NITE National Institute of Technology and Evaluation
- Cell morphology bacillus, spore formation: (-), motility: (-)
- B Colony morphology (Observation of colony morphology smeared on MRS agar medium and aerobically cultured at 37 ° C.
- Example 3 Evaluation of Immunomodulatory Ability of Lactobacillus Bacterial Strain
- the PIE cell line was stimulated with only the selected # 35 strain or the # 58 strain, and the immune response of the cells was analyzed.
- the analysis of the immune response of the cells was performed according to the following procedures [1] to [3].
- the PIE cell line was inoculated on a 12-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / well, and cultured in DMEM liquid medium for 3 days.
- the expression levels of various receptors (see FIG. 3) and the expression levels of various antiviral factors (see FIGS. 4 and 5) stimulated by the # 35 strain or the # 58 strain are based on the formula (stimulation of the # 35 strain or the # 58 strain). [Expression level of mRNA of various receptor genes or various antiviral factor genes / Expression level of mRNA of ⁇ -actin gene] / [Various receptor genes or various antiviral factor genes] by unstimulated strain (0 hours) Expression level of mRNA / expression level of ⁇ -actin gene mRNA]).
- Example 4.2 Evaluation of immunomodulatory ability of the Lactobacillus bacterial strain against a double-stranded RNA model virus 1 For the # 35 and # 58 strains, the # 35 strain or the # 58 strain before stimulating the PIE cell line with Poly I: C to evaluate the immunomodulatory ability against the double-stranded RNA model virus. The expression of antiviral factors was analyzed.
- A20 / TNFAIP3 is known to be a negative regulator that plays a particularly important role during virus infection, and it has been reported that rotavirus infection is significantly suppressed in cells in which A20 expression is knocked down.
- Tolllip Toll-interacting protein
- Tolllip is a negative regulator that interacts with IRAK-1 to inhibit phosphorylation and negatively control the signaling of TLR2 and TLR4, and is responsible for the expression of IFN- ⁇ and IFN- ⁇ .
- the analysis of the immune response of the cells was performed according to the following procedures [1] to [4].
- the PIE cell line was inoculated on a 12-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / well, and cultured in DMEM liquid medium for 3 days.
- PIE cell lines were similarly cultured in DMEM liquid medium free of these strains.
- Example 4 shows that when the # 35 strain or the # 58 strain is administered to cells infected with double-stranded RNA virus, it is a downregulator of antiviral factors such as A20 and Tollip. It is shown that the expression level of (negative regulator) is reduced and the expression enhancing effect of antiviral factors is exerted.
- Example 6 Confirmation of the virus infection reducing effect of the Lactobacillus strains Since the # 35 strain and the # 58 strain were shown to enhance the antiviral immune response, these strains have the effect of reducing the virus infection. To confirm that, a rotavirus infection test was performed.
- Test virus strain The livestock infectious group A rotavirus OSU strain using pigs as a host was distributed by the National Institute of Animal Health, Animal Health Research Division.
- the rotavirus infection test and the subsequent indirect fluorescent antibody method using the anti-rotavirus antibody were carried out according to the following procedures [1] to [7].
- the PIE cell line was inoculated on a 96-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / mL, and cultured in DMEM liquid medium for 8 days.
- the # 35 strain and the # 58 strain were added to the medium so as to have a multiplicity of infection (MOI) per well, respectively, and cultured (stimulated) for 48 hours.
- MOI multiplicity of infection
- PIE cell lines were similarly cultured in DMEM liquid medium free of these strains.
- the cells were washed 3 times with FCS-free DMEM liquid medium, and 100 ⁇ L (1 MOI) of the activated rotavirus solution prepared according to the method described in the above item [Preparation of activated rotavirus solution] was added to each well. Equivalent) was added at a time and incubated for 16 hours under the conditions of 5% CO 2 /20% O 2 and 37 ° C.
- the activated rotavirus solution was removed, 100 ⁇ L of an 80% acetone solution at 4 ° C. was added to each well, and the cells were incubated at 4 ° C.
- rotavirus A antibody anti-human RVAWa strain guinea pig serum, distributed by the National Institute of Animal Health, Animal Health Research Division. The primary antibody reaction was carried out. Specifically, rotavirus A antibody was diluted 800-fold with PBS, 50 ⁇ L was dispensed into each well, and then incubated at 37 ° C. for 40 minutes.
- a secondary antibody reaction was carried out using Alexa488-labeled anti-guinea pig IgG antibody (Cat. Ab150185, manufactured by abcam).
- such antibody was diluted 400-fold with PBS at 1: 400, 50 ⁇ L was dispensed into each well, and then incubated at 37 ° C. for 40 minutes.
- 50 ⁇ L of PBS containing 30% glycerin was dispensed into each well, and then a fluorescence signal derived from Alexa488 was emitted using a fluorescence microscope (IX70-FL manufactured by Olympus Corporation). Detected (see FIG. 8A) and calculated the percentage of cells positive for such fluorescence signal, i.e., rotavirus-positive (infected) cells (see FIG. 8B).
- Example 7 Evaluation of immunomodulatory ability of the Lactobacillus bacterial strain at the time of virus infection 1
- the PIE cell line was stimulated with the # 35 strain or the # 58 strain and then infected with rotavirus to infect the antivirus. Expression analysis of sex factors was performed.
- the rotavirus infection test and the subsequent analysis of the expression of antiviral factors were performed according to the following procedures [1] to [4].
- the PIE cell line was inoculated on a 96-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 5 ⁇ 10 4 cells / well, and cultured in DMEM liquid medium for 10 days.
- the # 35 strain and the # 58 strain were added to the medium so as to have 100 MOI per well, and cultured (stimulated) for 48 hours. As a control for unstimulating these strains, PIE cell lines were similarly cultured in DMEM liquid medium free of these strains.
- Example 8 Evaluation of immunomodulatory ability of the Lactobacillus bacterial strain at the time of virus infection 2
- the PIE cell line was stimulated with the # 35 strain or the # 58 strain, and then infected with rotavirus to infect the antivirus.
- the rotavirus infection test and the subsequent analysis of A20 and Tollip expression were performed according to the following procedures [1] to [4].
- the PIE cell line was inoculated on a 96-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / mL, and cultured in DMEM liquid medium for 8 days.
- the # 35 strain and the # 58 strain were added to the medium so as to have 100 MOI per well, and cultured (stimulated) for 48 hours. As a control for unstimulating these strains, PIE cell lines were similarly cultured in DMEM liquid medium free of these strains.
- Example 9 Confirmation of the effect of reducing the combined infection of rotavirus and ETEC by the lactobacillus genus bacterial strain In actual life, not only the infection of the virus alone but also the combined infection with the pathogenic bacterium occurs. Therefore, a complex infection system of rotavirus and entero toxigenic E. coli (ETEC) was constructed. First, a rotavirus infection was established between the infection of rotavirus alone and the combined infection of rotavirus and ETEC. We examined whether there was a difference in efficiency, and then analyzed whether the # 35 strain and the # 58 strain were effective against the combined infection of rotavirus and ETEC.
- ETEC entero toxigenic E. coli
- ETEC-containing liquid The ETEC strain (distributed by the National Institute of Animal Health, National Institute of Animal Health) was drawn on an agar medium containing 5% sheep defibered blood and cultured at 37 ° C. for 20 hours. The formed colonies are cultivated, inoculated into 5 mL of tryptone soya broth liquid medium (manufactured by Nippon BD), and left to stand at 37 ° C for 5 to 8 days for ciliary restoration culture. went. Then, the fungus was caught from the mycelial membrane forming part, inoculated into 11 mL of TSB, and cultured with shaking at 37 ° C. for 20 hours.
- the cells were collected by centrifugation, washed 3 times with PBS, and heat-sterilized at 100 ° C. for 15 minutes. Then, the cells were washed with PBS and suspended in DMEM liquid medium so that the cell concentration was 1.5 ⁇ 10 10 cells / mL to prepare an ETEC-containing solution.
- the rotavirus / ETEC combined infection test and the subsequent indirect immunofluorescence method using an anti-rotavirus antibody were carried out according to the following procedures [1] to [4].
- the PIE cell line was inoculated on a 96-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / well, and cultured in DMEM liquid medium for 8 days.
- the # 35 strain and the # 58 strain were added to the medium so as to have 100 MOI per well, and cultured (stimulated) for 48 hours.
- PIE cell lines were similarly cultured in DMEM liquid medium free of these strains.
- the cells were washed 3 times with FCS-free DMEM liquid medium, and the activated rotavirus solution prepared according to the method described in the item [Preparation of activated rotavirus solution] of Example 6 above was added to each well.
- Example 10 Evaluation of immunomodulatory ability of the Lactobacillus bacterial strain at the time of virus and bacterial infection 1
- the PIE cell line was stimulated with the # 35 strain or the # 58 strain and then converted into rotavirus. After infection, the expression of antiviral factors and down-regulatory factors of antiviral factors was analyzed.
- the virus / pathogenic bacterial complex infection test and the subsequent expression analysis of antiviral factors and downregulatory factors of antiviral factors were carried out according to the following procedures [1] to [4].
- the PIE cell line was inoculated on a 96-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 5 ⁇ 10 4 cells / well, and cultured in DMEM liquid medium for 10 days.
- the # 35 strain and the # 58 strain were added to the medium so as to have 100 MOI per well, and cultured (stimulated) for 48 hours. As a control for unstimulating these strains, PIE cell lines were similarly cultured in DMEM liquid medium free of these strains.
- Example 11 Selection of the Lactobacillus bacterial strain using wakame assimilation as an index Regarding the bacterial metabolites that assimilate seaweed, organic acids produced from algae hydrous carbon activate intestinal microorganisms in humans and marine invertebrates. (See “Front Immunol. 2014 Jan 14; 4: 512.” And “New Phytol. 2010 Oct; 188 (1): 82-97.”). Furthermore, as a prebiotic-related study, there is a report that metabolites obtained by microbial fermentation of red algae exert antioxidative, anticoagulant, and immunomodulatory effects (Reference "Phytomedicine. 2012 Jun 15; 19”. (8-9): 797-803. ").
- Lactobacillus salivarius strains that grow in a liquid medium containing wakame components
- Lactobacillus salivalius strain # 131 referred to as “the # 131 strain” in the present specification
- Lactobacillus salivalius strain # 71 referred to as “the # 71 strain”
- the # 131 strain is a Lactobacillus salivarius strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 1, similar to the # 35 strain and the # 58 strain, and is the # 35 strain and the # 58 strain. Has the same characteristics as. That is, the # 131 strain has the following characteristics.
- the # 131 strain will be placed in 2020 at the National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: 2-5-8 Kazusakamatari, Kisarazu City, Chiba Prefecture, 292-0818). It has been deposited internationally as the international deposit receipt number NITE ABP-03221 on May 19.
- A Cell morphology: bacillus, spore formation: (-), motility: (-)
- B Colony morphology (Observation of colony morphology smeared on MRS agar medium and aerobically cultured at 37 ° C.
- the # 71 strain is a Lactobacillus salivalius strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 1, similar to the # 35 strain and the # 58 strain, and is the # 35 strain and the # 58 strain. Has the same characteristics as. That is, the # 71 strain has the following characteristics.
- the # 71 strain will be placed in 2020 at the National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: 2-5-8 Kazusakamatari, Kisarazu City, Chiba Prefecture, 292-0818). It was deposited internationally as the international deposit receipt number NITE ABP-03220 on May 19.
- NITE National Institute of Technology and Evaluation
- Cell morphology bacillus, spore formation: (-), motility: (-)
- B Colony morphology (Observation of colony morphology smeared on MRS agar medium and aerobically cultured at 37 ° C.
- Example 12 Evaluation of the immunomodulatory ability of the # 131 strain against the double-stranded RNA model virus
- the # 131 strain has an antiviral immune response enhancing effect, similar to the # 35 strain and the # 58 strain.
- the PIE cell line was stimulated with the # 131 strain, and the expression of antiviral factors was analyzed.
- the analysis of the immune response of the cells was performed according to the following procedures [1] to [4].
- the PIE cell line was inoculated on a 12-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / well, and cultured in DMEM liquid medium for 3 days.
- the [2] present # 131 strain, in addition to the medium so as to be 5.0 ⁇ 10 7 cells / mL per well, were cultured for 48 hours (stimulation).
- the PIE cell line was similarly cultured in DMEM liquid medium free of the # 131 strain.
- the IFN- ⁇ expression level stimulated by the # 131 strain is the formula (before stimulating with Poly I: C, the expression level of the mRNA of the IFN- ⁇ gene when stimulated with the # 131 strain. / ⁇ -actin gene mRNA expression level] / Calculated based on [IFN- ⁇ gene mRNA expression level / ⁇ -actin gene mRNA expression level] when stimulated with Poly I: C.
- the expression levels of various antiviral factors when both the # 131 strain and Poly I: C were unstimulated were also calculated (see FIG. 16).
- the # 131 strain had the same effect of enhancing the expression of antiviral factors as the # 35 strain and the # 58 strain.
- the # 35 strain and the # 58 strain, and the # 131 strain and the # 71 strain are both Ligilactobacillus salivalius strains having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 1. It was suggested that the Ligilactobacillus salivarius strain having the 16S rRNA gene consisting of one nucleotide sequence has both antiviral and wakame assimilation properties.
- Example 13 Confirmation of virus infection reduction effect of this Lactobacillus bacterial strain (2) Even a strain of Lactobacillus spp. That is different from Lactobacillus salivarius has high sequence identity with the nucleotide sequence of the 16S rRNA gene of Lactobacillus salivalius that exerted an antiviral effect (that is, the nucleotide sequence of SEQ ID NO: 1). It was analyzed whether or not what had had an antiviral effect. Specifically, nine types of Lactobacillus plantarum strains included in the Lactobacillus spp.
- the # 16 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 52, and is the National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: ⁇ 292). -0818 It was deposited internationally as the international deposit receipt number NITE ABP-03474 on April 23, 2021 at Room 2-5-8 Kazusakamatari, Kisarazu City, Chiba Prefecture.
- the # 16 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- the # 6ML686 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 53, and is a National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: It has been internationally deposited as the international deposit receipt number NITE ABP-03466 on April 23, 2021 at Room 2-5-8 122, Kazusakamatari, Kisarazu City, Chiba Prefecture, 292-0818.
- the # 6ML686 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- A Cell morphology: bacillus, spore formation: (-), motility: (-)
- B Colony morphology (Observation of colony morphology smeared on MRS agar medium and aerobically cultured at 37 ° C.
- the # 6ML6109 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 53, and is a National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: It has been internationally deposited as the international deposit receipt number NITE ABP-03468 on April 23, 2021 at Room 2-5-8 122, Kazusakamatari, Kisarazu City, Chiba Prefecture, 292-0818.
- NITE National Institute of Technology and Evaluation
- the # 6ML6109 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- the # 6VG132 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 54, and is a National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: ⁇ 292). -0818 It has been internationally deposited as the international deposit receipt number NITE ABP-03467 on April 23, 2021 at Room 2-5-8 Kazusakamatari, Kisarazu City, Chiba Prefecture.
- the # 6VG132 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- the # 6VG141 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 55, and is a National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: ⁇ 292). -0818 It has been internationally deposited as the international deposit receipt number NITE ABP-03469 on April 23, 2021 at Room 2-5-8 Kazusakamatari, Kisarazu City, Chiba Prefecture.
- the # 6VG141 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- the # 2 CS82 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 56, and is a National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: It has been internationally deposited as the international deposit receipt number NITE ABP-03470 on April 23, 2021 at Room 2-5-8 122, Kazusakamatari, Kisarazu City, Chiba Prefecture, 292-0818.
- the # 2 CS82 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- A Cell morphology: bacillus, spore formation: (-), motility: (-)
- B Colony morphology (Observation of colony morphology smeared on MRS agar medium and aerobically cultured at 37 ° C.
- the # 1FeB18 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 56, and is a National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: It has been internationally deposited as the international deposit receipt number NITE ABP-03472 on April 23, 2021 at Room 2-5-8 122, Kazusakamatari, Kisarazu City, Chiba Prefecture, 292-0818.
- NITE National Institute of Technology and Evaluation
- the # 1 FeB18 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- the # 3 CS123 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 57, and is a National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: ⁇ 292). -0818 It was deposited internationally as the international deposit receipt number NITE ABP-03471 on April 23, 2021 at Room 2-5-8 Kazusakamatari, Kisarazu City, Chiba Prefecture.
- the # 3 CS123 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- the # 4FeB195 strain is a lactobacillus plantarum strain having a 16S rRNA gene consisting of the nucleotide sequence of SEQ ID NO: 58, and is a National Institute of Technology and Evaluation (NITE) Patent Microorganisms Depositary Center (NPMD) (Address: It has been internationally deposited as the international deposit receipt number NITE ABP-03473 on April 23, 2021 at Room 2-5-8 122, Kazusakamatari, Kisarazu City, Chiba Prefecture, 292-0818.
- NITE National Institute of Technology and Evaluation
- the # 4FeB195 strain has the following characteristics, similar to the above-mentioned four types of Lactobacillus salivalius strains (the # 35 strain, the # 58 strain, the # 131 strain, and the # 71 strain).
- Lactobacillus plantarum strains are all Lactobacillus genus bacterial strains having a 16S rRNA gene having at least 90% identity with the nucleotide sequence of SEQ ID NO: 1.
- the rotavirus infection test and subsequent analysis of rotavirus infection level and rotavirus-infected cells were performed according to the following procedures [1] to [7].
- the PIE1-3 cell line was inoculated on a 96-well plate (manufactured by Sumitomo Bakelite) coated with type I collagen so as to have 3 ⁇ 10 4 cells / mL, and cultured in DMEM liquid medium for 8 days. bottom.
- the above nine types of Lactobacillus plantarum strains were added to the medium so as to have 1 MOI per well, and cultured (stimulated) for 48 hours.
- PIE1-3 cell lines were similarly cultured in DMEM liquid medium free of these strains.
- the cells were washed 3 times with FCS-free DMEM liquid medium, and the activated rotavirus solution prepared according to the method described in the item [Preparation of activated rotavirus solution] of Example 6 above was added to each well. 100 ⁇ L per unit (equivalent to 1 MOI) was added and incubated for 12 hours under 5% CO 2 /20% O 2 and 37 ° C. conditions.
- RNA concentration and purity were measured with a NanoDrop ND-1000 spectrophotometer (manufactured by Thermo Fisher Scientific).
- cDNA was synthesized from the obtained total RNA using the Prime Script RT reagent Kit with gDNA Eraser (Perfect Real Time) (manufactured by Takara) according to the protocol attached to the product.
- the primer set sense primer and antisense primer shown in Table 2 below was used.
- the expression level of NSP5 by stimulation with the above 9 types of lactobacillus plantarum strains is the formula ([expression level of NSP5 gene mRNA / expression level of ⁇ -actin gene mRNA] by stimulation with the above 9 types of lactobacillus plantarum strains] / above. It was calculated based on [expression level of NSP5 gene mRNA / expression level of ⁇ -actin gene mRNA] by 9 kinds of lactobacillus plantarum strains unstimulated (see FIG. 17).
- the present invention contributes to the prevention or treatment of viral infections in the livestock industry and human medicine.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Microbiology (AREA)
- Genetics & Genomics (AREA)
- General Health & Medical Sciences (AREA)
- Zoology (AREA)
- Medicinal Chemistry (AREA)
- Biotechnology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Wood Science & Technology (AREA)
- Virology (AREA)
- Biomedical Technology (AREA)
- Veterinary Medicine (AREA)
- Molecular Biology (AREA)
- Public Health (AREA)
- Animal Behavior & Ethology (AREA)
- Pharmacology & Pharmacy (AREA)
- General Engineering & Computer Science (AREA)
- Mycology (AREA)
- Biochemistry (AREA)
- Epidemiology (AREA)
- Oncology (AREA)
- Communicable Diseases (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Tropical Medicine & Parasitology (AREA)
- Polymers & Plastics (AREA)
- Nutrition Science (AREA)
- Physiology (AREA)
- Food Science & Technology (AREA)
- Physics & Mathematics (AREA)
- Biophysics (AREA)
- Plant Pathology (AREA)
- Animal Husbandry (AREA)
- Medicines Containing Material From Animals Or Micro-Organisms (AREA)
Priority Applications (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2022526664A JPWO2021241728A1 (pl) | 2020-05-29 | 2021-05-28 | |
| EP21813425.2A EP4159839A1 (en) | 2020-05-29 | 2021-05-28 | Antiviral agent |
| US17/925,333 US20230263845A1 (en) | 2020-05-29 | 2021-05-28 | Antiviral Agent |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2020-093820 | 2020-05-29 | ||
| JP2020093820 | 2020-05-29 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2021241728A1 true WO2021241728A1 (ja) | 2021-12-02 |
Family
ID=78744833
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2021/020382 WO2021241728A1 (ja) | 2020-05-29 | 2021-05-28 | 抗ウイルス剤 |
Country Status (4)
| Country | Link |
|---|---|
| US (1) | US20230263845A1 (pl) |
| EP (1) | EP4159839A1 (pl) |
| JP (1) | JPWO2021241728A1 (pl) |
| WO (1) | WO2021241728A1 (pl) |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2023048197A1 (ja) * | 2021-09-22 | 2023-03-30 | アサヒバイオサイクル株式会社 | Lactobacillus plantarum株を含むトリの腸炎予防用組成物 |
Families Citing this family (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN117343880A (zh) * | 2023-12-05 | 2024-01-05 | 四川厌氧生物科技有限责任公司 | 一种唾液宿主关联乳杆菌及其应用 |
Citations (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2009511470A (ja) | 2005-10-06 | 2009-03-19 | プロビ エービー | ウイルス感染治療のための乳酸菌の使用 |
-
2021
- 2021-05-28 US US17/925,333 patent/US20230263845A1/en active Pending
- 2021-05-28 EP EP21813425.2A patent/EP4159839A1/en active Pending
- 2021-05-28 WO PCT/JP2021/020382 patent/WO2021241728A1/ja unknown
- 2021-05-28 JP JP2022526664A patent/JPWO2021241728A1/ja active Pending
Patent Citations (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2009511470A (ja) | 2005-10-06 | 2009-03-19 | プロビ エービー | ウイルス感染治療のための乳酸菌の使用 |
Non-Patent Citations (14)
| Title |
|---|
| ALTSCHUL SF ET AL., J MOL BIOL, vol. 215, 1990, pages 403 |
| ANIM SCI J, vol. 82, no. 2, pages 274 - 81 |
| D . Y. ZHANG , H. F. JI , S. X. WANG H. LIU , J. WANG , Y. M. WANG: "In vitro characterisation of two Lactobacillus strains and evaluation of their suitability as probiotics for growing-finishing pigs", ANIMAL PRODUCTION SCIENCE, vol. 59, no. 8, 19 September 2018 (2018-09-19), pages 1537 - 1545, XP009541688, ISSN: 1836-0939, DOI: 10.1071/AN17039 * |
| FISH SHELLFISH IMMUNOL, vol. 47, no. 2, pages 807 - 16 |
| FRONT IMMUNOL, vol. 4, 14 January 2014 (2014-01-14), pages 512 |
| KARLINALTSCHUL, PROC. NATL. ACAD. SCI. USA, vol. 87, 1990, pages 2264 - 2268 |
| MASUMIZU YUKI, ZHOU BINGHUI, KOBER A.K.M. HUMAYUN, ISLAM MD. AMINUL, IIDA HIKARU, IKEDA-OHTSUBO WAKAKO, SUDA YOSHIHITO, ALBARRACIN: "Isolation and Immunocharacterization of Lactobacillus salivarius from the Intestine of Wakame-Fed Pigs to Develop Novel "Immunosynbiotics"", MICROORGANISMS, MICROORGANISMS, vol. 7, no. 6, 6 June 2019 (2019-06-06), pages 167, XP055881076, ISSN: 2076-2607, DOI: 10.3390/microorganisms7060167 * |
| NEWPHYTOL, vol. 188, no. 1, October 2010 (2010-10-01), pages 82 - 97 |
| PHYTOMEDICINE, vol. 19, 15 June 2012 (2012-06-15), pages 797 - 803 |
| PROC NATL ACAD SCI USA, vol. 90, 1993, pages 5873 |
| RAFTIS E.J. ET AL.: "Unusual genome complexity in Lactobacillus salivarius JCM1046", BMC GENOMICS, vol. 15, 2014, pages 771, XP021195568, DOI: 10.1186/1471-2164-15-771 * |
| REJISH KUMAR V. J; BYEONG JOO SEO; MI RAN MUN; CHUL-JOONG KIM; INSUN LEE; HONGIK KIM; YONG-HA PARK: "Putative probioticspp. from porcine gastrointestinal tract inhibit transmissible gastroenteritis coronavirus and enteric bacterial pathogens", TROPICAL ANIMAL HEALTH AND PRODUCTION, KLUWER ACADEMIC PUBLISHERS, DO, vol. 42, no. 8, 10 July 2010 (2010-07-10), Do , pages 1855 - 1860, XP019857323, ISSN: 1573-7438, DOI: 10.1007/s11250-010-9648-5 * |
| SHIMAZU TOMOYUKI, BORJIGIN LIUSHIQI, KATOH KAZUO, ROH SANG‐GUN, KITAZAWA HARUKI, ABE KEIETSU, SUDA YOSHIHITO, SAITO HAYATO, KUNII : "Addition of Wakame seaweed ( Undaria pinnatifida ) stalk to animal feed enhances immune response and improves intestinal microflora in pigs", ANIMAL SCIENCE JOURNAL - NIHON CHIKUSAN GAKKAIHO, JAPANESE SOCIETY OF ZOOTECHNICAL SCIENCE., TOKYO, JP, vol. 90, no. 9, 1 September 2019 (2019-09-01), JP , pages 1248 - 1260, XP055881078, ISSN: 1344-3941, DOI: 10.1111/asj.13274 * |
| VET RES, vol. 42, 3 November 2011 (2011-11-03), pages 111 |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2023048197A1 (ja) * | 2021-09-22 | 2023-03-30 | アサヒバイオサイクル株式会社 | Lactobacillus plantarum株を含むトリの腸炎予防用組成物 |
Also Published As
| Publication number | Publication date |
|---|---|
| US20230263845A1 (en) | 2023-08-24 |
| JPWO2021241728A1 (pl) | 2021-12-02 |
| EP4159839A1 (en) | 2023-04-05 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US20220080003A1 (en) | Lactic acid bacterial strains | |
| Pender et al. | In ovo supplementation of probiotics and its effects on performance and immune-related gene expression in broiler chicks | |
| Guo et al. | Bacillus subtilis improves immunity and disease resistance in rabbits | |
| Sha et al. | Effects of lactic acid bacteria and the corresponding supernatant on the survival, growth performance, immune response and disease resistance of Litopenaeus vannamei | |
| Maeda et al. | Isolation of lactic acid bacteria from kuruma shrimp (Marsupenaeus japonicus) intestine and assessment of immunomodulatory role of a selected strain as probiotic | |
| Carasi et al. | Safety characterization and antimicrobial properties of kefir-isolated Lactobacillus kefiri | |
| KR101614262B1 (ko) | 신규 유산균, 그리고 신규 유산균을 함유하는 의약, 음식품 및 사료 | |
| Waititu et al. | Dietary supplementation with a nucleotide-rich yeast extract modulates gut immune response and microflora in weaned pigs in response to a sanitary challenge | |
| Gaudana et al. | Probiotic attributes of Lactobacillus strains isolated from food and of human origin | |
| Kobierecka et al. | In vitro characteristics of Lactobacillus spp. strains isolated from the chicken digestive tract and their role in the inhibition of Campylobacter colonization | |
| WO2021241728A1 (ja) | 抗ウイルス剤 | |
| Plumed-Ferrer et al. | Characterization of Lactococcus lactis isolates from bovine mastitis | |
| Dunislawska et al. | Transcriptome modulation by in ovo delivered Lactobacillus synbiotics in a range of chicken tissues | |
| US11690879B2 (en) | Dysosmobacter, a novel bacterial genus of the gastrointestinal microbiota and uses thereof | |
| Kobayashi et al. | Development of an in vitro immunobiotic evaluation system against rotavirus infection in bovine intestinal epitheliocytes | |
| JP4565057B2 (ja) | イムノグロブリンa誘導能の高い新規乳酸菌 | |
| KR101873682B1 (ko) | 면역 증강 효과를 갖는 신규한 락토바실러스 에시도필러스 (Lactobacillus acidophilus) KNU-02 균주 및 이의 용도 | |
| Chen et al. | Screening of lactic acid bacterial strains with antiviral activity against porcine epidemic diarrhea | |
| Feng et al. | Characterization and comparison of the adherence and immune modulation of two gut Lactobacillus strains isolated from Paralichthys olivaceus | |
| JP5646124B1 (ja) | 新規乳酸菌、並びに新規乳酸菌を含有する医薬、飲食品、及び飼料 | |
| KR101978455B1 (ko) | 락토바실러스 살리바리우스 DJ-sa-01 균주 및 이를 포함하는 면역 증강용 조성물 | |
| WO2022011271A2 (en) | Probiotics for use in the prevention or treatment of illness and/or symptoms associated with coronaviruses | |
| WO2024090413A1 (ja) | 乳酸菌、該乳酸菌由来の自然免疫活性化剤及び該乳酸菌を含有する食品 | |
| WO2024090414A1 (ja) | 乳酸菌、該乳酸菌由来の自然免疫活性化剤及び該乳酸菌を含有する食品 | |
| WO2024090415A1 (ja) | 乳酸菌、該乳酸菌由来の自然免疫活性化剤及び該乳酸菌を含有する食品 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 21813425 Country of ref document: EP Kind code of ref document: A1 |
|
| ENP | Entry into the national phase |
Ref document number: 2022526664 Country of ref document: JP Kind code of ref document: A |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2021813425 Country of ref document: EP Effective date: 20230102 |