KR20200075813A - 개선된 철-황 클러스터 전달을 갖는 세포 공장 - Google Patents
개선된 철-황 클러스터 전달을 갖는 세포 공장 Download PDFInfo
- Publication number
- KR20200075813A KR20200075813A KR1020207002518A KR20207002518A KR20200075813A KR 20200075813 A KR20200075813 A KR 20200075813A KR 1020207002518 A KR1020207002518 A KR 1020207002518A KR 20207002518 A KR20207002518 A KR 20207002518A KR 20200075813 A KR20200075813 A KR 20200075813A
- Authority
- KR
- South Korea
- Prior art keywords
- ala
- leu
- gly
- arg
- glu
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Granted
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/195—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria
- C07K14/24—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria from Enterobacteriaceae (F), e.g. Citrobacter, Serratia, Proteus, Providencia, Morganella, Yersinia
- C07K14/245—Escherichia (G)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/70—Vectors or expression systems specially adapted for E. coli
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/74—Vectors or expression systems specially adapted for prokaryotic hosts other than E. coli, e.g. Lactobacillus, Micromonospora
- C12N15/77—Vectors or expression systems specially adapted for prokaryotic hosts other than E. coli, e.g. Lactobacillus, Micromonospora for Corynebacterium; for Brevibacterium
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/10—Transferases (2.)
- C12N9/13—Transferases (2.) transferring sulfur containing groups (2.8)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/88—Lyases (4.)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/93—Ligases (6)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P17/00—Preparation of heterocyclic carbon compounds with only O, N, S, Se or Te as ring hetero atoms
- C12P17/16—Preparation of heterocyclic carbon compounds with only O, N, S, Se or Te as ring hetero atoms containing two or more hetero rings
- C12P17/167—Heterorings having sulfur atoms as ring heteroatoms, e.g. vitamin B1, thiamine nucleus and open chain analogs
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P17/00—Preparation of heterocyclic carbon compounds with only O, N, S, Se or Te as ring hetero atoms
- C12P17/18—Preparation of heterocyclic carbon compounds with only O, N, S, Se or Te as ring hetero atoms containing at least two hetero rings condensed among themselves or condensed with a common carbocyclic ring system, e.g. rifamycin
- C12P17/185—Heterocyclic compounds containing sulfur atoms as ring hetero atoms in the condensed system
- C12P17/186—Heterocyclic compounds containing sulfur atoms as ring hetero atoms in the condensed system containing a 2-oxo-thieno[3,4-d]imidazol nucleus, e.g. Biotin
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/40—Preparation of oxygen-containing organic compounds containing a carboxyl group including Peroxycarboxylic acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y201/00—Transferases transferring one-carbon groups (2.1)
- C12Y201/01—Methyltransferases (2.1.1)
- C12Y201/01197—Malonyl-CoA O-methyltransferase (2.1.1.197)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y203/00—Acyltransferases (2.3)
- C12Y203/01—Acyltransferases (2.3) transferring groups other than amino-acyl groups (2.3.1)
- C12Y203/01047—8-Amino-7-oxononanoate synthase (2.3.1.47)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y206/00—Transferases transferring nitrogenous groups (2.6)
- C12Y206/01—Transaminases (2.6.1)
- C12Y206/01062—Adenosylmethionine--8-amino-7-oxononanoate transaminase (2.6.1.62)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y208/00—Transferases transferring sulfur-containing groups (2.8)
- C12Y208/01—Sulfurtransferases (2.8.1)
- C12Y208/01006—Biotin synthase (2.8.1.6)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y208/00—Transferases transferring sulfur-containing groups (2.8)
- C12Y208/01—Sulfurtransferases (2.8.1)
- C12Y208/01008—Lipoyl synthase (2.8.1.8)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y301/00—Hydrolases acting on ester bonds (3.1)
- C12Y301/01—Carboxylic ester hydrolases (3.1.1)
- C12Y301/01085—Pimelyl-[acyl-carrier protein] methyl ester esterase (3.1.1.85)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y401/00—Carbon-carbon lyases (4.1)
- C12Y401/99—Other Carbon-Carbon Lyases (1.4.99)
- C12Y401/99017—Phosphomethylpyrimidine synthase (4.1.99.17)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y401/00—Carbon-carbon lyases (4.1)
- C12Y401/99—Other Carbon-Carbon Lyases (1.4.99)
- C12Y401/99019—2-Iminoacetate synthase (4.1.99.19)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y602/00—Ligases forming carbon-sulfur bonds (6.2)
- C12Y602/01—Acid-Thiol Ligases (6.2.1)
- C12Y602/01014—6-Carboxyhexanoate--CoA ligase (6.2.1.14)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y603/00—Ligases forming carbon-nitrogen bonds (6.3)
- C12Y603/03—Cyclo-ligases (6.3.3)
- C12Y603/03003—Dethiobiotin synthase (6.3.3.3)
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Biotechnology (AREA)
- Microbiology (AREA)
- Biomedical Technology (AREA)
- Molecular Biology (AREA)
- Medicinal Chemistry (AREA)
- Biophysics (AREA)
- General Chemical & Material Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Physics & Mathematics (AREA)
- Plant Pathology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Gastroenterology & Hepatology (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
Abstract
Description
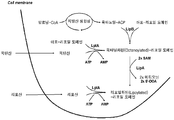
도 2 박테리아의 리포산 경로 및 리포일화된 리포일 도메인 (리포산 합성)을 일으키는 각각의 효소적 단계의 중간체를 나타내는 그림. 상기 경로에서의 주요한 효소는 LipA (리포산 신타아제) 및 LipB (옥타노일 단백질 ACP 운반 단백질: 단백질 트랜스퍼라제)를 포함하고, 기질은 SAM: S-아데노실-L-메티오닌. LipA는 리포에이트-단백질 리가아제 A; EC: 6.3.1.20이다.
도 3 박테리아의 티아민 경로 및 티아민 (THI); 티아민 모노포스페이트 (TMP) 및 티아민 디포스페이트 (TPP)의 합성을 일으키는 각각의 효소적 단계의 중간체를 나타내는 그림. 중간체의 약어: 5-아미노이미다졸 리보뉴클레오티드 (AIR: aminoimidazole ribonucleotide), 4-아미노-2-메틸-5-(포스포옥시메틸)피리미딘 (HMP-P: 4-amino-2-methyl- 5-(phosphooxymethyl)pyrimidine), 4-아미노-2-메틸-5-(디포스포메틸)피리미딘 (HMP-PP: 4-amino-2-methyl-5-(diphosphomethyl) pyrimidine), 1-데옥시-D-자일룰로스 5-포스페이트 (DXP: 1-deoxy-D-xylulose 5-phosphate), 디하이드로글리신 (DHG: dehydroglycine), 4-메틸-5-(2-포스포옥시에틸)티아졸 (THZ- P: 4-methyl-5-(2-phosphooxyethyl)thiazole), 아데노신 트리포스페이트 (ATP: adenosine triphosphate), 아데노신 모노포스페이트 (AMP: adenosine monophosphate), S-아데노실-L-메티오닌 (SAM: S-adenosyl-L-methionine), 환원된 니코틴아마이드 아데닌 디뉴클레오티드 포스페이트 (NADPH: reduced nicotinamide adenine dinucleotide phosphate), 니코틴아마이드 아데닌 디뉴클레오티드 포스페이트 (NADP+: nicotinamide adenine dinucleotide phosphate), 환원된 페레독신 (Fdx red: reduced ferredoxin), 산화된 페레독신 (Fdx ox: oxidized ferredoxin).
도 4 IPTG 유도성 bioB 발현 플라스미드를 포함하는 △bioB 균주 (대징균 BW25113) (오른쪽 패널); 및 IPTG 유도성 프레임시프트된 bioB (조기 종결 코돈) 발현 플라스미드를 포함하는 표준 균주 (대장균 BW25113) (왼쪽 패널)의 시간 경과에 따라 측정된 세포 밀도 (OD600에서 측정됨)의 그래픽 표현 (Graphical presentation). 0.1 g/L DTB, 50 ㎍/mL 카나마이신 및 0 (점), 0.01 (삼각형), 또는 0.1 (사각형) mM IPTG를 갖는 200 ㎕ mMOPS에서 성장된 4개의 생물학적 복제물에 대하여 OD620은 멀티스칸을 사용하여 측정되고, OD600으로 전환되었다. 각 지수 성장률 값은 인접한 상자에 표시된다.
도 5 275 rpm 진탕을 하며 37℃에서 20시간 동안의 인큐베이션한 후에 (하기 기재된 비오틴 정량화하는 방법과 같음), 0 또는 0.244 ㎍ 비오틴/mL까지 증가하는 농도의 비오틴이 보충된 40 ㎍/mL 제오신을 갖는 150 ㎕ mMOPS 상에서 성장된 플라스미드 pBS451을 포함하는 대장균 BS1011의 배양물의 최종 세포 밀도 (OD600에서 측정됨)를 나타낸 산포도의 그래픽 표현. 수직의 회색 점선은 비오틴 바이오어쎄이를 위한 0.024 내지 0.24 ㎍ 비오틴/L의 최적의 농도 범위를 식별한다.
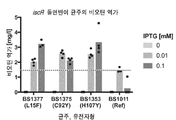
도 6 IPTG-유도성 bioB 발현 플라스미드 (pBS412)를 각각 포함하는 4개의 생물학적 복제물에서 아미노산 돌연변이를 갖는 돌연변이 IscR을 발현하는 3개의 상이한 iscR 돌연변이 균주 (BS1377, L15F), (BS1375, C92Y) 및 (BS1353, H107Y) 및 대장균 BW25113 bioB 균주 (BS1011, Ref) (균주에 대하여 표 1 참조)의 비오틴 생산을 나타내는 바 다이아그램. 균주들은 24시간 동안 37℃에서 275 rpm 진탕과 함께 0.1 g/l DTB 및 50 ㎍/mL 카나마이신을 갖는 400 ㎕ mMOPS에서 성장되었다. 바는 평균 비오틴 생산 값 (높이) 및 IPTG 유도 수준 (회색 음영)을 예시하고, 검은색 점은 개별적인 복제물 배양물로부터의 비오틴 생산을 나타내고, 수평의 점선은 참조 야생형 균주로부터의 최대 비오틴 생산을 표시한다. IPTG가 없이 배양된 경우 검출 가능한 수준의 비오틴을 생산한 균주는 없었다.
도 7 돌연변이 IscR을 발현하는 iscR 돌연변이 균주 (BS1353, H107Y), 및 대장균 BW25113 △bioB균주 (BS1011, Ref)의 세포 밀도 및 비오틴 생산의 그래픽 표현이고, 상기 각 균주는 IPTG-유도성 bioB 유전자 발현 플라스미드 (pBS412)를 포함한다. 상기 데이터는 25시간 동안 모니터링된 각각의 iscR H107Y 돌연변이 균주 (굵은 어두운 선) 및 표준 균주, 대장균 BW25113 △bioB 균주 (밝은 회색 점선)의 3개의 생물학적 복제물의 측정된 OD600 평균, 및 25시간 동안 모니터링된 각각의 iscR H107Y 돌연변이 균주 (굵은 어두운 점) 및 표준 균주, 대장균 BW25113 △bioB 균주 (밝은 회색 점)에 의한 비오틴 생산을 나타낸다. 상기 균주들은 250 mL 배플 진탕 플라스크 (baffled shake flask)에서 0.1 g/l DTB, 0.01 (A) 또는 0.5 mM IPTG (B) 및 50 ㎍/mL 카나마이신을 갖는 50 mL mMOPS에서 37℃에서 275 rpm 진탕과 함께 성장되었다. 성장률은 검정색 박스에 나타낸다.
도 8 동정된 돌연변이 균주의 iscR 유전자에서 뉴클레오티드 및 아미노산 서열 돌연변이의 위치를 나타내기 위해 주석이 달린 IscR 코딩 서열을 나타낸 그림
도 9 막대(sticks, WT 아미노산은 회색, 돌연변이 아미노산은 검정색으로 표시됨)로 표시된 L15F 및 H107Y iscR 돌연변이를 갖는 hya DNA 결합 부위 (검정색)에 결합된 IscR 이합체 (회색)의 결정 구조 (PDB entry 4HF1)를 나타낸 그림; 및 돌연변이된 잔기를 강조한 확대 이미지.
도 10 IPTG-유도성 bioB 발현 플라스미드 및 isc-오페론 (iscSUA-hscBA-fdx, iscR 유전자가 결핍된천연 대장균 isc 오페론 구조에 상응함) 또는 강한 리보좀 결합 부위 (RBS: strong ribosomal binding site)에 작동 가능하게 연결된 대장균 suf-오페론 (sufABCDSE)을 포함하는 플라스미드 및 미디엄 카피 넘버 플라스미드(p15A ori) 또는 대조군 플라스미드로부터의 T5 LacO 억제 프로모터를 포함하는 대장균 균주의 비오틴 생산을 나타내는 바 다이아그램. 상기 대조군 플라스미드는 suf- 또는 isc-오페론 대신에 IPTG-유도성 GFP를 코딩하는 유전자를 포함하였다. 각 균주의 생물학적 삼중물 (triplicates)은 100 μg/mL 암피실린 및 50 μg/mL 스펙티노마이신을 갖는 mMOPS에서 0.1 g/l (DTB)를 기질로서 제공하여 낮은 (0.01 mM IPTG) 및 높은 (0.1 mM IPTG) 유도 하에 배양되었다. 비오틴 생산이 성장 기반의 바이오어쎄이를 이용하여 평가된 후, 상기 균주들은 딥 웰 플레이트에서 24시간 동안 37℃에서 275 rpm으로 성장되었다. 바는 평균 비오틴 성장 값 (높이) 및 IPTG 유도 수준 (회색 음영)을 예시하고, 검은색 점은 개별적인 복제물로부터의 비오틴 생산을 나타내고, X표(crosses)는 OD600으로 측정된 각 균주의 종점(end-point, end) 세포 밀도를 나타낸다. 0.01 mM IPTG로 유도된 경우, 검출 가능한 수준의 비오틴을 생산한 균주는 없었다.
도 11 삼중불에서 수행된 4개의 상이한 샘플에서의 BioB 단백질 발현 수준 및 비오틴 생산의 상관관계의 그래픽 표현. 상기 균주는 pBS430 (bioB 프레임시프트 IPTG 유도성 플라스미드)을 갖는 BS1013 (대장균 BW25113, 백그라운드 균주), pBS412 (bioB IPTG 유도성 플라스미드)를 갖는 BS1011 (△bioB를 갖는 BS1013), pBS412를 갖는 BS1353 (iscR H107Y 돌연변이를 갖는 BS1011)이었다. 균주들은 0.1 g/l DTB 및 그래프에 나타낸 바와 같은 IPTG를 갖는 mMOPS에서 성장되었다.
도 12 IPTG-유도성 bioB 발현 플라스미드 및 다음 중 iscR의 게놈 변이체를 포함하는 대장균 △bioB 균주의 비오틴 생산을 나타내는 바 다이아그램: 야생형 (iscR WT), 22번 위치에서 종결 코돈으로 돌연변이된 E22* 글루탐산을 코딩하는 녹-아웃 돌연변이 (iscR KO), 92번 위치에서 시스테인으로부터 티로신으로의 치환을 코딩하는 돌연변이 (iscR C92Y). 바는 IPTG 유도의 주어진 수준 (회색의 음영)에서의 비오틴 생산 평균 값 (높이)을 예시하고, 점은 개별적인 복제물로부터의 비오틴 생산을 나타낸다. 각 균주의 생물학적 삼중물은 100 μg/mL 암피실린을 갖는 mMOPS에서 0.1 g/l DTB를 기질로서 제공하여 없는 (0 mM IPTG), 낮은 (0.01 mM IPTG) 및 낮은 (0.1 mM IPTG) 유도 하에 배양되었다. 비오틴 생산이 성장 기반의 바이오어쎄이를 이용하여 평가된 후, 각 균주는 딥 웰 플레이트에서 24시간 동안 37℃에서 275 rpm으로 성장되었다.
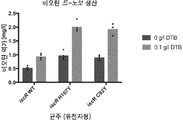
도 13 비오틴-오페론 플라스미드 및 다음 중 iscR의 게놈 변이체를 포함하는 대장균 △bioA △bioBFCD 균주에 의한 비오틴 생산을 나타내는 바 다이아그램: 야생형 (iscR WT), 돌연변이 iscR H107Y (107번 위치에서 히스티딘으로부터 티로신으로의 치환을 코딩함), 92번 위치에서 시스테인으로부터 티로신으로의 치환을 코딩하는 돌연변이 (iscR C92Y). 각 균주의 생물학적 사중물 (quadruplicate)은 0.1 데스티오비오틴 (DTB)을 기질로서 제공하거나 제공하지 않고 10 μg/mL 테트라사이클린을 갖는 mMOPS에서 배양되었다. 비오틴 생산이 성장 기반의 바이오어쎄이를 이용하여 평가된 후, 상기 균주들은 딥 웰 플레이트에서 24시간 동안 37℃에서 275 rpm으로 성장되었다. 바는 비오틴 생산 평균 값 (높이) 및 DTB의 공급 여부 (회색의 음영)를 예시하고, 검은색 점은 개별적인 복제물로부터의 비오틴 생산을 나타낸다.
도 14 IPTG 유도성 lipA (pBS993, 표 4 참조)를 각각 포함하는 3개의 생물학적 복제물에서 아미노산 돌연변이를 갖는 돌연변이 IscR을 발현하는 2개의 상이한 iscR 돌연변이 균주 (BS1375 C92Y) 및 (BS1353, H107Y) 및 대조군 균주 (BS1011, IscR WT) (균주에 대하여 표 1 참조)의 리포산 생산 및 생산 (X표) 24시간 후의 종점(최종) OD600을 나타내는 바 다이아그램. 균주들은 100 g/mL 암피실린, 0.1 mM 비오틴, 0.6 g/l 옥탄산 및 0.01 mM IPTG을 갖는 400 ㎕ mMOPS에서 24시간 동안 37℃에서 275 rpm 진탕과 함께 성장되었다. 바는 리포산 생산 평균 값 (높이)을 예시하고, 검은색 점은 개별적인 복제물 배양물로부터의 리포산 생산을 나타낸다. 종점 OD600이 동일하게 유지되더라도, 평균 리포산 생산은 1.79-배 증가하는 것을 볼 수 있다.
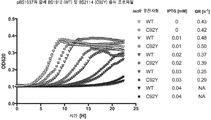
도 15 표준 균주 (대장균 BW25113), △lipA (WT, 삼각형); 및 돌연변이 iscR (C92Y)를 갖는 △lipA 균주 (C92Y, 사각형)로서, 상기 균주 모두 IPTG 유도성 lipA 발현 플라스미드 (pBS1037)를 포함하는 균주의 시간 경과에 따라 측정된, 세포 밀도 (OD620에서 측정됨)의 그래픽 표현. OD620은 0.6 g/L 옥탄산, 100 g/mL 암피실린 및 0 내지 0.04 mM IPTG (증가에 따라 회색 음영의 어둠이 증가함)을 갖는 200 ㎕ mMOPS에서 성장된 6개의 생물학적 균주 복제물에 대하여 멀티스칸을 이용하여 측정되었다. 각 생존율 (GR: growth rate)은 오른쪽에 나타낸다.
도 16 전체 티아민 경로 유전자, thiCEFSGHMD를 발현하는 플라스미드 (pBS140)를 각각 포함하는 4개의 생물학적 복제물에서 아미노산 돌연변이를 갖는 돌연변이 IscR을 발현하는 2개의 상이한 iscR 돌연변이 균주 (BS2019, C92Y) 및 (BS2020, H107Y) 및 대장균 BW25113 △thiP, thiL* 균주 (BS750, Ref) (균주에 대하여 표 5 참조)의 티아민 생산을 나타내는 바 다이아그램. 균주들은 50 ㎍/mL 카나마이신을 갖는 400 ㎕ mMOPS에서 24시간 동안 37℃에서 275 rpm 진탕과 함께 성장되었다. 바는 종점 OD600에 대하여 보정된 티오크롬 어쎄이 (티아민, TMP 및 TPP를 함유함)에 의해 측정된 바와 같은 상층액에서의 티아민 생산 평균 값 (높이)을 예시하고, 검은색 점은 개별적인 복제물 배양물로부터의 티아민 생산을 나타낸다. OD 정규화(normalized) 역가는 돌연변이 균주 (BS2019 및 BS2020)에서 표준 균주 (BS750)와 비교하여 1.43-배 향상된 것을 확인할 수 있다.
도 17 IPTG-유도성 BioB 과발현 플라스미드 pBS679 단독으로 (BS1937) 또는 FldA-Fpr의 항시적 과발현을 갖는 pBS1112 (BS2185) 또는 GFP의 항시적 과발현을 갖는 pBS1054 (BS2707)를 더한 대장균 △bioABFCD iscR H107Y (107번 위치에서 히스티딘으로부터 티로신으로의 치환을 코딩함) 균주에 의한 비오틴 생산을 나타내는 바 다이아그램. 각 균주는 100 ㎍/ml 암피실린, 기질로서 0.1 g/L 데스티오비오틴 (DTB), 및 0, 0.01, 0.025, 0.05, 0.075 또는 0.1 mM IPTG를 갖는 mMOPS에서 배양되었다. BS2185 및 BS2707을 위한 배지는 50 ㎍/ml 카나마이신을 포함하는 것을 제외하고는 동일하였다. 비오틴 생산이 성장 기반의 바이오어쎄이를 이용하여 평가된 후, 상기 균주들은 딥 웰 플레이트에서 24시간 동안 37℃에서 275 rpm 진탕과 함께 성장되었다. 바는 각각의 균주에 의한 비오틴 생산 값 (높이)을 예시한다: BS1937 (검은색 바); BS2185 (회색 바); 및 BS2707 (체크무늬 회색).
도 18 IPTG-유도성 BioB 과발현 플라스미드 pBS679를 포함하는 대장균 △bioABFCD iscR H107Y (107번 위치에서 히스티딘으로부터 티로신으로의 치환을 코딩함) 균주에 의한 비오틴 생산을 나타내는 바 다이아그램. BS2185는 FldA-Fpr의 항시적 과발현을 갖는 pBS1112를 더 포함한다. BS1937은 100 ㎍/ml 암피실린, 기질로서 0.1 g/L 데스티오비오틴 (DTB), 및 0.025 mM IPTG 유도를 갖는 mMOPS에서 배양되었다. BS2185를 위한 배지는 동일하나, 50 ㎍/ml 카나마이신을 더 포함하였다. 비오틴 생산이 성장 기반의 바이오어쎄이를 이용하여 평가된 후, 상기 균주들은 딥 웰 플레이트에서 24시간 동안 37℃에서 275 rpm과 함께 성장되었다. 어두운 회색 바는 비오틴 생산 평균 값 (높이) (BS1937 n=6 및 BS2185 n=8)을 예시하고, 밝은 회색 바는 종점 OD600을 예시한다. 검은색 점은 비오틴 생산 및 개별적인 복제물로부터의 종점 OD600을 나타낸다.
| 명칭 | 설명 |
| BS1013 | 하기 유전자형을 갖는 E. coli K-12 BW25113 모체 균주:rrnB3 △lacZ4787 hsdR514 △(araBAD)567 △(rhaBAD)568 rph-1 |
| BS1011 | E coli K-12 BW25113로부터 유래된 △bioB 1(JW0758-1) |
| BS1353 | iscR 유전자에 H107Y 돌연변이를 포함하는 BS1011 파생체(derivative) |
| BS1113 | IPTG-유도성 BioB 발현을 제공하는 pBS412 플라스미드를 포함하는 BS1011 파생체 |
| BS1375 | iscR 유전자에 C92Y 돌연변이를 포함하는 BS1011 파생체 |
| BS1377 | iscR 유전자에 L15F 돌연변이를 포함하는 BS1011 파생체 |
| 명칭 | 설명 |
| pBS412 | T5 lacO 억제 프로모터(repressed promoter)로부터의 BioB [서열번호 22] 과발현 플라스미드 (kanR, SC101) |
| pBS430 | T5 lacO 억제 프로모터 [서열번호 25]로부터의 bioB 1(kanR, SC101)에서 초기 프레임 시프트 돌연변이를 갖는 pBS412 |
| pBS451 | 항시적으로 발현되는 GFP [서열번호 287] (zeoR, p15A) |
| pBS281 | 미디엄 카피 넘버 플라스미드(medium copy number plasmid) (p15A ori)에서 복제된 IPTG 유도성 T5 프로모터로부터의 E. coli isc 오페론 (iscSUA-hscBA-fdx) |
| pBS282 | 미디엄 카피 넘버 플라스미드 (p15 ori)에서 복제된 IPTG 유도성 T5 프로모터로부터의 E. coli suf 오페론 (sufABCDSE) |
| pBS231 | IPTG 유도성 T5 프로모터로부터 sfGFP 단백질을 코딩하는 유전자를 발현하는 미디엄 카피 넘버 플라스미드 (p15 ori) |
| pBS936 | bio 오퍼레이터 위치에 "타입 9(type 9)" 돌연변이를 갖는 E coli 유래의 천연 비오틴-오페론 (Ifuku et al., 1993) |
| 명칭 | 설명 |
| BS1912 | E.coli K-12 BW25113로부터 유래된 △lipdA |
| BS2114 | iscR에서 H107Y 돌연변이를 포함하는 BS1912 파생체 |
| 명칭 | 설명 |
| pBS993 | AceF [서열번호 119] 발현의 추가적인 항시성 발현을 갖는 T5 lacO 억제 프로모터 [서열번호 234]로부터의 Lipid A [서열번호 103] 과발현 플라스미드 (kanR, SC101) |
| pBS1037 | AceF [서열번호 119] 발현의 낮은 RBS 강도 및 SC101 대신에 p15A 복제 기점을 갖는 pBS993 파생체 |
| pBS451 | 항시적으로 발현되는 GFP [서열번호 287] (zero, p15A) |
| 명칭 | 설명 |
| BS750 | 코딩된 TMP 키나아제에서 G133D 치환을 일으키는 천연 thiL 유전자: 코돈 133에서 GGT로부터 GAC로의 점 돌연변이를 포함하는 BS1013 파생체, BW25113 △thiP |
| BS2019 | iscR에 C92Y 돌연변이를 포함하는 BS750 파생체 |
| BS2020 | iscR에 H107Y 돌연변이를 포함하는 BS750 파생체 |
| 명칭 | 설명 |
| pBS140 | thiC 오페론 (apFAB46 프로모터 [서열번호 147] 및apFAB377 종결자 [서열번호 153]에 기능적으로 연결됨) 및 thiM 오페론 (apFAB71 프로모터 [서열번호 149] 및 apFAB378 종결자 [서열번호 152]에 기능적으로 연결됨)의 조합으로 구성된; 대장균 티아민 경로 유전자 thiCEFSGHMD를 포함하는 벡터 |
| pBS100 | pBS140의 구성에 사용되기 위한 빈 벡터 |
| pBS93 | pFAB70 프로모터 [서열번호 148] 및 ap FAB381 종결자 [서열번호 154]에 기능적으로 연결된 대장균에서의 발현에 최적화된 애기장대 (Arabidopsis thaliana) AT5G32470.1 포스파타아제 코돈을 코딩하는 합성 유전자 (synthetic gene)를 포함하는 벡터 |
| pBS209 | 대장균으로부터 추가적인 thiC를 갖는 pBS140에 기반한 플라스미드 |
| 명칭 | 설명 |
| BS1013 | 다음의 유전자형을 갖는 E. coli K-12 BW25113 모체 균주: rrnB3 △lacZ4787 hsdR514 △(araBAD)567 △(rhaBAD)568 rph-1 |
| BS1011 | E. coli K-12 BW25113로부터 유래된 △bioB (JW0758-1) |
| BS1353 | iscR에 H107Y 돌연변이를 포함하는 BS1011 파생체 |
| BS1615 | △bioAFCD의 추가적인 결실을 갖는 BS1011 파생체 |
| BS1937 | IPTG-유도성 BioB 발현을 제공하는 pBS679 플라스미드를 포함하는 BS1615 파생체 |
| BS2185 | IPTG-유도성 BioB 발현을 제공하는 pBS679 플라스미드 및 FldA-Fpr 항시적 발현을 제공하는 pBS1112를 포함하는 BS1615 파생체 |
| BS2707 | IPTG-유도성 BioB 발현을 제공하는 pBS679 플라스미드 및 GFP 항시적 발현을 제공하는 pBS1054를 포함하는 BS1615 파생체 |
| 명칭 | 설명 |
| pBS679 | T5 lacO 억제 프로모터 [서열번호 25]로부터의 BioB [서열번호 22] 과발현 플라스미드 (ampR, pSC101) 및 |
| pBS1054 | apFAB378 종결자 [서열번호 292]를 갖는 apFAB309 항시적 프로모터 [서열번호 291]로부터의 GFP [서열번호 276] 과발현 플라스미드 (kanR, pBR322) |
| pBS1112 | apFAB306 항시적 프로모터 (apFAB306-FldA-Fpr 유전자-apFAB378 종결자 [서열번호 288])로부터의 FldA-Fpr 과발현 플라스미드 (kanR, pBR322) |
Claims (18)
- 하기를 포함하는, 비오틴 또는 리포산 또는 티아민의 증가된 생산을 위한 유전자 변형 박테리아:
a) 돌연변이 IscR 폴리펩티드를 코딩하는 유전자 변형 내인성 iscR 유전자로서,
상기 돌연변이 IscR 폴리펩티드의 아미노산 서열은 서열번호 2, 4, 6, 8, 10, 12 및 14로 이루어진 군으로부터 선택된 서열과 적어도 80% 아미노산 서열 상동성을 갖는 것이고, 상기 아미노산 서열은 i) L15X, C92X, C98X, C104X, 및 H107X로 이루어진 군으로부터 선택된 적어도 하나의 아미노산 치환을 갖는 것이고; 상기 X는 서열번호 2, 4, 6, 8, 10, 12 및 14에 상응하는 아미노산 잔기 이외의 임의의 아미노산인 것인, 유전자; 및
b) 적어도 하나의 전이 유전자로서,
ii) 비오틴 신타아제 (EC 2.8.1.6) 활성을 갖는 폴리펩티드;
iii) 리포산 신타아제 (EC2.8.1.8) 활성을 갖는 폴리펩티드;
iv) HMP-P 신타아제 (EC 4.1.99.17) 활성을 갖는 폴리펩티드; 및
v) 티로신 리아제 (EC 4.1.99.19) 활성을 갖는 폴리펩티드로 이루어진 군으로부터 선택된 폴리펩티드를 코딩하는 적어도 하나의 전이 유전자. - 청구항 1에 있어서, 상기 돌연변이 IscR 폴리펩티드에서 상기 적어도 하나의 아미노산 치환은 다음으로 이루어진 군으로부터 선택되는 것인, 비오틴 또는 리포산 또는 티아민의 증가된 생산을 위한 유전자 변형 박테리아:
a. L15X, 상기 X는 F, Y, M 및 W 중 어느 하나임;
b. C92X, 상기 X는 Y, A, M, F 및 W 중 어느 하나임;
c. C98X, 상기 X는 A, V, I, L, F 및 W 중 어느 하나임;
d. C104X, 상기 X는 A V, I, L, F 및 W 중 어느 하나임; 및
e. H107X; 상기 X는 A, Y, V, I, 및 L 중 어느 하나임. - 청구항 1 또는 2에 있어서, 상기 비오틴 신타아제 활성을 갖는 폴리펩티드 (EC 2.8.1.6)를 코딩하는 적어도 하나의 전이 유전자는 다음으로 이루어진 군으로부터 선택된 하나 이상의 폴리펩티드를 코딩하는 추가적인 전이 유전자를 더 포함하고:
a. SAM (S-아데노실메티오닌)-의존적 메틸트랜스퍼라제 (EC 2.1.1.197) 활성을 갖는 폴리펩티드;
b. 7-케토-8-아미노펠라르곤산 (KAPA) 신타아제 (EC 2.3.1.47) 활성을 갖는 폴리펩티드;
c. 7,8-디아미노펠라르곤산 (DAPA) 신타아제 (EC:2.6.1.62 또는 EC2.6.1.105) 활성을 갖는 폴리펩티드; 및
d. 데티오비오틴 (DTB) 신타아제 (EC 6.3.3.3) 활성을 갖는 폴리펩티드;
e. 피멜로일-[아실-운반 단백질] 메틸 에스테르 에스테라제 (EC 3.1.1.85)를 갖는 폴리펩티드; 및
f. 6-카복시헥사노에이트-CoA 리가제 (EC 6.2.1.14) 활성을 갖는 폴리펩티드
상기 박테리아는 비오틴의 증가된 생산을 위한 것인 유전자 변형 박테리아. - 청구항 1 또는 2에 있어서, 상기 적어도 하나의 전이 유전자는 리포산 신타아제 (EC 2.8.1.8) 활성을 갖는 폴리펩티드를 코딩하고, 다음으로 이루어진 군으로부터 선택된 하나 이상의 폴리펩티드를 코딩하는 추가적인 전이 유전자를 더 포함하는 것이고:
a. 옥타노일트랜스퍼라제 (EC 2.3.1.181) 활성을 갖는 폴리펩티드;
b. 피루브산 탈수소효소 (EC 2.3.1.12)의 디하이드로리포일라이신-잔기 아세틸트랜스퍼라제 성분을 포함하는 폴리펩티드; 및
c. 리포에이트-단백질 리가아제 A (EC:6.3.1.20) 활성을 갖는 폴리펩티드;
상기 박테리아는 리포산의 증가된 생산을 위한 것인 유전자 변형 박테리아. - 청구항 1 또는 2에 있어서, 상기 HMP-P 신타아제 (EC 4.1.99.17) 활성을 갖는 폴리펩티드를 코딩하는 적어도 하나의 전이 유전자 및/또는 티로신 리아제 (EC 4.1.99.19) 활성을 갖는 폴리펩티드를 코딩하는 적어도 하나의 전이 유전자는 다음으로 이루어진 군으로부터 선택된 하나 이상의 폴리펩티드를 코딩하는 추가적인 전이 유전자를 더 포함하는 것이고:
a. ThiS 아데닐트랜스퍼라제 (EC2.7.7.73) 활성을 갖는 ThiF 폴리펩티드;
b. 티아민 포스페이트 신타아제 (EC 2.5.1.3) 활성을 갖는 ThiE 폴리펩티드;
c. 티아졸 신타아제 활성 (E.C.2.8.1.10) 활성을 갖는 ThiG 폴리펩티드;
d. 포스포하이드록시메틸피리미딘 키나아제 (EC 2.7.4.7) 활성을 갖는 ThiD 폴리펩티드;
e. 글리신 옥시다아제 (EC 1.4.3.19) 활성을 갖는 ThiO 폴리펩티드;
f. 황-운반 단백질 활성을 갖는 ThiS 폴리펩티드;
g. 하이드록시에틸티아졸 키나아제 (EC2.7.1.50) 활성을 갖는 ThiM 폴리펩티드; 및
h. 모노-포스페이트 포스파타아제 (E.C. 3.1.3.-) 활성을 갖는 폴리펩티드;
상기 박테리아는 티아민의 증가된 생산을 위한 것인 유전자 변형 박테리아. - 청구항 5에 있어서, 상기 박테리아는 다음을 코딩하는 추가적인 전이 유전자를 포함하는 것인 유전자 변형 박테리아:
a. HMP-P 신타아제 (EC 4.1.99.17) 활성을 갖는 ThiC 폴리펩티드;
b. 티로신 리아제 (EC 4.1.99.19) 활성을 갖는 ThiH 폴리펩티드 또는 글리신 옥시다아제 (EC 1.4.3.19) 활성을 갖는 ThiO 폴리펩티드;
c. ThiS 아데닐트랜스퍼라제 (EC 2.7.7.73) 활성을 갖는 ThiF 폴리펩티드;
d. 티아민 포스페이트 신타아제 (EC 2.5.1.3) 활성을 갖는 ThiE 폴리펩티드;
e. 티아졸 신타아제 (E.C.2.8.1.10) 활성을 갖는 ThiG 폴리펩티드;
f. 포스포하이드록시메틸피리미딘 키나아제 (EC 2.7.4.7) 활성을 갖는 ThiD 폴리펩티드;
g. 황-운반 단백질 활성을 갖는 ThiS 폴리펩티드; 및
h. 티아민 모노-포스페이트 포스파타아제 (E.C. 3.1.3.-) 활성을 갖는 폴리펩티드. - 청구항 1 내지 6 중 어느 한 항에 있어서, 상기 적어도 하나의 전이 유전자 및 상기 하나 이상의 추가적인 전이 유전자는 항시성 프로모터(constitutive promoter)에 작동 가능하게 연결된 것인 유전자 변형 박테리아.
- 청구항 1 내지 7 중 어느 한 항에 있어서, 상기 박테리아는 에셔리키아 (Escherichia), 바실러스 (Bacillus), 브레비박테리움 (Brevibacterium), 버크홀데리아 (Burkholderia), 캄필로박터 (Campylobacter), 코리네박테리움 (Corynebacterium), 슈도모나스 (Pseudomonas), 셀라티아 (Serratia), 락토바실러스 (Lactobacillus), 락토코커스 (Lactocooccus), 아시네토박터 (Acinetobacter), 슈도모나스 (Pseudomonas), 및 아세토박터 (Acetobacter)로 이루어진 군으로부터 선택된 박테리아의 속인 것인 유전자 변형 박테리아.
- 하기 단계를 포함하는 비오틴을 생산하는 방법:
a. 청구항 1 내지 3, 7 및 8 중 어느 한 항에 따른 유전자 변형 박테리아를 배양물을 생산하기 위한 증식 배지 내로 도입하는 단계로서, 여기에서 상기 적어도 하나의 전이 유전자는 비오틴 신타아제 (EC 2.8.1.6) 활성을 갖는 폴리펩티드를 코딩하는 것인 단계;
b. 상기 배양물을 배양하는 단계; 및
c. 상기 배양에 의해 생산된 비오틴을 회수하고, 선택적으로 회수된 비오틴을 정제하는 단계. - 리포산을 생산하는 방법으로서,
a. 청구항 1, 2, 4, 7 및 8 중 어느 한 항에 따른 유전자 변형 박테리아를 배양물을 생산하기 위한 증식 배지 내로 도입하는 단계로서, 여기에서 상기 적어도 하나의 전이 유전자는 리포산 신타아제 (EC 2.8.1.6) 활성을 갖는 폴리펩티드를 코딩하는 것인 단계;
b. 상기 배양물을 배양하는 단계; 및
c. 상기 배양에 의해 생산된 리포산을 회수하고, 선택적으로 회수된 리포산을 정제하는 단계를 포함하는 방법. - 티아민을 생산하는 방법으로서,
a. 청구항 1, 2, 및 5 내지 8 중 어느 한 항에 따른 유전자 변형 박테리아를 배양물을 생산하기 위한 증식 배지 내로 도입하는 단계로서, 여기에서 상기 적어도 하나의 전이 유전자는 HMP-P 신타아제 (EC 4.1.99.17) 활성을 갖는 폴리펩티드 및/또는 티로신 리아제 (EC 4.1.99.19) 활성을 갖는 폴리펩티드를 코딩하는 것인 단계;
b. 상기 배양물을 배양하는 단계; 및
c. 상기 배양에 의해 생산된 티아민을 회수하고, 선택적으로 회수된 티아민을 정제하는 단계를 포함하는 방법. - 청구항 9 내지 11 중 어느 한 항에 있어서, 상기 증식 배지는 글루코스, 말토스, 갈락토스, 프럭토스, 수크로스, 아라비노스, 자일로스, 라피노스, 만노스,및 락토스, 또는 이들의 임의의 조합으로부터 선택된 탄소원을 포함하는 것인, 비오틴, 리포산 및 티아민 중 어느 하나를 생산하는 방법.
- 유전자 변형 박테리아에서 비오틴, 리포산 또는 티아민 중 어느 하나의 생산을 증가시키기 위한 돌연변이 iscR 폴리펩티드를 코딩하는 유전적으로 변형된 유전자의 용도로서, 상기 박테리아는:
i. 비오틴 신타아제 (EC 2.8.1.6) 활성을 갖는 폴리펩티드;
ii. 리포산 신타아제 (EC 2.8.1.8) 활성을 갖는 폴리펩티드;
iii. HMP-P 신타아제 (EC 4.1.99.17) 활성을 갖는 폴리펩티드; 및
iv. 티로신 리아제 (EC 4.1.99.19) 활성을 갖는 폴리펩티드로 이루어진 군으로부터 선택된 폴리펩티드를 코딩하는 적어도 하나의 전이유전자를 포함하고 발현하는 것이고;
상기 유전적으로 변형된 유전자는 돌연변이 IscR 폴리펩티드를 코딩하는 내인성 iscR 유전자이고, 여기에서 상기 돌연변이 IscR 폴리펩티드의 아미노산 서열은 서열번호 2, 4, 6, 8, 10, 12 및 14와 적어도 80% 아미노산 서열 상동성을 갖는 것이고, 상기 아미노산 서열은: L15X, C92X, C98X, C104X, 및 H107X로 이루어진 군으로부터 선택된 적어도 하나의 아미산 치환을 갖는 것이고; 상기 X는 서열번호 2, 4, 6, 8, 10, 12 및 14에 상응하는 아미노산 잔기 이외의 임의의 아미노산인 것인 용도. - 청구항 13에 있어서, 상기 돌연변이 IscR 폴리펩티드에서 상기 적어도 하나의 아미노산 치환은 다음으로 이루어진 군으로부터 선택되는 것인, 유전자 변형 박테리아에서 비오틴, 리포산 또는 티아민 중 어느 하나의 생산을 증가시키기 위한 돌연변이 iscR 폴리펩티드를 코딩하는 유전적으로 변형된 유전자의 용도:
a. L15X, 상기 X는 F, Y, M 및 W 중 어느 하나임;
b. C92X, 상기 X는 Y, A, M, F 및 W 중 어느 하나임;
c. C98X, 상기 X는 A, V, I, L, F 및 W 중 어느 하나임;
d. C104X, 상기 X는 A V, I, L, F 및 W 중 어느 하나임; 및
e. H107X; 상기 X는 A, Y, V, I, 및 L 중 어느 하나임. - 비오틴, 리포산 또는 티아민 중 어느 하나의 증가된 생산을 위한, 청구항 1 내지 8 중 어느 한 항에 따른 유전자 변형 박테리아의 용도.
- 청구항 1 내지 8 중 어느 한 항에 있어서, 상기 박테리아는 하기 군으로부터 선택된 하나 이상의 유전자를 더 포함하는 것인 유전자 변형 박테리아:
a. 플라보독신/페레독신-NADP 환원 효소 (EC:1.18.1.2 및 EC 1.19.1.1)를 코딩하는 유전자;
b. 피루브산-플라보독신/페레독신 산화 환원 효소 (EC 1.2.7)를 코딩하는 유전자;
c. 플라보독신을 코딩하는 유전자;
d. 페레독신을 코딩하는 유전자; 및
e. 플라보독신 및 페레독신-NADP 환원 효소를 코딩하는 유전자;
여기서, 상기 하나 이상의 유전자는 상기 박테리아에서 상기 하나 이상의 유전자의 발현을 증가시킬 수 있는 비-천연(non-native) 프로모터에 작동 가능하게-연결되는 것이고, 상기 하나 이상의 유전자는 천연 유전자(native gene) 또는 전이 유전자일 수 있다. - 비오틴, 리포산 또는 티아민 중 어느 하나의 증가된 생산을 위한, 청구항 16에 따른 유전자 변형 박테리아의 용도.
- 청구항 9 내지 12 중 어느 한 항에 있어서, , 상기 유전자 변형 박테리아는 다음의 군으로부터 선택된 하나 이상의 유전자를 더 포함하는 것인 비오틴, 리포산 및 티아민 중 어느 하나를 생산하는 방법:
a. 플라보독신/페레독신-NADP 환원 효소 (EC:1.18.1.2 및 EC 1.19.1.1)를 코딩하는 유전자;
b. 피루브산-플라보독신/페레독신 산화 환원 효소 (EC 1.2.7)를 코딩하는 유전자;
c. 플라보독신을 코딩하는 유전자;
d. 페레독신을 코딩하는 유전자; 및
e. 플라보독신 및 페레독신-NADP 환원 효소;
상기 하나 이상의 유전자는 상기 박테리아에서 상기 하나 이상의 유전자의 발현을 증가시킬 수 있는 비-천연 프로모터에 작동 가능하게-연결된 것이고, 상기 하나 이상의 유전자는 천연 유전자 또는 전이 유전자일 수 있다.
Applications Claiming Priority (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP17181503.8 | 2017-07-14 | ||
| EP17181503 | 2017-07-14 | ||
| PCT/EP2018/068989 WO2019012058A1 (en) | 2017-07-14 | 2018-07-12 | CELL FACTORY HAVING IMPROVED DISTRIBUTION OF IRON SULFUR AMAS |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| KR20200075813A true KR20200075813A (ko) | 2020-06-26 |
| KR102760107B1 KR102760107B1 (ko) | 2025-01-31 |
Family
ID=59362984
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| KR1020207002518A Active KR102760107B1 (ko) | 2017-07-14 | 2018-07-12 | 개선된 철-황 클러스터 전달을 갖는 세포 공장 |
Country Status (9)
| Country | Link |
|---|---|
| US (1) | US11851461B2 (ko) |
| EP (1) | EP3652198B1 (ko) |
| JP (2) | JP2020527359A (ko) |
| KR (1) | KR102760107B1 (ko) |
| CN (1) | CN110869384A (ko) |
| AU (1) | AU2018300754B2 (ko) |
| BR (1) | BR112020000548A2 (ko) |
| CA (1) | CA3069650A1 (ko) |
| WO (1) | WO2019012058A1 (ko) |
Families Citing this family (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN108414772B (zh) * | 2018-03-28 | 2021-02-09 | 河南科技大学 | 一种用于研究细菌中类泛素系统的试剂盒及其应用 |
| EP3938518A4 (en) | 2019-03-12 | 2023-01-11 | Terra Bioworks, Inc. | EXPRESSION VECTOR |
| US20230041211A1 (en) * | 2019-12-20 | 2023-02-09 | Basf Se | Decreasing toxicity of terpenes and increasing the production potential in micro-organisms |
| EP4168566A1 (en) | 2020-06-18 | 2023-04-26 | Biosyntia ApS | Methods for producing biotin in genetically modified microorganisms |
| US20240225063A9 (en) | 2021-02-22 | 2024-07-11 | Symrise Ag | Biotechnological production of meat-like flavourings |
| WO2023285585A2 (en) | 2021-07-16 | 2023-01-19 | Biosyntia Aps | Microbial cell factories producing vitamin b compounds |
| GB202203725D0 (en) * | 2022-03-17 | 2022-05-04 | Univ Nottingham | Bio-manufacturing process |
| EP4555076A1 (en) * | 2022-07-15 | 2025-05-21 | Biosyntia ApS | Microbial cell factories producing thiamine |
| WO2025210159A2 (en) * | 2024-04-03 | 2025-10-09 | Biosyntia Aps | Cell factory for production of sulfurol |
| WO2025219247A1 (en) | 2024-04-15 | 2025-10-23 | Biosyntia Aps | Liposomal preparation encapsulating biotin |
Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2002085293A2 (en) * | 2001-04-20 | 2002-10-31 | Cargill, Incorporated | Production of alpha-lipoic acid |
| WO2007046943A2 (en) * | 2005-10-19 | 2007-04-26 | Stephen F. Austin State University | Method of producing biotin |
| WO2009091582A1 (en) * | 2008-01-17 | 2009-07-23 | Indigene Pharmaceuticals, Inc. | PRODUCTION OF R-α-LIPOIC ACID BY FERMENTATION USING GENETICALLY ENGINEERED MICROORGANISMS |
| WO2017103221A1 (en) * | 2015-12-18 | 2017-06-22 | Biosyntia Aps | A genetically modified bacterial cell factory for thiamine production |
Family Cites Families (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20060127993A1 (en) * | 2003-06-02 | 2006-06-15 | Goese Markus G | Thiamin production by fermentation |
| JP2011160673A (ja) * | 2010-02-04 | 2011-08-25 | Fujirebio Inc | 水素ガスの生産方法及びそのための微生物 |
| JP5950830B2 (ja) * | 2010-02-17 | 2016-07-13 | ビュータマックス・アドバンスド・バイオフューエルズ・エルエルシー | Fe−Sクラスター要求タンパク質の活性の改善 |
| CN104450762B (zh) * | 2013-09-17 | 2018-03-20 | 中国科学院广州生物医药与健康研究院 | α‑硫辛酸的生物合成方法、工程菌株及其制备方法 |
| CN106086052B (zh) | 2016-07-01 | 2019-11-01 | 福建师范大学 | 生产吡咯喹啉醌的细菌及其应用 |
| EP3683227A1 (en) * | 2019-01-16 | 2020-07-22 | Biosyntia ApS | Cell factories for improved production of compounds and proteins dependent on iron sulfur clusters |
-
2018
- 2018-07-12 EP EP18740207.8A patent/EP3652198B1/en active Active
- 2018-07-12 KR KR1020207002518A patent/KR102760107B1/ko active Active
- 2018-07-12 US US16/630,203 patent/US11851461B2/en active Active
- 2018-07-12 CN CN201880045992.6A patent/CN110869384A/zh active Pending
- 2018-07-12 BR BR112020000548-7A patent/BR112020000548A2/pt unknown
- 2018-07-12 AU AU2018300754A patent/AU2018300754B2/en active Active
- 2018-07-12 JP JP2020523046A patent/JP2020527359A/ja active Pending
- 2018-07-12 CA CA3069650A patent/CA3069650A1/en active Pending
- 2018-07-12 WO PCT/EP2018/068989 patent/WO2019012058A1/en not_active Ceased
-
2023
- 2023-04-05 JP JP2023061066A patent/JP7617658B2/ja active Active
Patent Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2002085293A2 (en) * | 2001-04-20 | 2002-10-31 | Cargill, Incorporated | Production of alpha-lipoic acid |
| WO2007046943A2 (en) * | 2005-10-19 | 2007-04-26 | Stephen F. Austin State University | Method of producing biotin |
| WO2009091582A1 (en) * | 2008-01-17 | 2009-07-23 | Indigene Pharmaceuticals, Inc. | PRODUCTION OF R-α-LIPOIC ACID BY FERMENTATION USING GENETICALLY ENGINEERED MICROORGANISMS |
| WO2017103221A1 (en) * | 2015-12-18 | 2017-06-22 | Biosyntia Aps | A genetically modified bacterial cell factory for thiamine production |
Also Published As
| Publication number | Publication date |
|---|---|
| JP2020527359A (ja) | 2020-09-10 |
| WO2019012058A1 (en) | 2019-01-17 |
| AU2018300754B2 (en) | 2023-02-23 |
| EP3652198A1 (en) | 2020-05-20 |
| CN110869384A (zh) | 2020-03-06 |
| EP3652198B1 (en) | 2021-10-06 |
| KR102760107B1 (ko) | 2025-01-31 |
| JP2023093532A (ja) | 2023-07-04 |
| BR112020000548A2 (pt) | 2020-07-21 |
| US11851461B2 (en) | 2023-12-26 |
| CA3069650A1 (en) | 2019-01-17 |
| JP7617658B2 (ja) | 2025-01-20 |
| US20230192778A1 (en) | 2023-06-22 |
| AU2018300754A1 (en) | 2020-01-16 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| KR102760107B1 (ko) | 개선된 철-황 클러스터 전달을 갖는 세포 공장 | |
| Liu et al. | CRISPR-assisted rational flux-tuning and arrayed CRISPRi screening of an L-proline exporter for L-proline hyperproduction | |
| CN101688190B (zh) | 组合物及甲硫氨酸的生产方法 | |
| US20230080311A1 (en) | Method of improving methyltransferase activity | |
| US10696992B2 (en) | Genetically modified bacterial cell factory for thiamine production | |
| KR20120114325A (ko) | 메티오닌 생산을 위한 균주 및 방법 | |
| Wenk et al. | Evolution-assisted engineering of E. coli enables growth on formic acid at ambient CO2 via the Serine Threonine Cycle | |
| Wenk et al. | An “energy‐auxotroph” Escherichia coli provides an in vivo platform for assessing NADH regeneration systems | |
| Flynn et al. | Decreased coenzyme A levels in ridA mutant strains of S almonella enterica result from inactivated serine hydroxymethyltransferase | |
| Hermes et al. | The role of the Saccharomyces cerevisiae lipoate protein ligase homologue, Lip3, in lipoic acid synthesis | |
| Christensen et al. | A novel amidotransferase required for lipoic acid cofactor assembly in Bacillus subtilis | |
| US20220127311A1 (en) | Cell factories for improved production of compounds and proteins dependent on iron sulfur clusters | |
| US20190071680A1 (en) | Microbial production of nicotinic acid riboside | |
| Ernst et al. | L‐2, 3‐diaminopropionate generates diverse metabolic stresses in Salmonella enterica | |
| Jojima et al. | Identification of a HAD superfamily phosphatase, HdpA, involved in 1, 3-dihydroxyacetone production during sugar catabolism in Corynebacterium glutamicum | |
| Buss et al. | Clustering of isochorismate synthase genes menF and entC and channeling of isochorismate in Escherichia coli | |
| Chakauya et al. | Pantothenate biosynthesis in higher plants: advances and challenges | |
| Rintala et al. | The ORF YNL274c (GOR1) codes for glyoxylate reductase in Saccharomyces cerevisiae | |
| US20240368659A1 (en) | Microbial cell factories producing vitamin b compounds | |
| EP4662301A1 (en) | Engineered obligate fermenting bacteria for the fermentative production of d-lactate and isobutanol | |
| CN117897476A (zh) | 生产维生素b化合物的微生物细胞工厂 | |
| JP2008200046A (ja) | トレハロースを含まないアミノ酸の製造方法 | |
| Wei et al. | Discovery and biochemical characterization of UDP-glucose dehydrogenase from Granulibacter bethesdensis | |
| Gómez-Coronado et al. | Implementation of the β-h drox aspartate c cle increases gro th performance of Pseudomonas putida on the PET monomer eth lene gl col | |
| Keasling et al. | Leveraging propionate-induced growth inhibition in Corynebacterium glutamicum to evolve improved methylmalonyl-CoA-dependent polyketide production |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PA0105 | International application |
St.27 status event code: A-0-1-A10-A15-nap-PA0105 |
|
| R18-X000 | Changes to party contact information recorded |
St.27 status event code: A-3-3-R10-R18-oth-X000 |
|
| P11-X000 | Amendment of application requested |
St.27 status event code: A-2-2-P10-P11-nap-X000 |
|
| P13-X000 | Application amended |
St.27 status event code: A-2-2-P10-P13-nap-X000 |
|
| PG1501 | Laying open of application |
St.27 status event code: A-1-1-Q10-Q12-nap-PG1501 |
|
| P11-X000 | Amendment of application requested |
St.27 status event code: A-2-2-P10-P11-nap-X000 |
|
| P13-X000 | Application amended |
St.27 status event code: A-2-2-P10-P13-nap-X000 |
|
| PA0201 | Request for examination |
St.27 status event code: A-1-2-D10-D11-exm-PA0201 |
|
| E902 | Notification of reason for refusal | ||
| PE0902 | Notice of grounds for rejection |
St.27 status event code: A-1-2-D10-D21-exm-PE0902 |
|
| P11-X000 | Amendment of application requested |
St.27 status event code: A-2-2-P10-P11-nap-X000 |
|
| P13-X000 | Application amended |
St.27 status event code: A-2-2-P10-P13-nap-X000 |
|
| E601 | Decision to refuse application | ||
| PE0601 | Decision on rejection of patent |
St.27 status event code: N-2-6-B10-B15-exm-PE0601 |
|
| P11-X000 | Amendment of application requested |
St.27 status event code: A-2-2-P10-P11-nap-X000 |
|
| P13-X000 | Application amended |
St.27 status event code: A-2-2-P10-P13-nap-X000 |
|
| PX0901 | Re-examination |
St.27 status event code: A-2-3-E10-E12-rex-PX0901 |
|
| E902 | Notification of reason for refusal | ||
| PE0902 | Notice of grounds for rejection |
St.27 status event code: A-1-2-D10-D21-exm-PE0902 |
|
| P11-X000 | Amendment of application requested |
St.27 status event code: A-2-2-P10-P11-nap-X000 |
|
| P13-X000 | Application amended |
St.27 status event code: A-2-2-P10-P13-nap-X000 |
|
| PX0701 | Decision of registration after re-examination |
St.27 status event code: A-3-4-F10-F13-rex-PX0701 |
|
| X701 | Decision to grant (after re-examination) | ||
| GRNT | Written decision to grant | ||
| PR0701 | Registration of establishment |
St.27 status event code: A-2-4-F10-F11-exm-PR0701 |
|
| PR1002 | Payment of registration fee |
St.27 status event code: A-2-2-U10-U12-oth-PR1002 Fee payment year number: 1 |
|
| PG1601 | Publication of registration |
St.27 status event code: A-4-4-Q10-Q13-nap-PG1601 |