WO2020162452A1 - CARライブラリおよびscFvの製造方法 - Google Patents
CARライブラリおよびscFvの製造方法 Download PDFInfo
- Publication number
- WO2020162452A1 WO2020162452A1 PCT/JP2020/004123 JP2020004123W WO2020162452A1 WO 2020162452 A1 WO2020162452 A1 WO 2020162452A1 JP 2020004123 W JP2020004123 W JP 2020004123W WO 2020162452 A1 WO2020162452 A1 WO 2020162452A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- amino acid
- acid sequence
- variable region
- chain variable
- seq
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/10—Processes for the isolation, preparation or purification of DNA or RNA
- C12N15/1034—Isolating an individual clone by screening libraries
- C12N15/1068—Template (nucleic acid) mediated chemical library synthesis, e.g. chemical and enzymatical DNA-templated organic molecule synthesis, libraries prepared by non ribosomal polypeptide synthesis [NRPS], DNA/RNA-polymerase mediated polypeptide synthesis
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K40/00—Cellular immunotherapy
- A61K40/10—Cellular immunotherapy characterised by the cell type used
- A61K40/11—T-cells, e.g. tumour infiltrating lymphocytes [TIL] or regulatory T [Treg] cells; Lymphokine-activated killer [LAK] cells
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K40/00—Cellular immunotherapy
- A61K40/30—Cellular immunotherapy characterised by the recombinant expression of specific molecules in the cells of the immune system
- A61K40/31—Chimeric antigen receptors [CAR]
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K40/00—Cellular immunotherapy
- A61K40/40—Cellular immunotherapy characterised by antigens that are targeted or presented by cells of the immune system
- A61K40/41—Vertebrate antigens
- A61K40/42—Cancer antigens
- A61K40/4202—Receptors, cell surface antigens or cell surface determinants
- A61K40/421—Immunoglobulin superfamily
- A61K40/4211—CD19 or B4
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K40/00—Cellular immunotherapy
- A61K40/40—Cellular immunotherapy characterised by antigens that are targeted or presented by cells of the immune system
- A61K40/41—Vertebrate antigens
- A61K40/42—Cancer antigens
- A61K40/4267—Cancer testis antigens, e.g. SSX, BAGE, GAGE or SAGE
- A61K40/4269—NY-ESO
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/705—Receptors; Cell surface antigens; Cell surface determinants
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/705—Receptors; Cell surface antigens; Cell surface determinants
- C07K14/70503—Immunoglobulin superfamily
- C07K14/7051—T-cell receptor (TcR)-CD3 complex
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/705—Receptors; Cell surface antigens; Cell surface determinants
- C07K14/70503—Immunoglobulin superfamily
- C07K14/70521—CD28, CD152
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/005—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies constructed by phage libraries
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2803—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/30—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants from tumour cells
- C07K16/3069—Reproductive system, e.g. ovaria, uterus, testes, prostate
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/62—DNA sequences coding for fusion proteins
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N5/00—Undifferentiated human, animal or plant cells, e.g. cell lines; Tissues; Cultivation or maintenance thereof; Culture media therefor
- C12N5/06—Animal cells or tissues; Human cells or tissues
- C12N5/0602—Vertebrate cells
- C12N5/0634—Cells from the blood or the immune system
- C12N5/0636—T lymphocytes
-
- C—CHEMISTRY; METALLURGY
- C40—COMBINATORIAL TECHNOLOGY
- C40B—COMBINATORIAL CHEMISTRY; LIBRARIES, e.g. CHEMICAL LIBRARIES
- C40B40/00—Libraries per se, e.g. arrays, mixtures
- C40B40/04—Libraries containing only organic compounds
- C40B40/06—Libraries containing nucleotides or polynucleotides, or derivatives thereof
- C40B40/08—Libraries containing RNA or DNA which encodes proteins, e.g. gene libraries
-
- C—CHEMISTRY; METALLURGY
- C40—COMBINATORIAL TECHNOLOGY
- C40B—COMBINATORIAL CHEMISTRY; LIBRARIES, e.g. CHEMICAL LIBRARIES
- C40B40/00—Libraries per se, e.g. arrays, mixtures
- C40B40/04—Libraries containing only organic compounds
- C40B40/10—Libraries containing peptides or polypeptides, or derivatives thereof
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/57—Medicinal preparations containing antigens or antibodies characterised by the type of response, e.g. Th1, Th2
- A61K2039/572—Medicinal preparations containing antigens or antibodies characterised by the type of response, e.g. Th1, Th2 cytotoxic response
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K2239/00—Indexing codes associated with cellular immunotherapy of group A61K40/00
- A61K2239/31—Indexing codes associated with cellular immunotherapy of group A61K40/00 characterized by the route of administration
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K2239/00—Indexing codes associated with cellular immunotherapy of group A61K40/00
- A61K2239/38—Indexing codes associated with cellular immunotherapy of group A61K40/00 characterised by the dose, timing or administration schedule
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K2239/00—Indexing codes associated with cellular immunotherapy of group A61K40/00
- A61K2239/46—Indexing codes associated with cellular immunotherapy of group A61K40/00 characterised by the cancer treated
- A61K2239/48—Blood cells, e.g. leukemia or lymphoma
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/10—Immunoglobulins specific features characterized by their source of isolation or production
- C07K2317/14—Specific host cells or culture conditions, e.g. components, pH or temperature
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/565—Complementarity determining region [CDR]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/60—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments
- C07K2317/62—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments comprising only variable region components
- C07K2317/622—Single chain antibody (scFv)
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/73—Inducing cell death, e.g. apoptosis, necrosis or inhibition of cell proliferation
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
- C07K2319/02—Fusion polypeptide containing a localisation/targetting motif containing a signal sequence
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
- C07K2319/03—Fusion polypeptide containing a localisation/targetting motif containing a transmembrane segment
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/30—Non-immunoglobulin-derived peptide or protein having an immunoglobulin constant or Fc region, or a fragment thereof, attached thereto
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/33—Fusion polypeptide fusions for targeting to specific cell types, e.g. tissue specific targeting, targeting of a bacterial subspecies
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2510/00—Genetically modified cells
Definitions
- the present invention relates to a CAR library and a method for producing scFv.
- T cells chimeric antigen receptor-expressing T cells, CAR-T cells
- scFv single chain Fv
- Screening for scFv is performed by a method using a hybridoma using B cells isolated from a living body, or a phage display method. Specifically, the former prepares a hybridoma library using B cells isolated from a living body after antigen immunization. Then, a hybridoma producing an antibody that binds to the target antigen is selected, and the sequence of the antigen recognition site of the antibody produced by the hybridoma is determined.
- each scFv is expressed in a phage after creating a scFv library containing a polynucleotide encoding the scFv. Then, it is carried out by selecting a phage that binds to the target antigen and identifying the polynucleotide in the phage.
- the present invention provides, for example, a CAR library used for screening an scFv that can function in CAR-T cells, and a method for producing an scFv using the CAR library.
- a chimeric antigen receptor (CAR) library of the present invention comprises a nucleic acid encoding a first chimeric antigen receptor (CAR),
- the first CAR comprises a first antigen binding domain, a first transmembrane domain, and a first intracellular signaling domain
- the first antigen binding domain comprises a first single chain antibody (scFv) that screens for binding to a target antigen
- the first scFv comprises a first heavy chain variable region and a first light chain variable region,
- the first heavy chain variable region and the first light chain variable region satisfy the following Condition 1 or Condition 2;
- Condition 1 Heavy chain complementarity determining regions (CDRH) 1, CDRH2, and CDRH3 in the first heavy chain variable region are respectively CDRH1, CDRH2 of the heavy chain variable region in the antibody that binds to the target antigen or an antigen-binding fragment thereof,
- the method for producing scFv of the present invention comprises a first expression step of expressing the CAR library of the present invention in immune cells, A first contacting step of contacting the immune cells obtained in the first expression step with the target antigen, A first selection step of selecting the first scFv of CAR expressed in immune cells bound to the target antigen in the first contact step as a first candidate scFv binding to the target antigen.
- a CAR library used for screening an scFv capable of functioning in CAR-T cells and a method for producing an scFv using the same can be provided.
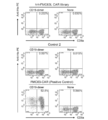
- FIG. 1 is a dot plot showing the results of flow cytometry in Example 1.
- FIG. 2 is a dot plot showing the results of flow cytometry in Example 1.
- FIG. 3 is a graph showing the ratio of A2/NY-ESO-1 157 tetramer positive cells in Example 1.
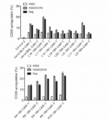
- FIG. 4 is a graph showing changes in the expression level of CD69 in Example 1.
- FIG. 5 is a graph showing changes in the expression level of CD69 in Example 1.
- FIG. 6 is a dot plot showing the results of flow cytometry in Example 1.
- FIG. 7A is a dot plot showing the results of flow cytometry in Example 1.
- FIG. 7B is a dot plot showing the results of flow cytometry in Example 1.
- FIG. 8 is a dot plot showing the results of flow cytometry in Example 2.
- FIG. 9 is a dot plot showing the results of flow cytometry in Example 2.
- FIG. 10 is a graph showing the ratio of A2/NY-ESO-1 157 tetramer positive cells in Example 2.
- FIG. 11 is a graph showing changes in the expression level of CD69 when the peptide concentration in Example 2 was 10 ⁇ g/mL.
- FIG. 12 is a graph showing changes in the expression level of CD69 in Example 2.
- FIG. 13 is a dot plot showing the results of flow cytometry in Example 3.
- FIG. 14 is a dot plot showing the results of flow cytometry in Example 4.
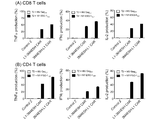
- FIG. 15 is a graph showing changes in the amount of cytokine produced in Example 4, (A) shows the results for CD8-positive T cells, and (B) shows the results for CD4-positive T cells.
- FIG. 16 is a graph showing changes in cytokine production in Example 5, (A) shows the results of T cells expressing 3M4E5LH CAR, and (B) is the T cells expressing L1-3M4E5H CAR. The result of is shown.
- FIG. 17 is a dot plot showing the results of flow cytometry in Example 6.
- FIG. 18 is a graph showing changes in cytokine production in Example 6, where (A) shows the results for CD8-positive T cells and (B) shows the results for CD4-positive T cells.
- FIG. 19 is a graph showing changes in cytokine production in Example 6, where (A) shows the results for CD8-positive T cells and (B) shows the results for CD4-positive T cells.
- FIG. 16 is a graph showing changes in cytokine production in Example 5, (A) shows the results of T cells expressing 3M4E5LH CAR, and (B) is the T cells expressing L1-3M4E5H CAR. The result of is shown.
- FIG. 20 is a graph showing changes in cytokine production in Example 7, (A) shows the results of T cells expressing 3M4E5LH CAR, and (B) shows T cells expressing 3M4E5H-L1 CAR. The result of is shown.
- FIG. 21 is a graph showing changes in cytokine production in Example 7, where (A) shows the results for CD8-positive T cells and (B) shows the results C for CD4-positive T cells.
- FIG. 22 is a dot plot showing the results of flow cytometry in Example 8.
- FIG. 23 is a graph showing changes in the amount of cytokine produced in Example 8, where (A) shows the results for CD8-positive T cells and (B) shows the results for CD4-positive T cells.
- FIG. 24 is a graph showing changes in the amount of cytokine produced in Example 8, where (A) shows the results for CD8-positive T cells and (B) shows the results for CD4-positive T cells.
- FIG. 25 is a dot plot showing the results of flow cytometry in Example 9.
- FIG. 26 is a graph showing changes in the expression level of CD69 in Example 9.
- FIG. 27 is a dot plot showing the results of flow cytometry in Example 10.
- FIG. 28 is a dot plot showing the results of flow cytometry in Example 10.
- FIG. 29 is a dot plot showing the results of flow cytometry in Example 10.
- FIG. 30 is a graph showing changes in the expression level of CD69 in Example 10.
- FIG. 31 is a graph showing the relative proportion of CD19 dimer positive cells in Example 10.
- FIG. 32 is a graph showing the relative value of the CD69 expression level when the target cell amount was changed in Example 10.
- FIG. 33 is a graph showing the ratio of cytokine-producing cells in Example 10.
- FIG. 34 is a diagram regarding the antitumor activity in the living body in Example 11.
- FIG. 35 is a graph showing the relationship between the index value of structural affinity and the index value of functional affinity in Example 12.
- FIG. 36 is a graph showing cytotoxicity in Example 13.
- FIG. 37 is a diagram regarding the overall survival rate in the living body in Example 14.
- FIG. 38 is a diagram regarding the overall survival rate in the living body in Example 14.
- FIG. 39 is a graph showing the growth ratio of cells in Example 15.
- FIG. 40 is a graph showing the ratio of cytokine-producing cells in Example 15.
- FIG. 40 is a graph showing the ratio of cytokine-producing cells in Example 15.
- FIG. 41 is a graph showing the cytotoxic activity in Example 15.
- FIG. 42 is a dot plot showing the distribution of CD45RA/CD62L-positive cells in CD8-positive CAR-T cells and CD4-positive CAR-T cells before and after co-culture in Example 15.
- 43 is a division ratio of each CAR-T cell in Example 15, a ratio of CD45RA-positive CD62L-positive CCR7-positive cells before and after co-culture, a ratio of CD45RA-negative CD62L-positive CCR7-positive, and CD45RA-positive CD62L-negative PD1-positive cell. It is a graph which shows and.
- FIG. 44 is a diagram for explaining the mechanism by which the screening method of the present invention in Example 15 can obtain scFv having an excellent antitumor effect in vivo .
- the CAR library of the present invention contains a nucleic acid encoding a first chimeric antigen receptor (CAR), and the first CAR comprises a first antigen-binding domain and a first transmembrane domain. And a first intracellular signaling domain, wherein the first antigen binding domain comprises a first single chain antibody (scFv) screening for binding to a target antigen, the first scFv being ,
- the first heavy chain variable region and the first light chain variable region, and the first heavy chain variable region and the first light chain variable region satisfy the following Condition 1 or Condition 2.
- the CAR library of the present invention is characterized by satisfying the following Condition 1 or Condition 2, and other configurations and conditions are not particularly limited.
- the heavy chain complementarity determining regions (CDRH) 1, CDRH2, and CDRH3 in the first heavy chain variable region are each an antibody or an antigen-binding fragment thereof that binds to the target antigen (hereinafter, also referred to as “antibody or the like”).
- Including CDRH1, CDRH2, and CDRH3 of the heavy chain variable region in Light chain complementarity determining regions (CDRL) 1, CDRL2, and CDRL3 in the first light chain variable region each include CDRL1, CDRL2, and CDRL3 in the light chain variable region of the first B cell receptor;
- the heavy chain complementarity determining regions (CDRH) 1, CDRH2, and CDRH3 in the first heavy chain variable region each include CDRH1, CDRH2, and CDRH3 in the heavy chain variable region of the first B cell receptor,
- Light chain complementarity determining regions (CDRL) 1, CDRL2, and CDRL3 in the first light chain variable region are respectively CDRL1, CDRL2 of the light chain variable region in the antibody or antigen-binding fragment thereof that binds to the target antigen, And CDRL3.
- the first CAR comprises the first antigen-binding domain, the first transmembrane domain, and the first intracellular signaling domain,
- the chain variable region and the first light chain variable region are characterized by satisfying the condition 1 or the condition 2, and other configurations and conditions are not particularly limited.
- the first CAR is a protein containing the first antigen-binding domain, the first transmembrane domain, and the first intracellular signal transduction domain.
- the first CAR has, for example, a structure similar to CAR including an extracellular domain that binds to an antigen, a transmembrane domain, and an intracellular signaling domain, and the extracellular domain that binds to the antigen is the target antigen.
- the first antigen binding domain that is screened for binding to is modified.
- the first CAR thus has a structure similar to that of the CAR.
- the first CAR that binds to the target antigen can induce a signal in the expressing cell via the first intracellular signaling domain, for example. Then, the first CAR that binds to the target antigen can activate, for example, its expressing cells (eg, proliferation, expression of activation marker, etc.). That is, according to the CAR library of the present invention, for example, the proportion of cells expressing the first CAR that binds to the target antigen in the CAR library-expressing cell group is increased by using the first screening method described below. be able to.
- the CAR library of the present invention for example, scFv binding to the target antigen can be screened with a smaller number or kinds of nucleic acids as compared with the phage display method.
- the first CAR that binds to the target antigen contains an scFv that functions in CAR-T cells because it can activate the cells expressing it in the first screening method described below, for example. I can say. Therefore, according to the CAR library of the present invention, for example, an scFv that can function in CAR-T cells can be screened.
- the first antigen-binding domain contains a first single-chain antibody (scFv) that screens for binding to the target antigen.
- the first scFv is, for example, a single-chain polypeptide derived from an antibody that binds to an antigen, and has a structure similar to scFv that retains the ability to bind to the antigen.
- the scFv is a polypeptide in which a variable region fragment (Fv) of a heavy chain (H chain) and a light chain (L chain) of an antibody that binds to the antigen is linked.
- one of a heavy chain variable region and a light chain variable region is a variable region such as an antibody that binds to the target antigen, and the other is B cell receptor, as described below.
- the CAR library of the present invention for example, compared with the phage display method in which the heavy chain variable region and the light chain variable region that bind to the target antigen are screened at a time, a smaller number or kind of nucleic acids can be used. , ScFv binding to the target antigen can be screened.
- the nucleic acid encoding the first CAR means, for example, a nucleic acid (polynucleotide) encoding the amino acid sequence of the first CAR.
- the nucleic acid may be composed of DNA, RNA, or both.
- the nucleic acid may be composed of a natural nucleic acid or a modified nucleic acid obtained by modifying the natural nucleic acid, and ENA (2'-O,4'-C-Ethylene-bridged Nucleic Acids) , LNA (Locked nucleic acid), PNA (Peptide Nucleic Acid), morpholino, and other artificial nucleic acids, or may be composed of two or more of these.
- the “domain” means, for example, in a polypeptide, a three-dimensionally structured or functionally integrated region.
- the polypeptide include peptides having a length of 10 amino acids or more.
- the target antigen is not particularly limited and can be any antigen.
- the target antigens include tumor antigens, virus antigens, bacterial antigens, parasite antigens, antigens associated with autoimmune diseases, sugar chain antigens and the like.
- the tumor antigen means, for example, a biomolecule having antigenicity such as an antigen newly expressed by a canceration of cells and an antigen whose expression level is increased.
- the tumor antigen may be, for example, a tumor-specific antigen or a tumor-associated antigen.
- the tumor antigen is, for example, 5T4, ⁇ 5 ⁇ 1-integrin, activated integrin ⁇ 7, 707-AP, ⁇ -fetoprotein (AFP), lectin-reactive AFP, ART-4, AURKA (AURORA A), B7H4, BAGE, ⁇ -.
- the virus antigens include, for example, adenoviruses such as adenovirus (adenovirus) ( Adenoviridae ); coronaviruses such as coronavirus ( Coronaviridae ); Ebolaviruses ( filoviridae ); Flaviviridae such as hepatitis C virus (HCV: hepatitis C virus), dengue virus (Dengue virus), Japanese encephalitis virus (Japanese encephalitis virus), West Nile virus (west Nile virus), yellow fever virus (yellow fever virus) ( Flaviviridae); B hepatitis virus (HVB: hepatitis B virus) hepadnaviridae such as (Hepadnaviridae); herpes simplex virus type 1 (HSV-1: herpes simplex virus -1), herpes simplex virus type 2 (HSV-2 : Herpes simplex virus-2), varicella zoster virus (VZV), human cytomegal
- the bacterial antigen can be, for example, tetanus (Clostridium tetani) Clostridium genus, etc. (Clostridium); E. (Escherichia coli) Esukerikia genus such as (Escherichia); Helicobacter pylori (Helicobacter pyloris) Helicobacter, such as (Helicobacter); Legionella pneumophila Firia (Legionella pneumophila) Legionella, such as (Legionella); Listeria monocytogenes (Listeria monocytogenes) Listeria such as (Listeria); Mycobacterium tuberculosis (Mycobacterium tuberculosis), Raikin (Mycobacterium leprae), Mycobacterium avium (Mycobacterium avium), Mycobacterium intracellulare et (Mycobacterium intracellulare), Mycobacterium kansasii (Mycobacterium kansasii), Mycobacterium
- Staphylococcus pneumococcal (Streptococcus pneumoniae), Streptococcus, such as Streptococcus pyogenes (Streptococcus pyogenes) (Streptococcus); Antigens derived from bacteria such as
- the parasite antigens include, for example, liver fluke ( Clonorchis sinensis ), Schistosoma japonicum ( Schistosoma japonicum ), Coleoptera ( Ascaris lumbricoides ), human pinworm ( Enterobius vermicularis ), Cysticercus cellulosae , and broad-segment cleft head.
- Antigens derived from parasites such as insects ( Diphyllobothrium latum ), Echinococcus ( Echinococcus ), Entamoeba histolytica , and Plasmodium are also known.
- the first scFv contains a first heavy chain variable region and a first light chain variable region.
- the first heavy chain variable region and the first light chain variable region have the same structure as the heavy chain variable region and the light chain variable region in the antibody molecule.
- the heavy chain variable region and the light chain variable region of an antibody molecule each have three complementarity determining regions (CDRs).
- CDR is also called a hypervariable domain. Even in the variable regions of the heavy chain and the light chain, CDR is a region in which the primary structure has a high variability, and the CDR is usually separated at three positions on the primary structure.
- the three CDRs in the heavy chain variable region are determined from the amino terminal (N-terminal) side in the amino acid sequence of the heavy chain variable region, namely, heavy chain CDR1 (CDRH1), heavy chain CDR2 (CDRH2), and heavy chain CDR3.
- the three CDRs in the light chain variable region are represented by (CDRH3), and the light chain CDR1 (CDRL1), the light chain CDR2 (CDRL2), and the light chain CDR3 (CDRL3 ). These sites are conformationally close to each other and determine their specificity for binding antigen.
- the first heavy chain variable region has CDRH1, CDRH2, and CDRH3.
- the first light chain variable region has CDRL1, CDRL2, and CDRL3. Further, the first heavy chain variable region and the first light chain variable region satisfy the above Condition 1 or Condition 2.
- CDRH1, CDRH2, and CDRH3 in the first heavy chain variable region use an antibody or the like that binds to the target antigen, and for CDRL1, CDRL2, and CDRL3 in the first light chain variable region, Screening for binding to the target antigen.
- Specific examples of the antibody or the antigen-binding fragment thereof can be referred to, for example, the description of the antibody and the antigen-binding fragment in the antibody and the like of the present invention described below.
- the antibody or the like that binds to the target antigen for example, a known antibody or the like can be used depending on the target antigen.
- a known antibody or the like can be used depending on the target antigen.
- 3M4E5 antibody can be used as an antibody against the complex of HLA-A*02:01 and NY-ESO-1 157-165 .
- an antibody against human CD19 for example, FMC63 antibody can be used as an antibody against human CD19.
- the antibody or the like that binds to the target antigen may be, for example, scFv obtained by the first screening method of the present invention described below, or an antibody obtained by immunizing the target antigen.
- CDRH1, CDRH2, and CDRH3 in the first heavy chain variable region may be, for example, a polypeptide consisting of the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the target antigen, respectively, It may be a polypeptide containing the amino acid sequences of CDRH1, CDRH2, and CDRH3 such as an antibody that binds to.
- the region other than CDRH1, CDRH2, and CDRH3, that is, the framework region (FR) may include, for example, the FR of the heavy chain variable region in the antibody or the like that binds to the target antigen. Good.
- the FRs are usually separated in four places on the primary structure.
- the four FRs in the heavy chain variable region are labeled from the N-terminal side in the amino acid sequence of the heavy chain variable region to heavy chain FR1 (FRH1), heavy chain FR2 (FRH2), heavy chain FR3 (FRH3). , And heavy chain FR4 (FRH4).
- the CDRHs and the FRHs are arranged in the order of FRH1, CDRH1, FRH2, CDRH2, FRH3, CDRH3, and FRH4 from the N-terminal side in the amino acid sequence of the heavy chain variable region.
- FRH1, FRH2, FRH3, and FRH4 in the first heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRH1, FRH2, FRH3, and FRH4, such as an antibody that binds to the target antigen. It may be a polypeptide containing the amino acid sequences of FRH1, FRH2, FRH3, and FRH4, such as an antibody that binds to the target antigen.
- the “antibody or the like that binds to the target antigen” in the description of each CDRH and the “antibody or the like that binds to the target antigen” in the description of each FRH are preferably the same antibody or the like.
- the first heavy chain variable region may include, for example, a heavy chain variable region in an antibody or the like that binds to the target antigen.
- the first heavy chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the heavy chain variable region in the antibody or the like that binds to the target antigen, or the heavy chain in the antibody or the like that binds to the target antigen. It may be a polypeptide containing the amino acid sequence of the variable region.
- the first light chain variable region is encoded by, for example, a VJ gene fragment formed by reconstructing a VJ gene fragment from a V gene fragment and a J gene fragment. Therefore, the first B cell receptor is, for example, a B cell receptor containing a polypeptide encoded by an artificial VJ gene fragment designed by artificially combining a V gene fragment and a J gene fragment. Good. In vivo B cells, for example, express the light chain variable region after VJ gene reconstruction. Therefore, the first B cell receptor may be, for example, an isolated B cell-derived B cell receptor. In this case, the first B cell receptor may be, for example, a human-derived B cell B cell receptor, and preferably a human peripheral blood B cell-derived B cell receptor. In the CAR library of the present invention, the first B cell receptor is preferably the light chain variable region of an isolated B cell-derived B cell receptor, because a CAR library can be prepared more easily.
- the CDRL1, CDRL2, and CDRL3 in the first light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRL1, CDRL2, and CDRL3 of the first B cell receptor, respectively. It may be a polypeptide comprising the amino acid sequences of CDRH1, CDRL2 and CDRL3 of the B cell receptor of.
- the FR may include, for example, the FR of the light chain variable region in the first B cell receptor.
- the FRs are usually separated in four places on the primary structure.
- four FRs in the light chain variable region are labeled from the N-terminal side in the amino acid sequence of the light chain variable region to a light chain FR1 (FRL1), a light chain FR2 (FRL2), a light chain FR3 (FRL3). , And light chain FR4 (FRL4).
- the CDRLs and the FRLs are arranged, for example, in the order of FRL1, CDRL1, FRL2, CDRL2, FRL3, CDRL3, and FRL4 from the N-terminal side in the amino acid sequence of the light chain variable region.
- FRL1, FRL2, FRL3, and FRL4 in the first light chain variable region may be, for example, a polypeptide consisting of the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the first B cell receptor, respectively.
- the polypeptide may include the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the first B cell receptor.
- the "first B cell receptor" in the description of each CDRL and the "first B cell receptor" in the description of each FRL are preferably the same B cell receptor.
- the first light chain variable region may include, for example, the light chain variable region in the first B cell receptor.
- the first light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region in the first B cell receptor, or the light chain in the first B cell receptor. It may be a polypeptide containing the amino acid sequence of the variable region.
- the CDRL1, CDRL2, and CDRL3 in the first light chain variable region are derived from an antibody or the like that binds to the target antigen, and CDRH1, CDRH2 in the first heavy chain variable region, And CDRH3 are screened for binding to the target antigen.
- the first heavy chain variable region is encoded by, for example, a VDJ gene fragment formed by reconstructing the VDJ gene fragment, V gene fragment, and J gene fragment. Therefore, the first B cell receptor is, for example, a B cell containing a polypeptide encoded by an artificial VDJ gene fragment designed by artificially combining a V gene fragment, a D gene fragment, and a J gene fragment. It may be a receptor. In vivo B cells, for example, express the heavy chain variable region after VDJ gene reconstruction. Therefore, the first B cell receptor may be, for example, an isolated B cell-derived B cell receptor.
- the first B cell receptor may be, for example, a human-derived B cell B cell receptor, and preferably a human peripheral blood B cell-derived B cell receptor.
- the first B cell receptor is preferably an isolated B cell-derived B cell receptor heavy chain variable region because a CAR library can be prepared more easily.
- CDRH1, CDRH2, and CDRH3 in the first heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRH1, CDRH2, and CDRH3 of the first B cell receptor, respectively. It may be a polypeptide comprising the amino acid sequences of CDRH1, CDRH2 and CDRH3 of the B cell receptor of.
- FRH1, FRH2, FRH3 and FRH4 in the first heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRH1, FRH2, FRH3 and FRH4 of the first B cell receptor, respectively.
- the polypeptide may include the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 of the first B cell receptor.
- the “first B cell receptor” in the description of each CDRH and the “first B cell receptor” in the description of each FRH are preferably the same B cell receptor.
- the first heavy chain variable region may include, for example, the heavy chain variable region in the first B cell receptor.
- the first heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequence of the heavy chain variable region in the first B cell receptor, or the heavy chain in the first B cell receptor. It may be a polypeptide containing the amino acid sequence of the variable region.
- CDRL1, CDRL2, and CDRL3 in the first light chain variable region may be, for example, a polypeptide consisting of the amino acid sequences of CDRL1, CDRL2, and CDRL3, such as an antibody that binds to the target antigen, or the target antigen, respectively. It may be a polypeptide containing the amino acid sequences of CDRL1, CDRL2, and CDRL3 such as an antibody that binds to.
- the FR may include, for example, the FR of the light chain variable region in the antibody or the like that binds to the target antigen.
- FRL1, FRL2, FRL3, and FRL4 in the first light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRL1, FRL2, FRL3, and FRL4, such as an antibody that binds to the target antigen. It may be a polypeptide containing the amino acid sequences of FRL1, FRL2, FRL3, and FRL4, such as an antibody that binds to the target antigen.
- the “antibody or the like that binds to the target antigen” in the description of each CDRL and the “antibody or the like that binds to the target antigen” in the description of each FRL are preferably the same antibody or the like.
- the first light chain variable region may include, for example, a light chain variable region in an antibody or the like that binds to the target antigen.
- the first light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region in the antibody or the like that binds to the target antigen, or the light chain in the antibody or the like that binds to the target antigen. It may be a polypeptide containing the amino acid sequence of the variable region.
- the first heavy chain variable region and the first light chain variable region are linked by, for example, a linker peptide (Fv linker peptide).
- the Fv linker peptide preferably does not, for example, inhibit the binding of the first scFv to the target antigen.
- the Fv linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the Fv linker peptide is composed of amino acids such as glycine and serine, and a specific example thereof is (GGGGS) n .
- the n is, for example, an integer of 1 to 6.
- the amino acid sequence of the Fv linker peptide includes, for example, a polypeptide having the amino acid sequence of SEQ ID NO: 1 or 2 below.
- Fv linker peptide 1 (SEQ ID NO: 1) GSTSGSGKPGSGEGSTKG
- Fv linker peptide 2 (SEQ ID NO: 2) GGGGSGGGGSGGGGS
- Nucleotide sequence encoding Fv linker peptide 1 (SEQ ID NO: 3) 5'-GGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGC-3'
- Nucleotide sequence encoding Fv linker peptide 2 (SEQ ID NO: 4) 5'-GGTGGAGGAGGCTCAGGAGGAGGTGGCTCTGGTGGTGGAGGCTCG-3'
- the first antigen-binding domain includes a first scFv, but the first binding domain may have a structure other than scFv that includes the heavy chain variable region and the light chain variable region.
- the first binding domain may be Fab, Fab′, F(ab′) 2 , variable region fragment (Fv), disulfide bond Fv, or the like.
- the first transmembrane domain is a domain that occupies a region that penetrates the cell membrane when, for example, the first CAR is expressed in cells.
- the first transmembrane domain may be, for example, a transmembrane domain of a transmembrane protein or an artificially designed artificial transmembrane domain.
- the transmembrane protein is, for example, ⁇ , ⁇ chain, CD3 ⁇ chain, CD28, CD3 ⁇ , CD45, CD4, CD5, CD8 (CD8 ⁇ or CD8 ⁇ ), CD9, CD16, CD22, CD27, of the T cell receptor.
- the variant having the same function is, for example, a polypeptide having an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added, and functions as a transmembrane domain.
- the “one or several” is, for example, 1 to 15, 1 to 10, 1 to 5, 1 or 2.
- the artificial transmembrane domain is a polypeptide mainly composed of hydrophobic amino acids such as leucine, isoleucine and valine.
- the artificial transmembrane domain may have, for example, tripeptides of phenylalanine, tryptophan, and valine at each end.
- the first transmembrane domain When the first transmembrane domain is a transmembrane domain of the transmembrane protein, the first transmembrane domain may be, for example, an N-terminal side or a C-terminal side continuous with the transmembrane domain of the transmembrane protein. It may contain one or several amino acids.
- the above-mentioned exemplification can be applied to the above-mentioned “one or several”.
- the first transmembrane domain is, for example, the transmembrane domain of CD28.
- the transmembrane domain of CD28 includes, for example, the following amino acid sequence from the 153rd position to the 179th position (SEQ ID NO: 5) in the amino acid sequence registered under NCBI accession number NP_006130.
- the NCBI accession number is a number assigned to the full-length amino acid sequence of the protein precursor (hereinafter the same).
- a transmembrane unit including an extracellular region on the N-terminal side and an intracellular region on the C-terminal side in addition to the transmembrane domain of CD28 may be used.
- CD28 transmembrane domain (SEQ ID NO: 5) FWVLVVVGGVLACYSLLVTVAFIIFWV
- Nucleotide sequence encoding the transmembrane domain of CD28 (SEQ ID NO: 6) 5'-TTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTG-3'
- CD28 transmembrane unit (SEQ ID NO: 7) IEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRS
- Nucleotide sequence encoding the transmembrane unit of CD28 (SEQ ID NO: 8) 5'-ATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCT-3 '
- the first intracellular signal transduction domain is, for example, a domain capable of transducing a signal into cells when the first antigen-binding domain in the same protein binds to the target antigen.
- the first signal transduction domain include cytoplasmic amino acid sequences of T cell receptor (TCR) complex and costimulatory molecule, and mutants having functions equivalent to these amino acid sequences.
- TCR T cell receptor
- the variant having the same function is, for example, a polypeptide consisting of an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added, and which functions as an intracellular signal transduction domain.
- the “one or several” the above description can be applied, for example.
- T cells are activated (primary activation) by antigen-dependent cytoplasmic signals such as intracellular signals mediated by the TCR complex. Then, this activation is enhanced by an antigen non-specific cytoplasmic signal such as an intracellular signal mediated by a costimulatory molecule (secondary activation). Therefore, the T cell has, for example, a domain (primary activation domain) that is activated through the TCR complex and a domain (secondary activation domain) that nonspecifically activates the antigen. Therefore, the first intracellular signaling domain has, for example, at least one of the primary activation domain and the secondary activation domain, and preferably has both.
- the primary activation domain can regulate primary activation by the TCR complex, for example.
- Examples of the primary activation domain that induces the primary activation include an intracellular signal transduction domain containing an immunoreceptor tyrosine-based activation motif (ITAM).
- examples of the primary activation domain that acts suppressively on the primary activation include an intracellular signal transduction domain containing an immunoreceptor tyrosine-based inhibitory motif (ITIM).
- the intracellular signal transduction domain including ITAM is, for example, an intracellular signal transduction domain such as CD3 ⁇ , FcR ⁇ , FcR ⁇ , CD3 ⁇ , CD3 ⁇ , CD5, CD22, CD79a, CD79b, CD66d or a mutation having a function equivalent to these. I can raise my body.
- the variant having the same function comprises, for example, an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added, and functions as an intracellular signal transduction domain including ITAM. Is. As for the “one or several”, the above description can be applied, for example.
- the intracellular signal transduction domain containing ITAM of CD3 ⁇ includes, for example, the amino acid sequence from the 52nd to the 164th amino acid sequence in the amino acid sequence registered under NCBI accession number NP_932170.1, or the amino acid sequence of SEQ ID NO: 9 below.
- the intracellular signal transduction domain including ITAM of Fc ⁇ RI ⁇ includes, for example, the 45th to 86th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_004097.1.
- the intracellular signal transduction domain containing ITAM of Fc ⁇ RI ⁇ includes, for example, amino acid sequences 201 to 244 in the amino acid sequence registered under NCBI accession number NP_000130.1.
- the intracellular signal transduction domain containing ITAM of CD3 ⁇ includes, for example, the 139th to 182nd amino acid sequences in the amino acid sequence registered under NCBI accession number NP_000064.1.
- the intracellular signal transduction domain containing ITAM of CD3 ⁇ includes, for example, the amino acid sequences 128 to 171 in the amino acid sequence registered under NCBI accession number NP_000723.1.
- the intracellular signal transduction domain containing ITAM of CD3 ⁇ includes, for example, the amino acid sequence at the 153rd to 207th positions in the amino acid sequence registered under NCBI accession number NP_000724.1.
- the intracellular signal transduction domain including ITAM of CD5 includes, for example, the 402nd to 495th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_055022.2.
- the intracellular signal transduction domain containing ITAM of CD22 includes, for example, the amino acid sequences at positions 707 to 847 in the amino acid sequence registered under NCBI accession number NP_001762.2.
- the intracellular signal transduction domain containing ITAM of CD79a includes, for example, the 166th to 226th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_001774.1.
- the intracellular signal transduction domain including ITAM of CD79b includes, for example, the 182nd to 229th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_000617.1.
- the intracellular signal transduction domain containing ITAM of CD66d includes, for example, the 177th to 252nd amino acid sequences in the amino acid sequence registered under NCBI accession number NP_001806.2.
- Intracellular signaling domain containing ITAM of CD3 ⁇ (SEQ ID NO: 9) RVKSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
- Nucleotide sequence encoding an intracellular signal transduction domain including ITAM of CD3 ⁇ (SEQ ID NO: 10) 5'-CGAGTGAAGAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCAGA-3 '
- the secondary activation domain is an intracellular signal transduction domain such as CD2, CD4, CD5, CD8 ⁇ , CD8 ⁇ , CD27, CD28, CD134 (OX40), CD137 (4-1BB), CD154, GITR, ICOS or the like.
- a mutant having the function of The variant having the same function is, for example, a polypeptide consisting of an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added, and which functions as an intracellular signal transduction domain.
- the intracellular signal transduction domain of CD2 includes, for example, the 236th to 351st amino acid sequences in the amino acid sequence registered under NCBI accession number NP_001758.2.
- the intracellular signal transduction domain of CD4 includes, for example, amino acid sequences 421 to 458 in the amino acid sequence registered under NCBI accession number NP_000607.1.
- the intracellular signal transduction domain of CD5 includes, for example, the 402nd to 495th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_055022.2.
- the intracellular signal transduction domain of CD8 ⁇ includes, for example, the 207th to 235th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_001759.3.
- the intracellular signal transduction domain of CD8 ⁇ includes, for example, the 196th to 210th amino acid sequences in the amino acid sequence registered under NCBI accession number AAA35664.1.
- the intracellular signal transduction domain of CD27 includes, for example, amino acid sequences 213 to 260 in the amino acid sequence registered under NCBI accession number M63928.1.
- the intracellular signal transduction domain of CD28 includes, for example, the 181 to 220th amino acid sequence (SEQ ID NO: 11) in the amino acid sequence registered under NCBI accession number NP_006130.1.
- the intracellular signal transduction domain of CD134 includes, for example, the 241st to 277th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_003318.1.
- the intracellular signal transduction domain of CD137 includes, for example, the amino acid sequence at positions 214 to 255 in the amino acid sequence registered under NCBI accession number NP — 001552.2.
- the intracellular signal transduction domain of GITR includes, for example, the 193rd to 241st amino acid sequences in the amino acid sequence registered under NCBI accession number NP_004186.1.
- the intracellular signal transduction domain of ICOS includes, for example, the 166th to 199th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_036224.1.
- Intracellular signaling domain of CD28 SEQ ID NO: 11
- Nucleotide sequence encoding the intracellular signal transduction domain of CD28 (SEQ ID NO: 12) 5'-CGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCT-3'
- the first CAR includes, for example, one or a plurality of first intracellular signaling domains.
- the first intracellular signaling domains may be the same or different.
- the first intracellular signaling domain preferably includes, for example, an intracellular signaling domain containing ITAM of CD3 ⁇ as the primary activation domain.
- the first intracellular signaling domain preferably includes, for example, at least one intracellular signaling domain selected from the group consisting of CD27, CD28 and CD137 as the secondary activation domain.
- the first intracellular signaling domain includes, for example, an intracellular signaling domain containing ITAM of CD3 ⁇ and at least one intracellular signaling domain selected from the group consisting of CD27, CD28 and CD137. including.
- the CAR library of the present invention is screened for scFvs that bind to the target antigen with a smaller number or type of nucleic acids as compared with the phage display method in the first screening method of the present invention described later, for example. it can.
- the intracellular signaling domain containing ITAM of CD3 ⁇ is the intracellular domain of CD28. It is preferably located on the C-terminal side of the signal transduction domain.
- the first intracellular signaling domain comprises an intracellular signaling domain containing ITAM of CD3 ⁇ and an intracellular signaling domain of CD28
- the first intracellular signaling domain further comprises CD134 and CD137. And at least one intracellular signal transduction domain thereof.
- the first antigen binding domain and the first transmembrane domain are directly or indirectly linked.
- the first antigen binding domain and the first transmembrane domain are preferably linked by a spacer peptide.
- the spacer peptide is composed of, for example, 1 to 300, 1 to 100, 10 to 100, and 25 to 50 amino acids.
- the spacer peptide promotes, for example, the binding between the first CAR and the target antigen, and, when the first CAR and the target antigen bind, a signal by the first intracellular signaling domain It preferably contains an amino acid that induces and/or enhances transduction (eg, cysteine, serine, threonine, etc.).
- the spacer peptide may be, for example, an immunoglobulin constant region such as CH1 region or L region, a partial region of CD4, CD8 ⁇ , CD8 ⁇ , or CD28.
- the first spacer peptide may be an artificially designed amino acid sequence.
- the partial region of CD8 ⁇ includes, for example, the hinge region of CD8 ⁇ , and specific examples thereof include the amino acid sequences at positions 118 to 178 in the amino acid sequence registered under NCBI accession number NP_001759.3.
- the partial region of CD8 ⁇ includes, for example, the 135th to 195th amino acid sequences in the amino acid sequence registered under NCBI accession number AAA35664.1.
- the partial region of CD4 includes, for example, the 315th to 396th amino acid sequences in the amino acid sequence registered under NCBI accession number NP_000607.1.
- the partial region of CD28 includes, for example, the 137th to 152nd amino acid sequences (SEQ ID NO: 13) in the amino acid sequence registered under NCBI accession number NP — 006130.1.
- the spacer peptide may be a part of each exemplified amino acid sequence.
- Nucleotide sequence encoding a part of CD28 (SEQ ID NO: 14) 5'-ATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCT-3'
- the domains may be linked by, for example, a linker peptide (domain linker peptide).
- domain linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the domain linker peptide is composed of amino acids such as glycine and serine, and (GGGGS) n is a specific example.
- the n is, for example, an integer of 1 to 6.
- Examples of the first CAR include a polypeptide having an amino acid sequence represented by the following formula (1) (SEQ ID NO: 15).
- V 1 and V 2 are the amino acid sequences of the heavy chain variable region and the light chain variable region, or the amino acid sequences of the light chain variable region and the heavy chain variable region, respectively.
- the amino acid sequence between V 1 and V 2 is the amino acid sequence of the Fv linker peptide
- the amino acid sequence on the C-terminal side of V 2 is the spacer peptide, transmembrane domain and intracellular signal transduction domain.
- the Fv linker peptide is Fv linker peptide 1, but it may be Fv linker peptide 2.
- V 1 [GSTSGSGKPGSGEGSTKG] - [V 2] - [IEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRS] - [RVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR] ⁇ (1)
- nucleic acid encoding the first CAR examples include a polynucleotide having a base sequence represented by the following formula (2) (SEQ ID NO: 16).
- N 1 and N 2 are the base sequences encoding the heavy chain variable region and the light chain variable region, or the base sequences encoding the light chain variable region and the heavy chain variable region, respectively.
- the base sequence between N 1 and N 2 is the base sequence encoding the Fv linker peptide
- amino acid sequence 3′ from N 2 is the spacer peptide, transmembrane domain and intracellular It is a nucleotide sequence encoding a signal transduction domain.
- the base sequence encoding the Fv linker peptide is the base sequence encoding the Fv linker peptide 1, but may be the base sequence encoding the Fv linker peptide 2.
- the CAR library of the present invention preferably contains, for example, a plurality of types of nucleic acids.
- the CAR library of the present invention is, for example, a mixture of plural kinds of nucleic acids.
- a part or all of the plurality of kinds of nucleic acids preferably encode different first CARs, and preferably encode different first antigen-binding domains.
- the regions other than the first antigen-binding domain in the first CAR have, for example, the same or different amino acid sequences.
- the types of the nucleic acids contained in the CAR library of the present invention are, for example, 1 ⁇ 10 5 to 1 ⁇ 10 7 types, 1 ⁇ 10 5 to 1 ⁇ 10 6 types, 1 ⁇ 10 6 to 5 ⁇ 10 6 types , Preferably about 2 ⁇ 10 6 types (eg, 1 ⁇ 10 6 to 3 ⁇ 10 6 types).
- about 1 ⁇ 10 8 kinds of nucleic acids are required for screening scFv binding to the target antigen.
- the CAR library of the present invention for example, even about 1 ⁇ 10 6 kinds of nucleic acids can be screened for scFv binding to the target antigen.
- scFv binding to the target antigen can be screened with a smaller number or kinds of nucleic acids as compared with the phage display method.
- the type of nucleic acid can also be referred to as, for example, nucleic acid heterogeneity.
- the heterogeneity can be measured, for example, by restriction enzyme mapping, a sequence such as Sanger method for each CDR and/or each FRH, and the like.
- the first CAR may have a signal peptide at the N-terminus, for example.
- the signal peptide include a signal peptide serving as a transport signal to the endoplasmic reticulum.
- the first CAR may have a tag, for example.
- the tag include a peptide tag and a protein tag.
- the tag include FLAG (registered trademark) tag, HA tag, His tag, Myc tag, V5 tag, and truncated NGFR (nerve growth factor receptor).
- the tag peptide is added to, for example, at least one of the N-terminus and the C-terminus of the first CAR.
- the tag When the tag is a truncated NGFR, the tag is arranged, for example, on the C-terminal side of the first CAR.
- the CAR library of the present invention can adjust the nucleic acid transfer efficiency so that, for example, one cell can be transferred with one type of nucleic acid encoding the first CAR.
- the first CAR and the tag may be linked by a linker peptide.
- the nucleic acid encoding the first CAR can be prepared by a conventional method, for example, based on the amino acid sequence of the first CAR.
- the nucleic acid encoding the first CAR is obtained by, for example, obtaining a base sequence encoding the amino acid sequence in a database in which the amino acid sequences of the above-mentioned domains are registered, and based on the base sequence, It can be prepared by biological methods and/or chemical synthetic methods.
- the base sequence of the nucleic acid may be codon-optimized depending on the origin of cells expressing the CAR library of the present invention.
- the nucleic acid encoding the first CAR may be introduced into an expression vector, for example.
- the expression vector can be prepared, for example, by ligating the nucleic acid encoding the first CAR to a ligation vector.
- the type of the ligation vector is not particularly limited, and examples thereof include retrovirus vectors such as oncoretrovirus vector, lentivirus vector, pseudotype vector; adenovirus vector, adeno-associated virus (AAV) vector, simian virus vector, vaccinia.
- retrovirus vectors such as oncoretrovirus vector, lentivirus vector, pseudotype vector; adenovirus vector, adeno-associated virus (AAV) vector, simian virus vector, vaccinia.
- retrovirus vectors such as oncoretrovirus vector, lentivirus vector, pseudotype vector
- adenovirus vector such as adeno-associated virus (AAV) vector
- simian virus vector vaccinia
- viral vectors such as viral vector
- the ligation vector examples include pUC, pCMV, pMX, and pELP.
- the ligation vector can be appropriately set, for example, according to the host into which the expression vector is introduced.
- the host is not particularly limited, and examples thereof include cultured cells of mammalian origin such as CHO cells, Jurkat cells, Jurkat76 cells, and immune cells.
- the immune cells include lymphocytes, granulocytes, macrophages and the like. Examples of the lymphocytes include T cells, NK cells, NKT cells, B cells and the like.
- the immune cells are, for example, cells isolated from a living body, immune cells derived from stem cells such as pluripotent stem cells, or cultured cells derived from immune cells.
- the T cells may be T cell-like cells. Examples of the T cell-like cells include cultured cells derived from T cells and the like, and specific examples thereof include Jurkat cells, Jurkat 76 cells and the like.
- the expression vector may have a regulatory sequence that regulates the expression of the nucleic acid encoding the first CAR and the expression of at least one of the first CAR encoded by the nucleic acid encoding the first CAR, for example. preferable.
- the regulatory sequence include a promoter, a terminator, an enhancer, a polyadenylation signal sequence, an origin of replication sequence (ori), and the like.
- the arrangement of the regulatory sequences in the expression vector is not particularly limited.
- the regulatory sequence is arranged so as to functionally regulate the expression of the nucleic acid encoding the first CAR and the expression of at least one of the first CAR encoded by the nucleic acid. However, it can be arranged based on a known method.
- the regulatory sequence for example, a sequence that the ligation vector has in advance may be used, the regulatory sequence may be further inserted into the ligation vector, or the ligation vector may have other regulatory sequences. It may be replaced with the regulatory sequence of.
- the expression vector may further have a selection marker coding sequence, for example.
- selection marker include drug resistance markers, fluorescent protein markers, enzyme markers, cell surface receptor markers, and the like.
- the first screening method of the present invention is obtained by the first expression step of expressing the CAR library of the present invention (hereinafter, also referred to as “first CAR library”) in immune cells, and the first expression step.
- a first contact step of contacting the target immune antigen with the target antigen, and the first scFv of CAR expressed in the immune cell bound to the target antigen in the first contact step is used as the target antigen.
- the first screening method of the present invention comprises the first expression step, the first contact step and the first selection step, wherein the CAR library of the present invention is used in the first expression step. And other configurations and conditions are not particularly limited.
- the first screening method of the present invention scFv that can function in CAR-T cells can be screened.
- an antibody or an antigen-binding fragment thereof that can bind to the target antigen can be prepared based on the amino acid sequences of the CDRs of the heavy chain variable region and the light chain variable region of scFv that bind to the target antigen. Therefore, the first screening method can also be referred to as, for example, a screening method for an antibody or an antigen-binding fragment thereof that binds to the target antigen.
- the description of the CAR library of the present invention can be applied to the first screening method of the present invention, for example.
- a new scFv can be screened. Therefore, the method for producing scFv of the present invention can also be referred to as, for example, a screening method for scFv.
- a chimeric antigen receptor (CAR) or the like that binds to a target antigen by the phage display method or the like
- a CAR or the like containing scFv is prepared.
- the effectiveness of the CAR and the like will be examined using immune cells and the like.
- the CAR library of the present invention and the immune cells are used, for example, screening of scFv that binds to the target antigen and efficacy of the scFv in immune cells The studies can be conducted simultaneously.
- the first screening method of the present invention as compared with, for example, the phage display method used to obtain an antibody that binds to a target antigen, and the like
- the scFv that can be used for CAR that effectively functions in the above can be easily screened.
- the first expression step is a step of expressing the CAR library of the present invention in immune cells. Specifically, in the first expression step, for example, the first CAR library is introduced into the immune cells. Then, in the first expression step, the immune cells are caused to express the first CAR library by, for example, culturing the immune cells into which the first CAR library has been introduced.
- the method for introducing the first CAR library is not particularly limited, and, for example, a known method for introducing a nucleic acid into cells can be used.
- the method for introducing the first CAR library includes, for example, a method using a nucleic acid introducing reagent such as liposome or cationic lipid; a method using a virus such as retrovirus or lentivirus;
- the period for culturing the immune cells is not particularly limited and is, for example, 6 hours to 30 days, 6 to 96 hours, 1 to 30 days.
- the culture temperature of the immune cells is, for example, 28 to 37°C.
- the first CAR library may be, for example, a nucleic acid encoding the first CAR or an expression vector into which the nucleic acid encoding the first CAR has been introduced.
- the immune cells are not particularly limited, and examples thereof include T cells, NK cells, NKT cells, B cells and the like.
- the immune cells include immune cells isolated from a living body, immune cells derived from stem cells such as pluripotent stem cells, or cultured cells derived from immune cells.
- the isolated immune cells may be, for example, immune cell-like cultured cells, and specific examples thereof include T cell-like cultured cells, NK cell-like cultured cells, NKT cell-like cultured cells, and B cell-like cultured cells. Examples include cells.
- the immune cells are, for example, immune cells isolated from a living body, and specific examples thereof include human peripheral blood-derived immune cells. Examples of the T cell-like cultured cells include Jurkat cells and Jurkat76 cells.
- the immune cells can be screened for scFvs that are more likely to function in CAR-T cells, for example, and in the first contacting step described below, immune cells expressing the first CAR that binds to the target antigen are selected.
- T cells or T cell-like cultured cells, NK cells, and NKT cells are preferable because they can be enriched.
- the number of the immune cells is not particularly limited, and may be, for example, 1 ⁇ 10 5 to 1 ⁇ 10 8 cells, 1 ⁇ 10 5 to 1 ⁇ 10 7 cells, 1 ⁇ 10 5 to 1 ⁇ 10 6 cells, 1 ⁇ 10 6 to 5 ⁇ 10 6 cells, 1 ⁇ 10 6 to 3 ⁇ 10 6 cells.
- the CAR library of the present invention is preferably expressed in immune cells so that, on average, one or less first CAR is expressed per immune cell.
- the first screening method of the present invention can suppress, for example, the selection of the non-specific first CAR that does not bind to the target antigen.
- the number of first CARs expressed per cell can be adjusted, for example, by changing the amount ratio of the number of cells to the vector to be introduced such as nucleic acid or virus. Specifically, in the first expression step, the number of first CARs expressed per cell can be reduced by reducing the ratio of the amount of vector to be introduced to a certain number of cells.
- the number of the first CAR expressed per cell can be increased by increasing the ratio of the amount of vector to be introduced to a certain number of cells.
- the efficiency of introducing the vector into the immune cells is not particularly limited and can be, for example, 10 to 60%, 10 to 50%, 20 to 40%, 25 to 35%.
- the immune cells (candidate immune cells) obtained in the first expression step are contacted with the target antigen.
- the contact between the candidate immune cells and the target antigen can be carried out, for example, by culturing in the presence of the candidate immune cells and the target antigen.
- the culture period is, for example, 6 hours to 30 days, 6 to 96 hours, or 1 to 30 days.
- the culture temperature is, for example, 28 to 37°C.
- the immune cells are T cells
- the candidate immune cells can also be referred to as candidate CAR-T, for example.
- the CAR expressed by the candidate immune cells may be, for example, one kind or plural kinds, but the former is preferable.
- the candidate immune cells are preferably candidate immune cells expressing CARs having different amino acid sequences, that is, a mixture of two or more candidate immune cells expressing different CARs.
- Examples of the target antigen to be brought into contact with the candidate immune cells include monomers of the target antigen, complexes of the target antigen, cells expressing the target antigen, and the like, and preferably cells expressing the target antigen. ..
- Examples of the complex of the target antigen include multimers (multimers) of the target antigen, and specific examples thereof include dimers and tetramers of the target antigen.
- the target antigen multimer can be prepared by, for example, a method of crosslinking the tagged target antigen with an antibody, a method of complexing the biotinylated target antigen with avidin, or the like.
- Examples of cells expressing the target antigen include cells that endogenously express the target antigen, cells that express the target antigen by introducing a nucleic acid encoding the target antigen, and the like.
- Examples of cells expressing the target antigen include cultured cells such as 293 cells, 293T cells, and K562 cells.
- a molecule that activates immune cells in addition to the target antigen, a so-called costimulatory molecule, may be contacted together.
- the costimulatory molecule include CD27, CD40, CD40L, CD80, CD83, CD86, OX40L, 4-1BBL, GITRL and ICOS.
- the costimulatory molecule include a monomer of the costimulatory molecule, a complex of the costimulatory molecule, a cell expressing the costimulatory molecule, and the like, and preferably a cell expressing the costimulatory molecule.
- the complex of the costimulatory molecule can be prepared, for example, in the same manner as the complex of the target antigen.
- the target antigen-expressing cells When the target antigen-expressing cells are used as the target antigen, the target antigen-expressing cells preferably also express a molecule that activates immune cells, a so-called co-stimulatory molecule.
- a molecule that activates immune cells examples include CD27, CD40, CD40L, CD80, CD83, CD86, OX40L, 4-1BBL, GITRL and ICOS.
- the target antigen-expressing cells may express one type of costimulatory molecule, or may express two or more types of costimulatory molecule.
- the target antigen-expressing cells are selected from the group consisting of CD80, CD83, CD40, and 4-1BBL, for example, because they can exhibit a function similar to that of antigen-presenting cells in a living body and can select CAR that functions in a living body. It is preferred to express at least one.
- the candidate immune cells are brought into contact with the target antigen in the first contact step. Therefore, a candidate immune cell expressing a CAR specific to the target antigen may be treated with the target antigen in the same manner as in vivo depending on the degree of affinity (binding) of CAR to the target antigen. It is possible to form immune synapse-like structures continuously or intermittently. As a result, the candidate immune cells expressing the CAR specific to the target antigen can survive, activate and/or derive from CAR depending on, for example, the degree of affinity (binding) of CAR to the target antigen. Or it can receive a growth signal.
- the candidate immune cells that have undergone the first contacting step have, for example, a degree of affinity (binding) of CAR to the target antigen. Correspondingly, it exhibits a phenotype of survival, expression of activation markers and/or increased cell number.
- candidate immune cells expressing such a phenotype are evaluated and selected. Therefore, in the first selection step, for example, CAR can be screened based on the phenotype (functionality) generated when the CAR is activated, in addition to the affinity of CAR expressed in the candidate immune cells. .. Therefore, the first screening method of the present invention can suitably screen for scFv that can be used for CAR that effectively functions in vivo or in immune cells by including the first contacting step.
- the first contact step may be performed multiple times.
- the plurality of times is, for example, 1 to 5 times, 2 to 3 times, and preferably 2 to 3 times.
- the candidate immune cells are recovered after the first first contacting step, and the second first contacting step is performed in the same manner as the first first contacting step.
- Contact step Then, collection and contact are repeated in the same manner up to a desired number of times.
- CAR expressed by a candidate immune cell that proliferates by carrying out the first contacting step a plurality of times is, for example, as described in the Examples below, the CAR expressed with respect to the cell expressing the CAR.
- the cells expressing the CAR are preferably grown in a stimulus-dependent manner on the target antigen, and the cells expressing the CAR are dependent on the target antigen. Activation such as specific cytotoxic activity can be induced. Therefore, the first screening method of the present invention can suitably screen, for example, an scFv that can be used for CAR that functions more favorably in vivo by performing the first contacting step a plurality of times.
- the candidate immune cells expressing CAR that recognizes the target antigen in the candidate immune cells are stimulated with the target antigen. Therefore, the first contacting step can also be referred to as, for example, an antigen stimulating step.
- the number of the candidate immune cells is not particularly limited, and may be, for example, 1 ⁇ 10 5 to 1 ⁇ 10 8 cells, 1 ⁇ 10 5 to 1 ⁇ 10 7 cells, 1 ⁇ 10 5 to 1 ⁇ 10 6 cells, 1 ⁇ 10 6 to 5 ⁇ 10 6 cells, 1 ⁇ 10 6 to 3 ⁇ 10 6 cells.
- the candidate immune cell (E) and the cell expressing the target antigen T Cell ratio (E:T) of 2) to 50:1, 5:1 to 40:1, 10:1 to 30:1, or about 20:1.
- the first scFv of the first CAR expressed in T cells bound to the target antigen in the first contact step is replaced with a first candidate scFv binding to the target antigen.
- the binding between the target antigen and the candidate immune cells can be evaluated directly or indirectly, for example.
- the binding between the target antigen and the candidate immune cells is indirect because, for example, when used as scFv of a chimeric antigen receptor, scFv capable of inducing activation of T cells and exertion of functionality can be obtained. You may carry out by evaluation.
- the direct evaluation method can be performed by, for example, a method for detecting binding between an antibody and an antigen such as surface plasmon resonance (SPR) or flow cytometry.
- the direct evaluation method can be performed using, for example, a labeled target antigen monomer or multimer.
- the labeled target antigen is brought into contact with the candidate immune cells.
- the candidate immune cell expresses CAR that binds to the target antigen
- the labeled target antigen and the candidate immune cell form a complex. Therefore, in the first selection step, for example, the candidate immune cells forming the complex containing the label can be evaluated as T cells bound to the target antigen.
- the candidate immune cells are mixed with a fluorescent-labeled multimer of a target antigen, and the candidate immune cells expressing a CAR specific to the target antigen are combined with the fluorescent label. To form a complex with the targeted target antigen.
- the resulting mixture is analyzed by flow cytometry, the candidate immune cells forming a complex with the fluorescently labeled target antigen, based on the signal from the fluorescent label, detected, evaluated. And single.
- the indirect evaluation method can be implemented as follows, for example.
- the T cell expressing CAR that binds to the target antigen is activated, for example, by binding to the target antigen. Therefore, in the indirect evaluation method, for example, the activation of the candidate immune cells is evaluated as an index.
- the expression of the activation marker is increased, cytokine and/or Increased production of chemokines, proliferation, etc. occur.
- the immune cells are T cells, NK cells, NKT cells or B cells, in each of the cells, for example, an increase in the expression of the following activation marker, an increase in the production amount of the following cytokine and/or chemokine, Proliferation occurs. Therefore, in the first selection step, for example, when it is determined that the candidate immune cells are activated based on any one or more indicators, the activated immune cells are treated as the target antigen. It can be evaluated as immune cells bound to.
- a reporter that increases the expression amount of mRNA or protein when the expression of the cytokine and/or chemokine is induced may be used.
- the proliferation can be evaluated, for example, by staining cells with a fluorescent cell staining dye such as Carboxyfluorescein succinimidyl ester (CFSE) and the like, and using the attenuation of fluorescence intensity as an index.
- a fluorescent cell staining dye such as Carboxyfluorescein succinimidyl ester (CFSE) and the like, and using the attenuation of fluorescence intensity as an index.
- -T cell activation marker cytokine such as CD69, CD107a-chemokine: IFN- ⁇ , IL-12, IL-2, TNF ⁇ , MIP-1 ⁇ etc.
- -NK cell activation marker cytokine such as CD69, CD107a-chemokine: INF- ⁇ , IL-12, etc.
- NKT cell activation marker CD69, CD107a, CD25, etc.
- cytokine, chemokine INF- ⁇ , IL-2, etc.
- B cell activation marker CD28, CD69,
- the first scFv of the first CAR expressed in the candidate immune cells bound to the target antigen is selected as the first candidate scFv binding to the target antigen.
- the selection of the first candidate scFv can be performed, for example, by selecting candidate immune cells that have bound to the target antigen and decoding the base sequence encoding scFv or CAR in the selected candidate immune cells.
- CDRH1, CDRH2, and CDRH3 in the heavy chain variable region and CDRL1, CDRL2, and CDRL3 in the light chain variable region in the first candidate scFv may be further specified.
- Identification of each CDR can be carried out by a known method with reference to, for example, genome information (for example, IMGT home page ⁇ http://www.imgt.org/>).
- a new scFv that binds to the target antigen can be screened.
- the first CAR in the first CAR library satisfies the condition 1
- a new light chain variable region that binds to the target antigen can be screened by the first screening method of the present invention.
- the first screening method of the present invention enables screening of a new heavy chain variable region that binds to the target antigen.
- the first screening method of the present invention is, for example, a method in which a new heavy chain variable region or light chain variable region of the first candidate scFv is added to an antigen or the like that binds to a target antigen in the first CAR library, respectively. Screening of the other region may be performed as the chain variable region or the light chain variable region.
- the first screening method of the present invention further includes, for example, a preparation step of preparing a second CAR library based on the first candidate scFv.
- the second CAR library contains, for example, a nucleic acid encoding the second CAR.
- the second CAR includes, for example, a second antigen binding domain, a second transmembrane domain, and a second intracellular signal transduction domain.
- the second antigen-binding domain also includes, for example, a second scFv that screens for binding to the target antigen.
- the nucleic acid encoding the second CAR means, for example, a nucleic acid (polynucleotide) encoding the amino acid sequence of the second CAR.
- the target antigen is the same as the target antigen in the first scFv.
- the second scFv has, for example, a structure similar to that of the first scFv.
- the second scFv includes, for example, a second heavy chain variable region and a second light chain variable region.
- the second heavy chain variable region has, for example, CDRH1, CDRH2, and CDRH3.
- the second light chain variable region has, for example, CDRL1, CDRL2, and CDRL3.
- the second heavy chain variable region and the second light chain variable region satisfy, for example, the following Condition 3 or Condition 4.
- CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region each comprise CDRH1, CDRH2, and CDRH3 in the heavy chain variable region of the second B cell receptor
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region include the light chain variable regions CDRL1, CDRL2, and CDRL3 in the first candidate scFv, respectively.
- CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region include CDRH1, CDRH2, and CDRH3 of the heavy chain variable region in the first candidate scFv, respectively
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region include CDRL1, CDRL2, and CDRL3, respectively, in the light chain variable region of the second B cell receptor.
- the condition 3 is, for example, a case where the second CAR library is prepared by using the first candidate scFv screened using the first CAR library that satisfies the condition 1.
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region use the first candidate scFv, and for CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region, Screening for binding to the target antigen.
- the second B cell receptor in the second heavy chain variable region is defined as, for example, “first B cell receptor” in the description of the first heavy chain variable region of the above Condition 2.
- the description can be incorporated by replacing "the first heavy chain variable region” with “the second heavy chain variable region” in the "cell receptor”.
- CDRH1, CDRH2 and CDRH3 in the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRH1, CDRH2 and CDRH3 of the second B cell receptor, respectively. It may be a polypeptide comprising the amino acid sequences of CDRH1, CDRH2 and CDRH3 of the B cell receptor of.
- FRH1, FRH2, FRH3 and FRH4 in the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRH1, FRH2, FRH3 and FRH4 of the second B cell receptor, respectively.
- the polypeptide may include the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 of the second B cell receptor.
- the "second B cell receptor" in the description of each CDRH and the "second B cell receptor" in the description of each FRH are preferably the same B cell receptor.
- the second heavy chain variable region may include, for example, the heavy chain variable region in the second B cell receptor.
- the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequence of the heavy chain variable region in the second B cell receptor, or the heavy chain in the second B cell receptor. It may be a polypeptide containing the amino acid sequence of the variable region.
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRL1, CDRL2, and CDRL3 of the first candidate scFv, respectively, or the first candidate. It may be a polypeptide containing the amino acid sequences of scFv CDRH1, CDRL2, and CDRL3.
- the FR may include, for example, the FR of the light chain variable region in the first candidate scFv.
- FRL1, FRL2, FRL3, and FRL4 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the first candidate scFv, respectively, It may be a polypeptide comprising the amino acid sequences of FRL1, FRL2, FRL3 and FRL4 of the first candidate scFv.
- the “first candidate scFv” in the description of each CDRL and the “first candidate scFv” in the description of each FRL are preferably the same scFv.
- the second light chain variable region may include, for example, the light chain variable region in the first candidate scFv.
- the second light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region of the first candidate scFv, or the amino acid of the light chain variable region of the first candidate scFv. It may be a polypeptide containing the sequence.
- condition 4 is a case where the second CAR library is prepared using the first candidate scFv screened using the first CAR library satisfying the condition 2, for example.
- condition 4 for example, CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region use the first candidate scFv, and for CDRL1, CDRL2, and CDRL3 in the second light chain variable region, Screening for binding to the target antigen.
- CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region may be, for example, a polypeptide consisting of the amino acid sequences of CDRH1, CDRH2, and CDRH3 of the first candidate scFv, respectively, or the first candidate. It may be a polypeptide containing the amino acid sequences of scFv CDRH1, CDRH2, and CDRH3.
- FRH1, FRH2, FRH3, and FRH4 in the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 of the first candidate scFv, respectively. It may be a polypeptide comprising the amino acid sequences of FRH1, FRH2, FRH3 and FRH4 of the first candidate scFv.
- the “first candidate scFv” in the description of each CDRH and the “first candidate scFv” in the description of each FRH are preferably the same scFv.
- the second heavy chain variable region may include, for example, the heavy chain variable region in the first candidate scFv.
- the second heavy chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the heavy chain variable region in the first candidate scFv, or the amino acid of the heavy chain variable region in the first candidate scFv. It may be a polypeptide containing the sequence.
- the second B cell receptor in the second light chain variable region is defined as, for example, “first B cell receptor” in the description of the first light chain variable region of the above Condition 1.
- the description can be incorporated by replacing “first light chain variable region” with “second light chain variable region” in “cell receptor”.
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRL1, CDRL2, and CDRL3 of the second B cell receptor, respectively. It may be a polypeptide comprising the amino acid sequences of CDRH1, CDRL2 and CDRL3 of the B cell receptor of.
- FRL1, FRL2, FRL3, and FRL4 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the second B cell receptor, respectively.
- the polypeptide may include the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the second B cell receptor. It is preferable that the “second B cell receptor” in the description of each CDRL and the “second B cell receptor” in the description of each FRL are the same B cell receptor.
- the second light chain variable region may include, for example, the light chain variable region of the second B cell receptor.
- the second light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region in the second B cell receptor, or the light chain in the second B cell receptor. It may be a polypeptide containing the amino acid sequence of the variable region.
- the second heavy chain variable region and the second light chain variable region are linked by, for example, the above-mentioned linker peptide (Fv linker peptide).
- the Fv linker peptide in the second scFv may be the same as or different from the Fv linker peptide in the first scFv, for example.
- the second antigen-binding domain includes a second scFv, but the second binding domain may have a structure other than scFv including the heavy chain variable region and the light chain variable region.
- the second binding domain may be Fab, Fab′, F(ab′) 2 , variable region fragment (Fv), disulfide bond Fv, or the like.
- the second transmembrane domain and the second intracellular signaling domain are, for example, the first transmembrane domain and the first intracellular signal in the first CAR library, respectively. It may be the same as or different from the transduction domain.
- the second transmembrane domain and the second intracellular signaling domain are, for example, the first transmembrane domain and the first intracellular signal in the CAR library of the present invention, respectively.
- the description of the transduction domain can be incorporated.
- the domains may be linked by, for example, a linker peptide (domain linker peptide).
- domain linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the domain linker peptide is composed of amino acids such as glycine and serine, and (GGGGS) n is a specific example.
- the n is, for example, an integer of 1 to 6.
- Examples of the second CAR include a polypeptide having the amino acid sequence represented by the formula (1) (SEQ ID NO: 15).
- V 1 and V 2 are the amino acid sequences of the heavy chain variable region and the light chain variable region, or the amino acid sequences of the light chain variable region and the heavy chain variable region, respectively.
- the order of the heavy chain variable region and the light chain variable region in the second CAR is, for example, that the second CAR library can be prepared more easily. The order and the reverse order are preferred.
- the heavy chain variable region and the light chain variable region are arranged in this order from the N-terminus in the first CAR, the light chain variable region and the heavy chain variable region and the heavy chain variable region are arranged in this order from the N-terminus in the second CAR.
- the chain variable regions are, for example, arranged in this order.
- the heavy chain variable region and the light chain variable region are arranged from the N-terminal in the second CAR. , For example, arranged in this order.
- nucleic acid encoding the second CAR examples include a polynucleotide having the base sequence represented by the formula (2) (SEQ ID NO: 16).
- N 1 and N 2 are the base sequences encoding the heavy chain variable region and the light chain variable region, or the base sequences encoding the light chain variable region and the heavy chain variable region, respectively.
- the second CAR library preferably contains, for example, a plurality of types of nucleic acids.
- the second CAR library is, for example, a mixture of plural kinds of nucleic acids.
- a part or all of the plurality of types of nucleic acids preferably encode different second CARs, and preferably encode different second antigen-binding domains.
- the regions of the second CAR other than the second antigen-binding domain have, for example, the same or different amino acid sequences.
- the types of the nucleic acids contained in the second CAR library are, for example, 1 ⁇ 10 5 to 1 ⁇ 10 7 types, 1 ⁇ 10 5 to 1 ⁇ 10 6 types, and 1 ⁇ 10 6 to 5 ⁇ 10 6 types. Yes, and preferably about 2 ⁇ 10 6 types (eg, 1 ⁇ 10 6 to 3 ⁇ 10 6 types).
- the second CAR may have a signal peptide at the N-terminus, for example.
- the second CAR may have a tag, for example.
- Regarding the signal peptide and tag for example, in the description of the signal peptide and tag in the first CAR, “first CAR” is replaced with “second CAR”, and the description can be incorporated.
- the nucleic acid encoding the second CAR can be prepared by a conventional method, for example, based on the amino acid sequence of the second CAR.
- the nucleic acid encoding the second CAR is obtained by, for example, obtaining a base sequence encoding the amino acid sequence in a database in which the amino acid sequence of each domain described above is registered, and based on the base sequence, a molecule It can be prepared by biological methods and/or chemical synthetic methods.
- the base sequence of the nucleic acid may be codon-optimized, for example, depending on the origin of cells expressing the second CAR library.
- the nucleic acid encoding the second CAR may be introduced into an expression vector, for example.
- an expression vector for example, the description of the expression vector in the CAR library of the present invention can be incorporated.
- the first screening method of the present invention is performed in the same manner except that the second CAR library prepared in the preparation step is used instead of the first CAR library in the same manner as in the expression step and the contact step, respectively. Perform the process and the selection process.
- the first screening method of the present invention further includes, for example, a second expression step of expressing the second CAR library in immune cells, and immune cells obtained in the second expression step.
- a second contact step of contacting the target antigen with the second contact step, and a second scFv of CAR expressed in immune cells bound to the target antigen in the second contact step Second selection step as the candidate scFv of.
- the immune cells in the first expression step, the first contact step, and the first selection step are the same as the immune cells in the second expression step, the second contact step, and the second selection step. However, it may be different.
- the immune cells are preferably T cells or T cell-like cells.
- the second expression step may be referred to as “first expression step” and “second expression step” and “first CAR library” and “second CAR.”
- the description can be incorporated by replacing “first CAR” with “second CAR” in “library”.
- the second contacting step is, for example, in the description of the first contacting step, the "first expressing step” is referred to as the “second expressing step”, and the “first contacting step” is referred to as the “second contacting step”.
- the description can be incorporated by replacing it with "process”.
- the second selection step is, for example, in the description of the first selection step, "first contact step” is referred to as “second contact step” and “first selection step” is referred to as “second selection step”. "Step”, “first scFv” to “second scFv”, “first candidate scFv” to “second candidate scFv”, “first CAR” to “second CAR” You can read it and use the explanation.
- the first screening method of the present invention can screen, for example, an scFv having a new heavy chain variable region and new light chain variable region that bind to the target antigen.
- the first screening method of the present invention may include a design step of designing an antibody or an antigen-binding fragment thereof based on the first candidate scFv or the second candidate scFv.
- a design step for example, in the first candidate scFv or the second candidate scFv, CDRH1, CDRH2, and CDRH3 in the heavy chain variable region and CDRL1, CDRL2, and CDRL3 in the light chain variable region are newly added. It can be carried out by transplanting the antibody or an antigen-binding fragment thereof.
- the designing step includes, for example, the amino acid sequences of CDRH1, CDRH2, and CDRH3 in the first candidate scFv or the second candidate scFv and the first candidate scFv or the second candidate scFv.
- the transplantation of CDRs in the design step may be performed for one CDR or a plurality of CDRs in the first candidate scFv or the second candidate scFv, for example, but Preference is given to carrying out the CDR.
- the types of the new antibody and the antigen-binding fragment thereof for example, the description of the types of the antibody of the present invention or the antigen-binding fragment thereof described below can be referred to.
- FRH1, FRH2, FRH3, and FRH4 in the heavy chain variable region and FRL1, FRL2, FRL3, and FRL4 in the light chain variable region are set.
- a new antibody or an antigen-binding fragment thereof may be transplanted.
- the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 in the first candidate scFv or the second candidate scFv and the first candidate scFv or the second candidate scFv are used.
- the FR transplantation in the design step may be performed for one FR or a plurality of FRs in the first candidate scFv or the second candidate scFv, for example. It is preferable to carry out for FR.
- the first screening method of the present invention includes the first expression step, but the present invention is not limited to this, and the first expression step may not be included.
- the first screening method of the present invention can be carried out from the first contact step using, for example, previously prepared immune cells expressing the CAR library of the present invention (CAR library cells).
- the chimeric antigen receptor (CAR) library cells of the present invention include cells expressing the CAR library of the present invention.
- the CAR library cells of the present invention are characterized in that they include cells expressing the CAR library of the present invention, and other configurations and conditions are not particularly limited.
- the first screening method of the present invention can be preferably carried out.
- the description of the CAR library of the present invention and the first screening method can be applied.
- the number of CAR library cells of the present invention is, for example, 1 ⁇ 10 5 to 1 ⁇ 10 8 cells, 1 ⁇ 10 5 to 1 ⁇ 10 7 cells, 1 ⁇ 10 5 to 1 ⁇ 10 6 cells, 1 ⁇ 10 6 to 5 ⁇ 10 6 cells, 1 ⁇ 10 6 to 3 ⁇ 10 6 cells.
- the CAR library cells preferably each express different CAR in the CAR library of the present invention.
- the CAR library cells may include, for example, cells that do not express the CAR library of the present invention, that is, cells into which the CAR library has not been introduced.
- the types of CAR expressed by the CAR library cells of the present invention are, for example, 1 ⁇ 10 4 to 1 ⁇ 10 7 types, 1 ⁇ 10 5 to 1 ⁇ 10 6 types, per 1 ⁇ 10 6 CAR library cells. There are ⁇ 10 6 to 5 ⁇ 10 6 types and 1 ⁇ 10 5 to 1 ⁇ 10 6 types.
- the CAR library cell of the present invention can be produced, for example, in the same manner as in the first expression step of the first screening method of the present invention.
- the CAR library cells of the present invention are, for example, preferably cells derived from immune cells, and are preferably derived from T cells, NK cells, and NKT cells.
- the antibody against the complex of HLA-A*02:01 and NY-ESO-1 157-165 (hereinafter, also referred to as “A2/NY-ESO-1 157 ”) or the antigen-binding fragment thereof of the present invention is as described above. Includes a heavy chain variable region (H) below and a light chain variable region (L) below.
- the antibody or the like of the present invention is characterized by containing the heavy chain variable region of (H) below and the light chain variable region of (L) below, and other configurations and conditions are not particularly limited.
- the antibody and the like of the present invention bind to A2/NY-ESO-1 157 .
- A2/NY-ESO-1 157 is known to be expressed in specific cancer cells such as lung cancer, malignant melanoma, synovial sarcoma, myeloma. Therefore, the antibody and the like of the present invention can be preferably used as, for example, a bispecific antibody against A2/NY-ESO-1 157- expressing cancer cells, a CAR antigen-binding domain of CAR-T cells, and the like.
- the description of the CAR library of the present invention, the first screening method and the like can be incorporated.
- (H) comprises heavy chain complementarity determining region (CDRH) 1, CDRH2, and CDRH3,
- CDRH1 is a polypeptide containing the following amino acid sequence (H1)
- CDRH2 is a polypeptide containing the following amino acid sequence (H2)
- CDRH3 is a polypeptide containing the amino acid sequence of (H3) below
- one or several amino acids are deleted, substituted, inserted and/or added (H2)
- amino acid sequence of one or several amino acids have been deleted, substituted, inserted and/or added (H3) of the following (H3-1), (H3-2) or (H3-3) Amino acid sequence (H3-1) Amino acid sequence (H3-3) having 80% or more identity to the amino acid sequence of any of the CDRH3s (H3-2) (H3-1) in Table 1A below.
- amino acid sequence of (H3-1) an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added
- (L) comprises light chain complementarity determining regions (CDRL) 1, CDRL2, and CDRL3,
- CDRL1 is a polypeptide containing the following amino acid sequence (L1)
- CDRL2 is a polypeptide containing the following amino acid sequence (L2)
- CDRL3 is a polypeptide containing the following amino acid sequence (L3) light chain variable region (L1) below (L1-1), (L1-2) or (L1-3) amino acid sequence (L1-1) below Amino acid sequence (L1-3) (L1-1) having 80% or more identity to the amino acid sequence (L1-2) (L1-1) of any one of CDRL1 in Table 1B.
- one or several amino acids are deleted, substituted, inserted and/or added (L2) The following (L2-1), (L2-2) or (L2-3) amino acid sequence (L2 -1)
- one or several amino acids are deleted, substituted, inserted and/or added (L3) of the following (L3-1), (L3-2) or (L3-3) Amino acid sequence (L3-1) Amino acid sequence (L3-3) having 80% or more identity with the amino acid sequence of any of CDRL3 (L3-2) (L3-1) of Table 1B below.
- HLA-A*02:01 means, for example, a class I antigen derived from the A*02:01 allele of human major histocompatibility complex (MHC).
- HLA-A*02:01 forms a complex with, for example, human ⁇ 2-microglobulin.
- the amino acid sequence of HLA-A*02:01 includes, for example, the amino acid sequence registered under NCBI Accession No. HG794376.
- the amino acid sequence of human ⁇ 2-microglobulin includes, for example, the amino acid sequence registered under NCBI Accession No. NM_004048.2.
- NY-ESO-1 157-165 means, for example, a peptide consisting of amino acids 157 to 165 (SLLMWITQC (SEQ ID NO: 206) in the amino acid sequence of NY-ESO-1 protein.
- ESO-1 is a cancer antigen expressed in cancers such as human lung cancer, malignant melanoma, synovial sarcoma, myeloma, etc.
- the amino acid sequence of NY-ESO-1 protein is, for example, NCBI accession number. The amino acid sequence registered in NM_001327.2 is listed.
- A2/NY-ESO-1 157 means, for example, a complex formed by HLA-A*02:01 and NY-ESO-1 157-165, and specifically, It means a complex in which NY-ESO-1 157-165 is bound to the peptide binding groove of HLA-A*02:01.
- A2/NY-ESO-1 157 is preferably a complex in which NY-ESO-1 157-165 is bound to the peptide binding groove of the complex of HLA-A*02:01 and human ⁇ 2-microglobulin. ..
- the antibody of the present invention may be, for example, a so-called “antibody” whose molecular structure is immunoglobulin, or an antigen-binding fragment thereof.
- the antibody of the present invention may have the heavy chain variable region and the light chain variable region described above.
- the present invention is an antibody, for example, its immunoglobulin class and isotype are not particularly limited.
- the immunoglobulin class include IgG, IgM, IgA, IgD, and IgE.
- Examples of the IgG include IgG1, IgG2, IgG3, IgG4 and the like.
- the antibody may be, for example, a monoclonal antibody, a polyclonal antibody, a recombinant antibody, a human (for example, fully human) antibody, a humanized antibody, a chimeric antibody, a multispecific antibody, or the like.
- the “antigen-binding fragment” in the present invention means a part of the antibody, for example, a partial fragment, which recognizes (binds) the A2/NY-ESO-1 157 .
- the antigen-binding fragment include Fab, Fab′, F(ab′) 2 , variable region fragment (Fv), disulfide bond Fv, single chain Fv (scFv), bispecific antibody and polymers thereof.
- Fv variable region fragment
- scFv single chain Fv
- bispecific antibody bispecific antibody and polymers thereof.
- the heavy chain variable region and the light chain variable region are linked via a linker.
- the order of the heavy chain variable region, the linker, and the light chain variable region may be from the N terminus to the heavy chain variable region, the linker, and the light chain variable region, or from the N terminus to the light chain. It may be the order of the chain region, the linker, and the heavy chain variable region.
- the antibody or the like of the present invention may have, for example, a constant region in addition to the heavy chain variable region and the light chain variable region described above, and the constant region is, for example, a human constant region or a mouse constant region.
- the heavy chain constant region includes, for example, the regions CH1, CH2, and CH3, and the light chain constant region includes, for example, the region CL.
- the antibody or the like of the present invention has the constant region, for example, the heavy chain variable region is bound to at least one of CH1, CH2, and CH3, and the light chain variable region is bound to the CL, The heavy chain variable region is directly linked to, for example, CH1.
- the heavy and light chains of an antibody molecule each have three complementarity determining regions (CDRs).
- CDR is also called a hypervariable domain. Even in the variable regions of the heavy chain and the light chain, CDR is a region in which the primary structure has a high variability, and the CDR is usually separated at three positions on the primary structure.
- the three CDRs in the heavy chain are referred to as the heavy chain CDR1 (CDRH1), the heavy chain CDR2 (CDRH2), and the heavy chain CDR3 (CDRH3) from the amino terminal side in the amino acid sequence of the heavy chain.
- the three CDRs in the chain are represented as light chain CDR1 (CDRL1), light chain CDR2 (CDRL2), and light chain CDR3 (CDRL3) from the amino terminal side in the amino acid sequence of the light chain. These sites are conformationally close to each other and determine their specificity for binding antigen.
- the CDRH1 is the CDRH1 in the (HA), (HB), or (HC).
- the CDRH2 is the CDRH2 in (HA), (HB) or (HC).
- the CDRH3 is the CDRH3 in (HA), (HB) or (HC).
- the CDRL1 is any one of CDLA1 of (LA) to (LQ) and (LR).
- the CDRL2 is any one of the above (LA) to (LQ) and (LR).
- the CDRL3 is any one of the above (LA) to (LQ) and (LR).
- identity is, for example, the degree of identity when the sequences to be compared are properly aligned with each other, and means the occurrence rate (%) of exact matches of amino acids between the sequences.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- the identity can be calculated with default parameters using analysis software such as BLAST and FASTA (the same applies hereinafter).
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the substitution of the amino acid may be, for example, a conservative substitution (hereinafter the same).
- the conservative substitution means substituting one or several amino acids with other amino acids and/or amino acid derivatives so as not to substantially change the function of the protein.
- the “substituted amino acid” and the “substituted amino acid” have similar properties and/or functions, for example.
- the hydrophobic and hydrophilic indices (hydropathy), chemical properties such as polarity and charge, or physical properties such as secondary structure are similar.
- Amino acids or amino acid derivatives having similar properties and/or functions are known in the art, for example.
- non-polar amino acids such as alanine, valine, isoleucine, leucine, proline, tryptophan, phenylalanine and methionine
- polar amino acids neutral amino acids
- neutral amino acids include glycine, serine and threonine.
- positively charged amino acids include arginine, histidine, lysine, etc.
- negatively charged amino acids include aspartic acid, glutamic acid.
- the combination of (H1-1), (H2-1) and (H3-1) is not particularly limited, and examples thereof include (HA) and (HB), respectively. Alternatively, it may be any combination of CDRH1 in (HC), CDRH2 in (HA), (HB) or (HC), and CDRH3 in (HA) or (HB).
- the combination of (H1-1), (H2-1) and (H3-1) is preferably CDRH1, CDRH2 and CDRH3 in the above (HA), (HB) or (HC).
- the combination of (L1-1), (L2-1) and (L3-1) is not particularly limited, and examples thereof include (LA) to (LQ), respectively. And any one CDRL1 in (LR), any one CDRL2 in (LA) to (LQ) and (LR) above, and any one CDRL3 in (LA) to (LQ) and (LR) above Can be any combination.
- the combination of (L1-1), (L2-1) and (L3-1) is preferably any one of CDRL1, CDRL2 and CDRL3 in (LA) to (LQ) and (LR).
- the combination of (H1-1), (H2-1) and (H3-1) with (L1-1), (L2-1) and (L3-1) is not particularly limited, and, for example, CDRH1 in the above (HA), (HB) or (HC), CDRH2 in the above (HA), (HB) or (HC), and CDRH3 in the above (HA), (HB) or (HC); Any one of CDRL1 in (LA) to (LQ) and (LR), any one of CDRL2 in (LA) to (LQ) and (LR), and (LA) to (LQ) and (LR) In any combination with CDRL3.
- the combination of (H1-1), (H2-1) and (H3-1) with (L1-1), (L2-1) and (L3-1) is preferably respectively the above ( HA), (HB) or (HC) in combination with CDRH1, CDRH2 and CDRH3 and any one of the above (LA) to (LQ) and (LR) CDRL1, CDRL2 and CDRL3, more preferably, It is a combination of Table 2 below.
- the antibody of the present invention and the like will be described more specifically with respect to the combination of the heavy chain variable region or heavy chain and the light chain variable region or light chain.
- the description of the heavy chain variable region and the heavy chain in each combination can be incorporated into each other. Further, the description of the light chain variable region and the light chain in each combination can be incorporated into each other.
- the underlined amino acid sequences and base sequences are the base sequences encoding the amino acid sequences corresponding to CDRs and the amino acid sequences corresponding to CDRs, unless otherwise specified.
- the combination of the heavy chain variable region and the light chain variable region is, for example, the combination of (1) to (18) described above.
- the antibodies and the like of the combination (1) are also referred to as, for example, antibody H1-3M4E5L group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A) below, and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A) below.
- the light chain variable region of (LA) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-A) below, and CDRL2 is (L2-A) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-A) below.
- H1-A Amino acid sequence of (H1-A1), (H1-A2) or (H1-A3) below (H1-A1) SEQ ID NO: 17 (GGSISSNY) (H1-A2) Amino acid of SEQ ID NO: 17 Amino acid sequence having 80% or more identity to the sequence (H1-A3) In the amino acid sequence of SEQ ID NO: 17, one or several amino acids are deleted, substituted, inserted and/or added
- H2-A The following (H2-A1), (H2-A2) or (H2-A3) amino acid sequence (H2-A1) SEQ ID NO: 18 (VSYSGST) amino acid sequence (H2-A2) SEQ ID NO: 18 amino acid Amino acid sequence (H2-A3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 18.
- H3-A Amino acid sequence of (H3-A1), (H3-A2) or (H3-A3) below (H3-A1) SEQ ID NO: 19 (ARESYYYYGMDV) (H3-A2) Amino acid of SEQ ID NO: 19 Amino acid sequence (H3-A3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 19
- (L1-A) Amino acid sequence of (L1-A1), (L1-A2) or (L1-A3) below (L1-A1) SEQ ID NO: 29 (SRDVGGYNY) (L1-A2) SEQ ID NO: 29 Amino acid sequence having 80% or more identity to the sequence (L1-A3) SEQ ID NO: 29, wherein one or several amino acids are deleted, substituted, inserted and/or added
- L2-A The following (L2-A1), (L2-A2) or (L2-A3) amino acid sequence (L2-A1) SEQ ID NO: 30 (DVI) amino acid sequence (L2-A2) SEQ ID NO: 30 Amino acid sequence having 80% or more identity to the sequence (L2-A3), wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 30.
- (L3-A) Amino acid sequence of the following (L3-A1), (L3-A2) or (L3-A3) (L3-A1) SEQ ID NO: 31 (WSFAGSYYV) (L3-A2) Amino acid of SEQ ID NO: 31 Amino acid sequence having 80% or more identity with the sequence (L3-A3) SEQ ID NO: 31, with one or several amino acids deleted, substituted, inserted and/or added
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide having the amino acid sequence of (H-A) below.
- the light chain variable region of (LA) includes, for example, a polypeptide having the amino acid sequence of (LA) below.
- H-A1 following (H-A1), (H -A2) or (H-A3) of the amino acid sequence (H-A1) of SEQ ID NO: 20 amino acid sequence SEQ ID NO 20: QVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSS (H-A2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 20 (H-A3) In the amino acid sequence of SEQ ID NO: 20, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- (LA) Amino acid sequence of the following (L-A1), (L-A2) or (L-A3) (L-A1) Amino acid sequence of SEQ ID NO: 32 SEQ ID NO: 32: QSELTQPRSVSGSPGQSVTISCTGT SRDVGGYNY VSWYQQHPGKAPKLIIH DVI ERSSGVPDRFSGSKSGNTASLTISGLQAEDEVLYYC FFA (L-A2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 32 (L-A3) In the amino acid sequence of SEQ ID NO: 32, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (H-A1) amino acid sequence is, for example, a sequence containing the (H1-A1) of CDRH1, (H2-A1) of CDRH2, and (H3-A1) of CDRH3.
- the amino acid sequence of (H-A2) includes, for example, (H1-A1) of CDRH1, (H2-A1) of CDRH2, and CDRH3 (H3-A1), and the amino acid sequence of SEQ ID NO: 20.
- the amino acid sequence may have an identity of 80% or more.
- the amino acid sequence of (H-A3) includes, for example, the amino acid sequences of (H1-A1) of CDRH1, (H2-A1) of CDRH2, and CDRH3 (H3-A1), and the amino acid sequence of SEQ ID NO: 20. In, it may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added.
- the above-mentioned (L-A1) amino acid sequence is, for example, a sequence including the CDL1 (L1-A1), CDRL2 (L2-A1), and CDRL3 (L3-A1) amino acid sequences.
- the amino acid sequence of (L-A2) includes, for example, the amino acid sequences of (L1-A1) of CDRL1, (L2-A1) of CDRL2, and (L3-A1) of CDRL3, and the amino acid of SEQ ID NO: 32. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-A3) includes, for example, the amino acid sequences of (L1-A1) of CDRL1, (L2-A1) of CDRL2, and (L3-A1) of CDRL3, and the amino acid of SEQ ID NO: 32. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-A1). Also known as "H1-3M4E5L”.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, respectively. It is 97% or more, 98% or more, and 99% or more.
- “1 or several” regarding substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, respectively.
- the numbers are 1 to 9, 1 to 8, 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LB) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-B) below, and CDRL2 is (L2-B) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-B) below.
- (L1-B) Amino acid sequence of (L1-B1), (L1-B2) or (L1-B3) below (L1-B1) SEQ ID NO: 33 (QSISSY) amino acid sequence (L1-B2) SEQ ID NO: 33 Amino acid sequence having 80% or more identity to the sequence (L1-B3) Amino acid sequence of SEQ ID NO: 33 with one or several amino acids deleted, substituted, inserted and/or added
- L2-B The following (L2-B1), (L2-B2) or (L2-B3) amino acid sequence (L2-B1) SEQ ID NO: 34 (AAS) amino acid sequence (L2-B2) SEQ ID NO: 34 amino acid Amino acid sequence having 80% or more identity to the sequence (L2-B3)
- amino acid sequence of SEQ ID NO: 34 one or several amino acids have been deleted, substituted, inserted and/or added
- (L3-B) The following (L3-B1), (L3-B2) or (L3-B3) amino acid sequence (L3-B1) SEQ ID NO: 35 (QQYYSTPQT) amino acid sequence (L3-B2) SEQ ID NO: 35 amino acid Amino acid sequence having 80% or more identity to the sequence (L3-B3), amino acid sequence of SEQ ID NO: 35 with one or several amino acids deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (HA).
- the light chain variable region of (LB) includes, for example, a polypeptide having the amino acid sequence of (LB) below.
- the above-mentioned (L-B1) amino acid sequence is, for example, a sequence including the CDL1 (L1-B1), CDRL2 (L2-B1), and CDRL3 (L3-B1) amino acid sequences.
- the amino acid sequence of (L-B2) includes, for example, (L1-B1) of CDRL1, (L2-B1) of CDRL2, and (L3-B1) of CDRL3, and the amino acid of SEQ ID NO: 36. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-B3) includes, for example, the amino acid sequences of (L1-B1) of CDRL1, (L2-B1) of CDRL2, and (L3-B1) of CDRL3, and the amino acid of SEQ ID NO: 36. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-B1)
- antibody of this combination hereinafter referred to as "antibody Also called H1-K52".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LC) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-C) below, and CDRL2 is (L2-C) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-C) below.
- (L1-C) The following (L1-C1), (L1-C2) or (L1-C3) amino acid sequence (L1-C1) SEQ ID NO:37 (QSISSY) amino acid sequence (L1-C2) SEQ ID NO:37 amino acid Amino acid sequence having 80% or more identity to the sequence (L1-C3) SEQ ID NO: 37 amino acid sequence with one or several amino acids deleted, substituted, inserted and/or added
- (L2-C) Amino acid sequence of the following (L2-C1), (L2-C2) or (L2-C3) (L2-C1) SEQ ID NO: 38 (AAS) (L2-C2) Amino acid of SEQ ID NO: 38 Amino acid sequence (L2-C3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 38.
- (L3-C) Amino acid sequence of (L3-C1), (L3-C2) or (L3-C3) below (L3-C1) SEQ ID NO: 39 (QQYESYRRS) (L3-C2) Amino acid of SEQ ID NO: 39 Amino acid sequence having 80% or more identity to the sequence (L3-C3) In the amino acid sequence of SEQ ID NO: 39, one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (HA).
- the light chain variable region of (LC) includes, for example, a polypeptide having the amino acid sequence of (LC) below.
- LC following (L-C1), (L -C2) or (L-C3) amino acid sequence (L-C1) SEQ ID NO: 40 amino acid sequence SEQ ID NO 40: DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKAGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYC QQYESYRRS FGQGTKVEIK (L-C2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 40 (L-C3) In the amino acid sequence of SEQ ID NO: 40, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-C1) amino acid sequence is, for example, a sequence including the CDL1 (L1-C1), CDRL2 (L2-C1), and CDRL3 (L3-C1) amino acid sequences.
- the amino acid sequence of (L-C2) includes, for example, the amino acid sequences of (L1-C1) of CDRL1, (L2-C1) of CDRL2, and (L3-C1) of CDRL3, and the amino acid of SEQ ID NO: 40. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-C3) includes, for example, the amino acid sequences of (L1-C1) of CDRL1, (L2-C1) of CDRL2, and (L3-C1) of CDRL3, and the amino acid of SEQ ID NO: 40. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-C1)
- antibody of this combination hereinafter referred to as "antibody Also called H1-K73".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LD) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-D) below, and CDRL2 is (L2-D) below.
- CDRL3 is a polypeptide containing the amino acid sequence of (L3-D) below.
- (L1-D) Amino acid sequence of the following (L1-D1), (L1-D2) or (L1-D3) (L1-D1) SEQ ID NO: 41 (QSISSY) (L1-D2) Amino acid of SEQ ID NO: 41 Amino acid sequence having 80% or more identity to the sequence (L1-D3), amino acid sequence of SEQ ID NO: 41 with one or several amino acids deleted, substituted, inserted and/or added
- L2-D The following (L2-D1), (L2-D2) or (L2-D3) amino acid sequence (L2-D1) SEQ ID NO: 42 (AAS) amino acid sequence (L2-D2) SEQ ID NO: 42 amino acid Amino acid sequence having 80% or more identity to the sequence (L2-D3) SEQ ID NO: 42, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L3-D) Amino acid sequence of the following (L3-D1), (L3-D2) or (L3-D3) (L3-D1) SEQ ID NO:43 (QQYNSYSRT) (L3-D2) Amino acid of SEQ ID NO:43 Amino acid sequence having 80% or more identity to the sequence (L3-D3) SEQ ID NO: 43 amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (HA).
- the light chain variable region of (LD) includes, for example, a polypeptide having the amino acid sequence of (L-DA) or (L-DB) below.
- L-DA following (L-DA1), (L -DA2) or (L-DA3) amino acid sequence (L-DA1) of SEQ ID NO: 44 amino acid sequence SEQ ID NO 44: DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDFATYYC QQYNSYSRT FGQGTKVEIK (L-DA2) Amino acid sequence having 80% or more identity with the amino acid sequence of SEQ ID NO:44 (L-DA3) In the amino acid sequence of SEQ ID NO:44, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- L-DB following (L-DB1), (L -DB2) or (L-DB3) of the amino acid sequence (L-DB1) amino acid sequence SEQ ID NO: 45
- SEQ ID NO 45 DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS RLESGVPSRFSGSGSGTDFTLTISCLQSEDFATYYC QQYNSYSRT FGQGTKVEIK
- L-DB2 Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 45 (L-DB3)
- amino acid sequence of SEQ ID NO: 45 one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-DA1) amino acid sequence is, for example, a sequence including the CDL1 (L1-D1), CDRL2 (L2-D1), and CDRL3 (L3-D1) amino acid sequences.
- the amino acid sequence of (L-DA2) includes, for example, the amino acid sequences of (L1-D1) of CDRL1, (L2-D1) of CDRL2, and (L3-D1) of CDRL3, and the amino acid of SEQ ID NO:44. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-DA3) includes, for example, the amino acid sequences of (L1-D1) of CDRL1, (L2-D1) of CDRL2, and (L3-D1) of CDRL3, and the amino acid of SEQ ID NO:44. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the above-mentioned (L-DB1) amino acid sequence is, for example, a sequence including the CDL1 (L1-D1), CDRL2 (L2-D1), and CDRL3 (L3-D1) amino acid sequences.
- the amino acid sequence of (L-DB2) includes, for example, the amino acid sequences of (L1-D1) of CDRL1, (L2-D1) of CDRL2, and (L3-D1) of CDRL3, and the amino acid of SEQ ID NO: 45. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-DB3) includes, for example, the amino acid sequences of (L1-D1) of CDRL1, (L2-D1) of CDRL2, and (L3-D1) of CDRL3, and the amino acid of SEQ ID NO: 45. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-DA1). Also called H1-K121”.
- the antibody of the present invention for example, the heavy chain variable region is the (H-A1), the light chain variable region is the (L-DB1), the antibody of this combination, Also referred to as "antibody H1-K124".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LE) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-E) below, and CDRL2 is (L2-E) below.
- CDRL3 is a polypeptide containing the amino acid sequence of (L3-E) below.
- (L1-E) The following (L1-E1), (L1-E2) or (L1-E3) amino acid sequence (L1-E1) SEQ ID NO: 46 (QSISSY) amino acid sequence (L1-E2) SEQ ID NO: 46 amino acid Amino acid sequence (L1-E3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 46.
- (L2-E) Amino acid sequence of the following (L2-E1), (L2-E2) or (L2-E3) (L2-E1) SEQ ID NO: 47 (AAS) (L2-E2) Amino acid of SEQ ID NO: 47 Amino acid sequence (L2-E3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 47.
- (L3-E) Amino acid sequence of the following (L3-E1), (L3-E2) or (L3-E3) (L3-E1) SEQ ID NO: 48 (QQYNSYSPCT) (L3-E2) Amino acid of SEQ ID NO: 48 Amino acid sequence having 80% or more identity to the sequence (L3-E3) SEQ ID NO: 48, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide having the amino acid sequence of (H-A).
- the light chain variable region of (LE) above includes, for example, a polypeptide having the amino acid sequence of (LE) below.
- the above-mentioned (L-E1) amino acid sequence is, for example, a sequence including the CDL1 (L1-E1), CDRL2 (L2-E1), and CDRL3 (L3-E1) amino acid sequences.
- the amino acid sequence of (L-E2) includes, for example, the amino acid sequences of (L1-E1) of CDRL1, (L2-E1) of CDRL2, and (L3-E1) of CDRL3, and the amino acid of SEQ ID NO: 49. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-E3) includes, for example, the amino acid sequences of (L1-E1) of CDRL1, (L2-E1) of CDRL2, and (L3-E1) of CDRL3, and the amino acid of SEQ ID NO:49. It may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is (H-A1)
- the light chain variable region is (L-E1)
- the antibody of this combination is referred to as "antibody Also called H1-K125".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LF) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-F) below, and CDRL2 is (L2-F) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-F) below.
- (L1-F) Amino acid sequence of the following (L1-F1), (L1-F2) or (L1-F3) (L1-F1) SEQ ID NO: 50 (QDISRY) (L1-F2) Amino acid of SEQ ID NO: 50 Amino acid sequence (L1-F3) having 80% or more identity with the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 50.
- L2-F The following (L2-F1), (L2-F2) or (L2-F3) amino acid sequence (L2-F1) SEQ ID NO: 51 (AAS) amino acid sequence (L2-F2) SEQ ID NO: 51 amino acid Amino acid sequence having 80% or more identity to the sequence (L2-F3) SEQ ID NO: 51, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L3-F) The following (L3-F1), (L3-F2) or (L3-F3) amino acid sequence (L3-F1) SEQ ID NO: 52 (QQYDNLIT) amino acid sequence (L3-F2) SEQ ID NO: 52 amino acid Amino acid sequence having 80% or more identity to the sequence (L3-F3) SEQ ID NO: 52, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LF) includes, for example, a polypeptide having the amino acid sequence of (LF) below.
- the above-mentioned (L-F1) amino acid sequence is, for example, a sequence containing the amino acid sequences of (L1-F1) of CDRL1, (L2-F1) of CDRL2, and (L3-F1) of CDRL3.
- the amino acid sequence of (L-F2) includes, for example, the amino acid sequences of (L1-F1) of CDRL1, (L2-F1) of CDRL2, and (L3-F1) of CDRL3, and the amino acid of SEQ ID NO: 53. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-F3) includes, for example, the amino acid sequences of (L1-F1) of CDRL1, (L2-F1) of CDRL2, and (L3-F1) of CDRL3, and the amino acid of SEQ ID NO: 53. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-F1). Also called H1-K131".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LG) includes CDRL1, CDRL2, and CDRL3, wherein CDRL1 is a polypeptide containing the amino acid sequence of (L1-G) below, and CDRL2 is (L2-G) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-G) below.
- (L1-G) Amino acid sequence (L1-G1), (L1-G2) or (L1-G3) amino acid sequence (L1-G1) SEQ ID NO: 54 (QDISRY) (L1-G2) SEQ ID NO: 54 Amino acid sequence having 80% or more identity to the sequence (L1-G3), amino acid sequence of SEQ ID NO: 54 with one or several amino acids deleted, substituted, inserted and/or added
- (L2-G) Amino acid sequence of the following (L2-G1), (L2-G2) or (L2-G3) (L2-G1) SEQ ID NO: 55 (AAS) (L2-G2) SEQ ID NO: 55 Amino acid sequence having 80% or more identity to the sequence (L2-G3) SEQ ID NO: 55, in which one or several amino acids are deleted, substituted, inserted and/or added
- (L3-G) Amino acid sequence of the following (L3-G1), (L3-G2) or (L3-G3) (L3-G1) SEQ ID NO: 56 (QQYNSYSRT) (L3-G2) Amino acid of SEQ ID NO: 56 Amino acid sequence having 80% or more identity to the sequence (L3-G3) In the amino acid sequence of SEQ ID NO: 56, one or several amino acids have been deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (HA).
- the light chain variable region of (LG) includes, for example, a polypeptide having the following amino acid sequence (L-G).
- the (L-G1) amino acid sequence is, for example, a sequence containing the CDL1 (L1-G1), CDRL2 (L2-G1), and CDRL3 (L3-G1) amino acid sequences.
- the amino acid sequence of (L-G2) includes, for example, (L1-G1) of CDRL1, (L2-G1) of CDRL2, and (L3-G1) of CDRL3, and the amino acid of SEQ ID NO: 57. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-G3) includes, for example, the amino acid sequences of (L1-G1) of CDRL1, (L2-G1) of CDRL2, and (L3-G1) of CDRL3, and the amino acid of SEQ ID NO: 57. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-G1)
- antibody of this combination hereinafter referred to as "antibody Also called H1-K145”.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LH) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-H) below, and CDRL2 is (L2-H) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-H) below.
- (L1-H) The following (L1-H1), (L1-H2) or (L1-H3) amino acid sequence (L1-H1) SEQ ID NO:58 (QSISSY) amino acid sequence (L1-H2) SEQ ID NO:58 amino acid Amino acid sequence (L1-H3) having 80% or more identity to the sequence, wherein the amino acid sequence of SEQ ID NO: 58 has one or several amino acids deleted, substituted, inserted and/or added
- (L2-H) The following (L2-H1), (L2-H2) or (L2-H3) amino acid sequence (L2-H1) SEQ ID NO:59 (AAS) amino acid sequence (L2-H2) SEQ ID NO:59 amino acid Amino acid sequence having 80% or more identity to the sequence (L2-H3) SEQ ID NO: 59, wherein one or several amino acids have been deleted, substituted, inserted and/or added
- (L3-H) Amino acid sequence of (L3-H1), (L3-H2) or (L3-H3) below (L3-H1) SEQ ID NO: 60 (QQYDNLIT) (L3-H2) Amino acid of SEQ ID NO: 60 Amino acid sequence having 80% or more identity to the sequence (L3-H3) SEQ ID NO: 60, in which one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LH) includes, for example, a polypeptide having the amino acid sequence of (LH) below.
- (LH) Amino acid sequence of (L-H1), (L-H2) or (L-H3) below (L-H1) Amino acid sequence of SEQ ID NO: 61
- SEQ ID NO: 61 DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTITSLQPDDFATYYC Q
- L-H2 Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 61 (L-H3)
- amino acid sequence of SEQ ID NO: 61 one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-H1) amino acid sequence is, for example, a sequence containing the amino acid sequences of (L1-H1) of CDRL1, (L2-H1) of CDRL2, and (L3-H1) of CDRL3.
- the amino acid sequence of (L-H2) includes, for example, the amino acid sequences of (L1-H1) of CDRL1, (L2-H1) of CDRL2, and (L3-H1) of CDRL3, and the amino acid of SEQ ID NO: 61. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-H3) includes, for example, (L1-H1) of CDRL1, (L2-H1) of CDRL2, and (L3-H1) of CDRL3, and the amino acid of SEQ ID NO: 61. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-H1)
- the antibody of this combination is referred to below as "antibody Also called H1-K151”.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LI) above includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-I) below, and CDRL2 is (L2-I) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-I) below.
- (L1-I) The following (L1-I1), (L1-I2) or (L1-I3) amino acid sequence (L1-I1) SEQ ID NO: 62 (QSVSSN) amino acid sequence (L1-I2) SEQ ID NO: 62 amino acid Amino acid sequence (L1-I3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 62.
- (L2-I) Amino acid sequence of the following (L2-I1), (L2-I2) or (L2-I3) (L2-I1) SEQ ID NO: 63 (GAS) (L2-I2) Amino acid of SEQ ID NO: 63 Amino acid sequence having 80% or more identity to the sequence (L2-I3) SEQ ID NO: 63, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L3-I) Amino acid sequence of (L3-I1), (L3-I2) or (L3-I3) below (L3-I1) SEQ ID NO: 64 (QQYNSYSRT) (L3-I2) Amino acid of SEQ ID NO: 64 Amino acid sequence having 80% or more identity to the sequence (L3-I3) SEQ ID NO: 64, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LI) includes, for example, a polypeptide having the amino acid sequence of (LI) below.
- (LI) Amino acid sequence of (L-I1), (L-I2) or (L-I3) below (L-I1) Amino acid sequence of SEQ ID NO: 65 SEQ ID NO: 65: EIVLTQSPATLSVSPGERATLSCRAS QSVSSN LAWYQQKPGQAPRLLIY GAS TRATGIPARFSGSGSGTEFTLTISRLEPEDFATYYC QQGTKSYS (L-I2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 65 (L-I3) In the amino acid sequence of SEQ ID NO: 65, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-I1) amino acid sequence is, for example, a sequence including the CDL1 (L1-I1), CDRL2 (L2-I1), and CDRL3 (L3-I1) amino acid sequences.
- the amino acid sequence of (L-I2) includes, for example, (L1-I1) of CDRL1, (L2-I1) of CDRL2, and (L3-I1) of CDRL3, and the amino acid of SEQ ID NO: 65. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-I3) includes, for example, the amino acid sequences of (L1-I1) of CDRL1, (L2-I1) of CDRL2, and (L3-I1) of CDRL3, and the amino acid of SEQ ID NO: 65. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-I1). Also called H1-K160".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the antibody etc. of the combination (10) are also referred to as, for example, antibody H1-K173 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LJ) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-J) below, and CDRL2 is (L2-J) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-J) below.
- (L1-J) Amino acid sequence of the following (L1-J1), (L1-J2) or (L1-J3) (L1-J1) SEQ ID NO: 66 (QSISSY) amino acid sequence (L1-J2) SEQ ID NO: 66 Amino acid sequence having 80% or more identity to the sequence (L1-J3), amino acid sequence of SEQ ID NO: 66 with one or several amino acids deleted, substituted, inserted and/or added
- (L2-J) Amino acid sequence of the following (L2-J1), (L2-J2) or (L2-J3) (L2-J1) SEQ ID NO:67 (AAS) (L2-J2) Amino acid of SEQ ID NO:67 Amino acid sequence (L2-J3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 67.
- (L3-J) Amino acid sequence of the following (L3-J1), (L3-J2) or (L3-J3) (L3-J1) SEQ ID NO:68 (QQYESYSRT) (L3-J2) Amino acid of SEQ ID NO:68 Amino acid sequence having 80% or more identity to the sequence (L3-J3) SEQ ID NO: 68, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (HA).
- the (LJ) light chain variable region includes, for example, a polypeptide consisting of the following (LJ) amino acid sequence.
- LJ amino acid sequence (L-J1) of SEQ ID NO: 69 amino acid sequence
- SEQ ID NO 69 DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKAGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYC QQYESYSRT FGQGTKVEIK
- L-J2 Amino acid sequence having 80% or more identity with the amino acid sequence of SEQ ID NO: 69 (L-J3)
- amino acid sequence of SEQ ID NO: 69 one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-J1) amino acid sequence is, for example, a sequence including the CDL1 (L1-J1), CDRL2 (L2-J1), and CDRL3 (L3-J1) amino acid sequences.
- the amino acid sequence of (L-J2) includes, for example, the amino acid sequences of (L1-J1) of CDRL1, (L2-J1) of CDRL2, and (L3-J1) of CDRL3, and the amino acid of SEQ ID NO: 69. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-J3) includes, for example, (L1-J1) of CDRL1, (L2-J1) of CDRL2, and (L3-J1) of CDRL3, and the amino acid of SEQ ID NO: 69. It may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-J1)
- antibody of this combination hereinafter referred to as "antibody Also called H1-K173".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the antibody etc. of the combination (11) are also referred to as, for example, antibody 3M4E5H-L1 group.
- the heavy chain variable region of (HB) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-B) below, and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-B), and CDRH3 is a polypeptide containing the amino acid sequence of the following (H3-B).
- the light chain variable region of (LK) above includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-K) below, and CDRL2 is (L2-K) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-K) below.
- H1-B Amino acid sequence of (H1-B1), (H1-B2) or (H1-B3) below (H1-B1) SEQ ID NO: 21 (GFTFSTYQ) (H1-B2) Amino acid of SEQ ID NO: 21 Amino acid sequence (H1-B3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 21.
- H2-B The following (H2-B1), (H2-B2) or (H2-B3) amino acid sequence (H2-B1) SEQ ID NO:22 (IVSSGGST) amino acid sequence (H2-B2) SEQ ID NO:22 amino acid Amino acid sequence having 80% or more identity with the sequence (H2-B3) SEQ ID NO: 22 with one or several amino acids deleted, substituted, inserted and/or added
- H3-B Amino acid sequence of (H3-B1), (H3-B2) or (H3-B3) below (H3-B1) SEQ ID NO:23 (AGELLPYYGMDV) (H3-B2) Amino acid of SEQ ID NO:23 Amino acid sequence having 80% or more identity to the sequence (H3-B3) In the amino acid sequence of SEQ ID NO: 23, one or several amino acids have been deleted, substituted, inserted and/or added
- (L1-K) The following (L1-K1), (L1-K2) or (L1-K3) amino acid sequence (L1-K1) SEQ ID NO: 70 (SSDVGGYDF) amino acid sequence (L1-K2) SEQ ID NO: 70 amino acid Amino acid sequence (L1-K3) having 80% or more identity to the sequence, wherein the amino acid sequence of SEQ ID NO: 70 has one or several amino acids deleted, substituted, inserted and/or added
- (L3-K) Amino acid sequence of (L3-K1), (L3-K2) or (L3-K3) below (L3-K1) SEQ ID NO: 72 (SSYAGSNSV) (L3-K2) Amino acid of SEQ ID NO: 72 Amino acid sequence having 80% or more identity to the sequence (L3-K3) SEQ ID NO: 72, wherein one or several amino acids are deleted, substituted, inserted and/or added
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HB) includes, for example, a polypeptide consisting of the following (H-B) amino acid sequence.
- the light chain variable region of (LK) includes, for example, a polypeptide having the amino acid sequence of (LK) below.
- (HB) Amino acid sequence of the following (H-B1), (H-B2) or (H-B3) (H-B1) Amino acid sequence of SEQ ID NO: 24 SEQ ID NO: 24: EVQLLESGGGLVQPGGSLRLSCAAS GFTFSTYQ MSWVRQAPGKGLEWV IV IVSSGGST AYADSVKGRFTISRDNSKNTLYLQVNTVLAGT (H-B2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 24 (H-B3) In the amino acid sequence of SEQ ID NO: 24, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- (LK) Amino acid sequence of the following (L-K1), (L-K2) or (L-K3) (L-K1) Amino acid sequence of SEQ ID NO: 73 SEQ ID NO: 73: QSALTQPPSASGSPGQSVTISCTGT SSDVGGYDF VSWYQQHPGEAPKLLVY DVN NRPSGVSNRFSGSKSGNTASLTISGLQAEDEGDYYC SSYAGSNSVL (L-K2) Amino acid sequence having 80% or more identity with the amino acid sequence of SEQ ID NO: 73 (L-K3) In the amino acid sequence of SEQ ID NO: 73, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (H-B1) amino acid sequence is, for example, a sequence including the (H1-B1) of CDRH1, (H2-B1) of CDRH2, and (H3-B1) of CDRH3.
- the amino acid sequence of (H-B2) includes, for example, the amino acid sequences of (H1-B1) of CDRH1, (H2-B1) of CDRH2, and CDRH3 (H3-B1), and the amino acid sequence of SEQ ID NO: 24.
- the amino acid sequence may have an identity of 80% or more.
- the amino acid sequence of (H-B3) includes, for example, (H1-B1) of CDRH1, (H2-B1) of CDRH2, and CDRH3 (H3-B1), and the amino acid sequence of SEQ ID NO: 24. In, it may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added.
- the above-mentioned (L-K1) amino acid sequence is, for example, a sequence including the CDL1 (L1-K1), CDRL2 (L2-K1), and CDRL3 (L3-K1) amino acid sequences.
- the amino acid sequence of (L-K2) includes, for example, (L1-K1) of CDRL1, (L2-K1) of CDRL2, and (L3-K1) of CDRL3, and the amino acid of SEQ ID NO: 73. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-K3) includes, for example, the amino acid sequences of (L1-K1) of CDRL1, (L2-K1) of CDRL2, and (L3-K1) of CDRL3, and the amino acid of SEQ ID NO: 73. It may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-B1) and the light chain variable region is the (L-K1). Also referred to as "3M4E5H-L1".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, respectively. It is 97% or more, 98% or more, and 99% or more.
- “1 or several” regarding substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, respectively.
- the numbers are 1 to 9, 1 to 8, 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the antibody or the like of the combination (12) is also referred to as, for example, antibody 3M4E5H-L66 group.
- the heavy chain variable region of (HB) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-B), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-B), and CDRH3 is a polypeptide containing the amino acid sequence of the above (H3-B).
- the light chain variable region of (LL) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-L) below, and CDRL2 is (L2-L) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-L) below.
- (L1-L) The following (L1-L1), (L1-L2) or (L1-L3) amino acid sequence (L1-L1) SEQ ID NO:74 (SSDVGGYEF) amino acid sequence (L1-L2) SEQ ID NO:74 amino acid Amino acid sequence having 80% or more identity to the sequence (L1-L3)
- amino acid sequence of SEQ ID NO: 74 one or several amino acids are deleted, substituted, inserted and/or added
- (L2-L) The following (L2-L1), (L2-L2) or (L2-L3) amino acid sequence (L2-L1) SEQ ID NO: 75 (DVI) amino acid sequence (L2-L2) SEQ ID NO: 75 amino acid Amino acid sequence (L2-L3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 75
- (L3-L) Amino acid sequence of the following (L3-L1), (L3-L2) or (L3-L3) (L3-L1) SEQ ID NO: 76 (SSYTSSSTYV) (L3-L2) Amino acid of SEQ ID NO: 76 Amino acid sequence having 80% or more identity to the sequence (L3-L3) SEQ ID NO: 76, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HB) includes, for example, a polypeptide consisting of the amino acid sequence of (H-B).
- the light chain variable region of (LL) includes, for example, a polypeptide having the amino acid sequence of (L-L) below.
- the above-mentioned (L-L1) amino acid sequence is, for example, a sequence including the CDL1 (L1-L1), CDRL2 (L2-L1), and CDRL3 (L3-L1) amino acid sequences.
- the amino acid sequence of (L-L2) includes, for example, the amino acid sequences of (L1-L1) of CDRL1, (L2-L1) of CDRL2, and (L3-L1) of CDRL3, and the amino acid of SEQ ID NO: 77. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-L3) includes, for example, the amino acid sequences of (L1-L1) of CDRL1, (L2-L1) of CDRL2, and (L3-L1) of CDRL3, and the amino acid of SEQ ID NO: 77. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-B1)
- the light chain variable region is the (L-L1)
- antibody of this combination hereinafter referred to as "antibody Also known as 3M4E5H-L66".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the antibody etc. of the combination (13) are also referred to as, for example, antibody 3M4E5H-L73 group.
- the heavy chain variable region of (HB) includes CDRH1, CDRH2, and CDRH3
- CDRH1 is a polypeptide containing the amino acid sequence of (H1-B)
- CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-B)
- CDRH3 is a polypeptide containing the amino acid sequence of (H3-B).
- the light chain variable region of (LM) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-M) below, and CDRL2 is (L2-M) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-M) below.
- (L1-M) Amino acid sequence of (L1-M1), (L1-M2) or (L1-M3) or (L1-M3) below (L1-M1), SEQ ID NO: 78 (GSDVGAYDY), (L1-M2) amino acid of SEQ ID NO: 78 Amino acid sequence (L1-M3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 78
- (L2-M) Amino acid sequence of the following (L2-M1), (L2-M2) or (L2-M3) (L2-M1) SEQ ID NO: 79 (DVS) amino acid sequence (L2-M2) SEQ ID NO: 79 Amino acid sequence (L2-M3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 79.
- (L3-M) Amino acid sequence (L3-M2), (L3-M2) or (L3-M2) or (L3-M3) amino acid sequence (L3-M1) SEQ ID NO: 80 (SSYSGSSTWV) (L3-M2) SEQ ID NO: 80 Amino acid sequence having 80% or more identity to the sequence (L3-M3) In the amino acid sequence of SEQ ID NO: 80, one or several amino acids have been deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HB) includes, for example, a polypeptide consisting of the amino acid sequence of (H-B).
- the light chain variable region of (LM) includes, for example, a polypeptide having the amino acid sequence of (LM) below.
- (LM) Amino acid sequence of (L-M1), (L-M2) or (L-M3) below (L-M1) Amino acid sequence of SEQ ID NO: 81
- SEQ ID NO: 81 QSALTQPASVSGSPGQSITISCTGT GSDVGAYDY VSWYQHHPGRAPRLIIR DVS VRPSGVPDRFSGSKSGNTASLTISGLQAEDEADVLC SSYSGSST
- L-M2 Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 81 (L-M3)
- amino acid sequence of SEQ ID NO: 81 one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-M1) amino acid sequence is, for example, a sequence including the CDL1 (L1-M1), CDRL2 (L2-M1), and CDRL3 (L3-M1) amino acid sequences.
- the amino acid sequence of (L-M2) includes, for example, the amino acid sequences of (L1-M1) of CDRL1, (L2-M1) of CDRL2, and (L3-M1) of CDRL3, and the amino acid of SEQ ID NO: 81. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-M3) includes, for example, the amino acid sequences of (L1-M1) of CDRL1, (L2-M1) of CDRL2, and (L3-M1) of CDRL3, and the amino acid of SEQ ID NO: 81. It may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-B1)
- the light chain variable region is the (L-M1)
- the antibody of this combination is referred to as "antibody Also known as 3M4E5H-L73".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the antibody etc. of the combination (14) are also referred to as, for example, antibody 3M4E5H-L80 group.
- the heavy chain variable region of (HB) includes CDRH1, CDRH2, and CDRH3
- CDRH1 is a polypeptide containing the amino acid sequence of (H1-B)
- CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-B)
- CDRH3 is a polypeptide containing the amino acid sequence of the above (H3-B).
- the light chain variable region of (LN) includes CDRL1, CDRL2, and CDRL3, wherein CDRL1 is a polypeptide containing the amino acid sequence of (L1-N) below, and CDRL2 is (L2-N) below.
- CDRL3 is a polypeptide containing the amino acid sequence of (L3-N) below.
- (L1-N) The following (L1-N1), (L1-N2) or (L1-N3) amino acid sequence (L1-N1) SEQ ID NO: 82 (SSDVGSYNL) amino acid sequence (L1-N2) SEQ ID NO: 82 amino acid Amino acid sequence having 80% or more identity to the sequence (L1-N3) In the amino acid sequence of SEQ ID NO: 82, one or several amino acids have been deleted, substituted, inserted and/or added
- L2-N The following (L2-N1), (L2-N2) or (L2-N3) amino acid sequence (L2-N1) SEQ ID NO:83 (DVS) amino acid sequence (L2-N2) SEQ ID NO:83 amino acid Amino acid sequence having 80% or more identity with the sequence (L2-N3) SEQ ID NO: 83, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L3-N) Amino acid sequence of the following (L3-N1), (L3-N2) or (L3-N3) (L3-N1) SEQ ID NO: 84 (SSYTSSSTFAV) (L3-N2) SEQ ID NO: 84 Amino acid sequence having 80% or more identity to the sequence (L3-N3) SEQ ID NO: 84, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HB) includes, for example, a polypeptide consisting of the amino acid sequence of (H-B).
- the light chain variable region of (LN) includes, for example, a polypeptide having the amino acid sequence of (L-N) below.
- (LN) Amino acid sequence of the following (L-N1), (L-N2) or (L-N3) (L-N1) Amino acid sequence of SEQ ID NO:85 SEQ ID NO:85: QSALTQPASVSGSPGQSITISCTGT SSDVGSYNL VSWYQQHPGKAPKLMIY DVS NRPSGVSYRFSGSKSGNTASLTISGLQAEDETFYYF SGT (L-N2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 85 (L-N3) In the amino acid sequence of SEQ ID NO: 85, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-N1) amino acid sequence is, for example, a sequence including the CDL1 (L1-N1), CDRL2 (L2-N1), and CDRL3 (L3-N1) amino acid sequences.
- the amino acid sequence of (L-N2) includes, for example, the amino acid sequences of (L1-N1) of CDRL1, (L2-N1) of CDRL2, and (L3-N1) of CDRL3, and the amino acid of SEQ ID NO:85. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-N3) includes, for example, the amino acid sequences of (L1-N1) of CDRL1, (L2-N1) of CDRL2, and (L3-N1) of CDRL3, and the amino acid of SEQ ID NO:85. It may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is (H-B1)
- the light chain variable region is (L-N1)
- the antibody of this combination is referred to as "antibody Also called 3M4E5H-L80”.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the antibody etc. of the combination (15) are also referred to as, for example, antibody 3M4E5H-L88 group.
- the heavy chain variable region of (HB) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-B), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-B), and CDRH3 is a polypeptide containing the amino acid sequence of the above (H3-B).
- the (LO) light chain variable region includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-O) below, and CDRL2 is (L2-O) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-O) below.
- (L1-O) Amino acid sequence of (L1-O1), (L1-O2) or (L1-O3) below (L1-O1) SEQ ID NO: 86 (SSDVGGYNY) (L1-O2) Amino acid of SEQ ID NO: 86 Amino acid sequence having 80% or more identity to the sequence (L1-O3) SEQ ID NO: 86 amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added
- (L3-O) Amino acid sequence of the following (L3-O1), (L3-O2) or (L3-O3) (L3-O1) SEQ ID NO: 88 (CSYAGGYYV) (L3-O2) Amino acid of SEQ ID NO: 88 Amino acid sequence having 80% or more identity to the sequence (L3-O3) SEQ ID NO: 88, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HB) includes, for example, a polypeptide consisting of the amino acid sequence of (H-B).
- the light chain variable region of (LO) includes, for example, a polypeptide having the amino acid sequence of (LO) below.
- the above-mentioned (L-O1) amino acid sequence is, for example, a sequence including the CDL1 (L1-O1), CDRL2 (L2-O1), and CDRL3 (L3-O1) amino acid sequences.
- the amino acid sequence of (L-O2) includes, for example, the amino acid sequences of (L1-O1) of CDRL1, (L2-O1) of CDRL2, and (L3-O1) of CDRL3, and the amino acid of SEQ ID NO: 89. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-O3) includes, for example, the amino acid sequences of (L1-O1) of CDRL1, (L2-O1) of CDRL2, and (L3-O1) of CDRL3, and the amino acid of SEQ ID NO: 89. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-B1) and the light chain variable region is the (L-O1). Also known as 3M4E5H-L88".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the antibody etc. of the combination (16) are also referred to as, for example, antibody 3M4E5H-L102 group.
- the heavy chain variable region of (HB) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-B), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-B), and CDRH3 is a polypeptide containing the amino acid sequence of the above (H3-B).
- the light chain variable region of (LP) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-P) below, and CDRL2 is (L2-P) below.
- CDRL3 is a polypeptide containing the amino acid sequence of (L3-P) below.
- (L1-P) Amino acid sequence of (L1-P1), (L1-P2) or (L1-P3) below (L1-P1) SEQ ID NO: 90 (SSDVGGYNY) (L1-P2) Amino acid of SEQ ID NO: 90 Amino acid sequence (L1-P3) having an identity of 80% or more with respect to the sequence, in which one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 90.
- (L2-P) Amino acid sequence of the following (L2-P1), (L2-P2) or (L2-P3) (L2-P1) SEQ ID NO: 91 (DVS) amino acid sequence (L2-P2) SEQ ID NO: 91 Amino acid sequence having 80% or more identity to the sequence (L2-P3), amino acid sequence of SEQ ID NO: 91 with one or several amino acids deleted, substituted, inserted and/or added
- (L3-P) Amino acid sequence of (L3-P1), (L3-P2) or (L3-P3) below (L3-P1) SEQ ID NO: 92 (SSYAGSGSTPFV) (L3-P2) Amino acid of SEQ ID NO: 92 Amino acid sequence (L3-P3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 92.
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HB) includes, for example, a polypeptide consisting of the amino acid sequence of (H-B).
- the light chain variable region of (LP) includes, for example, a polypeptide having the amino acid sequence of (LP) below.
- the above-mentioned (L-P1) amino acid sequence is, for example, a sequence including the CDL1 (L1-P1), CDRL2 (L2-P1), and CDRL3 (L3-P1) amino acid sequences.
- the amino acid sequence of (L-P2) includes, for example, the amino acid sequences of (L1-P1) of CDRL1, (L2-P1) of CDRL2, and (L3-P1) of CDRL3, and the amino acid of SEQ ID NO: 93. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-P3) includes, for example, the amino acid sequences of (L1-P1) of CDRL1, (L2-P1) of CDRL2, and (L3-P1) of CDRL3, and the amino acid of SEQ ID NO: 93. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-B1) and the light chain variable region is the (L-P1). Also known as 3M4E5H-L102".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the antibody etc. of the combination (17) are also referred to as, for example, antibody 3M4E5H-L124 group.
- the heavy chain variable region of (HB) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-B), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-B), and CDRH3 is a polypeptide containing the amino acid sequence of the above (H3-B).
- the light chain variable region of (LQ) includes CDRL1, CDRL2, and CDRL3, wherein CDRL1 is a polypeptide containing the amino acid sequence of (L1-Q) below, and CDRL2 is (L2-Q) below.
- CDRL3 is a polypeptide containing the amino acid sequence of (L3-Q) below.
- (L1-Q) Amino acid sequence (L1-Q1), (L1-Q2) or (L1-Q2) or (L1-Q3) amino acid sequence (L1-Q1) SEQ ID NO: 94 (SSDVGGYNY) (L1-Q2) SEQ ID NO: 94 Amino acid sequence having 80% or more identity to the sequence (L1-Q3) SEQ ID NO: 94, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L3-Q) Amino acid sequence of the following (L3-Q1), (L3-Q2) or (L3-Q3) (L3-Q1) SEQ ID NO: 96 (CSYAGRRYV) (L3-Q2) Amino acid of SEQ ID NO: 96 Amino acid sequence having 80% or more identity to the sequence (L3-Q3) SEQ ID NO: 96, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HB) includes, for example, a polypeptide consisting of the amino acid sequence of (H-B).
- the light chain variable region of (LQ) includes, for example, a polypeptide having the amino acid sequence of (LQ) below.
- the (L-Q1) amino acid sequence is, for example, a sequence including the CDL1 (L1-Q1), the CDRL2 (L2-Q1), and the CDRL3 (L3-Q1) amino acid sequences.
- the amino acid sequence of (L-Q2) includes, for example, the amino acid sequences of (L1-Q1) of CDRL1, (L2-Q1) of CDRL2, and (L3-Q1) of CDRL3, and the amino acid of SEQ ID NO: 97. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-Q3) includes, for example, the amino acid sequences of (L1-Q1) of CDRL1, (L2-Q1) of CDRL2, and (L3-Q1) of CDRL3, and the amino acid of SEQ ID NO: 97. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is (H-B1)
- the light chain variable region is (L-Q1)
- antibody of this combination is referred to as "antibody Also known as 3M4E5H-L124".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HC) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-C) below, and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-C), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-C) below.
- the light chain variable region of (LA) includes CDRL1, CDRL2, and CDRL3, wherein CDRL1 is a polypeptide containing the amino acid sequence of (L1-A), and CDRL2 is (L2-A). Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-A) above.
- H1-C Amino acid sequence of (H1-C1), (H1-C2) or (H1-C3) below (H1-C1) SEQ ID NO:25 (GGSISSYY) amino acid sequence (H1-C2) SEQ ID NO:25 amino acid Amino acid sequence having 80% or more identity to the sequence (H1-C3)
- amino acid sequence of SEQ ID NO: 25 one or several amino acids are deleted, substituted, inserted and/or added
- H2-C Amino acid sequence of the following (H2-C1), (H2-C2) or (H2-C3) (H2-C1) SEQ ID NO:26 (INHSGST) (H2-C2) Amino acid of SEQ ID NO:26 Amino acid sequence having 80% or more identity to the sequence (H2-C3) In the amino acid sequence of SEQ ID NO: 26, one or several amino acids have been deleted, substituted, inserted and/or added
- H3-C Amino acid sequence of (H3-C1), (H3-C2) or (H3-C3) below (H3-C1) SEQ ID NO: 27 (ARCPIYYYGMDV) (H3-C2) Amino acid of SEQ ID NO: 27 Amino acid sequence having 80% or more identity to the sequence (H3-C3) In the amino acid sequence of SEQ ID NO: 27, an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HC) includes, for example, a polypeptide consisting of the following (H-C) amino acid sequence.
- the light chain variable region of (LA) includes, for example, a polypeptide having the amino acid sequence of (LA).
- (HC) Amino acid sequence of the following (H-C1), (H-C2) or (H-C3) (H-C1) Amino acid sequence of SEQ ID NO: 28 SEQ ID NO: 28: QLQLQESGPGLVKPSQTLSLTCTVS GGSISSYY WSWIRQPPGKGLEWIGE INHSGST NYNPSLKSRVTISV DTSKNQFSLKLSSVTPIDTTVYYC (H-C2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 28 (H-C3) In the amino acid sequence of SEQ ID NO: 28, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (H-C1) amino acid sequence is, for example, a sequence containing the (H1-C1) of CDRH1, (H2-C1) of CDRH2, and (H3-C1) of CDRH3.
- the amino acid sequence of (H-C2) includes, for example, (H1-C1) of CDRH1, (H2-C1) of CDRH2, and CDRH3 (H3-C1), and the amino acid sequence of SEQ ID NO: 28.
- the amino acid sequence may have an identity of 80% or more.
- the amino acid sequence of (H-C3) includes, for example, the amino acid sequences of (H1-C1) of CDRH1, (H2-C1) of CDRH2, and CDRH3 (H3-C1), and the amino acid sequence of SEQ ID NO: 28. In, it may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added.
- the heavy chain variable region is the (H-C1) and the light chain variable region is the (L-A1). Also called H73-3M4E5L".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like is, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- amino acid sequences of SEQ ID NOs: 17 to 97 are, for example, human-derived amino acid sequences.
- the binding between the antibody etc. of the present invention and A2/NY-ESO-1 157 can be confirmed by a method for detecting the binding between the antibody and the antigen, such as surface plasmon resonance (SPR) or flow cytometry.
- SPR surface plasmon resonance
- the antibody etc. of the present invention may further have a labeling substance, for example.
- the labeling substance is not particularly limited, and examples thereof include a fluorescent substance, a dye, an isotope, an enzyme and the like.
- the fluorescent substance include pyrene, TAMRA, fluorescein, Cy3 dye, Cy5 dye, FAM dye, rhodamine dye, Texas red dye, JOE, MAX, HEX, TYE, and other fluorophores.
- Alexa dyes such as Alexa488 and Alexa647.
- the antibody etc. of the present invention may be modified with a low molecular weight compound such as a pharmaceutical compound.
- the low molecular weight compound include antitumor agents such as tubulin inhibitors.
- the antibody or the like of the present invention is, for example, directly or indirectly modified with the low molecular weight compound. In the latter case, the antibody or the like of the present invention is modified with the low molecular weight compound, for example, via a linker.
- the antibody or the like of the present invention may be immobilized on, for example, a carrier or a porous body.
- the carrier is not particularly limited, and examples thereof include a substrate, beads, and a container, and examples of the container include a microplate, a tube, and the like.
- the method for producing the antibody or the like of the present invention is not particularly limited, and for example, it can be produced by genetic engineering based on the aforementioned amino acid sequence information. Specifically, for example, it can be performed as follows.
- the present invention is not limited to this example.
- a vector containing a nucleic acid sequence encoding the amino acid sequence of each of the regions, the heavy chain, and/or the light chain of the antibody of the present invention is introduced into a host to obtain a transformant. Then, the transformant is cultured, a fraction containing an antibody that binds to A2/NY-ESO-1 157 is collected, and the antibody is isolated or purified from the obtained collected fraction.
- the vector examples include a vector containing a nucleic acid sequence encoding the heavy chain variable region, a vector containing a nucleic acid sequence encoding the light chain variable region, a vector containing a nucleic acid sequence encoding the heavy chain, the light chain Examples thereof include a vector containing a nucleic acid sequence encoding
- the host is not particularly limited as long as it can introduce the vector and can express the nucleic acid sequence in the vector.
- Examples of the host include mammalian cells such as HEK cells, CHO cells, COS cells, NSO cells and SP2/0 cells.
- the method of introducing the vector into the host is not particularly limited, and known methods can be adopted.
- the method for culturing the transformant is not particularly limited and can be appropriately determined according to the type of the host.
- the fraction containing the antibody can be collected as a liquid fraction by crushing the cultured transformant. Isolation or purification of the antibody is not particularly limited, and a known method can be adopted.
- the antibody is, for example, a monoclonal antibody.
- the monoclonal antibody include a monoclonal antibody obtained by immunizing an animal, a chimeric antibody, a humanized antibody, a human antibody (also referred to as a fully human antibody), and the like.
- the chimeric antibody is an antibody in which the variable region of an antibody derived from a non-human animal is linked to the constant region of a human antibody.
- the chimeric antibody can be produced, for example, as follows. First, for a monoclonal antibody derived from a non-human animal, a variable region (V region) gene that binds to the A2/NY-ESO-1 157 protein was prepared, and the variable region gene and the human antibody constant region (C region) were prepared. Region), and this is further ligated to an expression vector. Then, the cells transfected with the expression vector are cultured to recover the desired chimeric antibody secreted into the culture medium. Thereby, a chimeric antibody can be prepared.
- V region variable region
- C region human antibody constant region
- the animal from which the variable region gene is derived is not particularly limited, and examples thereof include rats and mice.
- the method for producing the chimeric antibody is not limited to this, and can be produced by referring to a known method such as the method described in JP-B-3-73280.
- the humanized antibody is an antibody in which only the CDR is derived from an animal other than human and the other regions are derived from human.
- the humanized antibody can be produced, for example, as follows. First, with respect to a monoclonal antibody derived from a non-human animal, the CDR gene is prepared, transplanted to a human antibody gene, for example, a constant region (CDR grafting), and further ligated to an expression vector. Then, by culturing the cells transfected with the expression vector, the humanized antibody transplanted with the CDR of interest is secreted into the culture medium. It can be prepared by recovering the secreted humanized antibody.
- the CDR-derived animal is not particularly limited, and examples thereof include a rat and a mouse.
- the method for producing the humanized antibody is not limited to this, and is produced by referring to known methods such as those described in Japanese Patent Publication No. 4-506458 and Japanese Patent Application Laid-Open No. 62-296890. it can.
- the human antibody can be produced, for example, by introducing a human antibody gene into an animal other than human.
- a transgenic animal for producing a human antibody can be used as the animal into which the human antibody gene is introduced.
- the type of animal is not particularly limited, and examples thereof include mice.
- the method for producing the human antibody is described in, for example, Nature Genetics, Vol. 7, pp. 13-21, 1994; Nature Genetics, Vol. 15, pp. 146-156, 1994; Manufactured by referring to a known method described in Table 7-509137; WO94/25585; Nature, Vol.368, p.856-859,1994; and Tokuhei 6-500233. it can.
- human antibody can also be produced, for example, by using the phage display method, for example, Marks, JJ.D. et al.: J. Mol. Biol., Vol.222, p.581-597, It can be produced by referring to a known method described in 1991.
- the antibody or the like of the present invention can also be prepared, for example, by immunizing an animal with an antigen.
- the antigen include A2/NY-ESO-1 157 protein.
- the antigen is preferably immunized multiple times.
- the peptide fragment may be, for example, a peptide fragment consisting of only an antigenic determinant (epitope) or a peptide fragment containing the antigenic determinant.
- Monoclonal antibodies obtained by immunizing the animals include, for example, ⁇ Current Protocols in Molecular Biology'' (John Wiley & Sons (1987)), Antibodies: A Laboratory Manual, Ed. )) and the like, and the like can be manufactured by referring to known methods. Specifically, for example, an animal is immunized with an antigen, and antibody-producing cells collected from the immunized animal are fused with myeloma cells (myeloma cells) lacking autoantibody-producing ability to prepare hybridomas. Subsequently, the hybridoma is screened for antibody-producing cells, and a single clone of the hybridoma is prepared by cloning.
- this hybridoma clone is administered to an animal, and a monoclonal antibody is purified from the obtained abdominal cavity.
- the hybridoma is cultured, and the monoclonal antibody is purified from the culture solution.
- a monoclonal antibody with uniform specificity can be stably supplied.
- the above-mentioned myeloma cells are preferably derived from, for example, mouse, rat, human or the like.
- the myeloma cells and the antibody-producing cells may be derived from the same species or different species, but are preferably the same species.
- the (H) heavy chain variable region and the (L) light chain variable region are the following (H) heavy chain variable region and (L) light chain variable region, respectively. May be.
- (H) comprises heavy chain complementarity determining region (CDRH) 1, CDRH2, and CDRH3,
- CDRH1 is a polypeptide containing the following amino acid sequence (H1)
- CDRH2 is a polypeptide containing the following amino acid sequence (H2)
- CDRH3 is a heavy chain variable region that is a polypeptide containing the following amino acid sequence (H3)
- H1 Amino acid sequence of (H1-1), (H1-2) or (H1-3) below (H1-1) Any amino acid sequence of CDRH1 under the following condition (H1) (H1-2) (H1- In the amino acid sequence (H1-3) (H1-1) having 80% or more identity with the amino acid sequence of 1), one or several amino acids are deleted, substituted, inserted and/or Or added amino acid sequence;
- H2 Amino acid sequence of (H2-1), (H2-2) or (H2-3) below (H2-1)
- (L) comprises light chain complementarity determining regions (CDRL) 1, CDRL2, and CDRL3,
- CDRL1 is a polypeptide containing the following amino acid sequence (L1)
- CDRL2 is a polypeptide containing the following amino acid sequence (L2)
- CDRL3 is a light chain variable region that is a polypeptide comprising the amino acid sequence of (L3) below:
- Examples of the heavy chain variable region that satisfies the above condition (H1) include the heavy chain variable regions of (HA), (HB), and (HC) above.
- X 7 is preferably absent, T or Y.
- X 4 , N, D or E is preferable.
- X 5 , Y, F or L is preferable.
- X 7 is preferably F or Y.
- Examples of the light chain variable region that satisfies the above condition (L1) include the light chain variable regions (LB) to (LJ) above.
- Examples of the light chain variable region that satisfies the above condition (L2) include the light chain variable regions (LA) and (LK) to (LQ) above.
- CDRH1 of the heavy chain variable region of (H) is GFTFSTYQ (SEQ ID NO: 21), and CDRH2 is IVSSGGST (SEQ ID NO: 22).
- CDRH3 is AGELLPYYGMDV (SEQ ID NO:23)
- CDRL1 of the light chain variable region of (L) above is SRDVGGGYNY (SEQ ID NO:29)
- CDRL2 is DVI (SEQ ID NO:30)
- CDRL3 is WSFAGSYYV. It is preferably an amino acid sequence other than (SEQ ID NO: 31).
- the antibody or antigen-binding fragment thereof against CD19 of the present invention includes the heavy chain variable region of (H) below and the light chain variable region of (L) below.
- the antibody or the like of the present invention is characterized by containing the heavy chain variable region of (H) below and the light chain variable region of (L) below, and other configurations and conditions are not particularly limited.
- the antibody or the like of the present invention binds to CD19. Further, CD19 is known to be expressed in specific cancer cells such as normal B cells and B cell lymphomas.
- the antibody and the like of the present invention can be preferably used as, for example, a bispecific antibody against a CD19-expressing cancer cell, an antigen-binding domain of CAR of CAR-T cell, and the like.
- the description of the CAR library of the present invention, the first screening method, the first antibody or the antigen-binding fragment thereof and the like can be incorporated.
- (H) comprises heavy chain complementarity determining region (CDRH) 1, CDRH2, and CDRH3,
- CDRH1 is a polypeptide containing the following amino acid sequence (H1)
- CDRH2 is a polypeptide containing the following amino acid sequence (H2)
- CDRH3 is a polypeptide containing the amino acid sequence of (H3) below
- one or several amino acids are deleted, substituted, inserted and/or added (H2)
- amino acid sequence of one or several amino acids have been deleted, substituted, inserted and/or added (H3) of the following (H3-1), (H3-2) or (H3-3) Amino acid sequence (H3-1)
- amino acid sequence of (H3-1) an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added
- (L) comprises light chain complementarity determining regions (CDRL) 1, CDRL2, and CDRL3,
- CDRL1 is a polypeptide containing the following amino acid sequence (L1)
- CDRL2 is a polypeptide containing the following amino acid sequence (L2)
- CDRL3 is a polypeptide containing the following amino acid sequence (L3) light chain variable region (L1) below (L1-1), (L1-2) or (L1-3) amino acid sequence (L1-1) below Amino acid sequence (L1-3) (L1-1) having 80% or more identity to the amino acid sequence (L1-2) (L1-1) of any of CDRL1s in Table 3B.
- one or several amino acids are deleted, substituted, inserted and/or added (L2) The following (L2-1), (L2-2) or (L2-3) amino acid sequence (L2 -1) An amino acid sequence (L2-3) (L2-1) having at least 80% identity with the amino acid sequence (L2-2) (L2-1) of any of CDRL2s in Table 3B below.
- amino acid sequence of (1) one or several amino acids are deleted, substituted, inserted and/or added (L3) of the following (L3-1), (L3-2) or (L3-3) Amino acid sequence (L3-1) An amino acid sequence (L3-3) having 80% or more identity to the amino acid sequence of any of the CDRL3s (L3-2) (L3-1) in Table 3B below.
- CD19 is, for example, a type I transmembrane glycoprotein having a molecular weight of 95 kDa.
- CD19 is known to be involved in, for example, B cell development, activation, regulation of differentiation, etc., and is mainly expressed in B cells.
- the amino acid sequence of human CD19 includes, for example, the amino acid sequence registered under NCBI accession number NM_001770.6.
- the amino acid sequence of mouse CD19 includes, for example, the amino acid sequence registered under NCBI accession number NM_009844.2.
- the CDRH1 is the CDRH1 in the (HA).
- the CDRH2 is the CDRH2 in the (HA).
- the CDRH3 is the CDRH3 in the above (HA).
- the CDRL1 is any one of (LA) to (LL) and (LM).
- the CDRL2 is any one of the above (LA) to (LL) and (LM).
- the CDRL3 is any one of the above (LA) to (LL) and (LM).
- the CDRL1, CDRL2, and CDRL3 may satisfy the same condition or different conditions in the above (LA) to (LL) and (LM), but preferably satisfy the same condition. .. When the CDRL1, CDRL2, and CDRL3 satisfy the same condition, the CDRL1, CDRL2, and CDRL3 satisfy any one of the (LA) to (LL) and (LM), CDRL1, CDRL2, and CDRL3.
- identity is, for example, the degree of identity when the sequences to be compared are properly aligned with each other, and means the occurrence rate (%) of exact matches of amino acids between the sequences.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- the identity can be calculated with default parameters using analysis software such as BLAST and FASTA (the same applies hereinafter).
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the substitution of the amino acid may be, for example, a conservative substitution (hereinafter the same).
- the conservative substitution means substituting one or several amino acids with other amino acids and/or amino acid derivatives so as not to substantially change the function of the protein.
- the “substituted amino acid” and the “substituted amino acid” have similar properties and/or functions, for example.
- the hydrophobic and hydrophilic indices (hydropathy), chemical properties such as polarity and charge, or physical properties such as secondary structure are similar.
- Amino acids or amino acid derivatives having similar properties and/or functions are known in the art, for example.
- non-polar amino acids such as alanine, valine, isoleucine, leucine, proline, tryptophan, phenylalanine and methionine
- polar amino acids neutral amino acids
- neutral amino acids include glycine, serine and threonine.
- positively charged amino acids include arginine, histidine, lysine, etc.
- negatively charged amino acids include aspartic acid, glutamic acid.
- the combination of (L1-1), (L2-1) and (L3-1) is not particularly limited, and examples thereof include (LA) to (LL), respectively. And any one CDRL1 in (LM), any one CDRL2 in (LA) to (LL) and (LM) above, and any one CDRL3 in (LA) to (LL) and (LM) above. Can be any combination.
- the combination of (L1-1), (L2-1) and (L3-1) is preferably any one of CDRL1, CDRL2 and CDRL3 in (LA) to (LL) and (LM).
- the antibody of the present invention and the like will be described more specifically with respect to the combination of the heavy chain variable region or heavy chain and the light chain variable region or light chain.
- the description of the heavy chain variable region and the heavy chain in each combination can be incorporated into each other. Further, the description of the light chain variable region and the light chain in each combination can be incorporated into each other.
- the underlined amino acid sequences and base sequences are the base sequences encoding the amino acid sequences corresponding to CDRs and the amino acid sequences corresponding to CDRs, unless otherwise specified.
- the combination of the heavy chain variable region and the light chain variable region is, for example, any of the above (HA) and (LA) to (LL) and (LM) as described above. Or one combination.
- the combination (LA) antibody and the like are also referred to as, for example, antibody 18H-L4 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A) below, and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A) below.
- the light chain variable region of (LA) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-A) below, and CDRL2 is (L2-A) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-A) below.
- H1-A Amino acid sequence of (H1-A1), (H1-A2) or (H1-A3) below (H1-A1) SEQ ID NO:216 (GFTFDDYA) (H1-A2) Amino acid of SEQ ID NO:216 Amino acid sequence having 80% or more identity to the sequence (H1-A3)
- amino acid sequence of SEQ ID NO: 216 one or several amino acids are deleted, substituted, inserted and/or added
- H2-A Amino acid sequence of the following (H2-A1), (H2-A2) or (H2-A3) (H2-A1) SEQ ID NO:217 (ISWNSGRI) (H2-A2) Amino acid of SEQ ID NO:217 Amino acid sequence having 80% or more identity with the sequence (H2-A3), wherein the amino acid sequence of SEQ ID NO: 217 has one or several amino acids deleted, substituted, inserted and/or added
- H3-A Amino acid sequence of (H3-A1), (H3-A2) or (H3-A3) (H3-A1) SEQ ID NO:218 (ARDQGYHYYDSAEHAFDI) (H3-A2) Amino acid of SEQ ID NO:218 Amino acid sequence having 80% or more identity to the sequence (H3-A3) SEQ ID NO: 218, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L1-A) Amino acid sequence of the following (L1-A1), (L1-A2) or (L1-A3) (L1-A1) SEQ ID NO: 220 (KLGDKY) (L1-A2) Amino acid of SEQ ID NO: 220 Amino acid sequence having 80% or more identity to the sequence (L1-A3) SEQ ID NO: 220, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L2-A) The following (L2-A1), (L2-A2) or (L2-A3) amino acid sequence (L2-A1) SEQ ID NO:221 (QDS) amino acid sequence (L2-A2) SEQ ID NO:221 amino acid Amino acid sequence having 80% or more identity to the sequence (L2-A3), amino acid sequence of SEQ ID NO: 221 in which one or several amino acids are deleted, substituted, inserted and/or added
- (L3-A) Amino acid sequence of (L3-A1), (L3-A2) or (L3-A3) below (L3-A1) SEQ ID NO: 222 (QAWDSSTHVV) (L3-A2) SEQ ID NO: 222 Amino acid sequence having 80% or more identity to the sequence (L3-A3) SEQ ID NO: 222, wherein one or several amino acids are deleted, substituted, inserted and/or added
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide having the amino acid sequence of (H-A) below.
- the light chain variable region of (LA) includes, for example, a polypeptide having the amino acid sequence of (LA) below.
- H-A1 following (H-A1), (H -A2) or (H-A3) of the amino acid sequence (H-A1) of SEQ ID NO: 219 amino acid sequence SEQ ID NO 219: EVQLVESGGGLVQPGRSLRLSCAAS GFTFDDYA MHWVRQAPGKGLEWVSG ISWNSGRI GYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYC ARDQGYHYYDSAEHAFDI WGQGTVVTVSS (H-A2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 219 (H-A3) In the amino acid sequence of SEQ ID NO: 219, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- LA the following (L-A1), (L -A2) or (L-A3) of the amino acid sequence (L-A1) of SEQ ID NO: 223 amino acid sequence SEQ ID NO 223: SYELTQPPSVSVSPGQTASITCSGD KLGDKY ACWYQQKPGQSPVLVIY QDS KRPSGIPERFSGSNSGNTATLTISGTQAMDEADYYC QAWDSSTHVV FGGGTKLTVL (L-A2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 223 (L-A3) In the amino acid sequence of SEQ ID NO: 223, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (H-A1) amino acid sequence is, for example, a sequence containing the (H1-A1) of CDRH1, (H2-A1) of CDRH2, and (H3-A1) of CDRH3.
- the amino acid sequence of (H-A2) includes, for example, (H1-A1) of CDRH1, (H2-A1) of CDRH2, and CDRH3 (H3-A1), and the amino acid sequence of SEQ ID NO: 219.
- the amino acid sequence may have an identity of 80% or more.
- the amino acid sequence of (H-A3) includes, for example, (H1-A1) of CDRH1, (H2-A1) of CDRH2, and CDRH3 (H3-A1), and the amino acid sequence of SEQ ID NO: 219. In, it may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added.
- the above-mentioned (L-A1) amino acid sequence is, for example, a sequence including the CDL1 (L1-A1), CDRL2 (L2-A1), and CDRL3 (L3-A1) amino acid sequences.
- the amino acid sequence of (L-A2) includes, for example, the amino acid sequences of (L1-A1) of CDRL1, (L2-A1) of CDRL2, and (L3-A1) of CDRL3, and the amino acid sequence of SEQ ID NO: 223. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-A3) includes, for example, the amino acid sequences of (L1-A1) of CDRL1, (L2-A1) of CDRL2, and (L3-A1) of CDRL3, and the amino acid of SEQ ID NO: 223. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-A1). Also known as 18H-L4".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, respectively. It is 97% or more, 98% or more, and 99% or more.
- “1 or several” regarding substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, respectively.
- the numbers are 1 to 9, 1 to 8, 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LB) antibody and the like are also referred to as, for example, antibody 18H-L7 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LB) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-B) below, and CDRL2 is (L2-B) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-B) below.
- (L1-B) Amino acid sequence of (L1-B1), (L1-B2) or (L1-B3) below (L1-B1) SEQ ID NO: 224 (SSDVGGYNY) (L1-B2) SEQ ID NO: 224 Amino acid sequence having 80% or more identity to the sequence (L1-B3) SEQ ID NO: 224, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L3-B) Amino acid sequence of (L3-B1), (L3-B2) or (L3-B3) below (L3-B1) SEQ ID NO: 226 (GTWDTSLTAVV) (L3-B2) SEQ ID NO: 226 Amino acid sequence having 80% or more identity to the sequence (L3-B3) SEQ ID NO: 226, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LB) includes, for example, a polypeptide having the amino acid sequence of (LB) below.
- (LB) Amino acid sequence of (L-B1), (L-B2) or (L-B3) below (L-B1) Amino acid sequence of SEQ ID NO: 227 SEQ ID NO: 227: QSALTQPRSVSGSPGQSVTISCTGT SSDVGGYNY VSWYQQHPGKAPKLMIY DVS NRPSGVSNRFSGSKSGNTASLTISGLQTGTELAVV GTWDD (L-B2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 227 (L-B3) In the amino acid sequence of SEQ ID NO: 227, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-B1) amino acid sequence is, for example, a sequence including the CDL1 (L1-B1), CDRL2 (L2-B1), and CDRL3 (L3-B1) amino acid sequences.
- the amino acid sequence of (L-B2) includes, for example, the amino acid sequences of (L1-B1) of CDRL1, (L2-B1) of CDRL2, and (L3-B1) of CDRL3, and the amino acid of SEQ ID NO: 227. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-B3) includes, for example, (L1-B1) of CDRL1, (L2-B1) of CDRL2, and (L3-B1) of CDRL3, and the amino acid sequence of SEQ ID NO: 227. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-B1)
- antibody of this combination hereinafter referred to as "antibody Also known as 18H-L7".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LC) antibody and the like are also referred to as, for example, antibody 18H-L9 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LC) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-C) below, and CDRL2 is (L2-C) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-C) below.
- (L1-C) The following (L1-C1), (L1-C2) or (L1-C3) amino acid sequence (L1-C1) SEQ ID NO:228 (WSNIGDDH) amino acid sequence (L1-C2) SEQ ID NO:228 amino acid Amino acid sequence having 80% or more identity to the sequence (L1-C3), amino acid sequence of SEQ ID NO: 228 in which one or several amino acids are deleted, substituted, inserted and/or added
- (L2-C) The following (L2-C1), (L2-C2) or (L2-C3) amino acid sequence (L2-C1) SEQ ID NO: 229 (DTS) amino acid sequence (L2-C2) SEQ ID NO: 229 amino acid Amino acid sequence having 80% or more identity to the sequence (L2-C3), amino acid sequence of SEQ ID NO: 229 in which one or several amino acids are deleted, substituted, inserted and/or added
- (L3-C) Amino acid sequence of the following (L3-C1), (L3-C2) or (L3-C3) (L3-C1) SEQ ID NO: 230 (GTWESSLSGVV) (L3-C2) Amino acid of SEQ ID NO: 230 Amino acid sequence having an identity of 80% or more (L3-C3) SEQ ID NO: 230 with one or several amino acids deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LC) includes, for example, a polypeptide having the amino acid sequence of (LC) below.
- (LC) Amino acid sequence of (L-C1), (L-C2) or (L-C3) below (L-C1) Amino acid sequence of SEQ ID NO: 231 SEQ ID NO: 231: QSVLTQPPSVSAAPGQKVTISCSGS WSNIGDDH VSWYQQFPGAAPKLLIY DTS KRPSRVADRFSGSKSGASATLAITGLQAGDEADYV GTFGSSL (L-C2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO:231 (L-C3) In the amino acid sequence of SEQ ID NO:231, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-C1) amino acid sequence is, for example, a sequence including the CDL1 (L1-C1), CDRL2 (L2-C1), and CDRL3 (L3-C1) amino acid sequences.
- the amino acid sequence of (L-C2) includes, for example, (L1-C1) of CDRL1, (L2-C1) of CDRL2, and (L3-C1) of CDRL3, and the amino acid sequence of SEQ ID NO: 231. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-C3) includes, for example, the amino acid sequences of (L1-C1) of CDRL1, (L2-C1) of CDRL2, and (L3-C1) of CDRL3, and the amino acid of SEQ ID NO: 231. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-C1)
- antibody of this combination hereinafter referred to as "antibody Also known as 18H-L9".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LC) antibody and the like are also referred to as, for example, antibody 18H-L13 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LD) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-D) below, and CDRL2 is (L2-D) below.
- CDRL3 is a polypeptide containing the amino acid sequence of (L3-D) below.
- (L1-D) Amino acid sequence of the following (L1-D1), (L1-D2) or (L1-D3) (L1-D1) SEQ ID NO:232 (SSDVGGYDY) (L1-D2) SEQ ID NO:232 Amino acid sequence (L1-D3) having 80% or more identity to the sequence, in which one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO:232
- L2-D The following (L2-D1), (L2-D2) or (L2-D3) amino acid sequence (L2-D1) SEQ ID NO:233 (DVT) amino acid sequence (L2-D2) SEQ ID NO:233 amino acid sequence Amino acid sequence having 80% or more identity to the sequence (L2-D3), amino acid sequence of SEQ ID NO: 233 in which one or several amino acids are deleted, substituted, inserted and/or added
- (L3-D) Amino acid sequence of (L3-D1), (L3-D2) or (L3-D3) below (L3-D1) SEQ ID NO: 234 (SSYTTSTTWV) amino acid sequence (L3-D2) SEQ ID NO: 234 Amino acid sequence having 80% or more identity to the sequence (L3-D3) SEQ ID NO: 234, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LD) includes, for example, a polypeptide having the amino acid sequence of (LD) below.
- (LD) Amino acid sequence of (L-D1), (L-D2) or (L-D3) below (L-D1) Amino acid sequence of SEQ ID NO: 235 SEQ ID NO: 235: QSALTQPRSVSGSPGQSVTISCTGT SSDVGGYNY VSWYQQHPGKAPKLMIY DVS NRPSGVSNRFSGSKSGNTASLTISGLQTGTELADVL GTWDADAVYYC GTWD (L-D2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 235 (L-D3) In the amino acid sequence of SEQ ID NO: 235, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-D1) amino acid sequence is, for example, a sequence including the CDL1 (L1-D1), CDRL2 (L2-D1), and CDRL3 (L3-D1) amino acid sequences.
- the amino acid sequence of (L-D2) includes, for example, the amino acid sequences of (L1-D1) of CDRL1, (L2-D1) of CDRL2, and (L3-D1) of CDRL3, and the amino acid sequence of SEQ ID NO: 235. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-D3) includes, for example, the amino acid sequences of (L1-D1) of CDRL1, (L2-D1) of CDRL2, and (L3-D1) of CDRL3, and the amino acid sequence of SEQ ID NO: 235. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-D1). Also known as 18H-L13".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LE) antibody and the like are also referred to as, for example, antibody 18H-L14 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LE) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-E) below, and CDRL2 is (L2-E) below.
- CDRL3 is a polypeptide containing the amino acid sequence of (L3-E) below.
- (L1-E) Amino acid sequence of (L1-E1), (L1-E2), or (L1-E2) or (L1-E3) below (L1-E1) SEQ ID NO:236 (TSDVGTTNY) (L1-E2) Amino acid of SEQ ID NO:236 Amino acid sequence (L1-E3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 236.
- (L2-E) Amino acid sequence of the following (L2-E1), (L2-E2) or (L2-E3) (L2-E1) SEQ ID NO:237 (DVT) (L2-E2) Amino acid of SEQ ID NO:237 Amino acid sequence having 80% or more identity to the sequence (L2-E3) SEQ ID NO: 237, in which one or several amino acids have been deleted, substituted, inserted and/or added
- (L3-E) Amino acid sequence of (L3-E1), (L3-E2) or (L3-E3) below (L3-E1) SEQ ID NO: 238 (SYAGSYTFVV) (L3-E2) Amino acid sequence of SEQ ID NO: 238 Amino acid sequence (L3-E3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 238.
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LE) above includes, for example, a polypeptide having the amino acid sequence of (LE) below.
- the above-mentioned (L-E1) amino acid sequence is, for example, a sequence including the CDL1 (L1-E1), CDRL2 (L2-E1), and CDRL3 (L3-E1) amino acid sequences.
- the amino acid sequence of (L-E2) includes, for example, the amino acid sequences of (L1-E1) of CDRL1, (L2-E1) of CDRL2, and (L3-E1) of CDRL3, and the amino acid of SEQ ID NO: 239. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-E3) includes, for example, (L1-E1) of CDRL1, (L2-E1) of CDRL2, and (L3-E1) of CDRL3, and the amino acid sequence of SEQ ID NO: 239. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is (H-A1)
- the light chain variable region is (L-E1)
- the antibody of this combination is referred to as "antibody Also known as 18H-L14".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LF) antibody and the like are also referred to as, for example, antibody 18H-L16 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LF) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-F) below, and CDRL2 is (L2-F) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-F) below.
- (L1-F) The following (L1-F1), (L1-F2) or (L1-F3) amino acid sequence (L1-F1) SEQ ID NO:240 (SSDVGVYNY) amino acid sequence (L1-F2) SEQ ID NO:240 amino acid Amino acid sequence (L1-F3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 240.
- (L2-F) The following (L2-F1), (L2-F2) or (L2-F3) amino acid sequence (L2-F1) SEQ ID NO: 241 (DVS) amino acid sequence (L2-F2) SEQ ID NO: 241 amino acid Amino acid sequence (L2-F3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 241.
- (L3-F) Amino acid sequence of (L3-F1), (L3-F2) or (L3-F3) below (L3-F1) SEQ ID NO:242 (AAWDDSLNGVV) (L3-F2) SEQ ID NO:242 Amino acid sequence having 80% or more identity to the sequence (L3-F3) SEQ ID NO: 242, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LF) includes, for example, a polypeptide having the amino acid sequence of (LF) below.
- (LF) Amino acid sequence of the following (L-F1), (L-F2) or (L-F3) (L-F1) Amino acid sequence of SEQ ID NO:243 SEQ ID NO:243: QSALTQPPSASGSPGQSVTISCTGT SSDVGVYNY VSWYQQHPGKAPKLMIY DVS KRPSGVPDRFSGSKSANTASLTISGLQAEGTQLVADEADYYV AA (L-F2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 243 (L-F3) In the amino acid sequence of SEQ ID NO: 243, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-F1) amino acid sequence is, for example, a sequence containing the amino acid sequences of (L1-F1) of CDRL1, (L2-F1) of CDRL2, and (L3-F1) of CDRL3.
- the amino acid sequence of (L-F2) includes, for example, the amino acid sequences of (L1-F1) of CDRL1, (L2-F1) of CDRL2, and (L3-F1) of CDRL3, and the amino acid sequence of SEQ ID NO: 243. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-F3) includes, for example, the amino acid sequences of (L1-F1) of CDRL1, (L2-F1) of CDRL2, and (L3-F1) of CDRL3, and the amino acid sequence of SEQ ID NO: 243. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-F1). Also known as 18H-L16".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LG) antibody and the like are also referred to as, for example, antibody 18H-L17 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3
- CDRH1 is a polypeptide containing the amino acid sequence of (H1-A)
- CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A)
- CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LG) includes CDRL1, CDRL2, and CDRL3, wherein CDRL1 is a polypeptide containing the amino acid sequence of (L1-G) below, and CDRL2 is (L2-G) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-G) below.
- (L1-G) Amino acid sequence of (L1-G1), (L1-G2) or (L1-G3) below (L1-G1), amino acid sequence of (L1-G1) SEQ ID NO: 244 (SSNIGNNY) (L1-G2), amino acid of SEQ ID NO: 244 Amino acid sequence having 80% or more identity to the sequence (L1-G3), amino acid sequence of SEQ ID NO: 244 in which one or several amino acids have been deleted, substituted, inserted and/or added
- L2-G The following (L2-G1), (L2-G2) or (L2-G3) amino acid sequence (L2-G1) SEQ ID NO:245 (DNV) amino acid sequence (L2-G2) SEQ ID NO:245 amino acid Amino acid sequence having 80% or more identity to the sequence (L2-G3) SEQ ID NO: 245, in which one or several amino acids are deleted, substituted, inserted and/or added
- (L3-G) Amino acid sequence of the following (L3-G1), (L3-G2) or (L3-G3) (L3-G1) SEQ ID NO:246 (AAWDDSLSAI) (L3-G2) SEQ ID NO:246 Amino acid sequence having 80% or more identity to the sequence (L3-G3)
- amino acid sequence of SEQ ID NO: 246 one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LG) includes, for example, a polypeptide having the following amino acid sequence (L-G).
- LG below (L-G1), (L -G2) or (L-G3) of the amino acid sequence (L-G1) SEQ ID NO: 247 amino acid sequence SEQ ID NO 247: QSVLTQPPSASGTPGQRVTISCSGS SSNIGNNY VCWYQHLPGTAPKLLIY DNV KRPSGIPDRFSGSKSGTSASLAISGLRSEDEADYYC AAWDDSLSAI FGGGTELTVL (L-G2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 247 (L-G3) In the amino acid sequence of SEQ ID NO: 247, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the (L-G1) amino acid sequence is, for example, a sequence containing the CDL1 (L1-G1), CDRL2 (L2-G1), and CDRL3 (L3-G1) amino acid sequences.
- the amino acid sequence of (L-G2) includes, for example, the amino acid sequences of (L1-G1) of CDRL1, (L2-G1) of CDRL2, and (L3-G1) of CDRL3, and the amino acid sequence of SEQ ID NO: 247. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-G3) includes, for example, (L1-G1) of CDRL1, (L2-G1) of CDRL2, and (L3-G1) of CDRL3, and the amino acid sequence of SEQ ID NO: 247. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-G1)
- antibody of this combination hereinafter referred to as "antibody Also known as 18H-L17”.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LH) antibody and the like are also referred to as, for example, antibody 18H-L22 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3
- CDRH1 is a polypeptide containing the amino acid sequence of (H1-A)
- CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A)
- CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LH) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-H) below, and CDRL2 is (L2-H) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-H) below.
- (L1-H) Amino acid sequence of (L1-H1), (L1-H2) or (L1-H3) or (L1-H3) below (L1-H1) SEQ ID NO: 248 (SSDVGGYNY) (L1-H2) SEQ ID NO: 248 Amino acid sequence (L1-H3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 248.
- (L2-H) The following (L2-H1), (L2-H2) or (L2-H3) amino acid sequence (L2-H1) SEQ ID NO: 249 (DVS) amino acid sequence (L2-H2) SEQ ID NO: 249 Amino acid sequence having 80% or more identity to the sequence (L2-H3), amino acid sequence of SEQ ID NO: 249 in which one or several amino acids are deleted, substituted, inserted and/or added
- (L3-H) Amino acid sequence of the following (L3-H1), (L3-H2) or (L3-H3) (L3-H1) SEQ ID NO:250 (HSYDSSLSHV) (L3-H2) Amino acid of SEQ ID NO:250 Amino acid sequence (L3-H3) having 80% or more identity to the sequence, wherein the amino acid sequence of SEQ ID NO: 250 has one or several amino acids deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LH) includes, for example, a polypeptide having the amino acid sequence of (LH) below.
- the above-mentioned (L-H1) amino acid sequence is, for example, a sequence containing the amino acid sequences of (L1-H1) of CDRL1, (L2-H1) of CDRL2, and (L3-H1) of CDRL3.
- the amino acid sequence of (L-H2) includes, for example, the amino acid sequences of (L1-H1) of CDRL1, (L2-H1) of CDRL2, and (L3-H1) of CDRL3, and the amino acid of SEQ ID NO:251. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-H3) includes, for example, (L1-H1) of CDRL1, (L2-H1) of CDRL2, and (L3-H1) of CDRL3, and the amino acid sequence of SEQ ID NO:251. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-H1)
- the antibody of this combination is referred to below as "antibody Also known as 18H-L22”.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LI) antibody and the like are also referred to as, for example, antibody H1-K4 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LI) above includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-I) below, and CDRL2 is (L2-I) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-I) below.
- L1-I Amino acid sequence (L1-I1), (L1-I2) or (L1-I3) amino acid sequence (L1-I1) SEQ ID NO:252 (QSVSSN) (L1-I2) SEQ ID NO:252 Amino acid sequence having 80% or more identity to the sequence (L1-I3) SEQ ID NO: 252, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L2-I) Amino acid sequence of the following (L2-I1), (L2-I2) or (L2-I3) (L2-I1) SEQ ID NO: 253 (GAS) (L2-I2) SEQ ID NO: 253 Amino acid sequence having 80% or more identity to the sequence (L2-I3) In the amino acid sequence of SEQ ID NO: 253, one or several amino acids have been deleted, substituted, inserted and/or added
- (L3-I) Amino acid sequence of (L3-I1), (L3-I2) or (L3-I3) below (L3-I1) SEQ ID NO: 254 (QQYNNWPPLYT) (L3-I2) Amino acid of SEQ ID NO: 254 Amino acid sequence having 80% or more identity to the sequence (L3-I3) SEQ ID NO: 254, in which one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LI) includes, for example, a polypeptide having the amino acid sequence of (LI) below.
- the above-mentioned (L-I1) amino acid sequence is, for example, a sequence including the CDL1 (L1-I1), CDRL2 (L2-I1), and CDRL3 (L3-I1) amino acid sequences.
- the amino acid sequence of (L-I2) includes, for example, the amino acid sequences of (L1-I1) of CDRL1, (L2-I1) of CDRL2, and (L3-I1) of CDRL3, and the amino acid sequence of SEQ ID NO: 255. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-I3) includes, for example, (L1-I1) of CDRL1, (L2-I1) of CDRL2, and (L3-I1) of CDRL3, and the amino acid of SEQ ID NO: 255. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-I1). Also called “H1-K4".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LJ) antibody and the like are also referred to as, for example, antibody 18H-K5 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3
- CDRH1 is a polypeptide containing the amino acid sequence of (H1-A)
- CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A)
- CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LJ) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-J) below, and CDRL2 is (L2-J) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-J) below.
- (L1-J) Amino acid sequence of the following (L1-J1), (L1-J2) or (L1-J3) (L1-J1) SEQ ID NO: 256 (QSVSSY) (L1-J2) Amino acid of SEQ ID NO: 256 Amino acid sequence having 80% or more identity to the sequence (L1-J3) In the amino acid sequence of SEQ ID NO: 256, one or several amino acids are deleted, substituted, inserted and/or added
- L2-J Amino acid sequence (L2-J1), (L2-J2) or (L2-J3) below (L2-J1) SEQ ID NO:257 (DAS) amino acid sequence (L2-J2) SEQ ID NO:257 Amino acid sequence (L2-J3) having 80% or more identity to the sequence, wherein one or several amino acids are deleted, substituted, inserted and/or added in the amino acid sequence of SEQ ID NO: 257
- (L3-J) Amino acid sequence of the following (L3-J1), (L3-J2) or (L3-J3) (L3-J1) SEQ ID NO:258 (QQSYSTLLYT) (L3-J2) Amino acid of SEQ ID NO:258 Amino acid sequence having 80% or more identity to the sequence (L3-J3) SEQ ID NO: 258, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the (LJ) light chain variable region includes, for example, a polypeptide consisting of the following (LJ) amino acid sequence.
- LJ amino acid sequence (L-J1) of SEQ ID NO: 259 amino acid sequence SEQ ID NO 259: EIVLTQSPGTLSLSPGERATLSCRAS QSVSSY LAWYQQKPGQAPRLLIY DAS NRATGIPARFSGSGSGTDFTLTISSLEPEDFATYYC QQSYSTLLYT FGQGTKLEIK (L-J2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO: 259 (L-J3) In the amino acid sequence of SEQ ID NO: 259, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-J1) amino acid sequence is, for example, a sequence including the CDL1 (L1-J1), CDRL2 (L2-J1), and CDRL3 (L3-J1) amino acid sequences.
- the amino acid sequence of (L-J2) includes, for example, the amino acid sequences of (L1-J1) of CDRL1, (L2-J1) of CDRL2, and (L3-J1) of CDRL3, and the amino acid of SEQ ID NO: 259. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-J3) includes, for example, (L1-J1) of CDRL1, (L2-J1) of CDRL2, and (L3-J1) of CDRL3, and the amino acid of SEQ ID NO: 259 It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-J1)
- antibody of this combination hereinafter referred to as "antibody Also known as 18H-K5".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, wherein CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LK) above includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-K) below, and CDRL2 is (L2-K) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-K) below.
- (L1-K) Amino acid sequence of the following (L1-K1), (L1-K2) or (L1-K3) (L1-K1) SEQ ID NO: 260 (QSVSSY) (L1-K2) Amino acid of SEQ ID NO: 260 Amino acid sequence having 80% or more identity to the sequence (L1-K3) SEQ ID NO: 260, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L3-K) Amino acid sequence of the following (L3-K1), (L3-K2) or (L3-K3) (L3-K1) SEQ ID NO:262 (QQYDSLPLT) (L3-K2) Amino acid of SEQ ID NO:262 Amino acid sequence (L3-K3) having 80% or more identity to the sequence, wherein the amino acid sequence of SEQ ID NO: 262 has one or several amino acids deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LK) includes, for example, a polypeptide having the amino acid sequence of (LK) below.
- the above-mentioned (L-K1) amino acid sequence is, for example, a sequence including the CDL1 (L1-K1), CDRL2 (L2-K1), and CDRL3 (L3-K1) amino acid sequences.
- the amino acid sequence of (L-K2) includes, for example, the amino acid sequences of (L1-K1) of CDRL1, (L2-K1) of CDRL2, and (L3-K1) of CDRL3, and the amino acid of SEQ ID NO:263. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-K3) includes, for example, the amino acid sequences of (L1-K1) of CDRL1, (L2-K1) of CDRL2, and (L3-K1) of CDRL3, and the amino acid of SEQ ID NO:263. It may be an amino acid sequence in which one or several amino acids are deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-K1)
- antibody of this combination hereinafter referred to as "antibody Also known as 18H-K6".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LL) antibody and the like are also referred to as, for example, antibody 18H-K9 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LL) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-L) below, and CDRL2 is (L2-L) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-L) below.
- (L1-L) The following (L1-L1), (L1-L2) or (L1-L3) amino acid sequence (L1-L1) SEQ ID NO:264 (QTISASS) amino acid sequence (L1-L2) SEQ ID NO:264 amino acid Amino acid sequence having 80% or more identity to the sequence (L1-L3)
- amino acid sequence of SEQ ID NO: 264 one or several amino acids are deleted, substituted, inserted and/or added
- (L2-L) The following (L2-L1), (L2-L2) or (L2-L3) amino acid sequence (L2-L1) SEQ ID NO:265 (GAS) amino acid sequence (L2-L2) SEQ ID NO:265 amino acid Amino acid sequence having 80% or more identity to the sequence (L2-L3) SEQ ID NO: 265, wherein one or several amino acids are deleted, substituted, inserted and/or added
- (L3-L) Amino acid sequence of the following (L3-L1), (L3-L2) or (L3-L3) (L3-L1) SEQ ID NO: 266 (QQFNEWPLT) (L3-L2) Amino acid of SEQ ID NO: 266 Amino acid sequence having 80% or more identity to the sequence (L3-L3) In the amino acid sequence of SEQ ID NO: 266, one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LL) includes, for example, a polypeptide having the amino acid sequence of (L-L) below.
- the above-mentioned (L-L1) amino acid sequence is, for example, a sequence including the CDL1 (L1-L1), CDRL2 (L2-L1), and CDRL3 (L3-L1) amino acid sequences.
- the amino acid sequence of (L-L2) includes, for example, the amino acid sequences of (L1-L1) of CDRL1, (L2-L1) of CDRL2, and (L3-L1) of CDRL3, and the amino acid sequence of SEQ ID NO: 267. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-L3) includes, for example, the amino acid sequences of (L1-L1) of CDRL1, (L2-L1) of CDRL2, and (L3-L1) of CDRL3, and the amino acid of SEQ ID NO: 267. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1)
- the light chain variable region is the (L-L1)
- the antibody of this combination is referred to as "antibody Also known as 18H-K9".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the combination (LM) antibody and the like are also referred to as, for example, antibody 18H-K10 group.
- the heavy chain variable region of (HA) includes CDRH1, CDRH2, and CDRH3, CDRH1 is a polypeptide containing the amino acid sequence of (H1-A), and CDRH2 is It is a polypeptide containing the amino acid sequence of (H2-A), and CDRH3 is a polypeptide containing the amino acid sequence of (H3-A).
- the light chain variable region of (LM) includes CDRL1, CDRL2, and CDRL3, where CDRL1 is a polypeptide containing the amino acid sequence of (L1-M) below, and CDRL2 is (L2-M) below. Is a polypeptide containing the amino acid sequence of, and CDRL3 is a polypeptide containing the amino acid sequence of (L3-M) below.
- (L1-M) Amino acid sequence of the following (L1-M1), (L1-M2) or (L1-M3) (L1-M1) SEQ ID NO:268 (QSVSSN) (L1-M2) SEQ ID NO:268 Amino acid sequence having 80% or more identity to the sequence (L1-M3)
- amino acid sequence of SEQ ID NO: 268 one or several amino acids are deleted, substituted, inserted and/or added
- L2-M The following (L2-M1), (L2-M2) or (L2-M3) amino acid sequence (L2-M1) SEQ ID NO:269 (GAS) amino acid sequence (L2-M2) SEQ ID NO:269 amino acid sequence Amino acid sequence having 80% or more identity to the sequence (L2-M3) In the amino acid sequence of SEQ ID NO: 269, one or several amino acids have been deleted, substituted, inserted and/or added
- L3-M Amino acid sequence (L3-M2), (L3-M2) or (L3-M2) or (L3-M3) amino acid sequence (L3-M1) SEQ ID NO:270 (QQYGSSPDIFT) (L3-M2) SEQ ID NO:270 Amino acid sequence having 80% or more identity to the sequence (L3-M3) SEQ ID NO: 270, wherein one or several amino acids are deleted, substituted, inserted and/or added
- “Identity” in each of the above-mentioned CDRs is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more, respectively.
- substitution and the like is, for example, 1 to 5, 1 to 4, 1 to 3, 1 or 2, 1 respectively.
- the heavy chain variable region of (HA) includes, for example, a polypeptide consisting of the amino acid sequence of (H-A).
- the light chain variable region of (LM) includes, for example, a polypeptide having the amino acid sequence of (LM) below.
- (LM) Amino acid sequence of the following (L-M1), (L-M2) or (L-M3) (L-M1) Amino acid sequence of SEQ ID NO: 271 SEQ ID NO: 271: EIVLTQSPATLSLSPGERATLSCRAS QSVSSN LAWYQQKLGQAPRLLIY GAS SRATGIPDRFSGSGSGTDFTLTISRLEPEDFAVDIC QQFG (L-M2) Amino acid sequence having 80% or more identity to the amino acid sequence of SEQ ID NO:271 (L-M3) In the amino acid sequence of SEQ ID NO:271, one or several amino acids are deleted or substituted , Inserted and/or added amino acid sequence
- the above-mentioned (L-M1) amino acid sequence is, for example, a sequence including the CDL1 (L1-M1), CDRL2 (L2-M1), and CDRL3 (L3-M1) amino acid sequences.
- the amino acid sequence of (L-M2) includes, for example, (L1-M1) of CDRL1, (L2-M1) of CDRL2, and (L3-M1) of CDRL3, and the amino acid of SEQ ID NO: 271. It may be an amino acid sequence having 80% or more identity with the sequence.
- the amino acid sequence of (L-M3) includes, for example, the amino acid sequences of (L1-M1) of CDRL1, (L2-M1) of CDRL2, and (L3-M1) of CDRL3, and the amino acid sequence of SEQ ID NO: 271. It may be an amino acid sequence in which one or several amino acids have been deleted, substituted, inserted and/or added in the sequence.
- the heavy chain variable region is the (H-A1) and the light chain variable region is the (L-M1). Also known as 18H-K10".
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99, respectively. % Or more.
- substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8 respectively. 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- amino acid sequences of SEQ ID NOs: 216 to 271 are, for example, human-derived amino acid sequences.
- the binding between the antibody and the like of the present invention and CD19 can be confirmed by a method for detecting the binding between the antibody and the antigen, such as surface plasmon resonance (SPR) and flow cytometry.
- a method for detecting the binding between the antibody and the antigen such as surface plasmon resonance (SPR) and flow cytometry.
- the antibody or the like of the present invention may be labeled with a labeling substance, or may be immobilized on a carrier, a porous body or the like.
- the above description can be applied to the labeling and immobilization.
- the method for producing the antibody or the like of the present invention is not particularly limited, and for example, it can be produced by genetic engineering based on the aforementioned amino acid sequence information. Specifically, for example, it can be performed as follows.
- the present invention is not limited to this example.
- a vector containing a nucleic acid sequence encoding the amino acid sequence of each region, the heavy chain, and/or the light chain of the antibody of the present invention is introduced into a host to obtain a transformant. Then, the transformant is cultured, a fraction containing an antibody that binds to CD19 is collected, and the antibody is isolated or purified from the obtained collected fraction.
- the antibody is, for example, a monoclonal antibody.
- the monoclonal antibody include a monoclonal antibody obtained by immunizing an animal, a chimeric antibody, a humanized antibody, a human antibody (also referred to as a fully human antibody), and the like. The above description can be applied to the chimeric antibody and the humanized antibody.
- the antibody etc. of the present invention can also be prepared, for example, by immunizing an animal with an antigen.
- the antigen include CD19 protein.
- the antigen is preferably immunized multiple times.
- the peptide fragment may be, for example, a peptide fragment consisting only of an antigenic determinant (epitope) or a peptide fragment containing the antigenic determinant. The above description can be applied to the method for producing the monoclonal antibody.
- the (L) light chain variable region may be the following (L) light chain variable region.
- (L) comprises light chain complementarity determining regions (CDRL) 1, CDRL2, and CDRL3,
- CDRL1 is a polypeptide containing the following amino acid sequence (L1)
- CDRL2 is a polypeptide containing the following amino acid sequence (L2)
- CDRL3 is a light chain variable region that is a polypeptide comprising the amino acid sequence of (L3) below:
- (L1) Amino acid sequence of (L1-1), (L1-2) or (L1-3) below (L1-1) Any amino acid sequence of CDRL1 under the following conditions (L1) to (L3) and (L4)
- the amino acid sequence of (L1-2) (L1-1) which has 80% or more identity with the amino acid sequence (L1-3) (L1-1), one or several amino acids are Amino acid sequences deleted, substituted, inserted and/or added;
- (L2) Amino acid sequence of (L2-1), (L2-2) or (L2-3) below (L2-1) Any amino acid sequence of CDRL2 under the following conditions (
- X 3 is preferably V or I.
- X 5 is preferably absent, Y or T.
- X 10 is preferably W or Y.
- X 1 is preferably S or T.
- X 2 is preferably V or I.
- X 6 is preferably Y or F.
- X 12 is preferably absent, Y or F.
- Examples of the light chain variable region satisfying the condition (L1) include the light chain variable region of the above (LA).
- Examples of the light chain variable region satisfying the condition (L2) include the light chain variable region of the above (LC).
- Examples of the light chain variable region that satisfies the condition (L3) include the light chain variable regions of (LB), (LE), (LF), (LG), and (LH) described above.
- Examples of the light chain variable region satisfying the condition (L4) include the light chain variable regions of (LI), (LJ), (LK), (LL), and (LM) described above.
- the coding gene of the present invention includes an antibody against the complex of HLA-A*02:01 and NY-ESO-1 157-165 or an antigen-binding fragment thereof or an antibody against CD19 or an antigen-binding fragment thereof (hereinafter, referred to as “the present invention”). Antibody or antigen-binding fragment thereof or “the antibody of the present invention”), and the amino acid sequence of the first antibody or antigen-binding fragment of the present invention or the second antibody of the present invention or A nucleic acid (polynucleotide) encoding an antigen-binding fragment is included.
- the above-mentioned antibody of the present invention and the like can be obtained.
- the sequence of the coding gene of the present invention is not particularly limited as long as it is a sequence encoding the amino acid sequence of the antibody of the present invention, and may be a sense sequence or an antisense sequence.
- the expression vector of the present invention is an expression vector of an antibody against A2/NY-ESO-1 157 or an antigen-binding fragment thereof or an antibody against CD19 or an antigen-binding fragment thereof, and contains the above-mentioned coding gene of the present invention.
- the coding gene is linked to the linking vector so that the antibody of the present invention or an antigen-binding fragment thereof can be expressed.
- the expression vector of the present invention is not particularly limited as long as it can express the antibody of the present invention and the like, and other configurations.
- the expression vector of the present invention may be, for example, an expression vector containing a nucleic acid sequence encoding the heavy chain variable region and a nucleic acid sequence encoding the light chain variable region, or may be a nucleic acid sequence encoding the heavy chain variable region. It may be a set of an expression vector containing the same and an expression vector containing a nucleic acid sequence encoding the light chain variable region.
- the expression vector of the present invention can be prepared, for example, by ligating the coding gene of the present invention to a ligation vector.
- the description of the expression vector in the CAR library of the present invention can be incorporated.
- the type of the ligation vector that ligates the coding gene is not particularly limited, and examples thereof include pUC and pMX.
- the ligation vector can be appropriately set, for example, according to the host into which the expression vector is introduced.
- the host is not particularly limited, and examples thereof include mammalian cells such as CHO cells and the like.
- the transformant of the present invention is a transformant that expresses the antibody or the like of the present invention, and contains a host and the above-mentioned coding gene of the present invention.
- the transformant of the present invention only needs to have the above-described coding gene of the present invention expressible.
- the transformant preferably has, for example, the expression vector of the present invention.
- the method of introducing the expression vector into the host is not particularly limited, and a known method can be adopted.
- the explanation of the CAR library of the present invention, the first screening method, the first antibody, etc., the second antibody of the present invention, etc. can be referred to.
- the first chimeric antigen receptor (CAR) of the present invention comprises an antigen binding domain, a transmembrane domain, and an intracellular signal transduction domain, and the antigen binding domain is the antibody of the present invention. Or an antigen-binding fragment thereof.
- the first CAR of the present invention is characterized in that the antigen-binding domain contains the first antibody of the present invention and the like, and other configurations and conditions are not particularly limited.
- the description of the CAR library of the present invention, the first screening method, the first antibody, etc., the second antibody, etc., genes, expression vectors, and transformants can be used.
- the first CAR of the present invention can be preferably used as, for example, CAR-T cell CAR against A2/NY-ESO-1 157- expressing cancer cells.
- the antigen-binding domain may be, for example, the first antibody of the present invention or an antigen-binding fragment of the first antibody of the present invention.
- the antigen-binding fragment is preferably a single chain antibody (scFv).
- scFv single chain antibody
- antigen-binding domain examples include the following (BD) polypeptide.
- BD Polypeptide
- BD2 consisting of the amino acid sequences of any of the following (BD1), (BD2) or (BD3) (BD1) SEQ ID NOS: 98 to 117 (BD2) SEQ ID NO: 98 to 117
- BD3 consisting of an amino acid sequence having 80% or more identity and binding to A2/NY-ESO-1 157 has one or several amino acid sequences of any of SEQ ID NOs: 98 to 117. Consisting of an amino acid sequence in which the amino acid of is deleted, substituted, inserted and/or added, and binds to A2/NY-ESO-1 157
- the amino acid sequences of SEQ ID NOs: 98 to 117 have, for example, the heavy chain variable region and the light chain variable region of the combinations (1) to (18) in the first antibody of the present invention.
- an amino acid sequence containing The amino acid sequences of the heavy chain variable region and the light chain variable region in the amino acid sequences of SEQ ID NOs: 98 to 117 are, for example, the heavy chain variable regions of the combinations (1) to (18) in the first antibody of the present invention and the like. Reference can be made to the amino acid sequence containing the light chain variable region.
- the amino acid sequences corresponding to CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and CDRL3 of the amino acid sequences of SEQ ID NOs: 98 to 117 are conserved, for example.
- the amino acid sequences of the CDRs in the amino acid sequences of SEQ ID NOs: 98 to 117 are, for example, amino acids containing the heavy chain variable region and the light chain variable region of the combinations (1) to (18) in the first antibody of the present invention.
- the amino acid sequence of each CDR in the sequence can be referred to.
- H1-3M4E5L SEQ ID NO:98
- QVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSELTQPRSVSGSPGQSVTISCTGT SRDVGGYNY VSWYQQHPGKAPKLIIH DVI ERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYC WSFAGSYYV FGTDVTVL
- K52-H1 DIQMTQSPSAMSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISSLQPEDFATYYC QQYYSTPQT FGPGTKVDIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSS
- K73-H1 (SEQ ID NO: 100) DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKAGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYC QQYESYRRS FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSS
- K121-H1 DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDFATYYC QQYNSYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSS
- K124-H1 SEQ ID NO: 102
- DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS RLESGVPSRFSGSGSGTDFTLTISCLQSEDFATYYC QQYNSYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSS
- K145-H1 SEQ ID NO: 105
- DIQMTQSPSSLSASVGDRVSITCRAS QDISRY LNWYQQKPGKAPKLLLY
- AAS RLESGVPSRFSGSGSGTDFTLTITSLQPDDFATYYC QQYNSYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSS
- K151-H1 SEQ ID NO: 106
- DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTITSLQPDDFATYYC QQYDNLIT FGQGTRLEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYYGMDV WGQGTTVTVSS
- K160-H1 EIVLTQSPATLSVSPGERATLSCRAS QSVSSN LAWYQQKPGQAPRLLIY GAS TRATGIPARFSGSGSGTEFTLTISRLEPEDFATYYC QQYNSYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSS
- K173-H1 DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKAGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYC QQYESYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSS
- Antigen binding domain of 3M4E5H-L1 (SEQ ID NO: 110) EVQLLESGGGLVQPGGSLRLSCAAS GFTFSTYQ MSWVRQAPGKGLEWVSG IVSSGGST AYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYC AGELLPYYGMDV WGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSALTQPPSASGSPGQSVTISCTGT SSDVGGYDF VSWYQQHPGEAPKLLVY DVN NRPSGVSNRFSGSKSGNTASLTISGLQAEDEGDYYC SSYAGSNSV FGTGTKVTVL
- Antigen-binding domain of H73-3M4E5L (SEQ ID NO: 117) QLQLQESGPGLVKPSQTLSLTCTVS GGSISSYY WSWIRQPPGKGLEWIGE INHSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARCPIYYYGMDV WGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSELTQPRSVSGSPGQSVTISCTGT SRDVGGYNY VSWYQQHPGKAPKLIIH DVI ERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYC WSFAGSYYV FGTDVTVL
- the polypeptide of (BD1) may be a polypeptide having an amino acid sequence in which the amino acid sequences of the heavy chain variable region and the light chain variable region in the amino acid sequences of SEQ ID NOs: 98 to 117 are exchanged.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99%, respectively. That is all.
- polypeptide (BD3) “one or several” regarding substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8, respectively. It is 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- transmembrane domain and the intracellular signal transduction domain for example, the description of the first transmembrane domain and the first intracellular signal transduction domain of the CAR library of the present invention can be applied, respectively.
- first CAR of the present invention include the following polypeptide (C).
- C A polypeptide (C2) consisting of the amino acid sequences of any of the following (C1), (C2) or (C3) (C1) SEQ ID NOS: 118 to 137;
- C3 C1 SEQ ID NOS: 118 to 137;
- a polypeptide comprising an amino acid sequence having 80% or more identity with the amino acid sequence of any one of SEQ ID NOS: 118 to 137
- a polypeptide comprising an amino acid sequence in which one amino acid has been deleted, substituted, inserted and/or added, and which binds to A2/NY-ESO-1 157
- H1-3M4E5L CAR (SEQ ID NO: 118) QVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSELTQPRSVSGSPGQSVTISCTGT SRDVGGYNY VSWYQQHPGKAPKLIIH DVI ERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYC WSFAGSYYV FGTDVTVLAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLD
- K52-H1 CAR (SEQ ID NO: 119) DIQMTQSPSAMSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISSLQPEDFATYYC QQYYSTPQT FGPGTKVDIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDV
- K73-H1 CAR (SEQ ID NO: 120) DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKAGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYC QQYESYRRS FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDK
- K121-H1 CAR (SEQ ID NO: 121) DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDFATYYC QQYNSYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRR
- K124-H1 CAR (SEQ ID NO: 122) DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKPGKAPKLLIY AAS RLESGVPSRFSGSGTDFTLTISCLQSEDFATYYC QQYNSYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEM
- K145-H1 CAR (SEQ ID NO: 125) DIQMTQSPSSLSASVGDRVSITCRAS QDISRY LNWYQQKPGKAPKLLLY AAS RLESGVPSRFSGSGTDFTLTITSLQPDDFATYYC QQYNSYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGG
- K160-H1 CAR (SEQ ID NO: 127) EIVLTQSPATLSVSPGERATLSCRAS QSVSSN LAWYQQKPGQAPRLLIY GAS TRATGIPARFSGSGSGTEFTLTISRLEPEDFATYYC QQYNSYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGK
- K173-H1 CAR (SEQ ID NO: 128) DIQMTQSPSSLSASVGDRVTITCRAS QSISSY LNWYQQKAGKAPKLLIY AAS SLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYC QQYESYSRT FGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVS GGSISSNY WSWIRQAPGKGLEWIGH VSYSGST NYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYC ARESYYYYGMDV WGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDK
- 3M4E5H-L1 CAR (SEQ ID NO: 130) EVQLLESGGGLVQPGGSLRLSCAAS GFTFSTYQ MSWVRQAPGKGLEWVSG IVSSGGST AYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYC AGELLPYYGMDV WGQGTTVTVSSEVQLLESGGGLVQPGGSLRLSCAAS GFTFSTYQ MSWVRQAPGKGLEWVSG IVSSGGST AYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYC AGELLPYYGMDV WGQGTTVTVSSQSALTQPPSASGSPGQSVTISCTGT SSDVGGYDF VSWYQQHPGEAPKLLVY DVN NRPSGVSNRFSGSKSGNTASLTISGLQAEDEGDYYC SSYAGSNSV FGTGTKVTVLAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFW
- the amino acid sequences of SEQ ID NOs: 118 to 137 are, for example, the heavy chain variable region and the light chain of the combinations (1) to (18) in the first antibody or the like of the present invention, respectively. It is an amino acid sequence containing a variable region.
- the amino acid sequences corresponding to CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and CDRL3 of the amino acid sequences of SEQ ID NOs: 118 to 137 are conserved, for example.
- amino acid sequences of the CDRs in the amino acid sequences of SEQ ID NOs: 118 to 137 are, for example, the heavy chain variable region and the light chain variable region of the combinations (1) to (18) in the first antibody of the present invention, respectively. Reference can be made to the containing amino acid sequence.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99%, respectively. That is all.
- polypeptide (C3) “one or several” regarding substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8, It is 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the method for producing the first CAR of the present invention is not particularly limited, and it can be produced genetically based on the above-mentioned amino acid sequence information, for example. Specifically, for example, it can be performed as follows.
- the present invention is not limited to this example.
- an expression vector containing the nucleic acid encoding the first CAR of the present invention is introduced into a host to obtain a transformant. Then, the transformant is cultured to obtain a transformant expressing the first CAR.
- Examples of the expression vector include a vector containing a nucleic acid encoding the first CAR, and the like, and the description of the expression vector can be applied, for example.
- the host is not particularly limited and, for example, the above description of the host can be applied.
- the second chimeric antigen receptor (CAR) of the present invention comprises an antigen-binding domain, a transmembrane domain, and an intracellular signal transduction domain, as described above, and the antigen-binding domain is the antibody of the present invention. Or an antigen-binding fragment thereof.
- the second CAR of the present invention is characterized in that the antigen-binding domain contains the second antibody of the present invention and the like, and other configurations and conditions are not particularly limited.
- the description of the CAR library of the present invention, the first screening method, the first antibody, etc., the second antibody, etc., genes, expression vectors, and transformants can be used.
- the second CAR of the present invention can be preferably used, for example, as CAR-T cell CAR against CD19-expressing cancer cells.
- the antigen-binding domain may be, for example, the second antibody of the present invention or an antigen-binding fragment of the second antibody of the present invention.
- the antigen-binding fragment is preferably a single chain antibody (scFv).
- scFv single chain antibody
- antigen-binding domain examples include the following polypeptide (bd).
- Bd Polypeptide (bd2) consisting of the amino acid sequences of any of the following (bd1), (bd2) or (bd3) polypeptide (bd1) SEQ ID NOS: 272 to 284, and any of the amino acid sequences of SEQ ID NOS: 272 to 284
- amino acid sequence of any one of the polypeptide (bd3) SEQ ID NOs: 272 to 284 which has an amino acid sequence having 80% or more identity to CD19, one or several amino acids are deleted,
- a polypeptide consisting of a substituted, inserted and/or added amino acid sequence, which binds to CD19
- the amino acid sequences of SEQ ID NOs: 272 to 284 are, for example, the heavy chain variable region and the light chain variable region of the combinations (LA) to (LM) in the second antibody of the present invention.
- the amino acid sequences of the heavy chain variable region and the light chain variable region in the amino acid sequences of SEQ ID NOs: 272 to 284 are, for example, the heavy chain variable regions of the combinations (LA) to (LM) in the second antibody of the present invention and the like. Reference can be made to the amino acid sequence containing the light chain variable region.
- amino acid sequences corresponding to CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and CDRL3 of the amino acid sequences of SEQ ID NOs: 272 to 284 are conserved, for example.
- the amino acid sequences of the CDRs in the amino acid sequences of SEQ ID NOs: 272 to 284 are, for example, amino acids containing the heavy chain variable region and the light chain variable region of the combinations (LA) to (LM) in the second antibody of the present invention.
- the amino acid sequence of each CDR in the sequence can be referred to.
- Antigen binding domain of L4-18H SYELTQPPSVSVSPGQTASITCSGD KLGDKY ACWYQQKPGQSPVLVIY QDS KRPSGIPERFSGSNSGNTATLTISGTQAMDEADYYC QAWDSSTHVV FGGGTKLTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAAS GFTFDDYA MHWVRQAPGKGLEWVSG ISWNSGRI GYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYC ARDQGYHYYDSAEHAFDI WGQGTVVTVSS
- K6-18H SEQ ID NO: 282
- EIVLTRSPATLSLSPGERATLSCRAS QSVSSY LAWYQQKPGQAPRLLIY DAS NRATGISARFSGSGSGTEFTLTISSLQSEDIATYYC QQYDSLPLT FGGGTKLEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAAS GFTFDDYA MHWVRQAPGKGLEWVSG ISWNSGRI GYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYC ARDQGYHYYDSAEHAFDI WGQGTVVTVSS
- the polypeptide (BD1) may be a polypeptide having an amino acid sequence in which the amino acid sequences of the heavy chain variable region and the light chain variable region in the amino acid sequences of SEQ ID NOs: 272 to 284 are exchanged.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99%, respectively. That is all.
- polypeptide (bd3) “one or several” regarding substitution and the like is, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8, respectively. It is 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- transmembrane domain and the intracellular signal transduction domain for example, the description of the first transmembrane domain and the first intracellular signal transduction domain of the CAR library of the present invention can be applied, respectively.
- the second CAR of the present invention include the following (CA) polypeptides.
- CA Polypeptide
- CA2 consisting of the amino acid sequences of any of the following (CA1), (CA2) or (CA3) polypeptides (CA1) SEQ ID NOs: 285 to 297, and any of the amino acid sequences of SEQ ID NOs: 285 to 297
- CA3 polypeptides which has an amino acid sequence having 80% or more identity to CD19, one or several amino acids are deleted,
- a polypeptide comprising a substituted, inserted and/or added amino acid sequence, which binds to CD19
- the amino acid sequences of SEQ ID NOs: 285 to 297 are, for example, the heavy chain variable region and the light chain of the combinations (LA) to (LM) in the second antibody of the present invention, respectively. It is an amino acid sequence containing a variable region.
- the amino acid sequences corresponding to CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and CDRL3 of the amino acid sequences of SEQ ID NOs: 285 to 297 are conserved, for example.
- amino acid sequences of the CDRs in the amino acid sequences of SEQ ID NOs: 285 to 297 include, for example, the heavy chain variable region and the light chain variable region of the combinations (LA) to (LM) in the second antibody of the present invention, respectively. Reference can be made to the containing amino acid sequence.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99%, respectively. That is all.
- polypeptide of (CA3) “one or several” regarding substitution and the like are, for example, 1 to 20, 1 to 15, 1 to 10, 1 to 9, 1 to 8, respectively. It is 1 to 7, 1 to 6, 1 to 5, 1 to 4, 1 to 3, 1 or 2, and 1.
- the method for producing the second CAR of the present invention is not particularly limited, and it can be produced genetically based on the above-mentioned amino acid sequence information, for example. Specifically, for example, it can be performed as follows.
- the present invention is not limited to this example.
- an expression vector containing the nucleic acid encoding the second CAR of the present invention is introduced into a host to obtain a transformant. Then, the transformant is cultured to obtain a transformant expressing the second CAR.
- Examples of the expression vector include, for example, a vector containing a nucleic acid encoding the second CAR, and the description of the expression vector can be applied, for example.
- the host is not particularly limited and, for example, the above description of the host can be applied.
- the nucleic acid of the present invention encodes the aforementioned chimeric antigen receptor of the present invention, as described above.
- the nucleic acid of the present invention is characterized in that it encodes the first chimeric antigen receptor or the second chimeric antigen receptor of the present invention (hereinafter, also collectively referred to as "chimeric antigen receptor of the present invention").
- the other configurations and conditions are not particularly limited.
- the nucleic acid of the present invention enables cells to express the CAR of the present invention, for example.
- the nucleic acid of the present invention is, for example, the CAR library of the present invention, the first screening method, the first antibody, etc., the first antibody, etc., the gene, the expression vector, the transformant, the first chimeric antigen receptor
- the descriptions of the two chimeric antigen receptors can be incorporated.
- the nucleic acid of the present invention can be prepared by a conventional method, for example, based on the amino acid sequence of the CAR of the present invention.
- the nucleic acid encoding the CAR of the present invention is obtained by, for example, obtaining a base sequence encoding the amino acid sequence in a database in which the amino acid sequences of the above-mentioned domains are registered, and based on the base sequence, a molecule It can be prepared by biological methods and/or chemical synthetic methods.
- the nucleotide sequence of the nucleic acid may be codon-optimized depending on the origin of cells expressing the CAR of the present invention.
- nucleic acid of the present invention include the polynucleotide (c) below.
- C Polynucleotide (nucleic acid) of the following (c1), (c2), (c3), (c4), (c5), or (c6)
- C1 Polynucleotide encoding a polypeptide consisting of the amino acid sequence of any of SEQ ID NOs: 118 to 137 (c2) Amino acid having 80% or more identity with the amino acid sequence of any of SEQ ID NOs: 118 to 137
- a polynucleotide (c3) comprising a sequence and encoding a polypeptide that binds to A2/NY-ESO-1 157 , wherein one or several amino acids are deleted or substituted in the amino acid sequence of any of SEQ ID NOs: 118 to 137;
- a polynucleotide (c4) comprising an inserted and/or added amino acid sequence and encoding a polypeptide that binds to A2/
- the polynucleotides (c1) to (c3) are nucleic acids encoding the polypeptides (C1) to (C3), respectively, and the description of the polypeptides (C1) to (C3) can be incorporated.
- the nucleotide sequences of SEQ ID NOs: 138 to 157 are, for example, the heavy chain variable region and the light chain variable region of the combinations (1) to (18) in the first antibody of the present invention.
- a polynucleotide encoding a polypeptide comprising an amino acid sequence comprising:
- the polynucleotides encoding the amino acid sequences corresponding to CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and CDRL3 of the nucleotide sequences of SEQ ID NOs: 138 to 157 are, for example, conserved. Has been done.
- nucleotide sequences encoding the amino acid sequences of the CDRs in the nucleotide sequences of SEQ ID NOs: 138 to 157 are, for example, the heavy chain variable regions of the combinations (1) to (18) in the first antibody of the present invention and the like, respectively.
- the amino acid sequence of each CDR in the amino acid sequence containing the light chain variable region can be referred to.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99%, respectively. That is all.
- polynucleotide (c6) “one or several” regarding substitution and the like are, for example, 1 to 280, 1 to 210, 1 to 140, 1 to 70, 1 to 63, 1 to 56, 1 to 49, 1 to 35, 1 to 28, 1 to 21, 1 to 14, 1 to 6, 1 to 5, 1 to 4, One to three, one or two, and one.
- K145-H1 CAR (SEQ ID NO:145) 5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGGGACAGAGTCTCCATCACTTGCCGGGCAAGT CAGGACATTAGCAGGTAT TTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGCTCTAT GCTGCATCC AGATTGGAAAGTGGGGTCCCATCCAGGTTCAGTGGCAGTGGATCTGGGACAGACTTCACTCTCACCATCACCAGCCTGCAGCCTGATGACTTTGCAACTTATTACTGC CAACAGTATAATAGTTATTCCCGGACG TTCGGCCAAGGGACCAAGGTGGAAATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCA
- the following polynucleotide (ca) can be mentioned.
- a polynucleotide encoding a polypeptide that binds to CD19 consisting of any of the nucleotide sequences of SEQ ID NOs: 298 to 310 (ca5) consisting of any of the nucleotide sequences of SEQ ID NOs: 298 to 310
- the nucleotide sequence of any one of SEQ ID NOs: 298 to 310 which has a nucleotide sequence having 80% or more identity and which encodes a polypeptide that binds to CD19 (ca6)
- one or several nucleotides A polynucleotide encoding a polypeptide which binds to CD19 and has a nucleotide sequence in which is deleted, substituted, inserted and/or added
- the polynucleotides (ca1) to (ca3) are nucleic acids encoding the polypeptides (CA1) to (CA3), respectively, and the description of the polypeptides (CA1) to (CA3) can be incorporated.
- the nucleotide sequences of SEQ ID NOs: 298 to 310 are, for example, the heavy chain variable region and the light chain variable region of the combination (LA) to (LM) in the second antibody of the present invention.
- a polynucleotide encoding a polypeptide comprising an amino acid sequence comprising:
- the polynucleotides encoding the amino acid sequences corresponding to CDRH1, CDRH2, CDRH3, CDRL1, CDRL2, and CDRL3 of the nucleotide sequences of SEQ ID NOs: 298 to 310 are, for example, conserved. Has been done.
- nucleotide sequences encoding the amino acid sequences of the CDRs in the nucleotide sequences of SEQ ID NOs: 298 to 310 are, for example, the heavy chain variable regions of the combinations (LA) to (LM) in the second antibody of the present invention and the like, respectively.
- the amino acid sequence of each CDR in the amino acid sequence containing the light chain variable region can be referred to.
- the “identity” is, for example, 80% or more, 85% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99%, respectively. That is all.
- polynucleotide (ca6) “one or several” regarding substitution and the like is, for example, 1 to 280, 1 to 210, 1 to 140, 1 to 70, 1 to 63, 1 to 56, 1 to 49, 1 to 35, 1 to 28, 1 to 21, 1 to 14, 1 to 6, 1 to 5, 1 to 4, One to three, one or two, and one.
- the nucleic acid of the present invention may be introduced into an expression vector, for example.
- an expression vector for example, the above description can be incorporated.
- the various polynucleotides can be produced, for example, by a known genetic engineering method or synthetic method.
- the cell of the present invention contains the chimeric antigen receptor of the present invention as described above.
- the cell of the present invention is characterized by containing the chimeric antigen receptor of the present invention, and the other constitution and conditions are not particularly limited.
- the cell of the present invention can treat cancer, for example.
- the cells of the present invention include, for example, the CAR library of the present invention, the first screening method, the first antibody, etc., the second antibody, etc., a gene, an expression vector, a transformant, a first chimeric antigen receptor, The description of the second chimeric antigen receptor, nucleic acid and the like can be incorporated.
- the cell of the present invention may express a CAR other than the CAR of the present invention.
- the cells are preferably T cells.
- the T cells may be, for example, the T cell-like cells.
- the method for producing a cell of the present invention includes the step of introducing the nucleic acid of the present invention into a cell.
- the method for producing a cell of the present invention is characterized in that the nucleic acid of the present invention is introduced into a cell in the introducing step, and other steps and conditions are not particularly limited.
- cells capable of treating cancer can be produced.
- the method for producing cells of the present invention includes, for example, the CAR library of the present invention, the first screening method, the first antibody and the like, the second antibody and the like, genes, expression vectors, transformants, and first chimeric antigens.
- the description of the receptor, the second chimeric antigen receptor, the nucleic acid, the cell and the like can be incorporated.
- the cells are T cells or T cell-like cells.
- the method of introducing the nucleic acid in the introduction step is not particularly limited, and for example, the description of the first expression step in the first screening method of the present invention can be incorporated.
- the method for producing a cell of the present invention may include, for example, an expression step of causing the cell to express the chimeric antigen receptor encoded by the nucleic acid after the introduction step.
- the expression step can be carried out, for example, by culturing the cells after the introduction step.
- the therapeutic drug for cancer includes the cell, gene, expression vector or nucleic acid of the present invention.
- the therapeutic agent of the present invention is characterized by containing the cell, gene, expression vector, or nucleic acid of the present invention, and other configurations and conditions are not particularly limited.
- the therapeutic agent of the present invention can treat cancer, for example.
- the cancer therapeutic agent of the present invention includes, for example, the CAR library of the present invention, the first screening method, the first antibody and the like, the second antibody and the like, genes, expression vectors, transformants, and first chimeric antigens.
- the explanation of the receptor, the second chimeric antigen receptor, the nucleic acid, the cell, the method for producing the cell and the like can be incorporated.
- the cancer to be treated is not particularly limited and can be appropriately determined depending on, for example, the type of chimeric antigen receptor to be expressed.
- the cancer therapeutic agent of the present invention is a cell, gene, expression vector or nucleic acid related to CAR that binds to A2/NY-ESO-1 157
- the cancer to be treated is, for example, malignant melanoma or lung cancer.
- Synovial sarcoma breast cancer, esophageal cancer, ovarian cancer, bladder cancer, neuroblastoma, myeloma and the like.
- the cancer to be treated is, for example, B cell lymphoma, acute lymphocytic leukemia, chronic lymphocytic. Examples include leukemia and multiple myeloma.
- the administration conditions of the therapeutic agent of the present invention are not particularly limited, and, for example, depending on the type of cancer to be targeted, the degree of cancer progression, the age of the patient, etc., the administration form, administration method, administration timing, and dose. Etc. can be set appropriately.
- the method of using the therapeutic agent of the present invention is not particularly limited, and for example, the therapeutic agent of the present invention may be administered to the administration subject.
- Examples of the administration target include cells, tissues or organs.
- Examples of the administration target include humans and non-human animals other than humans.
- Examples of the non-human animal include mammals such as mouse, rat, dog, monkey, rabbit, sheep, horse and pig.
- the administration may be, for example, in vivo or in vitro .
- the administration method is not particularly limited and can be appropriately determined depending on the administration subject, for example.
- Examples of the administration method include parenteral administration and oral administration.
- Examples of the parenteral administration include local administration, subcutaneous administration, intradermal administration, intramuscular administration, intraperitoneal administration, intravenous administration, intralymphatic administration, intratumoral administration and the like.
- the administration form of the therapeutic agent of the present invention is not particularly limited, and examples thereof include injections (injections), liquid solutions such as intravenous drip infusion, and the like.
- the compounding amount of the cells is not particularly limited.
- the administration conditions of the cells are not particularly limited.
- the administration condition of the cells is, for example, a single dose (total) for adult human male is, for example, 1 ⁇ 10 8 to 3 ⁇ 10 10 cells, and preferably Is 1 ⁇ 10 9 to 1 ⁇ 10 10 cells, and the administration frequency is, for example, once every 1 to 4 weeks.
- the compounding amount of the cells is contained at a concentration that can realize the exemplified administration conditions.
- the amount of the gene, expression vector, or nucleic acid compounded is not particularly limited.
- the administration conditions of the gene, expression vector, or nucleic acid are not particularly limited.
- the administration method is subcutaneous injection, intravenous injection, or the like, and in the case of systemic administration to humans, the dose (total) per administration is, for example, 5 to 5000 mg, 50 to 500 mg, and the administration frequency is, for example, 2 Once every eight to eight weeks.
- the therapeutic agent of the present invention may contain only one kind of the cells, genes, expression vectors, and nucleic acids, or may contain plural kinds thereof.
- the therapeutic agent of the present invention may further contain other additives.
- the amount of the additive compounded is not particularly limited as long as it does not interfere with the function of the therapeutic agent of the present invention.
- the additive is not particularly limited, and for example, a pharmaceutically acceptable additive is preferable.
- the kind of the additive is not particularly limited and can be appropriately selected depending on, for example, the kind of the administration subject. Examples of the additives include bases, stabilizers, preservatives and the like.
- the method for treating cancer of the present invention includes an administration step of administering the cell, gene, expression vector or nucleic acid of the present invention to an administration subject.
- the method for treating cancer of the present invention is characterized by administering the cell, gene, expression vector or nucleic acid of the present invention in the administration step, and other steps and conditions are not particularly limited.
- cancer can be treated.
- the method for treating cancer of the present invention includes, for example, the CAR library of the present invention, the first screening method, the first antibody and the like, the second antibody and the like, genes, expression vectors, transformants, and first chimeras.
- the description of the antigen receptor, the second chimeric antigen receptor, nucleic acid, cell, method for producing cell, therapeutic agent and the like can be incorporated.
- any one of the cells, genes, expression vectors, and nucleic acids of the present invention may be administered, or multiple types may be administered.
- the present invention is the antibody of the present invention or an antigen-binding fragment thereof for use in a method for treating cancer, or the antibody of the present invention or an antigen-binding fragment thereof for treating cancer.
- the present invention is also the use of the above-mentioned antibody of the present invention or an antigen-binding fragment thereof for the manufacture of a medicament for treating cancer.
- the present invention provides the cell, gene, expression vector, or nucleic acid of the present invention for use in a method for treating cancer, or the cell, gene, or expression of the present invention for treatment of cancer. It is a vector or nucleic acid.
- the present invention is also the use of the cell, gene, expression vector or nucleic acid according to the present invention for the manufacture of a medicament for treating cancer.
- the use of the present invention includes, for example, the CAR library of the present invention, the first screening method, the first antibody and the like, the second antibody and the like, the gene, the expression vector, the transformant, the first chimeric antigen receptor, The description of the second chimeric antigen receptor, nucleic acid, cell, method for producing cell, therapeutic agent, therapeutic method and the like can be incorporated.
- the bispecific antibody (BsAb) library of the present invention comprises a nucleic acid encoding a first bispecific antibody (BsAb), wherein the first BsAb comprises a first antigen-binding domain and a second antigen-binding domain.
- scFv single-chain antibody
- a second scFv for screening for binding to an antigen comprising a second heavy chain variable region and a second light chain variable region, said second heavy chain variable region and The second light chain variable region satisfies the following Condition 1 or Condition 2, and the first target antigen or the second target antigen is an immune cell activating receptor.
- the heavy chain complementarity determining regions (CDRH) 1, CDRH2, and CDRH3 in the second heavy chain variable region are respectively CDRH1 of the heavy chain variable region in the antibody or the antigen-binding fragment thereof that binds to the second target antigen.
- Light chain complementarity determining regions (CDRL) 1, CDRL2, and CDRL3 in the second light chain variable region each include CDRL1, CDRL2, and CDRL3 in the light chain variable region of the first B cell receptor;
- the heavy chain complementarity determining regions (CDRH) 1, CDRH2, and CDRH3 in the second heavy chain variable region each include CDRH1, CDRH2, and CDRH3 in the heavy chain variable region of the first B cell receptor,
- Light chain complementarity determining regions (CDRL) 1, CDRL2, and CDRL3 in the second light chain variable region are respectively CDRL1 of the light chain variable region in the antibody or the antigen-binding fragment thereof that binds to the second target antigen. , CDRL2, and CDRL3.
- the first BsAb in the BsAb library of the present invention, includes a first antigen-binding domain and a second antigen-binding domain, and the first antigen-binding domain is the first antigen-binding domain.
- a second scFv that screens for binding to a second target antigen and the second scFv is a second scFv that binds to a target antigen of A heavy chain variable region and a second light chain variable region, wherein the second heavy chain variable region and the second light chain variable region satisfy the condition 1 or the condition 2;
- the antigen or the second target antigen is an immune cell activating receptor, and other configurations and conditions are not particularly limited.
- the BsAb library of the present invention includes, for example, the CAR library of the present invention, the first screening method, the first antibody and the like, the second antibody and the like, genes, expression vectors, transformants, and first chimeric antigen receptors.
- the description of the second chimeric antigen receptor, nucleic acid, cell, method for producing cell, therapeutic agent, therapeutic method and the like can be incorporated.
- the first BsAb is a protein containing the first antigen-binding domain and the second antigen-binding domain.
- the first antigen-binding domain and the second antigen-binding domain respectively bind a first single-chain antibody (scFv) that binds to a first target antigen and a second target antigen to each other.
- scFv first single-chain antibody
- the second scFv has, for example, a structure similar to scFv, and the extracellular domain that binds to the antigen is changed to the second antigen binding domain that screens for binding to the second target antigen.
- the first target antigen or the second target antigen is an immune cell activating receptor. Therefore, when the first target antigen is the immune cell activating receptor, the first BsAb that binds to the second target antigen may be, for example, the second target antigen and the first target antigen.
- the immune cell activating receptor In the coexistence of immune cells capable of expressing (the immune cell activating receptor), a signal can be induced through the immune cell activating receptor of the immune cells, and as a result, the immune cells are activated (for example, , Proliferation, expression of activation markers, etc.).
- the first target antigen is the immune cell activating receptor
- the second screening method described below the first BsAb that activates the immune cells
- the scFv that binds to the second target antigen can be more easily screened as compared with, for example, the method using a hybridoma and the method using phage display.
- the first BsAb that binds to the second target antigen can activate the immune cells in, for example, the second screening method described below, and thus is a BsAb that can activate the immune cells. I can say. Therefore, according to the BsAb library of the present invention, for example, BsAbs capable of inducing the immune activity mediated by the immune cells can be screened for the cells expressing the second target antigen.
- the first BsAb that binds to the second target antigen may be, for example, the first target antigen and the second target antigen (the above-mentioned In the coexistence of immune cells capable of expressing immune cell activating receptors, a signal can be induced through the immune cell activating receptors of the immune cells, and as a result, immune cells can be activated (eg, proliferation, activity). Expression marker, etc.).
- the second target antigen is an immune cell activating receptor
- the second target antigen is an immune cell activating receptor
- the first BsAb that activates immune cells can be selected as the BsAb that binds to the second target antigen. Therefore, according to the BsAb library of the present invention, the scFv that binds to the second target antigen can be more easily screened as compared with, for example, the method using a hybridoma and the method using phage display.
- the first BsAb that binds to the second target antigen can be said to be a BsAb capable of activating immune cells, since it can activate the immune cells in the second screening method described later, for example. .. Therefore, according to the BsAb library of the present invention, for example, BsAb capable of inducing immune activity mediated by immune cells can be screened for cells expressing the first target antigen.
- the second antigen binding domain contains a second scFv for screening for binding to the target antigen.
- the second scFv is, for example, a single-chain polypeptide derived from an antibody that binds to an antigen, and has a structure similar to scFv that retains the ability to bind to the antigen.
- the scFv is a polypeptide in which a variable region fragment (Fv) of a heavy chain (H chain) and a light chain (L chain) of an antibody that binds to the antigen is linked.
- one of a heavy chain variable region and a light chain variable region is a variable region such as an antibody that binds to the second target antigen
- the other is A variable region derived from a B cell receptor, that is, a variable region whose binding to the second target antigen is unknown.
- the heavy chain variable region or the light chain variable region that binds to the second target antigen can be screened. ..
- the BsAb library of the present invention for example, as compared with the phage display method in which the heavy chain variable region and the light chain variable region that bind to the second target antigen are screened at once, a smaller number or type , The scFv that binds to the second target antigen can be screened.
- the nucleic acid encoding the first BsAb means, for example, a nucleic acid (polynucleotide) encoding the amino acid sequence of the first BsAb, and the above description can be applied.
- BsAb means, for example, an antibody or an antigen-binding fragment thereof in which two antigen-binding sites in an antibody molecule bind to different target antigens.
- the different target antigens mean, for example, different epitopes and may be epitopes of different antigens or different epitopes of the same antigen.
- the “BsAb” can adopt, for example, a structure of a known bispecific antibody, and specific examples thereof include a dual variable domain antibody (DVD-Ig (trademark)), a diabody, and a triabody.
- DVD-Ig dual variable domain antibody
- Tetrabody, tanda, flexibody which is a combination of scFv and diabody, tandem scFv (eg BiTE®, Micromet), DART® (MacroGenics), Fcab (trademark) or mAb 2 (trademark) (F-star), Fc engineering antibody (Xencor) or DuoBody (registered trademark) (Genmab), etc.
- tandem scFv eg BiTE®, Micromet
- DART® MicroGenics
- Fcab trademark
- mAb 2 trademark
- Fc engineering antibody Xencor
- DuoBody registered trademark
- the tandem scFv is, for example, a protein in which two scFvs are linked by a linker peptide (inter-Fv linker peptide), and the two scFvs have a heavy chain variable region and a light chain variable region, respectively, as a linker peptide (Fv linker peptide). ) Are connected by.
- the first target antigen is an immune cell activating receptor and a case where the second target antigen is an immune cell activating receptor will be described respectively.
- the immune cell activating receptor is, for example, expressed in the immune cell, and binds with a ligand or crosslinks a plurality of immune cell activating receptors.
- the immune cell activating receptor can be appropriately determined depending on, for example, the type of the immune cell.
- the immune cell activating receptor includes CD3 and the like. Examples of the CD3 include CD3 ⁇ , CD3 ⁇ , CD3 ⁇ , CD3 ⁇ , CD3 ⁇ , CD3 ⁇ or a complex thereof.
- the immune cell activating receptors include NK cell activating receptors.
- NK cell activation receptor examples include CD94/NKG2C, CD94/NKG2E, NKG2D/NKG2D and the like.
- examples of the immune cell activating receptor include CD3 and the NK cell activating receptor.
- examples of the immune cell activating receptor include CD19, CD79 ⁇ , CD79 ⁇ and the like.
- the second target antigen is not particularly limited, and for example, the description of the target antigen in the CAR library can be incorporated.
- the first scFv contains, for example, a first heavy chain variable region and a first light chain variable region.
- the first heavy chain variable region and the first light chain variable region have the same structure as the heavy chain variable region and the light chain variable region in the antibody molecule.
- the heavy chain variable region and the light chain variable region of an antibody molecule each have three complementarity determining regions (CDRs).
- CDR is also called a hypervariable domain. Even in the variable regions of the heavy chain and the light chain, CDR is a region in which the primary structure has a high variability, and the CDR is usually separated at three positions on the primary structure.
- the three CDRs in the heavy chain variable region are determined from the amino terminal (N-terminal) side in the amino acid sequence of the heavy chain variable region, namely, heavy chain CDR1 (CDRH1), heavy chain CDR2 (CDRH2), and heavy chain CDR3.
- the three CDRs in the light chain variable region are represented by (CDRH3), and the light chain CDR1 (CDRL1), the light chain CDR2 (CDRL2), and the light chain CDR3 (CDRL3 ). These sites are conformationally close to each other and determine their specificity for binding antigen.
- the first heavy chain variable region has CDRH1, CDRH2, and CDRH3.
- the first light chain variable region has CDRL1, CDRL2, and CDRL3.
- CDRH1, CDRH2, and CDRH3 in the first heavy chain variable region, and CDRL1, CDRL2, and CDRL3 in the first light chain variable region are an antibody that binds to an immune cell activating receptor such as CD3 or the like.
- An antigen-binding fragment is used. Specific examples of the antibody or the antigen-binding fragment thereof can be referred to, for example, the description of the antibody and the antigen-binding fragment in the antibody of the present invention described above.
- the antibody or the like that binds to the immune cell activating receptor for example, a known antibody or the like can be used depending on the type of the immune cell activating receptor.
- examples of the antibody against CD3 ⁇ include OKT3 antibody, UCHT1 antibody, L2K antibody, HIT3a antibody, 28F11 antibody, and 27H5 antibody.
- examples of the antibody against CD19 include FMC63 antibody and SJ25C1.
- the antibody or the like that binds to the immune cell activating receptor may be, for example, an antibody obtained by immunizing the immune cell activating receptor.
- the antibody or the like that binds to the immune cell activating receptor may be, for example, scFv obtained by the second screening method of the present invention described below.
- CDRH1, CDRH2, and CDRH3 in the first heavy chain variable region may each be, for example, a polypeptide consisting of the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the immune cell activating receptor.
- a polypeptide containing the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the immune cell activating receptor may be used.
- a region other than CDRH1, CDRH2, and CDRH3, that is, the framework region (FR) is, for example, a heavy chain variable region in an antibody or the like that binds to the immune cell activating receptor. It may include FR.
- the FRs are usually separated in four places on the primary structure.
- the four FRs in the heavy chain variable region are labeled from the N-terminal side in the amino acid sequence of the heavy chain variable region to heavy chain FR1 (FRH1), heavy chain FR2 (FRH2), heavy chain FR3 (FRH3). , And heavy chain FR4 (FRH4).
- the CDRHs and the FRHs are arranged in the order of FRH1, CDRH1, FRH2, CDRH2, FRH3, CDRH3, and FRH4 from the N-terminal side in the amino acid sequence of the heavy chain variable region. .. FRH1, FRH2, FRH3, and FRH4 in the first heavy chain variable region are each composed of, for example, an amino acid sequence of FRH1, FRH2, FRH3, and FRH4 of an antibody or the like that binds to the immune cell activating receptor.
- It may be a peptide or a polypeptide containing the amino acid sequences of FRH1, FRH2, FRH3, and FRH4, such as an antibody that binds to a preimmune cell activating receptor.
- the "antibody or the like that binds to an immune cell activating receptor” in the description of each CDRH and the “antibody or the like that binds to an immune cell activating receptor” in the description of each FRH may be the same antibody or the like. preferable.
- the first heavy chain variable region may include, for example, a heavy chain variable region in an antibody or the like that binds to the immune cell activating receptor.
- the first heavy chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the heavy chain variable region in an antibody or the like that binds to the immune cell activating receptor, or the immune cell activating receptor It may be a polypeptide containing the amino acid sequence of the heavy chain variable region of an antibody or the like that binds to.
- CDRL1, CDRL2, and CDRL3 in the first light chain variable region may each be, for example, a polypeptide consisting of the amino acid sequences of CDRL1, CDRL2, and CDRL3, such as an antibody that binds to the immune cell activating receptor.
- a polypeptide containing the amino acid sequences of CDRL1, CDRL2, and CDRL3, such as an antibody that binds to the immune cell activating receptor, may be used.
- the FR may include, for example, the FR of the light chain variable region in the antibody or the like that binds to the immune cell activating receptor.
- the FRs are usually separated in four places on the primary structure.
- four FRs in the light chain variable region are labeled from the N-terminal side in the amino acid sequence of the light chain variable region to a light chain FR1 (FRL1), a light chain FR2 (FRL2), a light chain FR3 (FRL3). , And light chain FR4 (FRL4).
- the CDRLs and the FRLs are arranged, for example, in the order of FRL1, CDRL1, FRL2, CDRL2, FRL3, CDRL3, and FRL4 from the N-terminal side in the amino acid sequence of the light chain variable region.
- FRL1, FRL2, FRL3, and FRL4 in the first light chain variable region are, for example, poly-amino acid sequences of FRL1, FRL2, FRL3, and FRL4, such as an antibody that binds to the immune cell activating receptor, respectively. It may be a peptide or a polypeptide containing the amino acid sequences of FRL1, FRL2, FRL3 and FRL4 such as an antibody that binds to the immune cell activating receptor.
- the “antibody or the like that binds to an immune cell activating receptor” in the description of each CDRL and the “antibody or the like that binds to an immune cell activating receptor” in the description of each FRL may be the same antibody or the like. preferable.
- the first light chain variable region may include, for example, a light chain variable region in an antibody or the like that binds to the immune cell activating receptor.
- the first light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region in an antibody or the like that binds to the immune cell activating receptor, or the immune cell activating receptor It may be a polypeptide containing the amino acid sequence of the light chain variable region of an antibody or the like that binds to.
- first linker peptide In the first scFv, the first heavy chain variable region and the first light chain variable region are linked by, for example, a first linker peptide (first Fv linker peptide).
- the first Fv linker peptide preferably does not, for example, inhibit the binding of the first scFv to the immune cell activating receptor.
- the first Fv linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the first Fv linker peptide is composed of amino acids such as glycine and serine, and a specific example thereof is (GGGGS) n .
- the n is, for example, an integer of 1 to 6.
- the amino acid sequence of the first Fv linker peptide includes, for example, a polypeptide having the amino acid sequence of SEQ ID NO: 1 or 2.
- the second scFv contains a second heavy chain variable region and a second light chain variable region.
- the second heavy chain variable region and the second light chain variable region have the same structure as the heavy chain variable region and the light chain variable region in the antibody molecule.
- the first scFv The explanation of can be applied.
- the second heavy chain variable region has CDRH1, CDRH2, and CDRH3.
- the second light chain variable region has CDRL1, CDRL2, and CDRL3. Further, the second heavy chain variable region and the second light chain variable region satisfy the condition 1 or the condition 2.
- CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region use an antibody or the like that binds to the second target antigen, and CDRL1, CDRL2 in the second light chain variable region, and CDRL3 is screened for binding to the second target antigen.
- Specific examples of the antibody or the antigen-binding fragment thereof can be referred to, for example, the description of the antibody and the antigen-binding fragment in the antibody of the present invention described above.
- the antibody or the like that binds to the second target antigen for example, a known antibody or the like can be used according to the second target antigen.
- a known antibody or the like can be used according to the second target antigen.
- 3M4E5 antibody can be used as an antibody against the complex of HLA-A*02:01 and NY-ESO-1 157-165 .
- FMC63 antibody can be used as an antibody against human CD19.
- the antibody or the like that binds to the second target antigen may be, for example, scFv obtained by the second screening method of the present invention described below, or may be obtained by immunizing with the second target antigen.
- Antibody may be used.
- CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region may each be, for example, a polypeptide consisting of the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the second target antigen. It may be a polypeptide containing the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the second target antigen.
- FRH1, FRH2, FRH3, and FRH4 in the second heavy chain variable region are, for example, polypeptides each comprising an amino acid sequence of FRH1, FRH2, FRH3, and FRH4, such as an antibody that binds to the second target antigen.
- it may be a polypeptide containing the amino acid sequences of FRH1, FRH2, FRH3, and FRH4, such as an antibody that binds to the second target antigen.
- the “antibody or the like that binds to the second target antigen” in the description of each CDRH and the “antibody or the like that binds to the second target antigen” in the description of each FRH are preferably the same antibody or the like.
- the second heavy chain variable region may include, for example, a heavy chain variable region in an antibody or the like that binds to the second target antigen.
- the second heavy chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the heavy chain variable region in an antibody or the like that binds to the second target antigen, or may bind to the second target antigen.
- the antibody may be a polypeptide containing the amino acid sequence of the heavy chain variable region.
- the second light chain variable region is encoded by, for example, a VJ gene fragment formed by reconstructing a VJ gene and a V gene fragment. Therefore, the first B cell receptor is, for example, a B cell receptor containing a polypeptide encoded by an artificial VJ gene fragment designed by artificially combining a V gene fragment and a J gene fragment. Good. In vivo B cells, for example, express the light chain variable region after VJ gene reconstruction. Therefore, the first B cell receptor may be, for example, an isolated B cell-derived B cell receptor. In this case, the first B cell receptor may be, for example, a human-derived B cell B cell receptor, and preferably a human peripheral blood B cell-derived B cell receptor. In the BsAb library of the present invention, the first B cell receptor is preferably the light chain variable region of the isolated B cell-derived B cell receptor because the BsAb library can be prepared more easily.
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRL1, CDRL2, and CDRL3 of the first B cell receptor, respectively. It may be a polypeptide comprising the amino acid sequences of CDRH1, CDRL2 and CDRL3 of the B cell receptor of.
- FRL1, FRL2, FRL3, and FRL4 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the first B cell receptor, respectively.
- the polypeptide may include the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the first B cell receptor.
- the "first B cell receptor" in the description of each CDRL and the "first B cell receptor" in the description of each FRL are preferably the same B cell receptor.
- the second light chain variable region may include, for example, the light chain variable region of the first B cell receptor.
- the second light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region in the first B cell receptor, or the light chain in the first B cell receptor. It may be a polypeptide containing the amino acid sequence of the variable region.
- the second heavy chain variable region and the second light chain variable region are linked by, for example, a second linker peptide (second Fv linker peptide).
- the second Fv linker peptide preferably does not, for example, inhibit the binding of the second scFv to the second target antigen.
- the second Fv linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the second Fv linker peptide is composed of amino acids such as glycine and serine, and a specific example thereof is (GGGGS) n .
- the n is, for example, an integer of 1 to 6.
- the amino acid sequence of the second Fv linker peptide includes, for example, a polypeptide having the amino acid sequence of SEQ ID NO: 1 or 2.
- the second Fv linker peptide may be the same as or different from the first Fv linker peptide, for example.
- the CDRL1, CDRL2, and CDRL3 in the second light chain variable region are derived from an antibody or the like that binds to the second target antigen, and CDRH1 in the second heavy chain variable region is , CDRH2, and CDRH3 are screened for binding to the second target antigen.
- the second heavy chain variable region is encoded by, for example, a VDJ gene fragment formed by reconstructing the VDJ gene fragment, V gene fragment, and J gene fragment. Therefore, the first B cell receptor is, for example, a B cell containing a polypeptide encoded by an artificial VDJ gene fragment designed by artificially combining a V gene fragment, a D gene fragment, and a J gene fragment. It may be a receptor. In vivo B cells, for example, express the heavy chain variable region after VDJ gene reconstruction. Therefore, the first B cell receptor may be, for example, an isolated B cell-derived B cell receptor.
- the first B cell receptor may be, for example, a human-derived B cell B cell receptor, and preferably a human peripheral blood B cell-derived B cell receptor.
- the first B cell receptor is a heavy chain variable region of an isolated B cell-derived B cell receptor because a BsAb library can be prepared more easily.
- CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRH1, CDRH2, and CDRH3 of the first B cell receptor, respectively. It may be a polypeptide comprising the amino acid sequences of CDRH1, CDRH2 and CDRH3 of the B cell receptor of.
- FRH1, FRH2, FRH3, and FRH4 in the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 of the first B cell receptor, respectively.
- the polypeptide may include the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 of the first B cell receptor.
- the “first B cell receptor” in the description of each CDRH and the “first B cell receptor” in the description of each FRH are preferably the same B cell receptor.
- the second heavy chain variable region may include, for example, the heavy chain variable region in the first B cell receptor.
- the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequence of the heavy chain variable region in the first B cell receptor, or the heavy chain in the first B cell receptor. It may be a polypeptide containing the amino acid sequence of the variable region.
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region may each be, for example, a polypeptide consisting of the amino acid sequences of CDRL1, CDRL2, and CDRL3 such as an antibody that binds to the second target antigen, It may be a polypeptide containing the amino acid sequences of CDRL1, CDRL2, and CDRL3, such as an antibody that binds to the second target antigen.
- the FR may include, for example, the FR of the light chain variable region in the antibody or the like that binds to the second target antigen.
- FRL1, FRL2, FRL3, and FRL4 in the second light chain variable region are, for example, polypeptides each comprising an amino acid sequence of FRL1, FRL2, FRL3, and FRL4, such as an antibody that binds to the second target antigen.
- it may be a polypeptide containing the amino acid sequences of FRL1, FRL2, FRL3, and FRL4, such as an antibody that binds to the second target antigen.
- the “antibody or the like that binds to the second target antigen” in the description of each CDRL and the “antibody or the like that binds to the second target antigen” in the description of each FRL are preferably the same antibody or the like.
- the second light chain variable region may include, for example, a light chain variable region in an antibody or the like that binds to the second target antigen.
- the second light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region in an antibody or the like that binds to the second target antigen, or may bind to the second target antigen.
- the antibody may be a polypeptide containing the amino acid sequence of the light chain variable region.
- the second heavy chain variable region and the second light chain variable region are linked by, for example, a second linker peptide (second Fv linker peptide).
- the second Fv linker peptide preferably does not, for example, inhibit the binding of the second scFv to the second target antigen.
- the description of the second Fv linker peptide can be referred to, for example, the description of the first Fv linker peptide.
- the second Fv linker peptide may be the same as or different from the first Fv linker peptide, for example.
- the first scFv and the second scFv are, for example, directly or indirectly linked.
- the connection of the first scFv and the second scFv can be appropriately designed, for example, according to the type of the first BsAb.
- the first BsAb is a tandem scFv
- the first scFv and the second scFv are linked by a linker peptide (inter-Fv linker peptide).
- the inter-Fv linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the inter-Fv linker peptide is composed of amino acids such as glycine and serine, and a specific example thereof is (GGGGS) n .
- the n is, for example, an integer of 1 to 6.
- the amino acid sequence of the inter-Fv linker peptide includes, for example, a polypeptide having the amino acid sequence of SEQ ID NO: 1 or 2.
- the first target antigen is not particularly limited, and for example, the description of the target antigen in the CAR library can be applied.
- the explanation of the first target antigen in “(1) First target antigen: immune cell activating receptor” of the BsAb library of the present invention can be applied.
- the first scFv contains, for example, a first heavy chain variable region and a first light chain variable region.
- the first heavy chain variable region and the first light chain variable region have the same structures as the heavy chain variable region and the light chain variable region in the antibody molecule, and the “(1) of the BsAb library of the present invention is used.
- the description of the first scFv in "First target antigen: immune cell activating receptor" can be incorporated.
- the first heavy chain variable region has, for example, CDRH1, CDRH2, and CDRH3.
- the first light chain variable region has, for example, CDRL1, CDRL2, and CDRL3.
- CDRH1, CDRH2, and CDRH3 in the first heavy chain variable region and CDRL1, CDRL2, and CDRL3 in the first light chain variable region are an antibody or an antigen-binding fragment thereof that binds to the first target antigen.
- Specific examples of the antibody or the antigen-binding fragment thereof can be referred to, for example, the description of the antibody and the antigen-binding fragment in the antibody of the present invention described above.
- the antibody or the like that binds to the first target antigen for example, an antibody or the like that binds to a known first target antigen can be used depending on the type of the first target antigen. Further, the antibody or the like that binds to the first target antigen may be, for example, an antibody obtained by immunizing the first target antigen. The antibody or the like that binds to the first target antigen may be, for example, scFv obtained by the second screening method of the present invention described below.
- CDRH1, CDRH2, and CDRH3 in the first heavy chain variable region may each be, for example, a polypeptide consisting of the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the first target antigen. It may be a polypeptide containing the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the first target antigen.
- FRH1, FRH2, FRH3, and FRH4 in the first heavy chain variable region are, for example, polypeptides each comprising an amino acid sequence of FRH1, FRH2, FRH3, and FRH4, such as an antibody that binds to the first target antigen.
- it may be a polypeptide containing the amino acid sequences of FRH1, FRH2, FRH3, and FRH4, such as an antibody that binds to the first target antigen.
- the “antibody or the like that binds to the first target antigen” in the description of each CDRH and the “antibody or the like that binds to the first target antigen” in the description of each FRH are preferably the same antibody or the like.
- the first heavy chain variable region may include, for example, a heavy chain variable region in an antibody or the like that binds to the first target antigen.
- the first heavy chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the heavy chain variable region in the antibody or the like that binds to the first target antigen, or may bind to the first target antigen.
- the antibody may be a polypeptide containing the amino acid sequence of the heavy chain variable region.
- CDRL1, CDRL2, and CDRL3 in the first light chain variable region may each be, for example, a polypeptide consisting of the amino acid sequences of CDRL1, CDRL2, and CDRL3, such as an antibody that binds to the first target antigen. It may be a polypeptide containing the amino acid sequences of CDRL1, CDRL2, and CDRL3, such as an antibody that binds to the first target antigen.
- FRL1, FRL2, FRL3, and FRL4 in the first light chain variable region are, for example, polypeptides comprising amino acid sequences of FRL1, FRL2, FRL3, and FRL4, such as an antibody that binds to the first target antigen, respectively.
- it may be a polypeptide containing the amino acid sequences of FRL1, FRL2, FRL3, and FRL4, such as an antibody that binds to the first target antigen.
- the “antibody or the like that binds to the first target antigen” in the description of each CDRL and the “antibody or the like that binds to the first target antigen” in the description of each FRL are preferably the same antibody or the like.
- the first light chain variable region may include, for example, the light chain variable region in an antibody or the like that binds to the first target antigen.
- the first light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region in an antibody or the like that binds to the first target antigen, or may bind to the first target antigen.
- the antibody may be a polypeptide containing the amino acid sequence of the light chain variable region.
- first linker peptide In the first scFv, the first heavy chain variable region and the first light chain variable region are linked by, for example, a first linker peptide (first Fv linker peptide).
- the first Fv linker peptide preferably does not, for example, inhibit the binding of the first scFv to the first target antigen.
- the first Fv linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the first Fv linker peptide is composed of amino acids such as glycine and serine, and a specific example thereof is (GGGGS) n .
- the n is, for example, an integer of 1 to 6.
- the amino acid sequence of the first Fv linker peptide includes, for example, a polypeptide having the amino acid sequence of SEQ ID NO: 1 or 2.
- the second scFv contains a second heavy chain variable region and a second light chain variable region.
- the second heavy chain variable region and the second light chain variable region have the same structure as the heavy chain variable region and the light chain variable region in the antibody molecule, and the structure thereof is, for example, in (1) above.
- the description of the first scFv can be incorporated.
- the second heavy chain variable region has CDRH1, CDRH2, and CDRH3.
- the second light chain variable region has CDRL1, CDRL2, and CDRL3. Further, the second heavy chain variable region and the second light chain variable region satisfy the condition 1 or the condition 2.
- the condition 1 is that CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region use an antibody or the like that binds to the immune cell activating receptor, and CDRL1, CDRL2 in the second light chain variable region, And CDRL3 are screened for binding to the immune cell activating receptor.
- Specific examples of the antibody or the antigen-binding fragment thereof can be referred to, for example, the description of the antibody and the antigen-binding fragment in the antibody of the present invention described above.
- CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region may each be, for example, a polypeptide consisting of the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the immune cell activating receptor.
- a polypeptide containing the amino acid sequences of CDRH1, CDRH2, and CDRH3, such as an antibody that binds to the immune cell activating receptor, may be used.
- FRH1, FRH2, FRH3, and FRH4 in the second heavy chain variable region are each composed of, for example, an amino acid sequence of FRH1, FRH2, FRH3, and FRH4 of an antibody or the like that binds to the immune cell activating receptor. It may be a peptide or a polypeptide containing the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 such as an antibody that binds to the immune cell activating receptor.
- the "antibody or the like that binds to an immune cell activating receptor" in the description of each CDRH and the "antibody or the like that binds to an immune cell activating receptor" in the description of each FRH may be the same antibody or the like. preferable.
- the second heavy chain variable region may include, for example, a heavy chain variable region in an antibody or the like that binds to the immune cell activating receptor.
- the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequence of the heavy chain variable region in the antibody or the like that binds to the immune cell activating receptor, or the immune cell activating receptor It may be a polypeptide containing the amino acid sequence of the heavy chain variable region of an antibody or the like that binds to.
- the second light chain variable region is encoded by, for example, a VJ gene fragment formed by reconstructing a VJ gene and a V gene fragment. Therefore, the first B cell receptor is, for example, a B cell receptor containing a polypeptide encoded by an artificial VJ gene fragment designed by artificially combining a V gene fragment and a J gene fragment. Good. In vivo B cells, for example, express the light chain variable region after VJ gene reconstruction. Therefore, the first B cell receptor may be, for example, an isolated B cell-derived B cell receptor. In this case, the first B cell receptor may be, for example, a human-derived B cell B cell receptor, and preferably a human peripheral blood B cell-derived B cell receptor. In the BsAb library of the present invention, the first B cell receptor is preferably the light chain variable region of the isolated B cell-derived B cell receptor because the BsAb library can be prepared more easily.
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRL1, CDRL2, and CDRL3 of the first B cell receptor, respectively. It may be a polypeptide comprising the amino acid sequences of CDRH1, CDRL2 and CDRL3 of the B cell receptor of.
- FRL1, FRL2, FRL3, and FRL4 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the first B cell receptor, respectively.
- the polypeptide may include the amino acid sequences of FRL1, FRL2, FRL3, and FRL4 of the first B cell receptor.
- the "first B cell receptor" in the description of each CDRL and the "first B cell receptor" in the description of each FRL are preferably the same B cell receptor.
- the second light chain variable region may include, for example, the light chain variable region of the first B cell receptor.
- the second light chain variable region may be, for example, a polypeptide consisting of the amino acid sequence of the light chain variable region in the first B cell receptor, or the light chain in the first B cell receptor. It may be a polypeptide containing the amino acid sequence of the variable region.
- first linker peptide In the first scFv, the first heavy chain variable region and the first light chain variable region are linked by, for example, a first linker peptide (first Fv linker peptide).
- the first Fv linker peptide preferably does not, for example, inhibit the binding of the first scFv to the first target antigen.
- the first Fv linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the first Fv linker peptide is composed of amino acids such as glycine and serine, and a specific example thereof is (GGGGS) n .
- the n is, for example, an integer of 1 to 6.
- the amino acid sequence of the first Fv linker peptide includes, for example, a polypeptide having the amino acid sequence of SEQ ID NO: 1 or 2.
- the CDRL1, CDRL2, and CDRL3 in the second light chain variable region are derived from an antibody or the like that binds to the immune cell activating receptor, and thus in the second heavy chain variable region.
- CDRH1, CDRH2, and CDRH3 are screened for binding to the immune cell activating receptor.
- the second heavy chain variable region is encoded by, for example, a VDJ gene fragment formed by reconstructing the VDJ gene fragment, V gene fragment, and J gene fragment. Therefore, the first B cell receptor is, for example, a B cell containing a polypeptide encoded by an artificial VDJ gene fragment designed by artificially combining a V gene fragment, a D gene fragment, and a J gene fragment. It may be a receptor. In vivo B cells, for example, express the heavy chain variable region after VDJ gene reconstruction. Therefore, the first B cell receptor may be, for example, an isolated B cell-derived B cell receptor.
- the first B cell receptor may be, for example, a human-derived B cell B cell receptor, and preferably a human peripheral blood B cell-derived B cell receptor.
- the first B cell receptor is a heavy chain variable region of an isolated B cell-derived B cell receptor because a BsAb library can be prepared more easily.
- CDRH1, CDRH2, and CDRH3 in the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRH1, CDRH2, and CDRH3 of the first B cell receptor, respectively. It may be a polypeptide comprising the amino acid sequences of CDRH1, CDRH2 and CDRH3 of the B cell receptor of.
- FRH1, FRH2, FRH3, and FRH4 in the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 of the first B cell receptor, respectively.
- the polypeptide may include the amino acid sequences of FRH1, FRH2, FRH3, and FRH4 of the first B cell receptor.
- the “first B cell receptor” in the description of each CDRH and the “first B cell receptor” in the description of each FRH are preferably the same B cell receptor.
- the second heavy chain variable region may include, for example, the heavy chain variable region in the first B cell receptor.
- the second heavy chain variable region may be, for example, a polypeptide comprising the amino acid sequence of the heavy chain variable region in the first B cell receptor, or the heavy chain in the first B cell receptor. It may be a polypeptide containing the amino acid sequence of the variable region.
- the antibody or the like that binds to the immune cell activating receptor in the second light chain variable region is, for example, “(1) first target antigen: immune cell activating receptor” of the BsAb library of the present invention.
- the description of the antibody and the like which bind to the immune cell activating receptor can be incorporated by reference.
- CDRL1, CDRL2, and CDRL3 in the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequences of CDRL1, CDRL2, and CDRL3, such as an antibody that binds to the immune cell activating receptor, respectively.
- a polypeptide containing the amino acid sequences of CDRL1, CDRL2, and CDRL3, such as an antibody that binds to the immune cell activating receptor, may be used.
- the FR may include, for example, the FR of the light chain variable region in the antibody or the like that binds to the immune cell activating receptor.
- FRL1, FRL2, FRL3, and FRL4 in the second light chain variable region are, for example, poly-amino acid sequences of FRL1, FRL2, FRL3, and FRL4, such as an antibody that binds to the immune cell activating receptor, respectively. It may be a peptide or a polypeptide containing the amino acid sequences of FRL1, FRL2, FRL3 and FRL4 such as an antibody that binds to the immune cell activating receptor.
- the “antibody or the like that binds to an immune cell activating receptor” in the description of each CDRL and the “antibody or the like that binds to an immune cell activating receptor” in the description of each FRL may be the same antibody or the like. preferable.
- the second light chain variable region may include, for example, a light chain variable region in an antibody or the like that binds to the immune cell activating receptor.
- the second light chain variable region may be, for example, a polypeptide comprising the amino acid sequence of the light chain variable region in an antibody or the like that binds to the immune cell activating receptor, or the immune cell activating receptor It may be a polypeptide containing the amino acid sequence of the light chain variable region of an antibody or the like that binds to.
- the second heavy chain variable region and the second light chain variable region are linked by, for example, a second linker peptide (second Fv linker peptide).
- the second Fv linker peptide preferably does not, for example, inhibit the binding of the second scFv to the immune cell activating receptor.
- the description of the second Fv linker peptide can be referred to, for example, the description of the first Fv linker peptide.
- the second Fv linker peptide may be the same as or different from the first Fv linker peptide, for example.
- the first scFv and the second scFv may be linked, for example, directly or indirectly, and when the third scFv and the fourth scFv coexist, they are associated (bound). You may form so that it may be formed.
- the connection of the first scFv and the second scFv can be appropriately designed, for example, according to the type of the first BsAb. As a specific example, when the first BsAb is a tandem scFv, the first scFv and the second scFv are linked by a linker peptide (inter-Fv linker peptide).
- the inter-Fv linker peptide is composed of, for example, 1 to 40, 1 to 18, 1 to 15, 1 to 7, 1 to 3, 1 or 2 amino acids.
- the inter-Fv linker peptide is composed of amino acids such as glycine and serine, and a specific example thereof is (GGGGS) n .
- the n is, for example, an integer of 1 to 6.
- the amino acid sequence of the inter-Fv linker peptide is, for example, a polypeptide comprising the amino acid sequence of SEQ ID NO: 1 or 2.
- charges of amino acid side chains can be used.
- the association can be induced, for example, by adding a domain of an amino acid having a positive charge to one binding domain and adding a domain of an amino acid having a negative charge to the other binding domain.
- a combination of a tag sequence and a polypeptide that recognizes the tag sequence can be used.
- the arrangement order of the first scFv and the second scFv in the first BsAb is, for example, from the N-terminal side, the first scFv and the second scFv are arranged in this order.
- the second scFv and the first scFv may be arranged in this order from the N-terminal side.
- the latter can be more easily prepared, so that the latter is preferable.
- Examples of the first BsAb include a polypeptide having an amino acid sequence represented by the following formula (3) (SEQ ID NO: 158).
- examples of V 1 , V 2 , V 3 and V 4 include the following combinations (V1) to (V8).
- V 1 , V 2 , V 3 , and V 4 are preferably (V7) or (V8) below.
- V 1 , V 2 , V 3 , and V 4 are preferably the following (V5) or (V6).
- the amino acid sequence between V 1 and V 2 is the amino acid sequence of the first Fv linker peptide
- the amino acid sequence between V 2 and V 3 is the amino acid sequence of the Fv linker peptide
- V 3 and V 4 is the amino acid sequence of the second Fv linker peptide.
- the first Fv linker peptide and the second Fv linker peptide are Fv linker peptide 1, but may be each independently Fv linker peptide 2.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- General Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Immunology (AREA)
- Engineering & Computer Science (AREA)
- Biochemistry (AREA)
- Molecular Biology (AREA)
- Zoology (AREA)
- Biomedical Technology (AREA)
- Medicinal Chemistry (AREA)
- Biophysics (AREA)
- Veterinary Medicine (AREA)
- Public Health (AREA)
- Animal Behavior & Ethology (AREA)
- Biotechnology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Wood Science & Technology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Epidemiology (AREA)
- Cell Biology (AREA)
- General Engineering & Computer Science (AREA)
- Toxicology (AREA)
- Gastroenterology & Hepatology (AREA)
- Microbiology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Physics & Mathematics (AREA)
- Plant Pathology (AREA)
- Hematology (AREA)
- Crystallography & Structural Chemistry (AREA)
- Bioinformatics & Computational Biology (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Virology (AREA)
- Gynecology & Obstetrics (AREA)
- Pharmacology & Pharmacy (AREA)
- Reproductive Health (AREA)
- Pregnancy & Childbirth (AREA)
Abstract
Description
前記第1のCARは、第1の抗原結合ドメインと、第1の膜貫通ドメインと、第1の細胞内シグナル伝達ドメインとを含み、
前記第1の抗原結合ドメインは、標的抗原への結合をスクリーニングする第1の一本鎖抗体(scFv)を含み、
前記第1のscFvは、第1の重鎖可変領域と、第1の軽鎖可変領域とを含み、
前記第1の重鎖可変領域および前記第1の軽鎖可変領域は、下記条件1または条件2を満たす;
条件1:
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、第1のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む;
条件2:
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、第1のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。
前記第1の発現工程で得られた免疫細胞と、前記標的抗原とを接触させる第1の接触工程と、
前記第1の接触工程において前記標的抗原と結合した免疫細胞に発現するCARの第1のscFvを、前記標的抗原に結合する第1の候補scFvとして選抜する第1の選抜工程とを含む。
本発明のCARライブラリは、前述のように、第1のキメラ抗原受容体(CAR)をコードする核酸を含み、前記第1のCARは、第1の抗原結合ドメインと、第1の膜貫通ドメインと、第1の細胞内シグナル伝達ドメインとを含み、前記第1の抗原結合ドメインは、標的抗原への結合をスクリーニングする第1の一本鎖抗体(scFv)を含み、前記第1のscFvは、第1の重鎖可変領域と、第1の軽鎖可変領域とを含み、前記第1の重鎖可変領域および前記第1の軽鎖可変領域は、下記条件1または条件2を満たす。本発明のCARライブラリは、下記条件1または条件2を満たすことが特徴であり、その他の構成および条件は、特に制限されない。
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片(以下、「抗体等」ともいう)における重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、第1のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む;
条件2:
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、第1のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。
GSTSGSGKPGSGEGSTKG
Fvリンカーペプチド2(配列番号2)
GGGGSGGGGSGGGGS
5'-GGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGC-3'
Fvリンカーペプチド2をコードする塩基配列(配列番号4)
5'-GGTGGAGGAGGCTCAGGAGGAGGTGGCTCTGGTGGTGGAGGCTCG-3'
FWVLVVVGGVLACYSLLVTVAFIIFWV
5'-TTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTG-3'
IEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRS
5'-ATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCT-3’
RVKSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
5'-CGAGTGAAGAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
RSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRS
5'-CGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCT-3'
IEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKP
5'-ATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCT-3'
[V1]-[GSTSGSGKPGSGEGSTKG]-[V2]-[IEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRS]-[RVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR] ・・・(1)
5'-[N1]-[GGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGC]-[N2]-[ATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCT]-[CGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA]-3' ・・・(2)
本発明の第1のスクリーニング方法は、前記本発明のCARライブラリ(以下、「第1のCARライブラリ」ともいう)を免疫細胞に発現させる第1の発現工程と、前記第1の発現工程で得られた免疫細胞と、前記標的抗原とを接触させる第1の接触工程と、前記第1の接触工程において前記標的抗原と結合した免疫細胞に発現するCARの第1のscFvを、前記標的抗原に結合する第1の候補scFvとして選抜する第1の選抜工程とを含む。本発明の第1のスクリーニング方法は、前記第1の発現工程、前記第1の接触工程および前記第1の選抜工程を含み、前記第1の発現工程において、前記本発明のCARライブラリを用いることが特徴であり、その他の構成および条件は、特に制限されない。本発明の第1のスクリーニング方法によれば、CAR-T細胞において機能しうるscFvをスクリーニングできる。また、前記標的抗原に結合するscFvの重鎖可変領域および軽鎖可変領域の各CDRのアミノ酸配列に基づき、例えば、前記標的抗原に結合可能な抗体またはその抗原結合断片を作製できる。このため、前記第1のスクリーニング方法は、例えば、前記標的抗原に結合する抗体またはその抗原結合断片のスクリーニング方法ということもできる。本発明の第1のスクリーニング方法は、例えば、前記本発明のCARライブラリの説明を援用できる。本発明のscFvの製造方法によれば、新たなscFvをスクリーニングすることができる。このため、本発明のscFvの製造方法は、例えば、scFvのスクリーニング方法ということもできる。
・T細胞
活性化マーカー:CD69、CD107a等
サイトカイン・ケモカイン:IFN-γ、IL-12、IL-2、TNFα、MIP-1β等
・NK細胞
活性化マーカー:CD69、CD107a等
サイトカイン・ケモカイン:INF-γ、IL-12等
・NKT細胞
活性化マーカー:CD69、CD107a、CD25等
サイトカイン・ケモカイン:INF-γ、IL-2等
・B細胞
活性化マーカー:CD28、CD69、CD80、CD138、B220等
サイトカイン・ケモカイン:CXCR4等
前記第1の発現工程におけるCARライブラリが前記条件1を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、第2のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、前記第1の候補scFvにおける軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。
条件4:
前記第1の発現工程におけるCARライブラリが前記条件2を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、前記第1の候補scFvにおける重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、第2のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む。
本発明のキメラ抗原受容体(CAR)ライブラリ細胞は、前記本発明のCARライブラリを発現する細胞を含む。本発明のCARライブラリ細胞は、前記本発明のCARライブラリを発現する細胞を含むことが特徴であり、その他の構成および条件は、特に制限されない。本発明のCARライブラリ細胞発現細胞によれば、前記本発明の第1のスクリーニング方法を好適に実施できる。本発明のCARライブラリ細胞は、前記本発明のCARライブラリおよび第1のスクリーニング方法の説明を援用できる。
本発明のHLA-A*02:01およびNY-ESO-1157-165の複合体(以下、「A2/NY-ESO-1157」ともいう)に対する抗体またはその抗原結合断片は、前述のように、下記(H)の重鎖可変領域と、下記(L)の軽鎖可変領域とを含む。本発明の抗体等は、下記(H)の重鎖可変領域と、下記(L)の軽鎖可変領域とを含むことが特徴であり、その他の構成および条件は、特に制限されない。本発明の抗体等は、A2/NY-ESO-1157に結合する。また、A2/NY-ESO-1157は、例えば、肺がん、悪性黒色腫、滑膜肉腫、骨髄腫等の特定のがん細胞に発現することが知られている。このため、本発明の抗体等は、例えば、A2/NY-ESO-1157発現がん細胞に対する二重特異性抗体、CAR-T細胞のCARの抗原結合ドメイン等として好適に使用できる。前記本発明のCARライブラリ、第1のスクリーニング方法等の説明を援用できる。
CDRH1が、下記(H1)のアミノ酸配列を含むポリペプチドであり、
CDRH2が、下記(H2)のアミノ酸配列を含むポリペプチドであり、
CDRH3が、下記(H3)のアミノ酸配列を含むポリペプチドである重鎖可変領域
(H1)下記(H1-1)、(H1-2)または(H1-3)のアミノ酸配列
(H1-1)下記表1AのCDRH1のいずれかのアミノ酸配列
(H1-2)(H1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-3)(H1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H2)下記(H2-1)、(H2-2)または(H2-3)のアミノ酸配列
(H2-1)下記表1AのCDRH2のいずれかのアミノ酸配列
(H2-2)(H2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-3)(H2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H3)下記(H3-1)、(H3-2)または(H3-3)のアミノ酸配列
(H3-1)下記表1AのCDRH3のいずれかのアミノ酸配列
(H3-2)(H3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-3)(H3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
CDRL1が、下記(L1)のアミノ酸配列を含むポリペプチドであり、
CDRL2が、下記(L2)のアミノ酸配列を含むポリペプチドであり、
CDRL3が、下記(L3)のアミノ酸配列を含むポリペプチドである軽鎖可変領域
(L1)下記(L1-1)、(L1-2)または(L1-3)のアミノ酸配列
(L1-1)下記表1BのCDRL1のいずれかのアミノ酸配列
(L1-2)(L1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-3)(L1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2)下記(L2-1)、(L2-2)または(L2-3)のアミノ酸配列
(L2-1)下記表1BのCDRL2のいずれかのアミノ酸配列
(L2-2)(L2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-3)(L2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3)下記(L3-1)、(L3-2)または(L3-3)のアミノ酸配列
(L3-1)下記表1BのCDRL3のいずれかのアミノ酸配列
(L3-2)(L3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-3)(L3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(1)の抗体等は、例えば、抗体H1-3M4E5L群ともいう。前記組合せ(1)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、下記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、下記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、下記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LA)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-A)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-A)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-A)のアミノ酸配列を含むポリペプチドである。
(H1-A1)配列番号17(GGSISSNY)のアミノ酸配列
(H1-A2)配列番号17のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-A3)配列番号17のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H2-A1)配列番号18(VSYSGST)のアミノ酸配列
(H2-A2)配列番号18のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-A3)配列番号18のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H3-A1)配列番号19(ARESYYYYGMDV)のアミノ酸配列
(H3-A2)配列番号19のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-A3)配列番号19のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L1-A1)配列番号29(SRDVGGYNY)のアミノ酸配列
(L1-A2)配列番号29のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-A3)配列番号29のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-A1)配列番号30(DVI)のアミノ酸配列
(L2-A2)配列番号30のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-A3)配列番号30のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-A1)配列番号31(WSFAGSYYV)のアミノ酸配列
(L3-A2)配列番号31のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-A3)配列番号31のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H-A1)配列番号20のアミノ酸配列
配列番号20:QVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
(H-A2)配列番号20のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H-A3)配列番号20のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-A1)配列番号32のアミノ酸配列
配列番号32:QSELTQPRSVSGSPGQSVTISCTGTSRDVGGYNYVSWYQQHPGKAPKLIIHDVIERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCWSFAGSYYVFGTGTDVTVL
(L-A2)配列番号32のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-A3)配列番号32のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(2)の抗体等は、例えば、抗体H1-K52群ともいう。前記組合せ(2)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LB)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-B)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-B)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-B)のアミノ酸配列を含むポリペプチドである。
(L1-B1)配列番号33(QSISSY)のアミノ酸配列
(L1-B2)配列番号33のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-B3)配列番号33のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-B1)配列番号34(AAS)のアミノ酸配列
(L2-B2)配列番号34のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-B3)配列番号34のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-B1)配列番号35(QQYYSTPQT)のアミノ酸配列
(L3-B2)配列番号35のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-B3)配列番号35のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-B1)配列番号36のアミノ酸配列
配列番号36:DIQMTQSPSAMSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQYYSTPQTFGPGTKVDIK
(L-B2)配列番号36のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-B3)配列番号36のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(3)の抗体等は、例えば、抗体H1-K73群ともいう。前記組合せ(3)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LC)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-C)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-C)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-C)のアミノ酸配列を含むポリペプチドである。
(L1-C1)配列番号37(QSISSY)のアミノ酸配列
(L1-C2)配列番号37のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-C3)配列番号37のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-C1)配列番号38(AAS)のアミノ酸配列
(L2-C2)配列番号38のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-C3)配列番号38のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-C1)配列番号39(QQYESYRRS)のアミノ酸配列
(L3-C2)配列番号39のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-C3)配列番号39のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-C1)配列番号40のアミノ酸配列
配列番号40:DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKAGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYCQQYESYRRSFGQGTKVEIK
(L-C2)配列番号40のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-C3)配列番号40のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(4)の抗体等は、例えば、抗体H1-K121-K124群ともいう。前記組合せ(4)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LD)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-D)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-D)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-D)のアミノ酸配列を含むポリペプチドである。
(L1-D1)配列番号41(QSISSY)のアミノ酸配列
(L1-D2)配列番号41のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-D3)配列番号41のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-D1)配列番号42(AAS)のアミノ酸配列
(L2-D2)配列番号42のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-D3)配列番号42のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-D1)配列番号43(QQYNSYSRT)のアミノ酸配列
(L3-D2)配列番号43のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-D3)配列番号43のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-DA1)配列番号44のアミノ酸配列
配列番号44:DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDFATYYCQQYNSYSRTFGQGTKVEIK
(L-DA2)配列番号44のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-DA3)配列番号44のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-DB1)配列番号45のアミノ酸配列
配列番号45:DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASRLESGVPSRFSGSGSGTDFTLTISCLQSEDFATYYCQQYNSYSRTFGQGTKVEIK
(L-DB2)配列番号45のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-DB3)配列番号45のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(5)の抗体等は、例えば、抗体H1-K125群ともいう。前記組合せ(5)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LE)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-E)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-E)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-E)のアミノ酸配列を含むポリペプチドである。
(L1-E1)配列番号46(QSISSY)のアミノ酸配列
(L1-E2)配列番号46のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-E3)配列番号46のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-E1)配列番号47(AAS)のアミノ酸配列
(L2-E2)配列番号47のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-E3)配列番号47のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-E1)配列番号48(QQYNSYSPCT)のアミノ酸配列
(L3-E2)配列番号48のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-E3)配列番号48のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-E1)配列番号49のアミノ酸配列
配列番号49:DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTEFTLTISSLQPDDFATYYCQQYNSYSPCTFGPGTKVDIK
(L-E2)配列番号49のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-E3)配列番号49のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(6)の抗体等は、例えば、抗体H1-K131群ともいう。前記組合せ(6)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LF)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-F)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-F)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-F)のアミノ酸配列を含むポリペプチドである。
(L1-F1)配列番号50(QDISRY)のアミノ酸配列
(L1-F2)配列番号50のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-F3)配列番号50のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-F1)配列番号51(AAS)のアミノ酸配列
(L2-F2)配列番号51のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-F3)配列番号51のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-F1)配列番号52(QQYDNLIT)のアミノ酸配列
(L3-F2)配列番号52のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-F3)配列番号52のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-F1)配列番号53のアミノ酸配列
配列番号53:DIQMTQSPSSLSASVGDRVSITCRASQDISRYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYDNLITFGQGTRLEIK
(L-F2)配列番号53のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-F3)配列番号53のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(7)の抗体等は、例えば、抗体H1-K145群ともいう。前記組合せ(7)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LG)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-G)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-G)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-G)のアミノ酸配列を含むポリペプチドである。
(L1-G1)配列番号54(QDISRY)のアミノ酸配列
(L1-G2)配列番号54のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-G3)配列番号54のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-G1)配列番号55(AAS)のアミノ酸配列
(L2-G2)配列番号55のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-G3)配列番号55のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-G1)配列番号56(QQYNSYSRT)のアミノ酸配列
(L3-G2)配列番号56のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-G3)配列番号56のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-G1)配列番号57のアミノ酸配列
配列番号57:DIQMTQSPSSLSASVGDRVSITCRASQDISRYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYNSYSRTFGQGTKVEIK
(L-G2)配列番号57のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-G3)配列番号57のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(8)の抗体等は、例えば、抗体H1-K151群ともいう。前記組合せ(8)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LH)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-H)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-H)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-H)のアミノ酸配列を含むポリペプチドである。
(L1-H1)配列番号58(QSISSY)のアミノ酸配列
(L1-H2)配列番号58のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-H3)配列番号58のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-H1)配列番号59(AAS)のアミノ酸配列
(L2-H2)配列番号59のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-H3)配列番号59のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-H1)配列番号60(QQYDNLIT)のアミノ酸配列
(L3-H2)配列番号60のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-H3)配列番号60のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-H1)配列番号61のアミノ酸配列
配列番号61:DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYDNLITFGQGTRLEIK
(L-H2)配列番号61のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-H3)配列番号61のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(9)の抗体等は、例えば、抗体H1-K160群ともいう。前記組合せ(9)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LI)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-I)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-I)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-I)のアミノ酸配列を含むポリペプチドである。
(L1-I1)配列番号62(QSVSSN)のアミノ酸配列
(L1-I2)配列番号62のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-I3)配列番号62のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-I1)配列番号63(GAS)のアミノ酸配列
(L2-I2)配列番号63のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-I3)配列番号63のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-I1)配列番号64(QQYNSYSRT)のアミノ酸配列
(L3-I2)配列番号64のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-I3)配列番号64のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-I1)配列番号65のアミノ酸配列
配列番号65:EIVLTQSPATLSVSPGERATLSCRASQSVSSNLAWYQQKPGQAPRLLIYGASTRATGIPARFSGSGSGTEFTLTISRLEPEDFATYYCQQYNSYSRTFGQGTKVEIK
(L-I2)配列番号65のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-I3)配列番号65のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(10)の抗体等は、例えば、抗体H1-K173群ともいう。前記組合せ(10)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LJ)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-J)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-J)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-J)のアミノ酸配列を含むポリペプチドである。
(L1-J1)配列番号66(QSISSY)のアミノ酸配列
(L1-J2)配列番号66のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-J3)配列番号66のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-J1)配列番号67(AAS)のアミノ酸配列
(L2-J2)配列番号67のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-J3)配列番号67のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-J1)配列番号68(QQYESYSRT)のアミノ酸配列
(L3-J2)配列番号68のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-J3)配列番号68のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-J1)配列番号69のアミノ酸配列
配列番号69:DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKAGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYCQQYESYSRTFGQGTKVEIK
(L-J2)配列番号69のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-J3)配列番号69のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(11)の抗体等は、例えば、抗体3M4E5H-L1群ともいう。前記組合せ(11)において、前記(HB)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、下記(H1-B)のアミノ酸配列を含むポリペプチドであり、CDRH2が、下記(H2-B)のアミノ酸配列を含むポリペプチドであり、CDRH3が、下記(H3-B)のアミノ酸配列を含むポリペプチドである。また、前記(LK)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-K)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-K)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-K)のアミノ酸配列を含むポリペプチドである。
(H1-B1)配列番号21(GFTFSTYQ)のアミノ酸配列
(H1-B2)配列番号21のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-B3)配列番号21のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H2-B1)配列番号22(IVSSGGST)のアミノ酸配列
(H2-B2)配列番号22のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-B3)配列番号22のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H3-B1)配列番号23(AGELLPYYGMDV)のアミノ酸配列
(H3-B2)配列番号23のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-B3)配列番号23のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L1-K1)配列番号70(SSDVGGYDF)のアミノ酸配列
(L1-K2)配列番号70のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-K3)配列番号70のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-K1)配列番号71(DVN)のアミノ酸配列
(L2-K2)配列番号71のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-K3)配列番号71のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-K1)配列番号72(SSYAGSNSV)のアミノ酸配列
(L3-K2)配列番号72のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-K3)配列番号72のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H-B1)配列番号24のアミノ酸配列
配列番号24:EVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
(H-B2)配列番号24のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H-B3)配列番号24のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-K1)配列番号73のアミノ酸配列
配列番号73:QSALTQPPSASGSPGQSVTISCTGTSSDVGGYDFVSWYQQHPGEAPKLLVYDVNNRPSGVSNRFSGSKSGNTASLTISGLQAEDEGDYYCSSYAGSNSVFGTGTKVTVL
(L-K2)配列番号73のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-K3)配列番号73のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(12)の抗体等は、例えば、抗体3M4E5H-L66群ともいう。前記組合せ(12)において、前記(HB)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-B)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-B)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-B)のアミノ酸配列を含むポリペプチドである。また、前記(LL)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-L)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-L)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-L)のアミノ酸配列を含むポリペプチドである。
(L1-L1)配列番号74(SSDVGGYEF)のアミノ酸配列
(L1-L2)配列番号74のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-L3)配列番号74のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-L1)配列番号75(DVI)のアミノ酸配列
(L2-L2)配列番号75のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-L3)配列番号75のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-L1)配列番号76(SSYTSSSTYV)のアミノ酸配列
(L3-L2)配列番号76のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-L3)配列番号76のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-L1)配列番号77のアミノ酸配列
配列番号77:QSALTQPASVSGSPGQSITISCTGTSSDVGGYEFVSWYQQHPGSAPKLIIYDVIERPFGVSYRFSASKSGNTASLTISGLQGEDEADYFCSSYTSSSTYVFGTGTKVTVL
(L-L2)配列番号77のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-L3)配列番号77のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(13)の抗体等は、例えば、抗体3M4E5H-L73群ともいう。前記組合せ(13)において、前記(HB)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-B)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-B)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-B)のアミノ酸配列を含むポリペプチドである。また、前記(LM)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-M)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-M)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-M)のアミノ酸配列を含むポリペプチドである。
(L1-M1)配列番号78(GSDVGAYDY)のアミノ酸配列
(L1-M2)配列番号78のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-M3)配列番号78のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-M1)配列番号79(DVS)のアミノ酸配列
(L2-M2)配列番号79のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-M3)配列番号79のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-M1)配列番号80(SSYSGSSTWV)のアミノ酸配列
(L3-M2)配列番号80のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-M3)配列番号80のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-M1)配列番号81のアミノ酸配列
配列番号81:QSALTQPASVSGSPGQSITISCTGTGSDVGAYDYVSWYQHHPGRAPRLIIRDVSVRPSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCSSYSGSSTWVFGGGTKLTVL
(L-M2)配列番号81のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-M3)配列番号81のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(14)の抗体等は、例えば、抗体3M4E5H-L80群ともいう。前記組合せ(14)において、前記(HB)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-B)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-B)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-B)のアミノ酸配列を含むポリペプチドである。また、前記(LN)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-N)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-N)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-N)のアミノ酸配列を含むポリペプチドである。
(L1-N1)配列番号82(SSDVGSYNL)のアミノ酸配列
(L1-N2)配列番号82のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-N3)配列番号82のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-N1)配列番号83(DVS)のアミノ酸配列
(L2-N2)配列番号83のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-N3)配列番号83のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-N1)配列番号84(SSYTSSSTFAV)のアミノ酸配列
(L3-N2)配列番号84のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-N3)配列番号84のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-N1)配列番号85のアミノ酸配列
配列番号85:QSALTQPASVSGSPGQSITISCTGTSSDVGSYNLVSWYQQHPGKAPKLMIYDVSNRPSGVSYRFSGSKSGNTASLTISGLQAEDEADYYCSSYTSSSTFAVFGGGTQLTVL
(L-N2)配列番号85のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-N3)配列番号85のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(15)の抗体等は、例えば、抗体3M4E5H-L88群ともいう。前記組合せ(15)において、前記(HB)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-B)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-B)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-B)のアミノ酸配列を含むポリペプチドである。また、前記(LO)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-O)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-O)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-O)のアミノ酸配列を含むポリペプチドである。
(L1-O1)配列番号86(SSDVGGYNY)のアミノ酸配列
(L1-O2)配列番号86のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-O3)配列番号86のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-O1)配列番号87(DVS)のアミノ酸配列
(L2-O2)配列番号87のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-O3)配列番号87のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-O1)配列番号88(CSYAGGYYV)のアミノ酸配列
(L3-O2)配列番号88のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-O3)配列番号88のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-O1)配列番号89のアミノ酸配列
配列番号89:QSALPQPASVSGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSGNTASLTVSGLQAEDEADYFCCSYAGGYYVFGTGTKLTVL
(L-O2)配列番号89のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-O3)配列番号89のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(16)の抗体等は、例えば、抗体3M4E5H-L102群ともいう。前記組合せ(16)において、前記(HB)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-B)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-B)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-B)のアミノ酸配列を含むポリペプチドである。また、前記(LP)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-P)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-P)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-P)のアミノ酸配列を含むポリペプチドである。
(L1-P1)配列番号90(SSDVGGYNY)のアミノ酸配列
(L1-P2)配列番号90のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-P3)配列番号90のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-P1)配列番号91(DVS)のアミノ酸配列
(L2-P2)配列番号91のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-P3)配列番号91のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-P1)配列番号92(SSYAGSGSTPFV)のアミノ酸配列
(L3-P2)配列番号92のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-P3)配列番号92のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-P1)配列番号93のアミノ酸配列
配列番号93:QSALTQPPSASGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSGNTASLTISGLQTEDEADYYCSSYAGSGSTPFVFGTGTKLTVL
(L-P2)配列番号93のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-P3)配列番号93のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(17)の抗体等は、例えば、抗体3M4E5H-L124群ともいう。前記組合せ(17)において、前記(HB)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-B)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-B)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-B)のアミノ酸配列を含むポリペプチドである。また、前記(LQ)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-Q)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-Q)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-Q)のアミノ酸配列を含むポリペプチドである。
(L1-Q1)配列番号94(SSDVGGYNY)のアミノ酸配列
(L1-Q2)配列番号94のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-Q3)配列番号94のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-Q1)配列番号95(DVS)のアミノ酸配列
(L2-Q2)配列番号95のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-Q3)配列番号95のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-Q1)配列番号96(CSYAGRRYV)のアミノ酸配列
(L3-Q2)配列番号96のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-Q3)配列番号96のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-Q1)配列番号97のアミノ酸配列
配列番号97:QSALTQPASVSGSPGQSITISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSRVPDRFAGSKSGNTASLTISGLQAEDEADYYCCSYAGRRYVFGTGTKLTVL
(L-Q2)配列番号97のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-Q3)配列番号97のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(18)の抗体等は、例えば、抗体H73-3M4E5L群ともいう。前記組合せ(18)において、前記(HC)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、下記(H1-C)のアミノ酸配列を含むポリペプチドであり、CDRH2が、下記(H2-C)のアミノ酸配列を含むポリペプチドであり、CDRH3が、下記(H3-C)のアミノ酸配列を含むポリペプチドである。また、前記(LA)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、前記(L1-A)のアミノ酸配列を含むポリペプチドであり、CDRL2が、前記(L2-A)のアミノ酸配列を含むポリペプチドであり、CDRL3が、前記(L3-A)のアミノ酸配列を含むポリペプチドである。
(H1-C1)配列番号25(GGSISSYY)のアミノ酸配列
(H1-C2)配列番号25のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-C3)配列番号25のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H2-C1)配列番号26(INHSGST)のアミノ酸配列
(H2-C2)配列番号26のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-C3)配列番号26のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H3-C1)配列番号27(ARCPIYYYGMDV)のアミノ酸配列
(H3-C2)配列番号27のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-C3)配列番号27のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H-C1)配列番号28のアミノ酸配列
配列番号28:QLQLQESGPGLVKPSQTLSLTCTVSGGSISSYYWSWIRQPPGKGLEWIGEINHSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARCPIYYYGMDVWGQGTTVTVSS
(H-C2)配列番号28のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H-C3)配列番号28のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
CDRH1が、下記(H1)のアミノ酸配列を含むポリペプチドであり、
CDRH2が、下記(H2)のアミノ酸配列を含むポリペプチドであり、
CDRH3が、下記(H3)のアミノ酸配列を含むポリペプチドである重鎖可変領域;
(H1)下記(H1-1)、(H1-2)または(H1-3)のアミノ酸配列
(H1-1)下記条件(H1)のCDRH1のいずれかのアミノ酸配列
(H1-2)(H1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-3)(H1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
(H2)下記(H2-1)、(H2-2)または(H2-3)のアミノ酸配列
(H2-1)下記条件(H1)のCDRH2のいずれかのアミノ酸配列
(H2-2)(H2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-3)(H2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
(H3)下記(H3-1)、(H3-2)または(H3-3)のアミノ酸配列
(H3-1)下記条件(H1)のCDRH3のいずれかのアミノ酸配列
(H3-2)(H3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-3)(H3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
条件(H1)
CDRH1は、GX1X2X3SX4X5X6(配列番号322)であり、
CDRH2は、X7X8X9X10SX11GST(配列番号323)であり、
CDRH3は、AX12X13X14X15X16GMDV(配列番号324)であり、
X1は、GまたはFであり、
X2は、SまたはTであり、
X3は、IまたはFであり、
X4は、SまたはTであり、
X5は、NまたはYであり、
X6は、YまたはQであり、
X7は、存在しないか、Iであり、
X8は、VまたはNであり、
X9は、SまたはHであり、
X10は、存在しないか、Yであり、
X11は、存在しないか、Gであり、
X12は、RまたはGであり、
X13は、EまたはCであり、
X14は、S、L、またはPであり、
X15は、Y、L、またはIであり、
X16は、YまたはPである。
CDRL1が、下記(L1)のアミノ酸配列を含むポリペプチドであり、
CDRL2が、下記(L2)のアミノ酸配列を含むポリペプチドであり、
CDRL3が、下記(L3)のアミノ酸配列を含むポリペプチドである軽鎖可変領域;
(L1)下記(L1-1)、(L1-2)または(L1-3)のアミノ酸配列
(L1-1)下記条件(L1)および(L2)のCDRL1のいずれかのアミノ酸配列
(L1-2)(L1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-3)(L1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
(L2)下記(L2-1)、(L2-2)または(L2-3)のアミノ酸配列
(L2-1)下記条件(L1)および(L2)のCDRL2のいずれかのアミノ酸配列
(L2-2)(L2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-3)(L2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
(L3)下記(L3-1)、(L3-2)または(L3-3)のアミノ酸配列
(L3-1)下記条件(L1)および(L2)のCDRL3のいずれかのアミノ酸配列
(L3-2)(L3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-3)(L3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
条件(L1)
CDRL1は、QX1X2SX3X4(配列番号313)であり、
CDRL2は、AAS(配列番号34)またはGAS(配列番号63)であり、
CDRL3は、QQYX5X6X7X8X9X10X11(配列番号314)であり、
X1は、SまたはDであり、
X2は、IまたはVであり、
X3は、SまたはRであり、
X4は、YまたはNであり、
X5は、Y、E、N、またはDであり、
X6は、SまたはNであり、
X7は、存在しないか、T、Y、またはSであり、
X8は、P、S、R、またはLであり、
X9は、Q、R、P、またはIであり、
X10は、存在しないか、SまたはCであり、
X11は、存在しないか、Tである;
条件(L2)
CDRL1は、X1X2DVGX3YX4X5(配列番号315)であり、
CDRL2は、DVN(配列番号71)、DVI(配列番号75)、またはDVS(配列番号79)であり、
CDRL3は、X6SX7X8X9X10X11X12X13X14X15V(配列番号316)であり、
X1は、SまたはGであり、
X2は、RまたはSであり、
X3は、G、A、またはSであり、
X4は、N、D、E、またはQであり、
X5は、Y、F、L、またはWであり、
X6は、W、S、またはCであり、
X7は、F、Y、またはWであり、
X8は、存在しないか、AまたはTであり、
X9は、存在しないか、Gであり、
X10は、存在しないか、Sであり、
X11は、GまたはSであり、
X12は、存在しないか、Sであり、
X13は、S、T、またはRであり、
X14は、Y、N、T、F、P、またはRであり、
X15は、Y、S、W、A、またはFである。
本発明のCD19に対する抗体またはその抗原結合断片(以下、「第2の抗体等」ともいう)は、下記(H)の重鎖可変領域と、下記(L)の軽鎖可変領域とを含む。本発明の抗体等は、下記(H)の重鎖可変領域と、下記(L)の軽鎖可変領域とを含むことが特徴であり、その他の構成および条件は、特に制限されない。本発明の抗体等は、CD19に結合する。また、CD19は、例えば、正常なB細胞およびB細胞系のリンパ腫等の特定のがん細胞に発現することが知られている。このため、本発明の抗体等は、例えば、CD19発現がん細胞に対する二重特異性抗体、CAR-T細胞のCARの抗原結合ドメイン等として好適に使用できる。前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体またはその抗原結合断片等の説明を援用できる。
CDRH1が、下記(H1)のアミノ酸配列を含むポリペプチドであり、
CDRH2が、下記(H2)のアミノ酸配列を含むポリペプチドであり、
CDRH3が、下記(H3)のアミノ酸配列を含むポリペプチドである重鎖可変領域
(H1)下記(H1-1)、(H1-2)または(H1-3)のアミノ酸配列
(H1-1)下記表3AのCDRH1のいずれかのアミノ酸配列
(H1-2)(H1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-3)(H1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H2)下記(H2-1)、(H2-2)または(H2-3)のアミノ酸配列
(H2-1)下記表3AのCDRH2のいずれかのアミノ酸配列
(H2-2)(H2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-3)(H2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H3)下記(H3-1)、(H3-2)または(H3-3)のアミノ酸配列
(H3-1)下記表3AのCDRH3のいずれかのアミノ酸配列
(H3-2)(H3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-3)(H3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
CDRL1が、下記(L1)のアミノ酸配列を含むポリペプチドであり、
CDRL2が、下記(L2)のアミノ酸配列を含むポリペプチドであり、
CDRL3が、下記(L3)のアミノ酸配列を含むポリペプチドである軽鎖可変領域
(L1)下記(L1-1)、(L1-2)または(L1-3)のアミノ酸配列
(L1-1)下記表3BのCDRL1のいずれかのアミノ酸配列
(L1-2)(L1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-3)(L1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2)下記(L2-1)、(L2-2)または(L2-3)のアミノ酸配列
(L2-1)下記表3BのCDRL2のいずれかのアミノ酸配列
(L2-2)(L2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-3)(L2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3)下記(L3-1)、(L3-2)または(L3-3)のアミノ酸配列
(L3-1)下記表3BのCDRL3のいずれかのアミノ酸配列
(L3-2)(L3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-3)(L3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LA)の抗体等は、例えば、抗体18H-L4群ともいう。前記組合せ(LA)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、下記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、下記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、下記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LA)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-A)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-A)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-A)のアミノ酸配列を含むポリペプチドである。
(H1-A1)配列番号216(GFTFDDYA)のアミノ酸配列
(H1-A2)配列番号216のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-A3)配列番号216のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H2-A1)配列番号217(ISWNSGRI)のアミノ酸配列
(H2-A2)配列番号217のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-A3)配列番号217のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H3-A1)配列番号218(ARDQGYHYYDSAEHAFDI)のアミノ酸配列
(H3-A2)配列番号218のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-A3)配列番号218のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L1-A1)配列番号220(KLGDKY)のアミノ酸配列
(L1-A2)配列番号220のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-A3)配列番号220のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-A1)配列番号221(QDS)のアミノ酸配列
(L2-A2)配列番号221のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-A3)配列番号221のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-A1)配列番号222(QAWDSSTHVV)のアミノ酸配列
(L3-A2)配列番号222のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-A3)配列番号222のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H-A1)配列番号219のアミノ酸配列
配列番号219:EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
(H-A2)配列番号219のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H-A3)配列番号219のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-A1)配列番号223のアミノ酸配列
配列番号223:SYELTQPPSVSVSPGQTASITCSGDKLGDKYACWYQQKPGQSPVLVIYQDSKRPSGIPERFSGSNSGNTATLTISGTQAMDEADYYCQAWDSSTHVVFGGGTKLTVL
(L-A2)配列番号223のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-A3)配列番号223のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LB)の抗体等は、例えば、抗体18H-L7群ともいう。前記組合せ(LB)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LB)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-B)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-B)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-B)のアミノ酸配列を含むポリペプチドである。
(L1-B1)配列番号224(SSDVGGYNY)のアミノ酸配列
(L1-B2)配列番号224のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-B3)配列番号224のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-B1)配列番号225(DVS)のアミノ酸配列
(L2-B2)配列番号225のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-B3)配列番号225のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-B1)配列番号226(GTWDTSLTAVV)のアミノ酸配列
(L3-B2)配列番号226のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-B3)配列番号226のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-B1)配列番号227のアミノ酸配列
配列番号227:QSALTQPRSVSGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSGVSNRFSGSKSGNTASLTISGLQTGDEADYYCGTWDTSLTAVVFGGGTELTVL
(L-B2)配列番号227のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-B3)配列番号227のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LC)の抗体等は、例えば、抗体18H-L9群ともいう。前記組合せ(LC)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LC)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-C)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-C)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-C)のアミノ酸配列を含むポリペプチドである。
(L1-C1)配列番号228(WSNIGDDH)のアミノ酸配列
(L1-C2)配列番号228のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-C3)配列番号228のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-C1)配列番号229(DTS)のアミノ酸配列
(L2-C2)配列番号229のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-C3)配列番号229のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-C1)配列番号230(GTWESSLSGVV)のアミノ酸配列
(L3-C2)配列番号230のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-C3)配列番号230のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-C1)配列番号231のアミノ酸配列
配列番号231:QSVLTQPPSVSAAPGQKVTISCSGSWSNIGDDHVSWYQQFPGAAPKLLIYDTSKRPSRVADRFSGSKSGASATLAITGLQAGDEADYYCGTWESSLSGVVFGGGTELTVL
(L-C2)配列番号231のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-C3)配列番号231のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LC)の抗体等は、例えば、抗体18H-L13群ともいう。前記組合せ(LD)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LD)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-D)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-D)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-D)のアミノ酸配列を含むポリペプチドである。
(L1-D1)配列番号232(SSDVGGYDY)のアミノ酸配列
(L1-D2)配列番号232のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-D3)配列番号232のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-D1)配列番号233(DVT)のアミノ酸配列
(L2-D2)配列番号233のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-D3)配列番号233のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-D1)配列番号234(SSYTTSTTWV)のアミノ酸配列
(L3-D2)配列番号234のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-D3)配列番号234のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-D1)配列番号235のアミノ酸配列
配列番号235:QSALTQPRSVSGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSGVSNRFSGSKSGNTASLTISGLQTGDEADYYCGTWDTSLTAVVFGGGTELTVL
(L-D2)配列番号235のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-D3)配列番号235のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LE)の抗体等は、例えば、抗体18H-L14群ともいう。前記組合せ(LE)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LE)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-E)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-E)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-E)のアミノ酸配列を含むポリペプチドである。
(L1-E1)配列番号236(TSDVGTTNY)のアミノ酸配列
(L1-E2)配列番号236のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-E3)配列番号236のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-E1)配列番号237(DVT)のアミノ酸配列
(L2-E2)配列番号237のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-E3)配列番号237のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-E1)配列番号238(SYAGSYTFVV)のアミノ酸配列
(L3-E2)配列番号238のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-E3)配列番号238のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-E1)配列番号239のアミノ酸配列
配列番号239:QSALTQPASVSGSPGQSITISCTGTTSDVGTTNYVSWYQQHPGKAPKLLIYDVTNRPSGVPDRFSGSKSANTASLTISGLQAEDEADYYCCSYAGSYTFVVFGGGTELTVL
(L-E2)配列番号239のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-E3)配列番号239のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LF)の抗体等は、例えば、抗体18H-L16群ともいう。前記組合せ(LF)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LF)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-F)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-F)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-F)のアミノ酸配列を含むポリペプチドである。
(L1-F1)配列番号240(SSDVGVYNY)のアミノ酸配列
(L1-F2)配列番号240のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-F3)配列番号240のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-F1)配列番号241(DVS)のアミノ酸配列
(L2-F2)配列番号241のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-F3)配列番号241のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-F1)配列番号242(AAWDDSLNGVV)のアミノ酸配列
(L3-F2)配列番号242のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-F3)配列番号242のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-F1)配列番号243のアミノ酸配列
配列番号243:QSALTQPPSASGSPGQSVTISCTGTSSDVGVYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSANTASLTISGLQAEDEADYYCAAWDDSLNGVVFGGGTQLTVL
(L-F2)配列番号243のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-F3)配列番号243のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LG)の抗体等は、例えば、抗体18H-L17群ともいう。前記組合せ(LG)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LG)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-G)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-G)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-G)のアミノ酸配列を含むポリペプチドである。
(L1-G1)配列番号244(SSNIGNNY)のアミノ酸配列
(L1-G2)配列番号244のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-G3)配列番号244のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-G1)配列番号245(DNV)のアミノ酸配列
(L2-G2)配列番号245のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-G3)配列番号245のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-G1)配列番号246(AAWDDSLSAI)のアミノ酸配列
(L3-G2)配列番号246のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-G3)配列番号246のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-G1)配列番号247のアミノ酸配列
配列番号247:QSVLTQPPSASGTPGQRVTISCSGSSSNIGNNYVCWYQHLPGTAPKLLIYDNVKRPSGIPDRFSGSKSGTSASLAISGLRSEDEADYYCAAWDDSLSAIFGGGTELTVL
(L-G2)配列番号247のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-G3)配列番号247のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LH)の抗体等は、例えば、抗体18H-L22群ともいう。前記組合せ(LH)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LH)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-H)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-H)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-H)のアミノ酸配列を含むポリペプチドである。
(L1-H1)配列番号248(SSDVGGYNY)のアミノ酸配列
(L1-H2)配列番号248のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-H3)配列番号248のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-H1)配列番号249(DVS)のアミノ酸配列
(L2-H2)配列番号249のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-H3)配列番号249のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-H1)配列番号250(HSYDSSLSHV)のアミノ酸配列
(L3-H2)配列番号250のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-H3)配列番号250のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-H1)配列番号251のアミノ酸配列
配列番号251:QSALTQPASVSGSPGQSITISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSGVPDRFSGSKSGTSASLAISGLQSEDEADYYCHSYDSSLSHVFGTGTKVTVL
(L-H2)配列番号251のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-H3)配列番号251のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LI)の抗体等は、例えば、抗体H1-K4群ともいう。前記組合せ(LI)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LI)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-I)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-I)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-I)のアミノ酸配列を含むポリペプチドである。
(L1-I1)配列番号252(QSVSSN)のアミノ酸配列
(L1-I2)配列番号252のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-I3)配列番号252のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-I1)配列番号253(GAS)のアミノ酸配列
(L2-I2)配列番号253のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-I3)配列番号253のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-I1)配列番号254(QQYNNWPPLYT)のアミノ酸配列
(L3-I2)配列番号254のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-I3)配列番号254のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-I1)配列番号255のアミノ酸配列
配列番号255:EIVLTQSPGTLSLSPGERATLSCRASQSVSSNLAWYQQKPGQAPRLLIYGASTRATGIPARFSGSGSGTEFTLTISSLQSEDFAVYYCQQYNNWPPLYTFGQGTKLEIK
(L-I2)配列番号255のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-I3)配列番号255のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LJ)の抗体等は、例えば、抗体18H-K5群ともいう。前記組合せ(LJ)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LJ)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-J)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-J)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-J)のアミノ酸配列を含むポリペプチドである。
(L1-J1)配列番号256(QSVSSY)のアミノ酸配列
(L1-J2)配列番号256のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-J3)配列番号256のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-J1)配列番号257(DAS)のアミノ酸配列
(L2-J2)配列番号257のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-J3)配列番号257のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-J1)配列番号258(QQSYSTLLYT)のアミノ酸配列
(L3-J2)配列番号258のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-J3)配列番号258のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-J1)配列番号259のアミノ酸配列
配列番号259:EIVLTQSPGTLSLSPGERATLSCRASQSVSSYLAWYQQKPGQAPRLLIYDASNRATGIPARFSGSGSGTDFTLTISSLEPEDFATYYCQQSYSTLLYTFGQGTKLEIK
(L-J2)配列番号259のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-J3)配列番号259のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LK)の抗体等は、例えば、抗体18H-K6群ともいう。前記組合せ(LK)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LK)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-K)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-K)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-K)のアミノ酸配列を含むポリペプチドである。
(L1-K1)配列番号260(QSVSSY)のアミノ酸配列
(L1-K2)配列番号260のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-K3)配列番号260のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-K1)配列番号261(DAS)のアミノ酸配列
(L2-K2)配列番号261のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-K3)配列番号261のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-K1)配列番号262(QQYDSLPLT)のアミノ酸配列
(L3-K2)配列番号262のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-K3)配列番号262のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-K1)配列番号263のアミノ酸配列
配列番号263:EIVLTRSPATLSLSPGERATLSCRASQSVSSYLAWYQQKPGQAPRLLIYDASNRATGISARFSGSGSGTEFTLTISSLQSEDIATYYCQQYDSLPLTFGGGTKLEIK
(L-K2)配列番号263のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-K3)配列番号263のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LL)の抗体等は、例えば、抗体18H-K9群ともいう。前記組合せ(LL)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LL)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-L)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-L)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-L)のアミノ酸配列を含むポリペプチドである。
(L1-L1)配列番号264(QTISASS)のアミノ酸配列
(L1-L2)配列番号264のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-L3)配列番号264のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-L1)配列番号265(GAS)のアミノ酸配列
(L2-L2)配列番号265のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-L3)配列番号265のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-L1)配列番号266(QQFNEWPLT)のアミノ酸配列
(L3-L2)配列番号266のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-L3)配列番号266のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-L1)配列番号267のアミノ酸配列
配列番号267:EIVLTQSPATLSLSPGERATVSCRPSQTISASSVAWYQQKAGQAPRLLIYGASSRATGIPDRFSGSGSGTDFTLTISRLEPEDFAVYYCQQFNEWPLTFGGGTKVEIK
(L-L2)配列番号267のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-L3)配列番号267のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
組合せ(LM)の抗体等は、例えば、抗体18H-K10群ともいう。前記組合せ(LM)において、前記(HA)の重鎖可変領域は、CDRH1、CDRH2、およびCDRH3を含み、CDRH1が、前記(H1-A)のアミノ酸配列を含むポリペプチドであり、CDRH2が、前記(H2-A)のアミノ酸配列を含むポリペプチドであり、CDRH3が、前記(H3-A)のアミノ酸配列を含むポリペプチドである。また、前記(LM)の軽鎖可変領域は、CDRL1、CDRL2、およびCDRL3を含み、CDRL1が、下記(L1-M)のアミノ酸配列を含むポリペプチドであり、CDRL2が、下記(L2-M)のアミノ酸配列を含むポリペプチドであり、CDRL3が、下記(L3-M)のアミノ酸配列を含むポリペプチドである。
(L1-M1)配列番号268(QSVSSN)のアミノ酸配列
(L1-M2)配列番号268のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-M3)配列番号268のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2-M1)配列番号269(GAS)のアミノ酸配列
(L2-M2)配列番号269のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-M3)配列番号269のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3-M1)配列番号270(QQYGSSPDIFT)のアミノ酸配列
(L3-M2)配列番号270のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-M3)配列番号270のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L-M1)配列番号271のアミノ酸配列
配列番号271:EIVLTQSPATLSLSPGERATLSCRASQSVSSNLAWYQQKLGQAPRLLIYGASSRATGIPDRFSGSGSGTDFTLTISRLEPEDFAVYYCQQYGSSPDIFTFGPGTKVDIK
(L-M2)配列番号271のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L-M3)配列番号271のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
CDRL1が、下記(L1)のアミノ酸配列を含むポリペプチドであり、
CDRL2が、下記(L2)のアミノ酸配列を含むポリペプチドであり、
CDRL3が、下記(L3)のアミノ酸配列を含むポリペプチドである軽鎖可変領域;
(L1)下記(L1-1)、(L1-2)または(L1-3)のアミノ酸配列
(L1-1)下記条件(L1)~(L3)および(L4)のCDRL1のいずれかのアミノ酸配列
(L1-2)(L1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-3)(L1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
(L2)下記(L2-1)、(L2-2)または(L2-3)のアミノ酸配列
(L2-1)下記条件(L1)~(L3)および(L4)のCDRL2のいずれかのアミノ酸配列
(L2-2)(L2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-3)(L2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
(L3)下記(L3-1)、(L3-2)または(L3-3)のアミノ酸配列
(L3-1)下記条件(L1)~(L3)および(L4)のCDRL3のいずれかのアミノ酸配列
(L3-2)(L3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-3)(L3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列;
条件(L1)
CDRL1は、KLGDKY(配列番号220)であり、
CDRL2は、QDS(配列番号221)であり、
CDRL3は、QAWDSSTHV(配列番号222)である;
条件(L2)
CDRL1は、WSNIGDDH(配列番号228)であり、
CDRL2は、DTS(配列番号229)であり、
CDRL3は、GTWESSLSGVV(配列番号230)である;
条件(L3)
CDRL1は、X1SX2X3GX4X5NY(配列番号317)であり、
CDRL2は、DX6X7(配列番号318)であり、
CDRL3は、X8X9X10X11X12SX13X14X15X16X17(配列番号319)であり、
X1は、SまたはTであり、
X2は、DまたはNであり、
X3は、V、I、L、またはAであり、
X4は、G、T、V、またはNであり、
X5は、存在しないか、Y、T、またはSであり、
X6は、VまたはNであり、
X7は、S、I、またはVであり、
X8は、存在しないか、G、A、またはHであり、
X9は、T、S、またはAであり、
X10は、W、Y、またはFであり、
X11は、AまたはDであり、
X12は、T、G、D、またはSであり、
X13は、LまたはYであり、
X14は、T、N、またはSであり、
X15は、A、F、G、またはHであり、
X15は、VまたはIであり、
X17は、存在しないか、Vである;
条件(L4)
CDRL1は、QX1X2SX3SX4(配列番号320)であり、
CDRL2は、GAS(配列番号253)またはDAS(配列番号257)であり、
CDRL3は、QQX5X6X7X8X9X10X11LX12T(配列番号321)であり、
X1は、S、T、N、またはQであり、
X2は、V、I、L、またはAであり、
X3は、存在しないか、A、I、V、L、またはGであり、
X4は、N、Y、またはSであり、
X5は、存在しないか、Sであり、
X6は、Y、F、またはWであり、
X7は、N、S、D、またはGであり、
X8は、N、T、S、またはEであり、
X9は、W、L、またはSであり、
X10は、存在しないか、Pであり、
X11は、存在しないか、PまたはDであり、
X12は、存在しないか、Y、F、またはWである。
本発明のコード遺伝子は、HLA-A*02:01およびNY-ESO-1157-165の複合体に対する抗体もしくはその抗原結合断片またはCD19に対する抗体もしくはその抗原結合断片(以下、あわせて「本発明の抗体またはその抗原結合断片」または「本発明の抗体等」ともいう)のコード遺伝子であり、前記本発明の第1の抗体もしくは抗原結合断片のアミノ酸配列または前記本発明の第2の抗体もしくは抗原結合断片をコードする核酸(ポリヌクレオチド)を含む。
本発明の第1のキメラ抗原受容体(CAR)は、前述のように、抗原結合ドメインと、膜貫通ドメインと、細胞内シグナル伝達ドメインとを含み、前記抗原結合ドメインは、前記本発明の抗体またはその抗原結合断片を含む。本発明の第1のCARは、前記抗原結合ドメインが、前記本発明の第1の抗体等を含むことが特徴であり、その他の構成および条件は、特に制限されない。本発明の第1のCARは、前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体等、第2の抗体等、遺伝子、発現ベクターおよび形質転換体の説明を援用できる。本発明の第1のCARは、例えば、A2/NY-ESO-1157発現がん細胞に対するCAR-T細胞のCARとして好適に使用できる。
(BD)下記(BD1)、(BD2)または(BD3)のポリペプチド
(BD1)配列番号98~117のいずれかのアミノ酸配列からなるポリペプチド
(BD2)配列番号98~117いずれかのアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列からなり、A2/NY-ESO-1157に結合するポリペプチド
(BD3)配列番号98~117のいずれかのアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列からなり、A2/NY-ESO-1157に結合するポリペプチド
QVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSELTQPRSVSGSPGQSVTISCTGTSRDVGGYNYVSWYQQHPGKAPKLIIHDVIERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCWSFAGSYYVFGTGTDVTVL
DIQMTQSPSAMSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQYYSTPQTFGPGTKVDIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKAGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYCQQYESYRRSFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDFATYYCQQYNSYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASRLESGVPSRFSGSGSGTDFTLTISCLQSEDFATYYCQQYNSYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTEFTLTISSLQPDDFATYYCQQYNSYSPCTFGPGTKVDIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
DIQMTQSPSSLSASVGDRVSITCRASQDISRYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYDNLITFGQGTRLEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
DIQMTQSPSSLSASVGDRVSITCRASQDISRYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYNSYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYDNLITFGQGTRLEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
EIVLTQSPATLSVSPGERATLSCRASQSVSSNLAWYQQKPGQAPRLLIYGASTRATGIPARFSGSGSGTEFTLTISRLEPEDFATYYCQQYNSYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKAGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYCQQYESYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSS
QSALTQPPSASGSPGQSVTISCTGTSSDVGGYDFVSWYQQHPGEAPKLLVYDVNNRPSGVSNRFSGSKSGNTASLTISGLQAEDEGDYYCSSYAGSNSVFGTGTKVTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
EVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSALTQPPSASGSPGQSVTISCTGTSSDVGGYDFVSWYQQHPGEAPKLLVYDVNNRPSGVSNRFSGSKSGNTASLTISGLQAEDEGDYYCSSYAGSNSVFGTGTKVTVL
QSALTQPASVSGSPGQSITISCTGTSSDVGGYEFVSWYQQHPGSAPKLIIYDVIERPFGVSYRFSASKSGNTASLTISGLQGEDEADYFCSSYTSSSTYVFGTGTKVTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
QSALTQPASVSGSPGQSITISCTGTGSDVGAYDYVSWYQHHPGRAPRLIIRDVSVRPSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCSSYSGSSTWVFGGGTKLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
QSALTQPASVSGSPGQSITISCTGTSSDVGSYNLVSWYQQHPGKAPKLMIYDVSNRPSGVSYRFSGSKSGNTASLTISGLQAEDEADYYCSSYTSSSTFAVFGGGTQLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
QSALPQPASVSGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSGNTASLTVSGLQAEDEADYFCCSYAGGYYVFGTGTKLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
QSALTQPPSASGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSGNTASLTISGLQTEDEADYYCSSYAGSGSTPFVFGTGTKLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
QSALTQPASVSGSPGQSITISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSRVPDRFAGSKSGNTASLTISGLQAEDEADYYCCSYAGRRYVFGTGTKLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
QLQLQESGPGLVKPSQTLSLTCTVSGGSISSYYWSWIRQPPGKGLEWIGEINHSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARCPIYYYGMDVWGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSELTQPRSVSGSPGQSVTISCTGTSRDVGGYNYVSWYQQHPGKAPKLIIHDVIERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCWSFAGSYYVFGTGTDVTVL
(C)下記(C1)、(C2)または(C3)のポリペプチド
(C1)配列番号118~137のいずれかのアミノ酸配列からなるポリペプチド
(C2)配列番号118~137のいずれかのアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列からなり、A2/NY-ESO-1157に結合するポリペプチド
(C3)配列番号118~137のいずれかのアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列からなり、A2/NY-ESO-1157に結合するポリペプチド
QVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSELTQPRSVSGSPGQSVTISCTGTSRDVGGYNYVSWYQQHPGKAPKLIIHDVIERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCWSFAGSYYVFGTGTDVTVLAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSAMSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISSLQPEDFATYYCQQYYSTPQTFGPGTKVDIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKAGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYCQQYESYRRSFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDFATYYCQQYNSYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASRLESGVPSRFSGSGSGTDFTLTISCLQSEDFATYYCQQYNSYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTEFTLTISSLQPDDFATYYCQQYNSYSPCTFGPGTKVDIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSSLSASVGDRVSITCRASQDISRYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYDNLITFGQGTRLEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSSLSASVGDRVSITCRASQDISRYLNWYQQKPGKAPKLLLYAASRLESGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYNSYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKPGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTITSLQPDDFATYYCQQYDNLITFGQGTRLEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
EIVLTQSPATLSVSPGERATLSCRASQSVSSNLAWYQQKPGQAPRLLIYGASTRATGIPARFSGSGSGTEFTLTISRLEPEDFATYYCQQYNSYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
DIQMTQSPSSLSASVGDRVTITCRASQSISSYLNWYQQKAGKAPKLLIYAASSLQSGVPSRFSGSGSGTDFTLTISCLQSEDVATYYCQQYESYSRTFGQGTKVEIKGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPPSASGSPGQSVTISCTGTSSDVGGYDFVSWYQQHPGEAPKLLVYDVNNRPSGVSNRFSGSKSGNTASLTISGLQAEDEGDYYCSSYAGSNSVFGTGTKVTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
EVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSQSALTQPPSASGSPGQSVTISCTGTSSDVGGYDFVSWYQQHPGEAPKLLVYDVNNRPSGVSNRFSGSKSGNTASLTISGLQAEDEGDYYCSSYAGSNSVFGTGTKVTVLAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPASVSGSPGQSITISCTGTSSDVGGYEFVSWYQQHPGSAPKLIIYDVIERPFGVSYRFSASKSGNTASLTISGLQGEDEADYFCSSYTSSSTYVFGTGTKVTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPASVSGSPGQSITISCTGTGSDVGAYDYVSWYQHHPGRAPRLIIRDVSVRPSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCSSYSGSSTWVFGGGTKLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPASVSGSPGQSITISCTGTSSDVGSYNLVSWYQQHPGKAPKLMIYDVSNRPSGVSYRFSGSKSGNTASLTISGLQAEDEADYYCSSYTSSSTFAVFGGGTQLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALPQPASVSGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSGNTASLTVSGLQAEDEADYFCCSYAGGYYVFGTGTKLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPPSASGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSGNTASLTISGLQTEDEADYYCSSYAGSGSTPFVFGTGTKLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPASVSGSPGQSITISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSRVPDRFAGSKSGNTASLTISGLQAEDEADYYCCSYAGRRYVFGTGTKLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QLQLQESGPGLVKPSQTLSLTCTVSGGSISSYYWSWIRQPPGKGLEWIGEINHSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARCPIYYYGMDVWGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSELTQPRSVSGSPGQSVTISCTGTSRDVGGYNYVSWYQQHPGKAPKLIIHDVIERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCWSFAGSYYVFGTGTDVTVLAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
本発明の第2のキメラ抗原受容体(CAR)は、前述のように、抗原結合ドメインと、膜貫通ドメインと、細胞内シグナル伝達ドメインとを含み、前記抗原結合ドメインは、前記本発明の抗体またはその抗原結合断片を含む。本発明の第2のCARは、前記抗原結合ドメインが、前記本発明の第2の抗体等を含むことが特徴であり、その他の構成および条件は、特に制限されない。本発明の第2のCARは、前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体等、第2の抗体等、遺伝子、発現ベクターおよび形質転換体の説明を援用できる。本発明の第2のCARは、例えば、CD19発現がん細胞に対するCAR-T細胞のCARとして好適に使用できる。
(bd)下記(bd1)、(bd2)または(bd3)のポリペプチド
(bd1)配列番号272~284のいずれかのアミノ酸配列からなるポリペプチド
(bd2)配列番号272~284のいずれかのアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列からなり、CD19に結合するポリペプチド
(bd3)配列番号272~284のいずれかのアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列からなり、CD19に結合するポリペプチド
SYELTQPPSVSVSPGQTASITCSGDKLGDKYACWYQQKPGQSPVLVIYQDSKRPSGIPERFSGSNSGNTATLTISGTQAMDEADYYCQAWDSSTHVVFGGGTKLTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
QSALTQPRSVSGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSGVSNRFSGSKSGNTASLTISGLQTGDEADYYCGTWDTSLTAVVFGGGTELTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
QSVLTQPPSVSAAPGQKVTISCSGSWSNIGDDHVSWYQQFPGAAPKLLIYDTSKRPSRVADRFSGSKSGASATLAITGLQAGDEADYYCGTWESSLSGVVFGGGTELTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
QSALTQPASVSGSPGQSITISCTGLSSDVGGYDYVSWYQQHPGIAPKLMIYDVTNRPSGVSSRFSGSKSGNTASLTISGLQAEDEADYYCSSYTTSTTWVFGAGTKLTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
QSALTQPASVSGSPGQSITISCTGTTSDVGTTNYVSWYQQHPGKAPKLLIYDVTNRPSGVPDRFSGSKSANTASLTISGLQAEDEADYYCCSYAGSYTFVVFGGGTELTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
QSALTQPPSASGSPGQSVTISCTGTSSDVGVYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSANTASLTISGLQAEDEADYYCAAWDDSLNGVVFGGGTQLTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
QSVLTQPPSASGTPGQRVTISCSGSSSNIGNNYVCWYQHLPGTAPKLLIYDNVKRPSGIPDRFSGSKSGTSASLAISGLRSEDEADYYCAAWDDSLSAIFGGGTELTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
QSALTQPASVSGSPGQSITISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSGVPDRFSGSKSGTSASLAISGLQSEDEADYYCHSYDSSLSHVFGTGTKVTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
EIVLTQSPGTLSLSPGERATLSCRASQSVSSNLAWYQQKPGQAPRLLIYGASTRATGIPARFSGSGSGTEFTLTISSLQSEDFAVYYCQQYNNWPPLYTFGQGTKLEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
EIVLTQSPGTLSLSPGERATLSCRASQSVSSYLAWYQQKPGQAPRLLIYDASNRATGIPARFSGSGSGTDFTLTISSLEPEDFATYYCQQSYSTLLYTFGQGTKLEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
EIVLTRSPATLSLSPGERATLSCRASQSVSSYLAWYQQKPGQAPRLLIYDASNRATGISARFSGSGSGTEFTLTISSLQSEDIATYYCQQYDSLPLTFGGGTKLEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
EIVLTQSPATLSLSPGERATVSCRPSQTISASSVAWYQQKAGQAPRLLIYGASSRATGIPDRFSGSGSGTDFTLTISRLEPEDFAVYYCQQFNEWPLTFGGGTKVEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
EIVLTQSPATLSLSPGERATLSCRASQSVSSNLAWYQQKLGQAPRLLIYGASSRATGIPDRFSGSGSGTDFTLTISRLEPEDFAVYYCQQYGSSPDIFTFGPGTKVDIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
(CA)下記(CA1)、(CA2)または(CA3)のポリペプチド
(CA1)配列番号285~297のいずれかのアミノ酸配列からなるポリペプチド
(CA2)配列番号285~297のいずれかのアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列からなり、CD19に結合するポリペプチド
(CA3)配列番号285~297のいずれかのアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列からなり、CD19に結合するポリペプチド
SYELTQPPSVSVSPGQTASITCSGDKLGDKYACWYQQKPGQSPVLVIYQDSKRPSGIPERFSGSNSGNTATLTISGTQAMDEADYYCQAWDSSTHVVFGGGTKLTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPRSVSGSPGQSVTISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSGVSNRFSGSKSGNTASLTISGLQTGDEADYYCGTWDTSLTAVVFGGGTELTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSVLTQPPSVSAAPGQKVTISCSGSWSNIGDDHVSWYQQFPGAAPKLLIYDTSKRPSRVADRFSGSKSGASATLAITGLQAGDEADYYCGTWESSLSGVVFGGGTELTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPASVSGSPGQSITISCTGLSSDVGGYDYVSWYQQHPGIAPKLMIYDVTNRPSGVSSRFSGSKSGNTASLTISGLQAEDEADYYCSSYTTSTTWVFGAGTKLTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPASVSGSPGQSITISCTGTTSDVGTTNYVSWYQQHPGKAPKLLIYDVTNRPSGVPDRFSGSKSANTASLTISGLQAEDEADYYCCSYAGSYTFVVFGGGTELTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPPSASGSPGQSVTISCTGTSSDVGVYNYVSWYQQHPGKAPKLMIYDVSKRPSGVPDRFSGSKSANTASLTISGLQAEDEADYYCAAWDDSLNGVVFGGGTQLTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSVLTQPPSASGTPGQRVTISCSGSSSNIGNNYVCWYQHLPGTAPKLLIYDNVKRPSGIPDRFSGSKSGTSASLAISGLRSEDEADYYCAAWDDSLSAIFGGGTELTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
QSALTQPASVSGSPGQSITISCTGTSSDVGGYNYVSWYQQHPGKAPKLMIYDVSNRPSGVPDRFSGSKSGTSASLAISGLQSEDEADYYCHSYDSSLSHVFGTGTKVTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
EIVLTQSPGTLSLSPGERATLSCRASQSVSSNLAWYQQKPGQAPRLLIYGASTRATGIPARFSGSGSGTEFTLTISSLQSEDFAVYYCQQYNNWPPLYTFGQGTKLEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
EIVLTQSPGTLSLSPGERATLSCRASQSVSSYLAWYQQKPGQAPRLLIYDASNRATGIPARFSGSGSGTDFTLTISSLEPEDFATYYCQQSYSTLLYTFGQGTKLEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
EIVLTRSPATLSLSPGERATLSCRASQSVSSYLAWYQQKPGQAPRLLIYDASNRATGISARFSGSGSGTEFTLTISSLQSEDIATYYCQQYDSLPLTFGGGTKLEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
EIVLTQSPATLSLSPGERATVSCRPSQTISASSVAWYQQKAGQAPRLLIYGASSRATGIPDRFSGSGSGTDFTLTISRLEPEDFAVYYCQQFNEWPLTFGGGTKVEIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
EIVLTQSPATLSLSPGERATLSCRASQSVSSNLAWYQQKLGQAPRLLIYGASSRATGIPDRFSGSGSGTDFTLTISRLEPEDFAVYYCQQYGSSPDIFTFGPGTKVDIKGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
本発明の核酸は、前述のように、前記本発明のキメラ抗原受容体をコードする。本発明の核酸は、前記本発明の第1のキメラ抗原受容体または第2のキメラ抗原受容体(以下、あわせて「本発明のキメラ抗原受容体」ともいう)をコードすることが特徴であり、その他の構成および条件は、特に制限されない。本発明の核酸によれば、例えば、細胞に本発明のCARを発現させることができる。本発明の核酸は、例えば、前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体等、第1の抗体等、遺伝子、発現ベクター、形質転換体、第1のキメラ抗原受容体第2のキメラ抗原受容体の説明を援用できる。
(c)下記(c1)、(c2)、(c3)、(c4)、(c5)、または(c6)のポリヌクレオチド(核酸)
(c1)配列番号118~137のいずれかのアミノ酸配列からなるポリペプチドをコードするポリヌクレオチド
(c2)配列番号118~137のいずれかのアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列からなり、A2/NY-ESO-1157に結合するポリペプチドをコードするポリヌクレオチド
(c3)配列番号118~137いずれかのアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列からなり、A2/NY-ESO-1157に結合するポリペプチドをコードするポリヌクレオチド
(c4)配列番号138~157のいずれかの塩基配列からなるポリヌクレオチド
(c5)配列番号138~157のいずれかの塩基配列に対して、80%以上の同一性を有する塩基配列からなり、A2/NY-ESO-1157に結合するポリペプチドをコードするポリヌクレオチド
(c6)配列番号138~157のいずれかの塩基配列において、1個または数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなり、A2/NY-ESO-1157に結合するポリペプチドをコードするポリヌクレオチド
5'-CAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGAGCGAGCTGACACAGCCTAGATCCGTGTCTGGCAGCCCTGGCCAGAGCGTGACCATCAGCTGTACCGGCACCAGCAGAGATGTGGGCGGCTACAACTACGTGTCCTGGTATCAGCAGCATCCCGGCAAGGCCCCCAAGCTGATCATCCACGACGTGATCGAGCGGAGCAGCGGCGTGCCCGATAGATTCAGCGGCAGCAAGAGCGGCAACACCGCCAGCCTGACAATCAGCGGACTGCAGGCCGAGGACGAGGCCGACTACTACTGTTGGAGCTTCGCCGGCAGCTACTACGTGTTCGGCACCGGCACCGATGTGACCGTGCTGGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCTGCCATGTCTGCATCTGTAGGAGACAGAGTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGCTGCATCCAGTTTGCAAAGTGGGGTCCCATCAAGGTTCAGTGGCAGTGGATCTGGGACAGATTTCACTCTCACCATCAGCAGTCTGCAGCCTGAAGATTTTGCAACTTACTATTGTCAGCAATATTATAGTACTCCTCAAACTTTCGGCCCTGGGACCAAAGTGGATATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGAGTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTATCAGCAGAAAGCAGGGAAAGCCCCTAAGCTTCTGATCTATGCTGCATCCAGTTTGCAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGATTTCACTCTCACCATCAGCTGCCTGCAGTCTGAAGATGTTGCAACCTATTACTGCCAACAGTATGAAAGTTATCGAAGGTCGTTCGGCCAAGGGACCAAGGTGGAAATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGAGTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGCTGCATCCAGTTTGCAAAGTGGGGTCCCATCAAGGTTCAGTGGCAGTGGATCTGGGACAGATTTCACTCTCACCATCAGCTGCCTGCAGTCTGAAGATTTTGCAACTTATTACTGCCAACAGTATAATAGTTATTCCCGGACGTTCGGCCAAGGGACCAAGGTGGAAATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGAGTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGCTGCATCCAGATTGGAAAGTGGGGTCCCATCAAGGTTCAGTGGCAGTGGATCTGGGACAGATTTCACTCTCACCATCAGCTGCCTGCAGTCTGAAGATTTTGCAACTTATTACTGCCAACAGTATAATAGTTATTCCCGGACGTTCGGCCAAGGGACCAAGGTGGAAATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGAGTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGCTCTATGCTGCATCCAGATTGGAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGAATTCACTCTCACCATCAGCAGCCTGCAGCCTGATGATTTTGCAACTTATTACTGCCAACAGTATAATAGTTATTCTCCGTGCACTTTCGGCCCTGGGACCAAAGTGGATATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGGGACAGAGTCTCCATCACTTGCCGGGCAAGTCAGGACATTAGCAGGTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGCTCTATGCTGCATCCAGATTGGAAAGTGGGGTCCCATCCAGGTTCAGTGGCAGTGGATCTGGGACAGACTTCACTCTCACCATCACCAGCCTGCAGCCTGATGACTTTGCAACTTATTACTGCCAACAGTATGATAATCTGATCACCTTCGGCCAAGGGACACGACTGGAGATTAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGGGACAGAGTCTCCATCACTTGCCGGGCAAGTCAGGACATTAGCAGGTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGCTCTATGCTGCATCCAGATTGGAAAGTGGGGTCCCATCCAGGTTCAGTGGCAGTGGATCTGGGACAGACTTCACTCTCACCATCACCAGCCTGCAGCCTGATGACTTTGCAACTTATTACTGCCAACAGTATAATAGTTATTCCCGGACGTTCGGCCAAGGGACCAAGGTGGAAATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGGGACAGAGTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAGCTCCTGATCTATGCTGCATCCAGTTTGCAAAGTGGGGTCCCATCAAGGTTCAGTGGCAGTGGATCTGGGACAGACTTCACTCTCACCATCACCAGCCTGCAGCCTGATGACTTTGCAACTTATTACTGCCAACAGTATGATAATCTGATCACCTTCGGCCAAGGGACACGACTGGAGATTAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GAAATTGTGTTGACGCAGTCTCCAGCCACCCTGTCTGTGTCTCCAGGGGAAAGAGCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCAACTTAGCCTGGTACCAGCAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGGTGCATCCACCAGGGCCACTGGTATCCCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGAGTTCACTCTCACCATCAGCAGACTGGAGCCTGAAGATTTTGCAACCTATTACTGCCAACAGTATAATAGTTATTCCCGGACGTTCGGCCAAGGGACCAAGGTGGAAATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GACATCCAGATGACCCAGTCTCCATCCTCCCTGTCTGCATCTGTAGGAGACAGAGTCACCATCACTTGCCGGGCAAGTCAGAGCATTAGCAGCTATTTAAATTGGTATCAGCAGAAAGCAGGGAAAGCCCCTAAGCTTCTGATCTATGCTGCATCCAGTTTGCAAAGTGGGGTCCCATCAAGGTTCAGCGGCAGTGGATCTGGGACAGATTTCACTCTCACCATCAGCTGCCTGCAGTCTGAAGATGTTGCAACCTATTACTGCCAACAGTATGAAAGTTATTCCCGGACGTTCGGCCAAGGGACCAAGGTGGAAATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTCCCTCCGCGTCCGGGTCTCCTGGACAGTCAGTCACCATCTCCTGCACTGGAACCAGCAGTGACGTTGGTGGTTATGACTTTGTCTCCTGGTACCAACAGCACCCAGGCGAAGCCCCCAAACTCCTCGTTTATGATGTCAATAACCGGCCCTCAGGGGTTTCTAATCGCTTCTCTGGCTCCAAGTCTGGCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGGCTGAGGACGAGGGTGACTATTACTGCAGCTCATATGCAGGCAGCAACAGCGTCTTCGGAACTGGGACCAAGGTCACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGTCTGCCCTGACTCAGCCTCCCTCCGCGTCCGGGTCTCCTGGACAGTCAGTCACCATCTCCTGCACTGGAACCAGCAGTGACGTTGGTGGTTATGACTTTGTCTCCTGGTACCAACAGCACCCAGGCGAAGCCCCCAAACTCCTCGTTTATGATGTCAATAACCGGCCCTCAGGGGTTTCTAATCGCTTCTCTGGCTCCAAGTCTGGCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGGCTGAGGACGAGGGTGACTATTACTGCAGCTCATATGCAGGCAGCAACAGCGTCTTCGGAACTGGGACCAAGGTCACCGTCCTAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGATCACCATCTCCTGCACTGGGACCAGCAGTGACGTTGGTGGTTATGAATTTGTCTCCTGGTACCAACAACACCCAGGCAGCGCCCCCAAACTCATTATTTATGACGTAATAGAGCGTCCCTTCGGTGTCTCCTATCGGTTCTCTGCCTCCAAGTCAGGCAACACGGCCTCCCTGACGATCTCTGGGCTCCAGGGTGAAGACGAGGCTGATTACTTCTGCAGCTCATATACAAGCAGCAGCACTTATGTCTTCGGAACTGGGACCAAGGTCACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGATCACCATCTCCTGCACTGGAACCGGCAGTGACGTTGGTGCTTATGACTATGTCTCCTGGTACCAACATCACCCAGGCAGAGCCCCCAGACTCATCATTCGTGATGTCAGTGTGCGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCAAGTCTGGCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGGCTGAGGACGAGGCTGATTATTACTGCTCCTCATATTCAGGCAGCAGCACTTGGGTGTTCGGCGGGGGGACCAAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGATCACCATCTCCTGCACTGGAACCAGCAGTGATGTTGGGAGTTATAACCTTGTCTCCTGGTACCAACAACACCCAGGCAAAGCCCCCAAACTCATGATTTATGATGTCAGTAATCGGCCCTCAGGGGTTTCTTATCGCTTCTCTGGCTCCAAGTCTGGCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGGCTGAGGACGAGGCTGATTATTACTGCAGCTCATATACAAGCAGCAGCACTTTCGCGGTGTTCGGCGGAGGGACCCAGCTGACCGTCCTCGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAATCTGCCCTGCCTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCAGTCACCATCTCCTGCACTGGAACCAGCAGTGATGTTGGTGGTTATAACTATGTCTCCTGGTACCAACAGCACCCAGGCAAAGCCCCCAAACTCATGATTTATGATGTCAGTAAGCGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGTTCCAAGTCTGGCAACACGGCCTCCCTGACCGTCTCTGGGCTCCAGGCTGAGGATGAGGCTGATTATTTCTGCTGTTCGTATGCAGGCGGCTATTATGTCTTCGGAACTGGGACCAAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTCCCTCCGCGTCCGGGTCTCCTGGACAGTCAGTCACCATCTCCTGCACTGGAACCAGCAGTGACGTTGGTGGTTATAACTATGTCTCCTGGTACCAACAGCACCCAGGCAAAGCCCCCAAACTCATGATTTATGATGTCAGTAAGCGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCAAGTCTGGCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGACTGAGGACGAGGCTGATTACTATTGCAGCTCATATGCAGGCAGCGGCAGCACCCCCTTTGTCTTCGGAACTGGGACCAAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGATCACCATCTCCTGCACTGGAACCAGCAGTGACGTTGGTGGTTATAACTATGTCTCCTGGTACCAACAACACCCAGGCAAAGCCCCCAAACTCATGATTTATGATGTCAGTAATCGGCCCTCAAGGGTCCCTGATCGCTTCGCTGGCTCCAAGTCTGGCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGGCTGAAGACGAGGCTGATTATTACTGCTGCTCATATGCCGGCAGACGTTATGTGTTCGGAACTGGGACCAAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGCTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCACAGACCCTGTCCCTCACCTGCACTGTCTCTGGTGGCTCCATCAGTAGTTACTACTGGAGCTGGATCCGGCAGCCCCCAGGGAAGGGACTGGAGTGGATTGGGGAAATCAATCATAGTGGAAGCACCAACTACAACCCGTCCCTCAAGAGTCGAGTCACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGGTGCCCTATCTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGAGCGAGCTGACACAGCCTAGATCCGTGTCTGGCAGCCCTGGCCAGAGCGTGACCATCAGCTGTACCGGCACCAGCAGAGATGTGGGCGGCTACAACTACGTGTCCTGGTATCAGCAGCATCCCGGCAAGGCCCCCAAGCTGATCATCCACGACGTGATCGAGCGGAGCAGCGGCGTGCCCGATAGATTCAGCGGCAGCAAGAGCGGCAACACCGCCAGCCTGACAATCAGCGGACTGCAGGCCGAGGACGAGGCCGACTACTACTGTTGGAGCTTCGCCGGCAGCTACTACGTGTTCGGCACCGGCACCGATGTGACCGTGCTGGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
(ca)下記(ca1)、(ca2)、(ca3)、(ca4)、(ca5)、または(ca6)のポリヌクレオチド(核酸)
(ca1)配列番号272~284のいずれかのアミノ酸配列からなるポリペプチドをコードするポリヌクレオチド
(ca2)配列番号272~284のいずれかのアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列からなり、CD19に結合するポリペプチドをコードするポリヌクレオチド
(ca3)配列番号272~284のいずれかのアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列からなり、CD19に結合するポリペプチドをコードするポリヌクレオチド
(ca4)配列番号298~310のいずれかの塩基配列からなるポリヌクレオチド
(ca5)配列番号298~310のいずれかの塩基配列に対して、80%以上の同一性を有する塩基配列からなり、CD19に結合するポリペプチドをコードするポリヌクレオチド
(ca6)配列番号298~310のいずれかの塩基配列において、1個または数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなり、CD19に結合するポリペプチドをコードするポリヌクレオチド
5'-TCCTATGAGCTGACTCAGCCACCCTCAGTGTCCGTGTCTCCAGGACAGACAGCCAGCATCACCTGCTCTGGAGATAAATTGGGGGATAAATATGCTTGCTGGTATCAGCAGAAGCCAGGCCAGTCCCCTGTGCTGGTCATCTATCAAGATAGCAAGCGGCCCTCAGGGATCCCTGAGCGATTCTCTGGCTCCAACTCTGGGAACACAGCCACTCTGACCATCAGCGGGACCCAGGCTATGGATGAGGCTGACTATTACTGTCAGGCGTGGGACAGCAGCACACATGTGGTATTCGGCGGAGGGACCAAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTCGCTCAGTGTCCGGGTCTCCTGGACAGTCAGTCACCATCTCCTGCACTGGAACCAGCAGTGATGTTGGTGGTTATAACTATGTCTCCTGGTACCAACAACACCCAGGCAAAGCCCCCAAACTCATGATTTATGATGTCAGTAATCGGCCCTCAGGGGTTTCTAATCGCTTCTCTGGCTCCAAGTCTGGCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGACTGGGGACGAGGCCGATTATTACTGCGGAACATGGGATACCAGCCTGACTGCTGTGGTATTCGGCGGAGGGACCGAGCTGACCGTCCTCGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGTGTTGACGCAGCCGCCCTCAGTGTCTGCGGCCCCAGGACAGAAGGTCACCATCTCCTGCTCTGGAAGCTGGTCCAACATTGGAGATGATCATGTCTCCTGGTACCAGCAGTTCCCAGGAGCAGCCCCCAAACTCCTCATTTATGACACTTCTAAGCGACCCTCACGCGTTGCTGACCGATTCTCTGGCTCCAAGTCTGGCGCGTCAGCCACCCTGGCCATCACTGGACTCCAGGCTGGGGACGAGGCCGACTATTATTGCGGAACATGGGAAAGCAGCCTGAGTGGTGTGGTTTTCGGCGGAGGGACCGAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGATCACCATCTCCTGCACTGGACTCAGCAGTGACGTTGGTGGTTATGACTATGTCTCCTGGTACCAACAACACCCAGGCATAGCCCCCAAACTCATGATTTATGATGTCACTAATCGGCCCTCAGGGGTTTCTAGTCGCTTCTCTGGCTCCAAGTCTGGCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGGCTGAGGACGAGGCTGATTATTACTGCAGCTCATATACAACCAGCACGACTTGGGTCTTCGGAGCTGGGACCAAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGATCACCATCTCCTGCACTGGAACCACCAGTGACGTTGGTACTACTAATTATGTCTCCTGGTACCAGCAACACCCAGGCAAAGCCCCCAAACTCCTAATTTATGATGTCACTAATCGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCAAGTCTGCCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGGCTGAGGACGAGGCTGATTATTACTGCTGCTCATATGCAGGCAGCTACACCTTCGTGGTATTCGGCGGAGGGACCGAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTCCCTCCGCGTCCGGGTCTCCTGGACAGTCAGTCACCATCTCCTGCACTGGAACCAGCAGTGACGTTGGTGTTTATAACTATGTCTCCTGGTACCAACAACACCCAGGCAAAGCCCCCAAACTCATGATTTATGATGTCAGTAAGCGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCAAGTCTGCCAACACGGCCTCCCTGACCATCTCTGGGCTCCAGGCTGAGGATGAGGCTGATTATTACTGTGCAGCATGGGATGACAGCCTGAATGGTGTGGTATTCGGCGGAGGCACCCAGCTGACCGTCCTCGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGTACTGACTCAACCACCCTCAGCGTCTGGGACCCCCGGGCAGAGGGTCACCATCTCTTGCTCTGGCAGCAGCTCCAACATCGGAAATAATTATGTATGCTGGTACCAACACCTCCCAGGAACGGCCCCCAAACTTCTCATTTATGACAATGTTAAGCGACCCTCAGGGATTCCTGACCGATTCTCTGGCTCCAAGTCTGGCACGTCAGCCTCCCTGGCCATCAGTGGGCTCCGGTCCGAGGATGAGGCTGATTATTACTGTGCAGCATGGGATGACAGCCTGAGTGCCATATTCGGCGGAGGGACCGAGCTGACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-CAGTCTGCCCTGACTCAGCCTGCCTCCGTGTCTGGGTCTCCTGGACAGTCGATCACCATCTCCTGCACTGGAACCAGCAGTGACGTTGGTGGTTATAACTATGTCTCCTGGTACCAACAACACCCAGGCAAAGCCCCCAAACTCATGATTTATGATGTCAGTAATCGGCCCTCAGGGGTCCCTGATCGCTTCTCTGGCTCCAAGTCTGGCACCTCAGCCTCCCTGGCCATCAGTGGGCTCCAGTCTGAGGATGAGGCTGATTATTACTGCCACTCCTATGACAGCAGCCTGAGTCATGTCTTCGGAACTGGGACCAAGGTCACCGTCCTAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GAAATTGTGTTGACGCAGTCTCCAGGCACCCTGTCTTTGTCTCCAGGGGAAAGAGCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCAACTTAGCCTGGTACCAGCAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGGTGCATCCACCAGGGCCACTGGTATCCCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGAGTTCACTCTCACCATCAGCAGCCTGCAGTCTGAAGATTTTGCAGTTTATTACTGTCAGCAGTATAATAACTGGCCTCCCTTGTACACTTTTGGCCAGGGGACCAAGCTGGAGATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GAAATTGTGTTGACGCAGTCTCCAGGCACCCTGTCTTTGTCTCCAGGGGAAAGAGCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCTACTTAGCCTGGTACCAACAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGATGCATCCAACAGGGCCACTGGCATCCCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTCACTCTCACCATCAGCAGCCTAGAGCCTGAAGATTTTGCAACTTACTACTGTCAACAGAGTTACAGTACCCTTTTGTACACTTTTGGCCAGGGGACCAAGCTGGAGATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GAAATTGTGTTGACACGGTCTCCAGCCACCCTGTCTTTGTCTCCAGGGGAAAGAGCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCTACTTAGCCTGGTACCAACAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGATGCCTCCAACAGGGCCACTGGTATTTCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGAGTTCACTCTCACCATCAGCAGCCTGCAGTCTGAAGATATTGCAACATATTACTGTCAACAGTATGATAGTCTCCCACTCACTTTCGGCGGAGGGACCAAGCTGGAGATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GAAATAGTGTTGACACAGTCTCCAGCCACCCTGTCTTTGTCTCCAGGGGAAAGAGCCACCGTCTCCTGTAGGCCCAGTCAGACCATTAGTGCCAGTTCCGTAGCCTGGTATCAGCAGAAAGCTGGCCAGGCTCCACGGCTCCTCATCTATGGTGCATCCAGCAGGGCCACTGGCATCCCAGACAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTCACTCTCACCATCAGCAGACTGGAGCCTGAAGATTTTGCAGTTTATTACTGTCAGCAATTTAATGAATGGCCTCTCACTTTCGGCGGAGGGACCAAGGTGGAAATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
5'-GAAATTGTGTTGACACAGTCTCCAGCCACCCTGTCTTTGTCTCCAGGGGAAAGAGCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCAACTTAGCCTGGTACCAGCAAAAACTTGGCCAGGCTCCCAGGCTCCTCATCTATGGTGCATCCAGCAGGGCCACTGGCATCCCAGACAGGTTCAGTGGCAGTGGGTCTGGGACAGACTTCACTCTCACCATCAGCAGACTGGAGCCTGAAGATTTTGCAGTGTATTACTGTCAGCAGTATGGTAGCTCACCCGATATATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAAAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCTGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
本発明の細胞は、前述のように、前記本発明のキメラ抗原受容体を含む。本発明の細胞は、前記本発明のキメラ抗原受容体を含むことが特徴であり、その他の構成および条件は、特に制限されない。本発明の細胞によれば、例えば、がんを治療できる。本発明の細胞は、例えば、前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体等、第2の抗体等、遺伝子、発現ベクター、形質転換体、第1のキメラ抗原受容体、第2のキメラ抗原受容体、核酸等の説明を援用できる。
本発明の細胞の製造方法は、前述のように、前記本発明の核酸を細胞に導入する導入工程を含む。本発明の細胞の製造方法は、前記導入工程において、前記本発明の核酸を細胞に導入することが特徴であり、その他の工程および条件は、特に制限されない。本発明の細胞の製造方法によれば、例えば、がんを治療可能な細胞を製造できる。本発明の細胞の製造方法は、例えば、前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体等、第2の抗体等、遺伝子、発現ベクター、形質転換体、第1のキメラ抗原受容体、第2のキメラ抗原受容体、核酸、細胞等の説明を援用できる。
本発明のがん治療薬(以下、「治療薬」ともいう)は、前記本発明の細胞、遺伝子、発現ベクター、または核酸を含む。本発明の治療薬は、前記本発明の細胞、遺伝子、発現ベクター、または核酸を含むことが特徴であり、その他の構成および条件は、特に制限されない。本発明の治療薬によれば、例えば、がんを治療できる。本発明のがん治療薬は、例えば、前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体等、第2の抗体等、遺伝子、発現ベクター、形質転換体、第1のキメラ抗原受容体、第2のキメラ抗原受容体、核酸、細胞、細胞の製造方法等の説明を援用できる。
本発明のがんの治療方法は、投与対象に、前記本発明の細胞、遺伝子、発現ベクター、または核酸を投与する投与工程を含む。本発明のがんの治療方法は、前記投与工程において、前記本発明の細胞、遺伝子、発現ベクター、または核酸を投与することが特徴であり、その他の工程および条件は、特に制限されない。本発明の治療方法によれば、がんを治療できる。本発明のがんの治療方法は、例えば、前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体等、第2の抗体等、遺伝子、発現ベクター、形質転換体、第1のキメラ抗原受容体、第2のキメラ抗原受容体、核酸、細胞、細胞の製造方法、治療薬等の説明を援用できる。
本発明は、がんの治療方法に使用するための、前記本発明の抗体またはその抗原結合断片、または、がんの治療のための、前記本発明の抗体またはその抗原結合断片である。また、本発明は、がんの治療用医薬の製造のための、前記本発明の抗体またはその抗原結合断片の使用である。さらに、本発明は、がんの治療方法に使用するための、前記本発明の細胞、遺伝子、発現ベクター、もしくは核酸、または、がんの治療のための、前記本発明の細胞、遺伝子、発現ベクター、もしくは核酸である。また、本発明は、がんの治療用医薬の製造のための、前記本発明の細胞、遺伝子、発現ベクター、または核酸の使用である。本発明の使用は、例えば、前記本発明のCARライブラリ、第1のスクリーニング方法、第1の抗体等、第2の抗体等、遺伝子、発現ベクター、形質転換体、第1のキメラ抗原受容体、第2のキメラ抗原受容体、核酸、細胞、細胞の製造方法、治療薬、治療方法等の説明を援用できる。
本発明の二重特異性抗体(BsAb)ライブラリは、第1の二重特異性抗体(BsAb)をコードする核酸を含み、前記第1のBsAbは、第1の抗原結合ドメインと、第2の抗原結合ドメインとを含み、前記第1の抗原結合ドメインは、第1の標的抗原に結合する第1の一本鎖抗体(scFv)を含み、前記第2の抗原結合ドメインは、第2の標的抗原への結合をスクリーニングする第2のscFvを含み、前記第2のscFvは、第2の重鎖可変領域と、第2の軽鎖可変領域とを含み、前記第2の重鎖可変領域および前記第2の軽鎖可変領域は、下記条件1または条件2を満たし、前記第1の標的抗原または前記第2の標的抗原は、免疫細胞活性化受容体である。
前記第2の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、前記第2の標的抗原に結合する抗体もしくはその抗原結合断片における重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、第1のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む;
条件2:
前記第2の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、第1のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、前記第2の標的抗原に結合する抗体もしくはその抗原結合断片における軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。
前記免疫細胞活性化受容体は、例えば、前記免疫細胞に発現し、かつ、リガンドとの結合または複数の免疫細胞活性化受容体の架橋により前記免疫細胞活性化受容体が発現する免疫細胞を活性化可能な受容体を意味する。前記免疫細胞活性化受容体は、例えば、前記免疫細胞の種類に応じて適宜決定できる。具体例として、前記免疫細胞がT細胞の場合、前記免疫細胞活性化受容体は、CD3等があげられる。前記CD3は、例えば、CD3γ、CD3δ、CD3ε、CD3ζ、CD3ηまたは、これらの複合体があげられる。前記免疫細胞がNK細胞の場合、前記免疫細胞活性化受容体は、NK細胞の活性化受容体等があげられる。前記NK細胞の活性化受容体は、例えば、CD94/NKG2C、CD94/NKG2E、NKG2D/NKG2D等があげられる。前記免疫細胞がNKT細胞の場合、前記免疫細胞活性化受容体は、CD3、前記NK細胞の活性化受容体等があげられる。前記免疫細胞がB細胞の場合、前記免疫細胞活性化受容体は、例えば、CD19、CD79α、CD79β等があげられる。
前記第1の標的抗原は、特に制限されず、例えば、前記CARライブラリにおける標的抗原の説明を援用できる。
[V1]-[GSTSGSGKPGSGEGSTKG]-[V2]-[SGSG]-[V3]-[GSTSGSGKPGSGEGSTKG]-[V4] ・・・(3)
(V2)V1:VL1、V2:VH1、V3:VH2、V4:VL2
(V3)V1:VH1、V2:VL1、V3:VL2、V4:VH2
(V4)V1:VL1、V2:VH1、V3:VL2、V4:VH2
(V5)V1:VH2、V2:VL2、V3:VH1、V5:VL1
(V6)V1:VH2、V2:VL2、V3:VL1、V4:VH1
(V7)V1:VL2、V2:VH2、V3:VH1、V4:VL1
(V8)V1:VL2、V2:VH2、V3:VL1、V4:VH1
VH1:第1の重鎖可変領域
VL1:第1の軽鎖可変領域
VH2:第2の重鎖可変領域
VL2:第2の軽鎖可変領域
5'-[N1]-[GGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGC]-[N2]-[AGCGGATCTGGC]-[N3]-[GGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGC]-[N4]-3’ ・・・(4)
(N2)N1:NL1、N2:NH1、N3:NH2、N4:NL2
(N3)N1:NH1、N2:NL1、N3:NL2、N4:NH2
(N4)N1:NL1、N2:NH1、N3:NL2、N4:NH2
(N5)N1:NH2、N2:NL2、N3:NH1、N4:NL1
(N6)N1:NH2、N2:NL2、N3:NL1、N4:NH1
(N7)N1:NL2、N2:NH2、N3:NH1、N4:NL1
(N8)N1:NL2、N2:NH2、N3:NL1、N4:NH1
NH1:第1の重鎖可変領域をコードする塩基配列
NL1:第1の軽鎖可変領域をコードする塩基配列
NH2:第2の重鎖可変領域をコードする塩基配列
NL2:第2の軽鎖可変領域をコードする塩基配列
本発明の第2のscFvのスクリーニング方法(以下、「第2のスクリーニング方法」ともいう)は、前記本発明のBsAbライブラリから第1のBsAbを準備する第1の準備工程と、
前記第1の標的抗原が免疫細胞活性化受容体の場合、
前記第1のBsAbと、前記第2の標的抗原と、前記免疫細胞活性化受容体を発現可能な免疫細胞とを接触させ、
前記第2の標的抗原が免疫細胞活性化受容体の場合、
前記第1のBsAbと、前記第1の標的抗原と、前記免疫細胞活性化受容体を発現可能な免疫細胞とを接触させる第1の接触工程と、
前記第1の接触工程において、前記第2の標的抗原と結合した第1のBsAbの第2のscFvを、前記第2の標的抗原に結合する第1の候補scFvとして選抜する第1の選抜工程とを含む。
・T細胞
活性化マーカー:CD69、CD107a等
サイトカイン・ケモカイン:IFN-γ、IL-12、IL-2、TNFα、MIP-1β等
・NK細胞
活性化マーカー:CD69、CD107a等
サイトカイン・ケモカイン:INF-γ、IL-12等
・NKT細胞
活性化マーカー:CD69、CD107a、CD25等
サイトカイン・ケモカイン:INF-γ、IL-2等
・B細胞
活性化マーカー:CD28、CD69、CD80、CD138、B220等
サイトカイン・ケモカイン:CXCR4等
前記第1の準備工程におけるBsAbライブラリが前記条件1を満たす場合、
前記第4の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、第2のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第4の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、前記第1の候補BsAbにおける軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む;
条件4:
前記第1の準備工程におけるBsAbライブラリが前記条件2を満たす場合、
前記第4の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、前記第1の候補BsAbにおける重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第4の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、第2のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む。
[V5]-[GSTSGSGKPGSGEGSTKG]-[V6]-[SGSG]-[V7]-[GSTSGSGKPGSGEGSTKG]-[V8] ・・・(5)
(V10)V5:VL3、V6:VH3、V7:VH4、V8:VL4
(V11)V5:VH3、V6:VL3、V7:VL4、V8:VH4
(V12)V5:VL3、V6:VH3、V7:VL4、V8:VH4
(V13)V5:VH4、V6:VL4、V7:VH3、V8:VL3
(V14)V5:VH4、V6:VL4、V7:VL3、V8:VH3
(V15)V5:VL4、V6:VH4、V7:VH3、V8:VL3
(V16)V5:VL4、V6:VH4、V7:VL3、V8:VH3
VH3:第3の重鎖可変領域
VL3:第3の軽鎖可変領域
VH4:第4の重鎖可変領域
VL4:第4の軽鎖可変領域
5'-[N5]-[GGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGC]-[N6]-[AGCGGATCTGGC]-[N7]-[GGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGC]-[N8]-3’・・・(6)
(N10)N5:NL3、N6:NH3、N7:NH4、N8:NL4
(N11)N5:NH3、N6:NL3、N7:NL4、N8:NH4
(N12)N5:NL3、N6:NH3、N7:NL4、N8:NH4
(N13)N5:NH4、N6:NL4、N7:NH3、N8:NL3
(N14)N5:NH4、N6:NL4、N7:NL3、N8:NH3
(N15)N5:NL4、N6:NH4、N7:NH3、N8:NL3
(N16)N5:NL4、N6:NH4、N7:NL3、N8:NH3
NH3:第3の重鎖可変領域をコードする塩基配列
NL3:第3の軽鎖可変領域をコードする塩基配列
NH4:第4の重鎖可変領域をコードする塩基配列
NL4:第4の軽鎖可変領域をコードする塩基配列
本発明のscFvの製造方法は、前述のように、第1のキメラ抗原受容体(CAR)ライブラリの発現細胞を動物に投与する第1の投与工程と、
前記動物において、標的抗原を発現する組織に集積した前記第1のCARライブラリ発現細胞を、前記標的抗原特異的なCARを発現する細胞として採取する第1の採取工程とを含み、
前記第1のCARライブラリは、第1のCARをコードする核酸を含み、
前記第1のCARは、第1の抗原結合ドメインと、第1の膜貫通ドメインと、第1の細胞内シグナル伝達ドメインとを含み、
前記第1の抗原結合ドメインは、標的抗原への結合をスクリーニングする第1の一本鎖抗体(scFv)を含み、
前記第1のscFvは、第1の重鎖可変領域と、第1の軽鎖可変領域とを含み、
前記第1の重鎖可変領域および前記第1の軽鎖可変領域は、下記条件1または条件2を満たす。
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、第1のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む。
(条件2)
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、第1のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。
前記第1のCARライブラリが前記条件1を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、第2のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、前記標的抗原特異的なCARを発現する細胞のscFvにおける軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む;
(条件4)
前記第1のCARライブラリが前記条件2を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、前記標的抗原特異的なCARを発現する細胞のscFvにおける重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、第2のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む。
本発明のCARライブラリを用いた本発明のスクリーニング方法により、A2/NY-ESO-1157に結合するscFvをスクリーニングできることを確認した。
3M4E5 CARは、A2/NY-ESO-1157特異的抗体(clone:3M4E5)を用いて調製した。具体的には、3M4E5の免疫グロブリン重鎖由来の重鎖可変領域をコードする核酸(配列番号162)および3M4E5の免疫グロブリン軽鎖由来の軽鎖可変領域をコードする核酸(配列番号163)を、Fvリンカーペプチド(GSTSGSGKPGSGEGSTKG:配列番号1)をコードする核酸(配列番号3)で連結することにより、3M4E5 scFv(をコードする核酸を調製した。3M4E5の重鎖可変領域、Fvリンカーペプチド、および3M4E5の軽鎖可変領域をこの順番で連結したscFvを、3M4E5HL scFvという。また、3M4E5の軽鎖可変領域、Fvリンカーペプチド、および3M4E5の重鎖可変領域をこの順番で連結したscFvを、3M4E5LH scFvという。
5’-GAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCA-3'
5’-CAGAGCGAGCTGACACAGCCTAGATCCGTGTCTGGCAGCCCTGGCCAGAGCGTGACCATCAGCTGTACCGGCACCAGCAGAGATGTGGGCGGCTACAACTACGTGTCCTGGTATCAGCAGCATCCCGGCAAGGCCCCCAAGCTGATCATCCACGACGTGATCGAGCGGAGCAGCGGCGTGCCCGATAGATTCAGCGGCAGCAAGAGCGGCAACACCGCCAGCCTGACAATCAGCGGACTGCAGGCCGAGGACGAGGCCGACTACTACTGTTGGAGCTTCGCCGGCAGCTACTACGTGTTCGGCACCGGCACCGATGTGACCGTGCTG-3'
EVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSGSTSGSGKPGSGEGSTKGQSELTQPRSVSGSPGQSVTISCTGTSRDVGGYNYVSWYQQHPGKAPKLIIHDVIERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCWSFAGSYYVFGTGTDVTVL
5’-GAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGAGCGAGCTGACACAGCCTAGATCCGTGTCTGGCAGCCCTGGCCAGAGCGTGACCATCAGCTGTACCGGCACCAGCAGAGATGTGGGCGGCTACAACTACGTGTCCTGGTATCAGCAGCATCCCGGCAAGGCCCCCAAGCTGATCATCCACGACGTGATCGAGCGGAGCAGCGGCGTGCCCGATAGATTCAGCGGCAGCAAGAGCGGCAACACCGCCAGCCTGACAATCAGCGGACTGCAGGCCGAGGACGAGGCCGACTACTACTGTTGGAGCTTCGCCGGCAGCTACTACGTGTTCGGCACCGGCACCGATGTGACCGTGCTG-3’
QSELTQPRSVSGSPGQSVTISCTGTSRDVGGYNYVSWYQQHPGKAPKLIIHDVIERSSGVPDRFSGSKSGNTASLTISGLQAEDEADYYCWSFAGSYYVFGTGTDVTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSS
5’-CAGAGCGAGCTGACACAGCCTAGATCCGTGTCTGGCAGCCCTGGCCAGAGCGTGACCATCAGCTGTACCGGCACCAGCAGAGATGTGGGCGGCTACAACTACGTGTCCTGGTATCAGCAGCATCCCGGCAAGGCCCCCAAGCTGATCATCCACGACGTGATCGAGCGGAGCAGCGGCGTGCCCGATAGATTCAGCGGCAGCAAGAGCGGCAACACCGCCAGCCTGACAATCAGCGGACTGCAGGCCGAGGACGAGGCCGACTACTACTGTTGGAGCTTCGCCGGCAGCTACTACGTGTTCGGCACCGGCACCGATGTGACCGTGCTGGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCA-3’
AAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRS
5’-GCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCT-3’
RVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
5’-CGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3’
B細胞精製キット(CD19-positive selection kit、MACS(登録商標)beads、Miltenyi Biotec社製)を用いて、ヒト末梢血単核球から1~3×106細胞のヒト末梢血B細胞を精製した。得られたCD19+B細胞を回収後、cDNA合成キット(SMARTer(商標)RACE cDNA amplification kit、Takara Bio社製)を用いて、RNAを抽出し、さらに、前記RNAからcDNAを合成した。
5’-CTAATACGACTCACTATAGGGCAAGCAGTGGTATCAACGCAGAGT-3’
3’IGHCプライマー(配列番号176)
5’-TGAGTTCCACGACACCGTCAC-3’
3’IGHMプライマー(配列番号177)
5’-TCCAGGACAAAGTGATGGAGTC-3’
5’-GTGTGGTGGTACGGGAATTCAAGCAGTGGTATCAACGCAGAGT-3’
3’IGHJ1プライマー(配列番号179)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTGAGGAGACGGTGACCAGGG-3’
3’IGHJ2プライマー(配列番号180)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTGAAGAGACGGTGACCATTGTC-3’
3’IGHJ3プライマー(配列番号181)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTGAGGAGACGGTGACCGTGGTC-3’
5’-AGGGTCAGAGGCCAAAGGATGG-3’
3’IGLCプライマー(配列番号183)
5’-CTTGGAGCTCCTCAGAGGAG-3’
3’IGKJ1プライマー(配列番号184)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTTTGATTTCCACCTTGGTCCC-3’
3’IGKJ2プライマー(配列番号185)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTTTGATCTCCAGCTTGGTCCC-3’
3’IGKJ3プライマー(配列番号186)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTTTGATATCCACTTTGGTCCC-3’
3’IGKJ4プライマー(配列番号187)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTTTAATCTCCAGTCGTGTCCC-3’
3’IGLJ1プライマー(配列番号188)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTAGGACGGTGACCTTGGTCCC-3’
3’IGLJ2プライマー(配列番号189)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTAGGACGGTCAGCTTGGTCCC-3’
3’IGLJ4プライマー(配列番号190)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTAAAATGATCAGCTGGGTTCC-3’
3’IGLJ5プライマー(配列番号191)
5’-CTTGCCAGAGCCGCTTGTAGAGCCTAGGACGGTCAGCTCGGTCCC-3’
3’IGLJ7プライマー(配列番号192)
5’-CTTGCCAGAGCCGCTTGTAGAGCCGAGGACGGTCAGCTGGGTGCC-3’
Plat-A細胞(国立大学法人東京大学医科学研究所 北村俊雄先生より分与)に、トランスフェクション試薬(TransIT293、Takara Bio社製)を用い、hH-3M4E4L CARライブラリの発現ベクターを導入し、エコトロピックレトロウイルス(Ecotropic retrovirus)を含む培養上清を調製した。前記Plat-A細胞は、10%ウシ胎児血清(FCS)、1μg/mLピューロマイシン、10μg/mLブラストサイジン含有DMEM培地で維持したものを使用した。つぎに、前記エコトロピックウイルスを含む培養上清を添加することにより、パッケージング細胞であるPG13細胞にhH-3M4E4L CARライブラリを導入し、hH-3M4E4L CARライブラリを含むGaLV-pseudotypedレトロウイルスを調製した。そして、GaLV-pseudotypedレトロウイルスをヒト末梢血T細胞に感染させることにより、CAR-Tを調製した。PG13細胞の培地の組成は、10%FCS含有DMEM培地とし、培養条件は、37℃、5%CO2とし、湿潤雰囲気下とした。以下、特に言及しない限り細胞の培養条件は同様とした。
レトロウイルスを用いて、HLA-A*02:01をコードする核酸と、CD80、CD83、CD40、および4-1BBLをコードする核酸とを、K562細胞(HLAおよびCD19の発現無し、ATCCより入手)に導入し、抗原提示細胞(APC、K562/A2)とした。
5'-ATGGCCGTCATGGCGCCCCGAACCCTCGTCCTGCTACTCTCGGGGGCTCTGGCCCTGACCCAGACCTGGGCGGGCTCTCACTCCATGAGGTATTTCTTCACATCCGTGTCCCGGCCCGGCCGCGGGGAGCCCCGCTTCATCGCAGTGGGCTACGTGGACGACACGCAGTTCGTGCGGTTCGACAGCGACGCCGCGAGCCAGAGGATGGAGCCGCGGGCGCCGTGGATAGAGCAGGAGGGTCCGGAGTATTGGGACGGGGAGACACGGAAAGTGAAGGCCCACTCACAGACTCACCGAGTGGACCTGGGGACCCTGCGCGGCTACTACAACCAGAGCGAGGCCGGTTCTCACACCGTCCAGAGGATGTATGGCTGCGACGTGGGGTCGGACTGGCGCTTCCTCCGCGGGTACCACCAGTACGCCTACGACGGCAAGGATTACATCGCCCTGAAAGAGGACCTGCGCTCTTGGACCGCGGCGGACATGGCAGCTCAGACCACCAAGCACAAGTGGGAGGCGGCCCATGTGGCGGAGCAGTTGAGAGCCTACCTGGAGGGCACGTGCGTGGAGTGGCTCCGCAGATACCTGGAGAACGGGAAGGAGACGCTGCAGCGCACGGACGCCCCCAAAACGCATATGACTCACCACGCTGTCTCTGACCATGAAGCCACCCTGAGGTGCTGGGCCCTGAGCTTCTACCCTGCGGAGATCACACTGACCTGGCAGCGGGATGGGGAGGACCAGACCCAGGACACGGAGCTCGTGGAGACCAGGCCTGCAGGGGATGGAACCTTCCAGAAGTGGGCGGCTGTGGTGGTGCCTTCTGGACAGGAGCAGAGATACACCTGCCATGTGCAGCATGAGGGTTTGCCCAAGCCCCTCACCCTGAGATGGGAGCCGTCTTCCCAGCCCACCATCCCCATCGTGGGCATCATTGCTGGCCTGGTTCTCTTTGGAGCTGTGATCACTGGAGCTGTGGTCGCTGCTGTGATGTGGAGGAGGAAGAGCTCAGATAGAAAAGGAGGGAGCTACTCTCAGGCTGCAAGCAGTGACAGTGCCCAGGGCTCTGATGTGTCTCTCACAGCTTGTAAAGTG-3'
5'-ATGGGCCACACACGGAGGCAGGGAACATCACCATCCAAGTGTCCATACCTCAATTTCTTTCAGCTCTTGGTGCTGGCTGGTCTTTCTCACTTCTGTTCAGGTGTTATCCACGTGACCAAGGAAGTGAAAGAAGTGGCAACGCTGTCCTGTGGTCACAATGTTTCTGTTGAAGAGCTGGCACAAACTCGCATCTACTGGCAAAAGGAGAAGAAAATGGTGCTGACTATGATGTCTGGGGACATGAATATATGGCCCGAGTACAAGAACCGGACCATCTTTGATATCACTAATAACCTCTCCATTGTGATCCTGGCTCTGCGCCCATCTGACGAGGGCACATACGAGTGTGTTGTTCTGAAGTATGAAAAAGACGCTTTCAAGCGGGAACACCTGGCTGAAGTGACGTTATCAGTCAAAGCTGACTTCCCTACACCTAGTATATCTGACTTTGAAATTCCAACTTCTAATATTAGAAGGATAATTTGCTCAACCTCTGGAGGTTTTCCAGAGCCTCACCTCTCCTGGTTGGAAAATGGAGAAGAATTAAATGCCATCAACACAACAGTTTCCCAAGATCCTGAAACTGAGCTCTATGCTGTTAGCAGCAAACTGGATTTCAATATGACAACCAACCACAGCTTCATGTGTCTCATCAAGTATGGACATTTAAGAGTGAATCAGACCTTCAACTGGAATACAACCAAGCAAGAGCATTTTCCTGATAACCTGCTCCCATCCTGGGCCATTACCTTAATCTCAGTAAATGGAATTTTTGTGATATGCTGCCTGACCTACTGCTTTGCCCCAAGATGCAGAGAGAGAAGGAGGAATGAGAGATTGAGAAGGGAAAGTGTACGCCCTGTA-3'
5'-ATGTCGCGCGGCCTCCAGCTTCTGCTCCTGAGCTGCGCCTACAGCCTGGCTCCCGCGACGCCGGAGGTGAAGGTGGCTTGCTCCGAAGATGTGGACTTGCCCTGCACCGCCCCCTGGGATCCGCAGGTTCCCTACACGGTCTCCTGGGTCAAGTTATTGGAGGGTGGTGAAGAGAGGATGGAGACACCCCAGGAAGACCACCTCAGGGGACAGCACTATCATCAGAAGGGGCAAAATGGTTCTTTCGACGCCCCCAATGAAAGGCCCTATTCCCTGAAGATCCGAAACACTACCAGCTGCAACTCGGGGACATACAGGTGCACTCTGCAGGACCCGGATGGGCAGAGAAACCTAAGTGGCAAGGTGATCTTGAGAGTGACAGGATGCCCTGCACAGCGTAAAGAAGAGACTTTTAAGAAATACAGAGCGGAGATTGTCCTGCTGCTGGCTCTGGTTATTTTCTACTTAACACTCATCATTTTCACTTGTAAGTTTGCACGGCTACAGAGTATCTTCCCAGATTTTTCTAAAGCTGGCATGGAACGAGCTTTTCTCCCAGTTACCTCCCCAAATAAGCATTTAGGGCTAGTGACTCCTCACAAGACAGAACTGGTA-3'
5'-ATGGTTCGTCTGCCTCTGCAGTGCGTCCTCTGGGGCTGCTTGCTGACCGCTGTCCATCCAGAACCACCCACTGCATGCAGAGAAAAACAGTACCTAATAAACAGTCAGTGCTGTTCTTTGTGCCAGCCAGGACAGAAACTGGTGAGTGACTGCACAGAGTTCACTGAAACGGAATGCCTTCCTTGCGGTGAAAGCGAATTCCTAGACACCTGGAACAGAGAGACACACTGCCACCAGCACAAATACTGCGACCCCAACCTAGGGCTTCGGGTCCAGCAGAAGGGCACCTCAGAAACAGACACCATCTGCACCTGTGAAGAAGGCTGGCACTGTACGAGTGAGGCCTGTGAGAGCTGTGTCCTGCACCGCTCATGCTCGCCCGGCTTTGGGGTCAAGCAGATTGCTACAGGGGTTTCTGATACCATCTGCGAGCCCTGCCCAGTCGGCTTCTTCTCCAATGTGTCATCTGCTTTCGAAAAATGTCACCCTTGGACAAGCTGTGAGACCAAAGACCTGGTTGTGCAACAGGCAGGCACAAACAAGACTGATGTTGTCTGTGGTCCCCAGGATCGGCTGAGAGCCCTGGTGGTGATCCCCATCATCTTCGGGATCCTGTTTGCCATCCTCTTGGTGCTGGTCTTTATCAAAAAGGTGGCCAAGAAGCCAACCAATAAGGCCCCCCACCCCAAGCAGGAACCCCAGGAGATCAATTTTCCCGACGATCTTCCTGGCTCCAACACTGCTGCTCCAGTGCAGGAGACTTTACATGGATGCCAACCGGTCACCCAGGAGGATGGCAAAGAGAGTCGCATCTCAGTGCAGGAGAGACAG-3'
5'-ATGGAATACGCCTCTGACGCTTCACTGGACCCCGAAGCCCCGTGGCCTCCCGCGCCCCGCGCTCGCGCCTGCCGCGTACTGCCTTGGGCCCTGGTCGCGGGGCTGCTGCTGCTGCTGCTGCTCGCTGCCGCCTGCGCCGTCTTCCTCGCCTGCCCCTGGGCCGTGTCCGGGGCTCGCGCCTCGCCCGGCTCCGCGGCCAGCCCGAGACTCCGCGAGGGTCCCGAGCTTTCGCCCGACGATCCCGCCGGCCTCTTGGACCTGCGGCAGGGCATGTTTGCGCAGCTGGTGGCCCAAAATGTTCTGCTGATCGATGGGCCCCTGAGCTGGTACAGTGACCCAGGCCTGGCAGGCGTGTCCCTGACGGGGGGCCTGAGCTACAAAGAGGACACGAAGGAGCTGGTGGTGGCCAAGGCTGGAGTCTACTATGTCTTCTTTCAACTAGAGCTGCGGCGCGTGGTGGCCGGCGAGGGCTCAGGCTCCGTTTCACTTGCGCTGCACCTGCAGCCACTGCGCTCTGCTGCTGGGGCCGCCGCCCTGGCTTTGACCGTGGACCTGCCACCCGCCTCCTCCGAGGCTCGGAACTCGGCCTTCGGTTTCCAGGGCCGCTTGCTGCACCTGAGTGCCGGCCAGCGCCTGGGCGTCCATCTTCACACTGAGGCCAGGGCACGCCATGCCTGGCAGCTTACCCAGGGCGCCACAGTCTTGGGACTCTTCCGGGTGACCCCCGAAATCCCAGCCGGACTCCCTTCACCGAGGTCGGAA-3'
前記(3)の第1の候補CAR-Tと、前記(4)で調製したAPCとを用いて、スクリーニングを行なった。具体的には、2×106第1の候補CAR-Tと、1×105APCとを各ウェル(24ウェルプレート)に播種後、7日間共培養した。前記APCは、20Gyでγ線処理後、1~10μg/mLでNY-ESO-1157-165ペプチド(NY-ESO-1157ペプチド、SLLMWITQC:配列番号206)をパルスし、前記パルス後にHLA-A*02:01に結合していないペプチドを除去した細胞を用いた。また、前記第1の候補CAR-TとAPCとの共培養に用いた培地の組成は、50μg/mLゲンタマイシン、10%ヒトAB血清(Sigma社製)、10IU/mL ヒトIL-2(Roche社製)、および10ng/mL ヒトIL-15(PeproTech社製)含有RPMI1640培地とした。前記共培養後、第1の候補CAR-Tを回収し、同様の条件でAPCとの共培養をさらに2回(合計3回)実施した。これにより、A2/NY-ESO-1157と結合するhH-3M4E4L CARを発現する第1の候補CAR-Tを富化した。
5'-ATCCCAGTGTGGTGGTACGGG-3'
リバースプライマー1(配列番号200)
5'-GGGTACATCACTTCGATTGC-3'
hH-3M4E4L CARライブラリの発現ベクターに代えて、pMX/H1-3M4E5L発現ベクターまたはpMX/H73-3M4E5L発現ベクターを用いた以外は、前記(3)と同様にして、GaLV-pseudotypedレトロウイルスを含む培養上清を調製した。つぎに、前記GaLV-pseudotypedレトロウイルスを含む培養上清を添加することにより、Jurkat76細胞にpMX/H1-3M4E5L発現ベクターまたはpMX/H73-3M4E5L発現ベクターを導入し、scFvとしてH1-3M4E5LまたはH73-3M4E5Lを含むCAR(H1-3M4E5L CARまたはH73-3M4E5L CAR)を発現させた。前記CAR発現後のJurkat76細胞を、前記PE標識anti-human NGFR mAbと反応させた。前記反応後、前記抗PEマイクロビーズを用いて、H1-3M4E5L CARおよびH73-3M4E5L CARを発現するJurkat76細胞(H1-3M4E5L CAR-TまたはH73-3M4E5L CAR T)を精製した。H1-3M4E5L CAR-TおよびH73-3M4E5L CAR Tについて、5μg/mL PE標識A2/NY-ESO-1157テトラマーで染色後、V450標識 anti-human NGFR mAbで染色した。前記染色後のH1-3M4E5L CAR-TおよびH73-3M4E5L CAR Tについて、フローサイトメトリーにより解析した。ポジティブコントロールは、pMX/H1-3M4E5L発現ベクターまたはpMX/H73-3M4E5L発現ベクターに代えて、タグ付加3M4E5LH CARをコードする核酸を含む発現ベクターを用いた以外は、同様に実施した。また、コントロール1は、A2/NY-ESO-1157テトラマーに代えて、A2/HIV-Gag77テトラマーを用いた以外は、同様にして実施した。コントロール2は、pMX/H1-3M4E5L発現ベクターまたはpMX/H73-3M4E5L発現ベクターを導入しなかった以外は、同様にして実施した。FSC+SSC+NGFR+細胞にゲートをかけた結果を図2に示す。
つぎに、前記(6)で得られたH1-3M4E5L CAR-TおよびH73-3M4E5L CAR Tと、3M4E5LH CARを発現するJurkat76細胞(3M4E5LH CAR-T)について、所定濃度(0.0004194304、0.001048576、0.00262144、0.0065536、0.016384、0.04096、0.1024、0.256、0.64、1.6、4、または10μg/mL)のPE標識A2/NY-ESO-1157テトラマーで染色後、V450標識 anti-human NGFR mAbで染色した以外は、前記(6)と同様にして、フローサイトメトリーにより解析した。そして、A2/NY-ESO-1157テトラマーの各濃度における、A2/NY-ESO-1157テトラマー陽性細胞の割合を算出した。この結果を、図3に示す。
つぎに、H1-3M4E5L CAR-TおよびH73-3M4E5L CAR-Tが、前記標的抗原との結合により、活性化するかを検討した。具体的には、3×105細胞のH1-3M4E5L CAR-TまたはH73-3M4E5L CAR-Tと、5×104細胞のT2細胞とを各ウェル(96ウェルプレート)に播種後、5時間共培養した。前記T2細胞(HLA-A*02:01発現細胞)は、所定濃度(0.0005、0.0015、0.004、0.01、0.04、0.122、0.366、1.11、3.33または10μg/mL)でNY-ESO-1157ペプチドをパルスした細胞(T2 + NY-ESO-1157)を用いた。また、前記共培養の培養液は、10%FCS含有RPMI1640培地とした。前記共培養後、H1-3M4E5L CAR-TおよびH73-3M4E5L CAR-Tを回収し、FITC標識anti-human CD69 mAb(clone:FN50)およびV450標識anti-human NGFR mAbで染色した。前記染色後のH1-3M4E5L CAR-TおよびH73-3M4E5L CAR-Tについて、前記フローサイトメーターを用いて、NGFR+細胞について、CD69の平均蛍光強度を測定した。ポジティブコントロールは、前記(6)で得られた3M4E5HL CAR-Tを用いた以外は、同様に実施した。また、コントロール1は、NY-ESO-1157ペプチドに代えて、HIV Gag77-85ペプチドをパルスしたT2細胞(T2 + HIV Gag77)を用いた以外は、同様にして実施した。コントロール2は、pMX/H1-3M4E5L発現ベクターを導入しなかった以外は、同様にして実施した。そして、各ペプチドの濃度について、コントロール2におけるCD69の平均蛍光強度を基準として、各サンプルのCD69の発現上昇の割合を算出した。これらの結果を図4および5に示す。
第1のCARライブラリからスクリーニングされた重鎖可変領域(HA)を用いて、第2のCARライブラリを調製した。具体的には、タグ付加3M4E5LH CARをコードする核酸が導入されたpMX発現ベクターについて、重鎖可変領域(3M4E5H)をコードする核酸を、重鎖可変領域(HA)をコードする核酸で置換した(3M4E5L-H1発現ベクター)。つぎに、3M4E5L-H1発現ベクターについて、軽鎖可変領域(3M4E5L)をコードする核酸を、前記(2)の軽鎖可変領域をコードする核酸のライブラリで置換した。これにより、第2のCARライブラリを含む発現ベクター(hK-H1 CARライブラリの発現ベクター)を調製した。なお、hK-H1 CARライブラリにおける重鎖可変領域の不均一性(heterogeneity)は、制限酵素マッピングおよび各領域をサンガー法によりシークエンスすることで、約1×106種類であることを確認した。
hH-3M4E4L CARライブラリの発現ベクターに代えて、hK-H1 CARライブラリの発現ベクターを用いた以外は、前記(3)と同様にして、hK-H1 CARライブラリが導入されたT細胞を精製し、これを第2の候補CAR-Tとした。
つぎに、前記第1の候補CAR-Tに代えて、前記第2の候補CAR-Tを用いた以外は、前記(5)と同様にして、A2/NY-ESO-1157と結合するCARを発現する第2の候補CAR-Tの存在を確認した(donor 1)。同様の試験を、異なる健常者由来のヒト末梢血B細胞由来の重鎖可変領域の核酸を含むライブラリを用いて実施した(donor 2)。また、ポジティブコントロールは、hK-H1 CARライブラリの発現ベクターに代えて、pMX/3M4E5L-H1発現ベクターを用いた以外は、同様に実施した。また、コントロール1は、A2/NY-ESO-1157テトラマーに代えて、A2/HIV-Gag77テトラマーを用いた以外は、同様にして実施した。コントロール2は、hH-3M4E5L CARライブラリの発現ベクターを導入しなかった以外は、同様にして実施した。FSC+SSC+NGFR+細胞にゲートをかけた結果を図6に示す。
hH-3M4E4L CARライブラリの発現ベクターのscFvをコードするポリヌクレオチドに代えて、前記(11)の各scFvをコードするポリヌクレオチドが導入されたpMX/CAR発現ベクターを用いた以外は、前記(3)と同様にして、GaLV-pseudotypedレトロウイルスを含む培養上清を調製した。つぎに、前記GaLV-pseudotypedレトロウイルスを含む培養上清を添加することにより、Jurkat76細胞に各scFvをコードするポリヌクレオチドが導入されたpMX/CAR発現ベクターを導入し、scFvとしてK52-H1、K73-H1、K121-H1、K124-H1、K125-H1、K131-H1、K145-H1、K151-H1、K160-H1、またはK173-H1を含むCARを発現させた。前記CAR発現後のJurkat76細胞を、前記PE標識anti-human NGFR mAbと反応させた。前記反応後、前記抗PEマイクロビーズを用いて、各CARを発現するJurkat76細胞を精製した。精製された各CARを発現するJurkat76細胞について、5μg/mL PE標識A2/NY-ESO-1157テトラマーで染色後、V450標識 anti-human NGFR mAbで染色した。前記染色後の各CARを発現するJurkat76細胞について、フローサイトメトリーにより解析した。また、各scFvを導入されたpMX/CAR発現ベクターに代えて、pMX/H1-3M4E5L発現ベクターを用いた以外は、同様に実施した。ポジティブコントロールは、各scFvを導入されたpMX/CAR発現ベクターに代えて、タグ付加3M4E5LH CARをコードする核酸を含む発現ベクターを用いた以外は、同様に実施した。また、コントロール1は、A2/NY-ESO-1157テトラマーに代えて、A2/HIV-Gag77テトラマーを用いた以外は、同様にして実施した。コントロール2は、各scFvを導入されたpMX/CAR発現ベクターを導入しなかった以外は、同様にして実施した。NGFR+細胞にゲートをかけた結果を図7(A)および(B)に示す。
本発明のCARライブラリを用いた本発明のスクリーニング方法により、A2/NY-ESO-1157に結合するscFvをスクリーニングできることを確認した。
前記実施例1(1)の3M4E5LH scFvをコードする核酸の5’側の軽鎖可変領域をコードする核酸を、前記ヒト末梢血B細胞由来の軽鎖可変領域の核酸を含むライブラリで置換することにより、hL-3M4E4H scFvライブラリを調製した。そして、3M4E5LH scFvをコードする核酸に代えて、hL-3M4E4H scFvライブラリを用いた以外は、前記実施例1(1)と同様にして、タグが付加されたhL-3M4E4H CARライブラリを調製した。タグが付加されたhL-3M4E4H CARライブラリを、pMX発現ベクターに導入し、hL-3M4E4H CARライブラリの発現ベクターを調製した。なお、hL-3M4E4H CARライブラリの重鎖可変領域の不均一性は、制限酵素マッピングおよび各領域をサンガー法によりシークエンスすることで確認した。
前記hH-3M4E4L CARライブラリの発現ベクターに代えて、hL-3M4E4H CARライブラリの発現ベクターを用いた以外は、前記実施例1(3)と同様にして、hL-3M4E4H CARライブラリが導入されたT細胞を精製し、これを第1の候補CAR-Tとした。
前記実施例1(3)の第1の候補CAR-Tに代えて、前記実施例2(2)の第1の候補CAR-Tを用いた以外は、前記実施例1(5)と同様にして、A2/NY-ESO-1157と結合するCARを発現する第1の候補CAR-Tの存在を確認した(donor 1)。同様の試験を、異なる健常者由来のヒト末梢血B細胞由来の重鎖可変領域の核酸を含むライブラリを用いて実施した(donor 2)。また、ポジティブコントロールは、hH-3M4E4L CARライブラリの発現ベクターに代えて、タグ付加3M4E5LH CARをコードする核酸を含む発現ベクターを用いた以外は、同様に実施した。また、コントロール1は、A2/NY-ESO-1157テトラマーに代えて、A2/HIV-Gag77テトラマーを用いた以外は、同様にして実施した。コントロール2は、hL-3M4E4H CARライブラリの発現ベクターを導入しなかった以外は、同様にして実施した。FSC+SSC+NGFR+細胞にゲートをかけた結果を図8に示す。
hH-3M4E4L CARライブラリの発現ベクターのscFvをコードするポリヌクレオチドに代えて、前記実施例2(3)の各scFvをコードするポリヌクレオチドが導入されたpMX/CAR発現ベクターを用いた以外は、前記実施例1(3)と同様にして、GaLV-pseudotypedレトロウイルスを含む培養上清を調製した。つぎに、前記GaLV-pseudotypedレトロウイルスを含む培養上清を添加することにより、Jurkat76細胞に各scFvをコードするポリヌクレオチドが導入されたpMX/CAR発現ベクターを導入し、scFvとしてL1-3M4E5、L66-3M4E5、L73-3M4E5、L80-3M4E5、L88-3M4E5、L102-3M4E5、またはL124-3M4E5を含むCARを発現させた。前記CAR発現後のJurkat76細胞を、前記PE標識anti-human NGFR mAbと反応させた。前記反応後、前記抗PEマイクロビーズを用いて、各CARを発現するJurkat76細胞を精製した。精製された各CARを発現するJurkat76細胞について、5μg/mL PE標識A2/NY-ESO-1157テトラマーで染色後、V450標識 anti-human NGFR mAbで染色した。前記染色後の各CARを発現するJurkat76細胞について、フローサイトメトリーにより解析した。ポジティブコントロールは、各scFvを導入されたpMX/CAR発現ベクターに代えて、タグ付加3M4E5LH CARをコードする核酸を含む発現ベクターを用いた以外は、同様に実施した。また、コントロール1は、A2/NY-ESO-1157テトラマーに代えて、A2/HIV-Gag77テトラマーを用いた以外は、同様にして実施した。コントロール2は、各scFvを導入されたpMX/CAR発現ベクターを導入しなかった以外は、同様にして実施した。NGFR+細胞にゲートをかけた結果を図9に示す。
つぎに、前記実施例2(4)で得られたL1-3M4E5、L66-3M4E5、L73-3M4E5、L80-3M4E5、L88-3M4E5、L102-3M4E5、またはL124-3M4E5を含むCARを発現するJurkat76細胞と、3M4E5LH CAR-Tについて、所定濃度(0.0004194304、0.001048576、0.00262144、0.0065536、0.016384、0.04096、0.1024、0.256、0.64、1.6、4、または10μg/mL)のPE標識A2/NY-ESO-1157テトラマーで染色後、V450標識 anti-human NGFR mAbで染色した以外は、前記(6)と同様にして、フローサイトメトリーにより解析した。そして、A2/NY-ESO-1157テトラマーの各濃度における、A2/NY-ESO-1157テトラマー陽性細胞の割合を算出した。この結果を、図10に示す。
つぎに、scFvとしてL1-3M4E5、L66-3M4E5、L73-3M4E5、L80-3M4E5、L88-3M4E5、L102-3M4E5、またはL124-3M4E5を発現するCAR-Tが、前記標的抗原との結合により、活性化するかを検討した。具体的には、3×105細胞の各CAR-Tと、5×104細胞のT2細胞とを各ウェル(96ウェルプレート)に播種後、5時間共培養した。前記T2細胞(HLA-A*02:01発現細胞)は、所定濃度(0.0005、0.0015、0.004、0.01、0.04、0.122、0.366、1.11、3.33または10μg/mL)でNY-ESO-1157-165ペプチドをパルスした細胞(T2 + NY-ESO-1157)を用いた。また、前記共培養の培養液は、10%FCS含有RPMI1640培地とした。前記共培養後、各CAR-Tを回収し、FITC標識anti-human CD69 mAb(clone:FN50)およびV450標識anti-human NGFR mAbで染色した。前記染色後の各CAR-Tについて、前記フローサイトメーターを用いて、NGFR+細胞について、CD69の平均蛍光強度を測定した。ポジティブコントロールは、pMX/H1-3M4E5L発現ベクターに代えて、タグ付加3M4E5LH CARをコードする核酸を含む発現ベクターを用いた以外は、同様に実施した。また、コントロール1は、NY-ESO-1157-165ペプチドに代えて、HIV Gag77-85ペプチドをパルスしたT2細胞(T2 + HIV Gag77)を用いた以外は、同様にして実施した。コントロール2は、前記発現ベクターを導入しなかった以外は、同様にして実施した。そして、コントロール2におけるCD69の平均蛍光強度を基準として、各サンプルのCD69の発現上昇の割合を算出した。これらの結果を図11に示す。
本発明のCARライブラリを用いた本発明のスクリーニング方法により、CD19に結合するscFvをスクリーニングできることを確認した。
前記A2/NY-ESO-1157特異的抗体の重鎖可変領域および軽鎖可変領域に代えて、ヒトCD19特異的抗体(clone:FMC63)の重鎖可変領域および軽鎖可変領域を用いた以外は、前記実施例1(1)と同様にして、FMC63HL scFvまたはFMC63LH scFvをコードする核酸を調製し、3’末端にタグを付加したタグ付加FMC63HL CARおよびタグ付加FMC63LH CARをコードする核酸を取得した。各核酸は、pMX発現ベクターに導入した。
EVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGVIWGSETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYYYGGSYAMDYWGQGTSVTVSSGSTSGSGKPGSGEGSTKGDIQMTQTTSSLSASLGDRVTISCRASQDISKYLNWYQQKPDGTVKLLIYHTSRLHSGVPSRFSGSGSGTDYSLTISNLEQEDIATYFCQQGNTLPYTFGGGTKLEIT
5'-GAAGTGAAACTGCAAGAGTCTGGCCCTGGACTGGTGGCCCCATCTCAGTCTCTGAGCGTGACCTGTACAGTCAGCGGAGTGTCCCTGCCTGATTACGGCGTGTCCTGGATCAGACAGCCTCCTCGGAAAGGCCTGGAATGGCTGGGAGTGATCTGGGGCAGCGAGACAACCTACTACAACAGCGCCCTGAAGTCCCGGCTGACCATCATCAAGGACAACTCCAAGAGCCAGGTGTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTATTGCGCCAAGCACTACTACTACGGCGGCAGCTACGCTATGGACTATTGGGGCCAGGGCACCAGCGTTACAGTGTCCTCGGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGATATCCAGATGACCCAGACAACAAGCAGCCTGAGCGCCAGCCTGGGCGATAGAGTGACCATCAGCTGTAGAGCCAGCCAGGACATCAGCAAGTACCTGAACTGGTATCAGCAGAAACCCGACGGCACCGTGAAGCTGCTGATCTACCACACCAGCAGACTGCACAGCGGCGTGCCAAGCAGATTTTCTGGCAGCGGCTCTGGCACCGACTACAGCCTGACCATCTCCAACCTGGAACAAGAGGATATCGCTACCTACTTCTGCCAGCAAGGCAACACCCTGCCTTACACCTTTGGCGGAGGCACCAAGCTGGAAATCACA-3'
DIQMTQTTSSLSASLGDRVTISCRASQDISKYLNWYQQKPDGTVKLLIYHTSRLHSGVPSRFSGSGSGTDYSLTISNLEQEDIATYFCQQGNTLPYTFGGGTKLEITGSTSGSGKPGSGEGSTKGEVKLQESGPGLVAPSQSLSVTCTVSGVSLPDYGVSWIRQPPRKGLEWLGVIWGSETTYYNSALKSRLTIIKDNSKSQVFLKMNSLQTDDTAIYYCAKHYYYGGSYAMDYWGQGTSVTVSS
5'-GATATCCAGATGACCCAGACAACAAGCAGCCTGAGCGCCAGCCTGGGCGATAGAGTGACCATCAGCTGTAGAGCCAGCCAGGACATCAGCAAGTACCTGAACTGGTATCAGCAGAAACCCGACGGCACCGTGAAGCTGCTGATCTACCACACCAGCAGACTGCACAGCGGCGTGCCAAGCAGATTTTCTGGCAGCGGCTCTGGCACCGACTACAGCCTGACCATCTCCAACCTGGAACAAGAGGATATCGCTACCTACTTCTGCCAGCAAGGCAACACCCTGCCTTACACCTTTGGCGGAGGCACCAAGCTGGAAATCACAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGAAACTGCAAGAGTCTGGCCCTGGACTGGTGGCCCCATCTCAGTCTCTGAGCGTGACCTGTACAGTCAGCGGAGTGTCCCTGCCTGATTACGGCGTGTCCTGGATCAGACAGCCTCCTCGGAAAGGCCTGGAATGGCTGGGAGTGATCTGGGGCAGCGAGACAACCTACTACAACAGCGCCCTGAAGTCCCGGCTGACCATCATCAAGGACAACTCCAAGAGCCAGGTGTTCCTGAAGATGAACAGCCTGCAGACCGACGACACCGCCATCTACTATTGCGCCAAGCACTACTACTACGGCGGCAGCTACGCTATGGACTATTGGGGCCAGGGCACCAGCGTTACAGTGTCCTCG-3'
前記実施例3(1)のFMC63HL scFvをコードする核酸の5’側の重鎖可変領域をコードする核酸を、前記ヒト末梢血B細胞由来の重鎖可変領域の核酸を含むライブラリで置換することにより、hH-FMC63L scFvライブラリを調製した。そして、3M4E5LH scFvをコードする核酸に代えて、hH-FMC63L scFvライブラリを用いた以外は、前記実施例1(1)と同様にして、タグが付加されたhH-FMC63L CARライブラリを調製した。タグが付加されたhH-FMC63L CARライブラリを、pMX発現ベクターに導入し、hH-FMC63L CARライブラリの発現ベクターを調製した。なお、hH-FMC63L CARライブラリの重鎖可変領域の不均一性は、制限酵素マッピングおよび各領域をサンガー法によりシークエンスすることで、約1×106種類であることを確認した。
前記hH-3M4E4L CARライブラリの発現ベクターに代えて、hH-FMC63L CARライブラリの発現ベクターを用いた以外は、前記実施例1(3)と同様にして、hH-FMC63L CARライブラリが導入されたT細胞を精製し、これを第1の候補CAR-Tとした。
レトロウイルスを用いて、ヒトCD19をコードする核酸を、K562細胞に導入し、抗原提示細胞(APC-CD19)とした。
5'-ATGCCACCTCCTCGCCTCCTCTTCTTCCTCCTCTTCCTCACCCCCATGGAAGTCAGGCCCGAGGAACCTCTAGTGGTGAAGGTGGAAGAGGGAGATAACGCTGTGCTGCAGTGCCTCAAGGGGACCTCAGATGGCCCCACTCAGCAGCTGACCTGGTCTCGGGAGTCCCCGCTTAAACCCTTCTTAAAACTCAGCCTGGGGCTGCCAGGCCTGGGAATCCACATGAGGCCCCTGGCCATCTGGCTTTTCATCTTCAACGTCTCTCAACAGATGGGGGGCTTCTACCTGTGCCAGCCGGGGCCCCCCTCTGAGAAGGCCTGGCAGCCTGGCTGGACAGTCAATGTGGAGGGCAGCGGGGAGCTGTTCCGGTGGAATGTTTCGGACCTAGGTGGCCTGGGCTGTGGCCTGAAGAACAGGTCCTCAGAGGGCCCCAGCTCCCCTTCCGGGAAGCTCATGAGCCCCAAGCTGTATGTGTGGGCCAAAGACCGCCCTGAGATCTGGGAGGGAGAGCCTCCGTGTCTCCCACCGAGGGACAGCCTGAACCAGAGCCTCAGCCAGGACCTCACCATGGCCCCTGGCTCCACACTCTGGCTGTCCTGTGGGGTACCCCCTGACTCTGTGTCCAGGGGCCCCCTCTCCTGGACCCATGTGCACCCCAAGGGGCCTAAGTCATTGCTGAGCCTAGAGCTGAAGGACGATCGCCCGGCCAGAGATATGTGGGTAATGGAGACGGGTCTGTTGTTGCCCCGGGCCACAGCTCAAGACGCTGGAAAGTATTATTGTCACCGTGGCAACCTGACCATGTCATTCCACCTGGAGATCACTGCTCGGCCAGTACTATGGCACTGGCTGCTGAGGACTGGTGGCTGGAAGGTCTCAGCTGTGACTTTGGCTTATCTGATCTTCTGCCTGTGTTCCCTTGTGGGCATTCTTCATCTTCAAAGAGCCCTGGTCCTGAGGAGGAAAAGAAAGCGAATGACTGACCCCACCAGGAGATTCTTCAAAGTGACGCCTCCCCCAGGAAGCGGGCCCCAGAACCAGTACGGGAACGTGCTGTCTCTCCCCACACCCACCTCAGGCCTCGGACGCGCCCAGCGTTGGGCCGCAGGCCTGGGGGGCACTGCCCCGTCTTATGGAAACCCGAGCAGCGACGTCCAGGCGGATGGAGCCTTGGGGTCCCGGAGCCCGCCGGGAGTGGGCCCAGAAGAAGAGGAAGGGGAGGGCTATGAGGAACCTGACAGTGAGGAGGACTCCGAGTTCTATGAGAACGACTCCAACCTTGGGCAGGACCAGCTCTCCCAGGATGGCAGCGGCTACGAGAACCCTGAGGATGAGCCCCTGGGTCCTGAGGATGAAGACTCCTTCTCCAACGCTGAGTCTTATGAGAACGAGGATGAAGAGCTGACCCAGCCGGTCGCCAGGACAATGGACTTCCTGAGCCCTCATGGGTCAGCCTGGGACCCCAGCCGGGAAGCAACCTCCCTGGGGTCCCAGTCCTATGAGGATATGAGAGGAATCCTGTATGCAGCCCCCCAGCTCCGCTCCATTCGGGGCCAGCCTGGACCCAATCATGAGGAAGATGCAGACTCTTATGAGAACATGGATAATCCCGATGGGCCAGACCCAGCCTGGGGAGGAGGGGGCCGCATGGGCACCTGGAGCACCAGG-3'
前記実施例3(3)の第1の候補CAR-Tと、前記実施例3(4)で調製したAPC-CD19とを用いて、スクリーニングを行なった。具体的には、2×106第1の候補CAR-Tと、1×105APC-CD19とを各ウェル(24ウェルプレート)に播種後、7日間共培養した。前記APC-CD19は、20Gyでγ線処理した細胞を用いた。また、前記第1の候補CAR-TとAPC-CD19との共培養に用いた培地の組成は、50μg/mLゲンタマイシン、10%ヒトAB血清(Sigma社製)、10IU/mL ヒトIL-2(Roche社製)、および10ng/mL ヒトIL-15(PeproTech社製)含有RPMI1640培地とした。前記共培養後、第1の候補CAR-Tを回収し、同様の条件でAPC-C19との共培養をさらに2回(合計3回)実施した。これにより、ヒトCD19と結合するhH-FMC63L CARを発現する第1の候補CAR-Tを富化した。
本発明のCARライブラリを用いた本発明のスクリーニング方法により得られたCARが、標的抗原に対して高い特異性を有することを確認した。
本発明のCARライブラリを用いた本発明のスクリーニング方法により得られたCARが、スクリーニングに用いた標的抗原に結合する抗体と異なる抗原認識性を示すことを確認した。
本発明のCARライブラリを用いた本発明のスクリーニング方法により得られたCARについて、重鎖可変領域と、軽鎖可変領域との順序を交換しても、抗原認識性を維持することを確認した。
本発明のCARライブラリを用いた本発明のスクリーニング方法により得られたCARが、スクリーニングに用いた標的抗原に結合する抗体と異なる抗原認識性を示すことおよびCARにおけるscFvの交差反応性を制御しながらスクリーニングできることを確認した。
前記実施例6と同様にして、3M4E5H-L1 CARを発現するCD8陽性T細胞およびCD4陽性T細胞を作製した。つぎに、3×105細胞のT細胞と、5×104細胞のT2細胞とを各ウェル(96ウェルプレート)に播種後、5時間共培養した。前記T2細胞(HLA-A*02:01発現細胞)は、10μg/mLでNY-ESO-1157ペプチドまたはNY-ESO-1157ペプチドの変異ペプチドをパルスした細胞(T2 + NY-ESO-1157)を用いた。前記変異ペプチドは、NY-ESO-1157ペプチドのアミノ酸配列において、各アミノ酸を1つずつアラニンに置換したペプチドである。前記共培養後、培養上清を回収し、前記実施例4と同様にして、CAR-TにおけるIL-2の産生量を測定した。コントロールは、3M4E5LH CARを発現するCD8陽性T細胞またはCD4陽性T細胞を用いた以外は、同様に実施した。また、コントロール1は、A2/NY-ESO-1157ペプチドに代えて、A2/HIV-Gag77ペプチドを用いた以外は、同様にして実施した。コントロール2は、発現ベクターを導入しなかったCD8陽性T細胞またはCD4陽性T細胞を用いた以外は、同様にして実施した。そして、コントロール2におけるサイトカインの産生量を基準として、各サンプルのサイトカインの産生増加の割合を算出した。これらの結果を図20に示す。
前記実施例7(1)で示したように、3M4E5LH CARは、NY-ESO-1157ペプチドの160~162番のアミノ酸を認識するのに対して、3M4E5H-L1 CARは、NY-ESO-1157ペプチドの158および160~162番、164番目のアミノ酸を認識する。3M4E5LH CARおよび3M4E5H-L1 CARの他のペプチドに対する交差反応性を検討するため、HLA-A*02:01に結合し、かつ3M4E5LH CARおよび3M4E5H-L1 CARと交差反応しうるペプチドを含むヒトタンパク質について、アルゴリズム(netMHC4.0、http://www.cbs.dtu.dk/services/NetMHC/)を用いて予測した。タンパク質のデータベースとしては、uniprotKB/swissprotを用いた。また、得られたタンパク質におけるペプチドにおいて、NY-ESO-1157ペプチドと相同性が高い相同ペプチド(homologous peptide)を、アルゴリズム(ScanProsit、https://prosite.expasy.org/scanprosite/)を用いて絞り込んだ。その結果、交差反応しうるペプチド(配列番号207:GLRMWIKQV)を含む、Lysophospholipase-like protein 1(LYPLAL1)タンパク質がヒットした。そこで、K562細胞に、LYPLAL1タンパク質を発現させ、3M4E5LH CARおよび3M4E5H-L1 CARの交差反応性を検討した。
5'-ATGGCGGCTGCATCTGGAAGCGTGCTGCAGAGATGTATCGTGTCCCCAGCCGGCAGACATAGCGCCAGCCTGATTTTTCTGCACGGCAGCGGCGATTCTGGCCAGGGACTGAGAATGTGGATCAAACAGGTGCTGAACCAGGACCTGACCTTCCAGCACATCAAGATCATCTACCCCACCGCTCCACCTCGGAGCTACACACCTATGAAGGGCGGCATCAGCAACGTTTGGTTCGACCGGTTCAAGATCACCAACGACTGCCCCGAGCACCTGGAATCCATCGACGTGATGTGTCAGGTGCTCACCGACCTGATCGACGAGGAAGTGAAGTCCGGCATCAAGAAGAACAGAATCCTGATCGGCGGCTTCAGCATGGGCGGCTGTATGGCCATTCACCTGGCCTACAGAAACCACCAGGATGTGGCTGGCGTGTTCGCCCTGAGCAGCTTTCTGAACAAAGCCAGCGCCGTGTATCAGGCCCTGCAGAAATCTAACGGCGTGCTGCCTGAGCTGTTCCAGTGTCATGGCACAGCCGATGAGCTGGTGCTGCACTCTTGGGCCGAAGAGACAAATAGCATGCTGAAAAGCCTGGGCGTGACCACCAAGTTCCACAGCTTCCCCAACGTGTACCACGAGCTGAGCAAGACCGAGCTGGACATCCTGAAACTGTGGATTCTGACCAAGCTGCCCGGCGAGATGGAAAAGCAGAAG-3'
5'-ATGCAGGCCGAAGGCCGGGGCACAGGGGGTTCGACGGGCGATGCTGATGGCCCAGGAGGCCCTGGCATTCCTGATGGCCCAGGGGGCAATGCTGGCGGCCCAGGAGAGGCGGGTGCCACGGGCGGCAGAGGTCCCCGGGGCGCAGGGGCAGCAAGGGCCTCGGGGCCGGGAGGAGGCGCCCCGCGGGGTCCGCATGGCGGCGCGGCTTCAGGGCTGAATGGATGCTGCAGATGCGGGGCCAGGGGGCCGGAGAGCCGCCTGCTTGAGTTCTACCTCGCCATGCCTTTCGCGACACCCATGGAAGCAGAGCTGGCCCGCAGGAGCCTGGCCCAGGATGCCCCACCGCTTCCCGTGCCAGGGGTGCTTCTGAAGGAGTTCACTGTGTCCGGCAACATACTGACTATCCGACTGACTGCTGCAGACCACCGCCAACTGCAGCTCTCCATCAGCTCCTGTCTCCAGCAGCTTTCCCTGTTGATGTGGATCACGCAGTGCTTTCTGCCCGTGTTTTTGGCTCAGCCTCCCTCAGGGCAGAGGCGC-3'
本発明のCARライブラリを用いた本発明のスクリーニング方法により得られたCARについて、重鎖可変領域と、軽鎖可変領域との順序を交換することで、抗原特異性が変化することを確認した。
本発明の第1のスクリーニング方法により、標的抗原における認識部位が異なるscFvをスクリーニングできることを確認した。
QSVLTQPPSVSGAPGQRVTISCTGSSSNLGAGYDVHWYQQLPGTAPKLLIYGSTTRPSGIPDRFSGSKSGTSATLGITGLQTGDEADYYCGTWDSSLSAGVFGGGTQLTVLGSTSGSGKPGSGEGSTKGEVQLLESGGGLVQPGGSLRLSCAASGFTFSTYQMSWVRQAPGKGLEWVSGIVSSGGSTAYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAGELLPYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
5'-CAGTCTGTGCTGACGCAGCCGCCCTCAGTGTCTGGGGCCCCAGGGCAGAGGGTCACCATCTCCTGCACTGGGAGCAGCTCCAACCTCGGGGCAGGTTATGATGTACACTGGTACCAGCAGCTTCCAGGAACAGCCCCCAAACTCCTCATCTATGGTAGCACCACTCGGCCCTCAGGGATTCCTGACCGATTCTCTGGCTCCAAGTCTGGCACGTCAGCCACCCTGGGCATCACCGGACTCCAGACTGGGGACGAGGCCGATTATTACTGCGGAACATGGGATAGCAGCCTGAGTGCTGGGGTGTTCGGCGGAGGCACCCAGCTGACCGTCCTCGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTGCTGGAATCTGGCGGCGGACTGGTGCAGCCTGGCGGATCTCTGAGACTGAGCTGTGCCGCCAGCGGCTTCACCTTCAGCACCTACCAGATGAGCTGGGTGCGCCAGGCCCCTGGCAAAGGACTGGAATGGGTGTCCGGCATCGTGTCCAGCGGCGGCTCTACAGCCTACGCCGATAGCGTGAAGGGCCGGTTCACCATCAGCCGGGACAACAGCAAGAACACCCTGTACCTGCAGATGAACAGCCTGAGAGCCGAGGACACCGCCGTGTACTATTGTGCCGGGGAGCTGCTGCCCTACTACGGCATGGATGTGTGGGGCCAGGGCACCACCGTGACAGTGTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
EIVLTQSPATLSVSPGERATLSCRASQSVSSNLAWYQQKPGQAPRLLIYGASTRATGIPARFSGSGSGTEFTLTISRLEPEDFAVYYCQQYNNWPPKFTFGPGTKVDIRGSTSGSGKPGSGEGSTKGQVQLQESGPGLVKPSDTLSLTCLVSGGSISSNYWSWIRQAPGKGLEWIGHVSYSGSTNYNPSLKSRVTISVDTSKNQFSLKLSSVTAADTAVYYCARESYYYYGMDVWGQGTTVTVSSAAAIEVMYPPPYLDNEKSNGTIIHVKGKHLCPSPLFPGPSKPFWVLVVVGGVLACYSLLVTVAFIIFWVRSKRSRLLHSDYMNMTPRRPGPTRKHYQPYAPPRDFAAYRSRVKFSRSADAPAYQQGQNQLYNELNLGRREEYDVLDKRRGRDPEMGGKPRRKNPQEGLYNELQKDKMAEAYSEIGMKGERRRGKGHDGLYQGLSTATKDTYDALHMQALPPR
5'-GAAATTGTGTTGACGCAGTCTCCAGCCACCCTGTCTGTGTCTCCAGGGGAAAGAGCCACCCTCTCCTGCAGGGCCAGTCAGAGTGTTAGCAGCAACTTAGCCTGGTACCAGCAGAAACCTGGCCAGGCTCCCAGGCTCCTCATCTATGGTGCATCCACCAGGGCCACTGGTATCCCAGCCAGGTTCAGTGGCAGTGGGTCTGGGACAGAGTTCACTCTCACCATCAGCAGACTGGAGCCTGAAGATTTTGCAGTGTATTACTGTCAGCAGTATAATAACTGGCCTCCGAAATTCACTTTCGGCCCTGGGACCAAAGTGGATATCAGAGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCCAGGTGCAGCTGCAGGAGTCGGGCCCAGGACTGGTGAAGCCTTCGGATACCCTGTCCCTCACCTGTCTTGTCTCTGGTGGCTCCATCAGTAGTAATTACTGGAGCTGGATCCGGCAGGCCCCAGGGAAGGGACTGGAGTGGATTGGACATGTCTCCTACAGTGGGAGCACCAACTACAACCCCTCCCTCAAGAGTCGAGTTACCATATCAGTAGACACGTCTAAGAACCAGTTCTCCCTGAAGCTGAGCTCTGTGACTGCCGCGGACACGGCCGTGTATTACTGTGCGAGAGAGTCCTACTACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGGTCACCGTCTCCTCAGCGGCCGCAATCGAAGTGATGTACCCCCCTCCCTACCTGGACAACGAGAAGTCCAACGGCACCATTATCCACGTGAAGGGAAAGCACCTGTGCCCCAGCCCTCTGTTCCCTGGCCCTAGCAAGCCTTTCTGGGTGCTGGTGGTCGTGGGCGGAGTGCTGGCCTGTTATAGCCTGCTCGTGACCGTGGCCTTCATCATCTTTTGGGTGCGCAGCAAGCGGAGCCGGCTGCTGCACAGCGACTACATGAACATGACCCCCAGACGGCCCGGACCCACCAGAAAGCACTACCAGCCTTACGCCCCTCCCAGAGACTTCGCCGCCTACAGATCTCGAGTGAAGTTCAGCAGAAGCGCCGACGCCCCTGCCTATCAGCAGGGCCAGAACCAGCTGTACAACGAGCTGAACCTGGGCAGACGGGAAGAGTACGACGTGCTGGACAAGCGGAGAGGCAGGGACCCTGAGATGGGCGGCAAGCCCAGAAGAAAGAACCCCCAGGAAGGCCTGTATAACGAACTGCAGAAAGACAAGATGGCCGAGGCCTACAGCGAGATCGGCATGAAGGGCGAGCGGAGAAGAGGCAAGGGCCACGATGGCCTGTACCAGGGCCTGAGCACCGCCACCAAGGACACCTATGACGCCCTGCACATGCAGGCCCTGCCCCCCAGA-3'
本発明のCARライブラリを用いた本発明のスクリーニング方法により、CD19に結合するscFvをスクリーニングできることを確認した。
ヒトCD19特異的抗体であるFMC63の重鎖可変領域および軽鎖可変領域に代えて、ヒトCD19特異的抗体(clone18(国際公開第2016/033570号参照))の重鎖可変領域および軽鎖可変領域を用いた以外は、前記実施例3(1)と同様にして、clone18 scFvまたはclone18 scFvをコードする核酸を調製した。各核酸は、pMX発現ベクターに導入した。なお、各scFvにおける重鎖可変領域および軽鎖可変領域のアミノ酸配列を下線で示す。
QSALTQPRSVSGFPGQSVTISCTGTTSDDVSWYQQHPGKAPQLMLYDVSKRPSGVPHRFSGSRSGRAASLIISGLQTEDEADYFCCSYAGRYNSVLFGGGTKLTVLGSTSGSGKPGSGEGSTKGEVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSGISWNSGRIGYADSVKGRFTISRDNAKNSLFLQMNSLRAEDTAVYYCARDQGYHYYDSAEHAFDIWGQGTVVTVSS
5'-CAATCTGCTCTGACACAGCCTAGAAGCGTGTCCGGCTTTCCTGGCCAGAGCGTGACCATCAGCTGTACCGGCACCACCTCCGATGACGTGTCCTGGTATCAGCAGCATCCTGGCAAAGCCCCTCAGCTGATGCTGTACGACGTGTCCAAAAGACCTAGCGGCGTGCCCCACAGATTCAGCGGATCTAGAAGTGGCAGAGCCGCCAGCCTGATCATCTCTGGACTGCAGACAGAGGACGAGGCCGACTACTTCTGCTGTAGCTACGCCGGCAGATACAACAGCGTGCTGTTTGGCGGCGGAACAAAGCTGACAGTGCTGGGCTCTACAAGCGGCTCTGGCAAGCCTGGATCTGGCGAGGGAAGCACCAAGGGCGAAGTGCAGCTTGTGGAATCTGGCGGAGGACTGGTGCAGCCTGGAAGAAGCCTGAGACTGTCTTGTGCCGCCAGCGGCTTCACCTTCGACGATTATGCCATGCACTGGGTCCGACAGGCCCCTGGAAAAGGCCTTGAATGGGTGTCCGGCATCTCTTGGAACAGCGGCAGAATCGGCTACGCCGACTCTGTGAAGGGCAGATTCACCATCAGCCGGGACAACGCCAAGAACAGCCTGTTCCTGCAGATGAACTCCCTGAGAGCCGAGGACACCGCCGTGTACTACTGTGCCAGAGATCAGGGCTACCACTACTACGACTCTGCCGAGCACGCCTTCGATATCTGGGGCCAGGGAACAGTGGTCACCGTTAGTTCT-3'
前記実施例10(1)のclone18LH scFvをコードする核酸の5’側の軽鎖可変領域をコードする核酸を、前記ヒト末梢血B細胞由来の軽鎖可変領域の核酸を含むライブラリで置換することにより、hK-clone18H scFvライブラリおよびhL-clone18H scFvライブラリを調製した。そして、3M4E5LH scFvをコードする核酸に代えて、hK-clone18H scFvライブラリおよびhL-clone18H scFvライブラリを用いた以外は、前記実施例1(2)と同様にして、hK-clone18H scFvライブラリおよびhL-clone18H scFvライブラリを調製した。つぎに、hK-clone18H scFvライブラリおよびhL-clone18H scFvライブラリを、実施例1(1)のヒトCD28の部分領域、膜貫通ドメインおよび細胞内シグナル伝達ドメインをコードする核酸と、ヒトCD3ζの細胞内シグナル伝達ドメインをコードする核酸とが、各ライブラリの3’端側にこの順番で配置されるようにpMX発現ベクターに導入し、hK-clone18H CARライブラリおよびhL-clone18H CARライブラリの発現ベクターを調製した。なお、hK-clone18H CARライブラリおよびhL-clone18H CARの軽鎖可変領域の不均一性は、制限酵素マッピングおよび各領域をサンガー法によりシークエンスすることで、約1×106種類であることを確認した。
前記hH-3M4E4L CARライブラリの発現ベクターに代えて、hK-clone18H CARライブラリおよびhL-clone18H CARライブラリの発現ベクターを用いた以外は、前記実施例1(3)と同様にして、hK-clone18H CARライブラリおよびhL-clone18H CARライブラリが導入されたT細胞を精製し、これを第1の候補CAR-Tとした。
前記実施例3(4)と同様にして、レトロウイルスを用いて、ヒトCD19をコードする核酸を、K562細胞に導入し、抗原提示細胞(APC-CD19)とした。
前記実施例10(3)の第1の候補CAR-Tと、前記実施例10(4)で調製したAPC-CD19とを用いた以外は、前記実施例3(5)と同様にして、ヒトCD19と結合するhK-clone18H CARまたはhL-clone18H CARを発現する第1の候補CAR-Tを富化した。
hK-clone18H CARライブラリおよびhL-clone18H CARライブラリの発現ベクターに代えて、各CARをコードするポリヌクレオチドを含む発現ベクターを用いた以外は、前記実施例10(3)と同様にして、GaLV-pseudotypedレトロウイルスを含む培養上清を調製した。つぎに、前記GaLV-pseudotypedレトロウイルスを含む培養上清を添加することにより、Jurkat76細胞に各CARをコードするポリヌクレオチドを含む発現ベクターを導入し、scFvとしてL4-18H、L7-18H、L9-18H、L13-18H、L14-18H、L16-18H、L17-18H、L22-18H、K4-18H、K5-18H、K6-18H、K9-18H、またはK10-18Hを含む各CAR(L4-18H CAR、L7-18H CAR、L9-18H CAR、L13-18H CAR、L14-18H CAR、L16-18H CAR、L17-18H CAR、L22-18H CAR、K4-18H CAR、K5-18H CAR、K6-18H CAR、K9-18H CAR、またはK10-18H CAR)を発現させた。前記CAR発現後のJurkat76細胞を、前記PE標識anti-human NGFR mAbと反応させた。前記反応後、前記抗PEマイクロビーズを用いて、各CARを発現するJurkat76細胞(L4-18H CAR-T、L7-18H CAR-T、L9-18H CAR-T、L13-18H CAR-T、L14-18H CAR-T、L16-18H CAR-T、L17-18H CAR-T、L22-18H CAR-T、K4-18H CAR-T、K5-18H CAR-T、K6-18H CAR-T、K9-18H CAR-T、またはK10-18H CAR-T)を精製した。各CARを発現するJurkat76細胞について、前記実施例10(5)と同様にして染色した。前記染色後の各CARを発現するJurkat76細胞について、フローサイトメトリーにより解析した。ポジティブコントロールは、各CARをコードするポリヌクレオチドを含む発現ベクターに代えて、タグ付加clone18LH CARまたはclone18HL CARをコードする核酸を含む発現ベクターを用いた以外は、同様に実施した。また、コントロール1は、CD19ダイマーで染色しなかった以外は同様にして実施した。コントロール2は、各CARをコードするポリヌクレオチドを含む発現ベクターを導入しなかった以外は、同様にして実施した。FSC+SSC+NGFR+細胞にゲートをかけた結果を図28および29に示す。
つぎに、前記実施例10(6)で得られた各CAR-Tが、前記標的抗原との結合により、活性化するかを検討した。具体的には、2×105細胞の各CAR-Tと、5×104細胞の抗原提示細胞(APC-CD19またはRaji細胞)とを各ウェル(96ウェルプレート)に播種後、5時間共培養した。前記抗原提示細胞(APC-CD19)は、前記実施例10(4)と同様にして調製した。また、前記共培養の培養液は、10%FCS含有RPMI1640培地とした。前記共培養後、各CAR-Tを回収し、FITC標識anti-human CD69 mAb(clone:FN50)およびV450標識anti-human NGFR mAbで染色した。前記染色後の各CAR-Tについて、前記フローサイトメーターを用いて、NGFR+細胞について、CD69の平均蛍光強度を測定した。ポジティブコントロールは、前記実施例10(6)で得られたclone18LH CAR-Tおよびclone18HL CAR-Tを用いた以外は、同様に実施した。また、コントロール1は、CD19をコードする核酸を導入していないK562細胞を用いた以外は、同様にして実施した。コントロール2は、各CARをコードするポリヌクレオチドを含む発現ベクターを導入しなかった以外は、同様にして実施した。そして、共培養前のCD69の平均蛍光強度を基準として、各サンプルのCD69の発現上昇の割合を算出した。これらの結果を図30に示す。
つぎに、前記実施例10(6)で得られたL4-18H CAR-T、L7-18H CAR-T、L16-18H CAR-T、L17-18H CAR-T、K4-18H CAR-T、K5-18H CAR-T、K6-18H CAR-T、およびK9-18H CAR-Tと、clone18LH CAR-Tが、前記標的抗原との親和性の違いを検討した。各CAR-Tについて、所定濃度(1.25、2.5、5、10、20、または40μg/mL)のPE標識CD19ダイマーで染色後、V450標識 anti-human NGFR mAbで染色した以外は、前記実施例10(6)と同様にして、フローサイトメトリーにより解析した。そして、CD19ダイマーの各濃度における、CD19ダイマー陽性細胞の割合を算出後、40μg/mLのCD19ダイマーで染色した際のCD19ダイマー陽性細胞の割合を基準として、各CD19ダイマーで染色した際のCD19ダイマー陽性細胞の相対割合を算出した。この結果を、図31に示す。
つぎに、前記(6)で得られたL4-18H CAR-T、L7-18H CAR-T、L16-18H CAR-T、L17-18H CAR-T、K4-18H CAR-T、K5-18H CAR-T、K6-18H CAR-T、およびK9-18H CAR-Tと、clone18LH CAR-Tが、前記標的抗原を発現する細胞に対する応答性の違いを検討した。具体的には、2×105細胞の各CAR-Tと、5×104細胞の抗原提示細胞(APC-CD19)とK562細胞との混合細胞とを各ウェル(96ウェルプレート)に播種後、5時間共培養した。前記混合細胞におけるAPC-CD19(A)とK562(K)との比率(A:K)は、所定比率(0:100、6.25:93.75、12.5:87.5、50:50、100:0)とした。前記共培養後、各CAR-Tを回収し、FITC標識anti-human CD69 mAb(clone:FN50)およびV450標識anti-human NGFR mAbで染色した。前記染色後の各CAR-Tについて、前記フローサイトメーターを用いて、NGFR+細胞について、CD69陽性細胞の割合を測定した。比率(A:K)=100:0におけるCD69陽性細胞の割合を基準として、各比率(A:K)で培養した際のCD69陽性細胞の相対割合(% Maximal responses)を算出した。また、これらの結果を図32に示す。
つぎに、前記(6)で得られたL4-18H CAR-T、L7-18H CAR-T、L16-18H CAR-T、L17-18H CAR-T、K4-18H CAR-T、K5-18H CAR-T、K6-18H CAR-T、およびK9-18H CAR-Tと、clone18LH CAR-Tが、前記標的抗原との結合により、サイトカイン産生するかを検討した。具体的には、3×105細胞の各CAR-Tと、5×104細胞の抗原提示細胞(APC-CD19またはRaji細胞)とを各ウェル(96ウェルプレート)に播種後、5時間共培養した。前記抗原提示細胞(APC-CD19)は、前記実施例10(4)と同様にして調製した。また、前記共培養の培養液は、10%FCS含有RPMI1640培地とした。前記培養後、5μmol/LとなるようにBFAを添加し、さらに5時間培養した。そして、培養後の細胞を、1%PFAで固定後、細胞膜の透過処理を行った。さらに、APC標識anti-human IL-2 mAb(clone:MQ1-17H12)、PE標識anti-human TNFα mAb(clone:Mab11)、PC7標識anti-human IFN-γ mAb(clone:B27)、FITC標識anti-human CD4 mAb (clone:OKT4)、PC5標識anti-human CD8a mAb (clone:B9.11)、およびV450標識anti-human NGFR mAb(clone:C40-1457)で染色した。前記染色後の細胞について、前記フローサイトメーターで測定した。そして、サイトカイン発現の割合を算出した。ポジティブコントロールは、前記実施例10(6)で得られたclone18LH CAR-Tを用いた以外は、同様に実施した。コントロール1は、CD19をコードする核酸を導入していないK562細胞を用いた以外は、同様にして実施した。コントロール2は、各CARをコードするポリヌクレオチドを含む発現ベクターを導入しなかった以外は、同様にして実施した。これらの結果を図33に示す。
本発明のCARライブラリを用いた本発明のスクリーニング方法により得られたscFvを含むCARが、in vivoで抗腫瘍効果を発揮することを確認した。
MEEENIVNGDRPRDLVFPGTAGLQLYQSLYKYSYITDGIIDAHTNEVISYAQIFETSCRLAVSLEKYGLDHNNVVAICSENNIHFFGPLIAALYQGIPMATSNDMYTEREMIGHLNISKPCLMFCSKKSLPFILKVQKHLDFLKKVIVIDSMYDINGVECVFSFVSRYTDHAFDPVKFNPKEFDPLERTALIMTSSGTTGLPKGVVISHRSITIRFVHSSDPIYGTRIAPDTSILAIAPFHHAFGLFTALAYFPVGLKIVMVKKFEGEFFLKTIQNYKIASIVVPPPIMVYLAKSPLVDEYNLSSLTEIACGGSPLGRDIADKVAKRLKVHGILQGYGLTETCSALILSPNDRELKKGAIGTPMPYVQVKVIDINTGKALGPREKGEICFKSQMLMKGYHNNPQATRDALDKDGWLHTGDLGYYDEDRFIYVVDRLKELIKYKGYQVAPAELENLLLQHPNISDAGVIGIPDEFAGQLPSACVVLEPGKTMTEKEVQDYIAELVTTTKHLRGGVVFIDSIPKGPTGKLMRNELRAIFAREQAKSKLRAKRSGSGATNFSLLKQAGDVEENPGPMDGPRLLLLLLLGVSLGGAKEACPTGLYTHSGECCKACNLGEGVAQPCGANQTVCEPCLDSVTFSDVVSATEPCKPCTECVGLQSMSAPCVEADDAVCRCAYGYYQDETTGRCEACRVCEAGSGLVFSCQDKQNTVCEECPDGTYSDEANHVDPCLPCTVCEDTERQLRECTRWADAECEEIPGRWITRSTPPEGSDSTAPSTQEPEAPPEQDLIASTVAGVVTTVMGSSQPVVTRGTTDNLIPVYCSILAAVVVGLVAYIAFKRWNS
5'-ATGGAAGAAGAGAACATCGTGAATGGCGATCGCCCTCGGGATCTGGTGTTCCCTGGCACAGCCGGCCTGCAGCTGTATCAGTCCCTGTATAAATACTCTTACATCACCGACGGAATCATCGACGCCCACACCAACGAGGTGATCTCCTATGCCCAGATTTTCGAAACAAGTTGCCGCCTGGCCGTGAGCCTGGAGAAGTATGGCCTGGATCACAACAACGTGGTGGCCATTTGCAGCGAGAACAACATCCACTTCTTCGGCCCTCTGATCGCTGCCCTATACCAGGGGATTCCAATGGCCACATCCAACGATATGTACACCGAGAGGGAGATGATCGGCCACCTGAACATCTCCAAGCCATGTCTGATGTTCTGTTCCAAGAAGTCCCTGCCATTCATCCTGAAGGTGCAGAAGCACCTGGACTTTCTCAAGAAGGTGATCGTGATCGACAGCATGTACGACATCAACGGCGTGGAGTGCGTGTTCAGTTTCGTGTCCCGGTACACCGATCATGCGTTCGATCCAGTGAAGTTCAACCCTAAAGAGTTTGATCCCCTGGAGAGAACCGCGCTGATCATGACATCCTCTGGAACAACCGGCCTGCCTAAGGGCGTGGTGATCAGCCACAGGAGCATCACCATCAGATTCGTCCACAGCAGCGATCCCATCTACGGCACCCGCATCGCCCCAGATACATCCATCCTGGCCATCGCCCCTTTCCACCACGCCTTCGGACTGTTTACCGCCCTGGCTTACTTTCCAGTGGGCCTGAAGATCGTGATGGTGAAAAAGTTTGAGGGCGAGTTCTTCCTGAAGACCATCCAGAACTACAAGATCGCTTCTATCGTGGTGCCTCCTCCAATCATGGTGTATCTGGCCAAGAGCCCTCTGGTGGATGAGTACAATCTGTCCAGCCTGACAGAGATCGCCTGTGGCGGCTCCCCTCTGGGCAGAGACATCGCCGACAAGGTGGCCAAGAGACTGAAGGTCCACGGCATCCTGCAGGGCTATGGCCTGACCGAGACCTGTAGCGCCCTGATCCTGAGCCCCAACGATAGAGAGCTGAAGAAGGGCGCCATCGGCACCCCTATGCCCTATGTCCAGGTGAAGGTGATTGACATCAACACCGGCAAAGCCCTGGGACCAAGAGAGAAGGGCGAGATTTGCTTCAAGAGCCAGATGCTGATGAAGGGCTACCACAACAACCCACAGGCCACCAGGGATGCCCTGGACAAGGACGGGTGGCTGCACACCGGCGATCTGGGCTACTACGACGAGGACAGATTCATCTATGTGGTGGATCGGCTGAAAGAACTCATCAAGTACAAGGGCTACCAGGTGGCCCCTGCCGAGCTGGAGAACTTGCTTCTGCAGCACCCTAACATCTCTGATGCCGGCGTCATCGGCATCCCAGACGAGTTTGCCGGCCAGCTGCCTTCCGCCTGTGTCGTGCTGGAGCCTGGCAAGACCATGACCGAGAAGGAGGTGCAGGATTATATCGCCGAGCTGGTGACCACCACCAAGCACCTGCGGGGCGGCGTGGTGTTCATCGACAGCATTCCGAAAGGCCCAACAGGCAAGCTGATGAGAAACGAGCTGAGGGCCATCTTTGCCCGCGAGCAGGCCAAGTCCAAGCTGAGGGCCAAGCGGTCCGGATCCGGAGCCACCAACTTCAGCCTGCTGAAGCAGGCCGGCGACGTGGAGGAGAACCCCGGCCCCATGGACGGGCCGCGCCTGCTGCTGTTGCTGCTTCTGGGGGTGTCCCTTGGAGGTGCCAAGGAGGCATGCCCCACAGGCCTGTACACACACAGCGGTGAGTGCTGCAAAGCCTGCAACCTGGGCGAGGGTGTGGCCCAGCCTTGTGGAGCCAACCAGACCGTGTGTGAGCCCTGCCTGGACAGCGTGACGTTCTCCGACGTGGTGAGCGCGACCGAGCCGTGCAAGCCGTGCACCGAGTGCGTGGGGCTCCAGAGCATGTCGGCGCCATGCGTGGAGGCCGACGACGCCGTGTGCCGCTGCGCCTACGGCTACTACCAGGATGAGACGACTGGGCGCTGCGAGGCGTGCCGCGTGTGCGAGGCGGGCTCGGGCCTCGTGTTCTCCTGCCAGGACAAGCAGAACACCGTGTGCGAGGAGTGCCCCGACGGCACGTATTCCGACGAGGCCAACCACGTGGACCCGTGCCTGCCCTGCACCGTGTGCGAGGACACCGAGCGCCAGCTCCGCGAGTGCACACGCTGGGCCGACGCCGAGTGCGAGGAGATCCCTGGCCGTTGGATTACACGGTCCACACCCCCAGAGGGCTCGGACAGCACAGCCCCCAGCACCCAGGAGCCTGAGGCACCTCCAGAACAAGACCTCATAGCCAGCACGGTGGCAGGTGTGGTGACCACAGTGATGGGCAGCTCCCAGCCCGTGGTGACCCGAGGCACCACCGACAACCTCATCCCTGTCTATTGCTCCATCCTGGCTGCTGTGGTTGTGGGTCTTGTGGCCTACATAGCCTTCAAGAGGTGGAACAGCTGA-3'
本発明のCARライブラリを用いた本発明のスクリーニング方法の第1の選抜工程および第2の選抜工程において、間接的な評価方法に基づく選抜、すなわち、標的抗原とCARとの結合により生じる候補免疫細胞の活性化指標の評価に基づく選抜により、CA-T細胞を機能的に活性化しうるscFvを含むCARをスクリーニング出来ることを確認した。
本発明のCARライブラリを用いた本発明のスクリーニング方法により得られたscFvを含むCARが、in vitroで細胞傷害活性を発揮することを確認した。
K=(LE-LS)/(LMax-LS)×100(%) ・・・(1)
K:細胞傷害性
LE:CPMの測定値
LS:ターゲット細胞のみの場合のCPMの測定値
LMax:ターゲット細胞のみのウェルに0.2%Triton-X含有PBSを添加した場合のCPMの測定値
本発明のCARライブラリを用いた本発明のスクリーニング方法により得られたscFvを含むCARが、in vivoで抗腫瘍効果を発揮することを確認した。
本発明のスクリーニング方法により、in vivoでの抗腫瘍効果に優れるscFvを取得できるメカニズムを検討した。
前記実施例10と同様にして、L17-18H CAR-Tと、clone18LH CAR-Tとを調製した。つぎに、20Gyでγ線照射したAPC-CD19(K562/CD19)またはRaji細胞(Raji)(T)と、L17-18H CAR-Tまたはclone18LH CAR-T(E)とについて、E:Tが、1:1または10:1となるように、前記CAR-T細胞を培養しているウェルに、標的細胞を添加した。前記標的細胞添加時を基準として、5日間培養した。前記培養では、10U/mLのIL-2および10ng/mLのIL-15を培地に添加した。さらに、培養5日目において、培養開始時と同じ条件で、各CAR-T細胞を再刺激した。そして、培養開始時から所定時(0、3、5、7または10日)における細胞数をカウントし、培養開始時の細胞数を1とした際の細胞数比を増殖比として算出した。これらの結果を図39に示す。
前記実施例15(1)の培養10日目のL17-18H CAR-Tおよびclone18LH CAR-Tのサイトカイン産生能について検討した。具体的には、3.0×105細胞のCAR-T細胞と、5.0×104個の標的細胞(K562細胞、Raji細胞、APC-CD19(K562/CD19)とを共培養した。2時間培養後、5μg/mLとなるように、Brefeldin A(BFA)を添加し、さらに、14~18時間培養した。そして、CAR-T細胞を回収後、1%パラホルムアルデヒド(PFA)で固定後、細胞膜の透過処理を行い、PE標識抗ヒトTNF-α抗体(clone MAb11)、APC標識抗ヒトIL-2抗体(clone MQ1-17H12)およびPC7標識抗ヒトIFN-γ抗体(clone B27)と、PC5標識抗ヒトCD8抗体(clone B9.11)、FITC標識抗CD4抗体(clone OKT4)およびV450標識anti-human NGFR(clone:C40-1457)で染色した。前記染色後のCAR-T細胞について、前記フローサイトメーターで測定し、各サイトカインを産生している細胞の割合を算出した。これらの結果を図40に示す。
前記実施例15(1)の培養10日目のL17-18H CAR-Tおよびclone18LH CAR-Tの細胞傷害活性について検討した。つぎに、5.0×103個の標的細胞であるAPC-CD19(K562/CD19)またはRaji細胞(Raji)およびK562細胞について、クロム51(51Cr)存在下で、1.5時間培養することにより、クロム51でラベルした。
4人の異なるドナー(ヒト)由来のT細胞を用いた以外は、前記実施例10と同様にして、L17-18H CAR-Tおよびclone18LH CAR-Tを調製した。得られたCAR-T細胞について、0.5μmol/L CFSEを含むPBS内で、15分間37℃でインキュベートし、CAR-T細胞をCFSEで標識した。つぎに、20Gyでγ線照射したRaji細胞(Raji)(T)と、L17-18H CAR-Tまたはclone18LH CAR-T(E)とについて、E:Tが、1:1または5:1となるように、それぞれをウェルに添加した。前記標的細胞添加時を基準として、3日間培養した。
Proliferation(%)=(control MFI-sample MFI)/control MFI×100 ・・・(2)
control MFI:K562細胞株とE:T=1:1の条件で共培養したCAR-T細胞のMFI
sample MFI:各CAR-T細胞のMFI
上記の実施形態および実施例の一部または全部は、以下の付記のように記載されうるが、以下には限られない。
(付記1)
第1のキメラ抗原受容体(CAR)をコードする核酸を含み、
前記第1のCARは、第1の抗原結合ドメインと、第1の膜貫通ドメインと、第1の細胞内シグナル伝達ドメインとを含み、
前記第1の抗原結合ドメインは、標的抗原への結合をスクリーニングする第1の一本鎖抗体(scFv)を含み、
前記第1のscFvは、第1の重鎖可変領域と、第1の軽鎖可変領域とを含み、
前記第1の重鎖可変領域および前記第1の軽鎖可変領域は、下記条件1または条件2を満たす、CARライブラリ;
条件1:
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、第1のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む;
条件2:
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、第1のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。
(付記2)
前記第1の細胞内シグナル伝達ドメインは、CD3ζの細胞内シグナル伝達ドメインを含む、付記1記載のCARライブラリ。
(付記3)
前記第1の膜貫通ドメインは、CD28の膜貫通ドメインを含む、付記1または2記載のCARライブラリ。
(付記4)
前記第1のB細胞受容体は、ヒト由来B細胞のB細胞受容体である、付記1から3のいずれかに記載のCARライブラリ。
(付記5)
付記1から4のいずれかに記載のCARライブラリを免疫細胞に発現させる第1の発現工程と、
前記第1の発現工程で得られた免疫細胞と、前記標的抗原とを接触させる第1の接触工程と、
前記第1の接触工程において前記標的抗原と結合した免疫細胞に発現するCARの第1のscFvを、前記標的抗原に結合する第1の候補scFvとして選抜する第1の選抜工程とを含む、scFvの製造方法(スクリーニング方法)。
(付記6)
前記第1の候補scFvに基づき、第2のCARライブラリを調製する調製工程を含み、
前記第2のCARライブラリは、第2のCARをコードする核酸を含み、
前記第2のCARは、第2の抗原結合ドメインと、第2の膜貫通ドメインと、第2の細胞内シグナル伝達ドメインとを含み、
前記第2の抗原結合ドメインは、前記標的抗原への結合をスクリーニングする第2のscFvを含み、
前記第2のscFvは、第2の重鎖可変領域と、第2の軽鎖可変領域とを含み、
前記第2の重鎖可変領域および前記第2の軽鎖可変領域は、下記条件3または条件4を満たし、
さらに、前記第2のCARライブラリを免疫細胞に発現させる第2の発現工程と、
前記第2の発現工程で得られた免疫細胞と、前記標的抗原とを接触させる第2の接触工程と、
前記第2の接触工程において前記標的抗原と結合した免疫細胞に発現するCARの第2のscFvを、前記標的抗原に結合する第2の候補scFvとして第2の選抜工程とを含む、付記5記載の製造方法(スクリーニング方法);
条件3:
前記第1の発現工程におけるCARライブラリが前記条件1を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、第2のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、前記第1の候補scFvにおける軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む;
条件4:
前記第1の発現工程におけるCARライブラリが前記条件2を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、前記第1の候補scFvにおける重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、第2のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む。
(付記7)
前記第2の接触工程を複数回実施する、付記6記載のscFvの製造方法(スクリーニング方法)。
(付記8)
前記第2の選抜工程において、前記標的抗原のモノマーまたはマルチマーを用いて、前記標的抗原に結合する免疫細胞を検出し、
検出された免疫細胞に発現するscFvを、前記第2の候補scFvとして選抜する、付記6または7記載のscFvの製造方法(スクリーニング方法)。
(付記9)
前記第2のB細胞受容体は、ヒト由来B細胞のB細胞受容体である、付記6から8のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記10)
前記第1の接触工程を複数回実施する、付記5から9のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記11)
前記第1の選抜工程において、前記標的抗原のモノマーまたはマルチマーを用いて、前記標的抗原に結合する免疫細胞を検出し、
検出された免疫細胞に発現するscFvを、前記第1の候補scFvとして選抜する、付記5から10のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記12)
前記免疫細胞は、T細胞である、付記5から11のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記13)
下記(H)の重鎖可変領域と、下記(L)の軽鎖可変領域とを含む、HLA-A*02:01およびNY-ESO-1157-165の複合体に対する抗体またはその抗原結合断片。
(H)重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3を含み、
CDRH1が、下記(H1)のアミノ酸配列を含むポリペプチドであり、
CDRH2が、下記(H2)のアミノ酸配列を含むポリペプチドであり、
CDRH3が、下記(H3)のアミノ酸配列を含むポリペプチドである重鎖可変領域
(H1)下記(H1-1)、(H1-2)または(H1-3)のアミノ酸配列
(H1-1)前記表1AのCDRH1のいずれかのアミノ酸配列
(H1-2)(H1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-3)(H1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H2)下記(H2-1)、(H2-2)または(H2-3)のアミノ酸配列
(H2-1)前記表1AのCDRH2のいずれかのアミノ酸配列
(H2-2)(H2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-3)(H2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H3)下記(H3-1)、(H3-2)または(H3-3)のアミノ酸配列
(H3-1)前記表1AのCDRH3のいずれかのアミノ酸配列
(H3-2)(H3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-3)(H3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L)軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3を含み、
CDRL1が、下記(L1)のアミノ酸配列を含むポリペプチドであり、
CDRL2が、下記(L2)のアミノ酸配列を含むポリペプチドであり、
CDRL3が、下記(L3)のアミノ酸配列を含むポリペプチドである軽鎖可変領域
(L1)下記(L1-1)、(L1-2)または(L1-3)のアミノ酸配列
(L1-1)前記表1BのCDRL1のいずれかのアミノ酸配列
(L1-2)(L1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-3)(L1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2)下記(L2-1)、(L2-2)または(L2-3)のアミノ酸配列
(L2-1)前記表1BのCDRL2のいずれかのアミノ酸配列
(L2-2)(L2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-3)(L2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3)下記(L3-1)、(L3-2)または(L3-3)のアミノ酸配列
(L3-1)前記表1BのCDRL3のいずれかのアミノ酸配列
(L3-2)(L3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-3)(L3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(付記14)
前記重鎖可変領域および前記軽鎖可変領域の組合せが、前記表2のいずれかの組合せである、付記13記載の抗体またはその抗原結合断片。
(付記15)
抗原結合ドメインと、膜貫通ドメインと、細胞内シグナル伝達ドメインとを含み、
前記抗原結合ドメインは、付記13または14記載の抗体またはその抗原結合断片を含む、キメラ抗原受容体。
(付記16)
前記抗原結合断片は、一本鎖抗体である、付記15記載のキメラ抗原受容体。
(付記17)
付記15または16記載のキメラ抗原受容体をコードする、核酸。
(付記18)
付記15または16記載のキメラ抗原受容体を含む、細胞。
(付記19)
前記細胞は、T細胞を含む、付記18記載の細胞。
(付記20)
付記17記載の核酸を細胞に導入する導入工程を含む、細胞の製造方法。
(付記21)
前記細胞は、T細胞を含む、付記20記載の細胞の製造方法。
(付記22)
第1の二重特異性抗体(BsAb)をコードする核酸を含み、
前記第1のBsAbは、第1の抗原結合ドメインと、第2の抗原結合ドメインとを含み、
前記第1の抗原結合ドメインは、第1の標的抗原に結合する第1の一本鎖抗体(scFv)を含み、
前記第2の抗原結合ドメインは、第2の標的抗原への結合をスクリーニングする第2のscFvを含み、
前記第2のscFvは、第2の重鎖可変領域と、第2の軽鎖可変領域とを含み、
前記第2の重鎖可変領域および前記第2の軽鎖可変領域は、下記条件1または条件2を満たし、
前記第1の標的抗原または前記第2の標的抗原は、免疫細胞活性化受容体である、BsAbライブラリ;
条件1:
前記第2の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、前記第2の標的抗原に結合する抗体もしくはその抗原結合断片における重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、第1のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む;
条件2:
前記第2の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、第1のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、前記第2の標的抗原に結合する抗体もしくはその抗原結合断片における軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。
(付記23)
前記第1のB細胞受容体は、ヒト由来B細胞のB細胞受容体である、付記22記載のBsAbライブラリ。
(付記24)
前記免疫細胞活性化受容体は、CD3である、付記22または23記載のBsAbライブラリ。
(付記25)
付記22から24のいずれかに記載のBsAbライブラリから第1のBsAbを準備する第1の準備工程と、
前記第1の標的抗原が免疫細胞活性化受容体の場合、
前記第1のBsAbと、前記第2の標的抗原と、前記免疫細胞活性化受容体を発現可能な免疫細胞とを接触させ、
前記第2の標的抗原が免疫細胞活性化受容体の場合、
前記第1のBsAbと、前記第1の標的抗原と、前記免疫細胞活性化受容体を発現可能な免疫細胞とを接触させる第1の接触工程と、
前記第1の接触工程において、前記第2の標的抗原と結合した第1のBsAbの第2のscFvを、前記第2の標的抗原に結合する第1の候補scFvとして選抜する第1の選抜工程とを含む、scFvの製造方法(スクリーニング方法)。
(付記26)
前記第1の候補scFvに基づき、第2のBsAbライブラリを調製する調製工程を含み、
前記第2のBsAbライブラリは、前記第2のBsAbをコードする核酸を含み、
前記第2のBsAbは、第3の抗原結合ドメインと、第4の抗原結合ドメインとを含み、
前記第3の抗原結合ドメインは、前記第1の標的抗原に結合する第3のscFvを含み、
前記第4の抗原結合ドメインは、前記第2の標的抗原への結合をスクリーニングする第4のscFvを含み、
前記第4のscFvは、第4の重鎖可変領域と、第4の軽鎖可変領域とを含み、
前記第4の重鎖可変領域および前記第4の軽鎖可変領域は、下記条件3または条件4を満たし、
さらに、前記第2のBsAbライブラリから第2のBsAbを準備する第2の準備工程と、
前記第1の標的抗原が前記免疫細胞活性化受容体の場合、
前記第2のBsAbと、前記第2の標的抗原と、前記免疫細胞活性化受容体を発現可能な免疫細胞とを接触させ、
前記第2の標的抗原が前記免疫細胞活性化受容体の場合、
前記第2のBsAbと、前記第1の標的抗原と、前記免疫細胞活性化受容体を発現可能な免疫細胞とを接触させる第2の接触工程と、
前記第2の接触工程において、前記第2の標的抗原と結合した第2のBsAbの第4のscFvを、前記第2の標的抗原に結合する第2の候補scFvとして選抜する第2の選抜工程とを含む、付記25記載のscFv製造方法(スクリーニング方法);
条件3:
前記第1の準備工程におけるBsAbライブラリが前記条件1を満たす場合、
前記第4の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、第2のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第4の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、前記第1の候補BsAbにおける軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む;
条件4:
前記第1の準備工程におけるBsAbライブラリが前記条件2を満たす場合、
前記第4の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、前記第1の候補BsAbにおける重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第4の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、第2のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む。
(付記27)
前記第2の接触工程および前記第2の選抜工程を1セットとし、前記セットを複数回実施する、付記26記載のscFvの製造方法(スクリーニング方法)。
(付記28)
前記第2の選抜工程において、前記免疫細胞の活性化分子の発現を検出し、
前記免疫細胞の活性化分子を誘導した第2のBsAbを、前記第2の候補BsAbとして選抜する、付記26または27記載のscFvの製造方法(スクリーニング方法)。
(付記29)
前記第2のB細胞受容体は、ヒト由来B細胞のB細胞受容体である、付記25から28のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記30)
前記第1の接触工程および前記第1の選抜工程を1セットし、前記セットを複数回実施する、付記25から29のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記31)
前記第1の選抜工程において、前記免疫細胞の活性化分子の発現を検出し、
前記免疫細胞の活性化分子を誘導した第1のBsAbを、前記第1の候補BsAbとして選抜する、付記25から30のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記32)
前記免疫細胞活性化受容体は、CD3である、付記25から31のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記33)
前記免疫細胞は、T細胞である、付記25から32のいずれかに記載のscFvの製造方法(スクリーニング方法)。
(付記34)
下記(H)の重鎖可変領域と、下記(L)の軽鎖可変領域とを含む、CD19に対する抗体またはその抗原結合断片。
(H)重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3を含み、
CDRH1が、下記(H1)のアミノ酸配列を含むポリペプチドであり、
CDRH2が、下記(H2)のアミノ酸配列を含むポリペプチドであり、
CDRH3が、下記(H3)のアミノ酸配列を含むポリペプチドである重鎖可変領域
(H1)下記(H1-1)、(H1-2)または(H1-3)のアミノ酸配列
(H1-1)前記表3AのCDRH1のいずれかのアミノ酸配列
(H1-2)(H1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H1-3)(H1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H2)下記(H2-1)、(H2-2)または(H2-3)のアミノ酸配列
(H2-1)前記表3AのCDRH2のいずれかのアミノ酸配列
(H2-2)(H2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H2-3)(H2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(H3)下記(H3-1)、(H3-2)または(H3-3)のアミノ酸配列
(H3-1)前記表3AのCDRH3のいずれかのアミノ酸配列
(H3-2)(H3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(H3-3)(H3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L)軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3を含み、
CDRL1が、下記(L1)のアミノ酸配列を含むポリペプチドであり、
CDRL2が、下記(L2)のアミノ酸配列を含むポリペプチドであり、
CDRL3が、下記(L3)のアミノ酸配列を含むポリペプチドである軽鎖可変領域
(L1)下記(L1-1)、(L1-2)または(L1-3)のアミノ酸配列
(L1-1)前記表3BのCDRL1のいずれかのアミノ酸配列
(L1-2)(L1-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L1-3)(L1-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L2)下記(L2-1)、(L2-2)または(L2-3)のアミノ酸配列
(L2-1)前記表3BのCDRL2のいずれかのアミノ酸配列
(L2-2)(L2-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L2-3)(L2-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(L3)下記(L3-1)、(L3-2)または(L3-3)のアミノ酸配列
(L3-1)前記表3BのCDRL3のいずれかのアミノ酸配列
(L3-2)(L3-1)のアミノ酸配列に対して、80%以上の同一性を有するアミノ酸配列
(L3-3)(L3-1)のアミノ酸配列において、1個または数個のアミノ酸が欠失、置換、挿入および/または付加されたアミノ酸配列
(付記35)
前記重鎖可変領域および前記軽鎖可変領域の組合せが、前記表2のいずれかの組合せである、付記34記載の抗体またはその抗原結合断片。
(付記36)
抗原結合ドメインと、膜貫通ドメインと、細胞内シグナル伝達ドメインとを含み、
前記抗原結合ドメインは、付記34または35記載の抗体またはその抗原結合断片を含む、キメラ抗原受容体。
(付記37)
前記抗原結合断片は、一本鎖抗体である、付記36記載のキメラ抗原受容体。
(付記38)
付記36または37記載のキメラ抗原受容体をコードする、核酸。
(付記39)
付記36または37記載のキメラ抗原受容体を含む、細胞。
(付記40)
前記細胞は、T細胞を含む、付記39記載の細胞。
(付記41)
付記38記載の核酸を細胞に導入する導入工程を含む、細胞の製造方法。
(付記42)
前記細胞は、T細胞を含む、付記41記載の細胞の製造方法。
(付記43)
第1のキメラ抗原受容体(CAR)ライブラリの発現細胞を動物に投与する第1の投与工程と、
前記動物において、標的抗原を発現する組織に集積した前記第1のCARライブラリ発現細胞を、前記標的抗原特異的なCARを発現する細胞として採取する第1の採取工程とを含み、
前記第1のCARライブラリは、第1のCARをコードする核酸を含み、
前記第1のCARは、第1の抗原結合ドメインと、第1の膜貫通ドメインと、第1の細胞内シグナル伝達ドメインとを含み、
前記第1の抗原結合ドメインは、標的抗原への結合をスクリーニングする第1の一本鎖抗体(scFv)を含み、
前記第1のscFvは、第1の重鎖可変領域と、第1の軽鎖可変領域とを含み、
前記第1の重鎖可変領域および前記第1の軽鎖可変領域は、下記条件1または条件2を満たす、scFvの製造方法:
(条件1)
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、第1のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む;
(条件2)
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、第1のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。
(付記44)
前記第1の投与工程において、複数種類の第1のCARライブラリ発現細胞を投与する、付記43記載の製造方法。
(付記45)
前記第1の投与工程後、前記標的抗原を前記動物に投与する第1の刺激工程を含む、付記43または44記載の製造方法。
(付記46)
前記標的抗原特異的なCARを発現する細胞を、前記標的抗原で再刺激する第1の再刺激工程を含む、付記43から45のいずれかに記載の製造方法。
(付記47)
前記第1の投与工程に先立ち、標的抗原を発現する細胞を前記動物に導入し、前記標的抗原を発現する組織を形成させる第1の形成工程を含む、付記43から46のいずれかに記載の製造方法。
(付記48)
発現する組織に集積した前記第1のCARライブラリ発現細胞を、前記標的抗原特異的なCARを発現する細胞として採取する、付記43から47のいずれかに記載の製造方法。
(付記49)
前記第1のB細胞受容体は、ヒト由来B細胞のB細胞受容体である、付記43から48のいずれかに記載の製造方法。
(付記50)
第2のCARライブラリの発現細胞を動物に投与する第2の投与工程と、
前記動物において、標的抗原を発現する組織に集積した前記第2のCARライブラリ発現細胞を、前記標的抗原特異的なCARを発現する細胞として採取する第2の採取工程とを含み、
前記第2のCARライブラリは、第2のCARをコードする核酸を含み、
前記第2のCARは、第2の抗原結合ドメインと、第2の膜貫通ドメインと、第2の細胞内シグナル伝達ドメインとを含み、
前記第2の抗原結合ドメインは、前記標的抗原への結合をスクリーニングする第2のscFvを含み、
前記第2のscFvは、第2の重鎖可変領域と、第2の軽鎖可変領域とを含み、
前記第2の重鎖可変領域および前記第2の軽鎖可変領域は、下記条件3または条件4を満たす、付記43から49のいずれかに記載の製造方法:
(条件3)
前記第1のCARライブラリが前記条件1を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、第2のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、前記標的抗原特異的なCARを発現する細胞のscFvにおける軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む;
(条件4)
前記第1のCARライブラリが前記条件2を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、前記標的抗原特異的なCARを発現する細胞のscFvにおける重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、第2のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む。
(付記51)
前記第2の投与工程において、複数種類の第2のCARライブラリ発現細胞を投与する、付記50記載の製造方法。
(付記52)
前記第2の投与工程後、前記標的抗原を前記動物に投与する第2の刺激工程を含む、付記50または51記載の製造方法。
(付記53)
前記標的抗原特異的なCARを発現する細胞を、前記標的抗原で再刺激する第1の再刺激工程を含む、付記50から52のいずれかに記載の製造方法。
(付記54)
前記第2の投与工程に先立ち、標的抗原を発現する細胞を前記動物に導入し、前記標的抗原を発現する組織を形成させる第2の形成工程を含む、付記50から53のいずれかに記載の製造方法。
(付記55)
前記第2の採取工程において、前記標的抗原を発現する組織を採取し、前記標的抗原を発現する組織に集積した前記第2のCARライブラリ発現細胞を、前記標的抗原特異的なCARを発現する細胞として採取する、付記50から54のいずれかに記載の製造方法。
(付記56)
前記第2のB細胞受容体は、ヒト由来B細胞のB細胞受容体である、付記50から55のいずれかに記載の製造方法。
(付記57)
前記発現細胞は、T細胞である、付記43から56のいずれかに記載の製造方法。
(付記58)
前記標的抗原は、腫瘍抗原である、付記43から57のいずれかに記載の製造方法。
(付記59)
前記動物は、非ヒト動物である、付記43から58のいずれかに記載の製造方法。
(付記60)
前記動物は、免疫抑制動物である、付記43から59のいずれかに記載の製造方法。
Claims (12)
- 第1のキメラ抗原受容体(CAR)をコードする核酸を含み、
前記第1のCARは、第1の抗原結合ドメインと、第1の膜貫通ドメインと、第1の細胞内シグナル伝達ドメインとを含み、
前記第1の抗原結合ドメインは、標的抗原への結合をスクリーニングする第1の一本鎖抗体(scFv)を含み、
前記第1のscFvは、第1の重鎖可変領域と、第1の軽鎖可変領域とを含み、
前記第1の重鎖可変領域および前記第1の軽鎖可変領域は、下記条件1または条件2を満たす、CARライブラリ;
条件1:
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、第1のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む;
条件2:
前記第1の重鎖可変領域における重鎖相補性決定領域(CDRH)1、CDRH2、およびCDRH3は、それぞれ、第1のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第1の軽鎖可変領域における軽鎖相補性決定領域(CDRL)1、CDRL2、およびCDRL3は、それぞれ、前記標的抗原に結合する抗体もしくはその抗原結合断片における軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む。 - 前記第1の細胞内シグナル伝達ドメインは、CD3ζの細胞内シグナル伝達ドメインを含む、請求項1記載のCARライブラリ。
- 前記第1の膜貫通ドメインは、CD28の膜貫通ドメインを含む、請求項1または2記載のCARライブラリ。
- 前記第1のB細胞受容体は、ヒト由来B細胞のB細胞受容体である、請求項1から3のいずれか一項に記載のCARライブラリ。
- 請求項1から4のいずれか一項に記載のCARライブラリを免疫細胞に発現させる第1の発現工程と、
前記第1の発現工程で得られた免疫細胞と、前記標的抗原とを接触させる第1の接触工程と、
前記第1の接触工程において前記標的抗原と結合した免疫細胞に発現するCARの第1のscFvを、前記標的抗原に結合する第1の候補scFvとして選抜する第1の選抜工程とを含む、scFvの製造方法。 - 前記第1の候補scFvに基づき、第2のCARライブラリを調製する調製工程を含み、
前記第2のCARライブラリは、第2のCARをコードする核酸を含み、
前記第2のCARは、第2の抗原結合ドメインと、第2の膜貫通ドメインと、第2の細胞内シグナル伝達ドメインとを含み、
前記第2の抗原結合ドメインは、前記標的抗原への結合をスクリーニングする第2のscFvを含み、
前記第2のscFvは、第2の重鎖可変領域と、第2の軽鎖可変領域とを含み、
前記第2の重鎖可変領域および前記第2の軽鎖可変領域は、下記条件3または条件4を満たし、
さらに、前記第2のCARライブラリを免疫細胞に発現させる第2の発現工程と、
前記第2の発現工程で得られた免疫細胞と、前記標的抗原とを接触させる第2の接触工程と、
前記第2の接触工程において前記標的抗原と結合した免疫細胞に発現するCARの第2のscFvを、前記標的抗原に結合する第2の候補scFvとして第2の選抜工程とを含む、請求項5記載の製造方法;
条件3:
前記第1の発現工程におけるCARライブラリが前記条件1を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、第2のB細胞受容体の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、前記第1の候補scFvにおける軽鎖可変領域のCDRL1、CDRL2、およびCDRL3を含む;
条件4:
前記第1の発現工程におけるCARライブラリが前記条件2を満たす場合、
前記第2の重鎖可変領域におけるCDRH1、CDRH2、およびCDRH3は、それぞれ、前記第1の候補scFvにおける重鎖可変領域のCDRH1、CDRH2、およびCDRH3を含み、
前記第2の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3は、それぞれ、第2のB細胞受容体の軽鎖可変領域におけるCDRL1、CDRL2、およびCDRL3を含む。 - 前記第2の接触工程を複数回実施する、請求項6記載のscFvの製造方法。
- 前記第2の選抜工程において、前記標的抗原のモノマーまたはマルチマーを用いて、前記標的抗原に結合する免疫細胞を検出し、
検出された免疫細胞に発現するscFvを、前記第2の候補scFvとして選抜する、請求項6または7記載のscFvの製造方法。 - 前記第2のB細胞受容体は、ヒト由来B細胞のB細胞受容体である、請求項6から8のいずれか一項に記載のscFvの製造方法。
- 前記第1の接触工程を複数回実施する、請求項5から9のいずれか一項に記載のscFvの製造方法。
- 前記第1の選抜工程において、前記標的抗原のモノマーまたはマルチマーを用いて、前記標的抗原に結合する免疫細胞を検出し、
検出された免疫細胞に発現するscFvを、前記第1の候補scFvとして選抜する、請求項5から10のいずれか一項に記載のscFvの製造方法。 - 前記免疫細胞は、T細胞である、請求項5から11のいずれか一項に記載のscFvの製造方法。
Priority Applications (8)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| AU2020218446A AU2020218446B2 (en) | 2019-02-04 | 2020-02-04 | "CAR LIBRARY AND scFv MANUFACTURING METHOD |
| EP20752298.8A EP3912992A4 (en) | 2019-02-04 | 2020-02-04 | Car library and production method for scfv |
| US17/428,214 US12241061B2 (en) | 2019-02-04 | 2020-02-04 | CAR library and scFv manufacturing method |
| CN202080012709.7A CN113423720A (zh) | 2019-02-04 | 2020-02-04 | CAR文库和scFv的制造方法 |
| JP2020531552A JP6821230B2 (ja) | 2019-02-04 | 2020-02-04 | CARライブラリおよびscFvの製造方法 |
| AU2023204079A AU2023204079B2 (en) | 2019-02-04 | 2023-06-27 | Car library and scFv manufacturing method |
| US19/035,459 US20250236864A1 (en) | 2019-02-04 | 2025-01-23 | CAR LIBRARY AND scFv MANUFACTURING METHOD |
| AU2025204510A AU2025204510A1 (en) | 2019-02-04 | 2025-06-17 | Car library and scFv manufacturing method |
Applications Claiming Priority (8)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2019-018269 | 2019-02-04 | ||
| JP2019018269 | 2019-02-04 | ||
| JP2019-094188 | 2019-05-18 | ||
| JP2019094188 | 2019-05-18 | ||
| JP2019-140121 | 2019-07-30 | ||
| JP2019140121 | 2019-07-30 | ||
| JP2019-184071 | 2019-10-04 | ||
| JP2019184071 | 2019-10-04 |
Related Child Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US17/428,214 A-371-Of-International US12241061B2 (en) | 2019-02-04 | 2020-02-04 | CAR library and scFv manufacturing method |
| US19/035,459 Continuation US20250236864A1 (en) | 2019-02-04 | 2025-01-23 | CAR LIBRARY AND scFv MANUFACTURING METHOD |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2020162452A1 true WO2020162452A1 (ja) | 2020-08-13 |
Family
ID=71948010
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2020/004123 Ceased WO2020162452A1 (ja) | 2019-02-04 | 2020-02-04 | CARライブラリおよびscFvの製造方法 |
Country Status (6)
| Country | Link |
|---|---|
| US (2) | US12241061B2 (ja) |
| EP (1) | EP3912992A4 (ja) |
| JP (4) | JP6821230B2 (ja) |
| CN (1) | CN113423720A (ja) |
| AU (3) | AU2020218446B2 (ja) |
| WO (1) | WO2020162452A1 (ja) |
Families Citing this family (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CA3214473A1 (en) * | 2021-04-07 | 2022-10-13 | Century Therapeutics, Inc. | Compositions and methods for generating alpha-beta t cells from induced pluripotent stem cells |
| CN117363722A (zh) * | 2023-12-07 | 2024-01-09 | 北京旌准医疗科技有限公司 | 一种用于检测igk和igl基因的引物组合及其应用 |
Citations (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JPS62296890A (ja) | 1986-03-27 | 1987-12-24 | メディカル リサーチ カウンスル | 組換えdna生産物及び製造法 |
| JPH0373280B2 (ja) | 1984-08-15 | 1991-11-21 | Shingijutsu Kaihatsu Jigyodan | |
| JPH04504365A (ja) | 1990-01-12 | 1992-08-06 | アブジェニックス インコーポレイテッド | 異種抗体の生成 |
| JPH04506458A (ja) | 1989-12-21 | 1992-11-12 | セルテック リミテッド | Cd3特異的組換え抗体 |
| JPH06500233A (ja) | 1990-08-29 | 1994-01-13 | ジェンファーム インターナショナル,インコーポレイティド | 異種免疫グロブリンを作る方法及びトランスジェニックマウス |
| WO1994025585A1 (en) | 1993-04-26 | 1994-11-10 | Genpharm International, Inc. | Transgenic non-human animals capable of producing heterologous antibodies |
| JPH07509137A (ja) | 1992-07-24 | 1995-10-12 | セル ジェネシス,インク. | 異種抗体の生産 |
| JP2016533507A (ja) * | 2013-09-30 | 2016-10-27 | エックス−ボディ インコーポレイテッド | 抗原受容体スクリーニングアッセイ |
| WO2018057051A1 (en) * | 2016-09-24 | 2018-03-29 | Abvitro Llc | Affinity-oligonucleotide conjugates and uses thereof |
| JP2018535652A (ja) * | 2015-09-24 | 2018-12-06 | アブビトロ, エルエルシー | 親和性−オリゴヌクレオチドコンジュゲートおよびその使用 |
Family Cites Families (11)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2013096828A1 (en) * | 2011-12-22 | 2013-06-27 | Sea Lane Biotechnologies, Llc | Surrogate binding proteins |
| JP6580579B2 (ja) | 2014-01-29 | 2019-09-25 | ユニバーシティー ヘルス ネットワーク | T細胞受容体を発現する細胞を生産する方法および組成物 |
| TWI805109B (zh) | 2014-08-28 | 2023-06-11 | 美商奇諾治療有限公司 | 對cd19具專一性之抗體及嵌合抗原受體 |
| GB201503742D0 (en) | 2015-03-05 | 2015-04-22 | Ucl Business Plc | Chimeric antigen receptor |
| CN119925616A (zh) | 2015-04-08 | 2025-05-06 | 诺华股份有限公司 | Cd20疗法、cd22疗法和与cd19嵌合抗原受体(car)表达细胞的联合疗法 |
| SG10201913915WA (en) | 2015-09-01 | 2020-03-30 | Univ California | Modular polypeptide libraries and methods of making and using same |
| US10222369B2 (en) | 2016-05-17 | 2019-03-05 | Chimera Bioengineering, Inc. | Methods for making novel antigen binding domains |
| EP3551197B1 (en) | 2016-12-12 | 2023-11-22 | Seattle Children's Hospital (DBA Seattle Children's Research Institute) | Chimeric transcription factor variants with augmented sensitivity to drug ligand induction of transgene expression in mammalian cells |
| JP7341900B2 (ja) | 2017-03-03 | 2023-09-11 | オブシディアン セラピューティクス, インコーポレイテッド | 免疫療法のためのcd19組成物及び方法 |
| JOP20180042A1 (ar) | 2017-04-24 | 2019-01-30 | Kite Pharma Inc | نطاقات ربط مولد ضد متوافقة مع البشر وطرق الاستخدام |
| US11241485B2 (en) | 2017-06-12 | 2022-02-08 | Obsidian Therapeutics, Inc. | PDE5 compositions and methods for immunotherapy |
-
2020
- 2020-02-04 US US17/428,214 patent/US12241061B2/en active Active
- 2020-02-04 WO PCT/JP2020/004123 patent/WO2020162452A1/ja not_active Ceased
- 2020-02-04 AU AU2020218446A patent/AU2020218446B2/en active Active
- 2020-02-04 JP JP2020531552A patent/JP6821230B2/ja active Active
- 2020-02-04 EP EP20752298.8A patent/EP3912992A4/en active Pending
- 2020-02-04 CN CN202080012709.7A patent/CN113423720A/zh active Pending
- 2020-12-24 JP JP2020214844A patent/JP7025059B2/ja active Active
-
2022
- 2022-02-03 JP JP2022015802A patent/JP7496093B2/ja active Active
-
2023
- 2023-06-27 AU AU2023204079A patent/AU2023204079B2/en active Active
-
2024
- 2024-05-16 JP JP2024080118A patent/JP2024105603A/ja active Pending
-
2025
- 2025-01-23 US US19/035,459 patent/US20250236864A1/en active Pending
- 2025-06-17 AU AU2025204510A patent/AU2025204510A1/en active Pending
Patent Citations (10)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JPH0373280B2 (ja) | 1984-08-15 | 1991-11-21 | Shingijutsu Kaihatsu Jigyodan | |
| JPS62296890A (ja) | 1986-03-27 | 1987-12-24 | メディカル リサーチ カウンスル | 組換えdna生産物及び製造法 |
| JPH04506458A (ja) | 1989-12-21 | 1992-11-12 | セルテック リミテッド | Cd3特異的組換え抗体 |
| JPH04504365A (ja) | 1990-01-12 | 1992-08-06 | アブジェニックス インコーポレイテッド | 異種抗体の生成 |
| JPH06500233A (ja) | 1990-08-29 | 1994-01-13 | ジェンファーム インターナショナル,インコーポレイティド | 異種免疫グロブリンを作る方法及びトランスジェニックマウス |
| JPH07509137A (ja) | 1992-07-24 | 1995-10-12 | セル ジェネシス,インク. | 異種抗体の生産 |
| WO1994025585A1 (en) | 1993-04-26 | 1994-11-10 | Genpharm International, Inc. | Transgenic non-human animals capable of producing heterologous antibodies |
| JP2016533507A (ja) * | 2013-09-30 | 2016-10-27 | エックス−ボディ インコーポレイテッド | 抗原受容体スクリーニングアッセイ |
| JP2018535652A (ja) * | 2015-09-24 | 2018-12-06 | アブビトロ, エルエルシー | 親和性−オリゴヌクレオチドコンジュゲートおよびその使用 |
| WO2018057051A1 (en) * | 2016-09-24 | 2018-03-29 | Abvitro Llc | Affinity-oligonucleotide conjugates and uses thereof |
Non-Patent Citations (8)
| Title |
|---|
| "Current Protocols in Molecular Biology", 1987, JOHN WILEY & SONS |
| "NCBI", Database accession no. No. NP_006130.1 |
| ED. HARLOWDAVID LANE: "Antibodies: A Laboratory Manual", 1988, COLD SPRING HARBOR LABORATORY |
| MARKS, J. D. ET AL., J. MOL. BIOL., vol. 222, 1991, pages 581 - 597 |
| NATURE GENETICS, vol. 15, 1997, pages 146 - 156 |
| NATURE GENETICS, vol. 7, 1994, pages 13 - 21 |
| NATURE, vol. 368, 1994, pages 856 - 859 |
| TOSHIMOTO OCHI, MASAKI MARUTA, HAJIME TANIMOTO, HIROSHI FUJIWARA, KATSUTO TAKENAKA, MASATAKA YASUKAWA: "007-1 A novel scFv screening technology by exploiting T cells in combination with CAR-based scFv library", PROGRAMS AND ABSTRACTS OF THE 23RD ANNUAL MEETING OF JAPANESE ASSOCIATION OF CANCER IMMUNOLOGY; AUGUST 21-23, 2019, 16 July 2019 (2019-07-16), JP, pages 172, XP009530041 * |
Also Published As
| Publication number | Publication date |
|---|---|
| JP7496093B2 (ja) | 2024-06-06 |
| EP3912992A4 (en) | 2022-11-23 |
| JP2022070899A (ja) | 2022-05-13 |
| AU2023204079A1 (en) | 2023-07-13 |
| US20250236864A1 (en) | 2025-07-24 |
| US20220290128A1 (en) | 2022-09-15 |
| AU2025204510A1 (en) | 2025-07-10 |
| AU2020218446A1 (en) | 2021-09-16 |
| CN113423720A (zh) | 2021-09-21 |
| JP7025059B2 (ja) | 2022-02-24 |
| JPWO2020162452A1 (ja) | 2021-02-18 |
| AU2023204079B2 (en) | 2025-03-20 |
| JP2024105603A (ja) | 2024-08-06 |
| JP6821230B2 (ja) | 2021-01-27 |
| JP2021063099A (ja) | 2021-04-22 |
| AU2020218446B2 (en) | 2023-05-25 |
| US12241061B2 (en) | 2025-03-04 |
| EP3912992A1 (en) | 2021-11-24 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US20250296970A1 (en) | Cd19-directed chimeric antigen receptors and uses thereof in immunotherapy | |
| JP2022137054A (ja) | NKp46結合タンパク質の可変領域 | |
| JP2020022508A (ja) | 細胞 | |
| US11459389B2 (en) | Monoclonal antibodies that bind human CD161 | |
| CA3086935A1 (en) | Anti-tigit antibodies and their use as therapeutics and diagnostics | |
| JP2022505921A (ja) | Cll1を標的とする抗体およびその応用 | |
| CA3032581A1 (en) | Treatment of cancer using a chimeric antigen receptor in combination with an inhibitor of a pro-m2 macrophage molecule | |
| CA3016287A1 (en) | Cells expressing multiple chimeric antigen receptor (car) molecules and uses therefore | |
| US20230028399A1 (en) | Bcma-directed cellular immunotherapy compositions and methods | |
| JP7496965B2 (ja) | Bcmaに特異的な抗体およびキメラ抗原受容体 | |
| US20250236864A1 (en) | CAR LIBRARY AND scFv MANUFACTURING METHOD | |
| JP2019533675A (ja) | モジュラー四価二重特異性抗体プラットフォーム | |
| CN111848809A (zh) | 靶向Claudin18.2的CAR分子、其修饰的免疫细胞及用途 | |
| JP2021500854A (ja) | Strep−タグ特異的結合タンパク質およびその使用 | |
| JP2024516748A (ja) | EGFRvIII結合タンパク質 | |
| WO2024193669A1 (zh) | 抗tnfr2/抗pd-1或ctla-4双特异性抗体及其应用 | |
| CA2976089C (en) | Humanized anti-muc1* antibodies |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| ENP | Entry into the national phase |
Ref document number: 2020531552 Country of ref document: JP Kind code of ref document: A |
|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 20752298 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2020218446 Country of ref document: AU Date of ref document: 20200204 Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 2020752298 Country of ref document: EP Effective date: 20210820 |
|
| WWG | Wipo information: grant in national office |
Ref document number: 17428214 Country of ref document: US |