EP1403641B1 - Verfahren zum Kalkulieren von Assoziations- und Dissoziationskonstanten mit einem Polymerchip zum Auffinden von ionischen Polymeren - Google Patents
Verfahren zum Kalkulieren von Assoziations- und Dissoziationskonstanten mit einem Polymerchip zum Auffinden von ionischen Polymeren Download PDFInfo
- Publication number
- EP1403641B1 EP1403641B1 EP20030019932 EP03019932A EP1403641B1 EP 1403641 B1 EP1403641 B1 EP 1403641B1 EP 20030019932 EP20030019932 EP 20030019932 EP 03019932 A EP03019932 A EP 03019932A EP 1403641 B1 EP1403641 B1 EP 1403641B1
- Authority
- EP
- European Patent Office
- Prior art keywords
- dna
- protein
- probe
- association
- double
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Expired - Lifetime
Links
- 238000010494 dissociation reaction Methods 0.000 title claims description 53
- 230000005593 dissociations Effects 0.000 title claims description 53
- 238000000034 method Methods 0.000 title claims description 45
- 229920000831 ionic polymer Polymers 0.000 title description 43
- 229920000642 polymer Polymers 0.000 title description 16
- 108020004414 DNA Proteins 0.000 claims description 110
- 108090000623 proteins and genes Proteins 0.000 claims description 82
- 102000004169 proteins and genes Human genes 0.000 claims description 80
- 239000000523 sample Substances 0.000 claims description 71
- 102000053602 DNA Human genes 0.000 claims description 38
- 238000000018 DNA microarray Methods 0.000 claims description 38
- 239000002904 solvent Substances 0.000 claims description 20
- 238000009739 binding Methods 0.000 claims description 14
- 230000027455 binding Effects 0.000 claims description 13
- 238000002198 surface plasmon resonance spectroscopy Methods 0.000 claims description 13
- 239000000758 substrate Substances 0.000 claims description 10
- 238000003556 assay Methods 0.000 claims description 4
- 235000018102 proteins Nutrition 0.000 description 64
- 102000052510 DNA-Binding Proteins Human genes 0.000 description 16
- 239000000126 substance Substances 0.000 description 13
- 108700020911 DNA-Binding Proteins Proteins 0.000 description 12
- FAPWRFPIFSIZLT-UHFFFAOYSA-M Sodium chloride Chemical compound [Na+].[Cl-] FAPWRFPIFSIZLT-UHFFFAOYSA-M 0.000 description 8
- 102000007315 Telomeric Repeat Binding Protein 1 Human genes 0.000 description 7
- 108010033711 Telomeric Repeat Binding Protein 1 Proteins 0.000 description 7
- 238000009396 hybridization Methods 0.000 description 7
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 7
- 230000004568 DNA-binding Effects 0.000 description 6
- 230000000295 complement effect Effects 0.000 description 5
- 238000005406 washing Methods 0.000 description 5
- 230000014509 gene expression Effects 0.000 description 4
- 230000003993 interaction Effects 0.000 description 4
- 230000003287 optical effect Effects 0.000 description 4
- 239000011780 sodium chloride Substances 0.000 description 4
- 239000010409 thin film Substances 0.000 description 4
- 108090001008 Avidin Proteins 0.000 description 3
- 101710096438 DNA-binding protein Proteins 0.000 description 3
- 102100036497 Telomeric repeat-binding factor 1 Human genes 0.000 description 3
- 238000004364 calculation method Methods 0.000 description 3
- 238000011161 development Methods 0.000 description 3
- 108091013637 double-stranded DNA binding proteins Proteins 0.000 description 3
- PCHJSUWPFVWCPO-UHFFFAOYSA-N gold Chemical compound [Au] PCHJSUWPFVWCPO-UHFFFAOYSA-N 0.000 description 3
- 239000010931 gold Substances 0.000 description 3
- 229910052737 gold Inorganic materials 0.000 description 3
- 239000003446 ligand Substances 0.000 description 3
- 239000000463 material Substances 0.000 description 3
- 238000005259 measurement Methods 0.000 description 3
- VLKZOEOYAKHREP-UHFFFAOYSA-N n-Hexane Chemical compound CCCCCC VLKZOEOYAKHREP-UHFFFAOYSA-N 0.000 description 3
- 230000008569 process Effects 0.000 description 3
- 108700014590 single-stranded DNA binding proteins Proteins 0.000 description 3
- 239000000243 solution Substances 0.000 description 3
- 238000001179 sorption measurement Methods 0.000 description 3
- CSCPPACGZOOCGX-UHFFFAOYSA-N Acetone Chemical compound CC(C)=O CSCPPACGZOOCGX-UHFFFAOYSA-N 0.000 description 2
- 108020004635 Complementary DNA Proteins 0.000 description 2
- 239000003298 DNA probe Substances 0.000 description 2
- KCXVZYZYPLLWCC-UHFFFAOYSA-N EDTA Chemical compound OC(=O)CN(CC(O)=O)CCN(CC(O)=O)CC(O)=O KCXVZYZYPLLWCC-UHFFFAOYSA-N 0.000 description 2
- ROHFNLRQFUQHCH-YFKPBYRVSA-N L-leucine Chemical compound CC(C)C[C@H](N)C(O)=O ROHFNLRQFUQHCH-YFKPBYRVSA-N 0.000 description 2
- ROHFNLRQFUQHCH-UHFFFAOYSA-N Leucine Natural products CC(C)CC(N)C(O)=O ROHFNLRQFUQHCH-UHFFFAOYSA-N 0.000 description 2
- NBIIXXVUZAFLBC-UHFFFAOYSA-N Phosphoric acid Chemical compound OP(O)(O)=O NBIIXXVUZAFLBC-UHFFFAOYSA-N 0.000 description 2
- WCUXLLCKKVVCTQ-UHFFFAOYSA-M Potassium chloride Chemical compound [Cl-].[K+] WCUXLLCKKVVCTQ-UHFFFAOYSA-M 0.000 description 2
- 229920001222 biopolymer Polymers 0.000 description 2
- 239000000872 buffer Substances 0.000 description 2
- 238000010804 cDNA synthesis Methods 0.000 description 2
- 210000000349 chromosome Anatomy 0.000 description 2
- 239000002299 complementary DNA Substances 0.000 description 2
- 239000000499 gel Substances 0.000 description 2
- 239000011521 glass Substances 0.000 description 2
- 150000002500 ions Chemical class 0.000 description 2
- 230000007246 mechanism Effects 0.000 description 2
- 230000035772 mutation Effects 0.000 description 2
- 239000002547 new drug Substances 0.000 description 2
- 102000039446 nucleic acids Human genes 0.000 description 2
- 108020004707 nucleic acids Proteins 0.000 description 2
- 150000007523 nucleic acids Chemical class 0.000 description 2
- 229920001308 poly(aminoacid) Polymers 0.000 description 2
- 230000006798 recombination Effects 0.000 description 2
- 238000005215 recombination Methods 0.000 description 2
- 238000011160 research Methods 0.000 description 2
- 230000009870 specific binding Effects 0.000 description 2
- JKMHFZQWWAIEOD-UHFFFAOYSA-N 2-[4-(2-hydroxyethyl)piperazin-1-yl]ethanesulfonic acid Chemical compound OCC[NH+]1CCN(CCS([O-])(=O)=O)CC1 JKMHFZQWWAIEOD-UHFFFAOYSA-N 0.000 description 1
- 208000030507 AIDS Diseases 0.000 description 1
- 241000894006 Bacteria Species 0.000 description 1
- 102100034330 Chromaffin granule amine transporter Human genes 0.000 description 1
- 230000004543 DNA replication Effects 0.000 description 1
- 241000701959 Escherichia virus Lambda Species 0.000 description 1
- LFQSCWFLJHTTHZ-UHFFFAOYSA-N Ethanol Chemical compound CCO LFQSCWFLJHTTHZ-UHFFFAOYSA-N 0.000 description 1
- 241000206602 Eukaryota Species 0.000 description 1
- 108010046914 Exodeoxyribonuclease V Proteins 0.000 description 1
- 239000007995 HEPES buffer Substances 0.000 description 1
- 102100039869 Histone H2B type F-S Human genes 0.000 description 1
- 101000641221 Homo sapiens Chromaffin granule amine transporter Proteins 0.000 description 1
- 101001035372 Homo sapiens Histone H2B type F-S Proteins 0.000 description 1
- XUJNEKJLAYXESH-REOHCLBHSA-N L-Cysteine Chemical compound SC[C@H](N)C(O)=O XUJNEKJLAYXESH-REOHCLBHSA-N 0.000 description 1
- HNDVDQJCIGZPNO-YFKPBYRVSA-N L-histidine Chemical compound OC(=O)[C@@H](N)CC1=CN=CN1 HNDVDQJCIGZPNO-YFKPBYRVSA-N 0.000 description 1
- 108060004795 Methyltransferase Proteins 0.000 description 1
- 206010028980 Neoplasm Diseases 0.000 description 1
- 108010047956 Nucleosomes Proteins 0.000 description 1
- OAICVXFJPJFONN-UHFFFAOYSA-N Phosphorus Chemical group [P] OAICVXFJPJFONN-UHFFFAOYSA-N 0.000 description 1
- 239000004743 Polypropylene Substances 0.000 description 1
- 239000004793 Polystyrene Substances 0.000 description 1
- 102000001218 Rec A Recombinases Human genes 0.000 description 1
- 108010055016 Rec A Recombinases Proteins 0.000 description 1
- VYPSYNLAJGMNEJ-UHFFFAOYSA-N Silicium dioxide Chemical compound O=[Si]=O VYPSYNLAJGMNEJ-UHFFFAOYSA-N 0.000 description 1
- 101710183280 Topoisomerase Proteins 0.000 description 1
- 108091023040 Transcription factor Proteins 0.000 description 1
- 102000040945 Transcription factor Human genes 0.000 description 1
- HCHKCACWOHOZIP-UHFFFAOYSA-N Zinc Chemical compound [Zn] HCHKCACWOHOZIP-UHFFFAOYSA-N 0.000 description 1
- PTFCDOFLOPIGGS-UHFFFAOYSA-N Zinc dication Chemical compound [Zn+2] PTFCDOFLOPIGGS-UHFFFAOYSA-N 0.000 description 1
- 229910000147 aluminium phosphate Inorganic materials 0.000 description 1
- 235000001014 amino acid Nutrition 0.000 description 1
- 150000001413 amino acids Chemical class 0.000 description 1
- 238000004458 analytical method Methods 0.000 description 1
- 239000007864 aqueous solution Substances 0.000 description 1
- 230000037429 base substitution Effects 0.000 description 1
- 230000008901 benefit Effects 0.000 description 1
- 230000015572 biosynthetic process Effects 0.000 description 1
- 108091006374 cAMP receptor proteins Proteins 0.000 description 1
- 201000011510 cancer Diseases 0.000 description 1
- 239000013522 chelant Substances 0.000 description 1
- 238000005352 clarification Methods 0.000 description 1
- 230000008878 coupling Effects 0.000 description 1
- 238000010168 coupling process Methods 0.000 description 1
- 238000005859 coupling reaction Methods 0.000 description 1
- 235000018417 cysteine Nutrition 0.000 description 1
- XUJNEKJLAYXESH-UHFFFAOYSA-N cysteine Natural products SCC(N)C(O)=O XUJNEKJLAYXESH-UHFFFAOYSA-N 0.000 description 1
- 230000003247 decreasing effect Effects 0.000 description 1
- 102000022788 double-stranded DNA binding proteins Human genes 0.000 description 1
- 239000003814 drug Substances 0.000 description 1
- 229940079593 drug Drugs 0.000 description 1
- 238000009509 drug development Methods 0.000 description 1
- 238000001035 drying Methods 0.000 description 1
- 230000000694 effects Effects 0.000 description 1
- 239000010408 film Substances 0.000 description 1
- 239000003574 free electron Substances 0.000 description 1
- HNDVDQJCIGZPNO-UHFFFAOYSA-N histidine Natural products OC(=O)C(N)CC1=CN=CN1 HNDVDQJCIGZPNO-UHFFFAOYSA-N 0.000 description 1
- 229910052739 hydrogen Inorganic materials 0.000 description 1
- 239000001257 hydrogen Substances 0.000 description 1
- 230000003100 immobilizing effect Effects 0.000 description 1
- 230000001900 immune effect Effects 0.000 description 1
- 238000012423 maintenance Methods 0.000 description 1
- 238000004519 manufacturing process Methods 0.000 description 1
- 239000003550 marker Substances 0.000 description 1
- 239000012528 membrane Substances 0.000 description 1
- 239000000203 mixture Substances 0.000 description 1
- 238000012544 monitoring process Methods 0.000 description 1
- 210000001623 nucleosome Anatomy 0.000 description 1
- 125000002467 phosphate group Chemical group [H]OP(=O)(O[H])O[*] 0.000 description 1
- 239000000049 pigment Substances 0.000 description 1
- 229920001184 polypeptide Polymers 0.000 description 1
- -1 polypropylene Polymers 0.000 description 1
- 229920001155 polypropylene Polymers 0.000 description 1
- 229920002223 polystyrene Polymers 0.000 description 1
- 239000001103 potassium chloride Substances 0.000 description 1
- 235000011164 potassium chloride Nutrition 0.000 description 1
- 238000007781 pre-processing Methods 0.000 description 1
- 108090000765 processed proteins & peptides Proteins 0.000 description 1
- 102000004196 processed proteins & peptides Human genes 0.000 description 1
- 238000012545 processing Methods 0.000 description 1
- 108020003175 receptors Proteins 0.000 description 1
- 102000005962 receptors Human genes 0.000 description 1
- 230000008439 repair process Effects 0.000 description 1
- 230000004044 response Effects 0.000 description 1
- 229920002477 rna polymer Polymers 0.000 description 1
- 239000012488 sample solution Substances 0.000 description 1
- 238000012216 screening Methods 0.000 description 1
- 230000019491 signal transduction Effects 0.000 description 1
- 239000000741 silica gel Substances 0.000 description 1
- 229910002027 silica gel Inorganic materials 0.000 description 1
- 229910001415 sodium ion Inorganic materials 0.000 description 1
- 229920001059 synthetic polymer Polymers 0.000 description 1
- 238000011282 treatment Methods 0.000 description 1
- 241001515965 unidentified phage Species 0.000 description 1
- 239000011701 zinc Substances 0.000 description 1
- 229910052725 zinc Inorganic materials 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C40—COMBINATORIAL TECHNOLOGY
- C40B—COMBINATORIAL CHEMISTRY; LIBRARIES, e.g. CHEMICAL LIBRARIES
- C40B30/00—Methods of screening libraries
- C40B30/04—Methods of screening libraries by measuring the ability to specifically bind a target molecule, e.g. antibody-antigen binding, receptor-ligand binding
Definitions
- the present invention relates to a method of calculating the association constant and the dissociation constant between a probe and a protein, a method of searching for the functionality of proteins using the calculation method, and a method of identifying proteins using the calculation method.
- the invention relates to a method of searching for the functionality of proteins by calculating the DNA-binding property of the proteins, and a method of identifying the proteins.
- DNA-binding proteins are considered to be among the important substances controlling or adjusting the expression of gene functions, the important substances controlling or adjusting the expression of gene functions, and clarification of their DNA binding properties is an important task.
- a DNA is a polymer formed by a sequence of four kinds of bases, and two DNAs can specifically bind to one another through their sequences. Because the nucleic acids of both such DNAs are negatively charged, hybridization is carried out in a highly ionic solvent. Conventional biochips take advantage of this fact to detect a sample DNA by immobilizing a probe DNA on a substrate.
- a DNA-binding protein cannot be identified by a conventional biochip. This is because a protein as a sample may be specifically adsorbed on a probe DNA on a biochip by its complementary DNA portion, and the protein may also be nonspecifically adsorbed on the probe DNA even though this is not supposed to happen, in light of the DNA. Because of such specific and nonspecific adsorptions that apparently occur on the probe DNA, it has been virtually impossible to identify sample proteins using a conventional biochip.
- association constant and the dissociation constant between ionic polymers are calculated based on a binding curve or dissociation curve obtained by monitoring the adsorption of a substance using an apparatus for measuring the strength of bonding between ionic polymers, such as a surface plasmon resonance measuring apparatus (see JP Patent Publication (Kohyo) No. 7-507865, for example).
- a ligand is immobilized on a sensor surface, and a sample containing a substance that acts on the ligand is added via a microchannel system. Minute mass changes due to the association or dissociation of molecules on the sensor surface are then monitored in real time in terms of changes in a surface plasmon resonance signal.
- This technique is effective in studying the interaction between biomolecules or the correlation between structure and function.
- Surface plasmon resonance is being used in a wide variety of areas including basic research, protein engineering, and the screening of new drugs.
- surface plasmon resonance is utilized include research into the mechanism of AIDS or cancer development, immunologic response, signal transduction, the ways in which receptors and ligands bind to one another, and the mechanism of gene expression control (see Chong L et al. A human telomeric protein. Science 1995 Dec 8; 270 (5242): 1663-7, for example).
- WO 02/18648 discloses a method of analysis of binding interactions, wherein a protein is bound to a nucleic acid chip. Fluorescence values are normalized versus the highest fluorescence intensity on a spot for the particular measurement. Selective washing steps to increase the specificity of binding are not disclosed.
- US 6,355,428 discloses a method for determining the relative binding affinity between DNA immobilized on a chip and DNA-binding proteins, wherein the second ionic polymer is labeled. Again, selective washing steps for increasing the specificity are not disclosed.
- the method using an ionic-polymer identifying chip makes it possible to determine the presence or absence of binding between double-stranded DNA and DNA-binding proteins or the relative association strengths for a large number of proteins simultaneously.
- This method is not capable of calculating the association and dissociation constants, which are absolute values.
- the invention was made upon the realization that the calculation of rough values of association and dissociation constants of many kinds of double-stranded DNA samples, which used to take a long time, can be made in a short time by using a specific protein identifying chip.
- the aforementioned need is solved by a method according to claim 1.
- the first to third ionic polymers may be selected from the group consisting of proteins, polyamino acids, DNA, RNA, and synthetic polymers.
- the invention provides a method of calculating association and dissociation constants between a probe and a protein, comprising:
- Examples of the method of measuring the association strength between a double-stranded DNA and a protein include a surface plasmon resonance method and a gel shift assay method.
- the surface plasmon resonance method is superior in accuracy and operability, and is suitable for the method of calculating the association and dissociation constants according to the invention.
- the invention also provides a method of searching for the functionality of proteins using the method of calculating association and dissociation constants as described above, wherein the binding property between the probe in which the first and second DNA are complementarily bound and the protein is evaluated.
- the material and shape of the substrate used in the present invention are not particularly limited, and the materials and shapes typically used for the substrates for biochips can be used.
- the material is selected from the group consisting of glass, silica gel, polystyrene, polypropylene, and membrane.
- the shape is preferably either plate-like or bead-like.
- the marker to be provided to the second DNA is selected from those typically used in the field of biochips. Particularly, a fluorescent substance is preferable from the viewpoint of optical processes in a read step.
- a fourth, sample polymer with an opposite ionic property should preferably be added so that they can be firmly immobilized, followed by a step of washing them with a nonionic solvent.
- the number of base sequences in the DNA should preferably be between 4 to 13 for each polymer.
- the complementary binding of the DNA comprising the first and second DNA is made stronger, and further the specific binding between the DNA chip comprising these polymers and the DNA-binding proteins as samples based on their chemical and physical structures is made stronger.
- the search for and identification of the functionality of the sample proteins can be made more easily and accurately.
- the nonionic solvent used in the invention is not particularly limited.
- it is a mixture of one or more kinds of substances selected from the group consisting of pure water, alcohol, acetone, and hexane.
- the ionic solvent is not particularly limited either. It may be an aqueous solution of sodium chloride or potassium chloride.
- Biopolymers such as DNA and proteins have a charge, which is negative in the case of DNA due to the phosphate group and either positive or negative in the case of proteins depending on the kind of amino acids.
- the double strand of DNA normally has a negative ion and is therefore repulsive, so that it does not hybridize in pure water. However, it can hybridize by hydrogen bonding in a system where a positive ion exists, as an Na + ion chelates with the phosphor group of DNA, for example. Proteins with a positive charge bind with DNA having a negative charge at the charge portion.
- the polymers with the same charge are dissociated when the ionic strength is low, whereas the polymers with opposite charges strongly bind to one another.
- the ionic strength is high, even polymers with the same charge can bind to one another, while polymers with different charges lose their association strength and are dissociated.
- the DNA-binding proteins as used herein refer to proteins that have affinity for DNA and that specifically or nonspecifically bind to base sequences.
- examples of such proteins include, mainly, 1) double-stranded DNA-binding proteins that control gene expression by causing changes in DNA structure, 2) single-stranded DNA-binding proteins that are indispensable for the process of duplication, recombination, or repair of DNA, 3) proteins involved with the maintenance of higher-order structures of chromosomes, 4) ATP-dependent Dnase, and 5) topoisomerases forming the DNA superhelix structure.
- proteins 1 are transcription factors, for which several structural motifs are known, such as helix-turn-helix that was found in the structures of lambda phage Cro protein and cAMP receptor proteins, zinc finger in which cysteine and histidine chelate a zinc ion, and leucine zipper in which two molecules of proteins comprising an alpha helix with leucine disposed on one side which are combined like a zipper.
- the proteins 2) are called SSB (single-stranded DNA-binding proteins), and they are found in various life forms ranging from bacteriophage to higher forms of life.
- the proteins 3) are represented by the histone proteins that exist in the chromosomes of eukaryotes, and they form a nucleosome structure.
- the proteins 4 include helicase that promote DNA replication (DnaB proteins, for example), recombination (RecA and RecBC proteins), and the unwinding of double-stranded DNA. Further, in the present invention, not only the above-mentioned proteins that inherently have DNA binding property but also proteins to which a DNA binding property has been given by physical or chemical treatments may be used.
- the above-mentioned various DNA-binding proteins can be used.
- the invention includes cases using not only one kind of DNA-binding proteins but two or more kinds of DNA-binding proteins.
- DNA-binding proteins characteristically have a positive charge at the DNA binding domain and bind to DNA or RNA with negative charges either specifically or nonspecifically.
- the double-stranded DNA to which these DNA-binding proteins have bound has increasing association strength with decreasing ionic strength.
- Fig. 1 shows a protein identifying DNA chip as an example of the polymer chip for identifying ionic polymers according to the invention.
- a first DNA 2 having a base sequence GCTA is immobilized on a substrate 1.
- a second DNA 3 labeled with a fluorescent substance and having a base sequence CGAT is added to an ionic solvent in the presence of the first DNA 2, they complimentarily bind to each other and form a double strand.
- a second DNA 6 labeled with the same fluorescent substance and having a base sequence GGTT is added to another DNA 5 having a base sequence CCAA, they complimentarily bind to each other and form a double strand.
- the protein 4 As a protein 4 that has not been identified is added to the DNA chip, the protein 4 binds to the double strand with the complementary binding of DNA 2 and DNA 3. However, the protein 4 does not bind to the double strand with the complementary biding of DNA 5 and DNA 6. Such phenomena can be measured with a fluorescence intensity meter and then analyzed in order to identify the protein.
- Fig. 2 shows a chart illustrating the process of identifying proteins using the protein identifying DNA chip shown in Fig. 1.
- a DNA chip is prepared, which comprises a substrate 1 on which DNA 2 and DNA 5 are immobilized (a).
- DNA 3 and DNA 6 that are labeled with a fluorescent substance are caused to complimentarily bind on the DNA chip in the presence of an ionic solvent (b).
- unidentified proteins 4 are added.
- the proteins 4 specifically bind to the double strands of DNAs 2 and 3 as they should, and they also nonspecifically bind to the double strands of DNAs 5 and 6 although they should not (c).
- the DNA chip is washed with an ionic solvent to remove the nonspecifically bound proteins (d).
- the DNA chip is further washed with a nonionic solvent to remove DNA 6 that is labeled with fluorescent substance (e).
- the remaining fluorescence is then read, and the protein is identified based on the result of reading (f).
- Fig. 3 shows the principle of measurement carried out by a surface plasmon resonance measuring apparatus available from Biacore.
- a gold thin film is formed at the bottom of a prism to a thickness of 50 nm, and the interface is irradiated with polarized light of wavelength 760 nm. This produces an energy wave called an evanescent wave on the thin film. Because this evanescent wave is utilized for the resonance of a free electron wave or a plasmon wave on the gold thin film, a loss of energy is observed at a certain angle of the reflected light. When the intensity of the reflected light is measured with a photodiode array, an optical valley is recognized, as indicated by I. This optical phenomena is the surface plasmon resonance, in which the loss angle varies depending on changes in the concentration of the solvent on the gold film surface.
- a synthesized oligo DNA is spotted on a substrate, and a complementary oligo DNA with a labeled terminal is hybridized, thus manufacturing a double-stranded DNA chip.
- a substance of which the association constant is to be measured is added onto the double-stranded DNA chip as a sample.
- the complementary DNA is dissociated if the sample has not bound to the double-stranded DNA.
- it can be determined to what degree the sample to be measured has bound to the double-stranded DNA.
- the values of dissociation constants KD for three or more kinds of samples on the chip are determined using surface plasmon measuring equipment, and an analytical curve is made.
- the dissociation constant KD values can be determined from the fluorescence intensity ratio using the analytical curve.
- TRF1 is introduced in the above-mentioned Non-Patent Document 1 (Chong L et al. A human telomeric protein. Science 1995 Dec 8; 270(5242): 1663-7).
- a biochip was fabricated as follows.
- probe DNA to be immobilized on a chip Four kinds of probe DNA to be immobilized on a chip were provided, and their respective 5' terminals were biotinylated. They were then spotted on six locations of an avidin-coated slide glass for each kind of probe using SPBIO (available from Hitachi Software Engineering Co., Ltd.). In the present example, four kinds of probe DNA were used, so that 24 spots were made.
- the sequences of the chip-immobilized probe DNA were 5'-GTTAGGGTTAGGG-3, 5'-GTTAAGGTCAGGG-3, 5'-GTTAAGGTTAGGG-3, and 5'-GTTAGGGCTAGGG-3. In the following, they will be referred to as an AGTT probe, an AATC probe, an AATT probe, and an AGCT probe.
- the concentration of the spotted probe DNA was adjusted to 1 mM with pure water.
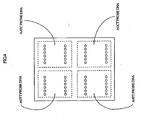
- An example of the thus spotted biochip is shown in Fig. 4.
- the 5'-terminal was labeled with a fluorescent pigment Cy5.
- a 5xSSC solution was prepared containing the sample DNA to a concentration of 100 nM, and 10 ⁇ l of the solution was added onto the prepared biochip to effect hybridization for 30 minutes in a humid environment so as to avoid drying.
- the chip was washed with 1xSSC, dried, and then read using a fluorescent scanner CRBIO2 (available from Hitachi Software Engineering Co., Ltd.).
- FIG. 5 An example of an image that has been read is shown in Fig. 5.
- darkened spots indicate the locations where the Cy5-labeled sample exists. Thus, the formation of a double-stranded DNA can be confirmed.
- TRF1 protein sample solution prepared with 5 mM of KPB (phosphoric acid) and 30 mM of NaCl to a TRF1 protein concentration of 110 ⁇ M was added onto the hybridized biochip, and hybridization was effected for 10 minutes.
- Fig. 6 shows an image that has been read after processing in Fig. 5.
- the double-stranded DNA hybridized with TRF1 does not dissociate even in pure water.
- the difference in fluorescent intensity among the four kinds of probes shown in Fig. 6 corresponds to the difference in association strength between the TRF1 protein and the double-stranded DNA chip.
- Fig. 7 shows a table of the fluorescent intensity ratios for the individual probe sequences.
- the above-described double-stranded DNA probes were then immobilized on an avidin-coated chip using the surface plasmon resonance measuring apparatus.
- the avidin coat was Sensor Chip SA available from Biacore.
- the immobilization buffer employed 10 mM HEPES pH7.4 + 3M EDTA + 150 mM NaCl.
- the flow volume during immobilization was 5 ⁇ l/min.
- the TRF1 protein was caused to flow on the immobilized chip in five different concentrations of 10 nM, 25 nM, 50 nM, 75 nM, and 100 nM.
- the TRF 1 binding amount was monitored using the surface plasmon measuring apparatus to prepare an association curve and a dissociation curve, based on which the dissociation constant (KD value) was calculated.
- the coupling buffer employed 10 mM HEFES pH 7.4 + 3M EDTA + 50 mM NaCl, and the flow volume was 20 ⁇ l/min.
- the accurate dissociation constants of the four kinds of probe are shown in the table of Fig. 7.
- FIG. 8 shows an analytical curve prepared based on the probes AGTT, AATC, and AATT.
- the vertical axis indicates the dissociation constant (KD value), while the horizontal axis indicates the fluorescent intensity ratio (T/H).
- the fluorescent intensity can read from the relevant spot on the biochip and compared with the analytical curve in order to calculate a rough dissociation constant.
- the fluorescent intensity of the AGCT probe is 0.437, as shown in Fig. 7, so that it can be expected that the dissociation constant lies roughly at the latter half of 10 -8 , based on the analytical curve of Fig. 8.
- the accurate dissociation constant KD value
- the association constant (KA value) and the dissociation constant (KD value), which are indicators of the structure and function of the mutation, can be comprehensively and effectively estimated by a simple method.
- the invention makes it possible to calculate rough association and dissociation constants in a number of mutants in which single base substitution such as SNP has occurred in an easy and quick manner.
- the invention can provide tools for drug development and/or medical examinations.
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Health & Medical Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Life Sciences & Earth Sciences (AREA)
- Biochemistry (AREA)
- Immunology (AREA)
- General Chemical & Material Sciences (AREA)
- Medicinal Chemistry (AREA)
- Molecular Biology (AREA)
- General Health & Medical Sciences (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
- Investigating Or Analysing Materials By Optical Means (AREA)
Claims (4)
- Verfahren zum Berechnen von Assoziations- und Dissoziationskonstanten zwischen einer Sonde und einem Protein, umfassend:einen ersten Schritt der Herstellung eines DNA-Chip zum Identifizieren von Proteinen, wobei der DNA-Chip ein Substrat umfaßt, auf dem eine erste DNA immobilisiert ist, wobei eine zweite DNA, die markiert ist, an die erste DNA in einem ionischen Lösungsmittel komplementär gebunden ist, um eine Sonde mit einer doppelsträngigen DNA zu bilden, wobei die Sonde dann getrocknet wird;einen zweiten Schritt, bei dem ein Protein als eine Probe dazu gebracht wird, auf der Sonde zu adsorbieren,einen dritten Schritt der Berechnung von genauen Assoziations- und Dissoziationskonstanten zwischen der Sonde mit einer doppelsträngigen DNA, bestehend aus der ersten und zweiten DNA, und dem Protein, unter Verwendung eines Verfahrens zum Messen der Assoziationsstärke zwischen einer doppelsträngigen DNA und einem Protein, wie etwa Oberflächenplasmonresonanzverfahren und Gel-Shift-Assays, undeinen vierten Schritt der Herstellung einer analytischen Kurve, basierend auf einer Korrelation zwischen der relativen Assoziationsstärke zwischen der doppelsträngigen DNA und dem Protein, erhalten in dem zweiten Schritt, und den genauen Assoziations- und Dissoziationskonstanten zwischen der doppelsträngigen DNA und dem Protein, erhalten in dem dritten Schritt, um die Assoziations- und Dissoziationskonstanten zwischen der Sonde mit der doppelsträngigen DNA, in der die erste und zweite DNA komplementär gebunden sind, und dem Protein zu berechnen,dadurch gekennzeichnet,
daß in dem zweiten Schritt, nachdem ein Protein als eine Probe dazu gebracht wird, auf der Sonde zu adsorbieren, der DNA-Chip dann mit einem ionischen Lösungsmittel gewaschen wird, um Proteine zu entfemen, die nicht-spezifisch gebunden sind, und daß der DNA-Chip dann mit einem nicht-ionischen Lösungsmittel gewaschen wird, um die zweite markierte DNA zu entfemen, außer da, wo Proteine spezifisch gebunden sind, und daß die Menge an verbleibender Markierung abgelesen wird. - Verfahren zum Berechnen von Assoziations- und Dissoziationskonstanten nach Anspruch 1, wobei das Verfahren zum Messen der Assoziationsstärke zwischen einer doppelsträngigen DNA und einem Protein ein Oberflächenplasmonverfahren oder ein Gel-Shift-Assay-Verfahren ist.
- Verfahren zum Untersuchen der Funktionalität von Proteinen unter Verwendung des Verfahrens zum Berechnen von Assoziations- und Dissoziationskonstanten nach einem der Ansprüche 1-2, wobei die Bindungseigenschaft zwischen der Sonde, in der die erste und zweite DNA komplementär gebunden sind, und dem Protein bewertet wird.
- Verfahren zum Identifizieren von Proteinen unter Verwendung des Verfahrens zum Berechnen von Assoziations- und Dissoziationskonstanten nach einem der Ansprüche 1-2, wobei die Bindungseigenschaft zwischen der Sonde, in der die erste und zweite DNA komplementär gebunden sind, und dem Protein bewertet wird.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2002281530A JP2004117201A (ja) | 2002-09-26 | 2002-09-26 | イオン性高分子同定用高分子チップを用いた結合定数および解離定数算出方法 |
| JP2002281530 | 2002-09-26 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| EP1403641A1 EP1403641A1 (de) | 2004-03-31 |
| EP1403641B1 true EP1403641B1 (de) | 2006-03-29 |
Family
ID=31973313
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| EP20030019932 Expired - Lifetime EP1403641B1 (de) | 2002-09-26 | 2003-09-02 | Verfahren zum Kalkulieren von Assoziations- und Dissoziationskonstanten mit einem Polymerchip zum Auffinden von ionischen Polymeren |
Country Status (3)
| Country | Link |
|---|---|
| EP (1) | EP1403641B1 (de) |
| JP (1) | JP2004117201A (de) |
| DE (1) | DE60304262T8 (de) |
Families Citing this family (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP4520873B2 (ja) * | 2005-02-02 | 2010-08-11 | セイコーインスツル株式会社 | 解離定数測定装置、解離定数測定方法、及び解離定数測定プログラム |
| CN101419214B (zh) * | 2007-10-23 | 2012-07-04 | 中国科学院上海药物研究所 | 基于分层原子加和模型的分子酸碱解离常数的预测方法 |
Family Cites Families (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US6420109B1 (en) * | 1998-09-11 | 2002-07-16 | Genelabs Technologies, Inc. | Nucleic acid ligand interaction assays |
| WO2002018648A2 (en) * | 2000-08-25 | 2002-03-07 | President And Fellows Of Harvard College | Analysis of binding interactions |
| JP2003284552A (ja) * | 2002-03-28 | 2003-10-07 | Hitachi Software Eng Co Ltd | イオン性高分子同定用高分子チップ及びイオン性高分子の同定方法 |
-
2002
- 2002-09-26 JP JP2002281530A patent/JP2004117201A/ja active Pending
-
2003
- 2003-09-02 EP EP20030019932 patent/EP1403641B1/de not_active Expired - Lifetime
- 2003-09-02 DE DE2003604262 patent/DE60304262T8/de active Active
Also Published As
| Publication number | Publication date |
|---|---|
| DE60304262T2 (de) | 2006-12-28 |
| EP1403641A1 (de) | 2004-03-31 |
| DE60304262T8 (de) | 2007-05-16 |
| JP2004117201A (ja) | 2004-04-15 |
| DE60304262D1 (de) | 2006-05-18 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| AU773291B2 (en) | UPA, a universal protein array system | |
| EP1307743B1 (de) | Kolloidzusammensetzungen für festphasen-biomolekulare analytische systeme | |
| JP5538726B2 (ja) | 表面処理されていない基質にプローブを固定するためのゾル−ゲルバイオチップ用のゾル組成物及びそのスクリーニング方法 | |
| US20180327824A1 (en) | Microarrays | |
| US20170370927A1 (en) | Method of measuring the affinity of biomolecules | |
| JP2002531098A (ja) | 表面上でのクローニング及びコピー作製方法 | |
| JP4425640B2 (ja) | Dna−結合タンパク質の検出法 | |
| Yingyongnarongkul et al. | Parallel and multiplexed bead-based assays and encoding strategies | |
| EP1403641B1 (de) | Verfahren zum Kalkulieren von Assoziations- und Dissoziationskonstanten mit einem Polymerchip zum Auffinden von ionischen Polymeren | |
| CA2686537A1 (en) | Nucleic acid chip for obtaining bind profile of single strand nucleic acid and unknown biomolecule, manufacturing method thereof and analysis method of unknown biomolecule using nucleic acid chip | |
| EP1348767A2 (de) | Polymer-chip und Verfahren zum Identifizieren eines ionischen Polymer | |
| WO2017059239A1 (en) | Methods for processing biopolymeric arrays | |
| Venton et al. | Screening combinatorial libraries | |
| JP2005521056A (ja) | アレイ | |
| US20250354988A1 (en) | Time-dependent profiling of binding interactions | |
| EP1974211B1 (de) | Molekulare identifizierung über membrankonstruierte zellen | |
| WO2025079662A1 (ja) | 生体分子間相互作用の新規解析法 | |
| WO2001036585A1 (en) | Sequence tag microarray and method for detection of multiple proteins through dna methods | |
| Cansız | Development of a sandwich-type dna array platform for the detection of label-free oligonucleotide targets | |
| AU2384999A (en) | Highly specific surfaces for biological reactions, method of preparation and utilization |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PUAI | Public reference made under article 153(3) epc to a published international application that has entered the european phase |
Free format text: ORIGINAL CODE: 0009012 |
|
| 17P | Request for examination filed |
Effective date: 20030902 |
|
| AK | Designated contracting states |
Kind code of ref document: A1 Designated state(s): AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HU IE IT LI LU MC NL PT RO SE SI SK TR |
|
| AX | Request for extension of the european patent |
Extension state: AL LT LV MK |
|
| 17Q | First examination report despatched |
Effective date: 20040811 |
|
| AKX | Designation fees paid |
Designated state(s): DE FR GB |
|
| GRAP | Despatch of communication of intention to grant a patent |
Free format text: ORIGINAL CODE: EPIDOSNIGR1 |
|
| GRAS | Grant fee paid |
Free format text: ORIGINAL CODE: EPIDOSNIGR3 |
|
| GRAA | (expected) grant |
Free format text: ORIGINAL CODE: 0009210 |
|
| AK | Designated contracting states |
Kind code of ref document: B1 Designated state(s): DE FR GB |
|
| REG | Reference to a national code |
Ref country code: GB Ref legal event code: FG4D |
|
| REF | Corresponds to: |
Ref document number: 60304262 Country of ref document: DE Date of ref document: 20060518 Kind code of ref document: P |
|
| ET | Fr: translation filed | ||
| PLBE | No opposition filed within time limit |
Free format text: ORIGINAL CODE: 0009261 |
|
| STAA | Information on the status of an ep patent application or granted ep patent |
Free format text: STATUS: NO OPPOSITION FILED WITHIN TIME LIMIT |
|
| 26N | No opposition filed |
Effective date: 20070102 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: FR Payment date: 20110928 Year of fee payment: 9 Ref country code: GB Payment date: 20110920 Year of fee payment: 9 Ref country code: DE Payment date: 20110923 Year of fee payment: 9 |
|
| GBPC | Gb: european patent ceased through non-payment of renewal fee |
Effective date: 20120902 |
|
| REG | Reference to a national code |
Ref country code: FR Ref legal event code: ST Effective date: 20130531 |
|
| REG | Reference to a national code |
Ref country code: DE Ref legal event code: R119 Ref document number: 60304262 Country of ref document: DE Effective date: 20130403 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: GB Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20120902 Ref country code: DE Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20130403 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: FR Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20121001 |