JP7044222B2 - 実質的にコアギュローゲンを含まないリムルスアメボサイトライセートを使用してエンドトキシンを検出する方法 - Google Patents
実質的にコアギュローゲンを含まないリムルスアメボサイトライセートを使用してエンドトキシンを検出する方法 Download PDFInfo
- Publication number
- JP7044222B2 JP7044222B2 JP2019504062A JP2019504062A JP7044222B2 JP 7044222 B2 JP7044222 B2 JP 7044222B2 JP 2019504062 A JP2019504062 A JP 2019504062A JP 2019504062 A JP2019504062 A JP 2019504062A JP 7044222 B2 JP7044222 B2 JP 7044222B2
- Authority
- JP
- Japan
- Prior art keywords
- lal
- coagulogen
- endotoxin
- sample
- color
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active
Links
- 239000002158 endotoxin Substances 0.000 title claims description 110
- 108010045487 coagulogen Proteins 0.000 title claims description 92
- 241000239218 Limulus Species 0.000 title claims description 19
- 238000000034 method Methods 0.000 claims description 75
- 239000003593 chromogenic compound Substances 0.000 claims description 44
- 238000009295 crossflow filtration Methods 0.000 claims description 40
- 239000000523 sample Substances 0.000 claims description 40
- 238000003556 assay Methods 0.000 claims description 39
- 102000004169 proteins and genes Human genes 0.000 claims description 33
- 108090000623 proteins and genes Proteins 0.000 claims description 33
- 239000000758 substrate Substances 0.000 claims description 31
- 239000003153 chemical reaction reagent Substances 0.000 claims description 27
- 239000000203 mixture Substances 0.000 claims description 26
- 108090000765 processed proteins & peptides Proteins 0.000 claims description 19
- 230000035945 sensitivity Effects 0.000 claims description 18
- 238000002415 sodium dodecyl sulfate polyacrylamide gel electrophoresis Methods 0.000 claims description 18
- 239000000872 buffer Substances 0.000 claims description 16
- 230000008859 change Effects 0.000 claims description 14
- 230000000694 effects Effects 0.000 claims description 14
- 239000007788 liquid Substances 0.000 claims description 14
- 239000012528 membrane Substances 0.000 claims description 14
- 238000002835 absorbance Methods 0.000 claims description 13
- 238000010186 staining Methods 0.000 claims description 10
- 102000004196 processed proteins & peptides Human genes 0.000 claims description 9
- 239000012472 biological sample Substances 0.000 claims description 8
- 238000003776 cleavage reaction Methods 0.000 claims description 8
- 230000007017 scission Effects 0.000 claims description 8
- 229920001184 polypeptide Polymers 0.000 claims description 7
- 239000004695 Polyether sulfone Substances 0.000 claims description 6
- 238000006911 enzymatic reaction Methods 0.000 claims description 6
- 229920006393 polyether sulfone Polymers 0.000 claims description 6
- 239000000243 solution Substances 0.000 claims description 6
- 239000007983 Tris buffer Substances 0.000 claims description 5
- LENZDBCJOHFCAS-UHFFFAOYSA-N tris Chemical group OCC(N)(CO)CO LENZDBCJOHFCAS-UHFFFAOYSA-N 0.000 claims description 5
- 239000007987 MES buffer Substances 0.000 claims description 4
- 230000002255 enzymatic effect Effects 0.000 claims description 4
- 229920002492 poly(sulfone) Polymers 0.000 claims description 4
- 229920012266 Poly(ether sulfone) PES Polymers 0.000 claims description 2
- 239000000706 filtrate Substances 0.000 claims 1
- 229920006008 lipopolysaccharide Polymers 0.000 description 24
- 238000011161 development Methods 0.000 description 22
- 238000000926 separation method Methods 0.000 description 17
- 102000004190 Enzymes Human genes 0.000 description 15
- 108090000790 Enzymes Proteins 0.000 description 15
- 229940088598 enzyme Drugs 0.000 description 15
- 241000894006 Bacteria Species 0.000 description 14
- TYMLOMAKGOJONV-UHFFFAOYSA-N 4-nitroaniline Chemical group NC1=CC=C([N+]([O-])=O)C=C1 TYMLOMAKGOJONV-UHFFFAOYSA-N 0.000 description 13
- 238000006243 chemical reaction Methods 0.000 description 13
- 238000005345 coagulation Methods 0.000 description 12
- 238000001262 western blot Methods 0.000 description 11
- 230000015271 coagulation Effects 0.000 description 10
- GZQKNULLWNGMCW-PWQABINMSA-N lipid A (E. coli) Chemical compound O1[C@H](CO)[C@@H](OP(O)(O)=O)[C@H](OC(=O)C[C@@H](CCCCCCCCCCC)OC(=O)CCCCCCCCCCCCC)[C@@H](NC(=O)C[C@@H](CCCCCCCCCCC)OC(=O)CCCCCCCCCCC)[C@@H]1OC[C@@H]1[C@@H](O)[C@H](OC(=O)C[C@H](O)CCCCCCCCCCC)[C@@H](NC(=O)C[C@H](O)CCCCCCCCCCC)[C@@H](OP(O)(O)=O)O1 GZQKNULLWNGMCW-PWQABINMSA-N 0.000 description 10
- 102000010911 Enzyme Precursors Human genes 0.000 description 8
- 108010062466 Enzyme Precursors Proteins 0.000 description 8
- 238000002360 preparation method Methods 0.000 description 8
- 239000000047 product Substances 0.000 description 8
- 230000035484 reaction time Effects 0.000 description 8
- 230000001580 bacterial effect Effects 0.000 description 7
- 230000001225 therapeutic effect Effects 0.000 description 7
- 239000000427 antigen Substances 0.000 description 6
- 108091007433 antigens Proteins 0.000 description 6
- 102000036639 antigens Human genes 0.000 description 6
- 238000009472 formulation Methods 0.000 description 6
- 241001529572 Chaceon affinis Species 0.000 description 5
- 238000004458 analytical method Methods 0.000 description 5
- 210000000601 blood cell Anatomy 0.000 description 5
- 210000004027 cell Anatomy 0.000 description 5
- 238000004040 coloring Methods 0.000 description 5
- 239000012678 infectious agent Substances 0.000 description 5
- 208000015181 infectious disease Diseases 0.000 description 5
- 150000002632 lipids Chemical class 0.000 description 5
- 239000012466 permeate Substances 0.000 description 5
- 238000011050 LAL assay Methods 0.000 description 4
- 241001465754 Metazoa Species 0.000 description 4
- FAPWRFPIFSIZLT-UHFFFAOYSA-M Sodium chloride Chemical compound [Na+].[Cl-] FAPWRFPIFSIZLT-UHFFFAOYSA-M 0.000 description 4
- 102000002262 Thromboplastin Human genes 0.000 description 4
- 108010000499 Thromboplastin Proteins 0.000 description 4
- LOHQECMUTAPWAC-UHFFFAOYSA-N coagulin Natural products C1C(C)=C(CO)C(=O)OC1C1(C)C(C2(C)CCC3C4(C(=O)CC=CC4=CCC43)C)(O)CCC24O1 LOHQECMUTAPWAC-UHFFFAOYSA-N 0.000 description 4
- 238000001914 filtration Methods 0.000 description 4
- 239000012634 fragment Substances 0.000 description 4
- 150000004676 glycans Chemical class 0.000 description 4
- 210000000987 immune system Anatomy 0.000 description 4
- 238000001727 in vivo Methods 0.000 description 4
- 238000005259 measurement Methods 0.000 description 4
- 230000003389 potentiating effect Effects 0.000 description 4
- 238000011002 quantification Methods 0.000 description 4
- 102000003712 Complement factor B Human genes 0.000 description 3
- 108090000056 Complement factor B Proteins 0.000 description 3
- 102000004127 Cytokines Human genes 0.000 description 3
- 108090000695 Cytokines Proteins 0.000 description 3
- 241000588722 Escherichia Species 0.000 description 3
- 241000239220 Limulus polyphemus Species 0.000 description 3
- 241000607142 Salmonella Species 0.000 description 3
- 206010040070 Septic Shock Diseases 0.000 description 3
- HEMHJVSKTPXQMS-UHFFFAOYSA-M Sodium hydroxide Chemical compound [OH-].[Na+] HEMHJVSKTPXQMS-UHFFFAOYSA-M 0.000 description 3
- 210000001744 T-lymphocyte Anatomy 0.000 description 3
- 108060008682 Tumor Necrosis Factor Proteins 0.000 description 3
- 230000004913 activation Effects 0.000 description 3
- 230000017531 blood circulation Effects 0.000 description 3
- 210000002421 cell wall Anatomy 0.000 description 3
- 150000001875 compounds Chemical class 0.000 description 3
- 238000005516 engineering process Methods 0.000 description 3
- 238000002955 isolation Methods 0.000 description 3
- 238000012454 limulus amebocyte lysate test Methods 0.000 description 3
- 238000004519 manufacturing process Methods 0.000 description 3
- 229920001542 oligosaccharide Polymers 0.000 description 3
- 150000002482 oligosaccharides Chemical class 0.000 description 3
- 239000007800 oxidant agent Substances 0.000 description 3
- 229920001282 polysaccharide Polymers 0.000 description 3
- 239000005017 polysaccharide Substances 0.000 description 3
- 230000004044 response Effects 0.000 description 3
- 231100000419 toxicity Toxicity 0.000 description 3
- 230000001988 toxicity Effects 0.000 description 3
- 102000003390 tumor necrosis factor Human genes 0.000 description 3
- -1 vaccines Substances 0.000 description 3
- CWRWJDAEKWYUJT-CGKXPTHNSA-N 1-(9S,13S,12-oxophytodienoyl)-2-(7Z,10Z,13Z)-hexadecatrienoyl-3-(beta-D-galactosyl)-sn-glycerol Chemical compound C([C@H](OC(=O)CCCCC\C=C/C\C=C/C\C=C/CC)COC(=O)CCCCCCC[C@@H]1[C@@H](C(=O)C=C1)C\C=C/CC)O[C@@H]1O[C@H](CO)[C@H](O)[C@H](O)[C@H]1O CWRWJDAEKWYUJT-CGKXPTHNSA-N 0.000 description 2
- ZTOBILYWTYHOJB-WBCGDKOGSA-N 3',6'-bis[[(2s,3r,4s,5r,6r)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy]spiro[2-benzofuran-3,9'-xanthene]-1-one Chemical compound O[C@@H]1[C@@H](O)[C@@H](O)[C@@H](CO)O[C@H]1OC1=CC=C2C3(C4=CC=CC=C4C(=O)O3)C3=CC=C(O[C@H]4[C@@H]([C@@H](O)[C@@H](O)[C@@H](CO)O4)O)C=C3OC2=C1 ZTOBILYWTYHOJB-WBCGDKOGSA-N 0.000 description 2
- KZZWQCKYLNIOBT-UHFFFAOYSA-N 5-amino-2-nitrobenzoic acid Chemical compound NC1=CC=C([N+]([O-])=O)C(C(O)=O)=C1 KZZWQCKYLNIOBT-UHFFFAOYSA-N 0.000 description 2
- AKASINKCIWHQEP-UHFFFAOYSA-N 7-amino-4-methoxychromen-2-one Chemical compound C1=C(N)C=CC2=C1OC(=O)C=C2OC AKASINKCIWHQEP-UHFFFAOYSA-N 0.000 description 2
- PAYRUJLWNCNPSJ-UHFFFAOYSA-N Aniline Chemical compound NC1=CC=CC=C1 PAYRUJLWNCNPSJ-UHFFFAOYSA-N 0.000 description 2
- 231100000699 Bacterial toxin Toxicity 0.000 description 2
- 241000606125 Bacteroides Species 0.000 description 2
- 241000588923 Citrobacter Species 0.000 description 2
- 241000124008 Mammalia Species 0.000 description 2
- 208000001145 Metabolic Syndrome Diseases 0.000 description 2
- 206010037660 Pyrexia Diseases 0.000 description 2
- 206010040047 Sepsis Diseases 0.000 description 2
- 201000000690 abdominal obesity-metabolic syndrome Diseases 0.000 description 2
- 150000001413 amino acids Chemical class 0.000 description 2
- 239000003242 anti bacterial agent Substances 0.000 description 2
- 229940088710 antibiotic agent Drugs 0.000 description 2
- 239000006286 aqueous extract Substances 0.000 description 2
- 239000007864 aqueous solution Substances 0.000 description 2
- 239000000688 bacterial toxin Substances 0.000 description 2
- 229960000074 biopharmaceutical Drugs 0.000 description 2
- 230000015572 biosynthetic process Effects 0.000 description 2
- 210000000748 cardiovascular system Anatomy 0.000 description 2
- 210000001175 cerebrospinal fluid Anatomy 0.000 description 2
- 235000009508 confectionery Nutrition 0.000 description 2
- GLNDAGDHSLMOKX-UHFFFAOYSA-N coumarin 120 Chemical compound C1=C(N)C=CC2=C1OC(=O)C=C2C GLNDAGDHSLMOKX-UHFFFAOYSA-N 0.000 description 2
- 230000009089 cytolysis Effects 0.000 description 2
- 230000034994 death Effects 0.000 description 2
- 235000014113 dietary fatty acids Nutrition 0.000 description 2
- 238000010790 dilution Methods 0.000 description 2
- 239000012895 dilution Substances 0.000 description 2
- DEKPYXUDJRABNK-UHFFFAOYSA-N dimethyl 5-aminobenzene-1,3-dicarboxylate Chemical compound COC(=O)C1=CC(N)=CC(C(=O)OC)=C1 DEKPYXUDJRABNK-UHFFFAOYSA-N 0.000 description 2
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 2
- 230000007893 endotoxin activity Effects 0.000 description 2
- 239000002095 exotoxin Substances 0.000 description 2
- 231100000776 exotoxin Toxicity 0.000 description 2
- 229930195729 fatty acid Natural products 0.000 description 2
- 239000000194 fatty acid Substances 0.000 description 2
- 150000004665 fatty acids Chemical class 0.000 description 2
- 238000004108 freeze drying Methods 0.000 description 2
- QRMZSPFSDQBLIX-UHFFFAOYSA-N homovanillic acid Chemical compound COC1=CC(CC(O)=O)=CC=C1O QRMZSPFSDQBLIX-UHFFFAOYSA-N 0.000 description 2
- WQYVRQLZKVEZGA-UHFFFAOYSA-N hypochlorite Chemical compound Cl[O-] WQYVRQLZKVEZGA-UHFFFAOYSA-N 0.000 description 2
- 230000002757 inflammatory effect Effects 0.000 description 2
- 230000004054 inflammatory process Effects 0.000 description 2
- 239000006166 lysate Substances 0.000 description 2
- 238000012986 modification Methods 0.000 description 2
- 230000004048 modification Effects 0.000 description 2
- 239000006225 natural substrate Substances 0.000 description 2
- 102000039446 nucleic acids Human genes 0.000 description 2
- 108020004707 nucleic acids Proteins 0.000 description 2
- 150000007523 nucleic acids Chemical class 0.000 description 2
- 230000003287 optical effect Effects 0.000 description 2
- 239000006201 parenteral dosage form Substances 0.000 description 2
- 108010005636 polypeptide C Proteins 0.000 description 2
- 230000008569 process Effects 0.000 description 2
- 238000000734 protein sequencing Methods 0.000 description 2
- 238000000746 purification Methods 0.000 description 2
- 239000002510 pyrogen Substances 0.000 description 2
- 239000002994 raw material Substances 0.000 description 2
- 230000009467 reduction Effects 0.000 description 2
- 230000036303 septic shock Effects 0.000 description 2
- 239000011780 sodium chloride Substances 0.000 description 2
- 239000007787 solid Substances 0.000 description 2
- 239000000126 substance Substances 0.000 description 2
- 235000000346 sugar Nutrition 0.000 description 2
- 230000009885 systemic effect Effects 0.000 description 2
- 238000012360 testing method Methods 0.000 description 2
- 229960005486 vaccine Drugs 0.000 description 2
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Chemical compound O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 2
- NNLZBVFSCVTSLA-XMABDTGBSA-N (4r,5r,6r)-6-[(1r)-1,2-dihydroxyethyl]-2,4,5-trihydroxyoxane-2-carboxylic acid Chemical compound OC[C@@H](O)[C@H]1OC(O)(C(O)=O)C[C@@H](O)[C@H]1O NNLZBVFSCVTSLA-XMABDTGBSA-N 0.000 description 1
- TZMSYXZUNZXBOL-UHFFFAOYSA-N 10H-phenoxazine Chemical compound C1=CC=C2NC3=CC=CC=C3OC2=C1 TZMSYXZUNZXBOL-UHFFFAOYSA-N 0.000 description 1
- OTLLEIBWKHEHGU-UHFFFAOYSA-N 2-[5-[[5-(6-aminopurin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy]-3,4-dihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy-3,5-dihydroxy-4-phosphonooxyhexanedioic acid Chemical compound C1=NC=2C(N)=NC=NC=2N1C(C(C1O)O)OC1COC1C(CO)OC(OC(C(O)C(OP(O)(O)=O)C(O)C(O)=O)C(O)=O)C(O)C1O OTLLEIBWKHEHGU-UHFFFAOYSA-N 0.000 description 1
- UJVBZCCNLAAMOV-UHFFFAOYSA-N 2h-1,2-benzothiazine Chemical compound C1=CC=C2C=CNSC2=C1 UJVBZCCNLAAMOV-UHFFFAOYSA-N 0.000 description 1
- VOUAQYXWVJDEQY-QENPJCQMSA-N 33017-11-7 Chemical compound OC(=O)CC[C@H](N)C(=O)N[C@@H](C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(=O)NCC(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](C(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)N[C@@H](CC(C)C)C(=O)NCC(=O)NCC(=O)NCC(=O)N1CCC[C@H]1C(=O)NCC(=O)N[C@@H](C)C(=O)NCC(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(=O)N1[C@H](C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(O)=O)C(=O)NCC(=O)N[C@@H](CO)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC(N)=O)C(O)=O)CCC1 VOUAQYXWVJDEQY-QENPJCQMSA-N 0.000 description 1
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 description 1
- PSGQCCSGKGJLRL-UHFFFAOYSA-N 4-methyl-2h-chromen-2-one Chemical compound C1=CC=CC2=C1OC(=O)C=C2C PSGQCCSGKGJLRL-UHFFFAOYSA-N 0.000 description 1
- 241000607534 Aeromonas Species 0.000 description 1
- 201000001320 Atherosclerosis Diseases 0.000 description 1
- 241000588807 Bordetella Species 0.000 description 1
- 241000589562 Brucella Species 0.000 description 1
- 108010075254 C-Peptide Proteins 0.000 description 1
- 241000589876 Campylobacter Species 0.000 description 1
- 101710140962 Capsid scaffolding protein Proteins 0.000 description 1
- 241000606161 Chlamydia Species 0.000 description 1
- 208000011231 Crohn disease Diseases 0.000 description 1
- 206010012735 Diarrhoea Diseases 0.000 description 1
- 241000588921 Enterobacteriaceae Species 0.000 description 1
- 241000588724 Escherichia coli Species 0.000 description 1
- 241000589601 Francisella Species 0.000 description 1
- 241000606790 Haemophilus Species 0.000 description 1
- 241000589989 Helicobacter Species 0.000 description 1
- 241000282412 Homo Species 0.000 description 1
- 206010061218 Inflammation Diseases 0.000 description 1
- 206010022489 Insulin Resistance Diseases 0.000 description 1
- 102000000589 Interleukin-1 Human genes 0.000 description 1
- 108010002352 Interleukin-1 Proteins 0.000 description 1
- 102000015696 Interleukins Human genes 0.000 description 1
- 108010063738 Interleukins Proteins 0.000 description 1
- 241000588748 Klebsiella Species 0.000 description 1
- 239000012480 LAL reagent Substances 0.000 description 1
- 241000589248 Legionella Species 0.000 description 1
- 208000007764 Legionnaires' Disease Diseases 0.000 description 1
- 241000239205 Merostomata Species 0.000 description 1
- 208000034486 Multi-organ failure Diseases 0.000 description 1
- 208000010718 Multiple Organ Failure Diseases 0.000 description 1
- 102000016943 Muramidase Human genes 0.000 description 1
- 108010014251 Muramidase Proteins 0.000 description 1
- 241000699670 Mus sp. Species 0.000 description 1
- 108010062010 N-Acetylmuramoyl-L-alanine Amidase Proteins 0.000 description 1
- 241000588653 Neisseria Species 0.000 description 1
- 241000588656 Neisseriaceae Species 0.000 description 1
- 208000008589 Obesity Diseases 0.000 description 1
- 206010057249 Phagocytosis Diseases 0.000 description 1
- 241000947836 Pseudomonadaceae Species 0.000 description 1
- 241000589516 Pseudomonas Species 0.000 description 1
- 102000012479 Serine Proteases Human genes 0.000 description 1
- 108010022999 Serine Proteases Proteins 0.000 description 1
- 241000607768 Shigella Species 0.000 description 1
- OUUQCZGPVNCOIJ-UHFFFAOYSA-M Superoxide Chemical compound [O-][O] OUUQCZGPVNCOIJ-UHFFFAOYSA-M 0.000 description 1
- 241001430183 Veillonellaceae Species 0.000 description 1
- 241000607598 Vibrio Species 0.000 description 1
- 241000607493 Vibrionaceae Species 0.000 description 1
- 239000002250 absorbent Substances 0.000 description 1
- 230000002745 absorbent Effects 0.000 description 1
- 238000000862 absorption spectrum Methods 0.000 description 1
- 239000002253 acid Substances 0.000 description 1
- 230000003213 activating effect Effects 0.000 description 1
- 230000006978 adaptation Effects 0.000 description 1
- 230000005875 antibody response Effects 0.000 description 1
- 238000003149 assay kit Methods 0.000 description 1
- 210000003719 b-lymphocyte Anatomy 0.000 description 1
- NFIKTSWJXJTUBZ-ZEQRLZLVSA-N benzyl n-[2-[(2s)-2-[[(2s)-5-(diaminomethylideneamino)-1-[(4-methyl-2-oxochromen-7-yl)amino]-1-oxopentan-2-yl]carbamoyl]pyrrolidin-1-yl]-2-oxoethyl]carbamate Chemical compound C([C@H]1C(=O)N[C@@H](CCCN=C(N)N)C(=O)NC2=CC=3OC(=O)C=C(C=3C=C2)C)CCN1C(=O)CNC(=O)OCC1=CC=CC=C1 NFIKTSWJXJTUBZ-ZEQRLZLVSA-N 0.000 description 1
- 125000001584 benzyloxycarbonyl group Chemical group C(=O)(OCC1=CC=CC=C1)* 0.000 description 1
- 230000004071 biological effect Effects 0.000 description 1
- 210000004369 blood Anatomy 0.000 description 1
- 239000008280 blood Substances 0.000 description 1
- 238000009835 boiling Methods 0.000 description 1
- 239000007853 buffer solution Substances 0.000 description 1
- 150000001720 carbohydrates Chemical group 0.000 description 1
- ZAIPMKNFIOOWCQ-UEKVPHQBSA-N cephalexin Chemical compound C1([C@@H](N)C(=O)N[C@H]2[C@@H]3N(C2=O)C(=C(CS3)C)C(O)=O)=CC=CC=C1 ZAIPMKNFIOOWCQ-UEKVPHQBSA-N 0.000 description 1
- 230000035605 chemotaxis Effects 0.000 description 1
- 238000007820 coagulation assay Methods 0.000 description 1
- 230000000295 complement effect Effects 0.000 description 1
- 230000004154 complement system Effects 0.000 description 1
- 238000010924 continuous production Methods 0.000 description 1
- 239000002537 cosmetic Substances 0.000 description 1
- 150000004775 coumarins Chemical class 0.000 description 1
- 230000007123 defense Effects 0.000 description 1
- 230000001419 dependent effect Effects 0.000 description 1
- 238000001514 detection method Methods 0.000 description 1
- 206010012601 diabetes mellitus Diseases 0.000 description 1
- 230000029087 digestion Effects 0.000 description 1
- 238000007865 diluting Methods 0.000 description 1
- 208000035475 disorder Diseases 0.000 description 1
- 229940079593 drug Drugs 0.000 description 1
- 239000003814 drug Substances 0.000 description 1
- 238000007905 drug manufacturing Methods 0.000 description 1
- 108010072542 endotoxin binding proteins Proteins 0.000 description 1
- 238000001952 enzyme assay Methods 0.000 description 1
- 208000010706 fatty liver disease Diseases 0.000 description 1
- 239000012530 fluid Substances 0.000 description 1
- GNBHRKFJIUUOQI-UHFFFAOYSA-N fluorescein Chemical compound O1C(=O)C2=CC=CC=C2C21C1=CC=C(O)C=C1OC1=CC(O)=CC=C21 GNBHRKFJIUUOQI-UHFFFAOYSA-N 0.000 description 1
- 235000013305 food Nutrition 0.000 description 1
- 210000001035 gastrointestinal tract Anatomy 0.000 description 1
- 230000009931 harmful effect Effects 0.000 description 1
- 150000002386 heptoses Chemical class 0.000 description 1
- 239000012510 hollow fiber Substances 0.000 description 1
- 230000003100 immobilizing effect Effects 0.000 description 1
- 230000028993 immune response Effects 0.000 description 1
- 230000005847 immunogenicity Effects 0.000 description 1
- 238000000338 in vitro Methods 0.000 description 1
- 238000011534 incubation Methods 0.000 description 1
- 230000028709 inflammatory response Effects 0.000 description 1
- 230000015788 innate immune response Effects 0.000 description 1
- 230000003993 interaction Effects 0.000 description 1
- 229940047122 interleukins Drugs 0.000 description 1
- 210000000936 intestine Anatomy 0.000 description 1
- 238000011835 investigation Methods 0.000 description 1
- 230000000366 juvenile effect Effects 0.000 description 1
- KYQCXUMVJGMDNG-SHUUEZRQSA-N keto-3-deoxy-D-manno-octulosonic acid Chemical compound OC[C@@H](O)[C@@H](O)[C@H](O)[C@H](O)CC(=O)C(O)=O KYQCXUMVJGMDNG-SHUUEZRQSA-N 0.000 description 1
- 150000002605 large molecules Chemical class 0.000 description 1
- IZWSFJTYBVKZNK-UHFFFAOYSA-N lauryl sulfobetaine Chemical compound CCCCCCCCCCCC[N+](C)(C)CCCS([O-])(=O)=O IZWSFJTYBVKZNK-UHFFFAOYSA-N 0.000 description 1
- 210000002751 lymph Anatomy 0.000 description 1
- 229960000274 lysozyme Drugs 0.000 description 1
- 239000004325 lysozyme Substances 0.000 description 1
- 235000010335 lysozyme Nutrition 0.000 description 1
- 229920002521 macromolecule Polymers 0.000 description 1
- 210000002540 macrophage Anatomy 0.000 description 1
- 230000014759 maintenance of location Effects 0.000 description 1
- 239000000463 material Substances 0.000 description 1
- 230000000813 microbial effect Effects 0.000 description 1
- 244000005700 microbiome Species 0.000 description 1
- 230000005012 migration Effects 0.000 description 1
- 238000013508 migration Methods 0.000 description 1
- 239000000178 monomer Substances 0.000 description 1
- 208000029744 multiple organ dysfunction syndrome Diseases 0.000 description 1
- 230000003472 neutralizing effect Effects 0.000 description 1
- 210000000440 neutrophil Anatomy 0.000 description 1
- 235000020824 obesity Nutrition 0.000 description 1
- 238000005457 optimization Methods 0.000 description 1
- 230000000242 pagocytic effect Effects 0.000 description 1
- 230000001717 pathogenic effect Effects 0.000 description 1
- 230000037361 pathway Effects 0.000 description 1
- 150000002978 peroxides Chemical class 0.000 description 1
- 230000008782 phagocytosis Effects 0.000 description 1
- UYWQUFXKFGHYNT-UHFFFAOYSA-N phenylmethyl ester of formic acid Natural products O=COCC1=CC=CC=C1 UYWQUFXKFGHYNT-UHFFFAOYSA-N 0.000 description 1
- 150000003904 phospholipids Chemical class 0.000 description 1
- 229920000642 polymer Polymers 0.000 description 1
- 230000002797 proteolythic effect Effects 0.000 description 1
- 238000003908 quality control method Methods 0.000 description 1
- 238000011158 quantitative evaluation Methods 0.000 description 1
- 230000009257 reactivity Effects 0.000 description 1
- 230000003252 repetitive effect Effects 0.000 description 1
- 238000011160 research Methods 0.000 description 1
- 230000000717 retained effect Effects 0.000 description 1
- 230000028327 secretion Effects 0.000 description 1
- 238000011896 sensitive detection Methods 0.000 description 1
- 238000012163 sequencing technique Methods 0.000 description 1
- 230000035939 shock Effects 0.000 description 1
- 238000002798 spectrophotometry method Methods 0.000 description 1
- 238000001228 spectrum Methods 0.000 description 1
- 239000008223 sterile water Substances 0.000 description 1
- 238000003860 storage Methods 0.000 description 1
- 125000001424 substituent group Chemical group 0.000 description 1
- 150000008163 sugars Chemical class 0.000 description 1
- 230000008718 systemic inflammatory response Effects 0.000 description 1
- 230000008280 toxic mechanism Effects 0.000 description 1
- 239000003053 toxin Substances 0.000 description 1
- 231100000765 toxin Toxicity 0.000 description 1
- 108700012359 toxins Proteins 0.000 description 1
- 230000001960 triggered effect Effects 0.000 description 1
- 238000004879 turbidimetry Methods 0.000 description 1
- 208000001072 type 2 diabetes mellitus Diseases 0.000 description 1
- 239000002435 venom Substances 0.000 description 1
- 231100000611 venom Toxicity 0.000 description 1
- 210000001048 venom Anatomy 0.000 description 1
- 238000001429 visible spectrum Methods 0.000 description 1
Images
Classifications
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N33/00—Investigating or analysing materials by specific methods not covered by groups G01N1/00 - G01N31/00
- G01N33/48—Biological material, e.g. blood, urine; Haemocytometers
- G01N33/50—Chemical analysis of biological material, e.g. blood, urine; Testing involving biospecific ligand binding methods; Immunological testing
- G01N33/579—Chemical analysis of biological material, e.g. blood, urine; Testing involving biospecific ligand binding methods; Immunological testing involving limulus lysate
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K1/00—General methods for the preparation of peptides, i.e. processes for the organic chemical preparation of peptides or proteins of any length
- C07K1/14—Extraction; Separation; Purification
- C07K1/34—Extraction; Separation; Purification by filtration, ultrafiltration or reverse osmosis
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/34—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving hydrolase
- C12Q1/37—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving hydrolase involving peptidase or proteinase
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2337/00—N-linked chromogens for determinations of peptidases and proteinases
- C12Q2337/10—Anilides
- C12Q2337/12—Para-Nitroanilides p-NA
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2400/00—Assays, e.g. immunoassays or enzyme assays, involving carbohydrates
- G01N2400/10—Polysaccharides, i.e. having more than five saccharide radicals attached to each other by glycosidic linkages; Derivatives thereof, e.g. ethers, esters
- G01N2400/50—Lipopolysaccharides; LPS
Landscapes
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Health & Medical Sciences (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Molecular Biology (AREA)
- Immunology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- Analytical Chemistry (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Zoology (AREA)
- Physics & Mathematics (AREA)
- Urology & Nephrology (AREA)
- Hematology (AREA)
- Wood Science & Technology (AREA)
- Biomedical Technology (AREA)
- Genetics & Genomics (AREA)
- Biophysics (AREA)
- Medicinal Chemistry (AREA)
- Pathology (AREA)
- General Physics & Mathematics (AREA)
- Cell Biology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- Food Science & Technology (AREA)
- Water Supply & Treatment (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
- Investigating Or Analysing Materials By The Use Of Chemical Reactions (AREA)
- Investigating Or Analysing Biological Materials (AREA)
Description
基質+H2O+酵素→ペプチド+pNA(黄色)
[0049]いくつかの実施形態において、グラム陰性菌エンドトキシンは、LAL中の酵素前駆体の活性化を間接的に触媒することが可能である。活性化の最初の速度は、存在するエンドトキシンの濃度によって決まる可能性がある。
タンジェンシャルフローろ過を使用したLALの精製
[0070]カブトガニの血液細胞(アメボサイト)から抽出したリムルスアメボサイトライセート(LAL)をロンザ(バーゼル、スイス)から入手した。コアギュローゲンを除去するために、タンジェンシャルフローろ過(TFF)用のホローファイバー修飾ポリエーテルスルホン(mPES)メンブランフィルター(スペクトラムラボラトリーズ(SpectrumLabs))にLALを供した。30kDa MWCO mPESフィルター、コールパーマー(Cole Parmer)マスターフレックスポンプチュービング(Masterflex Pump Tubing)並びに適したポンプ及び背圧弁を使用してTFFシステムをセットアップした。そのシステムをLAL試薬水(LRW)で洗い流し、その後、1N NaOHを使用して室温で1時間脱パイロジェン処理をした。脱パイロジェン処理後、LRWを使用して、システムをすすぎ、その後、そのシステムを、TFFバッファーを使用して平衡化した。
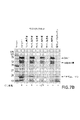
コアギュローゲンの定量
[0074]LAL保持液に対するSDS-PAGE/タンパク質ゲル染色を実施し、元のLAL中の20kDコアギュローゲンバンドのバンド密度を、あれば、TFF保持液中のコアギュローゲンバンドのバンド密度と比較することによって、LAL中の残存コアギュローゲンの半定量的評価を行った。例えば、図7A参照。
実質的にコアギュローゲンを含まないLALの安定性
[0079]実質的にコアギュローゲンを含まないLALの安定性を調査した。実質的にコアギュローゲンを含まないLALサンプルを1、2、3、4、5、6、7、8、9及び10週間保管した。示した期間の終わりに、実質的にコアギュローゲンを含まないLALサンプルを96ウェルプレート中のLAL発色試薬と混合し、405nmにおける吸光度を測定するインキュベートプレートリーダーに入れる。その反応を黄色の外観により経時的に自動的に監視した。
コアギュローゲン除去は分離を改善する
[0081]複数のオンセットmODにおける分離に対して、LALからコアギュローゲンを除去することが有する効果の調査を行った。通常のLAL対実質的にコアギュローゲンを含まないLALの比較は、実質的にコアギュローゲンを含まないLALを使用した場合に、それぞれのオンセットmODにおける分離の増加を示した。分離を増加させるために実質的にコアギュローゲンを含まないLALを使用することによって、アッセイの目標の感度を、より低いオンセットmOD設定においてより短い時間で達成することができる。配合物を調製するために実質的にコアギュローゲンを含まないLALを使用すると、0.005EU/mlの目標の感度を、368秒の分離で26分で達成できる。通常のLALを使用すると、0.005EU/mlの感度を50mODにおいて32分で達成するが、分離はわずか67秒である。
LAL及び実質的にコアギュローゲンを含まないLAL配合物の発色検出
[0083]LAL及び実質的にコアギュローゲンを含まないLALの両方のサンプルを、0.005EU/ml、0.05EU/ml、0.5EU/ml、5EU/ml及び50EU/mlの標準に対してLAL調製物中20%、30%、40%及び50%の濃度で試験した。さらに、さまざまな濃度のズイッタージェントも試験した。図3及び図4。このデータは、実質的にコアギュローゲンを含まないLAL及びズイッタージェントの濃度を高めると、LAL及び実質的にコアギュローゲンを含まないLAL配合物の両方に関してそれぞれ標準に対する反応速度が増加したことを示唆している。但し、LAL配合物により反応速度が増加するため、ブランクと最低の標準(0.005EU/ml)との間では分離が失われる一方で、すべての実質的にコアギュローゲンを含まないLAL配合物に関しては、少なくとも200秒の分離が維持されている。増加したズイッタージェントと配合した実質的にコアギュローゲンを含まないLALは、許容される分離で0.001EU/mlの感度を達成した。(データは示していない)。
実質的にコアギュローゲンを含まないLALと発色基質の比率
[0084]実質的にコアギュローゲンを含まないLALと発色基質の最適な比率を200mOD及び50mODにおいて調査した。図5A及び図5B。このデータは、70%のLAL調製物と30%の発色基質の比率が最速の反応時間になったことを示している。実質的にコアギュローゲンを含まないLALの3つの異なるロットを見ると、この観察結果が確認された。図6A。
本出願は、ASCII形式で電子的に提出された配列表を含有し、参照によりその全体が本明細書に組み込まれる。2017年9月5日に作成された上記ASCIIコピーは、0132-0023WO1_SL.txtという名前であり、1,055バイトのサイズである。
Claims (20)
- 発色アッセイを使用して、サンプル中のエンドトキシンを検出する方法であって、前記方法が、
a.前記サンプルをリムルスアメボサイトライセート(LAL)及び発色基質を含む試薬と接触させるステップと、
b.前記サンプル中のエンドトキシンの存在下における前記発色基質の変化によって生じる発色効果を測定するステップとを含み、
前記LALが、タンパク質染色を伴うSDS-PAGEにより測定される場合に、LAL中の総タンパク質に対して5%(wt/wt)未満のコアギュローゲンを含むものであり、
前記方法は、0.001EU/mL又は0.001EU/mL未満の感度を有する、方法。 - 前記発色基質の変化が、酵素反応により起こり、前記酵素反応が、ポリペプチドからの発色団の切断である、請求項1に記載の方法。
- 前記試薬が、水性液体である、請求項1又は2に記載の方法。
- 前記LAL、発色基質又はその両方が、凍結乾燥され、その後、前記サンプルと接触させるステップに先立って再構成される、請求項1~3のいずれか一項に記載の方法。
- 前記サンプルが、非経口剤形、ワクチン、抗生剤、治療用タンパク質、治療用核酸、治療用抗体、及びバイオテクノロジー製品からなる群から選択される、請求項1~4のいずれか一項に記載の方法。
- 前記LALが、SDS-PAGE及びタンパク質染色によって測定される場合に、前記LAL中の総タンパク質に対して2%(wt/wt)未満のコアギュローゲンを含む、請求項1~5のいずれか一項に記載の方法。
- 前記LALが、タンパク質染色を伴うSDS-PAGEによって測定される場合に、前記LAL中の総タンパク質に対して0.5%(wt/wt)未満のコアギュローゲンを含む、請求項1~6のいずれか一項に記載の方法。
- 発色アッセイを使用して、生物学的サンプル中のエンドトキシンを検出する方法であって、前記方法が、
a.前記生物学的サンプルをリムルスアメボサイトライセート(LAL)及びAc-Ile-Glu-Ala-Arg-pNA(配列番号:1)を含む水性試薬と接触させるステップと、
b.前記サンプル中のエンドトキシンの存在下におけるAc-Ile-Glu-Ala-Arg-pNA(配列番号:1)からのpNAの酵素切断によって生じる405nmにおける吸光度の変化を測定するステップとを含み、
前記LALが、タンパク質染色を伴うSDS-PAGEにより測定される場合に、LAL中の総タンパク質に対して5%(wt/wt)未満のコアギュローゲンを含むものであり、
前記方法は、0.001EU/mL又は0.001EU/mL未満の感度を有する、方法。 - 前記発色基質が、Ac-Ile-Glu-Ala-Arg-pNA(配列番号:1)である、請求項1~7のいずれか一項に記載の方法。
- タンパク質染色を伴うSDS-PAGEにより測定される場合に、LAL中の総タンパク質に対して5%(wt/wt)未満のコアギュローゲンを含むリムルスアメボサイトライセート(LAL)を生成する方法であって、前記方法が、
コアギュローゲンを含むLAL及びバッファーを含む組成物を20kDa~50kDaフィルターを使用したタンジェンシャルフローろ過(TFF)に供するステップであって、前記TFFが保持液及びろ液を生成するステップと、
前記保持液を回収し、それによりLAL中の総タンパク質に対して5%(wt/wt)未満のコアギュローゲンを含むLALを得るステップと
を含む、方法。 - 前記TFFが、350ml/分~600ml/分の流量で実施される、請求項10に記載の方法。
- 前記TFFが、350ml/分~500ml/分の流量で実施される、請求項11に記載の方法。
- LALを含む前記組成物が、バッファーである、請求項10~12のいずれか一項に記載の方法。
- 前記バッファーがトリスバッファー又はMESバッファーである、請求項13に記載の方法。
- 前記バッファーが6.0~9.0のpHを有する、請求項13又は14に記載の方法。
- 前記バッファーが7.0~8.0のpHを有する、請求項15に記載の方法。
- 前記TFFフィルターが、修飾ポリエーテルスルホン(mPES)、ポリスルホン(PS)及びポリエーテルスルホン(PES)から選択される膜を含む、請求項10~16のいずれか一項に記載の方法。
- 前記TFFフィルターが、修飾ポリエーテルスルホン(mPES)メンブランフィルターである、請求項16又は17に記載の方法。
- 前記LALが、タンパク質染色を伴うSDS-PAGEによって測定される場合に、前記LAL中の総タンパク質に対して2%(wt/wt)未満のコアギュローゲンを含む、請求項10~18のいずれか一項に記載の方法。
- 前記LALが、タンパク質染色を伴うSDS-PAGEによって測定される場合に、前記LAL中の総タンパク質に対して0.5%(wt/wt)未満のコアギュローゲンを含む、請求項10~19のいずれか一項に記載の方法。
Applications Claiming Priority (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US201662370493P | 2016-08-03 | 2016-08-03 | |
| US62/370,493 | 2016-08-03 | ||
| PCT/US2017/045137 WO2018026941A1 (en) | 2016-08-03 | 2017-08-02 | Method of detecting an endotoxin using limulus amebocyte lysate substantially free of coagulogen |
Publications (3)
| Publication Number | Publication Date |
|---|---|
| JP2019526052A JP2019526052A (ja) | 2019-09-12 |
| JP2019526052A5 JP2019526052A5 (ja) | 2020-09-10 |
| JP7044222B2 true JP7044222B2 (ja) | 2022-03-30 |
Family
ID=61069187
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| JP2019504062A Active JP7044222B2 (ja) | 2016-08-03 | 2017-08-02 | 実質的にコアギュローゲンを含まないリムルスアメボサイトライセートを使用してエンドトキシンを検出する方法 |
Country Status (8)
| Country | Link |
|---|---|
| US (2) | US11156614B2 (ja) |
| EP (2) | EP3943608B1 (ja) |
| JP (1) | JP7044222B2 (ja) |
| KR (1) | KR102561368B1 (ja) |
| CN (1) | CN109790561B (ja) |
| IL (1) | IL264464B2 (ja) |
| SG (1) | SG11201900767VA (ja) |
| WO (1) | WO2018026941A1 (ja) |
Families Citing this family (16)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US9458450B2 (en) | 2012-03-15 | 2016-10-04 | Flodesign Sonics, Inc. | Acoustophoretic separation technology using multi-dimensional standing waves |
| US10967298B2 (en) | 2012-03-15 | 2021-04-06 | Flodesign Sonics, Inc. | Driver and control for variable impedence load |
| US10704021B2 (en) | 2012-03-15 | 2020-07-07 | Flodesign Sonics, Inc. | Acoustic perfusion devices |
| US9950282B2 (en) | 2012-03-15 | 2018-04-24 | Flodesign Sonics, Inc. | Electronic configuration and control for acoustic standing wave generation |
| WO2015105955A1 (en) | 2014-01-08 | 2015-07-16 | Flodesign Sonics, Inc. | Acoustophoresis device with dual acoustophoretic chamber |
| US11021699B2 (en) | 2015-04-29 | 2021-06-01 | FioDesign Sonics, Inc. | Separation using angled acoustic waves |
| US11708572B2 (en) | 2015-04-29 | 2023-07-25 | Flodesign Sonics, Inc. | Acoustic cell separation techniques and processes |
| US11377651B2 (en) | 2016-10-19 | 2022-07-05 | Flodesign Sonics, Inc. | Cell therapy processes utilizing acoustophoresis |
| US11474085B2 (en) | 2015-07-28 | 2022-10-18 | Flodesign Sonics, Inc. | Expanded bed affinity selection |
| US11459540B2 (en) | 2015-07-28 | 2022-10-04 | Flodesign Sonics, Inc. | Expanded bed affinity selection |
| US11085035B2 (en) | 2016-05-03 | 2021-08-10 | Flodesign Sonics, Inc. | Therapeutic cell washing, concentration, and separation utilizing acoustophoresis |
| US11214789B2 (en) | 2016-05-03 | 2022-01-04 | Flodesign Sonics, Inc. | Concentration and washing of particles with acoustics |
| SG11201900767VA (en) | 2016-08-03 | 2019-02-27 | Lonza Walkersville Inc | Method of detecting an endotoxin using limulus amebocyte lysate substantially free of coagulogen |
| EP3529347A1 (en) | 2016-10-19 | 2019-08-28 | Flodesign Sonics, Inc. | Affinity cell extraction by acoustics |
| CN110192110B (zh) * | 2017-01-11 | 2023-01-10 | 隆萨沃克斯维尔股份有限公司 | 无凝固蛋白原的澄清的鲎阿米巴样细胞裂解物 |
| KR102439221B1 (ko) | 2017-12-14 | 2022-09-01 | 프로디자인 소닉스, 인크. | 음향 트랜스듀서 구동기 및 제어기 |
Citations (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2014014375A (ja) | 2007-11-12 | 2014-01-30 | Hiroshima Univ | エンドトキシンの濃度測定方法および濃度測定用キット |
Family Cites Families (17)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4322217A (en) | 1980-06-27 | 1982-03-30 | Mallinckrodt, Inc. | Process for preparing Limulus lysate |
| DK0426395T3 (ja) * | 1989-10-30 | 1997-03-03 | Biowhittaker Inc | |
| JP3122454B2 (ja) | 1990-09-27 | 2001-01-09 | 生化学工業株式会社 | カブトガニ・アメボサイト・ライセートの調製法 |
| US5208148A (en) | 1990-12-07 | 1993-05-04 | Molecular Probes, Inc. | Lipophilic fluorescent glycosidase substrates |
| US5576424A (en) | 1991-08-23 | 1996-11-19 | Molecular Probes, Inc. | Haloalkyl derivatives of reporter molecules used to analyze metabolic activity in cells |
| DE69229174T2 (de) | 1991-08-23 | 1999-09-16 | Molecular Probes Inc | Verwendung von haloalkylderivaten von reportermolekülen zur analyse der metabolischen aktivität in zellen |
| US5242805A (en) | 1991-08-23 | 1993-09-07 | Molecular Probes, Inc. | Long wavelength lipophilic fluorogenic glycosidase substrates |
| JP3242733B2 (ja) | 1993-02-26 | 2001-12-25 | 生化学工業株式会社 | エンドトキシン特異的測定剤 |
| US5830912A (en) | 1996-11-15 | 1998-11-03 | Molecular Probes, Inc. | Derivatives of 6,8-difluoro-7-hydroxycoumarin |
| US5773236A (en) | 1997-04-25 | 1998-06-30 | Molecule Probes, Inc. | Assay for glutathiane transferase using polyhaloaryl-substituted reporter molecules |
| WO2008114149A2 (en) | 2007-03-21 | 2008-09-25 | Id Biomedical Corporation Of Quebec | Chimeric antigens |
| US7846678B2 (en) | 2008-08-18 | 2010-12-07 | BioDtech, Inc. | Enhancing endotoxin detection |
| MY155541A (en) | 2010-10-29 | 2015-10-30 | Universiti Malaysia Terengganu | Process for the preparation of amoebocyte lysate from the haemolymph of the horseshoe crab |
| CN102305788A (zh) | 2010-11-22 | 2012-01-04 | 天津市一瑞生物工程有限公司 | 革兰阴性菌脂多糖(内毒素)比色检测试剂盒的制备及其使用方法 |
| SG11201900767VA (en) | 2016-08-03 | 2019-02-27 | Lonza Walkersville Inc | Method of detecting an endotoxin using limulus amebocyte lysate substantially free of coagulogen |
| CN110192110B (zh) * | 2017-01-11 | 2023-01-10 | 隆萨沃克斯维尔股份有限公司 | 无凝固蛋白原的澄清的鲎阿米巴样细胞裂解物 |
| US20190001330A1 (en) * | 2017-06-28 | 2019-01-03 | Lonza Walkersville, Inc. | Cartridge for endotoxin detection |
-
2017
- 2017-08-02 SG SG11201900767VA patent/SG11201900767VA/en unknown
- 2017-08-02 EP EP21183657.2A patent/EP3943608B1/en active Active
- 2017-08-02 JP JP2019504062A patent/JP7044222B2/ja active Active
- 2017-08-02 KR KR1020197005917A patent/KR102561368B1/ko active IP Right Grant
- 2017-08-02 CN CN201780048853.4A patent/CN109790561B/zh active Active
- 2017-08-02 EP EP17837616.6A patent/EP3494226B1/en active Active
- 2017-08-02 WO PCT/US2017/045137 patent/WO2018026941A1/en active Search and Examination
- 2017-08-02 IL IL264464A patent/IL264464B2/en unknown
- 2017-08-03 US US15/668,101 patent/US11156614B2/en active Active
-
2021
- 2021-09-23 US US17/482,918 patent/US11906521B2/en active Active
Patent Citations (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2014014375A (ja) | 2007-11-12 | 2014-01-30 | Hiroshima Univ | エンドトキシンの濃度測定方法および濃度測定用キット |
Non-Patent Citations (3)
| Title |
|---|
| James C. Hurley,Endotoxemia: Methods of detection and clinical correlates,Clinical Microbiology Reviews,1995年01月01日,vol 8, no.2,268-292 |
| Shin Nakamura et al.,Fractionation of limulus amebocyte lysate,Biochimica et Biophysica Acta,1982年10月05日,707,217-225 |
| Toshie Harada-Suzuki et al.,Further studies on the chromogenic substrate assay method for bacterial endotoxins using horseshoe crab (Tachypleus tridentatus) hemocyte lysate,Journal of Biochemistry,1982年12月01日,92,793-800 |
Also Published As
| Publication number | Publication date |
|---|---|
| KR102561368B1 (ko) | 2023-08-01 |
| SG11201900767VA (en) | 2019-02-27 |
| EP3943608B1 (en) | 2023-07-12 |
| EP3943608C0 (en) | 2023-07-12 |
| KR20190037284A (ko) | 2019-04-05 |
| EP3494226A4 (en) | 2020-03-18 |
| IL264464B1 (en) | 2023-10-01 |
| EP3943608A1 (en) | 2022-01-26 |
| WO2018026941A1 (en) | 2018-02-08 |
| IL264464B2 (en) | 2024-02-01 |
| US20220074940A1 (en) | 2022-03-10 |
| IL264464A (en) | 2019-02-28 |
| CN109790561B (zh) | 2024-02-06 |
| EP3494226A1 (en) | 2019-06-12 |
| EP3494226B1 (en) | 2021-09-22 |
| CN109790561A (zh) | 2019-05-21 |
| US11156614B2 (en) | 2021-10-26 |
| US20180038864A1 (en) | 2018-02-08 |
| JP2019526052A (ja) | 2019-09-12 |
| US11906521B2 (en) | 2024-02-20 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP7044222B2 (ja) | 実質的にコアギュローゲンを含まないリムルスアメボサイトライセートを使用してエンドトキシンを検出する方法 | |
| Wildfeuer et al. | Investigations on the specificity of the Limulus test for the detection of endotoxin | |
| US20220267824A1 (en) | Coagulogen-free clarified limulus amebocyte lysate | |
| JP5096166B2 (ja) | 生体サンプル中の微生物を分類する方法 | |
| Vanterpool et al. | Altered gingipain maturation in vimA-and vimE-defective isogenic mutants of Porphyromonas gingivalis | |
| US6391570B1 (en) | Methods and compositions for the detection of bacterial endotoxins | |
| Hodes et al. | Reaction of fungal products with amebocyte lysates of the Japanese horseshoe crab, Tachypleus tridentatus | |
| Vanhaecke et al. | Endotoxin testing | |
| Roth et al. | [6] Measurement of endotoxin levels in hemoglobin preparations | |
| Friberger | Endotoxin determination with Limulus lysate in clinical diagnosis and hygienic control | |
| Apicella et al. | Physicochemical properties of Neisseria meningitidis group X polysaccharide antigen | |
| JP2024507115A (ja) | ハイブリッドアメボサイト溶解物及びその使用 | |
| KR20200121096A (ko) | 리포폴리사카라이드 추출용 용매 및 이를 이용한 리포폴리사카라이드 추출 방법 | |
| Wu | A chromogenic Limulus amebocyte lysate test for determination of endotoxin in the chicken plasma | |
| McCartney | The Limulus Amoebocyte Lysate assay for bacterial endotoxins | |
| Al-Saffar et al. | Quantitative Detection of the Partially Purified Endotoxin Extracted from the Locally Isolated Salmonella typhimurium A3 | |
| Ito | Characterization of the Proteases Obtained from Streptomyces Cultures I. Biological Properties of Five Preparations Found as Cytotoxic Principles | |
| Wang | Fragmentation of the endotoxin complex from Serratia marcescens bizio and determination of the structure of its polysaccharide side chain | |
| Cáceres et al. | Studies about polymers, their interactions with the immune system and applications in immunotechnology in Uruguay: a systematic review Estudos sobre polímeros, suas interações com o sistema imunológico e aplicações em imunotecnologia no Uruguai: uma revisão sistemática |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| A529 | Written submission of copy of amendment under article 34 pct |
Free format text: JAPANESE INTERMEDIATE CODE: A529 Effective date: 20190318 |
|
| RD04 | Notification of resignation of power of attorney |
Free format text: JAPANESE INTERMEDIATE CODE: A7424 Effective date: 20190913 |
|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A523 Effective date: 20200731 |
|
| A621 | Written request for application examination |
Free format text: JAPANESE INTERMEDIATE CODE: A621 Effective date: 20200731 |
|
| A977 | Report on retrieval |
Free format text: JAPANESE INTERMEDIATE CODE: A971007 Effective date: 20210729 |
|
| A131 | Notification of reasons for refusal |
Free format text: JAPANESE INTERMEDIATE CODE: A131 Effective date: 20210817 |
|
| A521 | Request for written amendment filed |
Free format text: JAPANESE INTERMEDIATE CODE: A523 Effective date: 20211116 |
|
| TRDD | Decision of grant or rejection written | ||
| A01 | Written decision to grant a patent or to grant a registration (utility model) |
Free format text: JAPANESE INTERMEDIATE CODE: A01 Effective date: 20220215 |
|
| A61 | First payment of annual fees (during grant procedure) |
Free format text: JAPANESE INTERMEDIATE CODE: A61 Effective date: 20220302 |
|
| R150 | Certificate of patent or registration of utility model |
Ref document number: 7044222 Country of ref document: JP Free format text: JAPANESE INTERMEDIATE CODE: R150 |