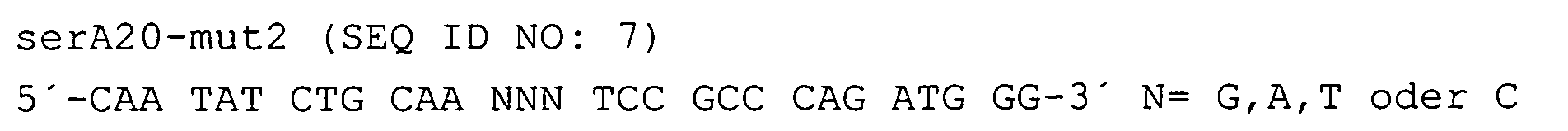EP1496111A2 - Variants of 3-Phosphoglycerate dehydrogenase with reduced inhibition by L-serine and genes encoding the same - Google Patents
Variants of 3-Phosphoglycerate dehydrogenase with reduced inhibition by L-serine and genes encoding the same Download PDFInfo
- Publication number
- EP1496111A2 EP1496111A2 EP04015524A EP04015524A EP1496111A2 EP 1496111 A2 EP1496111 A2 EP 1496111A2 EP 04015524 A EP04015524 A EP 04015524A EP 04015524 A EP04015524 A EP 04015524A EP 1496111 A2 EP1496111 A2 EP 1496111A2
- Authority
- EP
- European Patent Office
- Prior art keywords
- pgd
- amino acid
- sera
- amino acids
- codon
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Granted
Links
- 108010038555 Phosphoglycerate dehydrogenase Proteins 0.000 title claims description 56
- 108090000623 proteins and genes Proteins 0.000 title claims description 25
- 230000005764 inhibitory process Effects 0.000 title claims description 17
- MTCFGRXMJLQNBG-REOHCLBHSA-N (2S)-2-Amino-3-hydroxypropansäure Chemical compound OC[C@H](N)C(O)=O MTCFGRXMJLQNBG-REOHCLBHSA-N 0.000 title description 39
- 229960001153 serine Drugs 0.000 title description 20
- 150000001413 amino acids Chemical class 0.000 claims abstract description 49
- 108700028369 Alleles Proteins 0.000 claims abstract description 28
- 241000588724 Escherichia coli Species 0.000 claims abstract description 24
- 108020004705 Codon Proteins 0.000 claims abstract description 23
- 239000013598 vector Substances 0.000 claims abstract description 19
- KCRZDTROFIOPBP-UHFFFAOYSA-N phosphono 2,3-dihydroxypropanoate Chemical compound OCC(O)C(=O)OP(O)(O)=O KCRZDTROFIOPBP-UHFFFAOYSA-N 0.000 claims abstract description 15
- 150000001875 compounds Chemical class 0.000 claims abstract description 14
- 238000000034 method Methods 0.000 claims abstract description 13
- 230000037357 C1-metabolism Effects 0.000 claims abstract description 11
- MTCFGRXMJLQNBG-UHFFFAOYSA-N Serine Natural products OCC(N)C(O)=O MTCFGRXMJLQNBG-UHFFFAOYSA-N 0.000 claims abstract description 6
- 238000012262 fermentative production Methods 0.000 claims abstract description 3
- 230000000694 effects Effects 0.000 claims description 22
- 244000005700 microbiome Species 0.000 claims description 21
- 230000035772 mutation Effects 0.000 claims description 21
- DHMQDGOQFOQNFH-UHFFFAOYSA-N Glycine Chemical compound NCC(O)=O DHMQDGOQFOQNFH-UHFFFAOYSA-N 0.000 claims description 20
- 101150002295 serA gene Proteins 0.000 claims description 16
- 239000004471 Glycine Substances 0.000 claims description 10
- AYFVYJQAPQTCCC-UHFFFAOYSA-N Threonine Natural products CC(O)C(N)C(O)=O AYFVYJQAPQTCCC-UHFFFAOYSA-N 0.000 claims description 8
- 239000004473 Threonine Substances 0.000 claims description 8
- 230000008569 process Effects 0.000 claims description 5
- 238000002360 preparation method Methods 0.000 claims description 4
- 230000002255 enzymatic effect Effects 0.000 claims description 2
- 230000004060 metabolic process Effects 0.000 claims description 2
- 102000004190 Enzymes Human genes 0.000 abstract description 12
- 108090000790 Enzymes Proteins 0.000 abstract description 12
- 230000035945 sensitivity Effects 0.000 abstract description 5
- 230000000813 microbial effect Effects 0.000 abstract description 4
- 101710088194 Dehydrogenase Proteins 0.000 abstract 1
- 238000012258 culturing Methods 0.000 abstract 1
- LXJXRIRHZLFYRP-UHFFFAOYSA-N glyceraldehyde 3-phosphate Chemical compound O=CC(O)COP(O)(O)=O LXJXRIRHZLFYRP-UHFFFAOYSA-N 0.000 abstract 1
- 235000001014 amino acid Nutrition 0.000 description 26
- 229940024606 amino acid Drugs 0.000 description 24
- 238000004519 manufacturing process Methods 0.000 description 10
- 239000012634 fragment Substances 0.000 description 8
- 235000018102 proteins Nutrition 0.000 description 7
- 102000004169 proteins and genes Human genes 0.000 description 7
- GHOKWGTUZJEAQD-ZETCQYMHSA-N (D)-(+)-Pantothenic acid Chemical compound OCC(C)(C)[C@@H](O)C(=O)NCCC(O)=O GHOKWGTUZJEAQD-ZETCQYMHSA-N 0.000 description 6
- 108020004414 DNA Proteins 0.000 description 6
- QIVBCDIJIAJPQS-VIFPVBQESA-N L-tryptophane Chemical compound C1=CC=C2C(C[C@H](N)C(O)=O)=CNC2=C1 QIVBCDIJIAJPQS-VIFPVBQESA-N 0.000 description 6
- 230000015572 biosynthetic process Effects 0.000 description 6
- 210000004899 c-terminal region Anatomy 0.000 description 6
- 239000003112 inhibitor Substances 0.000 description 6
- 238000002703 mutagenesis Methods 0.000 description 6
- 231100000350 mutagenesis Toxicity 0.000 description 6
- 230000006696 biosynthetic metabolic pathway Effects 0.000 description 5
- FFEARJCKVFRZRR-BYPYZUCNSA-N L-methionine Chemical compound CSCC[C@H](N)C(O)=O FFEARJCKVFRZRR-BYPYZUCNSA-N 0.000 description 4
- VZXPDPZARILFQX-BYPYZUCNSA-N O-acetyl-L-serine Chemical compound CC(=O)OC[C@H]([NH3+])C([O-])=O VZXPDPZARILFQX-BYPYZUCNSA-N 0.000 description 4
- 210000004027 cell Anatomy 0.000 description 4
- 238000006243 chemical reaction Methods 0.000 description 4
- 238000012217 deletion Methods 0.000 description 4
- 230000037430 deletion Effects 0.000 description 4
- 229960004452 methionine Drugs 0.000 description 4
- 238000012986 modification Methods 0.000 description 4
- 230000004048 modification Effects 0.000 description 4
- 239000011713 pantothenic acid Substances 0.000 description 4
- 238000003752 polymerase chain reaction Methods 0.000 description 4
- 239000000047 product Substances 0.000 description 4
- 241000894006 Bacteria Species 0.000 description 3
- GHOKWGTUZJEAQD-UHFFFAOYSA-N Chick antidermatitis factor Natural products OCC(C)(C)C(O)C(=O)NCCC(O)=O GHOKWGTUZJEAQD-UHFFFAOYSA-N 0.000 description 3
- 108010074124 Escherichia coli Proteins Proteins 0.000 description 3
- XUJNEKJLAYXESH-REOHCLBHSA-N L-Cysteine Chemical compound SC[C@H](N)C(O)=O XUJNEKJLAYXESH-REOHCLBHSA-N 0.000 description 3
- 229960000723 ampicillin Drugs 0.000 description 3
- AVKUERGKIZMTKX-NJBDSQKTSA-N ampicillin Chemical compound C1([C@@H](N)C(=O)N[C@H]2[C@H]3SC([C@@H](N3C2=O)C(O)=O)(C)C)=CC=CC=C1 AVKUERGKIZMTKX-NJBDSQKTSA-N 0.000 description 3
- 238000010367 cloning Methods 0.000 description 3
- 229930027945 nicotinamide-adenine dinucleotide Natural products 0.000 description 3
- BOPGDPNILDQYTO-NNYOXOHSSA-N nicotinamide-adenine dinucleotide Chemical compound C1=CCC(C(=O)N)=CN1[C@H]1[C@H](O)[C@H](O)[C@@H](COP(O)(=O)OP(O)(=O)OC[C@@H]2[C@H]([C@@H](O)[C@@H](O2)N2C3=NC=NC(N)=C3N=C2)O)O1 BOPGDPNILDQYTO-NNYOXOHSSA-N 0.000 description 3
- 239000002773 nucleotide Substances 0.000 description 3
- 125000003729 nucleotide group Chemical group 0.000 description 3
- 229940055726 pantothenic acid Drugs 0.000 description 3
- 235000019161 pantothenic acid Nutrition 0.000 description 3
- 239000013612 plasmid Substances 0.000 description 3
- 239000013600 plasmid vector Substances 0.000 description 3
- 238000006467 substitution reaction Methods 0.000 description 3
- 229960004799 tryptophan Drugs 0.000 description 3
- LFLUCDOSQPJJBE-UHFFFAOYSA-K 3-phosphonatooxypyruvate(3-) Chemical compound [O-]C(=O)C(=O)COP([O-])([O-])=O LFLUCDOSQPJJBE-UHFFFAOYSA-K 0.000 description 2
- 241000186226 Corynebacterium glutamicum Species 0.000 description 2
- FFEARJCKVFRZRR-UHFFFAOYSA-N L-Methionine Natural products CSCCC(N)C(O)=O FFEARJCKVFRZRR-UHFFFAOYSA-N 0.000 description 2
- 229930195722 L-methionine Natural products 0.000 description 2
- JJIHLJJYMXLCOY-BYPYZUCNSA-N N-acetyl-L-serine Chemical compound CC(=O)N[C@@H](CO)C(O)=O JJIHLJJYMXLCOY-BYPYZUCNSA-N 0.000 description 2
- 108091028043 Nucleic acid sequence Proteins 0.000 description 2
- 108091034117 Oligonucleotide Proteins 0.000 description 2
- FAPWRFPIFSIZLT-UHFFFAOYSA-M Sodium chloride Chemical compound [Na+].[Cl-] FAPWRFPIFSIZLT-UHFFFAOYSA-M 0.000 description 2
- 241000319304 [Brevibacterium] flavum Species 0.000 description 2
- 238000013459 approach Methods 0.000 description 2
- 238000005119 centrifugation Methods 0.000 description 2
- 239000007795 chemical reaction product Substances 0.000 description 2
- 210000000349 chromosome Anatomy 0.000 description 2
- XUJNEKJLAYXESH-UHFFFAOYSA-N cysteine Natural products SCC(N)C(O)=O XUJNEKJLAYXESH-UHFFFAOYSA-N 0.000 description 2
- 229960002433 cysteine Drugs 0.000 description 2
- 239000013604 expression vector Substances 0.000 description 2
- 238000000855 fermentation Methods 0.000 description 2
- 230000004151 fermentation Effects 0.000 description 2
- 239000012467 final product Substances 0.000 description 2
- 230000001939 inductive effect Effects 0.000 description 2
- 238000003780 insertion Methods 0.000 description 2
- 230000037431 insertion Effects 0.000 description 2
- 230000037353 metabolic pathway Effects 0.000 description 2
- 239000002207 metabolite Substances 0.000 description 2
- 229930182817 methionine Natural products 0.000 description 2
- 238000012261 overproduction Methods 0.000 description 2
- 230000003647 oxidation Effects 0.000 description 2
- 238000007254 oxidation reaction Methods 0.000 description 2
- 230000001105 regulatory effect Effects 0.000 description 2
- 108091008146 restriction endonucleases Proteins 0.000 description 2
- 238000012163 sequencing technique Methods 0.000 description 2
- 238000002741 site-directed mutagenesis Methods 0.000 description 2
- 238000003786 synthesis reaction Methods 0.000 description 2
- MSTNYGQPCMXVAQ-RYUDHWBXSA-N (6S)-5,6,7,8-tetrahydrofolic acid Chemical compound C([C@H]1CNC=2N=C(NC(=O)C=2N1)N)NC1=CC=C(C(=O)N[C@@H](CCC(O)=O)C(O)=O)C=C1 MSTNYGQPCMXVAQ-RYUDHWBXSA-N 0.000 description 1
- PWKSKIMOESPYIA-UHFFFAOYSA-N 2-acetamido-3-sulfanylpropanoic acid Chemical compound CC(=O)NC(CS)C(O)=O PWKSKIMOESPYIA-UHFFFAOYSA-N 0.000 description 1
- OSJPPGNTCRNQQC-UWTATZPHSA-N 3-phospho-D-glyceric acid Chemical compound OC(=O)[C@H](O)COP(O)(O)=O OSJPPGNTCRNQQC-UWTATZPHSA-N 0.000 description 1
- OSJPPGNTCRNQQC-UWTATZPHSA-K 3-phosphonato-D-glycerate(3-) Chemical compound [O-]C(=O)[C@H](O)COP([O-])([O-])=O OSJPPGNTCRNQQC-UWTATZPHSA-K 0.000 description 1
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 description 1
- MSTNYGQPCMXVAQ-KIYNQFGBSA-N 5,6,7,8-tetrahydrofolic acid Chemical compound N1C=2C(=O)NC(N)=NC=2NCC1CNC1=CC=C(C(=O)N[C@@H](CCC(O)=O)C(O)=O)C=C1 MSTNYGQPCMXVAQ-KIYNQFGBSA-N 0.000 description 1
- 241000219195 Arabidopsis thaliana Species 0.000 description 1
- 244000063299 Bacillus subtilis Species 0.000 description 1
- 235000014469 Bacillus subtilis Nutrition 0.000 description 1
- 101100021490 Bacillus subtilis (strain 168) lnrK gene Proteins 0.000 description 1
- LHTRWNBZZJUFDX-MMFRDWCLSA-N C(CC[C@@H](C(=O)O)NC(=O)C1=CC=C(NCC2CNC=3N=C(N)NC(=O)C3N2)C=C1)(=O)O.NCC(=O)O Chemical compound C(CC[C@@H](C(=O)O)NC(=O)C1=CC=C(NCC2CNC=3N=C(N)NC(=O)C3N2)C=C1)(=O)O.NCC(=O)O LHTRWNBZZJUFDX-MMFRDWCLSA-N 0.000 description 1
- OKTJSMMVPCPJKN-UHFFFAOYSA-N Carbon Chemical compound [C] OKTJSMMVPCPJKN-UHFFFAOYSA-N 0.000 description 1
- LEVWYRKDKASIDU-QWWZWVQMSA-N D-cystine Chemical compound OC(=O)[C@H](N)CSSC[C@@H](N)C(O)=O LEVWYRKDKASIDU-QWWZWVQMSA-N 0.000 description 1
- 102000004594 DNA Polymerase I Human genes 0.000 description 1
- 108010017826 DNA Polymerase I Proteins 0.000 description 1
- 101100498063 Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) cysB gene Proteins 0.000 description 1
- 241000588921 Enterobacteriaceae Species 0.000 description 1
- 101100117984 Escherichia coli (strain K12) eamB gene Proteins 0.000 description 1
- 241001302584 Escherichia coli str. K-12 substr. W3110 Species 0.000 description 1
- 241000233866 Fungi Species 0.000 description 1
- 108010000445 Glycerate dehydrogenase Proteins 0.000 description 1
- 235000013878 L-cysteine Nutrition 0.000 description 1
- 239000004201 L-cysteine Substances 0.000 description 1
- LEVWYRKDKASIDU-IMJSIDKUSA-N L-cystine Chemical compound [O-]C(=O)[C@@H]([NH3+])CSSC[C@H]([NH3+])C([O-])=O LEVWYRKDKASIDU-IMJSIDKUSA-N 0.000 description 1
- 102000003960 Ligases Human genes 0.000 description 1
- 108090000364 Ligases Proteins 0.000 description 1
- 229910019142 PO4 Inorganic materials 0.000 description 1
- 101100408135 Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) phnA gene Proteins 0.000 description 1
- 241000700157 Rattus norvegicus Species 0.000 description 1
- 108700008625 Reporter Genes Proteins 0.000 description 1
- 240000004808 Saccharomyces cerevisiae Species 0.000 description 1
- 239000004098 Tetracycline Substances 0.000 description 1
- JLCPHMBAVCMARE-UHFFFAOYSA-N [3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[3-[[3-[[3-[[3-[[3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-[[5-(2-amino-6-oxo-1H-purin-9-yl)-3-hydroxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxyoxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(5-methyl-2,4-dioxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(6-aminopurin-9-yl)oxolan-2-yl]methoxy-hydroxyphosphoryl]oxy-5-(4-amino-2-oxopyrimidin-1-yl)oxolan-2-yl]methyl [5-(6-aminopurin-9-yl)-2-(hydroxymethyl)oxolan-3-yl] hydrogen phosphate Polymers Cc1cn(C2CC(OP(O)(=O)OCC3OC(CC3OP(O)(=O)OCC3OC(CC3O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c3nc(N)[nH]c4=O)C(COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3COP(O)(=O)OC3CC(OC3CO)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3ccc(N)nc3=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cc(C)c(=O)[nH]c3=O)n3cc(C)c(=O)[nH]c3=O)n3ccc(N)nc3=O)n3cc(C)c(=O)[nH]c3=O)n3cnc4c3nc(N)[nH]c4=O)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)n3cnc4c(N)ncnc34)O2)c(=O)[nH]c1=O JLCPHMBAVCMARE-UHFFFAOYSA-N 0.000 description 1
- 238000002835 absorbance Methods 0.000 description 1
- 238000009825 accumulation Methods 0.000 description 1
- 239000000654 additive Substances 0.000 description 1
- 230000002411 adverse Effects 0.000 description 1
- 238000000246 agarose gel electrophoresis Methods 0.000 description 1
- 229960003767 alanine Drugs 0.000 description 1
- 230000008856 allosteric binding Effects 0.000 description 1
- 230000003281 allosteric effect Effects 0.000 description 1
- 239000003242 anti bacterial agent Substances 0.000 description 1
- 229940088710 antibiotic agent Drugs 0.000 description 1
- 125000000613 asparagine group Chemical group N[C@@H](CC(N)=O)C(=O)* 0.000 description 1
- 230000001580 bacterial effect Effects 0.000 description 1
- UUQMNUMQCIQDMZ-UHFFFAOYSA-N betahistine Chemical compound CNCCC1=CC=CC=N1 UUQMNUMQCIQDMZ-UHFFFAOYSA-N 0.000 description 1
- 230000027455 binding Effects 0.000 description 1
- 230000003115 biocidal effect Effects 0.000 description 1
- 230000001486 biosynthesis of amino acids Effects 0.000 description 1
- 229940041514 candida albicans extract Drugs 0.000 description 1
- 229910052799 carbon Inorganic materials 0.000 description 1
- 230000003197 catalytic effect Effects 0.000 description 1
- 238000004113 cell culture Methods 0.000 description 1
- 230000008859 change Effects 0.000 description 1
- 229960005091 chloramphenicol Drugs 0.000 description 1
- WIIZWVCIJKGZOK-RKDXNWHRSA-N chloramphenicol Chemical compound ClC(Cl)C(=O)N[C@H](CO)[C@H](O)C1=CC=C([N+]([O-])=O)C=C1 WIIZWVCIJKGZOK-RKDXNWHRSA-N 0.000 description 1
- 239000000287 crude extract Substances 0.000 description 1
- 101150058227 cysB gene Proteins 0.000 description 1
- 101150111114 cysE gene Proteins 0.000 description 1
- 235000018417 cysteine Nutrition 0.000 description 1
- 150000001944 cysteine derivatives Chemical class 0.000 description 1
- 229960003067 cystine Drugs 0.000 description 1
- 230000003247 decreasing effect Effects 0.000 description 1
- 229950006137 dexfosfoserine Drugs 0.000 description 1
- VHJLVAABSRFDPM-QWWZWVQMSA-N dithiothreitol Chemical compound SC[C@@H](O)[C@H](O)CS VHJLVAABSRFDPM-QWWZWVQMSA-N 0.000 description 1
- 238000004520 electroporation Methods 0.000 description 1
- 238000006911 enzymatic reaction Methods 0.000 description 1
- 230000002068 genetic effect Effects 0.000 description 1
- 125000000291 glutamic acid group Chemical group N[C@@H](CCC(O)=O)C(=O)* 0.000 description 1
- 229960002449 glycine Drugs 0.000 description 1
- 238000010348 incorporation Methods 0.000 description 1
- 238000009776 industrial production Methods 0.000 description 1
- 230000010354 integration Effects 0.000 description 1
- 239000000543 intermediate Substances 0.000 description 1
- 238000002955 isolation Methods 0.000 description 1
- BPHPUYQFMNQIOC-NXRLNHOXSA-N isopropyl beta-D-thiogalactopyranoside Chemical compound CC(C)S[C@@H]1O[C@H](CO)[C@H](O)[C@H](O)[C@H]1O BPHPUYQFMNQIOC-NXRLNHOXSA-N 0.000 description 1
- 229960000318 kanamycin Drugs 0.000 description 1
- 229930027917 kanamycin Natural products 0.000 description 1
- SBUJHOSQTJFQJX-NOAMYHISSA-N kanamycin Chemical compound O[C@@H]1[C@@H](O)[C@H](O)[C@@H](CN)O[C@@H]1O[C@H]1[C@H](O)[C@@H](O[C@@H]2[C@@H]([C@@H](N)[C@H](O)[C@@H](CO)O2)O)[C@H](N)C[C@@H]1N SBUJHOSQTJFQJX-NOAMYHISSA-N 0.000 description 1
- 229930182823 kanamycin A Natural products 0.000 description 1
- 238000011068 loading method Methods 0.000 description 1
- 239000003550 marker Substances 0.000 description 1
- 230000007246 mechanism Effects 0.000 description 1
- 230000002503 metabolic effect Effects 0.000 description 1
- 125000002496 methyl group Chemical group [H]C([H])([H])* 0.000 description 1
- 235000015097 nutrients Nutrition 0.000 description 1
- 230000003287 optical effect Effects 0.000 description 1
- 239000010452 phosphate Substances 0.000 description 1
- BZQFBWGGLXLEPQ-REOHCLBHSA-N phosphoserine Chemical compound OC(=O)[C@@H](N)COP(O)(O)=O BZQFBWGGLXLEPQ-REOHCLBHSA-N 0.000 description 1
- 238000000746 purification Methods 0.000 description 1
- 239000011541 reaction mixture Substances 0.000 description 1
- 230000009467 reduction Effects 0.000 description 1
- 230000010076 replication Effects 0.000 description 1
- 210000003705 ribosome Anatomy 0.000 description 1
- 101150003830 serC gene Proteins 0.000 description 1
- IFGCUJZIWBUILZ-UHFFFAOYSA-N sodium 2-[[2-[[hydroxy-(3,4,5-trihydroxy-6-methyloxan-2-yl)oxyphosphoryl]amino]-4-methylpentanoyl]amino]-3-(1H-indol-3-yl)propanoic acid Chemical compound [Na+].C=1NC2=CC=CC=C2C=1CC(C(O)=O)NC(=O)C(CC(C)C)NP(O)(=O)OC1OC(C)C(O)C(O)C1O IFGCUJZIWBUILZ-UHFFFAOYSA-N 0.000 description 1
- 239000011780 sodium chloride Substances 0.000 description 1
- 238000010561 standard procedure Methods 0.000 description 1
- 239000007858 starting material Substances 0.000 description 1
- 239000000126 substance Substances 0.000 description 1
- 230000002195 synergetic effect Effects 0.000 description 1
- 238000012360 testing method Methods 0.000 description 1
- 229960002180 tetracycline Drugs 0.000 description 1
- 229930101283 tetracycline Natural products 0.000 description 1
- 235000019364 tetracycline Nutrition 0.000 description 1
- 150000003522 tetracyclines Chemical class 0.000 description 1
- 239000005460 tetrahydrofolate Substances 0.000 description 1
- 238000012546 transfer Methods 0.000 description 1
- 230000009466 transformation Effects 0.000 description 1
- 238000011426 transformation method Methods 0.000 description 1
- 101150044170 trpE gene Proteins 0.000 description 1
- 239000012137 tryptone Substances 0.000 description 1
- 238000012795 verification Methods 0.000 description 1
- 239000012138 yeast extract Substances 0.000 description 1
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/0004—Oxidoreductases (1.)
- C12N9/0006—Oxidoreductases (1.) acting on CH-OH groups as donors (1.1)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/04—Alpha- or beta- amino acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/04—Alpha- or beta- amino acids
- C12P13/06—Alanine; Leucine; Isoleucine; Serine; Homoserine
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y101/00—Oxidoreductases acting on the CH-OH group of donors (1.1)
- C12Y101/01—Oxidoreductases acting on the CH-OH group of donors (1.1) with NAD+ or NADP+ as acceptor (1.1.1)
- C12Y101/01095—Phosphoglycerate dehydrogenase (1.1.1.95)
Definitions
- the invention relates to variants of 3-phosphoglycerate dehydrogenase with reduced inhibition by L-serine and coding for it Genes.
- biosynthetic pathways are subject to wild-type strains a strict control that ensures that the amino acids be made only for personal use of the cell.
- important control mechanism in many biosyntheses the phenomenon of feedback inhibition (or end product inhibition).
- the enzyme of a biosynthetic pathway this is the preliminary enzymatic reaction of this Biosynthesis pathways catalyzed by the end product of Biosyntheseweges inhibited.
- the inhibition usually takes place through an allosteric Binding of the final product to the enzyme instead of and causes a conformational change to an inactive state. This ensures that when an accumulation of the final product in the cell further synthesis by inhibition of introductory step is prevented.
- the phosphoglycerate family of amino acids is defined by that they are amino acids involved in their biosynthesis derived from the 3-phosphoglyceric acid. Of the natural path of metabolism leads to it over the intermediates 3-Phospohydroxypyruvat and 3-phospho-Lserin to L-serine. L-serine can continue to glycine or over O-acetyl-serine be converted to L-cysteine. Also L-tryptophan is to be counted to this group, since it is also in derived from the biosynthesis of L-serine. Likewise, unnatural Amino acids described in US 2002/0039767 A1 Process to be assigned to the phosphoglycerate family.
- the initial step in the biosynthesis of amino acids of the phosphoglycerate family is the oxidation of D-3-phosphoglyceric acid to 3-phosphohydroxypyruvate and is catalyzed by the enzyme 3-phosphoglycerate dehydrogenase (PGD) [EC 1.1.1.95].
- PGD 3-phosphoglycerate dehydrogenase
- NAD + which is converted into NADH / H + , serves as the acceptor for the reduction equivalents formed during the reaction.
- PGD enzymes are known from a variety of organisms (e.g. Rattus norvegicus, Arabidopsis thaliana, Escherichia coli, Bacillus subtilis). The better characterized microbial representatives of feedback inhibition by L-serine.
- amino acid 1-340 of the E. coli sequence amino acid 1-340 of the E. coli sequence
- C-terminal portion has little similarity.
- the C-terminal Part is localized to the regulatory domain, the is responsible for serine inhibition (Peters-Wendisch et al., 2002, Appl. Microbiol. Biotechnol. 60: 437-441).
- PGD The best characterized PGD is that of Escherichia coli.
- the enzyme has been extensively biochemically studied (Dubrow & Pizer, 1977, J. Biol. Chem. 252, 1527-1538) and is subject to allosteric feedback inhibition by L-serine with an inhibitor constant K i of 5 ⁇ M.
- linker mutagenesis are usually problematic because by the incorporation or deletion of several residues the Structure of the protein is very much changed and so the overall activity or stability of the protein adversely affected becomes. In fact, most of them are described in EP0620853A Mutants a barely detectable activity.
- Object of the present invention was to variants of PGD of Escherichia coli which have an im Compared to Escherichia coli wild-type PGD decreased sensitivity against inhibition by serine.
- a PGD having an amino acid sequence which is characterized by being different from the Amino acid sequence of Escherichia coli wild-type PGD (SEQ ID NO: 2) with a methionine as position 1 differs thereby that at position 349 an amino acid except glycine or at position 372 an amino acid except threonine located.
- a PGD according to the invention may also have mutations in both Have positions of SEQ ID NO: 1.
- the invention further relates to a DNA sequence coding for a PGD according to the invention.
- This serA allele is different different from the gene of Escherichia coli PGD (serA gene, SEQ ID NO: 1) in that the codon 349 is a natural amino acid Exception of glycine or codon 372 for a natural amino acid coded with the exception of threonine.
- a serA allele according to the invention may also have a mutation have two codons mentioned.
- inventive serA alleles include such genes that are analyzed with the algorithm GAP (GCG Wisconsin Package, Genetics Computer Group (GCG) Madison, Wisconsin) a sequence identity greater than 30% if they have any of the above mutations exhibit. Particularly preferred is a sequence identity greater than 70%.
- proteins with a sequence identity greater than 40 % determined with the algorithm GAP in the sense of the present To conceive of the invention as proteins derived from E. coli PGD, provided they have a PGD activity and one of the above Have amino acid substitutions. Particularly preferred is a Sequence identity greater than 70%.
- genes according to the invention are allelic variants of the serA gene to be understood by deletion, insertion or Substitution of nucleotides from that shown in SEQ ID NO: 1 Derive sequence, with the enzymatic activity of the gene product more than 10% of the activity of the wild-type gene product and a mutation of codon 349 for the Amino acid glycine or codon 372 for the amino acid threonine, or a combination of the two mutations.
- PGD variants that have an amino acid substitution at position 349 or at position 372 or a combination thereof, can be generated using standard techniques of molecular biology become. For this purpose, mutations are made in the corresponding codons introduced the serA gene encoding the PGD. Appropriate Methods for introducing mutations at specific positions within a DNA fragment are known.
- the starting material for the mutagenesis is preferably the DNA of the E. coli serA gene.
- the serA gene to be mutated can chromosomally or extrachromosomally encoded. It is preferred However, the serA gene amplified by polymerase chain reaction and cloned into a vector.
- the DNA sequence is altered so that the encoded PGD undergoes an amino acid exchange at position 349 or 372, wherein position 1 is the starting methionine of SEQ ID NO: 1.
- any method can be used which allows to determine the activity of the enzyme in the presence of L-serine.
- the determination of PGD activity can be based on the method described by McKrickick and Pizer (1980, J. Bacteriol 141: 235-245).
- the enzyme activity is measured by the reverse reaction in an approach containing phosphohydroxypyruvate and NADH / H + .
- the reaction is started by enzyme addition and monitored by the decrease in absorbance at 340 nm, which is caused by oxidation of NADH / H + in a spectrophotometer.
- the inhibition of PGD activity is tested in the presence of various concentrations of L-serine in the reaction mixture.
- the catalytic activity of the various PGD variants is determined in the presence and absence of L-serine and used to determine the inhibitor constant K i .
- the K i describes that inhibitor concentration in which the activity is only 50% of the activity determined in the absence of the inhibitor.
- PGD enzymes according to the invention can, due to their feedback resistance for the production of amino acids of the phosphoglycerate family or of compounds resulting from the C1 metabolism derive, be used.
- inventive serA alleles expressed in a host strain are used.
- a serA allele according to the invention can be described under Control of the own promoter localized in front of the serA gene or by using other suitable promoter systems, which are known in the art, take place. This can be the corresponding allele, for example, under the control of a such promoter in either one or more copies located on the chromosome of the host organism.
- the strategies for the integration of genes into the chromosome are state of the art Technology.
- the serA allele to be expressed is preferred cloned into a vector, preferably a plasmid.
- the invention therefore also relates to a vector, characterized that he under a serA allelic according to the invention functional control of a promoter.
- vectors which already contain genetic elements (e.g. constitutive or regulatable promoters, terminators), either an ongoing or a controlled, allow inducible expression of the gene coding for the PGD. They are also located on an expression vector preferably other regulatory elements such as ribosomal Binding sites and termination sequences as well as sequences that code for selective markers and / or reporter genes.
- the expression Such selection marker facilitates identification of transformants. Suitable as selection markers Genes that are resistant to z. B. ampicillin, Tetracycline, chloramphenicol, kanamycin or other antibiotics encode.
- the plasmid vector should preferably contain an origin point of replication.
- plasmid vectors such as the E. coli vectors pACYC184, pUC18, pQE-70, pBR322, pSC101 and their derivatives.
- Suitable inducible promoters are, for example the lac, tac, trc, lambda PL, ara or tet promoter or sequences derived therefrom.
- Such vectors allow the direct production of inventive Microorganism strains with high production capacity from any microorganism strain, as such Plasmid also the lifting of other restrictions of the Metabolic pathway in a microorganism causes.
- a common transformation method e.g., electroporation
- the serA allele-containing plasmids according to the invention incorporated in microorganisms and, for example, by antibiotic resistance selected for plasmid-carrying clones.
- the invention thus also relates to processes for the preparation a microorganism strain according to the invention, characterized that in a microorganism strain an inventive Vector is introduced.
- a host organism for vectors of the invention all organisms suitable for the biosynthetic pathway for amino acids the phosphoglycerate family, recombinant Process are accessible and cultivable by fermentation are.
- Such microorganisms can be fungi, yeasts or bacteria be. Preference is given to bacteria of the phylogenetic group Eubacteria are used. Particularly preferred are microorganisms the family Enterobacteriaceae and in particular the Species Escherichia coli.
- the invention thus further relates to a microorganism strain, for the fermentative production of amino acids of Phosphoglycerate family or their derivatives or compounds, which derive from the C1 metabolism suitable is, characterized in that it is a PGD according to the invention has.
- Another object of the invention is the production of Amino acids of the phosphoglycerate family or of compounds, which derive from the C1 metabolism, by cultivation a microorganism strain according to the invention.
- the microorganism strain according to the invention for example cultured in a fermenter in a nutrient medium, the a suitable carbon, and a suitable source of energy, and other additives.
- the substances formed during the fermentation such as L-phosphoserine, L-serine, O-acetyl-L-serine, L-cysteine, Glycine, L-tryptophan, 1,2,4-triazol-2-yl-L-alanine, L-methionine or pantothenic acid can subsequently be purified become.
- the serA gene from Escherichia coli strain W3110 was amplified by the polymerase chain reaction.
- the oligonucleotides were used as specific primers and
- the resulting DNA fragment was digested with the restriction enzymes NdeI and HindIII digested and the 5 'overhangs with Klenow enzyme refilled. Subsequently, the DNA fragment was using purified by agarose gel electrophoresis and using the GeneClean method isolated (GeneClean Kit BI0101 P.O.Box 2284 La Jolla, California, 92038-2284). The cloning of the thus obtained serA fragment was inserted into the expression vector pQE-70 (Qiagen, Hilden, D). For this, the vector was first with SphI and BamHI cut and the 3 'overhang with Klenow enzyme or the 5 'overhang filled with Klenow enzyme.
- the vector fragment was purified and washed with ligated to the serA fragment.
- the resulting vector carries the Designation pFL209.
- the bacterial strain Escherichia coli JM109 / pFL209 was used in the DSMZ (German Collection for microorganisms and cell cultures GmbH, D-38142 Braunschweig) under the number DSM 15628 according to the Budapest Treaty deposited.
- Site-directed mutagenesis at codons 349 and 372 of the serA gene was performed by an inverse polymerase chain reaction.
- the template used was the vector pFL209 described in Example 1.
- the primers were and used.
- 100 ml LB medium (10 g / l tryptone, 5 g / l yeast extract, 10 g / l NaCl), which additionally contained 100 mg / l ampicillin , inoculated with a 2 ml overnight culture of the strains with the plasmid-coded serA alleles and incubated at 30 ° C and 150 rpm in a shaker.
- serA expression was induced by the addition of 0.4 mM isopropyl- ⁇ -thiogalactoside, and the culture was incubated for a further 3 hours.
- the cells were then harvested by centrifugation, washed and resuspended in 2 ml of buffer (100 mM K-phosphate pH 7.0, 10 mM MgCl 2 , 1 mM dithiothreitol). Cell disruption was performed using a French Press (Spectronic Instruments, Inc., Rochester, NY, USA) at a pressure of 18,000 psi. The crude extracts were clarified by centrifugation at 30,000 g and PGD activity was determined by the test of McKrickick and Pizer (1980, J. Bacteriol 141: 235-245). The following tables show the PGD activity of different mutants, as well as the corresponding inhibitor constants K i .
- a combination of codons 349 and 372 mutations should show whether the exchanges have a synergistic effect on feedback resistance.
- a unique HindII restriction site between the two mutation sites was used.
- HindIII-HindIII restriction of the serA20 allele vector isolated a 183 bp fragment corresponding to the 3 'end of the serA gene and containing the mutation at codon 372. This fragment was cloned into a similarly Hindllam-BamHI digested vector with the serA40 allele to give a clone which is a double mutant.
- the following table shows the associated enzyme data. Mutations at codons 349 and 372 allele mutation Activity [units / mg] Ki [mM] serA wildtype 0.05 ⁇ 0.1 serA2040 G349D, T372I 0.05 120
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Medicinal Chemistry (AREA)
- Molecular Biology (AREA)
- Biomedical Technology (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
- Enzymes And Modification Thereof (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
Description
Die Erfindung betrifft Varianten der 3-Phospoglyceratdehydrogenase mit reduzierter Hemmung durch L-Serin und dafür codierende Gene.The invention relates to variants of 3-phosphoglycerate dehydrogenase with reduced inhibition by L-serine and coding for it Genes.
Die Herstellung der zwanzig natürlichen, proteinogenen Aminosäuren wird heutzutage vorwiegend durch Fermentation von Mikroorganismen bewerkstelligt. Dabei wird ausgenützt, dass Mikroorganismen über entsprechende Biosynthesewege zur Synthese der natürlichen Aminosäuren verfügen.The production of the twenty natural, proteinogenic amino acids Nowadays, it is predominantly fermented by microorganisms accomplished. It exploits that microorganisms via appropriate biosynthetic pathways for synthesis of natural amino acids.
Solche Biosynthesewege unterliegen jedoch in Wildtyp-Stämmen einer strengen Kontrolle, die gewährleistet, dass die Aminosäuren nur zum Eigenbedarf der Zelle hergestellt werden. Ein wichtiger Kontrollmechanismus in vielen Biosynthesen ist beispielsweise das Phänomen der Feedback-Hemmung (oder Endprodukt-Hemmung). Dabei wird meist dasjenige Enzym eines Biosyntheseweges, das die einleitende enzymatische Reaktion dieses Biosynthesewegs katalysiert, durch das Endprodukt des Biosyntheseweges gehemmt. Die Hemmung findet meist durch eine allosterische Bindung des Endproduktes an das Enzym statt und bewirkt eine Konformationsänderung in einen inaktiven Zustand. So wird sichergestellt, dass bei einer Anhäufung des Endproduktes in der Zelle die weitere Synthese durch Hemmung des einleitenden Schrittes unterbunden wird.However, such biosynthetic pathways are subject to wild-type strains a strict control that ensures that the amino acids be made only for personal use of the cell. One For example, important control mechanism in many biosyntheses the phenomenon of feedback inhibition (or end product inhibition). Usually the enzyme of a biosynthetic pathway, this is the preliminary enzymatic reaction of this Biosynthesis pathways catalyzed by the end product of Biosyntheseweges inhibited. The inhibition usually takes place through an allosteric Binding of the final product to the enzyme instead of and causes a conformational change to an inactive state. This ensures that when an accumulation of the final product in the cell further synthesis by inhibition of introductory step is prevented.
Eine effiziente industrielle Produktion von Stoffwechselprodukten (wie z.B. Aminosäuren) ist deshalb nur möglich, wenn die Restriktionen durch Feedbackhemmung eines Stoffwechselweges aufgehoben werden können und damit Mikroorganismen verfügbar gemacht werden, die im Gegensatz zu Wildtyp-Organismen eine drastisch gesteigerte Produktionsleistung für die Herstellung des gewünschten Stoffwechselprodukts aufweisen.An efficient industrial production of metabolic products (such as amino acids) is therefore only possible if the restrictions by feedback inhibition of a metabolic pathway can be repealed and thus microorganisms available which, in contrast to wild-type organisms, a drastically increased production capacity for production have the desired metabolite.
Die Phosphoglycerat-Familie von Aminosäuren ist dadurch definiert, dass es sich um Aminosäuren handelt, die in ihrer Biosynthese von der 3-Phosphoglycerinsäure abgeleitet werden. Der natürliche Pfad des Stoffwechsels führt dabei zunächst über die Zwischenstufen 3-Phospohydroxypyruvat und 3-Phospho-Lserin zu L-Serin. L-Serin kann weiterhin zu Glycin bzw. über O-Acetyl-Serin zu L-Cystein umgesetzt werden. Auch L-Tryptophan ist zu dieser Gruppe zu zählen, da es sich ebenfalls in der Biosynthese von L-Serin ableitet. Ebenso sind unnatürliche Aminosäuren, die nach dem in US 2002/0039767 A1 beschrieben Verfahren hergestellt werden, der Phosphoglycerat-Familie zuzuordnen.The phosphoglycerate family of amino acids is defined by that they are amino acids involved in their biosynthesis derived from the 3-phosphoglyceric acid. Of the natural path of metabolism leads to it over the intermediates 3-Phospohydroxypyruvat and 3-phospho-Lserin to L-serine. L-serine can continue to glycine or over O-acetyl-serine be converted to L-cysteine. Also L-tryptophan is to be counted to this group, since it is also in derived from the biosynthesis of L-serine. Likewise, unnatural Amino acids described in US 2002/0039767 A1 Process to be assigned to the phosphoglycerate family.
Verbindungen, die sich aus dem C1-Stoffwechsel ableiten, zeigen ebenfalls eine Abhängigkeit von der Biosynthese der Aminosäuren der Phosphoglycerat-Familie. Dies beruht auf der Tatsache, dass bei der Konversion von L-Serin zu Glycin Tetrahydrofolat als C1-Gruppenakzeptor fungiert und das beladende Tetrahydrofolat als zentraler Methylgruppendonor im C1-Stoffwechsel an vielen Biosynthesen (z.B. L-Methionin, Nukleotide, Pantothensäure, etc.) beteiligt ist. Erfindungsgemäß sind Verbindungen, die sich aus dem C1-Stoffwechsel ableiten somit vorzugsweise Verbindungen, die in ihrer Biosynthese von einer C1-Gruppenübertragung über Tetrahydrofolsäure abhängen.Compounds derived from the C1 metabolism show also a dependence on the biosynthesis of the amino acids the phosphoglycerate family. This is based on the fact that in the conversion of L-serine to glycine tetrahydrofolate acts as a C1 group acceptor and the loading tetrahydrofolate as a central methyl group donor in C1 metabolism in many biosyntheses (e.g., L-methionine, nucleotides, pantothenic acid, etc.). According to the invention, compounds derive from the C1 metabolism thus preferably Compounds that in their biosynthesis of a C1 group transfer depend on tetrahydrofolic acid.
Der einleitende Schritt der Biosynthese von Aminosäuren der Phosphoglycerat-Familie ist die Oxidation der D-3-Phosphoglycerinsäure zu 3-Phosphohydroxypyruvat und wird durch das Enzym 3-Phosphoglyceratdehydrogenase (PGD) [EC 1.1.1.95] katalysiert. Als Akzeptor für die bei der Reaktion entstehenden Reduktionsäquivalente dient NAD+, das zu NADH/H+ umgesetzt wird.The initial step in the biosynthesis of amino acids of the phosphoglycerate family is the oxidation of D-3-phosphoglyceric acid to 3-phosphohydroxypyruvate and is catalyzed by the enzyme 3-phosphoglycerate dehydrogenase (PGD) [EC 1.1.1.95]. NAD + , which is converted into NADH / H + , serves as the acceptor for the reduction equivalents formed during the reaction.
PGD-Enzyme sind aus verschiedensten Organismen bekannt (z.B. Rattus norvegicus, Arabidopsis thaliana, Escherichia coli, Bacillus subtilis). Dabei unterliegen die besser charakterisierten mikrobiellen Vertreter einer Feedback-Hemmung durch L-Serin. PGD enzymes are known from a variety of organisms (e.g. Rattus norvegicus, Arabidopsis thaliana, Escherichia coli, Bacillus subtilis). The better characterized microbial representatives of feedback inhibition by L-serine.
Auf der Ebene der Aminosäuresequenz sind die mikrobiellen PGD-Enzyme im N-terminalen Teil (Aminosäure 1-340 der E. coli Sequenz) untereinander sehr ähnlich, wogegen der C-terminale Anteil nur geringe Ähnlichkeiten aufweist. Gerade in diesem C-terminalen Teil ist die regulatorische Domäne lokalisiert, die für die Serin-Hemmung verantwortlich ist (Peters-Wendisch et al., 2002, Appl. Microbiol. Biotechnol. 60:437-441).At the level of the amino acid sequence are the microbial PGD enzymes in the N-terminal part (amino acid 1-340 of the E. coli sequence) very similar to each other, whereas the C-terminal portion has little similarity. Especially in this C-terminal Part is localized to the regulatory domain, the is responsible for serine inhibition (Peters-Wendisch et al., 2002, Appl. Microbiol. Biotechnol. 60: 437-441).
Die am besten charakterisierte PGD ist diejenige aus Escherichia coli. Das Enzym wurde eingehend biochemisch untersucht (Dubrow & Pizer, 1977, J. Biol. Chem. 252, 1527-1538) und unterliegt einer allosterischen Feedback-Hemmung durch L-Serin mit einer Inhibitorkonstante Ki von 5 µM.The best characterized PGD is that of Escherichia coli. The enzyme has been extensively biochemically studied (Dubrow & Pizer, 1977, J. Biol. Chem. 252, 1527-1538) and is subject to allosteric feedback inhibition by L-serine with an inhibitor constant K i of 5 μM.
Diese Feedback-Hemmung steht einer effizienten Produktion von Aminosäuren der Phosphoglycerat-Familie im Wege und war deshalb bereits Ziel molekularbiologischer Ansätze.This feedback inhibition stands for an efficient production of Amino acids of the phosphoglycerate family in the way and was therefore already the goal of molecular biological approaches.
So wurden in der Schrift EP0620853A Varianten der Escherichia coli PGD mit verringerter Empfindlichkeit gegen Hemmung durch Serin beschrieben, die eine Modifikation in den C-terminalen 25% der Wildtyp-PGD, (d.h. Aminosäuren 307-410), bevorzugt eine Modifikation im Bereich der letzten 50 Reste (d.h. Aminosäuren 361-410) aufweisen. Die beschriebenen Mutanten wurden durch Linker-Mutagenese, d.h. durch einfache Nutzung von vorhandenen Restriktionsschnittstellen im E. coli serA-Gen und anschließendes Einsetzen von Oligonukleotid-Linkern einer Länge von 8-14 Basenpaaren erhalten.Thus, in the document EP0620853A variants of Escherichia coli PGD with reduced sensitivity to inhibition by Serine described a modification in the C-terminal 25% of the wild-type PGD (i.e., amino acids 307-410), preferably one Modification in the region of the last 50 residues (i.e., amino acids 361-410). The described mutants were by linker mutagenesis, i. by easy use of existing ones Restriction interfaces in the E. coli serA gene and subsequent insertion of oligonucleotide linkers of a length of 8-14 base pairs.
Solche Linker-Mutagenesen sind jedoch meist problematisch, da durch den Einbau bzw. die Deletion von mehreren Resten die Struktur des Proteins sehr stark verändert wird und so die Gesamtaktivität oder Stabilität des Proteins negativ beeinflusst wird. Tatsächlich zeigen die meisten in EP0620853A beschriebenen Mutanten eine kaum nachweisbare Aktivität. However, such linker mutagenesis are usually problematic because by the incorporation or deletion of several residues the Structure of the protein is very much changed and so the overall activity or stability of the protein adversely affected becomes. In fact, most of them are described in EP0620853A Mutants a barely detectable activity.
Auch in coryneformen Mikroorganismen wurden Mutagenesen am serA-Gen vorgenommen, die das Ziel einer Verringerung der Empfindlichkeit der PGD gegen Hemmung durch Serin erfüllen:
- Peters-Wendisch et al. (2002, Appl. Microbiol. Biotechnol. 60:437-441) beschreiben C-terminale Deletionen der PGD von Corynebacterium glutamicum. Auch hier führen die Deletionen zu teilweise starken Einbußen an Enzymaktivität.
- Die Anmeldung EP0943687A2 beschreibt einen Austausch des Glutaminsäure-Restes an Position 325 der PGD von Brevibacterium flavum. Dieser Rest liegt im Bezug zur Escherichia coli PGD in einem Alignment mit dem Algorithmus GAP des Programms GCG (GCG Wisconsin Package, Genetics Computer Group (GCG) Madison, Wisconsin) bereits in dem variablen C-terminalen Teil des Proteins und korreliert mit dem Asparaginrest 364 des Escherichia coli Proteins. Da diese Modifikation in dem variablen C-terminalen Teil der PGD liegen, sind keine Rückschlüsse auf das Escherichia coli Protein möglich.
- Peters-Wendisch et al. (2002, Appl.Microbiol.Biotechnol 60: 437-441) describe C-terminal deletions of PGD of Corynebacterium glutamicum. Again, the deletions lead to some strong losses of enzyme activity.
- The application EP0943687A2 describes an exchange of the glutamic acid residue at position 325 of the PGD of Brevibacterium flavum. This residue is already in the variable C-terminal part of the protein and correlates with the asparagine residue 364 in an alignment with the GAP algorithm of the GCG program (GCG Wisconsin Package, Genetics Computer Group (GCG) Madison, Wisconsin) in relation to Escherichia coli PGD of the Escherichia coli protein. Since these modifications are in the variable C-terminal part of the PGD, no conclusions are possible on the Escherichia coli protein.
Aufgabe der vorliegenden Erfindung war es, Varianten der PGD von Escherichia coli zur Verfügung zu stellen, die eine im Vergleich zur Escherichia coli-Wildtyp-PGD verringerte Empfindlichkeit gegen eine Hemmung durch Serin aufweisen.Object of the present invention was to variants of PGD of Escherichia coli which have an im Compared to Escherichia coli wild-type PGD decreased sensitivity against inhibition by serine.
Die Aufgabe wird gelöst durch eine PGD mit einer Aminosäuresequenz, die dadurch gekennzeichnet ist, dass sie sich von der Aminosäuresequenz der Escherichia coli Wildtyp PGD (SEQ ID NO: 2) mit einem Methionin als Position 1 dadurch unterscheidet, dass an Position 349 eine Aminosäure mit Ausnahme von Glycin oder an Position 372 eine Aminosäure mit Ausnahme von Threonin befindet.The object is achieved by a PGD having an amino acid sequence, which is characterized by being different from the Amino acid sequence of Escherichia coli wild-type PGD (SEQ ID NO: 2) with a methionine as position 1 differs thereby that at position 349 an amino acid except glycine or at position 372 an amino acid except threonine located.
Eine erfindungsgemäße PGD kann auch Mutationen an beiden genannten Positionen von SEQ ID NO: 1 aufweisen.A PGD according to the invention may also have mutations in both Have positions of SEQ ID NO: 1.
Die Erfindung betrifft ferner eine DNS-Sequenz codierend für eine erfindungsgemäße PGD. Dieses serA-Allel unterscheidet sich vom Gen der Escherichia coli PGD (serA-Gen, SEQ ID NO: 1) dadurch, dass das Codon 349 für eine natürliche Aminosäure mit Ausnahme von Glycin oder das Codon 372 für eine natürliche Aminosäure mit Ausnahme von Threonin codiert.The invention further relates to a DNA sequence coding for a PGD according to the invention. This serA allele is different different from the gene of Escherichia coli PGD (serA gene, SEQ ID NO: 1) in that the codon 349 is a natural amino acid Exception of glycine or codon 372 for a natural amino acid coded with the exception of threonine.
Ein erfindungsgemäßes serA-Allel kann auch eine Mutation an beiden genannten Codons aufweisen.A serA allele according to the invention may also have a mutation have two codons mentioned.
Im Rahmen der vorliegenden Erfindung sind als erfindungsgemäße serA-Allele auch solche Gene aufzufassen, die bei einer Analyse mit dem Algorithmus GAP (GCG Wisconsin Package, Genetics Computer Group (GCG) Madison, Wisconsin) eine Sequenzidentität von größer 30 % aufweisen, sofern sie eine der genannten Mutationen aufweisen. Besonders bevorzugt ist eine Sequenzidentität von größer 70 %.In the context of the present invention are as inventive serA alleles include such genes that are analyzed with the algorithm GAP (GCG Wisconsin Package, Genetics Computer Group (GCG) Madison, Wisconsin) a sequence identity greater than 30% if they have any of the above mutations exhibit. Particularly preferred is a sequence identity greater than 70%.
Ebenso sind Proteine mit einer Sequenzidentität von größer 40 % ermittelt mit dem Algorithmus GAP, im Sinne der vorliegenden Erfindung als von der E. coli PGD abgeleitete Proteine aufzufassen, sofern sie eine PGD Aktivität und einen der genannten Aminosäureaustausche aufweisen. Besonders bevorzugt ist eine Sequenzidentität von größer 70 %.Likewise, proteins with a sequence identity greater than 40 % determined with the algorithm GAP, in the sense of the present To conceive of the invention as proteins derived from E. coli PGD, provided they have a PGD activity and one of the above Have amino acid substitutions. Particularly preferred is a Sequence identity greater than 70%.
Ferner sind als erfindungsgemäße Gene Allelvarianten des serA-Gens zu verstehen, die sich durch Deletion, Insertion oder Substitution von Nukleotiden aus der in SEQ ID NO: 1 dargestellten Sequenz ableiten, wobei die enzymatische Aktivität des Genprodukts mehr als 10 % der Aktivität des Wildtyp Genprodukts entspricht und eine Mutation des Codons 349 für die Aminosäure Glycin bzw. des Codons 372 für die Aminosäure Threonin, oder eine Kombination der beiden Mutationen vorliegt.Furthermore, genes according to the invention are allelic variants of the serA gene to be understood by deletion, insertion or Substitution of nucleotides from that shown in SEQ ID NO: 1 Derive sequence, with the enzymatic activity of the gene product more than 10% of the activity of the wild-type gene product and a mutation of codon 349 for the Amino acid glycine or codon 372 for the amino acid threonine, or a combination of the two mutations.
PGD-Varianten, die einen Aminosäureaustausch an Position 349 oder an Position 372 oder eine Kombination davon besitzen, können mit Standardtechniken der Molekularbiologie erzeugt werden. Dazu werden an den entsprechenden Codons Mutationen in das für die PGD codierende serA-Gen eingebracht. Entsprechende Methoden zur Einführung von Mutationen an spezifischen Positionen innerhalb eines DNS-Fragmentes sind bekannt. PGD variants that have an amino acid substitution at position 349 or at position 372 or a combination thereof, can be generated using standard techniques of molecular biology become. For this purpose, mutations are made in the corresponding codons introduced the serA gene encoding the PGD. Appropriate Methods for introducing mutations at specific positions within a DNA fragment are known.
Als Ausgangsmaterial für die Mutagenese dient vorzugsweise die DNS des E. coli serA-Gens. Das zu mutierende serA-Gen kann chromosomal oder extrachromosomal codiert sein. Bevorzugt wird das serA-Gen jedoch durch Polymerase-Ketten-Reaktion amplifiziert und in einen Vektor kloniert. Durch Anwendung der vorgenannten Mutagenese-Methoden werden ein oder mehrere Nukleotide der DNS-Sequenz so verändert, dass die codierte PGD einen Aminosäureaustausch an Position 349 oder 372 aufweist, wobei Position 1 das Startmethionin aus SEQ ID NO: 1 ist.The starting material for the mutagenesis is preferably the DNA of the E. coli serA gene. The serA gene to be mutated can chromosomally or extrachromosomally encoded. It is preferred However, the serA gene amplified by polymerase chain reaction and cloned into a vector. By applying the aforementioned Mutagenesis methods are one or more nucleotides the DNA sequence is altered so that the encoded PGD undergoes an amino acid exchange at position 349 or 372, wherein position 1 is the starting methionine of SEQ ID NO: 1.
Diese Mutationen bewirken, dass die codierte PGD eine verminderte Empfindlichkeit gegen Hemmung durch L-Serin (=Feedback-Resistenz) besitzt. Dabei ist besonders vorteilhaft, dass die erfindungsgemäßen PGD-Varianten in Abwesenheit von L-Serin im Vergleich zur Wildtyp-PGD eine Aktivität von mehr als 10 %, vorzugsweise eine unveränderte Aktivität, aufweist.These mutations cause the encoded PGD to diminish one Sensitivity to inhibition by L-serine (= feedback resistance) has. It is particularly advantageous that the PGD variants according to the invention in the absence of L-serine in An activity of more than 10% compared to wild-type PGD, preferably an unchanged activity.
Zur Bestimmung des Ausmaßes der Feedback-Resistenz einer erfindungsgemäßen PGD-Variante kann jede Methode benützt werden, die es erlaubt, die Aktivität des Enzyms in Anwesenheit von L-Serin zu bestimmen. Beispielsweise kann die Bestimmung der PGD-Aktivität in Anlehnung an die von McKitrick und Pizer (1980, J.Bacteriol. 141:235-245) beschriebene Methode erfolgen. Die Enzymaktivität wird anhand der Rückreaktion in einem Ansatz, der Phosphohydroxypyruvat und NADH/H+ enthält, gemessen. Die Reaktion wird durch Enzymzugabe gestartet und über die Abnahme der Extinktion bei 340 nm, die durch Oxidation des NADH/H+ hervorgerufen wird, in einem Spektralphotometer verfolgt. Die Hemmung der PGD-Aktivität wird in Anwesenheit verschiedener Konzentrationen von L-Serin im Reaktionsansatz getestet. Die katalytische Aktivität der verschiedenen PGD-Varianten wird in An- und Abwesenheit von L-Serin bestimmt und daraus die Inhibitorkonstante Ki ermittelt. Der Ki beschreibt diejenige Inhibitorkonzentration, bei welcher die Aktivität nur noch 50 % der Aktivität beträgt, die in Abwesenheit des Inhibitors bestimmt wurde. To determine the extent of feedback resistance of a PGD variant according to the invention, any method can be used which allows to determine the activity of the enzyme in the presence of L-serine. For example, the determination of PGD activity can be based on the method described by McKrickick and Pizer (1980, J. Bacteriol 141: 235-245). The enzyme activity is measured by the reverse reaction in an approach containing phosphohydroxypyruvate and NADH / H + . The reaction is started by enzyme addition and monitored by the decrease in absorbance at 340 nm, which is caused by oxidation of NADH / H + in a spectrophotometer. The inhibition of PGD activity is tested in the presence of various concentrations of L-serine in the reaction mixture. The catalytic activity of the various PGD variants is determined in the presence and absence of L-serine and used to determine the inhibitor constant K i . The K i describes that inhibitor concentration in which the activity is only 50% of the activity determined in the absence of the inhibitor.
Erfindungsgemäße PGD-Enzyme können aufgrund ihrer Feedback-Resistenz zur Herstellung von Aminosäuren der Phosphoglycerat-Familie oder von Verbindungen, die sich aus dem C1-Stoffwechsel ableiten, verwendet werden. Hierfür werden die erfindungsgemäßen serA-Allele in einem Wirtsstamm exprimiert.PGD enzymes according to the invention can, due to their feedback resistance for the production of amino acids of the phosphoglycerate family or of compounds resulting from the C1 metabolism derive, be used. For this purpose, the inventive serA alleles expressed in a host strain.
Die Expression eines erfindungsgemäßen serA-Allels kann unter Kontrolle des eigenen, vor dem serA-Gen lokalisierten Promotors oder durch Verwendung anderer geeigneter Promotorsysteme, die dem Fachmann bekannt sind, erfolgen. Dabei kann sich das entsprechende Allel beispielsweise unter der Kontrolle eines solchen Promotors entweder in einer oder in mehreren Kopien auf dem Chromosom des Wirtsorganismus befinden. Die Strategien zur Integration von Genen in das Chromosom sind Stand der Technik. Bevorzugt wird das zu exprimierende serA-Allel jedoch in einen Vektor kloniert, vorzugsweise ein Plasmid.The expression of a serA allele according to the invention can be described under Control of the own promoter localized in front of the serA gene or by using other suitable promoter systems, which are known in the art, take place. This can be the corresponding allele, for example, under the control of a such promoter in either one or more copies located on the chromosome of the host organism. The strategies for the integration of genes into the chromosome are state of the art Technology. However, the serA allele to be expressed is preferred cloned into a vector, preferably a plasmid.
Die Erfindung betrifft daher auch einen Vektor, dadurch gekennzeichnet, dass er ein erfindungsgemäßes serA-Allel unter funktioneller Kontrolle eines Promotors enthält.The invention therefore also relates to a vector, characterized that he under a serA allelic according to the invention functional control of a promoter.
Zur Klonierung der erfindungsgemäßen serA-Allele können Vektoren verwendet werden, die bereits genetische Elemente (z.B. konstitutive oder regulierbare Promotoren, Terminatoren) enthalten, die entweder eine andauernde oder eine kontrollierte, induzierbare Expression des für die PGD codierenden Gens ermöglichen. Außerdem befinden sich auf einem Expressionsvektor vorzugsweise andere regulatorische Elemente wie ribosomale Bindungsstellen und Terminationssequenzen sowie Sequenzen, die für selektive Marker und/oder Reporter-Gene codieren. Die Expression derartiger Selektionsmarker erleichtert die Identifizierung von Transformanten. Als Selektionsmarker geeignet sind Gene, die für eine Resistenz gegenüber z. B. Ampicillin, Tetracyclin, Chloramphenicol, Kanamycin oder anderen Antibiotika codieren. Wenn das erfindungsgemäße serA-Allel extrachromosomal repliziert werden soll, sollte der Plasmidvektor vorzugsweise einen Ursprungspunkt der Replikation enthalten. Besonders bevorzugt sind Plasmidvektoren wie beispielsweise die E. coli-Vektoren pACYC184, pUC18, pQE-70, pBR322, pSC101 und ihre Derivate. Als induzierbare Promotoren eignen sich beispielsweise der lac-, tac-, trc-, lambda PL, ara- oder tet-Promotor oder davon abgeleitete Sequenzen.For cloning the serA alleles according to the invention, vectors which already contain genetic elements (e.g. constitutive or regulatable promoters, terminators), either an ongoing or a controlled, allow inducible expression of the gene coding for the PGD. They are also located on an expression vector preferably other regulatory elements such as ribosomal Binding sites and termination sequences as well as sequences that code for selective markers and / or reporter genes. The expression Such selection marker facilitates identification of transformants. Suitable as selection markers Genes that are resistant to z. B. ampicillin, Tetracycline, chloramphenicol, kanamycin or other antibiotics encode. When the serA allele according to the invention is extrachromosomal should be replicated, the plasmid vector should preferably contain an origin point of replication. Especially preferred are plasmid vectors such as the E. coli vectors pACYC184, pUC18, pQE-70, pBR322, pSC101 and their derivatives. Suitable inducible promoters are, for example the lac, tac, trc, lambda PL, ara or tet promoter or sequences derived therefrom.
Des weiteren sind Plasmidvektoren besonders bevorzugt, die bereits ein Gen/Allel enthalten, dessen Einsatz ebenfalls zu einer Überproduktion von Aminosäuren der Phosphoglycerat-Familie bzw. von Verbindungen, die sich aus dem C1-Stoffwechsel ableiten, führt, wie beispielsweise für die Produktion von:
- L-Serin (z.B. serB-, serC-, Exportcarrier-Gen wie beschreiben in DE10044831A1)
- N-Acetyl-Serin, O-Acetyl-Serin, Cystin, Cystein oder Cysteinderivaten (z.B. cysE-Allele wie beschrieben in WO97/15673, Efflux-Gene wie beschrieben in EP0885962A1, cysB-Gen wie beschrieben in DE19949579C1, yfiK-Gen wie beschrieben in DE 10232930A)
- L-Tryptophan (z.B. trpE-Allele wie beschrieben in EP0662143A)
- Pantothensäure (z.B. wie beschrieben in WO02061108)
- L-serine (eg serB, serC, export carrier gene as described in DE10044831A1)
- N-acetyl-serine, O-acetyl-serine, cystine, cysteine or cysteine derivatives (eg cysE alleles as described in WO97 / 15673, efflux genes as described in EP0885962A1, cysB gene as described in DE19949579C1, yfiK gene as described in DE 10232930A)
- L-tryptophan (eg trpE alleles as described in EP0662143A)
- Pantothenic acid (eg as described in WO02061108)
Solche Vektoren ermöglichen die direkte Herstellung von erfindungsgemäßen Mikroorganismenstämmen mit hoher Produktionsleistung aus einem beliebigen Mikroorganismenstamm, da ein solches Plasmid auch die Aufhebung von anderen Restriktionen des Stoffwechselweges in einem Mikroorganismus bewirkt.Such vectors allow the direct production of inventive Microorganism strains with high production capacity from any microorganism strain, as such Plasmid also the lifting of other restrictions of the Metabolic pathway in a microorganism causes.
Durch eine gängige Transformationsmethode (z.B. Elektroporation) werden die erfindungsgemäßen serA-Allel-haltigen Plasmide in Mikroorganismen eingebracht und beispielsweise mittels Antibiotika-Resistenz auf plasmid-tragende Klone selektiert.By a common transformation method (e.g., electroporation) become the serA allele-containing plasmids according to the invention incorporated in microorganisms and, for example, by antibiotic resistance selected for plasmid-carrying clones.
Die Erfindung betrifft somit auch Verfahren zur Herstellung eines erfindungsgemäßen Mikroorganismenstammes, dadurch gekennzeichnet, dass in einen Mikroorganismenstamm ein erfindungsgemäßer Vektor eingebracht wird. The invention thus also relates to processes for the preparation a microorganism strain according to the invention, characterized that in a microorganism strain an inventive Vector is introduced.
Es ist auch möglich Vektoren mit einem erfindungsgemäßen serA-Allel in Mikroorganismen einzubringen, die beispielsweise einzelne oder mehrere der oben genannten Gene/Allele bereits chromosomal exprimieren und bereits eine Überproduktion eines Stoffwechselprodukts aufweisen. In solchen Fällen kann durch das Einbringen eines erfindungsgemäßen serA-Allels die Produktionsleistung nochmals gesteigert werden.It is also possible vectors with a serA allele according to the invention in microorganisms, for example, single or more of the above genes / alleles already chromosomally express and already an overproduction of a Have metabolite. In such cases can through the introduction of a serA allele according to the invention the production output be increased again.
Generell sind als Wirtsorganismus für erfindungsgemäße Vektoren alle Organismen geeignet, die den Biosyntheseweg für Aminosäuren der Phosphoglycerat-Familie aufweisen, rekombinanten Verfahren zugänglich sind und durch Fermentation kultivierbar sind. Solche Mikroorganismen können Pilze, Hefen oder Bakterien sein. Bevorzugt kommen Bakterien der phylogenetischen Gruppe der Eubacteria zum Einsatz. Besonders bevorzugt sind Mikroorganismen der Familie Enterobacteriaceae und insbesondere der Art Escherichia coli.Generally, as a host organism for vectors of the invention all organisms suitable for the biosynthetic pathway for amino acids the phosphoglycerate family, recombinant Process are accessible and cultivable by fermentation are. Such microorganisms can be fungi, yeasts or bacteria be. Preference is given to bacteria of the phylogenetic group Eubacteria are used. Particularly preferred are microorganisms the family Enterobacteriaceae and in particular the Species Escherichia coli.
Die Erfindung betrifft somit ferner einen Mikroorganismenstamm, der zur fermentativen Herstellung von Aminosäuren der Phosphoglycerat-Familie oder deren Derivaten bzw. von Verbindungen, die sich aus dem C1 Stoffwechsel ableiten, geeignet ist, dadurch gekennzeichnet, dass er eine erfindungsgemäße PGD besitzt.The invention thus further relates to a microorganism strain, for the fermentative production of amino acids of Phosphoglycerate family or their derivatives or compounds, which derive from the C1 metabolism suitable is, characterized in that it is a PGD according to the invention has.
Ein weiterer Gegenstand der Erfindung ist die Herstellung von Aminosäuren der Phosphoglycerat-Familie bzw. von Verbindungen, die sich aus dem C1-Stoffwechsel ableiten, durch Kultivierung eines erfindungsgemäßen Mikroorganismenstammes.Another object of the invention is the production of Amino acids of the phosphoglycerate family or of compounds, which derive from the C1 metabolism, by cultivation a microorganism strain according to the invention.
Dazu wird der erfindungsgemäße Mikroorganismenstamm beispielsweise in einem Fermenter in einem Nährmedium kultiviert, das eine geeignete Kohlenstoff-, und eine geeignete Energiequelle, sowie andere Zusatzstoffe enthält.For this purpose, the microorganism strain according to the invention, for example cultured in a fermenter in a nutrient medium, the a suitable carbon, and a suitable source of energy, and other additives.
Die während der Fermentation gebildeten Substanzen wie beispielsweise L-Phosphoserin, L-Serin, O-Acetyl-L-Serin, L-Cystein, Glycin, L-Tryptophan, 1,2,4-Triazol-2-yl-L-alanin, L-Methionin oder Pantothensäure können anschließend aufgereinigt werden.The substances formed during the fermentation such as L-phosphoserine, L-serine, O-acetyl-L-serine, L-cysteine, Glycine, L-tryptophan, 1,2,4-triazol-2-yl-L-alanine, L-methionine or pantothenic acid can subsequently be purified become.
Die folgenden Beispiele dienen der weiteren Erläuterung der Erfindung. Sämtliche eingesetzten molekularbiologischen Verfahren, wie Polymerase-Kettenreaktion, Isolierung und Reinigung von DNS, Modifikation von DNS durch Restriktionsenzyme, Klenow-Fragment und Ligase, Transformation etc. wurden in der dem Fachmann bekannten, in der Literatur beschriebenen oder von den jeweiligen Herstellern empfohlenen Art und Weise durchgeführt.The following examples serve to further explain the Invention. All molecular biological procedures used, like polymerase chain reaction, isolation and purification DNA, modification of DNA by restriction enzymes, Klenow fragment and ligase, transformation, etc. were in the known in the art, described in the literature or recommended by the respective manufacturers way carried out.
Das serA-Gen aus Escherichia coli Stamm W3110 (American Type Culture Collection, ATCC27325) wurde mit Hilfe der Polymerase-Ketten-Reaktion amplifiziert. Als spezifische Primer dienten die Oligonukleotide und The serA gene from Escherichia coli strain W3110 (American Type Culture Collection, ATCC27325) was amplified by the polymerase chain reaction. The oligonucleotides were used as specific primers and
Das resultierende DNS-Fragment wurde mit den Restriktionsenzymen NdeI und HindIII verdaut und die 5'-Überhänge mit Klenow-Enzym aufgefüllt. Anschließend wurde das DNS-Fragment mit Hilfe einer Agarose-Gelelektrophorese gereinigt und mit der GeneClean-Methode isoliert (GeneClean Kit BI0101 P.O.Box 2284 La Jolla, California, 92038-2284). Die Klonierung des so erhaltenen serA-Fragments erfolgte in den Expressionsvektor pQE-70 (Qiagen, Hilden, D). Hierfür wurde der Vektor zunächst mit SphI und BamHI geschnitten und der 3'-Überhang mit Klenow-Enzym abgedaut bzw. der 5'-Überhang mit Klenow-Enzym aufgefüllt. Anschließend wurde das Vektorfragment gereinigt und mit dem serA-Fragment ligiert. Der resultierende Vektor trägt die Bezeichnung pFL209. Nach der Verifizierung des Konstrukts durch Sequenzierung wurde der Escherichia coli Stamm JM109 (Stratagene, Amsterdam, NL) transformiert und entsprechende Transformanten mit Ampicillin selektiert. Der Bakterienstamm Escherichia coli JM109 / pFL209 wurde bei der DSMZ (Deutsche Sammlung für Mikroorganismen und Zellkulturen GmbH, D-38142 Braunschweig) unter der Nummer DSM 15628 gemäß Budapester Vertrag hinterlegt.The resulting DNA fragment was digested with the restriction enzymes NdeI and HindIII digested and the 5 'overhangs with Klenow enzyme refilled. Subsequently, the DNA fragment was using purified by agarose gel electrophoresis and using the GeneClean method isolated (GeneClean Kit BI0101 P.O.Box 2284 La Jolla, California, 92038-2284). The cloning of the thus obtained serA fragment was inserted into the expression vector pQE-70 (Qiagen, Hilden, D). For this, the vector was first with SphI and BamHI cut and the 3 'overhang with Klenow enzyme or the 5 'overhang filled with Klenow enzyme. Subsequently, the vector fragment was purified and washed with ligated to the serA fragment. The resulting vector carries the Designation pFL209. After verification of the construct by sequencing the Escherichia coli strain JM109 (Stratagene, Amsterdam, NL) and corresponding Transformants selected with ampicillin. The bacterial strain Escherichia coli JM109 / pFL209 was used in the DSMZ (German Collection for microorganisms and cell cultures GmbH, D-38142 Braunschweig) under the number DSM 15628 according to the Budapest Treaty deposited.
Die ortsspezifische Mutagenese an den Codons 349 und 372 des serA-Gens wurde mittels einer inversen Polymerase-Ketten-Reaktion durchgeführt. Als Matrize diente der in Beispiel 1 beschriebene Vektor pFL209. Für die Mutagenese des Codons 349 wurden die Primer und verwendet.Site-directed mutagenesis at codons 349 and 372 of the serA gene was performed by an inverse polymerase chain reaction. The template used was the vector pFL209 described in Example 1. For the mutagenesis of codon 349, the primers were and used.
Das erhaltene PCR-Produkt wurde durch Ligation zirkularisiert
und in den E. coli-Stamm JM109 transformiert. Durch Sequenzierung
wurde schließlich die Mutation an Codon 349 bestimmt und
die Korrektheit der übrigen Sequenz überprüft.
Für die Mutagenese des Codons 372 wurde prinzipiell gleich
verfahren, jedoch wurden die Primer
und
verwendet.The resulting PCR product was circularized by ligation and transformed into E. coli strain JM109. Sequencing finally determined the mutation at codon 349 and checked the correctness of the remaining sequence.
In principle the same procedure was followed for the mutagenesis of codon 372, but the primers became and used.
Zur Bestimmung von PGD-Enzymaktivitäten und des Einflusses von
L-Serin auf die Aktivität wurden 100 ml LB-Medium (10 g/l
Trypton, 5 g/l Hefeextrakt, 10 g/l NaCl), das zusätzlich 100
mg/l Ampicillin enthielt, mit einer 2 ml Übernachtkultur der
Stämme mit den plasmidcodierten serA-Allelen beimpft und bei
30 °C und 150 rpm in einem Schüttler inkubiert. Bei einer optischen
Dichte von 1,0 wurde die serA-Expression durch Zugabe
von 0,4 mM Isopropyl-β-thiogalaktosid induziert und die Kultur
für weitere 3 Stunden inkubiert. Die Zellen wurden anschließend
durch Zentrifugation geerntet, gewaschen und in 2 ml Puffer
(100 mM K-Phosphat pH 7,0; 10 mM MgCl2; 1 mM Dithiothreitol)
resuspendiert. Der Zellaufschluss erfolgte mittels einer
French Press (Spectronic Instruments, Inc., Rochester, NY,
USA) bei einem Druck von 18 000 psi. Die Rohextrakte wurden
durch Zentrifugation bei 30 000 g geklärt und die PGD-Aktivität
mit dem Test von McKitrick und Pizer (1980,
J.Bacteriol. 141:235-245) bestimmt.
Die folgenden Tabellen zeigen die PGD-Aktivität verschiedener
Mutanten, sowie die entsprechenden Inhibitorkonstanten Ki.
The following tables show the PGD activity of different mutants, as well as the corresponding inhibitor constants K i .
Eine Kombination von Mutationen des Codons 349 bzw. 372 sollte
zeigen, ob die Austausche einen synergistischen Effekt auf die
Feedback-Resistenz haben. Hierfür wurde eine singuläre HindII
Restriktionsschnittstelle zwischen den beiden Mutationsorten
benützt. So wurde durch HindII-HindIII-Restriktion des Vektors
mit dem serA20-Allel ein 183 bp-Fragment isoliert das dem 3'-Ende
des serA-Gens entspricht und die Mutation an Codon 372
beinhaltet. Dieses Fragment wurde in einen ebenfalls HindII-BamHI-verdauten
Vektor mit dem serA40-Allel kloniert und so
ein Klon erhalten der eine Doppelmutante darstellt. Die folgende
Tabelle zeigt die zugehörigen Enzymdaten.
Claims (10)
Priority Applications (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP08152747A EP1950287B1 (en) | 2003-07-10 | 2004-07-01 | 3-Phosphoglycerate dehydrogenase variants whose inhibition by L-serine is reduced, and genes encoding them |
| EP07102448A EP1813669B1 (en) | 2003-07-10 | 2004-07-01 | 3-Phosphoglycerate dehydrogenase variants whose inhibition by L-serine is reduced, and genes encoding them |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| DE10331291 | 2003-07-10 | ||
| DE10331291A DE10331291A1 (en) | 2003-07-10 | 2003-07-10 | Variants of 3-phosphoglycerate dehydrogenase with reduced inhibition by L-serine and genes coding for it |
Related Child Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| EP07102448A Division EP1813669B1 (en) | 2003-07-10 | 2004-07-01 | 3-Phosphoglycerate dehydrogenase variants whose inhibition by L-serine is reduced, and genes encoding them |
Publications (3)
| Publication Number | Publication Date |
|---|---|
| EP1496111A2 true EP1496111A2 (en) | 2005-01-12 |
| EP1496111A3 EP1496111A3 (en) | 2005-03-09 |
| EP1496111B1 EP1496111B1 (en) | 2007-05-02 |
Family
ID=33441709
Family Applications (3)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| EP07102448A Expired - Lifetime EP1813669B1 (en) | 2003-07-10 | 2004-07-01 | 3-Phosphoglycerate dehydrogenase variants whose inhibition by L-serine is reduced, and genes encoding them |
| EP04015524A Expired - Lifetime EP1496111B1 (en) | 2003-07-10 | 2004-07-01 | Variants of 3-phosphoglycerate dehydrogenase with reduced inhibition by L-serine and genes encoding the same. |
| EP08152747A Expired - Lifetime EP1950287B1 (en) | 2003-07-10 | 2004-07-01 | 3-Phosphoglycerate dehydrogenase variants whose inhibition by L-serine is reduced, and genes encoding them |
Family Applications Before (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| EP07102448A Expired - Lifetime EP1813669B1 (en) | 2003-07-10 | 2004-07-01 | 3-Phosphoglycerate dehydrogenase variants whose inhibition by L-serine is reduced, and genes encoding them |
Family Applications After (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| EP08152747A Expired - Lifetime EP1950287B1 (en) | 2003-07-10 | 2004-07-01 | 3-Phosphoglycerate dehydrogenase variants whose inhibition by L-serine is reduced, and genes encoding them |
Country Status (7)
| Country | Link |
|---|---|
| US (1) | US7582460B2 (en) |
| EP (3) | EP1813669B1 (en) |
| JP (1) | JP4695855B2 (en) |
| CN (1) | CN100526457C (en) |
| AT (3) | ATE440946T1 (en) |
| DE (4) | DE10331291A1 (en) |
| ES (2) | ES2283906T3 (en) |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2007135188A2 (en) | 2006-05-24 | 2007-11-29 | Evonik Degussa Gmbh | Process for the preparation of l-methionine |
| WO2010076324A1 (en) * | 2008-12-31 | 2010-07-08 | Metabolic Explorer | Method for the preparation of diols |
| WO2012152664A1 (en) * | 2011-05-11 | 2012-11-15 | Wacker Chemie Ag | Method for producing l-cystine by fermentation under controlled oxygen saturation |
| WO2013171098A3 (en) * | 2012-05-18 | 2014-01-09 | Wacker Chemie Ag | Method for producing l-cysteine using a benzoic acid-containing culture medium |
Families Citing this family (16)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP5332237B2 (en) * | 2008-03-06 | 2013-11-06 | 味の素株式会社 | L-cysteine producing bacterium and method for producing L-cysteine |
| US8383372B2 (en) * | 2008-03-06 | 2013-02-26 | Ajinomoto Co., Inc. | L-cysteine producing bacterium and a method for producing L-cysteine |
| WO2011065469A1 (en) | 2009-11-30 | 2011-06-03 | 味の素株式会社 | L-cysteine-producing bacterium, and process for production of l-cysteine |
| EP2617808B1 (en) | 2010-09-14 | 2016-06-15 | Ajinomoto Co., Inc. | Sulfur-containing amino acid-producing bacterium and method for producing sulfur-containing amino acids |
| JP2014087259A (en) | 2011-02-22 | 2014-05-15 | Ajinomoto Co Inc | L-cysteine-producing bacterium, and production method of l-cysteine |
| US9234223B2 (en) | 2011-04-01 | 2016-01-12 | Ajinomoto Co., Inc. | Method for producing L-cysteine |
| WO2012137689A1 (en) | 2011-04-01 | 2012-10-11 | 味の素株式会社 | Method for producing l-cysteine |
| JP2014131487A (en) | 2011-04-18 | 2014-07-17 | Ajinomoto Co Inc | Method for producing l-cysteine |
| DE102011078481A1 (en) | 2011-06-30 | 2013-01-03 | Wacker Chemie Ag | Process for the fermentative production of natural L-cysteine |
| CN102433312A (en) * | 2011-11-30 | 2012-05-02 | 天津科技大学 | D-3-phosphoglycerate-dehydrogenase as well as coding gene and construction method thereof |
| DE102012216527A1 (en) | 2012-09-17 | 2014-03-20 | Wacker Chemie Ag | Process for the fermentative production of L-cysteine and derivatives of this amino acid |
| DE102013209274A1 (en) | 2013-05-17 | 2014-11-20 | Wacker Chemie Ag | Microorganism and method for fermentative overproduction of gamma-glutamylcysteine and derivatives of this dipeptide |
| CN103436504A (en) * | 2013-09-02 | 2013-12-11 | 江南大学 | Construction method and application of corynebacterium glutamicum SYPS-062 resistant to feedback inhibition on L-serine |
| KR20230025010A (en) | 2020-06-26 | 2023-02-21 | 와커 헤미 아게 | Improved Cysteine Producing Strains |
| JP2023547024A (en) | 2021-07-05 | 2023-11-09 | ワッカー ケミー アクチエンゲゼルシャフト | Method for enzymatic oxidation of sulfinic acid to sulfonic acid |
| CN118742558A (en) | 2022-03-01 | 2024-10-01 | 瓦克化学股份公司 | Improved cysteine producing strains |
Citations (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1993012235A1 (en) * | 1991-12-12 | 1993-06-24 | Wacker-Chemie Gmbh | Materials and methods for biosynthesis of serine and serine-related products |
| WO1994008031A1 (en) * | 1992-09-28 | 1994-04-14 | Consortium für elektrochemische Industrie GmbH | Microorganisms for the production of tryptophan and process for producing the same |
Family Cites Families (9)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| DE19539952A1 (en) | 1995-10-26 | 1997-04-30 | Consortium Elektrochem Ind | Process for the preparation of O-acetylserine, L-cysteine and L-cysteine-related products |
| DE19726083A1 (en) | 1997-06-19 | 1998-12-24 | Consortium Elektrochem Ind | Microorganisms and processes for the fermentative production of L-cysteine, L-cystine, N-acetyl-serine or thiazolidine derivatives |
| JP4066543B2 (en) * | 1998-01-12 | 2008-03-26 | 味の素株式会社 | Method for producing L-serine by fermentation |
| JP3997631B2 (en) | 1998-01-12 | 2007-10-24 | 味の素株式会社 | Method for producing L-serine by fermentation |
| DE19949579C1 (en) | 1999-10-14 | 2000-11-16 | Consortium Elektrochem Ind | Microorganism with deregulated cysteine metabolism, useful for high-level production of cysteine and its derivatives, has increased activity of the CysB transcription regulator |
| DE10044831A1 (en) | 2000-03-01 | 2002-04-04 | Forschungszentrum Juelich Gmbh | Improved process for the microbial production of L-serine and a suitable genetically modified microorganism |
| DE10046934A1 (en) | 2000-09-21 | 2002-04-18 | Consortium Elektrochem Ind | Process for the fermentative production of non-proteinogenic L-amino acids |
| AU2002243526A (en) | 2001-01-19 | 2002-08-12 | Basf Ag | Microorganisms and processes for enhanced production of pantothenate |
| DE10232930A1 (en) | 2002-07-19 | 2004-02-05 | Consortium für elektrochemische Industrie GmbH | Process for the fermentative production of amino acids and amino acid derivatives of the phosphoglycerate family |
-
2003
- 2003-07-10 DE DE10331291A patent/DE10331291A1/en not_active Withdrawn
-
2004
- 2004-07-01 EP EP07102448A patent/EP1813669B1/en not_active Expired - Lifetime
- 2004-07-01 AT AT08152747T patent/ATE440946T1/en active
- 2004-07-01 EP EP04015524A patent/EP1496111B1/en not_active Expired - Lifetime
- 2004-07-01 ES ES04015524T patent/ES2283906T3/en not_active Expired - Lifetime
- 2004-07-01 AT AT07102448T patent/ATE431847T1/en active
- 2004-07-01 ES ES07102448T patent/ES2325603T3/en not_active Expired - Lifetime
- 2004-07-01 DE DE502004009513T patent/DE502004009513D1/en not_active Expired - Lifetime
- 2004-07-01 DE DE502004003661T patent/DE502004003661D1/en not_active Expired - Lifetime
- 2004-07-01 DE DE502004009985T patent/DE502004009985D1/en not_active Expired - Lifetime
- 2004-07-01 AT AT04015524T patent/ATE361361T1/en active
- 2004-07-01 EP EP08152747A patent/EP1950287B1/en not_active Expired - Lifetime
- 2004-07-08 US US10/886,858 patent/US7582460B2/en active Active
- 2004-07-09 CN CNB2004100635275A patent/CN100526457C/en not_active Expired - Lifetime
- 2004-07-12 JP JP2004205113A patent/JP4695855B2/en not_active Expired - Lifetime
Patent Citations (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1993012235A1 (en) * | 1991-12-12 | 1993-06-24 | Wacker-Chemie Gmbh | Materials and methods for biosynthesis of serine and serine-related products |
| WO1994008031A1 (en) * | 1992-09-28 | 1994-04-14 | Consortium für elektrochemische Industrie GmbH | Microorganisms for the production of tryptophan and process for producing the same |
Non-Patent Citations (5)
| Title |
|---|
| AL-RABIEE REGINA ET AL: "The mechanism of velocity modulated allosteric regulation in D-3-phosphoglycerate dehydrogenase: Cross-linking adjacent regulatory domains with engineered disulfides mimics effector binding" JOURNAL OF BIOLOGICAL CHEMISTRY, Bd. 271, Nr. 22, 1996, Seiten 13013-13017, XP002295412 ISSN: 0021-9258 * |
| GRANT GREGORY A ET AL: "Probing the regulatory domain interface of D-3-phosphoglycerate dehydrogenase with engineered tryptophan residues" JOURNAL OF BIOLOGICAL CHEMISTRY, Bd. 273, Nr. 35, 28. August 1998 (1998-08-28), Seiten 22389-22394, XP002295411 ISSN: 0021-9258 * |
| GRANT GREGORY A ET AL: "Role of an interdomain Gly-Gly sequence at the regulatory-substrate domain interface in the regulation of Escherichia coli. D-3-phosphoglycerate dehydrogenase" BIOCHEMISTRY, Bd. 39, Nr. 24, 20. Juni 2000 (2000-06-20), Seiten 7316-7319, XP002312560 ISSN: 0006-2960 * |
| GRANT GREGORY A ET AL: "Specific interactions at the regulatory domain-substrate binding domain interface influence the cooperativity of inhibition and effector binding in Escherichia coli D-3-phosphoglycerate dehydrogenase" JOURNAL OF BIOLOGICAL CHEMISTRY, Bd. 276, Nr. 2, 12. Januar 2001 (2001-01-12), Seiten 1078-1083, XP002295413 ISSN: 0021-9258 * |
| SCHULLER D J ET AL: "THE ALLOSTERIC LIGAND SITE IN THE VMAX-TYPE COOPERATIVE ENZYME PHOSPHOGLYCERATE DEHYDROGENASE" NATURE STRUCTURAL BIOLOGY, NEW YORK, NY, US, Bd. 2, Nr. 1, 1. Januar 1995 (1995-01-01), Seiten 69-76, XP009036197 ISSN: 1072-8368 * |
Cited By (9)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2007135188A2 (en) | 2006-05-24 | 2007-11-29 | Evonik Degussa Gmbh | Process for the preparation of l-methionine |
| WO2007135188A3 (en) * | 2006-05-24 | 2008-03-06 | Basf Ag | Process for the preparation of l-methionine |
| WO2010076324A1 (en) * | 2008-12-31 | 2010-07-08 | Metabolic Explorer | Method for the preparation of diols |
| US9121041B2 (en) | 2008-12-31 | 2015-09-01 | Metabolic Explorer | Method for the preparation of diols |
| WO2012152664A1 (en) * | 2011-05-11 | 2012-11-15 | Wacker Chemie Ag | Method for producing l-cystine by fermentation under controlled oxygen saturation |
| CN103517989A (en) * | 2011-05-11 | 2014-01-15 | 瓦克化学股份公司 | Method for producing l-cystine by fermentation under controlled oxygen saturation |
| US9074230B2 (en) | 2011-05-11 | 2015-07-07 | Wacker Chemie Ag | Method for producing L-cystine by fermentation under controlled oxygen saturation |
| CN103517989B (en) * | 2011-05-11 | 2015-11-25 | 瓦克化学股份公司 | By the method for fermentative production CYSTINE under in check oxygen saturation |
| WO2013171098A3 (en) * | 2012-05-18 | 2014-01-09 | Wacker Chemie Ag | Method for producing l-cysteine using a benzoic acid-containing culture medium |
Also Published As
| Publication number | Publication date |
|---|---|
| US20050009162A1 (en) | 2005-01-13 |
| EP1813669A1 (en) | 2007-08-01 |
| DE502004009513D1 (en) | 2009-07-02 |
| EP1950287A2 (en) | 2008-07-30 |
| DE502004003661D1 (en) | 2007-06-14 |
| ES2283906T3 (en) | 2007-11-01 |
| EP1813669B1 (en) | 2009-05-20 |
| DE502004009985D1 (en) | 2009-10-08 |
| ATE440946T1 (en) | 2009-09-15 |
| EP1496111B1 (en) | 2007-05-02 |
| JP4695855B2 (en) | 2011-06-08 |
| EP1950287B1 (en) | 2009-08-26 |
| CN1609208A (en) | 2005-04-27 |
| ES2325603T3 (en) | 2009-09-09 |
| EP1496111A3 (en) | 2005-03-09 |
| CN100526457C (en) | 2009-08-12 |
| JP2005040134A (en) | 2005-02-17 |
| ATE431847T1 (en) | 2009-06-15 |
| EP1950287A3 (en) | 2008-08-13 |
| US7582460B2 (en) | 2009-09-01 |
| ATE361361T1 (en) | 2007-05-15 |
| DE10331291A1 (en) | 2005-02-17 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| EP1496111B1 (en) | Variants of 3-phosphoglycerate dehydrogenase with reduced inhibition by L-serine and genes encoding the same. | |
| EP1382684B1 (en) | Process for fermentative production of amino acids and amino acid derivatives of the phosphoglycerate family | |
| EP0387527B1 (en) | Process for the fermentative production of L-lysine | |
| DE102004001674B4 (en) | Process for the preparation of L-lysine using methanol-utilizing bacteria | |
| EP0858510B1 (en) | Process for preparing o-acetylserine, l-cysteine and l-cysteine-related products | |
| DE60207685T2 (en) | PROCESS FOR THE PREPARATION OF L-THREONINE | |
| EP1155139B2 (en) | Method for microbially producing l-valine | |
| EP1220940A1 (en) | Method for production of l-cysteine or l-cysteine derivatives by fermentation | |
| DE19855312A1 (en) | Process for the fermentative production of D-pantothenic acid using coryneform bacteria | |
| CA2262813A1 (en) | Novel strains of escherichia coli, methods of preparing the same and use thereof in fermentation processes for l-threonine production | |
| WO1995019442A1 (en) | Production of l-isoleucine by means of recombinant micro-organisms with deregulated threonine dehydratase | |
| EP1445310B1 (en) | Process for the fermentative production of L-methionine | |
| DE19846499A1 (en) | Production of microorganisms that overproduce pantothenic acid, useful as vitamin in e.g. foods or medicines, by overexpressing sequences that encode ketopantothenate reductase | |
| EP1570066B1 (en) | Feedback-resistant homoserine transsuccinylases with a modified c-terminal | |
| EP1924694B1 (en) | Method for producing amino acids using micro-organisms | |
| WO2015132213A1 (en) | Process for preparing terminal amino carboxylic acids and amino aldehydes by means of a recombinant microorganism | |
| EP1549754A2 (en) | Feedback-resistant homoserine transsuccinylases | |
| KR101755349B1 (en) | Microorganism producing L-threonine and process for producing L-threonine using the same | |
| DE102005049527B4 (en) | Process for the preparation of L-serine, gene sequence, vectors and microorganism | |
| DE10220234B4 (en) | Processes and microorganisms for the microbial production of pyruvate from carbohydrates and alcohols | |
| WO2004007705A1 (en) | Nucleotide sequences that encode deregulated phosphoglycerate dehydrogenases of coryneform bacteria and method for producing l-serine | |
| KR20160029964A (en) | A microorganism having enhanced L-lysine productivity and a method of producing L-lysine using the same | |
| DE10101501A1 (en) | Process for the fermentative production of pantothenic acid |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PUAI | Public reference made under article 153(3) epc to a published international application that has entered the european phase |
Free format text: ORIGINAL CODE: 0009012 |
|
| 17P | Request for examination filed |
Effective date: 20040701 |
|
| AK | Designated contracting states |
Kind code of ref document: A2 Designated state(s): AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HU IE IT LI LU MC NL PL PT RO SE SI SK TR |
|
| AX | Request for extension of the european patent |
Extension state: AL HR LT LV MK |
|
| PUAL | Search report despatched |
Free format text: ORIGINAL CODE: 0009013 |
|
| RTI1 | Title (correction) |
Free format text: VARIANTS OF 3-PHOSPHOGLYCERATE DEHYDROGENASE WITH REDUCED INHIBITION BY L-SERINE AND GENES ENCODING THE SAME |
|
| AK | Designated contracting states |
Kind code of ref document: A3 Designated state(s): AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HU IE IT LI LU MC NL PL PT RO SE SI SK TR |
|
| AX | Request for extension of the european patent |
Extension state: AL HR LT LV MK |
|
| 17Q | First examination report despatched |
Effective date: 20050221 |
|
| AKX | Designation fees paid |
Designated state(s): AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HU IE IT LI LU MC NL PL PT RO SE SI SK TR |
|
| GRAP | Despatch of communication of intention to grant a patent |
Free format text: ORIGINAL CODE: EPIDOSNIGR1 |
|
| RTI1 | Title (correction) |
Free format text: VARIANTS OF 3-PHOSPHOGLYCERATE DEHYDROGENASE WITH REDUCED INHIBITION BY L-SERINE AND GENES ENCODING THE SAME. |
|
| GRAS | Grant fee paid |
Free format text: ORIGINAL CODE: EPIDOSNIGR3 |
|
| GRAA | (expected) grant |
Free format text: ORIGINAL CODE: 0009210 |
|
| AK | Designated contracting states |
Kind code of ref document: B1 Designated state(s): AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HU IE IT LI LU MC NL PL PT RO SE SI SK TR |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: FI Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 |
|
| REG | Reference to a national code |
Ref country code: GB Ref legal event code: FG4D Free format text: NOT ENGLISH |
|
| RAP2 | Party data changed (patent owner data changed or rights of a patent transferred) |
Owner name: WACKER CHEMIE AG |
|
| REG | Reference to a national code |
Ref country code: CH Ref legal event code: EP |
|
| REG | Reference to a national code |
Ref country code: IE Ref legal event code: FG4D Free format text: LANGUAGE OF EP DOCUMENT: GERMAN |
|
| REF | Corresponds to: |
Ref document number: 502004003661 Country of ref document: DE Date of ref document: 20070614 Kind code of ref document: P |
|
| NLT2 | Nl: modifications (of names), taken from the european patent patent bulletin |
Owner name: WACKER CHEMIE AG Effective date: 20070516 |
|
| GBT | Gb: translation of ep patent filed (gb section 77(6)(a)/1977) |
Effective date: 20070613 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: SE Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070802 |
|
| ET | Fr: translation filed | ||
| REG | Reference to a national code |
Ref country code: ES Ref legal event code: FG2A Ref document number: 2283906 Country of ref document: ES Kind code of ref document: T3 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: PL Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 |
|
| REG | Reference to a national code |
Ref country code: IE Ref legal event code: FD4D |
|
| BERE | Be: lapsed |
Owner name: CONSORTIUM FUR ELEKTROCHEMISCHE INDUSTRIE G.M.B.H. Effective date: 20070731 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: IE Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 Ref country code: CZ Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 Ref country code: DK Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 Ref country code: BG Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070802 Ref country code: SI Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 Ref country code: PT Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20071002 |
|
| PLBE | No opposition filed within time limit |
Free format text: ORIGINAL CODE: 0009261 |
|
| STAA | Information on the status of an ep patent application or granted ep patent |
Free format text: STATUS: NO OPPOSITION FILED WITHIN TIME LIMIT |
|
| 26N | No opposition filed |
Effective date: 20080205 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: GR Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070803 Ref country code: MC Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20070731 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: RO Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: BE Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20070731 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: EE Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: CY Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: LU Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20070701 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: HU Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20071103 Ref country code: TR Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20070502 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: SK Payment date: 20140630 Year of fee payment: 11 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: NL Payment date: 20140721 Year of fee payment: 11 Ref country code: CH Payment date: 20140721 Year of fee payment: 11 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: ES Payment date: 20140728 Year of fee payment: 11 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: IT Payment date: 20140724 Year of fee payment: 11 |
|
| REG | Reference to a national code |
Ref country code: FR Ref legal event code: PLFP Year of fee payment: 12 |
|
| REG | Reference to a national code |
Ref country code: CH Ref legal event code: PL |
|
| REG | Reference to a national code |
Ref country code: SK Ref legal event code: MM4A Ref document number: E 3015 Country of ref document: SK Effective date: 20150701 |
|
| REG | Reference to a national code |
Ref country code: NL Ref legal event code: MM Effective date: 20150801 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: SK Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20150701 Ref country code: CH Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20150731 Ref country code: IT Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20150701 Ref country code: LI Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20150731 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: NL Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20150801 |
|
| REG | Reference to a national code |
Ref country code: FR Ref legal event code: PLFP Year of fee payment: 13 |
|
| REG | Reference to a national code |
Ref country code: ES Ref legal event code: FD2A Effective date: 20160831 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: ES Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20150702 |
|
| REG | Reference to a national code |
Ref country code: FR Ref legal event code: PLFP Year of fee payment: 14 |
|
| REG | Reference to a national code |
Ref country code: FR Ref legal event code: PLFP Year of fee payment: 15 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: AT Payment date: 20190722 Year of fee payment: 16 |
|
| REG | Reference to a national code |
Ref country code: AT Ref legal event code: MM01 Ref document number: 361361 Country of ref document: AT Kind code of ref document: T Effective date: 20200701 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: AT Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20200701 |
|
| P01 | Opt-out of the competence of the unified patent court (upc) registered |
Effective date: 20230502 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: GB Payment date: 20230719 Year of fee payment: 20 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: FR Payment date: 20230725 Year of fee payment: 20 Ref country code: DE Payment date: 20230719 Year of fee payment: 20 |
|
| REG | Reference to a national code |
Ref country code: DE Ref legal event code: R071 Ref document number: 502004003661 Country of ref document: DE |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: GB Free format text: LAPSE BECAUSE OF EXPIRATION OF PROTECTION Effective date: 20240630 |
|
| REG | Reference to a national code |
Ref country code: GB Ref legal event code: PE20 Expiry date: 20240630 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: GB Free format text: LAPSE BECAUSE OF EXPIRATION OF PROTECTION Effective date: 20240630 |
|
| REG | Reference to a national code |
Ref country code: DE Ref legal event code: R081 Ref document number: 502004003661 Country of ref document: DE Owner name: WACKER CHEMIE AG, DE Free format text: FORMER OWNER: WACKER CHEMIE AG, 81737 MUENCHEN, DE |