WO2009013750A2 - Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same - Google Patents
Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same Download PDFInfo
- Publication number
- WO2009013750A2 WO2009013750A2 PCT/IL2008/001024 IL2008001024W WO2009013750A2 WO 2009013750 A2 WO2009013750 A2 WO 2009013750A2 IL 2008001024 W IL2008001024 W IL 2008001024W WO 2009013750 A2 WO2009013750 A2 WO 2009013750A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- plant
- plants
- seq
- nos
- gene
- Prior art date
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8241—Phenotypically and genetically modified plants via recombinant DNA technology
- C12N15/8261—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield
- C12N15/8271—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance
- C12N15/8273—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance for drought, cold, salt resistance
-
- A—HUMAN NECESSITIES
- A01—AGRICULTURE; FORESTRY; ANIMAL HUSBANDRY; HUNTING; TRAPPING; FISHING
- A01H—NEW PLANTS OR NON-TRANSGENIC PROCESSES FOR OBTAINING THEM; PLANT REPRODUCTION BY TISSUE CULTURE TECHNIQUES
- A01H1/00—Processes for modifying genotypes ; Plants characterised by associated natural traits
- A01H1/12—Processes for modifying agronomic input traits, e.g. crop yield
- A01H1/122—Processes for modifying agronomic input traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance
-
- A—HUMAN NECESSITIES
- A01—AGRICULTURE; FORESTRY; ANIMAL HUSBANDRY; HUNTING; TRAPPING; FISHING
- A01H—NEW PLANTS OR NON-TRANSGENIC PROCESSES FOR OBTAINING THEM; PLANT REPRODUCTION BY TISSUE CULTURE TECHNIQUES
- A01H5/00—Angiosperms, i.e. flowering plants, characterised by their plant parts; Angiosperms characterised otherwise than by their botanic taxonomy
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/415—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from plants
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8241—Phenotypically and genetically modified plants via recombinant DNA technology
- C12N15/8242—Phenotypically and genetically modified plants via recombinant DNA technology with non-agronomic quality (output) traits, e.g. for industrial processing; Value added, non-agronomic traits
- C12N15/8243—Phenotypically and genetically modified plants via recombinant DNA technology with non-agronomic quality (output) traits, e.g. for industrial processing; Value added, non-agronomic traits involving biosynthetic or metabolic pathways, i.e. metabolic engineering, e.g. nicotine, caffeine
- C12N15/8247—Phenotypically and genetically modified plants via recombinant DNA technology with non-agronomic quality (output) traits, e.g. for industrial processing; Value added, non-agronomic traits involving biosynthetic or metabolic pathways, i.e. metabolic engineering, e.g. nicotine, caffeine involving modified lipid metabolism, e.g. seed oil composition
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8241—Phenotypically and genetically modified plants via recombinant DNA technology
- C12N15/8261—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8241—Phenotypically and genetically modified plants via recombinant DNA technology
- C12N15/8261—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield
- C12N15/8271—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y02—TECHNOLOGIES OR APPLICATIONS FOR MITIGATION OR ADAPTATION AGAINST CLIMATE CHANGE
- Y02A—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE
- Y02A40/00—Adaptation technologies in agriculture, forestry, livestock or agroalimentary production
- Y02A40/10—Adaptation technologies in agriculture, forestry, livestock or agroalimentary production in agriculture
- Y02A40/146—Genetically Modified [GMO] plants, e.g. transgenic plants
Definitions
- the present invention in some embodiments thereof, relates to isolated polypeptides and polynucleotides and more particularly, but not exclusively, to methods of using same for increasing tolerance of a plant to abiotic stress, growth, biomass, vigor and/or yield of a plant.
- Abiotic stress (ABS; also referred to as "environmental stress”) conditions such as salinity, drought, flood, suboptimal temperature and toxic chemical pollution, cause substantial damage to agricultural plants.
- Most plants have evolved strategies to protect themselves against these conditions. However, if the severity and duration of the stress conditions are too great, the effects on plant development, growth and yield are profound. Furthermore, most of the crop plants are highly susceptible to ABS and thus necessitate optimal growth conditions for commercial crop yields. Continuous exposure to stress causes major alterations in plant's metabolism which ultimately leads to cell death and consequently yield loss. Thus, despite extensive research and intensive crop- protection measures, losses due to abiotic stress conditions remain in the billions of dollars annually.
- Drought is a gradual phenomenon, which involves periods of abnormally dry weather that persists long enough to produce serious hydrologic imbalances such as crop damage and water supply shortage.
- drought can last many years and result in devastating effects on agriculture and water supplies.
- drought is not only the number one weather-related problem in agriculture, but it also ranks as one of the major natural disasters of all time, causing not only economic damage (e.g., losses from the US drought of 1988 exceeded $40 billion), but also loss of human lives, as in the 1984-1985 drought in the Horn of Africa which led to a famine that killed 750,000 people.
- drought is associated with increase susceptibility to various diseases.
- Soil salinity is thus one of the more important variables that determine whether a plant may thrive.
- sizable land areas are uncultivable due to naturally high soil salinity.
- Salt tolerance is of particular importance early in a plant's lifecycle, since evaporation from the soil surface causes upward water movement, and salt accumulates in the upper soil layer where the seeds are placed.
- germination normally takes place at a salt concentration which is higher than the mean salt level in the whole soil profile.
- Germination of many crops is sensitive to temperature. A gene that would enhance germination in hot conditions would be useful for crops that are planted late in the season or in hot climates.
- seedlings and mature plants that are exposed to excess heat may experience heat shock, which may arise in various organs, including leaves and particularly fruit, when transpiration is insufficient to overcome heat stress. Heat also damages cellular structures, including organelles and cytoskeleton, and impairs membrane function. Heat shock may produce a decrease in overall protein synthesis, accompanied by expression of heat shock proteins, e.g., chaperones, which are involved in refolding proteins denatured by heat.
- heat shock proteins e.g., chaperones
- Heat stress often accompanies conditions of low water availability. Heat itself is seen as an interacting stress and adds to the detrimental effects caused by water deficit conditions. Water Evaporative demand exhibits near exponential increases with increases in daytime temperatures and can result in high transpiration rates and low plant water potentials. High-temperature damage to pollen almost always occurs in conjunction with drought stress, and rarely occurs under well-watered conditions. Combined stress can alter plant metabolism in novel ways; therefore understanding the interaction between different stresses may be important for the development of strategies to enhance stress tolerance by genetic manipulation.
- Excessive chilling conditions e.g., low, but above freezing, temperatures affect crops of tropical origins, such as soybean, rice, maize, and cotton.
- Typical chilling damage includes wilting, necrosis, chlorosis or leakage of ions from cell membranes.
- the underlying mechanisms of chilling sensitivity are not completely understood yet, but probably involve the level of membrane saturation and other physiological deficiencies. For example, photoinliibition of photosynthesis (disruption of photosynthesis due to high light intensities) often occurs under clear atmospheric conditions subsequent to cold late summer/autumn nights. In addition, chilling may lead to yield losses and lower product quality through the delayed ripening of maize.
- Water deficit is a common component of many plant stresses. Water deficit occurs in plant cells when the whole plant transpiration rate exceeds the water uptake. In addition to drought, other stresses, such as salinity and low temperature, produce cellular dehydration.
- Salt and drought stress signal transduction consist of ionic and osmotic homeostasis signaling pathways.
- the ionic aspect of salt stress is signaled via the SOS pathway where a calcium-responsive SOS3-SOS2 protein kinase complex controls the expression and activity of ion transporters such as SOSl.
- the osmotic component of salt stress involves complex plant reactions that overlap with drought and/or cold stress responses.
- Abscisic acid biosynthesis is regulated by osmotic stress at multiple steps. Both ABA-dependent and -independent osmotic stress signaling first modify constitutively expressed transcription factors, leading to the expression of early response transcriptional activators, which then activate downstream stress tolerance effector genes.
- genes which increase tolerance to cold or salt stress can also improve drought stress protection, these include for example, the transcription factor AtCBF/DREBl, OsCDPK7 (Saijo et al. 2000, Plant J. 23: 319-327) or AVPl (a vacuolar pyrophosphatase-proton pump, Gaxiola et al. 2001, Proc. Natl. Acad. Sci. USA 98: 11444-11449).
- Developing stress-tolerant plants is a strategy that has the potential to solve or mediate at least some of these problems.
- traditional plant breeding strategies used to develop new lines of plants that exhibit tolerance to ABS are relatively inefficient since they are tedious, time consuming and of unpredictable outcome.
- limited germplasm resources for stress tolerance and incompatibility in crosses between distantly related plant species represent significant problems encountered in conventional breeding.
- the cellular processes leading to ABS tolerance are complex in nature and involve multiple mechanisms of cellular adaptation and numerous metabolic pathways.
- U.S. patents and patent applications also describe polynucleotides associated with stress tolerance and their use in generating stress tolerant plants.
- U.S. Pat. Nos. 5,296,462 and 5,356,816 describe transforming plants with polynucleotides encoding proteins involved in cold adaptation in Arabidopsis thaliana for promoting cold tolerance.
- U.S. Pat. No. 6,670,528 describes transforming plants with polynucleotides encoding polypeptides binding to stress responsive elements for promoting tolerance to abiotic stress.
- U.S. Pat. No. 6,720,477 describes transforming plants with a polynucleotide encoding a signal transduction stress-related protein, capable of increasing tolerance of the transformed plants to abiotic stress.
- U.S. Application Ser. Nos. 09/938842 and 10/342224 describe abiotic stress- related genes and their use to confer upon plants tolerance to abiotic stress.
- WO2004/104162 to Evogene Ltd teaches polynucleotide sequences and methods of utilizing same for increasing the tolerance of a plant to abiotic stresses and/or increasing the biomass of a plant.
- WO2007/020638 to Evogene Ltd. teaches polynucleotide sequences and methods of utilizing same for increasing the tolerance of a plant to abiotic stresses and/or increasing the biomass, vigor and/or yield of a plant.
- WO2007/049275 to Evogene Ltd. teaches isolated polypeptides, polynucleotides encoding same for increasing tolerance of a plant to abiotic stress, and/or for increasing biomass, vigor and/or yield of a plant.
- a method of increasing tolerance of a plant to abiotic stress comprising expressing within the plant an exogenous polynucleotide encoding a polypeptide comprising an amino acid sequence at least 90 % homologous to the amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202- 206, 208-211, 213-391, 1655, 961-1529, and 1660-1663, thereby increasing the tolerance of the plant to abiotic stress.
- a method of increasing tolerance of a plant to abiotic stress comprising expressing within the plant an exogenous polynucleotide encoding a polypeptide comprising an amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, and 1660- 1663, thereby increasing the tolerance of the plant to abiotic stress.
- a method of increasing biomass, growth rate, vigor and/or yield of a plant comprising expressing within the plant an exogenous polynucleotide encoding a polypeptide comprising an amino acid sequence at least 90 % homologous to the amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202- 206, 208-211, 213-391, 1655, 961-1529, and 1660-1663, thereby increasing the biomass, growth rate, vigor and/or yield of the plant.
- a method of increasing biomass, growth rate, vigor and/or yield of a plant comprising expressing within the plant an exogenous polynucleotide encoding a polypeptide comprising an amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, and 1660- 1663, thereby increasing the biomass, growth rate, vigor and/or yield of the plant.
- an isolated polynucleotide comprising a nucleic acid sequence at least 90 % identical to the nucleic acid sequence selected from the group consisting of SEQ ID NOs:1530, 1561, 1532, 1531, 1562, 1533, 1538, 1549, 1665, 1566, 1554, 1563, 1557, 1564, 1534, 1536, 1552, 1553, 1666, 1547, 1548, 1556, 1559, 1560, 1654, 1555, 1540, 1543, 1668, 1539, 1550, 1558, 1565, 1541, 1667, 1542, 1544, 1537, 1551, 1545, 1-200, 1653, 392-960, and 1656-1659.
- an isolated polynucleotide comprising a nucleic acid sequence selected from the group consisting of SEQ ID NOs:1530, 1561, 1532, 1531, 1562, 1533, 1538, 1549, 1665, 1566, 1554, 1563, 1557, 1564, 1534, 1536, 1552, 1553, 1666, 1547, 1548, 1556, 1559, 1560, 1654, 1555, 1540, 1543, 1668, 1539, 1550, 1558, 1565, 1541, 1667, 1542, 1544, 1537, 1551, 1545, 1-200, 1653, 392-960, and 1656-1659.
- a nucleic acid construct comprising the isolated polynucleotide and a promoter for directing transcription of the nucleic acid sequence.
- an isolated polypeptide comprising an amino acid sequence at least 90 % homologous to the amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, and 1660-1663.
- an isolated polypeptide comprising an amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961- 1529, and 1660-1663.
- a plant cell comprising an exogenous polypeptide comprising an amino acid sequence at least 90 % homologous to the amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, and 1660-1663.
- a plant cell comprising an exogenous polypeptide comprising an amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, and 1660-1663.
- a plant cell comprising an exogenous polynucleotide comprising a nucleic acid sequence at least 90 % homologous to the nucleic acid sequence selected from the group consisting of SEQ ID NOs:1530, 1561, 1532, 1531, 1562, 1533, 1538, 1549, 1665, 1566, 1554, 1563, 1557, 1564, 1534, 1536, 1552, 1553, 1666, 1547, 1548, 1556, 1559, 1560, 1654, 1555, 1540, 1543, 1668, 1539, 1550, 1558, 1565, 1541, 1667, 1542, 1544, 1537, 1551, 1545, 1-200, 1653, 392-960, and 1656-1659.
- a plant cell comprising an exogenous polynucleotide comprising a nucleic acid sequence selected from the group consisting of SEQ ID NOs: 1530, 1561, 1532, 1531, 1562, 1533, 1538, 1549, 1665, 1566, 1554, 1563, 1557, 1564, 1534, 1536, 1552, 1553, 1666, 1547, 1548, 1556, 1559, 1560, 1654, 1555, 1540, 1543, 1668, 1539, 1550, 1558, 1565, 1541, 1667, 1542, 1544, 1537, 1551, 1545, 1-200, 1653, 392-960, and 1656-1659.
- the nucleic acid sequence is selected from the group consisting of SEQ ID Nos:1530, 1561, 1532, 1531, 1562, 1533, 1538, 1549, 1665, 1566, 1554, 1563, 1557, 1564, 1534, 1536, 1552, 1553, 1666, 1547,
- the polynucleotide is selected from the group consisting of SEQ ID Nos:1530, 1561, 1532, 1531, 1562, 1533, 1538,
- the amino acid sequence is selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, and 1660-1663.
- the polypeptide is selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, and 1660-1663.
- the plant cell forms a part of a plant.
- the abiotic stress is selected from the group consisting of salinity, drought, water deprivation, low temperature, high temperature, heavy metal toxicity, anaerobiosis, nutrient deficiency, nutrient excess, atmospheric pollution and UV irradiation.
- the method further comprising growing the plant expressing the exogenous polynucleotide under the abiotic stress.
- FIG. 1 is a schematic illustration of the pGI binary plasmid used for expressing the isolated polynucleotide sequences of the invention.
- FIGs. 2a-b are images depicting visualization of root development of plants grown in transparent agar plates. The different transgenes were grown in transparent agar plates for 17 days and the plates were photographed every 2 days starting at day 7.
- Figure 2a An image of a photograph of plants taken following 12 days on agar plates.
- Figure 2b An image of root analysis in which the length of the root measured is represented by a red arrow.
- the present invention in some embodiments thereof, relates to isolated polypeptides and polynucleotides encoding same, and more particularly, but not exclusively, to methods of using same for increasing tolerance to abiotic stress, growth rate, yield, biomass and/or vigor of a plant.
- the present inventors While reducing the invention to practice, the present inventors have identified novel polypeptides and polynucleotides which can be used to increase tolerance to abiotic stress, and improve growth rate, biomass, yield and/or vigor of a plant.
- the present inventors have employed a bioinformatics approach which combines clustering and assembly of sequences from databases of the Arabidopsis, rice and other publicly available plant genomes, expressed sequence tags (ESTs), protein and pathway databases and QTL information with a digital expression profile ("electronic Northern Blot") and identified polynucleotides and polypeptides which can increase tolerance to abiotic stress, and improve growth, biomass, yield and vigor (SEQ ID NOs:l-200 and 1653 for polynucleotides; SEQ ID NOs:201-391 and 1655 for polypeptides; Table 1, Example 1).
- polynucleotide SEQ ID NOs: 1531, 1539, 1533, 1550, 1558, 1562, 1565, 1541, 1667, 1542, 1544, 1537, 1551 and 1545 were prepared.
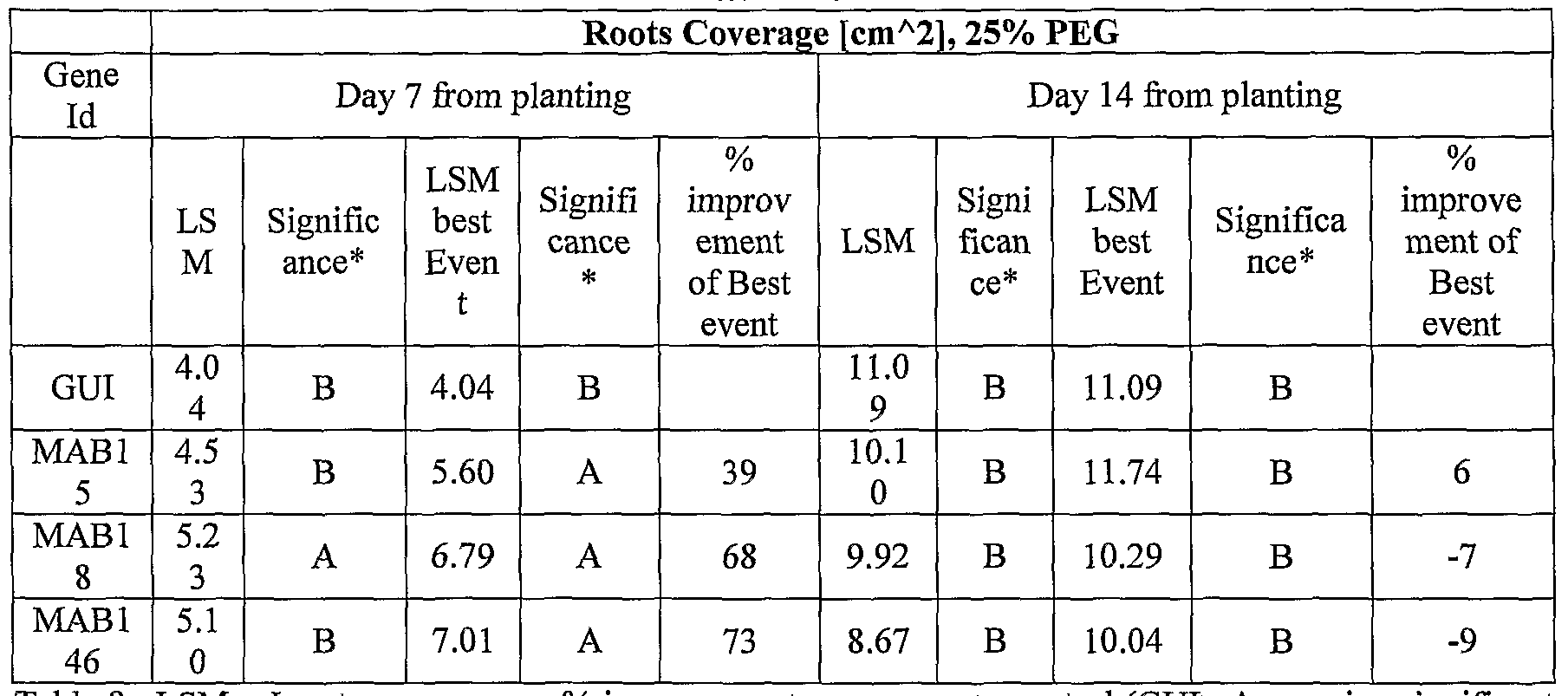
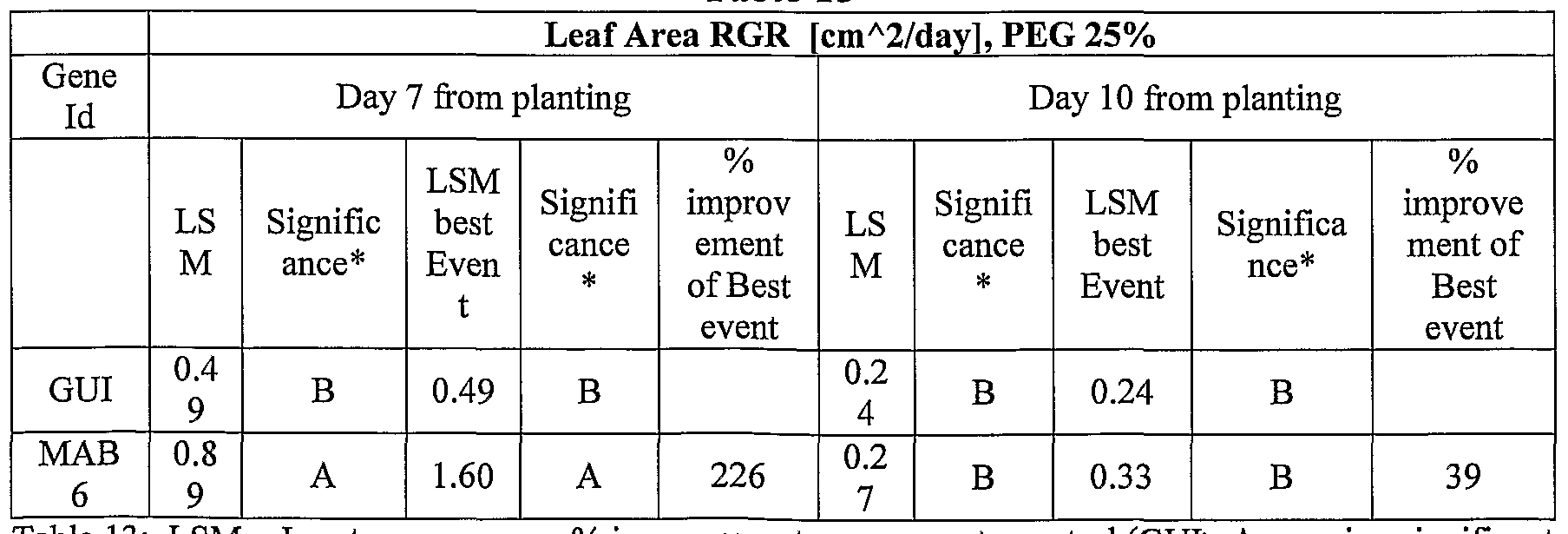
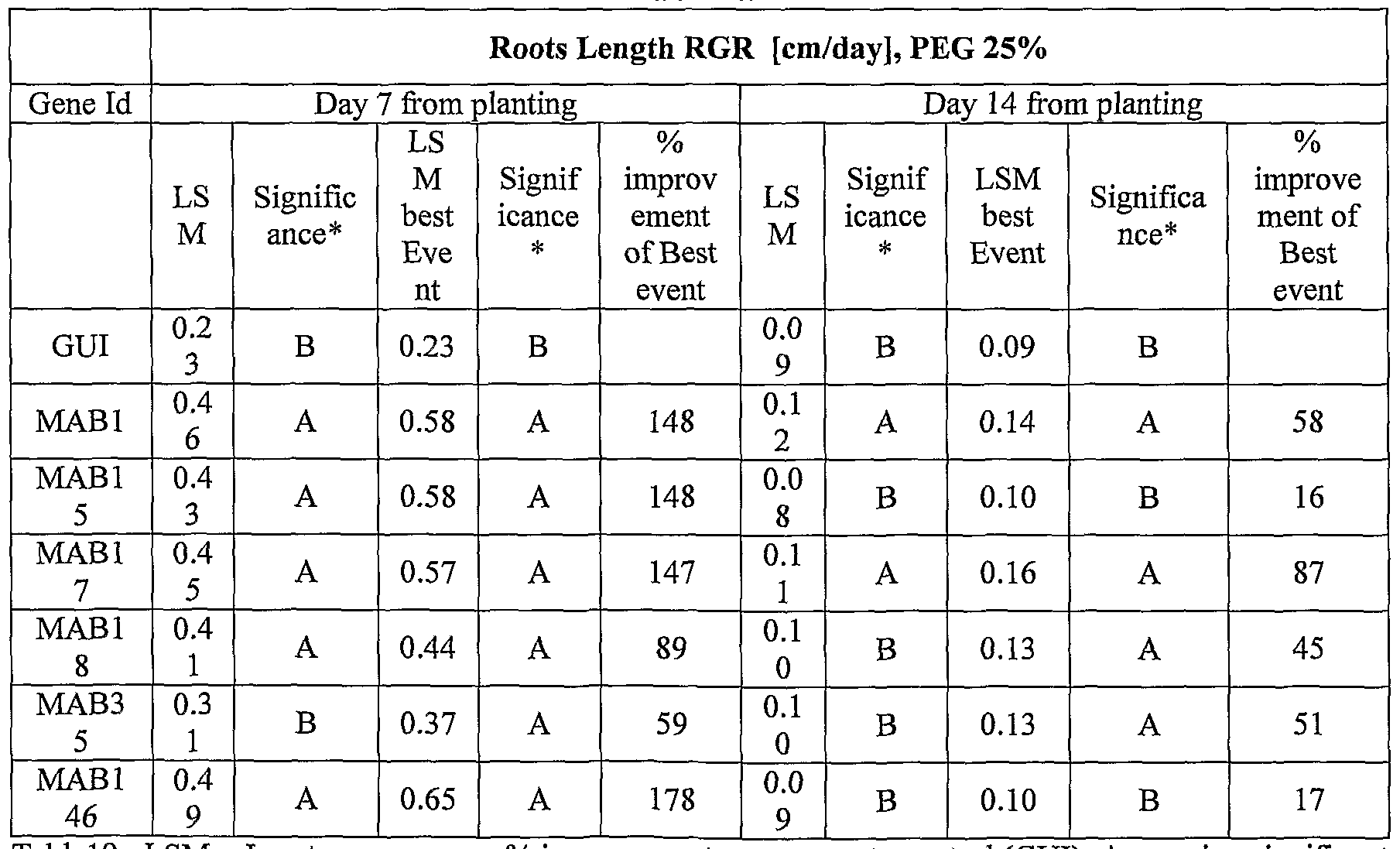
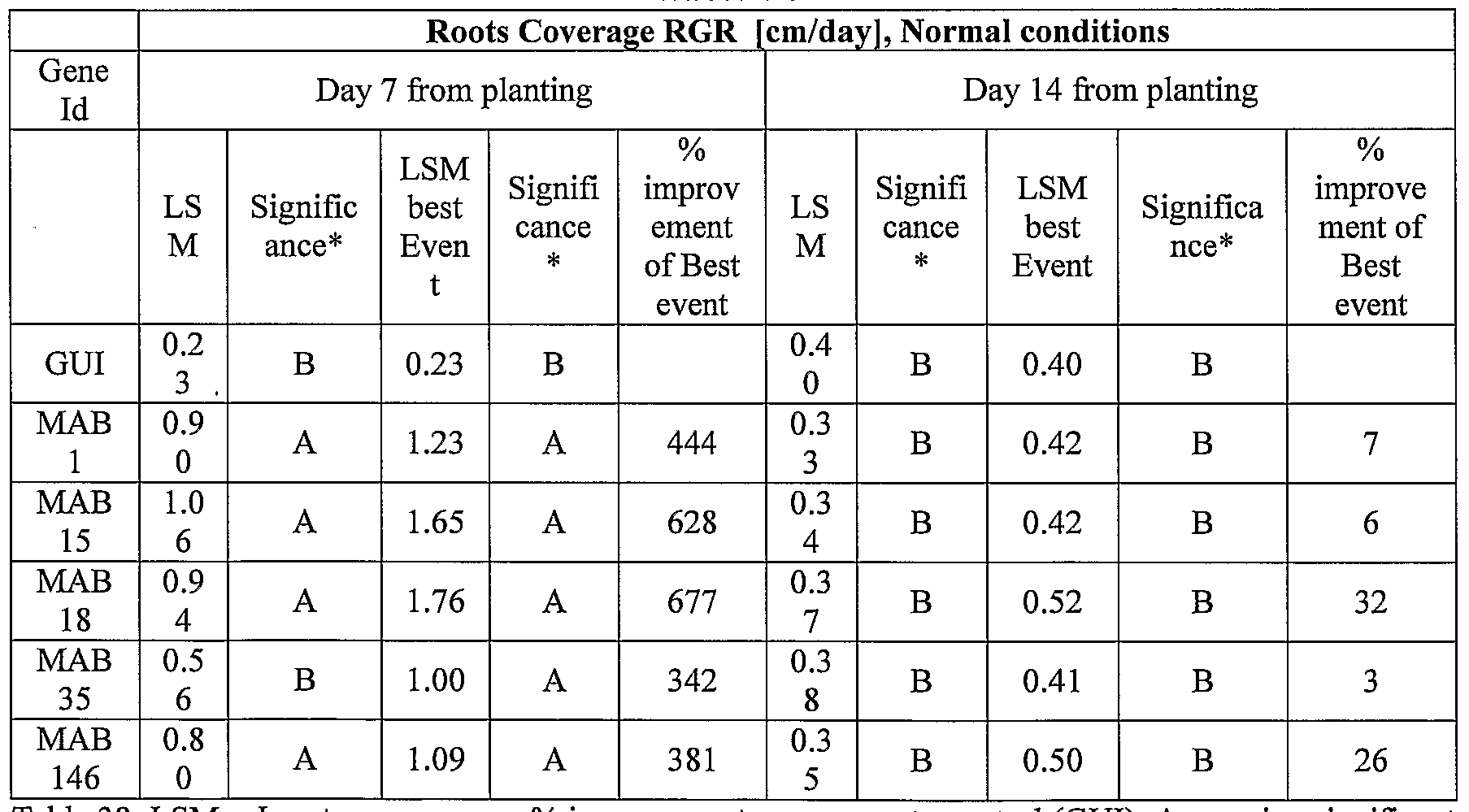
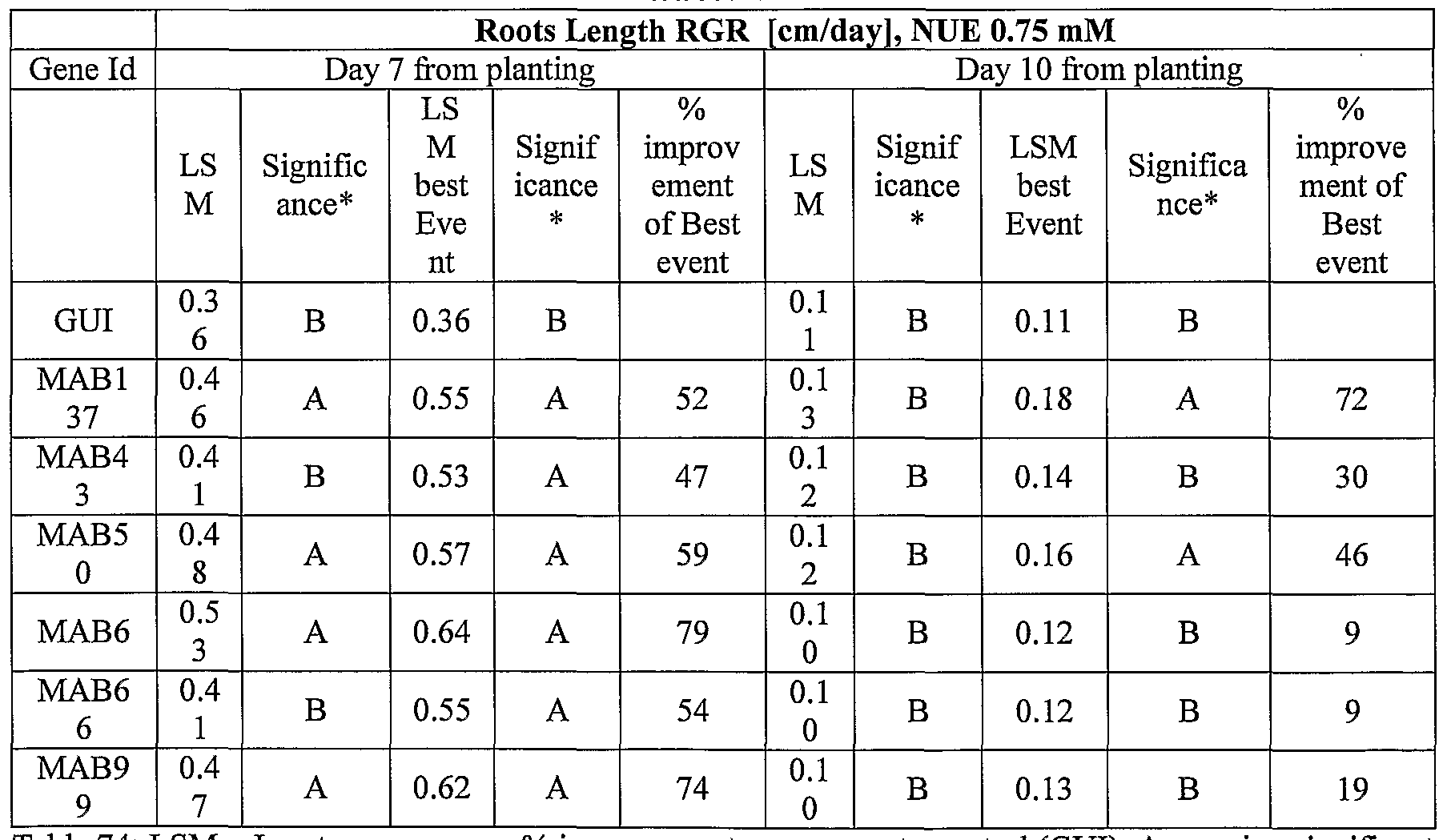
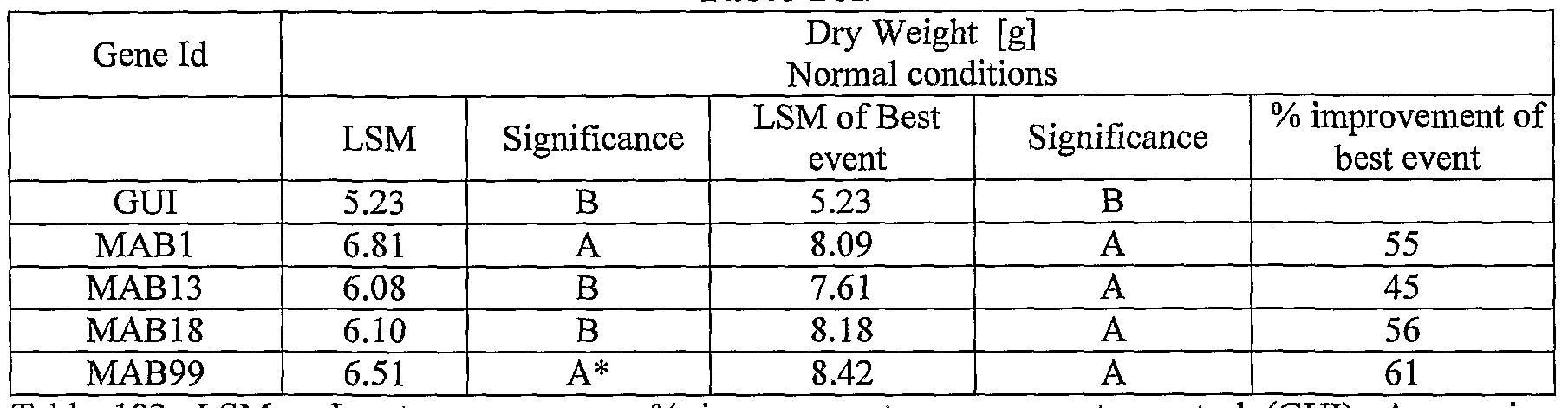
- transgenic plants exogenously expressing the cloned and/or optimized polynucleotides of the invention were generated. As shown in Tables 5-76, these plants exhibit increased seedling weight, root coverage, root length, and relative growth rate when grown under osmotic stress (in the presence of 25 % PEG), nitrogen deficiency (in the presence of 0.75 mM Nitrogen) or regular conditions.
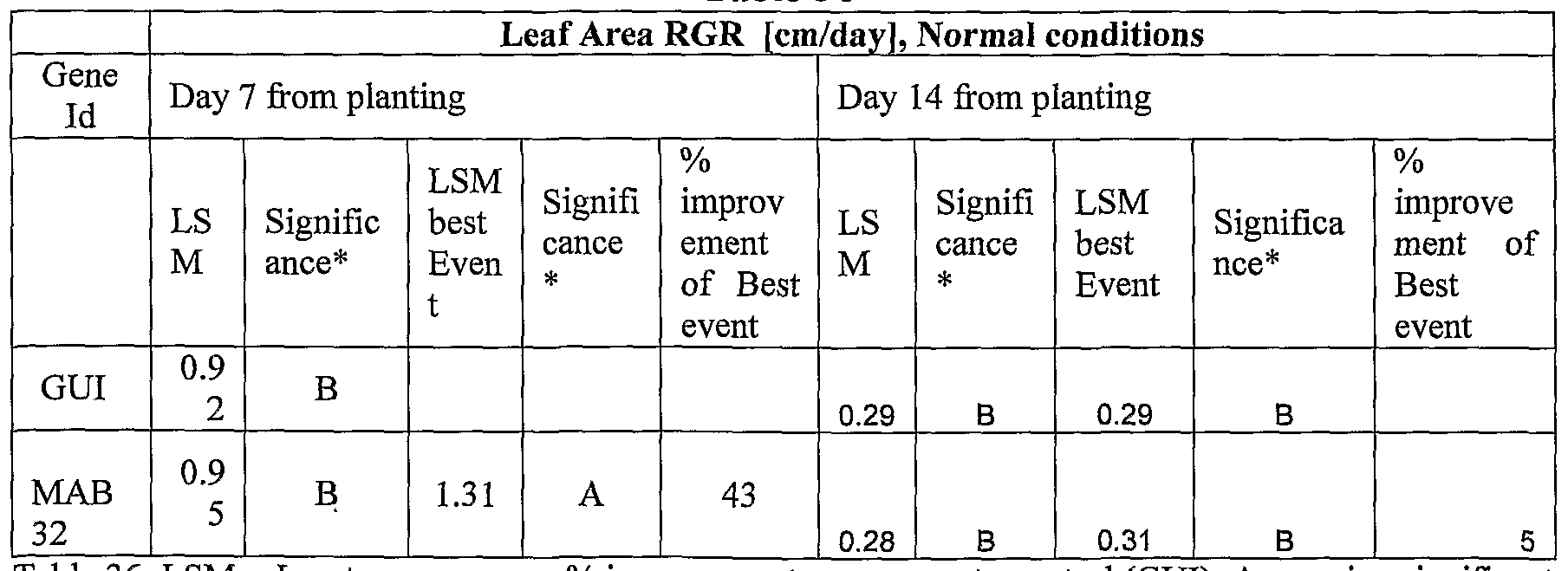
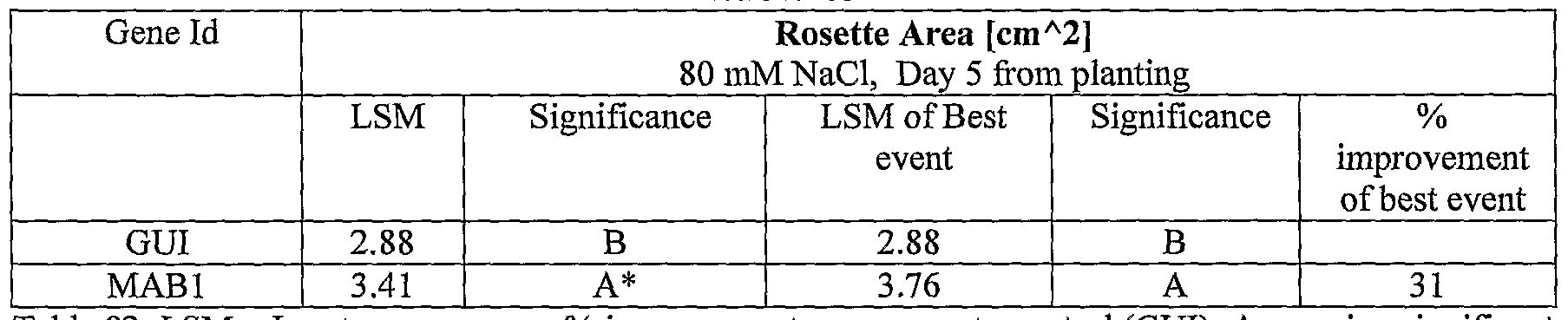
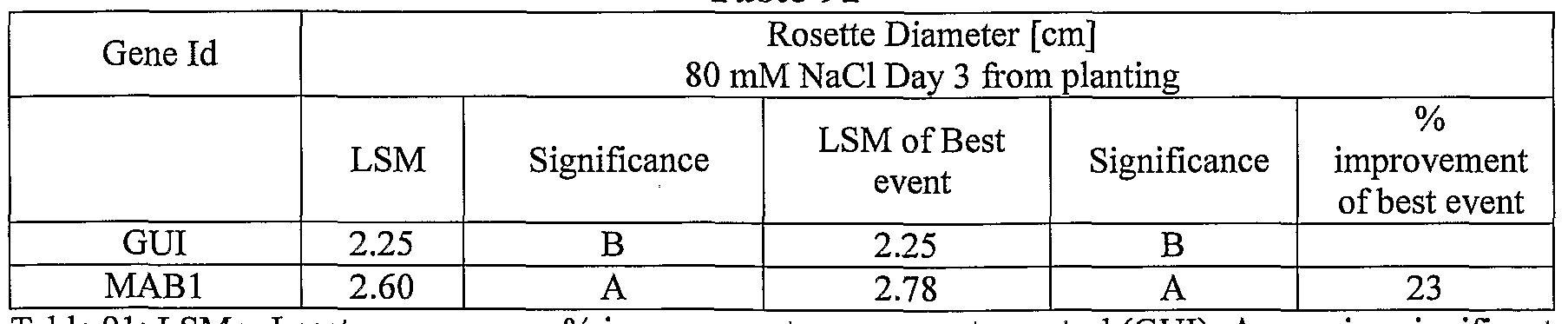
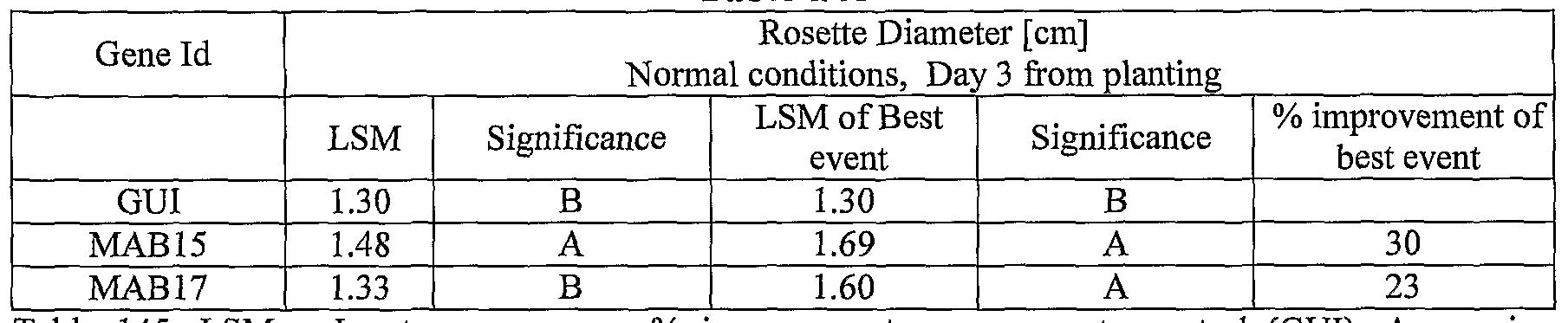
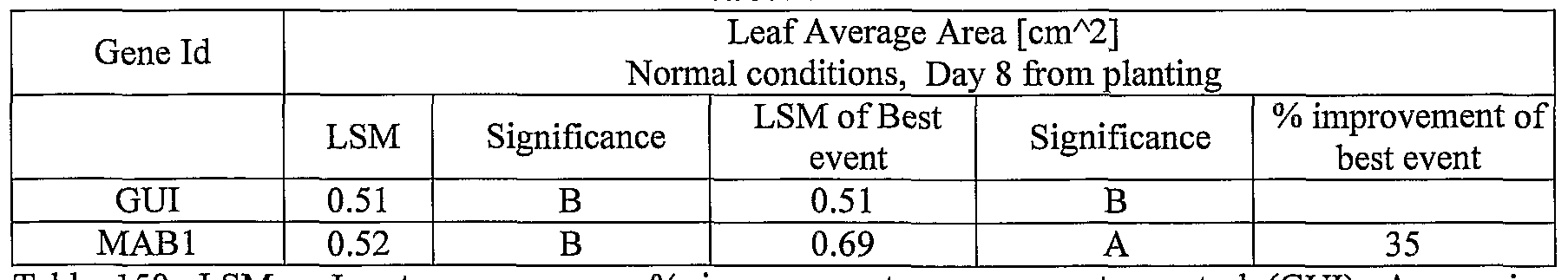
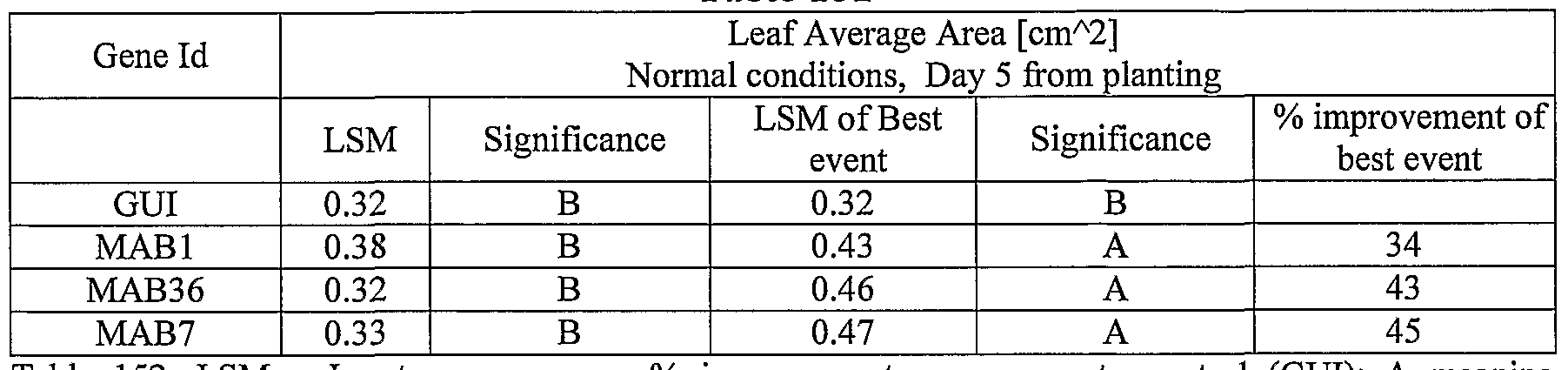
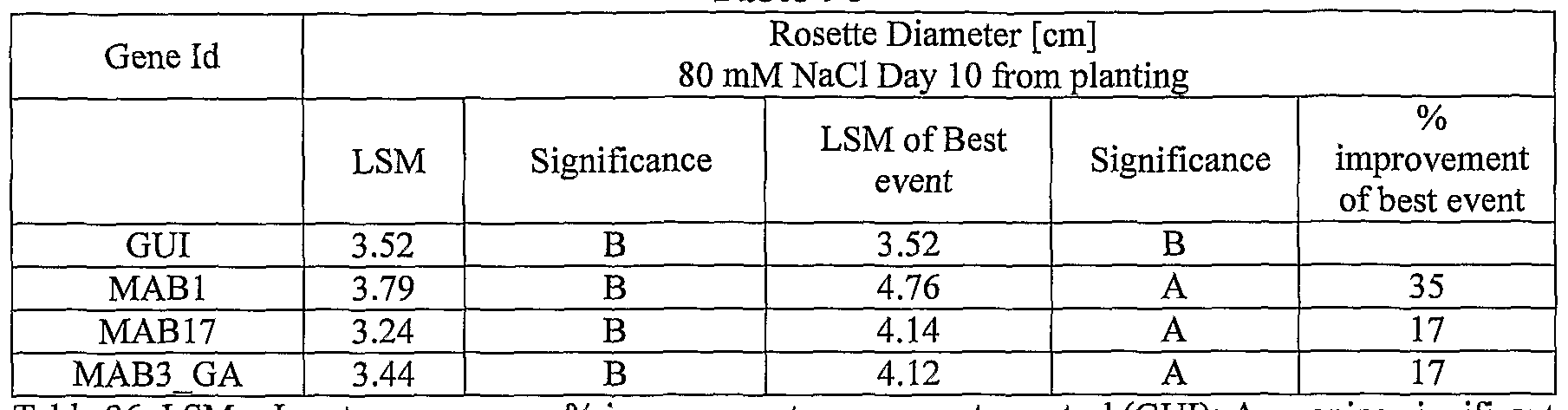
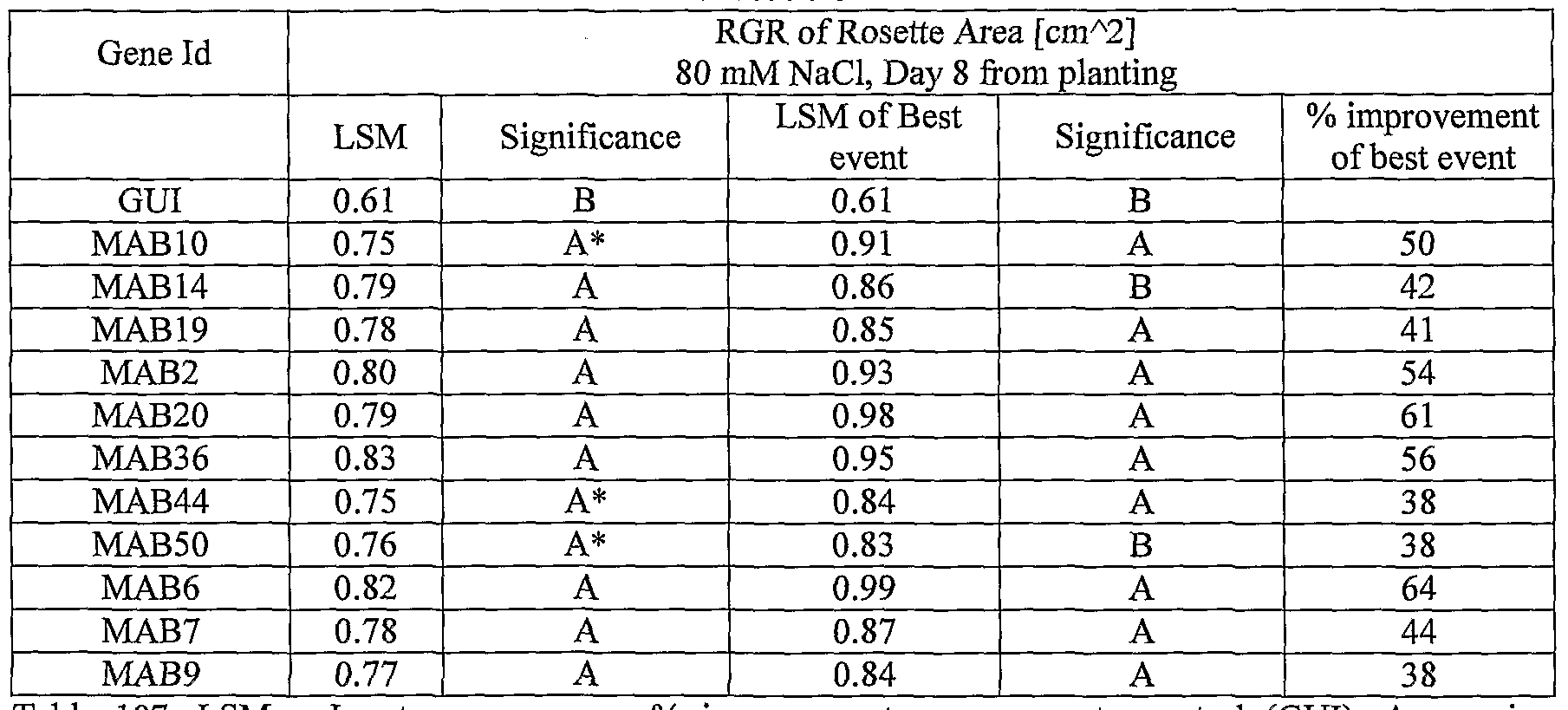
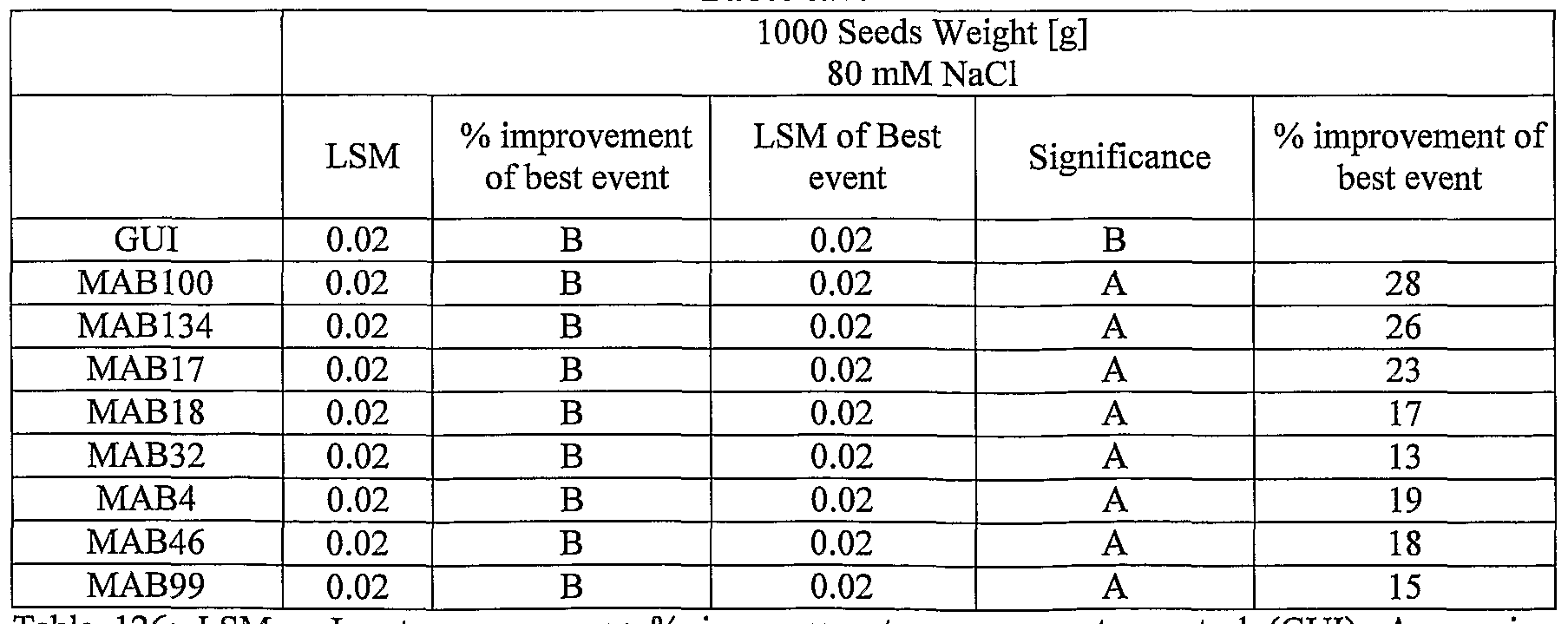
- plants exogenously expressing the polynucleotides of the invention exhibit increased rosette area, rosette diameter, leaf average area, relative growth rate of the above, plants biomass, plant seed yield, 1000 seed weight, and harvest index when grown under salinity stress or normal conditions.
- a method of increasing abiotic stress tolerance, growth rate, biomass, yield and/or vigor of a plant is effected by expressing within the plant an exogenous polynucleotide encoding a polypeptide comprising an amino acid sequence at least 60 % homologous to the amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, and 1660-1663.
- abiotic stress refers to any adverse effect on metabolism, growth, reproduction and/or viability of a plant. Accordingly, abiotic stress can be induced by suboptimal environmental growth conditions such as, for example, salinity, water deprivation, water deficit, drought, flooding, freezing, low or high temperature (e.g., chilling or excessive heat), toxic chemical pollution, heavy metal toxicity, anaerobiosis, nutrient deficiency, nutrient excess, atmospheric pollution or UV irradiation.
- suboptimal environmental growth conditions such as, for example, salinity, water deprivation, water deficit, drought, flooding, freezing, low or high temperature (e.g., chilling or excessive heat), toxic chemical pollution, heavy metal toxicity, anaerobiosis, nutrient deficiency, nutrient excess, atmospheric pollution or UV irradiation.
- abiotic stress tolerance refers to the ability of a plant to endure an abiotic stress without suffering a substantial alteration in metabolism, growth, productivity and/or viability.
- plant biomass refers to the amount (measured in grams of air-dry or dry tissue) of a tissue produced from the plant in a growing season, which could also determine or affect the plant yield or the yield per growing area.
- plant yield refers to the amount (as determined by weight, volume or size) or quantity (numbers) of tissue produced or harvested per plant or per growing season. Hence increased yield could affect the economic benefit one can obtain from the plant in a certain growing area and/or growing time.
- plant vigor refers to the amount (measured by weight) of tissue produced by the plant in a given time. Hence increase vigor could determine or affect the plant yield or the yield per growing time or growing area.
- increasing refers to at least about 2 %, at least about 3 %, at least about 4 %, at least about 5 %, at least about 10 %, at least about 15 %, at least about 20 %, at least about 30 %, at least about 40 %, at least about 50 %, at least about
- a native plant i.e., a plant not modified with the biomolecules (polynucleotide or polypeptides) of the invention, e.g., a non-transformed plant of the same species which is grown under the same growth conditions.
- exogenous polynucleotide refers to a heterologous nucleic acid sequence which may not be naturally expressed within the plant or which overexpression in the plant is desired.
- the exogenous polynucleotide may be introduced into the plant in a stable or transient manner, so as to produce a ribonucleic acid (RNA) molecule and/or a polypeptide molecule.
- RNA ribonucleic acid
- exogenous polynucleotide may comprise a nucleic acid sequence which is identical or partially homologous to an endogenous nucleic acid sequence of the plant.
- the exogenous polynucleotide of the invention encodes a polypeptide having an amino acid sequence at least about 60 %, at least about 65 %, at least about 70 %, at least about 75 %, at least about 80 %, at least about 81 %, at least about 82 %, at least about 83 %, at least about 84 %, at least about 85 %, at least about 86 %, at least about 87 %, at least about 88 %, at least about 89 %, at least about 90 %, at least about 91 %, at least about 92 %, at least about 93 %, at least about 94 %, at least about 95 %, at least about 96 %, at least about 97 %, at least about 98 %, at least about 99 %, or more say 100 % homologous to the amino acid sequence selected from the group consisting of SEQ ID NOs:201, 207, 212, 202-206, 208-211
- Homology can be determined using any homology comparison software, including for example, the BlastP or TBLASTN software of the National Center of Biotechnology Information (NCBI) such as by using default parameters, when starting from a polypeptide sequence; or the tBLASTX algorithm (available via the NCBI) such as by using default parameters, which compares the six- frame conceptual translation products of a nucleotide query sequence (both strands) against a protein sequence database.
- NCBI National Center of Biotechnology Information
- Homologous sequences include both orthologous and paralogous sequences.
- paralogous relates to gene-duplications within the genome of a species leading to paralogous genes.
- orthologous relates to homologous genes in different organisms due to ancestral relationship.
- One option to identify orthologues in monocot plant species is by performing a reciprocal blast search. This may be done by a first blast involving blasting the sequence-of-interest against any sequence database, such as the publicly available NCBI database which may be found at: Hypertext Transfer Protocol://World Wide Web (dot) ncbi (dot) nlm (dot) nih (dot) gov. If orthologues in rice were sought, the sequence-of- interest would be blasted against, for example, the 28,469 full-length cDNA clones from Oryza sativa Nipponbare available at NCBI. The blast results may be filtered.
- the ClustalW program may be used [Hypertext Transfer Protocol://World Wide Web (dot) ebi (dot) ac (dot) uk/Tools/clustalw2/index (dot) html], followed by a neighbor-joining tree (Hypertext Transfer Protocol://en (dot) wikipedia (dot) org/wiki/Neighbor-joining) which helps visualizing the clustering.
- the exogenous polynucleotide encodes a polypeptide consisting of the amino acid sequence set forth by SEQ ID NO:201, 207, 212, 202-206, 208-211, 213-391, 1655, 961-1529, 1660-1662 or 1663.
- the exogenous polynucleotide comprises a nucleic acid sequence which is at least about 60 %, at least about 65 %, at least about 70 %, at least about 75 %, at least about 80 %, at least about 81 %, at least about 82 %, at least about 83 %, at least about 84 %, at least about 85 %, at least about 86 %, at least about 87 %, at least about 88 %, at least about 89 %, at least about 90 %, at least about 91 %, at least about 92 %, at least about 93 %, at least about 93 %, at least about 94 %, at least about 95 %, at least about 96 %, at least about 97 %, at least about 98 %, at least about 99 %, e.g., 100 % identical to the nucleic acid sequence selected from the group consisting of SEQ ID NOs:1530, 1561
- the exogenous polynucleotide is at least about 60 %, at least about 65 %, at least about 70 %, at least about 75 %, at least about 80 %, at least about 81 %, at least about 82 %, at least about 83 %, at least about 84 %, at least about 85 %, at least about 86 %, at least about 87 %, at least about 88 %, at least about 89 %, at least about 90 %, at least about 91 %, at least about 92 %, at least about 93 %, at least about 93 %, at least about 94 %, at least about 95 %, at least about 96 %, at least about 97 %, at least about 98 %, at least about 99 %, e.g., 100 % identical to the polynucleotide selected from the group consisting of SEQ ID NOs: 1530, 1561, 1532, 1531,
- the exogenous polynucleotide is set forth by SEQ ID NO: 1530, 1561, 1532, 1531, 1562, 1533, 1538, 1549, 1665, 1566, 1554, 1563, 1557, 1564, 1534, 1536, 1552, 1553, 1666, 1547, 1548, 1556, 1559, 1560, 1654, 1555, 1540, 1543, 1668, 1539, 1550, 1558, 1565, 1541, 1667, 1542, 1544, 1537, 1551, 1545, 1-200, 1653, 392-960, and 1656-1658 or 1659.
- polynucleotide refers to a single or double stranded nucleic acid sequence which is isolated and provided in the form of an RNA sequence, a complementary polynucleotide sequence (cDNA), a genomic polynucleotide sequence and/or a composite polynucleotide sequences (e.g., a combination of the above).
- cDNA complementary polynucleotide sequence
- genomic polynucleotide sequence e.g., a combination of the above.
- composite polynucleotide sequences e.g., a combination of the above.
- complementary polynucleotide sequence refers to a sequence, which results from reverse transcription of messenger RNA using a reverse transcriptase or any other RNA dependent DNA polymerase. Such a sequence can be subsequently amplified in vivo or in vitro using a DNA dependent DNA polymerase.
- genomic polynucleotide sequence refers to a sequence derived (identified or isolated) from a chromosome and thus it represents a contiguous portion of a chromosome.
- composite polynucleotide sequence refers to a sequence, which is at least partially complementary and at least partially genomic.
- a composite sequence can include some exonal sequences required to encode the polypeptide of the present invention, as well as some intronic sequences interposing therebetween.
- the intronic sequences can be of any source, including of other genes, and typically will include conserved splicing signal sequences. Such intronic sequences may further include cis acting expression regulatory elements.
- Nucleic acid sequences encoding the polypeptides of the present invention may be optimized for expression.
- a non-limiting example of an optimized nucleic acid sequence is provided in SEQ ID NO: 1531, which encodes the polypeptide comprising the amino acid sequence set forth by SEQ ID NO:201.
- sequence modifications include, but are not limited to, an altered G/C content to more closely approach that typically found in the plant species of interest, and the removal of codons atypically found in the plant species commonly referred to as codon optimization.
- an optimized gene or nucleic acid sequence refers to a gene in which the nucleotide sequence of a native or naturally occurring gene has been modified in order to utilize statistically-preferred or statistically-favored codons within the plant.
- the nucleotide sequence typically is examined at the DNA level and the coding region optimized for expression in the plant species determined using any suitable procedure, for example as described in Sardana et al. (1996, Plant Cell Reports 15:677-681).
- the standard deviation of codon usage may be calculated by first finding the squared proportional deviation of usage of each codon of the native gene relative to that of highly expressed plant genes, followed by a calculation of the average squared deviation.
- a Table of codon usage from highly expressed genes of dicotyledonous plants is compiled using the data of Murray et al.
- the Codon Usage Database contains codon usage Tables for a number of different species, with each codon usage Table having been statistically determined based on the data present in Genbank.
- codon usage Tables for a particular species (for example, rice)
- a naturally- occurring nucleotide sequence encoding a protein of interest can be codon optimized for that particular plant species. This is effected by replacing codons that may have a low statistical incidence in the particular species genome with corresponding codons, in regard to an amino acid, that are statistically more favored.
- one or more less- favored codons may be selected to delete existing restriction sites, to create new ones at potentially useful junctions (5' and 3' ends to add signal peptide or termination cassettes, internal sites that might be used to cut and splice segments together to produce a correct full-length sequence), or to eliminate nucleotide sequences that may negatively effect mRNA stability or expression.
- codon optimization of the native nucleotide sequence may comprise determining which codons, within the native nucleotide sequence, are not statistically-favored with regards to a particular plant, and modifying these codons in accordance with a codon usage table of the particular plant to produce a codon optimized derivative.
- a modified nucleotide sequence may be fully or partially optimized for plant codon usage provided that the protein encoded by the modified nucleotide sequence is produced at a level higher than the protein encoded by the corresponding naturally occurring or native gene. Construction of synthetic genes by altering the codon usage is described in for example PCT Patent Application 93/07278.
- the invention encompasses nucleic acid sequences described hereinabove; fragments thereof, sequences hybridizable therewith, sequences homologous thereto, sequences encoding similar polypeptides with different codon usage, altered sequences characterized by mutations, such as deletion, insertion or substitution of one or more nucleotides, either naturally occurring or man induced, either randomly or in a targeted fashion.
- the invention provides an isolated polypeptide having an amino acid sequence at least about 60 %, at least about 65 %, at least about 70 %, at least about 75 %, at least about 80 %, at least about 81 %, at least about 82 %, at least about 83 %, at least about 84 %, at least about 85 %, at least about 86 %, at least about 87 %, at least about 88 %, at least about 89 %, at least about 90 %, at least about 91 %, at least about 92 %, at least about 93 %, at least about 93 %, at least about 94 %, at least about 95 %, at least about 96 %, at least about 97 %, at least about 98 %, at least about 99 %, or more say 100 % homologous to an amino acid sequence selected from the group consisting of SEQ ID NO:201, 207, 212, 202-206, 208-211, 213-391, 1655
- the invention also encompasses fragments of the above described polypeptides and polypeptides having mutations, such as deletions, insertions or substitutions of one or more amino acids, either naturally occurring or man induced, either randomly or in a targeted fashion.
- plant encompasses whole plants, ancestors and progeny of the plants and plant parts, including seeds, shoots, stems, roots (including tubers), and plant cells, tissues and organs.
- the plant may be in any form including suspension cultures, embryos, meristematic regions, callus tissue, leaves, gametophytes, sporophytes, pollen, and microspores.
- Plants that are particularly useful in the methods of the invention include all plants which belong to the superfamily Viridiplantae, in particular monocotyledonous and dicotyledonous plants including a fodder or forage legume, ornamental plant, food crop, tree, or shrub selected from the list comprising Acacia spp., Acer spp., Actinidia spp., Aesculus spp., Agathis australis, Albizia amara, Alsophila tricolor, Andropogon spp., Arachis spp, Areca catechu, Astelia fragrans, Astragalus cicer, Baikiaea plurijuga, Betula spp., Brassica.spp., Bruguiera gyninorrhiza, Burkea africana, Butea frondosa, Cadaba farinosa, Calliandra spp, Camellia sinensis, Carina indica, Capsicum spp., Cassia spp.,
- Echinochloa pyramidalis Ehraffia spp., Eleusine coracana, Eragrestis spp., Erythrina spp., Eucalypfus spp., Euclea schimperi, Eulalia vi/losa, Pagopyrum spp., Feijoa sellowlana, Fragaria spp., Flemingia spp, Freycinetia banksli, Geranium thunbergii, GinAgo biloba, Glycine javanica, Gliricidia spp, Gossypium hirsutum, Grevillea spp., Guibourtia coleosperma, Hedysarum spp., Hemaffhia altissima, Heteropogon contoffus, Hordeum vulgare, Hyparrhenia rufa, Hypericum erectum, Hypeffhelia dissolute, Indigo
- the plant used by the method of the invention is a crop plant such as rice, maize, wheat, barley, peanut, potato, sesame, olive tree, palm oil, banana, soybean, sunflower, canola, sugarcane, alfalfa, millet, leguminosae (bean, pea), flax, lupinus, rapeseed, tobacco, popular and cotton.
- a crop plant such as rice, maize, wheat, barley, peanut, potato, sesame, olive tree, palm oil, banana, soybean, sunflower, canola, sugarcane, alfalfa, millet, leguminosae (bean, pea), flax, lupinus, rapeseed, tobacco, popular and cotton.
- Expressing the exogenous polynucleotide of the invention within the plant can be effected by transforming one or more cells of the plant with the exogenous polynucleotide, followed by generating a mature plant from the transformed cells and cultivating the mature plant under conditions suitable for expressing the exogenous polynucleotide within the mature plant.
- the transformation is effected by introducing to the plant cell a nucleic acid construct which includes the exogenous polynucleotide of some embodiments of the invention and at least one promoter capable of directing transcription of the exogenous polynucleotide in the plant cell. Further details of suitable transformation approaches are provided hereinbelow.
- promoter refers to a region of DNA which lies upstream of the transcriptional initiation site of a gene to which RNA polymerase binds to initiate transcription of RNA.
- the promoter controls where (e.g., which portion of a plant) and/or when (e.g., at which stage or condition in the lifetime of an organism) the gene is expressed.
- the promoter is a constitutive promoter, a tissue-specific, or an abiotic stress-inducible promoter.
- Suitable constitutive promoters include, for example, CaMV 35S promoter (SEQ ID NO:1546; Odell et al., Nature 313:810-812, 1985); Arabidopsis At6669 promoter (SEQ ID NO: 1652; see PCT Publication No. WO04081173A2); maize Ubi 1 (Christensen et al., Plant Sol. Biol. 18:675-689, 1992); rice actin (McElroy et al., Plant Cell 2:163-171, 1990); pEMU (Last et al., Theor. Appl. Genet. 81:581-588, 1991); CaMV 19S (Nilsson et al., Physiol.
- tissue-specific promoters include, but not limited to, leaf-specific promoters [such as described, for example, by Yamamoto et al, Plant J. 12:255-265, 1997; Kwon et al., Plant Physiol.
- seed-preferred promoters [e.g., from seed specific genes (Simon, et al., Plant MoI. Biol. 5. 191, 1985; Scofield, et al., J. Biol. Chem. 262: 12202, 1987; Baszczynski, et al., Plant MoI.
- endosperm specific promoters e.g., wheat LMW and HMW 5 glutenin-1 (MoI Gen Genet 216:81-90, 1989; NAR 17:461-2), wheat a, b and g gliadins (EMBO3:1409-15, 1984), Barley ltrl promoter, barley Bl, C, D hordein (Theor Appl Gen 98:1253-62, 1999; Plant J 4:343-55, 1993; MoI Gen Genet 250:750- 60, 1996), Barley DOF (Mena et al., The Plant Journal, 116(1): 53- 62, 1998), Biz2 (EP99106056.7), Synthetic promoter (Vicente-Carbajosa et al., Plant J.
- KNOX Postma-Haarsma ef al, Plant MoI. Biol. 39:257-71, 1999
- rice oleosin Wild et at, J. Biochem., 123:386, 1998)]
- flower-specific promoters e.g., AtPRP4, chalene synthase (chsA) (Van der Meer, et al., Plant MoI. Biol. 15, 95- 109, 1990), LAT52 (Twell et al., MoI. Gen Genet. 217:240-245; 1989), apetala- 3].
- Suitable abiotic stress-inducible promoters include, but not limited to, salt- inducible promoters such as RD29A (Yamaguchi-Shinozalei et al., MoI. Gen. Genet. 236:331-340, 1993); drought-inducible promoters such as maize rabl7 gene promoter (PIa et. al., Plant MoI. Biol. 21:259-266, 1993), maize rab28 gene promoter (Busk et. al., Plant J. 11:1285-1295, 1997) and maize Ivr2 gene promoter (Pelleschi et. al., Plant MoL Biol. 39:373-380, 1999); heat-inducible promoters such as heat tomato hsp80-promoter from tomato (U.S. Pat. No. 5,187,267).
- salt- inducible promoters such as RD29A (Yamaguchi-Shinozalei et al., MoI.
- the nucleic acid construct of some embodiments of the invention can further include an appropriate selectable marker and/or an origin of replication.
- the nucleic acid construct utilized is a shuttle vector, which can propagate both in E. coli (wherein the construct comprises an appropriate selectable marker and origin of replication) and be compatible with propagation in cells.
- the construct according to the present invention can be, for example, a plasmid, a bacmid, a phagemid, a cosmid, a phage, a virus or an artificial chromosome.
- the nucleic acid construct of some embodiments of the invention can be utilized to stably or transiently transform plant cells, m stable transformation, the exogenous polynucleotide is integrated into the plant genome and as such it represents a stable and inherited trait.
- transient transformation the exogenous polynucleotide is expressed by the cell transformed but it is not integrated into the genome and as such it represents a transient trait.
- the Agrobacterium system includes the use of plasmid vectors that contain defined DNA segments that integrate into the plant genomic DNA. Methods of inoculation of the plant tissue vary depending upon the plant species and the Agrobacterium delivery system. A widely used approach is the leaf disc procedure which can be performed with any tissue explant that provides a good source for initiation of whole plant differentiation. See, e.g., Horsch et al. in Plant Molecular Biology Manual A5, Kluwer Academic Publishers, Dordrecht (1988) p. 1-9. A supplementary approach employs the Agrobacterium delivery system in combination with vacuum infiltration. The Agrobacterium system is especially viable in the creation of transgenic dicotyledonous plants.
- DNA transfer into plant cells There are various methods of direct DNA transfer into plant cells.
- electroporation the protoplasts are briefly exposed to a strong electric field.
- microinjection the DNA is mechanically injected directly into the cells using very small micropipettes.
- microparticle bombardment the DNA is adsorbed on microprojectiles such as magnesium sulfate crystals or tungsten particles, and the microprojectiles are physically accelerated into cells or plant tissues.
- Micropropagation is a process of growing new generation plants from a single piece of tissue that has been excised from a selected parent plant or cultivar. This process permits the mass reproduction of plants having the preferred tissue expressing the fusion protein.
- the new generation plants which are produced are genetically identical to, and have all of the characteristics of, the original plant.
- Micropropagation allows mass production of quality plant material in a short period of time and offers a rapid multiplication of selected cultivars in the preservation of the characteristics of the original transgenic or transformed plant.
- the advantages of cloning plants are the speed of plant multiplication and the quality and uniformity of plants produced.
- Micropropagation is a multi-stage procedure that requires alteration of culture medium or growth conditions between stages.
- the micropropagation process involves four basic stages: Stage one, initial tissue culturing; stage two, tissue culture multiplication; stage three, differentiation and plant formation; and stage four, greenhouse culturing and hardening.
- stage one initial tissue culturing
- stage two the initial tissue culture is established and certified contaminant-free.
- stage two the initial tissue culture is multiplied until a sufficient number of tissue samples are produced to meet production goals.
- stage three the tissue samples grown in stage two are divided and grown into individual plantlets.
- the transformed plantlets are transferred to a greenhouse for hardening where the plants' tolerance to light is gradually increased so that it can be grown in the natural environment.
- the transgenic plants are generated by transient transformation of leaf cells, meristematic cells or the whole plant. Transient transformation can be effected by any of the direct DNA transfer methods described above or by viral infection using modified plant viruses.
- Viruses that have been shown to be useful for the transformation of plant hosts include CaMV, Tobacco mosaic virus (TMV), brome mosaic virus (BMV) and Bean Common Mosaic Virus (BV or BCMV). Transformation of plants using plant viruses is described in U.S. Pat. No. 4,855,237 (bean golden mosaic virus; BGV), EP-A 67,553 (TMV), Japanese Published Application No. 63-14693 (TMV), EPA 194,809 (BV), EPA 278,667 (BV); and Gluzman, Y. et al., Communications in Molecular Biology:
- the virus used for transient transformations is avirulent and thus is incapable of causing severe symptoms such as reduced growth rate, mosaic, ring spots, leaf roll, yellowing, streaking, pox formation, tumor formation and pitting.
- a suitable avirulent virus may be a naturally occurring avirulent virus or an artificially attenuated virus.
- Virus attenuation may be effected by using methods well known in the art including, but not limited to, sub-lethal heating, chemical treatment or by directed mutagenesis techniques such as described, for example, by Kurihara and Watanabe (Molecular Plant Pathology 4:259-269, 2003), Galon et al. (1992), Atreya et al. (1992) and Huet et al. (1994).
- Suitable virus strains can be obtained from available sources such as, for example, the American Type culture Collection (ATCC) or by isolation from infected plants. Isolation of viruses from infected plant tissues can be effected by techniques well known in the art such as described, for example by Foster and Tatlor, Eds. "Plant Virology Protocols: From Virus Isolation to Transgenic Resistance (Methods in Molecular Biology (Humana Pr), VoI 81)", Humana Press, 1998. Briefly, tissues of an infected plant believed to contain a high concentration of a suitable virus, preferably young leaves and flower petals, are ground in a buffer solution (e.g., phosphate buffer solution) to produce a virus infected sap which can be used in subsequent inoculations.
- a buffer solution e.g., phosphate buffer solution
- the virus When the virus is a DNA virus, suitable modifications can be made to the virus itself. Alternatively, the virus can first be cloned into a bacterial plasmid for ease of constructing the desired viral vector with the foreign DNA. The virus can then be excised from the plasmid. If the virus is a DNA virus, a bacterial origin of replication can be attached to the viral DNA, which is then replicated by the bacteria. Transcription and translation of this DNA will produce the coat protein which will encapsidate the viral DNA. If the virus is an RNA virus, the virus is generally cloned as a cDNA and inserted into a plasmid. The plasmid is then used to make all of the constructions. The RNA virus is then produced by transcribing the viral sequence of the plasmid and translation of the viral genes to produce the coat protein(s) which encapsidate the viral RNA.
- a plant viral polynucleotide in which the native coat protein coding sequence has been deleted from a viral polynucleotide, a non-native plant viral coat protein coding sequence and a non-native promoter, preferably the subgenomic promoter of the non-native coat protein coding sequence, capable of expression in the plant host, packaging of the recombinant plant viral polynucleotide, and ensuring a systemic infection of the host by the recombinant plant viral polynucleotide, has been inserted.
- the coat protein gene may be inactivated by insertion of the non-native polynucleotide sequence within it, such that a protein is produced.
- the recombinant plant viral polynucleotide may contain one or more additional non-native subgenomic promoters.
- Each non-native subgenomic promoter is capable of transcribing or expressing adjacent genes or polynucleotide sequences in the plant host and incapable of recombination with each other and with native subgenomic promoters.
- Non-native (foreign) polynucleotide sequences may be inserted adjacent the native plant viral subgenomic promoter or the native and a non- native plant viral subgenomic promoters if more than one polynucleotide sequence is included.
- the non-native polynucleotide sequences are transcribed or expressed in the host plant under control of the subgenomic promoter to produce the desired products.
- a recombinant plant viral polynucleotide is provided as in the first embodiment except that the native coat protein coding sequence is placed adjacent one of the non-native coat protein subgenomic promoters instead of a non- native coat protein coding sequence.
- a recombinant plant viral polynucleotide in which the native coat protein gene is adjacent its subgenomic promoter and one or more non-native subgenomic promoters have been inserted into the viral polynucleotide.
- the inserted non-native subgenomic promoters are capable of transcribing or expressing adjacent genes in a plant host and are incapable of recombination with each other and with native subgenomic promoters.
- Non-native polynucleotide sequences may be inserted adjacent the non-native subgenomic plant viral promoters such that the sequences are transcribed or expressed in the host plant under control of the subgenomic promoters to produce the desired product.
- a recombinant plant viral polynucleotide is provided as in the third embodiment except that the native coat protein coding sequence is replaced by a non-native coat protein coding sequence.
- the viral vectors are encapsidated by the coat proteins encoded by the recombinant plant viral polynucleotide to produce a recombinant plant virus.
- the recombinant plant viral polynucleotide or recombinant plant virus is used to infect appropriate host plants.
- the recombinant plant viral polynucleotide is capable of replication in the host, systemic spread in the host, and transcription or expression of foreign gene(s) (exogenous polynucleotide) in the host to produce the desired protein.
- a technique for introducing exogenous polynucleotide sequences to the genome of the chloroplasts involves the following procedures. First, plant cells are chemically treated so as to reduce the number of chloroplasts per cell to about one. Then, the exogenous polynucleotide is introduced via particle bombardment into the cells with the aim of introducing at least one exogenous polynucleotide molecule into the chloroplasts. The exogenous polynucleotide is selected such that it is integratable into the chloroplast's genome via homologous recombination which is readily effected by enzymes inherent to the chloroplast.
- the exogenous polynucleotide includes, in addition to a gene of interest, at least one polynucleotide stretch which is derived from the chloroplast's genome.
- the exogenous polynucleotide includes a selectable marker, which serves by sequential selection procedures to ascertain that all or substantially all of the copies of the chloroplast genomes following such selection will include the exogenous polynucleotide. Further details relating to this technique are found in U.S. Pat. Nos. 4,945,050; and 5,693,507 which are incorporated herein by reference.
- a polypeptide can thus be produced by the protein expression system of the chloroplast and become integrated into the chloroplast's inner membrane.
- the present invention also envisages expressing a plurality of exogenous polynucleotides in a single host plant to thereby achieve superior effect on abiotic stress tolerance, growth, biomass, yield and/or vigor.
- Expressing a plurality of exogenous polynucleotides in a single host plant can be effected by co-introducing multiple nucleic acid constructs, each including a different exogenous polynucleotide, into a single plant cell.
- the transformed cell can then be regenerated into a mature plant using the methods described hereinabove.
- expressing a plurality of exogenous polynucleotides in a single host plant can be effected by co-introducing into a single plant-cell a single nucleic-acid construct including a plurality of different exogenous polynucleotides.
- Such a construct can be designed with a single promoter sequence which can transcribe a polycistronic messenger RNA including all the different exogenous polynucleotide sequences.
- the polynucleotide sequences can be inter-linked via an internal ribosome entry site (IRES) sequence which facilitates translation of polynucleotide sequences positioned downstream of the IRES sequence.
- IRES internal ribosome entry site
- a transcribed polycistronic RNA molecule encoding the different polypeptides described above will be translated from both the capped 5' end and the two internal IRES sequences of the polycistronic RNA molecule to thereby produce in the cell all different polypeptides.
- the construct can include several promoter sequences each linked to a different exogenous polynucleotide sequence.
- the plant cell transformed with the construct including a plurality of different exogenous polynucleotides can be regenerated into a mature plant, using the methods described hereinabove.
- expressing a plurality of exogenous polynucleotides can be effected by introducing different nucleic acid constructs, including different exogenous polynucleotides, into a plurality of plants.
- the regenerated transformed plants can then be cross-bred and resultant progeny selected for superior abiotic stress tolerance, growth, biomass, yield and/or vigor traits, using conventional plant breeding techniques.
- the plant expressing the exogenous polynucleotide(s) is grown under normal conditions.
- the method further comprising growing the plant expressing the exogenous polynucleotide(s) under the abiotic stress.
- the invention encompasses plants exogenously expressing (as described above) the polynucleotide(s) and/or polypeptide(s) of the invention.
- the level of the polypeptide encoded by the exogenous polynucleotide can be determined by methods well known in the art such as, activity assays, Western blots using antibodies capable of specifically binding the polypeptide, Enzyme-Linked Immunosorbent Assay (ELISA), radio-immuno-assays
- RNA transcribed from the exogenous polynucleotide are well known in the art and include, for example, Northern blot analysis, reverse transcription polymerase chain reaction (RT-PCR) analysis
- RNA-m situ hybridization including quantitative, semi-quantitative or real-time RT-PCR and RNA-m situ hybridization.
- polynucleotides and polypeptides described hereinabove can be used in a wide range of economical plants, in a safe and cost effective manner.
- the effect of the transgene (the exogenous polynucleotide encoding the polypeptide) on abiotic stress tolerance, growth, biomass, yield and/or vigor can be determined using known methods.
- Abiotic stress tolerance - Transformed (i.e., expressing the transgene) and non- transformed (wild type) plants are exposed to an abiotic stress condition, such as water deprivation, suboptimal temperature (low temperature, high temperature), nutrient deficiency, nutrient excess, a salt stress condition, osmotic stress, heavy metal toxicity, anaerobiosis, atmospheric pollution and UV irradiation.
- an abiotic stress condition such as water deprivation, suboptimal temperature (low temperature, high temperature), nutrient deficiency, nutrient excess, a salt stress condition, osmotic stress, heavy metal toxicity, anaerobiosis, atmospheric pollution and UV irradiation.
- Salinity tolerance assay - Transgenic plants with tolerance to high salt concentrations are expected to exhibit better germination, seedling vigor or growth in high salt.
- Salt stress can be effected in many ways such as, for example, by irrigating the plants with a hyperosmotic solution, by cultivating the plants hydroponically in a hyperosmotic growth solution (e.g., Hoagland solution with added salt), or by culturing the plants in a hyperosmotic growth medium [e.g., 50 % Murashige-Skoog medium (MS medium) with added salt].
- a hyperosmotic growth medium e.g., 50 % Murashige-Skoog medium (MS medium) with added salt.
- the salt concentration in the irrigation water, growth solution, or growth medium can be adjusted according to the specific characteristics of the specific plant cultivar or variety, so as to inflict a mild or moderate effect on the physiology and/or morphology of the plants (for guidelines as to appropriate concentration see, Bernstein and Kafkafi, Root Growth Under Salinity Stress In: Plant Roots, The Hidden Half 3rd ed. Waisel Y, Eshel A and Kafkafi U. (editors) Marcel Dekker Inc., New York, 2002, and reference therein).
- a salinity tolerance test can be performed by irrigating plants at different developmental stages with increasing concentrations of sodium chloride (for example 50 mM, 100 niM, 200 mM, 400 mM NaCl) applied from the bottom and from above to ensure even dispersal of salt. Following exposure to the stress condition the plants are frequently monitored until substantial physiological and/or morphological effects appear in wild type plants. Thus, the external phenotypic appearance, degree of wilting and overall success to reach maturity and yield progeny are compared between control and transgenic plants.
- sodium chloride for example 50 mM, 100 niM, 200 mM, 400 mM NaCl
- Quantitative parameters of tolerance measured include, but are not limited to, the average wet and dry weight, growth rate, leaf size, leaf coverage (overall leaf area), the weight of the seeds yielded, the average seed size and the number of seeds produced per plant.
- Transformed plants not exhibiting substantial physiological and/or morphological effects, or exhibiting higher biomass than wild-type plants, are identified as abiotic stress tolerant plants.
- Osmotic tolerance test - Osmotic stress assays including sodium chloride and
- PEG assays are conducted to determine if an osmotic stress phenotype was sodium chloride-specific or if it was a general osmotic stress related phenotype. Plants which are tolerant to osmotic stress may have more tolerance to drought and/or freezing. For salt and osmotic stress experiments, the medium is supplemented for example with 50 mM, 100 mM, 200 mM NaCl or 15 %, 20 % or 25 % PEG. See also Examples 6 and 7 of the Examples section which follows.
- Drought tolerance assay/Osmoticum assay - Tolerance to drought is performed to identify the genes conferring better plant survival after acute water deprivation.
- an osmotic stress produced by the non-ionic osmolyte sorbitol in the medium can be performed.
- Control and transgenic plants are germinated and grown in plant-agar plates for 4 days, after which they are transferred to plates containing 500 mM sorbitol. The treatment causes growth retardation, then both control and transgenic plants are compared, by measuring plant weight (wet and dry), yield, and by growth rates measured as time to flowering.
- soil-based drought screens are performed with plants overexpressing the polynucleotides detailed above. Seeds from control Arabidopsis plants, or other transgenic plants overexpressing the polypeptide of the invention are germinated and transferred to pots. Drought stress is obtained after irrigation is ceased accompanied by placing the pots on absorbent paper to enhance the soil-drying rate. Transgenic and control plants are compared to each other when the majority of the control plants develop severe wilting. Plants are re- watered after obtaining a significant fraction of the control plants displaying a severe wilting. Plants are ranked comparing to controls for each of two criteria: tolerance to the drought conditions and recovery (survival) following re-watering.
- Cold stress tolerance One way to analyze cold stress is as follows. Mature (25 day old) plants are transferred to 4 °C chambers for 1 or 2 weeks, with constitutive light. Later on plants are moved back to greenhouse. Two weeks later damages from chilling period, resulting in growth retardation and other phenotypes, are compared between control and transgenic plants, by measuring plant weight (wet and dry), and by comparing growth rates measured as time to flowering, plant size, yield, and the like.
- Heat stress tolerance One way to measure heat stress tolerance is by exposing the plants to temperatures above 34 °C for a certain period. Plant tolerance is examined after transferring the plants back to 22 0 C for recovery and evaluation after 5 days relative to internal controls (non-transgenic plants) or plants not exposed to neither cold or heat stress.
- Germination tests compare the percentage of seeds from transgenic plants that could complete the germination process to the percentage of seeds from control plants that are treated in the same manner. Normal conditions are considered for example, incubations at 22 °C under 22-hour light 2-hour dark daily cycles. Evaluation of germination and seedling vigor is conducted between 4 and 14 days after planting. The basal media is 50 % MS medium (Murashige and Skoog, 1962
- Germination is checked also at unfavorable conditions such as cold (incubating at temperatures lower than 10 0 C instead of 22 0 C) or using seed inhibition solutions that contain high concentrations of an osmolyte such as sorbitol (at concentrations of 50 mM,
- Plant vigor can be calculated by the increase in growth parameters such as leaf area, fiber length, rosette diameter, plant fresh weight and the like per time.
- the growth rate can be measured using digital analysis of growing plants. For example, images of plants growing in greenhouse on plot basis can be captured every 3 days and the rosette area can be calculated by digital analysis. Rosette area growth is calculated using the difference of rosette area between days of sampling divided by the difference in days between samples.
- Measurements of seed yield can be done by collecting the total seeds from 8-16 plants together, weighting them using analytical balance and dividing the total weight by the number of plants. Seed per growing area can be calculated in the same manner while taking into account the growing area given to a single plant. Increase seed yield per growing area could be achieved by increasing seed yield per plant, and/or by increasing number of plants capable of growing in a given area.
- Evaluation of the seed yield per plant can be done by measuring the amount
- Evaluation of growth rate can be done by measuring plant biomass produced, rosette area, leaf size or root length per time (can be measured in cm 2 per day of leaf area).
- Fiber length can be measured using fibrograph.
- the fibrograph system was used to compute length in terms of "Upper Half Mean” length.
- the upper half mean (UHM) is the average length of longer half of the fiber distribution.
- the fibrograph measures length in span lengths at a given percentage point (Hypertext Transfer Protocol://World
- the present invention is of high agricultural value for promoting the yield of commercially desired crops (e.g., biomass of vegetative organ such as poplar wood, or reproductive organ such as number of seeds or seed biomass).
- biomass of vegetative organ such as poplar wood, or reproductive organ such as number of seeds or seed biomass.
- reproductive organ such as number of seeds or seed biomass.
- compositions, methods or structure may include additional ingredients, steps and/or parts, but only if the additional ingredients, steps and/or parts do not materially alter the basic and novel characteristics of the claimed composition, method or structure.
- a compound or “at least one compound” may include a plurality of compounds, including mixtures thereof.
- various embodiments of this invention may be presented in a range format. It should be understood that the description in range format is merely for convenience and brevity and should not be construed as an inflexible limitation on the scope of the invention. Accordingly, the description of a range should be considered to have specifically disclosed all the possible subranges as well as individual numerical values within that range.
- a range such as from 1 to 6 should be considered to have specifically disclosed subranges such as from 1 to 3, from 1 to 4, from 1 to 5, from 2 to 4, from 2 to 6, from 3 to 6 etc., as well as individual numbers within that range, for example, 1, 2, 3, 4, 5, and 6. This applies regardless of the breadth of the range.
- a numerical range is indicated herein, it is meant to include any cited numeral (fractional or integral) within the indicated range.
- the phrases "ranging/ranges between" a first indicate number and a second indicate number and “ranging/ranges from” a first indicate number "to” a second indicate number are used herein interchangeably and are meant to include the first and second indicated numbers and all the fractional and integral numerals therebetween.
- method refers to manners, means, techniques and procedures for accomplishing a given task including, but not limited to, those manners, means, techniques and procedures either known to, or readily developed from known manners, means, techniques and procedures by practitioners of the chemical, pharmacological, biological, biochemical and medical arts.
- the present inventors have identified genes which increase abiotic stress- tolerance (ABST) and/or growth rate/yield/biomass/vigor, as follows.
- the genes were validated in vivo as previously described in WO2004/104162 to the present assignee. All nucleotide sequence datasets used here were originated from publicly available databases. Sequence data from 50 different species (mainly plant species) was introduced into a single, comprehensive database. Other information on gene expression, protein annotation, enzymes and pathways were also incorporated.
- Major databases used include: • Genomes o Arabidopsis genome [TAIR genome version 6 (Hypertext Transfer
- Microarray datasets were downloaded from o GEO (Hypertext Transfer Protocol. -//World Wide
- Database Assembly was performed to build a wide, rich, reliable annotated and easy to analyze database comprised of publicly available genomic mRNA, ESTs DNA sequences, data from various crops as well as gene expression, protein annotation and pathway data QTLs, and other relevant information.
- Database assembly is comprised of a toolbox of gene refining, structuring, annotation and analysis tools enabling to construct a tailored database for each gene discovery project.
- Gene refining and structuring tools enable to reliably detect splice variants and antisense transcripts, generating understanding of various potential phenotypic outcomes of a single gene.
- EST clustering and gene assembly For clustering and assembly of arabidopsis and rice genes the "genomic LEADS” version was employed. This tool allows most accurate clustering of ESTs and mRNA sequences on genome, and predicts gene structure as well as alternative splicing events and anti-sense transcription.
- Blast search Hypertext Transfer Protocol .-//World Wide Web (dot) ncbi (dot) nlm (dot) nih (dot) gov (dot) library (dot) vu (dot) edu (dot) au/BLAST/ ) against all plant UniProt (Hypertext Transfer Protocol://World Wide Web (dot) expasy (dot) uniprot (dot) org/) sequences was performed.
- Gene expression profiling Few data sources were exploited for gene expression profiling, namely microarray data and digital expression profile (see below). According to gene expression profile, a correlation analysis was performed to identify genes which are co-regulated under different development stages and environmental conditions.
- a digital expression profile summary was compiled for each cluster according to all keywords included in the sequence records comprising the cluster.
- Digital expression also known as electronic Northern Blot, is a tool that displays virtual expression profile based on the EST sequences forming the gene cluster.
- the tool can provide the expression profile of a cluster in terms of plant anatomy (in what tissues/organs is the gene expressed), developmental stage (the developmental stages at which a gene can be found) and profile of treatment (provides the physiological conditions under which a gene is expressed such as drought, cold, pathogen infection, etc).
- the digital expression Given a random distribution of ESTs in the different clusters, the digital expression provides a probability value that describes the probability of a cluster having a total of N ESTs to contain X ESTs from a certain collection of libraries.
- Orthologs and paralogs constitute two major types of homologs: The first evolved from a common ancestor by specialization, and the latter are related by duplication events. It is assumed that paralogs arising from ancient duplication events are likely to have diverged in function while true orthologs are more likely to retain identical function over evolutionary time.
- the present inventors have performed considerable work aimed at annotating sequences.
- ABST genes were identified to have a major impact on ABST when overexpressed in plants.
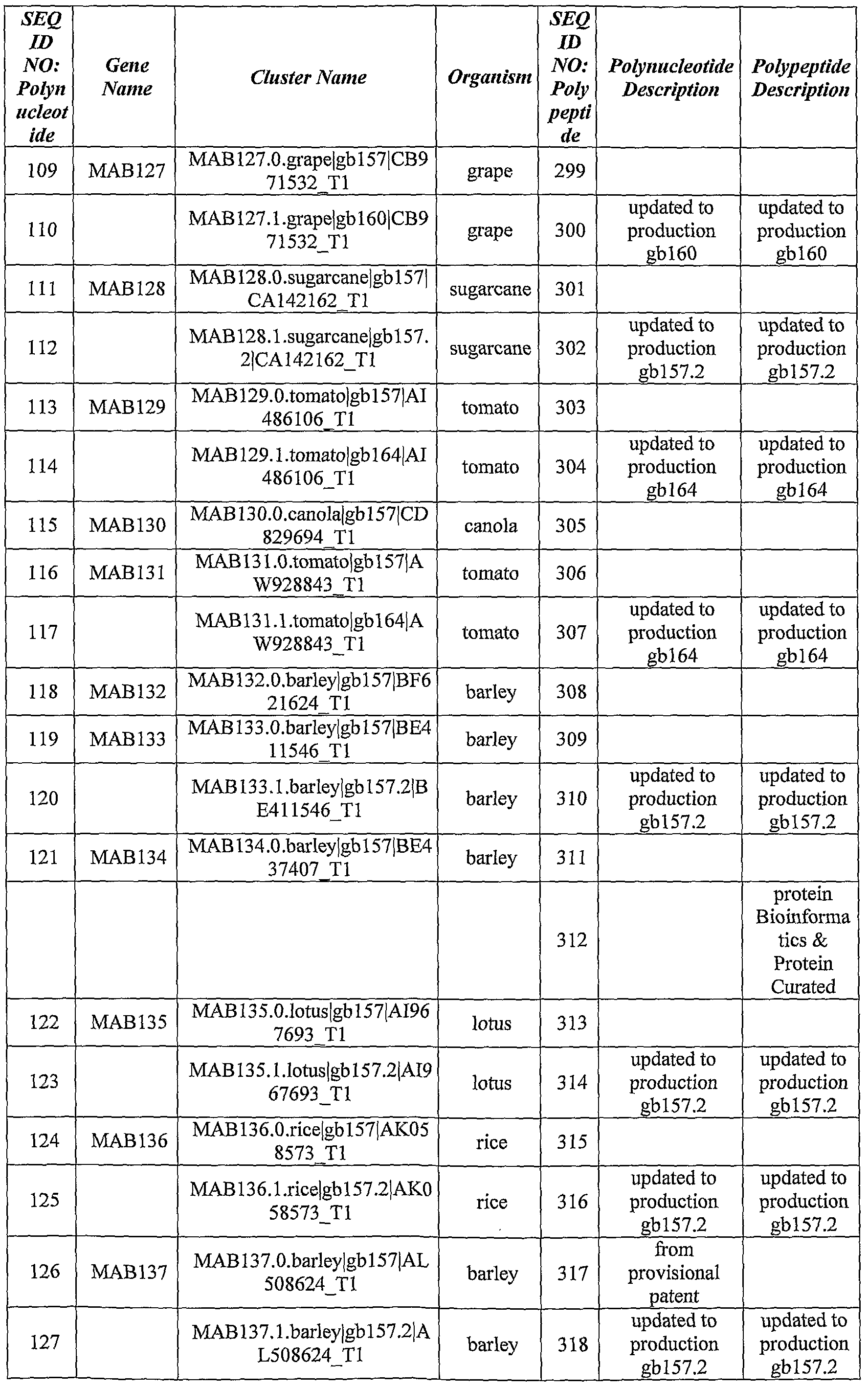
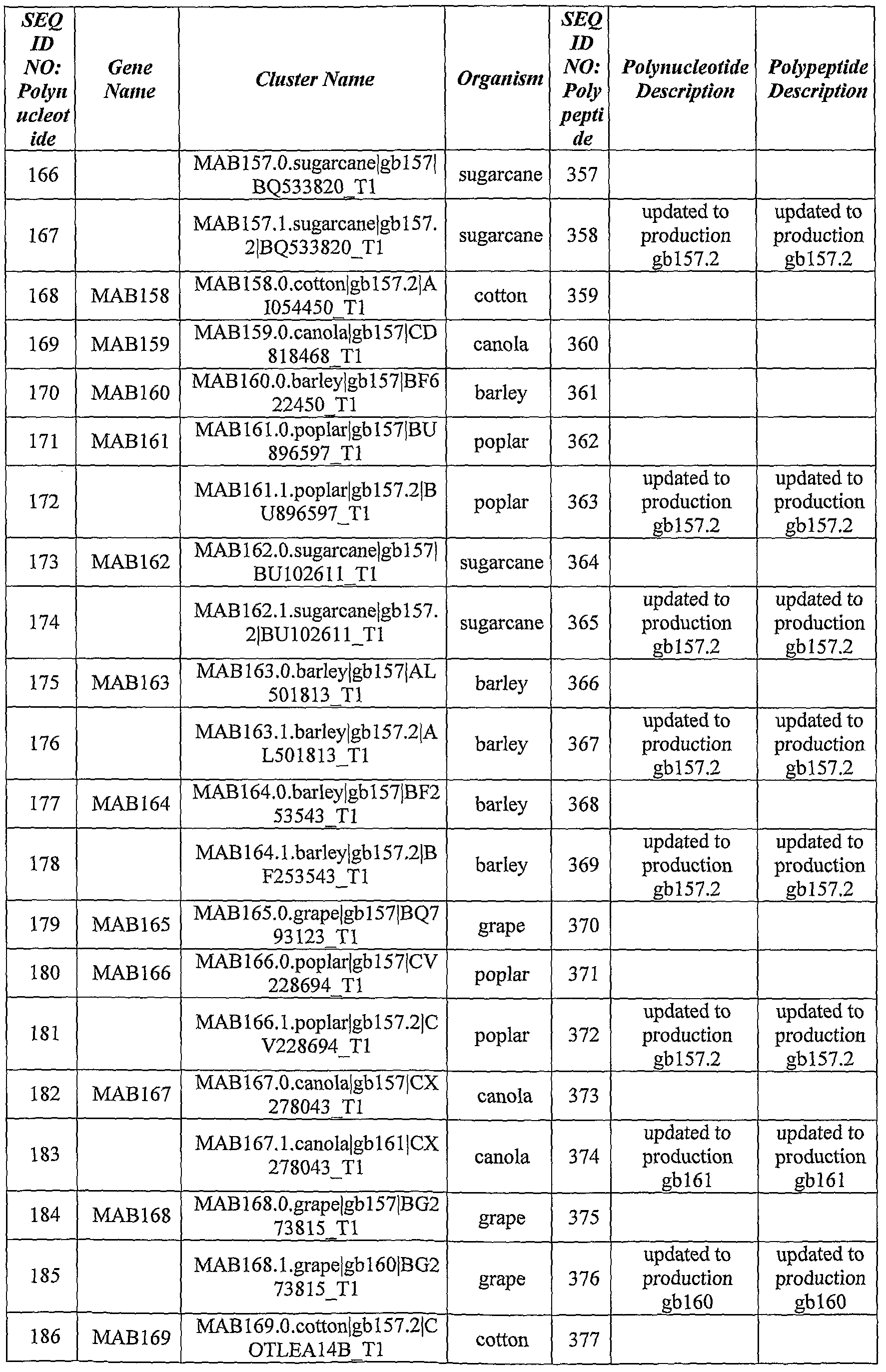
- the identified ABST genes, their curated polynucleotide and polypeptide sequences, as well as their updated sequences according to Genebank database are summarized in Table 1, hereinbelow.
- Table 1 Identi ed ABST Genes
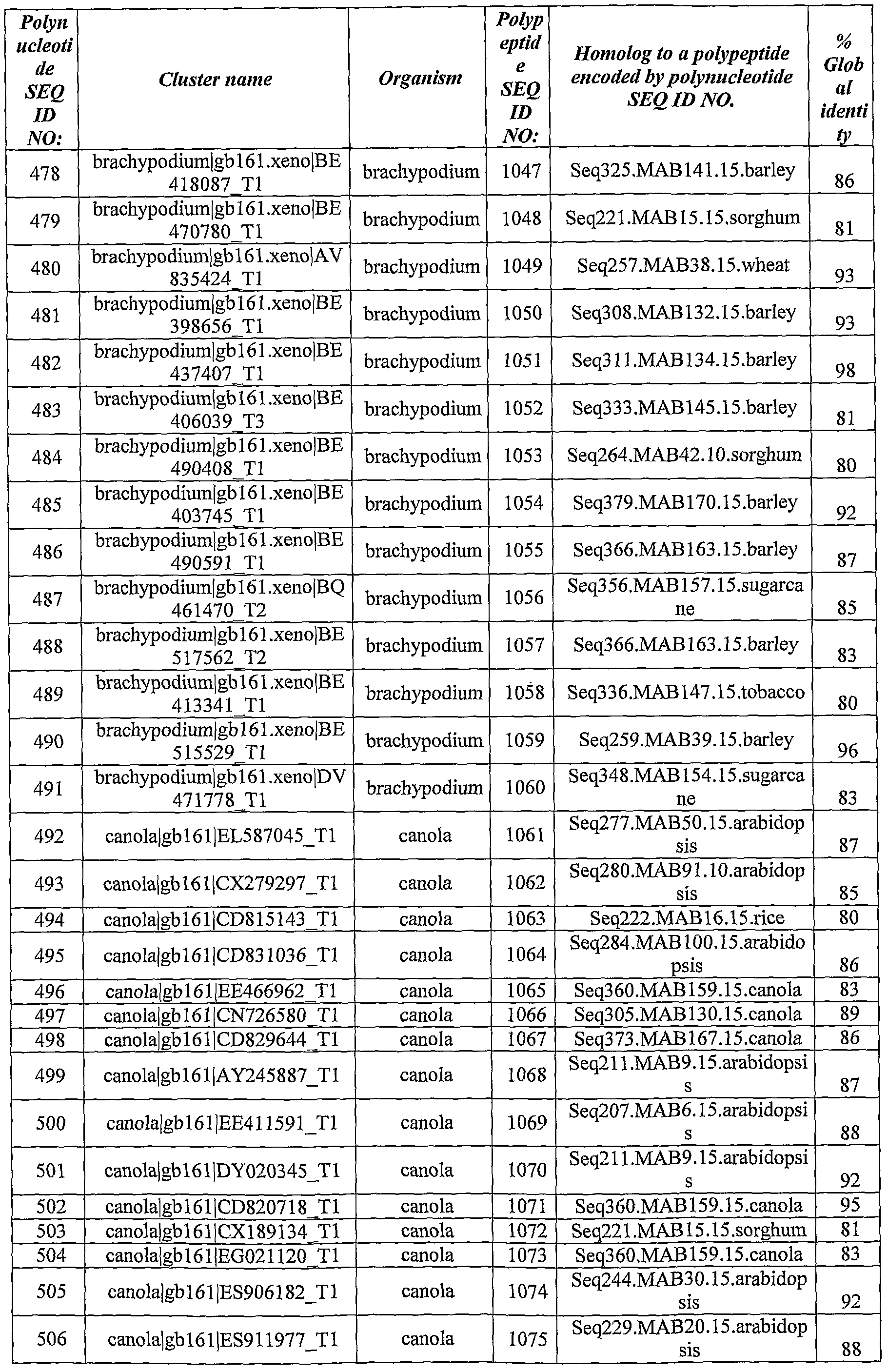
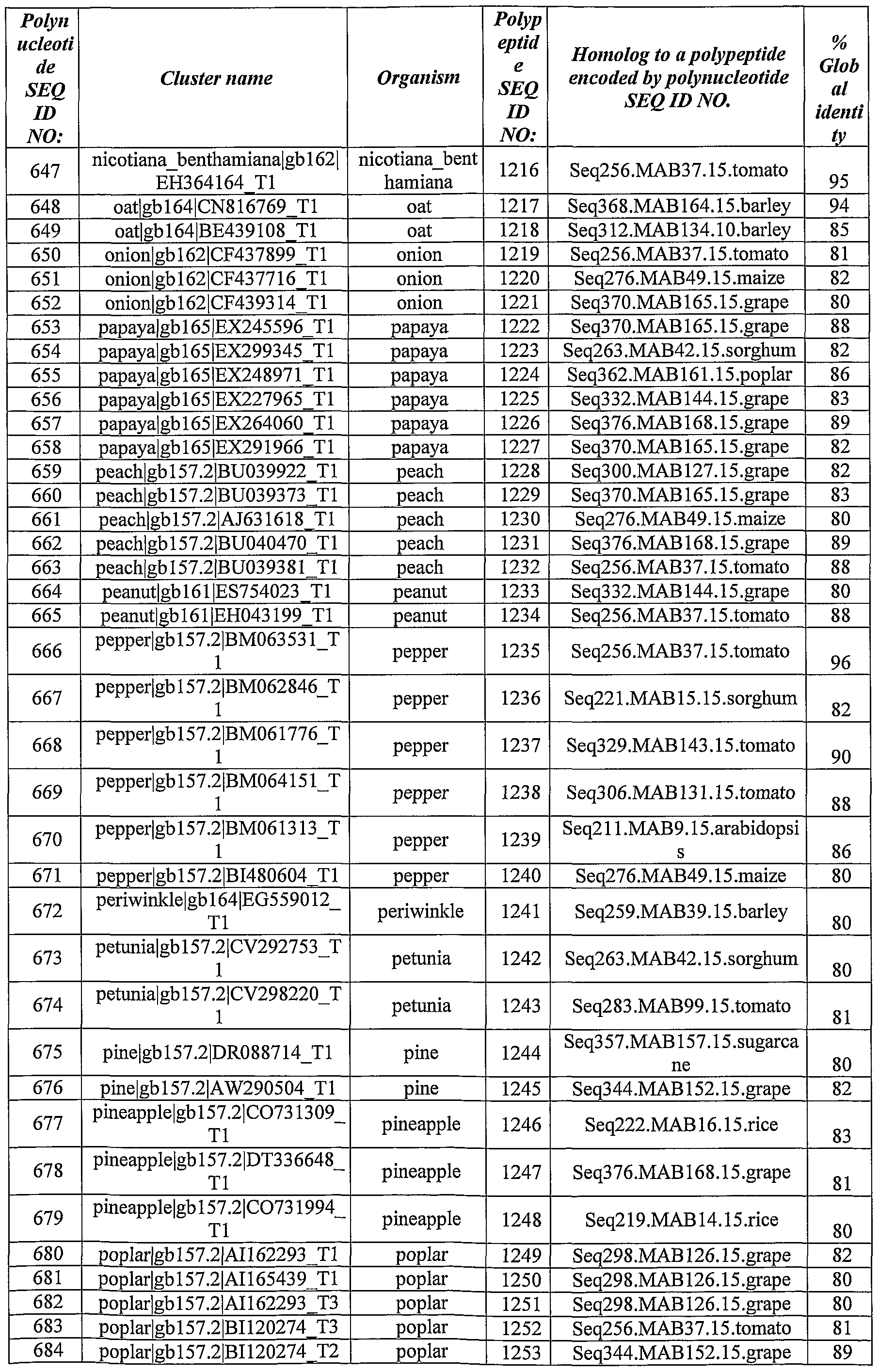
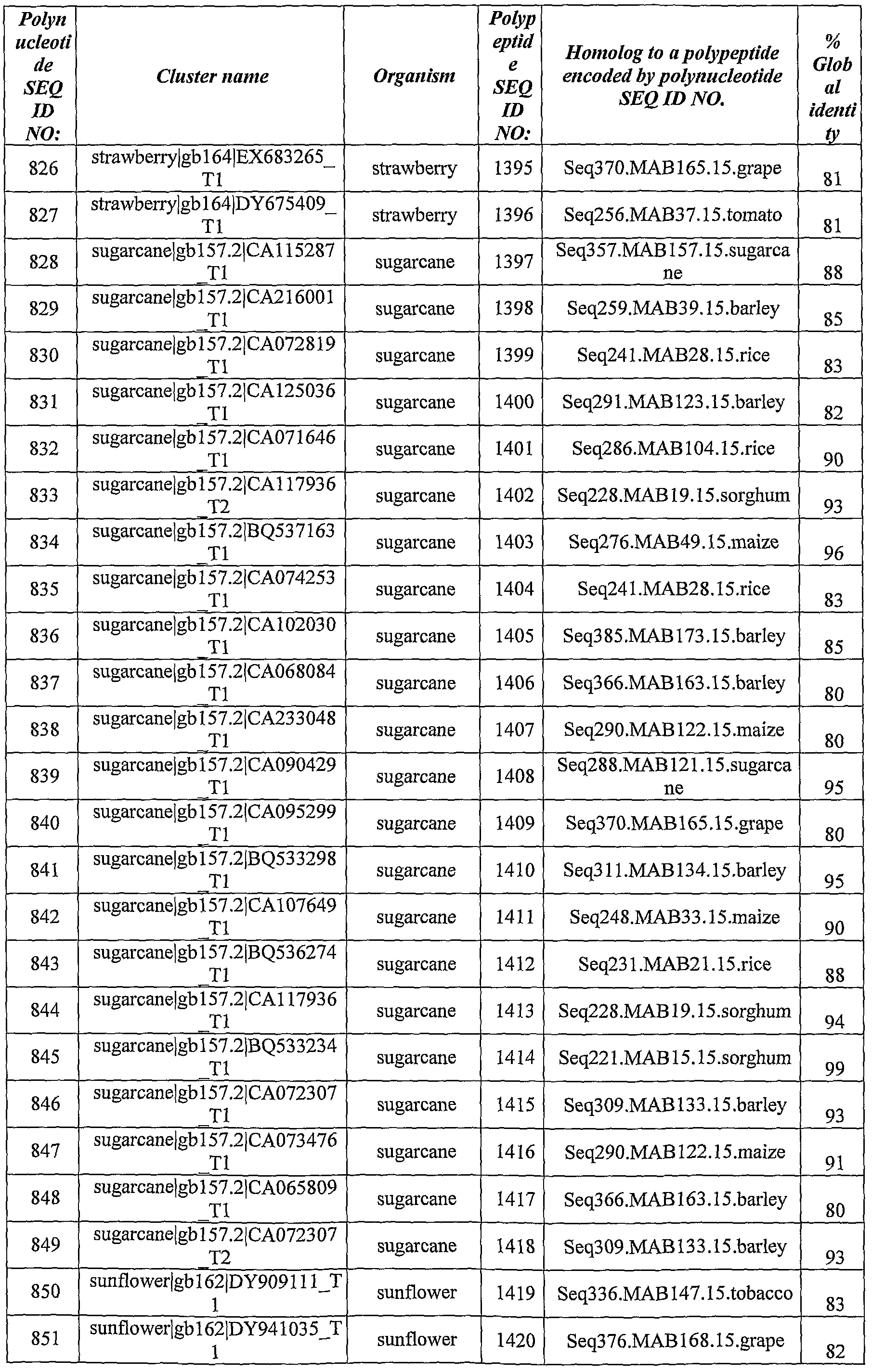
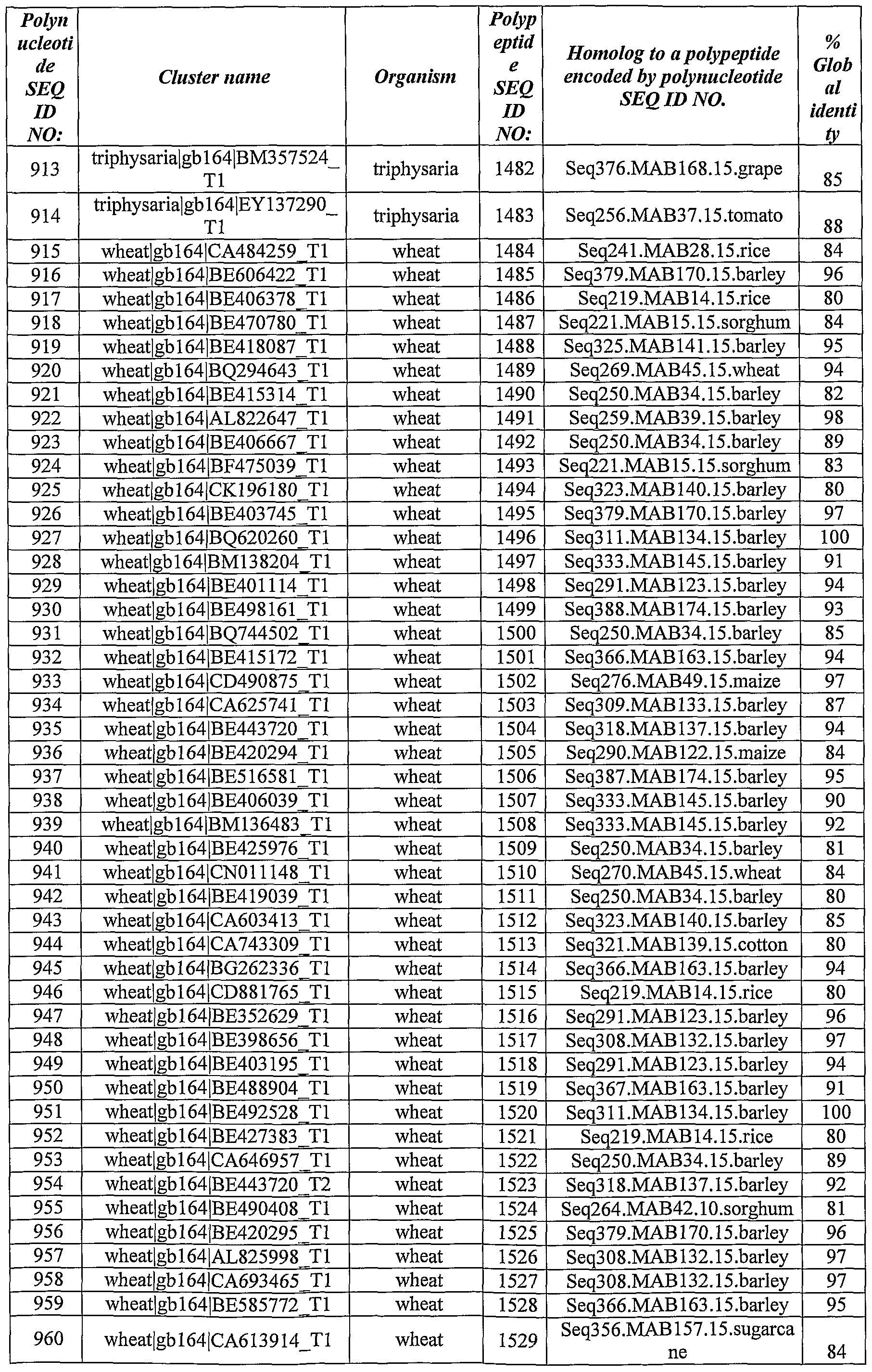
- Polynucleotides and polypeptides with significant homology to the identified ABST genes have been identified from the databases using BLAST software using the BlastX algorithm.
- the query nucleotide sequences were SEQ ID NOs: 1, 3, 5, 7, 9, 10, 11, 13, 15, 16, 17, 19, 21, 23, 25, 26, 28, 29, 30, 32, 34, 36, 37, 38, 40, 42, 44, 46, 48, 50, 52, 54, 55, 57, 59, 61, 63, 65, 67, 69, 71 ,73 ,75 ,77, 79, 81, 82, 84, 86, 88, 90, 91, 93, 94, 96, 98, 100, 101, 103, 105, 107, 109, 111, 113, 115, 116, 118, 119, 121, 122, 124, 126, 128, 130, 132, 134, 135, 138, 140, 142, 143, 145, 147, 149, 151,
- Table 2 *- Homology was calculated as % of identity over the aligned sequences.
- the query sequences were polynucleotide sequences SEQ ID NOs:l, 3, 5, 7, 9, 10, 11, 13, 15, 16, 17, 19, 21, 23, 25, 26, 28, 29, 30, 32, 34, 36, 37, 38, 40, 42, 44, 46, 48, 50, 52, 54, 55, 57, 59, 61, 63, 65, 67, 69, 71 ,73 ,75 ,77, 79, 81, 82, 84, 86, 88, 90, 91, 93, 94, 96, 98, 100, 101, 103, 105, 107, 109, 111, 113, 115, 116, 118, 119, 121, 122, 124, 126, 128, 130, 132, 134, 135, 138, 140, 142, 143, 145, 147, 149, 151, 153, 155, 157, 161, 163,
- DNA is designed in silico, based on the encoded amino-acid sequences of the ABST genes and using codon-usage Tables calculated from plant transcriptomes (example of such Tables can be found in the Codon Usage Database available online at Hypertext
- the optimized coding sequences are designed in a way that no changes are introduced in the encoded amino acid sequence while using codons preferred for expression in dicotyledonous plants (mainly tomato and Arabidopsis) and monocotyledonous plants such as maize. At least one silent mutation per 20 nucleotide base pairs is introduced in the sequence compared to the original sequences to avoid possible silencing when over- expressing the gene in the target crop.
- the following restriction enzymes sites are added- Sail, Xbal, BaniRI, Smal at the 5' end and Sad at the 3' end.
- the sequences synthesized by the supplier (GeneArt, Gmbh) are cloned in the pCR-Script plasmid. EXAMPLE 3
- Example 1 Selected genes from those presented in Example 1 were cloned into binary vectors for the generation of transgenic plants. For cloning, the full-length open reading frames (ORFs) were identified. EST clusters and in some cases mRNA sequences were analyzed to identify the entire open reading frame by comparing the results of several translation algorithms to known proteins from other plant species.
- ORFs open reading frames
- RNA extraction was performed using standard protocols described elsewhere (Sambrook J., E.F. Fritsch, and T. Maniatis. 1989. Molecular Cloning. A Laboratory Manual, 2nd Ed. Cold Spring Harbor Laboratory Press, New York.) which are well known to those skilled in the art.
- PCR products were purified using PCR purification kit (Qiagen) Usually, 2 sets of primers were prepared for the amplification of each gene, via nested PCR (meaning first amplifying the gene using external primers and then using the produced PCR product as a template for a second PCR reaction, where the internal set of primers are used). Alternatively, one or two of the internal primers were used for gene amplification, both in the first and the second PCR reactions (meaning only 2-3 primers were designed for a gene). To facilitate further cloning of the cDNAs, an 8-12 bp extension is added to the 5' of each internal primer. The primer extension includes an endonuclease restriction site.
- restriction sites are selected using two parameters: (a) the restriction site does not exist in the cDNA sequence; and (b) the restriction sites in the forward and reverse primers are designed such that the digested cDNA is inserted in the sense direction into the binary vector utilized for transformation.
- primers used for cloning ABST genes are provided. Table 3
- 69 iaoie j. rresente ⁇ are the cloned ABST genes and control gene(s) by the Gene Id number and the polynucleotide SEQ ID NO, Also presented are the primers and the restriction enzymes used to clone the ABST genes.
- PCR products were digested with the restriction endonucleases (Roche, Switzerland) according to the sites design in the primers (Table 3). Each digested PCR product was inserted into a high copy vector originated from pBlue-script KS plasmid vector (pBlue-script KS plasmid vector, Hypertext Transfer Protocol:// World Wide
- the cloned cDNA accompanied with the NOS terminator was introduced into the binary vectors pGI containing the At6669 promoter via digestion with appropriate restriction endonucleases.
- the cloned cDNA accompanied with the At6669 promoter was introduced into the pGI vector (that hasn't already contained the At6669 promoter).
- the insert was followed by single copy of the NOS terminator (SEQ ID NO: 1651).
- the digested products and the linearized plasmid vector were ligated using T4 DNA ligase enzyme (Roche, Switzerland).
- the pPI plasmid vector was constructed by inserting a synthetic poly-(A) signal sequence, originating from pGL3 basic plasmid vector (Promega, GenBank Accession No. U47295; nucleotides 4658-4811) into the HindIII restriction site of the binary vector pBI101.3 (Clontech, GenBank Accession No. U12640).
- pGI Figure 1 is similar to pPI, but the original gene in the back bone is GUS-Intron, rather than GUS.
- the Arabidopsis thaliana promoter sequence (set forth in SEQ ID NO: 1652) is inserted in the pPI binary vector, upstream to the cloned genes by using the restriction enzymes HindUl and Sail or BamHI (Roche), following by DNA ligation and binary plasmid extraction from positive E. coli colonies, as described above.
- the forward PCR primer was the primer set forth in SEQ ID NO: 1650 (from the At6669 promoter) and the reverse primer (derived from the specific cloned gene) was as follows:
- the reverse primer was SEQ ID NO:1570; for MAB14, the reverse primer was SEQ ID NO:1574; for MABlO, the reverse primer was SEQ ID NO:1577; for MAB25, the reverse primer was SEQ ID NO:1581; for MAB134, the reverse primer was SEQ ID NO:1585; for MAB99, the reverse primer was SEQ ID NO:1587; for MAB36, the reverse primer was SEQ ID NO: 1590; for MAB7, the reverse primer was SEQ ID NO: 1594; for MAB44, the reverse primer was SEQ ID NO: 1596; for MAB4, the reverse primer was SEQ ID NO:
- Synthetic sequences [such as of MAB 14, nucleotide SEQ ID NO:23, which encodes protein SEQ ID NO:219) of some of the cloned polynucleotides were ordered from a commercial supplier (Gene Art, GmbH). To optimize the coding sequence, codon-usage Tables calculated from plant transcriptomes were used [example of such Tables can be found in the Codon Usage Database available online at Hypertext Transfer Protocol ://World Wide Web (dot) kazusa (dot) or (dot) jp/codon/].
- the optimized coding sequences were designed in a way that no changes were introduced in the encoded amino acid sequence while using codons preferred for expression in dicotyledonous plants mainly tomato and Arabidopsis; and monocotyledonous plants such as maize. Such optimized sequences promote better translation rate and therefore higher protein expression levels. Parts of the sequences were ordered as the original sequences. To the optimized/non-optimized sequences flanking additional unique restriction enzymes sites were added to facilitate cloning genes in binary vectors.
- Promoters used Arabidopsis At6669 promoter (SEQ ID NO: 1652; which is SEQ ID NO:61 of WO04081173 to Evogene Ltd.).
- sequences of the cloned cDNAs are provided in SEQ ID NOs: 1530-1534, 1536-1545, 1547-1566, 1654, 1665, 1666, 1667 and 1668.
- the protein translation of the amplified cDNA sequence matched exactly that of the initial bioinformatics prediction of the protein sequences.
- the predicted polypeptide sequences of the cloned polynucleotides are provided in SEQ ID NOs:201, 212, 284, 213, 217, 317, 219, 221, 224, 225, 226, 227, 229, 237, 203, 247, 252, 205, 265, 267, 271, 277, 207, 208, 211, 283, 1655, 311, 334, and 254.
- Each of the binary vectors described in Example 3 above are used to transform Agrobacterium cells.
- Two additional binary constructs, having a GUS/Luciferase reporter gene replacing the ABST gene (positioned downstream of the At6669 promoter), are used as negative controls.
- the binary vectors are introduced to Agrobacterium tumefaciens GV301, or LB4404 competent cells (about 10 9 cells/mL) by electroporation.
- the electroporation is performed using a MicroPulser electroporator (Biorad), 0.2 cm cuvettes (Biorad) and EC-2 electroporation program (Biorad).
- the treated cells are cultured in LB liquid medium at 28 °C for 3 hours, then plated over LB agar supplemented with gentamycin (50 mg/L; for Agrobacterium strains GV301) or streptomycin (300 mg/L; for Agrobacterium strain LB4404) and kananiycin (50 mg/L) at 28 °C for 48 hours.
- Abrobacterium colonies which developed on the selective media were analyzed by PCR using the primers described above (Example 3) with respect to identification of positive binary vector colonies.
- the resulting PCR products are isolated and sequenced as described in Example 3 above, to verify that the correct ABST sequences are properly introduced to the Agrobacterium cells.
- T 0 plants Arabidopsis thaliana Columbia plants (T 0 plants) are transformed using the Floral Dip procedure described by Clough and Bent (10) and by Desfeux et al. (11), with minor modifications. Briefly, T 0 Plants are sown in 250 ml pots filled with wet peat- based growth mix. The pots are covered with aluminum foil and a plastic dome, kept at 4 °C for 3-4 days, then uncovered and incubated in a growth chamber at 18-24 °C under 16/8 hour light/dark cycles. The T 0 plants are ready for transformation six days before anthesis.
- the pellets comprising the Agrobacterium cells are re-suspended in a transformation medium containing half-strength (2.15 g/L) Murashige-Skoog (Duchefa); 0.044 ⁇ M benzylamino purine (Sigma); 112 ⁇ g/L B5 Gam strig vitamins (Sigma); 5 % sucrose; and 0.2 ml/L Silwet L-77 (OSI Specialists, CT) in double-distilled water, at pH of 5.7.
- Transformation of T 0 plants is performed by inverting each plant into an Agrobacterium suspension, such that the above ground plant tissue is submerged for 3-5 seconds.
- Each inoculated T 0 plant is immediately placed in a plastic tray, then covered with clear plastic dome to maintain humidity and is kept in the dark at room temperature for 18 hours, to facilitate infection and transformation.
- Transformed (transgenic) plants are then uncovered and transferred to a greenhouse for recovery and maturation.
- the transgenic To plants are grown in the greenhouse for 3-5 weeks until siliques are brown and dry. Seeds are harvested from plants and kept at room temperature until sowing.
- seeds collected from transgenic T 0 plants are surface-sterilized by soaking in 70 % ethanol for 1 minute, followed by soaking in 5 % sodium hypochloride and 0.05 % triton for 5 minutes.
- the surface-sterilized seeds are thoroughly washed in sterile distilled water then placed on culture plates containing half-strength Murashige-Skoog (Duchefa); 2 % sucrose; 0.8 % plant agar; 50 mM kanamycin; and 200 mM carbenicylin (Duchefa).
- the culture plates are incubated at 4 °C for 48 hours then transferred to a growth room at 25 °C for an additional week of incubation.
- T 1 Arabidopsis plants are transferred to a fresh culture plates for another week of incubation. Following incubation the T 1 plants are removed from culture plates and planted in growth mix contained in 250 ml pots. The transgenic plants are allowed to grow in a greenhouse to maturity. Seeds harvested from T 1 plants are cultured and grown to maturity as T 2 plants under the same conditions as used for culturing and growing the T 1 plants.
- PEG serves to simulate drought.
- Plants expressing the polynucleotides of the invention are compared to the average measurement of the control plants Mock- transgenic plants expressing the uidA reporter gene (GUS Intron - GUI) under the same promoter were used as control.
- Digital imaging - A laboratory image acquisition system, which consists of a digital reflex camera (Canon EOS 300D) attached with a 55 mm focal length lens (Canon EF-S series), mounted on a reproduction device (Kaiser RS), which included 4 light units (4 x 150 Watts light bulb) and located in a darkroom, was used for capturing images of plantlets sawn in square agar plates. The image capturing process was repeated every 7 days starting at day 0 till day
- An image analysis system was used, which consists of a personal desktop computer (Intel P4 3.0 GHz processor) and a public domain program - ImageJ 1.37 (Java based image processing program which was developed at the U. S National Institutes of Health and freely available on the internet at Hypertext Transfer Protocol://rsbweb (dot) nih (dot) gov/). Images were captured in resolution of 6 Mega Pixels (3072 x 2048 pixels) and stored in a low compression JPEG (Joint Photographic Experts Group standard) format. Next, analyzed data was saved to text files and processed using the JMP statistical analysis software (SAS institute).
- SAS institute JMP statistical analysis software
- RGR Relative Growth Rate
- Relative growth area rate ( ⁇ Area / ⁇ t) * (1/ Area t ⁇ ) ⁇ t is the current analyzed image day subtracted from the initial day (t-t ⁇ ).
- the relative growth area rate is in units of I/day and length growth rate is in units of I/day.
- Relative Growth Rate is determined by comparing the leaf area, root length and root coverage between each couple of sequential photographs, and results are used to resolve the effect of the gene introduced on plant vigor, under osmotic stress, as well as under optimal conditions.
- the effect of the gene introduced on biomass accumulation, under osmotic stress as well as under optimal conditions is determined by comparing the plants' fresh weight to control plants (GUI).
- results from the independent transformation events are evaluate for the overall influence of the gene (gene effect) and for each of the tested events (best event). Student's t test were applied, using significance of p ⁇ 0.05 or p ⁇ 0.1.
- the JMP statistics software package is used (Version 5.2.1, SAS Institute Inc., Gary, NC, USA).
- polynucleotide sequences of the invention were assayed for a number of desired traits.
- Tables 5-6 depict analyses of Leaf Area in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter under 25 %
- Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control, with A indicating a difference at a P ⁇ 0.05 level of significance and, A* a difference at a P ⁇ 0.1 level of significance.
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 7-9 depict analyses of Roots Coverage in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter under 25 %
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 10-11 depict analyses of Roots Length in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in 25% PEG. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 12-13 depict analyses of Leaf Area RGR in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in 25 % PEG. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- the SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above. Table 16
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 19-21 depict analyses of Roots Length RGR in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in 25% PEG. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Tables 22-23 depict analyses of Plant Fresh Weight in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in 25% PEG. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control. Table 22
- Tables 24-27 depict analyses of Leaf Area in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in normal conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- LSM Least square mean
- % improvement compare to control (GUI);
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 32-33 depict analyses of Roots Length in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in normal conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 34-36 depict analyses of Leaf Area RGR in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in normal conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Tables 37-41 depict analyses of Roots Coverage RGR in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in normal conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 42-46 depict analyses of Roots Length RGR in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in normal conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- LSM Least square mean
- % improvement compare to control (GUI);
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 47-48 depict analyses of Plant Fresh Weight in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in normal conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Assay 2 plant growth at Nitrogen deficiency under Tissue culture conditions —
- NUE Neuron Utilization Efficiency
- Plants expressing the polynucleotides of the invention are compared to the average measurement of the control plants (GUI- harboring the GUS gene under the same promoter) used in the same experiment. Digital imaging and statistical analysis - Parameters were measured and analyzed as described in Assay 1 above.
- Tables 49-53 depict analyses of Leaf Area in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in nitrogen deficient conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B) are significantly different from the control.
- LSM Least square mean
- % improvement compare to control (GUI)
- SEQ ID NOs. of the cloned genes (according to the Gene Id) which are exogenously expressed in the plants are provided in Table 3 above.
- Tables 54-57 depict analyses of Roots Coverage in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in nitrogen deficient conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Tables 58-61 depict analyses of Roots Length in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in nitrogen deficient conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Tables 65-69 depict analyses of Roots Coverage RGR in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in nitrogen deficient conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B, C) are significantly different from the control.
- Tables 70-74 depict analyses of Roots Length RGR in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in nitrogen deficient conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 3 ) are significantly different from the control.
- Tables 75-76 depict analyses of Plant Fresh Weight in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter in nitrogen deficient conditions. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control. Table 75
- ABS tolerance Yield and plant growth rate at high salinity concentration under greenhouse conditions - This assay follows the rosette area growth of plants grown in the greenhouse as well as seed yield at high salinity irrigation. Seeds were sown in agar media supplemented only with a selection agent (Kanamycm) and Hoagland solution under nursery conditions. The T 2 transgenic seedlings are then transplanted to 1.7 trays filled with peat and perlite. The trails were irrigated with tap water (provided from the pots' bottom). Half of the plants are irrigated with a salt solution (40-80 mM NaCl and 5 mM CaCl 2 ) to induce salinity stress (stress conditions).
- a salt solution 40-80 mM NaCl and 5 mM CaCl 2
- the other half of the plants are continued to be irrigated with tap water (normal conditions). All plants are grown in the greenhouse until plants reach the mature seeds stage, then harvested (the above ground tissue) and weighted (immediately or following drying in oven at 50 °C for 24 hour). High salinity conditions are achieved by irrigation with a solution containing 40-80 mM NaCl ("ABS" growth conditions) and are compared to regular growth conditions. The plants were analyzed for their overall size, growth rate, seed yield, and weight of 1,000 seeds, dry matter and harvest index (HI- seed yield / dry matter). Transgenic plants performance was compared to control plants grown in parallel under the same conditions. Mock- transgenic plants expressing the uidA reporter gene (GUS Intron - GUI) under the same promoter were used as control.
- GUS Intron - GUI uidA reporter gene
- the experiment is planned in nested randomized plot distribution. High salinity conditions are achieved by irrigation with a solution containing 40-80 mM NaCl ("ABS" growth conditions).
- Digital imaging - A laboratory image acquisition system which consists of a digital reflex camera (Canon EOS 300D) attached with a 55 mm focal length lens (Canon EF-S series), mounted on a reproduction device (Kaiser RS), which included 4 light units (4x150 Watts light bulb) was used for capturing images of plantlets.
- the image capturing process was repeated every 2-3 days starting at day 1 after sowing till day 10.
- the tubs were square shape include 1.7 liter trays. During the capture process, the trays were placed beneath the iron mount, while avoiding direct sun light and casting of shadows. This process was repeated every 2-3 days for up to 10 days.
- An image analysis system was used, which consists of a personal desktop computer (Intel P4 3.0 GHz processor) and a public domain program - ImageJ 1.37 (Java based image processing program which was developed at the U.
- Vegetative parameters analysis Using the digital analysis leaves data was calculated, including leaf Average area, Rosette diameter and rosette area.
- the Relative Growth Rate (RGR) for the rosette parameters was calculated according to Formula I as described in Example 6.
- RGR Relative Growth Rate
- the weight of 1000 seeds was determine as follows: seeds were scattered on a glass tray and a picture was taken. Each sample was weighted and then using the digital analysis, the number of seeds in each sample was calculated. 1000 seeds weight was calculated using formula II:
- Seed Weight number of seed in sample/ sample weight X 1000
- Tables 77-86 depict analyses of Rosette Area in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A 5 B 5 ) are significantly different from the control. Table 77
- Tables 87-96 depict analyses of Rosette Diameter in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 97-105 depict analyses of Leaf Average Area in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 106-111 depict analyses of RGR Rosette Area [cm ⁇ 2] of plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control. Table 106
- Tables 112-118 depict analyses of RGR of Rosette Diameter in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control. Table 112
- Tables 119-121 depict analyses of RGR of Leaf Average Area [cm ⁇ 2] in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Table 123 depicts analyses of Plot Dry weight (DW) in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 124-126 depict analyses of 1000 Seeds Weight in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A 5 B 3 ) are significantly different from the control.
- Table 130 depicts analyses of Harvest Index in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 131-140 depict analyses of Rosette Area in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Tables 141-148 depict analyses of Rosette Diameter in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 149-157 depict analyses of Leaf Average Area in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 158-166 depict analyses of RGR Rosette Area [cm A 2] of plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 167-175 depict analyses of RGR of Rosette Diameter in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 176-178 depict analyses of RGR of Leaf Average Area [cm ⁇ 2] in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Tables 179-180 depict analyses of RGR of Leaf Average Area [cm ⁇ 2] in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Tables 181-182 depict analyses of Plot Dry weight (DW) in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B 5 ) are significantly different from the control.
- Tables 183-185 depict analyses of 1000 Seeds Weight in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Tables 186-187 depict analyses of Seed Yield per Plant in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- Table 188 depicts analyses of Harvest Index in plants overexpressing the polynucleotides of the invention under the regulation of 6669 promoter. Each Table represents an independent experiment, using 4 independent events per gene. Genes not connected by same letter as the control (A, B,) are significantly different from the control.
- tomato M82 seeds were previously sterilized with Na-hipochloride 3 % + 2-3 drops of Tween 20 (Polysorbate 20). Seeds were washed 3 times with distilled sterile water. Seeds were then germinated in full strength
- Assay 1 Tomato field trial under regular and water deficient regimes -
- the tomato field trial was planned as a one source dripping irrigation (OSDI) system similar to a standard farmer field. Since water deficiency is applied in a relatively uniform manner, it allows measuring the effect of drought on small size populations of plants.
- the OSDI method was developed on the basis of the line source sprinklers irrigation system (Hanks et al. 1976 Soil Sci. Soc Am. J. 40 p. 426-429) with some significant modifications. Instead of sprinkler irrigation, dripping irrigation was used. In order to create a uniform and deep wet layer (at least 60 cm depth), and not the onion shape layer that is typically created by dripping irrigation, a low pressure compensating dripping irrigation system was used.
- Severe Stress 50 % of the optimal amount of water irrigation was applied once a day (at same time as regular irrigation is applied) All fertilizers were applied according to local standard protocols. Nitrogen was equally applied, as recommended, to all the treatments through the irrigation system.
- Each row 193 cm wide, contained two dripping irrigation lines creating coverage of six drippers per 1 sq. m.
- the irrigation control was performed separately for each treatment.
- the experiment was structured in a four randomized block design, eight plants per plot. The different water regimes were initiated only four weeks three transplantation, when plants initiated the flowering stage. Water availability in the soil was recorded using tensiometers (used to determine matric water potential ⁇ m which allows to evaluate the stress severeness).
- Assay 2 Tomato salt bath experiment - Transgenic tomato seeds are sown in trays containing growth denitrified media. Seedlings are germinated under nursery conditions. The experimental model used was 3 blocks random distributed, where 10 plants per events were sown in each block. At the stage of first true leaf, trays are transferred to different "tanks" containing growth solution of 300 mM NaCl. For normal treatment, a full Hoagland solution was applied. 5 events for each gene are evaluated while null segregating populations are used as negative controls. The experiment is performed for a period of 8 weeks, where parameters such as chlorophyll content (measured as SPAD units), plant biomass (FW and DW) are measured.
- parameters such as chlorophyll content (measured as SPAD units), plant biomass (FW and DW) are measured.
- Nguyen BD Brar DS, Bui BC, Nguyen TV, Pham LN, Nguyen HT (2003). Identification and mapping of the QTL for aluminum tolerance introgressed from the new source, ORYZA RUFIPOGON Griff, into indica rice ( Oryza sativa L.). Theor Appl Genet. 106:583-93.
- Floral dip a simplified method for Agrobacterium- mediated transformation of Arabidopsis thaliana. Plant J 16:735-43.
- File information is provided as: File name/byte size/date of creation/operating system/machine format.
- CD-ROMl (1 file of SEQUENCE LISTING):
Landscapes
- Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Biotechnology (AREA)
- Molecular Biology (AREA)
- Zoology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biomedical Technology (AREA)
- General Engineering & Computer Science (AREA)
- Wood Science & Technology (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- Biophysics (AREA)
- Physics & Mathematics (AREA)
- Plant Pathology (AREA)
- Microbiology (AREA)
- Cell Biology (AREA)
- Botany (AREA)
- Environmental Sciences (AREA)
- Developmental Biology & Embryology (AREA)
- Gastroenterology & Hepatology (AREA)
- Medicinal Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Oil, Petroleum & Natural Gas (AREA)
- Nutrition Science (AREA)
- Physiology (AREA)
- Breeding Of Plants And Reproduction By Means Of Culturing (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
- Agricultural Chemicals And Associated Chemicals (AREA)
Abstract
Description
Claims
Priority Applications (15)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN2008801094649A CN102037127A (en) | 2007-07-24 | 2008-07-24 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| BR122020022203-4A BR122020022203B1 (en) | 2007-07-24 | 2008-07-24 | method of increasing the growth rate of a plant |
| CA2694481A CA2694481C (en) | 2007-07-24 | 2008-07-24 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| EP08776651.5A EP2183371B1 (en) | 2007-07-24 | 2008-07-24 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| AU2008278654A AU2008278654B2 (en) | 2007-07-24 | 2008-07-24 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| BR122020022199-2A BR122020022199B1 (en) | 2007-07-24 | 2008-07-24 | METHOD OF INCREASING THE TOLERANCE OF A PLANT TO ABIOTIC STRESS, BIOMASS AND/OR PRODUCTIVITY OF A PLANT, AND CONSTRUCTION OF ISOLATED NUCLEIC ACID |
| BRPI0812742-5A BRPI0812742B1 (en) | 2007-07-24 | 2008-07-24 | method of increasing biomass, growth rate, seed productivity, nitrogen use efficiency, abiotic stress of a plant, root length, root cover, growth rate of the rosette area, and of the growth rate of the rosette diameter of a plant |
| ES08776651.5T ES2547305T3 (en) | 2007-07-24 | 2008-07-24 | Polynucleotides, polypeptides encoded by them, and methods of using them to increase tolerance to abiotic stress and / or biomass and / or yield in plants that express them |
| US12/669,975 US8686227B2 (en) | 2007-07-24 | 2008-07-24 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| ZA2010/01205A ZA201001205B (en) | 2007-07-24 | 2010-02-19 | Polynucleotides, polypeptides encoded thereby, and mthods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US14/071,715 US9518267B2 (en) | 2007-07-24 | 2013-11-05 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US15/278,086 US10155957B2 (en) | 2007-07-24 | 2016-09-28 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US16/154,833 US10961544B2 (en) | 2007-07-24 | 2018-10-09 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US16/218,559 US10995341B2 (en) | 2007-07-24 | 2018-12-13 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US17/177,309 US20210171974A1 (en) | 2007-07-24 | 2021-02-17 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US93504607P | 2007-07-24 | 2007-07-24 | |
| US60/935,046 | 2007-07-24 |
Related Child Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US12/669,975 A-371-Of-International US8686227B2 (en) | 2007-07-24 | 2008-07-24 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US14/071,715 Continuation US9518267B2 (en) | 2007-07-24 | 2013-11-05 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2009013750A2 true WO2009013750A2 (en) | 2009-01-29 |
| WO2009013750A3 WO2009013750A3 (en) | 2010-03-04 |
Family
ID=40281935
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/IL2008/001024 WO2009013750A2 (en) | 2007-07-24 | 2008-07-24 | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
Country Status (9)
| Country | Link |
|---|---|
| US (6) | US8686227B2 (en) |
| EP (2) | EP2183371B1 (en) |
| CN (1) | CN102037127A (en) |
| AU (1) | AU2008278654B2 (en) |
| BR (3) | BR122020022203B1 (en) |
| CA (3) | CA2694481C (en) |
| ES (2) | ES2685631T3 (en) |
| WO (1) | WO2009013750A2 (en) |
| ZA (1) | ZA201001205B (en) |
Cited By (37)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2010097343A1 (en) * | 2009-02-25 | 2010-09-02 | Basf Plant Science Company Gmbh | Plants having enhanced yield-related traits and a method for making the same |
| WO2010100595A2 (en) | 2009-03-02 | 2010-09-10 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US7910800B2 (en) | 2005-08-15 | 2011-03-22 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| WO2011135527A2 (en) | 2010-04-28 | 2011-11-03 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| WO2012007945A3 (en) * | 2010-07-12 | 2012-06-14 | The State Of Israel, Ministry Of Agriculture & Rural Development, Agricultural Research Organization, (A.R.O.), Volcani Center | Isolated polynucleotides and methods and plants using same for regulating plant acidity |
| EP2519097A2 (en) * | 2009-12-28 | 2012-11-07 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US8847008B2 (en) | 2008-05-22 | 2014-09-30 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant utility |
| US8937215B2 (en) | 2009-08-04 | 2015-01-20 | Evogene Ltd. | Polynucleotides and polypeptides for increasing desirable plant qualities |
| US8952218B2 (en) | 2008-12-29 | 2015-02-10 | Evogene Ltd. | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance, biomass and/or yield in plants expressing same |
| US8962915B2 (en) | 2004-06-14 | 2015-02-24 | Evogene Ltd. | Isolated polypeptides, polynucleotides encoding same, transgenic plants expressing same and methods of using same |
| EP2840141A1 (en) * | 2013-08-21 | 2015-02-25 | Industry-Academic Cooperation Foundation, Yonsei University | Gene implicated in abiotic stress tolerance and growth accelerating and use thereof |
| US9012728B2 (en) | 2004-06-14 | 2015-04-21 | Evogene Ltd. | Polynucleotides and polypeptides involved in plant fiber development and methods of using same |
| US9018445B2 (en) | 2008-08-18 | 2015-04-28 | Evogene Ltd. | Use of CAD genes to increase nitrogen use efficiency and low nitrogen tolerance to a plant |
| EP2440033B1 (en) * | 2009-06-10 | 2017-03-15 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US9631000B2 (en) | 2006-12-20 | 2017-04-25 | Evogene Ltd. | Polynucleotides and polypeptides involved in plant fiber development and methods of using same |
| US9670501B2 (en) | 2007-12-27 | 2017-06-06 | Evogene Ltd. | Isolated polypeptides, polynucleotides useful for modifying water user efficiency, fertilizer use efficiency, biotic/abiotic stress tolerance, yield and biomass in plants |
| US9745595B2 (en) | 2008-10-30 | 2017-08-29 | Evogene Ltd. | Methods of increasing biomass and/or growth rate of a plant under non-stress conditions |
| US9771598B2 (en) | 2012-12-26 | 2017-09-26 | Evogene Ltd. | Isolated polynucleotides and polypeptides, construct and plants comprising same and methods of using same for increasing nitrogen use efficiency of plants |
| US9834782B2 (en) | 2012-05-28 | 2017-12-05 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US9890389B2 (en) | 2012-12-25 | 2018-02-13 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency of plants |
| US9920330B2 (en) | 2012-02-29 | 2018-03-20 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US9920329B2 (en) | 2013-05-22 | 2018-03-20 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US9976157B2 (en) | 2011-08-23 | 2018-05-22 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10000768B2 (en) | 2011-11-21 | 2018-06-19 | Syngenta Participations Ag | Compositions and methods for increasing nematode resistance in plants |
| US10006042B2 (en) | 2013-08-27 | 2018-06-26 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10036031B2 (en) | 2007-04-09 | 2018-07-31 | Evogene Ltd. | Polynucleotides, polypeptides and methods for increasing oil content, growth rate and biomass of plants |
| US10113176B2 (en) | 2011-11-28 | 2018-10-30 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US10155957B2 (en) | 2007-07-24 | 2018-12-18 | Evogene Ltd. | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US10184132B2 (en) | 2003-05-22 | 2019-01-22 | Evogene Ltd. | Methods of increasing abiotic stress tolerance, yield and/or biomass in plants |
| US10260073B2 (en) | 2011-12-28 | 2019-04-16 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing yield of plants |
| US10457954B2 (en) | 2010-08-30 | 2019-10-29 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US10457952B2 (en) | 2010-12-22 | 2019-10-29 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for improving plant properties |
| US10760088B2 (en) | 2011-05-03 | 2020-09-01 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US10766935B2 (en) | 2015-12-28 | 2020-09-08 | Evogene Ltd. | Plant traits conferred by isolated polynucleotides and polypeptides |
| US10858403B2 (en) | 2014-08-27 | 2020-12-08 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10858665B2 (en) | 2012-08-27 | 2020-12-08 | Evogene Ltd. | Isolated polynucleotides, polypeptides and methods of using same for increasing abiotic stress tolerance, biomass and yield of plants |
| US10975383B2 (en) | 2014-05-28 | 2021-04-13 | Evogene Ltd. | Isolated polynucleotides, polypeptides and methods of using same for increasing abiotic stress tolerance, biomass and yield of plants |
Families Citing this family (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| KR101416506B1 (en) | 2012-08-10 | 2014-07-09 | 연세대학교 산학협력단 | Gene Implicated in Abiotic Stress Tolerance and Growth Accelerating and Use Thereof |
| CN111233988B (en) * | 2018-11-29 | 2021-11-30 | 上海交通大学 | Eggplant potassium ion channel protein SmAKT1, and coding gene and application thereof |
| CN114574499A (en) * | 2020-11-30 | 2022-06-03 | 华中农业大学 | Application of OsREP3 gene in controlling drought resistance of rice |
Citations (9)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5296462A (en) | 1992-11-19 | 1994-03-22 | Board Of Trustees Operating Michigan State University | Method and compositions using polypeptides of arabidopsis thaliana |
| US5356816A (en) | 1991-11-19 | 1994-10-18 | Board Of Trustees Operating Michigan State University | Method and compositions using polypeptides of arabidopsis thaliana |
| US20030056249A1 (en) | 2001-06-12 | 2003-03-20 | Simmons Carl R. | Anti-apoptosis genes and methods of use thereof |
| US6670528B1 (en) | 1998-10-14 | 2003-12-30 | Independent Administrative Institute, Japan International Research Center For Agricultural Sciences | Environmental stress-tolerant plants |
| US6720477B2 (en) | 2000-04-07 | 2004-04-13 | Basf Plant Science Gmbh | Signal transduction stress-related proteins and methods of use in plants |
| WO2004104162A2 (en) | 2003-05-22 | 2004-12-02 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| US20060183137A1 (en) | 2000-08-24 | 2006-08-17 | The Scripps Research Institute | Stress-regulated genes of plants, transgenic plants containing same, and methods of use |
| WO2007020638A2 (en) | 2005-08-15 | 2007-02-22 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| WO2007049275A2 (en) | 2005-10-24 | 2007-05-03 | Evogene Ltd. | Isolated polypeptides, polynucleotides encoding same, transgenic plants expressing same and methods of using same |
Family Cites Families (192)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| NL154600B (en) * | 1971-02-10 | 1977-09-15 | Organon Nv | METHOD FOR THE DETERMINATION AND DETERMINATION OF SPECIFIC BINDING PROTEINS AND THEIR CORRESPONDING BINDABLE SUBSTANCES. |
| NL154598B (en) | 1970-11-10 | 1977-09-15 | Organon Nv | PROCEDURE FOR DETERMINING AND DETERMINING LOW MOLECULAR COMPOUNDS AND PROTEINS THAT CAN SPECIFICALLY BIND THESE COMPOUNDS AND TEST PACKAGING. |
| NL154599B (en) | 1970-12-28 | 1977-09-15 | Organon Nv | PROCEDURE FOR DETERMINING AND DETERMINING SPECIFIC BINDING PROTEINS AND THEIR CORRESPONDING BINDABLE SUBSTANCES, AND TEST PACKAGING. |
| US3901654A (en) * | 1971-06-21 | 1975-08-26 | Biological Developments | Receptor assays of biologically active compounds employing biologically specific receptors |
| US3853987A (en) | 1971-09-01 | 1974-12-10 | W Dreyer | Immunological reagent and radioimmuno assay |
| US3867517A (en) | 1971-12-21 | 1975-02-18 | Abbott Lab | Direct radioimmunoassay for antigens and their antibodies |
| NL171930C (en) | 1972-05-11 | 1983-06-01 | Akzo Nv | METHOD FOR DETERMINING AND DETERMINING BITES AND TEST PACKAGING. |
| US3850578A (en) | 1973-03-12 | 1974-11-26 | H Mcconnell | Process for assaying for biologically active molecules |
| US3935074A (en) | 1973-12-17 | 1976-01-27 | Syva Company | Antibody steric hindrance immunoassay with two antibodies |
| US3996345A (en) | 1974-08-12 | 1976-12-07 | Syva Company | Fluorescence quenching with immunological pairs in immunoassays |
| US4034074A (en) | 1974-09-19 | 1977-07-05 | The Board Of Trustees Of Leland Stanford Junior University | Universal reagent 2-site immunoradiometric assay using labelled anti (IgG) |
| US3984533A (en) | 1975-11-13 | 1976-10-05 | General Electric Company | Electrophoretic method of detecting antigen-antibody reaction |
| US4098876A (en) * | 1976-10-26 | 1978-07-04 | Corning Glass Works | Reverse sandwich immunoassay |
| US4879219A (en) | 1980-09-19 | 1989-11-07 | General Hospital Corporation | Immunoassay utilizing monoclonal high affinity IgM antibodies |
| CA1192510A (en) | 1981-05-27 | 1985-08-27 | Lawrence E. Pelcher | Rna plant virus vector or portion thereof, a method of construction thereof, and a method of producing a gene derived product therefrom |
| US5504200A (en) * | 1983-04-15 | 1996-04-02 | Mycogen Plant Science, Inc. | Plant gene expression |
| JPS6054684A (en) | 1983-09-05 | 1985-03-29 | Teijin Ltd | Novel dna and hybrid dna |
| US5011771A (en) | 1984-04-12 | 1991-04-30 | The General Hospital Corporation | Multiepitopic immunometric assay |
| US4666828A (en) * | 1984-08-15 | 1987-05-19 | The General Hospital Corporation | Test for Huntington's disease |
| US4946674A (en) * | 1984-10-05 | 1990-08-07 | Bioferon Biochemische Substanzen Gmbh & Co. | Process for treatment of rheumatic diseases |
| US4945050A (en) * | 1984-11-13 | 1990-07-31 | Cornell Research Foundation, Inc. | Method for transporting substances into living cells and tissues and apparatus therefor |
| US5420034A (en) * | 1986-07-31 | 1995-05-30 | Calgene, Inc. | Seed-specific transcriptional regulation |
| US4943674A (en) | 1987-05-26 | 1990-07-24 | Calgene, Inc. | Fruit specific transcriptional factors |
| CA1288073C (en) | 1985-03-07 | 1991-08-27 | Paul G. Ahlquist | Rna transformation vector |
| US4683202A (en) * | 1985-03-28 | 1987-07-28 | Cetus Corporation | Process for amplifying nucleic acid sequences |
| US4801531A (en) * | 1985-04-17 | 1989-01-31 | Biotechnology Research Partners, Ltd. | Apo AI/CIII genomic polymorphisms predictive of atherosclerosis |
| US5569597A (en) | 1985-05-13 | 1996-10-29 | Ciba Geigy Corp. | Methods of inserting viral DNA into plant material |
| GB8608850D0 (en) | 1986-04-11 | 1986-05-14 | Diatech Ltd | Packaging system |
| JPS6314693A (en) | 1986-07-04 | 1988-01-21 | Sumitomo Chem Co Ltd | Plant virus rna vector |
| US5268463A (en) | 1986-11-11 | 1993-12-07 | Jefferson Richard A | Plant promoter α-glucuronidase gene construct |
| US5608142A (en) * | 1986-12-03 | 1997-03-04 | Agracetus, Inc. | Insecticidal cotton plants |
| EP0278667B1 (en) | 1987-02-09 | 1994-07-20 | Mycogen Plant Science, Inc. | Hybrid RNA virus |
| US5316931A (en) * | 1988-02-26 | 1994-05-31 | Biosource Genetics Corp. | Plant viral vectors having heterologous subgenomic promoters for systemic expression of foreign genes |
| US5693507A (en) | 1988-09-26 | 1997-12-02 | Auburn University | Genetic engineering of plant chloroplasts |
| US5495070A (en) * | 1988-10-04 | 1996-02-27 | Agracetus, Inc. | Genetically engineering cotton plants for altered fiber |
| US5597718A (en) * | 1988-10-04 | 1997-01-28 | Agracetus | Genetically engineering cotton plants for altered fiber |
| US5272057A (en) | 1988-10-14 | 1993-12-21 | Georgetown University | Method of detecting a predisposition to cancer by the use of restriction fragment length polymorphism of the gene for human poly (ADP-ribose) polymerase |
| US5302523A (en) | 1989-06-21 | 1994-04-12 | Zeneca Limited | Transformation of plant cells |
| US6329570B1 (en) | 1989-07-19 | 2001-12-11 | Calgene, Llc | Cotton modification using ovary-tissue transcriptional factors |
| US5192659A (en) * | 1989-08-25 | 1993-03-09 | Genetype Ag | Intron sequence analysis method for detection of adjacent and remote locus alleles as haplotypes |
| US5859330A (en) * | 1989-12-12 | 1999-01-12 | Epitope, Inc. | Regulated expression of heterologous genes in plants and transgenic fruit with a modified ripening phenotype |
| ES2187497T3 (en) | 1990-04-12 | 2003-06-16 | Syngenta Participations Ag | PROMOTERS PREFERREDLY IN FABRICS. |
| US5498830A (en) * | 1990-06-18 | 1996-03-12 | Monsanto Company | Decreased oil content in plant seeds |
| US5187267A (en) * | 1990-06-19 | 1993-02-16 | Calgene, Inc. | Plant proteins, promoters, coding sequences and use |
| US5399680A (en) * | 1991-05-22 | 1995-03-21 | The Salk Institute For Biological Studies | Rice chitinase promoter |
| DE69230290T2 (en) | 1991-08-27 | 2000-07-20 | Novartis Ag, Basel | PROTEINS WITH INSECTICIDAL PROPERTIES AGAINST HOMOPTERAN INSECTS AND THEIR USE IN PLANT PROTECTION |
| EP0612208B1 (en) | 1991-10-04 | 2004-09-15 | North Carolina State University | Pathogen-resistant transgenic plants |
| UA48104C2 (en) | 1991-10-04 | 2002-08-15 | Новартіс Аг | Dna fragment including sequence that codes an insecticide protein with optimization for corn, dna fragment providing directed preferable for the stem core expression of the structural gene of the plant related to it, dna fragment providing specific for the pollen expression of related to it structural gene in the plant, recombinant dna molecule, method for obtaining a coding sequence of the insecticide protein optimized for corn, method of corn plants protection at least against one pest insect |
| US5281521A (en) * | 1992-07-20 | 1994-01-25 | The Trustees Of The University Of Pennsylvania | Modified avidin-biotin technique |
| FR2697723B1 (en) * | 1992-11-06 | 1995-03-03 | Ungda | Use of polyether ionophoric antibiotics in industrial extraction or production of sweet products. |
| US5521708A (en) * | 1992-11-25 | 1996-05-28 | Canon Information & Systems, Inc. | Correlated color temperature detector |
| ZA939767B (en) | 1993-01-21 | 1994-09-14 | Univ North Carolina State | Nematode-resistant transgenic plants |
| EP0670670A4 (en) | 1993-09-30 | 1996-04-24 | Agracetus | Transgenic cotton plants producing heterologous peroxidase. |
| US5598515A (en) * | 1994-01-10 | 1997-01-28 | Gen Tech Corp. | System and method for reconstructing surface elements of solid objects in a three-dimensional scene from a plurality of two dimensional images of the scene |
| US5608144A (en) | 1994-08-12 | 1997-03-04 | Dna Plant Technology Corp. | Plant group 2 promoters and uses thereof |
| US7262055B2 (en) * | 1998-08-25 | 2007-08-28 | Gendaq Limited | Regulated gene expression in plants |
| US6310194B1 (en) * | 1994-09-26 | 2001-10-30 | Carnegie Institution Of Washington | Plant fatty acid hydroxylases |
| US5961466A (en) | 1995-01-03 | 1999-10-05 | Omnicorder Technologies, Inc. | Method of detection of cancerous lesions by their effect on the spatial distribution of modulation of temperature and homogeneity of tissue |
| US5659026A (en) * | 1995-03-24 | 1997-08-19 | Pioneer Hi-Bred International | ALS3 promoter |
| CA2221747A1 (en) | 1995-06-07 | 1996-12-19 | Kevin Mcbride | Cotton fiber transcriptional factors |
| JPH0967270A (en) * | 1995-08-31 | 1997-03-11 | Res Dev Corp Of Japan | Prevention and therapy for opacity of crystalline lens and medicine therefor |
| US6084153A (en) * | 1996-02-14 | 2000-07-04 | The Governors Of The University Of Alberta | Plants having enhanced nitrogen assimilation/metabolism |
| JPH1094392A (en) | 1996-09-20 | 1998-04-14 | Nisshinbo Ind Inc | Cotton gene |
| EP0905242A4 (en) | 1996-10-24 | 2001-11-07 | Japan Tobacco Inc | Method for controlling water content of plant |
| IL119831A (en) | 1996-12-15 | 2002-12-01 | Cognitens Ltd | Apparatus and method for 3d surface geometry reconstruction |
| CA2278796A1 (en) * | 1997-01-21 | 1998-07-23 | Monsanto Company | Strawberry promoters and genes |
| TR200000547T2 (en) * | 1997-08-27 | 2001-05-21 | Pioneer Hi-Bred International, Inc. | Genes encoding enzymes for lignin biosynthesis and their use. |
| US6201541B1 (en) * | 1997-12-11 | 2001-03-13 | Cognitens, Ltd. | System and method for “Stitching” a plurality of reconstructions of three-dimensional surface features of object(s) in a scene defined relative to respective coordinate systems to relate them to a common coordinate system |
| US20090093620A1 (en) * | 2000-09-05 | 2009-04-09 | David Kovalic | Annotated Plant Genes |
| ATE528401T1 (en) * | 1998-08-04 | 2011-10-15 | Cropdesign Nv | GENES INVOLVED IN TOLERANCE TO ENVIRONMENTAL STRESS |
| US6313375B1 (en) | 1998-08-13 | 2001-11-06 | Pioneer Hi-Bred International, Inc. | Maize aquaporins and uses thereof |
| US6313376B1 (en) | 1998-08-14 | 2001-11-06 | Pioneer Hi-Bred International, Inc. | Maize aquaporins and uses thereof |
| US6717034B2 (en) | 2001-03-30 | 2004-04-06 | Mendel Biotechnology, Inc. | Method for modifying plant biomass |
| US20050086718A1 (en) | 1999-03-23 | 2005-04-21 | Mendel Biotechnology, Inc. | Plant transcriptional regulators of abiotic stress |
| US7511190B2 (en) * | 1999-11-17 | 2009-03-31 | Mendel Biotechnology, Inc. | Polynucleotides and polypeptides in plants |
| EP1033405A3 (en) | 1999-02-25 | 2001-08-01 | Ceres Incorporated | Sequence-determined DNA fragments and corresponding polypeptides encoded thereby |
| WO2000066610A1 (en) * | 1999-04-30 | 2000-11-09 | Agritope, Inc. | Apple promoters for expression of transgenes in plants |
| US20040031072A1 (en) * | 1999-05-06 | 2004-02-12 | La Rosa Thomas J. | Soy nucleic acid molecules and other molecules associated with transcription plants and uses thereof for plant improvement |
| US20100293669A2 (en) * | 1999-05-06 | 2010-11-18 | Jingdong Liu | Nucleic Acid Molecules and Other Molecules Associated with Plants and Uses Thereof for Plant Improvement |
| US20110214206A1 (en) | 1999-05-06 | 2011-09-01 | La Rosa Thomas J | Nucleic acid molecules and other molecules associated with plants |
| US20030233670A1 (en) | 2001-12-04 | 2003-12-18 | Edgerton Michael D. | Gene sequences and uses thereof in plants |
| US8877916B2 (en) * | 2000-04-26 | 2014-11-04 | Ceres, Inc. | Promoter, promoter control elements, and combinations, and uses thereof |
| US6559363B1 (en) * | 1999-07-05 | 2003-05-06 | Toyo Boseki Kabushiki Kaisha | Cotton plants with improved cotton fiber characteristics and method for producing cotton fibers from these cotton plants |
| WO2001006006A1 (en) | 1999-07-19 | 2001-01-25 | Japan Science And Technology Corporation | Environmental stress resistance gene |
| US6472588B1 (en) | 1999-09-10 | 2002-10-29 | Texas Tech University | Transgenic cotton plants with altered fiber characteristics transformed with a sucrose phosphate synthase nucleic acid |
| US6359196B1 (en) * | 1999-09-23 | 2002-03-19 | Finn Lok | Germination-specific plant promoters |
| US6403862B1 (en) * | 1999-09-24 | 2002-06-11 | Pioneer Hi-Bred International, Inc. | Seed-preferred promoter from maize |
| IT1313518B1 (en) * | 1999-10-22 | 2002-07-24 | Meta Instr S R L | METHODS AND EQUIPMENT FOR MEASURING THE THREE-DIMENSIONAL DISTRIBUTION OF TEMPERATURES INSIDE DIELECTRIC MEDIA. |
| US6407315B1 (en) | 1999-11-02 | 2002-06-18 | Pioneer Hi-Bred International, Inc. | Seed-preferred promoter from barley |
| US6828476B1 (en) | 1999-12-02 | 2004-12-07 | The Regents Of The University Of California | Cotton transcription factors and their uses |
| GB2358752A (en) | 2000-01-31 | 2001-08-01 | Tricorder Technology Plc | Surface or volumetric data processing method and apparatus |
| JP3807721B2 (en) * | 2000-02-21 | 2006-08-09 | シャープ株式会社 | Image synthesizer |
| US20110131679A2 (en) * | 2000-04-19 | 2011-06-02 | Thomas La Rosa | Rice Nucleic Acid Molecules and Other Molecules Associated with Plants and Uses Thereof for Plant Improvement |
| US7834146B2 (en) | 2000-05-08 | 2010-11-16 | Monsanto Technology Llc | Recombinant polypeptides associated with plants |
| US20040181830A1 (en) * | 2001-05-07 | 2004-09-16 | Kovalic David K. | Nucleic acid molecules and other molecules associated with plants and uses thereof for plant improvement |
| US6701081B1 (en) * | 2000-06-06 | 2004-03-02 | Air Controls, Inc. | Dual camera mount for stereo imaging |
| TW519485B (en) * | 2000-09-20 | 2003-02-01 | Ind Tech Res Inst | Infrared 3D scanning system |
| US20020170088A1 (en) | 2000-11-03 | 2002-11-14 | The Regents Of The University Of California | Novel auxin binding proteins and uses thereof |
| JP2002164066A (en) | 2000-11-22 | 2002-06-07 | Mitsubishi Heavy Ind Ltd | Stacked heat exchanger |
| CA2430642A1 (en) * | 2000-12-01 | 2003-02-20 | John B. Ohlrogge | Plant seed specific promoters |
| CN1326996C (en) | 2000-12-08 | 2007-07-18 | 联邦科学及工业研究组织 | Modification of sucrose synthase gene expression in plant tissue and uses therefor |
| US7214786B2 (en) * | 2000-12-14 | 2007-05-08 | Kovalic David K | Nucleic acid molecules and other molecules associated with plants and uses thereof for plant improvement |
| US6801257B2 (en) | 2001-01-12 | 2004-10-05 | Cognitens Ltd. | Optical three-dimensional digital imaging and mensuration system for industrial applications |
| KR100350216B1 (en) | 2001-02-02 | 2002-08-28 | (주)제노마인 | Osmotic stress-inducible kinase functioning as a negative regulator in osmotic stress signaling pathway in plants |
| ATE540575T1 (en) * | 2001-03-16 | 2012-01-15 | Basf Plant Science Gmbh | REGULATORS OF SUGAR AND LIPID METABOLISM IN PLANTS |
| JP4739569B2 (en) * | 2001-04-09 | 2011-08-03 | パナソニック株式会社 | Driving assistance device |
| WO2002082988A2 (en) | 2001-04-16 | 2002-10-24 | The Johns Hopkins University | Method for imaging and spectroscopy of tumors and determination of the efficacy of anti-tumor drug therapies |
| AU2002302595B2 (en) | 2001-05-03 | 2006-07-13 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Freeze-tolerant eukaryotic cells |
| WO2003020025A2 (en) | 2001-08-31 | 2003-03-13 | The Dow Chemical Company | Nucleic acid compositions conferring insect control in plants |
| US7038111B2 (en) * | 2001-09-06 | 2006-05-02 | The Arizona Board Of Regents | Method for increasing stress tolerance in plants |
| DE10150918C2 (en) | 2001-10-18 | 2003-10-09 | Inframedic Ag | Process for the evaluation of thermal images of a female or male breast |
| US20050108791A1 (en) * | 2001-12-04 | 2005-05-19 | Edgerton Michael D. | Transgenic plants with improved phenotypes |
| AU2003233489B2 (en) | 2002-04-08 | 2008-10-02 | Pioneer Hi-Bred International, Inc. | Enhanced silk exsertion under stress |
| CN1653174A (en) * | 2002-05-08 | 2005-08-10 | 巴斯福植物科学有限公司 | Methods for increasing oil content in plants |
| US20030221218A1 (en) | 2002-05-17 | 2003-11-27 | The Regents Of The University Of California | Bioengineering cotton fiber properties |
| JP2005185101A (en) | 2002-05-30 | 2005-07-14 | National Institute Of Agrobiological Sciences | VEGETABLE FULL-LENGTH cDNA AND UTILIZATION THEREOF |
| ATE495259T1 (en) * | 2002-07-10 | 2011-01-15 | Basf Plant Science Gmbh | USING A GENE TO INCREASE OIL CONTENT IN PLANTS |
| WO2004053055A2 (en) | 2002-12-04 | 2004-06-24 | Monsanto Technology Llc | Transgenic maize with enhanced phenotype |
| WO2004058963A2 (en) | 2002-12-31 | 2004-07-15 | University Of Delhi | A novel gene osisap1 of rice confers tolerance to stresses and a method thereof |
| BRPI0408735A (en) | 2003-03-12 | 2006-03-07 | Evogene Ltd | isolated polynucleotide, nucleic acid construction, transgenic cell, transgenic organism, transgenic plant, method for producing a transgenic plant, method for expressing a polynucleotide of interest in a cell, and method for co-expressing two polynucleotides of interest in a cell |
| AU2004225483B2 (en) | 2003-03-28 | 2009-07-23 | Monsanto Technology, Llc | Novel plant promoters for use in early seed development |
| AU2004230490C1 (en) * | 2003-04-15 | 2012-08-16 | Basf Plant Science Gmbh | Nucleic acid sequences encoding proteins associated with abiotic stress response and plant cells and plants with increased tolerance to environmental stress |
| WO2004092367A1 (en) | 2003-04-16 | 2004-10-28 | Basf Plant Science Gmbh | Use of genes for increasing the oil content in plants |
| US7554007B2 (en) | 2003-05-22 | 2009-06-30 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants |
| EP1636333A4 (en) | 2003-06-19 | 2007-10-24 | Evogene Ltd | Nucleotide sequences for regulating gene expression in plant trichomes and constructs and methods utilizing same |
| JP4452876B2 (en) | 2003-08-06 | 2010-04-21 | 国立大学法人 香川大学 | Control of seed yield and dry weight of plants by gene transfer using LKP2 partial cDNA |
| US7884261B2 (en) * | 2004-06-30 | 2011-02-08 | CERES,Inc. | Nucleotide sequences and corresponding polypeptides conferring modulated plant growth rate and biomass in plants |
| US7803983B2 (en) * | 2004-06-30 | 2010-09-28 | Ceres, Inc. | Nucleotide sequences and corresponding polypeptides conferring modulated plant growth rate and biomass in plants |
| US20050054931A1 (en) | 2003-09-09 | 2005-03-10 | Clark David W. | Tracking clutter filter for spectral & audio doppler |
| US7989676B2 (en) * | 2006-08-31 | 2011-08-02 | Ceres, Inc. | Nucleotide sequences and corresponding polypeptides conferring modulated plant characteristics |
| US20060048240A1 (en) * | 2004-04-01 | 2006-03-02 | Nickolai Alexandrov | Sequence-determined DNA fragments and corresponding polypeptides encoded thereby |
| US20060107345A1 (en) * | 2003-09-30 | 2006-05-18 | Nickolai Alexandrov | Sequence-determined DNA fragments and corresponding polypeptides encoded thereby |
| US7569389B2 (en) | 2004-09-30 | 2009-08-04 | Ceres, Inc. | Nucleotide sequences and polypeptides encoded thereby useful for modifying plant characteristics |
| US20060143729A1 (en) * | 2004-06-30 | 2006-06-29 | Ceres, Inc. | Nucleotide sequences and polypeptides encoded thereby useful for modifying plant characteristics |
| US20060150283A1 (en) * | 2004-02-13 | 2006-07-06 | Nickolai Alexandrov | Sequence-determined DNA fragments and corresponding polypeptides encoded thereby |
| EP2302062A1 (en) | 2003-10-20 | 2011-03-30 | CropDesign N.V. | Identification of E2F target genes and uses thereof |
| US20050096515A1 (en) * | 2003-10-23 | 2005-05-05 | Geng Z. J. | Three-dimensional surface image guided adaptive therapy system |
| US7807874B2 (en) | 2003-12-10 | 2010-10-05 | Monsanto Technology Llc | Stress tolerant plants and methods thereof |
| WO2005084331A2 (en) | 2004-03-01 | 2005-09-15 | Syngenta Participations Ag | Sorghum gene expression profiling |
| US8049069B2 (en) | 2004-03-31 | 2011-11-01 | Commonwealth Scientific And Industrial Research Organisation | Genes involved in plant fibre development |
| AU2005229157B2 (en) | 2004-03-31 | 2011-07-21 | Commonwealth Scientific And Industrial Research Organisation | Genes involved in plant fibre development |
| DE602005027191D1 (en) * | 2004-04-23 | 2011-05-12 | Ceres Inc | NUCLEOTIDE SEQUENCES AND POLYPEPTIDES ENCODED TO MODIFY THE PERFORMANCE CHARACTERISTICS OF NITROGEN USE IN PLANTS |
| CA2564202A1 (en) | 2004-05-05 | 2005-11-17 | The Royal Veterinary And Agricultural University | Ammonium/ammonia transporter |
| CN101948846A (en) * | 2004-06-14 | 2011-01-19 | 伊沃基因有限公司 | Polynucleotides and polypeptides involved in plant fiber development and methods of using same |
| JP2008505684A (en) | 2004-07-07 | 2008-02-28 | リアル イメージング リミテッド | 3D thermal breast cancer detection |
| MX2007004884A (en) | 2004-10-22 | 2007-06-22 | Agrinomics Llc | Generation of plants with altered oil content. |
| WO2008069878A2 (en) | 2006-10-27 | 2008-06-12 | Ceres, Inc. | Modulating lignin in plants |
| WO2006076099A2 (en) * | 2004-12-08 | 2006-07-20 | Ceres, Inc. | Nucleotide sequences and corresponding polypeptides conferring modulated plant size and biomass in plants |
| US20080148432A1 (en) * | 2005-12-21 | 2008-06-19 | Mark Scott Abad | Transgenic plants with enhanced agronomic traits |
| PT1827078E (en) * | 2004-12-21 | 2014-05-26 | Monsanto Technology Llc | Transgenic plants with enhanced agronomic traits |
| EP1681128A1 (en) * | 2005-01-14 | 2006-07-19 | Siemens Aktiengesellschaft | Method and device for producing a hole |
| EP1882392A4 (en) * | 2005-05-10 | 2009-07-01 | Monsanto Technology Llc | Genes and uses for plant improvement |
| US20060288451A1 (en) | 2005-05-26 | 2006-12-21 | Monsanto Technology, L.L.C. | Elevation of oil in monocot plants |
| WO2006138012A1 (en) | 2005-06-17 | 2006-12-28 | Ceres Inc. | P450 substrates and methods related thereto |
| US20080301839A1 (en) | 2005-08-30 | 2008-12-04 | Ravanello Monica P | Transgenic plants with enhanced agronomic traits |
| CA2644675A1 (en) * | 2006-01-13 | 2007-07-26 | Greg Nadzan | Nucleotide sequences and corresponding polypeptides conferring improved nitrogen use efficiency characteristics in plants |
| EP2010661A2 (en) | 2006-03-24 | 2009-01-07 | BASF Plant Science GmbH | Proteins associated with abiotic stress response and homologs |
| WO2007113237A2 (en) | 2006-03-31 | 2007-10-11 | Basf Plant Science Gmbh | Plants having enhanced yield-related traits and a method for making the same |
| MX349479B (en) * | 2006-12-20 | 2017-07-31 | Evogene Ltd | Polynucleotides and polypeptides involved in plant fiber development and methods of using same. |
| MX2009010858A (en) | 2007-04-09 | 2009-11-02 | Evogene Ltd | Polynucleotides, polypeptides and methods for increasing oil content, growth rate and biomass of plants. |
| EP2573178A3 (en) | 2007-07-10 | 2013-07-24 | Monsanto Technology LLC | Transgenic plants with enhanced agronomic traits |
| BR122020022203B1 (en) | 2007-07-24 | 2021-04-20 | Evogene Ltd | method of increasing the growth rate of a plant |
| US8362325B2 (en) * | 2007-10-03 | 2013-01-29 | Ceres, Inc. | Nucleotide sequences and corresponding polypeptides conferring modulated plant characteristics |
| CN101977928B (en) | 2007-12-27 | 2014-12-10 | 伊沃基因有限公司 | Isolated polypeptides, polynucleotides useful for modifying water user efficiency, fertilizer use efficiency, biotic/abiotic stress tolerance, yield and biomass in plants |
| CA3148194A1 (en) | 2008-05-22 | 2009-11-26 | Evogene Ltd. | Isolated polynucleotides and peptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| EP2297336A1 (en) | 2008-05-29 | 2011-03-23 | Vib Vzw | Minichromosome maintenance complex interacting protein involved in cancer |
| BRPI0912898B1 (en) | 2008-08-18 | 2022-04-12 | Evogene Ltd | Method for increasing nitrogen use efficiency and/or nitrogen deficiency tolerance of a plant |
| EP2347014B1 (en) | 2008-10-30 | 2016-09-21 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficieny |
| MX340023B (en) | 2008-12-29 | 2016-06-22 | Evogene Ltd | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance, biomass and/or yield in plants expressing same. |
| CA3123543A1 (en) | 2009-03-02 | 2010-09-10 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| EP2440033B1 (en) | 2009-06-10 | 2017-03-15 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US8937215B2 (en) | 2009-08-04 | 2015-01-20 | Evogene Ltd. | Polynucleotides and polypeptides for increasing desirable plant qualities |
| US20110080674A1 (en) * | 2009-10-02 | 2011-04-07 | Joel Durand | Magnetic azimuth adjustment for tonearm |
| EP2519097B1 (en) | 2009-12-28 | 2016-03-02 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| AU2011246876B2 (en) * | 2010-04-28 | 2016-06-23 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| BR112013004851A2 (en) * | 2010-08-30 | 2016-06-07 | Evogene Ltd | method of increasing nitrogen use efficiency, yield, biomass, growth rate, vigor, oil content, fiber yield and / or abiotic stress tolerance of a plant, isolated polynucleotide, nucleic acid structure, isolated polypeptide, cell vegetable and transgenic plant |
| BR122021002248B1 (en) | 2010-12-22 | 2022-02-15 | Evogene Ltd | METHOD TO INCREASE TOLERANCE TO ABIOTIC STRESS, PRODUCTION, BIOMASS, AND/OR GROWTH RATE OF A PLANT |
| WO2012150598A2 (en) | 2011-05-03 | 2012-11-08 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| WO2013027223A2 (en) | 2011-08-23 | 2013-02-28 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| CN104254242A (en) | 2011-11-21 | 2014-12-31 | 先正达参股股份有限公司 | Compositions and methods for increasing nematode resistance in plants |
| WO2013080203A1 (en) | 2011-11-28 | 2013-06-06 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| BR122020022832B1 (en) | 2011-12-28 | 2021-08-17 | Evogene Ltd | METHOD TO INCREASE THE PRODUCTION, GROWTH RATE, BIOMASS, ENERGY AND/OR SEED PRODUCTION OF A PLANT COMPARED TO A NATIVE PLANT, AND, ISOLATED NUCLEIC ACID CONSTRUCTION |
| WO2013128448A1 (en) | 2012-02-29 | 2013-09-06 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| CA2873846A1 (en) | 2012-05-28 | 2013-12-05 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| BR122019028124B1 (en) | 2012-08-27 | 2022-08-09 | Evogene Ltd | METHOD TO INCREASE YIELD, GROWTH RATE, BIOMASS, VIGOR, PHOTOSYNTHETIC CAPACITY, NITROGEN USE EFFICIENCY, AND/OR TOLERANCE TO ABIOTIC STRESS OF A PLANT, METHOD FOR PRODUCING A HARVEST, NUCLEIC ACID CONSTRUCTION, AND, METHOD OF GROWING A CULTURE |
| BR112015015415B1 (en) | 2012-12-25 | 2022-08-16 | Evogene Ltd. | METHODS TO INCREASE NITROGEN USE EFFICIENCY, GROWTH RATE, BIOMASS, SEED YIELD, PHOTOSYNTHETIC CAPACITY AND/OR TOLERANCE TO ABIOTIC STRESS OF A PLANT, TO PRODUCE A CULTURE, TO GROW A CROP, AND, TO SELECT A PLANT |
| BR122020018366B1 (en) | 2012-12-26 | 2022-03-29 | Evogene Ltd | Method for increasing nitrogen use efficiency, yield, growth rate, biomass, vigor, photosynthetic capacity and/or abiotic stress tolerance of a plant, and isolated nucleic acid construct |
| CA2910097A1 (en) | 2013-05-22 | 2014-11-27 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| CA2916060A1 (en) | 2013-08-27 | 2015-03-05 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| AU2015265412B2 (en) | 2014-05-28 | 2021-03-25 | Evogene Ltd. | Isolated polynucleotides, polypeptides and methods of using same for increasing abiotic stress tolerance, biomass and yield of plants |
| US10858403B2 (en) | 2014-08-27 | 2020-12-08 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10766935B2 (en) | 2015-12-28 | 2020-09-08 | Evogene Ltd. | Plant traits conferred by isolated polynucleotides and polypeptides |
-
2008
- 2008-07-24 BR BR122020022203-4A patent/BR122020022203B1/en active IP Right Grant
- 2008-07-24 CA CA2694481A patent/CA2694481C/en active Active
- 2008-07-24 US US12/669,975 patent/US8686227B2/en active Active
- 2008-07-24 BR BRPI0812742-5A patent/BRPI0812742B1/en not_active IP Right Cessation
- 2008-07-24 CA CA3133548A patent/CA3133548A1/en active Pending
- 2008-07-24 ES ES15151271.2T patent/ES2685631T3/en active Active
- 2008-07-24 WO PCT/IL2008/001024 patent/WO2009013750A2/en active Application Filing
- 2008-07-24 ES ES08776651.5T patent/ES2547305T3/en active Active
- 2008-07-24 AU AU2008278654A patent/AU2008278654B2/en not_active Ceased
- 2008-07-24 EP EP08776651.5A patent/EP2183371B1/en not_active Not-in-force
- 2008-07-24 CN CN2008801094649A patent/CN102037127A/en active Pending
- 2008-07-24 CA CA3019282A patent/CA3019282C/en active Active
- 2008-07-24 EP EP15151271.2A patent/EP2910638B1/en not_active Not-in-force
- 2008-07-24 BR BR122020022199-2A patent/BR122020022199B1/en active IP Right Grant
-
2010
- 2010-02-19 ZA ZA2010/01205A patent/ZA201001205B/en unknown
-
2013
- 2013-11-05 US US14/071,715 patent/US9518267B2/en active Active
-
2016
- 2016-09-28 US US15/278,086 patent/US10155957B2/en not_active Expired - Fee Related
-
2018
- 2018-10-09 US US16/154,833 patent/US10961544B2/en active Active
- 2018-12-13 US US16/218,559 patent/US10995341B2/en active Active
-
2021
- 2021-02-17 US US17/177,309 patent/US20210171974A1/en not_active Abandoned
Patent Citations (9)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5356816A (en) | 1991-11-19 | 1994-10-18 | Board Of Trustees Operating Michigan State University | Method and compositions using polypeptides of arabidopsis thaliana |
| US5296462A (en) | 1992-11-19 | 1994-03-22 | Board Of Trustees Operating Michigan State University | Method and compositions using polypeptides of arabidopsis thaliana |
| US6670528B1 (en) | 1998-10-14 | 2003-12-30 | Independent Administrative Institute, Japan International Research Center For Agricultural Sciences | Environmental stress-tolerant plants |
| US6720477B2 (en) | 2000-04-07 | 2004-04-13 | Basf Plant Science Gmbh | Signal transduction stress-related proteins and methods of use in plants |
| US20060183137A1 (en) | 2000-08-24 | 2006-08-17 | The Scripps Research Institute | Stress-regulated genes of plants, transgenic plants containing same, and methods of use |
| US20030056249A1 (en) | 2001-06-12 | 2003-03-20 | Simmons Carl R. | Anti-apoptosis genes and methods of use thereof |
| WO2004104162A2 (en) | 2003-05-22 | 2004-12-02 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| WO2007020638A2 (en) | 2005-08-15 | 2007-02-22 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| WO2007049275A2 (en) | 2005-10-24 | 2007-05-03 | Evogene Ltd. | Isolated polypeptides, polynucleotides encoding same, transgenic plants expressing same and methods of using same |
Non-Patent Citations (1)
| Title |
|---|
| See also references of EP2183371A4 |
Cited By (113)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US10184132B2 (en) | 2003-05-22 | 2019-01-22 | Evogene Ltd. | Methods of increasing abiotic stress tolerance, yield and/or biomass in plants |
| US9834781B2 (en) | 2004-06-14 | 2017-12-05 | Evogene Ltd. | Polynucleotides and polypeptides involved in plant fiber development and methods of using same |
| US9012728B2 (en) | 2004-06-14 | 2015-04-21 | Evogene Ltd. | Polynucleotides and polypeptides involved in plant fiber development and methods of using same |
| US10774339B2 (en) | 2004-06-14 | 2020-09-15 | Evogene Ltd. | Polynucleotides and polypeptides involved in plant fiber development and methods of using same |
| US10533184B2 (en) | 2004-06-14 | 2020-01-14 | Evogene Ltd. | Isolated polypeptides, polynucleotides encoding same, transgenic plants expressing same and methods of using same |
| US8962915B2 (en) | 2004-06-14 | 2015-02-24 | Evogene Ltd. | Isolated polypeptides, polynucleotides encoding same, transgenic plants expressing same and methods of using same |
| US7910800B2 (en) | 2005-08-15 | 2011-03-22 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| US9487796B2 (en) | 2005-08-15 | 2016-11-08 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| US10829777B2 (en) | 2005-08-15 | 2020-11-10 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| US10214749B2 (en) | 2005-08-15 | 2019-02-26 | Evogene Ltd. | Methods of increasing abiotic stress tolerance and/or biomass in plants and plants generated thereby |
| US10844393B2 (en) | 2006-12-20 | 2020-11-24 | Evogene Ltd. | Polynucleotides and polypeptides involved in plant fiber development and methods of using same |
| US9631000B2 (en) | 2006-12-20 | 2017-04-25 | Evogene Ltd. | Polynucleotides and polypeptides involved in plant fiber development and methods of using same |
| US10036031B2 (en) | 2007-04-09 | 2018-07-31 | Evogene Ltd. | Polynucleotides, polypeptides and methods for increasing oil content, growth rate and biomass of plants |
| US10155957B2 (en) | 2007-07-24 | 2018-12-18 | Evogene Ltd. | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US10961544B2 (en) | 2007-07-24 | 2021-03-30 | Evogene Ltd. | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US10995341B2 (en) | 2007-07-24 | 2021-05-04 | Evogene Ltd. | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same |
| US10407690B2 (en) | 2007-12-27 | 2019-09-10 | Evogene Ltd. | Isolated polypeptides, polynucleotides useful for modifying water user efficiency, fertilizer use efficiency, biotic/abiotic stress tolerance, yield and biomass in plants |
| US9670501B2 (en) | 2007-12-27 | 2017-06-06 | Evogene Ltd. | Isolated polypeptides, polynucleotides useful for modifying water user efficiency, fertilizer use efficiency, biotic/abiotic stress tolerance, yield and biomass in plants |
| US10900048B2 (en) | 2008-05-22 | 2021-01-26 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant utility |
| US8847008B2 (en) | 2008-05-22 | 2014-09-30 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant utility |
| US10100326B2 (en) | 2008-05-22 | 2018-10-16 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant utility |
| US9714430B2 (en) | 2008-05-22 | 2017-07-25 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant utility |
| US9783818B2 (en) | 2008-08-18 | 2017-10-10 | Evogene Ltd. | Use of ADP/ATP transporter genes to increase nitrogen use efficiency and low nitrogen tolerance to a plant |
| US10208316B2 (en) | 2008-08-18 | 2019-02-19 | Evogene Ltd. | Use of UMP-CMP kinases for increasing nitrogen use efficiency and low nitrogen tolerance in plants |
| EP3072972A2 (en) | 2008-08-18 | 2016-09-28 | Evogene Ltd. | Isolated polypeptides and polynucleotides useful for increasing nitrogen use efficiency, abiotic stress tolerance, yield and biomass in plants |
| US11453887B2 (en) | 2008-08-18 | 2022-09-27 | Evogene Ltd. | Isolated polypeptides and polynucleotides useful for increasing nitrogen use efficiency, abiotic stress tolerance, yield and biomass in plants |
| EP3616504A2 (en) | 2008-08-18 | 2020-03-04 | Evogene Ltd. | Isolated polypeptides and polynucleotides useful for increasing nitrogen use efficiency, abiotic stress tolerance, yield and biomass in plants |
| US9018445B2 (en) | 2008-08-18 | 2015-04-28 | Evogene Ltd. | Use of CAD genes to increase nitrogen use efficiency and low nitrogen tolerance to a plant |
| US10829776B2 (en) | 2008-08-18 | 2020-11-10 | Evogene Ltd. | Use of MFS transporters for increasing biomass, growth rate, nitrogen use efficiency and low nitrogen tolerance in plants |
| US10793870B2 (en) | 2008-10-30 | 2020-10-06 | Evogene Ltd. | Methods of increasing biomass and/or growth rate of a plant under non-stress conditions |
| US9745595B2 (en) | 2008-10-30 | 2017-08-29 | Evogene Ltd. | Methods of increasing biomass and/or growth rate of a plant under non-stress conditions |
| US10975382B2 (en) | 2008-12-29 | 2021-04-13 | Evogene Ltd. | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance, biomass and/or yield in plants expressing same |
| US8952218B2 (en) | 2008-12-29 | 2015-02-10 | Evogene Ltd. | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance, biomass and/or yield in plants expressing same |
| US9260490B2 (en) | 2009-02-25 | 2016-02-16 | Basf Plant Science Company Gmbh | Plants having enhanced yield-related traits and a method for making the same |
| WO2010097343A1 (en) * | 2009-02-25 | 2010-09-02 | Basf Plant Science Company Gmbh | Plants having enhanced yield-related traits and a method for making the same |
| EP3000889A2 (en) | 2009-03-02 | 2016-03-30 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10597671B2 (en) | 2009-03-02 | 2020-03-24 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield, biomass, oil content and/or growth rate |
| WO2010100595A2 (en) | 2009-03-02 | 2010-09-10 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| EP3460062A3 (en) * | 2009-03-02 | 2019-05-08 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US8937220B2 (en) | 2009-03-02 | 2015-01-20 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield, biomass, vigor and/or growth rate of a plant |
| US9487795B2 (en) | 2009-03-02 | 2016-11-08 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield, biomass, oil content and/or growth rate |
| EP3626051A2 (en) | 2009-06-10 | 2020-03-25 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US11286495B2 (en) | 2009-06-10 | 2022-03-29 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US10006040B2 (en) | 2009-06-10 | 2018-06-26 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| EP3238534B1 (en) * | 2009-06-10 | 2019-11-13 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| EP2440033B1 (en) * | 2009-06-10 | 2017-03-15 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US10791690B2 (en) | 2009-06-10 | 2020-10-06 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| EP3238534A2 (en) | 2009-06-10 | 2017-11-01 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US10227607B2 (en) | 2009-08-04 | 2019-03-12 | Evogene Ltd. | MADS-box polynucleotides and polypeptides for increasing abiotic stress tolerance, yield, biomass, growth rate, and/or vigor in a plant |
| US9574200B2 (en) | 2009-08-04 | 2017-02-21 | Evogene Ltd. | Polynucleotides and polypeptides for increasing desirable plant qualities |
| US8937215B2 (en) | 2009-08-04 | 2015-01-20 | Evogene Ltd. | Polynucleotides and polypeptides for increasing desirable plant qualities |
| US11530418B2 (en) | 2009-08-04 | 2022-12-20 | Evogene Ltd. | Polynucleotides and polypeptides for increasing desirable plant qualities |
| US10883115B2 (en) | 2009-08-04 | 2021-01-05 | Evogene Ltd. | Protein kinase polynucleotides and polypeptides for increasing abiotic stress tolerance, yield, biomass, growth rate, and/or vigor in a plant |
| US10982224B2 (en) | 2009-12-28 | 2021-04-20 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| EP3056569A3 (en) * | 2009-12-28 | 2016-10-19 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US10351873B2 (en) | 2009-12-28 | 2019-07-16 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| EP2519097A4 (en) * | 2009-12-28 | 2013-11-13 | Evogene Ltd | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| EP2519097A2 (en) * | 2009-12-28 | 2012-11-07 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| WO2011135527A2 (en) | 2010-04-28 | 2011-11-03 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US11542522B2 (en) | 2010-04-28 | 2023-01-03 | Evogene Ltd. | Isolated polynucleotides and polypeptides for increasing plant yield and/or agricultural characteristics |
| US10689662B2 (en) | 2010-04-28 | 2020-06-23 | Evogene Ltd. | Isolated polynucleotides and polypeptides for increasing plant yield and/or agricultural characteristics |
| WO2012007945A3 (en) * | 2010-07-12 | 2012-06-14 | The State Of Israel, Ministry Of Agriculture & Rural Development, Agricultural Research Organization, (A.R.O.), Volcani Center | Isolated polynucleotides and methods and plants using same for regulating plant acidity |
| US9512188B2 (en) | 2010-07-12 | 2016-12-06 | The State Of Israel, Ministry Of Agriculture & Rural Development, Agricultural Research Organization (Aro) (Volcani Center) | Isolated polynucleotides and methods and plants using same for regulating plant acidity |
| US20130133106A1 (en) * | 2010-07-12 | 2013-05-23 | The State of Israel, Ministry of Agriculure & Rural Dvlp, Agricultural research Org.(A.R.O) | Isolated polynucleotides and methods and plants using same for regulating plant acidity |
| US11130957B2 (en) | 2010-08-30 | 2021-09-28 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US10457954B2 (en) | 2010-08-30 | 2019-10-29 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US10457952B2 (en) | 2010-12-22 | 2019-10-29 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for improving plant properties |
| US11111499B2 (en) | 2011-05-03 | 2021-09-07 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US10760088B2 (en) | 2011-05-03 | 2020-09-01 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US11293032B2 (en) | 2011-08-23 | 2022-04-05 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10501750B2 (en) | 2011-08-23 | 2019-12-10 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US9976157B2 (en) | 2011-08-23 | 2018-05-22 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10900052B2 (en) | 2011-11-21 | 2021-01-26 | Syngenta Participations Ag | Compositions and methods for increasing nematode resistance in plants |
| US10000768B2 (en) | 2011-11-21 | 2018-06-19 | Syngenta Participations Ag | Compositions and methods for increasing nematode resistance in plants |
| US11078492B2 (en) | 2011-11-28 | 2021-08-03 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US10113176B2 (en) | 2011-11-28 | 2018-10-30 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency, yield, growth rate, vigor, biomass, oil content, and/or abiotic stress tolerance |
| US11242538B2 (en) | 2011-12-28 | 2022-02-08 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing yield of plants |
| US10260073B2 (en) | 2011-12-28 | 2019-04-16 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing yield of plants |
| US9920330B2 (en) | 2012-02-29 | 2018-03-20 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US11326179B2 (en) | 2012-02-29 | 2022-05-10 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US11365421B2 (en) | 2012-02-29 | 2022-06-21 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US10253327B2 (en) | 2012-02-29 | 2019-04-09 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US10815492B2 (en) | 2012-02-29 | 2020-10-27 | Evogene Ltd. | Isolated polynucleotides and polypeptides and methods of using same for increasing plant yield, biomass, growth rate, vigor, oil content, abiotic stress tolerance of plants and nitrogen use efficiency |
| US9834782B2 (en) | 2012-05-28 | 2017-12-05 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10982222B2 (en) | 2012-05-28 | 2021-04-20 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US11512323B2 (en) | 2012-08-27 | 2022-11-29 | Evogene Ltd. | Isolated polynucleotides, polypeptides and methods of using same for increasing abiotic stress tolerance, biomass and yield of plants |
| US11485982B1 (en) | 2012-08-27 | 2022-11-01 | Evogene Ltd. | Isolated polynucleotides, polypeptides and methods of using same for increasing abiotic stress tolerance, biomass and yield of plants |
| US10858665B2 (en) | 2012-08-27 | 2020-12-08 | Evogene Ltd. | Isolated polynucleotides, polypeptides and methods of using same for increasing abiotic stress tolerance, biomass and yield of plants |
| US9890389B2 (en) | 2012-12-25 | 2018-02-13 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency of plants |
| US10597672B2 (en) | 2012-12-25 | 2020-03-24 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency of plants |
| US11352636B2 (en) | 2012-12-25 | 2022-06-07 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing nitrogen use efficiency of plants |
| US10501751B2 (en) | 2012-12-26 | 2019-12-10 | Evogene Ltd. | Isolated polynucleotides and polypeptides, construct and plants comprising same and methods of using same for increasing nitrogen use efficiency of plants |
| US9771598B2 (en) | 2012-12-26 | 2017-09-26 | Evogene Ltd. | Isolated polynucleotides and polypeptides, construct and plants comprising same and methods of using same for increasing nitrogen use efficiency of plants |
| US11453888B2 (en) | 2012-12-26 | 2022-09-27 | Evogene Ltd. | Isolated polynucleotides and polypeptides, construct and plants comprising same and methods of using same for increasing nitrogen use efficiency of plants |
| US11549122B2 (en) | 2013-05-22 | 2023-01-10 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10214748B2 (en) | 2013-05-22 | 2019-02-26 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US9920329B2 (en) | 2013-05-22 | 2018-03-20 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US11525141B2 (en) | 2013-05-22 | 2022-12-13 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US11560573B2 (en) | 2013-05-22 | 2023-01-24 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10889827B2 (en) | 2013-05-22 | 2021-01-12 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| EP2840141A1 (en) * | 2013-08-21 | 2015-02-25 | Industry-Academic Cooperation Foundation, Yonsei University | Gene implicated in abiotic stress tolerance and growth accelerating and use thereof |
| EP3091077A3 (en) * | 2013-08-21 | 2017-03-01 | Industry-Academic Cooperation Foundation, Yonsei University | Gene implicated in abiotic stress tolerance and growth accelerating and use thereof |
| US10774340B2 (en) | 2013-08-27 | 2020-09-15 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10006042B2 (en) | 2013-08-27 | 2018-06-26 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US11499161B2 (en) | 2013-08-27 | 2022-11-15 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10337023B2 (en) | 2013-08-27 | 2019-07-02 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10975383B2 (en) | 2014-05-28 | 2021-04-13 | Evogene Ltd. | Isolated polynucleotides, polypeptides and methods of using same for increasing abiotic stress tolerance, biomass and yield of plants |
| US11485761B2 (en) | 2014-08-27 | 2022-11-01 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US11472853B1 (en) | 2014-08-27 | 2022-10-18 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US11421004B2 (en) | 2014-08-27 | 2022-08-23 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10858403B2 (en) | 2014-08-27 | 2020-12-08 | Evogene Ltd. | Isolated polynucleotides and polypeptides, and methods of using same for increasing plant yield and/or agricultural characteristics |
| US10766935B2 (en) | 2015-12-28 | 2020-09-08 | Evogene Ltd. | Plant traits conferred by isolated polynucleotides and polypeptides |
| US11566053B2 (en) | 2015-12-28 | 2023-01-31 | Evogene Ltd. | Plant traits conferred by isolated polynucleotides and polypeptides |
Also Published As
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US10995341B2 (en) | Polynucleotides, polypeptides encoded thereby, and methods of using same for increasing abiotic stress tolerance and/or biomass and/or yield in plants expressing same | |
| US11453887B2 (en) | Isolated polypeptides and polynucleotides useful for increasing nitrogen use efficiency, abiotic stress tolerance, yield and biomass in plants | |
| US10407690B2 (en) | Isolated polypeptides, polynucleotides useful for modifying water user efficiency, fertilizer use efficiency, biotic/abiotic stress tolerance, yield and biomass in plants | |
| AU2017228711B2 (en) | Polynucleotides, Polypeptides Encoded Thereby, and Methods of Using Same for Increasing Abiotic Stress Tolerance and/or Biomass and/or Yield in Plants Expressing Same | |
| AU2014215945B2 (en) | Polynucleotides, Polypeptides Encoded Thereby, and Methods of Using Same for Increasing Abiotic Stress Tolerance and/or Biomass and/or Yield in Plants Expressing Same |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| WWE | Wipo information: entry into national phase |
Ref document number: 200880109464.9 Country of ref document: CN |
|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 08776651 Country of ref document: EP Kind code of ref document: A2 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2694481 Country of ref document: CA Ref document number: MX/A/2010/000975 Country of ref document: MX |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 12010500186 Country of ref document: PH |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2008278654 Country of ref document: AU |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 1301/DELNP/2010 Country of ref document: IN Ref document number: 2008776651 Country of ref document: EP |
|
| ENP | Entry into the national phase |
Ref document number: 2008278654 Country of ref document: AU Date of ref document: 20080724 Kind code of ref document: A |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 12669975 Country of ref document: US |
|
| ENP | Entry into the national phase |
Ref document number: PI0812742 Country of ref document: BR Kind code of ref document: A2 Effective date: 20100126 |