WO2015123783A1 - Cathepsin b-targeting probes - Google Patents
Cathepsin b-targeting probes Download PDFInfo
- Publication number
- WO2015123783A1 WO2015123783A1 PCT/CA2015/050136 CA2015050136W WO2015123783A1 WO 2015123783 A1 WO2015123783 A1 WO 2015123783A1 CA 2015050136 W CA2015050136 W CA 2015050136W WO 2015123783 A1 WO2015123783 A1 WO 2015123783A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- compound
- group
- lys
- pet
- amc
- Prior art date
Links
- 239000000523 sample Substances 0.000 title claims abstract description 72
- 102000005600 Cathepsins Human genes 0.000 title description 6
- 108010084457 Cathepsins Proteins 0.000 title description 6
- 150000001875 compounds Chemical class 0.000 claims abstract description 72
- 238000000799 fluorescence microscopy Methods 0.000 claims abstract description 11
- 238000003384 imaging method Methods 0.000 claims abstract description 8
- 238000000034 method Methods 0.000 claims description 15
- 239000000203 mixture Substances 0.000 claims description 13
- 229910052731 fluorine Inorganic materials 0.000 claims description 12
- 125000001424 substituent group Chemical group 0.000 claims description 11
- 229910052739 hydrogen Inorganic materials 0.000 claims description 10
- 229910052799 carbon Inorganic materials 0.000 claims description 9
- 239000007850 fluorescent dye Substances 0.000 claims description 9
- 229910052786 argon Inorganic materials 0.000 claims description 8
- 238000012879 PET imaging Methods 0.000 claims description 7
- JGTNAGYHADQMCM-UHFFFAOYSA-M 1,1,2,2,3,3,4,4,4-nonafluorobutane-1-sulfonate Chemical compound [O-]S(=O)(=O)C(F)(F)C(F)(F)C(F)(F)C(F)(F)F JGTNAGYHADQMCM-UHFFFAOYSA-M 0.000 claims description 6
- LBLYYCQCTBFVLH-UHFFFAOYSA-M 2-methylbenzenesulfonate Chemical compound CC1=CC=CC=C1S([O-])(=O)=O LBLYYCQCTBFVLH-UHFFFAOYSA-M 0.000 claims description 6
- AFVFQIVMOAPDHO-UHFFFAOYSA-N Methanesulfonic acid Chemical compound CS(O)(=O)=O AFVFQIVMOAPDHO-UHFFFAOYSA-N 0.000 claims description 6
- SRSXLGNVWSONIS-UHFFFAOYSA-M benzenesulfonate Chemical compound [O-]S(=O)(=O)C1=CC=CC=C1 SRSXLGNVWSONIS-UHFFFAOYSA-M 0.000 claims description 6
- 229940077388 benzenesulfonate Drugs 0.000 claims description 6
- CCIVGXIOQKPBKL-UHFFFAOYSA-M ethanesulfonate Chemical compound CCS([O-])(=O)=O CCIVGXIOQKPBKL-UHFFFAOYSA-M 0.000 claims description 6
- 125000004435 hydrogen atom Chemical group [H]* 0.000 claims description 6
- HLANOUYSOJLOPM-UHFFFAOYSA-N methoxy benzenesulfonate Chemical compound COOS(=O)(=O)C1=CC=CC=C1 HLANOUYSOJLOPM-UHFFFAOYSA-N 0.000 claims description 6
- 125000002496 methyl group Chemical group [H]C([H])([H])* 0.000 claims description 6
- UJJUJHTVDYXQON-UHFFFAOYSA-N nitro benzenesulfonate Chemical compound [O-][N+](=O)OS(=O)(=O)C1=CC=CC=C1 UJJUJHTVDYXQON-UHFFFAOYSA-N 0.000 claims description 6
- ITMCEJHCFYSIIV-UHFFFAOYSA-N triflic acid Chemical compound OS(=O)(=O)C(F)(F)F ITMCEJHCFYSIIV-UHFFFAOYSA-N 0.000 claims description 6
- 125000003118 aryl group Chemical group 0.000 claims description 5
- 125000003837 (C1-C20) alkyl group Chemical group 0.000 claims description 4
- OKTJSMMVPCPJKN-UHFFFAOYSA-N Carbon Chemical group [C] OKTJSMMVPCPJKN-UHFFFAOYSA-N 0.000 claims description 4
- 125000004429 atom Chemical group 0.000 claims description 4
- 125000001797 benzyl group Chemical group [H]C1=C([H])C([H])=C(C([H])=C1[H])C([H])([H])* 0.000 claims description 4
- 125000004432 carbon atom Chemical group C* 0.000 claims description 4
- 230000005670 electromagnetic radiation Effects 0.000 claims description 4
- 230000004807 localization Effects 0.000 claims description 3
- 239000008194 pharmaceutical composition Substances 0.000 claims description 3
- ZOXJGFHDIHLPTG-UHFFFAOYSA-N Boron Chemical group [B] ZOXJGFHDIHLPTG-UHFFFAOYSA-N 0.000 claims description 2
- 125000003545 alkoxy group Chemical group 0.000 claims description 2
- 229910052796 boron Inorganic materials 0.000 claims description 2
- 150000004677 hydrates Chemical class 0.000 claims description 2
- 238000002347 injection Methods 0.000 claims description 2
- 239000007924 injection Substances 0.000 claims description 2
- 239000007788 liquid Substances 0.000 claims description 2
- 125000004433 nitrogen atom Chemical group N* 0.000 claims description 2
- 150000003839 salts Chemical class 0.000 claims description 2
- 239000012453 solvate Substances 0.000 claims description 2
- 238000012636 positron electron tomography Methods 0.000 claims 2
- 239000003937 drug carrier Substances 0.000 claims 1
- 239000000758 substrate Substances 0.000 abstract description 29
- 102000004225 Cathepsin B Human genes 0.000 abstract description 27
- 108090000712 Cathepsin B Proteins 0.000 abstract description 27
- 238000002600 positron emission tomography Methods 0.000 abstract description 9
- 210000004027 cell Anatomy 0.000 description 55
- RTZKZFJDLAIYFH-UHFFFAOYSA-N Diethyl ether Chemical compound CCOCC RTZKZFJDLAIYFH-UHFFFAOYSA-N 0.000 description 50
- XEKOWRVHYACXOJ-UHFFFAOYSA-N Ethyl acetate Chemical compound CCOC(C)=O XEKOWRVHYACXOJ-UHFFFAOYSA-N 0.000 description 39
- IAZDPXIOMUYVGZ-WFGJKAKNSA-N Dimethyl sulfoxide Chemical compound [2H]C([2H])([2H])S(=O)C([2H])([2H])[2H] IAZDPXIOMUYVGZ-WFGJKAKNSA-N 0.000 description 38
- 239000007787 solid Substances 0.000 description 34
- 238000006243 chemical reaction Methods 0.000 description 25
- 239000000243 solution Substances 0.000 description 25
- 206010028980 Neoplasm Diseases 0.000 description 20
- AXKGIPZJYUNAIW-UHFFFAOYSA-N (4-aminophenyl)methanol Chemical compound NC1=CC=C(CO)C=C1 AXKGIPZJYUNAIW-UHFFFAOYSA-N 0.000 description 19
- 102000004190 Enzymes Human genes 0.000 description 17
- 108090000790 Enzymes Proteins 0.000 description 17
- 102000035195 Peptidases Human genes 0.000 description 17
- 108091005804 Peptidases Proteins 0.000 description 17
- 239000004365 Protease Substances 0.000 description 17
- 238000002474 experimental method Methods 0.000 description 17
- GLNDAGDHSLMOKX-UHFFFAOYSA-N coumarin 120 Chemical compound C1=C(N)C=CC2=C1OC(=O)C=C2C GLNDAGDHSLMOKX-UHFFFAOYSA-N 0.000 description 16
- IAZDPXIOMUYVGZ-UHFFFAOYSA-N Dimethylsulphoxide Chemical compound CS(C)=O IAZDPXIOMUYVGZ-UHFFFAOYSA-N 0.000 description 15
- 230000000694 effects Effects 0.000 description 14
- DLELKZFCVLJXKZ-GOTSBHOMSA-N Z-Arg-Arg-NHMec Chemical compound N([C@@H](CCCN=C(N)N)C(=O)N[C@@H](CCCN=C(N)N)C(=O)NC1=CC=2OC(=O)C=C(C=2C=C1)C)C(=O)OCC1=CC=CC=C1 DLELKZFCVLJXKZ-GOTSBHOMSA-N 0.000 description 13
- 230000015572 biosynthetic process Effects 0.000 description 13
- 238000003786 synthesis reaction Methods 0.000 description 13
- AZQWKYJCGOJGHM-UHFFFAOYSA-N 1,4-benzoquinone Chemical compound O=C1C=CC(=O)C=C1 AZQWKYJCGOJGHM-UHFFFAOYSA-N 0.000 description 12
- -1 2-cresyl Chemical group 0.000 description 12
- CSCPPACGZOOCGX-UHFFFAOYSA-N Acetone Chemical compound CC(C)=O CSCPPACGZOOCGX-UHFFFAOYSA-N 0.000 description 12
- XKRFYHLGVUSROY-UHFFFAOYSA-N Argon Chemical compound [Ar] XKRFYHLGVUSROY-UHFFFAOYSA-N 0.000 description 12
- 108090000765 processed proteins & peptides Proteins 0.000 description 12
- 239000011541 reaction mixture Substances 0.000 description 12
- DTQVDTLACAAQTR-UHFFFAOYSA-N trifluoroacetic acid Substances OC(=O)C(F)(F)F DTQVDTLACAAQTR-UHFFFAOYSA-N 0.000 description 12
- 238000001644 13C nuclear magnetic resonance spectroscopy Methods 0.000 description 11
- 239000002904 solvent Substances 0.000 description 11
- 238000005160 1H NMR spectroscopy Methods 0.000 description 10
- ZMANZCXQSJIPKH-UHFFFAOYSA-N Triethylamine Chemical compound CCN(CC)CC ZMANZCXQSJIPKH-UHFFFAOYSA-N 0.000 description 10
- 239000012131 assay buffer Substances 0.000 description 10
- 201000011510 cancer Diseases 0.000 description 10
- 239000013592 cell lysate Substances 0.000 description 10
- 238000004128 high performance liquid chromatography Methods 0.000 description 10
- XLYOFNOQVPJJNP-UHFFFAOYSA-N water Substances O XLYOFNOQVPJJNP-UHFFFAOYSA-N 0.000 description 10
- 108091003079 Bovine Serum Albumin Proteins 0.000 description 9
- 108010081642 CA 074 methyl ester Proteins 0.000 description 9
- OKKJLVBELUTLKV-UHFFFAOYSA-N Methanol Chemical compound OC OKKJLVBELUTLKV-UHFFFAOYSA-N 0.000 description 9
- 239000003795 chemical substances by application Substances 0.000 description 9
- XGWSRLSPWIEMLQ-YTFOTSKYSA-N methyl n-({(2s,3s)-3-[(propylamino)carbonyl]oxiran-2-yl}carbonyl)-l-isoleucyl-l-prolinate Chemical compound CCCNC(=O)[C@H]1O[C@@H]1C(=O)N[C@@H]([C@@H](C)CC)C(=O)N1[C@H](C(=O)OC)CCC1 XGWSRLSPWIEMLQ-YTFOTSKYSA-N 0.000 description 9
- FWBHETKCLVMNFS-UHFFFAOYSA-N 4',6-Diamino-2-phenylindol Chemical compound C1=CC(C(=N)N)=CC=C1C1=CC2=CC=C(C(N)=N)C=C2N1 FWBHETKCLVMNFS-UHFFFAOYSA-N 0.000 description 8
- CSJLBAMHHLJAAS-UHFFFAOYSA-N diethylaminosulfur trifluoride Chemical compound CCN(CC)S(F)(F)F CSJLBAMHHLJAAS-UHFFFAOYSA-N 0.000 description 8
- 239000000706 filtrate Substances 0.000 description 8
- 102000004196 processed proteins & peptides Human genes 0.000 description 8
- HEMHJVSKTPXQMS-UHFFFAOYSA-M sodium hydroxide Inorganic materials [OH-].[Na+] HEMHJVSKTPXQMS-UHFFFAOYSA-M 0.000 description 8
- YMWUJEATGCHHMB-UHFFFAOYSA-N Dichloromethane Chemical compound ClCCl YMWUJEATGCHHMB-UHFFFAOYSA-N 0.000 description 7
- KCXVZYZYPLLWCC-UHFFFAOYSA-N EDTA Chemical compound OC(=O)CN(CC(O)=O)CCN(CC(O)=O)CC(O)=O KCXVZYZYPLLWCC-UHFFFAOYSA-N 0.000 description 7
- 230000004913 activation Effects 0.000 description 7
- 229940098773 bovine serum albumin Drugs 0.000 description 7
- 239000012267 brine Substances 0.000 description 7
- 238000003776 cleavage reaction Methods 0.000 description 7
- 229940127573 compound 38 Drugs 0.000 description 7
- 238000010511 deprotection reaction Methods 0.000 description 7
- 238000001914 filtration Methods 0.000 description 7
- 238000006460 hydrolysis reaction Methods 0.000 description 7
- 239000006166 lysate Substances 0.000 description 7
- LGRLWUINFJPLSH-UHFFFAOYSA-N methanide Chemical compound [CH3-] LGRLWUINFJPLSH-UHFFFAOYSA-N 0.000 description 7
- PIDFDZJZLOTZTM-KHVQSSSXSA-N ombitasvir Chemical compound COC(=O)N[C@@H](C(C)C)C(=O)N1CCC[C@H]1C(=O)NC1=CC=C([C@H]2N([C@@H](CC2)C=2C=CC(NC(=O)[C@H]3N(CCC3)C(=O)[C@@H](NC(=O)OC)C(C)C)=CC=2)C=2C=CC(=CC=2)C(C)(C)C)C=C1 PIDFDZJZLOTZTM-KHVQSSSXSA-N 0.000 description 7
- 230000007017 scission Effects 0.000 description 7
- 239000011734 sodium Substances 0.000 description 7
- HPALAKNZSZLMCH-UHFFFAOYSA-M sodium;chloride;hydrate Chemical compound O.[Na+].[Cl-] HPALAKNZSZLMCH-UHFFFAOYSA-M 0.000 description 7
- NPRYCHLHHVWLQZ-TURQNECASA-N 2-amino-9-[(2R,3S,4S,5R)-4-fluoro-3-hydroxy-5-(hydroxymethyl)oxolan-2-yl]-7-prop-2-ynylpurin-8-one Chemical compound NC1=NC=C2N(C(N(C2=N1)[C@@H]1O[C@@H]([C@H]([C@H]1O)F)CO)=O)CC#C NPRYCHLHHVWLQZ-TURQNECASA-N 0.000 description 6
- YCEHPUBNERNXNP-UHFFFAOYSA-N 7-isocyanato-4-methylchromen-2-one Chemical compound C1=C(N=C=O)C=CC2=C1OC(=O)C=C2C YCEHPUBNERNXNP-UHFFFAOYSA-N 0.000 description 6
- 239000006144 Dulbecco’s modified Eagle's medium Substances 0.000 description 6
- YXFVVABEGXRONW-UHFFFAOYSA-N Toluene Chemical compound CC1=CC=CC=C1 YXFVVABEGXRONW-UHFFFAOYSA-N 0.000 description 6
- 238000005859 coupling reaction Methods 0.000 description 6
- KJOZJSGOIJQCGA-UHFFFAOYSA-N dichloromethane;2,2,2-trifluoroacetic acid Chemical compound ClCCl.OC(=O)C(F)(F)F KJOZJSGOIJQCGA-UHFFFAOYSA-N 0.000 description 6
- 239000000975 dye Substances 0.000 description 6
- 230000007062 hydrolysis Effects 0.000 description 6
- GUAQVFRUPZBRJQ-UHFFFAOYSA-N n-(3-aminopropyl)-2-methylprop-2-enamide Chemical compound CC(=C)C(=O)NCCCN GUAQVFRUPZBRJQ-UHFFFAOYSA-N 0.000 description 6
- 238000010898 silica gel chromatography Methods 0.000 description 6
- 230000007306 turnover Effects 0.000 description 6
- ISMDILRWKSYCOD-GNKBHMEESA-N C(C1=CC=CC=C1)[C@@H]1NC(OCCCCCCCCCCCNC([C@@H](NC(C[C@@H]1O)=O)C(C)C)=O)=O Chemical compound C(C1=CC=CC=C1)[C@@H]1NC(OCCCCCCCCCCCNC([C@@H](NC(C[C@@H]1O)=O)C(C)C)=O)=O ISMDILRWKSYCOD-GNKBHMEESA-N 0.000 description 5
- 229940126639 Compound 33 Drugs 0.000 description 5
- YGYAWVDWMABLBF-UHFFFAOYSA-N Phosgene Chemical compound ClC(Cl)=O YGYAWVDWMABLBF-UHFFFAOYSA-N 0.000 description 5
- 235000001014 amino acid Nutrition 0.000 description 5
- 150000001413 amino acids Chemical class 0.000 description 5
- 238000003556 assay Methods 0.000 description 5
- 108010085937 benzyloxycarbonyl-phenylalanylarginine-4-methylcoumaryl-7-amide Proteins 0.000 description 5
- 239000003153 chemical reaction reagent Substances 0.000 description 5
- 239000012043 crude product Substances 0.000 description 5
- 239000003480 eluent Substances 0.000 description 5
- 239000012216 imaging agent Substances 0.000 description 5
- 230000014759 maintenance of location Effects 0.000 description 5
- 239000012074 organic phase Substances 0.000 description 5
- 125000002924 primary amino group Chemical group [H]N([H])* 0.000 description 5
- 238000010992 reflux Methods 0.000 description 5
- 229910000030 sodium bicarbonate Inorganic materials 0.000 description 5
- 210000001519 tissue Anatomy 0.000 description 5
- 0 *C[C@@](C(N[C@@](CN)C(Nc1ccc(*)cc1*)=O)=O)NC(*)=O Chemical compound *C[C@@](C(N[C@@](CN)C(Nc1ccc(*)cc1*)=O)=O)NC(*)=O 0.000 description 4
- HSHNITRMYYLLCV-UHFFFAOYSA-N 4-methylumbelliferone Chemical compound C1=C(O)C=CC2=C1OC(=O)C=C2C HSHNITRMYYLLCV-UHFFFAOYSA-N 0.000 description 4
- KXDHJXZQYSOELW-UHFFFAOYSA-M Carbamate Chemical compound NC([O-])=O KXDHJXZQYSOELW-UHFFFAOYSA-M 0.000 description 4
- 108010016626 Dipeptides Proteins 0.000 description 4
- GKQLYSROISKDLL-UHFFFAOYSA-N EEDQ Chemical compound C1=CC=C2N(C(=O)OCC)C(OCC)C=CC2=C1 GKQLYSROISKDLL-UHFFFAOYSA-N 0.000 description 4
- 241000282326 Felis catus Species 0.000 description 4
- JUJWROOIHBZHMG-UHFFFAOYSA-N Pyridine Chemical compound C1=CC=NC=C1 JUJWROOIHBZHMG-UHFFFAOYSA-N 0.000 description 4
- 150000001298 alcohols Chemical class 0.000 description 4
- QRUDEWIWKLJBPS-UHFFFAOYSA-N benzotriazole Chemical compound C1=CC=C2N[N][N]C2=C1 QRUDEWIWKLJBPS-UHFFFAOYSA-N 0.000 description 4
- 239000012964 benzotriazole Substances 0.000 description 4
- 239000000872 buffer Substances 0.000 description 4
- 230000002255 enzymatic effect Effects 0.000 description 4
- 230000003100 immobilizing effect Effects 0.000 description 4
- 238000011534 incubation Methods 0.000 description 4
- 230000003834 intracellular effect Effects 0.000 description 4
- 239000012948 isocyanate Substances 0.000 description 4
- 150000002513 isocyanates Chemical class 0.000 description 4
- 210000003712 lysosome Anatomy 0.000 description 4
- 230000001868 lysosomic effect Effects 0.000 description 4
- VLKZOEOYAKHREP-UHFFFAOYSA-N n-Hexane Chemical class CCCCCC VLKZOEOYAKHREP-UHFFFAOYSA-N 0.000 description 4
- 235000018102 proteins Nutrition 0.000 description 4
- 108090000623 proteins and genes Proteins 0.000 description 4
- 102000004169 proteins and genes Human genes 0.000 description 4
- 239000000725 suspension Substances 0.000 description 4
- STBLNCCBQMHSRC-BATDWUPUSA-N (2s)-n-[(3s,4s)-5-acetyl-7-cyano-4-methyl-1-[(2-methylnaphthalen-1-yl)methyl]-2-oxo-3,4-dihydro-1,5-benzodiazepin-3-yl]-2-(methylamino)propanamide Chemical compound O=C1[C@@H](NC(=O)[C@H](C)NC)[C@H](C)N(C(C)=O)C2=CC(C#N)=CC=C2N1CC1=C(C)C=CC2=CC=CC=C12 STBLNCCBQMHSRC-BATDWUPUSA-N 0.000 description 3
- TVTJUIAKQFIXCE-HUKYDQBMSA-N 2-amino-9-[(2R,3S,4S,5R)-4-fluoro-3-hydroxy-5-(hydroxymethyl)oxolan-2-yl]-7-prop-2-ynyl-1H-purine-6,8-dione Chemical compound NC=1NC(C=2N(C(N(C=2N=1)[C@@H]1O[C@@H]([C@H]([C@H]1O)F)CO)=O)CC#C)=O TVTJUIAKQFIXCE-HUKYDQBMSA-N 0.000 description 3
- 206010006187 Breast cancer Diseases 0.000 description 3
- 208000026310 Breast neoplasm Diseases 0.000 description 3
- 102000004172 Cathepsin L Human genes 0.000 description 3
- 108090000624 Cathepsin L Proteins 0.000 description 3
- 102000010834 Extracellular Matrix Proteins Human genes 0.000 description 3
- 108010037362 Extracellular Matrix Proteins Proteins 0.000 description 3
- KDXKERNSBIXSRK-YFKPBYRVSA-N L-lysine Chemical compound NCCCC[C@H](N)C(O)=O KDXKERNSBIXSRK-YFKPBYRVSA-N 0.000 description 3
- 238000005481 NMR spectroscopy Methods 0.000 description 3
- BELBBZDIHDAJOR-UHFFFAOYSA-N Phenolsulfonephthalein Chemical compound C1=CC(O)=CC=C1C1(C=2C=CC(O)=CC=2)C2=CC=CC=C2S(=O)(=O)O1 BELBBZDIHDAJOR-UHFFFAOYSA-N 0.000 description 3
- 125000000217 alkyl group Chemical group 0.000 description 3
- 150000003862 amino acid derivatives Chemical class 0.000 description 3
- 125000003277 amino group Chemical group 0.000 description 3
- 230000003321 amplification Effects 0.000 description 3
- 150000001649 bromium compounds Chemical class 0.000 description 3
- KRKNYBCHXYNGOX-UHFFFAOYSA-N citric acid Chemical compound OC(=O)CC(O)(C(O)=O)CC(O)=O KRKNYBCHXYNGOX-UHFFFAOYSA-N 0.000 description 3
- 229940125782 compound 2 Drugs 0.000 description 3
- 229940125851 compound 27 Drugs 0.000 description 3
- 229940126214 compound 3 Drugs 0.000 description 3
- 229940125878 compound 36 Drugs 0.000 description 3
- 230000008878 coupling Effects 0.000 description 3
- 238000010168 coupling process Methods 0.000 description 3
- 235000018417 cysteine Nutrition 0.000 description 3
- XUJNEKJLAYXESH-UHFFFAOYSA-N cysteine Natural products SCC(N)C(O)=O XUJNEKJLAYXESH-UHFFFAOYSA-N 0.000 description 3
- 239000012039 electrophile Substances 0.000 description 3
- 210000002744 extracellular matrix Anatomy 0.000 description 3
- XLYOFNOQVPJJNP-ZSJDYOACSA-N heavy water Substances [2H]O[2H] XLYOFNOQVPJJNP-ZSJDYOACSA-N 0.000 description 3
- 230000002209 hydrophobic effect Effects 0.000 description 3
- 239000003112 inhibitor Substances 0.000 description 3
- 238000003199 nucleic acid amplification method Methods 0.000 description 3
- 229960003531 phenolsulfonphthalein Drugs 0.000 description 3
- 229940002612 prodrug Drugs 0.000 description 3
- 239000000651 prodrug Substances 0.000 description 3
- 230000002269 spontaneous effect Effects 0.000 description 3
- 239000000126 substance Substances 0.000 description 3
- 238000004809 thin layer chromatography Methods 0.000 description 3
- AOSZTAHDEDLTLQ-AZKQZHLXSA-N (1S,2S,4R,8S,9S,11S,12R,13S,19S)-6-[(3-chlorophenyl)methyl]-12,19-difluoro-11-hydroxy-8-(2-hydroxyacetyl)-9,13-dimethyl-6-azapentacyclo[10.8.0.02,9.04,8.013,18]icosa-14,17-dien-16-one Chemical compound C([C@@H]1C[C@H]2[C@H]3[C@]([C@]4(C=CC(=O)C=C4[C@@H](F)C3)C)(F)[C@@H](O)C[C@@]2([C@@]1(C1)C(=O)CO)C)N1CC1=CC=CC(Cl)=C1 AOSZTAHDEDLTLQ-AZKQZHLXSA-N 0.000 description 2
- IUSARDYWEPUTPN-OZBXUNDUSA-N (2r)-n-[(2s,3r)-4-[[(4s)-6-(2,2-dimethylpropyl)spiro[3,4-dihydropyrano[2,3-b]pyridine-2,1'-cyclobutane]-4-yl]amino]-3-hydroxy-1-[3-(1,3-thiazol-2-yl)phenyl]butan-2-yl]-2-methoxypropanamide Chemical compound C([C@H](NC(=O)[C@@H](C)OC)[C@H](O)CN[C@@H]1C2=CC(CC(C)(C)C)=CN=C2OC2(CCC2)C1)C(C=1)=CC=CC=1C1=NC=CS1 IUSARDYWEPUTPN-OZBXUNDUSA-N 0.000 description 2
- ZSQPDAOJXSYJNP-LBPRGKRZSA-N (2s)-2-amino-5-(diaminomethylideneamino)-n-(4-methyl-2-oxochromen-7-yl)pentanamide Chemical compound C1=C(NC(=O)[C@@H](N)CCCN=C(N)N)C=CC2=C1OC(=O)C=C2C ZSQPDAOJXSYJNP-LBPRGKRZSA-N 0.000 description 2
- NXLNNXIXOYSCMB-UHFFFAOYSA-N (4-nitrophenyl) carbonochloridate Chemical compound [O-][N+](=O)C1=CC=C(OC(Cl)=O)C=C1 NXLNNXIXOYSCMB-UHFFFAOYSA-N 0.000 description 2
- RTHHSXOVIJWFQP-UHFFFAOYSA-N 7-hydroxy-4-methyl-2-oxochromene-8-carbaldehyde Chemical compound O=CC1=C(O)C=CC2=C1OC(=O)C=C2C RTHHSXOVIJWFQP-UHFFFAOYSA-N 0.000 description 2
- PAYRUJLWNCNPSJ-UHFFFAOYSA-N Aniline Chemical compound NC1=CC=CC=C1 PAYRUJLWNCNPSJ-UHFFFAOYSA-N 0.000 description 2
- 206010008342 Cervix carcinoma Diseases 0.000 description 2
- 229940126657 Compound 17 Drugs 0.000 description 2
- 208000017891 HER2 positive breast carcinoma Diseases 0.000 description 2
- 241000282412 Homo Species 0.000 description 2
- 101000898449 Homo sapiens Cathepsin B Proteins 0.000 description 2
- 229930040373 Paraformaldehyde Natural products 0.000 description 2
- UIIMBOGNXHQVGW-UHFFFAOYSA-M Sodium bicarbonate Chemical compound [Na+].OC([O-])=O UIIMBOGNXHQVGW-UHFFFAOYSA-M 0.000 description 2
- 208000006105 Uterine Cervical Neoplasms Diseases 0.000 description 2
- 239000008351 acetate buffer Substances 0.000 description 2
- 238000013459 approach Methods 0.000 description 2
- 150000004657 carbamic acid derivatives Chemical class 0.000 description 2
- 230000001413 cellular effect Effects 0.000 description 2
- 201000010881 cervical cancer Diseases 0.000 description 2
- 238000012512 characterization method Methods 0.000 description 2
- 239000007795 chemical reaction product Substances 0.000 description 2
- 238000002512 chemotherapy Methods 0.000 description 2
- 229940125807 compound 37 Drugs 0.000 description 2
- 230000005284 excitation Effects 0.000 description 2
- 239000012091 fetal bovine serum Substances 0.000 description 2
- 238000002073 fluorescence micrograph Methods 0.000 description 2
- 125000000291 glutamic acid group Chemical group N[C@@H](CCC(O)=O)C(=O)* 0.000 description 2
- 102000053907 human CTSB Human genes 0.000 description 2
- 238000001727 in vivo Methods 0.000 description 2
- 239000000543 intermediate Substances 0.000 description 2
- 229910052943 magnesium sulfate Inorganic materials 0.000 description 2
- 230000007246 mechanism Effects 0.000 description 2
- VSPPONOIKZXUBJ-UHFFFAOYSA-N n,n-diethylethanamine;oxolane Chemical compound C1CCOC1.CCN(CC)CC VSPPONOIKZXUBJ-UHFFFAOYSA-N 0.000 description 2
- 230000000269 nucleophilic effect Effects 0.000 description 2
- 229920002866 paraformaldehyde Polymers 0.000 description 2
- 238000002360 preparation method Methods 0.000 description 2
- 239000000047 product Substances 0.000 description 2
- 230000002797 proteolythic effect Effects 0.000 description 2
- 238000000746 purification Methods 0.000 description 2
- UMJSCPRVCHMLSP-UHFFFAOYSA-N pyridine Natural products COC1=CC=CN=C1 UMJSCPRVCHMLSP-UHFFFAOYSA-N 0.000 description 2
- 230000009467 reduction Effects 0.000 description 2
- 102200042162 rs145415848 Human genes 0.000 description 2
- FDRCDNZGSXJAFP-UHFFFAOYSA-M sodium chloroacetate Chemical compound [Na+].[O-]C(=O)CCl FDRCDNZGSXJAFP-UHFFFAOYSA-M 0.000 description 2
- 239000006228 supernatant Substances 0.000 description 2
- 230000008685 targeting Effects 0.000 description 2
- 230000001225 therapeutic effect Effects 0.000 description 2
- FYSNRJHAOHDILO-UHFFFAOYSA-N thionyl chloride Chemical compound ClS(Cl)=O FYSNRJHAOHDILO-UHFFFAOYSA-N 0.000 description 2
- 229940086542 triethylamine Drugs 0.000 description 2
- RIOQSEWOXXDEQQ-UHFFFAOYSA-N triphenylphosphine Chemical compound C1=CC=CC=C1P(C=1C=CC=CC=1)C1=CC=CC=C1 RIOQSEWOXXDEQQ-UHFFFAOYSA-N 0.000 description 2
- ASGMFNBUXDJWJJ-JLCFBVMHSA-N (1R,3R)-3-[[3-bromo-1-[4-(5-methyl-1,3,4-thiadiazol-2-yl)phenyl]pyrazolo[3,4-d]pyrimidin-6-yl]amino]-N,1-dimethylcyclopentane-1-carboxamide Chemical compound BrC1=NN(C2=NC(=NC=C21)N[C@H]1C[C@@](CC1)(C(=O)NC)C)C1=CC=C(C=C1)C=1SC(=NN=1)C ASGMFNBUXDJWJJ-JLCFBVMHSA-N 0.000 description 1
- GLGNXYJARSMNGJ-VKTIVEEGSA-N (1s,2s,3r,4r)-3-[[5-chloro-2-[(1-ethyl-6-methoxy-2-oxo-4,5-dihydro-3h-1-benzazepin-7-yl)amino]pyrimidin-4-yl]amino]bicyclo[2.2.1]hept-5-ene-2-carboxamide Chemical compound CCN1C(=O)CCCC2=C(OC)C(NC=3N=C(C(=CN=3)Cl)N[C@H]3[C@H]([C@@]4([H])C[C@@]3(C=C4)[H])C(N)=O)=CC=C21 GLGNXYJARSMNGJ-VKTIVEEGSA-N 0.000 description 1
- IWZSHWBGHQBIML-ZGGLMWTQSA-N (3S,8S,10R,13S,14S,17S)-17-isoquinolin-7-yl-N,N,10,13-tetramethyl-2,3,4,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-3-amine Chemical compound CN(C)[C@H]1CC[C@]2(C)C3CC[C@@]4(C)[C@@H](CC[C@@H]4c4ccc5ccncc5c4)[C@@H]3CC=C2C1 IWZSHWBGHQBIML-ZGGLMWTQSA-N 0.000 description 1
- PFKFTWBEEFSNDU-UHFFFAOYSA-N 1,1'-Carbonyldiimidazole Substances C1=CN=CN1C(=O)N1C=CN=C1 PFKFTWBEEFSNDU-UHFFFAOYSA-N 0.000 description 1
- RYHBNJHYFVUHQT-UHFFFAOYSA-N 1,4-Dioxane Chemical compound C1COCCO1 RYHBNJHYFVUHQT-UHFFFAOYSA-N 0.000 description 1
- KQZLRWGGWXJPOS-NLFPWZOASA-N 1-[(1R)-1-(2,4-dichlorophenyl)ethyl]-6-[(4S,5R)-4-[(2S)-2-(hydroxymethyl)pyrrolidin-1-yl]-5-methylcyclohexen-1-yl]pyrazolo[3,4-b]pyrazine-3-carbonitrile Chemical compound ClC1=C(C=CC(=C1)Cl)[C@@H](C)N1N=C(C=2C1=NC(=CN=2)C1=CC[C@@H]([C@@H](C1)C)N1[C@@H](CCC1)CO)C#N KQZLRWGGWXJPOS-NLFPWZOASA-N 0.000 description 1
- KGNQDBQYEBMPFZ-UHFFFAOYSA-N 1-fluoro-4-iodobenzene Chemical compound FC1=CC=C(I)C=C1 KGNQDBQYEBMPFZ-UHFFFAOYSA-N 0.000 description 1
- RNAMYOYQYRYFQY-UHFFFAOYSA-N 2-(4,4-difluoropiperidin-1-yl)-6-methoxy-n-(1-propan-2-ylpiperidin-4-yl)-7-(3-pyrrolidin-1-ylpropoxy)quinazolin-4-amine Chemical compound N1=C(N2CCC(F)(F)CC2)N=C2C=C(OCCCN3CCCC3)C(OC)=CC2=C1NC1CCN(C(C)C)CC1 RNAMYOYQYRYFQY-UHFFFAOYSA-N 0.000 description 1
- QBWKPGNFQQJGFY-QLFBSQMISA-N 3-[(1r)-1-[(2r,6s)-2,6-dimethylmorpholin-4-yl]ethyl]-n-[6-methyl-3-(1h-pyrazol-4-yl)imidazo[1,2-a]pyrazin-8-yl]-1,2-thiazol-5-amine Chemical compound N1([C@H](C)C2=NSC(NC=3C4=NC=C(N4C=C(C)N=3)C3=CNN=C3)=C2)C[C@H](C)O[C@H](C)C1 QBWKPGNFQQJGFY-QLFBSQMISA-N 0.000 description 1
- TYMLOMAKGOJONV-UHFFFAOYSA-N 4-nitroaniline Chemical compound NC1=CC=C([N+]([O-])=O)C=C1 TYMLOMAKGOJONV-UHFFFAOYSA-N 0.000 description 1
- MZCNCJNOOWONIO-UHFFFAOYSA-N 5-chlorotetrazine Chemical compound ClC1=CN=NN=N1 MZCNCJNOOWONIO-UHFFFAOYSA-N 0.000 description 1
- QTBSBXVTEAMEQO-UHFFFAOYSA-M Acetate Chemical compound CC([O-])=O QTBSBXVTEAMEQO-UHFFFAOYSA-M 0.000 description 1
- 238000009010 Bradford assay Methods 0.000 description 1
- OBMZMSLWNNWEJA-XNCRXQDQSA-N C1=CC=2C(C[C@@H]3NC(=O)[C@@H](NC(=O)[C@H](NC(=O)N(CC#CCN(CCCC[C@H](NC(=O)[C@@H](CC4=CC=CC=C4)NC3=O)C(=O)N)CC=C)NC(=O)[C@@H](N)C)CC3=CNC4=C3C=CC=C4)C)=CNC=2C=C1 Chemical compound C1=CC=2C(C[C@@H]3NC(=O)[C@@H](NC(=O)[C@H](NC(=O)N(CC#CCN(CCCC[C@H](NC(=O)[C@@H](CC4=CC=CC=C4)NC3=O)C(=O)N)CC=C)NC(=O)[C@@H](N)C)CC3=CNC4=C3C=CC=C4)C)=CNC=2C=C1 OBMZMSLWNNWEJA-XNCRXQDQSA-N 0.000 description 1
- 229940123003 Cathepsin inhibitor Drugs 0.000 description 1
- 101000749287 Clitocybe nebularis Clitocypin Proteins 0.000 description 1
- 101000767029 Clitocybe nebularis Clitocypin-1 Proteins 0.000 description 1
- 229940127007 Compound 39 Drugs 0.000 description 1
- 229940094664 Cysteine protease inhibitor Drugs 0.000 description 1
- 101150029707 ERBB2 gene Proteins 0.000 description 1
- LFQSCWFLJHTTHZ-UHFFFAOYSA-N Ethanol Chemical compound CCO LFQSCWFLJHTTHZ-UHFFFAOYSA-N 0.000 description 1
- PXGOKWXKJXAPGV-UHFFFAOYSA-N Fluorine Chemical compound FF PXGOKWXKJXAPGV-UHFFFAOYSA-N 0.000 description 1
- 241000282414 Homo sapiens Species 0.000 description 1
- 101000983583 Homo sapiens Procathepsin L Proteins 0.000 description 1
- 101000852559 Homo sapiens Thioredoxin Proteins 0.000 description 1
- DGAQECJNVWCQMB-PUAWFVPOSA-M Ilexoside XXIX Chemical compound C[C@@H]1CC[C@@]2(CC[C@@]3(C(=CC[C@H]4[C@]3(CC[C@@H]5[C@@]4(CC[C@@H](C5(C)C)OS(=O)(=O)[O-])C)C)[C@@H]2[C@]1(C)O)C)C(=O)O[C@H]6[C@@H]([C@H]([C@@H]([C@H](O6)CO)O)O)O.[Na+] DGAQECJNVWCQMB-PUAWFVPOSA-M 0.000 description 1
- COLNVLDHVKWLRT-QMMMGPOBSA-N L-phenylalanine Chemical compound OC(=O)[C@@H](N)CC1=CC=CC=C1 COLNVLDHVKWLRT-QMMMGPOBSA-N 0.000 description 1
- KDXKERNSBIXSRK-UHFFFAOYSA-N Lysine Natural products NCCCCC(N)C(O)=O KDXKERNSBIXSRK-UHFFFAOYSA-N 0.000 description 1
- 239000004472 Lysine Substances 0.000 description 1
- 206010027476 Metastases Diseases 0.000 description 1
- 229930012538 Paclitaxel Natural products 0.000 description 1
- 101710176384 Peptide 1 Proteins 0.000 description 1
- 102000012479 Serine Proteases Human genes 0.000 description 1
- 108010022999 Serine Proteases Proteins 0.000 description 1
- 241000700584 Simplexvirus Species 0.000 description 1
- DHXVGJBLRPWPCS-UHFFFAOYSA-N Tetrahydropyran Chemical compound C1CCOCC1 DHXVGJBLRPWPCS-UHFFFAOYSA-N 0.000 description 1
- 102000006601 Thymidine Kinase Human genes 0.000 description 1
- 108020004440 Thymidine kinase Proteins 0.000 description 1
- 229920004890 Triton X-100 Polymers 0.000 description 1
- 239000013504 Triton X-100 Substances 0.000 description 1
- BIRZKUUPHCQOPK-GJZGRUSLSA-N Z-Arg-Arg Chemical compound NC(N)=NCCC[C@@H](C(O)=O)NC(=O)[C@H](CCCN=C(N)N)NC(=O)OCC1=CC=CC=C1 BIRZKUUPHCQOPK-GJZGRUSLSA-N 0.000 description 1
- LNUFLCYMSVYYNW-ZPJMAFJPSA-N [(2r,3r,4s,5r,6r)-2-[(2r,3r,4s,5r,6r)-6-[(2r,3r,4s,5r,6r)-6-[(2r,3r,4s,5r,6r)-6-[[(3s,5s,8r,9s,10s,13r,14s,17r)-10,13-dimethyl-17-[(2r)-6-methylheptan-2-yl]-2,3,4,5,6,7,8,9,11,12,14,15,16,17-tetradecahydro-1h-cyclopenta[a]phenanthren-3-yl]oxy]-4,5-disulfo Chemical compound O([C@@H]1[C@@H](COS(O)(=O)=O)O[C@@H]([C@@H]([C@H]1OS(O)(=O)=O)OS(O)(=O)=O)O[C@@H]1[C@@H](COS(O)(=O)=O)O[C@@H]([C@@H]([C@H]1OS(O)(=O)=O)OS(O)(=O)=O)O[C@@H]1[C@@H](COS(O)(=O)=O)O[C@H]([C@@H]([C@H]1OS(O)(=O)=O)OS(O)(=O)=O)O[C@@H]1C[C@@H]2CC[C@H]3[C@@H]4CC[C@@H]([C@]4(CC[C@@H]3[C@@]2(C)CC1)C)[C@H](C)CCCC(C)C)[C@H]1O[C@H](COS(O)(=O)=O)[C@@H](OS(O)(=O)=O)[C@H](OS(O)(=O)=O)[C@H]1OS(O)(=O)=O LNUFLCYMSVYYNW-ZPJMAFJPSA-N 0.000 description 1
- 239000002253 acid Substances 0.000 description 1
- 230000003213 activating effect Effects 0.000 description 1
- 238000010171 animal model Methods 0.000 description 1
- 230000001093 anti-cancer Effects 0.000 description 1
- 239000002543 antimycotic Substances 0.000 description 1
- 230000006907 apoptotic process Effects 0.000 description 1
- 239000008346 aqueous phase Substances 0.000 description 1
- 239000012298 atmosphere Substances 0.000 description 1
- 230000008901 benefit Effects 0.000 description 1
- KGNDCEVUMONOKF-UGPLYTSKSA-N benzyl n-[(2r)-1-[(2s,4r)-2-[[(2s)-6-amino-1-(1,3-benzoxazol-2-yl)-1,1-dihydroxyhexan-2-yl]carbamoyl]-4-[(4-methylphenyl)methoxy]pyrrolidin-1-yl]-1-oxo-4-phenylbutan-2-yl]carbamate Chemical compound C1=CC(C)=CC=C1CO[C@H]1CN(C(=O)[C@@H](CCC=2C=CC=CC=2)NC(=O)OCC=2C=CC=CC=2)[C@H](C(=O)N[C@@H](CCCCN)C(O)(O)C=2OC3=CC=CC=C3N=2)C1 KGNDCEVUMONOKF-UGPLYTSKSA-N 0.000 description 1
- 230000033228 biological regulation Effects 0.000 description 1
- 244000309464 bull Species 0.000 description 1
- 210000004899 c-terminal region Anatomy 0.000 description 1
- 239000012830 cancer therapeutic Substances 0.000 description 1
- 238000001460 carbon-13 nuclear magnetic resonance spectrum Methods 0.000 description 1
- 230000003197 catalytic effect Effects 0.000 description 1
- 125000002091 cationic group Chemical group 0.000 description 1
- 238000004113 cell culture Methods 0.000 description 1
- 230000030833 cell death Effects 0.000 description 1
- 230000030570 cellular localization Effects 0.000 description 1
- PBAYDYUZOSNJGU-UHFFFAOYSA-N chelidonic acid Natural products OC(=O)C1=CC(=O)C=C(C(O)=O)O1 PBAYDYUZOSNJGU-UHFFFAOYSA-N 0.000 description 1
- 229940125758 compound 15 Drugs 0.000 description 1
- 229940125833 compound 23 Drugs 0.000 description 1
- 229940125846 compound 25 Drugs 0.000 description 1
- 229940125877 compound 31 Drugs 0.000 description 1
- 230000021615 conjugation Effects 0.000 description 1
- 125000000332 coumarinyl group Chemical group O1C(=O)C(=CC2=CC=CC=C12)* 0.000 description 1
- 230000009089 cytolysis Effects 0.000 description 1
- 229940127089 cytotoxic agent Drugs 0.000 description 1
- 239000002254 cytotoxic agent Substances 0.000 description 1
- 230000034994 death Effects 0.000 description 1
- 230000001419 dependent effect Effects 0.000 description 1
- 239000000104 diagnostic biomarker Substances 0.000 description 1
- 238000002405 diagnostic procedure Methods 0.000 description 1
- WGLUMOCWFMKWIL-UHFFFAOYSA-N dichloromethane;methanol Chemical compound OC.ClCCl WGLUMOCWFMKWIL-UHFFFAOYSA-N 0.000 description 1
- IJKVHSBPTUYDLN-UHFFFAOYSA-N dihydroxy(oxo)silane Chemical compound O[Si](O)=O IJKVHSBPTUYDLN-UHFFFAOYSA-N 0.000 description 1
- 201000010099 disease Diseases 0.000 description 1
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 1
- 230000008030 elimination Effects 0.000 description 1
- 238000003379 elimination reaction Methods 0.000 description 1
- 238000006911 enzymatic reaction Methods 0.000 description 1
- 238000001952 enzyme assay Methods 0.000 description 1
- 150000002170 ethers Chemical class 0.000 description 1
- OAYLNYINCPYISS-UHFFFAOYSA-N ethyl acetate;hexane Chemical class CCCCCC.CCOC(C)=O OAYLNYINCPYISS-UHFFFAOYSA-N 0.000 description 1
- 238000011156 evaluation Methods 0.000 description 1
- 239000011737 fluorine Substances 0.000 description 1
- 239000007789 gas Substances 0.000 description 1
- 102000050937 human CTSL Human genes 0.000 description 1
- 102000056461 human TXN Human genes 0.000 description 1
- 239000005457 ice water Substances 0.000 description 1
- 238000000338 in vitro Methods 0.000 description 1
- 230000002779 inactivation Effects 0.000 description 1
- 238000010348 incorporation Methods 0.000 description 1
- 230000005764 inhibitory process Effects 0.000 description 1
- 229940030980 inova Drugs 0.000 description 1
- 230000002427 irreversible effect Effects 0.000 description 1
- 238000003367 kinetic assay Methods 0.000 description 1
- 150000002611 lead compounds Chemical class 0.000 description 1
- 238000010859 live-cell imaging Methods 0.000 description 1
- 239000003550 marker Substances 0.000 description 1
- 239000002609 medium Substances 0.000 description 1
- 239000012528 membrane Substances 0.000 description 1
- 230000009401 metastasis Effects 0.000 description 1
- 238000001000 micrograph Methods 0.000 description 1
- 238000000386 microscopy Methods 0.000 description 1
- 238000012986 modification Methods 0.000 description 1
- 230000004048 modification Effects 0.000 description 1
- 230000004660 morphological change Effects 0.000 description 1
- 230000009826 neoplastic cell growth Effects 0.000 description 1
- 229910052757 nitrogen Inorganic materials 0.000 description 1
- 230000003287 optical effect Effects 0.000 description 1
- AICOOMRHRUFYCM-ZRRPKQBOSA-N oxazine, 1 Chemical compound C([C@@H]1[C@H](C(C[C@]2(C)[C@@H]([C@H](C)N(C)C)[C@H](O)C[C@]21C)=O)CC1=CC2)C[C@H]1[C@@]1(C)[C@H]2N=C(C(C)C)OC1 AICOOMRHRUFYCM-ZRRPKQBOSA-N 0.000 description 1
- 229960001592 paclitaxel Drugs 0.000 description 1
- 230000035699 permeability Effects 0.000 description 1
- ISWSIDIOOBJBQZ-UHFFFAOYSA-N phenol group Chemical group C1(=CC=CC=C1)O ISWSIDIOOBJBQZ-UHFFFAOYSA-N 0.000 description 1
- 125000001997 phenyl group Chemical group [H]C1=C([H])C([H])=C(*)C([H])=C1[H] 0.000 description 1
- COLNVLDHVKWLRT-UHFFFAOYSA-N phenylalanine Natural products OC(=O)C(N)CC1=CC=CC=C1 COLNVLDHVKWLRT-UHFFFAOYSA-N 0.000 description 1
- 230000008569 process Effects 0.000 description 1
- 238000012545 processing Methods 0.000 description 1
- 239000000092 prognostic biomarker Substances 0.000 description 1
- 230000002035 prolonged effect Effects 0.000 description 1
- 230000001737 promoting effect Effects 0.000 description 1
- 238000000425 proton nuclear magnetic resonance spectrum Methods 0.000 description 1
- 150000003254 radicals Chemical class 0.000 description 1
- 230000002285 radioactive effect Effects 0.000 description 1
- HSSLDCABUXLXKM-UHFFFAOYSA-N resorufin Chemical compound C1=CC(=O)C=C2OC3=CC(O)=CC=C3N=C21 HSSLDCABUXLXKM-UHFFFAOYSA-N 0.000 description 1
- 230000004044 response Effects 0.000 description 1
- 230000000717 retained effect Effects 0.000 description 1
- 229920006395 saturated elastomer Polymers 0.000 description 1
- 210000002966 serum Anatomy 0.000 description 1
- 239000002002 slurry Substances 0.000 description 1
- 229910052708 sodium Inorganic materials 0.000 description 1
- 235000017557 sodium bicarbonate Nutrition 0.000 description 1
- 239000012279 sodium borohydride Substances 0.000 description 1
- 229910000033 sodium borohydride Inorganic materials 0.000 description 1
- TYXBZZBYMSVFPR-UHFFFAOYSA-M sodium;7-oxophenoxazin-3-olate Chemical compound [Na+].C1=CC(=O)C=C2OC3=CC([O-])=CC=C3N=C21 TYXBZZBYMSVFPR-UHFFFAOYSA-M 0.000 description 1
- 230000002459 sustained effect Effects 0.000 description 1
- RCINICONZNJXQF-MZXODVADSA-N taxol Chemical compound O([C@@H]1[C@@]2(C[C@@H](C(C)=C(C2(C)C)[C@H](C([C@]2(C)[C@@H](O)C[C@H]3OC[C@]3([C@H]21)OC(C)=O)=O)OC(=O)C)OC(=O)[C@H](O)[C@@H](NC(=O)C=1C=CC=CC=1)C=1C=CC=CC=1)O)C(=O)C1=CC=CC=C1 RCINICONZNJXQF-MZXODVADSA-N 0.000 description 1
- 238000012360 testing method Methods 0.000 description 1
- FPGGTKZVZWFYPV-UHFFFAOYSA-M tetrabutylammonium fluoride Chemical compound [F-].CCCC[N+](CCCC)(CCCC)CCCC FPGGTKZVZWFYPV-UHFFFAOYSA-M 0.000 description 1
- 238000004448 titration Methods 0.000 description 1
- CWMFRHBXRUITQE-UHFFFAOYSA-N trimethylsilylacetylene Chemical compound C[Si](C)(C)C#C CWMFRHBXRUITQE-UHFFFAOYSA-N 0.000 description 1
- 239000000439 tumor marker Substances 0.000 description 1
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K5/00—Peptides containing up to four amino acids in a fully defined sequence; Derivatives thereof
- C07K5/04—Peptides containing up to four amino acids in a fully defined sequence; Derivatives thereof containing only normal peptide links
- C07K5/06—Dipeptides
- C07K5/06086—Dipeptides with the first amino acid being basic
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K49/00—Preparations for testing in vivo
- A61K49/001—Preparation for luminescence or biological staining
- A61K49/0013—Luminescence
- A61K49/0017—Fluorescence in vivo
- A61K49/0019—Fluorescence in vivo characterised by the fluorescent group, e.g. oligomeric, polymeric or dendritic molecules
- A61K49/0021—Fluorescence in vivo characterised by the fluorescent group, e.g. oligomeric, polymeric or dendritic molecules the fluorescent group being a small organic molecule
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K49/00—Preparations for testing in vivo
- A61K49/001—Preparation for luminescence or biological staining
- A61K49/0013—Luminescence
- A61K49/0017—Fluorescence in vivo
- A61K49/0019—Fluorescence in vivo characterised by the fluorescent group, e.g. oligomeric, polymeric or dendritic molecules
- A61K49/0021—Fluorescence in vivo characterised by the fluorescent group, e.g. oligomeric, polymeric or dendritic molecules the fluorescent group being a small organic molecule
- A61K49/0028—Oxazine dyes
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K49/00—Preparations for testing in vivo
- A61K49/001—Preparation for luminescence or biological staining
- A61K49/0013—Luminescence
- A61K49/0017—Fluorescence in vivo
- A61K49/0019—Fluorescence in vivo characterised by the fluorescent group, e.g. oligomeric, polymeric or dendritic molecules
- A61K49/0021—Fluorescence in vivo characterised by the fluorescent group, e.g. oligomeric, polymeric or dendritic molecules the fluorescent group being a small organic molecule
- A61K49/0039—Coumarin dyes
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K49/00—Preparations for testing in vivo
- A61K49/001—Preparation for luminescence or biological staining
- A61K49/0013—Luminescence
- A61K49/0017—Fluorescence in vivo
- A61K49/0019—Fluorescence in vivo characterised by the fluorescent group, e.g. oligomeric, polymeric or dendritic molecules
- A61K49/0021—Fluorescence in vivo characterised by the fluorescent group, e.g. oligomeric, polymeric or dendritic molecules the fluorescent group being a small organic molecule
- A61K49/0041—Xanthene dyes, used in vivo, e.g. administered to a mice, e.g. rhodamines, rose Bengal
- A61K49/0043—Fluorescein, used in vivo
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K49/00—Preparations for testing in vivo
- A61K49/001—Preparation for luminescence or biological staining
- A61K49/0013—Luminescence
- A61K49/0017—Fluorescence in vivo
- A61K49/005—Fluorescence in vivo characterised by the carrier molecule carrying the fluorescent agent
- A61K49/0052—Small organic molecules
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K51/00—Preparations containing radioactive substances for use in therapy or testing in vivo
- A61K51/02—Preparations containing radioactive substances for use in therapy or testing in vivo characterised by the carrier, i.e. characterised by the agent or material covalently linked or complexing the radioactive nucleus
- A61K51/04—Organic compounds
- A61K51/08—Peptides, e.g. proteins, carriers being peptides, polyamino acids, proteins
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07D—HETEROCYCLIC COMPOUNDS
- C07D265/00—Heterocyclic compounds containing six-membered rings having one nitrogen atom and one oxygen atom as the only ring hetero atoms
- C07D265/28—1,4-Oxazines; Hydrogenated 1,4-oxazines
- C07D265/34—1,4-Oxazines; Hydrogenated 1,4-oxazines condensed with carbocyclic rings
- C07D265/38—[b, e]-condensed with two six-membered rings
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07D—HETEROCYCLIC COMPOUNDS
- C07D311/00—Heterocyclic compounds containing six-membered rings having one oxygen atom as the only hetero atom, condensed with other rings
- C07D311/02—Heterocyclic compounds containing six-membered rings having one oxygen atom as the only hetero atom, condensed with other rings ortho- or peri-condensed with carbocyclic rings or ring systems
- C07D311/04—Benzo[b]pyrans, not hydrogenated in the carbocyclic ring
- C07D311/06—Benzo[b]pyrans, not hydrogenated in the carbocyclic ring with oxygen or sulfur atoms directly attached in position 2
- C07D311/08—Benzo[b]pyrans, not hydrogenated in the carbocyclic ring with oxygen or sulfur atoms directly attached in position 2 not hydrogenated in the hetero ring
- C07D311/16—Benzo[b]pyrans, not hydrogenated in the carbocyclic ring with oxygen or sulfur atoms directly attached in position 2 not hydrogenated in the hetero ring substituted in position 7
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K5/00—Peptides containing up to four amino acids in a fully defined sequence; Derivatives thereof
- C07K5/04—Peptides containing up to four amino acids in a fully defined sequence; Derivatives thereof containing only normal peptide links
- C07K5/06—Dipeptides
- C07K5/06008—Dipeptides with the first amino acid being neutral
- C07K5/06017—Dipeptides with the first amino acid being neutral and aliphatic
- C07K5/06034—Dipeptides with the first amino acid being neutral and aliphatic the side chain containing 2 to 4 carbon atoms
- C07K5/06052—Val-amino acid
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K5/00—Peptides containing up to four amino acids in a fully defined sequence; Derivatives thereof
- C07K5/04—Peptides containing up to four amino acids in a fully defined sequence; Derivatives thereof containing only normal peptide links
- C07K5/06—Dipeptides
- C07K5/06008—Dipeptides with the first amino acid being neutral
- C07K5/06078—Dipeptides with the first amino acid being neutral and aromatic or cycloaliphatic
Definitions
- the present invention relates to compounds that are cathepsin B substrates.
- the compounds find particular use as imaging probes, for example in positron emission tomography (PET).
- PET positron emission tomography
- proteolytic network degrades the extracellular matrix (ECM) components producing a permissive region for cancer cells to invade.
- Cathepsin B (CTB, EC 3.4.22.1 ) is produced by a variety of tumor-associated cells and is a prominent member of the tumor promoting proteolytic network.' 3"51 Mounting evidence suggests that CTB may be a potential diagnostic and prognostic biomarker for various cancers.' 6"101 Interestingly, CTB appears to affect cancer progression dependent upon its localization with intracellular CTB involved in amplification of apoptosis. 1111 In contrast, aggressive cancers have high extracellular CTB activity at the invading edge of a tumor and in the local ECM secreted by a variety of cell types.
- CTB may protect against anticancer chemotherapies such as Taxol, meaning CTB and/or CTL activity may be a marker to predict response to chemotherapy.
- CTL cathepsin L
- 1131 Accurate and sensitive tools capable of assessing CTB's activity in cell culture, animal models of aggressive cancer or cancer patients are needed to validate this protease as a high priority cancer marker or therapeutic target.
- One object of the invention is to provide peptide-based agents that can be cleaved by cathepsin B.
- Preferred compounds yield a fluorophore and/or a PET imaging agent upon cleavage by cathepsin B become that then becomes
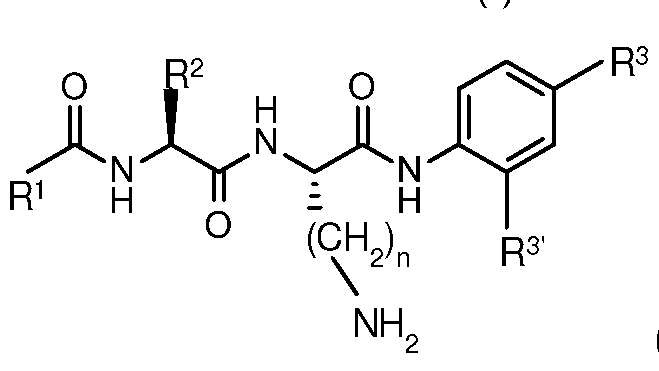
- a compound of the invention has formula (I):
- R 1 is PhCH 2 0-, CH 3 or PhCH 2 -;
- R 2 is PhCH 2 -, (CH 3 ) 2 CH- or H 2 N(CH 2 ) m - for m > 4;
- R 3 and R 3' are each H or -CH 2 OR 4 , and at least one of R 3 and R 3'
- R 4 is -CH 2 OR 4 ,wherein for each of R 3 and R 3 , R 4 is independently selected from the group consisting of:
- R 5 is H, -C(0)H, -CHX 2 , wherein each X is, independently of the other
- X selected from the group consisting of:
- R 7 is H, -C ⁇ CH, N 3 , -C(0)H, or -CHX 2 , wherein each X is, independently of the other X, selected from the group consisting of:
- R 8 is H, -C ⁇ CH, -C(0)H, N 3 , -CHF 2 , or -CH 2 F,
- R 9 and R 10 is, independently of the other, H,
- H F, CI, Br, I, -N0 2 , toluenesulfonate, methanesulfonate, trifluoromethansulfonate, perfluorobutanesulfonate, ethanesulfonate, benzenesulfonate, parachlorobenzenesulfonate, nitrobenzenesulfonate or methoxybenzenesulfonate, and both X are not simultaneously H,
- Y is selected from the group consisting of:
- Ar 1 is -C 6 H 5 in which 1 , 2 or 3 hydrogen atoms is optionally and independently substituted by -NO 2 , and
- N N where Z is selected from the group consisting of H, CI, and Br.
- R 1 1 is H or Ar 2 , in which Ar 2 is -C 6 H 5 in which 1 , 2 or 3 hydrogen atoms is optionally and independently substituted by F, and
- R 9 and R 10 are not simultaneously H
- the compound includes fluorescent dye i.e., a fluorescent dye molecule is incorporated into the compound to form a part thereof, as exemplified herein. It is understood that the dye is covalently linked to the remainder of the compound.
- fluorescent dyes include
- BODIPY borondipyrromethene core.
- a BODIPY portion of the compound can be covalently linked at the carbon atom of the dye located at the para-position with respect to the boron atom.
- the radiolabeled substituent comprises an 18 F atom.
- the radiolabeled substituent can be a C1 -C20 alkyl group or C5-C8 aromatic group in which the aromatic ring is made up of carbon atoms or carbon and nitrogen atoms.
- the ring is optionally substituted with a C1 -C20 alkyl or alkoxy group that is optionally substituted with said 18 F atom(s), and/or the ring bears one or more 18 F.
- X can be C N 2, 3, 4 5, 6
- R 3 is the same as R 3 , or they are different.
- m and n are each typically 4, 5, 6, 7, 8, 9 or 10.
- the values of m and n are independent of each other.
- the invention is a composition for use as an imaging probe, the composition including at least one compound having formula (I) as described above.
- the invention is a pharmaceutical composition for use in PET and/or fluorescence imaging, the composition including at least one compound having formula (I) as described above.
- the composition can be one in which the compound(s) is dispersed or dissolved in a liquid medium suitable for injection.
- the invention includes a method of PET and/or fluorescence imaging.
- the method includes administering to a patient in need thereof an effective amount of a composition(s) as described above, and scanning the subject with at PET and/or fluorescence imaging device.
- Such embodiments can be limited to a diagnostic method.
- the invention includes method for studying the localization of PET probes within a tissue of a subject comprising: administering to the subject an effective amount of a compound(s) as described above, subjecting a tissue of the subject to irradiation of an electromagnetic radiation, wherein the electromagnetic radiation is absorbed by the compound, detecting fluorescence of the compound within the tissue, wherein the fluorescence of the compound within the tissue is indicative of the presence of a PET probe within the tissue.
- Figure 1 illustrate the general strategy for obtaining probes of the invention
- Figure 2(b) is a schematic representation of a probe employing PABA extend into the CTB S2' recognition site, and Figure 2(c) shows two fluorogenic peptides of the invention
- Figure 3 shows fluorescent microscope images taken of HeLa cells treated for 2 hours with a) 40 ⁇ of (3) and b) 40 ⁇ of (4). Cells were treated overnight with CA-074Me and E64d followed by incubation with each probe for 2 hours. White bars represent 50 ⁇ ;
- Figure 4 is a bar graph showing results obtained with HeLa cell lysates incubated with Z-Arg-Arg-AMC, compounds 3 and 4 using untreated cell lysates and cell lysates prepared from cells treated with CA-074Me or E64D. Bars represent the relative activity (%) compared to Z-Arg-Arg-AMC incubated with untreated lysates adjusted to total protein concentration minus background RFU with error bars depicting standard deviation from 4 independent trials;
- Figure 5 illustrates a mechanism of cleavage of a particular agent by CTB followed by immobilization of a compound produced by said cleavage
- Figure 6 illustrates a general structure of an agent showing the CTB-targeting moiety and immobilizing compound, R 4 , connected to the targeting moiety via a linker;
- Figure 7 shows results obtained in a representative fluorescence microscopy experiment using the agent shown in Figure 5, to demonstrate the retention of the immobilized reporter molecule once activated by cathepsin B;
- FIG. 8 shows HPLC chromatogram of labelled bovine serum albumin (BSA) using Z-FK-PAB-CHF 2 HMC, compound 27.
- BSA bovine serum albumin
- One nanomolar of human Cathepsin B was activated with 1 nM human thioredoxin.
- Fifty microliters of activated Cathepsin B was then added to 100 ⁇ of 1 .5 mM probe and incubated for 1 hour at 37°C in 30 mM acetate buffer at pH 5.5.
- Labelled BSA was derivatized with 3.0 mM NaCNBH 3 for 1 hour at 37°C.
- Fifty to twenty five microliter aliquots were injected in the HPLC, and resolved on a size-exclusion column (95% H 2 0, 5% MeOH).
- Experiments show trapping of the QM reporter molecule by BSA once probe is activated by Cathepsin B: (a) BSA alone, (b) BSA and 27, and (c)
- Figure 9 shows results obtained in a representative fluorescence microscopy experiment using (a) 1 ⁇ compound 32, and (b) 1 ⁇ compound 33;
- Figure 10 shows images of Her2 positive breast cancer cells treated with (a) compound 32, (b) APMA and compound 32, (c) control, 4',6-diamidino-2- phenylindole (DAPI), (d) control, APMA and DAPI, (e) compound 33, (f) APMA and compound 33, (g) control, DAPI, and (h) control, APMA and DAPI. Images were taken using cells fixed with 4% paraformaldehyde;
- Figure 11 shows fluorescence microscope images of breast cancer cells treated with (a) DAPI that stains the nucleus blue (first column), (b) compound 36 that is a green after activation by Cathepsin B, and (c) lysotracker red DND99® that stains the lysosomes.
- Compound 36 is activated in the lysosomes since the green fluorescence is clearly overlapping with the red lysotracker dye.
- breast cancer cells were treated with DAPI and lysotracker dye to demonstrate no green autofluorescence can be detected in cells.
- Figure 12 shows results obtained under conditions described from Figure 11 , with images taken at different time points and 20 X resolution;
- Figure 13 demonstrates that the probe was specific to Cathepsin B, cells were pretreated with CA-074Me overnight prior to compound 32.
- Figure 14(a) shows an HPLC after compound 38 (100 ⁇ ) was incubated in assay buffer (30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 1 .0 mM DTT, 10% DMSO) at 37 °C for 90 minutes.
- assay buffer (30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 1 .0 mM DTT, 10% DMSO
- a 20 ⁇ iL aliquot was injected into an HPLC fitted with a C8 column. The largest peak in the chromatogram at ⁇ 5 minute retention time is the intact probe clearly demonstrating chemical stability of the probe candidate.
- (b) In a second experiment, all conditions were identical to the first except 0.3 nM of activated Cathepsin B was included in the assay buffer. After 90 minutes, a 20 ⁇ xL aliquot was injected into the HPLC that showed a 40% reduction in the amount of compound 38 and new reaction product peaks formed
- imaging probes having a peptide portion that provides high affinity and specificity to CTB, a self-destructive linker that
- the prodrug linker p- aminobenzyl alcohol (PABA), originally reported by Katzenellenbogan,' 171 can be coupled to peptides through its amino group, allowing conjugation of aniline-based fluorophores to the alcohol via a carbamate linkage. [18, 191
- the evaluation of probe candidates typically determines the signal
- Z-Arg-Arg-AMC is typically used for assaying CTB activity in cell lysates.
- the recognition of doubly cationic substrates like Z-Arg-Arg-AMC by CTB has been attributed to a glutamic acid residue at the top of the S2 binding pocket. 1261 CTL lacks the equivalent Glu residue and prefers hydrophobic amino acids at the P2 position of substrates, explaining the selectivity of Z-Arg-Arg-AMC.
- Z-Lys-Lys-AMC (2) was prepared and evaluated as a substrate of CTB.
- Z-Phe-Arg-AMC and Z-Arg-Arg-AMC were both excellent substrates of CTB.
- CTL efficiently hydrolyzed Z-Phe-Arg-AMC but had 1 % of CTB's relative activity towards Z-Arg-Arg-AMC.
- Peptide 1 was rapidly hydrolyzed by both enzymes.
- compound 2 was an efficient substrate for CTB but a poor substrate for CTL suggesting that probes reliant on Z-Lys-Lys- may be selectively hydrolyzed by CTB. It was then determined how the PABA spacer affects recognition and turnover by CTB and CTL.
- V 1 has been successfully used to image herpes simplex virus thymidine kinase activity in humans.
- CA-074Me the cell permeable, irreversible CTB inhibitor CA-074Me was used to treat both cell lines. As shown in Figure 3, cells inhibited with either CA-074Me or E64d were unable to efficiently activate 4 clearly demonstrating high CTB specificity of this probe in live cancer cells. In contrast, CA-074Me did not fully abolish intracellular fluorescence indicating that 3 was hydrolyzed inside the CTB-inhibited cells.
- cysteine cathepsin inhibitor E64d prevented the onset of intracellular fluorescence suggesting that 3 is efficiently recognized by another cysteine cathepsin such as Cathepsins F or S since they are E64 sensitive and known to efficiently hydrolyze Z-Phe-Arg-AMC.
- cell lysates were prepared from HeLa cells and incubated in the presence of Z-Arg-Arg-AMC, 3 and 4 ( Figure 4). As predicted by the kinetic assays, probe 4 was the most efficient substrate. The relative hydrolysis of probe 3 was found to be similar to Z-Arg-Arg-AMC. To ensure that CA-074Me and E64d fully abolished the intracellular CTB activity and determine if the inhibited cells could turnover each probe, lysates were prepared from cells treated overnight with each inhibitor.
- Compound 3 was thus established to be an efficient and cell permeable substrate of CTB, not well tolerated by living cells, and activated by other cellular proteases.
- Compound 4 was found to be cell permeable, highly efficient probe capable of selectively detecting CTB activity in cell lysates and living cells.
- the agent provides a substrate moiety having high affinity to the target protease, CTB, a self-destructive linker, and QM reporter molecule that becomes chemically activated and immobilized only after its release.
- a self-destructive linker enables the attachment of reporter molecules to the probe via chemically and metabolically stable carbamates and ethers.
- Linkers can extend out of the enzyme's active site thus enabling the attachment of bulky reporter molecules maintaining low K m values, and release of the QM outside of the enzyme active site will preserve the catalytic activity of the target enzyme resulting in amplification of signal.
- the immobilized compound containing a coumarin moiety, is fluorescent, and was found to be retained in HeLa cells.
- the difluoro compound can be e.g., conveniently 18 F-labeled making it useful as a PET imaging agent. The stable and prolonged retention of the reporter molecule that results from
- immobilization by nucleophilic attack by e.g., nearby proteins permits sensitive molecular imaging studies of protease activity in living models of disease.
- the substrate moiety is a dipeptide covalently linked at its carboxy-terminus to a self-immolative linker which in turn is covalently linked to a molecule to be released.
- the linker can be based on PABA, or it can be the m-substituted equivalent, or it can be both p- and m-substituted i.e., include two immobilizing molecules.
- the dipeptide includes either a large hydrophobic (e.g., phenylalanine) or positively charged amino acid derivative (denoted AAx) followed by (S)-lysine (Cbz- AAx-Lys) or other positively charged amino acid having side chain -(CH 2 )nNH 2 in which n is 4 (lysine) or greater.
- n is 4, 5, 6, 7, 8, 9 or 1 0.
- the conjugate acid of the side-chain amino group has a pK a that is at least 5, so is positively charged in biological systems in which the agent is to be used i.e., in an environment having pH >5.
- Figure 6 illustrates a general structure of an agent of the invention, as discussed above.
- the linker portion illustrated is p-substituted, but it can be o- substituted or the benzene ring can be tri-substituted having -(CH 2 )OR 4 substituents at o- and p-positions with respect to the amino group.
- probes 32 and 33 are sufficiently cell permeable and efficiently activated by cathepsin B, and that compound 33 in particular has high specificity towards CTB in HeLa cells.
- APMA is known to increase expression of Cathepsin B through protease truncation of the Her2 receptor. Dramatic increases in probe activation was observed in APMA-treated cells for compounds 32 and 33. See Figure 10.
- the invention thus includes embodiments that are probes which rely on a Cbz-AAx-Lys substrate (targeting) moiety in combination with the PABA self- immolative linker and a latent or activatable phenolic reporter that spontaneously produces a chemically reactive electrophile.
- Amino acid derivative x (AAx) can thus be an amino acid derivative having either an aromatic or amino containing groups.
- Probe 34 utilizes a fluorine as the leaving group to produce a reactive quinone methide. While compounds 35 and 36 utilize p-nitroaniline and chlorotetrazine, respectively, as a simultaneous quencher group (that turns BODIPY fluorescence off until it departs) and the leaving group to form the quinone methide. Compound 37 is a control probe that is unable to form a reactive quinone methide. Of these compounds, 36 was found to be the most suitable for cell imaging
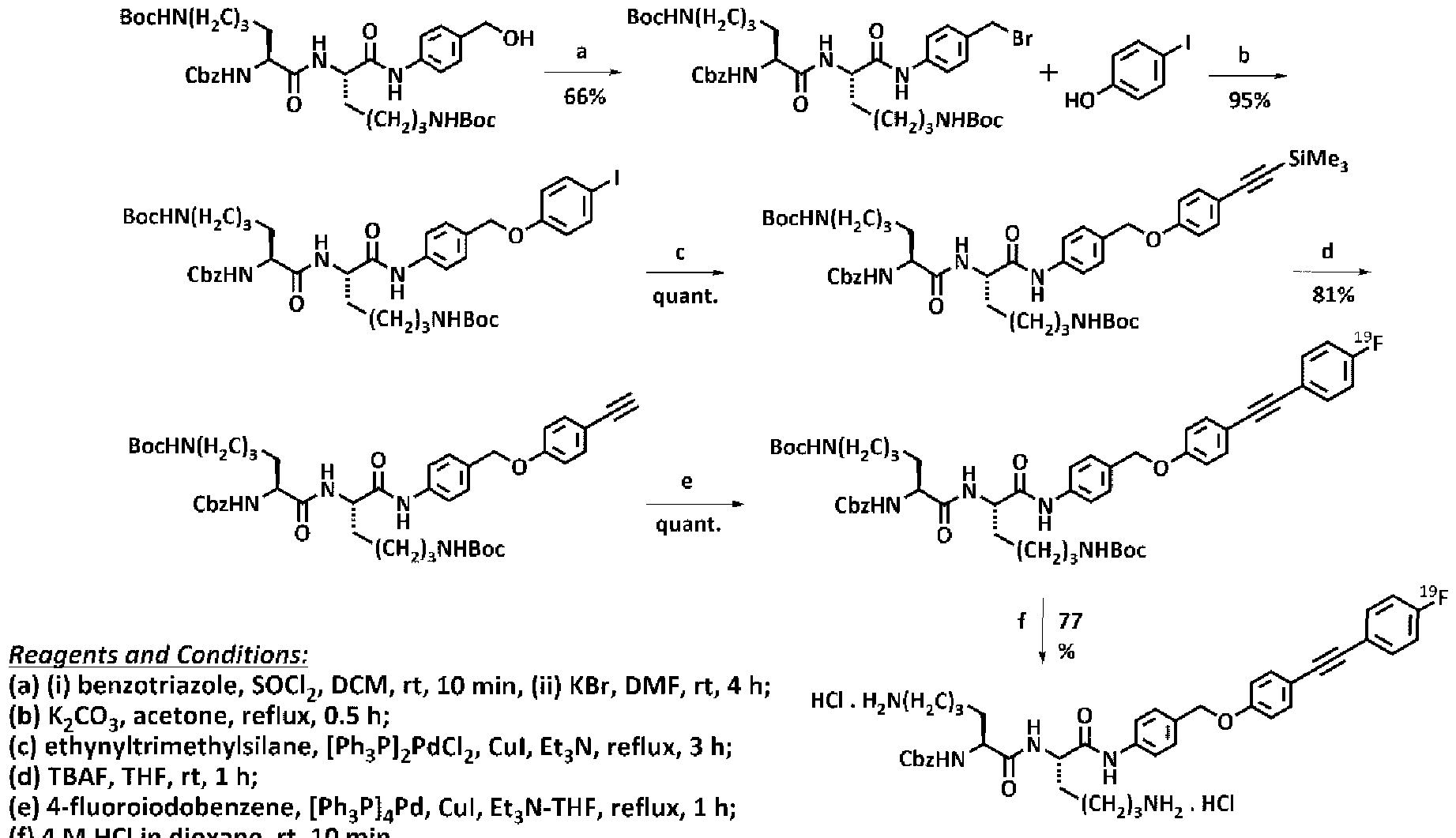
- the invention includes specific 19 F-labelled probes 38 and 39, radiolabeled so as to be useful as PET imaging agents:
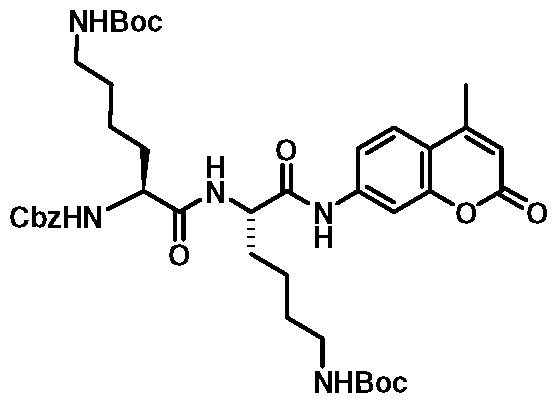
- Cbz-Lys-Lys-AMC (2) was easily prepared at gram scale levels using similar chemistry employed for the synthesis of 1 as shown in Scheme 1. Initially, a coupling reaction of Cbz-Lys-A/-£-Boc-Lys-A/-£-Boc-OH (11) which was available by the reaction of activated amino acid Cbz-Lys-A/-£-Boc-OSu (8) and H-Lys-A/-£-Boc- OH (10) [36] with 7-amino-4-methylcoumarin (AMC) (12) in presence of POCI 3 in pyridine at -1 5 °C following a similar literature procedure' 331 did not work.
- Reagents and conditions (a) NaHC0 3 , THF-H 2 0, rt, 1 6 h; (b) Et 3 N, DMF, 0 °C to rt, 16 h; (c) TFA-CH 2 CI 2 (1 :1 , v/v), ice-bath, 15 min.; (d) POCI 3 , pyridine, -1 5 °C, 1 h.
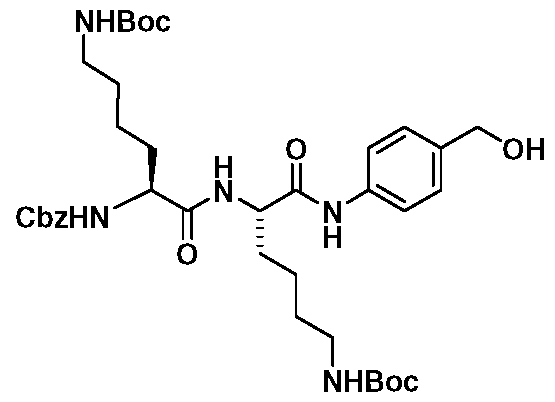
- Cbz-Lys-A/-£-Boc-Lys-A/-£-Boc-PAB-OH (18) was made by the reaction of 11 with PABA (14) using the same procedure as used for the synthesis of 15. Subsequent elaboration of 18 to carbamate Cbz-Lys-A/-£-Boc- Lys-A/-£-Boc-PABC-AMC (19) was made following the same strategy as used for the synthesis of 17.
- Reagents and conditions (a) NaHC0 3 , THF-H 2 0, rt, 16 h; (b) THF, EEDQ, rt, 16 h; (c) 15% phosgene in toluene, ⁇ 20 o C, 16 h; (d) THF, 80 °C, 2 h; (e) TFA-CH 2 CI 2 (1 :1 , v/v), ice-bath, 15 min; (f) Methanolic-HCI (0.5M), rt, 16 h; (g) Et 3 N, DMF, 0 °C to rt, 16 h.
- Tripeptide bromides Cbz-Phe-Lys-/V-£-Boc-PAB-Br (20) and Cbz-Lys-A/-£- Boc-Lys-/V-£-Boc-PAB-Br (21 ) were efficiently made from the corresponding alcohols 15 and 18, respectively by treating with KBr and SOCI 2 in presence of benzotriazole in DMF.
- Reagents and conditions (a) i) benzotriazole, SOCI 2 , CH 2 CI 2 , rt, 5 min; ii) KBr, DMF, rt, 4 h; (b) 22, acetone, K 2 C0 3 , reflux, 0.5 h; (c) DAST, CH 2 CI 2 , rt, 16 h; (d) TFA- CH 2 CI 2 (1 :1 , v/v), ice-bath, 15 min; (e) 29, DMF, K 2 C0 3 , rt, 16 h. Synthetic schemes for compounds 38 and 39 are shown in Schemes 4 and 5 respectively.
- TFA (3.TFA): To a solution of trifluoroacetic acid - DCM (1 :1 v/v, 2 ml_) at ice-bath temperature was added 17 (200 mg, 0.24 mmol) and stirred for 15 min. After reaction, diethyl ether was added to the reaction mixture to precipitate out the solids. The solids were centrifuged out and successively washed with diethyl ether (5 ml_ x 2) and ethyl acetate (5 ml_ x 2), and dried under high vacuum in dark to get the pure title compound 3.TFA. White solid, 204 mg (100%).
- HCI (3.HCI) A solution of 17 (100 mg, 0.12 mmol) in methanolic-HCI (0.5M, 5 ml_) was stirred at room temperature for 16 h. After reaction, solvents were removed in vacuo to dryness and the residue was taken in diethyl ether and stirred for 10 min. The suspension was centrifuged, the supernatant was discarded, and the process was repeated twice to get the pure title compound 3.HCI. White solid, 80 mg (87%).
- concentrations of fluorogenic peptide were varied between ⁇ 0.5 KM - 5 KM while maintaining a constant concentration of activated enzyme.
- the reaction mixture (150 ⁇ final volume) consisted of 30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 2.0 mM DTT, 10% DMSO, and 0.33 to 3.3 nM human cathepsin B or L in 96 well black plates. The samples were pre-warmed at 37°C for 30 min, and the reaction was initiated upon addition of the enzyme. The activity was monitored spectrophotometrically for the release of 7-amino-4-methyl-coumarin (AMC;
- the amount of AMC released from the reaction was determined by a standard curve in assay buffer. Incubation of the prodrug inspired probes in the assay buffer without enzyme demonstrated that the probes were stable for at least 6 hours with very little spontaneous hydrolysis of AMC as monitored by fluorescence or HPLC (data not shown).
- the assay to quantify active enzyme by titration with E64 was adapted from Barrett et a/.
- human cathepsin enzyme (-1 0 nM) in assay buffer (30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 1 .0 mM DTT) was inactivated with E64 (final concentrations ranging from 1 .5 to 7.0 nM) for 20 min at 37 °C.
- E64 final concentrations ranging from 1 .5 to 7.0 nM
- an aliquot of this mixture was added to a solution containing the substrate (30 ⁇ z-RR-AMC or z-FK- AMC in 10% DMSO 90% assay buffer) at 37 °C.
- the residual enzymatic activity was monitored spectrophotometrically for the release of 7-amino-4-methyl-coumarin and plotted versus the concentration of E64 to graphically determine the amount of active enzyme.
- the cell lysates and assay were adapted from Giusti et a/. [46] . Briefly, cells were grown to ⁇ 80% confluency and washed with Phenol Red free DMEM, treated with lyse buffer (10 mM Tris-HCI, pH 7.5, 1 50 mM NaCI, 1 % Triton X-100, 1 mM EDTA) and centrifuged (12,000 rpm) in a 1 .5 ml_ tube.
- 100 ⁇ _ of the lysis solution was added to a black 96 well plate and diluted with 100 ⁇ _ activation buffer (100 mM Acetate, pH 5.5, 5 mM DTT, 5 mM EDTA) followed by 15 minute incubation at 37 °C.
- 100 ⁇ _ activation buffer 100 mM Acetate, pH 5.5, 5 mM DTT, 5 mM EDTA
- a 100 ⁇ _ aliquot of the activated lysate was then added to a 50 ⁇ _ solution containing 300 ⁇ probe, in 30% DMSO and 70% buffer (100 mM acetate buffer, 5 mM EDTA, pH 5.5).
- the enzymatic reaction proceeded at 37 °C for 1 hour after which 50 ⁇ _ was withdrawn and added to 100 ⁇ _ of sodium choloroacetate (200 mM) to stop the reaction.
- the fluorescence at 460 nm was measured and normalized to total protein concentration (Bradford Assay) and expressed as a percentage of AMC released from experiments using Z-Arg-Arg- AMC.
- spontaneous AMC hydrolysis was determined in assay buffer without lysate while hydrolysis by other protease classes was assessed by pre- treating lysates with sodium chloroacetate (200 mM in assay buffer). Levels of free AMC were similar in both control reactions and therefore used as a blank which was subtracted from the total RFU in each experiment.
- MDA MB 231 breast cancer cells grown on cover slips were treated with PBS buffer (control) or compound 36 for 4h at 37°C. Results are shown in Figures 6 to 8. The compound turns green after Cathepsin B hydrolysis. Cells were then treated with DAPI (blue, to stain nucleus) and lysosome tracker (red) and incubated for another 30 minutes. Cells were then washed with PBS fixed with paraformaldehyde onto slides and fluorescence microscopy images were taken at 100X. The probe activation is clearly visualized in the lysosomes of cells.
- compound 38 a non-radioactive version of a potential Cathepsin B PET probe, in presence and absence of Cathepsin B was examined using HPLC.
- a fluorinated probe candidate In order to visualize Cathepsin B activity in vivo using PET, a fluorinated probe candidate must be an efficient substrates of the enzyme.
- compound 38 (100 ⁇ ) was incubated in assay buffer (30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 1 .0 mM DTT, 10% DMSO) at 37 °C for 90 minutes. A 20 ⁇ aliquot was injected into an HPLC fitted with a C8 column and detected at 254 nm.
Abstract
Compounds that are cathepsin B substrates and that are useful as imaging probes, for example in positron emission tomography (PET) or fluorescence imaging are described.
Description
CATHEPSIN B-TARGETING PROBES
Field
The present invention relates to compounds that are cathepsin B substrates. The compounds find particular use as imaging probes, for example in positron emission tomography (PET).
Background
The spread of cancer from a primary tumor to a secondary location, termed metastasis, is the cause of death in 90% of cancer patients.'11 A complex system of proteases called the proteolytic network degrades the extracellular matrix (ECM) components producing a permissive region for cancer cells to invade. [2, 31 Identifying legitimate therapeutic and diagnostic targets is challenging due to the number of different protease families operating in the tumor/tumor environment, the variety of tumor associated cell types which express and secrete proteases, and the numerous modes of enzyme regulation.
Cathepsin B (CTB, EC 3.4.22.1 ) is produced by a variety of tumor-associated cells and is a prominent member of the tumor promoting proteolytic network.'3"51 Mounting evidence suggests that CTB may be a potential diagnostic and prognostic biomarker for various cancers.'6"101 Interestingly, CTB appears to affect cancer progression dependent upon its localization with intracellular CTB involved in amplification of apoptosis.1111 In contrast, aggressive cancers have high extracellular CTB activity at the invading edge of a tumor and in the local ECM secreted by a variety of cell types.1121 Recent reports suggest CTB, together with the closely related enzyme cathepsin L (CTL, EC 3.4.22.15), may protect against anticancer chemotherapies such as Taxol, meaning CTB and/or CTL activity may be a marker to predict response to chemotherapy.1131 Accurate and sensitive tools capable of assessing CTB's activity in cell culture, animal models of aggressive cancer or cancer patients are needed to validate this protease as a high priority cancer marker or therapeutic target.
Discovering probes with sufficient membrane permeability, appropriate cellular localization, and rapid activation kinetics is a substantial task. Even more challenging is designing probes that are specific to a single protease due to the
substrate overlap often observed between related and unrelated proteases. Blum et al. have developed elegant mechanism-based optical probes that covalently label CTB's active site but unfortunately lack specificity since these probes were efficiently recognized by CTL.[14] The commercially available probe (Z-Arg-Arg)2-cresyl violet is an apparently CTB-specific substrate used for live cell studies'151 but fluorogenic intermediates of the hydrolysis reaction make it inappropriate for kinetic studies.'161 Efforts here focused on CTB-specific fluorescent probes suitable for kinetic studies and live cell imaging; ideally relatively simple, cell-permeable compounds, particularly peptide-based derivatives.
Summary
One object of the invention is to provide peptide-based agents that can be cleaved by cathepsin B. Preferred compounds yield a fluorophore and/or a PET imaging agent upon cleavage by cathepsin B become that then becomes
immobilized in a cellular milieu. Certain of the compounds are fluorescent while others can be labeled subsequent to immobilization.
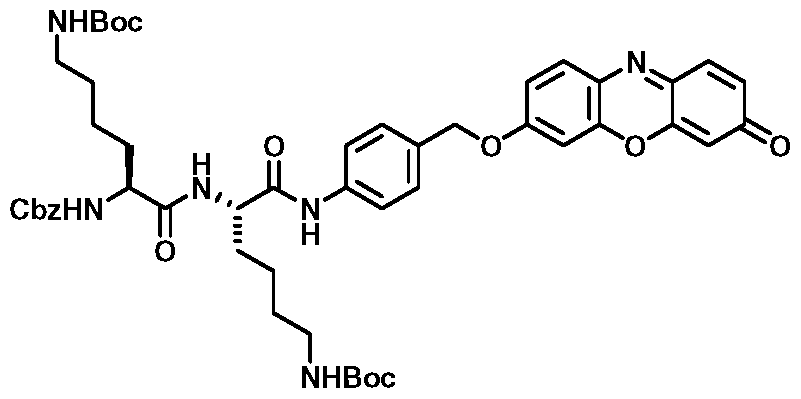
A compound of the invention has formula (I):
wherein:
R1 is PhCH20-, CH3 or PhCH2-;
R2 is PhCH2-, (CH3)2CH- or H2N(CH2)m- for m > 4;
R3 and R3' are each H or -CH2OR4, and at least one of R3 and R3'
is -CH2OR4,wherein for each of R3 and R3 , R4 is independently selected from the group consisting of:
wherein:
R5 is H, -C(0)H, -CHX2, wherein each X is, independently of the other
X, selected from the group consisting of:
H, F, CI, Br, I, -N02, toluenesulfonate, methanesulfonate, trifluoromethansulfonate, perfluorobutanesulfonate,
ethanesulfonate, benzenesulfonate,
parachlorobenzenesulfonate, nitrobenzenesulfonate or methoxybenzenesulfonate, and both X are not simultaneously
H,
wherein:
R7 is H, -C≡CH, N3, -C(0)H, or -CHX2, wherein each X is, independently of the other X, selected from the group consisting of:
H, F, CI, Br, I, -N02, toluenesulfonate, methanesulfonate, trifluoromethansulfonate, perfluorobutanesulfonate,
ethanesulfonate, benzenesulfonate,
parachlorobenzenesulfonate, nitrobenzenesulfonate or methoxybenzenesulfonate, and both X are not simultaneously H, and
wherein each of R9 and R10 is, independently of the other, H,
N3, -C(0)H, -CHX2, wherein each X is, independently of the other X, selected from the group consisting of:
H, F, CI, Br, I, -N02, toluenesulfonate, methanesulfonate, trifluoromethansulfonate, perfluorobutanesulfonate, ethanesulfonate, benzenesulfonate, parachlorobenzenesulfonate,
nitrobenzenesulfonate or methoxybenzenesulfonate, and both X are not simultaneously H,
-CH2Y, wherein Y is selected from the group consisting of:
-OH, -OC(O)NHAr1 , wherein Ar1 is -C6H5 in which 1 , 2 or 3 hydrogen atoms is optionally and independently substituted by -NO2, and
N=N
_ />— z
N N , where Z is selected from the group consisting of H, CI, and Br.
a fluorescent dye,
a radiolabeled substituent,
or -C≡CR11 wherein R1 1 is H or Ar2, in which Ar2 is -C6H5 in which 1 , 2 or 3 hydrogen atoms is optionally and independently substituted by F, and
R9 and R10 are not simultaneously H,
n > 4,
including salts, hydrates and solvates thereof.
In certain embodiments, the compound includes fluorescent dye i.e., a fluorescent dye molecule is incorporated into the compound to form a part thereof, as exemplified herein. It is understood that the dye is covalently linked to the remainder of the compound. Such dyes include
wherein one or more hydrogens bound to a carbon atom of the dye is optionally substituted with a methyl group. The structure on the left represents the
borondipyrromethene (BODIPY) core. A BODIPY portion of the compound can be covalently linked at the carbon atom of the dye located at the para-position with respect to the boron atom.
In embodiments, the radiolabeled substituent comprises an 18F atom. The radiolabeled substituent can be a C1 -C20 alkyl group or C5-C8 aromatic group in which the aromatic ring is made up of carbon atoms or carbon and nitrogen atoms. The ring is optionally substituted with a C1 -C20 alkyl or alkoxy group that is optionally substituted with said 18F atom(s), and/or the ring bears one or more 18F.
The radiolabeled substituent can be selected from the roup consisting of:
Where Y can be (CH2)m OR 0(CH2)m with m = 1 ,
Where X can be C N 2, 3, 4 5, 6
In embodiments, R3 is the same as R3 , or they are different.
The value of m and n are each typically 4, 5, 6, 7, 8, 9 or 10. The values of m and n are independent of each other.
In embodiments, the invention is a composition for use as an imaging probe, the composition including at least one compound having formula (I) as described above.
In embodiments, the invention is a pharmaceutical composition for use in PET and/or fluorescence imaging, the composition including at least one compound having formula (I) as described above. The composition can be one in which the compound(s) is dispersed or dissolved in a liquid medium suitable for injection.
The invention includes a method of PET and/or fluorescence imaging. The method includes administering to a patient in need thereof an effective amount of a composition(s) as described above, and scanning the subject with at PET and/or fluorescence imaging device. Such embodiments can be limited to a diagnostic method.
The invention includes method for studying the localization of PET probes within a tissue of a subject comprising: administering to the subject an effective amount of a compound(s) as described above, subjecting a tissue of the subject to irradiation of an electromagnetic radiation, wherein the electromagnetic radiation is absorbed by the compound, detecting fluorescence of the compound within the tissue, wherein the fluorescence of the compound within the tissue is indicative of the presence of a PET probe within the tissue.
Various embodiments of the invention are discussed throughout this specification. Any embodiment discussed with respect to one aspect of the invention applies to other aspects of the invention as well and vice versa. The embodiments in the detailed examples are understood to be embodiments of the invention that are applicable to all aspects of the invention.
The use of the word "a" or "an" when used in conjunction with the term "comprising" in the claims and/or the specification may mean "one," but it is also consistent with the meaning of "one or more," "at least one," and "one or more than one."
The term "about" is used to indicate that a value includes the standard deviation of error for the device or method being employed to determine the value.
The use of the term "or" in the claims is used to mean "and/or" unless explicitly indicated to refer to alternatives only or the alternatives are mutually exclusive.
As used in this application, the words "comprising", "having", "including" or "containing" and any of their forms, are inclusive or open-ended and do not exclude additional, unrecited elements or method steps.
It should be noted here, that when discussing radical portions of a molecule, such as "-R", "R", etc., the connecting bond may or may not be included in various contexts for the sake of convenience, and the skilled person understands this.
Other objects, features and advantages of the present invention will become apparent from the following detailed description. It should be understood, however, that the detailed description and the specific examples, while indicating specific embodiments of the invention, are given by way of illustration only, since various
changes and modifications within the spirit and scope of the invention will become apparent to those skilled in the art from this detailed description.
Brief Description of the Drawings
Some embodiments of the invention are described, by way of example, with reference to the accompanying drawings and figures. The particulars shown are by way of example and purposes of illustrative discussion of embodiments of the invention. The description taken with drawings and figures makes apparent to the skilled person how embodiments of the invention can be practiced.
Figure 1 illustrate the general strategy for obtaining probes of the invention;
Figure 2(b) is a schematic representation of a probe employing PABA extend into the CTB S2' recognition site, and Figure 2(c) shows two fluorogenic peptides of the invention;
Figure 3 shows fluorescent microscope images taken of HeLa cells treated for 2 hours with a) 40 μΜ of (3) and b) 40 μΜ of (4). Cells were treated overnight with CA-074Me and E64d followed by incubation with each probe for 2 hours. White bars represent 50 μηι;
Figure 4 is a bar graph showing results obtained with HeLa cell lysates incubated with Z-Arg-Arg-AMC, compounds 3 and 4 using untreated cell lysates and cell lysates prepared from cells treated with CA-074Me or E64D. Bars represent the relative activity (%) compared to Z-Arg-Arg-AMC incubated with untreated lysates adjusted to total protein concentration minus background RFU with error bars depicting standard deviation from 4 independent trials;
Figure 5 illustrates a mechanism of cleavage of a particular agent by CTB followed by immobilization of a compound produced by said cleavage;
Figure 6 illustrates a general structure of an agent showing the CTB-targeting moiety and immobilizing compound, R4, connected to the targeting moiety via a linker;
Figure 7 shows results obtained in a representative fluorescence microscopy experiment using the agent shown in Figure 5, to demonstrate the retention of the immobilized reporter molecule once activated by cathepsin B;
Figure 8 shows HPLC chromatogram of labelled bovine serum albumin (BSA) using Z-FK-PAB-CHF2 HMC, compound 27. One nanomolar of human Cathepsin B
was activated with 1 nM human thioredoxin. Fifty microliters of activated Cathepsin B was then added to 100 μΙ of 1 .5 mM probe and incubated for 1 hour at 37°C in 30 mM acetate buffer at pH 5.5. Labelled BSA was derivatized with 3.0 mM NaCNBH3 for 1 hour at 37°C. Fifty to twenty five microliter aliquots were injected in the HPLC, and resolved on a size-exclusion column (95% H20, 5% MeOH). Experiments show trapping of the QM reporter molecule by BSA once probe is activated by Cathepsin B: (a) BSA alone, (b) BSA and 27, and (c) 1 .0 mM 27;
Figure 9 shows results obtained in a representative fluorescence microscopy experiment using (a) 1 μΜ compound 32, and (b) 1 μΜ compound 33;
Figure 10 shows images of Her2 positive breast cancer cells treated with (a) compound 32, (b) APMA and compound 32, (c) control, 4',6-diamidino-2- phenylindole (DAPI), (d) control, APMA and DAPI, (e) compound 33, (f) APMA and compound 33, (g) control, DAPI, and (h) control, APMA and DAPI. Images were taken using cells fixed with 4% paraformaldehyde;
Figure 11 shows fluorescence microscope images of breast cancer cells treated with (a) DAPI that stains the nucleus blue (first column), (b) compound 36 that is a green after activation by Cathepsin B, and (c) lysotracker red DND99® that stains the lysosomes. Compound 36 is activated in the lysosomes since the green fluorescence is clearly overlapping with the red lysotracker dye. In a control experiment breast cancer cells were treated with DAPI and lysotracker dye to demonstrate no green autofluorescence can be detected in cells.
Figure 12 shows results obtained under conditions described from Figure 11 , with images taken at different time points and 20 X resolution;
Figure 13 demonstrates that the probe was specific to Cathepsin B, cells were pretreated with CA-074Me overnight prior to compound 32.
Figure 14(a) shows an HPLC after compound 38 (100 μΜ) was incubated in assay buffer (30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 1 .0 mM DTT, 10% DMSO) at 37 °C for 90 minutes. A 20 \iL aliquot was injected into an HPLC fitted with a C8 column. The largest peak in the chromatogram at ~ 5 minute retention time is the intact probe clearly demonstrating chemical stability of the probe candidate, (b) In a second experiment, all conditions were identical to the first except 0.3 nM of activated Cathepsin B was included in the assay buffer. After 90 minutes, a 20 \xL
aliquot was injected into the HPLC that showed a 40% reduction in the amount of compound 38 and new reaction product peaks formed.
Detailed Description
In one particular aspect, imaging probes having a peptide portion that provides high affinity and specificity to CTB, a self-destructive linker that
spontaneously releases a reporter upon its enzymatic deprotection and a latent fluorophore which becomes highly fluorescent once enzymatically freed from the intact probe. This is illustrated generally in Figure 1. The prodrug linker p- aminobenzyl alcohol (PABA), originally reported by Katzenellenbogan,'171 can be coupled to peptides through its amino group, allowing conjugation of aniline-based fluorophores to the alcohol via a carbamate linkage. [18, 191
The evaluation of probe candidates typically determines the signal
amplification after enzymatic activation as well as crude specificity studies by comparing the relative rates of probe turnover by competing proteases. Although these characteristics are critical in evaluating probe performance in vitro, such studies are largely qualitative making it difficult to compare lead probes to known substrates and existing imaging agents. In contrast, the specificity constant kcat/KM can be used quantitatively to rank structurally related probe candidates. In addition, the efficiency of lead compounds is evaluated by direct comparison to known substrates of the target protease and specificity assessed by comparing turnover rates by related proteases. Kinetic parameters may better predict probe performance in cellulo and possibly in vivo using benchmark kinetic values established by existing enzyme imaging agents used in humans.'20, 211
Benchmark kinetic values of M, /(cat and kcat/KM for a series of two-component fluorogenic peptides using recombinant CTB and the closely related CTL were established. Previous studies showed that carboxybenzyl (Z)-protected fluorogenic dipeptides like Z-Phe-Arg-AMC are efficient substrates of CTB and CTL.[22] It is known that CTB and CTL prefer an aromatic group at P3 (corresponding to pocket S3 in Figure 2(b), a large hydrophobic amino acid at P2 (S2 in Figure 2(b) and a positively charged substituent at the P1 position (S1 in Figure 2(b)). [23] The compound Z-Phe-Lys-AMC (1) was thus prepared and characterized as substrate for
each of CTB and CTL. Although the synthesis of 1 was recently described'241 no kinetic values for this compound as a substrate for CTB or CTL are known to have been reported.
Issues of substrate specificity are common with two-component fluorogenic peptides such as Z-Phe-Arg-AMC due to efficient recognition by serine proteases.'251 However, Z-Arg-Arg-AMC is typically used for assaying CTB activity in cell lysates. The recognition of doubly cationic substrates like Z-Arg-Arg-AMC by CTB has been attributed to a glutamic acid residue at the top of the S2 binding pocket.1261 CTL lacks the equivalent Glu residue and prefers hydrophobic amino acids at the P2 position of substrates, explaining the selectivity of Z-Arg-Arg-AMC. Z-Lys-Lys-AMC (2) was prepared and evaluated as a substrate of CTB.
PABA derived prodrugs have been seen to be efficiently processed by CTB to release a cytotoxic drug.1271 This suggests that PABA can be tolerated at ST. In addition, three-component peptides may offer higher specificity towards CTB because they occupy each of the core substrate binding positions S3 to S2' (Figure 2(b).1281 To evaluate the efficiency of CTB probes bearing the PABA spacer, Z-Phe- Lys-PABA-AMC (3) and Z-Lys-Lys-PABA-AMC (4) compound I in which) (Figure 2(c)) were prepared. It is noted here that 3 is compound (I) in which R1 = PhCH20-, R2 = PhCH2-, R3 = -CH2OR4 in which R4 is:
R3 = H, and n = 4: and compound 4 is compound (I) in which R1 = PhCH20-, R2 = H2N(CH2)m-, R3 = -CH2OR4 in which R4 is:
R3' = H, m = 4, and n = 4.
As shown in Table 1 , Z-Phe-Arg-AMC and Z-Arg-Arg-AMC were both excellent substrates of CTB. In contrast, CTL efficiently hydrolyzed Z-Phe-Arg-AMC but had 1 % of CTB's relative activity towards Z-Arg-Arg-AMC. Peptide 1 was rapidly
hydrolyzed by both enzymes. In agreement with the data obtained for Z-Arg-Arg- AMC, compound 2 was an efficient substrate for CTB but a poor substrate for CTL suggesting that probes reliant on Z-Lys-Lys- may be selectively hydrolyzed by CTB. It was then determined how the PABA spacer affects recognition and turnover by CTB and CTL. The introduction of PABA into 3 and 4 resulted in a lower apparent M value for CTB when compared to their corresponding two component peptides. This indicates that PABA results in higher affinity towards CTB and may improve probe performance in cellulo. The lower apparent KM value for 3 was partially offset by a lower /(cat, but an impressive specificity constant was maintained with kcat/KM = 61 mM"1 s"1 for CTB. Encouragingly, 4 sustained rapid turnover resulting in the highest specificity constant of all novel CTB substrates with kcat/KM = 231 mM"V1. For comparison purposes, 18F-FHBG with a kcat/kM = 43 mM"V1 has been successfully used to image herpes simplex virus thymidine kinase activity in humans. CTL recognition of 3 suffered dramatically as no reliable kinetic data were obtained by increasing the CTL concentration to 10 nM, and 4 was also a very poor substrate of CTL as expected from the data obtained from Z-Arg-Arg-AMC and 2.
TABLE 1
Kinetic parameters obtained for fluorogenic peptides as substrates of CTB and CTL
Cathepsin B Cathepsin L
cat M cat M
Probe (mivrV) (mMV)
Z-Phe-
0.09 ± 0.01 36 ± 2 400 ± 70 1 .52 ± 0.2 x 10 3 0.38 ± 0.01 256 ± 36 Arg-AMC
Z-Arg-
0.34 ± 0.06 56 ± 3 165 ± 37 0.024 ± 0.005 0.22 ± 0.01 9.2 ± 2.3 Arg-AMC
(1 ) 0.19 ± 0.03 26 ± 2 137 ± 13 0 59 ± °3 1 2 x 1 0 0.39 ± 0.01 661 ± 149
(2) 0.25 ± 0.01 37 ± 1 148 ± 5 0.092 ± 0.001 0.14 ± 0.06 1 .5 ± 0.6 <3> "fSt β1 ± 7 n.d.
(4) 0.16 ± 0.04 37 ± 2 231 ± 70 0.043 ± 0.01 0.1 1 ± 0.01 2.5 ± 0.8
All values were determined by an average of three trials ± standard deviation. Where indicated by n.d. no reliable kinetic data could be obtained using concentration of enzyme as high as 10 nM.
The activation of 3 and 4 could be visualized using fluorescent microscopy inside living HeLa cervical cancer cells'291 (see Figure 3). In agreement with enzyme kinetic data, 4 was the superior agent as strong fluorescence intensity was visible within 30 minutes of incubation. Similar results were observed when a Her2-positive breast cancer cell line derived from MD-MBA-231 called H2N[30] was treated with 3 and 4 (results not shown). Cell lines tested appeared to tolerate 4 based on the lack of observed changes in cell morphology, but HeLa and H2N cells incubated with 3 underwent rapid morphological changes reminiscent of cell death.
To evaluate the specificity of probe candidates towards CTB, the cell permeable, irreversible CTB inhibitor CA-074Me was used to treat both cell lines. As shown in Figure 3, cells inhibited with either CA-074Me or E64d were unable to efficiently activate 4 clearly demonstrating high CTB specificity of this probe in live cancer cells. In contrast, CA-074Me did not fully abolish intracellular fluorescence indicating that 3 was hydrolyzed inside the CTB-inhibited cells. However, the broad spectrum cysteine cathepsin inhibitor E64d prevented the onset of intracellular fluorescence suggesting that 3 is efficiently recognized by another cysteine
cathepsin such as Cathepsins F or S since they are E64 sensitive and known to efficiently hydrolyze Z-Phe-Arg-AMC.[31]
To determine if the enzyme kinetic values accurately predicted probe performance in a complex mixture, cell lysates were prepared from HeLa cells and incubated in the presence of Z-Arg-Arg-AMC, 3 and 4 (Figure 4). As predicted by the kinetic assays, probe 4 was the most efficient substrate. The relative hydrolysis of probe 3 was found to be similar to Z-Arg-Arg-AMC. To ensure that CA-074Me and E64d fully abolished the intracellular CTB activity and determine if the inhibited cells could turnover each probe, lysates were prepared from cells treated overnight with each inhibitor. CTB activity towards either Z-Arg-Arg-AMC or 4 was not observed in lysates from inhibited cells, demonstrating both inhibitors efficiently inactivated CTB and that probe 4 was highly specific towards this enzyme. In contrast, cell lysates treated with CA-074Me maintained about 35% of the original enzymatic activity towards 3 indicating hydrolysis by e.g., a competing proteases. In agreement with the live cell assays, lysates from HeLa cells treated with E64d or the cysteine protease inhibitor sodium chloroacetate were unable to hydrolyze 3 also indicating that the competing proteases are likely cysteine cathepsin enzymes.
Compound 3 was thus established to be an efficient and cell permeable substrate of CTB, not well tolerated by living cells, and activated by other cellular proteases. Compound 4 was found to be cell permeable, highly efficient probe capable of selectively detecting CTB activity in cell lysates and living cells.
The feasibility of use of a reactive quinone methide (QM) reporter molecule released after cleavage by CTB has also been established. This approach is illustrated in Figure 5. Compound 27 was prepared, 27 being a compound of formula (I) in which R1 = PhCH20-, R2 = PhCH2-, n=4, R3' = H, and R3 = -CH2OR4 in which R4 is:
and R5 is -CHF2 and R6 is H. Here, the agent provides a substrate moiety having high affinity to the target protease, CTB, a self-destructive linker, and QM reporter
molecule that becomes chemically activated and immobilized only after its release. Use of the self-destructive linker enables the attachment of reporter molecules to the probe via chemically and metabolically stable carbamates and ethers. Linkers can extend out of the enzyme's active site thus enabling the attachment of bulky reporter molecules maintaining low Km values, and release of the QM outside of the enzyme active site will preserve the catalytic activity of the target enzyme resulting in amplification of signal.
In this example, the immobilized compound, containing a coumarin moiety, is fluorescent, and was found to be retained in HeLa cells. The difluoro compound can be e.g., conveniently 18F-labeled making it useful as a PET imaging agent. The stable and prolonged retention of the reporter molecule that results from
immobilization by nucleophilic attack by e.g., nearby proteins, permits sensitive molecular imaging studies of protease activity in living models of disease.
The substrate moiety is a dipeptide covalently linked at its carboxy-terminus to a self-immolative linker which in turn is covalently linked to a molecule to be released. The linker can be based on PABA, or it can be the m-substituted equivalent, or it can be both p- and m-substituted i.e., include two immobilizing molecules. The dipeptide includes either a large hydrophobic (e.g., phenylalanine) or positively charged amino acid derivative (denoted AAx) followed by (S)-lysine (Cbz- AAx-Lys) or other positively charged amino acid having side chain -(CH2)nNH2 in which n is 4 (lysine) or greater. Preferably, n is 4, 5, 6, 7, 8, 9 or 1 0. In all of these cases, the conjugate acid of the side-chain amino group has a pKa that is at least 5, so is positively charged in biological systems in which the agent is to be used i.e., in an environment having pH >5.
Figure 6 illustrates a general structure of an agent of the invention, as discussed above. The linker portion illustrated is p-substituted, but it can be o- substituted or the benzene ring can be tri-substituted having -(CH2)OR4 substituents at o- and p-positions with respect to the amino group.
Experiments using HeLa cancer cells demonstrated that probe 27 is sufficiently cell permeable and efficiently activated by cathepsin B. Results are illustrated in Figure 7. It was thus demonstrated that the fluorescence was not washed out of cells while the control probe bearing 7-hydroxy-4-methyl coumarin
and incapable of chemical immobilization was readily wash out of the cells. In the experiments, the control probe was identical to the test compound but lacked the -CF2 group (i.e. 7-hydroxy-4-methyl coumarin). Cathespin B processing released a fluorophore incapable of chemically immobilizing inside the cells and was therefore readily washed out. Further evidence of the feasibility of this approach is given by the results of Figure 8 which show trapping of the QM reporter derived from molecule 27 by BSA once the probe is activated by Cathepsin B.
Compounds 32 and 33, probes from which resorufin is generated upon protease cleavage were prepared. Experiments using HeLa cancer cells
demonstrated that probes 32 and 33 are sufficiently cell permeable and efficiently activated by cathepsin B, and that compound 33 in particular has high specificity towards CTB in HeLa cells. See Figure 9. APMA is known to increase expression of Cathepsin B through protease truncation of the Her2 receptor. Dramatic increases in probe activation was observed in APMA-treated cells for compounds 32 and 33. See Figure 10.
The invention thus includes embodiments that are probes which rely on a Cbz-AAx-Lys substrate (targeting) moiety in combination with the PABA self- immolative linker and a latent or activatable phenolic reporter that spontaneously produces a chemically reactive electrophile. Amino acid derivative x (AAx) can thus be an amino acid derivative having either an aromatic or amino containing groups.
Four probes cathepsin B probes 34-37 the incorporate the fluorophore BODIPY were produced:
34: R = F
o
35: R = 0^—
36: R = °— /)— c
37: R = OH
Compounds 34, 35 and 36, once cleaved by Cathepsin B (Figure 5), the PABA decomposes to release BODIPY which then decomposes into a reactive quinone methide through spontaneous elimination of "HR". In the case of compound 37, the quinone methide does not form.
Probe 34 utilizes a fluorine as the leaving group to produce a reactive quinone methide. While compounds 35 and 36 utilize p-nitroaniline and chlorotetrazine, respectively, as a simultaneous quencher group (that turns BODIPY fluorescence off until it departs) and the leaving group to form the quinone methide. Compound 37 is
a control probe that is unable to form a reactive quinone methide. Of these compounds, 36 was found to be the most suitable for cell imaging
The invention includes specific 19F-labelled probes 38 and 39, radiolabeled so as to be useful as PET imaging agents:
In the case of compound 38, lacking the -CH2F group, a reactive electrophile is not generated upon cleavage of the compound by Cathepsin B. The probe is thus a non-immobilizing agent. Cleavage of compound 39 by Cathepsin B produces a PABA intermediate in analogy to that shown in Figure 5, which decomposes to a methide electrophile that can undergo nucleophilic attack by e.g., nearby proteins and become immobilized.
Methods
The synthesis of peptides 1 - 4, compound characterization, the conditions for the enzyme assays and the protocol to assay the HeLa cell lysates can be found in the Supporting Information. For the fluorescence microscopy experiments, -100,000 cells were seeded in a six well plate cultured in Dulbecco's modified Eagle medium (DMEM) with 1 0% fetal bovine serum (FBS) and 1 % antibiotics-antimycotics incubated at 37 °C in a 5% CO2 atmosphere. The cells were washed once with PBS and probes 3 and 4 (40 μΜ dissolved in Phenol Red free DMEM) were added and
incubated for 2 hours at 37 °C. Prior to florescence imaging the cells washed twice with PBS and the media replaced with Phenol Red free DMEM. Fluorescence was visualized and digitally captured using an inverted fluorescent microscope (Zeiss Axiovert 200, filter set with excitation maxima 365 nm and emission wavelength > 420 nm), digital camera attachment (Qlmaging QICAM Q21310), and Northern Eclipse software. For the inhibition experiments, cells were treated with CA074Me (50 μΜ) and E64D (50 μΜ) dissolved in serum free DMEM for 12 hours prior to probe addition.
Experimental
Reaction of H-Lys-A/-£-Boc-AMC (5)[32 34] with activated amino acid Cbz-Phe- OSu (6) in aqueous THF in presence of sodium bicarbonate'351 afforded the dipeptide Cbz-Phe-Lys-A/-£-Boc-AMC (7). Subsequent deprotection of the Boc group from 7 proceeded smoothly in a 50% TFA/CH2CI2 solution at ice-bath temperature that upon removal of the solvents in vacuo produced compound Cbz-Phe-Lys-AMC (1 ).
Similarly, Cbz-Lys-Lys-AMC (2) was easily prepared at gram scale levels using similar chemistry employed for the synthesis of 1 as shown in Scheme 1. Initially, a coupling reaction of Cbz-Lys-A/-£-Boc-Lys-A/-£-Boc-OH (11) which was available by the reaction of activated amino acid Cbz-Lys-A/-£-Boc-OSu (8) and H-Lys-A/-£-Boc- OH (10)[36] with 7-amino-4-methylcoumarin (AMC) (12) in presence of POCI3 in pyridine at -1 5 °C following a similar literature procedure'331 did not work. Instead, coupling of 5 with 8 in DMF in presence of triethyl amine'361 proceeded smoothly to give Cbz-Lys-A/-£-Boc-Lys-A/-£- Boc-AMC (9). Deprotection of Boc groups from 9 was performed using the same procedure as used for 7 to get compound 2 in pure form (Scheme 1).
Reagents and conditions: (a) NaHC03, THF-H20, rt, 1 6 h; (b) Et3N, DMF, 0 °C to rt, 16 h; (c) TFA-CH2CI2 (1 :1 , v/v), ice-bath, 15 min.; (d) POCI3, pyridine, -1 5 °C, 1 h.
The synthesis of the tripeptide was started with the reaction of 10 with 6 to get Cbz-Phe-Lys-A/-£-Boc-OH (13). Coupling of the linker p-aminobenzyl alcohol (PABA) (14) with 13 was carried out in presence of A/-ethoxycarbonyl-2-ethoxy-1 ,2- dihydroquinoline (EEDQ)[37] to give tripeptide Cbz-Phe-Lys-A/-£-Boc-PAB-OH (15). Several efforts for the preparation of AMC carbamate with 15 by activating it with p- nitrophenyl chloroformate (PNP),[35] 1 ,1 '-carbonyldiimidazole (CDI),[38] phosgene1391 were unsuccessful probably due to the low nucleophilicity of AMC. Efficient incorporation of AMC to yield carbamate Cbz-Phe-Lys-A/-£-Boc-PABC-AMC (17)
could only be accomplished following the reaction of 15 with 7-isocyanato-4- methylchromen-2-one (16) which was generated by refluxing of AMC 12 with phosgene.1401 On the other hand, Cbz-Lys-A/-£-Boc-Lys-A/-£-Boc-PAB-OH (18) was made by the reaction of 11 with PABA (14) using the same procedure as used for the synthesis of 15. Subsequent elaboration of 18 to carbamate Cbz-Lys-A/-£-Boc- Lys-A/-£-Boc-PABC-AMC (19) was made following the same strategy as used for the synthesis of 17. Finally, deprotection of Boc groups from both carbamates 17 and 19 with TFA/CH2CI2 furnished TFA salts of probes Cbz-Phe-Lys-PABC-AMC (3) and Cbz-Lys-Lys-PABC-AMC (4). In addition, deprotection of Boc from 17 with methanolic-HCI proceeded well to give HCI salt of probe 3 (Scheme 2).
SCHEME 2
3.TFA; X = TFA (100%)
3.HCI; X = HCI (87%)
Reagents and conditions: (a) NaHC03, THF-H20, rt, 16 h; (b) THF, EEDQ, rt, 16 h; (c) 15% phosgene in toluene, ^ 20oC, 16 h; (d) THF, 80 °C, 2 h; (e) TFA-CH2CI2 (1 :1 , v/v), ice-bath, 15 min; (f) Methanolic-HCI (0.5M), rt, 16 h; (g) Et3N, DMF, 0 °C to rt, 16 h.
Tripeptide bromides Cbz-Phe-Lys-/V-£-Boc-PAB-Br (20) and Cbz-Lys-A/-£- Boc-Lys-/V-£-Boc-PAB-Br (21 ) were efficiently made from the corresponding alcohols
15 and 18, respectively by treating with KBr and SOCI2 in presence of benzotriazole in DMF. Subsequent coupling reaction of 8-formyl-7-hydroxy-4-methylcoumarin (HMC) (22) with the corresponding bromides 20 or 21 proceeded well to get the respective products Cbz-Phe-Lys-A/-£-Boc-PAB-0-HMC (23) or Cbz-Lys-A/-£-Boc- Lys-A/-£-Boc-PAB-0-HMC (24) in good yields. Difluorination of 23 and 24 with diethylaminosulfur trifluoride (DAST) afforded Cbz-Phe-Lys-A/-£-Boc-PAB-0-DFHMC (25) and Cbz-Lys-A/-£-Boc-Lys-A/-£-Boc-PAB-0-DFHMC (26), respectively. Finally, deprotection of Boc groups from both HMC difluorides 25 and 26 with TFA/CH2CI2 furnished TFA salts of probes Cbz-Phe-Lys-PAB-O-DFHMC (27) and Cbz-Lys-Lys- PAB-O-DFHMC (28). On the other hand, coupling reactions of bromides 20 or 21 with sodium resorufin (29) in DMF yielded the corresponding ether Cbz-Phe-Lys-A/-£- Boc-PAB-O-res (30) or Cbz-Lys-A/-£-Boc-Lys-A/-£-Boc-PAB-0-res (31), respectively. Deprotection of 30 and 31 with 50% TFA resulted the final probes 32 and 33 in high yields (Scheme 3).
32; R1 = Ph 25; R1 = Ph
33; R' = -(CH2)3NHBoc
27; R1 = Ph
28; R1 = -(CH2)3NHBoc
Reagents and conditions: (a) i) benzotriazole, SOCI2, CH2CI2, rt, 5 min; ii) KBr, DMF, rt, 4 h; (b) 22, acetone, K2C03, reflux, 0.5 h; (c) DAST, CH2CI2, rt, 16 h; (d) TFA- CH2CI2 (1 :1 , v/v), ice-bath, 15 min; (e) 29, DMF, K2C03, rt, 16 h.
Synthetic schemes for compounds 38 and 39 are shown in Schemes 4 and 5 respectively.
SCHEME 4
4 M H n oxane, rt, 1 mn.
SCHEME 5
Reagents and Conditions:
(a) K2C03, acetone, reflux, 0.5 h;
(b) NaBH4, MeOH, rt, 2 h;
(c) (i) benzotriazole, SOCI2, DCM, rt, 10 min, (ii) KF, DMF, rt, 1.5 h;
(d) ethynyltrimethylsilane, [Ph3P]4Pd, Cul, Et3N-THF, reflux, 1.5 h;
(e) TBAF, THF, rt, 0.5 h;
(f) 4-fluoroiodobenzene, [Ph3P]4Pd, Cul, Et3N-THF, reflux, 1 h;
(g) 4 M HCI in dioxane, 10 min.
SCHEME 6
0 ^I t hNHj
General Remarks
1H and 13C NMR spectra were acquired in Lakehead University Instrumentation Laboratory (LUIL) on a Varian Unity Inova 500 MHz spectrometer in DMSO-d6 with TMS as the internal standard, where J (coupling constant) values were estimated in hertz (Hz). Microanalyses were measured for C, H, N in LUIL using Elementar Vario
EL and were within ±0.4% of theoretical values. Thin layer chromatography (TLC) and silica gel column chromatography were performed using TLC Silica Gel 60 F254
(EMD) and SiliaFlash®P60 (SiliCycle), respectively. Compounds 5,[32 34] 6,[43] 10,[41 ] 11 [36] and 13[42] were prepared according to literature procedures. Compound 8 was prepared using the same procedure as described for 6 and known in literature.'441 All other reagents and solvents were purchased from Sigma-Aldrich, Acros Organics, Alfa Aesar, ChemTech, and Fisher, and were used without further purification.
Synthesis and Characterization of Compounds
7
Cbz-Phe-Lys-W-£-Boc-AMC (7): To a solution of 5 (212 mg, 0.52 mmol) in THF - H20 (3.5:1 .5 v/v, 5 mL) and NaHC03 (44 mg, 0.52 mmol) was added 6 (206 mg, 0.52 mmol) in THF (3.5 mL) and stirred at room temperature for 16 h. The reaction mixture was then concentrated in vacuo and extracted with ethyl acetate (15 mL x 3). The combined organic phase was washed with brine (25 mL) and dried (Na2S04). After filtration, the solvent was removed from filtrate and the crude residue was purified by silica gel column chromatography using dichloromethane - methanol (9:1 v/v) as eluent to obtain the pure title compound 7. White solid, 288 mg (81 %); 1 H NMR (DMSO-de) δ 1 .28-1 .45 (m, 4H), 1 .35 (s, 9H), 1 .60-1 .70 (m, 1 H), 1 .70-1 .81 (m, 1 H), 2.41 (s, 3H), 2.75 (dd, J = 10.9, 13.7 Hz, 1 H), 2.83-2.95 (m, 2H), 3.05 (dd, J = 3.6, 15.3 Hz, 1 H), 4.31 -4.39 (m, 1 H), 4.39-4.46 (m, 1 H), 4.96 (s, 2H), 6.28 (d, J = 1 .3 Hz, 1 H), 6.79 (t, J = 5.6 Hz, 1 H), 7.16-7.36 (m, 10H), 7.47-7.54 (m, 2H), 7.74 (d, J = 8.8 Hz, 1 H), 7.79 (d, J = 1 .9 Hz, 1 H), 8.33 (d, J = 7.6 Hz, 1 H), 10.52 (s, 1 H). 13C NMR (DMSO-de) δ 18.5, 23.3, 28.7, 29.8, 32.2, 37.9, 54.2, 56.5, 65.7, 77.8, 106.2, 1 12.8, 1 15.6, 1 1 5.7, 126.4, 126.7, 127.3, 127.9, 128.2, 128.5, 1 28.8, 129.7, 137.5, 138.5, 142.7, 153.5, 154.1 , 156.0, 156.4, 160.5, 171 .9, 172.3. Anal. Calcd for C38H44N408: C, 66.65; H, 6.48; N, 8.18. Found: C, 66.40; H, 6.74; N, 8.10.
1
Cbz-Phe-Lys-AMC . TFA (1): To a solution of trifluoroacetic acid - DCM (1 :1 v/v, 2 mL) at ice-bath temperature was added 7 (200 mg, 0.29 mmol) and stirred for 15 min. After reaction, diethyl ether was added to the reaction mixture to precipitate out the solids. Then the solids were centrifuged out and successively washed with diethyl ether (5 mL x 2) and ethyl acetate (5 mL x 2), and dried under high vacuum in dark to get the pure title compound 1. White solid, 204mg (100%); 1 H NMR (DMSO- d6) δ 1 .23-1 .80 (m, 6H), 2.33 (s, 3H), 2.61 -2.78 (m, 3H), 2.94-3.04 (m, 1 H), 4.23-4.32 (m, 1 H), 4.32-4.43 (m, 1 H), 4.89 (s, 2H), 6.21 (d, J = 1 .2 Hz, 1 H), 7.08-7.31 (m, 10H), 7.45 (dd, J = 2.1 , 8.6 Hz, 1 H), 7.48 (d, J = 8.6 Hz, 1 H), 7.66 (d, J = 8.6 Hz, 1 H), 7.70-7.86 (m, 4H), 8.38 (d, J = 7.6 Hz, 1 H), 10.54 (s, 1 H). 13C NMR (DMSO-d6) δ 18.5, 22.9, 27.2, 31 .8, 37.8, 45.0, 54.1 , 56.5, 65.7, 1 06.2, 1 12.8, 1 15.6, 1 15.8, 1 17.7 (J = 300 Hz), 126.4, 1 26.7, 1 27.9, 128.2, 128.5, 128.8, 129.7, 137.5, 138.5, 142.6, 1 53.6, 154.1 , 156.4, 158.9 (J = 31 Hz), 160.5, 1 71 .8, 172.3. Anal. Calcd for C35H37F3N4O8: C, 60.1 7; H, 5.34; N, 8.02. Found: C, 59.82; H, 5.52; N, 7.95.
9
Cbz-Lys-/V-£-Boc-Lys-N-£-Boc-AMC (9): To a solution of 8 (592 mg, 1 .24 mmol) in dry DMF (5 mL) was added 5 (500 mg, 1 .24 mmol) and triethylamine (173 \iL, 1 .24 mmol) at 0 °C and stirred for 30 min and then at room temperature for 1 6 h. The reaction mixture was then evaporated in vacuo and the residue was dissolved in ethyl acetate (20 mL) and successively washed with 1 M citric acid (1 0 mL x 2), brine (1 5 mL) and dried (Na2S04). After filtration, the solvent was removed from filtrate and the crude residue was purified by silica gel column chromatography using ethyl
acetate - hexanes (4:1 v/v) as eluent to obtain the pure title compound 9. White solid, 935 mg (98%); 1H NMR (DMSO-d6) δ 1.29-1.44 (m, 6H), 1.37 (s, 18H), 1.45- 1.79 (m, 6H), 2.40 (s, 3H), 2.79-2.95 (m, 4H), 3.96-4.08 (m, 1H), 4.33-4.42 (m, 1H), 5.04 (s, 2H), 5.53- 5.61 (m, 1H), 6.28 (s, 1H), 6.77 (t, J= 5.0 Hz, 2H), 7.26-7.45 (m, 5H), 7.49 (dd, J= 2.0, 8.5 Hz, 1H), 7.72 (d, J= 8.5 Hz, 1H), 7.78 (d, J= 2.0 Hz, 1H), 8.14 (d, J= 7.6 Hz, 1H), 10.45 (s, 1H).13C NMR (DMSO-d6) δ 18.4, 23.2, 23.3, 24.9, 25.8, 28.7, 28.7, 29.7, 29.7, 32.1, 33.8, 54.0, 55.1, 65.9, 77.8, 77.8, 106.1, 112.8, 115.5, 115.7, 126.4, 128.2, 128.2, 128.8, 137.5, 142.6, 153.5, 154.1, 156.0, 156.5, 157.1 , 160.5, 172.0, 172.7. Anal. Calcd for C40H55N5O10: C, 62.73; H, 7.24; N, 9.14. Found: C, 63.12; H, 7.10; N,
Cbz-Lys-Lys-AMC .2TFA (2): To a solution of trifluoroacetic acid - DCM (1 :1 v/v, 1 mL) at ice-bath temperature was added 9 (100 mg, 0.13 mmol) and stirred for 15 min. After reaction, diethyl ether was added to the reaction mixture to precipitate out the solids. Then the solids were centrifuged out and successively washed with diethyl ether (5 mL x 2) and ethyl acetate (5 mL x 2), and dried under high vacuum in dark to get the pure title compound 2. White solid, 60 mg (58%).1H NMR (DMSO-d6) δ 1.28-1.83 (m, 12H), 2.41 (s, 3H), 2.69-2.84 (m, 4H), 3.98-4.11 (m, 1H), 4.34-4.47 (m, 1H), 5.04 (s, 2H), 6.28 (d, J= 1.2 Hz, 1H), 7.27-7.56 (m, 7H), 7.67-7.94 (m, 8H), 8.23-8.34 (m, 1H), 10.59 (s, 1H).13C NMR (DMSO-d6) δ 18.5, 22.9, 22.9, 27.1, 27.2, 31.7,31.7, 39.1,45.0, 53.9, 54.8, 65.9, 106.2, 114.1, 115.6, 115.7, 117.7 (J=300 Hz), 126.5, 128.2, 128.3, 128.8, 137.5, 142.6, 153.6, 154.1, 156.5, 158.7 (J= 31 Hz), 160.5, 171.9, 172.7. Anal. Calcd for C34H4i F6N5Oio.H20: C, 50.31 ; H, 5.34; N, 8.63. Found: C, 50.58; H, 5.35; N, 8.79
15
Cbz-Phe-Lys-W-£-Boc-PAB-OH (15): To a solution of 13 (0.83 g, 1 .57 mmol) and PABA (14) (232 mg, 1 .89 mmol) in THF (15 ml_) was added EEDQ (467 mg, 1 .89 mmol) and stirred at room temperature under argon for 16 h. After reaction, solvent was removed under reduced pressure and the residue was taken into diethyl ether (25 ml_) to form a suspension. The solid was filtered out and washed with diethyl ether (25 ml_ x 2) and dried under vacuum to get the pure title compound 15. Pale yellow solid, 0.93 g (94%). 1H NMR (DMSO-d6) δ 1 .20-1 .50 (m, 4H), 1 .36 (s, 9H), 1 .50-1 .60 (m, 1 H), 1 .60-1 .80 (m, 1 H), 2.74 (dd, J = 1 1 , 13.5 Hz, 1 H), 2.80-2.95 (m, 2H), 3.03 (dd, J = 5.5, 13.5 Hz, 1 H), 4.30-4.37 (m, 1 H), 4.37-4.50 (m, 1 H), 4.44 (d, J = 18 Hz, 2H), 4.96 (s, 2H), 5.12 (t, J = 5.5 Hz, 1 H), 6.79 (t, J = 5.5 Hz, 1 H), 7.1 5-7.35 (m, 12H), 7.51 (d, J = 8.5 Hz, 1 H), 7.56 (d, J = 8.0 Hz, 2H), 8.21 (d, J = 8.0 Hz, 1 H), 10.01 (S, 1 H). 13C NMR (DMSO-d6) δ 23.2, 28.7, 29.7, 32.5, 37.9, 40.5, 53.9, 56.5, 63.1 , 65.7, 77.8, 1 19.5, 126.7, 127.4, 127.9, 128.1 , 128.5, 128.8, 128.7, 137.5, 137.9, 1 38.0, 138.5, 156.0, 156.3, 170.8, 172.0. Anal. Calcd for C35H44N4O7: C, 66.44; H, 7.01 ; N, 8.85.
17
Cbz-Phe-Lys-W-E-Boc-PABC-AMC (17): AMC (12) (0.25 g, 1 .43 mmol) was taken in a 25 ml_ flask and a solution of 15% phosgene in toluene 10 ml_ was added to it. The reaction mixture was refluxed at 120 °C for 16 h under argon. After reaction, the mixture was cooled to room temperature and argon was bubbled into the solution for 10 minutes to remove excess phosgene gas. Then the solvents were removed under reduced pressure to dryness to get 7-isocyanato-4-methylchromen-2-one (16) as a
white powdered solid (ca. 0.29 g) which was used immediately without further purification. A solution of isocyanate 16 (ca. 0.29 g, 1 .43 mmol) and 15 (0.6 g, 0.95 mmol) in dry THF (5 mL) was stirred at 80 °C under argon for 2 h. After reaction, solvent was removed under reduced pressure to get the crude product 17 which contained some unreacted isocyanate 16 and alcohol derivative 15. To remove the isocyanate 16, the crude product was dissolved in dichloromethane (25 mL) and washed successively with 1 0% aqueous HCI solution (1 5 mL x 3), water (1 5 mL), saturated aqueous NaHC03 solution (15 mL), water (15 mL) and brine (15 mL), and dried (MgS04). After filtration, the solvents were removed from the filtrate and the solid was taken in ethyl acetate (20 mL) which dissolved unreacted alcohol derivative 15 only and the product was filtered out from the slurry. Repeating the procedure two more times afforded analytically pure title compound 17 as a white solid, 0.3 g (38%). The combined filtrates were evaporated under reduced pressure to recover the alcohol derivative 15 as a pale yellow solid, 0.25 g (42%). Compound 17: 1H NMR (DMSO-de) δ 1 .20-1 .50 (m, 4H), 1 .35 (s, 9H), 1 .60-1 .70 (m, 1 H), 1 .70-1 .80 (m, 1 H), 2.51 (s, 3H), 2.74 (dd, J = 9.5, 13.5 Hz, 1 H), 2.90 (d, J = 6 Hz, 2H), 3.05 (d, J = 13.5 Hz, 1 H), 4.34 (d, J = 6.5 Hz, 1 H), 4.41 (d, J = 6.5 Hz, 1 H), 4.96 (s, 2H), 5.14 (s, 2H), 6.24 (s, 1 H), 6.78 (s, 1 H), 7.15-7.35 (m, 10H), 7.40 (d, J = 8.5 Hz, 2H), 7.51 (d, J = 8.5 Hz, 1 H), 7.56 (s, 1 H), 7.64 (d, J = 8.5 Hz, 3H), 7.69 (d, J = 8.5 Hz, 1 H), 8.23 (d, J = 7.5 Hz, 1 H), 1 0.14 (s, 1 H), 1 0.27 (s, 1 H). 13C NMR (DMSO-d6) δ 18.4, 23.2, 28.7, 29.8, 32.4, 37.9, 45.0, 54.0, 54.0, 56.5, 65.7, 66.5, 77.8, 1 04.9, 104.9, 1 12.4, 1 14.7, 1 14.8, 1 1 9.6, 126.5, 126.7, 127.9, 128.1 , 128.5, 128.7, 1 29.6, 129.7, 131 .4, 137.5, 1 38.5, 139.4, 143.2, 153.6, 153.7, 154.3, 156.0, 156.3, 1 56.3, 160.5, 171 .1 , 172.0. MS (M + Na)+ 856. Anal. Calcd for C46H5iN5Oio.1 /2H20: C, 65.54; H, 6.22; N, 8.30. Found: C, 65.19;
3.TFA
Cbz-Phe-Lys-PABC-AMC . TFA (3.TFA): To a solution of trifluoroacetic acid - DCM (1 :1 v/v, 2 ml_) at ice-bath temperature was added 17 (200 mg, 0.24 mmol) and stirred for 15 min. After reaction, diethyl ether was added to the reaction mixture to precipitate out the solids. The solids were centrifuged out and successively washed with diethyl ether (5 ml_ x 2) and ethyl acetate (5 ml_ x 2), and dried under high vacuum in dark to get the pure title compound 3.TFA. White solid, 204 mg (100%). 1H NMR (DMSO-de) δ 1.30-1.84 (m, 6H), 2.39 (s, 3H), 2.73-2.83 (m, 3H), 3.05 (d, J = 9.9 Hz, 1H), 4.30-4.37 (m, 1H), 4.40-4.47 (m, 1H), 4.95 (s, 2H), 5.15 (s, 2H), 6.25 (s, 1H), 7.17-7.33 (m, 10H), 7.41 (d, J= 8.5 Hz, 3H), 7.53 (d, J= 8.9 Hz, 1H), 7.57 (bs, 1H), 7.66 (d, J= 8.9 Hz, 2H), 7.70 (d, J= 8.0 Hz, 1H), 7.78 (bs, 3H), 8.34 (d, J= 8.0 Hz, 1H), 10.21 (s, 1H), 10.29 (s, 1H).13C NMR (DMSO-d6) 18.0, 22.4, 26.8, 31.6, 37.4, 38.7, 53.4, 56.1, 65.3, 66.0, 104.5, 111.9, 114.3, 114.4, 119.2, 126.0, 126.3, 127.5, 127.7, 128.1, 128.3, 129.2, 129.3, 131.1, 137.0, 138.1, 138.9, 142.8, 153.2, 153.3, 153.9, 155.9, 160.1, 170.6, 171.8. Anal. Calcd for C43H44F3N5O10.I/2H2O: C, 60.27; H, 5.29; N, 8.17.
3.HCI
Cbz-Phe-Lys-PABC-AMC . HCI (3.HCI): A solution of 17 (100 mg, 0.12 mmol) in methanolic-HCI (0.5M, 5 ml_) was stirred at room temperature for 16 h. After reaction, solvents were removed in vacuo to dryness and the residue was taken in diethyl ether and stirred for 10 min. The suspension was centrifuged, the supernatant was discarded, and the process was repeated twice to get the pure title compound 3.HCI. White solid, 80 mg (87%).1H NMR (DMSO-d6) δ 1.31 -1.52 (m, 2H), 1.52-1.86 (m, 4H), 2.38 (s, 3H), 2.65-2.84 (m, 3H), 2.99-3.12 (m, 1H), 4.28-4.40 (m, 1H), 4.40- 4.51 (m, 1H), 4.95 (s, 2H), 5.14 (s, 2H), 6.23 (s, 1H), 7.16-7.36 (m, 10H), 7.37-7.46 (m, 3H), 7.51-7.60 (m, 2H), 7.66-7.73 (m, 3H), 8.03 (bs, 3H), 8.43 (d, J= 8.3 Hz, 1H), 10.31 (S, 1H), 10.37 (s, 1H).13C NMR (DMSO-d6) δ 18.5, 22.8, 27.0, 31.9, 37.8, 38.8, 53.9, 56.6, 65.7, 66.5, 105.0, 112.4, 114.7, 114.8, 119.6, 126.5, 126.7, 127.9,
128.1 , 128.5, 128.8, 1 29.6, 129.7, 131 .4, 138.6, 1 39.5, 143.2, 1 53.7, 154.3, 156.4, 160.5, 171 .1 , 172.2. Anal. Calcd for C41 H44CIN5O8.2H2O: C, 61 .07; H, 6.00; N, 8.69. Found: C, 60.85; H, 5.79; N, 8.69.
18
Cbz-Lys-W-E-Boc-Lys-W-E-Boc-PAB-OH (18): To a solution of 11 (3.71 g, 6.10 mmol) and PABA (14) (0.9 g, 7.32 mmol) in THF (50 mL) was added EEDQ (1 .81 g, 7.32 mmol) and stirred at room temperature under argon for 16 h. After reaction, solvent was removed under reduced pressure and the residue was taken into diethyl ether (50 mL) to form a suspension. The solid was filtered out and washed with diethyl ether (50 mL x 2) and dried under vacuum to get the pure title compound 18. White solid, 1 .42 g (33%). 1 H NMR (DMSO-d6) δ 1 .14-1 .75 (m, 12H), 1 .36 (s, 9H), 1 .37 (s, 9H), 2.88 (bs, 4H), 3.95-4.05 (m, 1 H), 4.31 -4.40 (m, 1 H), 4.44 (d, J = 5.6 Hz, 2H), 5.04 (s, 2H), 5.12 (t, J = 5.6 Hz, 1 H), 6.77 (t, J = 5.6 Hz, 2H), 7.24 (d, J = 8.0 Hz, 2H), 7.28-7.40 (m, 4H), 7.43 (d, J = 8.0 Hz, 1 H), 7.55 (d, J = 8.0 Hz, 2H), 8.05 (d, J = 8.0 Hz, 1 H), 9.96 (s, 1 H). 13C NMR (DMSO-d6) δ 23.2, 23.3, 28.7, 29.7, 29.7, 32.1 , 32.4, 53.8, 55.2, 63.1 , 65.9, 77.8, 1 1 9.4, 127.3, 128.2, 128.2, 128.8, 137.5, 137.9, 1 38.0, 156.0, 156.0, 156.5, 170.9, 172.5. Anal. Calcd for C37H55N5O9.I /2H2O: C, 61 .48; H, 7.81 ; N, 9.69. Found: C, 61 .50; H, 7.84; N, 9.79.
19
Cbz-Lys-/V-£-Boc-Lys-N-£-Boc-PABC-AMC (19): A solution of isocyanate 16 (ca. 2.29 g, 1 1 .43 mmol) and 18 (0.9 g, 1 .26 mmol) in dry THF (80 mL) was stirred at 80 °C under argon for 2 h. After reaction, solvent was removed under reduced pressure
and the residue was successively suspended in diethyl ether (100 mL), stirred for 10 min and filtered, and then repeated with ethyl acetate (50 mL x 2), stirred 10 min and filtered to get the pure title compound 19. Pale yellow solid, 0.96 g (83%).1H NMR (DMSO-de) δ 1.21 -1.45 (m, 8H), 1.34 (s, 9H), 1.37 (s, 9H), 1.45-1.77 (m, 4H), 2.39 (s, 3H), 2.81-2.94 (m, 4H), 3.98-4.04 (m, 1H), 4.34-4.41 (m, 1H), 5.01 (s, 2H), 5.16 (s, 2H), 6.23 (d, J= 1.2 Hz, 1H), 6.70-6.83 (m, 2H), 7.28-7.44 (m, 9H), 7.56 (d, J = 1.9 Hz, 1H), 7.63 (d, J= 8.4 Hz, 2H), 7.69 (d, J= 8.8 Hz, 1H), 8.04 (d, J= 7.8 Hz, 1H), 10.07 (s, 1H), 10.27 (s, 1H).13C NMR (DMSO-d6) δ 14.5, 18.5, 21.2, 23.2, 23.3, 28.7, 28.7, 29.7, 29.7, 32.1, 32.3, 53.8, 55.2, 60.2, 65.9, 66.5, 77.8, 104.9, 112.4, 114.7, 114.8, 119.6, 126.4, 128.2, 128.2, 128.8, 129.6, 131.4, 137.5, 139.4, 143.2, 153.6, 153.7, 154.3, 156.0, 156.0, 156.5, 160.5, 171.2, 172.6. Anal. Calcd for C48H62N60i2.1/2H20: 62.57; H, 6.96; N, 8.91.
4
Cbz-Lys-Lys-PABC-AMC .2TFA (4): To a solution of trifluoroacetic acid - DCM (1 :1 v/v, 2 mL) at ice-bath temperature was added 19 (200 mg, 0.22 mmol) and stirred for 15 min. After reaction, diethyl ether was added to the reaction mixture to precipitate out the solids and the suspension was centrifuged, discarding the supernatant. The solids obtained were dissolved in water (2 mL) washed with ethyl acetate (5 mL x 2), and the aqueous phase was freeze dried in dark to get the pure title compound 4. White solid, 48 mg (47%).1H NMR (DMSO-d6) δ 1.23-1.80 (m, 12H), 2.40 (s, 3H), 2.70-2.82 (m, 4H), 4.00-4.08 (m, 1H), 4.36-4.44 (m, 1H), 5.03 (s, 2H), 5.15 (s, 2H), 6.25 (d, J= 1.2 Hz, 1H), 7.28-7.48 (m, 9H), 7.57 (d, J= 2 Hz, 1H), 7.64 (d, J= 8.5 Hz, 2H), 7.67-7.81 (m, 7H), 8.17 (d, J= 7.9 Hz, 1H), 10.15 (s, 1H), 10.28 (S, 1 H).13C NMR (DMSO-d6) δ 18.5, 22.9, 22.9, 27.1 , 27.2, 31.7, 32.0, 39.1 , 45.0, 53.7, 54.9, 65.9, 66.4, 104.9, 112.4, 114.7, 114.8, 117.7 (J=300 Hz), 119.6, 126.5, 128.2, 128.3, 128.8, 129.6, 131.5, 137.5, 139.3, 143.2, 153.7, 153.7, 154.3,
156.5, 158.7 (J = 31 Hz), 160.5, 1 71 .1 , 1 72.5. Anal. Calcd for
C42H48F6N6Oi2.1/2H20: C, 53.00; H, 5.19; N, 8.83. Found: C, 52.62; H, 5.34; N, 8.84.
20
Representative Synthesis of Cbz-Phe-Lys-/V-£-Boc-PAB-Br (20): To a solution of 15 (3.16 g, 5 mmol) and benzotriazole (0.59 g, 5 mmol) in DCM (10 mL) was added thionyl chloride (0.4 mL, 5.50 mmol) and stirred at room temperature for 5 min. Then KBr (0.60 g, 5 mmol) followed by DMF (10 mL) were added to the reaction mixture and stirred for additional 4 hours. After reaction, water (25 mL) was added and extracted with ethyl acetate (50 mL x 2). The combined organic phase was successively washed with water (50 mL), brine (50 mL), and dried (Na2S04). After filtration, the filtrate was evaporated in vacuo, and the solids were washed several times with ethyl acetate to get the pure compound as a white solid 2.71 g (78%). [M - Br]+ 615.
23
Representative Synthesis of Cbz-Phe-Lys-W-E-Boc-PAB-O-HMC (23): To a solution of 20 (2.0 g, 2.88 mmol) and 22 (2.94 g, 14.40 mmol) in dry acetone (50 mL) was added anhydrous K2C03 (2.0 g, 14.40 mmol) and refluxed for 0.5 h. After reaction, acetone was removed and water (25 mL) was added and extracted with ethyl acetate (50 mL x 2). The combined organic phase was successively washed with water (50 mL), brine (50 mL), and dried (Na2S04). After filtration, the filtrate was evaporated in vacuo, and the crude product was purified by silica gel column chromatography using EtOAC : hexanes (4 :1 v/v) as eluents to get the pure title compound 23. Pale yellow solid, 1 .06 g (45%). 1H NMR (DMSO-d6) δ 1 .18-1 .40 (m,
13H), 1 .50-1 .60 (m, 1 H), 1 .60-1 .72 (m, 1 H), 2.36 (s, 3H), 2.69 (dd, J = 9.5, 13.5 Hz, 1 H), 2.79-2.89 (m, 2H), 2.98 (d, J = 13.5 Hz, 1 H), 4.24-4.31 (m, 1 H), 4.31 -4.39 (m, 1 H), 4.90 (S, 2H), 5.28 (s, 2H), 6.25 (s, 1 H), 6.70 (s, 1 H), 7.09-7.31 (m, 1 1 H), 7.38- 7.47 (m, 3H), 7.59 (d, J = 8.5 Hz, 2H), 7.92 (d, J = 8.5 Hz, 1 H), 8.15 (d, J = 8.5 Hz, 1 H), 1 0.01 (S, 1 H), 1 0.44
25
Representative Synthesis of Cbz-Phe-Lys-W-E-Boc-PAB-O-DFHMC (25): To a solution of 23 (0.50 g, 0.61 mmol) in dry CH2CI2 (8 mL) was added DAST (160 \iL, 1 .22 mmol) and stirred at room temperature for 1 6 h. The reaction mixture was quenched with ice-water and the organic phase was separated and washed with brine (5 mL), and dried (MgS04). After filtration, the filtrate was evaporated in vacuo, and the crude product was purified by silica gel column chromatography using EtOAC : hexanes (4 :1 v/v) as eluents to get the pure title compound 25. Pale yellow solid, 142 mg (28%). [M + 863.
27
Representative Synthesis of Cbz-Phe-Lys-PAB-O-DFHMC . TFA (27): To a solution of trifluoroacetic acid - DCM (1 :1 v/v, 0.5 mL) at ice-bath temperature was added 25 (100 mg, 0.12 mmol) and stirred for 10 min. After reaction, diethyl ether was added to the reaction mixture to precipitate out the solids. The solids were centrifuged out and washed with diethyl ether (2 mL x 2) and dried under high vacuum in dark to get the pure title compound 27. White solid, 52 mg (51 %).
31
Representative Synthesis of Cbz-Lys-/V-£-Boc-Lys-/V-£-Boc-PAB-0-res (31): To a solution of 21 (0.5 g, 0.64 mmol) and 29 (0.75 g, 3.20 mmol) in dry DMF (50 ml_) was added anhydrous K2C03 (0.44 g, 3.20 mmol) and stirred at room temperature for 16 h. After reaction, water (200 ml_) was added and extracted with ethyl acetate (100 ml_ x 2). The combined organic phase was successively washed with water (1 00 ml_ x 2), brine (100 ml_), and dried (Na2S04). After filtration, the filtrate was evaporated in vacuo, and the crude product was purified by silica gel column chromatography using CH2CI2 : MeOH (15 :1 v/v) as eluents to get the pure title compound 31. Red soli
33
Representative Synthesis of Cbz-Lys-Lys-PAB-O-res . 2TFA (33): To a solution of trifluoroacetic acid - DCM (1 :1 v/v, 2 mL) at ice-bath temperature was added 31 (200 mg, 0.22 mmol) and stirred for 15 min. After reaction, diethyl ether was added to the reaction mixture to precipitate out the solids. The solids were centrifuged out, dissolved in water (3 mL) and washed with ethyl acetate (5 mL x 2). The aqueous part was then evaporated using freeze dryer in dark to get the pure title compound 33. Red solid, 140 mg (68%).1H NMR (DMSO-d6) δ 1 .14-1 .71 (m, 12H), 2.54-2.77 (m, 4H), 3.88-4.02 (m, 1 H), 4.25-4.37 (m, 1 H), 4.95 (s, 2H), 5.15 (s, 2H), 6.20 (d, J = 1 .9 Hz, 1 H), 6.71 (dd, J = 1 .9, 9.8 Hz, 1 H), 7.04 (dd, J = 2.5, 8.8 Hz, 1 H), 7.1 1 (d, J = 2.5 Hz, 1 H), 7.21 -7.33 (m, 5H), 7.36 (d, J = 8.2 Hz, 3H), 7.46 (d, J = 9.8 Hz, 1 H),
7.57 (d, J = 8.2 Hz, 2H), 7.67 (bs, 6H), 7.71 (d, J = 8.8 Hz, 1 H), 8.1 3 (bs, 1 H), 10.1 0 (s, 1 H). Anal. Calcd for C43H46F6N6Oi i .4/5H20: C, C, 54.29; H, 5.04; N, 8.83. Found: C, 53.98; H, 4.88; N, 8.66.
Enzyme Kinetic Studies
To obtain the KM, kcat and kcat/KM parameters for each compound as a substrate of CTB or CTL, concentrations of fluorogenic peptide were varied between ~ 0.5 KM - 5 KM while maintaining a constant concentration of activated enzyme. The reaction mixture (150 μΙ final volume) consisted of 30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 2.0 mM DTT, 10% DMSO, and 0.33 to 3.3 nM human cathepsin B or L in 96 well black plates. The samples were pre-warmed at 37°C for 30 min, and the reaction was initiated upon addition of the enzyme. The activity was monitored spectrophotometrically for the release of 7-amino-4-methyl-coumarin (AMC;
excitation 380 nm; emission 460 nm). The amount of AMC released from the reaction was determined by a standard curve in assay buffer. Incubation of the prodrug inspired probes in the assay buffer without enzyme demonstrated that the probes were stable for at least 6 hours with very little spontaneous hydrolysis of AMC as monitored by fluorescence or HPLC (data not shown). The assay to quantify active enzyme by titration with E64 was adapted from Barrett et a/.[45] Briefly, human cathepsin enzyme (-1 0 nM) in assay buffer (30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 1 .0 mM DTT) was inactivated with E64 (final concentrations ranging from 1 .5 to 7.0 nM) for 20 min at 37 °C. Following the inactivation reaction, an aliquot of this mixture was added to a solution containing the substrate (30 μΜ z-RR-AMC or z-FK- AMC in 10% DMSO 90% assay buffer) at 37 °C. The residual enzymatic activity was monitored spectrophotometrically for the release of 7-amino-4-methyl-coumarin and plotted versus the concentration of E64 to graphically determine the amount of active enzyme.
Preparation of Cell Lysates from HeLa Cervical Cancer Cells
The cell lysates and assay were adapted from Giusti et a/.[46]. Briefly, cells were grown to ~ 80% confluency and washed with Phenol Red free DMEM, treated with lyse buffer (10 mM Tris-HCI, pH 7.5, 1 50 mM NaCI, 1 % Triton X-100, 1 mM EDTA) and centrifuged (12,000 rpm) in a 1 .5 ml_ tube. For the CTB activity assays,
100 μΙ_ of the lysis solution was added to a black 96 well plate and diluted with 100 μΙ_ activation buffer (100 mM Acetate, pH 5.5, 5 mM DTT, 5 mM EDTA) followed by 15 minute incubation at 37 °C. A 100 μΙ_ aliquot of the activated lysate was then added to a 50 μΙ_ solution containing 300 μΜ probe, in 30% DMSO and 70% buffer (100 mM acetate buffer, 5 mM EDTA, pH 5.5). The enzymatic reaction proceeded at 37 °C for 1 hour after which 50 μΙ_ was withdrawn and added to 100 μΙ_ of sodium choloroacetate (200 mM) to stop the reaction. The fluorescence at 460 nm was measured and normalized to total protein concentration (Bradford Assay) and expressed as a percentage of AMC released from experiments using Z-Arg-Arg- AMC. As a control, spontaneous AMC hydrolysis was determined in assay buffer without lysate while hydrolysis by other protease classes was assessed by pre- treating lysates with sodium chloroacetate (200 mM in assay buffer). Levels of free AMC were similar in both control reactions and therefore used as a blank which was subtracted from the total RFU in each experiment.
Fluorescent Probes
MDA MB 231 breast cancer cells grown on cover slips were treated with PBS buffer (control) or compound 36 for 4h at 37°C. Results are shown in Figures 6 to 8. The compound turns green after Cathepsin B hydrolysis. Cells were then treated with DAPI (blue, to stain nucleus) and lysosome tracker (red) and incubated for another 30 minutes. Cells were then washed with PBS fixed with paraformaldehyde onto slides and fluorescence microscopy images were taken at 100X. The probe activation is clearly visualized in the lysosomes of cells.
Radiolabeled Probes
Turnover of compound 38, a non-radioactive version of a potential Cathepsin B PET probe, in presence and absence of Cathepsin B was examined using HPLC. In order to visualize Cathepsin B activity in vivo using PET, a fluorinated probe candidate must be an efficient substrates of the enzyme. In the first experiment, compound 38 (100 μΜ) was incubated in assay buffer (30 mM acetate-NaOH, pH 5.5, 3.0 mM EDTA, 1 .0 mM DTT, 10% DMSO) at 37 °C for 90 minutes. A 20 μΐ aliquot was injected into an HPLC fitted with a C8 column and detected at 254 nm. In the second experiment, Cathepsin B (0.3 nM) was included in the assay buffer.
As can be seen in Figure 14, the largest peak in the chromatogram at ~ 5 minute retention time corresponds to intact compound 38 clearly demonstrating high chemical stability of the probe candidate. In a second experiment, a 20 μΙ_ aliquot was injected into the HPLC showing a 40% reduction in the total amount of compound 32. New reaction product peaks can be seen at longer retention times. This experiment clearly shows compound 38 is an efficient substrate of Cathepsin B. A series of HPLC experiments performed at varying concentrations of compound 32 allowed to estimate I M of 30,000 mM"1s"1.
References
[I ] M. B. Sporn, Lancet 1996, 347, 1377-1381 .
[2] L. A. Liotta, K. Tryggvason, S. Garbisa, I. Hart, C. M. Foltz, S. Shafie, Nature
1980, 284, 67-68.
[3] S. D. Mason, J. A. Joyce, Trends Cell Biol. 2011 , 21, 228-237.
[4] B. F. Sloane, S. Yan, I. Podgorski, B. E. Linebaugh, M. L. Cher, J. Mai, D.
Cavallo-Medved, M. Sameni, J. Dosescu, K. Moin, Semin. Cancer Biol. 2005,
15, 149-157.
[5] A. Premzl, V. Zavasnik-Bergant, V. Turk, J. Kos, Exp. Cell Res. 2003, 283, 206-214.
[6] A. T. Chan, Y. Baba, K. Shima, K. Nosho, D. C. Chung, K. E. Hung, U.
Mahmood, K. Madden, K. Poss, A. Ranieri, D. Shue, R. Kucherlapati, C. S.
Fuchs, S. Ogino, Cancer Epidemiol. Biomarkers Prev. 2010, 19, 2777-2785.
[7] M. R. Emmert-Buck, M. J. Roth, Z. Zhuang, E. Campo, J. Rozhin, B. F.
Sloane, L. A. Liotta, W. G. Stetler-Stevenson, Am. J. Pathol. 1994, 145, 1285-
1290.
[8] W. J. Kruszewski, R. Rzepko, J. Wojtacki, J. Skokowski, A. Kopacz, K.
Jaskiewicz, K. Drucis, Neoplasma 2004, 51, 38-43.
[9] M. A. Nouh, M. M. Mohamed, M. El-Shinawi, M. A. Shaalan, D. Cavallo- Medved, H. M. Khaled, B. F. Sloane, J. Transl. Med. 2011 , 9, 1 .
[10] B. C. Victor, A. Anbalagan, M. M. Mohamed, B. F. Sloane, D. Cavallo- Medved, Breast Cancer Res. 2011 , 13, R1 15.
[I I ] A. Spes, B. Sobotic, V. Turk, B. Turk, Biol. Chem. 2012, 393, 1417-1431 .
[12] M. M. Mohamed, B. F. Sloane, Nat. Rev. Cancer. 2006, 6, 764-775.
[13] T. Shree, O. C. Olson, B. T. Elie, J. C. Kester, A. L. Garfall, K. Simpson, K. M.
Bell-McGuinn, E. C. Zabor, E. Brogi, J. A. Joyce, Genes Dev. 2011 , 25, 2465- 2479.
[14] G. Blum, G. von Degenfeld, M. J. Merchant, H. M. Blau, M. Bogyo, Nat.
Chem. Biol. 2007, 3, 668-677.
[15] B. M. Creasy, C. B. Hartmann, F. K. White, K. L. McCoy, Cytometry A. 2007,
71, 1 14-123.
T. G. Terentyeva, W. Van Rossom, M. Van der Auweraer, K. Blank, J.
Hofkens, Bioconjug. Chem. 2011 , 22, 1932-1 938.
P. L. Carl, P. K. Chakravarty, J. A. Katzenellenbogen, J. Med. Chem. 1981 , 24, 479-480.
G. B. Jones, C. F. Crasto, J. E. Mathews, L. Xie, M. O. Mitchell, A. El-Shafey, A. V. D'Amico, G. J. Bubley, Bioorg. Med. Chem. 2006, 14, 418-425.
J. A. Richard, Y. Meyer, V. Jolivel, M. Massonneau, R. Dumeunier, D. Vaudry,
H. Vaudry, P. Y. Renard, A. Romieu, Bioconjug. Chem. 2008, 19, 1707-1718.
I. Penuelas, G. Mazzolini, J. F. Boan, B. Sangro, J. Marti-Climent, M. Ruiz, J. Ruiz, N. Satyamurthy, C. Qian, J. R. Barrio, M. E. Phelps, J. A. Richter, S. S. Gambhir, J. Prieto, Gastroenterology 2005, 128, 1 787-1795.
J. S. Fowler, R. R. MacGregor, A. P. Wolf, C. D. Arnett, S. L. Dewey, D.
Schlyer, D. Christman, J. Logan, M. Smith, S. Sachs, M. Aquilonius, P.
Bjurling, C. Halldin, P. Hartvig, K. L. Leenders, H. Lundqvist, L. Oreland, C. G. Stalnacke, B. and Langstrom, Science 1987, 235, 481 .
R. L. Melo, L. C. Alves, E. Del Nery, L. Juliano, M. A. Juliano, Anal. Biochem. 2001 , 293, 71 -77.
Y. Choe, F. Leonetti, D. C. Greenbaum, F. Lecaille, M. Bogyo, D. Bromme, J. A. Ellman, C. S. Craik, J. Biol. Chem. 2006, 281, 1 2824-12832.
S. Tian, K. Baek, I. Shin, Chem. Sci. 2013, 4, 947.
T. Morita, H. Kato, S. Iwanaga, K. Takada, T. Kimura, J. Biochem. 1977, 82, 1495-1498.
S. Hasnain, T. Hirama, C. P. Huber, P. Mason, J. S. Mort, J. Biol. Chem. 1993, 268, 235-240.
G. M. Dubowchik, R. A. Firestone, Bioorg. Med. Chem. Lett. 1998, 8, 3341 - 3346.
D. Turk, G. Guncar, M. Podobnik, B. Turk, Biol. Chem. 1998, 379, 137-147. M. Boshart, L. Gissmann, H. Ikenberg, A. Kleinheinz, W. Scheurlen, H. zur Hausen, EMBO J. 1984, 3, 1 151 -1 157.
J. M. du Manoir, G. Francia, S. Man, M. Mossoba, J. A. Medin, A. Viloria-Petit, D. J. Hicklin, U. Emmenegger, R. S. Kerbel, Clin. Cancer Res. 2006, 12, 904- 916.
B. Wang, G. P. Shi, P. M. Yao, Z. Li, H. A. Chapman, D. Bromme, J. Biol. Chem. 1998, 273, 32000-32008.
A. S. Madsen, C. A. Olsen, J. Med. Chem. 2012, 55, 5582-5590.
B. Heltweg, F. Dequiedt, B. L. Marshall, C. Brauch, M. Yoshida, N. Nishino, E. Verdin, M. Jung, J. Med. Chem. 2004, 47, 5235-5243.
B. J. Backes, J. L. Harris, F. Leonetti, C. S. Craik, J. A. Ellman, Nature Biotech. 2000, 18, 187-193.
K. A. Ajaj, M. L. Biniossek, F. Kratz, Bioconjugate Chem. 2009, 20, 390-396. A. M. Felix, D. Gillessen, W. Lergier, J. A. Meienhofer, A. Trzeciak, Eur. Pat. Appl. 1984, EP123951 A2.
G. M. Dubowchik, R. A. Firestone, L. Padilla, D. Willner, S. J. Hofstead, K. Mosure, J. O. Knipe, S. J. Lasch, P. A. Trail, Bioconjugate Chem. 2002, 13, 855-869.
Y. Gao, Y. Lam, J. Comb. Chem. 2010, 12, 69-74.
I. A. Mueller, F. Kratz, M. Jung, A. Warnecke, Tetrahedron Lett. 2010, 51, 4371 -4374.
A. Renslo, S. Mahajan, PCT Int. Appl., WO 2010/01 1 684.
S. Wiejak, E. Masiukiewicz, B. Rzeszotarska, Chem. Pharm. Bull. 1999, 47,
1489-1490.
P. Rauber, B. Walker, S. Stone, E. Shaw, Biochem. J. 1988, 250, 871 -876.
R. A. Firestone, G. M. Dubowchik, Eur. Pat. Appl. 1994, EP 624377 A2.
J. L. Kuhler, R. T. Brooks, W. K. Helms, M. Hodge, K. D. Holly, B. L. Inman,
0. A. Koriko, R. R. Patel, T. R. Shorter, Afr. J. Pure Appl. Chem. 2009, 3, 108- 1 15.
A. J. Barrett, A. A. Kembhavi, M. A. Brown, H. Kirschke, C. G. Knight, M. Tamai, and K. Hanada, Biochem. J. 1982, 201 , 189-1 98.
1. Giusti, S. D'Ascenzo, D. Millimaggi, G. Taraboletti, G. Carta, N. Franceschini, A. Pavan and V. Dolo. Neoplasia. 2008, 10, 481 -488.
Claims
1 . A compound of formul
wherein:
R1 is PhCH20-, CH3 or PhCH2-;
R2 is PhCH2-, (CH3)2CH- or H2N(CH2)m- for m > 4;
R3 and R3' are each H or -CH2OR4, and at least one of R3 and R3'
is -CH2OR4,wherein for each of R3 and R3 , R4 is independently selected from the group consisting of:
wherein:
R5 is H, -C(0)H, -CHX2, wherein each X is, independently of the other
X, selected from the group consisting of:
H, F, CI, Br, I, -N02, toluenesulfonate, methanesulfonate, trifluoromethansulfonate, perfluorobutanesulfonate,
ethanesulfonate, benzenesulfonate,
parachlorobenzenesulfonate, nitrobenzenesulfonate or methoxybenzenesulfonate, and both X are not simultaneously
H,
wherein:
R7 is H, -C≡CH, N3, -C(0)H, or -CHX2, wherein each X is, independently of the other X, selected from the group consisting of:
H, F, CI, Br, I, -N02, toluenesulfonate, methanesulfonate, trifluoromethansulfonate, perfluorobutanesulfonate,
ethanesulfonate, benzenesulfonate,
parachlorobenzenesulfonate, nitrobenzenesulfonate or methoxybenzenesulfonate, and both X are not simultaneously H, and
wherein each of R9 and R10 is, independently of the other, H,
N3, -C(0)H, -CHX2, wherein each X is, independently of the other X, selected from the group consisting of:
H, F, CI, Br, I, -N02, toluenesulfonate, methanesulfonate, trifluoromethansulfonate, perfluorobutanesulfonate, ethanesulfonate, benzenesulfonate, parachlorobenzenesulfonate,
nitrobenzenesulfonate or methoxybenzenesulfonate, and both X are not simultaneously H,
-CH2Y, wherein Y is selected from the group consisting of:
-OH, -OC(O)NHAr1 , wherein Ar1 is -C6H5 in which 1 , 2 or 3 hydrogen atoms is optionally and independently substituted by -NO2, and
, where Z is selected from the group consisting of H, CI, and Br.
a fluorescent dye,
a radiolabeled substituent,
or -C≡CR11 wherein R1 1 is H or Ar2, in which Ar2 is -C6H5 in which 1 , 2 or 3 hydrogen atoms is optionally and independently substituted by F, and
R9 and R10 are not simultaneously H,
n > 4,
including salts, hydrates and solvates thereof.
2. A compound of claim 1 , wherein the fluorescent dye is a substituent containing a borondipyrromethene (BODIPY) core.
3. A compound of claim 1 , wherein said fluorescent dye is
4. A compound of claim 2, wherein the BODIPY of the compound is covalently linked at the carbon located at the para-position with respect to the boron atom.
5. A compound of claim 1 , wherein the radiolabeled substituent comprises an 18F atom.
6. The compound of claim 5, wherein the radiolabeled substituent is a C1 -C20 alkyl group or C5-C8 aromatic group consisting of carbon atoms or carbon and nitrogen atoms, and optionally substituted with a C1 -C20 alkyl or alkoxy group substituted with said 18F atom(s).
7. The compound of claim 6, wherein the radiolabelled substituent is selected from the group consisting of:.
8. A compound as defined by any one of claims 1 to 7, wherein R3 is the same as R3'.
9. A compound as defined by any one of claims 1 to 7, wherein R3 is the same as R3 are different from each other.
10. A compound as defined by any one of claims 1 to 9, wherein m = 4, 5, 6, 7, 8, 9 or 1 0.
1 1 . A compound as defined by any one of claims 1 to 1 0, wherein n = 4, 5, 6, 7, 8, 9, or 1 0.
12. A composition for use as an imaging probe comprising at least one compound selected from the group consisting of compounds as defined in any of claims 1 to 1 1 .
13. A composition as defined by claim 12, for use in PET imaging, said compound comprising 18F as defined by any one of claims 1 and 5 to 1 1 .
14. A composition as defined by claim 12, for use in fluorescence imaging, said compound comprising a fluorescent dye.
15. A composition as defined by claim 14, wherein said compound is as defined by any one of claims 2 to 4.
16. A pharmaceutical composition for use in PET and/or fluorescence imaging comprising a compound as defined in any one of claims 1 to 1 1 , and a
pharmaceutically acceptable carrier.
17. The pharmaceutical composition according to claim 16 wherein the compound is dispersed or dissolved in a liquid medium suitable for injection.
18. A method of PET and/or fluorescence imaging comprising administering to a patient in need thereof an effective amount of the composition as defined by any one of claims 12 to 18, and scanning the subject with a PET and/or fluorescence imaging device.
19. A method for studying the localization of PET probes within a tissue of a subject comprising: administering to the subject an effective amount of a compound as defined by claim 1 3, subjecting a tissue of the subject to irradiation of an
electromagnetic radiation, wherein the electromagnetic radiation is absorbed by the compound, detecting fluorescence of the compound within the tissue, wherein the fluorescence of the compound within the tissue is indicative of the presence of a PET probe within the tissue.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US201461942751P | 2014-02-21 | 2014-02-21 | |
| US61/942,751 | 2014-02-21 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2015123783A1 true WO2015123783A1 (en) | 2015-08-27 |
Family
ID=53877486
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/CA2015/050136 WO2015123783A1 (en) | 2014-02-21 | 2015-02-23 | Cathepsin b-targeting probes |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2015123783A1 (en) |
Cited By (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN105859807A (en) * | 2016-04-12 | 2016-08-17 | 成都大学 | Beta-1,4-galactosyltransferase I substrate fluorescent molecular probe, and intermediate and preparation method thereof |
| EP3127913A1 (en) * | 2015-08-03 | 2017-02-08 | Ruprecht-Karls-Universität Heidelberg | Chromogenic and fluorogenic peptide substrates for the detection of serine protease activity |
-
2015
- 2015-02-23 WO PCT/CA2015/050136 patent/WO2015123783A1/en active Application Filing
Non-Patent Citations (3)
| Title |
|---|
| BAUDY, A.R. ET AL.: "Non-invasive optical imaging of muscle pathology in mdx mice using cathepsin caged near-infrared imaging", MOLECULAR IMAGING AND BIOLOGY, vol. 13, 2011, pages 462 - 470, XP019899005, ISSN: 1536-1632 * |
| CHOWDHURY, M.A. ET AL.: "Prodrug-inspired probes selective to cathepsin B over other cysteine cathepsins''.", JOURNAL OF MEDICINAL CHEMISTRY, vol. 57, 18 June 2014 (2014-06-18), pages 6092 - 6104, XP055220697, ISSN: 0022-2623 * |
| RYU, JU HEE ET AL.: "Cathepsin B-sensitive nanoprobe for in vivo tumor diagnosis''.", JOURNAL OF MATERIALS CHEMISTRY, vol. 21, 2011, pages 17631 - 17634, XP055220692, ISSN: 0959-9428 * |
Cited By (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| EP3127913A1 (en) * | 2015-08-03 | 2017-02-08 | Ruprecht-Karls-Universität Heidelberg | Chromogenic and fluorogenic peptide substrates for the detection of serine protease activity |
| WO2017021004A1 (en) * | 2015-08-03 | 2017-02-09 | Ruprecht-Karls-Universität Heidelberg | Chromogenic and fluorogenic peptide substrates for the detection of serine protease activity |
| CN105859807A (en) * | 2016-04-12 | 2016-08-17 | 成都大学 | Beta-1,4-galactosyltransferase I substrate fluorescent molecular probe, and intermediate and preparation method thereof |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Chowdhury et al. | Prodrug-inspired probes selective to cathepsin B over other cysteine cathepsins | |
| CN110143900B (en) | Methods of treatment using SIRT2 modulators | |
| Iwaszkiewicz-Grzes et al. | Synthesis and biological activity of mycophenolic acid-amino acid derivatives | |
| Brown et al. | Peptide length and leaving-group sterics influence potency of peptide phosphonate protease inhibitors | |
| JP5789254B2 (en) | Lysine compounds and their use in site-selective and functional group-selective modification of peptides and proteins | |
| PL171232B1 (en) | Method of obtaining novel derivatives of hydrazine | |
| CA2950612C (en) | Sulfonamide compounds and their use as stat5 inhibitors | |
| Kohl et al. | Design, characterization and cellular uptake studies of fluorescence-labeled prototypic cathepsin inhibitors | |
| WO2015123783A1 (en) | Cathepsin b-targeting probes | |
| US10383956B2 (en) | Fluorescent probe for detecting dipeptidyl peptidase IV | |
| US20150299250A1 (en) | Macrocyclic compounds and uses thereof | |
| NZ247837A (en) | Cyclic oxirane derivatives, medicaments and precursors | |
| CA3170840A1 (en) | Macrocyclic compounds, a process for their preparation and pharmaceu tical compositions containing them | |
| Huang et al. | Development of activity-based probes with tunable specificity for protein tyrosine phosphatase subfamilies | |
| RU2697519C1 (en) | Peptide agent comprising a psma-binding ligand based on a urea derivative, a method for production thereof and use thereof for preparing a conjugate with a drug and diagnostic agent | |
| FR2828884A1 (en) | HYDRAZINOPEPTOIDS AND THEIR USES IN THE TREATMENT OF CANCERS | |
| JP2018145126A (en) | Fluorescent probe for detection of carboxypeptidase activity | |
| Maezaki et al. | 3-Acyl-2-(N-cyanoimino) oxazolidine derivative, a new asymmetric acylating agent for racemic secondary alkyl amines | |
| US20100317831A1 (en) | Probe compounds for protein tyrosine phosphatase (ptp) and precursors thereof | |
| WO2008145830A2 (en) | Novel pro-fluorescent compounds | |
| US11241501B2 (en) | Anticancer prodrug for overcoming drug resistance | |
| Stranix et al. | Lysine sulfonamides as novel HIV-Protease inhibitors: optimization of the Nε-acyl-phenyl spacer | |
| PL227742B1 (en) | Derivatives of 1-aminalkanephosphonic acid diaryl esters, process for the preparation of derivative of 1-aminalkanephosphonic acid diaryl esters and their use | |
| CA2791535C (en) | Methods and systems for preparing irreversible inhibitors of protein tyrosine phosphatases | |
| Fuchs et al. | Subnanomolar Cathepsin S Inhibitors with High Selectivity: Optimizing Covalent Reversible α‐Fluorovinylsulfones and α‐Sulfonates as Potential Immunomodulators in Cancer |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 15751548 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 15751548 Country of ref document: EP Kind code of ref document: A1 |