EP2798069B1 - Porcine knob xenotype chimeric adenoviral vector for dendritic cell infection - Google Patents
Porcine knob xenotype chimeric adenoviral vector for dendritic cell infection Download PDFInfo
- Publication number
- EP2798069B1 EP2798069B1 EP12858372.1A EP12858372A EP2798069B1 EP 2798069 B1 EP2798069 B1 EP 2798069B1 EP 12858372 A EP12858372 A EP 12858372A EP 2798069 B1 EP2798069 B1 EP 2798069B1
- Authority
- EP
- European Patent Office
- Prior art keywords
- seq
- knob
- ad5luc1
- fiber
- immunizing antigen
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Not-in-force
Links
- 210000004443 dendritic cell Anatomy 0.000 title claims description 113
- 239000013598 vector Substances 0.000 title description 50
- 208000015181 infectious disease Diseases 0.000 title description 22
- 241001135569 Human adenovirus 5 Species 0.000 claims description 79
- 239000000835 fiber Substances 0.000 claims description 65
- 210000004027 cell Anatomy 0.000 claims description 64
- 108090000623 proteins and genes Proteins 0.000 claims description 48
- 239000000427 antigen Substances 0.000 claims description 39
- 108091007433 antigens Proteins 0.000 claims description 39
- 102000036639 antigens Human genes 0.000 claims description 39
- 241000282414 Homo sapiens Species 0.000 claims description 36
- 108090000765 processed proteins & peptides Proteins 0.000 claims description 27
- 230000003053 immunization Effects 0.000 claims description 26
- KFEUJDWYNGMDBV-LODBTCKLSA-N N-acetyllactosamine Chemical group O[C@@H]1[C@@H](NC(=O)C)[C@H](O)O[C@H](CO)[C@H]1O[C@H]1[C@H](O)[C@@H](O)[C@@H](O)[C@@H](CO)O1 KFEUJDWYNGMDBV-LODBTCKLSA-N 0.000 claims description 18
- 150000001413 amino acids Chemical class 0.000 claims description 18
- 102000004169 proteins and genes Human genes 0.000 claims description 16
- 230000003612 virological effect Effects 0.000 claims description 13
- 102000007563 Galectins Human genes 0.000 claims description 12
- 108010046569 Galectins Proteins 0.000 claims description 12
- 229940062780 lacto-n-neotetraose Drugs 0.000 claims description 12
- 238000000034 method Methods 0.000 claims description 12
- GUBGYTABKSRVRQ-QKKXKWKRSA-N Lactose Natural products OC[C@H]1O[C@@H](O[C@H]2[C@H](O)[C@@H](O)C(O)O[C@@H]2CO)[C@H](O)[C@@H](O)[C@H]1O GUBGYTABKSRVRQ-QKKXKWKRSA-N 0.000 claims description 11
- 239000008101 lactose Substances 0.000 claims description 11
- 150000001720 carbohydrates Chemical class 0.000 claims description 10
- 229960005486 vaccine Drugs 0.000 claims description 10
- 241001492394 Porcine adenovirus 4 Species 0.000 claims description 9
- IEQCXFNWPAHHQR-UHFFFAOYSA-N lacto-N-neotetraose Natural products OCC1OC(OC2C(C(OC3C(OC(O)C(O)C3O)CO)OC(CO)C2O)O)C(NC(=O)C)C(O)C1OC1OC(CO)C(O)C(O)C1O IEQCXFNWPAHHQR-UHFFFAOYSA-N 0.000 claims description 8
- GUBGYTABKSRVRQ-XLOQQCSPSA-N Alpha-Lactose Chemical compound O[C@@H]1[C@@H](O)[C@@H](O)[C@@H](CO)O[C@H]1O[C@@H]1[C@@H](CO)O[C@H](O)[C@H](O)[C@H]1O GUBGYTABKSRVRQ-XLOQQCSPSA-N 0.000 claims description 7
- RBMYDHMFFAVMMM-PLQWBNBWSA-N neolactotetraose Chemical compound O([C@H]1[C@H](O)[C@H]([C@@H](O[C@@H]1CO)O[C@@H]1[C@H]([C@H](O[C@H]([C@H](O)CO)[C@H](O)[C@@H](O)C=O)O[C@H](CO)[C@@H]1O)O)NC(=O)C)[C@@H]1O[C@H](CO)[C@H](O)[C@H](O)[C@H]1O RBMYDHMFFAVMMM-PLQWBNBWSA-N 0.000 claims description 7
- FWMNVWWHGCHHJJ-SKKKGAJSSA-N 4-amino-1-[(2r)-6-amino-2-[[(2r)-2-[[(2r)-2-[[(2r)-2-amino-3-phenylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]hexanoyl]piperidine-4-carboxylic acid Chemical compound C([C@H](C(=O)N[C@H](CC(C)C)C(=O)N[C@H](CCCCN)C(=O)N1CCC(N)(CC1)C(O)=O)NC(=O)[C@H](N)CC=1C=CC=CC=1)C1=CC=CC=C1 FWMNVWWHGCHHJJ-SKKKGAJSSA-N 0.000 claims description 6
- 206010028980 Neoplasm Diseases 0.000 claims description 6
- 201000011510 cancer Diseases 0.000 claims description 6
- 238000004113 cell culture Methods 0.000 claims description 6
- 108010065889 glycyl-leucyl-cysteinyl-threonyl-leucyl-valyl-alanyl-methionyl-leucine Proteins 0.000 claims description 6
- 108010061181 influenza matrix peptide (58-66) Proteins 0.000 claims description 6
- 235000014633 carbohydrates Nutrition 0.000 claims description 3
- 230000001131 transforming effect Effects 0.000 claims description 2
- 102000008198 Coxsackie and Adenovirus Receptor Like Membrane Protein Human genes 0.000 description 32
- 108010035601 Coxsackie and Adenovirus Receptor Like Membrane Protein Proteins 0.000 description 32
- 238000012546 transfer Methods 0.000 description 26
- 108060001084 Luciferase Proteins 0.000 description 25
- 239000005089 Luciferase Substances 0.000 description 25
- 230000000694 effects Effects 0.000 description 22
- 241000701161 unidentified adenovirus Species 0.000 description 20
- 210000001616 monocyte Anatomy 0.000 description 16
- 235000018102 proteins Nutrition 0.000 description 15
- 108020004414 DNA Proteins 0.000 description 13
- 108700019146 Transgenes Proteins 0.000 description 12
- 238000003556 assay Methods 0.000 description 12
- 235000001014 amino acid Nutrition 0.000 description 11
- 229940024606 amino acid Drugs 0.000 description 11
- 238000010494 dissociation reaction Methods 0.000 description 11
- 230000005593 dissociations Effects 0.000 description 11
- KFEUJDWYNGMDBV-UHFFFAOYSA-N (N-Acetyl)-glucosamin-4-beta-galaktosid Natural products OC1C(NC(=O)C)C(O)OC(CO)C1OC1C(O)C(O)C(O)C(CO)O1 KFEUJDWYNGMDBV-UHFFFAOYSA-N 0.000 description 10
- HESSGHHCXGBPAJ-UHFFFAOYSA-N N-acetyllactosamine Natural products CC(=O)NC(C=O)C(O)C(C(O)CO)OC1OC(CO)C(O)C(O)C1O HESSGHHCXGBPAJ-UHFFFAOYSA-N 0.000 description 10
- 238000001476 gene delivery Methods 0.000 description 10
- 239000013612 plasmid Substances 0.000 description 9
- 101710145505 Fiber protein Proteins 0.000 description 7
- 238000002474 experimental method Methods 0.000 description 7
- 239000002245 particle Substances 0.000 description 7
- 108090000978 Interleukin-4 Proteins 0.000 description 6
- 241000700605 Viruses Species 0.000 description 6
- 230000001404 mediated effect Effects 0.000 description 6
- 108010038196 saccharide-binding proteins Proteins 0.000 description 6
- 108090000331 Firefly luciferases Proteins 0.000 description 5
- 108700008625 Reporter Genes Proteins 0.000 description 5
- 230000001413 cellular effect Effects 0.000 description 5
- 239000002609 medium Substances 0.000 description 5
- 102000005962 receptors Human genes 0.000 description 5
- 108020003175 receptors Proteins 0.000 description 5
- 108010017213 Granulocyte-Macrophage Colony-Stimulating Factor Proteins 0.000 description 4
- 102000004457 Granulocyte-Macrophage Colony-Stimulating Factor Human genes 0.000 description 4
- AIYUHDOJVYHVIT-UHFFFAOYSA-M caesium chloride Chemical compound [Cl-].[Cs+] AIYUHDOJVYHVIT-UHFFFAOYSA-M 0.000 description 4
- 239000012634 fragment Substances 0.000 description 4
- 230000028993 immune response Effects 0.000 description 4
- 230000035800 maturation Effects 0.000 description 4
- 210000003819 peripheral blood mononuclear cell Anatomy 0.000 description 4
- 238000010361 transduction Methods 0.000 description 4
- 210000002845 virion Anatomy 0.000 description 4
- 108010029697 CD40 Ligand Proteins 0.000 description 3
- 102100032937 CD40 ligand Human genes 0.000 description 3
- 241000701022 Cytomegalovirus Species 0.000 description 3
- 102000003886 Glycoproteins Human genes 0.000 description 3
- 108090000288 Glycoproteins Proteins 0.000 description 3
- 101000961414 Homo sapiens Membrane cofactor protein Proteins 0.000 description 3
- 241000282567 Macaca fascicularis Species 0.000 description 3
- 102100039373 Membrane cofactor protein Human genes 0.000 description 3
- 241001529936 Murinae Species 0.000 description 3
- 230000001464 adherent effect Effects 0.000 description 3
- 230000004700 cellular uptake Effects 0.000 description 3
- 230000006957 competitive inhibition Effects 0.000 description 3
- 230000002950 deficient Effects 0.000 description 3
- 230000001419 dependent effect Effects 0.000 description 3
- 150000004676 glycans Chemical class 0.000 description 3
- 230000013595 glycosylation Effects 0.000 description 3
- 238000006206 glycosylation reaction Methods 0.000 description 3
- 238000011534 incubation Methods 0.000 description 3
- 239000003112 inhibitor Substances 0.000 description 3
- 230000005764 inhibitory process Effects 0.000 description 3
- 239000000203 mixture Substances 0.000 description 3
- SKOZFDIGKDPQBO-QMIVOQANSA-N n-[(2s,3r,4r,5r,6r)-4,5-dihydroxy-6-(hydroxymethyl)-2-phenylmethoxyoxan-3-yl]acetamide Chemical compound CC(=O)N[C@@H]1[C@@H](O)[C@@H](O)[C@@H](CO)O[C@@H]1OCC1=CC=CC=C1 SKOZFDIGKDPQBO-QMIVOQANSA-N 0.000 description 3
- FXUAIOOAOAVCGD-FKSUSPILSA-N swainsonine Chemical compound C1CC[C@H](O)[C@H]2[C@H](O)[C@H](O)CN21 FXUAIOOAOAVCGD-FKSUSPILSA-N 0.000 description 3
- 229960005566 swainsonine Drugs 0.000 description 3
- FXUAIOOAOAVCGD-UHFFFAOYSA-N swainsonine Natural products C1CCC(O)C2C(O)C(O)CN21 FXUAIOOAOAVCGD-UHFFFAOYSA-N 0.000 description 3
- 230000026683 transduction Effects 0.000 description 3
- 238000001890 transfection Methods 0.000 description 3
- 230000010415 tropism Effects 0.000 description 3
- 101000870242 Bacillus phage Nf Tail knob protein gp9 Proteins 0.000 description 2
- 241000588724 Escherichia coli Species 0.000 description 2
- 108050002220 Green fluorescent protein, GFP Proteins 0.000 description 2
- 102100028976 HLA class I histocompatibility antigen, B alpha chain Human genes 0.000 description 2
- 241000712431 Influenza A virus Species 0.000 description 2
- 102100037850 Interferon gamma Human genes 0.000 description 2
- 108010074328 Interferon-gamma Proteins 0.000 description 2
- ZDXPYRJPNDTMRX-VKHMYHEASA-N L-glutamine Chemical compound OC(=O)[C@@H](N)CCC(N)=O ZDXPYRJPNDTMRX-VKHMYHEASA-N 0.000 description 2
- 241000282553 Macaca Species 0.000 description 2
- 241000188845 Porcine adenovirus Species 0.000 description 2
- 210000001744 T-lymphocyte Anatomy 0.000 description 2
- 239000004473 Threonine Substances 0.000 description 2
- 102100033782 UDP-galactose translocator Human genes 0.000 description 2
- 108010075920 UDP-galactose translocator Proteins 0.000 description 2
- 238000004458 analytical method Methods 0.000 description 2
- 210000000612 antigen-presenting cell Anatomy 0.000 description 2
- 238000013459 approach Methods 0.000 description 2
- 230000015572 biosynthetic process Effects 0.000 description 2
- 210000004369 blood Anatomy 0.000 description 2
- 239000008280 blood Substances 0.000 description 2
- 210000004899 c-terminal region Anatomy 0.000 description 2
- 210000004978 chinese hamster ovary cell Anatomy 0.000 description 2
- 201000010099 disease Diseases 0.000 description 2
- 208000037265 diseases, disorders, signs and symptoms Diseases 0.000 description 2
- 230000002500 effect on skin Effects 0.000 description 2
- 238000000799 fluorescence microscopy Methods 0.000 description 2
- 238000001943 fluorescence-activated cell sorting Methods 0.000 description 2
- 239000005090 green fluorescent protein Substances 0.000 description 2
- 238000003306 harvesting Methods 0.000 description 2
- 230000006801 homologous recombination Effects 0.000 description 2
- 238000002744 homologous recombination Methods 0.000 description 2
- 238000000338 in vitro Methods 0.000 description 2
- 230000015788 innate immune response Effects 0.000 description 2
- 102000006495 integrins Human genes 0.000 description 2
- 108010044426 integrins Proteins 0.000 description 2
- 201000001441 melanoma Diseases 0.000 description 2
- 244000052769 pathogen Species 0.000 description 2
- 230000001717 pathogenic effect Effects 0.000 description 2
- 239000013641 positive control Substances 0.000 description 2
- 230000003389 potentiating effect Effects 0.000 description 2
- 238000002360 preparation method Methods 0.000 description 2
- 239000000047 product Substances 0.000 description 2
- 230000004044 response Effects 0.000 description 2
- 230000000638 stimulation Effects 0.000 description 2
- 239000006228 supernatant Substances 0.000 description 2
- 238000002198 surface plasmon resonance spectroscopy Methods 0.000 description 2
- 238000003786 synthesis reaction Methods 0.000 description 2
- 238000002255 vaccination Methods 0.000 description 2
- 238000001262 western blot Methods 0.000 description 2
- MTCFGRXMJLQNBG-REOHCLBHSA-N (2S)-2-Amino-3-hydroxypropansäure Chemical compound OC[C@H](N)C(O)=O MTCFGRXMJLQNBG-REOHCLBHSA-N 0.000 description 1
- 108091032973 (ribonucleotides)n+m Proteins 0.000 description 1
- JKMHFZQWWAIEOD-UHFFFAOYSA-N 2-[4-(2-hydroxyethyl)piperazin-1-yl]ethanesulfonic acid Chemical compound OCC[NH+]1CCN(CCS([O-])(=O)=O)CC1 JKMHFZQWWAIEOD-UHFFFAOYSA-N 0.000 description 1
- CXURGFRDGROIKG-UHFFFAOYSA-N 3,3-bis(chloromethyl)oxetane Chemical compound ClCC1(CCl)COC1 CXURGFRDGROIKG-UHFFFAOYSA-N 0.000 description 1
- 102100035793 CD83 antigen Human genes 0.000 description 1
- 108091008048 CMVpp65 Proteins 0.000 description 1
- 241000701101 Canine adenovirus 1 Species 0.000 description 1
- 241000701114 Canine adenovirus 2 Species 0.000 description 1
- 241000282465 Canis Species 0.000 description 1
- 108090000565 Capsid Proteins Proteins 0.000 description 1
- 241001133757 Carpentaria Species 0.000 description 1
- 102100023321 Ceruloplasmin Human genes 0.000 description 1
- 108091026890 Coding region Proteins 0.000 description 1
- 108020004705 Codon Proteins 0.000 description 1
- 208000035473 Communicable disease Diseases 0.000 description 1
- 241000699802 Cricetulus griseus Species 0.000 description 1
- 238000002965 ELISA Methods 0.000 description 1
- 238000012413 Fluorescence activated cell sorting analysis Methods 0.000 description 1
- 101150066002 GFP gene Proteins 0.000 description 1
- 208000032612 Glial tumor Diseases 0.000 description 1
- 206010018338 Glioma Diseases 0.000 description 1
- 239000007995 HEPES buffer Substances 0.000 description 1
- 102100028972 HLA class I histocompatibility antigen, A alpha chain Human genes 0.000 description 1
- 108010075704 HLA-A Antigens Proteins 0.000 description 1
- 108010058607 HLA-B Antigens Proteins 0.000 description 1
- 102000006354 HLA-DR Antigens Human genes 0.000 description 1
- 108010058597 HLA-DR Antigens Proteins 0.000 description 1
- 108010088652 Histocompatibility Antigens Class I Proteins 0.000 description 1
- 241000282412 Homo Species 0.000 description 1
- 101000946856 Homo sapiens CD83 antigen Proteins 0.000 description 1
- 101000603882 Homo sapiens Nuclear receptor subfamily 1 group I member 3 Proteins 0.000 description 1
- 108010001336 Horseradish Peroxidase Proteins 0.000 description 1
- 241000701024 Human betaherpesvirus 5 Species 0.000 description 1
- 241000701044 Human gammaherpesvirus 4 Species 0.000 description 1
- 108060003951 Immunoglobulin Proteins 0.000 description 1
- 102100022297 Integrin alpha-X Human genes 0.000 description 1
- 229930182816 L-glutamine Natural products 0.000 description 1
- AYFVYJQAPQTCCC-GBXIJSLDSA-N L-threonine Chemical compound C[C@@H](O)[C@H](N)C(O)=O AYFVYJQAPQTCCC-GBXIJSLDSA-N 0.000 description 1
- 102000043129 MHC class I family Human genes 0.000 description 1
- 108091054437 MHC class I family Proteins 0.000 description 1
- 241000701244 Mastadenovirus Species 0.000 description 1
- 102000012750 Membrane Glycoproteins Human genes 0.000 description 1
- 108010090054 Membrane Glycoproteins Proteins 0.000 description 1
- 241000701168 Murine adenovirus 1 Species 0.000 description 1
- 108700019961 Neoplasm Genes Proteins 0.000 description 1
- 102000048850 Neoplasm Genes Human genes 0.000 description 1
- 108010038807 Oligopeptides Proteins 0.000 description 1
- 102000015636 Oligopeptides Human genes 0.000 description 1
- 241000188266 Ovine adenovirus 7 Species 0.000 description 1
- 238000012408 PCR amplification Methods 0.000 description 1
- 238000010222 PCR analysis Methods 0.000 description 1
- 239000002033 PVDF binder Substances 0.000 description 1
- 102000003992 Peroxidases Human genes 0.000 description 1
- 108010008281 Recombinant Fusion Proteins Proteins 0.000 description 1
- 102000007056 Recombinant Fusion Proteins Human genes 0.000 description 1
- 239000006146 Roswell Park Memorial Institute medium Substances 0.000 description 1
- MTCFGRXMJLQNBG-UHFFFAOYSA-N Serine Natural products OCC(N)C(O)=O MTCFGRXMJLQNBG-UHFFFAOYSA-N 0.000 description 1
- AYFVYJQAPQTCCC-UHFFFAOYSA-N Threonine Natural products CC(O)C(N)C(O)=O AYFVYJQAPQTCCC-UHFFFAOYSA-N 0.000 description 1
- 238000002835 absorbance Methods 0.000 description 1
- 229940099550 actimmune Drugs 0.000 description 1
- 230000033289 adaptive immune response Effects 0.000 description 1
- 108010084938 adenovirus receptor Proteins 0.000 description 1
- 230000030741 antigen processing and presentation Effects 0.000 description 1
- 125000000613 asparagine group Chemical group N[C@@H](CC(N)=O)C(=O)* 0.000 description 1
- 230000001580 bacterial effect Effects 0.000 description 1
- 230000008901 benefit Effects 0.000 description 1
- 230000000903 blocking effect Effects 0.000 description 1
- 239000000872 buffer Substances 0.000 description 1
- 239000013592 cell lysate Substances 0.000 description 1
- 238000005119 centrifugation Methods 0.000 description 1
- 239000013000 chemical inhibitor Substances 0.000 description 1
- 239000013599 cloning vector Substances 0.000 description 1
- 150000001875 compounds Chemical class 0.000 description 1
- 238000010276 construction Methods 0.000 description 1
- 239000013078 crystal Substances 0.000 description 1
- 230000016396 cytokine production Effects 0.000 description 1
- 238000012217 deletion Methods 0.000 description 1
- 230000037430 deletion Effects 0.000 description 1
- 238000013461 design Methods 0.000 description 1
- 238000011161 development Methods 0.000 description 1
- 230000029087 digestion Effects 0.000 description 1
- 238000010790 dilution Methods 0.000 description 1
- 239000012895 dilution Substances 0.000 description 1
- 231100000673 dose–response relationship Toxicity 0.000 description 1
- 108010048367 enhanced green fluorescent protein Proteins 0.000 description 1
- 239000003623 enhancer Substances 0.000 description 1
- 239000003797 essential amino acid Substances 0.000 description 1
- 235000020776 essential amino acid Nutrition 0.000 description 1
- 238000000684 flow cytometry Methods 0.000 description 1
- 238000002073 fluorescence micrograph Methods 0.000 description 1
- 239000012737 fresh medium Substances 0.000 description 1
- 238000012239 gene modification Methods 0.000 description 1
- 230000005017 genetic modification Effects 0.000 description 1
- 235000013617 genetically modified food Nutrition 0.000 description 1
- ZDXPYRJPNDTMRX-UHFFFAOYSA-N glutamine Natural products OC(=O)C(N)CCC(N)=O ZDXPYRJPNDTMRX-UHFFFAOYSA-N 0.000 description 1
- 230000036541 health Effects 0.000 description 1
- 238000003384 imaging method Methods 0.000 description 1
- 238000003119 immunoblot Methods 0.000 description 1
- 102000018358 immunoglobulin Human genes 0.000 description 1
- 230000001939 inductive effect Effects 0.000 description 1
- 230000003993 interaction Effects 0.000 description 1
- 108010042414 interferon gamma-1b Proteins 0.000 description 1
- 238000001738 isopycnic centrifugation Methods 0.000 description 1
- 238000011005 laboratory method Methods 0.000 description 1
- 210000001821 langerhans cell Anatomy 0.000 description 1
- 229940087875 leukine Drugs 0.000 description 1
- 210000000265 leukocyte Anatomy 0.000 description 1
- 239000012139 lysis buffer Substances 0.000 description 1
- 238000004519 manufacturing process Methods 0.000 description 1
- 239000000463 material Substances 0.000 description 1
- 239000012528 membrane Substances 0.000 description 1
- 230000001617 migratory effect Effects 0.000 description 1
- 238000010369 molecular cloning Methods 0.000 description 1
- 239000000178 monomer Substances 0.000 description 1
- 230000035772 mutation Effects 0.000 description 1
- 210000004897 n-terminal region Anatomy 0.000 description 1
- 239000013642 negative control Substances 0.000 description 1
- 150000007523 nucleic acids Chemical group 0.000 description 1
- 239000002773 nucleotide Substances 0.000 description 1
- 125000003729 nucleotide group Chemical group 0.000 description 1
- 229920001542 oligosaccharide Polymers 0.000 description 1
- 150000002482 oligosaccharides Chemical class 0.000 description 1
- 210000001672 ovary Anatomy 0.000 description 1
- 230000037361 pathway Effects 0.000 description 1
- 108040007629 peroxidase activity proteins Proteins 0.000 description 1
- 230000002085 persistent effect Effects 0.000 description 1
- 229920002401 polyacrylamide Polymers 0.000 description 1
- 230000008488 polyadenylation Effects 0.000 description 1
- 229920001184 polypeptide Polymers 0.000 description 1
- 229920002981 polyvinylidene fluoride Polymers 0.000 description 1
- 238000002203 pretreatment Methods 0.000 description 1
- 230000037452 priming Effects 0.000 description 1
- 102000004196 processed proteins & peptides Human genes 0.000 description 1
- 230000000644 propagated effect Effects 0.000 description 1
- 238000000746 purification Methods 0.000 description 1
- 230000009467 reduction Effects 0.000 description 1
- 230000010076 replication Effects 0.000 description 1
- 230000003362 replicative effect Effects 0.000 description 1
- 108091008146 restriction endonucleases Proteins 0.000 description 1
- 108010038379 sargramostim Proteins 0.000 description 1
- 238000000926 separation method Methods 0.000 description 1
- 239000013605 shuttle vector Substances 0.000 description 1
- 210000003491 skin Anatomy 0.000 description 1
- 238000002415 sodium dodecyl sulfate polyacrylamide gel electrophoresis Methods 0.000 description 1
- 238000010561 standard procedure Methods 0.000 description 1
- 239000000126 substance Substances 0.000 description 1
- 239000000758 substrate Substances 0.000 description 1
- 230000008685 targeting Effects 0.000 description 1
- 230000001225 therapeutic effect Effects 0.000 description 1
- 238000002560 therapeutic procedure Methods 0.000 description 1
- 208000037816 tissue injury Diseases 0.000 description 1
- 239000013638 trimer Substances 0.000 description 1
- 210000004881 tumor cell Anatomy 0.000 description 1
- 230000003827 upregulation Effects 0.000 description 1
- 238000010200 validation analysis Methods 0.000 description 1
- 210000000605 viral structure Anatomy 0.000 description 1
- 239000013603 viral vector Substances 0.000 description 1
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/85—Vectors or expression systems specially adapted for eukaryotic hosts for animal cells
- C12N15/86—Viral vectors
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/51—Medicinal preparations containing antigens or antibodies comprising whole cells, viruses or DNA/RNA
- A61K2039/525—Virus
- A61K2039/5256—Virus expressing foreign proteins
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2710/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA dsDNA viruses
- C12N2710/00011—Details
- C12N2710/10011—Adenoviridae
- C12N2710/10041—Use of virus, viral particle or viral elements as a vector
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2710/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA dsDNA viruses
- C12N2710/00011—Details
- C12N2710/10011—Adenoviridae
- C12N2710/10311—Mastadenovirus, e.g. human or simian adenoviruses
- C12N2710/10322—New viral proteins or individual genes, new structural or functional aspects of known viral proteins or genes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2710/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA dsDNA viruses
- C12N2710/00011—Details
- C12N2710/10011—Adenoviridae
- C12N2710/10311—Mastadenovirus, e.g. human or simian adenoviruses
- C12N2710/10341—Use of virus, viral particle or viral elements as a vector
- C12N2710/10343—Use of virus, viral particle or viral elements as a vector viral genome or elements thereof as genetic vector
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2810/00—Vectors comprising a targeting moiety
- C12N2810/50—Vectors comprising as targeting moiety peptide derived from defined protein
- C12N2810/60—Vectors comprising as targeting moiety peptide derived from defined protein from viruses
- C12N2810/6009—Vectors comprising as targeting moiety peptide derived from defined protein from viruses dsDNA viruses
- C12N2810/6018—Adenoviridae
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y02—TECHNOLOGIES OR APPLICATIONS FOR MITIGATION OR ADAPTATION AGAINST CLIMATE CHANGE
- Y02A—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE
- Y02A50/00—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE in human health protection, e.g. against extreme weather
- Y02A50/30—Against vector-borne diseases, e.g. mosquito-borne, fly-borne, tick-borne or waterborne diseases whose impact is exacerbated by climate change
Description
- This application claims priority to
US Provisional Patent Application 61/576,116 filed 15 December 2011 - This work was made with the support of Grant 5R33AI076096-06 from the National Institutes of Health. The government of the United States of America may have certain rights in this work.
- The Sequence Listing, which is a part of the present disclosure, includes a computer readable form and a written sequence listing comprising nucleotide and/or amino acid sequences. The subject matter of the Sequence Listing is incorporated herein by reference in its entirety. The information recorded in computer readable form is identical to the written sequence listing.
- Dendritic cells (DCs) are an important component of innate immunity. DCs are potent antigen presenting cells (APCs) with the ability to initiate the primary immune response. Banchereau, J., et al., Annu. Rev. Immunol. 18: 767-811, 2000. In addition to their role in local innate immune responses, DCs play a crucial role in adaptive immune response by priming the immune response or by inducing tolerance. Humans have DCs with different phenotypes that are distributed throughout the body and reside at the site of potential pathogen entry or tissue injury, where they differentiate into mature DCs (Li, K., et al., Mol. Immunol. 48: 1121-1127, 2011). During maturation, DCs undergo phenotypic and functional changes allowing them to increase their antigen presenting and increase their expression of co-stimulatory molecules (Rea, D., et al., J. Immunol. 166: 5236-5244, 2001). Bacterial and viral compounds have been identified as major DC maturation signals (Sallusto, F. and Lanzavecchia, A., J. Exp. Med.182: 389-400, 1994; Hartmann, G., et al., Proc. Natl. Acad. Sci. USA. 96: 9305-9310, 1999; Sparwasser, T., et al., Eur. J. Immunol. 28: 2045-2054, 1998; Verdijk, R.M., et al., J. Immunol. 163: 57-61, 1999; Cella, M., et al., J. Exp. Med. 189: 821-829, 1999).
- Since DCs have a unique ability to prime an immune response, targeting them in immune intervention strategies against infectious diseases as well as cancer has shown promise (Palucka, K., et al., J. Immunol. 186: 1325-1331, 2011). Multiple approaches have been developed to deliver antigens to DCs for presentation including transfection with DNA or RNA and gene transfer via recombinant vectors (Fong, L., et al., Annu. Rev. Immunol. 18: 245-273, 2000; Gong, J., et al., Nat. Med. 3: 558-561, 1997). Genetic modification of DCs with recombinant viruses offers major advantages including persistent antigen presentation over time and exposure to potentially immune-activating viral components (Sloan, J.M., et al. Cancer Gene Ther. 9: 946-950, 2002). Clinical trials have shown treatment with adenovirus-based vectors are safe and with the development of transductionally targeted, selectively replicating vectors to be increasingly effective against diseases (Paul, C.P.L., et al., Cancer Biol. Ther. 7: 786-793, 2008). Paul et al. showed that replacing the fiber knob of Ad5 with certain non-human knobs enhanced infectivity of human glioma cell populations and primary tumor cells.
- Rieneke Van De Ven et al., (J. Immunother. 2009; 32(9): 895-906) disclose selective transduction of mature DC in human skin and lymph nodes by CD80 / CD86-targeted fiber-modified Adenovirus-5/3.
- C.P.L. Paul et al., (Cancer Biology & Therapy 7:5, 786-793; May 2008) characterizes infectivity of knob-modified adenoviral vectors in glioma.
- Infection of DCs with adenovirus is limited because human DCs lack the native adenovirus receptor, coxsackie-adenovirus receptor (CAR). Ad5 carrying subgroup B Ad fibers are more potent than classical Ad5 for gene transfer and expression in human DCs (Rea, D., et al., J. Immunol. 166: 5236-44, 2001). In order to achieve meaningful therapeutic efficacy of adenovirus-based therapies, new approaches for infection of human DCs are required
- The present inventors have developed dendritic cells comprising modified adenoviral vectors which can infect dendritic cells with much greater infectivity compared to wild type adenovirus, as defined in the claims. In various configurations, an adenoviral vector of the present teachings can be a chimeric adenovirus which comprises a fiber comprising a tail, a shaft and a knob, wherein the knob is a porcine knob. The tail is an Ad5 tail, and the shaft is an Ad5 shaft, so that a fiber of the present teachings comprises a human Ad5 tail, a human Ad5 shaft, and a porcine knob. In some embodiments a chimeric adenovirus of the present teachings can bind dendritic cells using a receptor other than a CAR receptor. In some embodiments, a chimeric adenovirus of the present teachings can bind dendritic cells using a receptor other than an integrin receptor. In some embodiments, a chimeric adenovirus of the present teachings can bind dendritic cells using a receptor other than either a CAR receptor or an integrin receptor. In some embodiments, a chimeric adenovirus of the present teachings can comprise a fiber comprising a knob, wherein the knob comprises a galectin domain. In some configurations, a galectin domain can bind to one or more carbohydrates, such as a carbohydate comprising lactose and N-acetyl-lactosamine units. In some configurations, a galectin domain comprised by a knob of the present teachings can bind a carbohydrate structure selected from the group consisting of Galß1-4GlcNAcß1-3Galß1-4GlcNAcß1-3Galß1-4GlcNAc [tri(Nacetyl-lactosamine)], GlcNAcα1-4Galß1-4GlcNAcß1-3Galß1-4GlcNAcß1-3Galß1-4GlcNAc, Galß1-4GlcNAcß1-3Galß1-4Glc (lacto-N-neotetraose), Galα1-4Galß1-4GlcNAcß1-3Galß1-4Glc and Galß1-4GlcNAcβ1-3Galß1-3GlcNAc. In some configurations, a chimeric adenovirus comprising a fiber comprising a knob of the present teachings can bind a cell-surface glycoprotein comprising a carbohydrate structure such as, without limitation, Galß1-4GlcNAcß1-3Galß1-4GlcNAcß1-3Galß1-4GlcNAc[tri(Nacetyl-lactosamine)], GlcNAcα1-4Galß1-4GlcNAcß1-3Galß1-4GlcNAcß1-3Galß1-4GlcNAc, Galß1-(lacto-N-neotetraose), Galα1-4Galß1-4GlcNAcß1-3Galß1-4Glc, Galß1-4GlcNAcß1-3Galß1-3GlcNAc or a combination thereof.
- In various embodiments, a galectin domain comprised by a knob of the present teachings can bind to Lacto-N-neotetraose with a dissociation constant of 193±9µM, 3-aminopropyl-lacto-N-neotetraose with a dissociation constant of 303±4µM, 2-azidoethyl-di(N-acetyl-lactosamine) with a dissociation constant of 309±9µM, or 2-aminoethyl-tri(N-acetyl-lactosamine) with a dissociation constant of 308±40µM.
- In various aspects, cellular uptake by a dendritic cell of DNA of a chimeric Ad5 of the present teachings can be greater than that of a wild-type Ad5. In various aspects, cellular uptake by a dendritic cell of DNA of a chimeric Ad5 of the present teachings can be as great, or greater than, that of an adenovirus comprising a knob comprising an RGD sequence. In various aspects, cellular uptake by a dendritic cell of DNA of a chimeric Ad5 can be as great, or greater than, that of an adenovirus comprising a knob comprising an adenovirus comprising a type 35 fiber.
- In some embodiments, a dendritic cell of the present teachings can comprise a chimeric adenovirus-5 (Ad5) viral genome, wherein the chimeric Ad5 genome encodes a) a fiber comprising a tail, a shaft and a knob, wherein the knob is a porcine knob; and b) a promoter operably linked to a heterologous sequence encoding an antigen peptide.
- In some embodiments, the present teachings include ex vivo cell cultures comprising a dendritic cell, in particular a human dendritic cell, wherein the dendritic cell comprises nucleic acid sequences encoding Ad5sequences encoding a modified tail that includes a porcine knob sequence, as described herein .
- In some embodiments, the present teachings include a vaccine comprising an Ad5 modified to comprise a porcine knob, as well as an antigen peptide sequence such as, for example, an antigen peptide consisting of a linear peptide of from at least 8 up to 15 amino acids, for example 9 amino acids such as a peptide sequence set forth in table 1.
- In some embodiments, a vaccine of the present teachings can comprise a dendritic cell comprising Ad5 sequences encoding a modified tail protein, such as a tail protein comprising a porcine knob. In some configurations, a vaccine can comprise dendritic cells autologous to a subject such as a human subject, in which dendritic cells obtained from the subject can be grown and/or infected with a modified Ad5 of the present teachings in a cell culture ex vivo. Such cells can be administered to a subject, such as, for example, the donor of the dendritic cells, using methods well known to skilled artisans.
- In some embodiments of the present teachings, a subject such as a human can receive a vaccination through administration of a modified Ad5 of the present teachings. In some embodiments, a subject such as a human can receive a vaccination through administration of dendritic cells infected with an Ad5 of the present teachings. In some configurations, the dendritic cells can be autologous dendritic cells.
- In various configurations, an antigen peptide sequence can comprise or consist of from about 8, at least 8 up to 15, or about 15, contiguous amino acids. In some configurations, an antigen peptide sequence can comprise or consist of 9 contiguous amino acids. In various aspects, a peptide sequence can be that of a protein fragment, wherein the protein is a pathogen protein or a cellular protein, such as, for example, a protein expressed by a cancer cell. In some aspects, an antigen can comprise an antigen peptide such as that of an HLA-A restricted peptide or HLA-B restricted peptide. In some aspects, an antigen peptide can comprise or consist of a sequence as set forth in Table 1.
Table 1: Antigen Peptide Sequences Name Source Sequence Identification CMVpp65 Cytomegalovirus NLVPMVATV SEQ ID NO: 1 EBV BMLF I Ebstein-Barr virus GLCTLVAML SEQ ID NO: 2 fluM1 Influenza A virus GILGFVFTL SEQ ID NO: 3 G209-2M human melanoma IMDQVPFSV SEQ ID NO: 4 G280-9V human melanoma YLEPGPVTV SEQ ID NO: 5 - Various embodiments of the present teachings include a dendritic cell comprising a chimeric adenovirus-5 (Ad5) viral genome, wherein said chimeric Ad5 genome encodes a) a fiber comprising a tail, a shaft and a knob, wherein the knob is a porcine knob; and b) a promoter operably linked to a heterologous sequence encoding an immunizing antigen. As used herein, an "immunizing antigen" is a protein, polypeptide or oligopeptide that can stimulate an immune response in a body such as a human body.
- In some embodiments, the present teachings include an ex vivo cell culture comprising a dendritic cell comprising a chimeric Ad5 genome which encodes a) a fiber comprising a tail, a shaft and a knob, wherein the knob is a porcine knob; and b) a promoter operably linked to a heterologous sequence encoding an immunizing antigen. For example, an antigen that can bind a corresponding MHC class I heavy chain or MHC class I-like antigen presenting molecule such as CD1 (Altamirano, M.M., et al., Proc. Nat'1 Acad. Sci. 98: 3288-3293, 2001). In some aspects, an immunizing antigen can be that of a peptide which can be presented by an MHC class I molecule.
- In some embodiments, the present teachings include vaccines, wherein a vaccine comprises a dendritic cell comprising a chimeric Ad5 genome which encodes a) a fiber comprising a tail, a shaft and a knob, wherein the knob is a porcine knob; and b) a promoter operably linked to a heterologous sequence encoding an immunizing antigen. In various configurations, an immunizing antigen can be a peptide comprising or consisting of about 8, from 8 to 15, or about 15 contiguous amino acids. In various configurations, an immunizing antigen can be a peptide comprising or consisting of 9 contiguous amino acids, or about 9 contiguous amino acids. In various configurations, an immunizing antigen can be a peptide comprising or consisting of a sequence selected from the group consisting of NLVPMVATV (SEQ ID NO: 1), GLCTLVAML (SEQ ID NO: 2), GILGFVFTL (SEQ ID NO: 3), IMDQVPFSV (SEQ ID NO: 4) and YLEPGPVTV (SEQ ID NO: 5).
- Various embodiments of the present teachings include a chimeric Ad5 comprising: a) a fiber comprising a tail, a shaft and a knob, wherein the knob is a porcine knob; and b) a promoter operably linked to a heterologous sequence encoding an immunizing antigen. In various configurations, an immunizing antigen can comprise or consist of a sequence of a protein expressed by a cell at a level associated with a disease. In various configurations, an immunizing antigen can comprise or consist of a sequence of a protein expressed by a cancer cell at a level associated with a cancerous phenotype. In various configurations, an immunizing antigen can comprise or consist of about 8, from 8 to 15, or about 15 contiguous amino acids. In various configurations, an immunizing antigen can comprise or consist of 9, or about 9 contiguous amino acids. In various configurations, an immunizing antigen can comprise or consist of a peptide having a sequence selected from the group consisting of NLVPMVATV (SEQ ID NO: 1), GLCTLVAML (SEQ ID NO: 2), GILGFVFTL (SEQ ID NO: 3), IMDQVPFSV (SEQ ID NO: 4) and YLEPGPVTV (SEQ ID NO: 5).
-
-
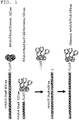
FIG. 1 illustrates the design of the Ad5Luc1-PK chimeric fiber. -
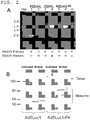
FIG. 2 illustrates molecular validation of Ad5Luc1-PK virions. -
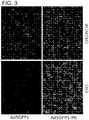
FIG. 3 illustrates fluorescence micrographs of CAR-negative CHO and CAR-positive CHO-hCAR cell lines. -
FIG. 4 illustrates that gene transfer of Ads-PK vectors is CAR-independent. -
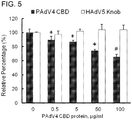
FIG. 5 illustrates that Ad5Luc1-PK uses carbohydrate binding domains for gene transfer. -
FIG. 6 illustrates that Ad5Luc1-PK-mediated gene delivery is mediated by glycans containing lactose. -
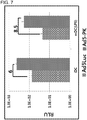
FIG. 7 illustrates Ad5Luc1-PK infectivity in murine dendritic cells. -
FIG. 8 illustrates Ad5Luc1-PK infectivity in Cynomolgus macaque dendritic cells. -
FIG. 9 illustrates Ad5Luc1-PK infectivity in human dendritic cells. -
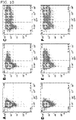
FIG. 10 illustrates Ad5GFP-PK infectivity in human dendritic cells. - The present inventors disclose a chimeric adenovirus and methods of transforming dendritic cells therewith. These methods, in various configurations, can enhance infectivity adenovirus towards human dendritic cells. In various embodiments, a porcine knob, which contains a galectin domain, is able to bind to carbohydrate moieties on the cell surface of dendritic cells. In some configurations, the carbohydrate moieties can comprise lactose and N-acetyl-lactosamine units. Furthermore, in some configurations the lactose and N-acetyl-lactosamine units can be Galß1-4GlcNAcß1-3Galß1-4GlcNAcß1-3Galß1-4GlcNAc [tri(Nacetyl-lactosamine)], GlcNAcα1-4Galß1-4GlcNAcß1-3Galß1-4GlcNAcß1-3Galß1-4GlcNAc, Galß1-4GlcNAcß1-3Galß1-4Glc (lacto-N-neotetraose), Galα1-4Galß1-4GlcNAcß1-3Galß1-4Glc, Galß1-4GlcNAcß1-3Galß1-3GlcNAc or a combination thereof. In various configurations the galectin domain can bind to Lacto-N-neotetraose with a dissociation constant of 193+9µM, to 3-aminopropyl-lacto-N-neotetraose with a dissociation constant of 303±4µM, to 2-azidoethyl-di(N-acetyl-lactosamine) with a dissociation constant of 309±9µM, or to 2-aminoethyl-tri(N-acetyl-lactosamine) with a dissociation constant of 308±40µM (the SPR response in µRIU).
- Monocytes and dendritic cells (DCs), such as freshly isolated human blood myeloid DCs, plasmacytoid DCs and monocyte-derived DCs lack CAR expression, but Langerhans cells and dermal DCs from skin express CAR. Furthermore, monocyte-derived DCs have lower CD46 expression then dermal DCs, Langerhans DCs, myeloid DCs, and plamacytoid DCs. Expression of CAR and CD46 (the subgroup C and B adenovirus receptors) on dendritic cell surfaces can be measured using FACS in cell lines. The correlation between infectivity enhancement and expression levels of CAR and CD46 can be determined.
- For example, infectivity of a panel of fiber-modified Ads that are CAR-independent can be compared in a variety of cancer cell types. The fiber-modified Ads can be examined to determine gene transfer to dendritic cell lines and compared with a tropism-modified Ad vector, Ad5/3, which encodes a fiber composed of the native Ad5 tail and shaft domains, but the fiber knob domain from Ad3.
- In some configurations, Ad5Luc1-PK and Ad5Luc1-CK1 fiber-modified adenovirus vectors of the present teachings can be Ad vectors with enhanced infectivity toward dendritic cells in comparison to an Ad5 comprising a wild-type knob. For example, three of the fiber-modified vectors, Ad5Luc1-PK, Ad5Luc1-CK1 and Ad5/3, can exhibit enhanced infectivity towards human dendritic cells compared to Ad5Luc1. In some configurations, Ad5Luc1-PK and Ad5Luc1-CK1 can have more than a 10-fold greater infectivity compared to that of Ad5/3. In some configurations, an Ad5Luc1-PK can have a greater infectivity compared to that of an Ad5 with a type 35 fiber described in Rea et al. (J. Immunol. 166: 5236-44, 2001).
- Methods and compositions described herein utilize laboratory techniques well known to skilled artisans. Such techniques can be found in laboratory manuals such as Sambrook, J., et al., Molecular Cloning: A Laboratory Manual, 3rd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 2001; Spector, D. L. et al., Cells: A Laboratory Manual, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1998; Harlow, E., Using Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1999.
- The following materials and methods are used in some experiments reported herein.
- Plasmid construction. A 1,750-bp region containing the PAdV-4 fiber knob and carbohydrate binding domains (amino acids 121-703) of the fiber protein was amplified from cell lysates containing wild type PAdV-4 virus obtained from the US Department of Agriculture National Veterinary Services Laboratory (Ames, Iowa) using the following primers: (PAd4 knob fwd) 5'-TGTGGACGGGGCCTGCTC-3'(SEQ ID: 6) and (PAd4 knob rev) 5'-TTTATTACAGTATCTGAGG-3'(SEQ ID: 7). Plasmid pSHAFT, a cloning vector containing the Ad5 fiber gene with the knob region deleted and replaced by a small linker containing Sma1 and EcoICR1 restriction sites (Krasnykh, V.N., et al., J. Virol. 70: 6839-46, 1996), was linearized by Sma1 and EcoICR1 digestion, leaving two blunt ends. Following gel purification, the PAdV-4 knob domain PCR product was ligated into linearized pSHAFT resulting in pSHAFT-PK and positive clones were screened for correct orientation via restriction enzyme digest. This plasmid contains the chimeric fiber gene encoding the complete Ad5 fiber shaft in-frame with the PAdV-4 knob domain. A stop codon and polyadenylation sequence is present at the 3' end. The chimeric fiber gene in pSHAFT was digested with Ncol and MunI to liberate the DNA fragment containing the carboxy terminus of the HAdV-5 shaft and the PAdV-4 knob domain. This fragment was ligated into the NcoI-MunI-digested fiber shuttle vector pNEB.PK.3.6 (Krasnykh, V.N., et al., J. Virol. 70: 6839-46, 1996), resulting in pNEB.PK.3.6-PK.
- Generation of recombinant adenovirus. The recombinant Ad5Luc1-PK genome containing the chimeric PAdV-4 fiber gene was derived by homologous recombination in E. coli BJ5183 with SwaI-linearized rescue plasmid pVK700 (Belousova, N., et al,. J. Virol. 76: 8621-31, 2002) and the fiber-containing PacI-KpnI-fragment of pNEB.PK.3.6-PK, essentially as described (Krasnykh, V., et al., J. Virol. 72: 1844-52, 1998). Plasmid pVK700 is derived from pTG3602 (Chartier, C., et al., J. Virol. 70: 4805-10, 1996), but contains an almost complete deletion of the fiber gene and contains a firefly luciferase reporter gene driven by the cytomegalovirus immediate early promoter in place of the E1 region. The recombinant genome of Ad5GFP1-PK containing the chimeric PAdV-4 fiber gene was derived by homologous recombination in E. coli BJ5183 with fiber shuttle plasmid pKan3.1-PK which contains the same chimeric fiber gene as pNEB.PK.3.6-PK described above, and SwaI-linearized rescue plasmid pVK900 (Murakami, M., et al., Virol. 407: 196-205, 2010). Plasmid pVK900 is a fiber-deleted HAdV-5 genome plasmid essentially the same as pVK700 except that EGFP is encoded in the E1 region (supplied by Victor Krasnykh, University of Texas MD Anderson Cancer Center). All genomic clones were sequenced and analyzed by PCR prior to transfection of HEK 293 cells. Ad5Luc1 is a replication defective E1-deleted Ad vector containing a firefly luciferase reporter gene driven by a cytomegalovirus promoter (Krasnykh, V., et al., J. Virol. 75: 4176-83, 2001). All vectors were propagated on HEK 293 cells and purified by equilibrium centrifugation in CsCl gradients by standard protocols. Viral particle concentration was determined at 260 nm by the method of Maizel et al. (Maizel, J.V., et al., Virol. 36: 115-25, 1968) by using a conversion factor of 1.1x1012 viral particles/absorbance unit.
- DC medium is RPMI + 2mM glutamine + HEPES + non-essential amino-acids + Pen/Strep + 1% AB sera.
- Peptide pulsing medium is Stemline + 2mM L-glutamine + Pen/Strep + 1% AB sera A-DC generation.
- 1) Fresh PBMC are isolated from blood, buffy coats or leukopheresis as instructed on protocol. Frozen PBMC (yield ∼ 2×108 cells/vial) are thawed, counted. Fresh or frozen PBMC are adjusted to 5x106/ml in DC media. Transfer 40ml (2x108 cells) to a T175 flask and incubate for 2h at 37°C.
- 2) Remove non-adherent cells and transfer to a 50ml conical tube. Wash T175 gently 2X with 25 ml PBS and transfer to another 50 ml conical tube.
- a. These PBL can be discarded or can be used as a source of T cells (see Primary CTL stimulation protocol).
- 3) To T175 ml flask add 30 ml DC media containing 100ng/ml GM-CSF (Leukine) and 20ng/ml IL-4 (CellGenix). Incubate cells at 37°
C 5% CO2. - 4) On
day 3 feed cells with 10ml DC medium containing 100ng/ml GM-CSF and 20ng/ml IL-4. - 5) Harvest cells on
day 6 by gently rocking flask back and forward; collect non-adherent and loosely adherent cells and transfer to a 50ml conical tube. Wash T175 flask gently 2X with 25ml PBS and transfer to another 50 ml conical tube. Spin at 1500 RPM for 5 min, aspirate supernatant, resuspend cells in DC medium (1-2 ml/tube), pool cells and count. DC are adjusted to 2x106/ml in DC media containing 200ng/ml GM-CSF and 40ng/ml IL-4 (2X concentration)
Yield: frozen PBMC ∼2-5x 106 DC/flask; from fresh cells ∼107 DC/flask.
B-CD40L/IFN-γ maturation - 6) Immature DC consist of cells grown in GM-CSF and IL-4; diluted 1:1 vol with DC media.
- 7) J558-muCD40L or K562-huCD40L are used for maturation. Cells are irradiated (5,000 RADS for J558 or 10,000RADS for K562), spun and resuspend in DC media at a 4x105 cells/ml in DC media.
- 8) Mix 1:1 vol of DC to CD40L-expressing cells, up to 4ml per well of a 6 well tray (Ultra-low #3471). Ratio of DC to CD40L cells is 5DC:1CD40L-expressing cell.
- 9) Add 100u/ml IFN-g (Actimmune). Incubate for 24-48h
- 10) Harvest DC, save undiluted supernatant for assessment of cytokine production. Wash cells once, and resuspend in Peptide pulsing media if cells are to be use in stimulation of T cells.
- PCR Analysis of the Fiber Region. Genomic DNA contained in Ad5Luc1, Ad5Luc1-PK and PAdV-4 viral particles was used as templates for PCR amplification of fiber genes using a HAdV-5-specific primer set: (fwd) 5'-CAGCTCCATCTCCTAACTGT-3'(SEQ ID: 8) and (rev) 5'-TTCTTGGGCAATGTATGAAA-3'(SEQ ID: 9) and a PAdV-4-specific primer set: (fwd) 5'-TGTGGACGGGGCCTGCTC-3' (SEQ ID: 10) and (rev) 5'-TTTATTACAGTATCTGAGG-3'(SEQ ID: 11). Wild type PAdV-4 virus was used as a positive control.
- Western Blot Analysis. Purified Ad virions (5.0 x 109) were diluted in Laemmli buffer and incubated at room temperature (unboiled samples) or 95°C (boiled samples) for 10 minutes and loaded onto a 4-20% gradient SDS-polyacrylamide gel (Bio-Rad, Hercules, CA). Following electrophoretic separation, Ad capsid proteins were electroblotted onto a PVDF membrane and detected with a 4D2 monoclonal anti-fiber tail primary antibody diluted 1/3000 (Lab Vision, Freemont CA). Immunoblots were developed by addition of a secondary horseradish peroxidase-conjugated anti-mouse immunoglobulin antibody at a 1/3000 dilution (Dako Corporation, Carpentaria, CA), followed by incubation with 3-3'-diaminobenzene peroxidase substrate, DAB, (Sigma Chemical Company, St. Louis, MO).
- Ad-Mediated Gene Transfer Assays. Cells were plated in 24-well plates and were transduced for 1 hour at 37°C with each Ad vector diluted to 100-300 viral particles/cell in 500 µL of transduction media containing 2% FBS. Following the incubation, virus-containing media was replaced with fresh media containing 2% FBS and cells were maintained at 37°C in an atmosphere containing 5% CO2. Cells were harvested 24 hours post-transduction in passive lysis buffer and gene transfer was determined using a luciferase activity assay system (Promega, Madison, WI). Fluorescence microscopy was performed with an inverted IX-70 microscope (Olympus) using a 20X objective. Cells were infected with Ad5GFP1-PK for 24 hours prior to imaging.
- For experiments assessing the competitive inhibition containing of vector binding to cells, recombinant fiber knob protein (Glasgow, J.N., et al., Virol. 324: 103-16, 2004) at 0.5, 5.0 and 50 µg/ml final concentration or recombinant PAdV-4 carbohydrate binding domain (CBD) protein [20] at 0.5, 5.0, 50 and 100 µg/ml was incubated with various cell lines at 37°C in media containing 2% FBS for 20 minutes prior to the addition of HAdV-5 vectors. Following transduction with Ad vectors, cells were rinsed with media to remove unbound virus and blocking proteins, and were maintained at 37°C in an atmosphere containing 5% CO2.
- To inhibit glycosylation of cellular proteins, chemical inhibitors of glycosylation were used. CHO-Pro5 cells were incubated with medium containing 10 µg/ml swainsonine (Sigma, Saint Louis, MO., S8195) and/or 1 µg/ml benzyl-α-GalNAc (Sigma B4894) for 24 hr at 37 °C, followed by addition of Ad vectors in media containing 2% FBS.
- The present teachings including descriptions provided in the Examples that are not intended to limit the scope of any claim or aspect. Unless specifically presented in the past tense, an example can be a prophetic or an actual example. The following non-limiting examples are provided to further illustrate the present teachings. Those of skill in the art should, in light of the present disclosure, appreciate that many changes can be made in the specific embodiments that are disclosed and still obtain a like or similar result without departing from the scope of the present teachings.
- This example illustrates the generation of a fiber-modified HAdV-5 vector containing the PAdV-4 knob and carbohydrate binding domains.
- The fiber protein of PAdV-4 NADC-1 is comprised of a homotrimer of 703 amino acids (
FIG. 1 ). Predicted functional domains include a tail domain (residues 1-37) containing a penton interaction sequence, a short shaft domain (residues 38-120) with six predicted triple beta-spiral repeats (van Raaij, M.J., et al. Virol. 262: 333-43, 1999) and a fiber knob domain homologous to other Ad fiber knob domains (residues 121-287) (Guardado-Calvo, P., et al. Acta. Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 65: 1149-52, 2009). This fiber also contains a unique C-terminal domain composed of two tandem carbohydrate binding domains (CBDs) (residues 393-681) that bind carbohydrates containing lactose and N-acetyl-lactosamine units (Guardado-Calvo, P., et al. J. Virol. 84: 10558-68, 2010). Almost all mastadenoviruses contain a conserved threonine-leucine-tryptophan-threonine (TLWT) (SEQ ID NO: 12) motif at the N-terminus of the fiber knob domain, and in human Ad2 and Ad5 a flexible region separating the shaft and the knob domains precedes this motif (van Raaij, M.J., et al. Nature. 401: 935-8, 1999). The modular fiber structure was used to substitute the coding region of the PAdV-4 knob and CBD domains for the HAdV-5 fiber knob sequence while retaining the TLWT motif common to both fibers (Renaut, L., et al. Virol. 321: 189-204, 2004). A recombinant E1-deleted HAdV-5 genome (Ad5Luc1-PK) containing the chimeric HAdV-5 shaft/PAdV-4 fiber gene and a firefly luciferase reporter gene controlled by the CMV immediate early promoter/enhancer were constructed. The Ad5Luc1-PK vector was rescued following transfection of HEK 293 cells and large-scale preparations of Ad5Luc1-PK and the Ad5Luc1 control vector were produced and purified by double CsCl gradient centrifugation. Ad5Luc1-PK viral particle concentration in full preparations ranged from 1.2 x 1011 to 1.25 x 1012 viral particles/ml, similar to that of the Ad5Luc1 control vector containing the HAdV-5 wild type fiber. The Ad5Luc1 vector is isogenic to Ad5Luc1-PK except for the fiber protein. - The fiber genotypes of Ad5Luc1 and Ad5Luc1-PK vectors were confirmed via diagnostic PCR using primer pairs specific for the fiber knob domain and genomes from purified virions as PCR templates. Genomic DNA from wild type PAdV-4 was used as a positive control. The expected PCR products were observed for the wild type HAdV-5 fiber knob domain (530 bp) and the PAdV-4 fiber knob and CBD domains (1,750 bp) (
FIG. 2A ). - SDS-PAGE was performed followed by western blot analysis on purified viral particles to verify that Ad5Luc1-PK virions contain correctly trimerized chimeric fiber proteins (
FIG. 2B ). Blots were probed with a monoclonal primary antibody (4D2) directed against the fiber tail domain common to both HAdV-5 and chimeric fiber molecules. In samples that were not heat denatured (FIG. 2B , unboiled) bands were observed at 183 kDa and an estimated 250 kDa, corresponding to trimers of the HAdV-5 fiber and chimeric fibers, respectively. Further, bands in boiled samples resolved at apparent molecular masses of 60 kDa for the wild type HAdV-5 fiber and 90 kDa for the chimeric fiber in Ad5Luc1-PK, representing fiber monomers. - This example illustrates that Ad5-PK vector infectivity is independent of CAR.
- High resolution crystal structure analysis has shown that the A-B loop in the N-terminal region of the PAdV-4 knob domain is structurally similar to the A-B loop in the CAR-binding domain in adenoviruses that use CAR as a primary receptor, but that only one CAR-binding residue is conserved (Guardado-Calvo, P., et al. J. Virol. 84: 10558-68, 2010). To investigate whether Ad5-PK vectors exhibit CAR-dependent tropism, gene transfer assays were performed in two cell lines with markedly different levels of CAR expression: CAR-deficient Chinese hamster ovary (CHO) cells and a CHO-derived cell line, CHO-hCAR, which stably expresses human CAR (Bergelson, J.M., et al. Science. 275: 1320-3, 1997). CHO and CHO-hCAR cells were infected with the Ad5GFP control vector and Ad5GFP1-PK, a vector isogenic to Ad5Luc1-PK except that the firefly luciferase reporter gene was replaced with green fluorescent protein, GFP. Fluorescence microscopy showed GFP expression in CHO-hCAR cells but not in CAR-deficient CHO cells (
FIG. 3 ), consistent with native HAdV-5 tropism. In contrast, Ad5Luc1-PK-mediated GFP gene delivery does not depend on CAR expression, as similar number of GFP-positive cells were observed in both the CHO and CHO-hCAR cell lines. Similar gene transfer assays were performed using luciferase-expressing Ad5Luc1 and Ad5Luc1-PK vectors to quantify differences in gene delivery based solely on CAR expression. Ad5Luc1 exhibited the expected CAR-dependent tropism as demonstrated by an 80-fold increase in luciferase activity in CHO-hCAR cells versus CHO cells (FIG. 4A ). In contrast, Ad5Luc1-PK provided robust gene delivery to both cell lines. In addition, competitive inhibition of CAR binding with recombinant HAdV-5 knob proteins (50 µg/ml) inhibited over 96% of Ad5Luc1 gene transfer to CHO-hCAR cells, but did not inhibit the gene transfer of Ad5Luc1-PK (FIG. 4B ). - This example illustrates that Ad5Luc1-PK uses carbohydrate binding domains for gene transfer.
- To determine whether the CBDs in the chimeric fiber of Ad5Luc1-PK participate in cellular attachment, competitive inhibition assays were performed using a recombinant protein consisting of the tandem PAdV-4 CBDs (residues 393-703 of the PAdV-4 fiber protein) or recombinant HAdV-5 fiber knob protein as a negative control. Addition of PAdV-4 CBD protein during infection caused a dose-dependent inhibition of Ad5Luc1-PK-mediated gene transfer with a maximum inhibition of 35% at 100 µg/ml (
FIG. 5 ), indicating that the CBDs in the chimeric fiber are responsible for cellular attachment during infection. - The CBDs within the PAdV-4 fiber protein bind to lactose, N-actyl-lactosamine and poly-N-acetyl-lactosamine in order of increasing affinity (Guardado-Calvo, P., et al., J. Virol. 84: 10558-68, 2010). However, whether the CBDs in the PAdV-4 chimeric fiber recognize these glycans and use them as a means for viral transduction is not known. Gene transfer assays were performed in the CHO-Lec8 cells which contains mutations in the UDP-galactose transporter/translocase (UGT) gene (Deutscher, S.L., et al., J. Biol. Chem. 261: 96-100, 1986; Stanley, P., Mol. Cell. Biol. 5: 923-29, 1985; Oelmann, S., et al., J. Biol. Chem. 276: 26291-300, 2001), and lack the ability to galactosylate glycoproteins and therefore produce glycoproteins with truncated carbohydrate chains that lack lactose, N-acetyl-lactosamine and poly-N-acetyl-lactosamine. The level of Ad5Luc1 gene delivery was unchanged between CHO-Lec8 cells and the control CHO-Pro5 cells that exhibits normal glycosylation (
Fig 6A ). In contrast, Ad5Luc1-PK gene delivery to CHO-Lec8 cells was reduced by 64% compared to the control CHO-Pro5 cells, showing that the presence of lactose-containing glycans at the cell surface is critical for Ad5Luc1-PK infectivity. - There are two major types of carbohydrate chains on glycoproteins: N-linked glycans linked to asparagine residues and O-linked glycans linked to serine or threonine (Li, H., and d'Anjou, M., Curr. Opin. Biotechnol. 20: 678-84, 2009; Schwarz, F., and Aebi, M., Curr. Opin. Struct. Biol. 21: 576-82, 2011; Dwek, R.A., Chem. Rev. 96: 683-720, 1996). To further investigate the nature of the glycans recognized during Ad5Luc1-PK infection, gene transfer assays were performed following incubation of CHO-Pro5 cells with inhibitors of N-linked glycan synthesis (swainsonine, 10 µg/ml), or O-linked glycan synthesis (benzyl-α-GalNAc, 1 µg/ml) (Elbein, A.D., et al., Proc. Natl. Acad. Sci. USA. 78: 7393-7, 1989; Kuan, S.F., et al., J. Biol. Chem. 264: 19271-7, 1989). The addition of these inhibitors singly or in combination to CHO-Pro5 cells did not alter levels of Ad5Luc1 gene transfer (
Fig. 6B ). In contrast, Ad5Luc1-PK gene transfer was blocked 35% by benzyl-α-GalNAc pre-treatment, with a minimal (<10%) reduction by swainsonine. Similar results were also observed in A549 cells pre-treated with these inhibitors, suggesting that O-linked cell-surface glycans may be preferred by Ad5Luc1-PK for infection. Collectively, these data show that the CBDs in the chimeric fiber protein of Ad5Luc1-PK directly participate in cellular attachment and that infection is highly dependent on the presence of lactose and/or N-acetyl-lactosamine-containing glycans, consistent with a novel, glycan-mediated cell entry pathway. - This example illustrates enhanced infectivity of Ad5Luc1-PK in murine dendritic cells.
- To analyze infectivity of Ad5Luc1-PK, gene transfer assays were performed in immature (iDC) and mature (LPS-treated) (mDC) murine dendritic cells. In these experiments, iDC and mDC were infected with the Ad5Luc1 and Ad5Luc1-PK. Ad5Luc1-PK demonstrated a 6-fold increase in luciferase activity in iDC compared to Ad5Luc1, while in mDC Ad5Luc1-PK demonstrated a 8.5 fold increase in luciferase activity as compared to Ad5Luc1 (
FIG. 7 ). - This example illustrates enhanced infectivity of Ad5Luc1-PK in Cynomolgus macaque dendritic cells.
- To analyze infectivity of Ad5Luc1-PK in Cynomolgus macaque dendritic cells, gene transfer assays were performed in C. macaque dendritic cells. In these experiments, C. macaque dendritic cells were infected with Ad5Luc1, Ad5/3 and Ad5Luc1-PK. Ad5Luc1-PK demonstrated an increase in luciferase activity compared to Ad5/3 and Ad5Luc1 (
FIG. 8 ). Ad5Luc1 (FIG. 7 ). - This example illustrates enhanced infectivity of Ad5Luc1-PK in human dendritic cells.
- To analyze infectivity of Ad5Luc1-PK in human dendritic cells, gene transfer assays were performed in immature (iDC) and mature (mDC) human dendritic cells. In these experiments, human dendritic cells were infected with Ad5Luc1, Ad5/3 and Ad5Luc1-PK. In iDC, Ad5/3 demonstrated a 6-fold increase in luciferase activity as compared to Ad5Luc1, while in mDC, Ad5/3 demonstrated a 30-fold increase in luciferase activity as compared to Ad5Luc1 (
FIG. 9 ). In iDC, Ad5Luc1-PK demonstrated an 8-fold increase in luciferase activity compared to Ad5Luc1, while in mDC, Ad5Luc1-PK demonstrated a 40-fold increase in luciferase activity compared to Ad5Luc1 (FIG. 9 ). - This example illustrates FACS analysis of enhanced infectivity of Ad5GFP1-PK in immature human dendritic cells.
- To analyze infectivity of Ad5GFP1-PK, gene transfer assays were performed in human dendritic cells. In these experiments, immature human dendritic cells were infected with the Ad5GFP control vector (
FIG. 10B, FIG. 10E ) and Ad5GFP1-PK (FIG. 10C, FIG 10F ), a vector isogenic to Ad5Luc1-PK except that the firefly luciferase reporter gene was replaced with green fluorescent protein, GFP. Uninfected immature human dendritic cells are shown inFIG. 10A and FIG. 10D . Flow cytometry was performed using standard procedures at 24 (FIG. 10 A, B, C ) and 48 (FIG. 10 D, E, F ) hours after infection. After 24 hours of infection, Ad5GFP infected about 10% of the dendritic cell population (FIG. 10B ) as compared to control (FIG. 10A ), while Ad5GFP-PK infected 56.5% of the dendritic cell population (FIG. 10C ) as compared to control (FIG. 10A ). After 48 hours of infection, Ad5GFP infected about 20% of the dendritic cell population (FIG. 10E ) as compared to control (FIG. 10D ), while Ad5GFP-PK infected 67.7% of the dendritic cell population (FIG. 10F ) as compared to control (FIG. 10D ). - This example illustrates that xeno-knob-modified adenovirus can have enhanced infectivity in human dendritic cells compared to other adenoviruses.
- To analyze the efficiency of gene delivery of xenotype-modified adenoviral vectors on dendritic cell lines in vitro, five xeno-knob fiber-modified vectors can be used. In vectors Ad5Luc1-MK, Ad5Luc1-PK, Ad5Luc1-CK1 and Ad5Luc1-CK2 the native Ad5 fiber knob domain can be replaced by the corresponding fiber knob domain from
mouse adenovirus type 1, porcine adenovirus, orcanine adenovirus type 1 or 2, respectively. In the Ad5Luc1-OvF, the entire Ad5 fiber can be replaced with the fiber from ovine adenovirus 7. A control vector, Ad5Luc1 comprising the native Ad5 fiber, and a fiber-modified vector, Ad5/3, comprising ahuman Ad serotype 3 fiber knob domain can be used in this example. Both of these vectors can serve as controls to which other fiber-modified Ad vectors can be compared. All Ad vectors can be isogenic except for the fiber gene. The luciferase transgene activity of fiber-modified vectors can be compared to the Ad5Luc1 control vector in dendritic cell lines; plasmacytoid DCs, monocyte-derived DCs, migratory DCs, and lymphoid DCs. Luciferase expression levels can be shown as relative light units (RLU) normalized to that of Ad5Luc1 (Ad5Luc1=100%). Three of the vectors, Ad5Luc1-MK, Ad5Luc1-OvF, and Ad5Luc1-CK2 can augment gene delivery less than 2.5-fold above Ad5Luc1 in dendritic cell lines. In various dendritic cells, the Ad5Luc1-CK1 vector can provide an increase in luciferase transgene activity of at least 7-fold compared to an Ad5Luc1 control. In various dendritic cells, the Ad5Luc1-CK1 vector can provide an increase in luciferase transgene activity of up to 50-fold compared to an Ad5Luc1 control. The Ad5Luc1-PK vector can have an increase in gene delivery from 10-fold up to 20-fold above Ad5Luc1 in the dendritic cell lines. An Ad5/3 vector can enhance gene delivery 14-fold up to 50-fold in dendritic cell lines compared to an Ad5Luc1. Infectivity levels of these two viral vectors (comprising the porcine knob or the canine knob) in the four dendritic cell lines can be similar to or can exceed that of Ad5/3, which had previously been shown to act as a successful fiber-modified vector for enhancement of viral infectivity in human dendritic cells. - This example demonstrates that xeno-knob modified infectivity can be independent of adenovirus native receptor, coxsackie-adenovirus-receptor (CAR).
- To analyze the infectivity independent of CAR on dendritic cell lines in vitro, Ad5Luc1-PK and Ad5Luc1 can be used, where the Ad5Luc1 can be used as a control. The luciferase transgene activity of Ad5Luc1-PK vector can be compared to the Ad5Luc1 control vector in two dendritic cell lines, monocyte-derived DCs and monocyte-derived DCs that constitutively express high levels of CAR. The luciferase expression levels can be shown as relative light units (RLU). After infection of Ad5Luc, into two dendritic cell lines, the luciferase transgene activity can increase up to 60 fold in monocyte-derived DCs that constitutively express CAR when compared to monocyte-derived dendritic cells. After infection of Ad5Luc1-PK, into monocyte-derived DCs, the luciferase transgene activity can be more then 60-fold increase when compared to Ad5Luc luciferase transgene activity in monocyte-derived DCs. Furthermore, no increase in luciferase transgene activity could (would) be observed in monocyte-derived cells that constitutively express CAR infected with Ad5Luc1-PK, as compared to monocyte-derived DCs infected with Ad5Luc1-PK.
- To further demonstrate that xeno-modified infectivity can be independent of CAR, an Ad5 Knob Inhibition assay can be performed. Monocyte-derived DCs that constitutively overexpress CAR can be infected with Ad5Luc or Ad5Luc1-PK where increasing amounts from 0-100 µg/mL of Ad5 Knob can be added to the infection mixture. The luciferase expression levels can be shown as relative light units (RLU). After infection of Ad5Luc into monocyte-derived DCs overexpressing CAR, there can be a 3-fold decrease in luciferase transgene activity when comparing infection with no Ad5 Knob present and infection with 10 µg/mL of Ad5 Knob present. Infection with 50 µg/mL or 100 µg/mL of Ad5 Knob present can further decrease the luciferase transgene activity. After infection of Ad5Luc1-PK into monocyte-derived DCs overexpressing CAR, there may be no effect on luciferase transgene activity when comparing infection with no Ad5 Knob present and infection with 10 µg/mL of Ad5 Knob present. Infection with increasing amounts of Ad5 Knob, such as 50 µg/mL or 100 µg/mL, may not affect luciferase transgene activity in DCs overexpressing CAR infected with Ad5Luc1-PK.
- This example demonstrates that the porcine knob can bind to lactose and N-acetyl-lactosamine units.
- The affinity of porcine adenovirus knob domain for Lacto-N-neotetraose, 3-aminopropyl-lacto-N-neotetraose, 2-azidoethyl-di(N-acetyl-lactosamine), and 2-aminoethyl-tri(N-acetyl-lactosamine) can be assessed by surface plasmon resonance experiments. The galectin can be bound to a sensor chip, and the oligosaccharides can be injected at various concentrations to monitor their binding to the galectin domain. Sensorgrams can be evaluated via steady-state analysis to yield the corresponding isotherms from which the dissociation constants can be calculated. Surface plasmon resonance response can be calculated in µRIU. Lacto-N-neotetraose binds weakly to the galectin domain with a dissociation constant of 193±9µM; while 3-aminopropyl-lacto-N-neotetraose, 2-azidoethyl-di(N-acetyl-lactosamine), 2-aminoethyl-tri(N-acetyl-lactosamine) bind stronger to the galectin domain with dissociation constants of 303±4µM, 309±9µM, and 308±40µM, respectively.
-
- <110> Washington University Curiel, David T
- <120> PORCINE KNOB XENOTYPE CHIMERIC ADENOVIRAL VECTOR FOR DENDRITIC CELL INFECTION
- <130> 1001-0560
- <150>
US 61/576,116
<151> 2011-12-15 - <160> 12
- <170> PatentIn version 3.5
- <210> 1
<211> 9
<212> PRT
<213>Human herpesvirus 5 - <400> 1
- <210> 2
<211> 9
<212> PRT
<213> Human herpesvirus 4 - <400> 2
- <210> 3
<211> 9
<212> PRT
<213> Influenza A virus - <400> 3
- <210> 4
<211> 9
<212> PRT
<213> Homo sapiens - <400> 4
- <210> 5
<211> 9
<212> PRT
<213> Homo sapiens - <400> 5
- <210> 6
<211> 18
<212> DNA
<213> Porcine adenovirus 4 - <400> 6
tgtggacggg gcctgctc 18 - <210> 7
<211> 19
<212> DNA
<213> Porcine adenovirus 4 - <400> 7
tttattacag tatctgagg 19 - <210> 8
<211> 20
<212> DNA
<213>Human adenovirus type 5 - <400> 8
cagctccatc tcctaactgt 20 - <210> 9
<211> 20
<212> DNA
<213>Human adenovirus type 5 - <400> 9
ttcttgggca atgtatgaaa 20 - <210> 10
<211> 18
<212> DNA
<213> Porcine adenovirus 4 - <400> 10
tgtggacggg gcctgctc 18 - <210> 11
<211> 19
<212> DNA
<213> Porcine adenovirus 4 - <400> 11
tttattacag tatctgagg 19 - <210> 12
<211> 4
<212> PRT
<213> Mastadenovirus 4 - <400> 12
Claims (14)
- A dendritic cell comprising:a) chimeric adenovirus-5 (Ad5) viral genome encoding:1) a fiber comprising a human Ad5 tail, a human Ad5 shaft and a porcine knob of porcine adenovirus type 4 (PAdV-4); and2) a promoter operably linked to a heterologous sequence encoding an immunizing antigen.
- A dendritic cell in accordance with claim 1, wherein the immunizing antigen consists of a portion of a protein expressed by a cancer cell at a level associated with a cancerous phenotype.
- A dendritic cell in accordance with claim 1, wherein the immunizing antigen consists of 8, from 8 to 15, or 15 contiguous amino acids.
- A dendritic cell in accordance with claim 1, wherein the immunizing antigen consists of 9 contiguous amino acids.
- A dendritic cell in accordance with claim 1, wherein the immunizing antigen consists of a sequence selected from the group consisting of NLVPMVATV (SEQ ID NO: 1), GLCTLVAML (SEQ ID NO: 2), GILGFVFTL (SEQ ID NO: 3), IMDQVPFSV (SEQ ID NO: 4), YLEPGPVTV (SEQ ID NO: 5).
- An ex vivo cell culture comprising a dendritic cell of claim 1.
- A vaccine comprising a dendritic cell of claim 1.
- A vaccine in accordance with claim 7, wherein the immunizing antigen is a peptide comprising 8, from 8 to 15, or 15 contiguous amino acids;
wherein, optionally, the immunizing antigen is a peptide comprising 9 contiguous amino acids. - A vaccine in accordance with claim 7, wherein the immunizing antigen is a peptide consisting of 8, from 8 to 15, or 15 contiguous amino acids;
wherein, optionally, the immunizing antigen is a peptide consisting of 9 contiguous amino acids. - A vaccine in accordance with claim 7, wherein the immunizing antigen is a peptide comprising a sequence selected from the group consisting of NLVPMVATV (SEQ ID NO: 1), GLCTLVAML (SEQ ID NO: 2), GILGFVFTL (SEQ ID NO: 3), IMDQVPFSV (SEQ ID NO: 4), YLEPGPVTV (SEQ ID NO: 5).
- A vaccine in accordance with claim 7, wherein the immunizing antigen is a peptide consisting of a sequence selected from the group consisting of NLVPMVATV (SEQ ID NO: 1), GLCTLVAML (SEQ ID NO: 2), GILGFVFTL (SEQ ID NO: 3), IMDQVPFSV (SEQ ID NO: 4), YLEPGPVTV (SEQ ID NO: 5).
- A method of transforming dendritic cells in a cell culture, the method comprising adding to a cell culture comprising dendritic cells, the chimeric adenovirus-5 (Ad5) comprising:1) a fiber comprising a human Ad5 tail, a human Ad5 shaft and a porcine knob of porcine adenovirus type 4 (PAdV-4); and2) a promoter operably linked to a heterologous sequence encoding an immunizing antigen.
- A method in accordance with claim 12, wherein the porcine knob comprises a galectin domain that binds to carbohydrates comprising lactose and N-acetyl-lactosamine units.
- A method in accordance with claim 12, wherein the porcine knob comprises a galectin domain that binds a carbohydrate structure selected from the group consisting of Galß1-4GlcNAcß1-3Galß1-4GlcNAcß1-3Galß1-4GlcNAc [tri(Nacetyl-lactosamine)], GlcNAcα1-4Galß1-4GlcNAcß1-3Galß1-4GlcNAcß1-3Galß1-4GlcNAc,Galß1-4GlcNAcß1-3Galß1-4Glc (lacto-N-neotetraose), Galα1-4Galß1-4GlcNAcß1-3Galß1-4Glc and Galß1-4GlcNAcß1-3Galß1-3GlcNAc.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US201161576116P | 2011-12-15 | 2011-12-15 | |
| PCT/US2012/069884 WO2013090806A2 (en) | 2011-12-15 | 2012-12-14 | Porcine knob xenotype chimeric adenoviral vector for dendritic cell infection |
Publications (3)
| Publication Number | Publication Date |
|---|---|
| EP2798069A2 EP2798069A2 (en) | 2014-11-05 |
| EP2798069A4 EP2798069A4 (en) | 2015-11-18 |
| EP2798069B1 true EP2798069B1 (en) | 2017-03-29 |
Family
ID=48613374
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| EP12858372.1A Not-in-force EP2798069B1 (en) | 2011-12-15 | 2012-12-14 | Porcine knob xenotype chimeric adenoviral vector for dendritic cell infection |
Country Status (3)
| Country | Link |
|---|---|
| US (1) | US9267153B2 (en) |
| EP (1) | EP2798069B1 (en) |
| WO (1) | WO2013090806A2 (en) |
Families Citing this family (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US9267153B2 (en) * | 2011-12-15 | 2016-02-23 | Washington University | Porcine knob xenotype chimeric adenoviral vector for dendritic cell infection |
| WO2014153204A1 (en) | 2013-03-14 | 2014-09-25 | Salk Institute For Biological Studies | Oncolytic adenovirus compositions |
| WO2017147269A1 (en) | 2016-02-23 | 2017-08-31 | Salk Institute For Biological Studies | Exogenous gene expression in therapeutic adenovirus for minimal impact on viral kinetics |
| AU2017222568B2 (en) | 2016-02-23 | 2020-09-10 | Salk Institute For Biological Studies | High throughput assay for measuring adenovirus replication kinetics |
| CN110062630A (en) | 2016-12-12 | 2019-07-26 | 萨克生物研究学院 | Cancer target synthesizes adenovirus and application thereof |
| WO2020033876A1 (en) * | 2018-08-10 | 2020-02-13 | Nantbio, Inc. | Cellular adjuvants for viral infection |
Family Cites Families (20)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5962311A (en) * | 1994-09-08 | 1999-10-05 | Genvec, Inc. | Short-shafted adenoviral fiber and its use |
| US7232899B2 (en) | 1996-09-25 | 2007-06-19 | The Scripps Research Institute | Adenovirus vectors, packaging cell lines, compositions, and methods for preparation and use |
| AU744252B2 (en) * | 1998-02-06 | 2002-02-21 | Uab Research Foundation, The | Adenovirus vector containing a heterologous peptide epitope in the hi loop of the fiber knob |
| WO2000067576A1 (en) * | 1999-05-12 | 2000-11-16 | The Uab Research Foundation | Infectivity-enhanced conditionally-replicative adenovirus and uses thereof |
| AU2001295193A1 (en) | 2000-05-01 | 2001-11-12 | Novartis Ag | Vectors for ocular transduction and use thereof for genetic therapy |
| EP1233059A1 (en) | 2001-02-09 | 2002-08-21 | ARTEMIS Pharmaceuticals GmbH | Influenza viruses with enhanced transcriptional and replicational capacities |
| AU2002250081A1 (en) * | 2001-02-14 | 2002-08-28 | Uab Research Foundation | Combined transductional and transcriptional targeting system for improved gene delivery |
| AU2002363322A1 (en) | 2001-10-26 | 2003-05-19 | Large Scale Biology Corporation | Endothelial cell derived hemotopoietic growth factor |
| AU2003247128A1 (en) | 2002-07-10 | 2004-02-02 | Transgene S.A. | Modified adenoviral fiber with ablated to cellular receptors |
| US20060062764A1 (en) | 2004-08-25 | 2006-03-23 | Seshidar-Reddy Police | Fiber-modified adenoviral vectors for enhanced transduction of tumor cells |
| US7741099B2 (en) | 2004-10-13 | 2010-06-22 | Beth Israel Deaconess Medical Center Inc. | Adenoviral vectors and uses thereof |
| WO2006119449A2 (en) * | 2005-05-04 | 2006-11-09 | Vectorlogics, Inc. | Modified adenovirus containing a stabilized antibody |
| JP2011501958A (en) | 2007-10-29 | 2011-01-20 | バージニア テック インテレクチュアル プロパティーズ インコーポレーテッド | Porcine DC-SIGN, ICAM-3 and LSECtin and their use |
| CA2705014A1 (en) * | 2007-11-08 | 2009-05-14 | Nestec S.A. | Use of oligosaccharides containing n-acetyllactosamine for maturation of immune responses in neonates |
| EP2325298B1 (en) | 2008-03-04 | 2016-10-05 | The Trustees Of The University Of Pennsylvania | SIMIAN ADENOVIRUSES SAdV-36, -42.1, -42.2, AND -44 AND USES THEREOF |
| EP2342321B1 (en) | 2008-09-17 | 2018-04-11 | Isogenis, Inc. | Construction of fully-deleted adenovirus-based gene delivery vectors and uses thereof |
| US20100083391A1 (en) * | 2008-09-26 | 2010-04-01 | Hamilton Thomas C | Cancer Compositions, Animal Models, and Methods of Use Thereof |
| CA2755192C (en) | 2009-03-20 | 2018-09-11 | Sangamo Biosciences, Inc. | Modification of cxcr4 using engineered zinc finger proteins |
| US8956828B2 (en) | 2009-11-10 | 2015-02-17 | Sangamo Biosciences, Inc. | Targeted disruption of T cell receptor genes using engineered zinc finger protein nucleases |
| US9267153B2 (en) * | 2011-12-15 | 2016-02-23 | Washington University | Porcine knob xenotype chimeric adenoviral vector for dendritic cell infection |
-
2012
- 2012-12-14 US US14/365,957 patent/US9267153B2/en active Active
- 2012-12-14 WO PCT/US2012/069884 patent/WO2013090806A2/en active Application Filing
- 2012-12-14 EP EP12858372.1A patent/EP2798069B1/en not_active Not-in-force
Non-Patent Citations (1)
| Title |
|---|
| None * |
Also Published As
| Publication number | Publication date |
|---|---|
| EP2798069A2 (en) | 2014-11-05 |
| WO2013090806A3 (en) | 2014-10-02 |
| WO2013090806A2 (en) | 2013-06-20 |
| EP2798069A4 (en) | 2015-11-18 |
| US9267153B2 (en) | 2016-02-23 |
| US20150017195A1 (en) | 2015-01-15 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| EP2798069B1 (en) | Porcine knob xenotype chimeric adenoviral vector for dendritic cell infection | |
| Sirena et al. | The human membrane cofactor CD46 is a receptor for species B adenovirus serotype 3 | |
| KR102044051B1 (en) | Batches of recombinant adenovirus with altered terminal ends | |
| EP3256140B1 (en) | Compositions for ebola virus vaccination | |
| US20100034774A1 (en) | Serotype of adenovirus and uses thereof | |
| MXPA06013570A (en) | Chimeric adenoviruses for use in cancer treatment. | |
| JP2024038194A (en) | Delivery of sialidase to cancer cells, immune cells and tumor microenvironment | |
| US9476061B2 (en) | Adenoviral vectors for transduction of vascular tissue | |
| Corredor et al. | The non-essential left end region of the fowl adenovirus 9 genome is suitable for foreign gene insertion/replacement | |
| JP2007527215A (en) | Adenovirus E1A / E1B complementation cell line | |
| WO2012163119A1 (en) | Construction and application of mutant type b human adenovirus ad11 having enhanced oncolytic power | |
| Fouletier-Dilling et al. | Novel compound enables high-level adenovirus transduction in the absence of an adenovirus-specific receptor | |
| Hemsath et al. | Ex vivo and in vivo CD46 receptor utilization by species D human adenovirus serotype 26 (HAdV26) | |
| Di Paolo et al. | Fiber shaft-chimeric adenovirus vectors lacking the KKTK motif efficiently infect liver cells in vivo | |
| US20100233125A1 (en) | Chimeric adenovirus, method for producing the same and pharmaceutical using the same | |
| CN115397448A (en) | Delivery of sialidases to cancer cells, immune cells and tumor microenvironment | |
| Perreau et al. | The conundrum between immunological memory to adenovirus and their use as vectors in clinical gene therapy | |
| van de Ven et al. | Selective transduction of mature DC in human skin and lymph nodes by CD80/CD86-targeted fiber-modified adenovirus-5/3 | |
| Fernandes et al. | Upstream bioprocess for adenovirus vectors | |
| JP2003504316A (en) | Adenovirus vectors for treating diseases | |
| JP2011505837A (en) | Vaccine against adenovirus serotype 14 | |
| WO2023047348A1 (en) | Stabilized corona virus spike protein fusion proteins | |
| Maguire | Analysis of targeting approaches intended to enhance the immunogenicity of adenovirally-vectored vaccine antigens | |
| Mercier et al. | 111. A Chimeric Adenovirus Vector Encoding Reovirus Attachment Protein Sigma 1 Targets Cells Expressing Junctional Adhesion Molecule 1 and Sialic Acid | |
| Strack | Transcriptional Targeting of Dendritic Cells as a New Therapeutic Vaccine against HIV-1 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PUAI | Public reference made under article 153(3) epc to a published international application that has entered the european phase |
Free format text: ORIGINAL CODE: 0009012 |
|
| 17P | Request for examination filed |
Effective date: 20140714 |
|
| AK | Designated contracting states |
Kind code of ref document: A2 Designated state(s): AL AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO RS SE SI SK SM TR |
|
| AX | Request for extension of the european patent |
Extension state: BA ME |
|
| R17P | Request for examination filed (corrected) |
Effective date: 20150330 |
|
| RBV | Designated contracting states (corrected) |
Designated state(s): AL AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO RS SE SI SK SM TR |
|
| DAX | Request for extension of the european patent (deleted) | ||
| RIC1 | Information provided on ipc code assigned before grant |
Ipc: C12N 5/0784 20100101ALI20151001BHEP Ipc: C12N 15/861 20060101AFI20151001BHEP Ipc: C12N 15/63 20060101ALI20151001BHEP Ipc: A61K 39/00 20060101ALI20151001BHEP |
|
| A4 | Supplementary search report drawn up and despatched |
Effective date: 20151016 |
|
| 17Q | First examination report despatched |
Effective date: 20160531 |
|
| GRAP | Despatch of communication of intention to grant a patent |
Free format text: ORIGINAL CODE: EPIDOSNIGR1 |
|
| INTG | Intention to grant announced |
Effective date: 20161028 |
|
| GRAS | Grant fee paid |
Free format text: ORIGINAL CODE: EPIDOSNIGR3 |
|
| STAA | Information on the status of an ep patent application or granted ep patent |
Free format text: STATUS: GRANT OF PATENT IS INTENDED |
|
| GRAA | (expected) grant |
Free format text: ORIGINAL CODE: 0009210 |
|
| STAA | Information on the status of an ep patent application or granted ep patent |
Free format text: STATUS: THE PATENT HAS BEEN GRANTED |
|
| AK | Designated contracting states |
Kind code of ref document: B1 Designated state(s): AL AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO RS SE SI SK SM TR |
|
| REG | Reference to a national code |
Ref country code: GB Ref legal event code: FG4D |
|
| REG | Reference to a national code |
Ref country code: CH Ref legal event code: EP |
|
| REG | Reference to a national code |
Ref country code: AT Ref legal event code: REF Ref document number: 879806 Country of ref document: AT Kind code of ref document: T Effective date: 20170415 |
|
| REG | Reference to a national code |
Ref country code: IE Ref legal event code: FG4D |
|
| RAP2 | Party data changed (patent owner data changed or rights of a patent transferred) |
Owner name: WASHINGTON UNIVERSITY |
|
| REG | Reference to a national code |
Ref country code: DE Ref legal event code: R096 Ref document number: 602012030603 Country of ref document: DE |
|
| REG | Reference to a national code |
Ref country code: DE Ref legal event code: R082 Ref document number: 602012030603 Country of ref document: DE Representative=s name: STOLMAR & PARTNER PATENTANWAELTE PARTG MBB, DE |
|
| REG | Reference to a national code |
Ref country code: CH Ref legal event code: NV Representative=s name: RENTSCH PARTNER AG, CH |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: FI Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: GR Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170630 Ref country code: HR Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: NO Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170629 Ref country code: LT Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| REG | Reference to a national code |
Ref country code: NL Ref legal event code: MP Effective date: 20170329 |
|
| REG | Reference to a national code |
Ref country code: AT Ref legal event code: MK05 Ref document number: 879806 Country of ref document: AT Kind code of ref document: T Effective date: 20170329 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: LV Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: BG Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170629 Ref country code: RS Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: SE Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: NL Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| REG | Reference to a national code |
Ref country code: CH Ref legal event code: PCAR Free format text: NEW ADDRESS: BELLERIVESTRASSE 203 POSTFACH, 8034 ZUERICH (CH) |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: IT Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: ES Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: SK Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: CZ Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: RO Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: AT Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: EE Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: PT Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170731 Ref country code: IS Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170729 Ref country code: SM Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 Ref country code: PL Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| REG | Reference to a national code |
Ref country code: FR Ref legal event code: PLFP Year of fee payment: 6 |
|
| REG | Reference to a national code |
Ref country code: DE Ref legal event code: R097 Ref document number: 602012030603 Country of ref document: DE |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: DK Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: MC Payment date: 20171214 Year of fee payment: 6 Ref country code: LU Payment date: 20171219 Year of fee payment: 6 |
|
| PLBE | No opposition filed within time limit |
Free format text: ORIGINAL CODE: 0009261 |
|
| STAA | Information on the status of an ep patent application or granted ep patent |
Free format text: STATUS: NO OPPOSITION FILED WITHIN TIME LIMIT |
|
| 26N | No opposition filed |
Effective date: 20180103 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: SI Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: MT Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20171214 |
|
| REG | Reference to a national code |
Ref country code: BE Ref legal event code: MM Effective date: 20171231 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: BE Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20171231 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: IE Payment date: 20181220 Year of fee payment: 7 Ref country code: DE Payment date: 20181210 Year of fee payment: 7 |
|
| PGFP | Annual fee paid to national office [announced via postgrant information from national office to epo] |
Ref country code: GB Payment date: 20181218 Year of fee payment: 7 Ref country code: FR Payment date: 20181221 Year of fee payment: 7 Ref country code: CH Payment date: 20181218 Year of fee payment: 7 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: HU Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT; INVALID AB INITIO Effective date: 20121214 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: MC Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20190102 Ref country code: LU Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20181214 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: CY Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: MK Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: TR Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| REG | Reference to a national code |
Ref country code: DE Ref legal event code: R119 Ref document number: 602012030603 Country of ref document: DE |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: AL Free format text: LAPSE BECAUSE OF FAILURE TO SUBMIT A TRANSLATION OF THE DESCRIPTION OR TO PAY THE FEE WITHIN THE PRESCRIBED TIME-LIMIT Effective date: 20170329 |
|
| REG | Reference to a national code |
Ref country code: CH Ref legal event code: PL |
|
| GBPC | Gb: european patent ceased through non-payment of renewal fee |
Effective date: 20191214 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: FR Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20191231 Ref country code: IE Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20191214 Ref country code: DE Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20200701 Ref country code: GB Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20191214 |
|
| PG25 | Lapsed in a contracting state [announced via postgrant information from national office to epo] |
Ref country code: LI Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20191231 Ref country code: CH Free format text: LAPSE BECAUSE OF NON-PAYMENT OF DUE FEES Effective date: 20191231 |