US11612604B2 - Methods of treating cancer with PLK4 inhibitors - Google Patents
Methods of treating cancer with PLK4 inhibitors Download PDFInfo
- Publication number
- US11612604B2 US11612604B2 US16/769,458 US201816769458A US11612604B2 US 11612604 B2 US11612604 B2 US 11612604B2 US 201816769458 A US201816769458 A US 201816769458A US 11612604 B2 US11612604 B2 US 11612604B2
- Authority
- US
- United States
- Prior art keywords
- substituted
- unsubstituted
- nhc
- positive
- cells
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Active, expires
Links
Images
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/33—Heterocyclic compounds
- A61K31/395—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins
- A61K31/535—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins having six-membered rings with at least one nitrogen and one oxygen as the ring hetero atoms, e.g. 1,2-oxazines
- A61K31/5375—1,4-Oxazines, e.g. morpholine
- A61K31/5377—1,4-Oxazines, e.g. morpholine not condensed and containing further heterocyclic rings, e.g. timolol
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/33—Heterocyclic compounds
- A61K31/395—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins
- A61K31/495—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins having six-membered rings with two or more nitrogen atoms as the only ring heteroatoms, e.g. piperazine or tetrazines
- A61K31/505—Pyrimidines; Hydrogenated pyrimidines, e.g. trimethoprim
- A61K31/506—Pyrimidines; Hydrogenated pyrimidines, e.g. trimethoprim not condensed and containing further heterocyclic rings
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K31/00—Medicinal preparations containing organic active ingredients
- A61K31/33—Heterocyclic compounds
- A61K31/395—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins
- A61K31/55—Heterocyclic compounds having nitrogen as a ring hetero atom, e.g. guanethidine or rifamycins having seven-membered rings, e.g. azelastine, pentylenetetrazole
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
- A61P35/02—Antineoplastic agents specific for leukemia
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
- A61P35/04—Antineoplastic agents specific for metastasis
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2800/00—Detection or diagnosis of diseases
- G01N2800/52—Predicting or monitoring the response to treatment, e.g. for selection of therapy based on assay results in personalised medicine; Prognosis
Definitions
- Centrioles play a role in cytokinesis. Centrioles are small cellular organelles whose distinguishing feature is an outer wall made up of a 9-fold symmetric array of stabilized microtubules. Centrioles have two critical functions in cells: (i) they template the formation of microtubule-based projections called cilia, and (2) they direct the assembly of pericentriolar material that nucleates and anchors microtubules to form centrosomes (Conduit et al., Nat Rev Mol Cell Biol. 16:611-624 (2015); Wong et al., Science. 348:1155-1160 (2014)).
- Centrosomes catalyze microtubule assembly to accelerate formation of the mitotic spindle that segregates the chromosomes during cell division and to ensure its bipolarity. Centriole duplication is tightly controlled to ensure that mitotic cells have precisely two centrosomes. Centrioles duplicate in S-phase, when a single daughter centriole forms adjacent to each mother centriole (Banterle et al., Annu. Rev. Cell. Dev. Biol., 33:23-49 (2017); Fu et al., Cold Spring Harb Perspect Biol. 7:a015800 (2015); Zitouni et al., Nat Rev Mol Cell Biol. 15:433-452 (2014)). Centriole duplication is controlled by the mitotic kinase PLK4 (Banterle et al.; Wong et al.; Zitouni et al.).
- Polo-like kinase is one regulator of centriole biogenesis. PLK4 overexpression may trigger centriole over duplication which can lead to cancer. PLK4 shares active site homology with other kinases, including Aurora kinases. To analyze the effect of centrosome removal in normal and cancer cells, centrinone, a potent specific PLK4 inhibitor was developed (Wong et al.) Centrinone treatment blocks centriole duplication, leading to progressive depelion of centrosomes as cells divide. Many cell lines with cancer-associated mutations (which frequently target the p53 circuit) continue to proliferate, aIbeit at reduced rates, following centrinone-mediated centrosome removal.
- Centrosome removal increases the amount of time that it takes to assemble as spindle and complete mitosis and is thought to stabilize p53 because it triggers the mitotic duration sensor (Fong et al.; Lambrus et al.; Meitinger et al.).
- the response of cells to centrosome removal via small molecule-mediated PLK4 inhibition was expected to depend on: (1) how well the cells can assemble a spindle that can accomplish chromosome segregation in the absence of centrosomes and, (2) if the cells are p53 positive, whether they can assemble a spindle fast enough to avoid activating the mitotic duration sensor.
- Genome-wide CRISPR/Cas9-based screens identified three genes whose inhibition allows RPE1 cells, which are p53 positive, to proliferate in the absence of centrosomes.
- Two, USP28 and 53BP1 were proteins whose loss inactivates the mitotic duration sensor (Fong et al.; Lambrus et al.; Meitinger et al.).
- TRIM37 The third protein whose knockdown facilitates proliferation in the absence of centrosomes is TRIM37 (Fong et al.; Meitinger et al.), an E3 ubiquitin ligase of the TRIpartite Motif (TRIM) protein family (Kallijarvi et al., Exp Cell Res. 308:146-155 (2005)).
- TAM TRIpartite Motif
- the TRIM37 knockout appeared to exert its effect by promoting the assembly of ectopic foci containing centrosomal components that are able to nucleate microtubules and promote spindle formation (Meitinger et al.), thereby reducing mitotic duration in cells that lack centrosomes.
- TRIM37 an elevated level of Chromogranin A
- Synaptophysin an elevated level of Synaptophysin
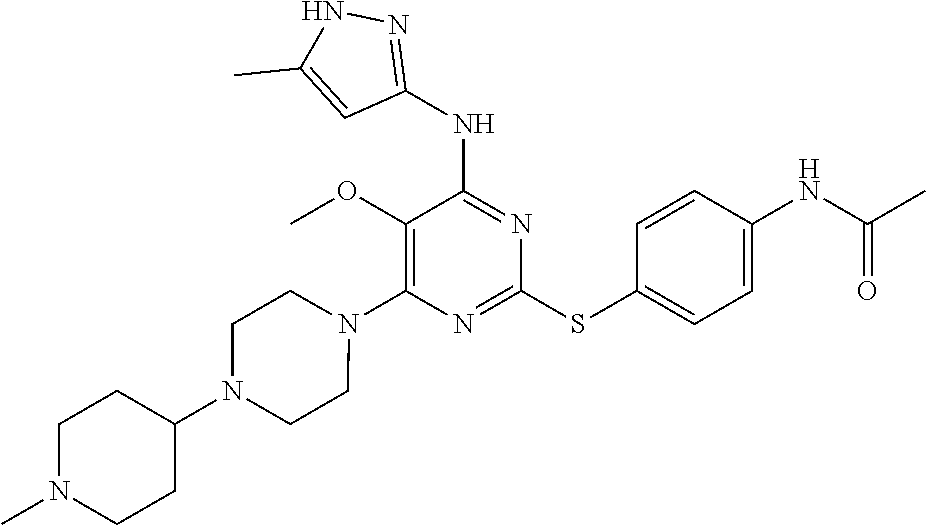
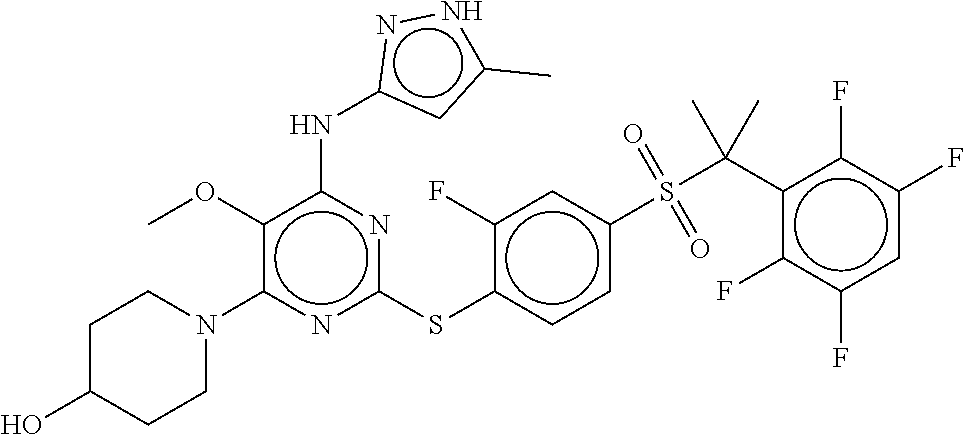
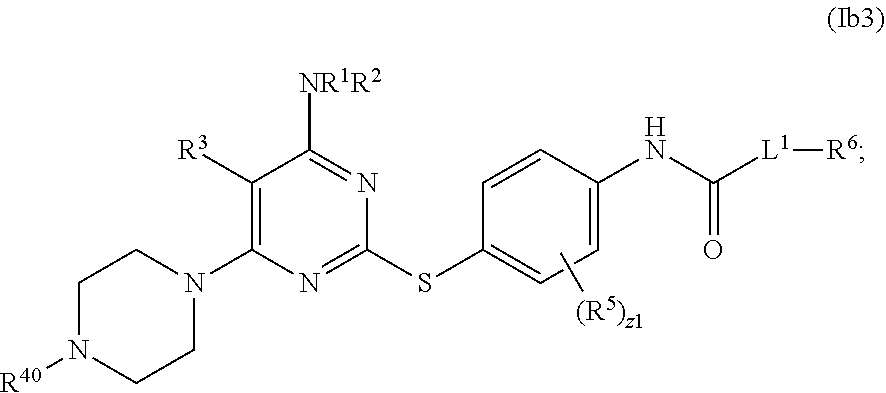
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the cancer is a cancer tumor.
- the cancer is a pediatric cancer.
- the pediatric cancer is a rhabdoid tumor, a neuroblastoma, an acute lymphoblastic leukemia tumor, or a brain and central nervous system tumor.
- the cancer is a neural crest-derived cancer.
- the neural crest-derived tumor is a small cell lung cancer, a melanoma, or a breast cancer.
- the cancer is neuroblastoma.
- the cancer is acute myeloid leukemia, prostate cancer, glioma, mesothelioma, osteosarcoma, breast cancer, Ewing's sarcoma, soft tissue cancer, or T cell lymphoma.
- the cancer is a rhabdoid tumor, basal cell carcinoma, small cell lung cancer, endometrial cancer, non-small cell lung cancer, ovarian cancer, thyroid cancer, kidney cancer, Hodgkin's lymphoma, stomach cancer, liver cancer, Burkitt lymphoma, giant cell tumor of bone, medulloblastoma, a urinary tract cancer, meningioma, bile duct cancer, melanoma, esophageal cancer, upper aerodigestive cancer, colorectal cancer, chondrosarcoma, multiple myeloma, B cell lymphoma, leukemia, diffuse large B cell lymphoma, or chronic myeloid leukemia.
- the cancer is a p53 positive cancer, including a wild type p53 positive cancer or a mutant p53 positive cancer.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- p53 positive cancer is a wild type p53 positive cancer.
- the p53 positive cancer is a mutant p53 positive cancer.
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the p53 positive cancer is a pediatric cancer.
- the pediatric cancer is a rhabdoid tumor, a neuroblastoma, an acute lymphoblastic leukemia tumor, or a brain and central nervous system tumor.
- the p53 positive cancer is a neural crest-derived cancer.
- the neural crest-derived tumor is a small cell lung cancer, a melanoma, or a breast cancer.
- the p53 positive cancer is neuroblastoma.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- the disclosure provides methods of treating a wild type p53 positive pediatric cancer in a subject in need thereof by administering to the subject an effective amount of a PLK4 inhibitor to treat the wild-type p53 positive pediatric cancer; wherein the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin, when compared to a control.
- the disclosure provides methods of treating a mutant p53 positive pediatric cancer in a subject in need thereof by administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive pediatric cancer; wherein the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin, when compared to a control.
- the pediatric cancer is a rhabdoid tumor, a neuroblastoma, an acute lymphoblastic leukemia tumor, or a brain and central nervous system tumor.
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- the disclosure provides methods of treating a wild type p53 positive, neural crest-derived cancer in a subject in need thereof by administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type p53 positive, neural crest-derived cancer; wherein the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin, when compared to a control.
- the disclosure provides methods of treating a mutant p53 positive, neural crest-derived cancer in a subject in need thereof by administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive, neural crest-derived cancer; wherein the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin, when compared to a control.
- the neural crest-derived tumor is a small cell lung cancer, a melanoma, or a breast cancer.
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- a wild-type p53 positive cancer in a subject in need thereof, the method comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild-type p53 positive cancer; wherein the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin, when compared to a control.
- the disclosure provides methods of treating a mutant p53 positive cancer in a subject in need thereof by administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive cancer; wherein the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin, when compared to a control.
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- a wild type p53 positive neuroblastoma in a subject in need thereof, the method comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type p53 positive neuroblastoma; wherein the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin, when compared to a control.
- the disclosure provides methods of treating a mutant p53 positive neuroblastoma in a subject in need thereof, the method comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive neuroblastoma; wherein the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin, when compared to a control.
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- TRIM37 a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level
- a biological sample obtained from the subject a biological sample obtained from the subject
- administering an effective amount of a PLK4 inhibitor to the subject to treat the cancer.
- the subject has an elevated level of TRIM37, Chromogranin A level, and/or a Synaptophysin.
- the subject has an elevated level of TRIM37, Chromogranin A level, and/or a Synaptophysin compared to a control.
- the biological sample is a tumor sample, such as a resected tumor sample, a tumor biopsy sample, a primary tumor sample, a resected primary tumor sample, a primary tumor biopsy sample, a metastatic tumor sample, a resected metastatic tumor sample, or a metastatic tumor biopsy sample.
- the biological sample is a blood sample, such as a peripheral blood sample.
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the cancer is a cancer tumor.
- the cancer is a pediatric cancer.
- the pediatric cancer is a rhabdoid tumor, a neuroblastoma, an acute lymphoblastic leukemia tumor, or a brain and central nervous system tumor.
- the cancer is a neural crest-derived cancer.
- the neural crest-derived tumor is a small cell lung cancer, a melanoma, or a breast cancer.
- the cancer is neuroblastoma.
- the cancer is acute myeloid leukemia, prostate cancer, glioma, mesothelioma, osteosarcoma, breast cancer, Ewing's sarcoma, soft tissue cancer, or T cell lymphoma.
- the cancer is a rhabdoid tumor, basal cell carcinoma, small cell lung cancer, endometrial cancer, non-small cell lung cancer, ovarian cancer, thyroid cancer, kidney cancer, Hodgkin's lymphoma, stomach cancer, liver cancer, Burkitt lymphoma, giant cell tumor of bone, medulloblastoma, a urinary tract cancer, meningioma, bile duct cancer, melanoma, esophageal cancer, upper aerodigestive cancer, colorectal cancer, chondrosarcoma, multiple myeloma, B cell lymphoma, leukemia, diffuse large B cell lymphoma, or chronic myeloid leukemia.
- the cancer is a p53 positive cancer, including a wild type p53 positive cancer or a mutant p53 positive cancer.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- a subject responsive to a PLK4 inhibitor by: (i) obtaining a biological sample from the subject; and (ii) measuring a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level, in the biological sample; wherein if the TRIM37 level, the Chromogranin A level, and/or the Synaptophysin level, is elevated, then the subject is identified as responsive to the PLK4 inhibitor.
- the TRIM37 level, the Chromogranin A level, and/or the Synaptophysin level is elevated when compared to a control.
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the biological sample is a tumor sample, such as a resected tumor sample, a tumor biopsy sample, a primary tumor sample, a resected primary tumor sample, a primary tumor biopsy sample, a metastatic tumor sample, a resected metastatic tumor sample, or a metastatic tumor biopsy sample.
- the biological sample is a blood sample, such as a peripheral blood sample.
- a subject for treatment with a PLK4 inhibitor by: (i) obtaining a biological sample from the subject; and (ii) measuring a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level, in the biological sample; wherein if the TRIM37 level, the Chromogranin A level, and/or the Synaptophysin level, is elevated, then the subject is selected for treatment with the PLK4 inhibitor.
- the TRIM37 level, the Chromogranin A level, and/or the Synaptophysin level is elevated when compared to a control.
- the PLK4 inhibitor is a compound of Formula (I), (Ia), (Ia1), (Ia2), (Ia3), (Ia4), (Ia5), (Ia6), (Ia7), (Ia8), (Ia9a), (Ia9c), (Ia9d), (Ia9f), (Ib), (Ib1), (Ib2), (Ib3), (Ib4), (Ib5), (Ib6), (Ib7), (IC), (I), (III), or (III).
- the PLK4 inhibitor is a compound set forth in Table 1.
- the PLK4 inhibitor is a compound set forth in Table 2.
- the biological sample is a tumor sample, such as a resected tumor sample, a tumor biopsy sample, a primary tumor sample, a resected primary tumor sample, a primary tumor biopsy sample, a metastatic tumor sample, a resected metastatic tumor sample, or a metastatic tumor biopsy sample.
- the biological sample is a blood sample, such as a peripheral blood sample.
- FIGS. 1 A- 1 C show sensitivity of cancer cell lines to PLK4 inhibition.
- FIG. 1 A provides graphs showing the results of passaging assays for the indicated cancer cell lines that monitor cell proliferation after the addition of DMSO (vehicle; black; upper line with sharp peaks and valleys or circle in upper right hand corner) or 150 nM centrinone (lower line) at day 0.
- the second graph in each pair shows a rescaled version of centrinone-treated curve.
- Cell lines in which the number of cells decreased to zero within 12 days following centrinone addition were considered sensitive to PLK4 inhibition (marked with the filled circles in the upper right hand corner, whereas cell lines that continued to proliferate were considered ‘Not Sensitive’ (marked with the empty circles in the upper right hand corner).
- FIG. 1 B provides graphs showing the results of passaging assays for CHP134 cells and CHP134 cells in which both endogenous PLK4 alleles have been engineered to express a G95L mutant PLK4 protein. Growth is shown after the addition of DMSO (vehicle; black; upper line) or 150 nM centrinone (lower line) at day 0.
- FIG. 1 C provides a graph plotting the percentage of apoptotic cells for each cancer cell line after treatment for four cell cycle durations with 150 nm centrinone, 500 nm centrinone B or DMSO (wherein control is left; centrinone is center; centrinone B is right).
- FIGS. 2 A-D show mitotic duration and anaphase failure rates following centrosome removal correlate with TRIM37 protein levels.
- FIG. 2 A shows the CHP134 neuroblastoma cell line has 4 copies of the TRIM37 gene. After CRISPR targeting, CHP134 clones with varying TRIM37 copy numbers were isolated and the levels of TRIM37 protein were measured by quantitative western blotting. Alpha-tubulin was used as a blotting control. Clones with TRIM37 levels between 12 and 64% of the levels in the WT cell line were isolated.
- FIG. 2 B and 2 C show analysis of mitosis by live cell filming in the CHP134 clones after 3 cell cycles in 150 nM centrinone and the duration of mitosis and rate of anaphase failure were measured for each clone.
- Graphs plot mean mitotic duration ( FIG. 2 B ) and the percentage of cells exhibiting anaphase failure ( FIG. 2 C ) versus measured TRIM37 protein level.
- FIG. 2 D presents data in which cellular proliferation was assessed for each of the CHP134 clones by performing an ATPlite assay after 5 days in 125 nM centrinone. Results are expressed as the percent of the value for the equivalent DMSO-treated control and are plotted versus measured TRIM37 protein level. Note that WT CHP134 cells exhibited a loss in proliferation comparable to that of the two highest mutant clones. Thus, for centrinone-treated CHP134 cells, mitotic success and cell viability both decreased as TRIM37 levels increased.
- FIGS. 3 A-B show sensitivity to centrosome removal correlates with TRIM37 copy number in breast cancer.
- FIG. 3 A is a graph plotting TRIM37 copy number versus expression level for breast cancer cell lines (data from the CCLE database; (Barretina et al., 2012)).
- FIG. 3 B are graphs showing the results of passaging assays for the indicated cancer cell lines that monitor cell proliferation after the addition of DMSO (vehicle; black) or 150 nM centrinone (grey) at day 0. The right graph in each pair shows the centrinone-treated curve without the control.
- FIGS. 4 A- 4 C show the quantification of p53 activity and TRIM37 protein levels.
- FIG. 4 A provides a graph shows relative p53 activity across the indicated panel of cell lines. The circles are color-coded to indicate which cell lines are p53 positive (p53+, CHP212, BT-16, HepG2, BT-12, RPE1, CHP134, MR32, SH—SY5Y, SK-N—SH) and negative (p53 ⁇ , A673, HeLa, SK-ES-1, SK-N-F1).
- FIG. 4 B provides a graph shows the relative amount of TRIM37 protein in each of the indicated cancer cell lines as measured by quantitative western blotting.
- FIG. 4 C is a graph shows the correlation between measured levels of TRIM37 mRNA (from the CCLE database; (Barretina et al., 2012)) and measured protein amount.
- FIGS. 5 A- 5 C shows PLK4 inhibition leads to mitotic failure and cell death due to activation of the mitotic duration sensor in cell lines expressing high levels of TRIM37.
- FIG. 5 A and FIG. 5 B are graphs plotting the distribution of mitotic phenotypes ( FIG. 5 A ) and mitotic duration ( FIG. 5 B ) for the indicated cell lines after growth for three cell cycle durations in DMSO or centrinone. Graph in FIG. 5 B is a 5-95% box-and-whiskers plot.
- FIG. 5 C provides an analysis of the mitotic duration sensor in the indicated cell lines. Vertical bars represent individual daughter cells. Bar height shows the time the mother cell spent in mitosis, and bar color indicates whether the daughter cell divided (grey), arrested, or died (black). Black dashed lines mark the mitotic duration threshold for each cell line. The majority of daughter cells whose mothers spent longer than the threshold in mitosis exhibit p53-dependent cell death or arrest.
- FIGS. 6 A- 6 B show that TRIM37 deletion suppresses the mitotic defects resulting from PLK4 inhibition in CHP134 cells and restores their ability to proliferate.
- FIG. 6 A provides graphs plotting the distribution of mitotic phenotypes (left panel) and the mitotic duration (right panel) after growth for three cell cycle durations in DMSO or centrinone for control CHP134 cells and CHP134 cells in which TRIM37 was deleted (TRIM37A), p53 was inhibited (TP53-sh), or both.
- Right graph is a 5-95% box-and-whiskers plot.
- FIG. 6 B provides graphs plotting the results of passaging assays that monitor the proliferation of wild-type and mutant CHP134 cell lines after addition of DMSO (vehicle) or centrinone at day 0.
- FIGS. 7 A- 7 D show that ligase activity of TRIM37 prevents the assembly of ectopic foci containing centrosomal proteins.
- FIG. 7 A provides immunofluorescence images of wild-type RPE1, RPE1 TRIM37 ⁇ and RPE1 TRIM37 ⁇ cells stably expressing wild-type TRIM37 or a ligase-inactive mutant TRIM37-C18R from the UbC-promoter. Cells were stained for DNA (blue) and with antibodies to the centrosomal protein Cep192 (green). Bar, 10 ⁇ m.
- FIG. 7 B provides graphs plotting the percentage of cells that have ectopic Cep192 foci around the centrosome as shown in FIG. 7 A .
- FIG. 7 A provides immunofluorescence images of wild-type RPE1, RPE1 TRIM37 ⁇ and RPE1 TRIM37 ⁇ cells stably expressing wild-type TRIM37 or a ligase-inactive mutant TRIM37
- FIG. 7 C provides an immunoblot showing the expression level of TRIM37 from the wild-type and C18R transgenes compared to the amount of endogenous TRIM37 in the control RPE1 and RPE1 TRIM37 ⁇ cell lines.
- FIG. 7 D provides a graph plotting mitotic duration for RPE1, RPE1 TRIM37 ⁇ and RPE1 TRIM37 ⁇ cells stably expressing wild-type and ligase-inactive mutant (C18R) TRIM37 from the UbC-promoter after three cell cycle durations in DMSO or centrinone.
- Graph is a 5-95% box-and-whiskers plot.
- FIG. 8 shows a plot of CHGA versus TRIM37 mRNA—Seq values for a number of cancer cell lines. Higher TRIM37 and CHGA values indicate sensitivity to a PLK4 inhibitor. PLK4i sensitivity was measured similarly to experiments in FIG. 3 B above.
- FIG. 9 shows a plot of CHGA versus TRIM37 mRNA—Seq values for a number of cell lines.
- the majority of PLK4i-sensitive lines express CHGA and high TRIM37.
- Small cell lung carcinoma cell lines are underlined and neuroblastoma (NB) cell lines are bolded.
- PLK4i sensitivity was measured similarly to experiments in FIG. 3 B above.
- PLK4 is used according to its common, ordinary meaning and refers to proteins of the same or similar names and functional fragments and homologs thereof.
- the term includes recombinant or naturally occurring forms of PLK4 (e.g. Polo-like Kinase 4; GI No: 160113150), or variants thereof that maintain PLK4 activity (e.g. within at least 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, or 100% activity compared to PLK4).
- inhibitor in reference to a protein-inhibitor interaction, means negatively affecting (e.g. decreasing) the activity or function of the protein relative to the activity or function of the protein in the absence of the inhibitor. Inhibition may refer to negatively affecting (e.g. decreasing) the concentration or level of the protein relative to the concentration or level of the protein in the absence of the inhibitor. Inhibition may refer reduction of a disease or symptoms of disease. Inhibition may refer to a reduction in the activity of a particular protein or nucleic acid target. Thus, inhibition includes, at least in part, partially or totally blocking stimulation, decreasing, preventing, or delaying activation, or inactivating, desensitizing, or down-regulating signal transduction or enzymatic activity or the amount of a protein.
- PLK4 inhibitor refers to a compound that selectively inhibits PLK4.
- PLK4 inhibitors cause depletion of centrioles, disruption in mitosis, induction of apoptosis, prevention of cell division, and inhibition of proliferation of PLK4-overexpressing tumor cells.
- Exemplary PLK4 inhibitors include compounds of Formula (I), Formula (Ia), Formula (Ia1), Formula (Ia2), Formula (Ia3), Formula (Ia4), Formula (Ia5), Formula (Ia6), Formula (Ia1), Formula (Ia.), Formula (Ia9a), Formula (Ia9b), Formula (Ia9c), Formula (Ia9d), Formula (Ia9f), Formula (Ib), Formula (Ib1), Formula (Ib2), Formula (Ib3), Formula (Ib4), Formula (Ib5), Formula (Ib6), Formula (Ib7), Formula (IC), Formula (II), and Formula (III).
- Other Exemplary PLK4 inhibitors include the compounds in Table 1 and Table 2.
- “Selective” or “selectively” or the like of a compound refers to the compound's ability to discriminate between molecular targets. “Specific”, “specifically”, “specificity”, or the like of a compound refers to the compound's ability to cause a particular action, such as inhibition, to a particular molecular target with minimal or no action to other proteins in the cell.
- TMM37 includes any of the recombinant or naturally-occurring forms of the E3 ubiquitin ligase of the TRIpartite Motif (TRIM) protein family Tripartite Motif Containing 37 (TRIM37) or variants or homologs thereof that maintain TRIM37 activity (e.g. within at least 50%, 80%, 90%, 95%, 96%, 97%, 98%, 99% or 100% activity compared to TRIM37).
- the variants or homologs have at least 90%, 95%, 96%, 97%, 98%, 99% or 100% nucleic acid sequence identity across the whole sequence or a portion of the sequence (e.g.
- the TRIM37 gene is substantially identical to the gene identified by the UniProtKB Reference Number TRI37_HUMAN,) 94972 or a variant or homolog having substantial identity thereto.
- the variants or homologs have at least 90%, 95%, 96%, 97%, 98%, 99% or 100% amino acid sequence identity across the whole sequence or a portion of the sequence (e.g. a 50, 100, 150 or 200 continuous amino acid portion) compared to a naturally occurring TRIM37 protein.
- the TRIM37 protein is substantially identical to the protein identified by the UniProtKB Reference Number 094972-TRI37_HUMAN or a variant or homolog having substantial identity thereto.
- the TRIM37 is a mutant TRIM37.
- the TRIM37 is TRIM37A.
- TAM37 levels as referred to herein is the level of TRIM37 expressed by a tumor.
- the TRIM37 levels can be measured by genes, mRNA, or proteins in a biological sample.
- TRIM37 is an elevated level of TRIM37 genes on a tumor or an elevated level of TRIM37 expressed (e.g., mRNA, proteins) by a tumor in a subject when compared to a control.
- TRIM37 levels can be measured from biological samples, such as a tumor sample (e.g., resected, biopsy) or a blood sample (e.g., peripheral blood), obtained from a subject.
- a tumor can be a primary tumor or a metastasis.
- a tumor as provided herein is a cellular mass including cancer cells and non-cancer cells.
- the non-cancer cells forming part of a tumor may be stromal cells, and immune cells (e.g., T cells, dendritic cells, B cells, macrophages).
- immune cells e.g., T cells, dendritic cells, B cells, macrophages.
- the elevated level of TRIM37 is expressed by a non-cancer cell (e.g., a stromal cell) or a cancer cell (e.g., a malignant T cell).
- TRIM37 levels can be detected at either the protein or the gene expression level.
- TRIM37 protein can for example be quantified by immunoblotting, immunohistochemistry (TIC) or flow cytometry with an antibody that detects TRIM37.
- TRIM37 gene expression can, for example, be quantified by multiple platforms such as reverse transcription polymerase chain reaction (rtPCR), Nanostring, RNA sequencing (RNA—Seq), also called whole transcriptome shotgun sequencing (WTSS), or in situ hybridization.
- TRIM37 gene copy number variation (CNV) (particularly gene amplification) can, for example, be quantified by array comparative genomic hybridization (array CGH), fluorescent in situ hybridization (FISH), genomic sequencing or quantitative polymerase chain reaction (qPCR).
- TRIM37 expression across and within tumor types that shows concordance when measured with either immunoblotting, rtPCR and/or RNA—Seq.
- TRIM37 expression across and within tumor types that shows concordance when measured with either IHC, Nanostring and/or by in situ hybridization.
- One skilled in the art will understand the importance of selecting a threshold of TRIM37 expression that constitutes elevated levels. Controls are also valuable for determining the significance of data. For example, if values for a given parameter are widely variant in controls, variation in test samples will not be considered as significant.
- the expression level of TRIM37 is assessed, the level is compared with a control expression level of TRIM37.
- control expression level is meant the expression level of TRIM37 from a sample or subject lacking cancer, a sample or subject at a selected stage of cancer or cancer state, or in the absence of a particular variable such as a therapeutic agent.
- the control level comprises a known amount of TRIM37. Such a known amount correlates with an average level of subjects lacking cancer, at a selected stage of cancer or cancer state, or in the absence of a particular variable such as a therapeutic agent.
- a control level also includes the expression level of TRIM37 from one or more selected samples or subjects as described herein.
- a control level includes an assessment of the expression level of TRIM37 in a sample from a subject that does not have cancer, is at a selected stage of cancer or cancer state, or has not received treatment for cancer.
- Another exemplary control level includes an assessment of the expression level of TRIM37 in samples taken from multiple subjects that do not have cancer, are at a selected stage of cancer, or have not received treatment for cancer.
- a threshold for elevated TRIM37 may be above the median expression level of a group of control samples. In some embodiments it may be above the first or third quartile of TRIM37 expression in a group of control samples.
- control level includes the expression level of TRIM37 in a sample or subject in the absence of a therapeutic agent
- control sample or subject is optionally the same sample or subject to be tested before or after treatment with a therapeutic agent or is a selected sample or subject in the absence of the therapeutic agent.
- a control level is an average expression level calculated from a number of subjects without a particular disease.
- a control level also includes a known control level or value known in the art.
- a control level is the TRIM37 expression level calculated from a biological sample from a subject with cancer prior to treatment.
- the elevated level of TRIM37 is calculated by determining the percentage of cells a biological sample that are positive for TRIM37.
- the cells may be tumor cells, tumor infiltrating cells, stromal cells, vasculature cells, or a composite thereof. In embodiments, the cells are tumor cells.
- the percentage of cells that are positive for TRIM37 may be greater than 1%, greater than 5%, greater than 10%, greater than 15%, greater than 20%, greater than 25%, greater than 30%, greater than 35%, greater than 40%, greater than 45%, greater than 50%, greater than 55%, greater than 60%, greater than 65%, greater than 70%, greater than 75%, greater than 80%, greater than 90%, or greater than 95%.
- the percentage of cells that are positive for TRIM37 is 1% or more.
- the percentage of cells that are positive for TRIM37 is 2% or more.
- the percentage of cells that are positive for TRIM37 is 3% or more.
- the percentage of cells that are positive for TRIM37 is 4% or more.
- the percentage of cells that are positive for TRIM37 is 5% or more. In aspects, the percentage of cells that are positive for TRIM37 is 6% or more. In aspects, the percentage of cells that are positive for TRIM37 is 7% or more. In aspects, the percentage of cells that are positive for TRIM37 is 8% or more. In aspects, the percentage of cells that are positive for TRIM37 is 9% or more. In aspects, the percentage of cells that are positive for TRIM37 is 10% or more. In aspects, the percentage of cells that are positive for TRIM37 is 11% or more. In aspects, the percentage of cells that are positive for TRIM37 is 12% or more. In aspects, the percentage of cells that are positive for TRIM37 is 13% or more.
- the percentage of cells that are positive for TRIM37 is 14% or more. In aspects, the percentage of cells that are positive for TRIM37 is 15% or more. In aspects, the percentage of cells that are positive for TRIM37 is 16% or more. In aspects, the percentage of cells that are positive for TRIM37 is 17% or more. In aspects, the percentage of cells that are positive for TRIM37 is 18% or more. In aspects, the percentage of cells that are positive for TRIM37 is 19% or more. In aspects, the percentage of cells that are positive for TRIM37 is 20% or more. In aspects, the percentage of cells that are positive for TRIM37 is 21% or more. In aspects, the percentage of cells that are positive for TRIM37 is 22% or more.
- the percentage of cells that are positive for TRIM37 is 23% or more. In aspects, the percentage of cells that are positive for TRIM37 is 24% or more. In aspects, the percentage of cells that are positive for TRIM37 is 25% or more. In aspects, the percentage of cells that are positive for TRIM37 is 26% or more. In aspects, the percentage of cells that are positive for TRIM37 is 27% or more. In aspects, the percentage of cells that are positive for TRIM37 is 28% or more. In aspects, the percentage of cells that are positive for TRIM37 is 29% or more. In aspects, the percentage of cells that are positive for TRIM37 is 30% or more. In aspects, the percentage of cells that are positive for TRIM37 is 31% or more.
- the percentage of cells that are positive for TRIM37 is 32% or more. In aspects, the percentage of cells that are positive for TRIM37 is 33% or more. In aspects, the percentage of cells that are positive for TRIM37 is 34% or more. In aspects, the percentage of cells that are positive for TRIM37 is 35% or more. In aspects, the percentage of cells that are positive for TRIM37 is 36% or more. In aspects, the percentage of cells that are positive for TRIM37 is 37% or more. In aspects, the percentage of cells that are positive for TRIM37 is 38% or more. In aspects, the percentage of cells that are positive for TRIM37 is 39% or more. In aspects, the percentage of cells that are positive for TRIM37 is 40% or more.
- the percentage of cells that are positive for TRIM37 is 41% or more. In aspects, the percentage of cells that are positive for TRIM37 is 42% or more. In aspects, the percentage of cells that are positive for TRIM37 is 43% or more. In aspects, the percentage of cells that are positive for TRIM37 is 44% or more. In aspects, the percentage of cells that are positive for TRIM37 is 45% or more. In aspects, the percentage of cells that are positive for TRIM37 is 46% or more. In aspects, the percentage of cells that are positive for TRIM37 is 47% or more. In aspects, the percentage of cells that are positive for TRIM37 is 48% or more. In aspects, the percentage of cells that are positive for TRIM37 is 49% or more.
- the percentage of cells that are positive for TRIM37 is 50% or more. In aspects, the percentage of cells that are positive for TRIM37 is 51% or more. In aspects, the percentage of cells that are positive for TRIM37 is 52% or more. In aspects, the percentage of cells that are positive for TRIM37 is 53% or more. In aspects, the percentage of cells that are positive for TRIM37 is 54% or more. In aspects, the percentage of cells that are positive for TRIM37 is 55% or more. In aspects, the percentage of cells that are positive for TRIM37 is 56% or more. In aspects, the percentage of cells that are positive for TRIM37 is 57% or more. In aspects, the percentage of cells that are positive for TRIM37 is 58% or more.
- the percentage of cells that are positive for TRIM37 is 59% or more. In aspects, the percentage of cells that are positive for TRIM37 is 60% or more. In aspects, the percentage of cells that are positive for TRIM37 is 61% or more. In aspects, the percentage of cells that are positive for TRIM37 is 62% or more. In aspects, the percentage of cells that are positive for TRIM37 is 63% or more. In aspects, the percentage of cells that are positive for TRIM37 is 64% or more. In aspects, the percentage of cells that are positive for TRIM37 is 65% or more. In aspects, the percentage of cells that are positive for TRIM37 is 66% or more. In aspects, the percentage of cells that are positive for TRIM37 is 67% or more.
- the percentage of cells that are positive for TRIM37 is 68% or more. In aspects, the percentage of cells that are positive for TRIM37 is 69% or more. In aspects, the percentage of cells that are positive for TRIM37 is 70% or more. In aspects, the percentage of cells that are positive for TRIM37 is 71% or more. In aspects, the percentage of cells that are positive for TRIM37 is 72% or more. In aspects, the percentage of cells that are positive for TRIM37 is 73% or more. In aspects, the percentage of cells that are positive for TRIM37 is 74% or more. In aspects, the percentage of cells that are positive for TRIM37 is 75% or more. In aspects, the percentage of cells that are positive for TRIM37 is 76% or more.
- the percentage of cells that are positive for TRIM37 is 77% or more. In aspects, the percentage of cells that are positive for TRIM37 is 78% or more. In aspects, the percentage of cells that are positive for TRIM37 is 79% or more. In aspects, the percentage of cells that are positive for TRIM37 is 80% or more. In aspects, the percentage of cells that are positive for TRIM37 is 81% or more. In aspects, the percentage of cells that are positive for TRIM37 is 82% or more. In aspects, the percentage of cells that are positive for TRIM37 is 83% or more. In aspects, the percentage of cells that are positive for TRIM37 is 84% or more. In aspects, the percentage of cells that are positive for TRIM37 is 85% or more.
- the percentage of cells that are positive for TRIM37 is 86% or more. In aspects, the percentage of cells that are positive for TRIM37 is 87% or more. In aspects, the percentage of cells that are positive for TRIM37 is 88% or more. In aspects, the percentage of cells that are positive for TRIM37 is 89% or more. In aspects, the percentage of cells that are positive for TRIM37 is 90% or more. In aspects, the percentage of cells that are positive for TRIM37 is 91% or more. In aspects, the percentage of cells that are positive for TRIM37 is 92% or more. In aspects, the percentage of cells that are positive for TRIM37 is 93% or more. In aspects, the percentage of cells that are positive for TRIM37 is 94% or more.
- the percentage of cells that are positive for TRIM37 is 95% or more. In aspects, the percentage of cells that are positive for TRIM37 is 96% or more. In aspects, the percentage of cells that are positive for TRIM37 is 97% or more. In aspects, the percentage of cells that are positive for TRIM37 is 98% or more. In aspects, the percentage of cells that are positive for TRIM37 is 99% or more.
- TRIM37 gene expression is used to assay for elevated TRIM37.
- quantitative rtPCR, Nanostring, and in situ hybridization are platforms to quantitate gene expression.
- Nanostring RNA is extracted from tumor samples and a known quantity of RNA is placed on the Nanostring machine for gene expression detection using gene specific probes. The number of counts of TRIM37 within a sample is determined and normalized to a set of “housekeeping” genes.
- levels of the TRIM37 gene and variations thereof can be used interchangeably with the term “copies of the TRIM37 gene.”
- the term “measuring the level of the TRIM37 gene” and variations thereof can be used interchangeably with the term “measuring copies of the TRIM37 gene.”
- Chromogranin A or “CHGA” as referred to herein includes any recombinant or naturally-occurring form of parathyroid secretory protein 1 (including homologs, isoforms, and functional fragments thereof) that maintain Chromogranin A activity (e.g. within at least 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, or 100% activity compared to wild type Chromogranin A).
- the Chromogranin A protein encoded by the CHGA gene has the amino acid sequence set forth in or corresponding to Entrez 1113, UniProt P10645, or RefSeq (protein) XP_011534672.1.
- the CHGA gene has the nucleic acid sequence set forth in RefSeq (mRNA) XM_011536370.1.
- the amino acid sequence or nucleic acid sequence is the sequence known at the time of filing of the present application.
- the Chromogranin A is a human Chromogranin A.
- the variants or homologs have at least 90%, 95%, 96%, 97%, 98%, 99% or 100% nucleic acid sequence identity across the whole sequence or a portion of the sequence (e.g. a 50, 100, 150 or 200 continuous nucleotide portion) compared to a naturally occurring Chromogranin A gene or mRNA.
- the variants or homologs have at least 90%, 95%, 96%, 97%, 98%, 99% or 100% amino acid sequence identity across the whole sequence or a portion of the sequence (e.g. a 50, 100, 150 or 200 continuous amino acid portion) compared to a naturally occurring Chromogranin A protein.
- Chromogranin A levels as referred to herein is the level of Chromogranin A expressed by a tumor.
- the Chromogranin A levels can be measured by genes, mRNA, or proteins in a biological sample.
- “An elevated level of Chromogranin A” as referred to herein is an elevated level of Chromogranin A genes on a tumor or an elevated level of Chromogranin A expressed (e.g., mRNA, proteins) by a tumor in a subject when compared to a control.
- Chromogranin A levels can be measured from biological samples, such as a tumor sample (e.g., resected, biopsy) or a blood sample (e.g., peripheral blood), obtained from a subject.
- a tumor can be a primary tumor or a metastasis.
- a tumor as provided herein is a cellular mass including cancer cells and non-cancer cells.
- the non-cancer cells forming part of a tumor may be stromal cells, and immune cells (e.g., T cells, dendritic cells, B cells, macrophages).
- immune cells e.g., T cells, dendritic cells, B cells, macrophages.
- the elevated level of Chromogranin A may be expressed by a non-cancer cell (e.g., a stromal cell) or a cancer cell (e.g., a malignant T cell).
- Chromogranin A levels can be detected at either the protein or gene expression level.
- Chromogranin A protein can be quantified by immunohistochemistry (IHC) or flow cytometry with an antibody that detects Chromogranin A.
- Chromogranin A gene expression can be quantified by multiple platforms such as real-time polymerase chain reaction (rtPCR), Nanostring, or in situ hybridization. There is a range of Chromogranin A expression across and within tumor types that shows concordance when measured with either IHC or by Nanostring.
- rtPCR real-time polymerase chain reaction
- Nanostring nanostring
- in situ hybridization There is a range of Chromogranin A expression across and within tumor types that shows concordance when measured with either IHC or by Nanostring.
- control expression level is meant the expression level of Chromogranin A from a sample or subject lacking cancer, a sample or subject at a selected stage of cancer or cancer state, or in the absence of a particular variable such as a therapeutic agent.
- control level comprises a known amount of Chromogranin A. Such a known amount correlates with an average level of subjects lacking cancer, at a selected stage of cancer or cancer state, or in the absence of a particular variable such as a therapeutic agent.
- a control level also includes the expression level of Chromogranin A from one or more selected samples or subjects as described herein.
- a control level includes an assessment of the expression level of Chromogranin A in a sample from a subject that does not have cancer, is at a selected stage of cancer or cancer state, or has not received treatment for cancer.
- Another exemplary control level includes an assessment of the expression level of Chromogranin A in samples taken from multiple subjects that do not have cancer, are at a selected stage of cancer, or have not received treatment for cancer.
- a threshold for elevated Chromogranin A may be above the median expression level of a group of control samples.
- control level includes the expression level of Chromogranin A in a sample or subject in the absence of a therapeutic agent
- the control sample or subject is optionally the same sample or subject to be tested before or after treatment with a therapeutic agent or is a selected sample or subject in the absence of the therapeutic agent.
- a control level is an average expression level calculated from a number of subjects without a particular disease.
- a control level also includes a known control level or value known in the art.
- the elevated level of Chromogranin A is calculated by determining the percentage of cells a biological sample that are positive for Chromogranin A.
- the cells may be tumor cells, tumor infiltrating cells, stromal cells, vasculature cells, or a composite thereof.
- the cells are tumor cells.
- the percentage of cells that are positive for Chromogranin A may be greater than 1%, greater than 5%, greater than 10%, greater than 15%, greater than 20%, greater than 25%, greater than 30%, greater than 35%, greater than 40%, greater than 45%, greater than 50%, greater than 55%, greater than 60%, greater than 65%, greater than 70%, greater than 75%, greater than 80%, greater than 90%, or greater than 95%.
- the percentage of cells that are positive for Chromogranin A is 1% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 2% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 3% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 4% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 5% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 6% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 7% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 8% or more.
- the percentage of cells that are positive for Chromogranin A is 9% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 10% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 11% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 12% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 13% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 14% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 15% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 16% or more.
- the percentage of cells that are positive for Chromogranin A is 17% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 18% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 19% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 20% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 21% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 22% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 23% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 24% or more.
- the percentage of cells that are positive for Chromogranin A is 25% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 26% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 27% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 28% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 29% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 30% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 31% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 32% or more.
- the percentage of cells that are positive for Chromogranin A is 33% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 34% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 35% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 36% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 37% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 38% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 39% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 40% or more.
- the percentage of cells that are positive for Chromogranin A is 41% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 42% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 43% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 44% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 45% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 46% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 47% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 48% or more.
- the percentage of cells that are positive for Chromogranin A is 49% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 50% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 51% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 52% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 53% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 54% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 55% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 56% or more.
- the percentage of cells that are positive for Chromogranin A is 57% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 58% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 59% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 60% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 61% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 62% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 63% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 64% or more.
- the percentage of cells that are positive for Chromogranin A is 65% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 66% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 67% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 68% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 69% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 70% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 71% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 72% or more.
- the percentage of cells that are positive for Chromogranin A is 73% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 74% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 75% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 76% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 77% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 78% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 79% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 80% or more.
- the percentage of cells that are positive for Chromogranin A is 81% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 82% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 83% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 84% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 85% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 86% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 87% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 88% or more.
- the percentage of cells that are positive for Chromogranin A is 89% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 90% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 91% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 92% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 93% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 94% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 95% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 96% or more.
- the percentage of cells that are positive for Chromogranin A is 97% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 98% or more. In aspects, the percentage of cells that are positive for Chromogranin A is 99% or more.
- Chromogranin A gene expression is used to assay for elevated Chromogranin A.
- quantitative rtPCR, Nanostring, and in situ hybridization are platforms to quantitate gene expression.
- Nanostring RNA is extracted from tumor samples and a known quantity of RNA is placed on the Nanostring machine for gene expression detection using gene specific probes. The number of counts of Chromogranin A within a sample is determined and normalized to a set of “housekeeping” genes.
- Synaptophysin or “SYP” as referred to herein includes any recombinant or naturally-occurring form of major synaptic vesicle protein p38 (including homologs, isoforms, and functional fragments thereof) that maintain Synaptophysin activity (e.g. within at least 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, or 100% activity compared to wild type Synaptophysin).
- the Synaptophysin protein encoded by the SYP gene has the amino acid sequence set forth in or corresponding to Entrez 6855, UniProt P08247, or RefSeq (protein) NP_003170.1.
- the amino acid sequence or nucleic acid sequence is the sequence known at the time of filing of the present application.
- the Synaptophysin is a human Synaptophysin.
- the variants or homologs have at least 90%, 95%, 96%, 97%, 98%, 99% or 100% nucleic acid sequence identity across the whole sequence or a portion of the sequence (e.g. a 50, 100, 150 or 200 continuous nucleotide portion) compared to a naturally occurring Synaptophysin gene or mRNA.
- the variants or homologs have at least 90%, 95%, 96%, 97%, 98%, 99% or 100% amino acid sequence identity across the whole sequence or a portion of the sequence (e.g. a 50, 100, 150 or 200 continuous amino acid portion) compared to a naturally occurring Synaptophysin protein.
- Synaptophysin levels as referred to herein is the level of Synaptophysin expressed by a tumor.
- the Synaptophysin levels can be measured by genes, mRNA, or proteins in a biological sample.
- “An elevated level of Synaptophysin” as referred to herein is an elevated level of Synaptophysin genes on a tumor or an elevated level of Synaptophysin expressed (e.g., mRNA, proteins) by a tumor in a subject when compared to a control.
- Synaptophysin levels can be measured from biological samples, such as a tumor sample (e.g., resected, biopsy) or a blood sample (e.g., peripheral blood), obtained from a subject.
- a tumor can be a primary tumor or a metastasis.
- a tumor as provided herein is a cellular mass including cancer cells and non-cancer cells.
- the non-cancer cells forming part of a tumor may be stromal cells, and immune cells (e.g., T cells, dendritic cells, B cells, macrophages).
- immune cells e.g., T cells, dendritic cells, B cells, macrophages.
- the elevated level of Synaptophysin may be expressed by a non-cancer cell (e.g., a stromal cell) or a cancer cell (e.g., a malignant T cell).
- Synaptophysin levels can be detected at either the protein or gene expression level.
- Synaptophysin protein can be quantified by immunohistochemistry (IHC) or flow cytometry with an antibody that detects Synaptophysin.
- Synaptophysin gene expression can be quantified by multiple platforms such as real-time polymerase chain reaction (rtPCR), Nanostring, or in situ hybridization. There is a range of Synaptophysin expression across and within tumor types that shows concordance when measured with either IHC or by Nanostring.
- rtPCR real-time polymerase chain reaction
- Nanostring nanostring
- in situ hybridization There is a range of Synaptophysin expression across and within tumor types that shows concordance when measured with either IHC or by Nanostring.
- control expression level is meant the expression level of Synaptophysin from a sample or subject lacking cancer, a sample or subject at a selected stage of cancer or cancer state, or in the absence of a particular variable such as a therapeutic agent.
- control level comprises a known amount of Synaptophysin. Such a known amount correlates with an average level of subjects lacking cancer, at a selected stage of cancer or cancer state, or in the absence of a particular variable such as a therapeutic agent.
- a control level also includes the expression level of Synaptophysin from one or more selected samples or subjects as described herein.
- a control level includes an assessment of the expression level of Synaptophysin in a sample from a subject that does not have cancer, is at a selected stage of cancer or cancer state, or has not received treatment for cancer.
- Another exemplary control level includes an assessment of the expression level of Synaptophysin in samples taken from multiple subjects that do not have cancer, are at a selected stage of cancer, or have not received treatment for cancer.
- a threshold for elevated Synaptophysin may be above the median expression level of a group of control samples.
- control level includes the expression level of Synaptophysin in a sample or subject in the absence of a therapeutic agent
- the control sample or subject is optionally the same sample or subject to be tested before or after treatment with a therapeutic agent or is a selected sample or subject in the absence of the therapeutic agent.
- a control level is an average expression level calculated from a number of subjects without a particular disease.
- a control level also includes a known control level or value known in the art.
- the elevated level of Synaptophysin is calculated by determining the percentage of cells in a biological sample that are positive for Synaptophysin.
- the cells may be tumor cells, tumor infiltrating cells, stromal cells, vasculature cells, or a composite thereof.
- the cells are tumor cells.
- the percentage of cells that are positive for Synaptophysin may be greater than 1%, greater than 5%, greater than 10%, greater than 15%, greater than 20%, greater than 25%, greater than 30%, greater than 35%, greater than 40%, greater than 45%, greater than 50%, greater than 55%, greater than 60%, greater than 65%, greater than 70%, greater than 75%, greater than 80%, greater than 90%, or greater than 95%.
- the percentage of cells that are positive for Synaptophysin is 1% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 2% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 3% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 4% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 5% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 6% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 7% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 8% or more.
- the percentage of cells that are positive for Synaptophysin is 9% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 10% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 11% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 12% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 13% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 14% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 15% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 16% or more.
- the percentage of cells that are positive for Synaptophysin is 17% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 18% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 19% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 20% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 21% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 22% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 23% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 24% or more.
- the percentage of cells that are positive for Synaptophysin is 25% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 26% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 27% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 28% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 29% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 30% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 31% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 32% or more.
- the percentage of cells that are positive for Synaptophysin is 33% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 34% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 35% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 36% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 37% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 38% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 39% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 40% or more.
- the percentage of cells that are positive for Synaptophysin is 41% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 42% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 43% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 44% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 45% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 46% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 47% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 48% or more.
- the percentage of cells that are positive for Synaptophysin is 49% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 50% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 51% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 52% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 53% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 54% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 55% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 56% or more.
- the percentage of cells that are positive for Synaptophysin is 57% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 58% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 59% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 60% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 61% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 62% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 63% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 64% or more.
- the percentage of cells that are positive for Synaptophysin is 65% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 66% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 67% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 68% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 69% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 70% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 71% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 72% or more.
- the percentage of cells that are positive for Synaptophysin is 73% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 74% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 75% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 76% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 77% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 78% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 79% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 80% or more.
- the percentage of cells that are positive for Synaptophysin is 81% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 82% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 83% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 84% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 85% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 86% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 87% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 88% or more.
- the percentage of cells that are positive for Synaptophysin is 89% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 90% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 91% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 92% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 93% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 94% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 95% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 96% or more.
- the percentage of cells that are positive for Synaptophysin is 97% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 98% or more. In aspects, the percentage of cells that are positive for Synaptophysin is 99% or more.
- Synaptophysin gene expression is used to assay for elevated Synaptophysin.
- quantitative rtPCR, Nanostring, and in situ hybridization are platforms to quantitate gene expression.
- Nanostring RNA is extracted from tumor samples and a known quantity of RNA is placed on the Nanostring machine for gene expression detection using gene specific probes. The number of counts of Synaptophysin within a sample is determined and normalized to a set of “housekeeping” genes.
- a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level refers to the following: (i) a TRIM37 level; (ii) a Chromogranin A level; (iii) a Synaptophysin level; (iv) a TRIM37 level and a Chromogranin A level; (v) a TRIM37 level and a Synaptophysin level; (vi) a Chromogranin A level and a Synaptophysin level; or (vii) a TRIM37 level, a Chromogranin A level, and a Synaptophysin level.
- a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level is (i). In embodiments, “a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level” is (ii). In embodiments, “a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level” is (iii). In embodiments, “a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level” is (iv).
- a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level is (v). In embodiments, “a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level” is (vi). In embodiments, “a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level” is (vii).
- an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin refers to the following: (i) an elevated level of TRIM37; (ii) an elevated level of Chromogranin A; (iii) an elevated level of Synaptophysin; (iv) an elevated level of TRIM37 and an elevated level of Chromogranin A; (v) an elevated level of TRIM37 and an elevated level of Synaptophysin; (vi) an elevated level of Chromogranin A and an elevated level of Synaptophysin; or (vii) an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin.
- an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin is (i). In embodiments, “an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin” is (ii). In embodiments, “an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin” is (iii). In embodiments, “an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin” is (iv).
- an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin is (v). In embodiments, “an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin” is (vi). In embodiments, “an elevated level of TRIM37, an elevated level of Chromogranin A, and/or an elevated level of Synaptophysin” is (vii).
- Subject responsive to a PLK4 inhibitor refers to a subject that responds to treatment when administered a PLK4 inhibitor.
- “Responsive” and “responds” indicate that: (i) a cancerous tumor does not grow in size or volume over time; (ii) a cancerous tumor decreases in size or volume over time; (iii) a cancerous tumor does not metastasize; or (iv) a combination of two or more of the foregoing.
- a subject responsive to a PLK4 inhibitor shows a decrease (i.e., reduction) in tumor size or volume after treatment compared to baseline or a control.
- a subject responsive to a PLK4 inhibitor shows at least a 5% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 10% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 15% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 20% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 25% reduction in tumor size or volume after treatment compared to baseline or a control.
- a subject responsive to a PLK4 inhibitor shows at least a 30% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 35% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 40% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 45% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 50% reduction in tumor size or volume after treatment compared to baseline or a control. In embodiments, a subject responsive to a PLK4 inhibitor shows at least a 60% reduction in tumor size or volume after treatment compared to baseline or a control.
- Biological sample refers to any biological sample taken from a subject. Biological samples include blood, plasma, serum, tumors, tissue, cells, and the like. In embodiments, the biological sample is a blood sample. In embodiments, the biological sample is a peripheral blood sample. In embodiments, the biological sample is a tumor sample. In embodiments, the biological sample is a primary tumor sample. In embodiments, the biological sample is a metastatic tumor sample. In embodiments, the biological sample is a resected tumor sample. In embodiments, the biological sample is a tumor biopsy sample. In embodiments, the biological sample is a resected tumor sample from a primary tumor. In embodiments, the biological sample is a resected tumor sample from a metastisic tumor.
- the biological sample is a tumor biopsy sample from a primary tumor. In embodiments, the biological sample is a tumor biopsy sample from a metastisic tumor. Biological samples can be taken from a subject by methods known in the art, and can be analyzed by methods known in the art.
- a “control” sample or value refers to a sample that serves as a reference, usually a known reference, for comparison to a test sample.
- a test sample can be taken from a patient suspected of having a given disease (cancer) and compared to samples from a known cancer patient, or a known normal (non-disease) individual.
- a control can also represent an average value gathered from a population of similar individuals, e.g., cancer patients or healthy individuals with a similar medical background, same age, weight, etc.
- a control value can also be obtained from the same individual, e.g., from an earlier-obtained sample, prior to disease, or prior to treatment.
- controls can be designed for assessment of any number of parameters.
- a control is a negative control.
- a control comprises the average amount of expression (e.g., protein or mRNA) of infiltration (e.g., number or percentage of cells in a population of cells) in a population of subjects (e.g., with cancer) or in a healthy or general population.
- the control comprises an average amount (e.g.
- control is a standard control. ne of skill in the art will understand which controls are valuable in a given situation and be able to analyze data based on comparisons to control values. Controls are also valuable for determining the significance of data. For example, if values for a given parameter are widely variant in controls, variation in test samples will not be considered as significant.
- a “p53 protein” or “p53” as referred to herein includes any of the recombinant or naturally-occurring forms of cellular tumor antigen p53 (p53) or variants or homologs thereof that maintain p53 activity (e.g. within at least 50%, 80%, 90%, 95%, 96%, 97%, 98%, 99% or 100% activity compared to p53).
- the variants or homologs have at least 90%, 95%, 96%, 97%, 98%, 99% or 100% amino acid sequence identity across the whole sequence or a portion of the sequence (e.g. a 50, 100, 150 or 200 continuous amino acid portion) compared to a naturally occurring p53.
- wild type p53 or wild type p53 protein refers to a p53 protein that is substantially identical to the protein identified by the UniProt reference number P04637 or a variant or homolog having substantial identity thereto.
- the p53 protein is a wild type p53 protein.
- mutant protein refers to a protein having aberrant biological activity compared to a non-mutant protein (e.g., a non-mutant p53 protein may be a protein identified by UniProt reference number P04637).
- a mutant protein may have increased or decreased biological activity or the mutant protein may have no detectable biological activity compared to the corresponding non-mutant protein (e.g. a non-mutant p53 protein identified by UniProt reference number P04637).
- a mutant protein may have biological activity distinct from the non-mutant protein (e.g. a non-mutant p53 protein identified by UniProt reference number P04637).
- Mutant proteins are encoded by DNA sequences (e.g., genes) including base pair insertions, deletions, or substitutions that are absent in the corresponding non-mutant protein and that result in the modulation (e.g., increased, decreased, loss of function, gain of function) of biological activity compared to the non-mutant protein.
- DNA sequences e.g., genes
- base pair insertions, deletions, or substitutions that are absent in the corresponding non-mutant protein and that result in the modulation (e.g., increased, decreased, loss of function, gain of function) of biological activity compared to the non-mutant protein.
- mutant refers to a p53 protein with aberrant biological activity compared to a non-mutant (wild type) p53 protein (e.g., the p53 protein identified by UniProt reference number P04637).
- the mutant p53 protein as referred to herein fails to act as a suppressor of cell division and may exist at elevated intracellular levels compared to a non-mutant p53 protein. Mutations in the p53 gene (e.g., the human p53 gene identified by Ensebl reference number ENSG00000141510) have been found to correlate with aggressive disease characteristics and metastasis.
- p53 positive or “p53+” as used herein refers to a cancer, tumor, or cell that expresses a p53 protein.
- Wild type p53 positive refers to a cancer, tumor, or cell that expresses a wild type p53 protein.
- Metant p53 positive refers to a cancer, tumor, or cell that expresses a mutant p53 protein.
- a cancer is identified as p53 positive when there is a positive test result of a biological sample by immunohistochemical (IHC) analysis.
- IHC immunohistochemical
- a cancer is identified as p53 positive when at least 1% of the cells from a biological sample test positive by IHC.
- a cancer is identified as p53 positive when at least 2% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 3% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 4% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 5% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 6% of the cells from a biological sample test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 7% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 8% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 9% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 10% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 11% of the cells from a biological sample test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 12% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 13% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 14% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 15% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 16% of the cells from a biological sample test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 17% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 18% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 19% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 20% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 21% of the cells from a biological sample test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 22% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 23% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 24% of the cells from a biological sample test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 25% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 26% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 27% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 28% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 29% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 30% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 31% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 32% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 33% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 34% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 35% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 36% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 37% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 38% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 39% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 40% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 41% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 42% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 43% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 44% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 45% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 46% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 47% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 48% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 49% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 50% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 51% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 52% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 53% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 54% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 55% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 56% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 57% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 58% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 59% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 60% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 61% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 62% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 63% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 64% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 65% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 66% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 67% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 68% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 69% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 70% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 71% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 72% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 73% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 74% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 75% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 76% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 77% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 78% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 79% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 80% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 81% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 82% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 83% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 84% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 85% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 86% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 87% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 88% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 89% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 90% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 91% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 92% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 93% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when at least 94% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 95% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 96% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 97% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 98% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when at least 99% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when about 1% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 5% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 10% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 25% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 50% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 55% to about 100% of the cells test positive by IHC analysis.
- a cancer is identified as p53 positive when about 60% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 65% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 70% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 75% to about 100% of the cells test positive by IHC analysis. In embodiments, a cancer is identified as p53 positive when about 80% to about 100% of the cells test positive by IHC analysis.
- Percentage of sequence identity is determined by comparing two optimally aligned sequences over a comparison window, wherein the portion of the polynucleotide or polypeptide sequence in the comparison window may comprise additions or deletions (i.e., gaps) as compared to the reference sequence (which does not comprise additions or deletions) for optimal alignment of the two sequences. The percentage is calculated by determining the number of positions at which the identical nucleic acid base or amino acid residue occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison and multiplying the result by 100 to yield the percentage of sequence identity.
- nucleic acids or polypeptide sequences refer to two or more sequences or subsequences that are the same or have a specified percentage of amino acid residues or nucleotides that are the same (i.e., 60% identity, optionally 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, or 99% identity over a specified region, e.g., of the entire polypeptide sequences of the invention or individual domains of the polypeptides of the invention), when compared and aligned for maximum correspondence over a comparison window, or designated region as measured using one of the following sequence comparison algorithms or by manual alignment and visual inspection.
- sequences are then said to be “substantially identical.”
- This definition also refers to the complement of a test sequence.
- the identity exists over a region that is at least about 50 nucleotides in length, or more preferably over a region that is 100 to 500 or 1000 or more nucleotides in length.
- sequence comparison typically one sequence acts as a reference sequence, to which test sequences are compared.
- test and reference sequences are entered into a computer, subsequence coordinates are designated, if necessary, and sequence algorithm program parameters are designated. Default program parameters can be used, or alternative parameters can be designated.
- sequence comparison algorithm then calculates the percent sequence identities for the test sequences relative to the reference sequence, based on the program parameters.
- a “comparison window”, as used herein, includes reference to a segment of any one of the number of contiguous positions selected from the group consisting of, e.g., a full length sequence or from 20 to 600, about 50 to about 200, or about 100 to about 150 amino acids or nucleotides in which a sequence may be compared to a reference sequence of the same number of contiguous positions after the two sequences are optimally aligned.
- Methods of alignment of sequences for comparison are well-known in the art.
- Optimal alignment of sequences for comparison can be conducted, e.g., by the local homology algorithm of Smith and Waterman (1970) Adv. Appl. Math. 2:482c, by the homology alignment algorithm of Needleman and Wunsch (1970) J. Mol.
- HSPs high scoring sequence pairs
- T is referred to as the neighborhood word score threshold (Altschul et al., supra). These initial neighborhood word hits act as seeds for initiating searches to find longer HSPs containing them. The word hits are extended in both directions along each sequence for as far as the cumulative alignment score can be increased. Cumulative scores are calculated using, for nucleotide sequences, the parameters M (reward score for a pair of matching residues; always >0) and N (penalty score for mismatching residues; always ⁇ 0). For amino acid sequences, a scoring matrix is used to calculate the cumulative score.
- Extension of the word hits in each direction are halted when: the cumulative alignment score falls off by the quantity X from its maximum achieved value; the cumulative score goes to zero or below, due to the accumulation of one or more negative-scoring residue alignments; or the end of either sequence is reached.
- the BLAST algorithm parameters W, T, and X determine the sensitivity and speed of the alignment.
- the BLAST algorithm also performs a statistical analysis of the similarity between two sequences (see, e.g., Karlin and Altschul (1993) Proc. Natl. Acad. Sci. USA 90:5873-5787).
- One measure of similarity provided by the BLAST algorithm is the smallest sum probability (P(N)), which provides an indication of the probability by which a match between two nucleotide or amino acid sequences would occur by chance.
- P(N) the smallest sum probability
- a nucleic acid is considered similar to a reference sequence if the smallest sum probability in a comparison of the test nucleic acid to the reference nucleic acid is less than about 0.2, more preferably less than about 0.01, and most preferably less than about 0.001.
- nucleic acid sequences or polypeptides are substantially identical is that the polypeptide encoded by the first nucleic acid is immunologically cross reactive with the antibodies raised against the polypeptide encoded by the second nucleic acid, as described below.
- a polypeptide is typically substantially identical to a second polypeptide, for example, where the two peptides differ only by conservative substitutions.
- Another indication that two nucleic acid sequences are substantially identical is that the two molecules or their complements hybridize to each other under stringent conditions, as described below.
- Yet another indication that two nucleic acid sequences are substantially identical is that the same primers can be used to amplify the sequence.
- substituent groups are specified by their conventional chemical formulae, written from left to right, they equally encompass the chemically identical substituents that would result from writing the structure from right to left, e.g., —CH 2 O— is equivalent to —OCH 2 —.
- alkyl by itself or as part of another substituent, means, unless otherwise stated, a straight (i.e., unbranched) or branched carbon chain (or carbon), or combination thereof, which may be fully saturated, mono- or polyunsaturated and can include mono-, di- and multivalent radicals, having the number of carbon atoms designated (i.e., C 1 -C 10 means one to ten carbons).
- Alkyl is an uncyclized chain.
- saturated hydrocarbon radicals include, but are not limited to, groups such as methyl, ethyl, n-propyl, isopropyl, n-butyl, t-butyl, isobutyl, sec-butyl, (cyclohexyl)methyl, homologs and isomers of, for example, n-pentyl, n-hexyl, n-heptyl, n-octyl, and the like.
- An unsaturated alkyl group is one having one or more double bonds or triple bonds.
- unsaturated alkyl groups include, but are not limited to, vinyl, 2-propenyl, crotyl, 2-isopentenyl, 2-(butadienyl), 2,4-pentadienyl, 3-(1,4-pentadienyl), ethynyl, 1- and 3-propynyl, 3-butynyl, and the higher homologs and isomers.
- An alkoxy is an alkyl attached to the remainder of the molecule via an oxygen linker (—O—).
- alkylene by itself or as part of another substituent, means, unless otherwise stated, a divalent radical derived from an alkyl, as exemplified, but not limited by, —CH 2 CH 2 CH 2 CH 2 —.
- an alkyl (or alkylene) group will have from 1 to 24 carbon atoms, with those groups having 10 or fewer carbon atoms being preferred herein.
- a “lower alkyl” or “lower alkylene” is a shorter chain alkyl or alkylene group, generally having eight or fewer carbon atoms.
- alkenylene by itself or as part of another substituent, means, unless otherwise stated, a divalent radical derived from an alkene.
- heteroalkyl by itself or in combination with another term, means, unless otherwise stated, a stable straight or branched chain, or combinations thereof, including at least one carbon atom and at least one heteroatom (e.g. selected from the group consisting of O, N, P, Si, and S, and wherein the nitrogen and sulfur atoms may optionally be oxidized, and the nitrogen heteroatom may optionally be quaternized).
- the heteroatom(s) O, N, P, S, B, As, and Si may be placed at any interior position of the heteroalkyl group or at the position at which the alkyl group is attached to the remainder of the molecule.
- Heteroalkyl is an uncyclized chain.
- Examples include, but are not limited to: —CH 2 —CH 2 —O—CH 3 , —CH 2 —CH 2 —NH—CH 3 , —CH 2 —CH 2 —N(CH 3 )—CH 3 , —CH 2 —S—CH 2 —CH 3 , —CH 2 —CH 2 , —S(O)—CH 3 , —CH 2 —CH 2 —S(O) 2 —CH 3 , —CH ⁇ CHO—CH 3 , —Si(CH 3 ) 3 , —CH 2 —CH ⁇ N—OCH 3 , —CH ⁇ CH—N(CH 3 )—CH 3 , —O—CH 3 , —O—CH 2 —CH 3 , and —CN. Up to two or three heteroatoms may be consecutive, such as, for example, —CH 2 —NH—OCH 3 and —CH 2 —O—Si(CH 3 ) 3 .
- heteroalkylene by itself or as part of another substituent, means, unless otherwise stated, a divalent radical derived from heteroalkyl, as exemplified, but not limited by, —CH 2 —CH 2 —S—CH 2 —CH 2 — and —CH 2 —S—CH 2 —CH 2 —NH—CH 2 —.
- heteroatoms can also occupy either or both of the chain termini (e.g., alkyleneoxy, alkylenedioxy, alkyleneamino, alkylenediamino, and the like).
- heteroalkyl groups include those groups that are attached to the remainder of the molecule through a heteroatom, such as —C(O)R′, —C(O)NR′, —NR′R′′, —OR′, —SR′, and/or —SO 2 R′.
- heteroalkyl is recited, followed by recitations of specific heteroalkyl groups, such as —NR′R′′ or the like, it will be understood that the terms heteroalkyl and —NR′R′′ are not redundant or mutually exclusive. Rather, the specific heteroalkyl groups are recited to add clarity. Thus, the term “heteroalkyl” should not be interpreted herein as excluding specific heteroalkyl groups, such as —NR′R′′ or the like.
- cycloalkyl and heterocycloalkyl mean, unless otherwise stated, cyclic versions of “alkyl” and “heteroalkyl,” respectively. Cycloalkyl and heteroalkyl are not aromatic. Additionally, for heterocycloalkyl, a heteroatom can occupy the position at which the heterocycle is attached to the remainder of the molecule. Examples of cycloalkyl include, but are not limited to, cyclopropyl, cyclobutyl, cyclopentyl, cyclohexyl, 1-cyclohexenyl, 3-cyclohexenyl, cycloheptyl, and the like.
- heterocycloalkyl examples include, but are not limited to, 1-(1,2,5,6-tetrahydropyridyl), 1-piperidinyl, 2-piperidinyl, 3-piperidinyl, 4-morpholinyl, 3-morpholinyl, tetrahydrofuran-2-yl, tetrahydrofuran-3-yl, tetrahydrothien-2-yl, tetrahydrothien-3-yl, 1-piperazinyl, 2-piperazinyl, and the like.
- a “cycloalkylene” and a “heterocycloalkylene,” alone or as part of another substituent, means a divalent radical derived from a cycloalkyl and heterocycloalkyl, respectively.
- halo or “halogen,” by themselves or as part of another substituent, mean, unless otherwise stated, a fluorine, chlorine, bromine, or iodine atom. Additionally, terms such as “haloalkyl” are meant to include monohaloalkyl and polyhaloalkyl.
- halo(C 1 -C 4 )alkyl includes, but is not limited to, fluoromethyl, difluoromethyl, trifluoromethyl, 2,2,2-trifluoroethyl, 4-chlorobutyl, 3-bromopropyl, and the like.
- acyl means, unless otherwise stated, —C(O)R where R is a substituted or unsubstituted alkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- aryl means, unless otherwise stated, a polyunsaturated, aromatic, hydrocarbon substituent, which can be a single ring or multiple rings (preferably from 1 to 3 rings) that are fused together (i.e., a fused ring aryl) or linked covalently.
- a fused ring aryl refers to multiple rings fused together wherein at least one of the fused rings is an aryl ring.
- heteroaryl refers to aryl groups (or rings) that contain at least one heteroatom such as N, O, or S, wherein the nitrogen and sulfur atoms are optionally oxidized, and the nitrogen atom(s) are optionally quaternized.
- heteroaryl includes fused ring heteroaryl groups (i.e., multiple rings fused together wherein at least one of the fused rings is a heteroaromatic ring).
- a 5,6-fused ring heteroarylene refers to two rings fused together, wherein one ring has 5 members and the other ring has 6 members, and wherein at least one ring is a heteroaryl ring.
- a 6,6-fused ring heteroarylene refers to two rings fused together, wherein one ring has 6 members and the other ring has 6 members, and wherein at least one ring is a heteroaryl ring.
- a 6,5-fused ring heteroarylene refers to two rings fused together, wherein one ring has 6 members and the other ring has 5 members, and wherein at least one ring is a heteroaryl ring.
- a heteroaryl group can be attached to the remainder of the molecule through a carbon or heteroatom.
- Non-limiting examples of aryl and heteroaryl groups include phenyl, naphthyl, pyrrolyl, pyrazolyl, pyridazinyl, triazinyl, pyrimidinyl, imidazolyl, pyrazinyl, purinyl, oxazolyl, isoxazolyl, thiazolyl, furyl, thienyl, pyridyl, pyrimidyl, benzothiazolyl, benzoxazoyl benzimidazolyl, benzofuran, isobenzofuranyl, indolyl, isoindolyl, benzothiophenyl, isoquinolyl, quinoxalinyl, quinolyl, 1-naphthyl, 2-naphthyl, 4-biphenyl, 1-pyrrolyl, 2-pyrrolyl, 3-pyrrolyl, 3-pyrazolyl, 2-imidazolyl, 4-imidazo
- arylene and heteroarylene independently or as part of another substituent, mean a divalent radical derived from an aryl and heteroaryl, respectively.
- a heteroaryl group substituent may be —O— bonded to a ring heteroatom nitrogen.
- a “fused ring aryl-heterocycloalkyl” is an aryl fused to a heterocycloalkyl.
- a “fused ring heteroaryl-heterocycloalkyl” is a heteroaryl fused to a heterocycloalkyl.
- a “fused ring heterocycloalkyl-cycloalkyl” is a heterocycloalkyl fused to a cycloalkyl.
- a “fused ring heterocycloalkyl-heterocycloalkyl” is a heterocycloalkyl fused to another heterocycloalkyl.
- Fused ring aryl-heterocycloalkyl, fused ring heteroaryl-heterocycloalkyl, fused ring heterocycloalkyl-cycloalkyl, or fused ring heterocycloalkyl-heterocycloalkyl may each independently be unsubstituted or substituted with one or more of the substituents described herein.
- Fused ring aryl-heterocycloalkyl, fused ring heteroaryl-heterocycloalkyl, fused ring heterocycloalkyl-cycloalkyl, or fused ring heterocycloalkyl-heterocycloalkyl may each independently be named according to the size of each of the fused rings.
- 6,5 aryl-heterocycloalkyl fused ring describes a 6 membered aryl moiety fused to a 5 membered heterocycloalkyl.
- Spirocyclic rings are two or more rings wherein adjacent rings are attached through a single atom.
- the individual rings within spirocyclic rings may be identical or different.
- Individual rings in spirocyclic rings may be substituted or unsubstituted and may have different substituents from other individual rings within a set of spirocyclic rings.
- Possible substituents for individual rings within spirocyclic rings are the possible substituents for the same ring when not part of spirocyclic rings (e.g. substituents for cycloalkyl or heterocycloalkyl rings).
- Spirocylic rings may be substituted or unsubstituted cycloalkyl, substituted or unsubstituted cycloalkylene, substituted or unsubstituted heterocycloalkyl or substituted or unsubstituted heterocycloalkylene and individual rings within a spirocyclic ring group may be any of the immediately previous list, including having all rings of one type (e.g. all rings being substituted heterocycloalkylene wherein each ring may be the same or different substituted heterocycloalkylene).
- heterocyclic spirocyclic rings means a spirocyclic rings wherein at least one ring is a heterocyclic ring and wherein each ring may be a different ring.
- substituted spirocyclic rings means that at least one ring is substituted and each substituent may optionally be different.
- oxo means an oxygen that is double bonded to a carbon atom.
- Substituents for the alkyl and heteroalkyl radicals can be one or more of a variety of groups selected from, but not limited to, —OR′, ⁇ O, ⁇ NR′, ⁇ N—OR′, —NR′R′′, —SR′, -halogen, —SiR′R′′R′′′, —OC(O)R′, —C(O)R′, —CO 2 R′, —CONR′R′′, —OC(O)NR′R′′, —NR′′C(O)R′, —NR′—C(O)NR′′R′′′, —NR′′C(O) 2 R′, —NR—C(NR′R′′R′′′) ⁇ NR′′′′, —NR—C(NR′R′′R′′′) ⁇ NR′′′′,
- R, R′, R′′, R′′′, and R′′′′ each preferably independently refer to hydrogen, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl (e.g., aryl substituted with 1-3 halogens), substituted or unsubstituted heteroaryl, substituted or unsubstituted alkyl, alkoxy, or thioalkoxy groups, or arylalkyl groups.
- aryl e.g., aryl substituted with 1-3 halogens
- substituted or unsubstituted heteroaryl substituted or unsubstituted alkyl, alkoxy, or thioalkoxy groups, or arylalkyl groups.
- each of the R groups is independently selected as are each R′, R′′, R′′′, and R′′′′ group when more than one of these groups is present.
- R′ and R′′ are attached to the same nitrogen atom, they can be combined with the nitrogen atom to form a 4-, 5-, 6-, or 7-membered ring.
- —NR′R′′ includes, but is not limited to, 1-pyrrolidinyl and 4-morpholinyl.
- alkyl is meant to include groups including carbon atoms bound to groups other than hydrogen groups, such as haloalkyl (e.g., —CF 3 and —CH 2 CF 3 ) and acyl (e.g., —C(O)CH 3 , —C(O)CF 3 , —C(O)CH 2 OCH 3 , and the like).
- haloalkyl e.g., —CF 3 and —CH 2 CF 3
- acyl e.g., —C(O)CH 3 , —C(O)CF 3 , —C(O)CH 2 OCH 3 , and the like.
- substituents for the aryl and heteroaryl groups are varied and are selected from, for example: —OR′, —NR′R′′, —SR′, -halogen, —SiR′R′′R′′′, —OC(O)R′, —C(O)R′, —CO 2 R′, —CONR′R′′, —OC(O)NR′R′′, —NR′′C(O)R′, —NR′—C(O)NR′′R′′′, —NR′′C(O) 2 R′, —NR—C(NR′R′′R′′′) ⁇ NR′′′′, —NR—C(NR′R′′) ⁇ NR′′′, —S(O)R′, —S(O) 2 R′, —S(O) 2 NR′R′′, —NRSO 2 R′, —NR′NR′′R′′′, —ONR′R′′, —NR′C(O)NR
- Substituents for rings may be depicted as substituents on the ring rather than on a specific atom of a ring (commonly referred to as a floating substituent).
- the substituent may be attached to any of the ring atoms (obeying the rules of chemical valency) and in the case of fused rings or spirocyclic rings, a substituent depicted as associated with one member of the fused rings or spirocyclic rings (a floating substituent on a single ring), may be a substituent on any of the fused rings or spirocyclic rings (a floating substituent on multiple rings).
- the multiple substituents may be on the same atom, same ring, different atoms, different fused rings, different spirocyclic rings, and each substituent may optionally be different.
- a point of attachment of a ring to the remainder of a molecule is not limited to a single atom (a floating substituent)
- the attachment point may be any atom of the ring and in the case of a fused ring or spirocyclic ring, any atom of any of the fused rings or spirocyclic rings while obeying the rules of chemical valency.
- a ring, fused rings, or spirocyclic rings contain one or more ring heteroatoms and the ring, fused rings, or spirocyclic rings are shown with one more floating substituents (including, but not limited to, points of attachment to the remainder of the molecule), the floating substituents may be bonded to the heteroatoms.
- the ring heteroatoms are shown bound to one or more hydrogens (e.g. a ring nitrogen with two bonds to ring atoms and a third bond to a hydrogen) in the structure or formula with the floating substituent, when the heteroatom is bonded to the floating substituent, the substituent will be understood to replace the hydrogen, while obeying the rules of chemical valency.
- Two or more substituents may optionally be joined to form aryl, heteroaryl, cycloalkyl, or heterocycloalkyl groups.
- Such so-called ring-forming substituents are typically, though not necessarily, found attached to a cyclic base structure.
- the ring-forming substituents may be attached to adjacent members of the base structure.
- two ring-forming substituents attached to adjacent members of a cyclic base structure create a fused ring structure.
- the ring-forming substituents may be attached to a single member of the base structure.
- two ring-forming substituents attached to a single member of a cyclic base structure create a spirocyclic structure.
- the ring-forming substituents may be attached to non-adjacent members of the base structure.
- Two of the substituents on adjacent atoms of the aryl or heteroaryl ring may optionally form a ring of the formula -T—C(O)—(CRR′) q —U—, wherein T and U are independently —NR—, —O—, —CRR′—, or a single bond, and q is an integer of from 0 to 3.
- two of the substituents on adjacent atoms of the aryl or heteroaryl ring may optionally be replaced with a substituent of the formula -A-(CH 2 ) r —B—, wherein A and B are independently —CRR′—, —O—, —NR—, —S—, —S(O)—, —S(O) 2 —, —S(O) 2 NR′—, or a single bond, and r is an integer of from 1 to 4.
- One of the single bonds of the new ring so formed may optionally be replaced with a double bond.
- two of the substituents on adjacent atoms of the aryl or heteroaryl ring may optionally be replaced with a substituent of the formula —(CRR′) s —X′—(C′′R′′R′′′) d —, where s and d are independently integers of from 0 to 3, and X′ is —O—, —NR′—, —S—, —S(O)—, —S(O) 2 —, or —S(O) 2 NR′—.
- R, R′, R′′, and R′′′ are preferably independently selected from hydrogen, substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, and substituted or unsubstituted heteroaryl.
- heteroatom or “ring heteroatom” are meant to include, oxygen (O), nitrogen (N), sulfur (S), phosphorus (P), Boron (B), Arsenic (As), and silicon (Si).
- a “substituent group,” as used herein, means a group selected from the following moieties: (A) oxo, halogen, —CF 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —SO 2 Cl, —SO 3 H, —SO 4 H, —SO 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHSO 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted heterocycloalkyl, unsubstituted aryl, unsubstituted heteroary
- a “size-limited substituent” or “size-limited substituent group,” as used herein, means a group selected from all of the substituents described above for a “substituent group,” wherein each substituted or unsubstituted alkyl is a substituted or unsubstituted C 1 -C 20 alkyl, each substituted or unsubstituted heteroalkyl is a substituted or unsubstituted 2 to 20 membered heteroalkyl, each substituted or unsubstituted cycloalkyl is a substituted or unsubstituted C 3 -C 8 cycloalkyl, and each substituted or unsubstituted heterocycloalkyl is a substituted or unsubstituted 3 to 8 membered heterocycloalkyl.
- a “lower substituent” or “lower substituent group,” as used herein, means a group selected from all of the substituents described above for a “substituent group,” wherein each substituted or unsubstituted alkyl is a substituted or unsubstituted C 1 -C 8 alkyl, each substituted or unsubstituted heteroalkyl is a substituted or unsubstituted 2 to 8 membered heteroalkyl, each substituted or unsubstituted cycloalkyl is a substituted or unsubstituted C 3 -C 7 cycloalkyl, and each substituted or unsubstituted heterocycloalkyl is a substituted or unsubstituted 3 to 7 membered heterocycloalkyl.
- Each substituted group described in the compounds herein may be substituted with at least one substituent group. More specifically, each substituted alkyl, substituted heteroalkyl, substituted cycloalkyl, substituted heterocycloalkyl, substituted aryl, substituted heteroaryl, substituted alkylene, substituted heteroalkylene, substituted cycloalkylene, substituted heterocycloalkylene, substituted arylene, and/or substituted heteroarylene described in the compounds herein may be substituted with at least one substituent group. At least one or all of these groups may be substituted with at least one size-limited substituent group. At least one or all of these groups may be substituted with at least one lower substituent group.
- Each substituted or unsubstituted alkyl may be a substituted or unsubstituted C 1 -C 20 alkyl
- each substituted or unsubstituted heteroalkyl may be a substituted or unsubstituted 2 to 20 membered heteroalkyl
- each substituted or unsubstituted cycloalkyl may be a substituted or unsubstituted C 3 -C 8 cycloalkyl
- each substituted or unsubstituted heterocycloalkyl may be a substituted or unsubstituted 3 to 8 membered heterocycloalkyl.
- Each substituted or unsubstituted alkylene may be a substituted or unsubstituted C 1 -C 20 alkylene
- each substituted or unsubstituted heteroalkylene may be a substituted or unsubstituted 2 to 20 membered heteroalkylene
- each substituted or unsubstituted cycloalkylene may be a substituted or unsubstituted C 3 -C 8 cycloalkylene
- each substituted or unsubstituted heterocycloalkylene may be a substituted or unsubstituted 3 to 8 membered heterocycloalkylene.
- Each substituted or unsubstituted alkyl may be a substituted or unsubstituted C 1 -C 8 alkyl
- each substituted or unsubstituted heteroalkyl may be a substituted or unsubstituted 2 to 8 membered heteroalkyl
- each substituted or unsubstituted cycloalkyl may be a substituted or unsubstituted C 3 -C 7 cycloalkyl
- each substituted or unsubstituted heterocycloalkyl may be a substituted or unsubstituted 3 to 7 membered heterocycloalkyl.
- Each substituted or unsubstituted alkylene may be a substituted or unsubstituted C 1 -C 8 alkylene
- each substituted or unsubstituted heteroalkylene may be a substituted or unsubstituted 2 to 8 membered heteroalkylene
- each substituted or unsubstituted cycloalkylene may be a substituted or unsubstituted C 3 -C 7 cycloalkylene
- each substituted or unsubstituted heterocycloalkylene may be a substituted or unsubstituted 3 to 7 membered heterocycloalkylene.
- Certain compounds herein possess asymmetric carbon atoms (optical or chiral centers) or double bonds; the enantiomers, racemates, diastereomers, tautomers, geometric isomers, stereoisometric forms that may be defined, in terms of absolute stereochemistry, as (R)- or (S)- or, as (D)- or (L)- for amino acids, and individual isomers are encompassed within the scope of the compounds described herein.
- the compounds described herein do not include those which are known in art to be too unstable to synthesize and/or isolate.
- the compounds described herein also are meant to include compounds in racemic and optically pure forms.
- Optically active (R)- and (S)-, or (D)- and (L)-isomers may be prepared using chiral synthons or chiral reagents, or resolved using conventional techniques.
- the compounds described herein contain olefinic bonds or other centers of geometric asymmetry, and unless specified otherwise, it is intended that the compounds include both E and Z geometric isomers.
- isomers refers to compounds having the same number and kind of atoms, and hence the same molecular weight, but differing in respect to the structural arrangement or configuration of the atoms.
- tautomer refers to one of two or more structural isomers which exist in equilibrium and which are readily converted from one isomeric form to another.
- structures depicted herein are also meant to include all stereochemical forms of the structure; i.e., the R and S configurations for each asymmetric center. Therefore, single stereochemical isomers as well as enantiomeric and diastereomeric mixtures of the present compounds, generally recognized as stable by those skilled in the art, are within the scope of the compounds described herein.
- structures depicted herein are also meant to include compounds which differ only in the presence of one or more isotopically enriched atoms.
- compounds having the present structures except for the replacement of a hydrogen by a deuterium or tritium, or the replacement of a carbon by 13 C- or 14 C-enriched carbon are within the scope of the compounds described herein.
- the compounds described herein may also contain unnatural proportions of atomic isotopes at one or more of the atoms that constitute such compounds.
- the compounds may be radiolabeled with radioactive isotopes, such as for example tritium (3H), iodine-125 ( 125 I), or carbon-14 ( 14 C). All isotopic variations of the compounds described herein, whether radioactive or not, are encompassed within the scope of the compounds described herein.
- R substituent
- R-substituted the group may be referred to as “R-substituted.”
- R-substituted the moiety is substituted with at least one R substituent and each R substituent is optionally different.
- a particular R group is present in the description of a chemical genus (such as Formula (I))
- a Roman decimal symbol may be used to distinguish each appearance of that particular R group.
- each R 13 substituent may be distinguished as R 13.1 , R 133.2 , R 13.3 , R 13.4 , etc., wherein each of R 13.1 , R 13.2 , R 13.3C , R 13.4 , etc. is defined within the scope of the definition of R 13 and optionally differently.
- Tozasertib is the compound having PubChem CID 5494449, and is also known as VX-680 and MK-0457.
- pharmaceutically acceptable salts is meant to include salts of the active compounds that are prepared with relatively nontoxic acids or bases, depending on the particular substituents found on the compounds described herein.
- base addition salts can be obtained by contacting the neutral form of such compounds with a sufficient amount of the desired base, either neat or in a suitable inert solvent.
- pharmaceutically acceptable base addition salts include sodium, potassium, calcium, ammonium, organic amino, or magnesium salt, or a similar salt.
- acid addition salts can be obtained by contacting the neutral form of such compounds with a sufficient amount of the desired acid, either neat or in a suitable inert solvent.
- Examples of pharmaceutically acceptable acid addition salts include those derived from inorganic acids like hydrochloric, hydrobromic, nitric, carbonic, monohydrogencarbonic, phosphoric, monohydrogenphosphoric, dihydrogenphosphoric, sulfuric, monohydrogensulfuric, hydriodic, or phosphorous acids and the like, as well as the salts derived from relatively nontoxic organic acids like acetic, propionic, isobutyric, maleic, malonic, benzoic, succinic, suberic, fumaric, lactic, mandelic, phthalic, benzenesulfonic, p-tolylsulfonic, citric, tartaric, oxalic, methanesulfonic, and the like.
- inorganic acids like hydrochloric, hydrobromic, nitric, carbonic, monohydrogencarbonic, phosphoric, monohydrogenphosphoric, dihydrogenphosphoric, sulfuric, monohydrogensulfuric, hydriodic,
- salts of amino acids such as arginate and the like, and salts of organic acids like glucuronic or galactunoric acids and the like (see, for example, Berge et al., “Pharmaceutical Salts”, Journal of Pharmaceutical Science, 1977, 66, 1-19).
- Compounds described herein may contain both basic and acidic functionalities that allow the compounds to be converted into either base or acid addition salts.
- the neutral forms of the compounds can be regenerated by contacting the salt with a base or acid and isolating the parent compound in the conventional manner.
- the parent form of the compound could differ from the various salt forms in certain physical properties, such as solubility in polar solvents.
- the compounds described herein may be provided in a prodrug form.
- Prodrugs of the compounds described herein include those compounds that readily undergo chemical or enzymatic changes under physiological conditions to provide the compounds described herein.
- prodrugs can be converted to the compounds described herein by chemical or biochemical methods in an ex vivo environment. For example, prodrugs can be slowly converted to the compounds described herein when placed in a transdermal patch reservoir with a suitable enzyme or chemical reagent.
- Certain compounds described herein can exist in unsolvated forms as well as solvated forms, including hydrated forms. In general, the solvated forms are equivalent to unsolvated forms and are encompassed within the scope of the compounds described herein. Certain compounds described herein may exist in multiple crystalline or amorphous forms.
- treating refers to any indicia of success in the therapy or amelioration of an injury, disease, pathology or condition, including any objective or subjective parameter such as abatement; remission; diminishing of symptoms or making the injury, pathology or condition more tolerable to the patient; slowing in the rate of degeneration or decline; making the final point of degeneration less debilitating; improving a patient's physical or mental well-being.
- the treatment of symptoms can be based on objective or subjective parameters; including the results of a physical examination, neuropsychiatric exams, and/or a psychiatric evaluation.
- treating a cancer tumor means preventing an increase in size or volume of the cancer tumor.
- the cancer tumor is a solid tumor.
- treating a cancer tumor includes decreasing the size of volume of a cancer tumor.
- treating a cancer tumor includes eliminating the cancer tumor altogether.
- a cancer tumor is eliminated when it is not detectable by an imaging test such as magnetic resonance imaging (MRI), a positron emission tomography (PET) scan, X-ray computed tomography (CT), ultrasound, or single-photon emission computed tomography (SPECT).
- treating a cancer tumor further comprises reducing or preventing metastasis of the cancer tumor.
- an “effective amount” is an amount sufficient to accomplish a stated purpose (e.g. achieve the effect for which it is administered, treat a disease, reduce enzyme activity, or reduce one or more symptoms of a disease or condition).
- An example of an “effective amount” is an amount sufficient to contribute to the treatment, prevention, or reduction of a symptom or symptoms of a disease, which herein is referred to as a “therapeutically effective amount.”
- a “reduction” of a symptom or symptoms means decreasing of the severity or frequency of the symptom(s), or elimination of the symptom(s).
- a “prophylactically effective amount” of a drug is an amount of a drug that, when administered to a subject, will have the intended prophylactic effect, e.g., preventing or delaying the onset (or reoccurrence) of an injury, disease, pathology or condition, or reducing the likelihood of the onset (or reoccurrence) of an injury, disease, pathology, or condition, or their symptoms.
- the full prophylactic effect does not necessarily occur by administration of one dose, and may occur only after administration of a series of doses.
- a prophylactically effective amount may be administered in one or more administrations.
- the therapeutically effective amount can be initially determined from cell culture assays.
- Target concentrations will be those concentrations of active compound(s) that are capable of achieving the methods described herein, as measured using the methods described herein or known in the art.
- therapeutically effective amounts for use in humans can also be determined from animal models.
- a dose for humans can be formulated to achieve a concentration that has been found to be effective in animals.
- the dosage in humans can be adjusted by monitoring compounds effectiveness and adjusting the dosage upwards or downwards, as described above. Adjusting the dose to achieve maximal efficacy in humans based on the methods described above and other methods is well within the capabilities of the ordinarily skilled artisan.
- Dosages may be varied depending upon the requirements of the patient and the compound being employed.
- the dose administered to a patient, in the context of the compounds described herein should be sufficient to effect a beneficial therapeutic response in the patient over time.
- the size of the dose also will be determined by the existence, nature, and extent of any adverse side-effects. Determination of the proper dosage for a particular situation is within the skill of the practitioner. Generally, treatment is initiated with smaller dosages which are less than the optimum dose of the compound. Thereafter, the dosage is increased by small increments until the optimum effect under circumstances is reached.
- an effective prophylactic or therapeutic treatment regimen can be planned that does not cause substantial toxicity and yet is effective to treat the clinical symptoms demonstrated by the particular patient.
- This planning should involve the careful choice of active compound by considering factors such as compound potency, relative bioavailability, patient body weight, presence and severity of adverse side effects, preferred mode of administration and the toxicity profile of the selected agent.
- “Pharmaceutically acceptable excipient” and “pharmaceutically acceptable carrier” refer to a substance that aids the administration of an active agent to and absorption by a subject and can be included in the compositions of the compounds described herein without causing a significant adverse toxicological effect on the patient.
- Non-limiting examples of pharmaceutically acceptable excipients include water, NaCl, normal saline solutions, lactated Ringer's, normal sucrose, normal glucose, binders, fillers, disintegrants, lubricants, coatings, sweeteners, flavors, salt solutions (such as Ringer's solution), alcohols, oils, gelatins, carbohydrates such as lactose, amylose or starch, fatty acid esters, hydroxymethylcellulose, polyvinyl pyrrolidine, and colors, and the like.
- Such preparations can be sterilized and, if desired, mixed with auxiliary agents such as lubricants, preservatives, stabilizers, wetting agents, emulsifiers, salts for influencing osmotic pressure, buffers, coloring, and/or aromatic substances and the like that do not deleteriously react with the compounds described herein.
- auxiliary agents such as lubricants, preservatives, stabilizers, wetting agents, emulsifiers, salts for influencing osmotic pressure, buffers, coloring, and/or aromatic substances and the like that do not deleteriously react with the compounds described herein.
- Pharmaceutically acceptable excipients are described in the Handbook of Pharmaceutical Excipients, 8 th Edition, published by the Pharmaceutical Press (2017), and the United States Food and Drug Administration Inactive Ingredient Database (July 2017), the disclosures of which are incorporated by reference herein in their entirety.
- One of skill in the art will recognize that other pharmaceutical excipients are useful in combination with the compounds described herein.
- preparation is intended to include the formulation of the active compound with encapsulating material as a carrier providing a capsule in which the active component with or without other carriers, is surrounded by a carrier, which is thus in association with it.
- carrier providing a capsule in which the active component with or without other carriers, is surrounded by a carrier, which is thus in association with it.
- cachets and lozenges are included. Tablets, powders, capsules, pills, cachets, and lozenges can be used as solid dosage forms suitable for oral administration.
- administering means oral administration, administration as a suppository, topical contact, intravenous, parenteral, intraperitoneal, intramuscular, intralesional, intrathecal, intranasal or subcutaneous administration, or the implantation of a slow-release device, e.g., a mini-osmotic pump, to a subject.
- Administration is by any route compatible with the selected compound preparation, including parenteral and transmucosal (e.g., buccal, sublingual, palatal, gingival, nasal, vaginal, rectal, or transdermal).
- Parenteral administration includes, e.g., intravenous, intramuscular, intra-arteriole, intradermal, subcutaneous, intraperitoneal, intraventricular, and intracranial.
- Other modes of delivery include, but are not limited to, the use of liposomal formulations, intravenous infusion, transdermal patches, etc.
- compositions disclosed herein can be delivered by transdermally, by a topical route, formulated as applicator sticks, solutions, suspensions, emulsions, gels, creams, ointments, pastes, jellies, paints, powders, and aerosols.
- Oral preparations include tablets, pills, powder, dragees, capsules, liquids, lozenges, cachets, gels, syrups, slurries, suspensions, etc., suitable for ingestion by the patient.
- Solid form preparations include powders, tablets, pills, capsules, cachets, suppositories, and dispersible granules.
- Liquid form preparations include solutions, suspensions, and emulsions, for example, water or water/propylene glycol solutions.
- the pharmaceutical compositions described herein may additionally include components to provide sustained release and/or comfort.
- Such components include high molecular weight, anionic mucomimetic polymers, gelling polysaccharides and finely-divided drug carrier substrates. These components are discussed in greater detail in U.S. Pat. Nos. 4,911,920; 5,403,841; 5,212,162; and 4,861,760. The entire contents of these patents are incorporated herein by reference in their entirety for all purposes.
- the compositions disclosed herein can also be delivered as microspheres for slow release in the body.
- microspheres can be administered via intradermal injection of drug-containing microspheres, which slowly release subcutaneously (see Rao, J. Biomater Sci. Polym. Ed. 7:623-645, 1995; as biodegradable and injectable gel formulations (see, e.g., Gao Pharm. Res. 12:857-863, 1995); or, as microspheres for oral administration (see, e.g., Eyles, J. Pharm. Pharmacol. 49:669-674, 1997).
- compositions of the compounds described herein can be delivered by the use of liposomes which fuse with the cellular membrane or are endocytosed, i.e., by employing receptor ligands attached to the liposome, that bind to surface membrane protein receptors of the cell resulting in endocytosis.
- liposomes particularly where the liposome surface carries receptor ligands specific for target cells, or are otherwise preferentially directed to a specific organ, one can focus the delivery of the compositions of the compounds described herein into the target cells in vivo.
- the compositions can also be delivered as nanoparticles.
- compositions may include compositions wherein the active ingredient (e.g. compounds described herein) is contained in a therapeutically effective amount, i.e., in an amount effective to achieve its intended purpose.
- the actual amount effective for a particular application will depend, inter alia, on the condition being treated.
- such compositions When administered in methods to treat a disease, such compositions will contain an amount of active ingredient effective to achieve the desired result, e.g., modulating the activity of a target molecule, and/or reducing, eliminating, or slowing the progression of disease symptoms.
- the dosage and frequency (single or multiple doses) administered to a mammal can vary depending upon a variety of factors, for example, whether the mammal suffers from another disease, and its route of administration; size, age, sex, health, body weight, body mass index, and diet of the recipient; nature and extent of symptoms of the disease being treated, kind of concurrent treatment, complications from the disease being treated or other health-related problems.
- Other therapeutic regimens or agents can be used in conjunction with the methods and compounds described herein. Adjustment and manipulation of established dosages (e.g., frequency and duration) are well within the ability of those skilled in the art.
- the compounds and complexes described herein can be used in combination with one another, with other active drugs known to be useful in treating a disease (e.g. anti-cancer agents) or with adjunctive agents that may not be effective alone, but may contribute to the efficacy of the active agent.
- active drugs e.g. anti-cancer agents
- adjunctive agents may not be effective alone, but may contribute to the efficacy of the active agent.
- co-administer it is meant that a composition described herein is administered at the same time, just prior to, or just after the administration of one or more additional therapies, for example an anti-cancer agent as described herein.
- additional therapies for example an anti-cancer agent as described herein.
- the compounds described herein can be administered alone or can be co-administered to the patient.
- Co-administration is meant to include simultaneous or sequential administration of the compound individually or in combination (more than one compound or agent).
- the preparations can also be combined, when desired, with other active substances (e.g. anti-cancer agents).
- Co-administration includes administering one active agent (e.g. a compound described herein) within 0.5, 1, 2, 4, 6, 8, 10, 12, 16, 20, or 24 hours of a second active agent (e.g. anti-cancer agents).
- Co-administration includes administering one active agent within 0.5, 1, 2, 4, 6, 8, 10, 12, 16, 20, or 24 hours of a second active agent.
- Co-administration includes administering two active agents simultaneously, approximately simultaneously (e.g., within about 1, 5, 10, 15, 20, or 30 minutes of each other), or sequentially in any order.
- Co-administration can be accomplished by co-formulation, i.e., preparing a single pharmaceutical composition including both active agents.
- the active agents can be formulated separately.
- the active and/or adjunctive agents may be linked or conjugated to one another.
- the compounds described herein may be combined with treatments for cancer such as chemotherapy or radiation therapy.
- a patient is a human.
- a patient is a human adult.
- a patient is a human child.
- “Child” refers to a human from the age of birth to 19 years. In embodiments, “child” refers to a human from the age of birth to 14 years.
- Disease or “condition” refer to a state of being or health status of a patient or subject capable of being treated with the compounds or methods provided herein. Disease as used herein may refer to cancer.
- cancer refers to all types of cancer, neoplasm, malignant or benign tumors found in mammals, including leukemia, carcinomas and sarcomas.
- exemplary cancers include acute myeloid leukemia (“AML”), chronic myelogenous leukemia (“CML”), and cancer of the brain, breast, pancreas, colon, liver, kidney, lung, non-small cell lung, melanoma, ovary, sarcoma, and prostate.
- AML acute myeloid leukemia
- CML chronic myelogenous leukemia
- cancer of the brain breast, pancreas, colon, liver, kidney, lung, non-small cell lung, melanoma, ovary, sarcoma, and prostate.
- Additional examples include, cervix cancers, stomach cancers, head & neck cancers, uterus cancers, mesothelioma, metastatic bone cancer, Medulloblastoma, Hodgkin's Disease, Non-Hodgkin's Lymphoma, multiple myeloma, neuroblastoma, ovarian cancer, rhabdomyosarcoma, primary thrombocytosis, primary macroglobulinemia, primary brain tumors, cancer, malignant pancreatic insulanoma, malignant carcinoid, urinary bladder cancer, premalignant skin lesions, testicular cancer, lymphomas, thyroid cancer, neuroblastoma, esophageal cancer, genitourinary tract cancer, malignant hypercalcemia, endometrial cancer, adrenal cortical cancer, and neoplasms of the endocrine and exocrine pancreas.
- sarcoma generally refers to a tumor which is made up of a substance like the embryonic connective tissue and is generally composed of closely packed cells embedded in a fibrillar or homogeneous substance.
- Sarcomas which can be treated with a combination of antineoplastic thiol-binding mitochondrial oxidant and an anticancer agent include a chondrosarcoma, fibrosarcoma, lymphosarcoma, melanosarcoma, myxosarcoma, osteosarcoma, Abemethy's sarcoma, adipose sarcoma, liposarcoma, alveolar soft part sarcoma, ameloblastic sarcoma, botryoid sarcoma, chloroma sarcoma, chorio carcinoma, embryonal sarcoma, Wilms' tumor sarcoma, endometrial sarcoma, stromal sarcoma, E
- carcinoma refers to a malignant new growth made up of epithelial cells tending to infiltrate the surrounding tissues and give rise to metastases.
- exemplary carcinomas which can be treated with a combination of antineoplastic thiol-binding mitochondrial oxidant and an anticancer agent include, for example, acinar carcinoma, acinous carcinoma, adenocystic carcinoma, adenoid cystic carcinoma, carcinoma adenomatosum, carcinoma of adrenal cortex, alveolar carcinoma, alveolar cell carcinoma, basal cell carcinoma, carcinoma basocellulare, basaloid carcinoma, basosquamous cell carcinoma, bronchioalveolar carcinoma, bronchiolar carcinoma, bronchogenic carcinoma, cerebriform carcinoma, cholangiocellular carcinoma, chorionic carcinoma, colloid carcinoma, comedo carcinoma, corpus carcinoma, cribriform carcinoma, carcinoma en cuirasse, carcinoma cutaneum, cylindrical carcinoma, cylindrical cell carcinoma, duct carcinoma, carcinoma durum, embryonal carcinoma, encephaloid carcinoma,
- neural crest cell refers to a temporary group of cells unique to vertebrates that arise from the embryonic ectoderm cell layer, and in turn give rise to a diverse cell lineage including, but not limited to, melanocytes, craniofacial cartilage and bone, smooth muscle, peripheral and enteric neurons, the adrenal medulla, and glia.
- neural crest-derived cancer refers to tumors derived from the neural crest.
- exemplary neural crest-derived cancers include small cell lung cancer, melanoma, breast cancer, medullary thyroid carcinoma, pheochromocytoma, and neuroblastoma.
- the neural crest-derived tumor is a small cell lung cancer, a melanoma, or a breast cancer.
- “pediatric cancer” or “childhood cancer” as used herein refers to any cancer occurring in a subject between birth and 19 years of age. In embodiments, “pediatric cancer” refers to cancers occurring in a subject between birth and 14 years of age. Exemplary pediatric cancers include acute lymphoblastic leukemia tumors, soft tissue sarcomas (e.g., rhabdomyosarcoma), rhabdoid tumors, neuroblastoma, kidney tumors, lymphoma, and brain and central nervous system tumors (e.g., astrocytoma, brain stem glioma, ependymoma, germ cell tumor, medulloblastoma). In aspects of the disclosure, the pediatric cancer is a rhabdoid tumor, a neuroblastoma, an acute lymphoblastic leukemia tumor, or a brain and central nervous system tumor.
- soft tissue sarcomas e.g., rhabdomyosarcoma
- Radioisotope therapy is used in accordance with its plain and ordinary meaning and refers to the use of ionizing radiation to kill or inhibit the growth or proliferation of cancer cells.
- Exemplary forms of radiation therapy include external beam radiation therapy, contact x-ray brachytherapy, sealed source radiotherapy, systemic radioisotope therapy, intraoperative radiotherapy, and deep inspiration breath-hold therapy.
- Systemic radioisotope therapy includes the delivery of radioisotopes (e.g., metaiodobenzylguanidine, iodine-131, lutetium-177, yttrium-90, strontium-89, samarium lexidronam) through infusion, injection, or ingestion.
- a radioisotope can be conjugated to an antibody (e.g., ZEVALIN®, BEXXAR®).
- “Differentiation therapy” refers to a cancer treatment in which compounds change the phenotype of cancer cells into normal cells.
- exemplary compounds include retinoids (e.g., 13-cis-retinoic acid, all-trans-retinoic acid).
- Tubulin binding drugs refers to compounds that target the mitotic spindle, and not the DNA, of cancer cells.
- Tubulin binding drugs include tubulin depolymerization inhibitors (e.g., paclitaxtel, epothilone, docetaxel, discodermolide) and tubulin polymerization inhibitors.
- Tubuling depolymerization inhibitors bind to the taxan site of tubulin.
- Tubulin polymerization inhibitors can bind to the vinca domain of tubulin.
- Such vinca domain-binding inhibitors include vinblastine, vincristine, vinoreIbine, vinfluine, dolastatins, halichondrins, hemiasterlins, cryptophysin 52.
- tubulin polymerization inhibitors can bind to the colchicine domain of tubulin.
- colchine domain-binding inhibitors include colchine, combrestatin, 2-methoxyestradiol, and methoxy benzenesulfonamide.
- Anti-cancer agent or “chemotherapeutic agent” is used in accordance with its plain and ordinary meaning and refers to a composition (e.g. compound, drug, antagonist, inhibitor, modulator) having antineoplastic properties or the ability to inhibit the growth or proliferation of cells.
- an anti-cancer agent is a chemotherapeutic.
- An anti-cancer agent may be an agent approved by the FDA or similar regulatory agency of a country other than the USA, for treating cancer. The methods described herein may further comprise administering a therapeutically effective amount of an anti-cancer or chemotherapeutic agent.
- anti-cancer agents include, but are not limited to, MEK (e.g. MEK1, MEK2, or MEK1 and MEK2) inhibitors (e.g. XL518, CI-1040, PD035901, selumetinib/AZD6244, GSK1120212/trametinib, GDC-0973, ARRY-162, ARRY-300, AZD8330, PD0325901, U0126, PD98059, TAK-733, PD318088, AS703026, BAY 869766), alkylating agents (e.g., cyclophosphamide, ifosfamide, chlorambucil, busulfan, melphalan, mechlorethamine, uramustine, thiotepa, nitrosoureas, nitrogen mustards (e.g., mechloroethamine, cyclophosphamide, chlorambucil, meiphalan), ethy
- TaxolTM i.e. paclitaxel
- TaxotereTM compounds comprising the taxane skeleton, Erbulozole (i.e. R-55104), Dolastatin 10 (i.e. DLS-10 and NSC-376128), Mivobulin isethionate (i.e. as CI-980), Vincristine, NSC-639829, Discodermolide (i.e. as NVP-XX-A-296), ABT-751 (Abbott, i.e. E-7010), Altorhyrtins (e.g. Altorhyrtin A and Altorhyrtin C), Spongistatins (e.g.
- Altorhyrtins e.g. Altorhyrtin A and Altorhyrtin C
- Spongistatins e.g.
- Epothilone E Epothilone F
- Epothilone B N-oxide Epothilone A N-oxide
- 16-aza-epothilone B Epothilone B
- 21-aminoepothilone B i.e. BMS-310705
- 21-hydroxyepothilone D i.e. Desoxyepothilone F and dEpoF
- 26-fluoroepothilone i.e. NSC-654663
- Soblidotin i.e. TZT-1027
- LS-4559-P Pulacia, i.e.
- LS-4577 LS-4578 (Pharmacia, i.e. LS-477-P), LS-4477 (Pharmacia), LS-4559 (Pharmacia), RPR-112378 (Aventis), Vincristine sulfate, DZ-3358 (Daiichi), FR-182877 (Fujisawa, i.e. WS-9885B), GS-164 (Takeda), GS-198 (Takeda), KAR-2 (Hungarian Academy of Sciences), BSF-223651 (BASF, i.e.
- ILX-651 and LU-223651 SAH-49960 (Lilly/Novartis), SDZ-268970 (Lilly/Novartis), AM-97 (Armad/Kyowa Hakko), AM-132 (Armad), AM-138 (Armad/Kyowa Hakko), IDN-5005 (Indena), Cryptophycin 52 (i.e. LY-355703), AC-7739 (Ajinomoto, i.e. AVE-8063A and CS-39.HCl), AC-7700 (Ajinomoto, i.e.
- T-900607 RPR-115781 (Aventis), Eleutherobins (such as Desmethyleleutherobin, Desaetyleleutherobin, lsoeleutherobin A, and Z-Eleutherobin), Caribaeoside, Caribaeolin, Halichondrin B, D-64131 (Asta Medica), D-68144 (Asta Medica), Diazonamide A, A-293620 (Abbott), NPI-2350 (Nereus), Taccalonolide A, TUB-245 (Aventis), A-259754 (Abbott), Diozostatin, ( ⁇ )-Phenylahistin (i.e.
- NSCL-96F03-7 D-68838 (Asta Medica), D-68836 (Asta Medica), Myoseverin B, D-43411 (Zentaris, i.e. D-81862), A-289099 (Abbott), A-318315 (Abbott), HTI-286 (i.e.
- SPA-110, trifluoroacetate salt) (Wyeth), D-82317 (Zentaris), D-82318 (Zentaris), SC-12983 (NCI), Resverastatin phosphate sodium, BPR-OY-007 (National Health Research Institutes), and SSR-250411 (Sanofi)), steroids (e.g., dexamethasone), finasteride, aromatase inhibitors, gonadotropin-releasing hormone agonists (GnRH) such as goserelin or leuprolide, adrenocorticosteroids (e.g., prednisone), progestins (e.g., hydroxyprogesterone caproate, megestrol acetate, medroxyprogesterone acetate), estrogens (e.g., diethlystiIbestrol, ethinyl estradiol), antiestrogen (e.g., tamoxifen), androgens (e
- gefitinib IressaTM
- erlotinib TarcevaTM
- cetuximab ErbituxTM
- lapatinib TykerbTM
- panitumumab VectibixTM
- vandetanib CaprelsaTM
- afatinib/BIBW2992 CI-1033/canertinib
- neratinib/HKI-272 CP-724714, TAK-285
- AST-1306 ARRY334543
- ARRY-380 AG-1478
- dacomitinib/PF299804 OSI-420/desmethyl erlotinib
- AZD8931 AEE788, pelitinib/EKB-569
- the PLK4 inhibitor is a compound of Formula (I) or a pharmaceutically acceptable salt thereof.
- the compound of Formula (I) is a compound of Formula (Ia), a compound of Formula (Ib), a compound of Formula (Ia1), a compound of Formula (Ia2), a compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), a compound of Formula (Ib
- the compound of Formula (Ia) is a compound of Formula (Ia1), a compound of Formula (Ia2), a compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), or a compound of Formula (Ia9f).
- the compound of Formula (Ib) is a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), or a compound of Formula (Ib7).
- the PLK4 inhibitor is a compound of Formula (Ia) or a pharmaceutically acceptable salt thereof:
- the PLK4 inhibitor is a compound of Formula (Ib) or a pharmaceutically acceptable salt thereof:
- L 1 is a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 13 —, —C(O)NR 13 —, —NR 13 C(O)—, —S(O) 2 —, —S(O)NR 13 —, substituted or unsubstituted alkylene, substituted or unsubstituted heteroalkylene, substituted or unsubstituted cycloalkylene, substituted or unsubstituted heterocycloalkylene, substituted or unsubstituted arylene, or substituted or unsubstituted heteroarylene.
- R 1 is hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 1A , —OR 1A , —NR 1A R 1B , —C(O)OR 1A , —C(O)NR 1A R 2B , —NO 2 , —SR 1A , —S(O) n1 R 1A , —S(O) n1 OR 1A , —S(O) n1 NR 1A R 1B , —NHNR 1A R 1B , —ONR 1A R 1B , —NHC(O)NHNR 1A R 1B , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted
- R 2 is hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 2A , —OR 2A , —NR 2A R 2B , —C(O)OR 2A , —C(O)NR 2A R 2B , —NO 2 , —SR 2A , —S(O) n2 R 2A , —S(O) n2 OR 2A , —S(O) n2 NR 2A R 2B , —NHNR 2A R 2B , —ONR 2A R 2B , —NHC(O)NHNR 2A R 2B , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted
- R 3 is hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3A , —OR 3A , —NR 3A R 3B , —C(O)OR 3A , —C(O)NR 3A R 3B , —NO 2 , —SR 3A , —S(O) n3 R 3A , —S(O) n3 OR 3A , —S(O) n3 NR 3A R 3B , —NHNR 3A R 3B , —ONR 3A R 3B , —NHC(O)NHNR 3A R 3B , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted
- R 4 is hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 4A , —OR 4A , —NR 4A R 4B , —C(O)OR 4A , —C(O)NR 4A R 4B , —NO 2 , —SR 4A , —S(O) n4 R 4A , —S(O) n4 OR 4A , —S(O) n4 NR 4A R 4B , —N—ONR 4A R 4B , ONR 4A R 4B , —NHC(O)NHNR 4A R 4B , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted
- R 3C -substituted or unsubstituted cycloalkyl substituted or unsubstituted heterocycloalkyl (e.g. R 3C -substituted or unsubstituted heterocycloalkyl), substituted or unsubstituted aryl (e.g. R 3C -substituted or unsubstituted aryl), or substituted or unsubstituted heteroaryl (e.g. R 3C -substituted or unsubstituted heteroaryl).
- substituted or unsubstituted heterocycloalkyl e.g. R 3C -substituted or unsubstituted heterocycloalkyl
- substituted or unsubstituted aryl e.g. R 3C -substituted or unsubstituted aryl
- substituted or unsubstituted heteroaryl e.g. R 3C -substi
- R 5 is independently hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 5A , —OR 5A , —NR 5A R 5B , —C(O)OR 5A , —C(O)NR 5A R 5B , —NO 2 , —SR 5A , —S(O) n5 R 5A , —S(O) n5 OR 5A , —S(O) n5 NR 5A R 5B , —NHNR 5A R 5B , —ONR 5A R 5B , —NHC(O)NHNR 5A R 5B , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substitute
- R 6 is hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —CHO, —OR 6A , —NR 6A R 6B , —C(O)OR 6A , —C(O)NR 6A R 6B , —NO 2 , —SR 6A , —S(O) n6 R 6A , —S(O) n6 OR 6A , —S(O) n6 NR 6A R 6B , —NHNR 6A R 6B , —ONR 6A R 6B , —NHC(O)NHNR 6A R 6B , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substitute
- R 1A , R 1B , R 2A , R 2B , R 3A , R 3B , R 4A , R 4B , R 5A , R 5B , R 6A , R 6B , and R 13 are independently hydrogen, oxo, halogen, —CF 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(NHC(NHC(O)
- the compound of Formula (Ib) is not tozasertib.
- R 4 is not a para-methyl piperidinyl.
- R 4 is not a para-ethyl piperidinyl.
- R 4 is not a para-propyl piperidinyl.
- R 4 is not a piperidinyl substituted with a methyl.
- R 4 is not a piperidinyl substituted with a ethyl.
- R 4 is not a piperidinyl substituted with a propyl.
- R 4 is not a piperidinyl substituted with an unsubstituted alkyl at the para position. In embodiments of Formula (Ib), R 4 is not a piperidinyl substituted with an unsubstituted alkyl. In embodiments of Formula (Ib), R 4 is not a piperidinyl substituted with a substituted or unsubstituted alkyl. In embodiments of Formula (Ib), R 4 is not a substituted or unsubstituted piperidinyl. In embodiments of Formula (Ib), R 4 is not a substituted heterocycloalkyl.
- R 4 is not a substituted or unsubstituted heterocycloalkyl. In embodiments of Formula (Ia), R 4 is not a para-methyl piperidinyl. In embodiments of Formula (Ib), R 4 is not a para-ethyl piperidinyl. In embodiments of Formula (Ib), R 4 is not a para-propyl piperidinyl. In embodiments of Formula (Ib) Formula (Ib), R 4 is not a piperidinyl substituted with a methyl. In embodiments of Formula (Ib), R 4 is not a piperidinyl substituted with a ethyl.
- R 4 is not a piperidinyl substituted with a propyl. In embodiments of Formula (Ia), R 4 is not a piperidinyl substituted with an unsubstituted alkyl at the para position. In embodiments of Formula (Ia), R 4 is not a piperidinyl substituted with an unsubstituted alkyl. In embodiments of Formula (Ia), R 4 is not a piperidinyl substituted with a substituted or unsubstituted alkyl. In embodiments of Formula (Ia), R 4 is not a substituted or unsubstituted piperidinyl.
- R 4 is not a substituted heterocycloalkyl. In embodiments of Formula (Ia), R 4 is not a substituted or unsubstituted heterocycloalkyl.
- the provisos set forth in the embodiments of the paragraph apply only when -L 1 -R 6 is cyclopropyl. The embodiments in this paragraph are equally applicable to all other appropriate formulae set forth herein.
- -L 1 -R 6 is not cyclopropyl. In embodiments of Formula (Ia), -L 1 -R 6 is not cyclopropyl. In embodiments of Formula (Ib), -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl. In embodiments of Formula (Ib), -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl substituted with substituted or unsubstituted alkyl. In embodiments of Formula (Ib), -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl substituted with unsubstituted alkyl.
- -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl substituted with methyl, ethyl or propyl. In embodiments of Formula (Ib), -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl substituted with para methyl, para ethyl or para propyl. In embodiments of Formula (Ia), -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl. In embodiments of Formula (Ia), -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl substituted with substituted or unsubstituted alkyl.
- -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl substituted with unsubstituted alkyl. In embodiments of Formula (Ia), -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl substituted with methyl, ethyl or propyl. In embodiments of Formula (Ia), -L 1 -R 6 is not cyclopropyl when R 4 is piperidinyl substituted with para methyl, para ethyl or para propyl. The embodiments in this paragraph are equally applicable to all other appropriate formulae set forth herein.
- R 1 is hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 1A , —OR 1A , —NR 1A R 1B , —C(O)OR 1A , or —C(O)NR 1A R 1B .
- R 1 is hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 1A , —OR 1A , —NR 1A R 1B , —C(O)OR 1A , or —C(O)NR 1A R 1B where R 1A and R 1B are independently hydrogen, oxo, halogen, —CF 3 , —OH, —NH 2 , —COOH, —CONH 2 , substituted or unsubstituted alkyl, or substituted or unsubstituted aryl.
- R 1 is hydrogen.
- R 1 is hydrogen or substituted or unsubstituted alkyl.
- R 1 may be substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 1 may be substituted or unsubstituted alkyl.
- R 1 may be substituted alkyl.
- R 1 may be unsubstituted alkyl.
- R 1 may be substituted or unsubstituted C 1 -C 20 alkyl.
- R 1 may be substituted C 1 -C 20 alkyl.
- R 1 may be substituted or unsubstituted C 1 -C 10 alkyl.
- R 1 may be substituted C 1 -C 10 alkyl.
- R 1 may be unsubstituted C 1 -C 10 alkyl.
- R 1 may be substituted or unsubstituted C 1 -C 5 alkyl.
- R 1 may be unsubstituted C 1 -C 5 alkyl.
- R 1 may be substituted C 1 -C 5 alkyl.
- R 1 may be methyl, substituted or unsubstituted ethyl, or substituted or unsubstituted propyl.
- R 1 may be methyl.
- R 1 may be ethyl.
- R 1 may be R 1C -substituted or unsubstituted alkyl.
- R 1 may be R 1C -substituted alkyl.
- R 1 may be R 1C -substituted or unsubstituted C 1 -C 20 alkyl.
- R 1 may be R 1C -substituted C 1 -C 20 alkyl.
- R 1 may be R 1C -substituted or unsubstituted C 1 -C 10 alkyl.
- R 1 may be R 1C -substituted C 1 -C 10 alkyl.
- R 1 may be R 1C -substituted or unsubstituted C 1 -C 5 alkyl.
- R 1 may be R 1C -substituted C 1 -C 5 alkyl.
- R 1 may be methyl, R 1C -substituted or unsubstituted ethyl, or R 1C -substituted or unsubstituted propyl.
- R 1 may be substituted or unsubstituted heteroalkyl.
- R 1 may be substituted heteroalkyl.
- R 1 may be unsubstituted heteroalkyl.
- R 1 may be substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 1 may be substituted 2 to 20 membered heteroalkyl.
- R 1 may be substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 1 may be substituted 2 to 10 membered heteroalkyl.
- R 1 may be substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 1 may be substituted 2 to 6 membered heteroalkyl.
- R 1 may be R 1C -substituted or unsubstituted heteroalkyl.
- R 1 may be R 1C -substituted heteroalkyl.
- R 1 may be R 1C -substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 1 may be R 1C -substituted 2 to 20 membered heteroalkyl.
- R 1 may be R 1C -substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 1 may be R 1C -substituted 2 to 10 membered heteroalkyl.
- R 1 may be R 1C -substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 1 may be R 1C -substituted 2 to 6 membered heteroalkyl.
- R 1 may be substituted or unsubstituted cycloalkyl.
- R 1 may be substituted cycloalkyl.
- R 1 may be unsubstituted cycloalkyl.
- R 1 may be substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 1 may be substituted 3 to 20 membered cycloalkyl.
- R 1 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 1 may be substituted 3 to 10 membered cycloalkyl.
- R 1 may be substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 1 may be substituted 3 to 6 membered cycloalkyl.
- R 1 may be R 1C -substituted or unsubstituted cycloalkyl.
- R 1 may be R 1C -substituted cycloalkyl.
- R 1 may be R 1C -substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 1 may be R 1C -substituted 3 to 20 membered cycloalkyl.
- R 1 may be R 1C -substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 1 may be R 1C -substituted 3 to 10 membered cycloalkyl.
- R 1 may be R 1C -substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 1 may be R 1C -substituted 3 to 6 membered cycloalkyl.
- R 1 may be substituted or unsubstituted heterocycloalkyl.
- R 1 may be substituted heterocycloalkyl.
- R 1 may be unsubstituted heterocycloalkyl.
- R 1 may be substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 1 may be substituted 3 to 20 membered heterocycloalkyl.
- R 1 may be substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 1 may be substituted 3 to 10 membered heterocycloalkyl.
- R 1 may be substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 1 may be substituted 3 to 6 membered heterocycloalkyl.
- R 1 may be R 1C -substituted or unsubstituted heterocycloalkyl.
- R 1 may be R 1C -substituted heterocycloalkyl.
- R 1 may be R 1C -substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 1 may be R 1C -substituted 3 to 20 membered heterocycloalkyl.
- R 1 may be R 1C -substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 1 may be R 1C -substituted 3 to 10 membered heterocycloalkyl.
- R 1 may be R 1C -substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 1 may be R 1C -substituted 3 to 6 membered heterocycloalkyl.
- R 1 may be substituted or unsubstituted aryl.
- R 1 may be substituted aryl.
- R 1 may be unsubstituted aryl.
- R 1 may be substituted or unsubstituted 5 to 20 membered aryl.
- R 1 may be substituted 5 to 20 membered aryl.
- R 1 may be substituted or unsubstituted 5 to 8 membered aryl (e.g. phenyl).
- R 1 may be substituted 5 to 8 membered aryl.
- R 1 may be substituted or unsubstituted 5 or 6 membered aryl.
- R 1 may be substituted 5 or 6 membered aryl.
- R 1 may be R 1C -substituted or unsubstituted aryl.
- R 1 may be R 1C -substituted aryl.
- R 1 may be R 1C -substituted or unsubstituted 5 to 20 membered aryl.
- R 1 may be R 1C -substituted 5 to 20 membered aryl.
- R 1 may be R 1C -substituted or unsubstituted 5 to 8 membered aryl.
- R 1 may be R 1C -substituted 5 to 8 membered aryl.
- R 1 may be R 1C -substituted or unsubstituted 5 or 6 membered aryl.
- R 1 may be R 1C -substituted 5 or 6 membered aryl (e.g. phenyl).
- R 1 may be substituted or unsubstituted heteroaryl.
- R 1 may be substituted heteroaryl.
- R 1 may be unsubstituted heteroaryl.
- R 1 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 1 may be substituted 5 to 20 membered heteroaryl.
- R 1 may be substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 1 may be substituted 5 to 8 membered heteroaryl.
- R 1 may be substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 1 may be substituted 5 or 6 membered heteroaryl.
- R 1 may be R 1C -substituted or unsubstituted heteroaryl.
- R 1 may be R 1C -substituted heteroaryl.
- R 1 may be R 1C -substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 1 may be R 1C -substituted 5 to 20 membered heteroaryl.
- R 1 may be R 1C -substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 1 may be R 1C -substituted 5 to 8 membered heteroaryl.
- R 1 may be R 1C -substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 1 may be R 1C -substituted 5 or 6 membered heteroaryl.
- R 1A may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 1D , —COR 1D , —NR 1D R 1E , —COOR 1D , —CONR 1D R 1E , —NO 2 , —SR 1D , —S(O) 2 R 1D , —S(O) 3 R 1D , —S(O) 4 R 1D , —S(O) 2 NR 1D R 1E , —NHNR 1D R 1E , —ONR 1D R 1E , —NHC(O)NHNR 1D R 1E , —NHC(O)NR 1D R 1E , —NHS(O) 2 R 1D , —NHC(O)R 1D , —NHC(O)—OR 1D , —
- R 1C is independently halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COR 1D , —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted hetero
- R 1C may independently be halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted heterocyclo
- R 1B may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted heterocycloalkyl
- R 1D and R 1E are independently hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted hetero
- R 2 may be hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 2A , —OR 2A , —NR 2A R 2B , —C(O)OR 2A , or —C(O)NR 2A R 2B .
- R 2 may be hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 2A , —OR 2A , —NR 2A R 2B , —C(O)OR 2A , or —C(O)NR 2A R 2B where R 2A and R 2B are independently hydrogen, oxo, halogen, —CF 3 , —OH, —NH 2 , —COOH, —CONH 2 , substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 2 may be substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 2 may be R 2A -substituted or unsubstituted cycloalkyl, R 2A -substituted or unsubstituted heterocycloalkyl, R 2A -substituted or unsubstituted aryl, or R 2A -substituted or unsubstituted heteroaryl.
- R 2 may be substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 2 may be R 2A -substituted or unsubstituted heterocycloalkyl, R 2A -substituted or unsubstituted aryl, or R 2A -substituted or unsubstituted heteroaryl.
- R 2 may be R 2C -substituted or unsubstituted alkyl, R 2C -substituted or unsubstituted heteroalkyl, R 2C -substituted or unsubstituted cycloalkyl, R 2C -substituted or unsubstituted heterocycloalkyl, R 2C -substituted or unsubstituted aryl, or R 2C -substituted or unsubstituted heteroaryl.
- R 2 is substituted with an alkyl, heteroalkyl, cycloalkyl, heterocycloalkyl, aryl or heteroaryl, wherein the alkyl, heteroalkyl, cycloalkyl, heterocycloalkyl, aryl or heteroaryl is substituted with a substituent selected from —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —C(O)OH, —C(O)NH 2 , —NO 2 , —SH, substituted or unsubstituted alkyl, and substituted or unsubstituted heteroalkyl.
- R 2 may be substituted or unsubstituted alkyl.
- R 2 may be substituted alkyl.
- R 2 may be unsubstituted alkyl.
- R 2 may be substituted or unsubstituted C 1 -C 20 alkyl.
- R 2 may be substituted C 1 -C 20 alkyl.
- R 2 may be substituted or unsubstituted C 1 -C 10 alkyl.
- R 2 may be substituted C 1 -C 10 alkyl.
- R 2 may be unsubstituted C 1 -C 10 alkyl.
- R 2 may be substituted or unsubstituted C 1 -C 5 alkyl.
- R 2 may be unsubstituted C 1 -C 5 alkyl.
- R 2 may be substituted C 1 -C 5 alkyl.
- R 2 may be methyl, substituted or unsubstituted ethyl, or substituted or unsubstituted propyl.
- R 2 may be methyl.
- R 2 may be ethyl.
- R 2 may be R 2C -substituted or unsubstituted alkyl.
- R 2 may be R 2 -substituted alkyl.
- R 2 may be R 2C -substituted or unsubstituted C 1 -C 20 alkyl.
- R 2 may be R 2C -substituted C 1 -C 20 alkyl.
- R 2 may be R 2C -substituted or unsubstituted C 1 -C 10 alkyl.
- R 2 may be R 2C -substituted C 1 -C 10 alkyl.
- R 2 may be R 2C -substituted or unsubstituted C 1 -C 5 alkyl.
- R 2 may be R 2 -substituted C 1 -C 5 alkyl.
- R 2 may be methyl, R 2C -substituted or unsubstituted ethyl, or R 2C -substituted or unsubstituted propyl.
- R 2 may be substituted or unsubstituted heteroalkyl.
- R 2 may be substituted heteroalkyl.
- R 2 may be unsubstituted heteroalkyl.
- R 2 may be substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 2 may be substituted 2 to 20 membered heteroalkyl.
- R 2 may be unsubstituted 2 to 20 membered heteroalkyl.
- R 2 may be substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 2 may be substituted 2 to 10 membered heteroalkyl.
- R 2 may be unsubstituted 2 to 10 membered heteroalkyl.
- R 2 may be substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 2 may be substituted 2 to 6 membered heteroalkyl.
- R 2 may be unsubstituted 2 to 6 membered heteroalkyl.
- R 2 may be R 2C -substituted or unsubstituted heteroalkyl.
- R 2 may be R 2C -substituted heteroalkyl.
- R 2 may be R 2C -substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 2 may be R 2C -substituted 2 to 20 membered heteroalkyl.
- R 2 may be R 2C -substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 2 may be R 2C -substituted 2 to 10 membered heteroalkyl.
- R 2 may be R 2C -substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 2 may be R 2C -substituted 2 to 6 membered heteroalkyl.
- R 2 may be substituted or unsubstituted cycloalkyl.
- R 2 may be substituted cycloalkyl.
- R 2 may be unsubstituted cycloalkyl.
- R 2 may be substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 2 may be substituted 3 to 20 membered cycloalkyl.
- R 2 may be unsubstituted 3 to 20 membered cycloalkyl.
- R 2 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 2 may be substituted 3 to 10 membered cycloalkyl.
- R 2 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 2 may be unsubstituted 3 to 6 membered cycloalkyl.
- R 2 may be substituted 3 to 6 membered cycloalkyl.
- R 2 may be R 2C -substituted or unsubstituted cycloalkyl.
- R 2 may be R 2C -substituted cycloalkyl.
- R 2 may be R 2C -substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 2 may be R 2C -substituted 3 to 20 membered cycloalkyl.
- R 2 may be R 2C -substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 2 may be R 2C -substituted 3 to 10 membered cycloalkyl.
- R 2 may be R 2C -substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 2 may be R 2C -substituted 3 to 6 membered cycloalkyl.
- R 2 may be substituted or unsubstituted heterocycloalkyl.
- R 2 may be substituted heterocycloalkyl.
- R 2 may be unsubstituted heterocycloalkyl.
- R 2 may be substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 2 may be substituted 3 to 20 membered heterocycloalkyl.
- R 2 may be unsubstituted 3 to 20 membered heterocycloalkyl.
- R 2 may be substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 2 may be substituted 3 to 10 membered heterocycloalkyl.
- R 2 may be unsubstituted 3 to 10 membered heterocycloalkyl.
- R 2 may be substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 2 may be substituted 3 to 6 membered heterocycloalkyl.
- R 2 may be unsubstituted 3 to 6 membered heterocycloalkyl.
- R 2 may be R 2C -substituted or unsubstituted heterocycloalkyl.
- R 2 may be R 2C -substituted heterocycloalkyl.
- R 2 may be R 2C -substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 2 may be R 2C -substituted 3 to 20 membered heterocycloalkyl.
- R 2 may be R 2C -substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 2 may be R 2C -substituted 3 to 10 membered heterocycloalkyl.
- R 2 may be R 2C -substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 2 may be R 2C -substituted 3 to 6 membered heterocycloalkyl.
- R 2 may be substituted or unsubstituted aryl.
- R 2 may be substituted aryl.
- R 2 may be unsubstituted aryl.
- R 2 may be substituted or unsubstituted 5 to 20 membered aryl.
- R 2 may be substituted 5 to 20 membered aryl.
- R 2 may be unsubstituted 5 to 20 membered aryl.
- R 2 may be substituted or unsubstituted 5 to 8 membered aryl (e.g. phenyl).
- R 2 may be substituted 5 to 8 membered aryl.
- R 2 may be substituted or unsubstituted 5 or 6 membered aryl.
- R 2 may be unsubstituted 5 to 8 membered aryl (e.g. phenyl). R 2 may be substituted 5 or 6 membered aryl. R 2 may be unsubstituted 5 or 6 membered aryl.
- R 2 may be R 2C -substituted or unsubstituted aryl.
- R 2 may be R 2C -substituted aryl.
- R 2 may be R 2C -substituted or unsubstituted 5 to 20 membered aryl.
- R 2 may be R 2C -substituted 5 to 20 membered aryl.
- R 2 may be R 2C -substituted or unsubstituted 5 to 8 membered aryl.
- R 2 may be R 2C -substituted 5 to 8 membered aryl.
- R 2 may be R 2C -substituted or unsubstituted 5 or 6 membered aryl.
- R 2 may be R 2C -substituted 5 or 6 membered aryl (e.g. phenyl).
- R 2 may be substituted or unsubstituted heteroaryl.
- R 2 may be substituted heteroaryl.
- R 2 may be unsubstituted heteroaryl.
- R 2 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 2 may be substituted 5 to 20 membered heteroaryl.
- R 2 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 2 may be substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 2 may be substituted 5 to 8 membered heteroaryl.
- R 2 may be unsubstituted 5 to 8 membered heteroaryl.
- R 2 may be substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 2 may be substituted 5 or 6 membered heteroaryl.
- R 2 may be unsubstituted 5 or 6 membered heteroaryl.
- R 2 may be unsubstitute
- R 2 may be R 2C -substituted or unsubstituted heteroaryl.
- R 2 may be R 2 -substituted heteroaryl.
- R 2 may be R 2C -substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 2 may be R 2C -substituted 5 to 20 membered heteroaryl.
- R 2 may be R 2C -substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 2 may be R 2C -substituted 5 to 8 membered heteroaryl.
- R 2 may be R 2C -substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 2 may be R 2C -substituted 5 or 6 membered heteroaryl.
- R 2A may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COR 2C , —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 2C -substituted or unsubstituted alkyl, R 2C -substituted or unsubstituted
- R 2C is independently hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 2D , —COR 2D , —NR 2D R 2E , —COOR 2D , —CONR 2D R 2E , —NO 2 , —SH, —S(O) 2 R 2D , —S(O) 3 R 2D , —S(O) 4 R 2D , —S(O) 2 NR 2D R 2E , —NHNR 2D R 2E , —ONR 2D R 2E , —NHC(O)NHNR 2D R 2E , —NHC(O)NR 2D R 2E , —NHC(O)NR 2D R 2E , —NHS(O) 2 R 2D , —NHC(O)R 2D , —NHC
- R 2C may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COR 2D , —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 2D -substituted or unsubstituted alkyl, R 2D -substituted or unsubstituted
- R 2C may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 2D -substituted or unsubstituted alkyl, R 2D -substituted or unsubstituted heteroalky
- R 2B may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted heterocycloalkyl
- R 2D and R 2E are independently hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted hetero
- R 2 may be R 2A -substituted or unsubstituted heteroaryl where R 2A is —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —C(O)OH, —C(O)NH 2 , —NO 2 , —SH, substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl.
- R 2 may be R 2A -substituted or unsubstituted heteroaryl where R 2A is —CF 3 , —CN, —OH, —NH 2 , —C(O)OH, —C(O)NH 2 , —NO 2 , or substituted or unsubstituted alkyl.
- R 2A may be substituted or unsubstituted C 1 -C 5 alkyl.
- R 2A may be methyl.
- R 2 may be substituted or unsubstituted furanyl, substituted or unsubstituted pyrroyl, substituted or unsubstituted thiophenyl, substituted or unsubstituted imidazoyl, substituted or unsubstituted pyrazolyl, substituted or unsubstituted oxazoyl, substituted or unsubstituted isoxazolyl, substituted or unsubstituted thiazolyl, substituted or unsubstituted pyridinyl, substituted or unsubstituted pyrazinyl, substituted or unsubstituted pyrimidinyl, or substituted or unsubstituted pyridazinyl.
- R 2 may be substituted or unsubstituted furanyl.
- R 2 may be substituted or unsubstituted pyrroyl.
- R 2 may be substituted or unsubstituted thiophenyl.
- R 2 may be substituted or unsubstituted imidazoyl.
- R 2 may be substituted or unsubstituted pyrazolyl.
- R 2 may be substituted or unsubstituted oxazoyl.
- R 2 may be substituted or unsubstituted isoxazolyl.
- R 2 may be substituted or unsubstituted thiazolyl.
- R 2 may be substituted or unsubstituted pyridinyl.
- R 2 may be substituted or unsubstituted pyrazinyl.
- R 2 may be substituted or unsubstituted pyrimidinyl.
- R 2 may be substituted or unsubstituted pyridaziny
- R 2 may be R 2A -substituted or unsubstituted furanyl, substituted or unsubstituted pyrroyl, substituted or unsubstituted thiophenyl, substituted or unsubstituted imidazoyl, substituted or unsubstituted pyrazolyl, substituted or unsubstituted oxazoyl, substituted or unsubstituted isoxazolyl, substituted or unsubstituted thiazolyl, substituted or unsubstituted pyridinyl, substituted or unsubstituted pyrazinyl, substituted or unsubstituted pyrimidinyl, or substituted or unsubstituted pyridazinyl.
- R 2 may be R 2A -substituted or unsubstituted furanyl.
- R 2 may be R 2A -substituted or unsubstituted pyrrolyl.
- R 2 may be R 2A -substituted or unsubstituted thiophenyl.
- R 2 may be R 2A -substituted or unsubstituted imidazoyl.
- R 2 may be R 2A -substituted or unsubstituted pyrazolyl.
- R 2 may be R 2A -substituted or unsubstituted pyrazolyl where R 2A is —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —C(O)OH, —C(O)NH 2 , —NO 2 , —SH, substituted or unsubstituted alkyl, or substituted or unsubstituted heteroalkyl.
- R 2 may be R 2A -substituted or unsubstituted oxazoyl.
- R 2 may be R 2A -substituted or unsubstituted isoxazolyl.
- R 2 may be R 2A -substituted or unsubstituted thiazolyl.
- R 2 may be R 2A -substituted or unsubstituted pyridinyl.
- R 2 may be R 2A -substituted or unsubstituted pyrazinyl.
- R 2 may be R 2A -substituted or unsubstituted pyrimidinyl.
- R 2 may be R 2A -substituted or unsubstituted pyridazinyl.
- R 2 may be 5-methyl-1H-pyrazolyl.
- R 3 may be hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3A , —OR 3A , —NR 3A R 3B , —C(O)OR 3A , —C(O)NR 3A R 3B , —NO 2 , —SR 3A , —S(O) n3 R 3A , —S(O) n3 OR 3A , —S(O) n3 NR 3A R 3B , —NHNR 3A R 3B , —ONR 3A R 3B , —NHC(O)NHNR 3A R 3B , substituted or unsubstituted alkyl, or optionally combined with R 4 to form a substituted or unsubstituted cycloalkyl where R 3A and R 3B are independently hydrogen, oxo, halogen,
- R 3 may be hydrogen, halogen, —OR 3A , or substituted or unsubstituted alkyl, where R 3A is as defined herein.
- R 3 may be hydrogen, halogen, —OR 3A , or R 3C -substituted or unsubstituted alkyl, where R 3A and R 3C are as defined herein.
- R 3 may be hydrogen, halogen, —OR 3A , or substituted or unsubstituted alkyl where R 3A is substituted or unsubstituted alkyl.
- R 3 may be hydrogen, halogen, —OR 3A , or R 3C -substituted or unsubstituted alkyl where R 3A is substituted or unsubstituted alkyl and R 3C is as defined herein.
- R 3 may be hydrogen, halogen, —OR 3A , or substituted or unsubstituted alkyl where R 3A is R 3C -substituted or unsubstituted alkyl.
- R 3 may be hydrogen, halogen, —OR 3A , or R 3C -substituted or unsubstituted alkyl where R 3A is R 3C -substituted or unsubstituted alkyl and R 3C is as defined herein.
- R 3 may be —Cl, —I, or —Br.
- R 3 may be —Cl.
- R 3 may be —Br.
- R 3 may be —I.
- R 3 may be —F.
- R 3 may be —OR 3A , where R 3A is as defined herein.
- R 3 may be —OR 3A where R 3A is substituted or unsubstituted alkyl.
- R 3 may be —OR 3A where R 3A is substituted or unsubstituted C 1 -C 5 alkyl.
- R 3 may be —OR 3A where R 3A is R 3C -substituted or unsubstituted C 1 -C 5 alkyl.
- R 3 may be —OCH 3 .
- R 3 may be —OCH 2 CH 3 .
- R 3 may be —OR 3A where R 3A is substituted or unsubstituted C 5 -C 6 aryl.
- R 3 may be substituted or unsubstituted alkyl.
- R 3 may be substituted alkyl.
- R 3 may be unsubstituted alkyl.
- R 3 may be substituted or unsubstituted C 1 -C 20 alkyl.
- R 3 may be substituted C 1 -C 20 alkyl.
- R 3 may be substituted or unsubstituted C 1 -C 10 alkyl.
- R 3 may be substituted C 1 -C 10 alkyl.
- R 3 may be unsubstituted C 1 -C 10 alkyl.
- R 3 may be substituted or unsubstituted C 1 -C 5 alkyl.
- R 3 may be unsubstituted C 1 -C 5 alkyl.
- R 3 may be substituted C 1 -C 5 alkyl.
- R 3 may be methyl, substituted or unsubstituted ethyl, or substituted or unsubstituted propyl.
- R 3 may be methyl.
- R 3 may be ethyl.
- R 3 may be R 3C -substituted or unsubstituted alkyl.
- R 3 may be R 3 -substituted alkyl.
- R 3 may be R 3C -substituted or unsubstituted C 1 -C 20 alkyl.
- R 3 may be R 3 -substituted C 1 -C 20 alkyl.
- R 3 may be R 3C -substituted or unsubstituted C 1 -C 10 alkyl.
- R 3 may be R 3 -substituted C 1 -C 10 alkyl.
- R 3 may be R 3C -substituted or unsubstituted C 1 -C 5 alkyl.
- R 3 may be R 3 -substituted C 1 -C 5 alkyl.
- R 3 may be methyl, R 3C -substituted or unsubstituted ethyl, or R 3C -substituted or unsubstituted propyl.
- R 3 may be substituted or unsubstituted heteroalkyl.
- R 3 may be substituted heteroalkyl.
- R 3 may be unsubstituted heteroalkyl.
- R 3 may be substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 3 may be substituted 2 to 20 membered heteroalkyl.
- R 3 may be substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 3 may be substituted 2 to 10 membered heteroalkyl.
- R 3 may be unsubstituted 2 to 10 membered heteroalkyl.
- R 3 may be substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 3 may be substituted 2 to 6 membered heteroalkyl.
- R 3 may be unsubstituted 2 to 6 membered heteroalkyl.
- R 3 may be R 3C -substituted or unsubstituted heteroalkyl.
- R 3 may be R 3 -substituted heteroalkyl.
- R 3 may be R 3C -substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 3 may be R 3C -substituted 2 to 20 membered heteroalkyl.
- R 3 may be R 3C -substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 3 may be R 3C -substituted 2 to 10 membered heteroalkyl.
- R 3 may be R 3C -substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 3 may be R 3C -substituted 2 to 6 membered heteroalkyl.
- R 3 may be substituted or unsubstituted cycloalkyl.
- R 3 may be substituted cycloalkyl.
- R 3 may be unsubstituted cycloalkyl.
- R 3 may be substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 3 may be substituted 3 to 20 membered cycloalkyl.
- R 3 may be unsubstituted 3 to 20 membered cycloalkyl.
- R 3 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 3 may be substituted 3 to 10 membered cycloalkyl.
- R 3 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 3 may be unsubstituted 3 to 6 membered cycloalkyl.
- R 3 may be substituted 3 to 6 membered cycloalkyl.
- R 3 may be R 3C -substituted or unsubstituted cycloalkyl.
- R 3 may be R 3 -substituted cycloalkyl.
- R 3 may be R 3C -substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 3 may be R 3C -substituted 3 to 20 membered cycloalkyl.
- R 3 may be R 3C -substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 3 may be R 3C -substituted 3 to 10 membered cycloalkyl.
- R 3 may be R 3C -substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 3 may be R 3C -substituted 3 to 6 membered cycloalkyl.
- R 3 may be substituted or unsubstituted heterocycloalkyl.
- R 3 may be substituted heterocycloalkyl.
- R 3 may be unsubstituted heterocycloalkyl.
- R 3 may be substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 3 may be substituted 3 to 20 membered heterocycloalkyl.
- R 3 may be unsubstituted 3 to 20 membered heterocycloalkyl.
- R 3 may be substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 3 may be substituted 3 to 10 membered heterocycloalkyl.
- R 3 may be unsubstituted 3 to 10 membered heterocycloalkyl.
- R 3 may be substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 3 may be substituted 3 to 6 membered heterocycloalkyl.
- R 3 may be unsubstituted 3 to 6 membered heterocycloalkyl.
- R 3 may be R 3C -substituted or unsubstituted heterocycloalkyl.
- R 3 may be R 3C -substituted heterocycloalkyl.
- R 3 may be R 3C -substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 3 may be R 3C -substituted 3 to 20 membered heterocycloalkyl.
- R 3 may be R 3C -substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 3 may be R 3C -substituted 3 to 10 membered heterocycloalkyl.
- R 3 may be R 3C -substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 3 may be R 3C -substituted 3 to 6 membered heterocycloalkyl.
- R 3 may be substituted or unsubstituted aryl.
- R 3 may be substituted aryl.
- R 3 may be unsubstituted aryl.
- R 3 may be substituted or unsubstituted 5 to 20 membered aryl.
- R 3 may be substituted 5 to 20 membered aryl.
- R 3 may be unsubstituted 5 to 20 membered aryl.
- R 3 may be substituted or unsubstituted 5 to 8 membered aryl (e.g. phenyl).
- R 3 may be substituted 5 to 8 membered aryl.
- R 3 may be substituted or unsubstituted 5 or 6 membered aryl.
- R 3 may be unsubstituted 5 to 8 membered aryl (e.g. phenyl). R 3 may be substituted 5 or 6 membered aryl. R 3 may be unsubstituted 5 or 6 membered aryl.
- R 3 may be R 3C -substituted or unsubstituted aryl.
- R 3 may be R 3 -substituted aryl.
- R 3 may be R 3C -substituted or unsubstituted 5 to 20 membered aryl.
- R 3 may be R 3C -substituted 5 to 20 membered aryl.
- R 3 may be R 3C -substituted or unsubstituted 5 to 8 membered aryl.
- R 3 may be R 3C -substituted 5 to 8 membered aryl.
- R 3 may be R 3C -substituted or unsubstituted 5 or 6 membered aryl.
- R 3 may be R 3C -substituted 5 or 6 membered aryl (e.g. phenyl).
- R 3 may be substituted or unsubstituted heteroaryl.
- R 3 may be substituted heteroaryl.
- R 3 may be unsubstituted heteroaryl.
- R 3 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 3 may be substituted 5 to 20 membered heteroaryl.
- R 3 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 3 may be substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 3 may be substituted 5 to 8 membered heteroaryl.
- R 3 may be unsubstituted 5 to 8 membered heteroaryl.
- R 3 may be substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 3 may be substituted 5 or 6 membered heteroaryl.
- R 3 may be unsubstituted 5 or 6 membered heteroaryl.
- R 3 may be unsubstitute
- R 3 may be R 3C -substituted or unsubstituted heteroaryl.
- R 3 may be R 3 -substituted heteroaryl.
- R 3 may be R 3C -substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 3 may be R 3C -substituted 5 to 20 membered heteroaryl.
- R 3 may be R 3C -substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 3 may be R 3C -substituted 5 to 8 membered heteroaryl.
- R 3 may be R 3C -substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 3 may be R 3C -substituted 5 or 6 membered heteroaryl.
- R 3A may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3C , —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 3C -substituted or unsubstituted alkyl, R 3C -substituted or unsubstituted
- R 3C is independently hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3D , —OR 3D , —NR 3D R 3E , —COOR 3D , —CONR 3D R 3E , —NO 2 , —SR 3D , —S(O) 2 R 3D , —S(O) 3 R 3D , —S(O) 4 R 3D , —S(O) 2 NR 3D R 3E , —NHNR 3D R 3E , —ONR 3D R 3E , —NHC(O)NHNR 3D R 3E , —NHC(O)NR 3D R 3E , —NHC(O)NR 3D R 3E , —NHS(O) 2 R 3D , —NHC(O)R 3D ,
- R 3C may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3D , —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 3D -substituted or unsubstituted alkyl, R 3D -substituted or unsubstituted
- R 3B may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl,
- R 3D is independently hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COR 3F , —NH 2 , —COH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 3F -substituted or unsubstituted heteroalkyl, R 3F -
- R 3D may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstitute
- R 3E and R 3F are independently hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloal
- R 3 and R 4 may together form a substituted or unsubstituted cycloalkyl.
- R 3 and R 4 may together form a R 3C -substituted or unsubstituted cycloalkyl.
- R 3 and R 4 may together form an unsubstituted cycloalkyl.
- R 3 and R 4 may together form an unsubstituted C 3 -C 6 cycloalkyl.
- R 3 and R 4 may together form an unsubstituted saturated C 3 -C 6 cycloalkyl.
- R 3 and R 4 may together form an C 3 -C 6 unsubstituted unsaturated cycloalkyl.
- R 3 and R 4 may together form an R 3C -substituted saturated C 3 -C 6 cycloalkyl where R 3C is as defined herein.
- R 3C may be halogen, —COR 3D , unsubstituted C 1 -C 5 alkyl, substituted or unsubstituted cycloalkyl, or substituted or unsubstituted heterocycloalkyl.
- R 3C may be methyl, unsubstituted ethyl, or unsubstituted propyl.
- R 3C may be substituted or unsubstituted C 3 -C 6 cycloalkyl.
- R 3C may be unsubstituted C 3 -C 6 cycloalkyl.
- R 3C may be substituted C 3 -C 6 cycloalkyl.
- R 3C may be R 3D -substituted cycloalkyl, where R 3D is as defined herein.
- R 3C may be substituted or unsubstituted C 3 -C 6 heterocycloalkyl.
- R 3C may be unsubstituted C 3 -C 6 heterocycloalkyl.
- R 3C may be substituted C 3 -C 6 heterocycloalkyl.
- R 3C may be R 3D -substituted heterocycloalkyl, where R 3D is as defined herein.
- R 3D may be —COR 3E , substituted or unsubstituted alkyl or substituted or unsubstituted heterocycloalkyl.
- R 4 may be substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 4 may be substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 4 may be substituted or unsubstituted alkyl.
- R 4 may be substituted alkyl.
- R 4 may be unsubstituted alkyl.
- R 4 may be substituted or unsubstituted C 1 -C 20 alkyl.
- R 4 may be substituted C 1 -C 20 alkyl.
- R 4 may be substituted or unsubstituted C 1 -C 10 alkyl.
- R 4 may be substituted C 1 -C 10 alkyl.
- R 4 may be unsubstituted C 1 -C 10 alkyl.
- R 4 may be substituted or unsubstituted C 1 -C 5 alkyl.
- R 4 may be substituted C 1 -C 5 alkyl.
- R 4 may be unsubstituted C 1 -C 5 alkyl.
- R 4 may be methyl, substituted or unsubstituted ethyl, or substituted or unsubstituted propyl.
- R 4 may be R 40 -substituted or unsubstituted alkyl.
- R 4 may be R 40 -substituted alkyl.
- R 4 may be R 40 -substituted or unsubstituted C 1 -C 20 alkyl.
- R 4 may be R 40 -substituted C 1 -C 20 alkyl.
- R 4 may be R 40 -substituted or unsubstituted C 1 -C 10 alkyl.
- R 4 may be R 40 -substituted C 1 -C 10 alkyl.
- R 4 may be R 40 -substituted or unsubstituted C 1 -C 5 alkyl.
- R 4 may be R 40 -substituted C 1 -C 5 alkyl.
- R 4 may be methyl, R 40 -substituted or unsubstituted ethyl, or R 40 -substituted or unsubstituted propyl.
- R 4 may be substituted or unsubstituted heteroalkyl.
- R 4 may be substituted heteroalkyl.
- R 4 may be unsubstituted heteroalkyl.
- R 4 may be substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 4 may be substituted 2 to 20 membered heteroalkyl.
- R 4 may be substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 4 may be substituted 2 to 10 membered heteroalkyl.
- R 4 may be unsubstituted 2 to 10 membered heteroalkyl.
- R 4 may be substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 4 may be substituted 2 to 6 membered heteroalkyl.
- R 4 may be unsubstituted 2 to 6 membered heteroalkyl.
- R 4 may be R 40 -substituted or unsubstituted heteroalkyl.
- R 4 may be R 40 -substituted heteroalkyl.
- R 4 may be R 40 -substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 4 may be R 40 -substituted 2 to 20 membered heteroalkyl.
- R 4 may be R 40 -substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 4 may be R 40 -substituted 2 to 10 membered heteroalkyl.
- R 4 may be R 40 -substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 4 may be R 40 -substituted 2 to 6 membered heteroalkyl.
- R 4 may be substituted or unsubstituted cycloalkyl.
- R 4 may be substituted cycloalkyl.
- R 4 may be unsubstituted cycloalkyl.
- R 4 may be substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 4 may be substituted 3 to 20 membered cycloalkyl.
- R 4 may be unsubstituted 3 to 20 membered cycloalkyl.
- R 4 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 4 may be substituted 3 to 10 membered cycloalkyl.
- R 4 may be unsubstituted 3 to 10 membered cycloalkyl.
- R 4 may be substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 4 may be substituted 3 to 6 membered cycloalkyl.
- R 4 may be unsubstituted 3 to 6 membered cycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted cycloalkyl.
- R 4 may be R 40 -substituted cycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 4 may be R 40 -substituted 3 to 20 membered cycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 4 may be R 40 -substituted 3 to 10 membered cycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 4 may be R 40 -substituted 3 to 6 membered cycloalkyl.
- R 4 may be substituted or unsubstituted heterocycloalkyl.
- R 4 may be substituted heterocycloalkyl.
- R 4 may be unsubstituted heterocycloalkyl.
- R 4 may be substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 4 may be substituted 3 to 20 membered heterocycloalkyl.
- R 4 may be unsubstituted 3 to 20 membered heterocycloalkyl.
- R 4 may be substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 4 may be substituted 3 to 10 membered heterocycloalkyl.
- R 4 may be unsubstituted 3 to 10 membered heterocycloalkyl.
- R 4 may be substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 4 may be substituted 3 to 6 membered heterocycloalkyl.
- R 4 may be unsubstituted 3 to 6 membered heterocycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted heterocycloalkyl.
- R 4 may be R 40 -substituted heterocycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 4 may be R 40 -substituted 3 to 20 membered heterocycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 4 may be R 40 -substituted 3 to 10 membered heterocycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 4 may be R 40 -substituted 3 to 6 membered heterocycloalkyl.
- R 4 may be substituted or unsubstituted aryl.
- R 4 may be substituted aryl.
- R 4 may be unsubstituted aryl.
- R 4 may be substituted or unsubstituted 5 to 20 membered aryl.
- R 4 may be substituted 5 to 20 membered aryl.
- R 4 may be substituted or unsubstituted 5 to 8 membered aryl.
- R 4 may be substituted 5 to 8 membered aryl.
- R 4 may be unsubstituted 5 to 8 membered aryl.
- R 4 may be substituted or unsubstituted 5 to 8 membered aryl.
- R 4 may be substituted or unsubstituted 5 or 6 membered aryl.
- R 4 may be substituted 5 or 6 membered aryl (e.g. phenyl).
- R 4 may be unsubstituted 5 or 6 membered aryl.
- R 4 may be R 40 -substituted or unsubstituted aryl.
- R 4 may be R 40 -substituted aryl.
- R 4 may be R 40 -substituted or unsubstituted 5 to 20 membered aryl.
- R 4 may be R 40 -substituted 5 to 20 membered aryl.
- R 4 may be R 40 -substituted or unsubstituted 5 to 8 membered aryl.
- R 4 may be R 40 -substituted 5 to 8 membered aryl.
- R 4 may be R 40 -substituted or unsubstituted 5 or 6 membered aryl.
- R 4 may be R 40 -substituted 5 or 6 membered aryl (e.g. phenyl).
- R 4 may be substituted or unsubstituted aryl.
- R 4 may be substituted heteroaryl.
- R 4 may be unsubstituted heteroaryl.
- R 4 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 4 may be substituted 5 to 20 membered aryl.
- R 4 may be unsubstituted 5 to 20 membered heteroaryl.
- R 4 may be substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 4 may be substituted 5 to 8 membered heteroaryl.
- R 4 may be unsubstituted 5 to 8 membered heteroaryl.
- R 4 may be substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 4 may be substituted 5 or 6 membered heteroaryl.
- R 4 may be unsubstituted 5 or 6 membered heteroaryl.
- R 4 may be R 40 -substituted or unsubstituted heteroaryl.
- R 4 may be R 40 -substituted heteroaryl.
- R 4 may be R 40 -substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 4 may be R 40 -substituted 5 to 20 membered heteroaryl.
- R 4 may be R 40 -substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 4 may be R 40 -substituted 5 to 8 membered heteroaryl.
- R 4 may be R 40 -substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 4 may be R 40 -substituted 5 or 6 membered heteroaryl.
- R 4 may be R 40 -substituted or unsubstituted cycloalkyl, R 40 -substituted or unsubstituted heterocycloalkyl, R 40 -substituted or unsubstituted aryl, or R 40 -substituted or unsubstituted heteroaryl.
- R 4 may be R 40 -substituted or unsubstituted cycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted aryl.
- R 4 may be R 40 -substituted or unsubstituted heteroaryl.
- R 4 may be R 40 -substituted or unsubstituted heterocycloalkyl.
- R 4 may be substituted or unsubstituted 5 to 8 membered heterocycloalkyl.
- R 4 may be R 40 -substituted or unsubstituted 5 to 8 membered heterocycloalkyl.
- R 4 may be substituted or unsubstituted 5 to 8 membered heterocycloalkyl having at least one ring nitrogen.
- R 4 may be R 4 -substituted or unsubstituted 5 to 8 membered heterocycloalkyl having at least one ring nitrogen.
- R 4 may be substituted or unsubstituted piperidinyl, substituted or unsubstituted piperazinyl, substituted or unsubstituted morpholino, or substituted or unsubstituted pyrrolidinyl.
- R 4 may be substituted or unsubstituted piperidinyl.
- R 4 may be substituted or unsubstituted piperazinyl.
- R 4 may be substituted or unsubstituted morpholino.
- R 4 may be substituted or unsubstituted pyrrolidinyl.
- R 4 may be R 40 -substituted or unsubstituted piperidinyl, R 40 -substituted or unsubstituted piperazinyl, R 40 -substituted or unsubstituted morpholino, or R 40 -substituted or unsubstituted pyrrolidinyl.
- R 4 may be R 40 -substituted or unsubstituted piperidinyl.
- R 4 may be R 40 -substituted or unsubstituted piperazinyl.
- R 4 may be R 40 -substituted or unsubstituted morpholino.
- R 4 may be R 40 -substituted or unsubstituted pyrrolidinyl.
- R 40 is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 40A , —OR 40A , —NR 40A R 40B , —C(O)OR 40A , —C(O)NR 40A R 40B , —NO 2 , —SR 40A , —S(O) 2 R 40A , —S(O) 2 OR 40A , —S(O) 2 NR 40A R 40B , —NHNR 40A R 40B , —ONR 40A R 40B , —NHC(O)NHNR 40A R 40B , R 40A -substituted or unsubstituted alkyl, R 40A -substituted or unsubstituted heteroalkyl, R 40A -substituted or unsubstituted cyclo
- R 40 may independently be oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 40A , —OR 40A , —NR 40A R 40B , —C(O)OR 40A , —C(O)NR 40A R 40B , —NO 2 , —SR 40A , —S(O) 2 R 40A , —S(O) 2 OR 40A , —S(O) 2 NR 40A R 40B , —NHNR 40A R 40B , —ONR 40A R 40B , —NHC(O)NHNR 40A R 40B , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted heterocycloalkyl, unsubstituted aryl, or unsubstit
- R 40 may independently be oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —C(O)OH, —C(O)NH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 H, —S(O) 2 OH, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted heterocycloalkyl, unsubstituted aryl, or unsubstituted heteroaryl.
- R 40A is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 41 , —NR 41 R 40C , —COR 41 , —COOR 41 , —CONR 41 R 40C , —NO 2 , —SR 41 , —S(O) 2 R 41 , —S(O) 3 R 41 , —S(O) 2 NR 41 R 40C , S(O) 4 R 41 , —NHNR 41 R 40C , ONR 41 R 40C , —NHC(O)NHNR 41 R 40C , —NHC(O)NR 41 R 40C , —NHS(O) 2 R 41 , —NHC(O)R 41 , —NHC(O)—OR 41 , —NHOR 41 , —OCF 3 , —OCHF 2 , R
- R 40A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COR 41 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 41 -substituted or unsubstituted alkyl, R 41 -substituted or unsubstituted hetero
- R 40A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalky
- R 41 is independently hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 41A , —OR 41A , —NR 41A R 41B , —C(O)OR 41A , —C(O)NR 41A R 41B , —NO 2 , —SR 41A , —S(O) 2 R 41A , —S(O) 2 OR 41A , —S(O) 2 NR 41A R 41B , —NR 41A R 41B , —ONR 41A R 41B , —NHC(O)NHNR 41A R 41B , R 42 -substituted or unsubstituted alkyl, R 42 -substituted or unsubstituted heteroalkyl, R 42 -substituted or unsubstituted cycloalkyl, R 42
- R 41 may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl
- R 42 is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 43 -substituted or unsubstituted alkyl, R 43 -substituted or unsubstituted
- R 42 may independently be oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, un
- R 43 is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 44 -substituted or unsubstituted alkyl, R 44 -substituted or unsubstituted
- R 41A is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 41C -substituted or unsubstituted heteroalkyl, R 41C -substituted or
- R 41C is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 41D -substituted or unsubstituted alkyl, R 41D -substituted or un
- R 41B is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 41E -substituted or unsubstituted alkyl, R 41E -substituted or un
- R 41E is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 41F -substituted or unsubstituted alkyl, R 41F -substituted or unsubsti
- R 40B is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 45 , —NR 45 R 40D , —COR 45 , —COOR 45 , —CONR 45 R 40D , —NO 2 , —SR 45 , —S(O) 2 R 45 , —S(O) 3 R 45 , —S(O) 4 R 45 , —S(O) 2 NR 45 R 40D , —NHNR 45 R 40D , —ONR 45 R 40D , —NHC(O)NHNR 45 R 40D , —NHC(O)NR 45 R 40D , —NHS(O) 2 R 45 , —NHC(O)R 45 , —NHC(O)—OR 45 , —NHOR 45 , —OCF 3 , —OCHF
- R 40B may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COR 45 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 45 -substituted or unsubstituted heteroalkyl, R 45 -substituted or unsubstituted
- R 40B may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —COH, —COCH 3 , —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalky
- R 45 is independently hydrogen, halogen, oxo, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 45A , —OR 45A , —NR 45A R 45B , —C(O)OR 45A , —C(O)NR 45A R 45B , —NO 2 , —SR 45A , —S(O) 2 R 45A , —S(O) 2 OR 45A , —S(O) 2 NR 45A R 45B , —NHNR 45A R 45B , —ONR 45A R 45B , —NHC(O)NHNR 45A R 45B , R 46 -substituted or unsubstituted alkyl, R 46 -substituted or unsubstituted heteroalkyl, R 46 -substituted or unsubstituted cycloal
- R 45A is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 45C -substituted or unsubstituted heteroalkyl, R 45C -substituted or
- R 45C is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 45D -substituted or unsubstituted alkyl, R 45D -substituted or un
- R 45B is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 45E -substituted or unsubstituted alkyl, R 45E -substituted or un
- R 45E is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 45F -substituted or unsubstituted alkyl, R 45F -substituted or unsubsti
- R 46 is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 47 -substituted or unsubstituted alkyl, R 47 -substituted or unsubstituted
- R 40C , R 40D , R 41D , R 41F , and R 45D are independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsub
- R 45F , R 44 , and R 47 are independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstitute
- R 4 may be —NR 4A R 4B .
- R 4A and R 4B are as defined herein.
- R 4A may be hydrogen, R 41 -substituted or unsubstituted alkyl, or R 41 -substituted or unsubstituted heteroalkyl, where R 41 is as defined herein.
- R 4A may be hydrogen, R 41 -substituted or unsubstituted alkyl, or R 41 -substituted or unsubstituted heteroalkyl, where R 41 is hydrogen, halogen, —CF 3 , —OR 41A , —NR 41A R 41B , R 42 -substituted or unsubstituted alkyl, R 42 -substituted or unsubstituted heteroalkyl, or R 42 -substituted or unsubstituted aryl.
- R 4A may be hydrogen, R 41 -substituted or unsubstituted alkyl, or R 41 -substituted or unsubstituted heteroalkyl, where R 41 is hydrogen, halogen, —CF 3 , —OR 41A , —NR 41A R 41B , R 42 -substituted or unsubstituted alkyl, R 42 -substituted or unsubstituted heteroalkyl, or R 42 -substituted or unsubstituted aryl, R 41A and R 41B are independently hydrogen or unsubstituted C 1 -C 5 alkyl, and R 42 is as described herein.
- R 4B may be hydrogen, R 45 -substituted or unsubstituted alkyl, or R 45 -substituted or unsubstituted heteroalkyl, where R 45 is as defined herein.
- R 4B may be hydrogen, R 45 -substituted or unsubstituted alkyl, or R 45 -substituted or unsubstituted heteroalkyl, where R 45 is hydrogen, halogen, CF 3 , —OR 45A , —NR 45A R 45B , R 46 -substituted or unsubstituted alkyl, R 46 -substituted or unsubstituted heteroalkyl, or R 46 -substituted or unsubstituted aryl.
- R 4B may be hydrogen, R 45 -substituted or unsubstituted alkyl, or R 45 -substituted or unsubstituted heteroalkyl, where R 45 is hydrogen, halogen, CF 3 , —OR 45A , —NR 45A R 45B , R 46 -substituted or unsubstituted alkyl, R 46 -substituted or unsubstituted heteroalkyl, or R 46 -substituted or unsubstituted aryl, R 45A and R 45B are independently hydrogen or unsubstituted C 1 -C 5 alkyl and R 46 is as defined herein.
- R 4A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 41 , —NR 41 R 40C , —COR 41 , —COOR 41 , —CONR 41 R 40C , —NO 2 , —SR 41 , —S(O) 2 R 41 , —S(O) 3 R 41 , —S(O) 4 R 41 , —NHNR 41 R 40C , ONR 41 R 40C , —NHC(O)NHNR 41 R 40C , —NHC(O)NR 41 R 40C , —NHS(O) 2 R 41 , —NHC(O)R 41 , —NHC(O)—OR 41 , —NHOR 41 , —OCF 3 , —OCHF 2 , R 41 -substituted or unsubsti
- R 4A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COR 41 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 41 -substituted or unsubstituted alkyl, R 41 -substituted or unsubstituted hetero
- R 4A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 41 -substituted or unsubstituted alkyl, R 41 -substituted or unsub
- R 4B may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 45 , —NR 45 R 4D , —COR 45 , —COOR 45 , —CONR 45 R 40D , —NO 2 , —SR 45 , —S(O) 2 R 45 , —S(O) 3 R 45 , —S(O) 4 R 45 , —S(O) 2 NR 45 R 40D , —NHNR 45 R 40D , —ONR 45 R 40D , —NHC(O)NHNR 45 R 40D , —NHC(O)NR 45 R 40D , —NHS(O) 2 R 45 , —NHC(O)R 45 , —NHC(O)—OR 45 , —NHOR 45 , —OCF 3 , —OC
- R 4B may independently be oxo, halogen, —CF 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 45 -substituted or unsubstituted alkyl, R 45 -substituted or unsubstituted heteroalkyl, R 45 -substituted or unsubstituted cycloal
- the PLK4 inhibitor is the compound of Formula (Ia1) or a pharmaceutically acceptable salt thereof:
- L 1 , z1, R 3 , R 5 , R 6 , R 40A , and R 40B are as described herein.
- the PLK4 inhibitor is the compound of Formula (Ib1) or a pharmaceutically acceptable salt thereof:
- L 1 , z1, R 3 , R 5 , R 6 , R 40A , and R 40B are as described herein.
- R 5 may independently be hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 5A , —OR 5A , —NR 5A R 5B , —C(O)OR 5A , —C(O)NR 5A R 5B , —NO 2 , —SR 5A , —S(O) n5 R 5A , —S(O) n5 OR 5A , —S(O) n5 NR 5A R 5B , —NHNR 5A R 5B , —ONR 5A R 5B or —NHC(O)NHNR 5A R 5B .
- R 5 may independently be substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 5 may be hydrogen or halogen.
- R 5 may be hydrogen or —Cl, —I, or —Br.
- R 5 may be hydrogen or —Cl or —F.
- R 5 may be hydrogen.
- R 5 may be —Cl.
- R 5 may be —I.
- R 5 may be —Br.
- R 5 may be —F.
- the symbol z1 may be 1, 2, or 3.
- the symbol z1 may be 1 or 2.
- R 5 may be hydrogen or —Cl or —F where the symbol z1 is 1 or 2.
- R 5 may be substituted or unsubstituted alkyl.
- R 5 may be substituted alkyl.
- R 5 may be unsubstituted alkyl.
- R 5 may be substituted or unsubstituted C 1 -C 20 alkyl.
- R 5 may be substituted C 1 -C 20 alkyl.
- R 5 may be substituted or unsubstituted C 1 -C 10 alkyl.
- R 5 may be substituted C 1 -C 10 alkyl.
- R 5 may be unsubstituted C 1 -C 10 alkyl.
- R 5 may be substituted or unsubstituted C 1 -C 5 alkyl.
- R 5 may be unsubstituted C 1 -C 5 alkyl.
- R 5 may be substituted C 1 -C 5 alkyl.
- R 5 may be methyl, substituted or unsubstituted ethyl, or substituted or unsubstituted propyl.
- R 5 may be methyl.
- R 5 may be ethyl.
- R 5 may be R 5C -substituted or unsubstituted alkyl.
- R 5 may be R 5C -substituted alkyl.
- R 5 may be R 5C -substituted or unsubstituted C 1 -C 20 alkyl.
- R 5 may be R 5C -substituted C 1 -C 20 alkyl.
- R 5 may be R 5C -substituted or unsubstituted C 1 -C 10 alkyl.
- R 5 may be R 5C -substituted C 1 -C 10 alkyl.
- R 5 may be R 5C -substituted or unsubstituted C 1 -C 5 alkyl.
- R 5 may be R 5C -substituted C 1 -C 5 alkyl.
- R 5 may be methyl, R 5C -substituted or unsubstituted ethyl, or R 5C -substituted or unsubstituted propyl.
- R 5 may be substituted or unsubstituted heteroalkyl.
- R 5 may be substituted heteroalkyl.
- R 5 may be unsubstituted heteroalkyl.
- R 5 may be substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 5 may be substituted 2 to 20 membered heteroalkyl.
- R 5 may be substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 5 may be substituted 2 to 10 membered heteroalkyl.
- R 5 may be substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 5 may be substituted 2 to 6 membered heteroalkyl.
- R 5 may be R 5C -substituted or unsubstituted heteroalkyl.
- R 5 may be R 5C -substituted heteroalkyl.
- R 5 may be R 5C -substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 5 may be R 5C -substituted 2 to 20 membered heteroalkyl.
- R 5 may be R 5C -substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 5 may be R 5C -substituted 2 to 10 membered heteroalkyl.
- R 5 may be R 5C -substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 5 may be R 5C -substituted 2 to 6 membered heteroalkyl.
- R 5 may be substituted or unsubstituted cycloalkyl.
- R 5 may be substituted cycloalkyl.
- R 5 may be unsubstituted cycloalkyl.
- R 5 may be substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 5 may be substituted 3 to 20 membered cycloalkyl.
- R 5 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 5 may be substituted 3 to 10 membered cycloalkyl.
- R 5 may be substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 5 may be substituted 3 to 6 membered cycloalkyl.
- R 5 may be R 5C -substituted or unsubstituted cycloalkyl.
- R 5 may be R 5C -substituted cycloalkyl.
- R 5 may be R 5C -substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 5 may be R 5C -substituted 3 to 20 membered cycloalkyl.
- R 5 may be R 5C -substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 5 may be R 5C -substituted 3 to 10 membered cycloalkyl.
- R 5 may be R 5C -substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 5 may be R 5C -substituted 3 to 6 membered cycloalkyl.
- R 5 may be substituted or unsubstituted heterocycloalkyl.
- R 5 may be substituted heterocycloalkyl.
- R 5 may be unsubstituted heterocycloalkyl.
- R 5 may be substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 5 may be substituted 3 to 20 membered heterocycloalkyl.
- R 5 may be substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 5 may be substituted 3 to 10 membered heterocycloalkyl.
- R 5 may be substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 5 may be substituted 3 to 6 membered heterocycloalkyl.
- R 5 may be R 5C -substituted or unsubstituted heterocycloalkyl.
- R 5 may be R 5C -substituted heterocycloalkyl.
- R 5 may be R 5C -substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 5 may be R 5C -substituted 3 to 20 membered heterocycloalkyl.
- R 5 may be R 5C -substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 5 may be R 5C -substituted 3 to 10 membered heterocycloalkyl.
- R 5 may be R 5C -substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 5 may be R 5C -substituted 3 to 6 membered heterocycloalkyl.
- R 5 may be substituted or unsubstituted aryl.
- R 5 may be substituted aryl.
- R 5 may be unsubstituted aryl.
- R 5 may be substituted or unsubstituted 5 to 20 membered aryl.
- R 5 may be substituted 5 to 20 membered aryl.
- R 5 may be substituted or unsubstituted 5 to 8 membered aryl (e.g. phenyl).
- R 5 may be substituted 5 to 8 membered aryl.
- R 5 may be substituted or unsubstituted 5 or 6 membered aryl.
- R 5 may be substituted 5 or 6 membered aryl.
- R 5 may be R 5C -substituted or unsubstituted aryl.
- R 5 may be R 5 -substituted aryl.
- R 5 may be R 5C -substituted or unsubstituted 5 to 20 membered aryl.
- R 5 may be R 5C -substituted 5 to 20 membered aryl.
- R 5 may be R 5C -substituted or unsubstituted 5 to 8 membered aryl.
- R 5 may be R 5C -substituted 5 to 8 membered aryl.
- R 5 may be R 5C -substituted or unsubstituted 5 or 6 membered aryl.
- R 5 may be R 5C -substituted 5 or 6 membered aryl (e.g. phenyl).
- R 5 may be substituted or unsubstituted heteroaryl.
- R 5 may be substituted heteroaryl.
- R 5 may be unsubstituted heteroaryl.
- R 5 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 5 may be substituted 5 to 20 membered heteroaryl.
- R 5 may be substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 5 may be substituted 5 to 8 membered heteroaryl.
- R 5 may be substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 5 may be substituted 5 or 6 membered heteroaryl.
- R 5 may be R 5C -substituted or unsubstituted heteroaryl.
- R 5 may be R 5 -substituted heteroaryl.
- R 5 may be R 5C -substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 5 may be R 5C -substituted 5 to 20 membered heteroaryl.
- R 5 may be R 5C -substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 5 may be R 5C -substituted 5 to 8 membered heteroaryl.
- R 5 may be R 5C -substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 5 may be R 5C -substituted 5 or 6 membered heteroaryl.
- R 5A may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 5C -substituted or unsubstituted alkyl, R 5 -substituted or unsubsti
- R 5C is independently halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COR 5D , —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 5D -substituted or unsubstituted alkyl, R 5D -substit
- R 5B may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl,
- R 5D is independently halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COR 5E , —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 5E -substituted or unsubstituted heteroalkyl, R 5E -substi
- R 5E is independently halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubsti
- R 6 may be hydrogen, —NH 2 , —CF 3 , —NR 6A R 6B , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 6 may be hydrogen, —CF 3 , —NR 6A R 6B , R 60 -substituted or unsubstituted alkyl, R 60 -substituted or unsubstituted heteroalkyl, R 60 -substituted or unsubstituted cycloalkyl, R 60 -substituted or unsubstituted heterocycloalkyl, R 60 -substituted or unsubstituted aryl, or R 60 -substituted or unsubstituted heteroaryl.
- R 6 may be substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 6 may be substituted or unsubstituted alkyl.
- R 6 may be substituted alkyl.
- R 6 may be unsubstituted alkyl.
- R 6 may be substituted or unsubstituted C 1 -C 20 alkyl.
- R 6 may be substituted C 1 -C 20 alkyl.
- R 6 may be substituted or unsubstituted C 1 -C 10 alkyl.
- R 6 may be substituted C 1 -C 10 alkyl.
- R 6 may be unsubstituted C 1 -C 10 alkyl.
- R 6 may be substituted or unsubstituted C 1 -C 5 alkyl.
- R 6 may be substituted C 1 -C 5 alkyl.
- R 6 may be unsubstituted C 1 -C 5 alkyl.
- R 6 may be methyl, substituted or unsubstituted ethyl, or substituted or unsubstituted propyl.
- R 6 may be hydrogen, methyl, ethyl or propyl.
- R 6 may be hydrogen.
- R 6 may be methyl.
- R 6 may be R 60 -substituted or unsubstituted alkyl.
- R 6 may be R 60 -substituted alkyl.
- R 6 may be R 60 -substituted or unsubstituted C 1 -C 20 alkyl.
- R 6 may be R 60 -substituted C 1 -C 20 alkyl.
- R 6 may be R 60 -substituted or unsubstituted C 1 -C 10 alkyl.
- R 6 may be R 60 -substituted C 1 -C 10 alkyl.
- R 6 may be R 60 -substituted or unsubstituted C 1 -C 5 alkyl.
- R 6 may be R 60 -substituted C 1 -C 5 alkyl.
- R 6 may be methyl, R 60 -substituted or unsubstituted ethyl, or R 60 -substituted or unsubstituted propyl.
- R 6 may be substituted or unsubstituted heteroalkyl.
- R 6 may be substituted heteroalkyl.
- R 6 may be unsubstituted heteroalkyl.
- R 6 may be substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 6 may be substituted 2 to 20 membered heteroalkyl.
- R 6 may be unsubstituted 2 to 20 membered heteroalkyl.
- R 6 may be substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 6 may be substituted 2 to 10 membered heteroalkyl.
- R 6 may be unsubstituted 2 to 10 membered heteroalkyl.
- R 6 may be substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 6 may be substituted 2 to 6 membered heteroalkyl.
- R 6 may be unsubstituted 2 to 6 membered heteroalkyl.
- R 6 may be R 60 -substituted or unsubstituted heteroalkyl.
- R 6 may be R 60 -substituted heteroalkyl.
- R 6 may be R 60 -substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 6 may be R 60 -substituted 2 to 20 membered heteroalkyl.
- R 6 may be R 60 -substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 6 may be R 60 -substituted 2 to 10 membered heteroalkyl.
- R 6 may be R 60 -substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 6 may be R 60 -substituted 2 to 6 membered heteroalkyl.
- R 6 may be substituted or unsubstituted cycloalkyl.
- R 6 may be substituted cycloalkyl.
- R 6 may be unsubstituted cycloalkyl.
- R 6 may be substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 6 may be substituted 3 to 20 membered cycloalkyl.
- R 6 may be unsubstituted 3 to 20 membered cycloalkyl.
- R 6 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 6 may be substituted 3 to 10 membered cycloalkyl.
- R 6 may be unsubstituted 3 to 10 membered cycloalkyl.
- R 6 may be substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 6 may be substituted 3 to 6 membered cycloalkyl.
- R 6 may be unsubstituted 3 to 6 membered cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted cycloalkyl.
- R 6 may be R 60 -substituted cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 6 may be R 60 -substituted 3 to 20 membered cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 6 may be R 60 -substituted 3 to 10 membered cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 6 may be R 60 -substituted 3 to 6 membered cycloalkyl.
- R 6 may be R 60 -substituted 3
- R 6 may be substituted or unsubstituted heterocycloalkyl.
- R 6 may be substituted heterocycloalkyl.
- R 6 may be unsubstituted heterocycloalkyl.
- R 6 may be substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 6 may be substituted 3 to 20 membered heterocycloalkyl.
- R 6 may be unsubstituted 3 to 20 membered heterocycloalkyl.
- R 6 may be substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 6 may be substituted 3 to 10 membered heterocycloalkyl.
- R 6 may be unsubstituted 3 to 10 membered heterocycloalkyl.
- R 6 may be substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 6 may be substituted 3 to 6 membered heterocycloalkyl.
- R 6 may be unsubstituted 3 to 6 membered heterocycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted heterocycloalkyl.
- R 6 may be R 60 -substituted heterocycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 6 may be R 60 -substituted 3 to 20 membered heterocycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 6 may be R 60 -substituted 3 to 10 membered heterocycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 6 may be R 60 -substituted 3 to 6 membered heterocycloalkyl.
- R 6 may be substituted or unsubstituted aryl.
- R 6 may be substituted aryl.
- R 6 may be unsubstituted aryl.
- R 6 may be substituted or unsubstituted 5 to 20 membered aryl.
- R 6 may be substituted 5 to 20 membered aryl.
- R 6 may be substituted or unsubstituted 5 to 8 membered aryl.
- R 6 may be substituted 5 to 8 membered aryl.
- R 6 may be unsubstituted 5 to 8 membered aryl.
- R 6 may be substituted or unsubstituted 5 or 6 membered aryl.
- R 6 may be substituted 5 or 6 membered aryl.
- R 6 may be substituted 5 or 6 membered aryl (e.g. phenyl).
- R 6 may be unsubstituted 5 or 6 membered aryl (e.g. phenyl).
- R 6 may be R 60 -substituted or unsubstituted aryl.
- R 6 may be R 60 -substituted aryl.
- R 6 may be R 60 -substituted or unsubstituted 5 to 20 membered aryl.
- R 6 may be R 60 -substituted 5 to 20 membered aryl.
- R 6 may be R 60 -substituted or unsubstituted 5 to 8 membered aryl.
- R 6 may be R 60 -substituted 5 to 8 membered aryl.
- R 6 may be R 60 -substituted or unsubstituted 5 or 6 membered aryl.
- R 6 may be R 60 -substituted 5 or 6 membered aryl (e.g. phenyl).
- R 6 may be substituted or unsubstituted heteroaryl.
- R 6 may be substituted heteroaryl.
- R 6 may be unsubstituted heteroaryl.
- R 6 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 6 may be substituted 5 to 20 membered heteroaryl.
- R 6 may be unsubstituted 5 to 20 membered heteroaryl.
- R 6 may be substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 6 may be substituted 5 to 8 membered heteroaryl.
- R 6 may be unsubstituted 5 to 8 membered heteroaryl.
- R 6 may be substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 6 may be substituted 5 or 6 membered heteroaryl.
- R 6 may be unsubstituted 5 or 6 membered heteroaryl.
- R 6 may be substituted 5 or 6 membered
- R 6 may be R 60 -substituted or unsubstituted heteroaryl.
- R 6 may be R 60 -substituted heteroaryl.
- R 6 may be R 60 -substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 6 may be R 60 -substituted 5 to 20 membered heteroaryl.
- R 6 may be R 60 -substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 6 may be R 60 -substituted 5 to 8 membered heteroaryl.
- R 6 may be R 60 -substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 6 may be R 60 -substituted 5 or 6 membered heteroaryl.
- R 6 may be R 60 -substituted or unsubstituted cycloalkyl, R 60 -substituted or unsubstituted heterocycloalkyl, R 60 -substituted or unsubstituted aryl, or R 60 -substituted or unsubstituted heteroaryl.
- R 6 may be R 60 -substituted or unsubstituted C 3 cycloalkyl.
- R 6 may be unsubstituted C 3 cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted C 4 cycloalkyl.
- R 6 may be unsubstituted C 4 cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted CS cycloalkyl.
- R 6 may be unsubstituted C 5 cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted saturated C 3 cycloalkyl.
- R 6 may be unsubstituted saturated C 3 cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted saturated C 4 cycloalkyl.
- R 6 may be R 60 -substituted or unsubstituted saturated C 5 cycloalkyl.
- R 6 may be unsubstituted saturated C 5 cycloalkyl.
- R 6 may R 60 -substituted or unsubstituted aryl where R 60 is as defined herein, R 60A is hydrogen, halogen, —NO 2 , —CF 3 , —CN, —COR 61 , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or unsubstituted heteroalkyl, or R 61 -substituted or unsubstituted aryl, where R 61 is as defined herein and R 60B is hydrogen, halogen, or unsubstituted alkyl.
- R 6 may be R 60 -substituted or unsubstituted heteroaryl where R 60 is halogen, —CF 3 , —NR 60A R 60B , —NO 2 , —OR 60A , COOR 60A , —COR 61 , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or unsubstituted heteroalkyl, or R 61 -substituted or unsubstituted aryl, where R 61 is —NR 61A R 61B , and R 60A , R 60B , R 61A , and R 61B are independently hydrogen or unsubstituted C 1 -C 5 unsubstituted alkyl.
- R 6 may be substituted or unsubstituted thiophenyl, substituted or unsubstituted thiazolyl, substituted or unsubstituted imidazolyl, or substituted or unsubstituted oxazolyl.
- R 6 may be substituted or unsubstituted thiophenyl.
- R 6 may be substituted or unsubstituted thiazolyl.
- R 6 may be substituted or unsubstituted imidazolyl.
- R 6 may be substituted or unsubstituted oxazolyl.
- R 6 may be R 60 -substituted or unsubstituted thiophenyl, R 60 -substituted or unsubstituted thiazolyl, R 60 -substituted or unsubstituted imidazolyl, or R 60 -substituted or unsubstituted oxazolyl.
- R 6 may be R 60 -substituted or unsubstituted thiophenyl.
- R 6 may be R 60 -substituted or unsubstituted thiazolyl.
- R 6 may be R 60 -substituted or unsubstituted imidazolyl.
- R 6 may be R 60 -substituted or unsubstituted oxazolyl.
- R 60 is independently hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 61 , —OR 60A , —NR 60A R 60B , —C(O)OR 60A , —C(O)NR 60A R 60B , —NO 2 , —SR 60A , —S(O) 2 H, —S(O) 2 OH, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNR 60A R 60B , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or unsubstituted heteroalkyl, R 61 -substituted or unsubstituted cycloalkyl, R 61 -substituted or unsubstitute
- R 60 may independently be oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COH, —COCH 3 , —OH, —NH 2 , —C(O)OH, —C(O)NH 2 , —NO 2 , —SH, —S(O) 2 H, —S(O) 2 OH, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted heterocycloalkyl, unsubstituted aryl, or unsubstituted heteroaryl.
- R 60A is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 61 , —NR 61 R 60C , —COOR 61 , —CONR 61 R 60C , —COR 61 , —NO 2 , —SR 61 , —S(O) 2 R 61 , —S(O) 3 R 61 , —S(O) 4 R 61 , —S(O) 2 NR 61 R 60C , —NHNR 61 R 60C , —ONR 61 R 60C , —NHC(O)NHNR 61 R 60C , —NHC(O)NR 61 R 60C , —NHC(O)NR 61 R 60C , —NHS(O) 2 R 61 , —NHC(O)R 61
- R 60A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COR 61 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or unsubstit
- R 60A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalky
- R 61 is independently hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 61A , —OR 61A , —NR 61A R 61B , —C(O)OR 61A , —C(O)NR 61A R 61B , —NO 2 , —SR 61A , —S(O) 2 R 61A , —S(O) 2 OR 61A , —S(O) 2 NR 61A R 61B , —NHNR 61A R 61B , —ONR 61A R 61B , —NHC(O)NHNR 61A R 61B , R 62 -substituted or unsubstituted alkyl, R 62 -substituted or unsubstituted heteroalkyl, R 62
- R 61 is independently hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 61A , —OR 61A , —NR 61A R 61B , —C(O)OR 61A , —C(O)NR 61A R 61B , —NO 2 , —SR 61A , —S(O) 2 H, —S(O) 2 OH, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNR 61A R 61B , R 62 -substituted or unsubstituted alkyl, R 62 -substituted or unsubstituted heteroalkyl, R 62 -substituted or unsubstituted cycloalkyl, R 62 -
- R 61 may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalky
- R 62 is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 63 -substituted or unsubstituted alkyl, R 63 -substituted or unsubsti
- R 62 may independently be oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl,
- R 63 is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 64 -substituted or unsubstituted alkyl, R 64 -substituted or unsubstitute
- R 61A is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 61C -substituted or unsubstituted heteroalkyl, R 61C -substit
- R 61A may independently be hydrogen or unsubstituted C 1 -C 5 alkyl.
- R 61A may independently be hydrogen.
- R 61A may independently be methyl or unsubstituted ethyl.
- R 61A may independently be methyl.
- R 61C is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 61D -substituted or unsubstituted alkyl, R 61D -substituted or
- R 61B is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 61E -substituted or unsubstituted alkyl, R 61E -substitute
- R 61B may independently be hydrogen or unsubstituted C 1 -C 5 alkyl.
- R 61B may independently be hydrogen.
- R 61B may independently be methyl or unsubstituted ethyl.
- R 61B may independently be methyl.
- R 61E is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 61F -substituted or unsubstituted alkyl, R 61F -substituted or
- R 60B is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 65 , —NR 65 R 60D , COOR 65 , CONR 65 R 60D , —COR 65 , —NO 2 , —SH, —S(O) 2 R 65 , —S(O) 3 R 65 , —S(O) 4 R 65 , —S(O) 2 NR 65 R 60D , —NHNR 65 R 60D , —ONR 65 R 60D , —NHC(O)NHNR 65 R 60D , —NHC(O)NR 65 R 60D , —NHS(O) 2 R 65 , —NHC(O)R 65 , —NHC(O)—OR 65 , —NHOR 65 , —OCF 3 , —OCHF 2 , R 65
- R 60B may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COR 65 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 65 -substituted or unsubstituted heteroalkyl, R 65 -substituted or unsubstituted
- R 60B may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalky
- R 65 is independently hydrogen, halogen, oxo, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 65A , —OR 65A , —NR 65A R 65B , —C(O)OR 65A , C(O)NR 65A R 65B , —NO 2 , —SR 65A , —S(O) 2 H, —S(O) 2 OH, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNR 65A R 65B , R 66 -substituted or unsubstituted alkyl, R 66 -substituted or unsubstituted heteroalkyl, R 66 -substituted or unsubstituted cycloalkyl, R 66 -substituted or unsub
- R 65A is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 65C -substituted or unsubstituted heteroalkyl, R 65C -substituted or
- R 65C is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 65D -substituted or unsubstituted alkyl, R 65D -substituted or unsubsti
- R 65B is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 65E -substituted or unsubstituted alkyl, R 65E -substituted or un
- R 65E is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 65F -substituted or unsubstituted alkyl, R 65F -substituted or unsubsti
- R 66 is independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 67 -substituted or unsubstituted alkyl, R 67 -substituted or unsubsti
- R 60C , R 60D , R 61D , R 61F , R 65D , R 65F , R 64 , and R 67 are independently oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2
- R 6A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 61 , —NR 61 R 60C , —COOR 61 , —CONR 61 R 60C , —COR 61 , —NO 2 , —SR 61 , —S(O) 2 R 61 , —S(O) 3 R 61 , —S(O) 4 R 61 , —S(O) 2 NR 61 R 60C , —NHNR 61 R 60C , —ONR 61 R 60C , —NHC(O)NHNR 61 R 60C , —NHC(O)NR 61 R 60C , —NHS(O)NR 61 R 60C , —NHS(O) 2 R 61 , —NHC(O)R 61
- R 6A may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or
- R 6B may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OR 65 , —NR 65 R 60D , COOR 65 , CONR 65 R 60D , —COR 65 , —NO 2 , —SH, —S(O) 2 R 65 , —S(O) 3 R 65 , —S(O) 4 R 65 , —S(O) 2 NR 65 R 60D , —NHNR 65 R 60D , —ONR 65 R 60D , —NHC(O)NHNR 65 R 60D , —NHC(O)NR 65 R 60D , —NHS(O) 2 R 65 , —NHC(O)R 65 , —NHC(O)—OR 65 , —NHOR 65 , —OCF 3 , —OCHF 2 , R 65
- R 6B may independently be oxo, halogen, —CF 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —COH, —COCH 3 , —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 65 -substituted or unsubstituted alkyl, R 65 -substituted or unsubstituted heteroalkyl, R 65 -substituted or unsubstituted cycloal
- the compound of Formula (Ia) is:
- R 3 , R 4 , R 5 , z1, R 13 , and R 60 are as described herein.
- the symbol z3 is an integer of 0, 1, 2, 3, 4, or 5.
- the symbol z3 may be 1, 2, or 3.
- the symbol z3 may be 1.
- the symbol z3 may be 2.
- the symbol z3 may be 3.
- R 60 may independently be hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 61 , —OR 60A , —NR 60A R 60B , —C(O)OR 60A , —C(O)NR 60A R 60B , —NO 2 , —SR 60A , —S(O) 2 H, —S(O) 2 OH, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNR 60A R 60B , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or unsubstituted heteroalkyl, R 61 -substituted or unsubstituted cycloalkyl, R 61 -substituted or unsubstit
- R 60A may independently be hydrogen, halogen, —NO 2 , —CF 3 , —CN, —COR 61 , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or unsubstituted heteroalkyl, or R 61 -substituted or unsubstituted aryl.
- R 61 may independently be hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 61A , —OR 61A , —NR 61A R 61B , —C(O)OR 61A , —C(O)NR 61A R 61B , —NO 2 , —SR 61A , —S(O) 2 H, —S(O) 2 OH, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNR 61A R 61B , R 62 -substituted or unsubstituted alkyl, R 62 -substituted or unsubstituted heteroalkyl, R 62 -substituted or unsubstituted cycloalkyl, R 62
- R 62 may independently be hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalky
- the compound of Formula (Ia) is:
- the compound of Formula (Ia) is:
- R 60 are independently hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 61 , —OR 60A , —NR 60A R 60B , —C(O)OR 60A , —C(O)NR 60A R 60B , —NO 2 , —SR 60A , —S(O) 2 H, —S(O) 2 OH, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNR 60A R 60B , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or unsubstituted heteroalkyl, R 61 -substituted or un
- R 60A may independently be hydrogen, halogen, —NO 2 , —CF 3 , —CN, —COR 61 , R 61 -substituted or unsubstituted alkyl, R 61 -substituted or unsubstituted heteroalkyl, or R 61 -substituted or unsubstituted aryl; R 61 is independently hydrogen, halogen, or unsubstituted alkyl.
- R 60B may independently be hydrogen halogen, or unsubstituted alkyl.
- the PLK4 inhibitor is the compound of Formula: (Ia2) or a pharmaceutically acceptable salt thereof:
- Ring A is cycloalkyl or heterocycloalkyl.
- the compound of Formula (Ia2) is:
- the PLK4 inhibitor is the compound of Formula (Ia3) or a pharmaceutically acceptable salt thereof:
- the PLK4 inhibitor is the compound of Formula (Ia4) or a pharmaceutically acceptable salt thereof:
- the PLK4 inhibitor is the compound of Formula (Ia5) or a pharmaceutically acceptable salt thereof:
- the PLK4 inhibitor is the compound of Formula (Ia6) or a pharmaceutically acceptable salt thereof:
- the PLK4 inhibitor is the compound of Formula (Ib2) or a pharmaceutically acceptable salt thereof:
- Ring A is cycloalkyl or heterocycloalkyl.
- the compound of Formula (Ib2) is:
- the compound of Formula (Ib2) is not tozasertib.
- the PLK4 inhibitor is the compound of Formula (Ib3) or a pharmaceutically acceptable salt thereof:
- the compound of Formula (Ib3) is not tozasertib.
- the PLK4 inhibitor is the compound of Formula (Ib4) or a pharmaceutically acceptable salt thereof:
- the PLK4 inhibitor is the compound of Formula (Ib5) or a pharmaceutically acceptable salt thereof:
- the PLK4 inhibitor is the compound of Formula (Ib6) or a pharmaceutically acceptable salt thereof:
- L 1 may be a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 13 —, —C(O)NR 13 —, —NR 13 C(O)—, —S(O) 2 —, —S(O)NR 13 —.
- L 1 may be a bond, substituted or unsubstituted alkylene, substituted or unsubstituted heteroalkylene, substituted or unsubstituted cycloalkylene, substituted or unsubstituted heterocycloalkylene, substituted or unsubstituted arylene, or substituted or unsubstituted heteroarylene.
- L 1 may be a bond, substituted or unsubstituted alkylene, or substituted or unsubstituted heteroalkylene.
- L 1 may be a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 13 —, —C(O)NR 13 —, —NR 13 C(O)—, —S(O) 2 —, —S(O)NR 13 —, substituted or unsubstituted alkylene, or substituted or unsubstituted heteroalkylene.
- L 1 may be a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 13 —, —C(O)NR 13 —, —NR 13 C(O)—, —S(O) 2 —, —S(O)NR 13 —, substituted or unsubstituted alkylene, or substituted or unsubstituted heteroalkylene where R 13 is hydrogen or substituted or unsubstituted alkyl.
- L 1 may be a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 13 —, —C(O)NR 13 —, —NR 13 C(O)—, —S(O) 2 —, —S(O)NR 13 —, R 13 -substituted or unsubstituted alkylene, or R 13 -substituted or unsubstituted heteroalkylene where R 13 is hydrogen or substituted or unsubstituted alkyl. R 13 may be unsubstituted alkyl.
- L 1 may be a bond or substituted or unsubstituted alkylene.
- L 1 may be a bond or R 13 -substituted or unsubstituted alkylene.
- L 1 may be a bond or R 13 -substituted or unsubstituted alkylene where R 13 is hydrogen or substituted or unsubstituted alkyl.
- L 1 may be a bond or substituted or unsubstituted C 1 -C 5 alkylene.
- L 1 may be a bond or R 13 -substituted or unsubstituted C 1 -C 5 alkylene.
- L 1 may be a bond or R 13 -substituted or unsubstituted C 1 -C 5 alkylene where R 13 is hydrogen or substituted or unsubstituted C 1 -C 5 alkyl.
- L 1 may be R 13 -substituted or unsubstituted alkylene where R 13 is hydrogen, halogen, or substituted or unsubstituted alkyl.
- L 1 may be unsubstituted alkylene.
- L 1 may be unsubstituted methylene.
- L 1 may be R 13 -substituted methylene.
- L 1 may be methylene substituted with one R 13 -substituent.
- the R 13 -substituent may be halogen.
- L 1 may be methylene substituted with one fluoro substituent.
- L 1 may be methylene independently substituted with two R 13 -substituents.
- L 1 may be methylene independently substituted with halogen substituents.
- the R 13 -substituent may be alkyl.
- L 1 may be methylene mono substituted with substituted or unsubstituted alkyl.
- L 1 may be methylene disubstituted with substituted or unsubstituted alkyl.
- the two independent R 13 -substitutents combine to form a substituted or unsubstituted cycloalkyl, or a substituted or unsubstituted heterocycloalkyl.
- L 1 may be methylene singly substituted with halogen or unsubstituted alkyl. In embodiments, L 1 may be methylene independently disubstituted with halogen or unsubstituted alkyl. In embodiments, L 1 may be methylene singly substituted with halogen or substituted alkyl. In embodiments, L 1 may be methylene independently disubstituted with halogen or substituted alkyl.
- R 13 may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 13A -substituted or unsubstituted heteroalkyl, R 13A -substituted or unsubstituted cyclo
- R 13A is independently halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubstituted heterocycloalkyl, unsub
- L 1 is a substituted or unsubstituted cycloalkylene or heterocycloalkylene
- a single carbon of the cycloalkylene or heterocycloalkylene is connected (bonded) to both the sulfonyl moiety and R 6 .
- L 1 is:
- L 1 may be substituted or unsubstituted C 1 -C 5 cycloalkylene. In embodiments, L 1 may be substituted or unsubstituted cyclopropylene. In embodiments, L 1 may be substituted or unsubstituted cyclobutylene. In embodiments, L 1 may be substituted or unsubstituted cyclopentylene. In embodiments, L may be unsubstituted cyclopropylene. In embodiments, L 1 may be unsubstituted cyclobutylene. In embodiments, L 1 may be unsubstituted cyclopentylene. In embodiments, L may be C 1 -C 5 cycloalkylene substituted with halogen, or substituted or unsubstituted alkyl.
- the PLK4 inhibitor is the compound of Formula (Ib7) or a pharmaceutically acceptable salt thereof:
- the compound of Formula (Ia7) is:
- the PLK4 inhibitor is the compound of Formula (Ib8) or a pharmaceutically acceptable salt thereof:
- the compound of Formula (Ia8) is:
- the PLK4 inhibitor is the compound of Formula (Ia9a) or a pharmaceutically acceptable salt thereof.
- R 3A is substituted or unsubstituted alkyl
- R 4C is substituted or unsubstituted heterocycloalkyl
- R 2 is substituted or unsubstituted heteroaryl. In embodiments, R 2 is unsubstituted heteroaryl. In embodiments, R 2 is substituted heteroaryl. In embodiments, R 2 is R 2C -substituted heteroaryl, wherein R 2C is as defined herein. R 2 may be R 2C -substituted or unsubstituted 5 to 20 membered heteroaryl. R 2 may be R 2C -substituted 5 to 20 membered heteroaryl. R 2 may be R 2C -substituted or unsubstituted 5 to 8 membered heteroaryl. R 2 may be R 2C -substituted 5 to 8 membered heteroaryl. R 2 may be R 2C -substituted or unsubstituted 5 or 6 membered heteroaryl. R 2 may be R 2C -substituted 5 or 6 membered heteroaryl. R 2 may be R 2C -substi
- R 2 is substituted or unsubstituted pyrazolyl. In embodiments, R 2 is unsubstituted pyrazolyl. In embodiments, R 2 is substituted pyrazolyl. In embodiments, R 2 is pyrazolyl substituted with substituted or unsubstituted alkyl. In embodiments, R 2 is pyrazolyl substituted with unsubstituted alkyl. In embodiments, R 2 is pyrazolyl substituted with unsubstituted lower alkyl. In embodiments, R 2 is pyrazolyl substituted methyl, ethyl or propyl. In embodiments, R 2 is methyl substituted pyrazolyl.
- R 2 is R 2C -substituted pyrazolyl. In embodiments, R 2 is R 2C -substituted pyrazolyl, wherein R 2C is substituted or unsubstituted alkyl. In embodiments, R 2 is R 2C -substituted pyrazolyl, wherein R 2C is unsubstituted alkyl. In embodiments, R 2 is R 2C -substituted pyrazolyl, wherein R 2C is unsubstituted lower alkyl. In embodiments, R 2 is R 2C -substituted pyrazolyl, wherein R 2C is unsubstituted methyl, ethyl or propyl. In embodiments, R 2 is R 2C -substituted pyrazolyl, wherein R 2C is unsubstituted methyl.
- the PLK4 inhibitor is the compound of Formula (Ia9b) or a pharmaceutically acceptable salt thereof:
- R 3A is substituted or unsubstituted alkyl. In embodiments R 3A is unsubstituted alkyl. In embodiments, R 3A is unsubstituted lower alkyl. In embodiments, R 3A is methyl, ethyl or propyl. In embodiments, R 3A is methyl. In embodiments, R 3A is R 3C -substituted or unsubstituted alkyl, wherein R 3C is as defined herein.
- the PLK4 inhibitor is the compound of Formula (Ia9c) or a pharmaceutically acceptable salt thereof:
- R 4C is substituted or unsubstituted heterocycloalkyl. In embodiments, R 4C is unsubstituted heterocycloalkyl. In embodiments, R 4C is substituted heterocycloalkyl. In embodiments, R 3C is R 40 -substituted heterocycloalkyl, wherein R 40 is as defined herein.
- the PLK4 inhibitor is the compound of Formula (Ia9d) or a pharmaceutically acceptable salt thereof:
- R 40 may be substituted or unsubstituted alkyl or substituted or unsubstituted heteroalkyl (e.g. —CH 2 C(O)N(CH 3 ) 2 ).
- R 5 and z1 are as defined herein.
- R 5 is halogen, and z1 is 1.
- R 5 is fluoro.
- R 6 is substituted or unsubstituted aryl. In embodiments, R 6 is unsubstituted aryl. R 6 is aryl independently substituted one or more times with halogen or —NO 2 .
- the PLK4 inhibitor is the compound of Formula (Ia9f) or a pharmaceutically acceptable salt thereof:
- R 60 and z5 are as defined herein.
- R 60 is halogen or —NO 2
- z5 is 1 or 2.
- z5 is 1.
- z5 is 2.
- the PLK4 inhibitor is the compound of Formula (Ib7) or a pharmaceutically acceptable salt thereof:
- the compound of Formula (Ib7) is:
- the compound of Formula (Ib7) is not tozasertib.
- the PLK4 inhibitor is the compound of Formula (IC) or a pharmaceutically acceptable salt thereof:
- R 3 , R 4 , R 1C , R 5 and R 13 are as defined herein, including all embodiments thereof.
- R 60.1 and R 60.2 are independently as defined for R 60 . In embodiments, R 60.1 and R 60.2 are different.
- R 13 is hydrogen.
- R 3 is methoxy (—O—CH 3 ).
- R 5 is halogen (e.g. F).
- R 1C is substituted or unsubstituted alkyl (e.g. methyl).
- R 60.1 is halogen (e.g. F).
- R 60.2 is nitro (—NO 2 ).
- R 4 is substituted or unsubstituted heterocycloalkyl (e.g. unsubstituted morphilino or unsubstituted piperidinyl). In embodiments, R 4 is substituted or unsubstituted heterocycloalkyl wherein the point of attachment to the remainder of the molecule is a ring nitrogen.
- the PLK4 inhibitor is the compound of Formula (II) or a pharmaceutically acceptable salt thereof:
- L 2 is independently a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 14 —, —C(O)NR 14 —, —NR 14 C(O)—, —S(O)—, —S(O) 2 —, —S(O)NR 14 —, substituted or unsubstituted alkylene, substituted or unsubstituted heteroalkylene, substituted or unsubstituted cycloalkylene, substituted or unsubstituted heterocycloalkylene, substituted or unsubstituted arylene, or substituted or unsubstituted heteroarylene.
- R 7 is independently hydrogen, oxo, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 7A , —OR 7A , —NR 7A R 7B , —C(O)OR 7A , —C(O)NR 7A R 7B , —NO 2 , —SR 7A , —S(O) n7 R 7A , —S(O) 7 OR 7A , —S(O) n7 NR 7A R 7B , —NHNR 7A R 7B , —ONR 7A R 7B , —NHC(O)NHNR 7A R 7B , -L 1 -R 6 , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or un
- R 7A , R 7B , R 13 , and R 14 are independently hydrogen, oxo, halogen, —CF 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted
- L 1 , R 1 , R 2 , and R 6 of the compound of formula (II) are as described hereinabove for compounds having formula (I), including all embodiments thereof.
- R 3 of the compound of formula (II) is as described hereinabove for compounds of formula (I) with the proviso that R 3 is not hydrogen.
- R 3 may be halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3A , —OR 3A , —NR 3A R 3B , —C(O)OR 3A , —C(O)NR 3A R 3B , —NO 2 , —SR 3A , —S(O) n3 R 3A , —S(O) n3 OR 3A , —S(O) n3 NR 3A R 3B , —NHNR 3A R 3B , —ONR 3A R 3B , —NHC(O)NHNR 3A R 3B , or substituted or unsubstituted alkyl, where R 3A , R 3B
- R 3 may be halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3A , —OR 3A , —NR 3A R 3B , —C(O)OR 3A , —C(O)NR 3A R 3B , —NO 2 , —SR 3A , —S(O) n3 R 3A , —S(O) n3 OR 3A , —S(O) n3 NR 3A R 3B , —NR 3A R 3B , —ONR 3A R 3B , —NHC(O)NHNR 3A R 3B , or substituted or unsubstituted alkyl, where R 3A , R 3B , and R 3C are independently hydrogen, oxo, halogen, —CF 3 , —OH, —NH 2 , —COOH,
- R 3 may be halogen, —OR 3A , or substituted or unsubstituted alkyl.
- R 3 may be halogen, —OR 3A , or substituted or unsubstituted alkyl where f substituted or unsubstituted alkyl.
- R 3 may be —OR 3A .
- R 3 may be —OR 3A where R 3A is substituted or unsubstituted alkyl
- R 4 of the compound of formula (II) is as described hereinabove for compounds of formula (I) with the proviso that R 4 is not hydrogen or substituted or unsubstituted alkyl.
- the compound of formula (II) may be compound having one of the formulae Ia, Ia1, Ia2, Ia3, Ia4, Ia5, Ia6, Ia7, Ia8, Ib, Ib1, Ib2, Ib3, Ib4, Ib5, Ib6, or Ib7.
- R 7 may independently be hydrogen, halogen, —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 7A , —OR 7A , —NR 7A R 7B , —C(O)OR 7A , —C(O)NR 7A R 7B , —NO 2 , —SR 7A , —S(O) n7 R 7A , —S(O) n7 OR 7A , —S(O) n7 NR 7A R 7B , —NHNR 7A R 7B , —ONR 7A R 7B , or —NHC(O)NHNR 7A R 7B .
- R 7 may independently be substituted or unsubstituted alkyl, substituted or unsubstituted heteroalkyl, substituted or unsubstituted cycloalkyl, substituted or unsubstituted heterocycloalkyl, substituted or unsubstituted aryl, or substituted or unsubstituted heteroaryl.
- R 7 may be hydrogen or halogen.
- R 7 may be hydrogen or —Cl, —I, or —Br.
- R 7 may be hydrogen or —Cl or —F.
- R 7 may be hydrogen.
- R 7 may be —Cl.
- R 7 may be —I.
- R 7 may be —I.
- R 7 may be —F.
- the symbol z1 may be 1, 2, or 3.
- the symbol z1 may be 1 or 2.
- R 7 may be hydrogen or —Cl or —F where the symbol z1 is 1 or 2.
- R 7 may be substituted or unsubstituted alkyl.
- R 7 may be substituted alkyl.
- R 7 may be unsubstituted alkyl.
- R 7 may be substituted or unsubstituted C 1 -C 20 alkyl.
- R 7 may be substituted C 1 -C 20 alkyl.
- R 7 may be substituted or unsubstituted C 1 -C 10 alkyl.
- R 7 may be substituted C 1 -C 10 alkyl.
- R 7 may be unsubstituted C 1 -C 10 alkyl.
- R 7 may be substituted or unsubstituted C 1 -C 5 alkyl.
- R 7 may be unsubstituted C 1 -C 5 alkyl.
- R 7 may be substituted C 1 -C 5 alkyl.
- R 7 may be methyl, substituted or unsubstituted ethyl, or substituted or unsubstituted propyl.
- R 7 may be methyl.
- R 7 may be ethyl.
- R 7 may be R 7C -substituted or unsubstituted alkyl.
- R 7 may be R 7C -substituted alkyl.
- R 7 may be R 7C -substituted or unsubstituted C 1 -C 20 alkyl.
- R 7 may be R 7C -substituted C 1 -C 20 alkyl.
- R 7 may be R 7C -substituted or unsubstituted C 1 -C 10 alkyl.
- R 7 may be R 7C -substituted C 1 -C 10 alkyl.
- R 7 may be R 7C -substituted or unsubstituted C 1 -C 5 alkyl.
- R 7 may be R 7C -substituted C 1 -C 5 alkyl.
- R 7 may be methyl, R 7C -substituted or unsubstituted ethyl, or R 7C -substituted or unsubstituted propyl.
- R 7 may be substituted or unsubstituted heteroalkyl.
- R 7 may be substituted heteroalkyl.
- R 7 may be unsubstituted heteroalkyl.
- R 7 may be substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 7 may be substituted 2 to 20 membered heteroalkyl.
- R 7 may be substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 7 may be substituted 2 to 10 membered heteroalkyl.
- R 7 may be substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 7 may be substituted 2 to 6 membered heteroalkyl.
- R 7 may be R 7C -substituted or unsubstituted heteroalkyl.
- R 7 may be R 7C -substituted heteroalkyl.
- R 7 may be R 7C -substituted or unsubstituted 2 to 20 membered heteroalkyl.
- R 7 may be R 7C -substituted 2 to 20 membered heteroalkyl.
- R 7 may be R 7C -substituted or unsubstituted 2 to 10 membered heteroalkyl.
- R 7 may be R 7C -substituted 2 to 10 membered heteroalkyl.
- R 7 may be R 7C -substituted or unsubstituted 2 to 6 membered heteroalkyl.
- R 7 may be R 7C -substituted 2 to 6 membered heteroalkyl.
- R 7 may be substituted or unsubstituted cycloalkyl.
- R 7 may be substituted cycloalkyl.
- R 7 may be unsubstituted cycloalkyl.
- R 7 may be substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 7 may be substituted 3 to 20 membered cycloalkyl.
- R 7 may be substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 7 may be substituted 3 to 10 membered cycloalkyl.
- R 7 may be substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 7 may be substituted 3 to 6 membered cycloalkyl.
- R 7 may be R 7C -substituted or unsubstituted cycloalkyl.
- R 7 may be R 7C -substituted cycloalkyl.
- R 7 may be R 7C -substituted or unsubstituted 3 to 20 membered cycloalkyl.
- R 7 may be R 7C -substituted 3 to 20 membered cycloalkyl.
- R 7 may be R 7C -substituted or unsubstituted 3 to 10 membered cycloalkyl.
- R 7 may be R 7C -substituted 3 to 10 membered cycloalkyl.
- R 7 may be R 7C -substituted or unsubstituted 3 to 6 membered cycloalkyl.
- R 7 may be R 7C -substituted 3 to 6 membered cycloalkyl.
- R 7 may be substituted or unsubstituted heterocycloalkyl.
- R 7 may be substituted heterocycloalkyl.
- R 7 may be unsubstituted heterocycloalkyl.
- R 7 may be substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 7 may be substituted 3 to 20 membered heterocycloalkyl.
- R 7 may be substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 7 may be substituted 3 to 10 membered heterocycloalkyl.
- R 7 may be substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 7 may be substituted 3 to 6 membered heterocycloalkyl.
- R 7 may be R 7C -substituted or unsubstituted heterocycloalkyl.
- R 7 may be R 7C -substituted heterocycloalkyl.
- R 7 may be R 7C -substituted or unsubstituted 3 to 20 membered heterocycloalkyl.
- R 7 may be R 7C -substituted 3 to 20 membered heterocycloalkyl.
- R 7 may be R 7C -substituted or unsubstituted 3 to 10 membered heterocycloalkyl.
- R 7 may be R 7C -substituted 3 to 10 membered heterocycloalkyl.
- R 7 may be R 7C -substituted or unsubstituted 3 to 6 membered heterocycloalkyl.
- R 7 may be R 7C -substituted 3 to 6 membered heterocycloalkyl.
- R 7 may be substituted or unsubstituted aryl.
- R 7 may be substituted aryl.
- R 7 may be unsubstituted aryl.
- R 7 may be substituted or unsubstituted 5 to 20 membered aryl.
- R 7 may be substituted 5 to 20 membered aryl.
- R 7 may be substituted or unsubstituted 5 to 8 membered aryl (e.g. phenyl).
- R 7 may be substituted 5 to 8 membered aryl.
- R 7 may be substituted or unsubstituted 5 or 6 membered aryl.
- R 7 may be substituted 5 or 6 membered aryl.
- R 7 may be R 7C -substituted or unsubstituted aryl.
- R 7 may be R 7C -substituted aryl.
- R 7 may be R 7C -substituted or unsubstituted 5 to 20 membered aryl.
- R 7 may be R 7C -substituted 5 to 20 membered aryl.
- R 7 may be R 7C -substituted or unsubstituted 5 to 8 membered aryl.
- R 7 may be R 7C -substituted 5 to 8 membered aryl.
- R 7 may be R 7C -substituted or unsubstituted 5 or 6 membered aryl.
- R 7 may be R 7C -substituted 5 or 6 membered aryl (e.g. phenyl).
- R 7 may be substituted or unsubstituted heteroaryl.
- R 7 may be substituted heteroaryl.
- R 7 may be unsubstituted heteroaryl.
- R 7 may be substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 7 may be substituted 5 to 20 membered heteroaryl.
- R 7 may be substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 7 may be substituted 5 to 8 membered heteroaryl.
- R 7 may be substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 7 may be substituted 5 or 6 membered heteroaryl.
- R 7 may be R 7C -substituted or unsubstituted heteroaryl.
- R 7 may be R 7C -substituted heteroaryl.
- R 7 may be R 7C -substituted or unsubstituted 5 to 20 membered heteroaryl.
- R 7 may be R 7C -substituted 5 to 20 membered heteroaryl.
- R 7 may be R 7C -substituted or unsubstituted 5 to 8 membered heteroaryl.
- R 7 may be R 7C -substituted 5 to 8 membered heteroaryl.
- R 7 may be R 7C -substituted or unsubstituted 5 or 6 membered heteroaryl.
- R 7 may be R 7C -substituted 5 or 6 membered heteroaryl.
- R 7A is independently hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 7C -substituted or unsubstituted alkyl, R 7C -substituted or unsubstituted heteroalkyl, R 7C
- R 7C is independently halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COR 7D , —NR 7D R 7E , —COOR 7D , —CONR 7D R 7E , —NO 2 , —SR 7D , —S(O) 2 R 7D , —S(O) 3 R 7D , —S(O) 4 R 7D , —S(O) 2 NR 7D R 7E , —NHNR 7D R 7E , —ONR 7D R 7E , —NHC(O)NHNR 7D R 7E , —NHC(O)NR 7D R 7E , —NHS(O)NR 7D R 7E , —NHS(O) 2 R 7D , —NHC(O)R 7D , —NHC(O
- R 7C may independently be halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —C(O)H, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 7D -substituted or unsubstituted alkyl, R 7D -substituted or un
- R 7B and R 7E may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cyclo
- R 7D is independently hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COR 7F , —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 7F -substituted or unsubstituted heteroalkyl, R 7F -
- R 7F is independently halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubsti
- R 7 may be -L 1 -R 6 , where L 1 and R 6 are independently as described herein.
- R 7 may be -L 1 -R 6 , where L 1 and R 6 are independently as described herein and L 2 is a bond, —SO 2 , or —NHC(O)—.
- L 2 may independently be a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 14 —, —C(O)NR 14 —, —NR 14 C(O)—, —S(O)—, —S(O) 2 —, —S(O)NR 14 —.
- L 2 may independently be a bond, —S(O) 2 —, or —NR 14 C(O)—, where R 14 is hydrogen or unsubstituted C 1 -C 5 alkyl.
- L 2 may independently be a bond, substituted or unsubstituted alkylene, substituted or unsubstituted heteroalkylene, substituted or unsubstituted cycloalkylene, substituted or unsubstituted heterocycloalkylene, substituted or unsubstituted arylene, or substituted or unsubstituted heteroarylene.
- L 2 may independently be a bond, substituted or unsubstituted alkylene, or substituted or unsubstituted heteroalkylene.
- L 2 may independently be a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 14 —, —C(O)NR 14 —, —NR 14 C(O)—, —S(O) 2 —, —S(O)NR 14 —, substituted or unsubstituted alkylene, or substituted or unsubstituted heteroalkylene.
- L 2 may independently be a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 14 —, —C(O)NR 14 —, —NR 14 C(O)—, —S(O) 2 —, —S(O)NR 14 —, substituted or unsubstituted alkylene, or substituted or unsubstituted heteroalkylene where R 14 is hydrogen or substituted or unsubstituted alkyl.
- L 2 may independently be a bond, —C(O)—, —C(O)O—, —O—, —S—, —NR 14 —, —C(O)NR 14 —, —NR 14 C(O)—, —S(O) 2 —, —S(O)NR 14 —, R 14 -substituted or unsubstituted alkylene, or R 14 -substituted or unsubstituted heteroalkylene where R 14 is hydrogen or substituted or unsubstituted alkyl. R 14 may be unsubstituted alkyl.
- L 2 may independently be a bond or substituted or unsubstituted alkylene.
- L 2 may independently be a bond or R 14 -substituted or unsubstituted alkylene.
- L 2 may independently be a bond or R 14 -substituted or unsubstituted alkylene where R 14 is hydrogen or substituted or unsubstituted alkyl.
- L 2 may independently be a bond or substituted or unsubstituted C 1 -C 5 alkylene.
- L 2 may independently be a bond or R 14 -substituted or unsubstituted C 1 -C 5 alkylene.
- L 2 may independently be a bond or R 14 -substituted or unsubstituted C 1 -C 5 alkylene where R 14 is hydrogen or substituted or unsubstituted C 1 -C 5 alkyl.
- L 2 may independently be R 14 -substituted or unsubstituted alkylene where R 14 is hydrogen, halogen, or substituted or unsubstituted alkyl.
- L 2 may independently be a bond.
- L 2 may independently be bond, —SO 2 , or —NHC(O)— and R 7 may be L 1 -R 6 , where L 1 and R 6 are as described herein.
- R 14 may independently be hydrogen, halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —COH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 Cl, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , R 14A -substituted or unsubstituted heteroalkyl, R 14A -substit
- R 14A is independently halogen, oxo, N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —OH, —NH 2 , —COOH, —CONH 2 , —COH, —COCH 3 , —NO 2 , —SH, —S(O) 2 C, —S(O) 3 H, —S(O) 4 H, —S(O) 2 NH 2 , —NHNH 2 , —ONH 2 , —NHC(O)NHNH 2 , —NHC(O)NH 2 , —NHS(O) 2 H, —NHC(O)H, —NHC(O)—OH, —NHOH, —OCF 3 , —OCHF 2 , unsubstituted alkyl, unsubstituted heteroalkyl, unsubstituted cycloalkyl, unsubsti
- the compound of Formula (II) is:
- the compound of Formula (II) is:
- z2a is 0, 1, 2, 3, or 4, and the compound is not tozasertib.
- z2a may be 0, 1, or 2.
- z2a may be 0.
- z2a may be 1.
- z2a may be 2.
- z2a may be 3.
- z2a may be 4.
- R 1 , R 2 , and R 4 are as described hereinabove for compounds of formula (I).
- L 2 , R 7 , and z2 are as described hereinabove for first aspect of the compound having formula (II).
- R 7 may be -L 1 -R 6 , where L 1 and R 6 are independently as described herein. The compound is not tozasertib.
- R 3 of the second aspect of the compound of formula (II) is as described hereinabove for compounds of formula (I) with the proviso that R 3 is not hydrogen, halogen, or substituted or unsubstituted alkyl.
- R 3 may be —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3A , —OR 3A , —NR 3A R 3B , —C(O)OR 3A , —C(O)NR 3A R 3B , —NO 2 , —SR 3A , —S(O) n3 R 3A , —S(O) n3 OR 3A , —S(O) n3 NR 3A R 3B , —NHNR 3A R 3B , —ONR 3A R 3B , —NHC(O)NHNR 3A R 3B , where R 3A
- R 3 may be —N 3 , —CF 3 , —CCl 3 , —CBr 3 , —CI 3 , —CN, —COR 3A , —OR 3A , —NR 3A R 3B , —C(O)OR 3A , —C(O)NR 3A R 3B , —NO 2 , —SR 3A , —S(O) n3 R 3A , —S(O) n3 OR 3A , —S(O) n3 NR 3A R 3B , —NHNR 3A R 3B , —ONR 3A R 3B , —NHC(O)NHNR 3A R 3B , where R 3A , R 3B , R 3C , are independently hydrogen, oxo, halogen, —CF 3 , —OH, —NH 2 , —COOH, —CONH 2 , substituted or unsubstituted
- R 3 may be —OR 3A .
- R 3 may be —OR 3A where R 3A is substituted or unsubstituted alkyl.
- the compound of Formula (II) is:
- the compound of Formula (II) is:
- z2a is 0, 1, 2, 3, or 4, and the compound is not tozasertib.
- z2a may be 0, 1, or 2.
- z2a may be 0.
- z2a may be 1.
- z2a may be 2.
- z2a may be 3.
- z2a may be 4.
- the PLK4 inhibitor is the compound of Formula (III) or a pharmaceutically acceptable salt thereof.
- R 1 , R 2 , R 3 , and R 4 are as described hereinabove for formula (I). In embodiments, R 3 , and R 4 are as set forth hereinabove for formula (II) (including the first and second aspects).
- R 8 is unsubstituted C 1 -C 5 alkyl. R 8 may be methyl, ethyl, or propyl. R 8 may be methyl.
- the PLK4 inhibitor is a compound set forth in Table 1. In embodiments, the PLK4 inhibitor is a compound set forth in Table 1 having activity indicated as “XXX.” In embodiments, the PLK4 inhibitor is a compound set forth in Table 1 having the formula (I). In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (I) and having activity indicated as “XXX.” In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (I) and having activity indicated as “XX”. In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (I) and having activity indicated as “X”.
- the PLK4 inhibitor is a compound set forth in Table 1, having the formula (II) (including the first or second aspect). In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (II) (including the first or second aspect) and having activity indicated as “XXX”. In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (II) (including the first or second aspect) and having activity indicated as “XX”. In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (II) (including the first or second aspect) and having activity indicated as “X”. In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (III).
- the PLK4 inhibitor is a compound set forth in Table 1, having the formula (III) and having activity indicated as “XXX”. In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (III) and having activity indicated as “XX”. In embodiments, the PLK4 inhibitor is a compound set forth in Table 1, having the formula (III) and having activity indicated as “X”.
- the PLK4 inhibitor is centrinone or a pharmaceutically acceptable salt thereof.
- Centrinone has PubChem CID 91801159.
- the PLK4 inhibitor is centrinone B or a pharmaceutically acceptable salt thereof.
- Centrinone B has PubChem CID 118704753.
- the compound is not a compound set forth in U.S. Pat. No. 8,455,507, which is herein incorporated by reference in its entirety. In embodiments the compound is not a compound set forth in U.S. Pat. No. 7,531,536 which is herein incorporated by reference in its entirety. In embodiments the compound is not a compound set forth in U.S. Pat. No. 7,951,820 which is herein incorporated by reference in its entirety.
- Table 1 exemplifies compounds of Formula (I), (II), and (III).
- Activity denoted by X indicates the compound has IC 50 value greater than about 1 ⁇ M.
- Activity denoted by XX indicates the compound has IC 50 value of about 100 nM to about 1 ⁇ M.
- Activity denoted by XXX indicates compound has IC 50 value less than about 100 nM. All compounds were confirmed by mass spectroscopy and by 1 H and 13 C NMR.
- the compounds described herein, including the compounds in Table 1 and Table 2 may be synthesized by methods known in the art.
- the compounds described herein, including the compounds in Table 1 and Table 2 are synthesized by the methods set forth in WO 2016/166604 published on 20 Oct. 2016, the disclosure of which is incorporated by reference herein in its entirety.
- compositions of the compounds described herein are pharmaceutical compositions of the compounds described herein.
- a pharmaceutical composition that includes a pharmaceutically acceptable excipient and a compound (e.g. formula (I), (II) (including the first or second aspect), (III), or a compound of Table 1 or Table 2) as described herein.
- the compound may be a compound having the formula (I) as described herein.
- the compound may be a compound having the formula (II) (including the first or second aspect) as described herein.
- the compound may be a compound having the formula (III) as described herein.
- the compound may be a compound set forth in Table 1.
- the compound may be a compound set forth in Table 2.
- the pharmaceutical composition may include a second agent in a therapeutically effective amount.
- the pharmaceutical composition may include a second agent where the second agent treats cancer.
- the second agent may be an anti-cancer agent as described herein.
- the pharmaceutical composition may be prepared and administered in a wide variety of dosage formulations.
- Compounds described herein e.g. formula (I), (II) (including the first or second aspect), (III) or a compound of Table 1 or Table 2) may be administered orally, rectally, or by injection (e.g. intravenously, intramuscularly, intracutaneously, subcutaneously, intraduodenally, or intraperitoneally).
- pharmaceutically acceptable carriers can be either solid or liquid.
- Solid form preparations include powders, tablets, pills, capsules, cachets, suppositories, and dispersible granules.
- a solid carrier may be one or more substance that may also act as diluents, flavoring agents, binders, preservatives, tablet disintegrating agents, or an encapsulating material.
- the carrier may be a finely divided solid in a mixture with the finely divided active component.
- the active component may be mixed with the carrier having the necessary binding properties in suitable proportions and compacted in the shape and size desired.
- the powders and tablets preferably contain from 5% to 70% of the active compound.
- Suitable carriers are magnesium carbonate, magnesium stearate, talc, sugar, lactose, pectin, dextrin, starch, gelatin, tragacanth, methylcellulose, sodium carboxymethylcellulose, a low melting wax, cocoa butter, and the like.
- preparation is intended to include the formulation of the active compound with encapsulating material as a carrier providing a capsule in which the active component with or without other carriers, is surrounded by a carrier, which is thus in association with it.
- carrier providing a capsule in which the active component with or without other carriers, is surrounded by a carrier, which is thus in association with it.
- cachets and lozenges are included. Tablets, powders, capsules, pills, cachets, and lozenges can be used as solid dosage forms suitable for oral administration.
- a low melting wax such as a mixture of fatty acid glycerides or cocoa butter
- the active component is dispersed homogeneously therein, as by stirring.
- the molten homogeneous mixture is then poured into convenient sized molds, allowed to cool, and thereby to solidify.
- Liquid form preparations include solutions, suspensions, and emulsions, for example, water or water/propylene glycol solutions.
- liquid preparations can be formulated in solution in aqueous polyethylene glycol solution.
- Aqueous solutions suitable for oral use can be prepared by dissolving the active component in water and adding suitable colorants, flavors, stabilizers, and thickening agents as desired.
- Aqueous suspensions suitable for oral use can be made by dispersing the finely divided active component in water with viscous material, such as natural or synthetic gums, resins, methylcellulose, sodium carboxymethylcellulose, and other well-known suspending agents.
- solid form preparations that are intended to be converted, shortly before use, to liquid form preparations for oral administration.
- liquid forms include solutions, suspensions, and emulsions.
- These preparations may contain, in addition to the active component, colorants, flavors, stabilizers, buffers, artificial and natural sweeteners, dispersants, thickeners, solubilizing agents, and the like.
- the pharmaceutical preparation is preferably in unit dosage form.
- the preparation is subdivided into unit doses containing appropriate quantities of the active component.
- the unit dosage form can be a packaged preparation, the package containing discrete quantities of preparation, such as packeted tablets, capsules, and powders in vials or ampoules.
- the unit dosage form can be a capsule, tablet, cachet, or lozenge itself, or it can be the appropriate number of any of these in packaged form.
- the quantity of active component in a unit dose preparation may be varied or adjusted from 0.1 mg to 10000 mg according to the particular application and the potency of the active component.
- the composition can, if desired, also contain other compatible therapeutic agents.
- Some compounds may have limited solubility in water and therefore may require a surfactant or other appropriate co-solvent in the composition.
- co-solvents include: Polysorbate 20, 60, and 80; Pluronic F-68, F-84, and P-103; cyclodextrin; and polyoxyl 35 castor oil.
- co-solvents are typically employed at a level between about 0.01% and about 2% by weight. Viscosity greater than that of simple aqueous solutions may be desirable to decrease variability in dispensing the formulations, to decrease physical separation of components of a suspension or emulsion of formulation, and/or otherwise to improve the formulation.
- Such viscosity building agents include, for example, polyvinyl alcohol, polyvinyl pyrrolidone, methyl cellulose, hydroxy propyl methylcellulose, hydroxyethyl cellulose, carboxymethyl cellulose, hydroxy propyl cellulose, chondroitin sulfate and salts thereof, hyaluronic acid and salts thereof, and combinations of the foregoing.
- Such agents are typically employed at a level between about 0.01% and about 2% by weight.
- the pharmaceutical compositions may additionally include components to provide sustained release and/or comfort.
- Such components include high molecular weight, anionic mucomimetic polymers, gelling polysaccharides, and finely-divided drug carrier substrates. These components are discussed in greater detail in U.S. Pat. Nos. 4,911,920; 5,403,841; 5,212,162; and 4,861,760. The entire contents of these patents are incorporated herein by reference in their entirety for all purposes.
- the pharmaceutical composition may be intended for intravenous use.
- the pharmaceutically acceptable excipient can include buffers to adjust the pH to a desirable range for intravenous use.
- buffers including salts of inorganic acids such as phosphate, borate, and sulfate are known.
- the pharmaceutical composition may include compositions wherein the active ingredient is contained in a therapeutically effective amount, i.e., in an amount effective to achieve its intended purpose.
- a therapeutically effective amount i.e., in an amount effective to achieve its intended purpose.
- the actual amount effective for a particular application will depend, inter alia, on the condition being treated.
- the dosage and frequency (single or multiple doses) of compounds administered can vary depending upon a variety of factors, including route of administration; size, age, sex, health, body weight, body mass index, and diet of the recipient; nature and extent of symptoms of the disease being treated; presence of other diseases or other health-related problems; kind of concurrent treatment; and complications from any disease or treatment regimen.
- Other therapeutic regimens or agents can be used in conjunction with the methods and compounds disclosed herein.
- the therapeutically effective amounts can be initially determined from cell culture assays.
- Target concentrations will be those concentrations of active compound(s) that are capable of increasing the extent of cancer cell death as measured, for example, using methods known in the art.
- Therapeutically effective amounts for use in humans may be determined from animal models. For example, a dose for humans can be formulated to achieve a concentration that has been found to be effective in animals. The dosage in humans can be adjusted by monitoring response of the cancer to the treatment and adjusting the dosage upwards or downwards, as described above.
- Dosages may be varied depending upon the requirements of the subject and the compound being employed.
- the dose administered to a subject should be sufficient to effect a beneficial therapeutic response in the subject over time.
- the size of the dose also will be determined by the existence, nature, and extent of any adverse side effects. Generally, treatment is initiated with smaller dosages, which are less than the optimum dose of the compound. Thereafter, the dosage is increased by small increments until the optimum effect under circumstances is reached.
- the disclosure provides methods of treating cancer in a subject in need thereof, the method comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the cancer; wherein the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin.
- the subject has an elevated level of TRIM37.
- the subject has an elevated level of Chromogranin A.
- the subject has an elevated level of Synaptophysin.
- the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A.
- the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin.
- the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin.
- the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level.
- the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin, when compared to a control.
- the subject has an elevated level of TRIM37 when compared to a control.
- the subject has an elevated level of Chromogranin A when compared to a control.
- the subject has an elevated level of Synaptophysin when compared to a control.
- the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level when compared to a control. In embodiments, the methods are for treating a cancer tumor.
- the PLK4 inhibitor is compound of Formula (Ia), a compound of Formula (Ib), a compound of Formula (Ia1), a compound of Formula (Ia2), a compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), a compound of Formula (Ib7), a compound of Formula (IC), a compound of Formula (II), a compound of Formula (III), a
- the PLK4 inhibitor is compound set forth in Table 1. In embodiments, the PLK4 inhibitor is compound set forth in Table 2. In embodiments, the PLK4 inhibitor is centrinone. In embodiments, the PLK4 inhibitor is centrinone B.
- the cancer is a cancer tumor, such as a primary tumor or a metastatic cancer tumor. In embodiments, the cancer is a pediatric cancer. In embodiments, the pediatric cancer is a rhabdoid tumor, a neuroblastoma, an acute lymphoblastic leukemia tumor, or a brain and central nervous system tumor. In embodiments, the cancer is a neural crest-derived cancer.
- the neural crest-derived tumor is a small cell lung cancer, a melanoma, or a breast cancer.
- the cancer is neuroblastoma.
- the cancer is acute myeloid leukemia, prostate cancer, glioma, mesothelioma, osteosarcoma, breast cancer, Ewing's sarcoma, soft tissue cancer, or T cell lymphoma.
- the T cell lymphoma is extranodal T cell lymphoma, cutaneous T cell lymphoma, anaplastic large cell lymphoma, or angioimmunoblastic T cell lymphoma.
- the cancer is a rhabdoid tumor, basal cell carcinoma, small cell lung cancer, endometrial cancer, non-small cell lung cancer, ovarian cancer, thyroid cancer, kidney cancer, Hodgkin's lymphoma, stomach cancer, liver cancer, Burkitt lymphoma, giant cell tumor of bone, medulloblastoma, a urinary tract cancer, meningioma, bile duct cancer, melanoma, esophageal cancer, upper aerodigestive cancer, colorectal cancer, chondrosarcoma, multiple myeloma, B cell lymphoma, leukemia, diffuse large B cell lymphoma, or chronic myeloid leukemia.
- the cancer is a p53 positive cancer, including a wild type p53 positive cancer or a mutant p53 positive cancer.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- the chemotherapeutic agent comprises a tubule polymerization inhibitor, such as vincristine, vinblastine, vinoreIbine, vinfluine, a dolastatin, a halichondrin, a hemiasterline, cryptophysin 52, or a combination of two or more thereof.
- the chemotherapeutic agent comprises cyclophosphamide, ifosfamide, cisplatin, carboplatin, vincristine, doxorubicin, etoposide, topotecan, busulfan, melphalan, or a combination of two or more thereof.
- the chemotherapeutic agent comprises differentiation therapy, wherein the chemotherapeutic agent is retinoic acid (e.g., all-trans-retinoic acid, 13-cis-retinoic acid), arsenic trioxide, or a combination thereof.
- the disclosure provides methods of treating a p53 positive cancer in a subject in need thereof, the method comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the p53 positive cancer.
- the methods comprise treating a wild type p53 positive cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type p53 positive cancer.
- the methods comprise treating a mutant p53 positive cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive cancer.
- the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin, when compared to a control.
- the subject has an elevated level of TRIM37 when compared to a control. In embodiments, the subject has an elevated level of Chromogranin A when compared to a control. In embodiments, the subject has an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level when compared to a control.
- the methods are for treating a cancer tumor.
- the PLK4 inhibitor is compound of Formula (Ia), a compound of Formula (Ib), a compound of Formula (Ia1), a compound of Formula (Ia2), a compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound
- the PLK4 inhibitor is compound set forth in Table 1. In embodiments, the PLK4 inhibitor is compound set forth in Table 2. In embodiments, the PLK4 inhibitor is centrinone. In embodiments, the PLK4 inhibitor is centrinone B.
- the cancer is a cancer tumor, such as a primary tumor or a metastatic cancer tumor. In embodiments, the cancer is a pediatric cancer. In embodiments, the pediatric cancer is a rhabdoid tumor, a neuroblastoma, an acute lymphoblastic leukemia tumor, or a brain and central nervous system tumor. In embodiments, the cancer is a neural crest-derived cancer.
- the neural crest-derived tumor is a small cell lung cancer, a melanoma, or a breast cancer.
- the cancer is neuroblastoma.
- the cancer is acute myeloid leukemia, prostate cancer, glioma, mesothelioma, osteosarcoma, breast cancer, Ewing's sarcoma, soft tissue cancer, or T cell lymphoma.
- the T cell lymphoma is extranodal T cell lymphoma, cutaneous T cell lymphoma, anaplastic large cell lymphoma, or angioimmunoblastic T cell lymphoma.
- the cancer is a rhabdoid tumor, basal cell carcinoma, small cell lung cancer, endometrial cancer, non-small cell lung cancer, ovarian cancer, thyroid cancer, kidney cancer, Hodgkin's lymphoma, stomach cancer, liver cancer, Burkitt lymphoma, giant cell tumor of bone, medulloblastoma, a urinary tract cancer, meningioma, bile duct cancer, melanoma, esophageal cancer, upper aerodigestive cancer, colorectal cancer, chondrosarcoma, multiple myeloma, B cell lymphoma, leukemia, diffuse large B cell lymphoma, or chronic myeloid leukemia.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- the chemotherapeutic agent comprises a tubule polymerization inhibitor, such as vincristine, vinblastine, vinoreIbine, vinfluine, a dolastatin, a halichondrin, a hemiasterline, cryptophysin 52, or a combination of two or more thereof.
- the chemotherapeutic agent comprises cyclophosphamide, ifosfamide, cisplatin, carboplatin, vincristine, doxorubicin, etoposide, topotecan, busulfan, melphalan, or a combination of two or more thereof.
- the chemotherapeutic agent comprises differentiation therapy, wherein the chemotherapeutic agent is retinoic acid (e.g., all-trans-retinoic acid, 13-cis-retinoic acid), arsenic trioxide, or a combination thereof.
- retinoic acid e.g., all-trans-retinoic acid, 13-cis-retinoic acid
- arsenic trioxide or a combination thereof.
- the disclosure provides methods of treating neuroblastoma in a subject in need thereof, the method comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the neuroblastoma; wherein the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin.
- the disclosure provides methods of treating a wild type or mutant p53 positive neuroblastoma in a subject in need thereof, the method comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type or mutant p53 positive neuroblastoma.
- the disclosure provides methods of treating a wild type or mutant p53 positive neuroblastoma in a subject in need thereof, the method comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type or mutant p53 positive neuroblastoma; wherein the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin.
- the subject has an elevated level of TRIM37.
- the subject has an elevated level of Chromogranin A.
- the subject has an elevated level of Synaptophysin.
- the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A.
- the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin. In embodiments, the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin. In embodiments, the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level. In embodiments, the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of TRIM37 when compared to a control. In embodiments, the subject has an elevated level of Chromogranin A when compared to a control.
- the subject has an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level when compared to a control.
- the PLK4 inhibitor is compound of Formula (Ia), a compound of Formula (Ib), a compound of Formula (Ia1), a compound of Formula (Ia2), a compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), a compound of Formula (Ib7), a compound of Formula (IC), a compound of Formula (II), a compound of Formula (III), a
- the PLK4 inhibitor is compound set forth in Table 1. In embodiments, the PLK4 inhibitor is compound set forth in Table 2. In embodiments, the PLK4 inhibitor is centrinone. In embodiments, the PLK4 inhibitor is centrinone B. In embodiments, the subject is a human. In embodiments, the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- the chemotherapeutic agent comprises a tubule polymerization inhibitor, such as vincristine, vinblastine, vinoreIbine, vinfluine, a dolastatin, a halichondrin, a hemiasterline, cryptophysin 52, or a combination of two or more thereof.
- the chemotherapeutic agent comprises cyclophosphamide, ifosfamide, cisplatin, carboplatin, vincristine, doxorubicin, etoposide, topotecan, busulfan, melphalan, or a combination of two or more thereof.
- the chemotherapeutic agent comprises differentiation therapy, wherein the chemotherapeutic agent is retinoic acid (e.g., all-trans-retinoic acid, 13-cis-retinoic acid), arsenic trioxide, or a combination thereof.
- the disclosure provides methods of treating small cell lung cancer, neuroblastoma, prostate cancer, glioma, mesothelioma, osteosarcoma, breast cancer, Ewing's sarcoma, soft tissue cancer, or T-Cell lymphoma in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor; wherein the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin.
- the disclosure provides methods of treating a wild type or mutant p53 positive small cell lung cancer, a wild type or mutant p53 positive neuroblastoma, a wild type or mutant p53 positive prostate cancer, a wild type or mutant p53 positive glioma, a wild type or mutant p53 positive mesothelioma, a wild type or mutant p53 positive osteosarcoma, a wild type or mutant p53 positive breast cancer, a wild type or mutant p53 positive Ewing's sarcoma, a wild type or mutant p53 positive soft tissue cancer, or a wild type or mutant p53 positive T-Cell lymphoma in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor.
- the disclosure provides methods of treating a wild type or mutant p53 positive small cell lung cancer, a wild type or mutant p53 positive neuroblastoma, a wild type or mutant p53 positive prostate cancer, a wild type or mutant p53 positive glioma, a wild type or mutant p53 positive mesothelioma, a wild type or mutant p53 positive osteosarcoma, a wild type or mutant p53 positive breast cancer, a wild type or mutant p53 positive Ewing's sarcoma, a wild type or mutant p53 positive soft tissue cancer, or a wild type or mutant p53 positive T-Cell lymphoma in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor; wherein the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin.
- the subject has an elevated level of TRIM37. In embodiments, the subject has an elevated level of Chromogranin A. In embodiments, the subject has an elevated level of Synaptophysin. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin. In embodiments, the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin. In embodiments, the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level.
- the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin, when compared to a control. In embodiments, the subject has an elevated level of TRIM37 when compared to a control. In embodiments, the subject has an elevated level of Chromogranin A when compared to a control. In embodiments, the subject has an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin when compared to a control.
- the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level when compared to a control.
- the PLK4 inhibitor is compound of Formula (Ia), a compound of Formula (Ib), a 5 compound of Formula (Ia1), a compound of Formula (Ia2), a compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), a compound of Formula (Ib7), a compound of Formula (IC), a compound of Formula (II), a compound of Formula (IC),
- the PLK4 inhibitor is compound set forth in Table 1. In embodiments, the PLK4 inhibitor is compound set forth in Table 2. In embodiments, the PLK4 inhibitor is centrinone. In embodiments, the PLK4 inhibitor is centrinone B. In embodiments, the cancer is a primary cancer. In embodiments, the cancer is metastatic cancer. In aspects, the methods comprise treating small cell lung cancer, a wild type p53 positive small cell lung cancer, or a mutant p53 positive small cell lung cancer. In aspects, the methods comprise treating neuroblastoma, a wild type p53 positive neuroblastoma, or a mutant p53 positive neuroblastoma.
- the methods comprise treating prostate cancer, a wild type p53 positive prostate cancer, or a mutant p53 positive prostate cancer.
- the methods comprise treating glioma, a wild type p53 positive glioma, or a mutant p53 positive glioma.
- the methods comprise treating mesothelioma, a wild type p53 positive mesothelioma, or a mutant p53 positive mesothelioma.
- the methods comprise treating osteosarcoma, a wild type p53 positive osteosarcoma, or a mutant p53 positive osteosarcoma.
- the methods comprise treating breast cancer, a wild type p53 positive breast cancer, or a mutant p53 positive breast cancer.
- the methods comprise treating Ewing's sarcoma, a wild type p53 positive Ewing's sarcoma, or a mutant p53 positive Ewing's sarcoma.
- the methods comprise treating soft tissue cancer, a wild type p53 positive soft tissue cancer, or a mutant p53 positive soft tissue cancer.
- the methods comprise treating T cell lymphoma, a wild type p53 positive T cell lymphoma, or a mutant p53 positive T cell lymphoma.
- the T cell lymphoma is extranodal T cell lymphoma.
- the T cell lymphoma is cutaneous T cell lymphoma. In aspects, the T cell lymphoma is anaplastic large cell lymphoma. In aspects, the T cell lymphoma is angioimmunoblastic T cell lymphoma. In embodiments, the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- the chemotherapeutic agent comprises a tubule polymerization inhibitor, such as vincristine, vinblastine, vinoreIbine, vinfluine, a dolastatin, a halichondrin, a hemiasterline, cryptophysin 52, or a combination of two or more thereof.
- the chemotherapeutic agent comprises cyclophosphamide, ifosfamide, cisplatin, carboplatin, vincristine, doxorubicin, etoposide, topotecan, busulfan, melphalan, or a combination of two or more thereof.
- the chemotherapeutic agent comprises differentiation therapy, wherein the chemotherapeutic agent is retinoic acid (e.g., all-trans-retinoic acid, 13-cis-retinoic acid), arsenic trioxide, or a combination thereof.
- the disclosure provides methods of treating a pediatric cancer or a neural crest-derived caner in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor; wherein the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin.
- the disclosure provides methods of treating a wild type or mutant a pediatric cancer or a wild-type or mutant neural crest-derived caner in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor.
- the disclosure provides methods of treating a wild type or mutant a pediatric cancer or a wild-type or mutant neural crest-derived caner in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor; wherein the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin.
- the subject has an elevated level of TRIM37.
- the subject has an elevated level of Chromogranin A.
- the subject has an elevated level of Synaptophysin.
- the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A.
- the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin. In embodiments, the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin. In embodiments, the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level. In embodiments, the subject has an elevated level of TRIM37, Chromogranin A, and/or Synaptophysin, when compared to a control. In embodiments, the subject has an elevated level of TRIM37 when compared to a control. In embodiments, the subject has an elevated level of Chromogranin A when compared to a control.
- the subject has an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Chromogranin A when compared to a control. In embodiments, the subject has an elevated level of TRIM37 and an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control. In embodiments, the subject has an elevated level of TRIM37, an elevated level of Chromogranin A, and an elevated level of Synaptophysin level when compared to a control.
- the PLK4 inhibitor is compound of Formula (Ia), a compound of Formula (Ib), a compound of Formula (Ia1), a compound of Formula (Ia2), a 5 compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), a compound of Formula (Ib7), a compound of Formula (IC), a compound of Formula (II), a compound of Formula (IC),
- the PLK4 inhibitor is compound set forth in Table 1. In embodiments, the PLK4 inhibitor is compound set forth in Table 2. In embodiments, the PLK4 inhibitor is centrinone. In embodiments, the PLK4 inhibitor is centrinone B. In embodiments, the cancer is a primary cancer. In embodiments, the cancer is metastatic cancer. In aspects, the methods comprise treating pediatric cancer, a wild type p53 positive pediatric cancer, or a mutant p53 positive pediatric cancer. In embodiments, the pediatric cancer is a rhabdoid tumor, a neuroblastoma, an acute lymphoblastic leukemia tumor, or a brain and central nervous system tumor.
- the methods comprise treating neural crest-derived cancer, a wild type p53 positive neural crest-derived cancer, or a mutant p53 positive neural crest-derived cancer.
- the neural crest-derived tumor is a small cell lung cancer, a melanoma, or a breast cancer.
- the methods further comprising administering a chemotherapeutic agent, radiation therapy, or a combination thereof.
- the chemotherapeutic agent comprises a tubule polymerization inhibitor, such as vincristine, vinblastine, vinoreIbine, vinfluine, a dolastatin, a halichondrin, a hemiasterline, cryptophysin 52, or a combination of two or more thereof.
- the chemotherapeutic agent comprises cyclophosphamide, ifosfamide, cisplatin, carboplatin, vincristine, doxorubicin, etoposide, topotecan, busulfan, melphalan, or a combination of two or more thereof.
- the chemotherapeutic agent comprises differentiation therapy, wherein the chemotherapeutic agent is retinoic acid (e.g., all-trans-retinoic acid, 13-cis-retinoic acid), arsenic trioxide, or a combination thereof.
- the disclosure provides methods of treating cancer in a subject in need thereof, the method comprising: (i) measuring a TRIM37 level, Chromogranin A level and/or Synaptophysin level in a biological sample obtained from the subject; and (ii) administering an effective amount of a PLK4 inhibitor to the subject to treat the cancer.
- the methods comprise measuring: (i) the gene level, (ii) the mRNA level, (iii) the protein level, or (iv) a combination of two or more of the gene, mRNA, and protein levels of: (a) TRIM37, (b) Chromogranin A, (c) Synaptophysin, or (d) a combination of two or more of TRIM37, Chromogranin A, and Synaptophysin.
- the methods comprise measuring a TRIM37 gene level in the biological sample.
- the methods comprise measuring a TRIM37 mRNA level in the biological sample.
- the methods comprise measuring a TRIM37 protein level in the biological sample.
- the methods comprise measuring a Chromogranin A gene level in the biological sample. In embodiments, the methods comprise measuring a Chromogranin A mRNA level in the biological sample. In embodiments, the methods comprise measuring a Chromogranin A protein level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin gene level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin mRNA level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin protein level in the biological sample. In embodiments, the subject has an elevated TRIM37 level. In embodiments, the subject has an elevated Chromogranin A level. In embodiments, the subject has an elevated Synaptophysin level.
- the subject has an elevated TRIM37 level and an elevated Chromogranin A level. In embodiments, the subject has an elevated TRIM37 level and an elevated Synaptophysin level. In embodiments, the subject has an elevated Chromogranin A level and an elevated Synaptophysin level. In embodiments, the subject has an elevated TRIM37 level, an elevated Chromogranin A level, and an elevated Synaptophysin level. In embodiments, the subject has an elevated TRIM37 level when compared to a control. In embodiments, the subject has an elevated Chromogranin A level when compared to a control. In embodiments, the subject has an elevated Synaptophysin level when compared to a control.
- the subject has an elevated TRIM37 level and an elevated Chromogranin A level when compared to a control. In embodiments, the subject has an elevated TRIM37 level and an elevated Synaptophysin level when compared to a control. In embodiments, the subject has an elevated Chromogranin A level and an elevated Synaptophysin level when compared to a control. In embodiments, the subject has an elevated TRIM37 level, an elevated Chromogranin A level, and an elevated Synaptophysin level when compared to a control. In embodiments, the subject has a wild type or mutant p53 positive cancer. In embodiments, the biological sample is a plasma sample. In embodiments, the biological sample is a serum sample.
- the biological sample is a tissue sample. In embodiments, the biological sample is a cell sample. In embodiments, the biological sample is a blood sample. In embodiments, the biological sample is a peripheral blood sample. In embodiments, the biological sample is a tumor sample. In embodiments, the biological sample is a primary tumor sample. In embodiments, the biological sample is a metastatic tumor sample. In embodiments, the biological sample is a resected tumor sample. In embodiments, the biological sample is a tumor biopsy sample. In embodiments, the biological sample is a resected primary tumor sample. In embodiments, the biological sample is a resected metastatic tumor sample. In embodiments, the biological sample is a primary tumor biopsy sample.
- the biological sample is a metastatic tumor biopsy sample.
- the PLK4 inhibitor is compound of Formula (Ia), a compound of Formula (Ib), a compound of Formula (Ia1), a compound of Formula (Ia2), a compound of 10 Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), a compound of Formula (Ib7), a compound of Formula (IC)
- the PLK4 inhibitor is compound set forth in Table 1. In embodiments, the PLK4 inhibitor is compound set forth in Table 2. In embodiments, the PLK4 inhibitor is centrinone. In embodiments, the PLK4 inhibitor is centrinone B. In embodiments, the cancer is a cancer tumor. In embodiments, the cancer tumor is a primary cancer tumor. In embodiments, the cancer tumor is a metastatic cancer tumor.
- the disclosure provides methods to identify a subject responsive to a PLK4 inhibitor, the method comprising: (i) obtaining a biological sample from the subject; and (ii) measuring a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level in the biological sample; wherein if the TRIM37 level, the Chromogranin A level, and/or the Synaptophysin level is elevated the subject is identified as responsive to the PLK4 inhibitor.
- the disclosure provides methods to identify a subject responsive to a PLK4 inhibitor, the method comprising: (i) obtaining a biological sample from the subject; (ii) measuring a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level in the biological sample; (iii) analyzing the biological sample for p53; wherein if the subject has an elevated TRIM37 level a Chromogranin A level, and/or a Synaptophysin level and is p53 positive, the subject is identified as responsive to the PLK4 inhibitor.
- the methods comprise measuring: (i) the gene level, (ii) the mRNA level, (iii) the protein level, or (iv) a combination of two or more of the gene, mRNA, and protein levels of: (a) TRIM37, (b) Chromogranin A, (c) Synaptophysin, or (d) a combination of two or more of TRIM37, Chromogranin A, and Synaptophysin.
- the methods comprise measuring a TRIM37 gene level in the biological sample.
- the methods comprise measuring a TRIM37 mRNA level in the biological sample.
- the methods comprise measuring a TRIM37 protein level in the biological sample.
- the methods comprise measuring a Chromogranin A gene level in the biological sample. In embodiments, the methods comprise measuring a Chromogranin A mRNA level in the biological sample. In embodiments, the methods comprise measuring a Chromogranin A protein level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin gene level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin mRNA level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin protein level in the biological sample. In embodiments, the TRIM37 level is elevated. In embodiments, the Chromogranin A level is elevated. In embodiments, the Synaptophysin level is elevated.
- the TRIM37 level is elevated and the Chromogranin A level is elevated. In embodiments, the TRIM37 level is elevated and the Synaptophysin level is elevated. In embodiments, the Chromogranin A level is elevated and the Synaptophysin level is elevated. In embodiments, the TRIM37 level is elevated, the Chromogranin A level is elevated, and the Synaptophysin level is elevated. In embodiments, the TRIM37 level is elevated when compared to a control. In embodiments, the Chromogranin A level is elevated when compared to a control. In embodiments, the Synaptophysin level is elevated when compared to a control.
- the TRIM37 level is elevated and the Chromogranin A level is elevated when compared to a control. In embodiments, the TRIM37 level is elevated and the Synaptophysin level is elevated when compared to a control. In embodiments, the Chromogranin A level is elevated and the Synaptophysin level is elevated when compared to a control. In embodiments, the TRIM37 level is elevated, the Chromogranin A level is elevated, and the Synaptophysin level is elevated when compared to a control.
- the disclosure provides methods to identify a subject responsive to a PLK4 inhibitor, the method comprising: (i) obtaining a biological sample from the subject; (ii) analyzing the biological sample for mutant or wild type p53; wherein if the subject is p53 positive, the subject is identified as responsive to the PLK4 inhibitor. In embodiments, the methods further comprising administering an effective amount of a PLK4 inhibitor to the subject.
- the biological sample is a plasma sample.
- the biological sample is a serum sample.
- the biological sample is a tissue sample.
- the biological sample is a cell sample.
- the biological sample is a blood sample.
- the biological sample is a peripheral blood sample.
- the biological sample is a tumor sample. In embodiments, the biological sample is a primary tumor sample. In embodiments, the biological sample is a metastatic tumor sample. In embodiments, the biological sample is a resected tumor sample. In embodiments, the biological sample is a tumor biopsy sample. In embodiments, the biological sample is a resected primary tumor sample. In embodiments, the biological sample is a resected metastatic tumor sample. In embodiments, the biological sample is a primary tumor biopsy sample. In embodiments, the biological sample is a metastatic tumor biopsy sample.
- the PLK4 inhibitor is compound of Formula (Ia), a compound of Formula (Ib), a compound of Formula (Ia1), a compound of Formula (Ia2), a compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), a compound of Formula (Ib7), a compound of Formula (IC), a compound of Formula (II), a compound of Formula (III), a
- the PLK4 inhibitor is compound set forth in Table 1. In embodiments, the PLK4 inhibitor is compound set forth in Table 2. In embodiments, the PLK4 inhibitor is centrinone. In embodiments, the PLK4 inhibitor is centrinone B. In embodiments, the subject has cancer. In embodiments, the cancer is a cancer tumor. In embodiments, the cancer tumor is a primary cancer tumor. In embodiments, the cancer tumor is a metastatic cancer tumor.
- the disclosure provides methods of selecting a subject for treatment with a PLK4 inhibitor, the method comprising: (i) obtaining a biological sample from the subject; and (ii) measuring a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level in the biological sample; wherein if the TRIM37 level, the Chromogranin A level, and/or the Synaptophysin level is elevated, the subject is selected for treatment with the PLK4 inhibitor.
- the disclosure provides methods of selecting a subject for treatment with a PLK4 inhibitor, the method comprising: (i) obtaining a biological sample from the subject; (ii) measuring a TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level in the biological sample; and (iii) analyzing the biological sample for p53; wherein if the subject has an elevated TRIM37 level, a Chromogranin A level, and/or a Synaptophysin level and is p53 positive, the subject is identified as responsive to the PLK4 inhibitor.
- the methods comprise measuring: (i) the gene level, (ii) the mRNA level, (iii) the protein level, or (iv) a combination of two or more of the gene, mRNA, and protein levels of: (a) TRIM37, (b) Chromogranin A, (c) Synaptophysin, or (d) a combination of two or more of TRIM37, Chromogranin A, and Synaptophysin.
- the methods comprise measuring a TRIM37 gene level in the biological sample.
- the methods comprise measuring a TRIM37 mRNA level in the biological sample.
- the methods comprise measuring a TRIM37 protein level in the biological sample.
- the methods comprise measuring a Chromogranin A gene level in the biological sample. In embodiments, the methods comprise measuring a Chromogranin A mRNA level in the biological sample. In embodiments, the methods comprise measuring a Chromogranin A protein level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin gene level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin mRNA level in the biological sample. In embodiments, the methods comprise measuring a Synaptophysin protein level in the biological sample. In embodiments, the TRIM37 level is elevated. In embodiments, the Chromogranin A level is elevated. In embodiments, the Synaptophysin level is elevated.
- the TRIM37 level is elevated and the Chromogranin A level is elevated. In embodiments, the TRIM37 level is elevated and the Synaptophysin level is elevated. In embodiments, the Chromogranin A level is elevated and the Synaptophysin level is elevated. In embodiments, the TRIM37 level is elevated, the Chromogranin A level is elevated, and the Synaptophysin level is elevated. In embodiments, the TRIM37 level is elevated when compared to a control. In embodiments, the Chromogranin A level is elevated when compared to a control. In embodiments, the Synaptophysin level is elevated when compared to a control.
- the TRIM37 level is elevated and the Chromogranin A level is elevated when compared to a control. In embodiments, the TRIM37 level is elevated and the Synaptophysin level is elevated when compared to a control. In embodiments, the Chromogranin A level is elevated and the Synaptophysin level is elevated when compared to a control. In embodiments, the TRIM37 level is elevated, the Chromogranin A level is elevated, and the Synaptophysin level is elevated when compared to a control.
- the disclosure provides methods of selecting a subject for treatment with a PLK4 inhibitor, the method comprising: (i) obtaining a biological sample from the subject; (ii) analyzing the biological sample for wild type or mutant p53; wherein if the subject is p53 positive, the subject is identified as responsive to the PLK4 inhibitor. In embodiments, the methods further comprising administering an effective amount of a PLK4 inhibitor to the subject.
- the biological sample is a plasma sample.
- the biological sample is a serum sample.
- the biological sample is a tissue sample.
- the biological sample is a cell sample.
- the biological sample is a blood sample.
- the biological sample is a peripheral blood sample.
- the biological sample is a tumor sample. In embodiments, the biological sample is a primary tumor sample. In embodiments, the biological sample is a metastatic tumor sample. In embodiments, the biological sample is a resected tumor sample. In embodiments, the biological sample is a tumor biopsy sample. In embodiments, the biological sample is a resected primary tumor sample. In embodiments, the biological sample is a resected metastatic tumor sample. In embodiments, the biological sample is a primary tumor biopsy sample. In embodiments, the biological sample is a metastatic tumor biopsy sample.
- the PLK4 inhibitor is compound of Formula (Ia), a compound of Formula (Ib), a compound of Formula (Ia1), a compound of Formula (Ia2), a compound of Formula (Ia3), a compound of Formula (Ia4), a compound of Formula (Ia5), a compound of Formula (Ia6), a compound of Formula (Ia7), a compound of Formula (Ia8), a compound of Formula (Ia9a), a compound of Formula (Ia9b), a compound of Formula (Ia9c), a compound of Formula (Ia9d), a compound of Formula (Ia9f), a compound of Formula (Ib1), a compound of Formula (Ib2), a compound of Formula (Ib3), a compound of Formula (Ib4), a compound of Formula (Ib5), a compound of Formula (Ib6), a compound of Formula (Ib7), a compound of Formula (IC), a compound of Formula (II), a compound of Formula (III), a
- the PLK4 inhibitor is compound set forth in Table 1. In embodiments, the PLK4 inhibitor is compound set forth in Table 2. In embodiments, the PLK4 inhibitor is centrinone. In embodiments, the PLK4 inhibitor is centrinone B. In embodiments, the subject has cancer. In embodiments, the cancer is a cancer tumor. In embodiments, the cancer tumor is a primary cancer tumor. In embodiments, the cancer tumor is a metastatic cancer tumor.
- a method of treating cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 1 wherein the subject has an elevated level of Chromogranin A; an elevated level of Synaptophysin; or an elevated level of Chromogranin A and an elevated level of Synaptophysin.
- a method of treating cancer in a subject in need thereof comprising: (i) measuring a TRIM37 level in a biological sample obtained from the subject; and (ii) administering an effective amount of a PLK4 inhibitor to the subject to treat the cancer.
- Embodiment 3 comprising measuring copies of the TRIM37 gene, measuring the level of the TRIM37 mRNA, measuring the level of the TRIM37 protein, or a combination of two or more thereof.
- Embodiment 3 or 4 wherein the biological sample obtained from the subject has an elevated level of TRIM37 when compared to a control.
- Embodiments 3 to 5 further comprising measuring a Chromogranin A level; a Synaptophysin level; or a Chromogranin A level and a Synaptophysin level.
- Embodiment 12 wherein the blood sample is a peripheral blood sample.
- Embodiment 19 The method of Embodiment 19, wherein the neural crest-derived cancer is small cell lung cancer.
- Embodiment 19 wherein the neural crest-derived cancer is melanoma.
- Embodiment 19 wherein the neural crest-derived cancer is breast cancer.
- T cell lymphoma is extranodal T cell lymphoma, cutaneous T cell lymphoma, anaplastic large cell lymphoma, or angioimmunoblastic T cell lymphoma.
- Embodiment 37 wherein the cancer is a wild-type p53 positive cancer.
- Embodiment 37 wherein the cancer is a mutant p53 positive cancer.
- a method of treating a p53 positive cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the p53 positive cancer.
- Embodiment 40 wherein the p53 positive cancer is a wild type p53 positive cancer.
- Embodiment 40 wherein the p53 positive cancer is a mutant p53 positive cancer.
- a method of treating a wild type p53 positive pediatric cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type p53 positive pediatric cancer.
- a method of treating a mutant p53 positive pediatric cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive pediatric cancer.
- a method of treating a wild-type p53 positive pediatric cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild-type p53 positive pediatric cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 45 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a mutant p53 positive pediatric cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive pediatric cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 47 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a pediatric cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the pediatric cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 49 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- Embodiments 43 to 50 wherein the pediatric cancer is acute lymphoblastic leukemia, neuroblastoma, a rhabdoid tumor, or tumor of the brain and central nervous system.
- a method of treating a wild type p53 positive, neural crest-derived cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type p53 positive, neural crest-derived cancer.
- a method of treating a mutant type p53 positive, neural crest-derived cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive, neural crest-derived cancer.
- a method of treating a wild type p53 positive, neural crest-derived cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type p53 positive, neural crest-derived cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 54 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a mutant p53 positive, neural crest-derived cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive, neural crest-derived cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 56 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a neural crest-derived cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the neural crest-derived cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 58 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- the neural crest-derived cancer is small cell lung cancer, breast cancer, or melanoma.
- a method of treating a wild-type p53 positive cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild-type p53 positive cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 61 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a mutant p53 positive cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 63 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a wild type p53 positive neuroblastoma in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type p53 positive neuroblastoma; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 65 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a mutant p53 positive neuroblastoma in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive neuroblastoma; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 67 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a wild type or mutant p53 positive neuroblastoma in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type or mutant p53 positive neuroblastoma.
- a method of treating a wild type p53 positive small cell lung cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type p53 positive small cell lung cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 70 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a mutant p53 positive small cell lung cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the mutant p53 positive small cell lung cancer; wherein the subject has an elevated level of TRIM37 when compared to a control.
- Embodiment 72 wherein the subject has an elevated level of Chromogranin A when compared to a control; an elevated level of Synaptophysin when compared to a control; or an elevated level of Chromogranin A and an elevated level of Synaptophysin when compared to a control.
- a method of treating a wild type or mutant p53 positive small cell lung cancer in a subject in need thereof comprising administering to the subject an effective amount of a PLK4 inhibitor to treat the wild type or mutant p53 positive small cell lung cancer.
- Embodiment 75 wherein the chemotherapeutic agent is a tubule polymerization inhibitor.
- tubule polymerization inhibitor is vincristine, vinblastine, vinoreIbine, vinfluine, a dolastatin, a halichondrin, a hemiasterline, cryptophysin 52, or a combination of two or more thereof.
- the chemotherapeutic agent comprises cyclophosphamide, ifosfamide, cisplatin, carboplatin, vincristine, doxorubicin, etoposide, topotecan, busulfan, melphalan, or a combination of two or more thereof.
- Embodiment 79 wherein the differentiation therapy comprises retinoic acid, arsenic trioxide, or a combination thereof.
- Embodiment 80 wherein the retinoic acid is all-trans-retinoic acid, 13-cis-retinoic acid, or a combination thereof.
- a method to identify a subject responsive to a PLK4 inhibitor comprising: (i) obtaining a biological sample from the subject; and (ii) measuring a TRIM37 level in the biological sample; wherein if the TRIM37 level is elevated when compared to a control, the subject is identified as responsive to the PLK4 inhibitor.
- Embodiment 83 further comprising measuring a Chromogranin A level; measuring a Synaptophysin level; or measuring a Chromogranin A and a Synaptophysin level.
- a method of selecting a subject for treatment with a PLK4 inhibitor comprising: (i) obtaining a biological sample from the subject; and (ii) measuring a TRIM37 level in the biological sample; wherein if the TRIM37 level is elevated when compared to a control, the subject is selected for treatment with the PLK4 inhibitor.
- Embodiment 85 further comprising measuring a Chromogranin A level; measuring a Synaptophysin level; or measuring a Chromogranin A and a Synaptophysin level.
- the tumor sample is a resected tumor or a tumor biopsy sample.
- Embodiment 89 or 90 wherein the tumor sample is a primary tumor sample or a metastisic tumor sample.
- R 3 , R 4 , R 5 , z1, R 13 , and R 60 areas defined in the specification.
- R 3 , R 4 , R 5 , z1, z3, and R 60 are as defined in the specification.
- R 3 , R 4 , R 5 , z1, z3, and R 60 are as defined in the specification.
- L 1 , z1, R 1 , R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- L 1 , z1, R 1 , R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- L 1 , z1, R 1 , R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- L 1 , z1, R 1 , R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- R 1 , R 2 , R 3 , R 4 , R, and R 6 are as defined in the specification.
- R 3A is substituted or unsubstituted alkyl
- R 4C is substituted or unsubstituted heterocycloalkyl, and the remaining substituents are as defined in the specification.
- L 1 , z1, R 3 , R 5 , R 6 , R 4A , and R 40B are as defined in the specification.
- L 1 , z1, z4, R 1 , R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- L 1 , z1, R, R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- L 1 , z1, R, R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- L 1 , z1, R 1 , R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- L 1 , z1, R 1 , R 2 , R 5 , R 6 , and R 40 are as defined in the specification.
- centrosome removal would trigger the mitotic duration sensor in these cell lines, enhancing the effect of centrosome removal on proliferation.
- six neuroblastoma cells lines were collected (SH—SY5Y, CHP134, IMR32, CHP212, SK-N—SH, and SK-N-F1) along with 5 other control cell lines derived from tumors from children or adolescents (1 hepatocellular carcinoma, HepG2; 2 Rhabdoid Tumor (ATRT), BT-12 and BT-16; and 2 Ewing Sarcoma SK-ES-1 and A673).
- Caspase assays comparing RPE1 cells, which senesce but do not die in the presence of centrinone, and two other ‘Not Sensitive’ cell lines (BT-12 and BT-16), to three of the ‘Sensitive’ neuroblastoma cell lines, revealed high levels of caspase activity in the ‘Sensitive’ cell lines compared to relatively low levels in the cell lines that were ‘Not Sensitive’ ( FIG. 1 C ). Comparable levels of caspase activity were also detected in cells treated with centrinone B, a related PLK4 inhibitor, suggesting that the effect the result of inhibiting PLK4, rather than an off target effect.
- both PLK4 alleles in one of the neuroblastoma lines, CHP134 was engineered to express a G95L mutant PLK4 protein.
- the CHP134 cells harboring the G95L mutant PLK4 proliferated in centrinone at a rate comparable to non-mutant cells in DMSO ( FIG. 1 B ) demonstrating that the cell death in the presence of centrinone is due to PLK4 inhibition.
- FIG. 2 A shows the CHP134 neuroblastoma cell line has four (4) copies of the TRIM37 gene.
- CHP134 clones with varying TRIM37 copy numbers were isolated and the levels of TRIM37 protein were measured by quantitative western blotting. Alpha-tubulin was used as a blotting control. Clones with TRIM37 levels between 12 and 64% of the levels in the WT cell line were isolated.
- FIG. 2 B and 2 C show analysis of mitosis by live cell filming in the CHP134 clones after 3 cell cycles in 150 nM centrinone and the duration of mitosis and rate of anaphase failure were measured for each clone.
- Graphs plot mean mitotic duration ( FIG. 2 B ) and the percentage of cells exhibiting anaphase failure ( FIG. 2 C ) versus measured TRIM37 protein level.
- FIG. 2 D presents data in which cellular proliferation was assessed for each of the CHP134 clones by performing an ATPlite assay after 5 days in 125 nM centrinone. Results are expressed as the percent of the value for the equivalent DMSO-treated control and are plotted versus measured TRIM37 protein level. Note that WT CHP134 cells exhibited a loss in proliferation comparable to that of the two highest mutant clones. Thus, for centrinone-treated CHP134 cells, mitotic success and cell viability both decreased as TRIM37 levels increased.
- FIG. 3 A is a graph plotting TRIM37 copy number versus expression level for breast cancer cell lines (data from the CCLE database; (Barretina et al., 2012)).
- FIG. 3 B are graphs showing the results of passaging assays for the indicated cancer cell lines that monitor cell proliferation after the addition of DMSO (vehicle; black) or 150 nM centrinone (grey) at day 0. The right graph in each pair shows the centrinone-treated curve without the control.
- p53 activity and TRIM37 protein levels across the same panel of cell lines as in FIG. 1 was examined.
- cells were challenged by incubation with an inhibitor of Mdm2, an ubiquitin ligase that continuously targets p53 for proteasomal degradation.
- a value of 1 means that there was no cell proliferation after addition of Mdm2 inhibitor indicating fully functional p53, and a value of 0 means that proliferation was comparable in the presence and absence of Mdm2 inhibitor, indicating lack of functional p53.
- This analysis revealed that p53 was functional in RPE1 cells as well as 5 of the neuroblastoma cell lines (SH—SY5Y, CHP134, MR32, CHP212 and SK-N—SH) and 3 of the control cell lines (HepG2, BT-12, and BT-16) and was inactive in HeLa cells as well as 1 of the neuroblastoma cell lines (SK-N-F1) and two of the control cell lines (A673 and SK-ES-1) ( FIG. 4 A ).
- TRIM37 protein levels were measured across the same panel of cell lines by quantitative western blotting ( FIG. 4 B ).
- the p53+ cell lines black in FIG. 4 B
- the five p53+ neuroblastoma cell lines all expressed more TRIM37 than the four p53+ control cell lines.
- the four p53 ⁇ cell lines FIG. 4 B
- the one p53-neuroblastoma cell line that we analyzed (SK-N-F1) expressed substantially more TRIM37 than the three p53 ⁇ control cell lines.
- the inventors measured mitotic duration to determine if they would be likely to die due to tripping the mitotic duration sensor.
- the mean duration of mitosis was similar for the five p53+ cell lines (Range 32.1 to 41.8 minutes, black bars in FIG. 5 B ).
- the mean duration of mitosis increased substantially (Range 69.4 to 490.3 minutes, FIG. 5 B ).
- the three cell lines with the low TRIM37 levels exhibited the lowest mean mitotic durations after centrinone treatment (Range 69.4 to 152.8) and the three cell lines with high TRIM37 levels exhibited the highest mean mitotic durations (Range 221.1 to 490.3 minutes).
- TRIM37 deletion completely suppressed the anaphase failure and indicated the increase in mitotic duration observed in centrinone-treated CHP134 cells ( FIG. 6 A ), and enabled their proliferation in a 12-day passaging assay ( FIG. 6 B ).
- shRNA-mediated knockdown of p53 which abrogates the mitotic duration sensor, did not prevent the anaphase failure or increase in mitotic duration resulting from centrinone-mediated centrosome loss.
- it allowed CHP134 cells to proliferate in the 12-day passaging assay presumably because it enabled daughter cells born after prolonged mitosis to continue to divide.
- CHP134 cells in which TRIM37 was deleted and p53 was knocked down also exhibited robust proliferation in centrinone ( FIG. 6 A , B).
- Cep192 is an E3 ubiquitin ligase of the TRIpartite Motif (TRIM) protein family. To determine if its ligase activity is important for its activity in antagonizing the formation of centrosome-like foci, the formation of foci containing the centrosomal protein Cep192 was examined in control RPE1 cells, RPE1 TRIM37 ⁇ cells, along with RPE1 TRIM37 ⁇ cells in which expression of wild-type TRIM37 or a TRIM37 mutant with a residue change previously shown to its ubiquitin ligase activity (TRIM37-C18R; (Bhatnagaar et al., 2014)) was reconstituted at levels comparable to those in unperturbed cells by expression under the UBC promoter ( FIG.
Landscapes
- Health & Medical Sciences (AREA)
- Veterinary Medicine (AREA)
- Chemical & Material Sciences (AREA)
- Medicinal Chemistry (AREA)
- Pharmacology & Pharmacy (AREA)
- Life Sciences & Earth Sciences (AREA)
- Animal Behavior & Ethology (AREA)
- General Health & Medical Sciences (AREA)
- Public Health (AREA)
- Epidemiology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Organic Chemistry (AREA)
- Oncology (AREA)
- Hematology (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
Abstract
Description
wherein the substituents are defined herein. In embodiments, the PLK4 inhibitor is a compound of Formula (Ib) or a pharmaceutically acceptable salt thereof:
Wherein R3, R4, R5, z1, R13, and R60 are as described herein. The symbol z3 is an integer of 0, 1, 2, 3, 4, or 5. The symbol z3 may be 1, 2, or 3. The symbol z3 may be 1. The symbol z3 may be 2. The symbol z3 may be 3.
Wherein R3, R4, R5, z1, z3, and R60 are as described herein. R60 are independently hydrogen, halogen, —N3, —CF3, —CCl3, —CBr3, —CI3, —CN, —COR61, —OR60A, —NR60AR60B, —C(O)OR60A, —C(O)NR60AR60B, —NO2, —SR60A, —S(O)2H, —S(O)2OH, —S(O)2NH2, —NHNH2, —ONH2, —NHC(O)NHNR60AR60B, R61-substituted or unsubstituted alkyl, R61-substituted or unsubstituted heteroalkyl, R61-substituted or unsubstituted cycloalkyl, R61-substituted or unsubstituted heterocycloalkyl, R61-substituted or unsubstituted aryl, or R61-substituted or unsubstituted heteroaryl. R60A may independently be hydrogen, halogen, —NO2, —CF3, —CN, —COR61, R61-substituted or unsubstituted alkyl, R61-substituted or unsubstituted heteroalkyl, or R61-substituted or unsubstituted aryl; R61 is independently hydrogen, halogen, or unsubstituted alkyl. R60B may independently be hydrogen halogen, or unsubstituted alkyl.
where L1, z1, R1, R2, R5, R6, and R40 are as described herein. The symbol z4 is an integer of 0, 1, 2, 3, 4, 5, 6, or 7. Ring A is cycloalkyl or heterocycloalkyl. In embodiments, the compound of Formula (Ia2) is:
where L1, z1, R1, R2, R5, R6, and R40 are as described herein. In embodiments, the compound of Formula (Ia3) is:
where L1, z1, R1, R2, R5, R6, and R40 are as described herein and the symbol z4 is an integer of 0, 1, 2, 3, or 4. In embodiments, the compound of Formula (Ia4) is:
where L1, z1, z4, R1, R2, R5, R6, and R40 are as described herein. In embodiments, the compound of Formula (Ia5) is:
where L1, z1, R1, R2, R5, R6, and R40 are as described herein and the symbol z4 is an integer of 0, 1, 2, 3, or 4. In embodiments, the compound of Formula (Ia6) is:
where L1, z1, z4, R1, R2, R5, R6, and R40 are as described herein and the compound is not tozasertib. Ring A is cycloalkyl or heterocycloalkyl. In embodiments, the compound of Formula (Ib2) is:
where L1, z1, R1, R2, R5, R6, and R40 are as described herein. In embodiments, the compound of Formula (Ib3) is:
where L1, z1, R1, R2, R5, R6, and R40 are as described herein and the symbol z4 is an integer of 0, 1, 2, 3, or 4. In embodiments, the compound of Formula (Ib4) is:
where L1, z1, R1, R2, R5, R6, and R40 are as described herein and the symbol z4 is an integer of 0, 1, 2, 3, 4, or 5. In embodiments, the compound of Formula (Ib5) is:
where L1, z1, R1, R2, R5, R6, and R40 are as described herein and the symbol z4 is an integer of 0, 1, 2, 3, or 4. In embodiments, the compound of Formula (Ib6) is:
where z1, R1, R2, R3, R4, R5, and R6 are as described herein. In embodiments, the compound of Formula (Ia7) is:
where z1, R1, R2, R3, R4, R5, and R6 are as described herein. In embodiments, the compound of Formula (Ia8) is:
wherein R3A is substituted or unsubstituted alkyl, and R4C is substituted or unsubstituted heterocycloalkyl.
wherein R60 and z5 are as defined herein. In embodiments, R60 is halogen or —NO2, and z5 is 1 or 2. In embodiments, z5 is 1. In embodiments, z5 is 2.
wherein z1, R1, R2, R3, R4, R5, and R6 are as described herein. In embodiments, the compound of Formula (Ib7) is:
wherein z2a is 0, 1, 2, 3, or 4, and the compound is not tozasertib. z2a may be 0, 1, or 2. z2a may be 0. z2a may be 1. z2a may be 2. z2a may be 3. z2a may be 4.
wherein z2a is 0, 1, 2, 3, or 4, and the compound is not tozasertib. z2a may be 0, 1, or 2. z2a may be 0. z2a may be 1. z2a may be 2. z2a may be 3. z2a may be 4.
| TABLE 1 | |
| Activity | |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
X |
|
|
X |
|
|
X |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
X |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
X |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
X |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
X |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
X |
|
|
X |
|
|
X |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
X |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
X |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XX |
|
|
XXX |
|
|
XX |
|
|
XX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
|
|
XXX |
| TABLE 2 | |
| Compound Names for exemplified compounds of formula(I), (II), and (III). | |
| All compounds were confirmed by mass spectro scopy and by 1H and 13C NMR. | |
| No. | Chemical Name |
| 1 | N-(4-((4,5-dimethyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 2 | N-(4-((4-((5-methyl-1H-pyrazol-3-yl)amino)-6,7-dihydro-5H-cyclopenta[d]pyrimidin- |
| 2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 3 | N-(4-((4-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 4 | N-(4-((5-(hydroxymethyl)-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin- |
| 1-yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 5 | N-(4-((5-ethyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 6 | N-(4-((5-ethyl-4-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 7 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 8 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 9 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 10 | N-(4-((4-(3,4-dimethylpiperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 11 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 12 | N-(4-((4-(4-(2-methoxyethyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 13 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 14 | N-(4-((4-(4-isopropylpiperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 15 | N-(4-((4-(dimethylamino)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 16 | N-(4-((4-(ethyl(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 17 | N-(4-((5-methoxy-4-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 18 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6,7-dihydro-5H- |
| cyclopenta[d]pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 19 | N-(4-((4-((2-(dimethylamino)ethyl)(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 20 | N-(4-((4-((2-(dimethylamino)ethyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 21 | N-(4-((4-((2-methoxyethyl)(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 22 | N-(4-((4-((5-ethyl-1H-pyrazol-3-yl)amino)-5-methyl-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 23 | N-(4-((4-(1H-imidazol-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin- |
| 2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 24 | N-(4-((4-(diethylamino)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 25 | N-(4-((4-chloro-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 26 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 27 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 28 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 29 | N-(4-((5-methyl-4-(methyl(2-(methylamino)ethyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 30 | N-(4-((4-((1H-pyrazol-3-yl)amino)-5-methyl-6-(4-methylpiperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 31 | N-(4-((4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1-yl)-5- |
| propoxypyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 32 | N-(4-((4-(diethylamino)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 33 | N-(4-((4-(dimethylamino)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin- |
| 2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 34 | N-(4-((4-(ethyl(methyl)amino)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 35 | N-(4-((5-(2-methoxyethoxy)-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4- |
| methylpiperazin-1-yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 36 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 37 | N-(4-((5-methoxy-4-((2-methoxyethyl)(methyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 38 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 39 | N-(4-((4-((2-(dimethylamino)ethyl)(methyl)amino)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 40 | N-(4-((4-((2-(dimethylamino)ethyl)amino)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 41 | N-(4-((4-(diethylamino)-6-((5-methyl-1H-pyrazol-3-yl)amino)-5-propoxypyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 42 | N-(4-((4-(isopropyl(methyl)amino)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 43 | N-(4-((4-(isopropylamino)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin- |
| 2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 44 | N-(4-((5-methoxy-4-((2-methoxyethyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 45 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(methylamino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 46 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 47 | N-(4-((5-methoxy-4-(methyl(2-(methylamino)ethyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 48 | N-(4-((4-((2-methoxyethyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 49 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 50 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 51 | N-(4-((4-(4-ethylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 52 | N-(4-((4-(4-isopropylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 53 | N-(4-((4-(ethylamino)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 54 | N-(4-((4-(isopropyl(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 55 | N-(4-((5-methoxy-4-(4-(2-methoxyethyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 56 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6,7-dihydro-5H- |
| cyclopenta[d]pyrimidin-2-yl)thio)phenyl)acetamide | |
| 57 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6,7-dihydro-5H- |
| cyclopenta[d]pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 58 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(methylamino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 59 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 60 | N-(4-((4-((2-methoxyethyl)(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 61 | N-(4-((4-((2-methoxyethyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 62 | N-(4-((4-(4-(tert-butyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 63 | N-(4-((4-(dimethylamino)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 64 | N-(4-((4-(ethyl(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 65 | N-(4-((4-(ethylamino)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 66 | N-(4-((4-(isopropylamino)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 67 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)butyramide | |
| 68 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 69 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 70 | N-(4-((4-(diethylamino)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 71 | N-(4-((4-(dimethylamino)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin- |
| 2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 72 | N-(4-((4-(ethyl(methyl)amino)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 73 | N-(4-((5-methoxy-4-((2-methoxyethyl)(methyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 74 | N-(4-((5-methoxy-4-((2-methoxyethyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 75 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 76 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 77 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide N-(4-((4- | |
| (diethylamino)-5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- | |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 78 | N-(4-((4-(dimethylamino)-5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 79 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 80 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 81 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 82 | N-(4-((5-ethoxy-4-(ethyl(methyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 83 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 84 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 85 | N-(4-((5-methoxy-4-(4-(2-methoxyethyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 86 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 87 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-ethoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 88 | N-(4-((4-(diethylamino)-5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 89 | N-(4-((4-(dimethylamino)-5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 90 | N-(4-((5-ethoxy-4-((2-methoxyethyl)(methyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 91 | N-(4-((5-ethoxy-4-((2-methoxyethyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 92 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 93 | N-(4-((5-ethoxy-4-(4-(2-methoxyethyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 94 | N-(4-((5-ethoxy-4-(ethyl(methyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 95 | 3-(5-fluoro-2-methylpyridin-4-yl)-6-(4-(2-(piperidin-1-yl)ethoxy)phenyl)pyrazolo[1,5- |
| α]pyrimidine | |
| 96 | N-(4-((4-(4-(3-(dimethylamino)propanoyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 97 | N-(4-((4-(4-(cyclopropylmethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 98 | N-(4-((4-(4-(dimethylglycyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 99 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 100 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 101 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(1-methylpiperidin-4- |
| yl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 102 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2- |
| (methylamino)ethyl)piperazin-1-yl)pyrimidin-2- | |
| yl)thio)phenyl)cyclopropanecarboxamide | |
| 103 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-phenylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 104 | N-(4-((5-methoxy-4-(4-(2-methoxyacetyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 105 | N-(4-((5-methoxy-4-(4-(3-methoxyl)ropanoyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide(1:1:1:1:1:1:1:1:1:1) | |
| 106 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 107 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-ethoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 108 | N-(4-((4-(diethylamino)-5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 109 | N-(4-((4-(diethylamino)-5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 110 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 111 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 112 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 113 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 114 | N-(4-((5-ethoxy-4-(4-(2-methoxyethyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 115 | N-(4-((5-ethoxy-4-(ethyl(methyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 116 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 117 | N-(4-((4-(4-(3-(dimethylamino)propanoyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 118 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-ethoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 119 | N-(4-((5-ethoxy-4-(4-(2-methoxyacetyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 120 | N-(4-((5-ethoxy-4-(4-(2-methoxyethyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 121 | N-(4-((5-ethoxy-4-(4-(3-methoxyl)ropanoyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 122 | N-(4-((5-ethoxy-4-(ethyl(methyl)amino)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 123 | N-(4-((5-ethyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6,7-dihydro-5H- |
| cyclopenta[d]pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 124 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-4- |
| ylamino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 125 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 126 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclobutanecarboxamide | |
| 127 | N-(4-((4-(4-(cyclopropylmethyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 128 | N-(4-((4-(4-(cyclopropylmethyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 129 | N-(4-((4-(4-(dimethylglycyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 130 | N-(4-((4-(4-(dimethylglycyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 131 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(1-methylpiperidin-4- |
| yl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 132 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(1-methylpiperidin-4- |
| yl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 133 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(piperidin-4-yl)piperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 134 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 135 | N-(4-((5-ethoxy-4-(4-(2-methoxyacetyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 136 | N-(4-((5-ethoxy-4-(4-(3-methoxyl)ropanoyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 137 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-((1-methylpiperidin-4- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 138 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 139 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 140 | N-(4-((4-(4-(2-methoxyethyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 141 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 142 | N-(4-((4-(dimethylamino)-5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 143 | N-(4-((4-(dimethylamino)-5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 144 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(piperidin-4-yl)piperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 145 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 146 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(1-methylpiperidin-4- |
| yl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 147 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 148 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 149 | N-(4-((4-(4-(2-methoxyacetyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 150 | N-(4-((4-(4-(2-methoxyethyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 151 | N-(4-((4-(4-(3-(dimethylamino)propanoyl)piperazin-1-yl)-5-ethoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 152 | N-(4-((4-(4-(3-(dimethylamino)propanoyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 153 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 154 | N-(4-((4-(isopropyl(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 155 | N-(4-((5-(ethylthio)-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 156 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 157 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(1-methylpiperidin-4- |
| yl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 158 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 159 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 160 | N-(4-((4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1-yl)-5- |
| (methylthio)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 161 | N-(4-((4-(4-(2-methoxyacetyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 162 | N-(4-((4-(4-hydroxypiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 163 | N-(4-((4-(diethylamino)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 164 | N-(4-((4-(dimethylamino)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 165 | N-(4-((4-(ethyl(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 166 | N-(4-((4-(isopropyl(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopentanecarboxamide | |
| 167 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)cyclopentanecarboxamide | |
| 168 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(pyrrolidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 169 | methyl(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)carbamate | |
| 170 | N-(4-((4-(azepan-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 171 | N-(4-((5-chloro-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 172 | N-(4-((5-chloro-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 173 | 1-methyl-3-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)urea | |
| 174 | 2-((4-aminophenyl)thio)-5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-6-(piperidin-1- |
| yl)pyrimidin-4-amine | |
| 175 | 5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-2-(methylthio)-6-(piperidin-1-yl)pyrimidin-4- |
| amine | |
| 176 | 5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-2-(phenylthio)-6-(piperidin-1-yl)pyrimidin-4- |
| amine | |
| 177 | 5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-6-(piperidin-1-yl)-2-(p-tolylthio)pyrimidin-4- |
| amine | |
| 178 | N-(4-((4-(3,6-dihydropyridin-1(2H)-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 179 | N-(4-((4-(4,4-difluoropiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 180 | N-(4-((4-(4-fluoro-3,6-dihydropyridin-1(2H)-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 181 | N-(4-((4-(4-fluoropiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 182 | N-(4-((5-chloro-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 183 | N-(4-((5-ethoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(1-methylpiperidin-4- |
| yl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 184 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(1-methylpiperidin-4- |
| yl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 185 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 186 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperidin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 187 | 2-((4-(dimethylamino)phenyl)thio)-5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| (piperidin-1-yl)pyrimidin-4-amine | |
| 188 | 5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-2-((4-(methylamino)phenyl)thio)-6-(piperidin- |
| 1-yl)pyrimidin-4-amine | |
| 189 | N-(4-((4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 190 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 191 | N-(4-((4-(4-acetylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 192 | N-(4-((4-(azepan-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 193 | N-(4-((4-(diethylamino)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 194 | N-(4-((4-(dimethylamino)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin- |
| 2-yl)thio)phenyl)acetamide | |
| 195 | N-(4-((4-(ethyl(methyl)amino)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 196 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-methylpiperidin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 197 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperazin-1-yl)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 198 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(pyrrolidin-1-yl)pyrimidin- |
| 2-yl)thio)phenyl)acetamide | |
| 199 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 200 | N-(4-((5-methoxy-4-(4-(2-methoxyethyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 201 | N-(4-((4-(3,6-dihydropyridin-1(2H)-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 202 | N-(4-((4-(4,4-difluoropiperidin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 203 | N-(4-((4-(4-acetylpiperidin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 204 | N-(4-((4-(4-aminopiperidin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 205 | N-(4-((4-(4-ethylpiperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 206 | N-(4-((4-(4-fluoro-3,6-dihydropyridin-1(21/)-yl)-5-methoxy-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 207 | N-(4-((4-(4-fluoropiperidin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 208 | N-(4-((4-(4-hydroxypiperidin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 209 | N-(4-((4-(isopropyl(methyl)amino)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 210 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((2-(thiazol-2-yl)propan-2-yl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 211 | N-(4-((5-methoxy-4-(4-(2-methoxyacetyl)piperazin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 212 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(2-methylpiperidin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 213 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-5-methyl-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 214 | N-(4-((4-(4-(dimethylamino)piperidin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 215 | N-(4-((4-(4-ethylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 216 | N-(4-((4-(4-isobutyrylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 217 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(2-methylpiperidin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 218 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-propionylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 219 | 1-(5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)-2-(methylthio)pyrimidin-4- |
| yl)piperidin-4-ol | |
| 220 | 5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-2-(methylthio)-6-morpholinopyrimidin-4- |
| amine | |
| 221 | N-(4-((4-(4-(dimethylglycyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 222 | N-(4-((4-(4-(dimethylglycyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 223 | N-(4-((4-(4-acetylpiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 224 | N-(4-((4-(4-isobutyrylpiperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 225 | N-(4-((4-(4-isopropylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 226 | N-(4-((4-(4-isopropylpiperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 227 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(methylglycyl)piperazin- |
| 1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 228 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(methylamino)piperidin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 229 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(methylglycyl)piperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 230 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-propionylpiperazin-1- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 231 | N-(4-((4-(4-(dimethylamino)piperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 232 | N-(4-((4-(4-aminopiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 233 | N-(4-((4-(4-hydroxy-4-methylpiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 234 | N-(4-((4-(diethylamino)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 235 | 1-(4-((4-(4-hydroxypiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)-3-methylurea | |
| 236 | 1-methyl-3-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| morpholinopyrimidin-2-yl)thio)phenyl)urea | |
| 237 | 1-(5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)-2-(phenylthio)pyrimidin-4- |
| yl)piperidin-4-ol | |
| 238 | 5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-6-(piperidin-1-yl)-2-(p-tolyloxy)pyrimidin-4- |
| amine | |
| 239 | 5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholino-2-(phenylthio)pyrimidin-4- |
| amine | |
| 240 | N4-isopropyl-N4,5-dimethyl-N6-(5-methyl-1H-pyrazol-3-yl)-2-(phenylthio)pyrimidine- |
| 4,6-diamine | |
| 241 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 242 | 1-(4-((4-(4-(2-methoxyacetyl)piperazin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)-3-methylurea | |
| 243 | 1-(4-((4-(isopropyl(methyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)-3-methylurea | |
| 244 | 1-(4-(2((4-aminophenyl)thio)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperazin-1-yl)-2-methoxyethan-1-one | |
| 245 | 2-methoxy-1-(4-(5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)-2- |
| (phenylthio)pyrimidin-4-yl)piperazin-1-yl)ethan-1-one | |
| 246 | 1-(2-((4-aminophenyl)thio)-5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin- |
| 4-yl)piperidin-4-ol | |
| 247 | 6-(1-(2-(dimethylamino)ethyl)piperidin-4-yl)-2-((4-(ethylsulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 248 | 6-(4-(2-(8-azabicyclo[3.2.1]octan-8-yl)ethyl)piperazin-1-yl)-2-((4- |
| (benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| amine | |
| 249 | 6-(4-(2-(9-azabicyclo[3.2.1]nonan-9-yl)ethyl)piperidin-1-yl)-2-((4- |
| (ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| |
|
| 250 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((2-(pyrimidin-4-yl)propan-2-yl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 251 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-((2- |
| (dimethylamino)ethyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 252 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-ethoxy-2-((4- |
| (ethylsulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 253 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((oxazol-2-ylmethyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 254 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((thiazol-2-ylmethyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 255 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-(phenylsulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 256 | 2-((4-((2,4-difluorobenzyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 257 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-ethoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| (piperazin-1-yl)pyrimidin-4-amine | |
| 258 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-6-(4-(2- |
| (methyl(phenyl)amino)ethyl)piperazin-1-yl)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| amine | |
| 259 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5- |
| isopropoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 260 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5- |
| methoxy-N-(1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 261 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5- |
| methyl-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 262 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-N-(5- |
| methyl-1H-pyrazol-3-yl)-5-propoxypyrimidin-4-amine | |
| 263 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-N-(5- |
| methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 264 | 2-((4-(benzylsulfonyl)phenyl)thio)-N4,N4-diethyl-5-methoxy-N6-(5-methyl-1H-pyrazol- |
| 3-yl)pyrimidine-4,6-diamine | |
| 265 | 2-((4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| (piperidin-1-yl)pyrimidin-4-amine | |
| 266 | 2-((4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| morpholinopyrimidin-4-amine | |
| 267 | 5-ethoxy-2-((4-(ethylsulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)-6-(piperazin- |
| 1-yl)pyrimidin-4-amine | |
| 268 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-((2- |
| fluorobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 269 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((3-nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 270 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((4-nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 271 | N4,N4-diethyl-2-((4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N6-(5-methyl-1H-pyrazol-3- |
| yl)pyrimidine-4,6-diamine | |
| 272 | 1-(3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)phenyl)ethan-1-one | |
| 273 | 1-(2-(4-(2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperazin-1-yl)ethyl)piperidin-4-ol | |
| 274 | 3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)phenol | |
| 275 | 2-((4-((2,3-difluorobenzyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 276 | 2-((4-((2,5-difluorobenzyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 277 | 2((4-((2-fluorobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)- |
| 6-morpholinopyrimidin-4-amine | |
| 278 | 2-((4-((3-(benzyloxy)benzyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 279 | 2-((4-(([1,1′-biphenyl]-3-ylmethyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 280 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2- |
| (piperidin-1-yl)ethyl)piperazin-1-yl)pyrimidin-4-amine | |
| 281 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(4-(2-(2,6-dimethylpiperidin-1-yl)ethyl)piperazin- |
| 1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 282 | 5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholino-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 283 | 6-(4-(2-(8-azabicyclo[3.2.1]octan-8-yl)ethyl)piperazin-1-yl)-2-((4- |
| (benzylsulfonyl)phenyl)thio)-5-ethoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| amine | |
| 284 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((2-nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 285 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((3-(trifluoromethyl)benzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 286 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-5-ethoxy-2-((4- |
| (ethylsulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 287 | N2-(4-(benzylsulfonyl)phenyl)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5- |
| methoxy-N4-(5-methyl-1H-pyrazol-3-yl)pyrimidine-2,4-diamine | |
| 288 | 3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzonitrile | |
| 289 | (R)-2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(2- |
| methylmorpholino)pyrimidin-4-amine | |
| 290 | 2-((4-(benzylsulfonyl)-2-fluorophenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 291 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(1- |
| methylpiperidin-4-yl)piperazin-1-yl)pyrimidin-4-amine | |
| 292 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2- |
| (4-methylpiperazin-1-yl)ethyl)piperidin-1-yl)pyrimidin-4-amine | |
| 293 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2- |
| (pyrrolidin-1-yl)ethyl)piperazin-1-yl)pyrimidin-4-amine | |
| 294 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(4- |
| methylpiperazin-1-yl)piperidin-1-yl)pyrimidin-4-amine | |
| 295 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4- |
| (pyridin-4-ylmethyl)piperazin-1-yl)pyrimidin-4-amine | |
| 296 | 6-(4-(1-benzylpiperidin-4-yl)piperazin-1-yl)-2-((4-(benzylsulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 297 | 6-(4-(2-(azepan-1-yl)ethyl)piperazin-1-yl)-2-((4-(benzylsulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 298 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2-((4-((2- |
| methoxybenzyl)sulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 299 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2-((4-((3- |
| methoxybenzyl)sulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- |
|
| 300 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((3-methylbenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 301 | 6-(4-benzylpiperazin-1-yl)-2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 302 | methyl 3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl- |
| 1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzoate | |
| 303 | 3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzamide | |
| 304 | N-(2-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)phenyl)acetamide | |
| 305 | 2-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzonitrile | |
| 306 | (R)-2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(3- |
| methylmorpholino)pyrimidin-4-amine | |
| 307 | (S)-2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(2- |
| methylmorpholino)pyrimidin-4-amine | |
| 308 | (S)-2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(3- |
| methylmorpholino)pyrimidin-4-amine | |
| 309 | 2-((4-((2-(dimethylamino)benzyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4- |
|
| 310 | 2-((4-(([1,1′-biphenyl]-2-ylmethyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 311 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(8-oxa-3-azabicyclo[3.2.1]octan-3-yl)-5-methoxy- |
| N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 312 | 6-(4-(2-(azetidin-1-yl)ethyl)piperazin-1-yl)-2-((4-(benzylsulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 313 | 6-(4-(2-(benzyl(methyl)amino)ethyl)piperazin-1-yl)-5-methoxy-2-((4-((3- |
| methoxybenzyl)sulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 314 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-((2-fluoro-3- |
| nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 315 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-((3- |
| fluorobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 316 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((2-(trifluoromethyl)benzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 317 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((2-methylbenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 318 | 2-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzoic acid | |
| 319 | 3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzoic acid | |
| 320 | methyl 2-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl- |
| 1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzoate | |
| 321 | 2-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)-N,N- | |
| dimethylbenzamide | |
| 322 | 2-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzamide | |
| 323 | 3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)-N,N- | |
| dimethylbenzamide | |
| 324 | N-(3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)phenyl)acetamide | |
| 325 | (R)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(2-methylmorpholino)-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 326 | (R)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(3-methylmorpholino)-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 327 | (S)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(2-methylmorpholino)-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 328 | (S)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(3-methylmorpholino)-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 329 | 2-((4-((3-(dimethylamino)benzyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 330 | 2-((4-(benzylsulfonyl)-2-chlorophenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 331 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4- |
| (piperidin-4-yl)piperazin-1-yl)pyrimidin-4-amine | |
| 332 | 6-(4-(2-(8-azabicyclo[3.2.1]octan-8-yl)ethyl)piperidin-1-yl)-2-((4- |
| (benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| amine | |
| 333 | 2-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)-6-nitrobenzonitrile | |
| 334 | 1-(2-((4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-ol | |
| 335 | (5)-2-((2-fluoro-4-((3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 336 | 2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 337 | 2-((2-fluoro-4-((3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 338 | 2-((4-((2-chloro-3-nitrobenzyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 339 | 2-((4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-(piperidin-1-yl)pyrimidin-4-amine | |
| 340 | 2-((4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 341 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((2-fluoro-3- |
| nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 342 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 343 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-((2-fluoro-5- |
| nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 344 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-((3-fluoro-5- |
| nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 345 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2-((4-((2-methoxy-3- |
| nitrobenzyl)sulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 346 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2-((4-((2-methoxy-5- |
| nitrobenzyl)sulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 347 | 2-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)-6-nitrophenol | |
| 348 | (R)-2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 349 | (R)-2-((2-fluoro-4-((3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 350 | (S)-2-((2-chloro-4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 351 | 2-((2-chloro-4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| (4-methylpiperazin-1-yl)pyrimidin-4-amine | |
| 352 | 2-((2-chloro-4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| morpholinopyrimidin-4-amine | |
| 353 | 2-((2-chloro-4-(ethylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 354 | 2-((2-chloro-4-(ethylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperidin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 355 | 2-((2-chloro-4-(methylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 356 | 2-((2-chloro-4-(methylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperidin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 357 | 6-(3-oxa-8-azabicyclo[3.2.1]octan-8-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-2- |
| ((4-((3-nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 358 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4- |
| (methylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| amine | |
| 359 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-(ethylsulfonyl)-2- |
| fluorophenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 360 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-2-((2-fluoro-4- |
| (methylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| amine | |
| 361 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-2-((4-(ethylsulfonyl)-2- |
| fluorophenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 362 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((2-(pyrazin-2-yl)propan-2-yl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 363 | (R)-(4-(5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-yl)morpholin-3-yl)methanol | |
| 364 | (R)-2-((2-chloro-4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 365 | (R)-2-((4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 366 | (R)-6-(3-ethylmorpholino)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 367 | (S)-2-((4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 368 | (S)-2-((4-(ethylsulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 369 | 2-((2-chloro-4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| (piperidin-1-yl)pyrimidin-4-amine | |
| 370 | 2-((2-fluoro-4-(methylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)- |
| 6-(piperidin-1-yl)pyrimidin-4-amine | |
| 371 | 2-((2-fluoro-4-(methylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)- |
| 6-morpholinopyrimidin-4-amine | |
| 372 | 2-((4-((2,3-difluorobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-6-(piperidin-1-yl)pyrimidin-4-amine | |
| 373 | 2-((4-(ethylsulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| (piperidin-1-yl)pyrimidin-4-amine | |
| 374 | 2-((4-(ethylsulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| morpholinopyrimidin-4-amine | |
| 375 | 5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-2-((4-(methylsulfonyl)phenyl)thio)-6- |
| (piperidin-1-yl)pyrimidin-4-amine | |
| 376 | 6-(3-ethylmorpholino)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 377 | 1-(2-((2-fluoro-4-((1-(2-fluoro-3-nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)-N,N-dimethylpiperidine- | |
| 4-carboxamide compound | |
| 378 | 1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)-N,N-dimethylpiperidine- | |
| 4-carboxamide | |
| 379 | 1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidine-4- | |
| carboxamide | |
| 380 | 2,2,2-trifluoro-N-(1-(2-((2-fluoro-4-((1-(2-fluoro-3- |
| nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- | |
| yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 381 | 2,2,2-trifluoro-N-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4- | |
| yl)piperidin-4-yl)acetamide | |
| 382 | 2,2,2-trifluoro-N-(2-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4- | |
| yl)piperidin-4-yl)ethyl)acetamide | |
| 383 | 2-(1-(2-((2-fluoro-4-((1-(2-fluoro-3-nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 384 | 2-(1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)- |
| 5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4- | |
| yl)acetamide | |
| 385 | 2-(1-(2-((2-fluoro-4 (2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 386 | N-(1-(2-((2-fluoro-4-((1-(2-fluoro-3-nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 387 | N-(1-(2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 388 | N-(2-(1-(2-((2-fluoro-4-((1-(2-fluoro-3-nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)- |
| 5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4- | |
| yl)ethyl)acetamide | |
| 389 | N-(2-(1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4- | |
| yl)piperidin-4-yl)ethyl)acetamide | |
| 390 | N-(2-(1-(2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-6- |
| ((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)ethyl)acetamide | |
| 391 | 1-(4-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)- |
| 5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperazin-1-yl)ethan-1- | |
| one | |
| 392 | 1-(4-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperazin-1-yl)ethan-1- | |
| one | |
| 393 | 2-((2-fluoro-4-((1-(2,3,5,6-tetrafluorophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 394 | 2-((2-fluoro-4-((1-(2-fluoro-3-nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(piperazin-1-yl)pyrimidin-4-amine | |
| 395 | 2-((2-fluoro-4-((1-(2-fluoro-3-nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 396 | 2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(piperazin-1-yl)pyrimidin-4-amine | |
| 397 | 2-((4-((2-(1H-imidazol-2-yl)propan-2-yl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 398 | 6-(3-aminopyrrolidin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 399 | 6-(3-aminopyrrolidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- |
|
| 400 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((1-(2-fluoro-3- |
| nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 401 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-2-((2-fluoro-4-((1-(2-fluoro-3- |
| nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 402 | 6-(4-(2-aminoethyl)piperidin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6- |
| tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 403 | 6-(4-(aminomethyl)piperidin-1-yl)-2-((2-fluoro-4-((1-(2-fluoro-3- |
| nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 404 | 6-(4-(aminomethyl)piperidin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6- |
| tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 405 | 6-(4-(aminomethyl)piperidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan- |
| 2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 406 | 6-(4-aminopiperidin-1-yl)-2-((2-fluoro-4-((1-(2-fluoro-3- |
| nitrophenyl)cyclopropyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 407 | 6-(4-aminopiperidin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 408 | 6-(4-aminopiperidin-1-yl)-2-((2-fluoro-4-((2-fluoro-3- |
| nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 409 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((2,3,6-trifluorobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 410 | 2-fluoro-3-(((3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| morpholinopyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzonitrile | |
| 411 | (S)-(4-(5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)-2-((4-((3- |
| nitrobenzyl)sulfonyl)phenyl)thio)pyrimidin-4-yl)morpholin-3-yl)methanol | |
| 412 | (S)-2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 413 | (S)-2-((2-fluoro-4-(methylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 414 | (S)-2-((4-((2,3-difluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)-6-(3-methylmorpholino)pyrimidin-4-amine | |
| 415 | 2-((2-fluoro-4-((2,3,5,6-tetrafluorobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 416 | 2-((2-fluoro-4-((2-fluoro-3-(trifluoromethyl)benzyl)sulfonyl)phenyl)thio)-5-methoxy-N- |
| (5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 417 | 2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)-6-(piperidin-1-yl)pyrimidin-4-amine | |
| 418 | 2-((4-((2,3-difluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-(piperidin-1-yl)pyrimidin-4-amine | |
| 419 | 2-((4-((2,3-difluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 420 | 2-((4-((2,3-difluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 421 | 2-((4-((3-bromo-2-fluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 422 | 2-((4-((3-chloro-2-fluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 423 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((2,3,6- |
| trifluorobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 424 | methyl 2-fluoro-3-(((3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| morpholinopyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzoate | |
| 425 | 2-fluoro-3-(((3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| morpholinopyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)-N,N-dimethylbenzamide | |
| 426 | 2-fluoro-3-(((3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| morpholinopyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benz amide | |
| 427 | 3-fluoro-N-(2-fluoro-3-nitrophenyl)-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3- |
| yl)amino)-6-morpholinopyrimidin-2-yl)thio)benzamide | |
| 428 | (S)-(4-(2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)morpholin-3-yl)methanol | |
| 429 | (E)-2-((2-fluoro-4-(2-fluoro-3-nitrostyryl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 430 | 2-((2-fluoro-4-((2-fluoro-3-iodobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 431 | 2-((2-fluoro-4-((2-fluoro-3-nitrophenyl)ethynyl)phenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 432 | 2-((2-fluoro-4-((3-nitrophenyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 433 | 2-((2-fluoro-4-((fluoro(2-fluoro-3-nitrophenyl)methyl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 434 | 2-((2-fluoro-4-(2-fluoro-3-nitrophenethyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 435 | 2-((4-((3-amino-2-fluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 436 | 2-((4-((3-brom o-2-fluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-(piperidin-1-yl)pyrimidin-4-amine | |
| 437 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((fluoro(2-fluoro-3- |
| nitrophenyl)methyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 438 | 2-fluoro-3-nitrophenyl 3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| morpholinopyrimidin-2-yl)thio)benzoate | |
| 439 | 2-(2-fluoro-3-nitrophenyl)-N-(3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3- |
| yl)amino)-6-morpholinopyrimidin-2-yl)thio)phenyl)acetamide | |
| 440 | 2-fluoro-N-(3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| morpholinopyrimidin-2-yl)thio)phenyl)-3-nitrobenzamide | |
| 441 | 3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin- |
| 2-yl)thio)-N-(3-nitrophenyl)benzenesulfonamide | |
| 442 | 2-fluoro-3-(((3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| (piperidin-1-yl)pyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzonitrile | |
| 443 | 2-fluoro-3-(fluoro((3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6- |
| morpholinopyrimidin-2-yl)thio)phenyl)sulfonyl)methyl)benzonitrile | |
| 444 | 3-(((4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)-3-fluorophenyl)sulfonyl)methyl)-2- | |
| fluorobenzonitrile | |
| 445 | 2-(2-fluoro-3-nitrophenyl)-1-(3-fluoro-4-((5-methoxy-4-((5-methyl-1H-pyrazol-3- |
| yl)amino)-6-morpholinopyrimidin-2-yl)thio)phenyl)ethan-1-one | |
| 446 | ((3S)-4-(2-((2-fluoro-4-((fluoro(2-fluoro-3-nitrophenyl)methyl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)morpholin-3-yl)methanol | |
| 447 | (R)-( 1-(2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-2-yl)methanol | |
| 448 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(piperidin-1-yl)pyrimidin-4-amine | |
| 449 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 450 | 2-((2-fluoro-4-(7-nitrobenzofuran-2-yl)phenyl)thio)-5-methoxy-N-(5-methyl-1 H- |
| pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 451 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3- |
| nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 452 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-2-((2-fluoro-4-((2-fluoro-3- |
| nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 453 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-2-((2-fluoro-4-((fluoro(2-fluoro-3- |
| nitrophenyl)methyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 454 | (S)-(4-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)morpholin-3-yl)methanol | |
| 455 | 2-((2-fluoro-4-((2-(2,3,5,6-tetrafluoro-4-methylphenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- | |
| morpholinopyrimidin-4-amine | |
| 456 | 2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 457 | 2-((2-fluoro-4-((2-fluoro-3-nitrophenethyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 458 | 2-((2-fluoro-4-((fluoro(2,3,5,6-tetrafluorophenyl)methyl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 459 | 2-((4-((2-(2,5-difluoro-3-nitrophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 460 | 2-((4-((2-(3-bromo-2-fluorophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 461 | 2-((4-((2-(3-bromo-2-fluorophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 462 | 2-((4-((2-(3-chloro-2,5-difluorophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 463 | 2-((4-((2-(3-chloro-2-fluorophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 464 | 2-((4-((2-(3-chloro-2-fluorophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 465 | 2-((4-((difluoro(2,3,5,6-tetrafluorophenyl)methyl)sulfonyl)-2-fluorophenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 466 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6-tetrafluoro-4- |
| methylphenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol- | |
| 3-yl)pyrimidin-4-amine | |
| 467 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6- |
| tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 468 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((fluoro(2,3,5,6- |
| tetrafluorophenyl)methyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 469 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3- |
| nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 470 | 2-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)-N,N- | |
| dimethyl acetamide | |
| 471 | N-(2-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)- |
| 5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4- | |
| yl)ethyl)acetamide | |
| 472 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-(4-(2-((4-methoxybenzyl)(methyl)amino)ethyl)piperidin-1-yl)-N-(5-methyl- | |
| 1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 473 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-methylpiperazin-1-yl)pyrimidin-4-amine | |
| 474 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(piperazin-1-yl)pyrimidin-4-amine | |
| 475 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N4,N4-dimethyl-N6-(5-methyl-1H-pyrazol-3-yl)pyrimidine-4,6-diamine | |
| 476 | 2-((4-((2-(2,5-difluoro-3-nitrophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-6-(4- |
| (2-(dimethylamino)ethyl)piperidin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 477 | 2-((4-((2-(3-chloro-2,5-difluorophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-6-(4- |
| (2-(dimethylamino)ethyl)piperidin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 478 | 2-((4-((2-(3-chloro-2-fluorophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperidin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 479 | 2-((4-((3-chloro-2,5,6-trifluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 480 | 6-(3-(dimethylamino)piperidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan- |
| 2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 481 | 6-(4-((dimethylamino)methyl)piperidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3- |
| nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 482 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6- |
| tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 483 | 6-(4-(3-(dimethylamino)propyl)piperidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3- |
| nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 484 | 6-(4-(dimethylamino)piperidin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6- |
| tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 485 | 6-(4-(dimethylamino)piperidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan- |
| 2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 486 | 2-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetic acid | |
| 487 | methyl 2-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4- | |
| yl)piperidin-4-yl)acetate | |
| 488 | 2-(dimethylamino)-1-(4-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4- | |
| yl)piperazin-1-yl)ethan-1-one | |
| 489 | 2-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)ethan-1-ol | |
| 490 | 2-((2-fluoro-4-(((perfluorophenyl)methyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 491 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-(3-methoxypyrrolidin-1-yl)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| amine | |
| 492 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-(4-(2-methoxyethyl)piperidin-1-yl)-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 493 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2-(methylamino)ethyl)piperidin-1- | |
| yl)pyrimidin-4-amine | |
| 494 | 2-((2-fluoro-4-((2-(perfluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5- |
| methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 495 | 2-((4-((2-(3-chloro-2,5,6-trifluorophenyl)propan-2-yl)sulfonyl)-2-fluorophenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 496 | 2-((4-((3-chloro-2,5,6-trifluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 497 | 2-((4-((3-chloro-2,5,6-trifluorobenzyl)sulfonyl)-2-fluorophenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperidin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 498 | 6-(3-(dimethylamino)pyrrolidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3- |
| nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 499 | 6-(4-(2-aminoethyl)piperidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 500 | (S)-N-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)- |
| 5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)pyrrolidin-3- | |
| yl) |
|
| 501 | 2-(1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)-N,N- | |
| dimethylacetamide | |
| 502 | 2-(4-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperazin-1-yl)-N,N- | |
| dimethylacetamide | |
| 503 | N-(1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 504 | N-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 505 | 1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)pyrrolidin-3-ol | |
| 506 | (R)-6-(3-(dimethylamino)pyrrolidin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6- |
| tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 507 | (R)-6-(3-(dimethylamino)pyrrolidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3- |
| nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 508 | (S)-6-(3-(dimethylamino)pyrrolidin-1-yl)-2-((2-fluoro-4-((2-(2,3,5,6- |
| tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 509 | (S)-6-(3-(dimethylamino)pyrrolidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3- |
| nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 510 | 2-((4-(tert-butylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 511 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((2- |
| (perfluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 512 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-2-((2-fluoro-4-((2- |
| (perfluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- | |
| pyrazol-3-yl)pyrimidin-4-amine | |
| 513 | 6-(4-(dimethylamino)piperidin-1-yl)-2-((2-fluoro-4-((2-(perfluorophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 514 | 1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl acetate | |
| 515 | 1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)pyrrolidin-3-yl acetate | |
| 516 | 1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)-N,N-dimethylpiperidine- | |
| 4-carboxamide | |
| 517 | 1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidine-4- | |
| carboxamide | |
| 518 | N-(1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)-N- | |
| methyl acetamide | |
| 519 | N-(1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)pyrrolidin-3- | |
| yl)acetamide | |
| 520 | N-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)-N- | |
| methylacetamide | |
| 521 | 1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-ol | |
| 522 | 1-(2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)pyrrolidin-3-ol | |
| 523 | 2-((2-fluoro-4-((2-(2,3,5,6-tetrafluorophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-(4-methoxyl)iperidin-1-yl)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 524 | 2-((2-fluoro-4-((4-methoxybenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-(piperidin-1-yl)pyrimidin-4-amine | |
| 525 | 2-((2-fluoro-4-((4-methoxybenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H- |
| pyrazol-3-yl)-6-morpholinopyrimidin-4-amine | |
| 526 | 5-methoxy-2-((4-((4-methoxybenzyl)sulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3- |
| yl)-6-(piperidin-1-yl)pyrimidin-4-amine | |
| 527 | 5-methoxy-2-((4-((4-methoxybenzyl)sulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3- |
| yl)-6-morpholinopyrimidin-4-amine | |
| 528 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((2-fluoro-4-((2-(2,3,6-trifluoro-5- |
| nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 529 | 2-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)-N- | |
| methyl acetamide | |
| 530 | 2-(1-(2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 531 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2-((1,1,1-trifluoropropan-2- | |
| yl)amino)ethyl)piperidin-1-yl)pyrimidin-4-amine | |
| 532 | 2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2-yl)sulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(methylamino)piperidin-1-yl)pyrimidin-4- | |
| amine | |
| 533 | 2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-6-(piperidin- |
| 1-yl)-N-(1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 534 | 2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-6-morpholino- |
| N-(1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 535 | 2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)-6-(4-methylpiperazin-1-yl)pyrimidin-4-amine | |
| 536 | 2-((2-fluoro-4-((2-fluoro-3-nitrobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl- |
| 1H-pyrazol-3-yl)-6-(piperazin-1-yl)pyrimidin-4-amine | |
| 537 | 2-((4-((2-cyclopropylpropan-2-yl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 538 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((2-(pyridin-3-yl)propan-2-yl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 539 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-24-((42-(thiophen-2-yl)propan-2-yl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 540 | 6-(4-aminopiperidin-1-yl)-2-((2-fluoro-4-((2-(2-fluoro-3-nitrophenyl)propan-2- |
| yl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 541 | 1-(5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)-2-(p-tolylthio)pyrimidin-4- |
| yl)piperidin-4-ol | |
| 542 | 2-((4-aminophenyl)thio)-5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| morpholinopyrimidin-4-amine | |
| 543 | 2((4-aminophenyl)thio)-N4-isopropyl-N4,5-dimethyl-N6-(5-methyl-1H-pyrazol-3- |
| yl)pyrimidine-4,6-diamine | |
| 544 | N4-isopropyl-N4,5-dimethyl-N6-(5-methyl-1H-pyrazol-3-yl)-2-(methylthio)pyrimidine- |
| 4,6-diamine | |
| 545 | N-(4-((4-((2-hydroxyethyl)(isopropyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 546 | N-(4-((4-(ethyl(isopropyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 547 | N-(4-((4-(isopropyl(2-methoxyethyl)amino)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 548 | N-(4-((5-chloro-2-(4-hydroxypiperidin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)thio)phenyl)acetamide | |
| 549 | N-(4-((5-chloro-4-(4-hydroxypiperidin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 550 | 2-methoxy-1-(4-(5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)-2-(p- |
| tolylthio)pyrimidin-4-yl)piperazin-1-yl)ethan-1-one | |
| 551 | 5-methyl-N-(5-methyl-1H-pyrazol-3-yl)-6-morpholino-2-(p-tolylthio)pyrimidin-4- |
| amine | |
| 552 | N-(4-((4-(4-hydroxypiperidin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 553 | N-(4-((4-(4-hydroxypiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 554 | N-(4-((5-ethoxy-4-(4-hydroxypiperidin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 555 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2- |
| morpholinoethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 556 | 2-methoxy-1-(4-(5-methyl-6-((5-methyl-1H-pyrazol-3-yl)amino)-2- |
| (methylthio)pyrimidin-4-yl)piperazin-1-yl)ethan-1-one | |
| 557 | 1-(5-ethoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)-2-(methylthio)pyrimidin-4- |
| yl)piperidin-4-ol | |
| 558 | 5-ethoxy-N-(5-methyl-1H-pyrazol-3-yl)-2-(methylthio)-6-(piperidin-1-yl)pyrimidin-4- |
| amine | |
| 559 | 5-ethoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-methylpiperazin-1-yl)-2- |
| (methylthio)pyrimidin-4-amine | |
| 560 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-ethoxy-N-(5-methyl-1H-pyrazol-3-yl)- |
| 2-(methylthio)pyrimidin-4-amine | |
| 561 | N4-isopropyl-N4,5-dimethyl-N6-(5-methyl-1H-pyrazol-3-yl)-2-(p-tolylthio)pyrimidine- |
| 4,6-diamine | |
| 562 | (S)-N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(3- |
| methylmorpholino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 563 | N-(4-((4-(3-hydroxy-8-azabicyclo[3.2.1]octan-8-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 564 | N-(4-((4-(3-hydroxy-8-azabicyclo[3.2.1]octan-8-yl)-5-methyl-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 565 | N-(4-((4-(4-(2-(dimethylamino)ethoxy)piperidin-1-yl)-5-methyl-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 566 | N-(4-((4-(4-hydroxy-2-methylpiperidin-1-yl)-5-methyl-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 567 | N-(4-((5-ethoxy-4-(4-hydroxypiperidin-1-yl)-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)cyclopropanecarboxamide | |
| 568 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2- |
| morpholinoethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 569 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-(piperidin-1- |
| yl)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 570 | 2-((4-aminophenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N- |
| (5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 571 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-(methylthio)pyrimidin-4-amine | |
| 572 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-(phenylthio)pyrimidin-4-amine | |
| 573 | (R)-N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(3- |
| methylmorpholino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 574 | N-(4-((4-(3-hydroxyl)yrrolidin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 575 | N-(4-((4-(4-(2-(dimethylamino)ethoxy)piperidin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 576 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)-2,2,2-trifluoroacetamide | |
| 577 | N-(4-((4-(4-(2-acetamidoethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 578 | N-(4-((4-(4-(3-(dimethylamino)propyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 579 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2- |
| (methylamino)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 580 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-(piperidin-1- |
| yl)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 581 | 1-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)-3-methylurea | |
| 582 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methyl-N-(5-methyl-1H-pyrazol-3-yl)- |
| 2-(methylthio)pyrimidin-4-amine | |
| 583 | (R)-N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(3- |
| methylmorpholino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 584 | (S)-N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(3- |
| methylmorpholino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 585 | 2-(4-(2-((4-ac etamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperazin-1-yl)-N,N-dimethylacetamide | |
| 586 | 4-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)-N,N-dimethylpiperazine-1-carboxamide | |
| 587 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-((1-methylpiperidin-4- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 588 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2- |
| morpholinoethyl)piperidin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 589 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-((1-methylpiperidin-4- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 590 | N-(4-((5-methyl-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-4- |
| ylamino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 591 | 1-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)urea | |
| 592 | N-(3-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 593 | N-(3-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)phenyl)acetamide | |
| 594 | N-(4-((4-((5-methyl-1H-pyrazol-3-yl)amino)-1′-pivaloyl-5,6- |
| dihydrospiro[cyclopenta[d]pyrimidine-7,4′-piperidin]-2-yl)thio)phenyl)acetamide | |
| 595 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(1,2,3,6-tetrahydropyridin-4- |
| yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 596 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-morpholino-2- |
| oxoethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 597 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-oxo-2-(piperidin-1- |
| yl)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 598 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(piperidin-4- |
| ylamino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 599 | N-(545-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-morpholinopyrimidin-2- |
| yl)thio)pyridin-2-yl) |
|
| 600 | 2,2-dimethyl-1-(4-((5-methyl-1H-pyrazol-3-yl)amino)-2-((4-nitrophenyl)thio)-5,6- |
| dihydrospiro[cyclopenta[d]pyrimidine-7,4′-piperidin]-1′-yl)propan-1-one | |
| 601 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-nitrophenyl)thio)pyrimidin-4-amine | |
| 602 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-N-(5-ethyl-1H-pyrazol-3-yl)-5-methoxy- |
| 2-(methylthio)pyrimidin-4-amine | |
| 603 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-N-(5-ethyl-1H-pyrazol-3-yl)-5-methyl-2- |
| (methylthio)pyrimidin-4-amine | |
| 604 | N-(5-methyl-1H-pyrazol-3-yl)-2-((4-nitrophenyl)thio)-5,6- |
| dihydrospiro[cyclopenta[d]pyrimidine-7,4′-piperidin]-4-amine | |
| 605 | 2-(4-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)-3,6-dihydropyridin-1(2H)-yl)-N,N-dimethylacetamide | |
| 606 | N-(4-((1′-(2-(dimethylamino)ethyl)-4-((5-methyl-1H-pyrazol-3-yl)amino)-5,6- |
| dihydrospiro[cyclopenta[d]pyrimidine-7,4′-piperidin]-2-yl)thio)phenyl)acetamide | |
| 607 | N-(4-((4-(1-(2-(dimethylamino)ethyl)-1,2,3,6-tetrahydropyridin-4-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 608 | N-(4-((4-(4-(2-(diethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 609 | N-(4-((4-(4-(2-(dimethylamino)ethyl)-2-methylpiperazin-1-yl)-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 610 | N-(4-((4-(4-(2-(dimethylamino)ethyl)-3-methylpiperazin-1-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 611 | N-(4-((4-(4-(2-(ethyl(methyl)amino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 612 | N-(4-((4-(4-(2-aminoethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 613 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-(piperazin-1- |
| yl)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 614 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-(pyrrolidin-1- |
| yl)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 615 | N-(4-((5-methoxy-4-(1-methyl-1,2,3,6-tetrahydropyridin-4-yl)-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 616 | 1′-(2-(dimethylamino)ethyl)-N-(5-methyl-1H-pyrazol-3-yl)-2-((4-nitrophenyl)thio)-5,6- |
| dihydrospiro[cyclopenta[d]pyrimidine-7,4′-piperidin]-4-amine | |
| 617 | 2-((4-aminophenyl)thio)-1′-(2-(dimethylamino)ethyl)-N-(5-methyl-1H-pyrazol-3-yl)- |
| 5,6-dihydrospiro[cyclopenta[d]pyrimidine-7,4′-piperidin]-4-amine 2-(1-(2-((4- | |
| acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-4- | |
| yl)piperidin-4-yl)acetic acid | |
| 618 | 2-(3-((6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2- |
| (methylthio)pyrimidin-4-yl)amino)-1H-pyrazol-5-yl)acetic acid | |
| 619 | 2-(4-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperazin-1-yl)acetic acid | |
| 620 | methyl 2-(1-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetate | |
| 621 | methyl 2-(3-((6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2- |
| (methylthio)pyrimidin-4-yl)amino)-1H-pyrazol-5-yl)acetate | |
| 622 | 2-(1-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperidin-4-yl)acetamide | |
| 623 | 2-(3-((6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2- |
| (methylthio)pyrimidin-4-yl)amino)-1H-pyrazol-5-yl)-N-methylacetamide | |
| 624 | 2-(4-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperazin-1-yl)acetamide | |
| 625 | 4-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperazine-1-carboxamide | |
| 626 | N-(4-((4-(4-(2-(4-hydroxypiperidin-1-yl)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl- |
| 1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 627 | N-(4-((4-(4-( N-hydroxycarbamimidoyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 628 | N-(4-((4-(8-(2-(dimethylamino)ethyl)-3,8-diazabicyclo[3.2.1]octan-3-yl)-5-methoxy-6- |
| ((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 629 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-(4-methylpiperazin-1- |
| yl)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 630 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-(piperidin-1- |
| yl)ethyl)piperidin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 631 | methyl 2-(4-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperazin-1-yl)acetate | |
| 632 | methyl 4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)benzoate | |
| 633 | 2-(1-(2-((4-acetamidophenyl)thio)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-4-yl)piperidin-4-yl)-N,N-dimethylacetamide | |
| 634 | 4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)-N,N-dimethylbenzamide | |
| 635 | N-(4-((4-(1-(2-(dimethylamino)ethyl)piperidin-4-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 636 | N-(4-((4-(4-(2-(azepan-1-yl)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 637 | N-(4-((4-(4-(2-(azocan-1-yl)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 638 | N-(4-((4-(4-(2-(dimethylamino)ethyl)-3,5-dimethylpiperazin-1-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 639 | N-(4-((4-(4-(2-guanidinoethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 640 | N-(4-((4-(4-carbamimidoylpiperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol-3- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 641 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-oxo-2-(piperidin-1- |
| yl)ethyl)piperidin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 642 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2-ureidoethyl)piperazin- |
| 1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 643 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(8-(2-(4-methylpiperazin-1- |
| yl)ethyl)-3,8-diazabicyclo[3.2.1]octan-3-yl)-pyrimidin-2-yl)thio)phenyl)acetamide | |
| 644 | N-(4-((5-methoxy-4-(4-(2-(methoxy(methyl)amino)ethyl)piperazin-1-yl)-6-((5-methyl- |
| 1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 645 | 1-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| 6-((5 pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)ethan-1-one | |
| 646 | 2-(3-((6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2- |
| (methylthio)pyrimidin-4-yl)amino)-1H-pyrazol-5-yl)-N-(3-fluorophenyl)acetamide | |
| 647 | 4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)-N-methylbenzamide | |
| 648 | N-(4-((4-(1-(2-(dimethylamino)ethyl)piperidin-4-yl)-5-methyl-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 649 | N-(4-((4-(4-(2-((1R,5S)-8-azabicyclo[3.2.1]octan-8-yl)ethyl)piperazin-1-yl)-5-methoxy- |
| 6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 650 | N-(4-((4-(4-(2-(9-azabicyclo[3.3.1]nonan-9-yl)ethyl)piperazin-1-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 651 | N-(4-((4-(4-(2-(benzyl(methyl)amino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl- |
| 1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 652 | N-(4-((4-(4-(2-(dimethylamino)ethyl)-2-methylpiperazin-1-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 653 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methylthiazol-2- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 654 | N-(4-((4-(8-(2-(azocan-1-yl)ethyl)-3,8-diazabicyclo[3.2.1]octan-3-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 655 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2- |
| (methylsulfonyl)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 656 | N-(4-((5-methoxy-4-(4-(2-(methyl(phenyl)amino)ethyl)piperazin-1-yl)-6-((5-methyl- |
| 1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 657 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)methanesulfonamide | |
| 658 | 5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2-(4-methylpiperazin-1- |
| yl)ethyl)piperazin-1-yl)-2-((4-(methylsulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 659 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-(methylsulfinyl)phenyl)thio)pyrimidin-4-amine | |
| 660 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-(methylsulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 661 | 2-(3-((6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2- |
| (methylthio)pyrimidin-4-yl)amino)-1H-pyrazol-5-yl)-N-(3-fluorophenyl)acetamide | |
| 662 | 4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H-pyrazol- |
| 3-yl)amino)pyrimidin-2-yl)thio)-N-methylbenzamide | |
| 663 | N-(4-((4-(1-(2-(dimethylamino)ethyl)piperidin-4-yl)-5-methyl-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 664 | N-(4-((4-(4-(2-((1R,5S)-8-azabicyclo[3.2.1]octan-8-yl)ethyl)piperazin-1-yl)-5-methoxy- |
| 6-((5-methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 665 | N-(4-((4-(4-(2-(9-azabicyclo[3.3.1]nonan-9-yl)ethyl)piperazin-1-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 666 | N-(4-((4-(4-(2-(benzyl(methyl)amino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl- |
| 1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 667 | N-(4-((4-(4-(2-(dimethylamino)ethyl)-2-methylpiperazin-1-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 668 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methylthiazol-2- |
| yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 669 | N-(4-((4-(8-(2-(azocan-1-yl)ethyl)-3,8-diazabicyclo[3.2.1]octan-3-yl)-5-methoxy-6-((5- |
| methyl-1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 670 | N-(4-((5-methoxy-4-((5-methyl-1H-pyrazol-3-yl)amino)-6-(4-(2- |
| (methylsulfonyl)ethyl)piperazin-1-yl)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 671 | N-(4-((5-methoxy-4-(4-(2-(methyl(phenyl)amino)ethyl)piperazin-1-yl)-6-((5-methyl- |
| 1H-pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)acetamide | |
| 672 | N-(4-((4-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-6-((5-methyl-1H- |
| pyrazol-3-yl)amino)pyrimidin-2-yl)thio)phenyl)methanesulfonamide | |
| 673 | 5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2-(4-methylpiperazin-1- |
| yl)ethyl)piperazin-1-yl)-2-((4-(methylsulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 674 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-(methylsulfinyl)phenyl)thio)pyrimidin-4-amine | |
| 675 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-(methylsulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 676 | 2-((4-((2-chlorobenzyl)sulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 677 | 2-((4-((3-chlorobenzyl)sulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 678 | 2-((4-((4-chlorobenzyl)sulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1- |
| yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 679 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4- |
| methylpiperazin-1-yl)pyrimidin-4-amine | |
| 680 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 681 | 2-((4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2-(4- |
| methylpiperazin-1-yl)ethyl)piperazin-1-yl)pyrimidin-4-amine | |
| 682 | 2-((4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4- |
| methylpiperazin-1-yl)pyrimidin-4-amine | |
| 683 | 2-((4-(ethylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| (piperazin-1-yl)pyrimidin-4-amine | |
| 684 | 6-(4-(2-(9-azabicyclo[3.3.1]nonan-9-yl)ethyl)piperazin-1-yl)-2-((4- |
| (benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4- | |
| amine | |
| 685 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-((4- |
| fluorobenzyl)sulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin- | |
| 4-amine | |
| 686 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-2-((4-(isopropylsulfonyl)phenyl)thio)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 687 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-2-((4-((4- |
| methoxybenzyl)sulfonyl)phenyl)thio)-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 688 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((pyridin-3-ylmethyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 689 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((pyridin-4-ylmethyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 690 | 6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-((thiophen-2-ylmethyl)sulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 691 | 6-(4-(2-(dimethylamino)ethyl)piperidin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- |
| yl)-2-((4-(methylsulfonyl)phenyl)thio)pyrimidin-4-amine | |
| 692 | 2-((4-(((1H-imidazol-2-yl)methyl)sulfonyl)phenyl)thio)-6-(4-(2- |
| (dimethylamino)ethyl)piperazin-1-yl)-5-methoxy-N-(5-methyl-1H-pyrazol-3- | |
| yl)pyrimidin-4-amine | |
| 693 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-6-methyl-N-(5-methyl-1H-pyrazol-3- |
| yl)pyrimidin-4-amine | |
| 694 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6-(4-(2- |
| (4-methylpiperazin-1-yl)ethyl)piperazin-1-yl)pyrimidin-4-amine | |
| 695 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| (piperidin-1-yl)pyrimidin-4-amine | |
| 696 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N-(5-methyl-1H-pyrazol-3-yl)-6- |
| morpholinopyrimidin-4-amine | |
| 697 | 2-((4-(benzylsulfonyl)phenyl)thio)-5-methoxy-N4,N4-dimethyl-N6-(5-methyl-1H- |
| pyrazol-3-yl)pyrimidine-4,6-diamine | |
| 698 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(1-(2-(dimethylamino)ethyl)piperidin-4-yl)-5- |
| methoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 699 | 2-((4-(benzylsulfonyl)phenyl)thio)-6-(4-(2-(dimethylamino)ethyl)piperazin-1-yl)-5- |
| ethoxy-N-(5-methyl-1H-pyrazol-3-yl)pyrimidin-4-amine | |
| 700 | 2-((4-(ethylsulfonyl)phenyl)thio)-5-methoxy-6-methyl-N-(5-methyl-1H-pyrazol-3- |
| yl)pyrimidin-4-amine | |
wherein R3A is substituted or unsubstituted alkyl, and R4C is substituted or unsubstituted heterocycloalkyl, and the remaining substituents are as defined in the specification.
Claims (20)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US16/769,458 US11612604B2 (en) | 2017-12-06 | 2018-12-06 | Methods of treating cancer with PLK4 inhibitors |
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US201762595498P | 2017-12-06 | 2017-12-06 | |
| US201862650957P | 2018-03-30 | 2018-03-30 | |
| US16/769,458 US11612604B2 (en) | 2017-12-06 | 2018-12-06 | Methods of treating cancer with PLK4 inhibitors |
| PCT/US2018/064243 WO2019113311A1 (en) | 2017-12-06 | 2018-12-06 | Methods of treating cancer with plk4 inhibitors |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| US20200383990A1 US20200383990A1 (en) | 2020-12-10 |
| US11612604B2 true US11612604B2 (en) | 2023-03-28 |
Family
ID=66751248
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US16/769,458 Active 2039-06-04 US11612604B2 (en) | 2017-12-06 | 2018-12-06 | Methods of treating cancer with PLK4 inhibitors |
Country Status (3)
| Country | Link |
|---|---|
| US (1) | US11612604B2 (en) |
| EP (1) | EP3720560A4 (en) |
| WO (1) | WO2019113311A1 (en) |
Families Citing this family (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| BR112022011052A2 (en) | 2019-12-06 | 2022-08-16 | Univ Health Network | Treatment for acute myeloid leukemia or myelodysplastic syndrome |
| CN112225729B (en) * | 2020-11-04 | 2022-02-01 | 四川大学华西医院 | Pyrimidine derivative, preparation method and application thereof and pharmaceutical composition |
| JP7717842B2 (en) | 2021-05-11 | 2025-08-04 | オリック ファーマシューティカルズ,インク. | Polo-like kinase 4 inhibitor |
| JP2025506772A (en) * | 2022-02-23 | 2025-03-13 | リペア セラピューティクス インコーポレイテッド | Polo-like kinase 4 (PLK4) inhibitors, pharmaceutical compositions, and methods for preparing and using the same - Patents.com |
| WO2023183525A1 (en) * | 2022-03-23 | 2023-09-28 | The Trustees Of Columbia University In The City Of New York | Compounds, targets, and methods for modulating lytic granule convergence in cytotoxic cells to promote bystander killing in cellular therapies |
| CN116850289B (en) * | 2022-07-29 | 2025-10-28 | 东北大学 | Application of PLK4-targeted drugs in the treatment of platinum-resistant tumors |
| CN115976214A (en) * | 2022-12-29 | 2023-04-18 | 重庆医科大学附属永川医院 | Application of PLK4 gene in the treatment of acute myeloid leukemia |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20030004161A1 (en) | 2000-12-21 | 2003-01-02 | David Bebbington | Pyrazole compounds useful as protein kinase inhititors |
| WO2012121662A1 (en) | 2011-03-04 | 2012-09-13 | Agency For Science, Technology And Research | Novel pharmaceutical combinations and methods for treating cancer |
| WO2016166604A1 (en) | 2015-04-17 | 2016-10-20 | Ludwig Institute For Cancer Research Ltd | Plk4 inhibitors |
Family Cites Families (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4911920A (en) | 1986-07-30 | 1990-03-27 | Alcon Laboratories, Inc. | Sustained release, comfort formulation for glaucoma therapy |
| FR2588189B1 (en) | 1985-10-03 | 1988-12-02 | Merck Sharp & Dohme | LIQUID-GEL PHASE TRANSITION PHARMACEUTICAL COMPOSITION |
| JP2594486B2 (en) | 1991-01-15 | 1997-03-26 | アルコン ラボラトリーズ インコーポレイテッド | Topical ophthalmic composition |
| US5212162A (en) | 1991-03-27 | 1993-05-18 | Alcon Laboratories, Inc. | Use of combinations gelling polysaccharides and finely divided drug carrier substrates in topical ophthalmic compositions |
| WO2002022605A1 (en) | 2000-09-15 | 2002-03-21 | Vertex Pharmaceuticals Incorporated | Pyrazole compounds useful as protein kinase inhibitors |
| NZ580343A (en) | 2007-04-13 | 2012-03-30 | Vertex Pharma | Aminopyrimidines useful as kinase inhibitors |
-
2018
- 2018-12-06 EP EP18886861.6A patent/EP3720560A4/en active Pending
- 2018-12-06 US US16/769,458 patent/US11612604B2/en active Active
- 2018-12-06 WO PCT/US2018/064243 patent/WO2019113311A1/en not_active Ceased
Patent Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20030004161A1 (en) | 2000-12-21 | 2003-01-02 | David Bebbington | Pyrazole compounds useful as protein kinase inhititors |
| WO2012121662A1 (en) | 2011-03-04 | 2012-09-13 | Agency For Science, Technology And Research | Novel pharmaceutical combinations and methods for treating cancer |
| WO2016166604A1 (en) | 2015-04-17 | 2016-10-20 | Ludwig Institute For Cancer Research Ltd | Plk4 inhibitors |
| CA2982890A1 (en) | 2015-04-17 | 2016-10-20 | Ludwig Institute For Cancer Research Ltd | Plk4 inhibitors |
Non-Patent Citations (29)
| Title |
|---|
| Bebbington, D. et al. (Jul. 1, 2009, e-published May 3, 2009). "The discovery of the potent aurora inhibitor MK-0457 (VX-680)," Bioorg Med Chem Lett 19(13):3586-3592. |
| Coelho et al., Open Biology (2015), 5:150209, 15 pages. * |
| Engstrom, P.F. et al. (Apr. 15, 1982). "Combination chemotherapy of advanced colorectal cancer utilizing 5-fluorouracil, semustine, dacarbazine, vincristine, and hydroxyurea: a phase III trial by the Eastern Cooperative Oncology Group (EST: 4275)," Cancer 49(8):1555-1560. |
| Extended European Search Report dated Dec. 3, 2021, for EP Patent Application No. 18886861.6, 25 pages. |
| Fiskus, W. et al. (Sep. 2012, e-published Jul. 24, 2012). "Co-treatmentwith vorinostat synergistically enhances activity of Aurora kinase inhibitor against human breast cancer cells," Breast Cancer Res Treat 135(2):433-444. |
| Giles, F.J. et al. (Jan. 2013). "MK-0457, an Aurora kinase and BCR-ABL inhibitor, is active in patients with BCR-ABL T315I leukemia," Leukemia 27(1):113-117. |
| Ham, J. et al. (Feb. 8, 2016). "Exploitation of the Apoptosis-Primed State of MYCN-Amplified Neuroblastoma to Develop a Potent and Specific Targeted Therapy Combination," Cancer Cell 29(2): 159-172. |
| Hu, C.E. et al. (Mar. 2017, e-published Jan. 16, 2017). "TRIM37 promotes epithelial-mesenchymal transition in colorectal cancer," Mol Med Rep 15(3):1057-1062. |
| International Search Report dated Feb. 27, 2019, for PCT Application No. PCT/US2018/064243, filed Dec. 6, 2018, 3 pages. |
| Jiang, J. et al. (Feb. 2016, e-published Sep. 22, 2015). "TRIM37 promoted the growth and migration of the pancreatic cancer cells," Tumour Biol 37(2):2629-2634. |
| Kidd, M. et al. (Feb. 2006). "Q RT-PCR detection of chromogranin A: a new standard in the identification of neuroendocrine tumor disease," Annals of Surgery 243(2):273-280. |
| Marsella, J.M. et al. (Sep. 1997). "Susceptibility of p53-deficient mice to induction of mesothelioma by crocidolite asbestos fibers," Environmental Health Perspectives 105 Suppl 5(Suppl 5):1069-1072. |
| Meitinger, F. et al. (2018). "Abstract 4130: TRIM37 expression levels dictate susceptibility to centrosome removal, supporting Plk4 inhibition as a potential new strategy for targeting neuroblastoma," Cancer Research, Proceedings: AACR Annual Meeting Apr. 14-18, 2018, Chicago, IL, located at <https://cancerres.aacrjournals.org/content/78/13_supplement/4130>, 4 pages. |
| Meitinger, F. et al. (Sep. 2020, e-published Sep. 9, 2020). "TRIM37 controls cancer-specific vulnerability to PLK4 inhibition," Nature 585(7825):440-446. |
| Michaelis, M. et al. (Sep. 30, 2014). "Aurora kinases as targets in drug-resistant neuroblastoma cells," PLoS One 9(9):e108758. |
| Okabe, S. et al. (Nov. 2010, e-published Jun. 20, 2010). "Efficacy of MK-0457 and in combination with vorinostat against Philadelphia chromosome positive acute lymphoblastic leukemia cells," Ann Hematol 89(11):1081-1087. |
| Partial Supplementary European Search Report dated Aug. 4, 2021, for EP Patent Application No. 18886861.6, 25 pages. |
| Pirker, C. et al. (Dec. 2010). "Response of experimental malignant melanoma models to the pan-Aurora kinase inhibitor VE-465," Exp Dermatol 19(12):1040-1047. |
| Recchia, F. et al. (May 2007, e-published Sep. 6, 2006). "Phase II study of interleukin-2 and 13-cis-retinoic acid as maintenance therapy in metastatic colorectal cancer," Cancer Immunol Immunother 56(5):699-708. |
| Ries, L. et al. "Cancer Incidence and Survival among Children and Adolescents: United States SEER Program 1975-1995," located at https://seer.cancer.gov/archive/publications/childhood/childhood-monograph.pdf, retrieved on Jan. 22, 2018, 65 pages. |
| Schubbert, S. et al. (Dec. 1, 2014, e-published Oct. 6, 2014). "Targeting the MYC and PI3K pathways eliminates leukemia-initiating cells in T-cell acute lymphoblastic leukemia," Cancer Res 74(23):7048-7059. |
| Soda, M. et al. (Aug. 2, 2007, e-published Jul. 11, 2007). "Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer," Nature 448(7153):561-566. |
| Sotlar, K. et al. (Mar. 2001). "Human papillomavirus type 16-associated primary squamous cell carcinoma of the rectum," Gastroenterology 120(4):988-994. |
| Tavanti, E. et al. (Nov. 12, 2013, e-published Oct. 15, 2013). "Preclinical validation of Aurora kinases-targeting drugs in osteosarcoma," Br J Cancer 109(10):2607-2618. |
| Winter, G.E. et al. (Oct. 2011, e-published Jul. 18, 2011). "An integrated chemical biology approach identifies specific vulnerability of Ewing's sarcoma to combined inhibition of Aurora kinases A and B," Mol Cancer Ther 10(10):1846-1856. |
| Wong et al., Science (2015), 348(6239), pp. 1155-1160. * |
| Written Opinion dated Feb. 27, 2019, for PCT Application No. PCT/US2018/064243, filed Dec. 6, 2018, 23 pages. |
| Yao, R. et al. (Jan. 2014. e-published Oct. 29, 2013). "VX680 suppresses the growth of HepG2 cells and enhances the chemosensitivity to cisplatin," Oncol Lett 7(1):121-124. |
| Yoshida, K. et al. (Dec. 15, 2011, e-published Sep. 29, 2011). "Vincristine potentiates the anti-proliferative effect of an aurora kinase inhibitor, VE-465, in myeloid leukemia cells," Biochem Pharmacol 82(12):1884-1890. |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2019113311A1 (en) | 2019-06-13 |
| EP3720560A1 (en) | 2020-10-14 |
| US20200383990A1 (en) | 2020-12-10 |
| EP3720560A4 (en) | 2022-01-05 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11964941B2 (en) | 5-Bromo-indirubins | |
| US11612604B2 (en) | Methods of treating cancer with PLK4 inhibitors | |
| US9895373B2 (en) | Substituted pyrazolo[3,4-D]pyrimidines and uses thereof | |
| US11730736B2 (en) | Anticancer agents | |
| US12576084B2 (en) | Methods for treating testicular and ovarian adrenal rest tumors | |
| US10752612B2 (en) | PLK4 inhibitors | |
| US11505523B2 (en) | HDAC8 inhibitors for treating cancer | |
| US12551482B2 (en) | Aurora kinase inhibitors | |
| US20220274964A1 (en) | 5-bromo-indirubins | |
| US10570124B2 (en) | Deoxycytidine kinase binding compounds | |
| US11840523B2 (en) | IRE1α inhibitors and uses thereof | |
| US12590056B2 (en) | IGF2BP2 inhibitors and uses thereof | |
| US11213510B2 (en) | Thioindirubins |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| FEPP | Fee payment procedure |
Free format text: ENTITY STATUS SET TO UNDISCOUNTED (ORIGINAL EVENT CODE: BIG.); ENTITY STATUS OF PATENT OWNER: SMALL ENTITY |
|
| FEPP | Fee payment procedure |
Free format text: ENTITY STATUS SET TO SMALL (ORIGINAL EVENT CODE: SMAL); ENTITY STATUS OF PATENT OWNER: SMALL ENTITY |
|
| STPP | Information on status: patent application and granting procedure in general |
Free format text: APPLICATION DISPATCHED FROM PREEXAM, NOT YET DOCKETED |
|
| STPP | Information on status: patent application and granting procedure in general |
Free format text: DOCKETED NEW CASE - READY FOR EXAMINATION |
|
| STPP | Information on status: patent application and granting procedure in general |
Free format text: NON FINAL ACTION MAILED |
|
| STPP | Information on status: patent application and granting procedure in general |
Free format text: RESPONSE TO NON-FINAL OFFICE ACTION ENTERED AND FORWARDED TO EXAMINER |
|
| STPP | Information on status: patent application and granting procedure in general |
Free format text: NON FINAL ACTION MAILED |
|
| STPP | Information on status: patent application and granting procedure in general |
Free format text: RESPONSE TO NON-FINAL OFFICE ACTION ENTERED AND FORWARDED TO EXAMINER |
|
| STPP | Information on status: patent application and granting procedure in general |
Free format text: NOTICE OF ALLOWANCE MAILED -- APPLICATION RECEIVED IN OFFICE OF PUBLICATIONS |
|
| AS | Assignment |
Owner name: LUDWI INSTITUTE FOR CANCER RESEARCH LTD., SWITZERLAND Free format text: ASSIGNMENT OF ASSIGNORS INTEREST;ASSIGNORS:OEGEMA, KAREN;SHIAU, ANDREW;DESAI, ARSHAD;AND OTHERS;REEL/FRAME:061711/0799 Effective date: 20181130 |
|
| STCF | Information on status: patent grant |
Free format text: PATENTED CASE |